class: center, middle, inverse, title-slide # Animated graphics in R ### Abhijit Dasgupta, PhD --- layout: true <div class="my-header"> <span>BIOF 339: Practical R</span></div> --- class: middle,inverse # Dynamic graphics --- ## Dynamic graphics in R There are several package in R that provide dynamic graphics meant to be consumed on the web. Many of these are wrappers around well-known Javascript libraries like D3.js, leaflet.js and others These packages have mostly come under the umbrella of of the `htmlwidgets` package, which allows these HTML-based graphics to be displayed through R and R Markdown <iframe src="https://www.htmlwidgets.org" height="300" width="100%"></iframe> --- ## Dynamic graphics in R There are several broad categories of dynamic graphs General purpose: <table class="table" style="margin-left: auto; margin-right: auto;"> <thead> <tr> <th style="text-align:left;"> Package </th> <th style="text-align:left;"> Description </th> </tr> </thead> <tbody> <tr> <td style="text-align:left;"> r2d3 </td> <td style="text-align:left;"> Interface for D3.js, requires D3.js code </td> </tr> <tr> <td style="text-align:left;"> plotly </td> <td style="text-align:left;"> Interface with plot.ly, direct conversion from ggplot2 </td> </tr> <tr> <td style="text-align:left;"> highcharter </td> <td style="text-align:left;"> Using the Highcharts.js package </td> </tr> <tr> <td style="text-align:left;"> dygraphs </td> <td style="text-align:left;"> For time series or longitudinal data </td> </tr> </tbody> </table> Maps: ```r tribble( ~Package, ~Description, "leaflet", "Maps using OpenStreetMaps") %>% kable() %>% kable_styling() ``` <table class="table" style="margin-left: auto; margin-right: auto;"> <thead> <tr> <th style="text-align:left;"> Package </th> <th style="text-align:left;"> Description </th> </tr> </thead> <tbody> <tr> <td style="text-align:left;"> leaflet </td> <td style="text-align:left;"> Maps using OpenStreetMaps </td> </tr> </tbody> </table> --- ## Dynamic graphics in R Networks: ```r tribble(~Package, ~Description, 'networkD3', 'Dynamic network visualizations using D3', 'visNetwork', 'Interface to the vis.js pacman::p_load') %>% kable() %>% kable_styling() ``` <table class="table" style="margin-left: auto; margin-right: auto;"> <thead> <tr> <th style="text-align:left;"> Package </th> <th style="text-align:left;"> Description </th> </tr> </thead> <tbody> <tr> <td style="text-align:left;"> networkD3 </td> <td style="text-align:left;"> Dynamic network visualizations using D3 </td> </tr> <tr> <td style="text-align:left;"> visNetwork </td> <td style="text-align:left;"> Interface to the vis.js pacman::p_load </td> </tr> </tbody> </table> --- class: middle, inverse # Plotly https://plotly.com/graphing-libraries --- ## Plotly Plotly is a company that developed the `plotly.js` graphing pacman::p_load, as well as packages for R and Python. For the R package, it developed a turnkey method to convert `ggplot2` graphics into interactive graphs. .pull-left[ ```r plt <- ggplot(penguins, aes(bill_length_mm, body_mass_g, color = species))+ geom_point() plt ``` 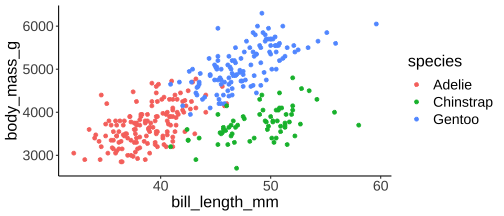<!-- --> ] .pull-right[ ```r ggplotly(plt) ``` <div id="htmlwidget-f7c5df5315fce64de5a1" style="width:504px;height:288px;" class="plotly html-widget"></div> <script type="application/json" data-for="htmlwidget-f7c5df5315fce64de5a1">{"x":{"data":[{"x":[39.1,39.5,40.3,null,36.7,39.3,38.9,39.2,34.1,42,37.8,37.8,41.1,38.6,34.6,36.6,38.7,42.5,34.4,46,37.8,37.7,35.9,38.2,38.8,35.3,40.6,40.5,37.9,40.5,39.5,37.2,39.5,40.9,36.4,39.2,38.8,42.2,37.6,39.8,36.5,40.8,36,44.1,37,39.6,41.1,37.5,36,42.3,39.6,40.1,35,42,34.5,41.4,39,40.6,36.5,37.6,35.7,41.3,37.6,41.1,36.4,41.6,35.5,41.1,35.9,41.8,33.5,39.7,39.6,45.8,35.5,42.8,40.9,37.2,36.2,42.1,34.6,42.9,36.7,35.1,37.3,41.3,36.3,36.9,38.3,38.9,35.7,41.1,34,39.6,36.2,40.8,38.1,40.3,33.1,43.2,35,41,37.7,37.8,37.9,39.7,38.6,38.2,38.1,43.2,38.1,45.6,39.7,42.2,39.6,42.7,38.6,37.3,35.7,41.1,36.2,37.7,40.2,41.4,35.2,40.6,38.8,41.5,39,44.1,38.5,43.1,36.8,37.5,38.1,41.1,35.6,40.2,37,39.7,40.2,40.6,32.1,40.7,37.3,39,39.2,36.6,36,37.8,36,41.5],"y":[3750,3800,3250,null,3450,3650,3625,4675,3475,4250,3300,3700,3200,3800,4400,3700,3450,4500,3325,4200,3400,3600,3800,3950,3800,3800,3550,3200,3150,3950,3250,3900,3300,3900,3325,4150,3950,3550,3300,4650,3150,3900,3100,4400,3000,4600,3425,2975,3450,4150,3500,4300,3450,4050,2900,3700,3550,3800,2850,3750,3150,4400,3600,4050,2850,3950,3350,4100,3050,4450,3600,3900,3550,4150,3700,4250,3700,3900,3550,4000,3200,4700,3800,4200,3350,3550,3800,3500,3950,3600,3550,4300,3400,4450,3300,4300,3700,4350,2900,4100,3725,4725,3075,4250,2925,3550,3750,3900,3175,4775,3825,4600,3200,4275,3900,4075,2900,3775,3350,3325,3150,3500,3450,3875,3050,4000,3275,4300,3050,4000,3325,3500,3500,4475,3425,3900,3175,3975,3400,4250,3400,3475,3050,3725,3000,3650,4250,3475,3450,3750,3700,4000],"text":["bill_length_mm: 39.1<br />body_mass_g: 3750<br />species: Adelie","bill_length_mm: 39.5<br />body_mass_g: 3800<br />species: Adelie","bill_length_mm: 40.3<br />body_mass_g: 3250<br />species: Adelie","bill_length_mm: NA<br />body_mass_g: NA<br />species: Adelie","bill_length_mm: 36.7<br />body_mass_g: 3450<br />species: Adelie","bill_length_mm: 39.3<br />body_mass_g: 3650<br />species: Adelie","bill_length_mm: 38.9<br />body_mass_g: 3625<br />species: Adelie","bill_length_mm: 39.2<br />body_mass_g: 4675<br />species: Adelie","bill_length_mm: 34.1<br />body_mass_g: 3475<br />species: Adelie","bill_length_mm: 42.0<br />body_mass_g: 4250<br />species: Adelie","bill_length_mm: 37.8<br />body_mass_g: 3300<br />species: Adelie","bill_length_mm: 37.8<br />body_mass_g: 3700<br />species: Adelie","bill_length_mm: 41.1<br />body_mass_g: 3200<br />species: Adelie","bill_length_mm: 38.6<br />body_mass_g: 3800<br />species: Adelie","bill_length_mm: 34.6<br />body_mass_g: 4400<br />species: Adelie","bill_length_mm: 36.6<br />body_mass_g: 3700<br />species: Adelie","bill_length_mm: 38.7<br />body_mass_g: 3450<br />species: Adelie","bill_length_mm: 42.5<br />body_mass_g: 4500<br />species: Adelie","bill_length_mm: 34.4<br />body_mass_g: 3325<br />species: Adelie","bill_length_mm: 46.0<br />body_mass_g: 4200<br />species: Adelie","bill_length_mm: 37.8<br />body_mass_g: 3400<br />species: Adelie","bill_length_mm: 37.7<br />body_mass_g: 3600<br />species: Adelie","bill_length_mm: 35.9<br />body_mass_g: 3800<br />species: Adelie","bill_length_mm: 38.2<br />body_mass_g: 3950<br />species: Adelie","bill_length_mm: 38.8<br />body_mass_g: 3800<br />species: Adelie","bill_length_mm: 35.3<br />body_mass_g: 3800<br />species: Adelie","bill_length_mm: 40.6<br />body_mass_g: 3550<br />species: Adelie","bill_length_mm: 40.5<br />body_mass_g: 3200<br />species: Adelie","bill_length_mm: 37.9<br />body_mass_g: 3150<br />species: Adelie","bill_length_mm: 40.5<br />body_mass_g: 3950<br />species: Adelie","bill_length_mm: 39.5<br />body_mass_g: 3250<br />species: Adelie","bill_length_mm: 37.2<br />body_mass_g: 3900<br />species: Adelie","bill_length_mm: 39.5<br />body_mass_g: 3300<br />species: Adelie","bill_length_mm: 40.9<br />body_mass_g: 3900<br />species: Adelie","bill_length_mm: 36.4<br />body_mass_g: 3325<br />species: Adelie","bill_length_mm: 39.2<br />body_mass_g: 4150<br />species: Adelie","bill_length_mm: 38.8<br />body_mass_g: 3950<br />species: Adelie","bill_length_mm: 42.2<br />body_mass_g: 3550<br />species: Adelie","bill_length_mm: 37.6<br />body_mass_g: 3300<br />species: Adelie","bill_length_mm: 39.8<br />body_mass_g: 4650<br />species: Adelie","bill_length_mm: 36.5<br />body_mass_g: 3150<br />species: Adelie","bill_length_mm: 40.8<br />body_mass_g: 3900<br />species: Adelie","bill_length_mm: 36.0<br />body_mass_g: 3100<br />species: Adelie","bill_length_mm: 44.1<br />body_mass_g: 4400<br />species: Adelie","bill_length_mm: 37.0<br />body_mass_g: 3000<br />species: Adelie","bill_length_mm: 39.6<br />body_mass_g: 4600<br />species: Adelie","bill_length_mm: 41.1<br />body_mass_g: 3425<br />species: Adelie","bill_length_mm: 37.5<br />body_mass_g: 2975<br />species: Adelie","bill_length_mm: 36.0<br />body_mass_g: 3450<br />species: Adelie","bill_length_mm: 42.3<br />body_mass_g: 4150<br />species: Adelie","bill_length_mm: 39.6<br />body_mass_g: 3500<br />species: Adelie","bill_length_mm: 40.1<br />body_mass_g: 4300<br />species: Adelie","bill_length_mm: 35.0<br />body_mass_g: 3450<br />species: Adelie","bill_length_mm: 42.0<br />body_mass_g: 4050<br />species: Adelie","bill_length_mm: 34.5<br />body_mass_g: 2900<br />species: Adelie","bill_length_mm: 41.4<br />body_mass_g: 3700<br />species: Adelie","bill_length_mm: 39.0<br />body_mass_g: 3550<br />species: Adelie","bill_length_mm: 40.6<br />body_mass_g: 3800<br />species: Adelie","bill_length_mm: 36.5<br />body_mass_g: 2850<br />species: Adelie","bill_length_mm: 37.6<br />body_mass_g: 3750<br />species: Adelie","bill_length_mm: 35.7<br />body_mass_g: 3150<br />species: Adelie","bill_length_mm: 41.3<br />body_mass_g: 4400<br />species: Adelie","bill_length_mm: 37.6<br />body_mass_g: 3600<br />species: Adelie","bill_length_mm: 41.1<br />body_mass_g: 4050<br />species: Adelie","bill_length_mm: 36.4<br />body_mass_g: 2850<br />species: Adelie","bill_length_mm: 41.6<br />body_mass_g: 3950<br />species: Adelie","bill_length_mm: 35.5<br />body_mass_g: 3350<br />species: Adelie","bill_length_mm: 41.1<br />body_mass_g: 4100<br />species: Adelie","bill_length_mm: 35.9<br />body_mass_g: 3050<br />species: Adelie","bill_length_mm: 41.8<br />body_mass_g: 4450<br />species: Adelie","bill_length_mm: 33.5<br />body_mass_g: 3600<br />species: Adelie","bill_length_mm: 39.7<br />body_mass_g: 3900<br />species: Adelie","bill_length_mm: 39.6<br />body_mass_g: 3550<br />species: Adelie","bill_length_mm: 45.8<br />body_mass_g: 4150<br />species: Adelie","bill_length_mm: 35.5<br />body_mass_g: 3700<br />species: Adelie","bill_length_mm: 42.8<br />body_mass_g: 4250<br />species: Adelie","bill_length_mm: 40.9<br />body_mass_g: 3700<br />species: Adelie","bill_length_mm: 37.2<br />body_mass_g: 3900<br />species: Adelie","bill_length_mm: 36.2<br />body_mass_g: 3550<br />species: Adelie","bill_length_mm: 42.1<br />body_mass_g: 4000<br />species: Adelie","bill_length_mm: 34.6<br />body_mass_g: 3200<br />species: Adelie","bill_length_mm: 42.9<br />body_mass_g: 4700<br />species: Adelie","bill_length_mm: 36.7<br />body_mass_g: 3800<br />species: Adelie","bill_length_mm: 35.1<br />body_mass_g: 4200<br />species: Adelie","bill_length_mm: 37.3<br />body_mass_g: 3350<br />species: Adelie","bill_length_mm: 41.3<br />body_mass_g: 3550<br />species: Adelie","bill_length_mm: 36.3<br />body_mass_g: 3800<br />species: Adelie","bill_length_mm: 36.9<br />body_mass_g: 3500<br />species: Adelie","bill_length_mm: 38.3<br />body_mass_g: 3950<br />species: Adelie","bill_length_mm: 38.9<br />body_mass_g: 3600<br />species: Adelie","bill_length_mm: 35.7<br />body_mass_g: 3550<br />species: Adelie","bill_length_mm: 41.1<br />body_mass_g: 4300<br />species: Adelie","bill_length_mm: 34.0<br />body_mass_g: 3400<br />species: Adelie","bill_length_mm: 39.6<br />body_mass_g: 4450<br />species: Adelie","bill_length_mm: 36.2<br />body_mass_g: 3300<br />species: Adelie","bill_length_mm: 40.8<br />body_mass_g: 4300<br />species: Adelie","bill_length_mm: 38.1<br />body_mass_g: 3700<br />species: Adelie","bill_length_mm: 40.3<br />body_mass_g: 4350<br />species: Adelie","bill_length_mm: 33.1<br />body_mass_g: 2900<br />species: Adelie","bill_length_mm: 43.2<br />body_mass_g: 4100<br />species: Adelie","bill_length_mm: 35.0<br />body_mass_g: 3725<br />species: Adelie","bill_length_mm: 41.0<br />body_mass_g: 4725<br />species: Adelie","bill_length_mm: 37.7<br />body_mass_g: 3075<br />species: Adelie","bill_length_mm: 37.8<br />body_mass_g: 4250<br />species: Adelie","bill_length_mm: 37.9<br />body_mass_g: 2925<br />species: Adelie","bill_length_mm: 39.7<br />body_mass_g: 3550<br />species: Adelie","bill_length_mm: 38.6<br />body_mass_g: 3750<br />species: Adelie","bill_length_mm: 38.2<br />body_mass_g: 3900<br />species: Adelie","bill_length_mm: 38.1<br />body_mass_g: 3175<br />species: Adelie","bill_length_mm: 43.2<br />body_mass_g: 4775<br />species: Adelie","bill_length_mm: 38.1<br />body_mass_g: 3825<br />species: Adelie","bill_length_mm: 45.6<br />body_mass_g: 4600<br />species: Adelie","bill_length_mm: 39.7<br />body_mass_g: 3200<br />species: Adelie","bill_length_mm: 42.2<br />body_mass_g: 4275<br />species: Adelie","bill_length_mm: 39.6<br />body_mass_g: 3900<br />species: Adelie","bill_length_mm: 42.7<br />body_mass_g: 4075<br />species: Adelie","bill_length_mm: 38.6<br />body_mass_g: 2900<br />species: Adelie","bill_length_mm: 37.3<br />body_mass_g: 3775<br />species: Adelie","bill_length_mm: 35.7<br />body_mass_g: 3350<br />species: Adelie","bill_length_mm: 41.1<br />body_mass_g: 3325<br />species: Adelie","bill_length_mm: 36.2<br />body_mass_g: 3150<br />species: Adelie","bill_length_mm: 37.7<br />body_mass_g: 3500<br />species: Adelie","bill_length_mm: 40.2<br />body_mass_g: 3450<br />species: Adelie","bill_length_mm: 41.4<br />body_mass_g: 3875<br />species: Adelie","bill_length_mm: 35.2<br />body_mass_g: 3050<br />species: Adelie","bill_length_mm: 40.6<br />body_mass_g: 4000<br />species: Adelie","bill_length_mm: 38.8<br />body_mass_g: 3275<br />species: Adelie","bill_length_mm: 41.5<br />body_mass_g: 4300<br />species: Adelie","bill_length_mm: 39.0<br />body_mass_g: 3050<br />species: Adelie","bill_length_mm: 44.1<br />body_mass_g: 4000<br />species: Adelie","bill_length_mm: 38.5<br />body_mass_g: 3325<br />species: Adelie","bill_length_mm: 43.1<br />body_mass_g: 3500<br />species: Adelie","bill_length_mm: 36.8<br />body_mass_g: 3500<br />species: Adelie","bill_length_mm: 37.5<br />body_mass_g: 4475<br />species: Adelie","bill_length_mm: 38.1<br />body_mass_g: 3425<br />species: Adelie","bill_length_mm: 41.1<br />body_mass_g: 3900<br />species: Adelie","bill_length_mm: 35.6<br />body_mass_g: 3175<br />species: Adelie","bill_length_mm: 40.2<br />body_mass_g: 3975<br />species: Adelie","bill_length_mm: 37.0<br />body_mass_g: 3400<br />species: Adelie","bill_length_mm: 39.7<br />body_mass_g: 4250<br />species: Adelie","bill_length_mm: 40.2<br />body_mass_g: 3400<br />species: Adelie","bill_length_mm: 40.6<br />body_mass_g: 3475<br />species: Adelie","bill_length_mm: 32.1<br />body_mass_g: 3050<br />species: Adelie","bill_length_mm: 40.7<br />body_mass_g: 3725<br />species: Adelie","bill_length_mm: 37.3<br />body_mass_g: 3000<br />species: Adelie","bill_length_mm: 39.0<br />body_mass_g: 3650<br />species: Adelie","bill_length_mm: 39.2<br />body_mass_g: 4250<br />species: Adelie","bill_length_mm: 36.6<br />body_mass_g: 3475<br />species: Adelie","bill_length_mm: 36.0<br />body_mass_g: 3450<br />species: Adelie","bill_length_mm: 37.8<br />body_mass_g: 3750<br />species: Adelie","bill_length_mm: 36.0<br />body_mass_g: 3700<br />species: Adelie","bill_length_mm: 41.5<br />body_mass_g: 4000<br />species: Adelie"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(248,118,109,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(248,118,109,1)"}},"hoveron":"points","name":"Adelie","legendgroup":"Adelie","showlegend":true,"xaxis":"x","yaxis":"y","hoverinfo":"text","frame":null},{"x":[46.5,50,51.3,45.4,52.7,45.2,46.1,51.3,46,51.3,46.6,51.7,47,52,45.9,50.5,50.3,58,46.4,49.2,42.4,48.5,43.2,50.6,46.7,52,50.5,49.5,46.4,52.8,40.9,54.2,42.5,51,49.7,47.5,47.6,52,46.9,53.5,49,46.2,50.9,45.5,50.9,50.8,50.1,49,51.5,49.8,48.1,51.4,45.7,50.7,42.5,52.2,45.2,49.3,50.2,45.6,51.9,46.8,45.7,55.8,43.5,49.6,50.8,50.2],"y":[3500,3900,3650,3525,3725,3950,3250,3750,4150,3700,3800,3775,3700,4050,3575,4050,3300,3700,3450,4400,3600,3400,2900,3800,3300,4150,3400,3800,3700,4550,3200,4300,3350,4100,3600,3900,3850,4800,2700,4500,3950,3650,3550,3500,3675,4450,3400,4300,3250,3675,3325,3950,3600,4050,3350,3450,3250,4050,3800,3525,3950,3650,3650,4000,3400,3775,4100,3775],"text":["bill_length_mm: 46.5<br />body_mass_g: 3500<br />species: Chinstrap","bill_length_mm: 50.0<br />body_mass_g: 3900<br />species: Chinstrap","bill_length_mm: 51.3<br />body_mass_g: 3650<br />species: Chinstrap","bill_length_mm: 45.4<br />body_mass_g: 3525<br />species: Chinstrap","bill_length_mm: 52.7<br />body_mass_g: 3725<br />species: Chinstrap","bill_length_mm: 45.2<br />body_mass_g: 3950<br />species: Chinstrap","bill_length_mm: 46.1<br />body_mass_g: 3250<br />species: Chinstrap","bill_length_mm: 51.3<br />body_mass_g: 3750<br />species: Chinstrap","bill_length_mm: 46.0<br />body_mass_g: 4150<br />species: Chinstrap","bill_length_mm: 51.3<br />body_mass_g: 3700<br />species: Chinstrap","bill_length_mm: 46.6<br />body_mass_g: 3800<br />species: Chinstrap","bill_length_mm: 51.7<br />body_mass_g: 3775<br />species: Chinstrap","bill_length_mm: 47.0<br />body_mass_g: 3700<br />species: Chinstrap","bill_length_mm: 52.0<br />body_mass_g: 4050<br />species: Chinstrap","bill_length_mm: 45.9<br />body_mass_g: 3575<br />species: Chinstrap","bill_length_mm: 50.5<br />body_mass_g: 4050<br />species: Chinstrap","bill_length_mm: 50.3<br />body_mass_g: 3300<br />species: Chinstrap","bill_length_mm: 58.0<br />body_mass_g: 3700<br />species: Chinstrap","bill_length_mm: 46.4<br />body_mass_g: 3450<br />species: Chinstrap","bill_length_mm: 49.2<br />body_mass_g: 4400<br />species: Chinstrap","bill_length_mm: 42.4<br />body_mass_g: 3600<br />species: Chinstrap","bill_length_mm: 48.5<br />body_mass_g: 3400<br />species: Chinstrap","bill_length_mm: 43.2<br />body_mass_g: 2900<br />species: Chinstrap","bill_length_mm: 50.6<br />body_mass_g: 3800<br />species: Chinstrap","bill_length_mm: 46.7<br />body_mass_g: 3300<br />species: Chinstrap","bill_length_mm: 52.0<br />body_mass_g: 4150<br />species: Chinstrap","bill_length_mm: 50.5<br />body_mass_g: 3400<br />species: Chinstrap","bill_length_mm: 49.5<br />body_mass_g: 3800<br />species: Chinstrap","bill_length_mm: 46.4<br />body_mass_g: 3700<br />species: Chinstrap","bill_length_mm: 52.8<br />body_mass_g: 4550<br />species: Chinstrap","bill_length_mm: 40.9<br />body_mass_g: 3200<br />species: Chinstrap","bill_length_mm: 54.2<br />body_mass_g: 4300<br />species: Chinstrap","bill_length_mm: 42.5<br />body_mass_g: 3350<br />species: Chinstrap","bill_length_mm: 51.0<br />body_mass_g: 4100<br />species: Chinstrap","bill_length_mm: 49.7<br />body_mass_g: 3600<br />species: Chinstrap","bill_length_mm: 47.5<br />body_mass_g: 3900<br />species: Chinstrap","bill_length_mm: 47.6<br />body_mass_g: 3850<br />species: Chinstrap","bill_length_mm: 52.0<br />body_mass_g: 4800<br />species: Chinstrap","bill_length_mm: 46.9<br />body_mass_g: 2700<br />species: Chinstrap","bill_length_mm: 53.5<br />body_mass_g: 4500<br />species: Chinstrap","bill_length_mm: 49.0<br />body_mass_g: 3950<br />species: Chinstrap","bill_length_mm: 46.2<br />body_mass_g: 3650<br />species: Chinstrap","bill_length_mm: 50.9<br />body_mass_g: 3550<br />species: Chinstrap","bill_length_mm: 45.5<br />body_mass_g: 3500<br />species: Chinstrap","bill_length_mm: 50.9<br />body_mass_g: 3675<br />species: Chinstrap","bill_length_mm: 50.8<br />body_mass_g: 4450<br />species: Chinstrap","bill_length_mm: 50.1<br />body_mass_g: 3400<br />species: Chinstrap","bill_length_mm: 49.0<br />body_mass_g: 4300<br />species: Chinstrap","bill_length_mm: 51.5<br />body_mass_g: 3250<br />species: Chinstrap","bill_length_mm: 49.8<br />body_mass_g: 3675<br />species: Chinstrap","bill_length_mm: 48.1<br />body_mass_g: 3325<br />species: Chinstrap","bill_length_mm: 51.4<br />body_mass_g: 3950<br />species: Chinstrap","bill_length_mm: 45.7<br />body_mass_g: 3600<br />species: Chinstrap","bill_length_mm: 50.7<br />body_mass_g: 4050<br />species: Chinstrap","bill_length_mm: 42.5<br />body_mass_g: 3350<br />species: Chinstrap","bill_length_mm: 52.2<br />body_mass_g: 3450<br />species: Chinstrap","bill_length_mm: 45.2<br />body_mass_g: 3250<br />species: Chinstrap","bill_length_mm: 49.3<br />body_mass_g: 4050<br />species: Chinstrap","bill_length_mm: 50.2<br />body_mass_g: 3800<br />species: Chinstrap","bill_length_mm: 45.6<br />body_mass_g: 3525<br />species: Chinstrap","bill_length_mm: 51.9<br />body_mass_g: 3950<br />species: Chinstrap","bill_length_mm: 46.8<br />body_mass_g: 3650<br />species: Chinstrap","bill_length_mm: 45.7<br />body_mass_g: 3650<br />species: Chinstrap","bill_length_mm: 55.8<br />body_mass_g: 4000<br />species: Chinstrap","bill_length_mm: 43.5<br />body_mass_g: 3400<br />species: Chinstrap","bill_length_mm: 49.6<br />body_mass_g: 3775<br />species: Chinstrap","bill_length_mm: 50.8<br />body_mass_g: 4100<br />species: Chinstrap","bill_length_mm: 50.2<br />body_mass_g: 3775<br />species: Chinstrap"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(0,186,56,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(0,186,56,1)"}},"hoveron":"points","name":"Chinstrap","legendgroup":"Chinstrap","showlegend":true,"xaxis":"x","yaxis":"y","hoverinfo":"text","frame":null},{"x":[46.1,50,48.7,50,47.6,46.5,45.4,46.7,43.3,46.8,40.9,49,45.5,48.4,45.8,49.3,42,49.2,46.2,48.7,50.2,45.1,46.5,46.3,42.9,46.1,44.5,47.8,48.2,50,47.3,42.8,45.1,59.6,49.1,48.4,42.6,44.4,44,48.7,42.7,49.6,45.3,49.6,50.5,43.6,45.5,50.5,44.9,45.2,46.6,48.5,45.1,50.1,46.5,45,43.8,45.5,43.2,50.4,45.3,46.2,45.7,54.3,45.8,49.8,46.2,49.5,43.5,50.7,47.7,46.4,48.2,46.5,46.4,48.6,47.5,51.1,45.2,45.2,49.1,52.5,47.4,50,44.9,50.8,43.4,51.3,47.5,52.1,47.5,52.2,45.5,49.5,44.5,50.8,49.4,46.9,48.4,51.1,48.5,55.9,47.2,49.1,47.3,46.8,41.7,53.4,43.3,48.1,50.5,49.8,43.5,51.5,46.2,55.1,44.5,48.8,47.2,null,46.8,50.4,45.2,49.9],"y":[4500,5700,4450,5700,5400,4550,4800,5200,4400,5150,4650,5550,4650,5850,4200,5850,4150,6300,4800,5350,5700,5000,4400,5050,5000,5100,4100,5650,4600,5550,5250,4700,5050,6050,5150,5400,4950,5250,4350,5350,3950,5700,4300,4750,5550,4900,4200,5400,5100,5300,4850,5300,4400,5000,4900,5050,4300,5000,4450,5550,4200,5300,4400,5650,4700,5700,4650,5800,4700,5550,4750,5000,5100,5200,4700,5800,4600,6000,4750,5950,4625,5450,4725,5350,4750,5600,4600,5300,4875,5550,4950,5400,4750,5650,4850,5200,4925,4875,4625,5250,4850,5600,4975,5500,4725,5500,4700,5500,4575,5500,5000,5950,4650,5500,4375,5850,4875,6000,4925,null,4850,5750,5200,5400],"text":["bill_length_mm: 46.1<br />body_mass_g: 4500<br />species: Gentoo","bill_length_mm: 50.0<br />body_mass_g: 5700<br />species: Gentoo","bill_length_mm: 48.7<br />body_mass_g: 4450<br />species: Gentoo","bill_length_mm: 50.0<br />body_mass_g: 5700<br />species: Gentoo","bill_length_mm: 47.6<br />body_mass_g: 5400<br />species: Gentoo","bill_length_mm: 46.5<br />body_mass_g: 4550<br />species: Gentoo","bill_length_mm: 45.4<br />body_mass_g: 4800<br />species: Gentoo","bill_length_mm: 46.7<br />body_mass_g: 5200<br />species: Gentoo","bill_length_mm: 43.3<br />body_mass_g: 4400<br />species: Gentoo","bill_length_mm: 46.8<br />body_mass_g: 5150<br />species: Gentoo","bill_length_mm: 40.9<br />body_mass_g: 4650<br />species: Gentoo","bill_length_mm: 49.0<br />body_mass_g: 5550<br />species: Gentoo","bill_length_mm: 45.5<br />body_mass_g: 4650<br />species: Gentoo","bill_length_mm: 48.4<br />body_mass_g: 5850<br />species: Gentoo","bill_length_mm: 45.8<br />body_mass_g: 4200<br />species: Gentoo","bill_length_mm: 49.3<br />body_mass_g: 5850<br />species: Gentoo","bill_length_mm: 42.0<br />body_mass_g: 4150<br />species: Gentoo","bill_length_mm: 49.2<br />body_mass_g: 6300<br />species: Gentoo","bill_length_mm: 46.2<br />body_mass_g: 4800<br />species: Gentoo","bill_length_mm: 48.7<br />body_mass_g: 5350<br />species: Gentoo","bill_length_mm: 50.2<br />body_mass_g: 5700<br />species: Gentoo","bill_length_mm: 45.1<br />body_mass_g: 5000<br />species: Gentoo","bill_length_mm: 46.5<br />body_mass_g: 4400<br />species: Gentoo","bill_length_mm: 46.3<br />body_mass_g: 5050<br />species: Gentoo","bill_length_mm: 42.9<br />body_mass_g: 5000<br />species: Gentoo","bill_length_mm: 46.1<br />body_mass_g: 5100<br />species: Gentoo","bill_length_mm: 44.5<br />body_mass_g: 4100<br />species: Gentoo","bill_length_mm: 47.8<br />body_mass_g: 5650<br />species: Gentoo","bill_length_mm: 48.2<br />body_mass_g: 4600<br />species: Gentoo","bill_length_mm: 50.0<br />body_mass_g: 5550<br />species: Gentoo","bill_length_mm: 47.3<br />body_mass_g: 5250<br />species: Gentoo","bill_length_mm: 42.8<br />body_mass_g: 4700<br />species: Gentoo","bill_length_mm: 45.1<br />body_mass_g: 5050<br />species: Gentoo","bill_length_mm: 59.6<br />body_mass_g: 6050<br />species: Gentoo","bill_length_mm: 49.1<br />body_mass_g: 5150<br />species: Gentoo","bill_length_mm: 48.4<br />body_mass_g: 5400<br />species: Gentoo","bill_length_mm: 42.6<br />body_mass_g: 4950<br />species: Gentoo","bill_length_mm: 44.4<br />body_mass_g: 5250<br />species: Gentoo","bill_length_mm: 44.0<br />body_mass_g: 4350<br />species: Gentoo","bill_length_mm: 48.7<br />body_mass_g: 5350<br />species: Gentoo","bill_length_mm: 42.7<br />body_mass_g: 3950<br />species: Gentoo","bill_length_mm: 49.6<br />body_mass_g: 5700<br />species: Gentoo","bill_length_mm: 45.3<br />body_mass_g: 4300<br />species: Gentoo","bill_length_mm: 49.6<br />body_mass_g: 4750<br />species: Gentoo","bill_length_mm: 50.5<br />body_mass_g: 5550<br />species: Gentoo","bill_length_mm: 43.6<br />body_mass_g: 4900<br />species: Gentoo","bill_length_mm: 45.5<br />body_mass_g: 4200<br />species: Gentoo","bill_length_mm: 50.5<br />body_mass_g: 5400<br />species: Gentoo","bill_length_mm: 44.9<br />body_mass_g: 5100<br />species: Gentoo","bill_length_mm: 45.2<br />body_mass_g: 5300<br />species: Gentoo","bill_length_mm: 46.6<br />body_mass_g: 4850<br />species: Gentoo","bill_length_mm: 48.5<br />body_mass_g: 5300<br />species: Gentoo","bill_length_mm: 45.1<br />body_mass_g: 4400<br />species: Gentoo","bill_length_mm: 50.1<br />body_mass_g: 5000<br />species: Gentoo","bill_length_mm: 46.5<br />body_mass_g: 4900<br />species: Gentoo","bill_length_mm: 45.0<br />body_mass_g: 5050<br />species: Gentoo","bill_length_mm: 43.8<br />body_mass_g: 4300<br />species: Gentoo","bill_length_mm: 45.5<br />body_mass_g: 5000<br />species: Gentoo","bill_length_mm: 43.2<br />body_mass_g: 4450<br />species: Gentoo","bill_length_mm: 50.4<br />body_mass_g: 5550<br />species: Gentoo","bill_length_mm: 45.3<br />body_mass_g: 4200<br />species: Gentoo","bill_length_mm: 46.2<br />body_mass_g: 5300<br />species: Gentoo","bill_length_mm: 45.7<br />body_mass_g: 4400<br />species: Gentoo","bill_length_mm: 54.3<br />body_mass_g: 5650<br />species: Gentoo","bill_length_mm: 45.8<br />body_mass_g: 4700<br />species: Gentoo","bill_length_mm: 49.8<br />body_mass_g: 5700<br />species: Gentoo","bill_length_mm: 46.2<br />body_mass_g: 4650<br />species: Gentoo","bill_length_mm: 49.5<br />body_mass_g: 5800<br />species: Gentoo","bill_length_mm: 43.5<br />body_mass_g: 4700<br />species: Gentoo","bill_length_mm: 50.7<br />body_mass_g: 5550<br />species: Gentoo","bill_length_mm: 47.7<br />body_mass_g: 4750<br />species: Gentoo","bill_length_mm: 46.4<br />body_mass_g: 5000<br />species: Gentoo","bill_length_mm: 48.2<br />body_mass_g: 5100<br />species: Gentoo","bill_length_mm: 46.5<br />body_mass_g: 5200<br />species: Gentoo","bill_length_mm: 46.4<br />body_mass_g: 4700<br />species: Gentoo","bill_length_mm: 48.6<br />body_mass_g: 5800<br />species: Gentoo","bill_length_mm: 47.5<br />body_mass_g: 4600<br />species: Gentoo","bill_length_mm: 51.1<br />body_mass_g: 6000<br />species: Gentoo","bill_length_mm: 45.2<br />body_mass_g: 4750<br />species: Gentoo","bill_length_mm: 45.2<br />body_mass_g: 5950<br />species: Gentoo","bill_length_mm: 49.1<br />body_mass_g: 4625<br />species: Gentoo","bill_length_mm: 52.5<br />body_mass_g: 5450<br />species: Gentoo","bill_length_mm: 47.4<br />body_mass_g: 4725<br />species: Gentoo","bill_length_mm: 50.0<br />body_mass_g: 5350<br />species: Gentoo","bill_length_mm: 44.9<br />body_mass_g: 4750<br />species: Gentoo","bill_length_mm: 50.8<br />body_mass_g: 5600<br />species: Gentoo","bill_length_mm: 43.4<br />body_mass_g: 4600<br />species: Gentoo","bill_length_mm: 51.3<br />body_mass_g: 5300<br />species: Gentoo","bill_length_mm: 47.5<br />body_mass_g: 4875<br />species: Gentoo","bill_length_mm: 52.1<br />body_mass_g: 5550<br />species: Gentoo","bill_length_mm: 47.5<br />body_mass_g: 4950<br />species: Gentoo","bill_length_mm: 52.2<br />body_mass_g: 5400<br />species: Gentoo","bill_length_mm: 45.5<br />body_mass_g: 4750<br />species: Gentoo","bill_length_mm: 49.5<br />body_mass_g: 5650<br />species: Gentoo","bill_length_mm: 44.5<br />body_mass_g: 4850<br />species: Gentoo","bill_length_mm: 50.8<br />body_mass_g: 5200<br />species: Gentoo","bill_length_mm: 49.4<br />body_mass_g: 4925<br />species: Gentoo","bill_length_mm: 46.9<br />body_mass_g: 4875<br />species: Gentoo","bill_length_mm: 48.4<br />body_mass_g: 4625<br />species: Gentoo","bill_length_mm: 51.1<br />body_mass_g: 5250<br />species: Gentoo","bill_length_mm: 48.5<br />body_mass_g: 4850<br />species: Gentoo","bill_length_mm: 55.9<br />body_mass_g: 5600<br />species: Gentoo","bill_length_mm: 47.2<br />body_mass_g: 4975<br />species: Gentoo","bill_length_mm: 49.1<br />body_mass_g: 5500<br />species: Gentoo","bill_length_mm: 47.3<br />body_mass_g: 4725<br />species: Gentoo","bill_length_mm: 46.8<br />body_mass_g: 5500<br />species: Gentoo","bill_length_mm: 41.7<br />body_mass_g: 4700<br />species: Gentoo","bill_length_mm: 53.4<br />body_mass_g: 5500<br />species: Gentoo","bill_length_mm: 43.3<br />body_mass_g: 4575<br />species: Gentoo","bill_length_mm: 48.1<br />body_mass_g: 5500<br />species: Gentoo","bill_length_mm: 50.5<br />body_mass_g: 5000<br />species: Gentoo","bill_length_mm: 49.8<br />body_mass_g: 5950<br />species: Gentoo","bill_length_mm: 43.5<br />body_mass_g: 4650<br />species: Gentoo","bill_length_mm: 51.5<br />body_mass_g: 5500<br />species: Gentoo","bill_length_mm: 46.2<br />body_mass_g: 4375<br />species: Gentoo","bill_length_mm: 55.1<br />body_mass_g: 5850<br />species: Gentoo","bill_length_mm: 44.5<br />body_mass_g: 4875<br />species: Gentoo","bill_length_mm: 48.8<br />body_mass_g: 6000<br />species: Gentoo","bill_length_mm: 47.2<br />body_mass_g: 4925<br />species: Gentoo","bill_length_mm: NA<br />body_mass_g: NA<br />species: Gentoo","bill_length_mm: 46.8<br />body_mass_g: 4850<br />species: Gentoo","bill_length_mm: 50.4<br />body_mass_g: 5750<br />species: Gentoo","bill_length_mm: 45.2<br />body_mass_g: 5200<br />species: Gentoo","bill_length_mm: 49.9<br />body_mass_g: 5400<br />species: Gentoo"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(97,156,255,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(97,156,255,1)"}},"hoveron":"points","name":"Gentoo","legendgroup":"Gentoo","showlegend":true,"xaxis":"x","yaxis":"y","hoverinfo":"text","frame":null}],"layout":{"margin":{"t":28.7853881278539,"r":7.30593607305936,"b":56.2889165628892,"l":69.4063926940639},"plot_bgcolor":"rgba(255,255,255,1)","paper_bgcolor":"rgba(255,255,255,1)","font":{"color":"rgba(0,0,0,1)","family":"","size":14.6118721461187},"xaxis":{"domain":[0,1],"automargin":true,"type":"linear","autorange":false,"range":[30.725,60.975],"tickmode":"array","ticktext":["40","50","60"],"tickvals":[40,50,60],"categoryorder":"array","categoryarray":["40","50","60"],"nticks":null,"ticks":"outside","tickcolor":"rgba(51,51,51,1)","ticklen":3.65296803652968,"tickwidth":0.66417600664176,"showticklabels":true,"tickfont":{"color":"rgba(77,77,77,1)","family":"","size":18.5969281859693},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"y","title":{"text":"bill_length_mm","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis":{"domain":[0,1],"automargin":true,"type":"linear","autorange":false,"range":[2520,6480],"tickmode":"array","ticktext":["3000","4000","5000","6000"],"tickvals":[3000,4000,5000,6000],"categoryorder":"array","categoryarray":["3000","4000","5000","6000"],"nticks":null,"ticks":"outside","tickcolor":"rgba(51,51,51,1)","ticklen":3.65296803652968,"tickwidth":0.66417600664176,"showticklabels":true,"tickfont":{"color":"rgba(77,77,77,1)","family":"","size":18.5969281859693},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x","title":{"text":"body_mass_g","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"shapes":[{"type":"rect","fillcolor":null,"line":{"color":null,"width":0,"linetype":[]},"yref":"paper","xref":"paper","x0":0,"x1":1,"y0":0,"y1":1}],"showlegend":true,"legend":{"bgcolor":"rgba(255,255,255,1)","bordercolor":"transparent","borderwidth":1.88976377952756,"font":{"color":"rgba(0,0,0,1)","family":"","size":18.5969281859693},"y":0.84251968503937},"annotations":[{"text":"species","x":1.02,"y":1,"showarrow":false,"ax":0,"ay":0,"font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363},"xref":"paper","yref":"paper","textangle":-0,"xanchor":"left","yanchor":"bottom","legendTitle":true}],"hovermode":"closest","barmode":"relative"},"config":{"doubleClick":"reset","showSendToCloud":false},"source":"A","attrs":{"184107d24951f":{"x":{},"y":{},"colour":{},"type":"scatter"}},"cur_data":"184107d24951f","visdat":{"184107d24951f":["function (y) ","x"]},"highlight":{"on":"plotly_click","persistent":false,"dynamic":false,"selectize":false,"opacityDim":0.2,"selected":{"opacity":1},"debounce":0},"shinyEvents":["plotly_hover","plotly_click","plotly_selected","plotly_relayout","plotly_brushed","plotly_brushing","plotly_clickannotation","plotly_doubleclick","plotly_deselect","plotly_afterplot","plotly_sunburstclick"],"base_url":"https://plot.ly"},"evals":[],"jsHooks":[]}</script> ] --- ## Plotly You can do some customization on the tooltips (what shows up when you put your mouse over a point). ```r p <- ggplot(data = diamonds, aes(x = cut, fill = clarity))+ geom_bar(position = 'dodge') *ggplotly(p, tooltip = c('cut')) ``` <div id="htmlwidget-95c28ac76de6c1bc88ef" style="width:864px;height:360px;" class="plotly html-widget"></div> <script type="application/json" data-for="htmlwidget-95c28ac76de6c1bc88ef">{"x":{"data":[{"orientation":"v","width":[0.1125,0.1125,0.1125,0.1125,0.112500000000001],"base":[0,0,0,0,0],"x":[0.60625,1.60625,2.60625,3.60625,4.60625],"y":[210,96,84,205,146],"text":["cut: Fair","cut: Good","cut: Very Good","cut: Premium","cut: Ideal"],"type":"bar","marker":{"autocolorscale":false,"color":"rgba(68,1,84,1)","line":{"width":1.88976377952756,"color":"transparent"}},"name":"I1","legendgroup":"I1","showlegend":true,"xaxis":"x","yaxis":"y","hoverinfo":"text","frame":null},{"orientation":"v","width":[0.1125,0.1125,0.1125,0.1125,0.112500000000001],"base":[0,0,0,0,0],"x":[0.71875,1.71875,2.71875,3.71875,4.71875],"y":[466,1081,2100,2949,2598],"text":["cut: Fair","cut: Good","cut: Very Good","cut: Premium","cut: Ideal"],"type":"bar","marker":{"autocolorscale":false,"color":"rgba(70,51,126,1)","line":{"width":1.88976377952756,"color":"transparent"}},"name":"SI2","legendgroup":"SI2","showlegend":true,"xaxis":"x","yaxis":"y","hoverinfo":"text","frame":null},{"orientation":"v","width":[0.1125,0.1125,0.1125,0.1125,0.112500000000001],"base":[0,0,0,0,0],"x":[0.83125,1.83125,2.83125,3.83125,4.83125],"y":[408,1560,3240,3575,4282],"text":["cut: Fair","cut: Good","cut: Very Good","cut: Premium","cut: Ideal"],"type":"bar","marker":{"autocolorscale":false,"color":"rgba(54,92,141,1)","line":{"width":1.88976377952756,"color":"transparent"}},"name":"SI1","legendgroup":"SI1","showlegend":true,"xaxis":"x","yaxis":"y","hoverinfo":"text","frame":null},{"orientation":"v","width":[0.1125,0.1125,0.1125,0.1125,0.112500000000001],"base":[0,0,0,0,0],"x":[0.94375,1.94375,2.94375,3.94375,4.94375],"y":[261,978,2591,3357,5071],"text":["cut: Fair","cut: Good","cut: Very Good","cut: Premium","cut: Ideal"],"type":"bar","marker":{"autocolorscale":false,"color":"rgba(39,127,142,1)","line":{"width":1.88976377952756,"color":"transparent"}},"name":"VS2","legendgroup":"VS2","showlegend":true,"xaxis":"x","yaxis":"y","hoverinfo":"text","frame":null},{"orientation":"v","width":[0.1125,0.1125,0.1125,0.112500000000001,0.112500000000001],"base":[0,0,0,0,0],"x":[1.05625,2.05625,3.05625,4.05625,5.05625],"y":[170,648,1775,1989,3589],"text":["cut: Fair","cut: Good","cut: Very Good","cut: Premium","cut: Ideal"],"type":"bar","marker":{"autocolorscale":false,"color":"rgba(31,161,135,1)","line":{"width":1.88976377952756,"color":"transparent"}},"name":"VS1","legendgroup":"VS1","showlegend":true,"xaxis":"x","yaxis":"y","hoverinfo":"text","frame":null},{"orientation":"v","width":[0.1125,0.1125,0.1125,0.112500000000001,0.112500000000001],"base":[0,0,0,0,0],"x":[1.16875,2.16875,3.16875,4.16875,5.16875],"y":[69,286,1235,870,2606],"text":["cut: Fair","cut: Good","cut: Very Good","cut: Premium","cut: Ideal"],"type":"bar","marker":{"autocolorscale":false,"color":"rgba(74,193,109,1)","line":{"width":1.88976377952756,"color":"transparent"}},"name":"VVS2","legendgroup":"VVS2","showlegend":true,"xaxis":"x","yaxis":"y","hoverinfo":"text","frame":null},{"orientation":"v","width":[0.1125,0.1125,0.1125,0.112500000000001,0.112500000000001],"base":[0,0,0,0,0],"x":[1.28125,2.28125,3.28125,4.28125,5.28125],"y":[17,186,789,616,2047],"text":["cut: Fair","cut: Good","cut: Very Good","cut: Premium","cut: Ideal"],"type":"bar","marker":{"autocolorscale":false,"color":"rgba(159,218,58,1)","line":{"width":1.88976377952756,"color":"transparent"}},"name":"VVS1","legendgroup":"VVS1","showlegend":true,"xaxis":"x","yaxis":"y","hoverinfo":"text","frame":null},{"orientation":"v","width":[0.1125,0.1125,0.1125,0.112500000000001,0.112500000000001],"base":[0,0,0,0,0],"x":[1.39375,2.39375,3.39375,4.39375,5.39375],"y":[9,71,268,230,1212],"text":["cut: Fair","cut: Good","cut: Very Good","cut: Premium","cut: Ideal"],"type":"bar","marker":{"autocolorscale":false,"color":"rgba(253,231,37,1)","line":{"width":1.88976377952756,"color":"transparent"}},"name":"IF","legendgroup":"IF","showlegend":true,"xaxis":"x","yaxis":"y","hoverinfo":"text","frame":null}],"layout":{"margin":{"t":33.5342465753425,"r":7.30593607305936,"b":61.0377750103777,"l":69.4063926940639},"plot_bgcolor":"rgba(255,255,255,1)","paper_bgcolor":"rgba(255,255,255,1)","font":{"color":"rgba(0,0,0,1)","family":"","size":14.6118721461187},"xaxis":{"domain":[0,1],"automargin":true,"type":"linear","autorange":false,"range":[0.4,5.6],"tickmode":"array","ticktext":["Fair","Good","Very Good","Premium","Ideal"],"tickvals":[1,2,3,4,5],"categoryorder":"array","categoryarray":["Fair","Good","Very Good","Premium","Ideal"],"nticks":null,"ticks":"outside","tickcolor":"rgba(51,51,51,1)","ticklen":3.65296803652968,"tickwidth":0.66417600664176,"showticklabels":true,"tickfont":{"color":"rgba(77,77,77,1)","family":"","size":18.5969281859693},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"y","title":{"text":"cut","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis":{"domain":[0,1],"automargin":true,"type":"linear","autorange":false,"range":[-253.55,5324.55],"tickmode":"array","ticktext":["0","1000","2000","3000","4000","5000"],"tickvals":[0,1000,2000,3000,4000,5000],"categoryorder":"array","categoryarray":["0","1000","2000","3000","4000","5000"],"nticks":null,"ticks":"outside","tickcolor":"rgba(51,51,51,1)","ticklen":3.65296803652968,"tickwidth":0.66417600664176,"showticklabels":true,"tickfont":{"color":"rgba(77,77,77,1)","family":"","size":18.5969281859693},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x","title":{"text":"count","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"shapes":[{"type":"rect","fillcolor":null,"line":{"color":null,"width":0,"linetype":[]},"yref":"paper","xref":"paper","x0":0,"x1":1,"y0":0,"y1":1}],"showlegend":true,"legend":{"bgcolor":"rgba(255,255,255,1)","bordercolor":"transparent","borderwidth":1.88976377952756,"font":{"color":"rgba(0,0,0,1)","family":"","size":18.5969281859693},"y":0.874015748031496},"annotations":[{"text":"clarity","x":1.02,"y":1,"showarrow":false,"ax":0,"ay":0,"font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363},"xref":"paper","yref":"paper","textangle":-0,"xanchor":"left","yanchor":"bottom","legendTitle":true}],"hovermode":"closest","barmode":"relative"},"config":{"doubleClick":"reset","showSendToCloud":false},"source":"A","attrs":{"184106d4e8047":{"x":{},"fill":{},"type":"bar"}},"cur_data":"184106d4e8047","visdat":{"184106d4e8047":["function (y) ","x"]},"highlight":{"on":"plotly_click","persistent":false,"dynamic":false,"selectize":false,"opacityDim":0.2,"selected":{"opacity":1},"debounce":0},"shinyEvents":["plotly_hover","plotly_click","plotly_selected","plotly_relayout","plotly_brushed","plotly_brushing","plotly_clickannotation","plotly_doubleclick","plotly_deselect","plotly_afterplot","plotly_sunburstclick"],"base_url":"https://plot.ly"},"evals":[],"jsHooks":[]}</script> --- ## Plotly You can do linked plots in plotly, so interactions in one plot are reflected in a second plot. This is called _brushing_. The key here is to use `highlight_key`, which allows a data frame to be shared between multiple plots at the same time. ```r d <- highlight_key(penguins) plt1 <- ggplot(d, aes(x = bill_length_mm, y = body_mass_g))+geom_point() plt2 <- ggplot(d, aes(x = bill_length_mm, y = flipper_length_mm))+geom_point() subplot(plt1, plt2) %>% layout(title = "Click and drag to select points") %>% highlight("plotly_selected") ``` <div id="htmlwidget-ffae68c3d6fbd06732ec" style="width:648px;height:288px;" class="plotly html-widget"></div> <script type="application/json" data-for="htmlwidget-ffae68c3d6fbd06732ec">{"x":{"data":[{"x":[39.1,39.5,40.3,null,36.7,39.3,38.9,39.2,34.1,42,37.8,37.8,41.1,38.6,34.6,36.6,38.7,42.5,34.4,46,37.8,37.7,35.9,38.2,38.8,35.3,40.6,40.5,37.9,40.5,39.5,37.2,39.5,40.9,36.4,39.2,38.8,42.2,37.6,39.8,36.5,40.8,36,44.1,37,39.6,41.1,37.5,36,42.3,39.6,40.1,35,42,34.5,41.4,39,40.6,36.5,37.6,35.7,41.3,37.6,41.1,36.4,41.6,35.5,41.1,35.9,41.8,33.5,39.7,39.6,45.8,35.5,42.8,40.9,37.2,36.2,42.1,34.6,42.9,36.7,35.1,37.3,41.3,36.3,36.9,38.3,38.9,35.7,41.1,34,39.6,36.2,40.8,38.1,40.3,33.1,43.2,35,41,37.7,37.8,37.9,39.7,38.6,38.2,38.1,43.2,38.1,45.6,39.7,42.2,39.6,42.7,38.6,37.3,35.7,41.1,36.2,37.7,40.2,41.4,35.2,40.6,38.8,41.5,39,44.1,38.5,43.1,36.8,37.5,38.1,41.1,35.6,40.2,37,39.7,40.2,40.6,32.1,40.7,37.3,39,39.2,36.6,36,37.8,36,41.5,46.1,50,48.7,50,47.6,46.5,45.4,46.7,43.3,46.8,40.9,49,45.5,48.4,45.8,49.3,42,49.2,46.2,48.7,50.2,45.1,46.5,46.3,42.9,46.1,44.5,47.8,48.2,50,47.3,42.8,45.1,59.6,49.1,48.4,42.6,44.4,44,48.7,42.7,49.6,45.3,49.6,50.5,43.6,45.5,50.5,44.9,45.2,46.6,48.5,45.1,50.1,46.5,45,43.8,45.5,43.2,50.4,45.3,46.2,45.7,54.3,45.8,49.8,46.2,49.5,43.5,50.7,47.7,46.4,48.2,46.5,46.4,48.6,47.5,51.1,45.2,45.2,49.1,52.5,47.4,50,44.9,50.8,43.4,51.3,47.5,52.1,47.5,52.2,45.5,49.5,44.5,50.8,49.4,46.9,48.4,51.1,48.5,55.9,47.2,49.1,47.3,46.8,41.7,53.4,43.3,48.1,50.5,49.8,43.5,51.5,46.2,55.1,44.5,48.8,47.2,null,46.8,50.4,45.2,49.9,46.5,50,51.3,45.4,52.7,45.2,46.1,51.3,46,51.3,46.6,51.7,47,52,45.9,50.5,50.3,58,46.4,49.2,42.4,48.5,43.2,50.6,46.7,52,50.5,49.5,46.4,52.8,40.9,54.2,42.5,51,49.7,47.5,47.6,52,46.9,53.5,49,46.2,50.9,45.5,50.9,50.8,50.1,49,51.5,49.8,48.1,51.4,45.7,50.7,42.5,52.2,45.2,49.3,50.2,45.6,51.9,46.8,45.7,55.8,43.5,49.6,50.8,50.2],"y":[3750,3800,3250,null,3450,3650,3625,4675,3475,4250,3300,3700,3200,3800,4400,3700,3450,4500,3325,4200,3400,3600,3800,3950,3800,3800,3550,3200,3150,3950,3250,3900,3300,3900,3325,4150,3950,3550,3300,4650,3150,3900,3100,4400,3000,4600,3425,2975,3450,4150,3500,4300,3450,4050,2900,3700,3550,3800,2850,3750,3150,4400,3600,4050,2850,3950,3350,4100,3050,4450,3600,3900,3550,4150,3700,4250,3700,3900,3550,4000,3200,4700,3800,4200,3350,3550,3800,3500,3950,3600,3550,4300,3400,4450,3300,4300,3700,4350,2900,4100,3725,4725,3075,4250,2925,3550,3750,3900,3175,4775,3825,4600,3200,4275,3900,4075,2900,3775,3350,3325,3150,3500,3450,3875,3050,4000,3275,4300,3050,4000,3325,3500,3500,4475,3425,3900,3175,3975,3400,4250,3400,3475,3050,3725,3000,3650,4250,3475,3450,3750,3700,4000,4500,5700,4450,5700,5400,4550,4800,5200,4400,5150,4650,5550,4650,5850,4200,5850,4150,6300,4800,5350,5700,5000,4400,5050,5000,5100,4100,5650,4600,5550,5250,4700,5050,6050,5150,5400,4950,5250,4350,5350,3950,5700,4300,4750,5550,4900,4200,5400,5100,5300,4850,5300,4400,5000,4900,5050,4300,5000,4450,5550,4200,5300,4400,5650,4700,5700,4650,5800,4700,5550,4750,5000,5100,5200,4700,5800,4600,6000,4750,5950,4625,5450,4725,5350,4750,5600,4600,5300,4875,5550,4950,5400,4750,5650,4850,5200,4925,4875,4625,5250,4850,5600,4975,5500,4725,5500,4700,5500,4575,5500,5000,5950,4650,5500,4375,5850,4875,6000,4925,null,4850,5750,5200,5400,3500,3900,3650,3525,3725,3950,3250,3750,4150,3700,3800,3775,3700,4050,3575,4050,3300,3700,3450,4400,3600,3400,2900,3800,3300,4150,3400,3800,3700,4550,3200,4300,3350,4100,3600,3900,3850,4800,2700,4500,3950,3650,3550,3500,3675,4450,3400,4300,3250,3675,3325,3950,3600,4050,3350,3450,3250,4050,3800,3525,3950,3650,3650,4000,3400,3775,4100,3775],"text":["bill_length_mm: 39.1<br />body_mass_g: 3750","bill_length_mm: 39.5<br />body_mass_g: 3800","bill_length_mm: 40.3<br />body_mass_g: 3250","bill_length_mm: NA<br />body_mass_g: NA","bill_length_mm: 36.7<br />body_mass_g: 3450","bill_length_mm: 39.3<br />body_mass_g: 3650","bill_length_mm: 38.9<br />body_mass_g: 3625","bill_length_mm: 39.2<br />body_mass_g: 4675","bill_length_mm: 34.1<br />body_mass_g: 3475","bill_length_mm: 42.0<br />body_mass_g: 4250","bill_length_mm: 37.8<br />body_mass_g: 3300","bill_length_mm: 37.8<br />body_mass_g: 3700","bill_length_mm: 41.1<br />body_mass_g: 3200","bill_length_mm: 38.6<br />body_mass_g: 3800","bill_length_mm: 34.6<br />body_mass_g: 4400","bill_length_mm: 36.6<br />body_mass_g: 3700","bill_length_mm: 38.7<br />body_mass_g: 3450","bill_length_mm: 42.5<br />body_mass_g: 4500","bill_length_mm: 34.4<br />body_mass_g: 3325","bill_length_mm: 46.0<br />body_mass_g: 4200","bill_length_mm: 37.8<br />body_mass_g: 3400","bill_length_mm: 37.7<br />body_mass_g: 3600","bill_length_mm: 35.9<br />body_mass_g: 3800","bill_length_mm: 38.2<br />body_mass_g: 3950","bill_length_mm: 38.8<br />body_mass_g: 3800","bill_length_mm: 35.3<br />body_mass_g: 3800","bill_length_mm: 40.6<br />body_mass_g: 3550","bill_length_mm: 40.5<br />body_mass_g: 3200","bill_length_mm: 37.9<br />body_mass_g: 3150","bill_length_mm: 40.5<br />body_mass_g: 3950","bill_length_mm: 39.5<br />body_mass_g: 3250","bill_length_mm: 37.2<br />body_mass_g: 3900","bill_length_mm: 39.5<br />body_mass_g: 3300","bill_length_mm: 40.9<br />body_mass_g: 3900","bill_length_mm: 36.4<br />body_mass_g: 3325","bill_length_mm: 39.2<br />body_mass_g: 4150","bill_length_mm: 38.8<br />body_mass_g: 3950","bill_length_mm: 42.2<br />body_mass_g: 3550","bill_length_mm: 37.6<br />body_mass_g: 3300","bill_length_mm: 39.8<br />body_mass_g: 4650","bill_length_mm: 36.5<br />body_mass_g: 3150","bill_length_mm: 40.8<br />body_mass_g: 3900","bill_length_mm: 36.0<br />body_mass_g: 3100","bill_length_mm: 44.1<br />body_mass_g: 4400","bill_length_mm: 37.0<br />body_mass_g: 3000","bill_length_mm: 39.6<br />body_mass_g: 4600","bill_length_mm: 41.1<br />body_mass_g: 3425","bill_length_mm: 37.5<br />body_mass_g: 2975","bill_length_mm: 36.0<br />body_mass_g: 3450","bill_length_mm: 42.3<br />body_mass_g: 4150","bill_length_mm: 39.6<br />body_mass_g: 3500","bill_length_mm: 40.1<br />body_mass_g: 4300","bill_length_mm: 35.0<br />body_mass_g: 3450","bill_length_mm: 42.0<br />body_mass_g: 4050","bill_length_mm: 34.5<br />body_mass_g: 2900","bill_length_mm: 41.4<br />body_mass_g: 3700","bill_length_mm: 39.0<br />body_mass_g: 3550","bill_length_mm: 40.6<br />body_mass_g: 3800","bill_length_mm: 36.5<br />body_mass_g: 2850","bill_length_mm: 37.6<br />body_mass_g: 3750","bill_length_mm: 35.7<br />body_mass_g: 3150","bill_length_mm: 41.3<br />body_mass_g: 4400","bill_length_mm: 37.6<br />body_mass_g: 3600","bill_length_mm: 41.1<br />body_mass_g: 4050","bill_length_mm: 36.4<br />body_mass_g: 2850","bill_length_mm: 41.6<br />body_mass_g: 3950","bill_length_mm: 35.5<br />body_mass_g: 3350","bill_length_mm: 41.1<br />body_mass_g: 4100","bill_length_mm: 35.9<br />body_mass_g: 3050","bill_length_mm: 41.8<br />body_mass_g: 4450","bill_length_mm: 33.5<br />body_mass_g: 3600","bill_length_mm: 39.7<br />body_mass_g: 3900","bill_length_mm: 39.6<br />body_mass_g: 3550","bill_length_mm: 45.8<br />body_mass_g: 4150","bill_length_mm: 35.5<br />body_mass_g: 3700","bill_length_mm: 42.8<br />body_mass_g: 4250","bill_length_mm: 40.9<br />body_mass_g: 3700","bill_length_mm: 37.2<br />body_mass_g: 3900","bill_length_mm: 36.2<br />body_mass_g: 3550","bill_length_mm: 42.1<br />body_mass_g: 4000","bill_length_mm: 34.6<br />body_mass_g: 3200","bill_length_mm: 42.9<br />body_mass_g: 4700","bill_length_mm: 36.7<br />body_mass_g: 3800","bill_length_mm: 35.1<br />body_mass_g: 4200","bill_length_mm: 37.3<br />body_mass_g: 3350","bill_length_mm: 41.3<br />body_mass_g: 3550","bill_length_mm: 36.3<br />body_mass_g: 3800","bill_length_mm: 36.9<br />body_mass_g: 3500","bill_length_mm: 38.3<br />body_mass_g: 3950","bill_length_mm: 38.9<br />body_mass_g: 3600","bill_length_mm: 35.7<br />body_mass_g: 3550","bill_length_mm: 41.1<br />body_mass_g: 4300","bill_length_mm: 34.0<br />body_mass_g: 3400","bill_length_mm: 39.6<br />body_mass_g: 4450","bill_length_mm: 36.2<br />body_mass_g: 3300","bill_length_mm: 40.8<br />body_mass_g: 4300","bill_length_mm: 38.1<br />body_mass_g: 3700","bill_length_mm: 40.3<br />body_mass_g: 4350","bill_length_mm: 33.1<br />body_mass_g: 2900","bill_length_mm: 43.2<br />body_mass_g: 4100","bill_length_mm: 35.0<br />body_mass_g: 3725","bill_length_mm: 41.0<br />body_mass_g: 4725","bill_length_mm: 37.7<br />body_mass_g: 3075","bill_length_mm: 37.8<br />body_mass_g: 4250","bill_length_mm: 37.9<br />body_mass_g: 2925","bill_length_mm: 39.7<br />body_mass_g: 3550","bill_length_mm: 38.6<br />body_mass_g: 3750","bill_length_mm: 38.2<br />body_mass_g: 3900","bill_length_mm: 38.1<br />body_mass_g: 3175","bill_length_mm: 43.2<br />body_mass_g: 4775","bill_length_mm: 38.1<br />body_mass_g: 3825","bill_length_mm: 45.6<br />body_mass_g: 4600","bill_length_mm: 39.7<br />body_mass_g: 3200","bill_length_mm: 42.2<br />body_mass_g: 4275","bill_length_mm: 39.6<br />body_mass_g: 3900","bill_length_mm: 42.7<br />body_mass_g: 4075","bill_length_mm: 38.6<br />body_mass_g: 2900","bill_length_mm: 37.3<br />body_mass_g: 3775","bill_length_mm: 35.7<br />body_mass_g: 3350","bill_length_mm: 41.1<br />body_mass_g: 3325","bill_length_mm: 36.2<br />body_mass_g: 3150","bill_length_mm: 37.7<br />body_mass_g: 3500","bill_length_mm: 40.2<br />body_mass_g: 3450","bill_length_mm: 41.4<br />body_mass_g: 3875","bill_length_mm: 35.2<br />body_mass_g: 3050","bill_length_mm: 40.6<br />body_mass_g: 4000","bill_length_mm: 38.8<br />body_mass_g: 3275","bill_length_mm: 41.5<br />body_mass_g: 4300","bill_length_mm: 39.0<br />body_mass_g: 3050","bill_length_mm: 44.1<br />body_mass_g: 4000","bill_length_mm: 38.5<br />body_mass_g: 3325","bill_length_mm: 43.1<br />body_mass_g: 3500","bill_length_mm: 36.8<br />body_mass_g: 3500","bill_length_mm: 37.5<br />body_mass_g: 4475","bill_length_mm: 38.1<br />body_mass_g: 3425","bill_length_mm: 41.1<br />body_mass_g: 3900","bill_length_mm: 35.6<br />body_mass_g: 3175","bill_length_mm: 40.2<br />body_mass_g: 3975","bill_length_mm: 37.0<br />body_mass_g: 3400","bill_length_mm: 39.7<br />body_mass_g: 4250","bill_length_mm: 40.2<br />body_mass_g: 3400","bill_length_mm: 40.6<br />body_mass_g: 3475","bill_length_mm: 32.1<br />body_mass_g: 3050","bill_length_mm: 40.7<br />body_mass_g: 3725","bill_length_mm: 37.3<br />body_mass_g: 3000","bill_length_mm: 39.0<br />body_mass_g: 3650","bill_length_mm: 39.2<br />body_mass_g: 4250","bill_length_mm: 36.6<br />body_mass_g: 3475","bill_length_mm: 36.0<br />body_mass_g: 3450","bill_length_mm: 37.8<br />body_mass_g: 3750","bill_length_mm: 36.0<br />body_mass_g: 3700","bill_length_mm: 41.5<br />body_mass_g: 4000","bill_length_mm: 46.1<br />body_mass_g: 4500","bill_length_mm: 50.0<br />body_mass_g: 5700","bill_length_mm: 48.7<br />body_mass_g: 4450","bill_length_mm: 50.0<br />body_mass_g: 5700","bill_length_mm: 47.6<br />body_mass_g: 5400","bill_length_mm: 46.5<br />body_mass_g: 4550","bill_length_mm: 45.4<br />body_mass_g: 4800","bill_length_mm: 46.7<br />body_mass_g: 5200","bill_length_mm: 43.3<br />body_mass_g: 4400","bill_length_mm: 46.8<br />body_mass_g: 5150","bill_length_mm: 40.9<br />body_mass_g: 4650","bill_length_mm: 49.0<br />body_mass_g: 5550","bill_length_mm: 45.5<br />body_mass_g: 4650","bill_length_mm: 48.4<br />body_mass_g: 5850","bill_length_mm: 45.8<br />body_mass_g: 4200","bill_length_mm: 49.3<br />body_mass_g: 5850","bill_length_mm: 42.0<br />body_mass_g: 4150","bill_length_mm: 49.2<br />body_mass_g: 6300","bill_length_mm: 46.2<br />body_mass_g: 4800","bill_length_mm: 48.7<br />body_mass_g: 5350","bill_length_mm: 50.2<br />body_mass_g: 5700","bill_length_mm: 45.1<br />body_mass_g: 5000","bill_length_mm: 46.5<br />body_mass_g: 4400","bill_length_mm: 46.3<br />body_mass_g: 5050","bill_length_mm: 42.9<br />body_mass_g: 5000","bill_length_mm: 46.1<br />body_mass_g: 5100","bill_length_mm: 44.5<br />body_mass_g: 4100","bill_length_mm: 47.8<br />body_mass_g: 5650","bill_length_mm: 48.2<br />body_mass_g: 4600","bill_length_mm: 50.0<br />body_mass_g: 5550","bill_length_mm: 47.3<br />body_mass_g: 5250","bill_length_mm: 42.8<br />body_mass_g: 4700","bill_length_mm: 45.1<br />body_mass_g: 5050","bill_length_mm: 59.6<br />body_mass_g: 6050","bill_length_mm: 49.1<br />body_mass_g: 5150","bill_length_mm: 48.4<br />body_mass_g: 5400","bill_length_mm: 42.6<br />body_mass_g: 4950","bill_length_mm: 44.4<br />body_mass_g: 5250","bill_length_mm: 44.0<br />body_mass_g: 4350","bill_length_mm: 48.7<br />body_mass_g: 5350","bill_length_mm: 42.7<br />body_mass_g: 3950","bill_length_mm: 49.6<br />body_mass_g: 5700","bill_length_mm: 45.3<br />body_mass_g: 4300","bill_length_mm: 49.6<br />body_mass_g: 4750","bill_length_mm: 50.5<br />body_mass_g: 5550","bill_length_mm: 43.6<br />body_mass_g: 4900","bill_length_mm: 45.5<br />body_mass_g: 4200","bill_length_mm: 50.5<br />body_mass_g: 5400","bill_length_mm: 44.9<br />body_mass_g: 5100","bill_length_mm: 45.2<br />body_mass_g: 5300","bill_length_mm: 46.6<br />body_mass_g: 4850","bill_length_mm: 48.5<br />body_mass_g: 5300","bill_length_mm: 45.1<br />body_mass_g: 4400","bill_length_mm: 50.1<br />body_mass_g: 5000","bill_length_mm: 46.5<br />body_mass_g: 4900","bill_length_mm: 45.0<br />body_mass_g: 5050","bill_length_mm: 43.8<br />body_mass_g: 4300","bill_length_mm: 45.5<br />body_mass_g: 5000","bill_length_mm: 43.2<br />body_mass_g: 4450","bill_length_mm: 50.4<br />body_mass_g: 5550","bill_length_mm: 45.3<br />body_mass_g: 4200","bill_length_mm: 46.2<br />body_mass_g: 5300","bill_length_mm: 45.7<br />body_mass_g: 4400","bill_length_mm: 54.3<br />body_mass_g: 5650","bill_length_mm: 45.8<br />body_mass_g: 4700","bill_length_mm: 49.8<br />body_mass_g: 5700","bill_length_mm: 46.2<br />body_mass_g: 4650","bill_length_mm: 49.5<br />body_mass_g: 5800","bill_length_mm: 43.5<br />body_mass_g: 4700","bill_length_mm: 50.7<br />body_mass_g: 5550","bill_length_mm: 47.7<br />body_mass_g: 4750","bill_length_mm: 46.4<br />body_mass_g: 5000","bill_length_mm: 48.2<br />body_mass_g: 5100","bill_length_mm: 46.5<br />body_mass_g: 5200","bill_length_mm: 46.4<br />body_mass_g: 4700","bill_length_mm: 48.6<br />body_mass_g: 5800","bill_length_mm: 47.5<br />body_mass_g: 4600","bill_length_mm: 51.1<br />body_mass_g: 6000","bill_length_mm: 45.2<br />body_mass_g: 4750","bill_length_mm: 45.2<br />body_mass_g: 5950","bill_length_mm: 49.1<br />body_mass_g: 4625","bill_length_mm: 52.5<br />body_mass_g: 5450","bill_length_mm: 47.4<br />body_mass_g: 4725","bill_length_mm: 50.0<br />body_mass_g: 5350","bill_length_mm: 44.9<br />body_mass_g: 4750","bill_length_mm: 50.8<br />body_mass_g: 5600","bill_length_mm: 43.4<br />body_mass_g: 4600","bill_length_mm: 51.3<br />body_mass_g: 5300","bill_length_mm: 47.5<br />body_mass_g: 4875","bill_length_mm: 52.1<br />body_mass_g: 5550","bill_length_mm: 47.5<br />body_mass_g: 4950","bill_length_mm: 52.2<br />body_mass_g: 5400","bill_length_mm: 45.5<br />body_mass_g: 4750","bill_length_mm: 49.5<br />body_mass_g: 5650","bill_length_mm: 44.5<br />body_mass_g: 4850","bill_length_mm: 50.8<br />body_mass_g: 5200","bill_length_mm: 49.4<br />body_mass_g: 4925","bill_length_mm: 46.9<br />body_mass_g: 4875","bill_length_mm: 48.4<br />body_mass_g: 4625","bill_length_mm: 51.1<br />body_mass_g: 5250","bill_length_mm: 48.5<br />body_mass_g: 4850","bill_length_mm: 55.9<br />body_mass_g: 5600","bill_length_mm: 47.2<br />body_mass_g: 4975","bill_length_mm: 49.1<br />body_mass_g: 5500","bill_length_mm: 47.3<br />body_mass_g: 4725","bill_length_mm: 46.8<br />body_mass_g: 5500","bill_length_mm: 41.7<br />body_mass_g: 4700","bill_length_mm: 53.4<br />body_mass_g: 5500","bill_length_mm: 43.3<br />body_mass_g: 4575","bill_length_mm: 48.1<br />body_mass_g: 5500","bill_length_mm: 50.5<br />body_mass_g: 5000","bill_length_mm: 49.8<br />body_mass_g: 5950","bill_length_mm: 43.5<br />body_mass_g: 4650","bill_length_mm: 51.5<br />body_mass_g: 5500","bill_length_mm: 46.2<br />body_mass_g: 4375","bill_length_mm: 55.1<br />body_mass_g: 5850","bill_length_mm: 44.5<br />body_mass_g: 4875","bill_length_mm: 48.8<br />body_mass_g: 6000","bill_length_mm: 47.2<br />body_mass_g: 4925","bill_length_mm: NA<br />body_mass_g: NA","bill_length_mm: 46.8<br />body_mass_g: 4850","bill_length_mm: 50.4<br />body_mass_g: 5750","bill_length_mm: 45.2<br />body_mass_g: 5200","bill_length_mm: 49.9<br />body_mass_g: 5400","bill_length_mm: 46.5<br />body_mass_g: 3500","bill_length_mm: 50.0<br />body_mass_g: 3900","bill_length_mm: 51.3<br />body_mass_g: 3650","bill_length_mm: 45.4<br />body_mass_g: 3525","bill_length_mm: 52.7<br />body_mass_g: 3725","bill_length_mm: 45.2<br />body_mass_g: 3950","bill_length_mm: 46.1<br />body_mass_g: 3250","bill_length_mm: 51.3<br />body_mass_g: 3750","bill_length_mm: 46.0<br />body_mass_g: 4150","bill_length_mm: 51.3<br />body_mass_g: 3700","bill_length_mm: 46.6<br />body_mass_g: 3800","bill_length_mm: 51.7<br />body_mass_g: 3775","bill_length_mm: 47.0<br />body_mass_g: 3700","bill_length_mm: 52.0<br />body_mass_g: 4050","bill_length_mm: 45.9<br />body_mass_g: 3575","bill_length_mm: 50.5<br />body_mass_g: 4050","bill_length_mm: 50.3<br />body_mass_g: 3300","bill_length_mm: 58.0<br />body_mass_g: 3700","bill_length_mm: 46.4<br />body_mass_g: 3450","bill_length_mm: 49.2<br />body_mass_g: 4400","bill_length_mm: 42.4<br />body_mass_g: 3600","bill_length_mm: 48.5<br />body_mass_g: 3400","bill_length_mm: 43.2<br />body_mass_g: 2900","bill_length_mm: 50.6<br />body_mass_g: 3800","bill_length_mm: 46.7<br />body_mass_g: 3300","bill_length_mm: 52.0<br />body_mass_g: 4150","bill_length_mm: 50.5<br />body_mass_g: 3400","bill_length_mm: 49.5<br />body_mass_g: 3800","bill_length_mm: 46.4<br />body_mass_g: 3700","bill_length_mm: 52.8<br />body_mass_g: 4550","bill_length_mm: 40.9<br />body_mass_g: 3200","bill_length_mm: 54.2<br />body_mass_g: 4300","bill_length_mm: 42.5<br />body_mass_g: 3350","bill_length_mm: 51.0<br />body_mass_g: 4100","bill_length_mm: 49.7<br />body_mass_g: 3600","bill_length_mm: 47.5<br />body_mass_g: 3900","bill_length_mm: 47.6<br />body_mass_g: 3850","bill_length_mm: 52.0<br />body_mass_g: 4800","bill_length_mm: 46.9<br />body_mass_g: 2700","bill_length_mm: 53.5<br />body_mass_g: 4500","bill_length_mm: 49.0<br />body_mass_g: 3950","bill_length_mm: 46.2<br />body_mass_g: 3650","bill_length_mm: 50.9<br />body_mass_g: 3550","bill_length_mm: 45.5<br />body_mass_g: 3500","bill_length_mm: 50.9<br />body_mass_g: 3675","bill_length_mm: 50.8<br />body_mass_g: 4450","bill_length_mm: 50.1<br />body_mass_g: 3400","bill_length_mm: 49.0<br />body_mass_g: 4300","bill_length_mm: 51.5<br />body_mass_g: 3250","bill_length_mm: 49.8<br />body_mass_g: 3675","bill_length_mm: 48.1<br />body_mass_g: 3325","bill_length_mm: 51.4<br />body_mass_g: 3950","bill_length_mm: 45.7<br />body_mass_g: 3600","bill_length_mm: 50.7<br />body_mass_g: 4050","bill_length_mm: 42.5<br />body_mass_g: 3350","bill_length_mm: 52.2<br />body_mass_g: 3450","bill_length_mm: 45.2<br />body_mass_g: 3250","bill_length_mm: 49.3<br />body_mass_g: 4050","bill_length_mm: 50.2<br />body_mass_g: 3800","bill_length_mm: 45.6<br />body_mass_g: 3525","bill_length_mm: 51.9<br />body_mass_g: 3950","bill_length_mm: 46.8<br />body_mass_g: 3650","bill_length_mm: 45.7<br />body_mass_g: 3650","bill_length_mm: 55.8<br />body_mass_g: 4000","bill_length_mm: 43.5<br />body_mass_g: 3400","bill_length_mm: 49.6<br />body_mass_g: 3775","bill_length_mm: 50.8<br />body_mass_g: 4100","bill_length_mm: 50.2<br />body_mass_g: 3775"],"key":["1","2","3","4","5","6","7","8","9","10","11","12","13","14","15","16","17","18","19","20","21","22","23","24","25","26","27","28","29","30","31","32","33","34","35","36","37","38","39","40","41","42","43","44","45","46","47","48","49","50","51","52","53","54","55","56","57","58","59","60","61","62","63","64","65","66","67","68","69","70","71","72","73","74","75","76","77","78","79","80","81","82","83","84","85","86","87","88","89","90","91","92","93","94","95","96","97","98","99","100","101","102","103","104","105","106","107","108","109","110","111","112","113","114","115","116","117","118","119","120","121","122","123","124","125","126","127","128","129","130","131","132","133","134","135","136","137","138","139","140","141","142","143","144","145","146","147","148","149","150","151","152","153","154","155","156","157","158","159","160","161","162","163","164","165","166","167","168","169","170","171","172","173","174","175","176","177","178","179","180","181","182","183","184","185","186","187","188","189","190","191","192","193","194","195","196","197","198","199","200","201","202","203","204","205","206","207","208","209","210","211","212","213","214","215","216","217","218","219","220","221","222","223","224","225","226","227","228","229","230","231","232","233","234","235","236","237","238","239","240","241","242","243","244","245","246","247","248","249","250","251","252","253","254","255","256","257","258","259","260","261","262","263","264","265","266","267","268","269","270","271","272","273","274","275","276","277","278","279","280","281","282","283","284","285","286","287","288","289","290","291","292","293","294","295","296","297","298","299","300","301","302","303","304","305","306","307","308","309","310","311","312","313","314","315","316","317","318","319","320","321","322","323","324","325","326","327","328","329","330","331","332","333","334","335","336","337","338","339","340","341","342","343","344"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(0,0,0,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(0,0,0,1)"}},"hoveron":"points","set":"SharedData2edbd762","showlegend":false,"xaxis":"x","yaxis":"y","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[39.1,39.5,40.3,null,36.7,39.3,38.9,39.2,34.1,42,37.8,37.8,41.1,38.6,34.6,36.6,38.7,42.5,34.4,46,37.8,37.7,35.9,38.2,38.8,35.3,40.6,40.5,37.9,40.5,39.5,37.2,39.5,40.9,36.4,39.2,38.8,42.2,37.6,39.8,36.5,40.8,36,44.1,37,39.6,41.1,37.5,36,42.3,39.6,40.1,35,42,34.5,41.4,39,40.6,36.5,37.6,35.7,41.3,37.6,41.1,36.4,41.6,35.5,41.1,35.9,41.8,33.5,39.7,39.6,45.8,35.5,42.8,40.9,37.2,36.2,42.1,34.6,42.9,36.7,35.1,37.3,41.3,36.3,36.9,38.3,38.9,35.7,41.1,34,39.6,36.2,40.8,38.1,40.3,33.1,43.2,35,41,37.7,37.8,37.9,39.7,38.6,38.2,38.1,43.2,38.1,45.6,39.7,42.2,39.6,42.7,38.6,37.3,35.7,41.1,36.2,37.7,40.2,41.4,35.2,40.6,38.8,41.5,39,44.1,38.5,43.1,36.8,37.5,38.1,41.1,35.6,40.2,37,39.7,40.2,40.6,32.1,40.7,37.3,39,39.2,36.6,36,37.8,36,41.5,46.1,50,48.7,50,47.6,46.5,45.4,46.7,43.3,46.8,40.9,49,45.5,48.4,45.8,49.3,42,49.2,46.2,48.7,50.2,45.1,46.5,46.3,42.9,46.1,44.5,47.8,48.2,50,47.3,42.8,45.1,59.6,49.1,48.4,42.6,44.4,44,48.7,42.7,49.6,45.3,49.6,50.5,43.6,45.5,50.5,44.9,45.2,46.6,48.5,45.1,50.1,46.5,45,43.8,45.5,43.2,50.4,45.3,46.2,45.7,54.3,45.8,49.8,46.2,49.5,43.5,50.7,47.7,46.4,48.2,46.5,46.4,48.6,47.5,51.1,45.2,45.2,49.1,52.5,47.4,50,44.9,50.8,43.4,51.3,47.5,52.1,47.5,52.2,45.5,49.5,44.5,50.8,49.4,46.9,48.4,51.1,48.5,55.9,47.2,49.1,47.3,46.8,41.7,53.4,43.3,48.1,50.5,49.8,43.5,51.5,46.2,55.1,44.5,48.8,47.2,null,46.8,50.4,45.2,49.9,46.5,50,51.3,45.4,52.7,45.2,46.1,51.3,46,51.3,46.6,51.7,47,52,45.9,50.5,50.3,58,46.4,49.2,42.4,48.5,43.2,50.6,46.7,52,50.5,49.5,46.4,52.8,40.9,54.2,42.5,51,49.7,47.5,47.6,52,46.9,53.5,49,46.2,50.9,45.5,50.9,50.8,50.1,49,51.5,49.8,48.1,51.4,45.7,50.7,42.5,52.2,45.2,49.3,50.2,45.6,51.9,46.8,45.7,55.8,43.5,49.6,50.8,50.2],"y":[181,186,195,null,193,190,181,195,193,190,186,180,182,191,198,185,195,197,184,194,174,180,189,185,180,187,183,187,172,180,178,178,188,184,195,196,190,180,181,184,182,195,186,196,185,190,182,179,190,191,186,188,190,200,187,191,186,193,181,194,185,195,185,192,184,192,195,188,190,198,190,190,196,197,190,195,191,184,187,195,189,196,187,193,191,194,190,189,189,190,202,205,185,186,187,208,190,196,178,192,192,203,183,190,193,184,199,190,181,197,198,191,193,197,191,196,188,199,189,189,187,198,176,202,186,199,191,195,191,210,190,197,193,199,187,190,191,200,185,193,193,187,188,190,192,185,190,184,195,193,187,201,211,230,210,218,215,210,211,219,209,215,214,216,214,213,210,217,210,221,209,222,218,215,213,215,215,215,216,215,210,220,222,209,207,230,220,220,213,219,208,208,208,225,210,216,222,217,210,225,213,215,210,220,210,225,217,220,208,220,208,224,208,221,214,231,219,230,214,229,220,223,216,221,221,217,216,230,209,220,215,223,212,221,212,224,212,228,218,218,212,230,218,228,212,224,214,226,216,222,203,225,219,228,215,228,216,215,210,219,208,209,216,229,213,230,217,230,217,222,214,null,215,222,212,213,192,196,193,188,197,198,178,197,195,198,193,194,185,201,190,201,197,181,190,195,181,191,187,193,195,197,200,200,191,205,187,201,187,203,195,199,195,210,192,205,210,187,196,196,196,201,190,212,187,198,199,201,193,203,187,197,191,203,202,194,206,189,195,207,202,193,210,198],"text":["bill_length_mm: 39.1<br />flipper_length_mm: 181","bill_length_mm: 39.5<br />flipper_length_mm: 186","bill_length_mm: 40.3<br />flipper_length_mm: 195","bill_length_mm: NA<br />flipper_length_mm: NA","bill_length_mm: 36.7<br />flipper_length_mm: 193","bill_length_mm: 39.3<br />flipper_length_mm: 190","bill_length_mm: 38.9<br />flipper_length_mm: 181","bill_length_mm: 39.2<br />flipper_length_mm: 195","bill_length_mm: 34.1<br />flipper_length_mm: 193","bill_length_mm: 42.0<br />flipper_length_mm: 190","bill_length_mm: 37.8<br />flipper_length_mm: 186","bill_length_mm: 37.8<br />flipper_length_mm: 180","bill_length_mm: 41.1<br />flipper_length_mm: 182","bill_length_mm: 38.6<br />flipper_length_mm: 191","bill_length_mm: 34.6<br />flipper_length_mm: 198","bill_length_mm: 36.6<br />flipper_length_mm: 185","bill_length_mm: 38.7<br />flipper_length_mm: 195","bill_length_mm: 42.5<br />flipper_length_mm: 197","bill_length_mm: 34.4<br />flipper_length_mm: 184","bill_length_mm: 46.0<br />flipper_length_mm: 194","bill_length_mm: 37.8<br />flipper_length_mm: 174","bill_length_mm: 37.7<br />flipper_length_mm: 180","bill_length_mm: 35.9<br />flipper_length_mm: 189","bill_length_mm: 38.2<br />flipper_length_mm: 185","bill_length_mm: 38.8<br />flipper_length_mm: 180","bill_length_mm: 35.3<br />flipper_length_mm: 187","bill_length_mm: 40.6<br />flipper_length_mm: 183","bill_length_mm: 40.5<br />flipper_length_mm: 187","bill_length_mm: 37.9<br />flipper_length_mm: 172","bill_length_mm: 40.5<br />flipper_length_mm: 180","bill_length_mm: 39.5<br />flipper_length_mm: 178","bill_length_mm: 37.2<br />flipper_length_mm: 178","bill_length_mm: 39.5<br />flipper_length_mm: 188","bill_length_mm: 40.9<br />flipper_length_mm: 184","bill_length_mm: 36.4<br />flipper_length_mm: 195","bill_length_mm: 39.2<br />flipper_length_mm: 196","bill_length_mm: 38.8<br />flipper_length_mm: 190","bill_length_mm: 42.2<br />flipper_length_mm: 180","bill_length_mm: 37.6<br />flipper_length_mm: 181","bill_length_mm: 39.8<br />flipper_length_mm: 184","bill_length_mm: 36.5<br />flipper_length_mm: 182","bill_length_mm: 40.8<br />flipper_length_mm: 195","bill_length_mm: 36.0<br />flipper_length_mm: 186","bill_length_mm: 44.1<br />flipper_length_mm: 196","bill_length_mm: 37.0<br />flipper_length_mm: 185","bill_length_mm: 39.6<br />flipper_length_mm: 190","bill_length_mm: 41.1<br />flipper_length_mm: 182","bill_length_mm: 37.5<br />flipper_length_mm: 179","bill_length_mm: 36.0<br />flipper_length_mm: 190","bill_length_mm: 42.3<br />flipper_length_mm: 191","bill_length_mm: 39.6<br />flipper_length_mm: 186","bill_length_mm: 40.1<br />flipper_length_mm: 188","bill_length_mm: 35.0<br />flipper_length_mm: 190","bill_length_mm: 42.0<br />flipper_length_mm: 200","bill_length_mm: 34.5<br />flipper_length_mm: 187","bill_length_mm: 41.4<br />flipper_length_mm: 191","bill_length_mm: 39.0<br />flipper_length_mm: 186","bill_length_mm: 40.6<br />flipper_length_mm: 193","bill_length_mm: 36.5<br />flipper_length_mm: 181","bill_length_mm: 37.6<br />flipper_length_mm: 194","bill_length_mm: 35.7<br />flipper_length_mm: 185","bill_length_mm: 41.3<br />flipper_length_mm: 195","bill_length_mm: 37.6<br />flipper_length_mm: 185","bill_length_mm: 41.1<br />flipper_length_mm: 192","bill_length_mm: 36.4<br />flipper_length_mm: 184","bill_length_mm: 41.6<br />flipper_length_mm: 192","bill_length_mm: 35.5<br />flipper_length_mm: 195","bill_length_mm: 41.1<br />flipper_length_mm: 188","bill_length_mm: 35.9<br />flipper_length_mm: 190","bill_length_mm: 41.8<br />flipper_length_mm: 198","bill_length_mm: 33.5<br />flipper_length_mm: 190","bill_length_mm: 39.7<br />flipper_length_mm: 190","bill_length_mm: 39.6<br />flipper_length_mm: 196","bill_length_mm: 45.8<br />flipper_length_mm: 197","bill_length_mm: 35.5<br />flipper_length_mm: 190","bill_length_mm: 42.8<br />flipper_length_mm: 195","bill_length_mm: 40.9<br />flipper_length_mm: 191","bill_length_mm: 37.2<br />flipper_length_mm: 184","bill_length_mm: 36.2<br />flipper_length_mm: 187","bill_length_mm: 42.1<br />flipper_length_mm: 195","bill_length_mm: 34.6<br />flipper_length_mm: 189","bill_length_mm: 42.9<br />flipper_length_mm: 196","bill_length_mm: 36.7<br />flipper_length_mm: 187","bill_length_mm: 35.1<br />flipper_length_mm: 193","bill_length_mm: 37.3<br />flipper_length_mm: 191","bill_length_mm: 41.3<br />flipper_length_mm: 194","bill_length_mm: 36.3<br />flipper_length_mm: 190","bill_length_mm: 36.9<br />flipper_length_mm: 189","bill_length_mm: 38.3<br />flipper_length_mm: 189","bill_length_mm: 38.9<br />flipper_length_mm: 190","bill_length_mm: 35.7<br />flipper_length_mm: 202","bill_length_mm: 41.1<br />flipper_length_mm: 205","bill_length_mm: 34.0<br />flipper_length_mm: 185","bill_length_mm: 39.6<br />flipper_length_mm: 186","bill_length_mm: 36.2<br />flipper_length_mm: 187","bill_length_mm: 40.8<br />flipper_length_mm: 208","bill_length_mm: 38.1<br />flipper_length_mm: 190","bill_length_mm: 40.3<br />flipper_length_mm: 196","bill_length_mm: 33.1<br />flipper_length_mm: 178","bill_length_mm: 43.2<br />flipper_length_mm: 192","bill_length_mm: 35.0<br />flipper_length_mm: 192","bill_length_mm: 41.0<br />flipper_length_mm: 203","bill_length_mm: 37.7<br />flipper_length_mm: 183","bill_length_mm: 37.8<br />flipper_length_mm: 190","bill_length_mm: 37.9<br />flipper_length_mm: 193","bill_length_mm: 39.7<br />flipper_length_mm: 184","bill_length_mm: 38.6<br />flipper_length_mm: 199","bill_length_mm: 38.2<br />flipper_length_mm: 190","bill_length_mm: 38.1<br />flipper_length_mm: 181","bill_length_mm: 43.2<br />flipper_length_mm: 197","bill_length_mm: 38.1<br />flipper_length_mm: 198","bill_length_mm: 45.6<br />flipper_length_mm: 191","bill_length_mm: 39.7<br />flipper_length_mm: 193","bill_length_mm: 42.2<br />flipper_length_mm: 197","bill_length_mm: 39.6<br />flipper_length_mm: 191","bill_length_mm: 42.7<br />flipper_length_mm: 196","bill_length_mm: 38.6<br />flipper_length_mm: 188","bill_length_mm: 37.3<br />flipper_length_mm: 199","bill_length_mm: 35.7<br />flipper_length_mm: 189","bill_length_mm: 41.1<br />flipper_length_mm: 189","bill_length_mm: 36.2<br />flipper_length_mm: 187","bill_length_mm: 37.7<br />flipper_length_mm: 198","bill_length_mm: 40.2<br />flipper_length_mm: 176","bill_length_mm: 41.4<br />flipper_length_mm: 202","bill_length_mm: 35.2<br />flipper_length_mm: 186","bill_length_mm: 40.6<br />flipper_length_mm: 199","bill_length_mm: 38.8<br />flipper_length_mm: 191","bill_length_mm: 41.5<br />flipper_length_mm: 195","bill_length_mm: 39.0<br />flipper_length_mm: 191","bill_length_mm: 44.1<br />flipper_length_mm: 210","bill_length_mm: 38.5<br />flipper_length_mm: 190","bill_length_mm: 43.1<br />flipper_length_mm: 197","bill_length_mm: 36.8<br />flipper_length_mm: 193","bill_length_mm: 37.5<br />flipper_length_mm: 199","bill_length_mm: 38.1<br />flipper_length_mm: 187","bill_length_mm: 41.1<br />flipper_length_mm: 190","bill_length_mm: 35.6<br />flipper_length_mm: 191","bill_length_mm: 40.2<br />flipper_length_mm: 200","bill_length_mm: 37.0<br />flipper_length_mm: 185","bill_length_mm: 39.7<br />flipper_length_mm: 193","bill_length_mm: 40.2<br />flipper_length_mm: 193","bill_length_mm: 40.6<br />flipper_length_mm: 187","bill_length_mm: 32.1<br />flipper_length_mm: 188","bill_length_mm: 40.7<br />flipper_length_mm: 190","bill_length_mm: 37.3<br />flipper_length_mm: 192","bill_length_mm: 39.0<br />flipper_length_mm: 185","bill_length_mm: 39.2<br />flipper_length_mm: 190","bill_length_mm: 36.6<br />flipper_length_mm: 184","bill_length_mm: 36.0<br />flipper_length_mm: 195","bill_length_mm: 37.8<br />flipper_length_mm: 193","bill_length_mm: 36.0<br />flipper_length_mm: 187","bill_length_mm: 41.5<br />flipper_length_mm: 201","bill_length_mm: 46.1<br />flipper_length_mm: 211","bill_length_mm: 50.0<br />flipper_length_mm: 230","bill_length_mm: 48.7<br />flipper_length_mm: 210","bill_length_mm: 50.0<br />flipper_length_mm: 218","bill_length_mm: 47.6<br />flipper_length_mm: 215","bill_length_mm: 46.5<br />flipper_length_mm: 210","bill_length_mm: 45.4<br />flipper_length_mm: 211","bill_length_mm: 46.7<br />flipper_length_mm: 219","bill_length_mm: 43.3<br />flipper_length_mm: 209","bill_length_mm: 46.8<br />flipper_length_mm: 215","bill_length_mm: 40.9<br />flipper_length_mm: 214","bill_length_mm: 49.0<br />flipper_length_mm: 216","bill_length_mm: 45.5<br />flipper_length_mm: 214","bill_length_mm: 48.4<br />flipper_length_mm: 213","bill_length_mm: 45.8<br />flipper_length_mm: 210","bill_length_mm: 49.3<br />flipper_length_mm: 217","bill_length_mm: 42.0<br />flipper_length_mm: 210","bill_length_mm: 49.2<br />flipper_length_mm: 221","bill_length_mm: 46.2<br />flipper_length_mm: 209","bill_length_mm: 48.7<br />flipper_length_mm: 222","bill_length_mm: 50.2<br />flipper_length_mm: 218","bill_length_mm: 45.1<br />flipper_length_mm: 215","bill_length_mm: 46.5<br />flipper_length_mm: 213","bill_length_mm: 46.3<br />flipper_length_mm: 215","bill_length_mm: 42.9<br />flipper_length_mm: 215","bill_length_mm: 46.1<br />flipper_length_mm: 215","bill_length_mm: 44.5<br />flipper_length_mm: 216","bill_length_mm: 47.8<br />flipper_length_mm: 215","bill_length_mm: 48.2<br />flipper_length_mm: 210","bill_length_mm: 50.0<br />flipper_length_mm: 220","bill_length_mm: 47.3<br />flipper_length_mm: 222","bill_length_mm: 42.8<br />flipper_length_mm: 209","bill_length_mm: 45.1<br />flipper_length_mm: 207","bill_length_mm: 59.6<br />flipper_length_mm: 230","bill_length_mm: 49.1<br />flipper_length_mm: 220","bill_length_mm: 48.4<br />flipper_length_mm: 220","bill_length_mm: 42.6<br />flipper_length_mm: 213","bill_length_mm: 44.4<br />flipper_length_mm: 219","bill_length_mm: 44.0<br />flipper_length_mm: 208","bill_length_mm: 48.7<br />flipper_length_mm: 208","bill_length_mm: 42.7<br />flipper_length_mm: 208","bill_length_mm: 49.6<br />flipper_length_mm: 225","bill_length_mm: 45.3<br />flipper_length_mm: 210","bill_length_mm: 49.6<br />flipper_length_mm: 216","bill_length_mm: 50.5<br />flipper_length_mm: 222","bill_length_mm: 43.6<br />flipper_length_mm: 217","bill_length_mm: 45.5<br />flipper_length_mm: 210","bill_length_mm: 50.5<br />flipper_length_mm: 225","bill_length_mm: 44.9<br />flipper_length_mm: 213","bill_length_mm: 45.2<br />flipper_length_mm: 215","bill_length_mm: 46.6<br />flipper_length_mm: 210","bill_length_mm: 48.5<br />flipper_length_mm: 220","bill_length_mm: 45.1<br />flipper_length_mm: 210","bill_length_mm: 50.1<br />flipper_length_mm: 225","bill_length_mm: 46.5<br />flipper_length_mm: 217","bill_length_mm: 45.0<br />flipper_length_mm: 220","bill_length_mm: 43.8<br />flipper_length_mm: 208","bill_length_mm: 45.5<br />flipper_length_mm: 220","bill_length_mm: 43.2<br />flipper_length_mm: 208","bill_length_mm: 50.4<br />flipper_length_mm: 224","bill_length_mm: 45.3<br />flipper_length_mm: 208","bill_length_mm: 46.2<br />flipper_length_mm: 221","bill_length_mm: 45.7<br />flipper_length_mm: 214","bill_length_mm: 54.3<br />flipper_length_mm: 231","bill_length_mm: 45.8<br />flipper_length_mm: 219","bill_length_mm: 49.8<br />flipper_length_mm: 230","bill_length_mm: 46.2<br />flipper_length_mm: 214","bill_length_mm: 49.5<br />flipper_length_mm: 229","bill_length_mm: 43.5<br />flipper_length_mm: 220","bill_length_mm: 50.7<br />flipper_length_mm: 223","bill_length_mm: 47.7<br />flipper_length_mm: 216","bill_length_mm: 46.4<br />flipper_length_mm: 221","bill_length_mm: 48.2<br />flipper_length_mm: 221","bill_length_mm: 46.5<br />flipper_length_mm: 217","bill_length_mm: 46.4<br />flipper_length_mm: 216","bill_length_mm: 48.6<br />flipper_length_mm: 230","bill_length_mm: 47.5<br />flipper_length_mm: 209","bill_length_mm: 51.1<br />flipper_length_mm: 220","bill_length_mm: 45.2<br />flipper_length_mm: 215","bill_length_mm: 45.2<br />flipper_length_mm: 223","bill_length_mm: 49.1<br />flipper_length_mm: 212","bill_length_mm: 52.5<br />flipper_length_mm: 221","bill_length_mm: 47.4<br />flipper_length_mm: 212","bill_length_mm: 50.0<br />flipper_length_mm: 224","bill_length_mm: 44.9<br />flipper_length_mm: 212","bill_length_mm: 50.8<br />flipper_length_mm: 228","bill_length_mm: 43.4<br />flipper_length_mm: 218","bill_length_mm: 51.3<br />flipper_length_mm: 218","bill_length_mm: 47.5<br />flipper_length_mm: 212","bill_length_mm: 52.1<br />flipper_length_mm: 230","bill_length_mm: 47.5<br />flipper_length_mm: 218","bill_length_mm: 52.2<br />flipper_length_mm: 228","bill_length_mm: 45.5<br />flipper_length_mm: 212","bill_length_mm: 49.5<br />flipper_length_mm: 224","bill_length_mm: 44.5<br />flipper_length_mm: 214","bill_length_mm: 50.8<br />flipper_length_mm: 226","bill_length_mm: 49.4<br />flipper_length_mm: 216","bill_length_mm: 46.9<br />flipper_length_mm: 222","bill_length_mm: 48.4<br />flipper_length_mm: 203","bill_length_mm: 51.1<br />flipper_length_mm: 225","bill_length_mm: 48.5<br />flipper_length_mm: 219","bill_length_mm: 55.9<br />flipper_length_mm: 228","bill_length_mm: 47.2<br />flipper_length_mm: 215","bill_length_mm: 49.1<br />flipper_length_mm: 228","bill_length_mm: 47.3<br />flipper_length_mm: 216","bill_length_mm: 46.8<br />flipper_length_mm: 215","bill_length_mm: 41.7<br />flipper_length_mm: 210","bill_length_mm: 53.4<br />flipper_length_mm: 219","bill_length_mm: 43.3<br />flipper_length_mm: 208","bill_length_mm: 48.1<br />flipper_length_mm: 209","bill_length_mm: 50.5<br />flipper_length_mm: 216","bill_length_mm: 49.8<br />flipper_length_mm: 229","bill_length_mm: 43.5<br />flipper_length_mm: 213","bill_length_mm: 51.5<br />flipper_length_mm: 230","bill_length_mm: 46.2<br />flipper_length_mm: 217","bill_length_mm: 55.1<br />flipper_length_mm: 230","bill_length_mm: 44.5<br />flipper_length_mm: 217","bill_length_mm: 48.8<br />flipper_length_mm: 222","bill_length_mm: 47.2<br />flipper_length_mm: 214","bill_length_mm: NA<br />flipper_length_mm: NA","bill_length_mm: 46.8<br />flipper_length_mm: 215","bill_length_mm: 50.4<br />flipper_length_mm: 222","bill_length_mm: 45.2<br />flipper_length_mm: 212","bill_length_mm: 49.9<br />flipper_length_mm: 213","bill_length_mm: 46.5<br />flipper_length_mm: 192","bill_length_mm: 50.0<br />flipper_length_mm: 196","bill_length_mm: 51.3<br />flipper_length_mm: 193","bill_length_mm: 45.4<br />flipper_length_mm: 188","bill_length_mm: 52.7<br />flipper_length_mm: 197","bill_length_mm: 45.2<br />flipper_length_mm: 198","bill_length_mm: 46.1<br />flipper_length_mm: 178","bill_length_mm: 51.3<br />flipper_length_mm: 197","bill_length_mm: 46.0<br />flipper_length_mm: 195","bill_length_mm: 51.3<br />flipper_length_mm: 198","bill_length_mm: 46.6<br />flipper_length_mm: 193","bill_length_mm: 51.7<br />flipper_length_mm: 194","bill_length_mm: 47.0<br />flipper_length_mm: 185","bill_length_mm: 52.0<br />flipper_length_mm: 201","bill_length_mm: 45.9<br />flipper_length_mm: 190","bill_length_mm: 50.5<br />flipper_length_mm: 201","bill_length_mm: 50.3<br />flipper_length_mm: 197","bill_length_mm: 58.0<br />flipper_length_mm: 181","bill_length_mm: 46.4<br />flipper_length_mm: 190","bill_length_mm: 49.2<br />flipper_length_mm: 195","bill_length_mm: 42.4<br />flipper_length_mm: 181","bill_length_mm: 48.5<br />flipper_length_mm: 191","bill_length_mm: 43.2<br />flipper_length_mm: 187","bill_length_mm: 50.6<br />flipper_length_mm: 193","bill_length_mm: 46.7<br />flipper_length_mm: 195","bill_length_mm: 52.0<br />flipper_length_mm: 197","bill_length_mm: 50.5<br />flipper_length_mm: 200","bill_length_mm: 49.5<br />flipper_length_mm: 200","bill_length_mm: 46.4<br />flipper_length_mm: 191","bill_length_mm: 52.8<br />flipper_length_mm: 205","bill_length_mm: 40.9<br />flipper_length_mm: 187","bill_length_mm: 54.2<br />flipper_length_mm: 201","bill_length_mm: 42.5<br />flipper_length_mm: 187","bill_length_mm: 51.0<br />flipper_length_mm: 203","bill_length_mm: 49.7<br />flipper_length_mm: 195","bill_length_mm: 47.5<br />flipper_length_mm: 199","bill_length_mm: 47.6<br />flipper_length_mm: 195","bill_length_mm: 52.0<br />flipper_length_mm: 210","bill_length_mm: 46.9<br />flipper_length_mm: 192","bill_length_mm: 53.5<br />flipper_length_mm: 205","bill_length_mm: 49.0<br />flipper_length_mm: 210","bill_length_mm: 46.2<br />flipper_length_mm: 187","bill_length_mm: 50.9<br />flipper_length_mm: 196","bill_length_mm: 45.5<br />flipper_length_mm: 196","bill_length_mm: 50.9<br />flipper_length_mm: 196","bill_length_mm: 50.8<br />flipper_length_mm: 201","bill_length_mm: 50.1<br />flipper_length_mm: 190","bill_length_mm: 49.0<br />flipper_length_mm: 212","bill_length_mm: 51.5<br />flipper_length_mm: 187","bill_length_mm: 49.8<br />flipper_length_mm: 198","bill_length_mm: 48.1<br />flipper_length_mm: 199","bill_length_mm: 51.4<br />flipper_length_mm: 201","bill_length_mm: 45.7<br />flipper_length_mm: 193","bill_length_mm: 50.7<br />flipper_length_mm: 203","bill_length_mm: 42.5<br />flipper_length_mm: 187","bill_length_mm: 52.2<br />flipper_length_mm: 197","bill_length_mm: 45.2<br />flipper_length_mm: 191","bill_length_mm: 49.3<br />flipper_length_mm: 203","bill_length_mm: 50.2<br />flipper_length_mm: 202","bill_length_mm: 45.6<br />flipper_length_mm: 194","bill_length_mm: 51.9<br />flipper_length_mm: 206","bill_length_mm: 46.8<br />flipper_length_mm: 189","bill_length_mm: 45.7<br />flipper_length_mm: 195","bill_length_mm: 55.8<br />flipper_length_mm: 207","bill_length_mm: 43.5<br />flipper_length_mm: 202","bill_length_mm: 49.6<br />flipper_length_mm: 193","bill_length_mm: 50.8<br />flipper_length_mm: 210","bill_length_mm: 50.2<br />flipper_length_mm: 198"],"key":["1","2","3","4","5","6","7","8","9","10","11","12","13","14","15","16","17","18","19","20","21","22","23","24","25","26","27","28","29","30","31","32","33","34","35","36","37","38","39","40","41","42","43","44","45","46","47","48","49","50","51","52","53","54","55","56","57","58","59","60","61","62","63","64","65","66","67","68","69","70","71","72","73","74","75","76","77","78","79","80","81","82","83","84","85","86","87","88","89","90","91","92","93","94","95","96","97","98","99","100","101","102","103","104","105","106","107","108","109","110","111","112","113","114","115","116","117","118","119","120","121","122","123","124","125","126","127","128","129","130","131","132","133","134","135","136","137","138","139","140","141","142","143","144","145","146","147","148","149","150","151","152","153","154","155","156","157","158","159","160","161","162","163","164","165","166","167","168","169","170","171","172","173","174","175","176","177","178","179","180","181","182","183","184","185","186","187","188","189","190","191","192","193","194","195","196","197","198","199","200","201","202","203","204","205","206","207","208","209","210","211","212","213","214","215","216","217","218","219","220","221","222","223","224","225","226","227","228","229","230","231","232","233","234","235","236","237","238","239","240","241","242","243","244","245","246","247","248","249","250","251","252","253","254","255","256","257","258","259","260","261","262","263","264","265","266","267","268","269","270","271","272","273","274","275","276","277","278","279","280","281","282","283","284","285","286","287","288","289","290","291","292","293","294","295","296","297","298","299","300","301","302","303","304","305","306","307","308","309","310","311","312","313","314","315","316","317","318","319","320","321","322","323","324","325","326","327","328","329","330","331","332","333","334","335","336","337","338","339","340","341","342","343","344"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(0,0,0,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(0,0,0,1)"}},"hoveron":"points","set":"SharedData2edbd762","showlegend":false,"xaxis":"x2","yaxis":"y2","hoverinfo":"text","_isNestedKey":false,"frame":null}],"layout":{"xaxis":{"domain":[0,0.48],"automargin":true,"type":"linear","autorange":false,"range":[30.725,60.975],"tickmode":"array","ticktext":["40","50","60"],"tickvals":[40,50,60],"categoryorder":"array","categoryarray":["40","50","60"],"nticks":null,"ticks":"outside","tickcolor":"rgba(51,51,51,1)","ticklen":3.65296803652968,"tickwidth":0.66417600664176,"showticklabels":true,"tickfont":{"color":"rgba(77,77,77,1)","family":"","size":18.5969281859693},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"y","hoverformat":".2f"},"xaxis2":{"domain":[0.52,1],"automargin":true,"type":"linear","autorange":false,"range":[30.725,60.975],"tickmode":"array","ticktext":["40","50","60"],"tickvals":[40,50,60],"categoryorder":"array","categoryarray":["40","50","60"],"nticks":null,"ticks":"outside","tickcolor":"rgba(51,51,51,1)","ticklen":3.65296803652968,"tickwidth":0.66417600664176,"showticklabels":true,"tickfont":{"color":"rgba(77,77,77,1)","family":"","size":18.5969281859693},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"y2","hoverformat":".2f"},"yaxis2":{"domain":[0,1],"automargin":true,"type":"linear","autorange":false,"range":[169.05,233.95],"tickmode":"array","ticktext":["170","180","190","200","210","220","230"],"tickvals":[170,180,190,200,210,220,230],"categoryorder":"array","categoryarray":["170","180","190","200","210","220","230"],"nticks":null,"ticks":"outside","tickcolor":"rgba(51,51,51,1)","ticklen":3.65296803652968,"tickwidth":0.66417600664176,"showticklabels":true,"tickfont":{"color":"rgba(77,77,77,1)","family":"","size":18.5969281859693},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x2","hoverformat":".2f"},"yaxis":{"domain":[0,1],"automargin":true,"type":"linear","autorange":false,"range":[2520,6480],"tickmode":"array","ticktext":["3000","4000","5000","6000"],"tickvals":[3000,4000,5000,6000],"categoryorder":"array","categoryarray":["3000","4000","5000","6000"],"nticks":null,"ticks":"outside","tickcolor":"rgba(51,51,51,1)","ticklen":3.65296803652968,"tickwidth":0.66417600664176,"showticklabels":true,"tickfont":{"color":"rgba(77,77,77,1)","family":"","size":18.5969281859693},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x","hoverformat":".2f"},"annotations":[],"shapes":[{"type":"rect","fillcolor":null,"line":{"color":null,"width":0,"linetype":[]},"yref":"paper","xref":"paper","x0":0,"x1":0.48,"y0":0,"y1":1},{"type":"rect","fillcolor":null,"line":{"color":null,"width":0,"linetype":[]},"yref":"paper","xref":"paper","x0":0.52,"x1":1,"y0":0,"y1":1}],"images":[],"margin":{"t":32.4383561643836,"r":7.30593607305936,"b":59.9418845994189,"l":60.1079286010793},"plot_bgcolor":"rgba(255,255,255,1)","paper_bgcolor":"rgba(255,255,255,1)","font":{"color":"rgba(0,0,0,1)","family":"","size":14.6118721461187},"showlegend":false,"legend":{"bgcolor":"rgba(255,255,255,1)","bordercolor":"transparent","borderwidth":1.88976377952756,"font":{"color":"rgba(0,0,0,1)","family":"","size":18.5969281859693}},"hovermode":"closest","barmode":"relative","dragmode":"select","title":"Click and drag to select points"},"attrs":{"1841017398d2b":{"x":{},"y":{},"type":"scatter"},"1841075eb0af7":{"x":{},"y":{},"type":"scatter"}},"source":"A","config":{"doubleClick":"reset","showSendToCloud":false},"highlight":{"on":"plotly_selected","off":"plotly_deselect","persistent":false,"dynamic":false,"color":null,"selectize":false,"defaultValues":null,"opacityDim":0.2,"selected":{"opacity":1},"debounce":0,"ctGroups":["SharedData2edbd762"]},"subplot":true,"shinyEvents":["plotly_hover","plotly_click","plotly_selected","plotly_relayout","plotly_brushed","plotly_brushing","plotly_clickannotation","plotly_doubleclick","plotly_deselect","plotly_afterplot","plotly_sunburstclick"],"base_url":"https://plot.ly"},"evals":[],"jsHooks":[]}</script> --- ## Plotly You can also do brushing over multiple plots drawn on the same dataset .left-column70[ ```r highlight_key(iris) %>% GGally::ggpairs(aes(colour = Species), columns = 1:4) %>% ggplotly(tooltip = c("x", "y", "colour")) %>% highlight("plotly_selected") ``` <div id="htmlwidget-39b02b8d9a3cbbd3cd3c" style="width:504px;height:360px;" class="plotly html-widget"></div> <script type="application/json" data-for="htmlwidget-39b02b8d9a3cbbd3cd3c">{"x":{"data":[{"x":[4.3,4.30704500978474,4.31409001956947,4.32113502935421,4.32818003913894,4.33522504892368,4.34227005870842,4.34931506849315,4.35636007827789,4.36340508806262,4.37045009784736,4.37749510763209,4.38454011741683,4.39158512720157,4.3986301369863,4.40567514677104,4.41272015655577,4.41976516634051,4.42681017612524,4.43385518590998,4.44090019569472,4.44794520547945,4.45499021526419,4.46203522504892,4.46908023483366,4.4761252446184,4.48317025440313,4.49021526418787,4.4972602739726,4.50430528375734,4.51135029354207,4.51839530332681,4.52544031311155,4.53248532289628,4.53953033268102,4.54657534246575,4.55362035225049,4.56066536203523,4.56771037181996,4.5747553816047,4.58180039138943,4.58884540117417,4.5958904109589,4.60293542074364,4.60998043052838,4.61702544031311,4.62407045009785,4.63111545988258,4.63816046966732,4.64520547945205,4.65225048923679,4.65929549902153,4.66634050880626,4.673385518591,4.68043052837573,4.68747553816047,4.69452054794521,4.70156555772994,4.70861056751468,4.71565557729941,4.72270058708415,4.72974559686888,4.73679060665362,4.74383561643836,4.75088062622309,4.75792563600783,4.76497064579256,4.7720156555773,4.77906066536204,4.78610567514677,4.79315068493151,4.80019569471624,4.80724070450098,4.81428571428571,4.82133072407045,4.82837573385519,4.83542074363992,4.84246575342466,4.84951076320939,4.85655577299413,4.86360078277886,4.8706457925636,4.87769080234834,4.88473581213307,4.89178082191781,4.89882583170254,4.90587084148728,4.91291585127202,4.91996086105675,4.92700587084149,4.93405088062622,4.94109589041096,4.94814090019569,4.95518590998043,4.96223091976517,4.9692759295499,4.97632093933464,4.98336594911937,4.99041095890411,4.99745596868885,5.00450097847358,5.01154598825832,5.01859099804305,5.02563600782779,5.03268101761252,5.03972602739726,5.046771037182,5.05381604696673,5.06086105675147,5.0679060665362,5.07495107632094,5.08199608610567,5.08904109589041,5.09608610567515,5.10313111545988,5.11017612524462,5.11722113502935,5.12426614481409,5.13131115459883,5.13835616438356,5.1454011741683,5.15244618395303,5.15949119373777,5.1665362035225,5.17358121330724,5.18062622309198,5.18767123287671,5.19471624266145,5.20176125244618,5.20880626223092,5.21585127201566,5.22289628180039,5.22994129158513,5.23698630136986,5.2440313111546,5.25107632093933,5.25812133072407,5.26516634050881,5.27221135029354,5.27925636007828,5.28630136986301,5.29334637964775,5.30039138943249,5.30743639921722,5.31448140900196,5.32152641878669,5.32857142857143,5.33561643835616,5.3426614481409,5.34970645792564,5.35675146771037,5.36379647749511,5.37084148727984,5.37788649706458,5.38493150684932,5.39197651663405,5.39902152641879,5.40606653620352,5.41311154598826,5.42015655577299,5.42720156555773,5.43424657534247,5.4412915851272,5.44833659491194,5.45538160469667,5.46242661448141,5.46947162426614,5.47651663405088,5.48356164383562,5.49060665362035,5.49765166340509,5.50469667318982,5.51174168297456,5.5187866927593,5.52583170254403,5.53287671232877,5.5399217221135,5.54696673189824,5.55401174168297,5.56105675146771,5.56810176125245,5.57514677103718,5.58219178082192,5.58923679060665,5.59628180039139,5.60332681017613,5.61037181996086,5.6174168297456,5.62446183953033,5.63150684931507,5.6385518590998,5.64559686888454,5.65264187866928,5.65968688845401,5.66673189823875,5.67377690802348,5.68082191780822,5.68786692759295,5.69491193737769,5.70195694716243,5.70900195694716,5.7160469667319,5.72309197651663,5.73013698630137,5.73718199608611,5.74422700587084,5.75127201565558,5.75831702544031,5.76536203522505,5.77240704500978,5.77945205479452,5.78649706457926,5.79354207436399,5.80058708414873,5.80763209393346,5.8146771037182,5.82172211350294,5.82876712328767,5.83581213307241,5.84285714285714,5.84990215264188,5.85694716242661,5.86399217221135,5.87103718199609,5.87808219178082,5.88512720156556,5.89217221135029,5.89921722113503,5.90626223091976,5.9133072407045,5.92035225048924,5.92739726027397,5.93444227005871,5.94148727984344,5.94853228962818,5.95557729941292,5.96262230919765,5.96966731898239,5.97671232876712,5.98375733855186,5.9908023483366,5.99784735812133,6.00489236790607,6.0119373776908,6.01898238747554,6.02602739726027,6.03307240704501,6.04011741682975,6.04716242661448,6.05420743639922,6.06125244618395,6.06829745596869,6.07534246575342,6.08238747553816,6.0894324853229,6.09647749510763,6.10352250489237,6.1105675146771,6.11761252446184,6.12465753424657,6.13170254403131,6.13874755381605,6.14579256360078,6.15283757338552,6.15988258317025,6.16692759295499,6.17397260273973,6.18101761252446,6.1880626223092,6.19510763209393,6.20215264187867,6.20919765166341,6.21624266144814,6.22328767123288,6.23033268101761,6.23737769080235,6.24442270058708,6.25146771037182,6.25851272015656,6.26555772994129,6.27260273972603,6.27964774951076,6.2866927592955,6.29373776908024,6.30078277886497,6.30782778864971,6.31487279843444,6.32191780821918,6.32896281800391,6.33600782778865,6.34305283757339,6.35009784735812,6.35714285714286,6.36418786692759,6.37123287671233,6.37827788649706,6.3853228962818,6.39236790606654,6.39941291585127,6.40645792563601,6.41350293542074,6.42054794520548,6.42759295499022,6.43463796477495,6.44168297455969,6.44872798434442,6.45577299412916,6.46281800391389,6.46986301369863,6.47690802348337,6.4839530332681,6.49099804305284,6.49804305283757,6.50508806262231,6.51213307240705,6.51917808219178,6.52622309197652,6.53326810176125,6.54031311154599,6.54735812133072,6.55440313111546,6.5614481409002,6.56849315068493,6.57553816046967,6.5825831702544,6.58962818003914,6.59667318982387,6.60371819960861,6.61076320939335,6.61780821917808,6.62485322896282,6.63189823874755,6.63894324853229,6.64598825831703,6.65303326810176,6.6600782778865,6.66712328767123,6.67416829745597,6.6812133072407,6.68825831702544,6.69530332681018,6.70234833659491,6.70939334637965,6.71643835616438,6.72348336594912,6.73052837573386,6.73757338551859,6.74461839530333,6.75166340508806,6.7587084148728,6.76575342465753,6.77279843444227,6.77984344422701,6.78688845401174,6.79393346379648,6.80097847358121,6.80802348336595,6.81506849315068,6.82211350293542,6.82915851272016,6.83620352250489,6.84324853228963,6.85029354207436,6.8573385518591,6.86438356164384,6.87142857142857,6.87847358121331,6.88551859099804,6.89256360078278,6.89960861056752,6.90665362035225,6.91369863013699,6.92074363992172,6.92778864970646,6.93483365949119,6.94187866927593,6.94892367906067,6.9559686888454,6.96301369863014,6.97005870841487,6.97710371819961,6.98414872798434,6.99119373776908,6.99823874755382,7.00528375733855,7.01232876712329,7.01937377690802,7.02641878669276,7.03346379647749,7.04050880626223,7.04755381604697,7.0545988258317,7.06164383561644,7.06868884540117,7.07573385518591,7.08277886497065,7.08982387475538,7.09686888454012,7.10391389432485,7.11095890410959,7.11800391389433,7.12504892367906,7.1320939334638,7.13913894324853,7.14618395303327,7.153228962818,7.16027397260274,7.16731898238748,7.17436399217221,7.18140900195695,7.18845401174168,7.19549902152642,7.20254403131116,7.20958904109589,7.21663405088063,7.22367906066536,7.2307240704501,7.23776908023483,7.24481409001957,7.25185909980431,7.25890410958904,7.26594911937378,7.27299412915851,7.28003913894325,7.28708414872798,7.29412915851272,7.30117416829746,7.30821917808219,7.31526418786693,7.32230919765166,7.3293542074364,7.33639921722114,7.34344422700587,7.35048923679061,7.35753424657534,7.36457925636008,7.37162426614481,7.37866927592955,7.38571428571429,7.39275929549902,7.39980430528376,7.40684931506849,7.41389432485323,7.42093933463797,7.4279843444227,7.43502935420744,7.44207436399217,7.44911937377691,7.45616438356164,7.46320939334638,7.47025440313112,7.47729941291585,7.48434442270059,7.49138943248532,7.49843444227006,7.50547945205479,7.51252446183953,7.51956947162427,7.526614481409,7.53365949119374,7.54070450097847,7.54774951076321,7.55479452054795,7.56183953033268,7.56888454011742,7.57592954990215,7.58297455968689,7.59001956947163,7.59706457925636,7.6041095890411,7.61115459882583,7.61819960861057,7.6252446183953,7.63228962818004,7.63933463796478,7.64637964774951,7.65342465753425,7.66046966731898,7.66751467710372,7.67455968688845,7.68160469667319,7.68864970645793,7.69569471624266,7.7027397260274,7.70978473581213,7.71682974559687,7.7238747553816,7.73091976516634,7.73796477495108,7.74500978473581,7.75205479452055,7.75909980430528,7.76614481409002,7.77318982387476,7.78023483365949,7.78727984344423,7.79432485322896,7.8013698630137,7.80841487279844,7.81545988258317,7.82250489236791,7.82954990215264,7.83659491193738,7.84363992172211,7.85068493150685,7.85772994129159,7.86477495107632,7.87181996086106,7.87886497064579,7.88590998043053,7.89295499021526,7.9,7.9,7.89295499021526,7.88590998043053,7.87886497064579,7.87181996086106,7.86477495107632,7.85772994129159,7.85068493150685,7.84363992172211,7.83659491193738,7.82954990215264,7.82250489236791,7.81545988258317,7.80841487279844,7.8013698630137,7.79432485322896,7.78727984344423,7.78023483365949,7.77318982387476,7.76614481409002,7.75909980430528,7.75205479452055,7.74500978473581,7.73796477495108,7.73091976516634,7.7238747553816,7.71682974559687,7.70978473581213,7.7027397260274,7.69569471624266,7.68864970645793,7.68160469667319,7.67455968688845,7.66751467710372,7.66046966731898,7.65342465753425,7.64637964774951,7.63933463796478,7.63228962818004,7.6252446183953,7.61819960861057,7.61115459882583,7.6041095890411,7.59706457925636,7.59001956947163,7.58297455968689,7.57592954990215,7.56888454011742,7.56183953033268,7.55479452054795,7.54774951076321,7.54070450097847,7.53365949119374,7.526614481409,7.51956947162427,7.51252446183953,7.50547945205479,7.49843444227006,7.49138943248532,7.48434442270059,7.47729941291585,7.47025440313112,7.46320939334638,7.45616438356164,7.44911937377691,7.44207436399217,7.43502935420744,7.4279843444227,7.42093933463797,7.41389432485323,7.40684931506849,7.39980430528376,7.39275929549902,7.38571428571429,7.37866927592955,7.37162426614481,7.36457925636008,7.35753424657534,7.35048923679061,7.34344422700587,7.33639921722114,7.3293542074364,7.32230919765166,7.31526418786693,7.30821917808219,7.30117416829746,7.29412915851272,7.28708414872798,7.28003913894325,7.27299412915851,7.26594911937378,7.25890410958904,7.25185909980431,7.24481409001957,7.23776908023483,7.2307240704501,7.22367906066536,7.21663405088063,7.20958904109589,7.20254403131116,7.19549902152642,7.18845401174168,7.18140900195695,7.17436399217221,7.16731898238748,7.16027397260274,7.153228962818,7.14618395303327,7.13913894324853,7.1320939334638,7.12504892367906,7.11800391389433,7.11095890410959,7.10391389432485,7.09686888454012,7.08982387475538,7.08277886497065,7.07573385518591,7.06868884540117,7.06164383561644,7.0545988258317,7.04755381604697,7.04050880626223,7.03346379647749,7.02641878669276,7.01937377690802,7.01232876712329,7.00528375733855,6.99823874755382,6.99119373776908,6.98414872798434,6.97710371819961,6.97005870841487,6.96301369863014,6.9559686888454,6.94892367906067,6.94187866927593,6.93483365949119,6.92778864970646,6.92074363992172,6.91369863013699,6.90665362035225,6.89960861056752,6.89256360078278,6.88551859099804,6.87847358121331,6.87142857142857,6.86438356164384,6.8573385518591,6.85029354207436,6.84324853228963,6.83620352250489,6.82915851272016,6.82211350293542,6.81506849315068,6.80802348336595,6.80097847358121,6.79393346379648,6.78688845401174,6.77984344422701,6.77279843444227,6.76575342465753,6.7587084148728,6.75166340508806,6.74461839530333,6.73757338551859,6.73052837573386,6.72348336594912,6.71643835616438,6.70939334637965,6.70234833659491,6.69530332681018,6.68825831702544,6.6812133072407,6.67416829745597,6.66712328767123,6.6600782778865,6.65303326810176,6.64598825831703,6.63894324853229,6.63189823874755,6.62485322896282,6.61780821917808,6.61076320939335,6.60371819960861,6.59667318982387,6.58962818003914,6.5825831702544,6.57553816046967,6.56849315068493,6.5614481409002,6.55440313111546,6.54735812133072,6.54031311154599,6.53326810176125,6.52622309197652,6.51917808219178,6.51213307240705,6.50508806262231,6.49804305283757,6.49099804305284,6.4839530332681,6.47690802348337,6.46986301369863,6.46281800391389,6.45577299412916,6.44872798434442,6.44168297455969,6.43463796477495,6.42759295499022,6.42054794520548,6.41350293542074,6.40645792563601,6.39941291585127,6.39236790606654,6.3853228962818,6.37827788649706,6.37123287671233,6.36418786692759,6.35714285714286,6.35009784735812,6.34305283757339,6.33600782778865,6.32896281800391,6.32191780821918,6.31487279843444,6.30782778864971,6.30078277886497,6.29373776908024,6.2866927592955,6.27964774951076,6.27260273972603,6.26555772994129,6.25851272015656,6.25146771037182,6.24442270058708,6.23737769080235,6.23033268101761,6.22328767123288,6.21624266144814,6.20919765166341,6.20215264187867,6.19510763209393,6.1880626223092,6.18101761252446,6.17397260273973,6.16692759295499,6.15988258317025,6.15283757338552,6.14579256360078,6.13874755381605,6.13170254403131,6.12465753424657,6.11761252446184,6.1105675146771,6.10352250489237,6.09647749510763,6.0894324853229,6.08238747553816,6.07534246575342,6.06829745596869,6.06125244618395,6.05420743639922,6.04716242661448,6.04011741682975,6.03307240704501,6.02602739726027,6.01898238747554,6.0119373776908,6.00489236790607,5.99784735812133,5.9908023483366,5.98375733855186,5.97671232876712,5.96966731898239,5.96262230919765,5.95557729941292,5.94853228962818,5.94148727984344,5.93444227005871,5.92739726027397,5.92035225048924,5.9133072407045,5.90626223091976,5.89921722113503,5.89217221135029,5.88512720156556,5.87808219178082,5.87103718199609,5.86399217221135,5.85694716242661,5.84990215264188,5.84285714285714,5.83581213307241,5.82876712328767,5.82172211350294,5.8146771037182,5.80763209393346,5.80058708414873,5.79354207436399,5.78649706457926,5.77945205479452,5.77240704500978,5.76536203522505,5.75831702544031,5.75127201565558,5.74422700587084,5.73718199608611,5.73013698630137,5.72309197651663,5.7160469667319,5.70900195694716,5.70195694716243,5.69491193737769,5.68786692759295,5.68082191780822,5.67377690802348,5.66673189823875,5.65968688845401,5.65264187866928,5.64559686888454,5.6385518590998,5.63150684931507,5.62446183953033,5.6174168297456,5.61037181996086,5.60332681017613,5.59628180039139,5.58923679060665,5.58219178082192,5.57514677103718,5.56810176125245,5.56105675146771,5.55401174168297,5.54696673189824,5.5399217221135,5.53287671232877,5.52583170254403,5.5187866927593,5.51174168297456,5.50469667318982,5.49765166340509,5.49060665362035,5.48356164383562,5.47651663405088,5.46947162426614,5.46242661448141,5.45538160469667,5.44833659491194,5.4412915851272,5.43424657534247,5.42720156555773,5.42015655577299,5.41311154598826,5.40606653620352,5.39902152641879,5.39197651663405,5.38493150684932,5.37788649706458,5.37084148727984,5.36379647749511,5.35675146771037,5.34970645792564,5.3426614481409,5.33561643835616,5.32857142857143,5.32152641878669,5.31448140900196,5.30743639921722,5.30039138943249,5.29334637964775,5.28630136986301,5.27925636007828,5.27221135029354,5.26516634050881,5.25812133072407,5.25107632093933,5.2440313111546,5.23698630136986,5.22994129158513,5.22289628180039,5.21585127201566,5.20880626223092,5.20176125244618,5.19471624266145,5.18767123287671,5.18062622309198,5.17358121330724,5.1665362035225,5.15949119373777,5.15244618395303,5.1454011741683,5.13835616438356,5.13131115459883,5.12426614481409,5.11722113502935,5.11017612524462,5.10313111545988,5.09608610567515,5.08904109589041,5.08199608610567,5.07495107632094,5.0679060665362,5.06086105675147,5.05381604696673,5.046771037182,5.03972602739726,5.03268101761252,5.02563600782779,5.01859099804305,5.01154598825832,5.00450097847358,4.99745596868885,4.99041095890411,4.98336594911937,4.97632093933464,4.9692759295499,4.96223091976517,4.95518590998043,4.94814090019569,4.94109589041096,4.93405088062622,4.92700587084149,4.91996086105675,4.91291585127202,4.90587084148728,4.89882583170254,4.89178082191781,4.88473581213307,4.87769080234834,4.8706457925636,4.86360078277886,4.85655577299413,4.84951076320939,4.84246575342466,4.83542074363992,4.82837573385519,4.82133072407045,4.81428571428571,4.80724070450098,4.80019569471624,4.79315068493151,4.78610567514677,4.77906066536204,4.7720156555773,4.76497064579256,4.75792563600783,4.75088062622309,4.74383561643836,4.73679060665362,4.72974559686888,4.72270058708415,4.71565557729941,4.70861056751468,4.70156555772994,4.69452054794521,4.68747553816047,4.68043052837573,4.673385518591,4.66634050880626,4.65929549902153,4.65225048923679,4.64520547945205,4.63816046966732,4.63111545988258,4.62407045009785,4.61702544031311,4.60998043052838,4.60293542074364,4.5958904109589,4.58884540117417,4.58180039138943,4.5747553816047,4.56771037181996,4.56066536203523,4.55362035225049,4.54657534246575,4.53953033268102,4.53248532289628,4.52544031311155,4.51839530332681,4.51135029354207,4.50430528375734,4.4972602739726,4.49021526418787,4.48317025440313,4.4761252446184,4.46908023483366,4.46203522504892,4.45499021526419,4.44794520547945,4.44090019569472,4.43385518590998,4.42681017612524,4.41976516634051,4.41272015655577,4.40567514677104,4.3986301369863,4.39158512720157,4.38454011741683,4.37749510763209,4.37045009784736,4.36340508806262,4.35636007827789,4.34931506849315,4.34227005870842,4.33522504892368,4.32818003913894,4.32113502935421,4.31409001956947,4.30704500978474,4.3,4.3],"y":[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,5.02214699197813e-18,1.59236334172339e-17,2.68251198424897e-17,2.77555756156289e-17,1.37661170765172e-17,0,1.04212650444057e-18,1.19436129296936e-17,4.91047626067952e-18,0,3.01470878201652e-18,1.39546026067296e-17,3.57575754572411e-17,2.57061785391324e-17,8.89049674821983e-18,0,0,6.95532230381219e-17,6.93050213060118e-17,2.96826475596003e-17,1.00727057205579e-18,1.19087569973099e-17,6.74725214963231e-17,1.1501297735128e-16,1.23878613829981e-16,6.93960419732176e-17,9.11990148237314e-17,1.28859460693789e-16,1.32845925377115e-16,1.03072915888659e-16,1.35777375164426e-16,1.58580515652841e-16,1.52683418884197e-16,1.54572742167064e-16,1.84564882912693e-16,7.55500186601383e-17,9.11015499607296e-17,1.22864899062134e-16,1.44667871912643e-16,1.66470844763152e-16,1.23052725853989e-16,9.52345413077431e-17,1.22607360405501e-16,1.9817158493394e-16,2.19974557784452e-16,1.23379976375384e-16,1.10827372736505e-16,1.38777878078145e-16,1.3315252583442e-16,6.77436072828871e-17,9.09621262311956e-17,8.74765329928126e-17,5.55111512312578e-17,5.55111512312578e-17,9.88524553415037e-17,6.38681545925024e-17,1.79901290907424e-17,7.62568751954184e-18,5.1231633220565e-17,5.55111512312578e-17,2.80899290711732e-17,0,5.41621664942292e-18,7.08251352009494e-17,1.27406165934889e-16,8.82882151004941e-17,0,0,1.0800470095178e-17,6.05361909519576e-18,0,0,0,0,0,0,0,0,1.75860637704339e-17,1.61221146103114e-17,0,0,0,0,0,0,0,0,6.78559367525416e-18,1.0068495515119e-17,0,0,0,0,0,0,0,0,0,0,0,0,0,3.77443636031199e-18,1.30796528300625e-17,2.1781664048081e-18,0,0,0,0,0,0,7.11494929936107e-18,1.81661844861984e-17,1.81395484831306e-18,0,0,0,0,0,0,0,0,0,1.17271510022084e-16,2.02606508276005e-16,2.78174613640682e-16,4.00641103952288e-16,7.38587183135251e-16,1.14896321677315e-15,1.78914429812288e-15,2.76495097579255e-15,4.19320728925276e-15,6.61333727565924e-15,1.04613126077533e-14,1.62897043245078e-14,2.48915820520263e-14,3.74173899546449e-14,5.8704491869487e-14,9.08136716649467e-14,1.38297078740258e-13,2.06874113834396e-13,3.11283100072294e-13,4.77968720746789e-13,7.23992976188553e-13,1.08003136476993e-12,1.58312882081629e-12,2.3795151086404e-12,3.57866861598763e-12,5.31075384423196e-12,7.76400212469733e-12,1.12176420809073e-11,1.67287176075871e-11,2.46525238826734e-11,3.5857844144099e-11,5.14001906062098e-11,7.40415967776878e-11,1.08205284080562e-10,1.56306141399908e-10,2.22936311409038e-10,3.13505616903919e-10,4.4962872778474e-10,6.44148441364361e-10,9.12465360387898e-10,1.27675771142395e-09,1.77051480064651e-09,2.51286410764443e-09,3.53036497158407e-09,4.90612238862031e-09,6.73806044687497e-09,9.28889859598065e-09,1.29290479992003e-08,1.78199726243003e-08,2.43058491191019e-08,3.27822249875613e-08,4.48656390127408e-08,6.12646290367026e-08,8.28746840060533e-08,1.109982976443e-07,1.47650031410993e-07,1.995814475492e-07,2.67473578301081e-07,3.55267330423093e-07,4.67468688502831e-07,6.16631825800541e-07,8.18045487271995e-07,1.07642384455844e-06,1.40448696029623e-06,1.81649159199391e-06,2.37309689154378e-06,3.09104688734889e-06,3.99525634340275e-06,5.12320132233775e-06,6.53489798320235e-06,8.42058773523181e-06,1.07734148461158e-05,1.36842048903371e-05,1.72537342524602e-05,2.17798729371969e-05,2.75664588465454e-05,3.46578934934782e-05,4.32805914070438e-05,5.36818154152171e-05,6.69896709294729e-05,8.3318885310522e-05,0.000102983956134408,0.000126498298331752,0.000154730682173028,0.000190302109837553,0.000232693325379889,0.000282887107986215,0.000341940722605073,0.000413301608429019,0.000499754576070899,0.000601037492459244,0.000719007401944926,0.000855635845938929,0.00102111611522097,0.00121448085588383,0.00143728365038259,0.00169268162805242,0.00198657403733727,0.00233608934839894,0.00273424845690945,0.00318566199316614,0.00369511922572534,0.0042814032141455,0.00495505846024811,0.00571002976393804,0.00655255129910043,0.00748900648515755,0.00856214725748036,0.00975746016991333,0.0110757564322938,0.0125242291467235,0.0141199291820635,0.0159117621641438,0.0178638517531702,0.0199829578514071,0.0222756657933406,0.0247850739579746,0.0275201727109328,0.0304521295921115,0.0335849799124515,0.0369223325564482,0.0405343808359278,0.0443684232144798,0.048412066323137,0.05266404318698,0.0571363829343619,0.0618693810213739,0.0667933200181362,0.0719006366741757,0.0771829961128537,0.082659522316567,0.0883054678194522,0.0940779163162582,0.0999624035250797,0.105943845369191,0.112020757775016,0.118144215128315,0.124292196205171,0.130446346556428,0.136584838051448,0.14266590311761,0.148676297282236,0.154599044034482,0.160417727787576,0.166087158498287,0.171585731575429,0.176925174413171,0.182096837806915,0.187093297265822,0.191848462805082,0.196412200505378,0.200796905701498,0.20500790554435,0.209040607455964,0.212883049707972,0.216601840552749,0.220215940458855,0.223745734112598,0.227210595103176,0.230660942653852,0.234134890276146,0.237661870802475,0.24127193940771,0.245055729041931,0.249027336987266,0.253211115565283,0.257636316427382,0.262357737756768,0.267485585556989,0.27296019008926,0.278799718008996,0.285020386486772,0.291722264342989,0.298900412215107,0.306487809199709,0.314480742249643,0.322872928519874,0.331773204861097,0.34103728911715,0.350625066866474,0.360509126843619,0.370677708552897,0.381110641558705,0.391691643171896,0.402379147301527,0.413130764975761,0.423889122351575,0.434572187454015,0.445136401104692,0.455540418447746,0.465743930735662,0.475602449638224,0.48514226192016,0.494347268984643,0.503195413810198,0.511631277066041,0.519562192951398,0.527098253304393,0.534246324923226,0.541016996272716,0.547356763578811,0.55333718985744,0.559051726866273,0.564541391642729,0.569850406610259,0.575023117944408,0.580185669020055,0.585403116285688,0.590738595417642,0.596285522842156,0.602214872365891,0.608538484796591,0.615320386383836,0.622623300780423,0.630662525796753,0.639495467120669,0.649075248703433,0.659443031821837,0.670636446549689,0.683003224766195,0.696290004029568,0.710479045073503,0.725570258192401,0.741645617923657,0.75882121037335,0.776814098803347,0.7955824734783,0.815079904300484,0.835379950118662,0.856310437359097,0.877697510562037,0.899461388731854,0.921519188904802,0.943786023938476,0.966057135690827,0.988232318528252,1.01021428183188,1.03185623494685,1.05290927231027,1.0733627062222,1.09312501934501,1.11210670097511,1.12998308217192,1.14669031384652,1.16228986909844,1.17671823772205,1.18991573191053,1.20133519484263,1.21136050091836,1.21998725169373,1.22719321316637,1.23280244717625,1.23658121430498,1.2389240419527,1.23984654459563,1.23936836743949,1.23720514323324,1.23355461660342,1.22865240036676,1.2225464538886,1.21528733699739,1.20657685779707,1.19688482181882,1.18628739757544,1.17484773508307,1.16255419196159,1.14942333427728,1.13571877743196,1.1215024186752,1.10683523733113,1.09170995005438,1.07626637266278,1.06060505481901,1.04477489523049,1.02882307190724,1.01279762897796,0.996779396625618,0.980802285898652,0.964898001702712,0.949113880918398,0.933515064844472,0.918093387910821,0.902867060171735,0.887852959607035,0.87312797643806,0.858686458707843,0.844506605950202,0.830596130543193,0.816961868695549,0.803717173067409,0.790762681617046,0.778098817239408,0.765726537747002,0.753681728365397,0.741999726615821,0.730603612530492,0.719489735497536,0.708653951877653,0.698156675803699,0.687953288359587,0.678002114376213,0.668295338248377,0.658824883318442,0.649659608062432,0.640701109993922,0.631939592131762,0.623365220762666,0.614988367358886,0.606810832168403,0.598779882088708,0.590886650376946,0.583122546027826,0.575506298677956,0.568009688555495,0.560612480966763,0.553309200681906,0.546095165330095,0.538993758610154,0.531969164429461,0.525018610192539,0.518139462419462,0.511337790684076,0.504616588113662,0.497956430530263,0.491354261784246,0.484806748931685,0.478320850808876,0.471880515187846,0.465473339300539,0.45909208421561,0.452728977483651,0.446370151636633,0.439997533466835,0.433599225923292,0.427162925893987,0.420665593431491,0.414074624416134,0.407388865303944,0.400594512513625,0.393677905607654,0.386583359101276,0.379310479319655,0.371865022555584,0.36423771039219,0.356417337625579,0.34832450589634,0.340026511182669,0.331523058341844,0.322815106034853,0.313878048402364,0.304704930214177,0.295353493194222,0.285835354759425,0.276163289686038,0.266323448880213,0.256367886983996,0.246330916072009,0.236233034433003,0],"text":["density: 2.362330e-01<br />Sepal.Length: 4.300000","density: 2.463309e-01<br />Sepal.Length: 4.307045","density: 2.563679e-01<br />Sepal.Length: 4.314090","density: 2.663234e-01<br />Sepal.Length: 4.321135","density: 2.761633e-01<br />Sepal.Length: 4.328180","density: 2.858354e-01<br />Sepal.Length: 4.335225","density: 2.953535e-01<br />Sepal.Length: 4.342270","density: 3.047049e-01<br />Sepal.Length: 4.349315","density: 3.138780e-01<br />Sepal.Length: 4.356360","density: 3.228151e-01<br />Sepal.Length: 4.363405","density: 3.315231e-01<br />Sepal.Length: 4.370450","density: 3.400265e-01<br />Sepal.Length: 4.377495","density: 3.483245e-01<br />Sepal.Length: 4.384540","density: 3.564173e-01<br />Sepal.Length: 4.391585","density: 3.642377e-01<br />Sepal.Length: 4.398630","density: 3.718650e-01<br />Sepal.Length: 4.405675","density: 3.793105e-01<br />Sepal.Length: 4.412720","density: 3.865834e-01<br />Sepal.Length: 4.419765","density: 3.936779e-01<br />Sepal.Length: 4.426810","density: 4.005945e-01<br />Sepal.Length: 4.433855","density: 4.073889e-01<br />Sepal.Length: 4.440900","density: 4.140746e-01<br />Sepal.Length: 4.447945","density: 4.206656e-01<br />Sepal.Length: 4.454990","density: 4.271629e-01<br />Sepal.Length: 4.462035","density: 4.335992e-01<br />Sepal.Length: 4.469080","density: 4.399975e-01<br />Sepal.Length: 4.476125","density: 4.463702e-01<br />Sepal.Length: 4.483170","density: 4.527290e-01<br />Sepal.Length: 4.490215","density: 4.590921e-01<br />Sepal.Length: 4.497260","density: 4.654733e-01<br />Sepal.Length: 4.504305","density: 4.718805e-01<br />Sepal.Length: 4.511350","density: 4.783209e-01<br />Sepal.Length: 4.518395","density: 4.848067e-01<br />Sepal.Length: 4.525440","density: 4.913543e-01<br />Sepal.Length: 4.532485","density: 4.979564e-01<br />Sepal.Length: 4.539530","density: 5.046166e-01<br />Sepal.Length: 4.546575","density: 5.113378e-01<br />Sepal.Length: 4.553620","density: 5.181395e-01<br />Sepal.Length: 4.560665","density: 5.250186e-01<br />Sepal.Length: 4.567710","density: 5.319692e-01<br />Sepal.Length: 4.574755","density: 5.389938e-01<br />Sepal.Length: 4.581800","density: 5.460952e-01<br />Sepal.Length: 4.588845","density: 5.533092e-01<br />Sepal.Length: 4.595890","density: 5.606125e-01<br />Sepal.Length: 4.602935","density: 5.680097e-01<br />Sepal.Length: 4.609980","density: 5.755063e-01<br />Sepal.Length: 4.617025","density: 5.831225e-01<br />Sepal.Length: 4.624070","density: 5.908867e-01<br />Sepal.Length: 4.631115","density: 5.987799e-01<br />Sepal.Length: 4.638160","density: 6.068108e-01<br />Sepal.Length: 4.645205","density: 6.149884e-01<br />Sepal.Length: 4.652250","density: 6.233652e-01<br />Sepal.Length: 4.659295","density: 6.319396e-01<br />Sepal.Length: 4.666341","density: 6.407011e-01<br />Sepal.Length: 4.673386","density: 6.496596e-01<br />Sepal.Length: 4.680431","density: 6.588249e-01<br />Sepal.Length: 4.687476","density: 6.682953e-01<br />Sepal.Length: 4.694521","density: 6.780021e-01<br />Sepal.Length: 4.701566","density: 6.879533e-01<br />Sepal.Length: 4.708611","density: 6.981567e-01<br />Sepal.Length: 4.715656","density: 7.086540e-01<br />Sepal.Length: 4.722701","density: 7.194897e-01<br />Sepal.Length: 4.729746","density: 7.306036e-01<br />Sepal.Length: 4.736791","density: 7.419997e-01<br />Sepal.Length: 4.743836","density: 7.536817e-01<br />Sepal.Length: 4.750881","density: 7.657265e-01<br />Sepal.Length: 4.757926","density: 7.780988e-01<br />Sepal.Length: 4.764971","density: 7.907627e-01<br />Sepal.Length: 4.772016","density: 8.037172e-01<br />Sepal.Length: 4.779061","density: 8.169619e-01<br />Sepal.Length: 4.786106","density: 8.305961e-01<br />Sepal.Length: 4.793151","density: 8.445066e-01<br />Sepal.Length: 4.800196","density: 8.586865e-01<br />Sepal.Length: 4.807241","density: 8.731280e-01<br />Sepal.Length: 4.814286","density: 8.878530e-01<br />Sepal.Length: 4.821331","density: 9.028671e-01<br />Sepal.Length: 4.828376","density: 9.180934e-01<br />Sepal.Length: 4.835421","density: 9.335151e-01<br />Sepal.Length: 4.842466","density: 9.491139e-01<br />Sepal.Length: 4.849511","density: 9.648980e-01<br />Sepal.Length: 4.856556","density: 9.808023e-01<br />Sepal.Length: 4.863601","density: 9.967794e-01<br />Sepal.Length: 4.870646","density: 1.012798e+00<br />Sepal.Length: 4.877691","density: 1.028823e+00<br />Sepal.Length: 4.884736","density: 1.044775e+00<br />Sepal.Length: 4.891781","density: 1.060605e+00<br />Sepal.Length: 4.898826","density: 1.076266e+00<br />Sepal.Length: 4.905871","density: 1.091710e+00<br />Sepal.Length: 4.912916","density: 1.106835e+00<br />Sepal.Length: 4.919961","density: 1.121502e+00<br />Sepal.Length: 4.927006","density: 1.135719e+00<br />Sepal.Length: 4.934051","density: 1.149423e+00<br />Sepal.Length: 4.941096","density: 1.162554e+00<br />Sepal.Length: 4.948141","density: 1.174848e+00<br />Sepal.Length: 4.955186","density: 1.186287e+00<br />Sepal.Length: 4.962231","density: 1.196885e+00<br />Sepal.Length: 4.969276","density: 1.206577e+00<br />Sepal.Length: 4.976321","density: 1.215287e+00<br />Sepal.Length: 4.983366","density: 1.222546e+00<br />Sepal.Length: 4.990411","density: 1.228652e+00<br />Sepal.Length: 4.997456","density: 1.233555e+00<br />Sepal.Length: 5.004501","density: 1.237205e+00<br />Sepal.Length: 5.011546","density: 1.239368e+00<br />Sepal.Length: 5.018591","density: 1.239847e+00<br />Sepal.Length: 5.025636","density: 1.238924e+00<br />Sepal.Length: 5.032681","density: 1.236581e+00<br />Sepal.Length: 5.039726","density: 1.232802e+00<br />Sepal.Length: 5.046771","density: 1.227193e+00<br />Sepal.Length: 5.053816","density: 1.219987e+00<br />Sepal.Length: 5.060861","density: 1.211361e+00<br />Sepal.Length: 5.067906","density: 1.201335e+00<br />Sepal.Length: 5.074951","density: 1.189916e+00<br />Sepal.Length: 5.081996","density: 1.176718e+00<br />Sepal.Length: 5.089041","density: 1.162290e+00<br />Sepal.Length: 5.096086","density: 1.146690e+00<br />Sepal.Length: 5.103131","density: 1.129983e+00<br />Sepal.Length: 5.110176","density: 1.112107e+00<br />Sepal.Length: 5.117221","density: 1.093125e+00<br />Sepal.Length: 5.124266","density: 1.073363e+00<br />Sepal.Length: 5.131311","density: 1.052909e+00<br />Sepal.Length: 5.138356","density: 1.031856e+00<br />Sepal.Length: 5.145401","density: 1.010214e+00<br />Sepal.Length: 5.152446","density: 9.882323e-01<br />Sepal.Length: 5.159491","density: 9.660571e-01<br />Sepal.Length: 5.166536","density: 9.437860e-01<br />Sepal.Length: 5.173581","density: 9.215192e-01<br />Sepal.Length: 5.180626","density: 8.994614e-01<br />Sepal.Length: 5.187671","density: 8.776975e-01<br />Sepal.Length: 5.194716","density: 8.563104e-01<br />Sepal.Length: 5.201761","density: 8.353800e-01<br />Sepal.Length: 5.208806","density: 8.150799e-01<br />Sepal.Length: 5.215851","density: 7.955825e-01<br />Sepal.Length: 5.222896","density: 7.768141e-01<br />Sepal.Length: 5.229941","density: 7.588212e-01<br />Sepal.Length: 5.236986","density: 7.416456e-01<br />Sepal.Length: 5.244031","density: 7.255703e-01<br />Sepal.Length: 5.251076","density: 7.104790e-01<br />Sepal.Length: 5.258121","density: 6.962900e-01<br />Sepal.Length: 5.265166","density: 6.830032e-01<br />Sepal.Length: 5.272211","density: 6.706364e-01<br />Sepal.Length: 5.279256","density: 6.594430e-01<br />Sepal.Length: 5.286301","density: 6.490752e-01<br />Sepal.Length: 5.293346","density: 6.394955e-01<br />Sepal.Length: 5.300391","density: 6.306625e-01<br />Sepal.Length: 5.307436","density: 6.226233e-01<br />Sepal.Length: 5.314481","density: 6.153204e-01<br />Sepal.Length: 5.321526","density: 6.085385e-01<br />Sepal.Length: 5.328571","density: 6.022149e-01<br />Sepal.Length: 5.335616","density: 5.962855e-01<br />Sepal.Length: 5.342661","density: 5.907386e-01<br />Sepal.Length: 5.349706","density: 5.854031e-01<br />Sepal.Length: 5.356751","density: 5.801857e-01<br />Sepal.Length: 5.363796","density: 5.750231e-01<br />Sepal.Length: 5.370841","density: 5.698504e-01<br />Sepal.Length: 5.377886","density: 5.645414e-01<br />Sepal.Length: 5.384932","density: 5.590517e-01<br />Sepal.Length: 5.391977","density: 5.533372e-01<br />Sepal.Length: 5.399022","density: 5.473568e-01<br />Sepal.Length: 5.406067","density: 5.410170e-01<br />Sepal.Length: 5.413112","density: 5.342463e-01<br />Sepal.Length: 5.420157","density: 5.270983e-01<br />Sepal.Length: 5.427202","density: 5.195622e-01<br />Sepal.Length: 5.434247","density: 5.116313e-01<br />Sepal.Length: 5.441292","density: 5.031954e-01<br />Sepal.Length: 5.448337","density: 4.943473e-01<br />Sepal.Length: 5.455382","density: 4.851423e-01<br />Sepal.Length: 5.462427","density: 4.756024e-01<br />Sepal.Length: 5.469472","density: 4.657439e-01<br />Sepal.Length: 5.476517","density: 4.555404e-01<br />Sepal.Length: 5.483562","density: 4.451364e-01<br />Sepal.Length: 5.490607","density: 4.345722e-01<br />Sepal.Length: 5.497652","density: 4.238891e-01<br />Sepal.Length: 5.504697","density: 4.131308e-01<br />Sepal.Length: 5.511742","density: 4.023791e-01<br />Sepal.Length: 5.518787","density: 3.916916e-01<br />Sepal.Length: 5.525832","density: 3.811106e-01<br />Sepal.Length: 5.532877","density: 3.706777e-01<br />Sepal.Length: 5.539922","density: 3.605091e-01<br />Sepal.Length: 5.546967","density: 3.506251e-01<br />Sepal.Length: 5.554012","density: 3.410373e-01<br />Sepal.Length: 5.561057","density: 3.317732e-01<br />Sepal.Length: 5.568102","density: 3.228729e-01<br />Sepal.Length: 5.575147","density: 3.144807e-01<br />Sepal.Length: 5.582192","density: 3.064878e-01<br />Sepal.Length: 5.589237","density: 2.989004e-01<br />Sepal.Length: 5.596282","density: 2.917223e-01<br />Sepal.Length: 5.603327","density: 2.850204e-01<br />Sepal.Length: 5.610372","density: 2.787997e-01<br />Sepal.Length: 5.617417","density: 2.729602e-01<br />Sepal.Length: 5.624462","density: 2.674856e-01<br />Sepal.Length: 5.631507","density: 2.623577e-01<br />Sepal.Length: 5.638552","density: 2.576363e-01<br />Sepal.Length: 5.645597","density: 2.532111e-01<br />Sepal.Length: 5.652642","density: 2.490273e-01<br />Sepal.Length: 5.659687","density: 2.450557e-01<br />Sepal.Length: 5.666732","density: 2.412719e-01<br />Sepal.Length: 5.673777","density: 2.376619e-01<br />Sepal.Length: 5.680822","density: 2.341349e-01<br />Sepal.Length: 5.687867","density: 2.306609e-01<br />Sepal.Length: 5.694912","density: 2.272106e-01<br />Sepal.Length: 5.701957","density: 2.237457e-01<br />Sepal.Length: 5.709002","density: 2.202159e-01<br />Sepal.Length: 5.716047","density: 2.166018e-01<br />Sepal.Length: 5.723092","density: 2.128830e-01<br />Sepal.Length: 5.730137","density: 2.090406e-01<br />Sepal.Length: 5.737182","density: 2.050079e-01<br />Sepal.Length: 5.744227","density: 2.007969e-01<br />Sepal.Length: 5.751272","density: 1.964122e-01<br />Sepal.Length: 5.758317","density: 1.918485e-01<br />Sepal.Length: 5.765362","density: 1.870933e-01<br />Sepal.Length: 5.772407","density: 1.820968e-01<br />Sepal.Length: 5.779452","density: 1.769252e-01<br />Sepal.Length: 5.786497","density: 1.715857e-01<br />Sepal.Length: 5.793542","density: 1.660872e-01<br />Sepal.Length: 5.800587","density: 1.604177e-01<br />Sepal.Length: 5.807632","density: 1.545990e-01<br />Sepal.Length: 5.814677","density: 1.486763e-01<br />Sepal.Length: 5.821722","density: 1.426659e-01<br />Sepal.Length: 5.828767","density: 1.365848e-01<br />Sepal.Length: 5.835812","density: 1.304463e-01<br />Sepal.Length: 5.842857","density: 1.242922e-01<br />Sepal.Length: 5.849902","density: 1.181442e-01<br />Sepal.Length: 5.856947","density: 1.120208e-01<br />Sepal.Length: 5.863992","density: 1.059438e-01<br />Sepal.Length: 5.871037","density: 9.996240e-02<br />Sepal.Length: 5.878082","density: 9.407792e-02<br />Sepal.Length: 5.885127","density: 8.830547e-02<br />Sepal.Length: 5.892172","density: 8.265952e-02<br />Sepal.Length: 5.899217","density: 7.718300e-02<br />Sepal.Length: 5.906262","density: 7.190064e-02<br />Sepal.Length: 5.913307","density: 6.679332e-02<br />Sepal.Length: 5.920352","density: 6.186938e-02<br />Sepal.Length: 5.927397","density: 5.713638e-02<br />Sepal.Length: 5.934442","density: 5.266404e-02<br />Sepal.Length: 5.941487","density: 4.841207e-02<br />Sepal.Length: 5.948532","density: 4.436842e-02<br />Sepal.Length: 5.955577","density: 4.053438e-02<br />Sepal.Length: 5.962622","density: 3.692233e-02<br />Sepal.Length: 5.969667","density: 3.358498e-02<br />Sepal.Length: 5.976712","density: 3.045213e-02<br />Sepal.Length: 5.983757","density: 2.752017e-02<br />Sepal.Length: 5.990802","density: 2.478507e-02<br />Sepal.Length: 5.997847","density: 2.227567e-02<br />Sepal.Length: 6.004892","density: 1.998296e-02<br />Sepal.Length: 6.011937","density: 1.786385e-02<br />Sepal.Length: 6.018982","density: 1.591176e-02<br />Sepal.Length: 6.026027","density: 1.411993e-02<br />Sepal.Length: 6.033072","density: 1.252423e-02<br />Sepal.Length: 6.040117","density: 1.107576e-02<br />Sepal.Length: 6.047162","density: 9.757460e-03<br />Sepal.Length: 6.054207","density: 8.562147e-03<br />Sepal.Length: 6.061252","density: 7.489006e-03<br />Sepal.Length: 6.068297","density: 6.552551e-03<br />Sepal.Length: 6.075342","density: 5.710030e-03<br />Sepal.Length: 6.082387","density: 4.955058e-03<br />Sepal.Length: 6.089432","density: 4.281403e-03<br />Sepal.Length: 6.096477","density: 3.695119e-03<br />Sepal.Length: 6.103523","density: 3.185662e-03<br />Sepal.Length: 6.110568","density: 2.734248e-03<br />Sepal.Length: 6.117613","density: 2.336089e-03<br />Sepal.Length: 6.124658","density: 1.986574e-03<br />Sepal.Length: 6.131703","density: 1.692682e-03<br />Sepal.Length: 6.138748","density: 1.437284e-03<br />Sepal.Length: 6.145793","density: 1.214481e-03<br />Sepal.Length: 6.152838","density: 1.021116e-03<br />Sepal.Length: 6.159883","density: 8.556358e-04<br />Sepal.Length: 6.166928","density: 7.190074e-04<br />Sepal.Length: 6.173973","density: 6.010375e-04<br />Sepal.Length: 6.181018","density: 4.997546e-04<br />Sepal.Length: 6.188063","density: 4.133016e-04<br />Sepal.Length: 6.195108","density: 3.419407e-04<br />Sepal.Length: 6.202153","density: 2.828871e-04<br />Sepal.Length: 6.209198","density: 2.326933e-04<br />Sepal.Length: 6.216243","density: 1.903021e-04<br />Sepal.Length: 6.223288","density: 1.547307e-04<br />Sepal.Length: 6.230333","density: 1.264983e-04<br />Sepal.Length: 6.237378","density: 1.029840e-04<br />Sepal.Length: 6.244423","density: 8.331889e-05<br />Sepal.Length: 6.251468","density: 6.698967e-05<br />Sepal.Length: 6.258513","density: 5.368182e-05<br />Sepal.Length: 6.265558","density: 4.328059e-05<br />Sepal.Length: 6.272603","density: 3.465789e-05<br />Sepal.Length: 6.279648","density: 2.756646e-05<br />Sepal.Length: 6.286693","density: 2.177987e-05<br />Sepal.Length: 6.293738","density: 1.725373e-05<br />Sepal.Length: 6.300783","density: 1.368420e-05<br />Sepal.Length: 6.307828","density: 1.077341e-05<br />Sepal.Length: 6.314873","density: 8.420588e-06<br />Sepal.Length: 6.321918","density: 6.534898e-06<br />Sepal.Length: 6.328963","density: 5.123201e-06<br />Sepal.Length: 6.336008","density: 3.995256e-06<br />Sepal.Length: 6.343053","density: 3.091047e-06<br />Sepal.Length: 6.350098","density: 2.373097e-06<br />Sepal.Length: 6.357143","density: 1.816492e-06<br />Sepal.Length: 6.364188","density: 1.404487e-06<br />Sepal.Length: 6.371233","density: 1.076424e-06<br />Sepal.Length: 6.378278","density: 8.180455e-07<br />Sepal.Length: 6.385323","density: 6.166318e-07<br />Sepal.Length: 6.392368","density: 4.674687e-07<br />Sepal.Length: 6.399413","density: 3.552673e-07<br />Sepal.Length: 6.406458","density: 2.674736e-07<br />Sepal.Length: 6.413503","density: 1.995814e-07<br />Sepal.Length: 6.420548","density: 1.476500e-07<br />Sepal.Length: 6.427593","density: 1.109983e-07<br />Sepal.Length: 6.434638","density: 8.287468e-08<br />Sepal.Length: 6.441683","density: 6.126463e-08<br />Sepal.Length: 6.448728","density: 4.486564e-08<br />Sepal.Length: 6.455773","density: 3.278222e-08<br />Sepal.Length: 6.462818","density: 2.430585e-08<br />Sepal.Length: 6.469863","density: 1.781997e-08<br />Sepal.Length: 6.476908","density: 1.292905e-08<br />Sepal.Length: 6.483953","density: 9.288899e-09<br />Sepal.Length: 6.490998","density: 6.738060e-09<br />Sepal.Length: 6.498043","density: 4.906122e-09<br />Sepal.Length: 6.505088","density: 3.530365e-09<br />Sepal.Length: 6.512133","density: 2.512864e-09<br />Sepal.Length: 6.519178","density: 1.770515e-09<br />Sepal.Length: 6.526223","density: 1.276758e-09<br />Sepal.Length: 6.533268","density: 9.124654e-10<br />Sepal.Length: 6.540313","density: 6.441484e-10<br />Sepal.Length: 6.547358","density: 4.496287e-10<br />Sepal.Length: 6.554403","density: 3.135056e-10<br />Sepal.Length: 6.561448","density: 2.229363e-10<br />Sepal.Length: 6.568493","density: 1.563061e-10<br />Sepal.Length: 6.575538","density: 1.082053e-10<br />Sepal.Length: 6.582583","density: 7.404160e-11<br />Sepal.Length: 6.589628","density: 5.140019e-11<br />Sepal.Length: 6.596673","density: 3.585784e-11<br />Sepal.Length: 6.603718","density: 2.465252e-11<br />Sepal.Length: 6.610763","density: 1.672872e-11<br />Sepal.Length: 6.617808","density: 1.121764e-11<br />Sepal.Length: 6.624853","density: 7.764002e-12<br />Sepal.Length: 6.631898","density: 5.310754e-12<br />Sepal.Length: 6.638943","density: 3.578669e-12<br />Sepal.Length: 6.645988","density: 2.379515e-12<br />Sepal.Length: 6.653033","density: 1.583129e-12<br />Sepal.Length: 6.660078","density: 1.080031e-12<br />Sepal.Length: 6.667123","density: 7.239930e-13<br />Sepal.Length: 6.674168","density: 4.779687e-13<br />Sepal.Length: 6.681213","density: 3.112831e-13<br />Sepal.Length: 6.688258","density: 2.068741e-13<br />Sepal.Length: 6.695303","density: 1.382971e-13<br />Sepal.Length: 6.702348","density: 9.081367e-14<br />Sepal.Length: 6.709393","density: 5.870449e-14<br />Sepal.Length: 6.716438","density: 3.741739e-14<br />Sepal.Length: 6.723483","density: 2.489158e-14<br />Sepal.Length: 6.730528","density: 1.628970e-14<br />Sepal.Length: 6.737573","density: 1.046131e-14<br />Sepal.Length: 6.744618","density: 6.613337e-15<br />Sepal.Length: 6.751663","density: 4.193207e-15<br />Sepal.Length: 6.758708","density: 2.764951e-15<br />Sepal.Length: 6.765753","density: 1.789144e-15<br />Sepal.Length: 6.772798","density: 1.148963e-15<br />Sepal.Length: 6.779843","density: 7.385872e-16<br />Sepal.Length: 6.786888","density: 4.006411e-16<br />Sepal.Length: 6.793933","density: 2.781746e-16<br />Sepal.Length: 6.800978","density: 2.026065e-16<br />Sepal.Length: 6.808023","density: 1.172715e-16<br />Sepal.Length: 6.815068","density: 0.000000e+00<br />Sepal.Length: 6.822114","density: 0.000000e+00<br />Sepal.Length: 6.829159","density: 0.000000e+00<br />Sepal.Length: 6.836204","density: 0.000000e+00<br />Sepal.Length: 6.843249","density: 0.000000e+00<br />Sepal.Length: 6.850294","density: 0.000000e+00<br />Sepal.Length: 6.857339","density: 0.000000e+00<br />Sepal.Length: 6.864384","density: 0.000000e+00<br />Sepal.Length: 6.871429","density: 0.000000e+00<br />Sepal.Length: 6.878474","density: 1.813955e-18<br />Sepal.Length: 6.885519","density: 1.816618e-17<br />Sepal.Length: 6.892564","density: 7.114949e-18<br />Sepal.Length: 6.899609","density: 0.000000e+00<br />Sepal.Length: 6.906654","density: 0.000000e+00<br />Sepal.Length: 6.913699","density: 0.000000e+00<br />Sepal.Length: 6.920744","density: 0.000000e+00<br />Sepal.Length: 6.927789","density: 0.000000e+00<br />Sepal.Length: 6.934834","density: 0.000000e+00<br />Sepal.Length: 6.941879","density: 2.178166e-18<br />Sepal.Length: 6.948924","density: 1.307965e-17<br />Sepal.Length: 6.955969","density: 3.774436e-18<br />Sepal.Length: 6.963014","density: 0.000000e+00<br />Sepal.Length: 6.970059","density: 0.000000e+00<br />Sepal.Length: 6.977104","density: 0.000000e+00<br />Sepal.Length: 6.984149","density: 0.000000e+00<br />Sepal.Length: 6.991194","density: 0.000000e+00<br />Sepal.Length: 6.998239","density: 0.000000e+00<br />Sepal.Length: 7.005284","density: 0.000000e+00<br />Sepal.Length: 7.012329","density: 0.000000e+00<br />Sepal.Length: 7.019374","density: 0.000000e+00<br />Sepal.Length: 7.026419","density: 0.000000e+00<br />Sepal.Length: 7.033464","density: 0.000000e+00<br />Sepal.Length: 7.040509","density: 0.000000e+00<br />Sepal.Length: 7.047554","density: 0.000000e+00<br />Sepal.Length: 7.054599","density: 1.006850e-17<br />Sepal.Length: 7.061644","density: 6.785594e-18<br />Sepal.Length: 7.068689","density: 0.000000e+00<br />Sepal.Length: 7.075734","density: 0.000000e+00<br />Sepal.Length: 7.082779","density: 0.000000e+00<br />Sepal.Length: 7.089824","density: 0.000000e+00<br />Sepal.Length: 7.096869","density: 0.000000e+00<br />Sepal.Length: 7.103914","density: 0.000000e+00<br />Sepal.Length: 7.110959","density: 0.000000e+00<br />Sepal.Length: 7.118004","density: 0.000000e+00<br />Sepal.Length: 7.125049","density: 1.612211e-17<br />Sepal.Length: 7.132094","density: 1.758606e-17<br />Sepal.Length: 7.139139","density: 0.000000e+00<br />Sepal.Length: 7.146184","density: 0.000000e+00<br />Sepal.Length: 7.153229","density: 0.000000e+00<br />Sepal.Length: 7.160274","density: 0.000000e+00<br />Sepal.Length: 7.167319","density: 0.000000e+00<br />Sepal.Length: 7.174364","density: 0.000000e+00<br />Sepal.Length: 7.181409","density: 0.000000e+00<br />Sepal.Length: 7.188454","density: 0.000000e+00<br />Sepal.Length: 7.195499","density: 6.053619e-18<br />Sepal.Length: 7.202544","density: 1.080047e-17<br />Sepal.Length: 7.209589","density: 0.000000e+00<br />Sepal.Length: 7.216634","density: 0.000000e+00<br />Sepal.Length: 7.223679","density: 8.828822e-17<br />Sepal.Length: 7.230724","density: 1.274062e-16<br />Sepal.Length: 7.237769","density: 7.082514e-17<br />Sepal.Length: 7.244814","density: 5.416217e-18<br />Sepal.Length: 7.251859","density: 0.000000e+00<br />Sepal.Length: 7.258904","density: 2.808993e-17<br />Sepal.Length: 7.265949","density: 5.551115e-17<br />Sepal.Length: 7.272994","density: 5.123163e-17<br />Sepal.Length: 7.280039","density: 7.625688e-18<br />Sepal.Length: 7.287084","density: 1.799013e-17<br />Sepal.Length: 7.294129","density: 6.386815e-17<br />Sepal.Length: 7.301174","density: 9.885246e-17<br />Sepal.Length: 7.308219","density: 5.551115e-17<br />Sepal.Length: 7.315264","density: 5.551115e-17<br />Sepal.Length: 7.322309","density: 8.747653e-17<br />Sepal.Length: 7.329354","density: 9.096213e-17<br />Sepal.Length: 7.336399","density: 6.774361e-17<br />Sepal.Length: 7.343444","density: 1.331525e-16<br />Sepal.Length: 7.350489","density: 1.387779e-16<br />Sepal.Length: 7.357534","density: 1.108274e-16<br />Sepal.Length: 7.364579","density: 1.233800e-16<br />Sepal.Length: 7.371624","density: 2.199746e-16<br />Sepal.Length: 7.378669","density: 1.981716e-16<br />Sepal.Length: 7.385714","density: 1.226074e-16<br />Sepal.Length: 7.392759","density: 9.523454e-17<br />Sepal.Length: 7.399804","density: 1.230527e-16<br />Sepal.Length: 7.406849","density: 1.664708e-16<br />Sepal.Length: 7.413894","density: 1.446679e-16<br />Sepal.Length: 7.420939","density: 1.228649e-16<br />Sepal.Length: 7.427984","density: 9.110155e-17<br />Sepal.Length: 7.435029","density: 7.555002e-17<br />Sepal.Length: 7.442074","density: 1.845649e-16<br />Sepal.Length: 7.449119","density: 1.545727e-16<br />Sepal.Length: 7.456164","density: 1.526834e-16<br />Sepal.Length: 7.463209","density: 1.585805e-16<br />Sepal.Length: 7.470254","density: 1.357774e-16<br />Sepal.Length: 7.477299","density: 1.030729e-16<br />Sepal.Length: 7.484344","density: 1.328459e-16<br />Sepal.Length: 7.491389","density: 1.288595e-16<br />Sepal.Length: 7.498434","density: 9.119901e-17<br />Sepal.Length: 7.505479","density: 6.939604e-17<br />Sepal.Length: 7.512524","density: 1.238786e-16<br />Sepal.Length: 7.519569","density: 1.150130e-16<br />Sepal.Length: 7.526614","density: 6.747252e-17<br />Sepal.Length: 7.533659","density: 1.190876e-17<br />Sepal.Length: 7.540705","density: 1.007271e-18<br />Sepal.Length: 7.547750","density: 2.968265e-17<br />Sepal.Length: 7.554795","density: 6.930502e-17<br />Sepal.Length: 7.561840","density: 6.955322e-17<br />Sepal.Length: 7.568885","density: 0.000000e+00<br />Sepal.Length: 7.575930","density: 0.000000e+00<br />Sepal.Length: 7.582975","density: 8.890497e-18<br />Sepal.Length: 7.590020","density: 2.570618e-17<br />Sepal.Length: 7.597065","density: 3.575758e-17<br />Sepal.Length: 7.604110","density: 1.395460e-17<br />Sepal.Length: 7.611155","density: 3.014709e-18<br />Sepal.Length: 7.618200","density: 0.000000e+00<br />Sepal.Length: 7.625245","density: 4.910476e-18<br />Sepal.Length: 7.632290","density: 1.194361e-17<br />Sepal.Length: 7.639335","density: 1.042127e-18<br />Sepal.Length: 7.646380","density: 0.000000e+00<br />Sepal.Length: 7.653425","density: 1.376612e-17<br />Sepal.Length: 7.660470","density: 2.775558e-17<br />Sepal.Length: 7.667515","density: 2.682512e-17<br />Sepal.Length: 7.674560","density: 1.592363e-17<br />Sepal.Length: 7.681605","density: 5.022147e-18<br />Sepal.Length: 7.688650","density: 0.000000e+00<br />Sepal.Length: 7.695695","density: 0.000000e+00<br />Sepal.Length: 7.702740","density: 0.000000e+00<br />Sepal.Length: 7.709785","density: 0.000000e+00<br />Sepal.Length: 7.716830","density: 0.000000e+00<br />Sepal.Length: 7.723875","density: 0.000000e+00<br />Sepal.Length: 7.730920","density: 0.000000e+00<br />Sepal.Length: 7.737965","density: 0.000000e+00<br />Sepal.Length: 7.745010","density: 0.000000e+00<br />Sepal.Length: 7.752055","density: 0.000000e+00<br />Sepal.Length: 7.759100","density: 0.000000e+00<br />Sepal.Length: 7.766145","density: 0.000000e+00<br />Sepal.Length: 7.773190","density: 0.000000e+00<br />Sepal.Length: 7.780235","density: 0.000000e+00<br />Sepal.Length: 7.787280","density: 0.000000e+00<br />Sepal.Length: 7.794325","density: 0.000000e+00<br />Sepal.Length: 7.801370","density: 0.000000e+00<br />Sepal.Length: 7.808415","density: 0.000000e+00<br />Sepal.Length: 7.815460","density: 0.000000e+00<br />Sepal.Length: 7.822505","density: 0.000000e+00<br />Sepal.Length: 7.829550","density: 0.000000e+00<br />Sepal.Length: 7.836595","density: 0.000000e+00<br />Sepal.Length: 7.843640","density: 0.000000e+00<br />Sepal.Length: 7.850685","density: 0.000000e+00<br />Sepal.Length: 7.857730","density: 0.000000e+00<br />Sepal.Length: 7.864775","density: 0.000000e+00<br />Sepal.Length: 7.871820","density: 0.000000e+00<br />Sepal.Length: 7.878865","density: 0.000000e+00<br />Sepal.Length: 7.885910","density: 0.000000e+00<br />Sepal.Length: 7.892955","density: 0.000000e+00<br />Sepal.Length: 7.900000","density: 0.000000e+00<br />Sepal.Length: 7.900000","density: 0.000000e+00<br />Sepal.Length: 7.892955","density: 0.000000e+00<br />Sepal.Length: 7.885910","density: 0.000000e+00<br />Sepal.Length: 7.878865","density: 0.000000e+00<br />Sepal.Length: 7.871820","density: 0.000000e+00<br />Sepal.Length: 7.864775","density: 0.000000e+00<br />Sepal.Length: 7.857730","density: 0.000000e+00<br />Sepal.Length: 7.850685","density: 0.000000e+00<br />Sepal.Length: 7.843640","density: 0.000000e+00<br />Sepal.Length: 7.836595","density: 0.000000e+00<br />Sepal.Length: 7.829550","density: 0.000000e+00<br />Sepal.Length: 7.822505","density: 0.000000e+00<br />Sepal.Length: 7.815460","density: 0.000000e+00<br />Sepal.Length: 7.808415","density: 0.000000e+00<br />Sepal.Length: 7.801370","density: 0.000000e+00<br />Sepal.Length: 7.794325","density: 0.000000e+00<br />Sepal.Length: 7.787280","density: 0.000000e+00<br />Sepal.Length: 7.780235","density: 0.000000e+00<br />Sepal.Length: 7.773190","density: 0.000000e+00<br />Sepal.Length: 7.766145","density: 0.000000e+00<br />Sepal.Length: 7.759100","density: 0.000000e+00<br />Sepal.Length: 7.752055","density: 0.000000e+00<br />Sepal.Length: 7.745010","density: 0.000000e+00<br />Sepal.Length: 7.737965","density: 0.000000e+00<br />Sepal.Length: 7.730920","density: 0.000000e+00<br />Sepal.Length: 7.723875","density: 0.000000e+00<br />Sepal.Length: 7.716830","density: 0.000000e+00<br />Sepal.Length: 7.709785","density: 0.000000e+00<br />Sepal.Length: 7.702740","density: 0.000000e+00<br />Sepal.Length: 7.695695","density: 5.022147e-18<br />Sepal.Length: 7.688650","density: 1.592363e-17<br />Sepal.Length: 7.681605","density: 2.682512e-17<br />Sepal.Length: 7.674560","density: 2.775558e-17<br />Sepal.Length: 7.667515","density: 1.376612e-17<br />Sepal.Length: 7.660470","density: 0.000000e+00<br />Sepal.Length: 7.653425","density: 1.042127e-18<br />Sepal.Length: 7.646380","density: 1.194361e-17<br />Sepal.Length: 7.639335","density: 4.910476e-18<br />Sepal.Length: 7.632290","density: 0.000000e+00<br />Sepal.Length: 7.625245","density: 3.014709e-18<br />Sepal.Length: 7.618200","density: 1.395460e-17<br />Sepal.Length: 7.611155","density: 3.575758e-17<br />Sepal.Length: 7.604110","density: 2.570618e-17<br />Sepal.Length: 7.597065","density: 8.890497e-18<br />Sepal.Length: 7.590020","density: 0.000000e+00<br />Sepal.Length: 7.582975","density: 0.000000e+00<br />Sepal.Length: 7.575930","density: 6.955322e-17<br />Sepal.Length: 7.568885","density: 6.930502e-17<br />Sepal.Length: 7.561840","density: 2.968265e-17<br />Sepal.Length: 7.554795","density: 1.007271e-18<br />Sepal.Length: 7.547750","density: 1.190876e-17<br />Sepal.Length: 7.540705","density: 6.747252e-17<br />Sepal.Length: 7.533659","density: 1.150130e-16<br />Sepal.Length: 7.526614","density: 1.238786e-16<br />Sepal.Length: 7.519569","density: 6.939604e-17<br />Sepal.Length: 7.512524","density: 9.119901e-17<br />Sepal.Length: 7.505479","density: 1.288595e-16<br />Sepal.Length: 7.498434","density: 1.328459e-16<br />Sepal.Length: 7.491389","density: 1.030729e-16<br />Sepal.Length: 7.484344","density: 1.357774e-16<br />Sepal.Length: 7.477299","density: 1.585805e-16<br />Sepal.Length: 7.470254","density: 1.526834e-16<br />Sepal.Length: 7.463209","density: 1.545727e-16<br />Sepal.Length: 7.456164","density: 1.845649e-16<br />Sepal.Length: 7.449119","density: 7.555002e-17<br />Sepal.Length: 7.442074","density: 9.110155e-17<br />Sepal.Length: 7.435029","density: 1.228649e-16<br />Sepal.Length: 7.427984","density: 1.446679e-16<br />Sepal.Length: 7.420939","density: 1.664708e-16<br />Sepal.Length: 7.413894","density: 1.230527e-16<br />Sepal.Length: 7.406849","density: 9.523454e-17<br />Sepal.Length: 7.399804","density: 1.226074e-16<br />Sepal.Length: 7.392759","density: 1.981716e-16<br />Sepal.Length: 7.385714","density: 2.199746e-16<br />Sepal.Length: 7.378669","density: 1.233800e-16<br />Sepal.Length: 7.371624","density: 1.108274e-16<br />Sepal.Length: 7.364579","density: 1.387779e-16<br />Sepal.Length: 7.357534","density: 1.331525e-16<br />Sepal.Length: 7.350489","density: 6.774361e-17<br />Sepal.Length: 7.343444","density: 9.096213e-17<br />Sepal.Length: 7.336399","density: 8.747653e-17<br />Sepal.Length: 7.329354","density: 5.551115e-17<br />Sepal.Length: 7.322309","density: 5.551115e-17<br />Sepal.Length: 7.315264","density: 9.885246e-17<br />Sepal.Length: 7.308219","density: 6.386815e-17<br />Sepal.Length: 7.301174","density: 1.799013e-17<br />Sepal.Length: 7.294129","density: 7.625688e-18<br />Sepal.Length: 7.287084","density: 5.123163e-17<br />Sepal.Length: 7.280039","density: 5.551115e-17<br />Sepal.Length: 7.272994","density: 2.808993e-17<br />Sepal.Length: 7.265949","density: 0.000000e+00<br />Sepal.Length: 7.258904","density: 5.416217e-18<br />Sepal.Length: 7.251859","density: 7.082514e-17<br />Sepal.Length: 7.244814","density: 1.274062e-16<br />Sepal.Length: 7.237769","density: 8.828822e-17<br />Sepal.Length: 7.230724","density: 0.000000e+00<br />Sepal.Length: 7.223679","density: 0.000000e+00<br />Sepal.Length: 7.216634","density: 1.080047e-17<br />Sepal.Length: 7.209589","density: 6.053619e-18<br />Sepal.Length: 7.202544","density: 0.000000e+00<br />Sepal.Length: 7.195499","density: 0.000000e+00<br />Sepal.Length: 7.188454","density: 0.000000e+00<br />Sepal.Length: 7.181409","density: 0.000000e+00<br />Sepal.Length: 7.174364","density: 0.000000e+00<br />Sepal.Length: 7.167319","density: 0.000000e+00<br />Sepal.Length: 7.160274","density: 0.000000e+00<br />Sepal.Length: 7.153229","density: 0.000000e+00<br />Sepal.Length: 7.146184","density: 1.758606e-17<br />Sepal.Length: 7.139139","density: 1.612211e-17<br />Sepal.Length: 7.132094","density: 0.000000e+00<br />Sepal.Length: 7.125049","density: 0.000000e+00<br />Sepal.Length: 7.118004","density: 0.000000e+00<br />Sepal.Length: 7.110959","density: 0.000000e+00<br />Sepal.Length: 7.103914","density: 0.000000e+00<br />Sepal.Length: 7.096869","density: 0.000000e+00<br />Sepal.Length: 7.089824","density: 0.000000e+00<br />Sepal.Length: 7.082779","density: 0.000000e+00<br />Sepal.Length: 7.075734","density: 6.785594e-18<br />Sepal.Length: 7.068689","density: 1.006850e-17<br />Sepal.Length: 7.061644","density: 0.000000e+00<br />Sepal.Length: 7.054599","density: 0.000000e+00<br />Sepal.Length: 7.047554","density: 0.000000e+00<br />Sepal.Length: 7.040509","density: 0.000000e+00<br />Sepal.Length: 7.033464","density: 0.000000e+00<br />Sepal.Length: 7.026419","density: 0.000000e+00<br />Sepal.Length: 7.019374","density: 0.000000e+00<br />Sepal.Length: 7.012329","density: 0.000000e+00<br />Sepal.Length: 7.005284","density: 0.000000e+00<br />Sepal.Length: 6.998239","density: 0.000000e+00<br />Sepal.Length: 6.991194","density: 0.000000e+00<br />Sepal.Length: 6.984149","density: 0.000000e+00<br />Sepal.Length: 6.977104","density: 0.000000e+00<br />Sepal.Length: 6.970059","density: 3.774436e-18<br />Sepal.Length: 6.963014","density: 1.307965e-17<br />Sepal.Length: 6.955969","density: 2.178166e-18<br />Sepal.Length: 6.948924","density: 0.000000e+00<br />Sepal.Length: 6.941879","density: 0.000000e+00<br />Sepal.Length: 6.934834","density: 0.000000e+00<br />Sepal.Length: 6.927789","density: 0.000000e+00<br />Sepal.Length: 6.920744","density: 0.000000e+00<br />Sepal.Length: 6.913699","density: 0.000000e+00<br />Sepal.Length: 6.906654","density: 7.114949e-18<br />Sepal.Length: 6.899609","density: 1.816618e-17<br />Sepal.Length: 6.892564","density: 1.813955e-18<br />Sepal.Length: 6.885519","density: 0.000000e+00<br />Sepal.Length: 6.878474","density: 0.000000e+00<br />Sepal.Length: 6.871429","density: 0.000000e+00<br />Sepal.Length: 6.864384","density: 0.000000e+00<br />Sepal.Length: 6.857339","density: 0.000000e+00<br />Sepal.Length: 6.850294","density: 0.000000e+00<br />Sepal.Length: 6.843249","density: 0.000000e+00<br />Sepal.Length: 6.836204","density: 0.000000e+00<br />Sepal.Length: 6.829159","density: 0.000000e+00<br />Sepal.Length: 6.822114","density: 1.172715e-16<br />Sepal.Length: 6.815068","density: 2.026065e-16<br />Sepal.Length: 6.808023","density: 2.781746e-16<br />Sepal.Length: 6.800978","density: 4.006411e-16<br />Sepal.Length: 6.793933","density: 7.385872e-16<br />Sepal.Length: 6.786888","density: 1.148963e-15<br />Sepal.Length: 6.779843","density: 1.789144e-15<br />Sepal.Length: 6.772798","density: 2.764951e-15<br />Sepal.Length: 6.765753","density: 4.193207e-15<br />Sepal.Length: 6.758708","density: 6.613337e-15<br />Sepal.Length: 6.751663","density: 1.046131e-14<br />Sepal.Length: 6.744618","density: 1.628970e-14<br />Sepal.Length: 6.737573","density: 2.489158e-14<br />Sepal.Length: 6.730528","density: 3.741739e-14<br />Sepal.Length: 6.723483","density: 5.870449e-14<br />Sepal.Length: 6.716438","density: 9.081367e-14<br />Sepal.Length: 6.709393","density: 1.382971e-13<br />Sepal.Length: 6.702348","density: 2.068741e-13<br />Sepal.Length: 6.695303","density: 3.112831e-13<br />Sepal.Length: 6.688258","density: 4.779687e-13<br />Sepal.Length: 6.681213","density: 7.239930e-13<br />Sepal.Length: 6.674168","density: 1.080031e-12<br />Sepal.Length: 6.667123","density: 1.583129e-12<br />Sepal.Length: 6.660078","density: 2.379515e-12<br />Sepal.Length: 6.653033","density: 3.578669e-12<br />Sepal.Length: 6.645988","density: 5.310754e-12<br />Sepal.Length: 6.638943","density: 7.764002e-12<br />Sepal.Length: 6.631898","density: 1.121764e-11<br />Sepal.Length: 6.624853","density: 1.672872e-11<br />Sepal.Length: 6.617808","density: 2.465252e-11<br />Sepal.Length: 6.610763","density: 3.585784e-11<br />Sepal.Length: 6.603718","density: 5.140019e-11<br />Sepal.Length: 6.596673","density: 7.404160e-11<br />Sepal.Length: 6.589628","density: 1.082053e-10<br />Sepal.Length: 6.582583","density: 1.563061e-10<br />Sepal.Length: 6.575538","density: 2.229363e-10<br />Sepal.Length: 6.568493","density: 3.135056e-10<br />Sepal.Length: 6.561448","density: 4.496287e-10<br />Sepal.Length: 6.554403","density: 6.441484e-10<br />Sepal.Length: 6.547358","density: 9.124654e-10<br />Sepal.Length: 6.540313","density: 1.276758e-09<br />Sepal.Length: 6.533268","density: 1.770515e-09<br />Sepal.Length: 6.526223","density: 2.512864e-09<br />Sepal.Length: 6.519178","density: 3.530365e-09<br />Sepal.Length: 6.512133","density: 4.906122e-09<br />Sepal.Length: 6.505088","density: 6.738060e-09<br />Sepal.Length: 6.498043","density: 9.288899e-09<br />Sepal.Length: 6.490998","density: 1.292905e-08<br />Sepal.Length: 6.483953","density: 1.781997e-08<br />Sepal.Length: 6.476908","density: 2.430585e-08<br />Sepal.Length: 6.469863","density: 3.278222e-08<br />Sepal.Length: 6.462818","density: 4.486564e-08<br />Sepal.Length: 6.455773","density: 6.126463e-08<br />Sepal.Length: 6.448728","density: 8.287468e-08<br />Sepal.Length: 6.441683","density: 1.109983e-07<br />Sepal.Length: 6.434638","density: 1.476500e-07<br />Sepal.Length: 6.427593","density: 1.995814e-07<br />Sepal.Length: 6.420548","density: 2.674736e-07<br />Sepal.Length: 6.413503","density: 3.552673e-07<br />Sepal.Length: 6.406458","density: 4.674687e-07<br />Sepal.Length: 6.399413","density: 6.166318e-07<br />Sepal.Length: 6.392368","density: 8.180455e-07<br />Sepal.Length: 6.385323","density: 1.076424e-06<br />Sepal.Length: 6.378278","density: 1.404487e-06<br />Sepal.Length: 6.371233","density: 1.816492e-06<br />Sepal.Length: 6.364188","density: 2.373097e-06<br />Sepal.Length: 6.357143","density: 3.091047e-06<br />Sepal.Length: 6.350098","density: 3.995256e-06<br />Sepal.Length: 6.343053","density: 5.123201e-06<br />Sepal.Length: 6.336008","density: 6.534898e-06<br />Sepal.Length: 6.328963","density: 8.420588e-06<br />Sepal.Length: 6.321918","density: 1.077341e-05<br />Sepal.Length: 6.314873","density: 1.368420e-05<br />Sepal.Length: 6.307828","density: 1.725373e-05<br />Sepal.Length: 6.300783","density: 2.177987e-05<br />Sepal.Length: 6.293738","density: 2.756646e-05<br />Sepal.Length: 6.286693","density: 3.465789e-05<br />Sepal.Length: 6.279648","density: 4.328059e-05<br />Sepal.Length: 6.272603","density: 5.368182e-05<br />Sepal.Length: 6.265558","density: 6.698967e-05<br />Sepal.Length: 6.258513","density: 8.331889e-05<br />Sepal.Length: 6.251468","density: 1.029840e-04<br />Sepal.Length: 6.244423","density: 1.264983e-04<br />Sepal.Length: 6.237378","density: 1.547307e-04<br />Sepal.Length: 6.230333","density: 1.903021e-04<br />Sepal.Length: 6.223288","density: 2.326933e-04<br />Sepal.Length: 6.216243","density: 2.828871e-04<br />Sepal.Length: 6.209198","density: 3.419407e-04<br />Sepal.Length: 6.202153","density: 4.133016e-04<br />Sepal.Length: 6.195108","density: 4.997546e-04<br />Sepal.Length: 6.188063","density: 6.010375e-04<br />Sepal.Length: 6.181018","density: 7.190074e-04<br />Sepal.Length: 6.173973","density: 8.556358e-04<br />Sepal.Length: 6.166928","density: 1.021116e-03<br />Sepal.Length: 6.159883","density: 1.214481e-03<br />Sepal.Length: 6.152838","density: 1.437284e-03<br />Sepal.Length: 6.145793","density: 1.692682e-03<br />Sepal.Length: 6.138748","density: 1.986574e-03<br />Sepal.Length: 6.131703","density: 2.336089e-03<br />Sepal.Length: 6.124658","density: 2.734248e-03<br />Sepal.Length: 6.117613","density: 3.185662e-03<br />Sepal.Length: 6.110568","density: 3.695119e-03<br />Sepal.Length: 6.103523","density: 4.281403e-03<br />Sepal.Length: 6.096477","density: 4.955058e-03<br />Sepal.Length: 6.089432","density: 5.710030e-03<br />Sepal.Length: 6.082387","density: 6.552551e-03<br />Sepal.Length: 6.075342","density: 7.489006e-03<br />Sepal.Length: 6.068297","density: 8.562147e-03<br />Sepal.Length: 6.061252","density: 9.757460e-03<br />Sepal.Length: 6.054207","density: 1.107576e-02<br />Sepal.Length: 6.047162","density: 1.252423e-02<br />Sepal.Length: 6.040117","density: 1.411993e-02<br />Sepal.Length: 6.033072","density: 1.591176e-02<br />Sepal.Length: 6.026027","density: 1.786385e-02<br />Sepal.Length: 6.018982","density: 1.998296e-02<br />Sepal.Length: 6.011937","density: 2.227567e-02<br />Sepal.Length: 6.004892","density: 2.478507e-02<br />Sepal.Length: 5.997847","density: 2.752017e-02<br />Sepal.Length: 5.990802","density: 3.045213e-02<br />Sepal.Length: 5.983757","density: 3.358498e-02<br />Sepal.Length: 5.976712","density: 3.692233e-02<br />Sepal.Length: 5.969667","density: 4.053438e-02<br />Sepal.Length: 5.962622","density: 4.436842e-02<br />Sepal.Length: 5.955577","density: 4.841207e-02<br />Sepal.Length: 5.948532","density: 5.266404e-02<br />Sepal.Length: 5.941487","density: 5.713638e-02<br />Sepal.Length: 5.934442","density: 6.186938e-02<br />Sepal.Length: 5.927397","density: 6.679332e-02<br />Sepal.Length: 5.920352","density: 7.190064e-02<br />Sepal.Length: 5.913307","density: 7.718300e-02<br />Sepal.Length: 5.906262","density: 8.265952e-02<br />Sepal.Length: 5.899217","density: 8.830547e-02<br />Sepal.Length: 5.892172","density: 9.407792e-02<br />Sepal.Length: 5.885127","density: 9.996240e-02<br />Sepal.Length: 5.878082","density: 1.059438e-01<br />Sepal.Length: 5.871037","density: 1.120208e-01<br />Sepal.Length: 5.863992","density: 1.181442e-01<br />Sepal.Length: 5.856947","density: 1.242922e-01<br />Sepal.Length: 5.849902","density: 1.304463e-01<br />Sepal.Length: 5.842857","density: 1.365848e-01<br />Sepal.Length: 5.835812","density: 1.426659e-01<br />Sepal.Length: 5.828767","density: 1.486763e-01<br />Sepal.Length: 5.821722","density: 1.545990e-01<br />Sepal.Length: 5.814677","density: 1.604177e-01<br />Sepal.Length: 5.807632","density: 1.660872e-01<br />Sepal.Length: 5.800587","density: 1.715857e-01<br />Sepal.Length: 5.793542","density: 1.769252e-01<br />Sepal.Length: 5.786497","density: 1.820968e-01<br />Sepal.Length: 5.779452","density: 1.870933e-01<br />Sepal.Length: 5.772407","density: 1.918485e-01<br />Sepal.Length: 5.765362","density: 1.964122e-01<br />Sepal.Length: 5.758317","density: 2.007969e-01<br />Sepal.Length: 5.751272","density: 2.050079e-01<br />Sepal.Length: 5.744227","density: 2.090406e-01<br />Sepal.Length: 5.737182","density: 2.128830e-01<br />Sepal.Length: 5.730137","density: 2.166018e-01<br />Sepal.Length: 5.723092","density: 2.202159e-01<br />Sepal.Length: 5.716047","density: 2.237457e-01<br />Sepal.Length: 5.709002","density: 2.272106e-01<br />Sepal.Length: 5.701957","density: 2.306609e-01<br />Sepal.Length: 5.694912","density: 2.341349e-01<br />Sepal.Length: 5.687867","density: 2.376619e-01<br />Sepal.Length: 5.680822","density: 2.412719e-01<br />Sepal.Length: 5.673777","density: 2.450557e-01<br />Sepal.Length: 5.666732","density: 2.490273e-01<br />Sepal.Length: 5.659687","density: 2.532111e-01<br />Sepal.Length: 5.652642","density: 2.576363e-01<br />Sepal.Length: 5.645597","density: 2.623577e-01<br />Sepal.Length: 5.638552","density: 2.674856e-01<br />Sepal.Length: 5.631507","density: 2.729602e-01<br />Sepal.Length: 5.624462","density: 2.787997e-01<br />Sepal.Length: 5.617417","density: 2.850204e-01<br />Sepal.Length: 5.610372","density: 2.917223e-01<br />Sepal.Length: 5.603327","density: 2.989004e-01<br />Sepal.Length: 5.596282","density: 3.064878e-01<br />Sepal.Length: 5.589237","density: 3.144807e-01<br />Sepal.Length: 5.582192","density: 3.228729e-01<br />Sepal.Length: 5.575147","density: 3.317732e-01<br />Sepal.Length: 5.568102","density: 3.410373e-01<br />Sepal.Length: 5.561057","density: 3.506251e-01<br />Sepal.Length: 5.554012","density: 3.605091e-01<br />Sepal.Length: 5.546967","density: 3.706777e-01<br />Sepal.Length: 5.539922","density: 3.811106e-01<br />Sepal.Length: 5.532877","density: 3.916916e-01<br />Sepal.Length: 5.525832","density: 4.023791e-01<br />Sepal.Length: 5.518787","density: 4.131308e-01<br />Sepal.Length: 5.511742","density: 4.238891e-01<br />Sepal.Length: 5.504697","density: 4.345722e-01<br />Sepal.Length: 5.497652","density: 4.451364e-01<br />Sepal.Length: 5.490607","density: 4.555404e-01<br />Sepal.Length: 5.483562","density: 4.657439e-01<br />Sepal.Length: 5.476517","density: 4.756024e-01<br />Sepal.Length: 5.469472","density: 4.851423e-01<br />Sepal.Length: 5.462427","density: 4.943473e-01<br />Sepal.Length: 5.455382","density: 5.031954e-01<br />Sepal.Length: 5.448337","density: 5.116313e-01<br />Sepal.Length: 5.441292","density: 5.195622e-01<br />Sepal.Length: 5.434247","density: 5.270983e-01<br />Sepal.Length: 5.427202","density: 5.342463e-01<br />Sepal.Length: 5.420157","density: 5.410170e-01<br />Sepal.Length: 5.413112","density: 5.473568e-01<br />Sepal.Length: 5.406067","density: 5.533372e-01<br />Sepal.Length: 5.399022","density: 5.590517e-01<br />Sepal.Length: 5.391977","density: 5.645414e-01<br />Sepal.Length: 5.384932","density: 5.698504e-01<br />Sepal.Length: 5.377886","density: 5.750231e-01<br />Sepal.Length: 5.370841","density: 5.801857e-01<br />Sepal.Length: 5.363796","density: 5.854031e-01<br />Sepal.Length: 5.356751","density: 5.907386e-01<br />Sepal.Length: 5.349706","density: 5.962855e-01<br />Sepal.Length: 5.342661","density: 6.022149e-01<br />Sepal.Length: 5.335616","density: 6.085385e-01<br />Sepal.Length: 5.328571","density: 6.153204e-01<br />Sepal.Length: 5.321526","density: 6.226233e-01<br />Sepal.Length: 5.314481","density: 6.306625e-01<br />Sepal.Length: 5.307436","density: 6.394955e-01<br />Sepal.Length: 5.300391","density: 6.490752e-01<br />Sepal.Length: 5.293346","density: 6.594430e-01<br />Sepal.Length: 5.286301","density: 6.706364e-01<br />Sepal.Length: 5.279256","density: 6.830032e-01<br />Sepal.Length: 5.272211","density: 6.962900e-01<br />Sepal.Length: 5.265166","density: 7.104790e-01<br />Sepal.Length: 5.258121","density: 7.255703e-01<br />Sepal.Length: 5.251076","density: 7.416456e-01<br />Sepal.Length: 5.244031","density: 7.588212e-01<br />Sepal.Length: 5.236986","density: 7.768141e-01<br />Sepal.Length: 5.229941","density: 7.955825e-01<br />Sepal.Length: 5.222896","density: 8.150799e-01<br />Sepal.Length: 5.215851","density: 8.353800e-01<br />Sepal.Length: 5.208806","density: 8.563104e-01<br />Sepal.Length: 5.201761","density: 8.776975e-01<br />Sepal.Length: 5.194716","density: 8.994614e-01<br />Sepal.Length: 5.187671","density: 9.215192e-01<br />Sepal.Length: 5.180626","density: 9.437860e-01<br />Sepal.Length: 5.173581","density: 9.660571e-01<br />Sepal.Length: 5.166536","density: 9.882323e-01<br />Sepal.Length: 5.159491","density: 1.010214e+00<br />Sepal.Length: 5.152446","density: 1.031856e+00<br />Sepal.Length: 5.145401","density: 1.052909e+00<br />Sepal.Length: 5.138356","density: 1.073363e+00<br />Sepal.Length: 5.131311","density: 1.093125e+00<br />Sepal.Length: 5.124266","density: 1.112107e+00<br />Sepal.Length: 5.117221","density: 1.129983e+00<br />Sepal.Length: 5.110176","density: 1.146690e+00<br />Sepal.Length: 5.103131","density: 1.162290e+00<br />Sepal.Length: 5.096086","density: 1.176718e+00<br />Sepal.Length: 5.089041","density: 1.189916e+00<br />Sepal.Length: 5.081996","density: 1.201335e+00<br />Sepal.Length: 5.074951","density: 1.211361e+00<br />Sepal.Length: 5.067906","density: 1.219987e+00<br />Sepal.Length: 5.060861","density: 1.227193e+00<br />Sepal.Length: 5.053816","density: 1.232802e+00<br />Sepal.Length: 5.046771","density: 1.236581e+00<br />Sepal.Length: 5.039726","density: 1.238924e+00<br />Sepal.Length: 5.032681","density: 1.239847e+00<br />Sepal.Length: 5.025636","density: 1.239368e+00<br />Sepal.Length: 5.018591","density: 1.237205e+00<br />Sepal.Length: 5.011546","density: 1.233555e+00<br />Sepal.Length: 5.004501","density: 1.228652e+00<br />Sepal.Length: 4.997456","density: 1.222546e+00<br />Sepal.Length: 4.990411","density: 1.215287e+00<br />Sepal.Length: 4.983366","density: 1.206577e+00<br />Sepal.Length: 4.976321","density: 1.196885e+00<br />Sepal.Length: 4.969276","density: 1.186287e+00<br />Sepal.Length: 4.962231","density: 1.174848e+00<br />Sepal.Length: 4.955186","density: 1.162554e+00<br />Sepal.Length: 4.948141","density: 1.149423e+00<br />Sepal.Length: 4.941096","density: 1.135719e+00<br />Sepal.Length: 4.934051","density: 1.121502e+00<br />Sepal.Length: 4.927006","density: 1.106835e+00<br />Sepal.Length: 4.919961","density: 1.091710e+00<br />Sepal.Length: 4.912916","density: 1.076266e+00<br />Sepal.Length: 4.905871","density: 1.060605e+00<br />Sepal.Length: 4.898826","density: 1.044775e+00<br />Sepal.Length: 4.891781","density: 1.028823e+00<br />Sepal.Length: 4.884736","density: 1.012798e+00<br />Sepal.Length: 4.877691","density: 9.967794e-01<br />Sepal.Length: 4.870646","density: 9.808023e-01<br />Sepal.Length: 4.863601","density: 9.648980e-01<br />Sepal.Length: 4.856556","density: 9.491139e-01<br />Sepal.Length: 4.849511","density: 9.335151e-01<br />Sepal.Length: 4.842466","density: 9.180934e-01<br />Sepal.Length: 4.835421","density: 9.028671e-01<br />Sepal.Length: 4.828376","density: 8.878530e-01<br />Sepal.Length: 4.821331","density: 8.731280e-01<br />Sepal.Length: 4.814286","density: 8.586865e-01<br />Sepal.Length: 4.807241","density: 8.445066e-01<br />Sepal.Length: 4.800196","density: 8.305961e-01<br />Sepal.Length: 4.793151","density: 8.169619e-01<br />Sepal.Length: 4.786106","density: 8.037172e-01<br />Sepal.Length: 4.779061","density: 7.907627e-01<br />Sepal.Length: 4.772016","density: 7.780988e-01<br />Sepal.Length: 4.764971","density: 7.657265e-01<br />Sepal.Length: 4.757926","density: 7.536817e-01<br />Sepal.Length: 4.750881","density: 7.419997e-01<br />Sepal.Length: 4.743836","density: 7.306036e-01<br />Sepal.Length: 4.736791","density: 7.194897e-01<br />Sepal.Length: 4.729746","density: 7.086540e-01<br />Sepal.Length: 4.722701","density: 6.981567e-01<br />Sepal.Length: 4.715656","density: 6.879533e-01<br />Sepal.Length: 4.708611","density: 6.780021e-01<br />Sepal.Length: 4.701566","density: 6.682953e-01<br />Sepal.Length: 4.694521","density: 6.588249e-01<br />Sepal.Length: 4.687476","density: 6.496596e-01<br />Sepal.Length: 4.680431","density: 6.407011e-01<br />Sepal.Length: 4.673386","density: 6.319396e-01<br />Sepal.Length: 4.666341","density: 6.233652e-01<br />Sepal.Length: 4.659295","density: 6.149884e-01<br />Sepal.Length: 4.652250","density: 6.068108e-01<br />Sepal.Length: 4.645205","density: 5.987799e-01<br />Sepal.Length: 4.638160","density: 5.908867e-01<br />Sepal.Length: 4.631115","density: 5.831225e-01<br />Sepal.Length: 4.624070","density: 5.755063e-01<br />Sepal.Length: 4.617025","density: 5.680097e-01<br />Sepal.Length: 4.609980","density: 5.606125e-01<br />Sepal.Length: 4.602935","density: 5.533092e-01<br />Sepal.Length: 4.595890","density: 5.460952e-01<br />Sepal.Length: 4.588845","density: 5.389938e-01<br />Sepal.Length: 4.581800","density: 5.319692e-01<br />Sepal.Length: 4.574755","density: 5.250186e-01<br />Sepal.Length: 4.567710","density: 5.181395e-01<br />Sepal.Length: 4.560665","density: 5.113378e-01<br />Sepal.Length: 4.553620","density: 5.046166e-01<br />Sepal.Length: 4.546575","density: 4.979564e-01<br />Sepal.Length: 4.539530","density: 4.913543e-01<br />Sepal.Length: 4.532485","density: 4.848067e-01<br />Sepal.Length: 4.525440","density: 4.783209e-01<br />Sepal.Length: 4.518395","density: 4.718805e-01<br />Sepal.Length: 4.511350","density: 4.654733e-01<br />Sepal.Length: 4.504305","density: 4.590921e-01<br />Sepal.Length: 4.497260","density: 4.527290e-01<br />Sepal.Length: 4.490215","density: 4.463702e-01<br />Sepal.Length: 4.483170","density: 4.399975e-01<br />Sepal.Length: 4.476125","density: 4.335992e-01<br />Sepal.Length: 4.469080","density: 4.271629e-01<br />Sepal.Length: 4.462035","density: 4.206656e-01<br />Sepal.Length: 4.454990","density: 4.140746e-01<br />Sepal.Length: 4.447945","density: 4.073889e-01<br />Sepal.Length: 4.440900","density: 4.005945e-01<br />Sepal.Length: 4.433855","density: 3.936779e-01<br />Sepal.Length: 4.426810","density: 3.865834e-01<br />Sepal.Length: 4.419765","density: 3.793105e-01<br />Sepal.Length: 4.412720","density: 3.718650e-01<br />Sepal.Length: 4.405675","density: 3.642377e-01<br />Sepal.Length: 4.398630","density: 3.564173e-01<br />Sepal.Length: 4.391585","density: 3.483245e-01<br />Sepal.Length: 4.384540","density: 3.400265e-01<br />Sepal.Length: 4.377495","density: 3.315231e-01<br />Sepal.Length: 4.370450","density: 3.228151e-01<br />Sepal.Length: 4.363405","density: 3.138780e-01<br />Sepal.Length: 4.356360","density: 3.047049e-01<br />Sepal.Length: 4.349315","density: 2.953535e-01<br />Sepal.Length: 4.342270","density: 2.858354e-01<br />Sepal.Length: 4.335225","density: 2.761633e-01<br />Sepal.Length: 4.328180","density: 2.663234e-01<br />Sepal.Length: 4.321135","density: 2.563679e-01<br />Sepal.Length: 4.314090","density: 2.463309e-01<br />Sepal.Length: 4.307045","density: 2.362330e-01<br />Sepal.Length: 4.300000","density: 2.362330e-01<br />Sepal.Length: 4.300000"],"key":["1","2","3","4","5","6","7","8","9","10","11","12","13","14","15","16","17","18","19","20","21","22","23","24","25","26","27","28","29","30","31","32","33","34","35","36","37","38","39","40","41","42","43","44","45","46","47","48","49","50"],"type":"scatter","mode":"lines","line":{"width":1.88976377952756,"color":"rgba(0,0,0,1)","dash":"solid"},"fill":"toself","fillcolor":"rgba(248,118,109,1)","hoveron":"points","set":"SharedData91833e3a","name":"setosa","legendgroup":"setosa","showlegend":true,"xaxis":"x","yaxis":"y4","hoverinfo":"text","_isSimpleKey":true,"_isNestedKey":false,"frame":null},{"x":[4.3,4.30704500978474,4.31409001956947,4.32113502935421,4.32818003913894,4.33522504892368,4.34227005870842,4.34931506849315,4.35636007827789,4.36340508806262,4.37045009784736,4.37749510763209,4.38454011741683,4.39158512720157,4.3986301369863,4.40567514677104,4.41272015655577,4.41976516634051,4.42681017612524,4.43385518590998,4.44090019569472,4.44794520547945,4.45499021526419,4.46203522504892,4.46908023483366,4.4761252446184,4.48317025440313,4.49021526418787,4.4972602739726,4.50430528375734,4.51135029354207,4.51839530332681,4.52544031311155,4.53248532289628,4.53953033268102,4.54657534246575,4.55362035225049,4.56066536203523,4.56771037181996,4.5747553816047,4.58180039138943,4.58884540117417,4.5958904109589,4.60293542074364,4.60998043052838,4.61702544031311,4.62407045009785,4.63111545988258,4.63816046966732,4.64520547945205,4.65225048923679,4.65929549902153,4.66634050880626,4.673385518591,4.68043052837573,4.68747553816047,4.69452054794521,4.70156555772994,4.70861056751468,4.71565557729941,4.72270058708415,4.72974559686888,4.73679060665362,4.74383561643836,4.75088062622309,4.75792563600783,4.76497064579256,4.7720156555773,4.77906066536204,4.78610567514677,4.79315068493151,4.80019569471624,4.80724070450098,4.81428571428571,4.82133072407045,4.82837573385519,4.83542074363992,4.84246575342466,4.84951076320939,4.85655577299413,4.86360078277886,4.8706457925636,4.87769080234834,4.88473581213307,4.89178082191781,4.89882583170254,4.90587084148728,4.91291585127202,4.91996086105675,4.92700587084149,4.93405088062622,4.94109589041096,4.94814090019569,4.95518590998043,4.96223091976517,4.9692759295499,4.97632093933464,4.98336594911937,4.99041095890411,4.99745596868885,5.00450097847358,5.01154598825832,5.01859099804305,5.02563600782779,5.03268101761252,5.03972602739726,5.046771037182,5.05381604696673,5.06086105675147,5.0679060665362,5.07495107632094,5.08199608610567,5.08904109589041,5.09608610567515,5.10313111545988,5.11017612524462,5.11722113502935,5.12426614481409,5.13131115459883,5.13835616438356,5.1454011741683,5.15244618395303,5.15949119373777,5.1665362035225,5.17358121330724,5.18062622309198,5.18767123287671,5.19471624266145,5.20176125244618,5.20880626223092,5.21585127201566,5.22289628180039,5.22994129158513,5.23698630136986,5.2440313111546,5.25107632093933,5.25812133072407,5.26516634050881,5.27221135029354,5.27925636007828,5.28630136986301,5.29334637964775,5.30039138943249,5.30743639921722,5.31448140900196,5.32152641878669,5.32857142857143,5.33561643835616,5.3426614481409,5.34970645792564,5.35675146771037,5.36379647749511,5.37084148727984,5.37788649706458,5.38493150684932,5.39197651663405,5.39902152641879,5.40606653620352,5.41311154598826,5.42015655577299,5.42720156555773,5.43424657534247,5.4412915851272,5.44833659491194,5.45538160469667,5.46242661448141,5.46947162426614,5.47651663405088,5.48356164383562,5.49060665362035,5.49765166340509,5.50469667318982,5.51174168297456,5.5187866927593,5.52583170254403,5.53287671232877,5.5399217221135,5.54696673189824,5.55401174168297,5.56105675146771,5.56810176125245,5.57514677103718,5.58219178082192,5.58923679060665,5.59628180039139,5.60332681017613,5.61037181996086,5.6174168297456,5.62446183953033,5.63150684931507,5.6385518590998,5.64559686888454,5.65264187866928,5.65968688845401,5.66673189823875,5.67377690802348,5.68082191780822,5.68786692759295,5.69491193737769,5.70195694716243,5.70900195694716,5.7160469667319,5.72309197651663,5.73013698630137,5.73718199608611,5.74422700587084,5.75127201565558,5.75831702544031,5.76536203522505,5.77240704500978,5.77945205479452,5.78649706457926,5.79354207436399,5.80058708414873,5.80763209393346,5.8146771037182,5.82172211350294,5.82876712328767,5.83581213307241,5.84285714285714,5.84990215264188,5.85694716242661,5.86399217221135,5.87103718199609,5.87808219178082,5.88512720156556,5.89217221135029,5.89921722113503,5.90626223091976,5.9133072407045,5.92035225048924,5.92739726027397,5.93444227005871,5.94148727984344,5.94853228962818,5.95557729941292,5.96262230919765,5.96966731898239,5.97671232876712,5.98375733855186,5.9908023483366,5.99784735812133,6.00489236790607,6.0119373776908,6.01898238747554,6.02602739726027,6.03307240704501,6.04011741682975,6.04716242661448,6.05420743639922,6.06125244618395,6.06829745596869,6.07534246575342,6.08238747553816,6.0894324853229,6.09647749510763,6.10352250489237,6.1105675146771,6.11761252446184,6.12465753424657,6.13170254403131,6.13874755381605,6.14579256360078,6.15283757338552,6.15988258317025,6.16692759295499,6.17397260273973,6.18101761252446,6.1880626223092,6.19510763209393,6.20215264187867,6.20919765166341,6.21624266144814,6.22328767123288,6.23033268101761,6.23737769080235,6.24442270058708,6.25146771037182,6.25851272015656,6.26555772994129,6.27260273972603,6.27964774951076,6.2866927592955,6.29373776908024,6.30078277886497,6.30782778864971,6.31487279843444,6.32191780821918,6.32896281800391,6.33600782778865,6.34305283757339,6.35009784735812,6.35714285714286,6.36418786692759,6.37123287671233,6.37827788649706,6.3853228962818,6.39236790606654,6.39941291585127,6.40645792563601,6.41350293542074,6.42054794520548,6.42759295499022,6.43463796477495,6.44168297455969,6.44872798434442,6.45577299412916,6.46281800391389,6.46986301369863,6.47690802348337,6.4839530332681,6.49099804305284,6.49804305283757,6.50508806262231,6.51213307240705,6.51917808219178,6.52622309197652,6.53326810176125,6.54031311154599,6.54735812133072,6.55440313111546,6.5614481409002,6.56849315068493,6.57553816046967,6.5825831702544,6.58962818003914,6.59667318982387,6.60371819960861,6.61076320939335,6.61780821917808,6.62485322896282,6.63189823874755,6.63894324853229,6.64598825831703,6.65303326810176,6.6600782778865,6.66712328767123,6.67416829745597,6.6812133072407,6.68825831702544,6.69530332681018,6.70234833659491,6.70939334637965,6.71643835616438,6.72348336594912,6.73052837573386,6.73757338551859,6.74461839530333,6.75166340508806,6.7587084148728,6.76575342465753,6.77279843444227,6.77984344422701,6.78688845401174,6.79393346379648,6.80097847358121,6.80802348336595,6.81506849315068,6.82211350293542,6.82915851272016,6.83620352250489,6.84324853228963,6.85029354207436,6.8573385518591,6.86438356164384,6.87142857142857,6.87847358121331,6.88551859099804,6.89256360078278,6.89960861056752,6.90665362035225,6.91369863013699,6.92074363992172,6.92778864970646,6.93483365949119,6.94187866927593,6.94892367906067,6.9559686888454,6.96301369863014,6.97005870841487,6.97710371819961,6.98414872798434,6.99119373776908,6.99823874755382,7.00528375733855,7.01232876712329,7.01937377690802,7.02641878669276,7.03346379647749,7.04050880626223,7.04755381604697,7.0545988258317,7.06164383561644,7.06868884540117,7.07573385518591,7.08277886497065,7.08982387475538,7.09686888454012,7.10391389432485,7.11095890410959,7.11800391389433,7.12504892367906,7.1320939334638,7.13913894324853,7.14618395303327,7.153228962818,7.16027397260274,7.16731898238748,7.17436399217221,7.18140900195695,7.18845401174168,7.19549902152642,7.20254403131116,7.20958904109589,7.21663405088063,7.22367906066536,7.2307240704501,7.23776908023483,7.24481409001957,7.25185909980431,7.25890410958904,7.26594911937378,7.27299412915851,7.28003913894325,7.28708414872798,7.29412915851272,7.30117416829746,7.30821917808219,7.31526418786693,7.32230919765166,7.3293542074364,7.33639921722114,7.34344422700587,7.35048923679061,7.35753424657534,7.36457925636008,7.37162426614481,7.37866927592955,7.38571428571429,7.39275929549902,7.39980430528376,7.40684931506849,7.41389432485323,7.42093933463797,7.4279843444227,7.43502935420744,7.44207436399217,7.44911937377691,7.45616438356164,7.46320939334638,7.47025440313112,7.47729941291585,7.48434442270059,7.49138943248532,7.49843444227006,7.50547945205479,7.51252446183953,7.51956947162427,7.526614481409,7.53365949119374,7.54070450097847,7.54774951076321,7.55479452054795,7.56183953033268,7.56888454011742,7.57592954990215,7.58297455968689,7.59001956947163,7.59706457925636,7.6041095890411,7.61115459882583,7.61819960861057,7.6252446183953,7.63228962818004,7.63933463796478,7.64637964774951,7.65342465753425,7.66046966731898,7.66751467710372,7.67455968688845,7.68160469667319,7.68864970645793,7.69569471624266,7.7027397260274,7.70978473581213,7.71682974559687,7.7238747553816,7.73091976516634,7.73796477495108,7.74500978473581,7.75205479452055,7.75909980430528,7.76614481409002,7.77318982387476,7.78023483365949,7.78727984344423,7.79432485322896,7.8013698630137,7.80841487279844,7.81545988258317,7.82250489236791,7.82954990215264,7.83659491193738,7.84363992172211,7.85068493150685,7.85772994129159,7.86477495107632,7.87181996086106,7.87886497064579,7.88590998043053,7.89295499021526,7.9,7.9,7.89295499021526,7.88590998043053,7.87886497064579,7.87181996086106,7.86477495107632,7.85772994129159,7.85068493150685,7.84363992172211,7.83659491193738,7.82954990215264,7.82250489236791,7.81545988258317,7.80841487279844,7.8013698630137,7.79432485322896,7.78727984344423,7.78023483365949,7.77318982387476,7.76614481409002,7.75909980430528,7.75205479452055,7.74500978473581,7.73796477495108,7.73091976516634,7.7238747553816,7.71682974559687,7.70978473581213,7.7027397260274,7.69569471624266,7.68864970645793,7.68160469667319,7.67455968688845,7.66751467710372,7.66046966731898,7.65342465753425,7.64637964774951,7.63933463796478,7.63228962818004,7.6252446183953,7.61819960861057,7.61115459882583,7.6041095890411,7.59706457925636,7.59001956947163,7.58297455968689,7.57592954990215,7.56888454011742,7.56183953033268,7.55479452054795,7.54774951076321,7.54070450097847,7.53365949119374,7.526614481409,7.51956947162427,7.51252446183953,7.50547945205479,7.49843444227006,7.49138943248532,7.48434442270059,7.47729941291585,7.47025440313112,7.46320939334638,7.45616438356164,7.44911937377691,7.44207436399217,7.43502935420744,7.4279843444227,7.42093933463797,7.41389432485323,7.40684931506849,7.39980430528376,7.39275929549902,7.38571428571429,7.37866927592955,7.37162426614481,7.36457925636008,7.35753424657534,7.35048923679061,7.34344422700587,7.33639921722114,7.3293542074364,7.32230919765166,7.31526418786693,7.30821917808219,7.30117416829746,7.29412915851272,7.28708414872798,7.28003913894325,7.27299412915851,7.26594911937378,7.25890410958904,7.25185909980431,7.24481409001957,7.23776908023483,7.2307240704501,7.22367906066536,7.21663405088063,7.20958904109589,7.20254403131116,7.19549902152642,7.18845401174168,7.18140900195695,7.17436399217221,7.16731898238748,7.16027397260274,7.153228962818,7.14618395303327,7.13913894324853,7.1320939334638,7.12504892367906,7.11800391389433,7.11095890410959,7.10391389432485,7.09686888454012,7.08982387475538,7.08277886497065,7.07573385518591,7.06868884540117,7.06164383561644,7.0545988258317,7.04755381604697,7.04050880626223,7.03346379647749,7.02641878669276,7.01937377690802,7.01232876712329,7.00528375733855,6.99823874755382,6.99119373776908,6.98414872798434,6.97710371819961,6.97005870841487,6.96301369863014,6.9559686888454,6.94892367906067,6.94187866927593,6.93483365949119,6.92778864970646,6.92074363992172,6.91369863013699,6.90665362035225,6.89960861056752,6.89256360078278,6.88551859099804,6.87847358121331,6.87142857142857,6.86438356164384,6.8573385518591,6.85029354207436,6.84324853228963,6.83620352250489,6.82915851272016,6.82211350293542,6.81506849315068,6.80802348336595,6.80097847358121,6.79393346379648,6.78688845401174,6.77984344422701,6.77279843444227,6.76575342465753,6.7587084148728,6.75166340508806,6.74461839530333,6.73757338551859,6.73052837573386,6.72348336594912,6.71643835616438,6.70939334637965,6.70234833659491,6.69530332681018,6.68825831702544,6.6812133072407,6.67416829745597,6.66712328767123,6.6600782778865,6.65303326810176,6.64598825831703,6.63894324853229,6.63189823874755,6.62485322896282,6.61780821917808,6.61076320939335,6.60371819960861,6.59667318982387,6.58962818003914,6.5825831702544,6.57553816046967,6.56849315068493,6.5614481409002,6.55440313111546,6.54735812133072,6.54031311154599,6.53326810176125,6.52622309197652,6.51917808219178,6.51213307240705,6.50508806262231,6.49804305283757,6.49099804305284,6.4839530332681,6.47690802348337,6.46986301369863,6.46281800391389,6.45577299412916,6.44872798434442,6.44168297455969,6.43463796477495,6.42759295499022,6.42054794520548,6.41350293542074,6.40645792563601,6.39941291585127,6.39236790606654,6.3853228962818,6.37827788649706,6.37123287671233,6.36418786692759,6.35714285714286,6.35009784735812,6.34305283757339,6.33600782778865,6.32896281800391,6.32191780821918,6.31487279843444,6.30782778864971,6.30078277886497,6.29373776908024,6.2866927592955,6.27964774951076,6.27260273972603,6.26555772994129,6.25851272015656,6.25146771037182,6.24442270058708,6.23737769080235,6.23033268101761,6.22328767123288,6.21624266144814,6.20919765166341,6.20215264187867,6.19510763209393,6.1880626223092,6.18101761252446,6.17397260273973,6.16692759295499,6.15988258317025,6.15283757338552,6.14579256360078,6.13874755381605,6.13170254403131,6.12465753424657,6.11761252446184,6.1105675146771,6.10352250489237,6.09647749510763,6.0894324853229,6.08238747553816,6.07534246575342,6.06829745596869,6.06125244618395,6.05420743639922,6.04716242661448,6.04011741682975,6.03307240704501,6.02602739726027,6.01898238747554,6.0119373776908,6.00489236790607,5.99784735812133,5.9908023483366,5.98375733855186,5.97671232876712,5.96966731898239,5.96262230919765,5.95557729941292,5.94853228962818,5.94148727984344,5.93444227005871,5.92739726027397,5.92035225048924,5.9133072407045,5.90626223091976,5.89921722113503,5.89217221135029,5.88512720156556,5.87808219178082,5.87103718199609,5.86399217221135,5.85694716242661,5.84990215264188,5.84285714285714,5.83581213307241,5.82876712328767,5.82172211350294,5.8146771037182,5.80763209393346,5.80058708414873,5.79354207436399,5.78649706457926,5.77945205479452,5.77240704500978,5.76536203522505,5.75831702544031,5.75127201565558,5.74422700587084,5.73718199608611,5.73013698630137,5.72309197651663,5.7160469667319,5.70900195694716,5.70195694716243,5.69491193737769,5.68786692759295,5.68082191780822,5.67377690802348,5.66673189823875,5.65968688845401,5.65264187866928,5.64559686888454,5.6385518590998,5.63150684931507,5.62446183953033,5.6174168297456,5.61037181996086,5.60332681017613,5.59628180039139,5.58923679060665,5.58219178082192,5.57514677103718,5.56810176125245,5.56105675146771,5.55401174168297,5.54696673189824,5.5399217221135,5.53287671232877,5.52583170254403,5.5187866927593,5.51174168297456,5.50469667318982,5.49765166340509,5.49060665362035,5.48356164383562,5.47651663405088,5.46947162426614,5.46242661448141,5.45538160469667,5.44833659491194,5.4412915851272,5.43424657534247,5.42720156555773,5.42015655577299,5.41311154598826,5.40606653620352,5.39902152641879,5.39197651663405,5.38493150684932,5.37788649706458,5.37084148727984,5.36379647749511,5.35675146771037,5.34970645792564,5.3426614481409,5.33561643835616,5.32857142857143,5.32152641878669,5.31448140900196,5.30743639921722,5.30039138943249,5.29334637964775,5.28630136986301,5.27925636007828,5.27221135029354,5.26516634050881,5.25812133072407,5.25107632093933,5.2440313111546,5.23698630136986,5.22994129158513,5.22289628180039,5.21585127201566,5.20880626223092,5.20176125244618,5.19471624266145,5.18767123287671,5.18062622309198,5.17358121330724,5.1665362035225,5.15949119373777,5.15244618395303,5.1454011741683,5.13835616438356,5.13131115459883,5.12426614481409,5.11722113502935,5.11017612524462,5.10313111545988,5.09608610567515,5.08904109589041,5.08199608610567,5.07495107632094,5.0679060665362,5.06086105675147,5.05381604696673,5.046771037182,5.03972602739726,5.03268101761252,5.02563600782779,5.01859099804305,5.01154598825832,5.00450097847358,4.99745596868885,4.99041095890411,4.98336594911937,4.97632093933464,4.9692759295499,4.96223091976517,4.95518590998043,4.94814090019569,4.94109589041096,4.93405088062622,4.92700587084149,4.91996086105675,4.91291585127202,4.90587084148728,4.89882583170254,4.89178082191781,4.88473581213307,4.87769080234834,4.8706457925636,4.86360078277886,4.85655577299413,4.84951076320939,4.84246575342466,4.83542074363992,4.82837573385519,4.82133072407045,4.81428571428571,4.80724070450098,4.80019569471624,4.79315068493151,4.78610567514677,4.77906066536204,4.7720156555773,4.76497064579256,4.75792563600783,4.75088062622309,4.74383561643836,4.73679060665362,4.72974559686888,4.72270058708415,4.71565557729941,4.70861056751468,4.70156555772994,4.69452054794521,4.68747553816047,4.68043052837573,4.673385518591,4.66634050880626,4.65929549902153,4.65225048923679,4.64520547945205,4.63816046966732,4.63111545988258,4.62407045009785,4.61702544031311,4.60998043052838,4.60293542074364,4.5958904109589,4.58884540117417,4.58180039138943,4.5747553816047,4.56771037181996,4.56066536203523,4.55362035225049,4.54657534246575,4.53953033268102,4.53248532289628,4.52544031311155,4.51839530332681,4.51135029354207,4.50430528375734,4.4972602739726,4.49021526418787,4.48317025440313,4.4761252446184,4.46908023483366,4.46203522504892,4.45499021526419,4.44794520547945,4.44090019569472,4.43385518590998,4.42681017612524,4.41976516634051,4.41272015655577,4.40567514677104,4.3986301369863,4.39158512720157,4.38454011741683,4.37749510763209,4.37045009784736,4.36340508806262,4.35636007827789,4.34931506849315,4.34227005870842,4.33522504892368,4.32818003913894,4.32113502935421,4.31409001956947,4.30704500978474,4.3,4.3],"y":[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,5.53779886116776e-06,6.40416954535529e-06,7.36822045174752e-06,8.43968848724364e-06,9.73172715334422e-06,1.11666816467295e-05,1.27472275747377e-05,1.46480213056304e-05,1.67621235120386e-05,1.90872393051559e-05,2.1839517251353e-05,2.49225241602315e-05,2.83077657177351e-05,3.22549062855983e-05,3.67052599839225e-05,4.15834354269031e-05,4.71901641944785e-05,5.35492824672334e-05,6.05069325562704e-05,6.83951231354485e-05,7.73899448934126e-05,8.72123932160415e-05,9.82047559672582e-05,0.000110799596955379,0.000124525581828109,0.000139698048665437,0.000157156625695408,0.000176143288797289,0.000196886661454971,0.000220844801989423,0.000246843419850846,0.000274934476827275,0.00030748352130782,0.000342725803244075,0.000380699878831881,0.000424187753667769,0.000471481782030738,0.000522341871662442,0.000579854221435357,0.000642688180852341,0.00071012236725678,0.000785467532903885,0.00086811828833334,0.000956633521343985,0.00105441680234973,0.00116205918931259,0.00127708990917776,0.00140280984119352,0.0015416212997474,0.00168963165689392,0.00184976854096712,0.00202702255783144,0.00221559538307223,0.00241768588657684,0.00264182672862297,0.00287973150691791,0.00313242249082656,0.00341311305223292,0.00371034461349305,0.0040242650752487,0.00437155338739811,0.00473933298121548,0.00512692838788716,0.0055513734259544,0.00600210440422062,0.00647608416815474,0.00699017675654067,0.00753734831993242,0.00811147227451343,0.00872861468878362,0.00938664907437282,0.0100755583221858,0.0108098907912664,0.0115939318327272,0.0124129208943297,0.013279096822157,0.0142047408671995,0.0151694730660432,0.0161823856984628,0.0172653592109901,0.0183915306095002,0.0195659967160335,0.0208217882522168,0.0221247487626701,0.0234751576021162,0.0249186149227192,0.0264129581425322,0.0279583607073463,0.0295978018421357,0.0312969374056474,0.0330504428041505,0.0348974413057741,0.0368131648563155,0.0387860682035864,0.0408505167425434,0.0429925928575294,0.0451940473831211,0.0474837149019207,0.049859487426997,0.0522961247523911,0.0548163286223206,0.0574303710135435,0.0601059646028068,0.0628592840691221,0.0657130997765717,0.0686282807712683,0.0716143186822665,0.0747060987272322,0.0778581304892896,0.081073327914184,0.084397770079423,0.0877803851698542,0.0912210418699153,0.094766083423363,0.0983693848920881,0.10202759378264,0.105778352059324,0.109588780160586,0.113449997250789,0.11739119883935,0.121391564788067,0.125437637485961,0.129550717428658,0.133720307508332,0.137929599341836,0.142192840593152,0.146507596485751,0.150855190228256,0.155243934794275,0.15967671884875,0.164134707540358,0.168621648227386,0.173142604749896,0.177680482169384,0.182236045228546,0.186813069910285,0.191398232530036,0.195991085113056,0.200590389800029,0.205188760379515,0.209785711314461,0.214373230584503,0.218950008793876,0.223516189329755,0.228058945548249,0.232579483189835,0.237080436033276,0.24154622081481,0.245977008605333,0.250379702308628,0.254738022028232,0.25904776227803,0.263321681855565,0.267544756787808,0.271705482866587,0.275823753677708,0.279887529716789,0.283875720459968,0.287816210184484,0.291701332564385,0.295498959437034,0.299245293660371,0.302937985413511,0.306533423937304,0.310075846018891,0.313565223170725,0.316957367263566,0.320293373990353,0.323576770776944,0.326770654848085,0.329905346528965,0.332989928853072,0.335995476689315,0.338941573794116,0.341842074448403,0.344676068646298,0.347453566071196,0.350191835298124,0.352877379889857,0.355512832993723,0.358117023543107,0.360682691589912,0.363208153311303,0.365711566686027,0.368190263689413,0.370641917026345,0.373081562075113,0.3755091558514,0.377925708428478,0.380340644717097,0.382754487341029,0.385175354221263,0.387604908717903,0.390043583642798,0.392505700524403,0.394987655651695,0.397488261177103,0.400025402721424,0.4025942560532,0.40519014948981,0.407832011641758,0.410517402518923,0.413236688248149,0.416007619274932,0.418833006170278,0.42169729571721,0.424615289872961,0.42759694581356,0.430620451754705,0.433696452097492,0.436842825582754,0.440031830594189,0.443269369798349,0.446580837267833,0.44993355812398,0.453328748147021,0.456797762852249,0.460304608322655,0.463849054382238,0.467458095176796,0.471102297759232,0.474778734629989,0.478506203078079,0.482264788081849,0.486048751845701,0.489869423393253,0.493714465283429,0.497576926279719,0.501461926139751,0.505362031102015,0.509270928455117,0.513189170232398,0.51711111520508,0.521033026589443,0.524952744042574,0.528863195527479,0.532765063741167,0.536655366736091,0.54052259723567,0.54437343011004,0.548205812021236,0.552001327959955,0.555773784640384,0.559522877034202,0.563223498889753,0.566895274098544,0.570538646574504,0.574129079879045,0.577683999292961,0.581207256640105,0.584676744562572,0.588105490001081,0.591501225036381,0.594845581773732,0.59814597560582,0.601413920027696,0.604635485217252,0.6078122801149,0.610958949067356,0.614066086406883,0.61713022966549,0.620168037074579,0.62317416677645,0.626141537547253,0.629087390016486,0.632009571977069,0.634899223752033,0.637772635887158,0.64062975072727,0.643461728291299,0.646282539540988,0.649092413425725,0.651885983099469,0.654671792665136,0.657450696077555,0.660219807750385,0.662983276559121,0.665742123949574,0.668494080145198,0.671240273960499,0.673981682959823,0.676715195648651,0.679439158094492,0.682155119725999,0.684858358927769,0.687543105176351,0.690213127701103,0.692862246207371,0.695477355955961,0.698067254802627,0.700625528932862,0.703126490872819,0.70558796660532,0.70800566342336,0.710334083743475,0.712605186842191,0.714818003229768,0.716904965826068,0.718911493805579,0.720838655772169,0.722610173864202,0.724267986486848,0.725823122391271,0.727194482981948,0.72841265927825,0.729503605364454,0.730386250282641,0.731070951288904,0.731603782769687,0.73190913040623,0.731967978436602,0.731851495096787,0.731495082305359,0.730841825217349,0.729992250105785,0.728897977798773,0.727457178366328,0.725802804418329,0.723907055432578,0.721618533894712,0.719104046364174,0.716359443288253,0.713182206525,0.709772419670515,0.706129972808099,0.702066417302663,0.697749191398772,0.693199705858529,0.688258828691818,0.68304697647736,0.677610154172342,0.671821031135682,0.665753078561322,0.659474347079839,0.652889652468609,0.646029383496874,0.638979096692703,0.631673951103434,0.624108736162626,0.616380170850845,0.608449953513877,0.600287931498436,0.591994170019036,0.583551393460218,0.574917643368689,0.566187495941875,0.557357892143141,0.548389788303787,0.539362963751061,0.530279341089343,0.521122964228785,0.511944739336463,0.502748208911136,0.493546708215249,0.484362372237684,0.475197380306746,0.466085375962622,0.457034736042062,0.448038778006455,0.4391424561071,0.430354681435756,0.421653285052706,0.413086094824619,0.404675153075075,0.396377520172204,0.388236524002971,0.380297424875177,0.372492932936148,0.364855965860802,0.357461976421508,0.350217688492403,0.343141436996107,0.336342375378218,0.329701647041993,0.323220705806168,0.317042357736823,0.311024567423593,0.305167223832947,0.299596120779698,0.294197488414144,0.288955071432169,0.283971673677727,0.279167076302806,0.274509097817422,0.270076937579248,0.265822582286045,0.261700738811153,0.257768159066462,0.254004935726867,0.25035661792289,0.246860104012984,0.243517414544982,0.240269632332082,0.237137436863364,0.23413728477969,0.231210790380987,0.228366664462072,0.225627788148511,0.222941182706209,0.220308032815848,0.217749877495578,0.215223501637447,0.212727918283788,0.210273151817083,0.207832585076591,0.20540428580057,0.202985520045085,0.20056454629691,0.198139909869649,0.195701219920203,0.193244153661533,0.190770618011004,0.188266928448255,0.185730237397825,0.18316771910015,0.180565816782168,0.177919017467428,0.17524060461307,0.172519518233316,0.169745360715518,0.16693730617498,0.164088087312378,0.16118208092602,0.158243157320085,0.155268124796154,0.152237487047085,0.149177793110732,0.146089280322887,0.142951458618618,0.139790787957902,0.13660769934799,0.133390380467218,0.130156277225853,0.126907929958321,0.123641299474879,0.120366930916894,0.117087665791459,0.11380567446815,0.110527648192424,0.1072546515364,0.103993077130116,0.100749321264164,0.0975205411965655,0.0943151696456409,0.0911429790198272,0.0879952337377271,0.0848802369263089,0.0818145683190496,0.0787819230931022,0.0757884918852442,0.0728605685741298,0.0699730347389523,0.0671283059756821,0.0643645677317698,0.0616471528212422,0.0589763159459285,0.0563948598131293,0.0538669747418122,0.0513899348379012,0.0490030601248597,0.0466782683780716,0.0444070105904893,0.0422236778536096,0.0401097541510464,0.0380505357460816,0.0360745398499074,0.0341737924643758,0.0323275301826757,0.0305579258138225,0.0288677280436686,0.0272305781237553,0.0256622545533909,0.0241757267693571,0.0227397986770331,0.0213641532032293,0.0200709370808062,0.0188250833254695,0.0176307450707457,0.0165178154649016,0.0154484446600066,0.0144224336881668,0.01347447184235,0.012566339529308,0.0116973920019591,0.0108949133746835,0.0101318541603815,0.00940365840808473,0.00873109176196335,0.00809666611689249,0.00749279854973014,0.00693468702357511,0.00641272583466607,0.00591716512764903,0.00545858788726778,0.00503362688296027,0.00463115719648675,0.00425805351816488,0.00391565748358002,0.00359216263701482,0.00329156216367968,0.00301854512761234,0.00276120142148778,0.00252136865086939,0.00230592081867588,0.00210330029768608,0.00191380430876107,0.00174553945684702,0.00158763788372632,0.00143998699422743,0.00130929844918229,0.00118750329263047,0.00107388696829395,0],"text":["density: 1.073887e-03<br />Sepal.Length: 4.300000","density: 1.187503e-03<br />Sepal.Length: 4.307045","density: 1.309298e-03<br />Sepal.Length: 4.314090","density: 1.439987e-03<br />Sepal.Length: 4.321135","density: 1.587638e-03<br />Sepal.Length: 4.328180","density: 1.745539e-03<br />Sepal.Length: 4.335225","density: 1.913804e-03<br />Sepal.Length: 4.342270","density: 2.103300e-03<br />Sepal.Length: 4.349315","density: 2.305921e-03<br />Sepal.Length: 4.356360","density: 2.521369e-03<br />Sepal.Length: 4.363405","density: 2.761201e-03<br />Sepal.Length: 4.370450","density: 3.018545e-03<br />Sepal.Length: 4.377495","density: 3.291562e-03<br />Sepal.Length: 4.384540","density: 3.592163e-03<br />Sepal.Length: 4.391585","density: 3.915657e-03<br />Sepal.Length: 4.398630","density: 4.258054e-03<br />Sepal.Length: 4.405675","density: 4.631157e-03<br />Sepal.Length: 4.412720","density: 5.033627e-03<br />Sepal.Length: 4.419765","density: 5.458588e-03<br />Sepal.Length: 4.426810","density: 5.917165e-03<br />Sepal.Length: 4.433855","density: 6.412726e-03<br />Sepal.Length: 4.440900","density: 6.934687e-03<br />Sepal.Length: 4.447945","density: 7.492799e-03<br />Sepal.Length: 4.454990","density: 8.096666e-03<br />Sepal.Length: 4.462035","density: 8.731092e-03<br />Sepal.Length: 4.469080","density: 9.403658e-03<br />Sepal.Length: 4.476125","density: 1.013185e-02<br />Sepal.Length: 4.483170","density: 1.089491e-02<br />Sepal.Length: 4.490215","density: 1.169739e-02<br />Sepal.Length: 4.497260","density: 1.256634e-02<br />Sepal.Length: 4.504305","density: 1.347447e-02<br />Sepal.Length: 4.511350","density: 1.442243e-02<br />Sepal.Length: 4.518395","density: 1.544844e-02<br />Sepal.Length: 4.525440","density: 1.651782e-02<br />Sepal.Length: 4.532485","density: 1.763075e-02<br />Sepal.Length: 4.539530","density: 1.882508e-02<br />Sepal.Length: 4.546575","density: 2.007094e-02<br />Sepal.Length: 4.553620","density: 2.136415e-02<br />Sepal.Length: 4.560665","density: 2.273980e-02<br />Sepal.Length: 4.567710","density: 2.417573e-02<br />Sepal.Length: 4.574755","density: 2.566225e-02<br />Sepal.Length: 4.581800","density: 2.723058e-02<br />Sepal.Length: 4.588845","density: 2.886773e-02<br />Sepal.Length: 4.595890","density: 3.055793e-02<br />Sepal.Length: 4.602935","density: 3.232753e-02<br />Sepal.Length: 4.609980","density: 3.417379e-02<br />Sepal.Length: 4.617025","density: 3.607454e-02<br />Sepal.Length: 4.624070","density: 3.805054e-02<br />Sepal.Length: 4.631115","density: 4.010975e-02<br />Sepal.Length: 4.638160","density: 4.222368e-02<br />Sepal.Length: 4.645205","density: 4.440701e-02<br />Sepal.Length: 4.652250","density: 4.667827e-02<br />Sepal.Length: 4.659295","density: 4.900306e-02<br />Sepal.Length: 4.666341","density: 5.138993e-02<br />Sepal.Length: 4.673386","density: 5.386697e-02<br />Sepal.Length: 4.680431","density: 5.639486e-02<br />Sepal.Length: 4.687476","density: 5.897632e-02<br />Sepal.Length: 4.694521","density: 6.164715e-02<br />Sepal.Length: 4.701566","density: 6.436457e-02<br />Sepal.Length: 4.708611","density: 6.712831e-02<br />Sepal.Length: 4.715656","density: 6.997303e-02<br />Sepal.Length: 4.722701","density: 7.286057e-02<br />Sepal.Length: 4.729746","density: 7.578849e-02<br />Sepal.Length: 4.736791","density: 7.878192e-02<br />Sepal.Length: 4.743836","density: 8.181457e-02<br />Sepal.Length: 4.750881","density: 8.488024e-02<br />Sepal.Length: 4.757926","density: 8.799523e-02<br />Sepal.Length: 4.764971","density: 9.114298e-02<br />Sepal.Length: 4.772016","density: 9.431517e-02<br />Sepal.Length: 4.779061","density: 9.752054e-02<br />Sepal.Length: 4.786106","density: 1.007493e-01<br />Sepal.Length: 4.793151","density: 1.039931e-01<br />Sepal.Length: 4.800196","density: 1.072547e-01<br />Sepal.Length: 4.807241","density: 1.105276e-01<br />Sepal.Length: 4.814286","density: 1.138057e-01<br />Sepal.Length: 4.821331","density: 1.170877e-01<br />Sepal.Length: 4.828376","density: 1.203669e-01<br />Sepal.Length: 4.835421","density: 1.236413e-01<br />Sepal.Length: 4.842466","density: 1.269079e-01<br />Sepal.Length: 4.849511","density: 1.301563e-01<br />Sepal.Length: 4.856556","density: 1.333904e-01<br />Sepal.Length: 4.863601","density: 1.366077e-01<br />Sepal.Length: 4.870646","density: 1.397908e-01<br />Sepal.Length: 4.877691","density: 1.429515e-01<br />Sepal.Length: 4.884736","density: 1.460893e-01<br />Sepal.Length: 4.891781","density: 1.491778e-01<br />Sepal.Length: 4.898826","density: 1.522375e-01<br />Sepal.Length: 4.905871","density: 1.552681e-01<br />Sepal.Length: 4.912916","density: 1.582432e-01<br />Sepal.Length: 4.919961","density: 1.611821e-01<br />Sepal.Length: 4.927006","density: 1.640881e-01<br />Sepal.Length: 4.934051","density: 1.669373e-01<br />Sepal.Length: 4.941096","density: 1.697454e-01<br />Sepal.Length: 4.948141","density: 1.725195e-01<br />Sepal.Length: 4.955186","density: 1.752406e-01<br />Sepal.Length: 4.962231","density: 1.779190e-01<br />Sepal.Length: 4.969276","density: 1.805658e-01<br />Sepal.Length: 4.976321","density: 1.831677e-01<br />Sepal.Length: 4.983366","density: 1.857302e-01<br />Sepal.Length: 4.990411","density: 1.882669e-01<br />Sepal.Length: 4.997456","density: 1.907706e-01<br />Sepal.Length: 5.004501","density: 1.932442e-01<br />Sepal.Length: 5.011546","density: 1.957012e-01<br />Sepal.Length: 5.018591","density: 1.981399e-01<br />Sepal.Length: 5.025636","density: 2.005645e-01<br />Sepal.Length: 5.032681","density: 2.029855e-01<br />Sepal.Length: 5.039726","density: 2.054043e-01<br />Sepal.Length: 5.046771","density: 2.078326e-01<br />Sepal.Length: 5.053816","density: 2.102732e-01<br />Sepal.Length: 5.060861","density: 2.127279e-01<br />Sepal.Length: 5.067906","density: 2.152235e-01<br />Sepal.Length: 5.074951","density: 2.177499e-01<br />Sepal.Length: 5.081996","density: 2.203080e-01<br />Sepal.Length: 5.089041","density: 2.229412e-01<br />Sepal.Length: 5.096086","density: 2.256278e-01<br />Sepal.Length: 5.103131","density: 2.283667e-01<br />Sepal.Length: 5.110176","density: 2.312108e-01<br />Sepal.Length: 5.117221","density: 2.341373e-01<br />Sepal.Length: 5.124266","density: 2.371374e-01<br />Sepal.Length: 5.131311","density: 2.402696e-01<br />Sepal.Length: 5.138356","density: 2.435174e-01<br />Sepal.Length: 5.145401","density: 2.468601e-01<br />Sepal.Length: 5.152446","density: 2.503566e-01<br />Sepal.Length: 5.159491","density: 2.540049e-01<br />Sepal.Length: 5.166536","density: 2.577682e-01<br />Sepal.Length: 5.173581","density: 2.617007e-01<br />Sepal.Length: 5.180626","density: 2.658226e-01<br />Sepal.Length: 5.187671","density: 2.700769e-01<br />Sepal.Length: 5.194716","density: 2.745091e-01<br />Sepal.Length: 5.201761","density: 2.791671e-01<br />Sepal.Length: 5.208806","density: 2.839717e-01<br />Sepal.Length: 5.215851","density: 2.889551e-01<br />Sepal.Length: 5.222896","density: 2.941975e-01<br />Sepal.Length: 5.229941","density: 2.995961e-01<br />Sepal.Length: 5.236986","density: 3.051672e-01<br />Sepal.Length: 5.244031","density: 3.110246e-01<br />Sepal.Length: 5.251076","density: 3.170424e-01<br />Sepal.Length: 5.258121","density: 3.232207e-01<br />Sepal.Length: 5.265166","density: 3.297016e-01<br />Sepal.Length: 5.272211","density: 3.363424e-01<br />Sepal.Length: 5.279256","density: 3.431414e-01<br />Sepal.Length: 5.286301","density: 3.502177e-01<br />Sepal.Length: 5.293346","density: 3.574620e-01<br />Sepal.Length: 5.300391","density: 3.648560e-01<br />Sepal.Length: 5.307436","density: 3.724929e-01<br />Sepal.Length: 5.314481","density: 3.802974e-01<br />Sepal.Length: 5.321526","density: 3.882365e-01<br />Sepal.Length: 5.328571","density: 3.963775e-01<br />Sepal.Length: 5.335616","density: 4.046752e-01<br />Sepal.Length: 5.342661","density: 4.130861e-01<br />Sepal.Length: 5.349706","density: 4.216533e-01<br />Sepal.Length: 5.356751","density: 4.303547e-01<br />Sepal.Length: 5.363796","density: 4.391425e-01<br />Sepal.Length: 5.370841","density: 4.480388e-01<br />Sepal.Length: 5.377886","density: 4.570347e-01<br />Sepal.Length: 5.384932","density: 4.660854e-01<br />Sepal.Length: 5.391977","density: 4.751974e-01<br />Sepal.Length: 5.399022","density: 4.843624e-01<br />Sepal.Length: 5.406067","density: 4.935467e-01<br />Sepal.Length: 5.413112","density: 5.027482e-01<br />Sepal.Length: 5.420157","density: 5.119447e-01<br />Sepal.Length: 5.427202","density: 5.211230e-01<br />Sepal.Length: 5.434247","density: 5.302793e-01<br />Sepal.Length: 5.441292","density: 5.393630e-01<br />Sepal.Length: 5.448337","density: 5.483898e-01<br />Sepal.Length: 5.455382","density: 5.573579e-01<br />Sepal.Length: 5.462427","density: 5.661875e-01<br />Sepal.Length: 5.469472","density: 5.749176e-01<br />Sepal.Length: 5.476517","density: 5.835514e-01<br />Sepal.Length: 5.483562","density: 5.919942e-01<br />Sepal.Length: 5.490607","density: 6.002879e-01<br />Sepal.Length: 5.497652","density: 6.084500e-01<br />Sepal.Length: 5.504697","density: 6.163802e-01<br />Sepal.Length: 5.511742","density: 6.241087e-01<br />Sepal.Length: 5.518787","density: 6.316740e-01<br />Sepal.Length: 5.525832","density: 6.389791e-01<br />Sepal.Length: 5.532877","density: 6.460294e-01<br />Sepal.Length: 5.539922","density: 6.528897e-01<br />Sepal.Length: 5.546967","density: 6.594743e-01<br />Sepal.Length: 5.554012","density: 6.657531e-01<br />Sepal.Length: 5.561057","density: 6.718210e-01<br />Sepal.Length: 5.568102","density: 6.776102e-01<br />Sepal.Length: 5.575147","density: 6.830470e-01<br />Sepal.Length: 5.582192","density: 6.882588e-01<br />Sepal.Length: 5.589237","density: 6.931997e-01<br />Sepal.Length: 5.596282","density: 6.977492e-01<br />Sepal.Length: 5.603327","density: 7.020664e-01<br />Sepal.Length: 5.610372","density: 7.061300e-01<br />Sepal.Length: 5.617417","density: 7.097724e-01<br />Sepal.Length: 5.624462","density: 7.131822e-01<br />Sepal.Length: 5.631507","density: 7.163594e-01<br />Sepal.Length: 5.638552","density: 7.191040e-01<br />Sepal.Length: 5.645597","density: 7.216185e-01<br />Sepal.Length: 5.652642","density: 7.239071e-01<br />Sepal.Length: 5.659687","density: 7.258028e-01<br />Sepal.Length: 5.666732","density: 7.274572e-01<br />Sepal.Length: 5.673777","density: 7.288980e-01<br />Sepal.Length: 5.680822","density: 7.299923e-01<br />Sepal.Length: 5.687867","density: 7.308418e-01<br />Sepal.Length: 5.694912","density: 7.314951e-01<br />Sepal.Length: 5.701957","density: 7.318515e-01<br />Sepal.Length: 5.709002","density: 7.319680e-01<br />Sepal.Length: 5.716047","density: 7.319091e-01<br />Sepal.Length: 5.723092","density: 7.316038e-01<br />Sepal.Length: 5.730137","density: 7.310710e-01<br />Sepal.Length: 5.737182","density: 7.303863e-01<br />Sepal.Length: 5.744227","density: 7.295036e-01<br />Sepal.Length: 5.751272","density: 7.284127e-01<br />Sepal.Length: 5.758317","density: 7.271945e-01<br />Sepal.Length: 5.765362","density: 7.258231e-01<br />Sepal.Length: 5.772407","density: 7.242680e-01<br />Sepal.Length: 5.779452","density: 7.226102e-01<br />Sepal.Length: 5.786497","density: 7.208387e-01<br />Sepal.Length: 5.793542","density: 7.189115e-01<br />Sepal.Length: 5.800587","density: 7.169050e-01<br />Sepal.Length: 5.807632","density: 7.148180e-01<br />Sepal.Length: 5.814677","density: 7.126052e-01<br />Sepal.Length: 5.821722","density: 7.103341e-01<br />Sepal.Length: 5.828767","density: 7.080057e-01<br />Sepal.Length: 5.835812","density: 7.055880e-01<br />Sepal.Length: 5.842857","density: 7.031265e-01<br />Sepal.Length: 5.849902","density: 7.006255e-01<br />Sepal.Length: 5.856947","density: 6.980673e-01<br />Sepal.Length: 5.863992","density: 6.954774e-01<br />Sepal.Length: 5.871037","density: 6.928622e-01<br />Sepal.Length: 5.878082","density: 6.902131e-01<br />Sepal.Length: 5.885127","density: 6.875431e-01<br />Sepal.Length: 5.892172","density: 6.848584e-01<br />Sepal.Length: 5.899217","density: 6.821551e-01<br />Sepal.Length: 5.906262","density: 6.794392e-01<br />Sepal.Length: 5.913307","density: 6.767152e-01<br />Sepal.Length: 5.920352","density: 6.739817e-01<br />Sepal.Length: 5.927397","density: 6.712403e-01<br />Sepal.Length: 5.934442","density: 6.684941e-01<br />Sepal.Length: 5.941487","density: 6.657421e-01<br />Sepal.Length: 5.948532","density: 6.629833e-01<br />Sepal.Length: 5.955577","density: 6.602198e-01<br />Sepal.Length: 5.962622","density: 6.574507e-01<br />Sepal.Length: 5.969667","density: 6.546718e-01<br />Sepal.Length: 5.976712","density: 6.518860e-01<br />Sepal.Length: 5.983757","density: 6.490924e-01<br />Sepal.Length: 5.990802","density: 6.462825e-01<br />Sepal.Length: 5.997847","density: 6.434617e-01<br />Sepal.Length: 6.004892","density: 6.406298e-01<br />Sepal.Length: 6.011937","density: 6.377726e-01<br />Sepal.Length: 6.018982","density: 6.348992e-01<br />Sepal.Length: 6.026027","density: 6.320096e-01<br />Sepal.Length: 6.033072","density: 6.290874e-01<br />Sepal.Length: 6.040117","density: 6.261415e-01<br />Sepal.Length: 6.047162","density: 6.231742e-01<br />Sepal.Length: 6.054207","density: 6.201680e-01<br />Sepal.Length: 6.061252","density: 6.171302e-01<br />Sepal.Length: 6.068297","density: 6.140661e-01<br />Sepal.Length: 6.075342","density: 6.109589e-01<br />Sepal.Length: 6.082387","density: 6.078123e-01<br />Sepal.Length: 6.089432","density: 6.046355e-01<br />Sepal.Length: 6.096477","density: 6.014139e-01<br />Sepal.Length: 6.103523","density: 5.981460e-01<br />Sepal.Length: 6.110568","density: 5.948456e-01<br />Sepal.Length: 6.117613","density: 5.915012e-01<br />Sepal.Length: 6.124658","density: 5.881055e-01<br />Sepal.Length: 6.131703","density: 5.846767e-01<br />Sepal.Length: 6.138748","density: 5.812073e-01<br />Sepal.Length: 6.145793","density: 5.776840e-01<br />Sepal.Length: 6.152838","density: 5.741291e-01<br />Sepal.Length: 6.159883","density: 5.705386e-01<br />Sepal.Length: 6.166928","density: 5.668953e-01<br />Sepal.Length: 6.173973","density: 5.632235e-01<br />Sepal.Length: 6.181018","density: 5.595229e-01<br />Sepal.Length: 6.188063","density: 5.557738e-01<br />Sepal.Length: 6.195108","density: 5.520013e-01<br />Sepal.Length: 6.202153","density: 5.482058e-01<br />Sepal.Length: 6.209198","density: 5.443734e-01<br />Sepal.Length: 6.216243","density: 5.405226e-01<br />Sepal.Length: 6.223288","density: 5.366554e-01<br />Sepal.Length: 6.230333","density: 5.327651e-01<br />Sepal.Length: 6.237378","density: 5.288632e-01<br />Sepal.Length: 6.244423","density: 5.249527e-01<br />Sepal.Length: 6.251468","density: 5.210330e-01<br />Sepal.Length: 6.258513","density: 5.171111e-01<br />Sepal.Length: 6.265558","density: 5.131892e-01<br />Sepal.Length: 6.272603","density: 5.092709e-01<br />Sepal.Length: 6.279648","density: 5.053620e-01<br />Sepal.Length: 6.286693","density: 5.014619e-01<br />Sepal.Length: 6.293738","density: 4.975769e-01<br />Sepal.Length: 6.300783","density: 4.937145e-01<br />Sepal.Length: 6.307828","density: 4.898694e-01<br />Sepal.Length: 6.314873","density: 4.860488e-01<br />Sepal.Length: 6.321918","density: 4.822648e-01<br />Sepal.Length: 6.328963","density: 4.785062e-01<br />Sepal.Length: 6.336008","density: 4.747787e-01<br />Sepal.Length: 6.343053","density: 4.711023e-01<br />Sepal.Length: 6.350098","density: 4.674581e-01<br />Sepal.Length: 6.357143","density: 4.638491e-01<br />Sepal.Length: 6.364188","density: 4.603046e-01<br />Sepal.Length: 6.371233","density: 4.567978e-01<br />Sepal.Length: 6.378278","density: 4.533287e-01<br />Sepal.Length: 6.385323","density: 4.499336e-01<br />Sepal.Length: 6.392368","density: 4.465808e-01<br />Sepal.Length: 6.399413","density: 4.432694e-01<br />Sepal.Length: 6.406458","density: 4.400318e-01<br />Sepal.Length: 6.413503","density: 4.368428e-01<br />Sepal.Length: 6.420548","density: 4.336965e-01<br />Sepal.Length: 6.427593","density: 4.306205e-01<br />Sepal.Length: 6.434638","density: 4.275969e-01<br />Sepal.Length: 6.441683","density: 4.246153e-01<br />Sepal.Length: 6.448728","density: 4.216973e-01<br />Sepal.Length: 6.455773","density: 4.188330e-01<br />Sepal.Length: 6.462818","density: 4.160076e-01<br />Sepal.Length: 6.469863","density: 4.132367e-01<br />Sepal.Length: 6.476908","density: 4.105174e-01<br />Sepal.Length: 6.483953","density: 4.078320e-01<br />Sepal.Length: 6.490998","density: 4.051901e-01<br />Sepal.Length: 6.498043","density: 4.025943e-01<br />Sepal.Length: 6.505088","density: 4.000254e-01<br />Sepal.Length: 6.512133","density: 3.974883e-01<br />Sepal.Length: 6.519178","density: 3.949877e-01<br />Sepal.Length: 6.526223","density: 3.925057e-01<br />Sepal.Length: 6.533268","density: 3.900436e-01<br />Sepal.Length: 6.540313","density: 3.876049e-01<br />Sepal.Length: 6.547358","density: 3.851754e-01<br />Sepal.Length: 6.554403","density: 3.827545e-01<br />Sepal.Length: 6.561448","density: 3.803406e-01<br />Sepal.Length: 6.568493","density: 3.779257e-01<br />Sepal.Length: 6.575538","density: 3.755092e-01<br />Sepal.Length: 6.582583","density: 3.730816e-01<br />Sepal.Length: 6.589628","density: 3.706419e-01<br />Sepal.Length: 6.596673","density: 3.681903e-01<br />Sepal.Length: 6.603718","density: 3.657116e-01<br />Sepal.Length: 6.610763","density: 3.632082e-01<br />Sepal.Length: 6.617808","density: 3.606827e-01<br />Sepal.Length: 6.624853","density: 3.581170e-01<br />Sepal.Length: 6.631898","density: 3.555128e-01<br />Sepal.Length: 6.638943","density: 3.528774e-01<br />Sepal.Length: 6.645988","density: 3.501918e-01<br />Sepal.Length: 6.653033","density: 3.474536e-01<br />Sepal.Length: 6.660078","density: 3.446761e-01<br />Sepal.Length: 6.667123","density: 3.418421e-01<br />Sepal.Length: 6.674168","density: 3.389416e-01<br />Sepal.Length: 6.681213","density: 3.359955e-01<br />Sepal.Length: 6.688258","density: 3.329899e-01<br />Sepal.Length: 6.695303","density: 3.299053e-01<br />Sepal.Length: 6.702348","density: 3.267707e-01<br />Sepal.Length: 6.709393","density: 3.235768e-01<br />Sepal.Length: 6.716438","density: 3.202934e-01<br />Sepal.Length: 6.723483","density: 3.169574e-01<br />Sepal.Length: 6.730528","density: 3.135652e-01<br />Sepal.Length: 6.737573","density: 3.100758e-01<br />Sepal.Length: 6.744618","density: 3.065334e-01<br />Sepal.Length: 6.751663","density: 3.029380e-01<br />Sepal.Length: 6.758708","density: 2.992453e-01<br />Sepal.Length: 6.765753","density: 2.954990e-01<br />Sepal.Length: 6.772798","density: 2.917013e-01<br />Sepal.Length: 6.779843","density: 2.878162e-01<br />Sepal.Length: 6.786888","density: 2.838757e-01<br />Sepal.Length: 6.793933","density: 2.798875e-01<br />Sepal.Length: 6.800978","density: 2.758238e-01<br />Sepal.Length: 6.808023","density: 2.717055e-01<br />Sepal.Length: 6.815068","density: 2.675448e-01<br />Sepal.Length: 6.822114","density: 2.633217e-01<br />Sepal.Length: 6.829159","density: 2.590478e-01<br />Sepal.Length: 6.836204","density: 2.547380e-01<br />Sepal.Length: 6.843249","density: 2.503797e-01<br />Sepal.Length: 6.850294","density: 2.459770e-01<br />Sepal.Length: 6.857339","density: 2.415462e-01<br />Sepal.Length: 6.864384","density: 2.370804e-01<br />Sepal.Length: 6.871429","density: 2.325795e-01<br />Sepal.Length: 6.878474","density: 2.280589e-01<br />Sepal.Length: 6.885519","density: 2.235162e-01<br />Sepal.Length: 6.892564","density: 2.189500e-01<br />Sepal.Length: 6.899609","density: 2.143732e-01<br />Sepal.Length: 6.906654","density: 2.097857e-01<br />Sepal.Length: 6.913699","density: 2.051888e-01<br />Sepal.Length: 6.920744","density: 2.005904e-01<br />Sepal.Length: 6.927789","density: 1.959911e-01<br />Sepal.Length: 6.934834","density: 1.913982e-01<br />Sepal.Length: 6.941879","density: 1.868131e-01<br />Sepal.Length: 6.948924","density: 1.822360e-01<br />Sepal.Length: 6.955969","density: 1.776805e-01<br />Sepal.Length: 6.963014","density: 1.731426e-01<br />Sepal.Length: 6.970059","density: 1.686216e-01<br />Sepal.Length: 6.977104","density: 1.641347e-01<br />Sepal.Length: 6.984149","density: 1.596767e-01<br />Sepal.Length: 6.991194","density: 1.552439e-01<br />Sepal.Length: 6.998239","density: 1.508552e-01<br />Sepal.Length: 7.005284","density: 1.465076e-01<br />Sepal.Length: 7.012329","density: 1.421928e-01<br />Sepal.Length: 7.019374","density: 1.379296e-01<br />Sepal.Length: 7.026419","density: 1.337203e-01<br />Sepal.Length: 7.033464","density: 1.295507e-01<br />Sepal.Length: 7.040509","density: 1.254376e-01<br />Sepal.Length: 7.047554","density: 1.213916e-01<br />Sepal.Length: 7.054599","density: 1.173912e-01<br />Sepal.Length: 7.061644","density: 1.134500e-01<br />Sepal.Length: 7.068689","density: 1.095888e-01<br />Sepal.Length: 7.075734","density: 1.057784e-01<br />Sepal.Length: 7.082779","density: 1.020276e-01<br />Sepal.Length: 7.089824","density: 9.836938e-02<br />Sepal.Length: 7.096869","density: 9.476608e-02<br />Sepal.Length: 7.103914","density: 9.122104e-02<br />Sepal.Length: 7.110959","density: 8.778039e-02<br />Sepal.Length: 7.118004","density: 8.439777e-02<br />Sepal.Length: 7.125049","density: 8.107333e-02<br />Sepal.Length: 7.132094","density: 7.785813e-02<br />Sepal.Length: 7.139139","density: 7.470610e-02<br />Sepal.Length: 7.146184","density: 7.161432e-02<br />Sepal.Length: 7.153229","density: 6.862828e-02<br />Sepal.Length: 7.160274","density: 6.571310e-02<br />Sepal.Length: 7.167319","density: 6.285928e-02<br />Sepal.Length: 7.174364","density: 6.010596e-02<br />Sepal.Length: 7.181409","density: 5.743037e-02<br />Sepal.Length: 7.188454","density: 5.481633e-02<br />Sepal.Length: 7.195499","density: 5.229612e-02<br />Sepal.Length: 7.202544","density: 4.985949e-02<br />Sepal.Length: 7.209589","density: 4.748371e-02<br />Sepal.Length: 7.216634","density: 4.519405e-02<br />Sepal.Length: 7.223679","density: 4.299259e-02<br />Sepal.Length: 7.230724","density: 4.085052e-02<br />Sepal.Length: 7.237769","density: 3.878607e-02<br />Sepal.Length: 7.244814","density: 3.681316e-02<br />Sepal.Length: 7.251859","density: 3.489744e-02<br />Sepal.Length: 7.258904","density: 3.305044e-02<br />Sepal.Length: 7.265949","density: 3.129694e-02<br />Sepal.Length: 7.272994","density: 2.959780e-02<br />Sepal.Length: 7.280039","density: 2.795836e-02<br />Sepal.Length: 7.287084","density: 2.641296e-02<br />Sepal.Length: 7.294129","density: 2.491861e-02<br />Sepal.Length: 7.301174","density: 2.347516e-02<br />Sepal.Length: 7.308219","density: 2.212475e-02<br />Sepal.Length: 7.315264","density: 2.082179e-02<br />Sepal.Length: 7.322309","density: 1.956600e-02<br />Sepal.Length: 7.329354","density: 1.839153e-02<br />Sepal.Length: 7.336399","density: 1.726536e-02<br />Sepal.Length: 7.343444","density: 1.618239e-02<br />Sepal.Length: 7.350489","density: 1.516947e-02<br />Sepal.Length: 7.357534","density: 1.420474e-02<br />Sepal.Length: 7.364579","density: 1.327910e-02<br />Sepal.Length: 7.371624","density: 1.241292e-02<br />Sepal.Length: 7.378669","density: 1.159393e-02<br />Sepal.Length: 7.385714","density: 1.080989e-02<br />Sepal.Length: 7.392759","density: 1.007556e-02<br />Sepal.Length: 7.399804","density: 9.386649e-03<br />Sepal.Length: 7.406849","density: 8.728615e-03<br />Sepal.Length: 7.413894","density: 8.111472e-03<br />Sepal.Length: 7.420939","density: 7.537348e-03<br />Sepal.Length: 7.427984","density: 6.990177e-03<br />Sepal.Length: 7.435029","density: 6.476084e-03<br />Sepal.Length: 7.442074","density: 6.002104e-03<br />Sepal.Length: 7.449119","density: 5.551373e-03<br />Sepal.Length: 7.456164","density: 5.126928e-03<br />Sepal.Length: 7.463209","density: 4.739333e-03<br />Sepal.Length: 7.470254","density: 4.371553e-03<br />Sepal.Length: 7.477299","density: 4.024265e-03<br />Sepal.Length: 7.484344","density: 3.710345e-03<br />Sepal.Length: 7.491389","density: 3.413113e-03<br />Sepal.Length: 7.498434","density: 3.132422e-03<br />Sepal.Length: 7.505479","density: 2.879732e-03<br />Sepal.Length: 7.512524","density: 2.641827e-03<br />Sepal.Length: 7.519569","density: 2.417686e-03<br />Sepal.Length: 7.526614","density: 2.215595e-03<br />Sepal.Length: 7.533659","density: 2.027023e-03<br />Sepal.Length: 7.540705","density: 1.849769e-03<br />Sepal.Length: 7.547750","density: 1.689632e-03<br />Sepal.Length: 7.554795","density: 1.541621e-03<br />Sepal.Length: 7.561840","density: 1.402810e-03<br />Sepal.Length: 7.568885","density: 1.277090e-03<br />Sepal.Length: 7.575930","density: 1.162059e-03<br />Sepal.Length: 7.582975","density: 1.054417e-03<br />Sepal.Length: 7.590020","density: 9.566335e-04<br />Sepal.Length: 7.597065","density: 8.681183e-04<br />Sepal.Length: 7.604110","density: 7.854675e-04<br />Sepal.Length: 7.611155","density: 7.101224e-04<br />Sepal.Length: 7.618200","density: 6.426882e-04<br />Sepal.Length: 7.625245","density: 5.798542e-04<br />Sepal.Length: 7.632290","density: 5.223419e-04<br />Sepal.Length: 7.639335","density: 4.714818e-04<br />Sepal.Length: 7.646380","density: 4.241878e-04<br />Sepal.Length: 7.653425","density: 3.806999e-04<br />Sepal.Length: 7.660470","density: 3.427258e-04<br />Sepal.Length: 7.667515","density: 3.074835e-04<br />Sepal.Length: 7.674560","density: 2.749345e-04<br />Sepal.Length: 7.681605","density: 2.468434e-04<br />Sepal.Length: 7.688650","density: 2.208448e-04<br />Sepal.Length: 7.695695","density: 1.968867e-04<br />Sepal.Length: 7.702740","density: 1.761433e-04<br />Sepal.Length: 7.709785","density: 1.571566e-04<br />Sepal.Length: 7.716830","density: 1.396980e-04<br />Sepal.Length: 7.723875","density: 1.245256e-04<br />Sepal.Length: 7.730920","density: 1.107996e-04<br />Sepal.Length: 7.737965","density: 9.820476e-05<br />Sepal.Length: 7.745010","density: 8.721239e-05<br />Sepal.Length: 7.752055","density: 7.738994e-05<br />Sepal.Length: 7.759100","density: 6.839512e-05<br />Sepal.Length: 7.766145","density: 6.050693e-05<br />Sepal.Length: 7.773190","density: 5.354928e-05<br />Sepal.Length: 7.780235","density: 4.719016e-05<br />Sepal.Length: 7.787280","density: 4.158344e-05<br />Sepal.Length: 7.794325","density: 3.670526e-05<br />Sepal.Length: 7.801370","density: 3.225491e-05<br />Sepal.Length: 7.808415","density: 2.830777e-05<br />Sepal.Length: 7.815460","density: 2.492252e-05<br />Sepal.Length: 7.822505","density: 2.183952e-05<br />Sepal.Length: 7.829550","density: 1.908724e-05<br />Sepal.Length: 7.836595","density: 1.676212e-05<br />Sepal.Length: 7.843640","density: 1.464802e-05<br />Sepal.Length: 7.850685","density: 1.274723e-05<br />Sepal.Length: 7.857730","density: 1.116668e-05<br />Sepal.Length: 7.864775","density: 9.731727e-06<br />Sepal.Length: 7.871820","density: 8.439688e-06<br />Sepal.Length: 7.878865","density: 7.368220e-06<br />Sepal.Length: 7.885910","density: 6.404170e-06<br />Sepal.Length: 7.892955","density: 5.537799e-06<br />Sepal.Length: 7.900000","density: 5.537799e-06<br />Sepal.Length: 7.900000","density: 6.404170e-06<br />Sepal.Length: 7.892955","density: 7.368220e-06<br />Sepal.Length: 7.885910","density: 8.439688e-06<br />Sepal.Length: 7.878865","density: 9.731727e-06<br />Sepal.Length: 7.871820","density: 1.116668e-05<br />Sepal.Length: 7.864775","density: 1.274723e-05<br />Sepal.Length: 7.857730","density: 1.464802e-05<br />Sepal.Length: 7.850685","density: 1.676212e-05<br />Sepal.Length: 7.843640","density: 1.908724e-05<br />Sepal.Length: 7.836595","density: 2.183952e-05<br />Sepal.Length: 7.829550","density: 2.492252e-05<br />Sepal.Length: 7.822505","density: 2.830777e-05<br />Sepal.Length: 7.815460","density: 3.225491e-05<br />Sepal.Length: 7.808415","density: 3.670526e-05<br />Sepal.Length: 7.801370","density: 4.158344e-05<br />Sepal.Length: 7.794325","density: 4.719016e-05<br />Sepal.Length: 7.787280","density: 5.354928e-05<br />Sepal.Length: 7.780235","density: 6.050693e-05<br />Sepal.Length: 7.773190","density: 6.839512e-05<br />Sepal.Length: 7.766145","density: 7.738994e-05<br />Sepal.Length: 7.759100","density: 8.721239e-05<br />Sepal.Length: 7.752055","density: 9.820476e-05<br />Sepal.Length: 7.745010","density: 1.107996e-04<br />Sepal.Length: 7.737965","density: 1.245256e-04<br />Sepal.Length: 7.730920","density: 1.396980e-04<br />Sepal.Length: 7.723875","density: 1.571566e-04<br />Sepal.Length: 7.716830","density: 1.761433e-04<br />Sepal.Length: 7.709785","density: 1.968867e-04<br />Sepal.Length: 7.702740","density: 2.208448e-04<br />Sepal.Length: 7.695695","density: 2.468434e-04<br />Sepal.Length: 7.688650","density: 2.749345e-04<br />Sepal.Length: 7.681605","density: 3.074835e-04<br />Sepal.Length: 7.674560","density: 3.427258e-04<br />Sepal.Length: 7.667515","density: 3.806999e-04<br />Sepal.Length: 7.660470","density: 4.241878e-04<br />Sepal.Length: 7.653425","density: 4.714818e-04<br />Sepal.Length: 7.646380","density: 5.223419e-04<br />Sepal.Length: 7.639335","density: 5.798542e-04<br />Sepal.Length: 7.632290","density: 6.426882e-04<br />Sepal.Length: 7.625245","density: 7.101224e-04<br />Sepal.Length: 7.618200","density: 7.854675e-04<br />Sepal.Length: 7.611155","density: 8.681183e-04<br />Sepal.Length: 7.604110","density: 9.566335e-04<br />Sepal.Length: 7.597065","density: 1.054417e-03<br />Sepal.Length: 7.590020","density: 1.162059e-03<br />Sepal.Length: 7.582975","density: 1.277090e-03<br />Sepal.Length: 7.575930","density: 1.402810e-03<br />Sepal.Length: 7.568885","density: 1.541621e-03<br />Sepal.Length: 7.561840","density: 1.689632e-03<br />Sepal.Length: 7.554795","density: 1.849769e-03<br />Sepal.Length: 7.547750","density: 2.027023e-03<br />Sepal.Length: 7.540705","density: 2.215595e-03<br />Sepal.Length: 7.533659","density: 2.417686e-03<br />Sepal.Length: 7.526614","density: 2.641827e-03<br />Sepal.Length: 7.519569","density: 2.879732e-03<br />Sepal.Length: 7.512524","density: 3.132422e-03<br />Sepal.Length: 7.505479","density: 3.413113e-03<br />Sepal.Length: 7.498434","density: 3.710345e-03<br />Sepal.Length: 7.491389","density: 4.024265e-03<br />Sepal.Length: 7.484344","density: 4.371553e-03<br />Sepal.Length: 7.477299","density: 4.739333e-03<br />Sepal.Length: 7.470254","density: 5.126928e-03<br />Sepal.Length: 7.463209","density: 5.551373e-03<br />Sepal.Length: 7.456164","density: 6.002104e-03<br />Sepal.Length: 7.449119","density: 6.476084e-03<br />Sepal.Length: 7.442074","density: 6.990177e-03<br />Sepal.Length: 7.435029","density: 7.537348e-03<br />Sepal.Length: 7.427984","density: 8.111472e-03<br />Sepal.Length: 7.420939","density: 8.728615e-03<br />Sepal.Length: 7.413894","density: 9.386649e-03<br />Sepal.Length: 7.406849","density: 1.007556e-02<br />Sepal.Length: 7.399804","density: 1.080989e-02<br />Sepal.Length: 7.392759","density: 1.159393e-02<br />Sepal.Length: 7.385714","density: 1.241292e-02<br />Sepal.Length: 7.378669","density: 1.327910e-02<br />Sepal.Length: 7.371624","density: 1.420474e-02<br />Sepal.Length: 7.364579","density: 1.516947e-02<br />Sepal.Length: 7.357534","density: 1.618239e-02<br />Sepal.Length: 7.350489","density: 1.726536e-02<br />Sepal.Length: 7.343444","density: 1.839153e-02<br />Sepal.Length: 7.336399","density: 1.956600e-02<br />Sepal.Length: 7.329354","density: 2.082179e-02<br />Sepal.Length: 7.322309","density: 2.212475e-02<br />Sepal.Length: 7.315264","density: 2.347516e-02<br />Sepal.Length: 7.308219","density: 2.491861e-02<br />Sepal.Length: 7.301174","density: 2.641296e-02<br />Sepal.Length: 7.294129","density: 2.795836e-02<br />Sepal.Length: 7.287084","density: 2.959780e-02<br />Sepal.Length: 7.280039","density: 3.129694e-02<br />Sepal.Length: 7.272994","density: 3.305044e-02<br />Sepal.Length: 7.265949","density: 3.489744e-02<br />Sepal.Length: 7.258904","density: 3.681316e-02<br />Sepal.Length: 7.251859","density: 3.878607e-02<br />Sepal.Length: 7.244814","density: 4.085052e-02<br />Sepal.Length: 7.237769","density: 4.299259e-02<br />Sepal.Length: 7.230724","density: 4.519405e-02<br />Sepal.Length: 7.223679","density: 4.748371e-02<br />Sepal.Length: 7.216634","density: 4.985949e-02<br />Sepal.Length: 7.209589","density: 5.229612e-02<br />Sepal.Length: 7.202544","density: 5.481633e-02<br />Sepal.Length: 7.195499","density: 5.743037e-02<br />Sepal.Length: 7.188454","density: 6.010596e-02<br />Sepal.Length: 7.181409","density: 6.285928e-02<br />Sepal.Length: 7.174364","density: 6.571310e-02<br />Sepal.Length: 7.167319","density: 6.862828e-02<br />Sepal.Length: 7.160274","density: 7.161432e-02<br />Sepal.Length: 7.153229","density: 7.470610e-02<br />Sepal.Length: 7.146184","density: 7.785813e-02<br />Sepal.Length: 7.139139","density: 8.107333e-02<br />Sepal.Length: 7.132094","density: 8.439777e-02<br />Sepal.Length: 7.125049","density: 8.778039e-02<br />Sepal.Length: 7.118004","density: 9.122104e-02<br />Sepal.Length: 7.110959","density: 9.476608e-02<br />Sepal.Length: 7.103914","density: 9.836938e-02<br />Sepal.Length: 7.096869","density: 1.020276e-01<br />Sepal.Length: 7.089824","density: 1.057784e-01<br />Sepal.Length: 7.082779","density: 1.095888e-01<br />Sepal.Length: 7.075734","density: 1.134500e-01<br />Sepal.Length: 7.068689","density: 1.173912e-01<br />Sepal.Length: 7.061644","density: 1.213916e-01<br />Sepal.Length: 7.054599","density: 1.254376e-01<br />Sepal.Length: 7.047554","density: 1.295507e-01<br />Sepal.Length: 7.040509","density: 1.337203e-01<br />Sepal.Length: 7.033464","density: 1.379296e-01<br />Sepal.Length: 7.026419","density: 1.421928e-01<br />Sepal.Length: 7.019374","density: 1.465076e-01<br />Sepal.Length: 7.012329","density: 1.508552e-01<br />Sepal.Length: 7.005284","density: 1.552439e-01<br />Sepal.Length: 6.998239","density: 1.596767e-01<br />Sepal.Length: 6.991194","density: 1.641347e-01<br />Sepal.Length: 6.984149","density: 1.686216e-01<br />Sepal.Length: 6.977104","density: 1.731426e-01<br />Sepal.Length: 6.970059","density: 1.776805e-01<br />Sepal.Length: 6.963014","density: 1.822360e-01<br />Sepal.Length: 6.955969","density: 1.868131e-01<br />Sepal.Length: 6.948924","density: 1.913982e-01<br />Sepal.Length: 6.941879","density: 1.959911e-01<br />Sepal.Length: 6.934834","density: 2.005904e-01<br />Sepal.Length: 6.927789","density: 2.051888e-01<br />Sepal.Length: 6.920744","density: 2.097857e-01<br />Sepal.Length: 6.913699","density: 2.143732e-01<br />Sepal.Length: 6.906654","density: 2.189500e-01<br />Sepal.Length: 6.899609","density: 2.235162e-01<br />Sepal.Length: 6.892564","density: 2.280589e-01<br />Sepal.Length: 6.885519","density: 2.325795e-01<br />Sepal.Length: 6.878474","density: 2.370804e-01<br />Sepal.Length: 6.871429","density: 2.415462e-01<br />Sepal.Length: 6.864384","density: 2.459770e-01<br />Sepal.Length: 6.857339","density: 2.503797e-01<br />Sepal.Length: 6.850294","density: 2.547380e-01<br />Sepal.Length: 6.843249","density: 2.590478e-01<br />Sepal.Length: 6.836204","density: 2.633217e-01<br />Sepal.Length: 6.829159","density: 2.675448e-01<br />Sepal.Length: 6.822114","density: 2.717055e-01<br />Sepal.Length: 6.815068","density: 2.758238e-01<br />Sepal.Length: 6.808023","density: 2.798875e-01<br />Sepal.Length: 6.800978","density: 2.838757e-01<br />Sepal.Length: 6.793933","density: 2.878162e-01<br />Sepal.Length: 6.786888","density: 2.917013e-01<br />Sepal.Length: 6.779843","density: 2.954990e-01<br />Sepal.Length: 6.772798","density: 2.992453e-01<br />Sepal.Length: 6.765753","density: 3.029380e-01<br />Sepal.Length: 6.758708","density: 3.065334e-01<br />Sepal.Length: 6.751663","density: 3.100758e-01<br />Sepal.Length: 6.744618","density: 3.135652e-01<br />Sepal.Length: 6.737573","density: 3.169574e-01<br />Sepal.Length: 6.730528","density: 3.202934e-01<br />Sepal.Length: 6.723483","density: 3.235768e-01<br />Sepal.Length: 6.716438","density: 3.267707e-01<br />Sepal.Length: 6.709393","density: 3.299053e-01<br />Sepal.Length: 6.702348","density: 3.329899e-01<br />Sepal.Length: 6.695303","density: 3.359955e-01<br />Sepal.Length: 6.688258","density: 3.389416e-01<br />Sepal.Length: 6.681213","density: 3.418421e-01<br />Sepal.Length: 6.674168","density: 3.446761e-01<br />Sepal.Length: 6.667123","density: 3.474536e-01<br />Sepal.Length: 6.660078","density: 3.501918e-01<br />Sepal.Length: 6.653033","density: 3.528774e-01<br />Sepal.Length: 6.645988","density: 3.555128e-01<br />Sepal.Length: 6.638943","density: 3.581170e-01<br />Sepal.Length: 6.631898","density: 3.606827e-01<br />Sepal.Length: 6.624853","density: 3.632082e-01<br />Sepal.Length: 6.617808","density: 3.657116e-01<br />Sepal.Length: 6.610763","density: 3.681903e-01<br />Sepal.Length: 6.603718","density: 3.706419e-01<br />Sepal.Length: 6.596673","density: 3.730816e-01<br />Sepal.Length: 6.589628","density: 3.755092e-01<br />Sepal.Length: 6.582583","density: 3.779257e-01<br />Sepal.Length: 6.575538","density: 3.803406e-01<br />Sepal.Length: 6.568493","density: 3.827545e-01<br />Sepal.Length: 6.561448","density: 3.851754e-01<br />Sepal.Length: 6.554403","density: 3.876049e-01<br />Sepal.Length: 6.547358","density: 3.900436e-01<br />Sepal.Length: 6.540313","density: 3.925057e-01<br />Sepal.Length: 6.533268","density: 3.949877e-01<br />Sepal.Length: 6.526223","density: 3.974883e-01<br />Sepal.Length: 6.519178","density: 4.000254e-01<br />Sepal.Length: 6.512133","density: 4.025943e-01<br />Sepal.Length: 6.505088","density: 4.051901e-01<br />Sepal.Length: 6.498043","density: 4.078320e-01<br />Sepal.Length: 6.490998","density: 4.105174e-01<br />Sepal.Length: 6.483953","density: 4.132367e-01<br />Sepal.Length: 6.476908","density: 4.160076e-01<br />Sepal.Length: 6.469863","density: 4.188330e-01<br />Sepal.Length: 6.462818","density: 4.216973e-01<br />Sepal.Length: 6.455773","density: 4.246153e-01<br />Sepal.Length: 6.448728","density: 4.275969e-01<br />Sepal.Length: 6.441683","density: 4.306205e-01<br />Sepal.Length: 6.434638","density: 4.336965e-01<br />Sepal.Length: 6.427593","density: 4.368428e-01<br />Sepal.Length: 6.420548","density: 4.400318e-01<br />Sepal.Length: 6.413503","density: 4.432694e-01<br />Sepal.Length: 6.406458","density: 4.465808e-01<br />Sepal.Length: 6.399413","density: 4.499336e-01<br />Sepal.Length: 6.392368","density: 4.533287e-01<br />Sepal.Length: 6.385323","density: 4.567978e-01<br />Sepal.Length: 6.378278","density: 4.603046e-01<br />Sepal.Length: 6.371233","density: 4.638491e-01<br />Sepal.Length: 6.364188","density: 4.674581e-01<br />Sepal.Length: 6.357143","density: 4.711023e-01<br />Sepal.Length: 6.350098","density: 4.747787e-01<br />Sepal.Length: 6.343053","density: 4.785062e-01<br />Sepal.Length: 6.336008","density: 4.822648e-01<br />Sepal.Length: 6.328963","density: 4.860488e-01<br />Sepal.Length: 6.321918","density: 4.898694e-01<br />Sepal.Length: 6.314873","density: 4.937145e-01<br />Sepal.Length: 6.307828","density: 4.975769e-01<br />Sepal.Length: 6.300783","density: 5.014619e-01<br />Sepal.Length: 6.293738","density: 5.053620e-01<br />Sepal.Length: 6.286693","density: 5.092709e-01<br />Sepal.Length: 6.279648","density: 5.131892e-01<br />Sepal.Length: 6.272603","density: 5.171111e-01<br />Sepal.Length: 6.265558","density: 5.210330e-01<br />Sepal.Length: 6.258513","density: 5.249527e-01<br />Sepal.Length: 6.251468","density: 5.288632e-01<br />Sepal.Length: 6.244423","density: 5.327651e-01<br />Sepal.Length: 6.237378","density: 5.366554e-01<br />Sepal.Length: 6.230333","density: 5.405226e-01<br />Sepal.Length: 6.223288","density: 5.443734e-01<br />Sepal.Length: 6.216243","density: 5.482058e-01<br />Sepal.Length: 6.209198","density: 5.520013e-01<br />Sepal.Length: 6.202153","density: 5.557738e-01<br />Sepal.Length: 6.195108","density: 5.595229e-01<br />Sepal.Length: 6.188063","density: 5.632235e-01<br />Sepal.Length: 6.181018","density: 5.668953e-01<br />Sepal.Length: 6.173973","density: 5.705386e-01<br />Sepal.Length: 6.166928","density: 5.741291e-01<br />Sepal.Length: 6.159883","density: 5.776840e-01<br />Sepal.Length: 6.152838","density: 5.812073e-01<br />Sepal.Length: 6.145793","density: 5.846767e-01<br />Sepal.Length: 6.138748","density: 5.881055e-01<br />Sepal.Length: 6.131703","density: 5.915012e-01<br />Sepal.Length: 6.124658","density: 5.948456e-01<br />Sepal.Length: 6.117613","density: 5.981460e-01<br />Sepal.Length: 6.110568","density: 6.014139e-01<br />Sepal.Length: 6.103523","density: 6.046355e-01<br />Sepal.Length: 6.096477","density: 6.078123e-01<br />Sepal.Length: 6.089432","density: 6.109589e-01<br />Sepal.Length: 6.082387","density: 6.140661e-01<br />Sepal.Length: 6.075342","density: 6.171302e-01<br />Sepal.Length: 6.068297","density: 6.201680e-01<br />Sepal.Length: 6.061252","density: 6.231742e-01<br />Sepal.Length: 6.054207","density: 6.261415e-01<br />Sepal.Length: 6.047162","density: 6.290874e-01<br />Sepal.Length: 6.040117","density: 6.320096e-01<br />Sepal.Length: 6.033072","density: 6.348992e-01<br />Sepal.Length: 6.026027","density: 6.377726e-01<br />Sepal.Length: 6.018982","density: 6.406298e-01<br />Sepal.Length: 6.011937","density: 6.434617e-01<br />Sepal.Length: 6.004892","density: 6.462825e-01<br />Sepal.Length: 5.997847","density: 6.490924e-01<br />Sepal.Length: 5.990802","density: 6.518860e-01<br />Sepal.Length: 5.983757","density: 6.546718e-01<br />Sepal.Length: 5.976712","density: 6.574507e-01<br />Sepal.Length: 5.969667","density: 6.602198e-01<br />Sepal.Length: 5.962622","density: 6.629833e-01<br />Sepal.Length: 5.955577","density: 6.657421e-01<br />Sepal.Length: 5.948532","density: 6.684941e-01<br />Sepal.Length: 5.941487","density: 6.712403e-01<br />Sepal.Length: 5.934442","density: 6.739817e-01<br />Sepal.Length: 5.927397","density: 6.767152e-01<br />Sepal.Length: 5.920352","density: 6.794392e-01<br />Sepal.Length: 5.913307","density: 6.821551e-01<br />Sepal.Length: 5.906262","density: 6.848584e-01<br />Sepal.Length: 5.899217","density: 6.875431e-01<br />Sepal.Length: 5.892172","density: 6.902131e-01<br />Sepal.Length: 5.885127","density: 6.928622e-01<br />Sepal.Length: 5.878082","density: 6.954774e-01<br />Sepal.Length: 5.871037","density: 6.980673e-01<br />Sepal.Length: 5.863992","density: 7.006255e-01<br />Sepal.Length: 5.856947","density: 7.031265e-01<br />Sepal.Length: 5.849902","density: 7.055880e-01<br />Sepal.Length: 5.842857","density: 7.080057e-01<br />Sepal.Length: 5.835812","density: 7.103341e-01<br />Sepal.Length: 5.828767","density: 7.126052e-01<br />Sepal.Length: 5.821722","density: 7.148180e-01<br />Sepal.Length: 5.814677","density: 7.169050e-01<br />Sepal.Length: 5.807632","density: 7.189115e-01<br />Sepal.Length: 5.800587","density: 7.208387e-01<br />Sepal.Length: 5.793542","density: 7.226102e-01<br />Sepal.Length: 5.786497","density: 7.242680e-01<br />Sepal.Length: 5.779452","density: 7.258231e-01<br />Sepal.Length: 5.772407","density: 7.271945e-01<br />Sepal.Length: 5.765362","density: 7.284127e-01<br />Sepal.Length: 5.758317","density: 7.295036e-01<br />Sepal.Length: 5.751272","density: 7.303863e-01<br />Sepal.Length: 5.744227","density: 7.310710e-01<br />Sepal.Length: 5.737182","density: 7.316038e-01<br />Sepal.Length: 5.730137","density: 7.319091e-01<br />Sepal.Length: 5.723092","density: 7.319680e-01<br />Sepal.Length: 5.716047","density: 7.318515e-01<br />Sepal.Length: 5.709002","density: 7.314951e-01<br />Sepal.Length: 5.701957","density: 7.308418e-01<br />Sepal.Length: 5.694912","density: 7.299923e-01<br />Sepal.Length: 5.687867","density: 7.288980e-01<br />Sepal.Length: 5.680822","density: 7.274572e-01<br />Sepal.Length: 5.673777","density: 7.258028e-01<br />Sepal.Length: 5.666732","density: 7.239071e-01<br />Sepal.Length: 5.659687","density: 7.216185e-01<br />Sepal.Length: 5.652642","density: 7.191040e-01<br />Sepal.Length: 5.645597","density: 7.163594e-01<br />Sepal.Length: 5.638552","density: 7.131822e-01<br />Sepal.Length: 5.631507","density: 7.097724e-01<br />Sepal.Length: 5.624462","density: 7.061300e-01<br />Sepal.Length: 5.617417","density: 7.020664e-01<br />Sepal.Length: 5.610372","density: 6.977492e-01<br />Sepal.Length: 5.603327","density: 6.931997e-01<br />Sepal.Length: 5.596282","density: 6.882588e-01<br />Sepal.Length: 5.589237","density: 6.830470e-01<br />Sepal.Length: 5.582192","density: 6.776102e-01<br />Sepal.Length: 5.575147","density: 6.718210e-01<br />Sepal.Length: 5.568102","density: 6.657531e-01<br />Sepal.Length: 5.561057","density: 6.594743e-01<br />Sepal.Length: 5.554012","density: 6.528897e-01<br />Sepal.Length: 5.546967","density: 6.460294e-01<br />Sepal.Length: 5.539922","density: 6.389791e-01<br />Sepal.Length: 5.532877","density: 6.316740e-01<br />Sepal.Length: 5.525832","density: 6.241087e-01<br />Sepal.Length: 5.518787","density: 6.163802e-01<br />Sepal.Length: 5.511742","density: 6.084500e-01<br />Sepal.Length: 5.504697","density: 6.002879e-01<br />Sepal.Length: 5.497652","density: 5.919942e-01<br />Sepal.Length: 5.490607","density: 5.835514e-01<br />Sepal.Length: 5.483562","density: 5.749176e-01<br />Sepal.Length: 5.476517","density: 5.661875e-01<br />Sepal.Length: 5.469472","density: 5.573579e-01<br />Sepal.Length: 5.462427","density: 5.483898e-01<br />Sepal.Length: 5.455382","density: 5.393630e-01<br />Sepal.Length: 5.448337","density: 5.302793e-01<br />Sepal.Length: 5.441292","density: 5.211230e-01<br />Sepal.Length: 5.434247","density: 5.119447e-01<br />Sepal.Length: 5.427202","density: 5.027482e-01<br />Sepal.Length: 5.420157","density: 4.935467e-01<br />Sepal.Length: 5.413112","density: 4.843624e-01<br />Sepal.Length: 5.406067","density: 4.751974e-01<br />Sepal.Length: 5.399022","density: 4.660854e-01<br />Sepal.Length: 5.391977","density: 4.570347e-01<br />Sepal.Length: 5.384932","density: 4.480388e-01<br />Sepal.Length: 5.377886","density: 4.391425e-01<br />Sepal.Length: 5.370841","density: 4.303547e-01<br />Sepal.Length: 5.363796","density: 4.216533e-01<br />Sepal.Length: 5.356751","density: 4.130861e-01<br />Sepal.Length: 5.349706","density: 4.046752e-01<br />Sepal.Length: 5.342661","density: 3.963775e-01<br />Sepal.Length: 5.335616","density: 3.882365e-01<br />Sepal.Length: 5.328571","density: 3.802974e-01<br />Sepal.Length: 5.321526","density: 3.724929e-01<br />Sepal.Length: 5.314481","density: 3.648560e-01<br />Sepal.Length: 5.307436","density: 3.574620e-01<br />Sepal.Length: 5.300391","density: 3.502177e-01<br />Sepal.Length: 5.293346","density: 3.431414e-01<br />Sepal.Length: 5.286301","density: 3.363424e-01<br />Sepal.Length: 5.279256","density: 3.297016e-01<br />Sepal.Length: 5.272211","density: 3.232207e-01<br />Sepal.Length: 5.265166","density: 3.170424e-01<br />Sepal.Length: 5.258121","density: 3.110246e-01<br />Sepal.Length: 5.251076","density: 3.051672e-01<br />Sepal.Length: 5.244031","density: 2.995961e-01<br />Sepal.Length: 5.236986","density: 2.941975e-01<br />Sepal.Length: 5.229941","density: 2.889551e-01<br />Sepal.Length: 5.222896","density: 2.839717e-01<br />Sepal.Length: 5.215851","density: 2.791671e-01<br />Sepal.Length: 5.208806","density: 2.745091e-01<br />Sepal.Length: 5.201761","density: 2.700769e-01<br />Sepal.Length: 5.194716","density: 2.658226e-01<br />Sepal.Length: 5.187671","density: 2.617007e-01<br />Sepal.Length: 5.180626","density: 2.577682e-01<br />Sepal.Length: 5.173581","density: 2.540049e-01<br />Sepal.Length: 5.166536","density: 2.503566e-01<br />Sepal.Length: 5.159491","density: 2.468601e-01<br />Sepal.Length: 5.152446","density: 2.435174e-01<br />Sepal.Length: 5.145401","density: 2.402696e-01<br />Sepal.Length: 5.138356","density: 2.371374e-01<br />Sepal.Length: 5.131311","density: 2.341373e-01<br />Sepal.Length: 5.124266","density: 2.312108e-01<br />Sepal.Length: 5.117221","density: 2.283667e-01<br />Sepal.Length: 5.110176","density: 2.256278e-01<br />Sepal.Length: 5.103131","density: 2.229412e-01<br />Sepal.Length: 5.096086","density: 2.203080e-01<br />Sepal.Length: 5.089041","density: 2.177499e-01<br />Sepal.Length: 5.081996","density: 2.152235e-01<br />Sepal.Length: 5.074951","density: 2.127279e-01<br />Sepal.Length: 5.067906","density: 2.102732e-01<br />Sepal.Length: 5.060861","density: 2.078326e-01<br />Sepal.Length: 5.053816","density: 2.054043e-01<br />Sepal.Length: 5.046771","density: 2.029855e-01<br />Sepal.Length: 5.039726","density: 2.005645e-01<br />Sepal.Length: 5.032681","density: 1.981399e-01<br />Sepal.Length: 5.025636","density: 1.957012e-01<br />Sepal.Length: 5.018591","density: 1.932442e-01<br />Sepal.Length: 5.011546","density: 1.907706e-01<br />Sepal.Length: 5.004501","density: 1.882669e-01<br />Sepal.Length: 4.997456","density: 1.857302e-01<br />Sepal.Length: 4.990411","density: 1.831677e-01<br />Sepal.Length: 4.983366","density: 1.805658e-01<br />Sepal.Length: 4.976321","density: 1.779190e-01<br />Sepal.Length: 4.969276","density: 1.752406e-01<br />Sepal.Length: 4.962231","density: 1.725195e-01<br />Sepal.Length: 4.955186","density: 1.697454e-01<br />Sepal.Length: 4.948141","density: 1.669373e-01<br />Sepal.Length: 4.941096","density: 1.640881e-01<br />Sepal.Length: 4.934051","density: 1.611821e-01<br />Sepal.Length: 4.927006","density: 1.582432e-01<br />Sepal.Length: 4.919961","density: 1.552681e-01<br />Sepal.Length: 4.912916","density: 1.522375e-01<br />Sepal.Length: 4.905871","density: 1.491778e-01<br />Sepal.Length: 4.898826","density: 1.460893e-01<br />Sepal.Length: 4.891781","density: 1.429515e-01<br />Sepal.Length: 4.884736","density: 1.397908e-01<br />Sepal.Length: 4.877691","density: 1.366077e-01<br />Sepal.Length: 4.870646","density: 1.333904e-01<br />Sepal.Length: 4.863601","density: 1.301563e-01<br />Sepal.Length: 4.856556","density: 1.269079e-01<br />Sepal.Length: 4.849511","density: 1.236413e-01<br />Sepal.Length: 4.842466","density: 1.203669e-01<br />Sepal.Length: 4.835421","density: 1.170877e-01<br />Sepal.Length: 4.828376","density: 1.138057e-01<br />Sepal.Length: 4.821331","density: 1.105276e-01<br />Sepal.Length: 4.814286","density: 1.072547e-01<br />Sepal.Length: 4.807241","density: 1.039931e-01<br />Sepal.Length: 4.800196","density: 1.007493e-01<br />Sepal.Length: 4.793151","density: 9.752054e-02<br />Sepal.Length: 4.786106","density: 9.431517e-02<br />Sepal.Length: 4.779061","density: 9.114298e-02<br />Sepal.Length: 4.772016","density: 8.799523e-02<br />Sepal.Length: 4.764971","density: 8.488024e-02<br />Sepal.Length: 4.757926","density: 8.181457e-02<br />Sepal.Length: 4.750881","density: 7.878192e-02<br />Sepal.Length: 4.743836","density: 7.578849e-02<br />Sepal.Length: 4.736791","density: 7.286057e-02<br />Sepal.Length: 4.729746","density: 6.997303e-02<br />Sepal.Length: 4.722701","density: 6.712831e-02<br />Sepal.Length: 4.715656","density: 6.436457e-02<br />Sepal.Length: 4.708611","density: 6.164715e-02<br />Sepal.Length: 4.701566","density: 5.897632e-02<br />Sepal.Length: 4.694521","density: 5.639486e-02<br />Sepal.Length: 4.687476","density: 5.386697e-02<br />Sepal.Length: 4.680431","density: 5.138993e-02<br />Sepal.Length: 4.673386","density: 4.900306e-02<br />Sepal.Length: 4.666341","density: 4.667827e-02<br />Sepal.Length: 4.659295","density: 4.440701e-02<br />Sepal.Length: 4.652250","density: 4.222368e-02<br />Sepal.Length: 4.645205","density: 4.010975e-02<br />Sepal.Length: 4.638160","density: 3.805054e-02<br />Sepal.Length: 4.631115","density: 3.607454e-02<br />Sepal.Length: 4.624070","density: 3.417379e-02<br />Sepal.Length: 4.617025","density: 3.232753e-02<br />Sepal.Length: 4.609980","density: 3.055793e-02<br />Sepal.Length: 4.602935","density: 2.886773e-02<br />Sepal.Length: 4.595890","density: 2.723058e-02<br />Sepal.Length: 4.588845","density: 2.566225e-02<br />Sepal.Length: 4.581800","density: 2.417573e-02<br />Sepal.Length: 4.574755","density: 2.273980e-02<br />Sepal.Length: 4.567710","density: 2.136415e-02<br />Sepal.Length: 4.560665","density: 2.007094e-02<br />Sepal.Length: 4.553620","density: 1.882508e-02<br />Sepal.Length: 4.546575","density: 1.763075e-02<br />Sepal.Length: 4.539530","density: 1.651782e-02<br />Sepal.Length: 4.532485","density: 1.544844e-02<br />Sepal.Length: 4.525440","density: 1.442243e-02<br />Sepal.Length: 4.518395","density: 1.347447e-02<br />Sepal.Length: 4.511350","density: 1.256634e-02<br />Sepal.Length: 4.504305","density: 1.169739e-02<br />Sepal.Length: 4.497260","density: 1.089491e-02<br />Sepal.Length: 4.490215","density: 1.013185e-02<br />Sepal.Length: 4.483170","density: 9.403658e-03<br />Sepal.Length: 4.476125","density: 8.731092e-03<br />Sepal.Length: 4.469080","density: 8.096666e-03<br />Sepal.Length: 4.462035","density: 7.492799e-03<br />Sepal.Length: 4.454990","density: 6.934687e-03<br />Sepal.Length: 4.447945","density: 6.412726e-03<br />Sepal.Length: 4.440900","density: 5.917165e-03<br />Sepal.Length: 4.433855","density: 5.458588e-03<br />Sepal.Length: 4.426810","density: 5.033627e-03<br />Sepal.Length: 4.419765","density: 4.631157e-03<br />Sepal.Length: 4.412720","density: 4.258054e-03<br />Sepal.Length: 4.405675","density: 3.915657e-03<br />Sepal.Length: 4.398630","density: 3.592163e-03<br />Sepal.Length: 4.391585","density: 3.291562e-03<br />Sepal.Length: 4.384540","density: 3.018545e-03<br />Sepal.Length: 4.377495","density: 2.761201e-03<br />Sepal.Length: 4.370450","density: 2.521369e-03<br />Sepal.Length: 4.363405","density: 2.305921e-03<br />Sepal.Length: 4.356360","density: 2.103300e-03<br />Sepal.Length: 4.349315","density: 1.913804e-03<br />Sepal.Length: 4.342270","density: 1.745539e-03<br />Sepal.Length: 4.335225","density: 1.587638e-03<br />Sepal.Length: 4.328180","density: 1.439987e-03<br />Sepal.Length: 4.321135","density: 1.309298e-03<br />Sepal.Length: 4.314090","density: 1.187503e-03<br />Sepal.Length: 4.307045","density: 1.073887e-03<br />Sepal.Length: 4.300000","density: 1.073887e-03<br />Sepal.Length: 4.300000"],"key":["51","52","53","54","55","56","57","58","59","60","61","62","63","64","65","66","67","68","69","70","71","72","73","74","75","76","77","78","79","80","81","82","83","84","85","86","87","88","89","90","91","92","93","94","95","96","97","98","99","100"],"type":"scatter","mode":"lines","line":{"width":1.88976377952756,"color":"rgba(0,0,0,1)","dash":"solid"},"fill":"toself","fillcolor":"rgba(0,186,56,1)","hoveron":"points","set":"SharedData91833e3a","name":"versicolor","legendgroup":"versicolor","showlegend":true,"xaxis":"x","yaxis":"y4","hoverinfo":"text","_isSimpleKey":true,"_isNestedKey":false,"frame":null},{"x":[4.3,4.30704500978474,4.31409001956947,4.32113502935421,4.32818003913894,4.33522504892368,4.34227005870842,4.34931506849315,4.35636007827789,4.36340508806262,4.37045009784736,4.37749510763209,4.38454011741683,4.39158512720157,4.3986301369863,4.40567514677104,4.41272015655577,4.41976516634051,4.42681017612524,4.43385518590998,4.44090019569472,4.44794520547945,4.45499021526419,4.46203522504892,4.46908023483366,4.4761252446184,4.48317025440313,4.49021526418787,4.4972602739726,4.50430528375734,4.51135029354207,4.51839530332681,4.52544031311155,4.53248532289628,4.53953033268102,4.54657534246575,4.55362035225049,4.56066536203523,4.56771037181996,4.5747553816047,4.58180039138943,4.58884540117417,4.5958904109589,4.60293542074364,4.60998043052838,4.61702544031311,4.62407045009785,4.63111545988258,4.63816046966732,4.64520547945205,4.65225048923679,4.65929549902153,4.66634050880626,4.673385518591,4.68043052837573,4.68747553816047,4.69452054794521,4.70156555772994,4.70861056751468,4.71565557729941,4.72270058708415,4.72974559686888,4.73679060665362,4.74383561643836,4.75088062622309,4.75792563600783,4.76497064579256,4.7720156555773,4.77906066536204,4.78610567514677,4.79315068493151,4.80019569471624,4.80724070450098,4.81428571428571,4.82133072407045,4.82837573385519,4.83542074363992,4.84246575342466,4.84951076320939,4.85655577299413,4.86360078277886,4.8706457925636,4.87769080234834,4.88473581213307,4.89178082191781,4.89882583170254,4.90587084148728,4.91291585127202,4.91996086105675,4.92700587084149,4.93405088062622,4.94109589041096,4.94814090019569,4.95518590998043,4.96223091976517,4.9692759295499,4.97632093933464,4.98336594911937,4.99041095890411,4.99745596868885,5.00450097847358,5.01154598825832,5.01859099804305,5.02563600782779,5.03268101761252,5.03972602739726,5.046771037182,5.05381604696673,5.06086105675147,5.0679060665362,5.07495107632094,5.08199608610567,5.08904109589041,5.09608610567515,5.10313111545988,5.11017612524462,5.11722113502935,5.12426614481409,5.13131115459883,5.13835616438356,5.1454011741683,5.15244618395303,5.15949119373777,5.1665362035225,5.17358121330724,5.18062622309198,5.18767123287671,5.19471624266145,5.20176125244618,5.20880626223092,5.21585127201566,5.22289628180039,5.22994129158513,5.23698630136986,5.2440313111546,5.25107632093933,5.25812133072407,5.26516634050881,5.27221135029354,5.27925636007828,5.28630136986301,5.29334637964775,5.30039138943249,5.30743639921722,5.31448140900196,5.32152641878669,5.32857142857143,5.33561643835616,5.3426614481409,5.34970645792564,5.35675146771037,5.36379647749511,5.37084148727984,5.37788649706458,5.38493150684932,5.39197651663405,5.39902152641879,5.40606653620352,5.41311154598826,5.42015655577299,5.42720156555773,5.43424657534247,5.4412915851272,5.44833659491194,5.45538160469667,5.46242661448141,5.46947162426614,5.47651663405088,5.48356164383562,5.49060665362035,5.49765166340509,5.50469667318982,5.51174168297456,5.5187866927593,5.52583170254403,5.53287671232877,5.5399217221135,5.54696673189824,5.55401174168297,5.56105675146771,5.56810176125245,5.57514677103718,5.58219178082192,5.58923679060665,5.59628180039139,5.60332681017613,5.61037181996086,5.6174168297456,5.62446183953033,5.63150684931507,5.6385518590998,5.64559686888454,5.65264187866928,5.65968688845401,5.66673189823875,5.67377690802348,5.68082191780822,5.68786692759295,5.69491193737769,5.70195694716243,5.70900195694716,5.7160469667319,5.72309197651663,5.73013698630137,5.73718199608611,5.74422700587084,5.75127201565558,5.75831702544031,5.76536203522505,5.77240704500978,5.77945205479452,5.78649706457926,5.79354207436399,5.80058708414873,5.80763209393346,5.8146771037182,5.82172211350294,5.82876712328767,5.83581213307241,5.84285714285714,5.84990215264188,5.85694716242661,5.86399217221135,5.87103718199609,5.87808219178082,5.88512720156556,5.89217221135029,5.89921722113503,5.90626223091976,5.9133072407045,5.92035225048924,5.92739726027397,5.93444227005871,5.94148727984344,5.94853228962818,5.95557729941292,5.96262230919765,5.96966731898239,5.97671232876712,5.98375733855186,5.9908023483366,5.99784735812133,6.00489236790607,6.0119373776908,6.01898238747554,6.02602739726027,6.03307240704501,6.04011741682975,6.04716242661448,6.05420743639922,6.06125244618395,6.06829745596869,6.07534246575342,6.08238747553816,6.0894324853229,6.09647749510763,6.10352250489237,6.1105675146771,6.11761252446184,6.12465753424657,6.13170254403131,6.13874755381605,6.14579256360078,6.15283757338552,6.15988258317025,6.16692759295499,6.17397260273973,6.18101761252446,6.1880626223092,6.19510763209393,6.20215264187867,6.20919765166341,6.21624266144814,6.22328767123288,6.23033268101761,6.23737769080235,6.24442270058708,6.25146771037182,6.25851272015656,6.26555772994129,6.27260273972603,6.27964774951076,6.2866927592955,6.29373776908024,6.30078277886497,6.30782778864971,6.31487279843444,6.32191780821918,6.32896281800391,6.33600782778865,6.34305283757339,6.35009784735812,6.35714285714286,6.36418786692759,6.37123287671233,6.37827788649706,6.3853228962818,6.39236790606654,6.39941291585127,6.40645792563601,6.41350293542074,6.42054794520548,6.42759295499022,6.43463796477495,6.44168297455969,6.44872798434442,6.45577299412916,6.46281800391389,6.46986301369863,6.47690802348337,6.4839530332681,6.49099804305284,6.49804305283757,6.50508806262231,6.51213307240705,6.51917808219178,6.52622309197652,6.53326810176125,6.54031311154599,6.54735812133072,6.55440313111546,6.5614481409002,6.56849315068493,6.57553816046967,6.5825831702544,6.58962818003914,6.59667318982387,6.60371819960861,6.61076320939335,6.61780821917808,6.62485322896282,6.63189823874755,6.63894324853229,6.64598825831703,6.65303326810176,6.6600782778865,6.66712328767123,6.67416829745597,6.6812133072407,6.68825831702544,6.69530332681018,6.70234833659491,6.70939334637965,6.71643835616438,6.72348336594912,6.73052837573386,6.73757338551859,6.74461839530333,6.75166340508806,6.7587084148728,6.76575342465753,6.77279843444227,6.77984344422701,6.78688845401174,6.79393346379648,6.80097847358121,6.80802348336595,6.81506849315068,6.82211350293542,6.82915851272016,6.83620352250489,6.84324853228963,6.85029354207436,6.8573385518591,6.86438356164384,6.87142857142857,6.87847358121331,6.88551859099804,6.89256360078278,6.89960861056752,6.90665362035225,6.91369863013699,6.92074363992172,6.92778864970646,6.93483365949119,6.94187866927593,6.94892367906067,6.9559686888454,6.96301369863014,6.97005870841487,6.97710371819961,6.98414872798434,6.99119373776908,6.99823874755382,7.00528375733855,7.01232876712329,7.01937377690802,7.02641878669276,7.03346379647749,7.04050880626223,7.04755381604697,7.0545988258317,7.06164383561644,7.06868884540117,7.07573385518591,7.08277886497065,7.08982387475538,7.09686888454012,7.10391389432485,7.11095890410959,7.11800391389433,7.12504892367906,7.1320939334638,7.13913894324853,7.14618395303327,7.153228962818,7.16027397260274,7.16731898238748,7.17436399217221,7.18140900195695,7.18845401174168,7.19549902152642,7.20254403131116,7.20958904109589,7.21663405088063,7.22367906066536,7.2307240704501,7.23776908023483,7.24481409001957,7.25185909980431,7.25890410958904,7.26594911937378,7.27299412915851,7.28003913894325,7.28708414872798,7.29412915851272,7.30117416829746,7.30821917808219,7.31526418786693,7.32230919765166,7.3293542074364,7.33639921722114,7.34344422700587,7.35048923679061,7.35753424657534,7.36457925636008,7.37162426614481,7.37866927592955,7.38571428571429,7.39275929549902,7.39980430528376,7.40684931506849,7.41389432485323,7.42093933463797,7.4279843444227,7.43502935420744,7.44207436399217,7.44911937377691,7.45616438356164,7.46320939334638,7.47025440313112,7.47729941291585,7.48434442270059,7.49138943248532,7.49843444227006,7.50547945205479,7.51252446183953,7.51956947162427,7.526614481409,7.53365949119374,7.54070450097847,7.54774951076321,7.55479452054795,7.56183953033268,7.56888454011742,7.57592954990215,7.58297455968689,7.59001956947163,7.59706457925636,7.6041095890411,7.61115459882583,7.61819960861057,7.6252446183953,7.63228962818004,7.63933463796478,7.64637964774951,7.65342465753425,7.66046966731898,7.66751467710372,7.67455968688845,7.68160469667319,7.68864970645793,7.69569471624266,7.7027397260274,7.70978473581213,7.71682974559687,7.7238747553816,7.73091976516634,7.73796477495108,7.74500978473581,7.75205479452055,7.75909980430528,7.76614481409002,7.77318982387476,7.78023483365949,7.78727984344423,7.79432485322896,7.8013698630137,7.80841487279844,7.81545988258317,7.82250489236791,7.82954990215264,7.83659491193738,7.84363992172211,7.85068493150685,7.85772994129159,7.86477495107632,7.87181996086106,7.87886497064579,7.88590998043053,7.89295499021526,7.9,7.9,7.9,7.89295499021526,7.88590998043053,7.87886497064579,7.87181996086106,7.86477495107632,7.85772994129159,7.85068493150685,7.84363992172211,7.83659491193738,7.82954990215264,7.82250489236791,7.81545988258317,7.80841487279844,7.8013698630137,7.79432485322896,7.78727984344423,7.78023483365949,7.77318982387476,7.76614481409002,7.75909980430528,7.75205479452055,7.74500978473581,7.73796477495108,7.73091976516634,7.7238747553816,7.71682974559687,7.70978473581213,7.7027397260274,7.69569471624266,7.68864970645793,7.68160469667319,7.67455968688845,7.66751467710372,7.66046966731898,7.65342465753425,7.64637964774951,7.63933463796478,7.63228962818004,7.6252446183953,7.61819960861057,7.61115459882583,7.6041095890411,7.59706457925636,7.59001956947163,7.58297455968689,7.57592954990215,7.56888454011742,7.56183953033268,7.55479452054795,7.54774951076321,7.54070450097847,7.53365949119374,7.526614481409,7.51956947162427,7.51252446183953,7.50547945205479,7.49843444227006,7.49138943248532,7.48434442270059,7.47729941291585,7.47025440313112,7.46320939334638,7.45616438356164,7.44911937377691,7.44207436399217,7.43502935420744,7.4279843444227,7.42093933463797,7.41389432485323,7.40684931506849,7.39980430528376,7.39275929549902,7.38571428571429,7.37866927592955,7.37162426614481,7.36457925636008,7.35753424657534,7.35048923679061,7.34344422700587,7.33639921722114,7.3293542074364,7.32230919765166,7.31526418786693,7.30821917808219,7.30117416829746,7.29412915851272,7.28708414872798,7.28003913894325,7.27299412915851,7.26594911937378,7.25890410958904,7.25185909980431,7.24481409001957,7.23776908023483,7.2307240704501,7.22367906066536,7.21663405088063,7.20958904109589,7.20254403131116,7.19549902152642,7.18845401174168,7.18140900195695,7.17436399217221,7.16731898238748,7.16027397260274,7.153228962818,7.14618395303327,7.13913894324853,7.1320939334638,7.12504892367906,7.11800391389433,7.11095890410959,7.10391389432485,7.09686888454012,7.08982387475538,7.08277886497065,7.07573385518591,7.06868884540117,7.06164383561644,7.0545988258317,7.04755381604697,7.04050880626223,7.03346379647749,7.02641878669276,7.01937377690802,7.01232876712329,7.00528375733855,6.99823874755382,6.99119373776908,6.98414872798434,6.97710371819961,6.97005870841487,6.96301369863014,6.9559686888454,6.94892367906067,6.94187866927593,6.93483365949119,6.92778864970646,6.92074363992172,6.91369863013699,6.90665362035225,6.89960861056752,6.89256360078278,6.88551859099804,6.87847358121331,6.87142857142857,6.86438356164384,6.8573385518591,6.85029354207436,6.84324853228963,6.83620352250489,6.82915851272016,6.82211350293542,6.81506849315068,6.80802348336595,6.80097847358121,6.79393346379648,6.78688845401174,6.77984344422701,6.77279843444227,6.76575342465753,6.7587084148728,6.75166340508806,6.74461839530333,6.73757338551859,6.73052837573386,6.72348336594912,6.71643835616438,6.70939334637965,6.70234833659491,6.69530332681018,6.68825831702544,6.6812133072407,6.67416829745597,6.66712328767123,6.6600782778865,6.65303326810176,6.64598825831703,6.63894324853229,6.63189823874755,6.62485322896282,6.61780821917808,6.61076320939335,6.60371819960861,6.59667318982387,6.58962818003914,6.5825831702544,6.57553816046967,6.56849315068493,6.5614481409002,6.55440313111546,6.54735812133072,6.54031311154599,6.53326810176125,6.52622309197652,6.51917808219178,6.51213307240705,6.50508806262231,6.49804305283757,6.49099804305284,6.4839530332681,6.47690802348337,6.46986301369863,6.46281800391389,6.45577299412916,6.44872798434442,6.44168297455969,6.43463796477495,6.42759295499022,6.42054794520548,6.41350293542074,6.40645792563601,6.39941291585127,6.39236790606654,6.3853228962818,6.37827788649706,6.37123287671233,6.36418786692759,6.35714285714286,6.35009784735812,6.34305283757339,6.33600782778865,6.32896281800391,6.32191780821918,6.31487279843444,6.30782778864971,6.30078277886497,6.29373776908024,6.2866927592955,6.27964774951076,6.27260273972603,6.26555772994129,6.25851272015656,6.25146771037182,6.24442270058708,6.23737769080235,6.23033268101761,6.22328767123288,6.21624266144814,6.20919765166341,6.20215264187867,6.19510763209393,6.1880626223092,6.18101761252446,6.17397260273973,6.16692759295499,6.15988258317025,6.15283757338552,6.14579256360078,6.13874755381605,6.13170254403131,6.12465753424657,6.11761252446184,6.1105675146771,6.10352250489237,6.09647749510763,6.0894324853229,6.08238747553816,6.07534246575342,6.06829745596869,6.06125244618395,6.05420743639922,6.04716242661448,6.04011741682975,6.03307240704501,6.02602739726027,6.01898238747554,6.0119373776908,6.00489236790607,5.99784735812133,5.9908023483366,5.98375733855186,5.97671232876712,5.96966731898239,5.96262230919765,5.95557729941292,5.94853228962818,5.94148727984344,5.93444227005871,5.92739726027397,5.92035225048924,5.9133072407045,5.90626223091976,5.89921722113503,5.89217221135029,5.88512720156556,5.87808219178082,5.87103718199609,5.86399217221135,5.85694716242661,5.84990215264188,5.84285714285714,5.83581213307241,5.82876712328767,5.82172211350294,5.8146771037182,5.80763209393346,5.80058708414873,5.79354207436399,5.78649706457926,5.77945205479452,5.77240704500978,5.76536203522505,5.75831702544031,5.75127201565558,5.74422700587084,5.73718199608611,5.73013698630137,5.72309197651663,5.7160469667319,5.70900195694716,5.70195694716243,5.69491193737769,5.68786692759295,5.68082191780822,5.67377690802348,5.66673189823875,5.65968688845401,5.65264187866928,5.64559686888454,5.6385518590998,5.63150684931507,5.62446183953033,5.6174168297456,5.61037181996086,5.60332681017613,5.59628180039139,5.58923679060665,5.58219178082192,5.57514677103718,5.56810176125245,5.56105675146771,5.55401174168297,5.54696673189824,5.5399217221135,5.53287671232877,5.52583170254403,5.5187866927593,5.51174168297456,5.50469667318982,5.49765166340509,5.49060665362035,5.48356164383562,5.47651663405088,5.46947162426614,5.46242661448141,5.45538160469667,5.44833659491194,5.4412915851272,5.43424657534247,5.42720156555773,5.42015655577299,5.41311154598826,5.40606653620352,5.39902152641879,5.39197651663405,5.38493150684932,5.37788649706458,5.37084148727984,5.36379647749511,5.35675146771037,5.34970645792564,5.3426614481409,5.33561643835616,5.32857142857143,5.32152641878669,5.31448140900196,5.30743639921722,5.30039138943249,5.29334637964775,5.28630136986301,5.27925636007828,5.27221135029354,5.26516634050881,5.25812133072407,5.25107632093933,5.2440313111546,5.23698630136986,5.22994129158513,5.22289628180039,5.21585127201566,5.20880626223092,5.20176125244618,5.19471624266145,5.18767123287671,5.18062622309198,5.17358121330724,5.1665362035225,5.15949119373777,5.15244618395303,5.1454011741683,5.13835616438356,5.13131115459883,5.12426614481409,5.11722113502935,5.11017612524462,5.10313111545988,5.09608610567515,5.08904109589041,5.08199608610567,5.07495107632094,5.0679060665362,5.06086105675147,5.05381604696673,5.046771037182,5.03972602739726,5.03268101761252,5.02563600782779,5.01859099804305,5.01154598825832,5.00450097847358,4.99745596868885,4.99041095890411,4.98336594911937,4.97632093933464,4.9692759295499,4.96223091976517,4.95518590998043,4.94814090019569,4.94109589041096,4.93405088062622,4.92700587084149,4.91996086105675,4.91291585127202,4.90587084148728,4.89882583170254,4.89178082191781,4.88473581213307,4.87769080234834,4.8706457925636,4.86360078277886,4.85655577299413,4.84951076320939,4.84246575342466,4.83542074363992,4.82837573385519,4.82133072407045,4.81428571428571,4.80724070450098,4.80019569471624,4.79315068493151,4.78610567514677,4.77906066536204,4.7720156555773,4.76497064579256,4.75792563600783,4.75088062622309,4.74383561643836,4.73679060665362,4.72974559686888,4.72270058708415,4.71565557729941,4.70861056751468,4.70156555772994,4.69452054794521,4.68747553816047,4.68043052837573,4.673385518591,4.66634050880626,4.65929549902153,4.65225048923679,4.64520547945205,4.63816046966732,4.63111545988258,4.62407045009785,4.61702544031311,4.60998043052838,4.60293542074364,4.5958904109589,4.58884540117417,4.58180039138943,4.5747553816047,4.56771037181996,4.56066536203523,4.55362035225049,4.54657534246575,4.53953033268102,4.53248532289628,4.52544031311155,4.51839530332681,4.51135029354207,4.50430528375734,4.4972602739726,4.49021526418787,4.48317025440313,4.4761252446184,4.46908023483366,4.46203522504892,4.45499021526419,4.44794520547945,4.44090019569472,4.43385518590998,4.42681017612524,4.41976516634051,4.41272015655577,4.40567514677104,4.3986301369863,4.39158512720157,4.38454011741683,4.37749510763209,4.37045009784736,4.36340508806262,4.35636007827789,4.34931506849315,4.34227005870842,4.33522504892368,4.32818003913894,4.32113502935421,4.31409001956947,4.30704500978474,4.3,4.3],"y":[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0.151883789230172,0.155982797475983,0.160066182672139,0.164111548521534,0.168127879921537,0.172114000589417,0.176030427071674,0.179899666036438,0.183721507186772,0.18745258073997,0.191111744474456,0.194707158577107,0.198196855250299,0.201587166455011,0.20489901141782,0.20809679632842,0.211166487703532,0.214145409496401,0.217008599873917,0.219715574386802,0.222322590836791,0.224818497184747,0.227132659546243,0.229341361012744,0.23144435452103,0.233354139476226,0.235152333234642,0.236843794409481,0.238358645425626,0.239749297507048,0.241036440114495,0.242169014959607,0.243170384857192,0.244075607861851,0.244851914955401,0.245496832350468,0.246056746226576,0.246515025909718,0.246849320297213,0.247112941760192,0.247301872778977,0.247382031418282,0.247408241279367,0.247381698819536,0.247274757464667,0.247129175852389,0.24694909117617,0.246723536073591,0.246475018484015,0.246210550095523,0.245930424805315,0.245647416553221,0.24536628616116,0.245093098294744,0.24484010045504,0.244605137251987,0.244394980107753,0.244229375137892,0.244095489617779,0.243996589773515,0.243966049721106,0.243977848454035,0.244032575090917,0.244169355599021,0.244359559517496,0.24459956683785,0.24492325358583,0.245312019907966,0.245753995215946,0.246275347867701,0.246870524979436,0.247519076929249,0.24823842376453,0.24903669510494,0.249885738211826,0.25079442401079,0.251783227344773,0.252818096000847,0.253900490968273,0.255060539155868,0.256260641634202,0.257500363510043,0.2588047285659,0.260146510856983,0.261521475024568,0.262946179246127,0.264406146937061,0.265893353708243,0.267418097381443,0.268975642922845,0.270555716574031,0.272164283498083,0.273804086174922,0.275463512966463,0.277145515169352,0.278859320264438,0.280592018122629,0.282344039920288,0.284131678713312,0.285939779478601,0.28776855293904,0.289635420519414,0.291529214648135,0.293447619796058,0.295407794584765,0.297404861144887,0.299432255438226,0.301505862729572,0.303629478268654,0.305790358023098,0.308001401966444,0.310278384274956,0.312600100216604,0.31497436990523,0.317432561397588,0.319942741497927,0.322505538451653,0.325171164492484,0.327895084572129,0.330677650939838,0.333564116773037,0.336522262249831,0.339543564711528,0.342665463699164,0.345871506836418,0.349143069870023,0.352508084083198,0.355967461645615,0.359492295211585,0.363100231595007,0.366809451257684,0.370581583320182,0.374424210595117,0.378370945665253,0.382375746288783,0.38643822733864,0.390601249146422,0.394816507846433,0.399082529804228,0.403429235504698,0.407826880756541,0.41226708605858,0.416768755902902,0.421317042356718,0.425899072848218,0.430525181722025,0.435191122716467,0.439882079162714,0.444602428103342,0.449354223250722,0.45412304355114,0.458909745676328,0.463718948638536,0.468538453224135,0.473367849432177,0.478210766223371,0.483059146000204,0.487912467584,0.492771603601101,0.497633174914839,0.502496455297136,0.507361249861723,0.512226368388334,0.517091506215681,0.521956326528494,0.526820432031003,0.531683989796536,0.536546823688282,0.541408673158369,0.546269897417604,0.551130434995866,0.55598966500493,0.560847892628919,0.565705060830347,0.570559382773844,0.575411066166738,0.580259977388083,0.585102517510555,0.589938150832293,0.594767172643419,0.599583788918251,0.604385041849984,0.60917297918076,0.613940122083762,0.618677484461162,0.6233914242803,0.628074519331111,0.63270572848926,0.637299738113105,0.64185234780526,0.646321812841631,0.650736684049223,0.655095653113932,0.659339941901247,0.663504088572992,0.667591452827986,0.671540167288758,0.675371687654631,0.679103163029813,0.682675314108709,0.686085128555167,0.689370308310649,0.692480814867927,0.695376695720867,0.69812329447619,0.700686861528158,0.702978090953793,0.705096726035914,0.707032079388067,0.70863441286361,0.710043587791214,0.71125836628005,0.712118365181817,0.71274929727074,0.713169812582039,0.713243687765896,0.713044628846136,0.712624455180569,0.711876850494511,0.710816585591404,0.709530645166763,0.707946173641771,0.70601644355504,0.703863080487186,0.701447726704552,0.698664400858826,0.695666171094249,0.692447593495334,0.688850511802533,0.685053636992895,0.681058261478023,0.676731851974842,0.672204380702485,0.667499708139834,0.662526121675981,0.657354930100766,0.652032422236593,0.64650213075937,0.640788969674552,0.63495370767734,0.628967800796369,0.622824652032726,0.616590545807502,0.610256457662065,0.603800496584617,0.597285191494066,0.590712262347018,0.584060733838902,0.577380520933765,0.570673604684397,0.563940942885323,0.557205960980675,0.550472371260926,0.543754762426708,0.537064744494809,0.530400721621303,0.523782342345476,0.517223109130473,0.510710138132622,0.504261055899975,0.497901901367238,0.491604374832305,0.48537882601863,0.479270877242603,0.47323474519938,0.467270989500631,0.461445819220271,0.455698229774187,0.450027556824586,0.444488426191821,0.439038306314378,0.433664800099945,0.42840878960531,0.423248269476424,0.418159748913574,0.413170149288235,0.408276697474594,0.403447030861422,0.398695531203258,0.394034642418165,0.389426564098179,0.384875807362036,0.380404069551908,0.375972322970548,0.371579661410726,0.367247047283448,0.362941676483231,0.358661633693482,0.354415192412697,0.350184810565587,0.345966111154635,0.341758900436148,0.337553778072277,0.333347631957241,0.329135935861215,0.324910771754723,0.320673531336746,0.316419018614867,0.312135281512074,0.307830641748202,0.303502116217011,0.299129874374179,0.294730541941986,0.29030379163904,0.285824420047609,0.281313209081043,0.276771554571447,0.272178672482079,0.267549015831231,0.262888839495242,0.25818320469051,0.253439099820377,0.248667241521551,0.243858775882836,0.239014211914932,0.234147256843622,0.229254282937511,0.224332218618019,0.219395360507271,0.21444350884456,0.209474489279381,0.20450006114988,0.199520913490938,0.19454143822714,0.18956721287995,0.184599027803504,0.179647513867601,0.174714879918576,0.169799744022014,0.164915930855691,0.160068047333408,0.155249302551177,0.150473428373575,0.145753452336163,0.141073960237367,0.136445309211854,0.131894777390876,0.127395358463024,0.12295097348774,0.118608401832332,0.114326507897369,0.110105889576493,0.105999856438437,0.101968514474121,0.0980065428187591,0.0941610693380349,0.0904081146758742,0.0867310999604286,0.0831684356639713,0.0797160331736187,0.0763445272180602,0.0730816911100521,0.0699461257732566,0.0668947854272969,0.0639435241643656,0.0611352283797713,0.0584129038892684,0.0557798257620215,0.0533036005244585,0.05091360367144,0.0486097914633586,0.0464558313371378,0.0443963336913046,0.0424217934416901,0.0405820672674232,0.0388465786893763,0.0371936158597537,0.0356594241042941,0.0342373055002746,0.0328942228179617,0.0316534598362204,0.03053043149952,0.0294820302790104,0.0285196078204377,0.0276782246790724,0.0269062667114983,0.0262044991490923,0.0256245758536195,0.0251081924729893,0.0246549205965098,0.0243061169889188,0.0240221639300852,0.0237947987064159,0.0236531923538912,0.0235765599653672,0.023549496233854,0.0235907179737696,0.0236946151547261,0.0238406454870978,0.0240389868791545,0.0242952228805582,0.0245858236230266,0.0249144906823016,0.0252937810369798,0.0256994301118005,0.0261308900284788,0.026603151755869,0.0270937014038363,0.0276019269405277,0.0281347947717319,0.0286799158350638,0.029234607388324,0.0298001127959579,0.0303698176257656,0.030941178647793,0.0315120187816374,0.0320772584325272,0.0326366118104684,0.0331867022862867,0.0337200216077904,0.0342404768705577,0.0347455389069324,0.0352217618485373,0.0356789024487187,0.0361165628169865,0.0365139650517759,0.0368865166933485,0.0372344133850663,0.037537813327462,0.0378082496050371,0.0380499775023888,0.0382458308492825,0.0384008720230228,0.0385243907800886,0.0386031890426207,0.0386341538224932,0.0386320906908075,0.038588564704172,0.0384915431860657,0.0383613096093104,0.0381944708558858,0.0379702984937617,0.0377140044386688,0.0374257007562136,0.0370810416353148,0.0367052061459677,0.0362997231623523,0.0358467427645587,0.0353618142159837,0.0348505889468976,0.0343011876480706,0.0337209011448199,0.0331184517439181,0.0324870158150377,0.03182762677519,0.0311507583836298,0.0304534471943792,0.0297328883057896,0.0289998372631922,0.0282538342063992,0.0274908469626051,0.0267204149457648,0.0259428770428294,0.0251564755727228,0.024367203586124,0.0235756780029368,0.0227832628612626,0.0219926894155833,0.0212043309078029,0.0204211915928654,0.0196452306274446,0.0188754168824322,0.0181150810320156,0.0173675441867351,0.0166294429108546,0.015903352550305,0.0151956284290577,0.0144999229031726,0.0138172437626986,0.0131581350098461,0.012512905537619,0.0118816540957972,0.0112761727743939,0.0106869213370807,0.0101127455190614,0.00956349986884765,0.00903329735783636,0.00851863250279689,0.00802704060394208,0.00755677064972911,0.00710194708643542,0.00666765109854049,0.00625632187037595,0.00585989119427852,0.00548105264404746,0.00512614790137884,0.00478524027611393,0.00445885341340397,0.00415669617251763,0.00386738300532311,0.0035908281607772,0.00333569233492729,0.00309333280038231,0.00286241318179878,0.00264910536152288,0.00244866024424731,0.00225827290848804,0.00208200846096196,0.00191831008114173,0.00176329167794655,0.00161930885643972,0.00148728093636749,0.00136261253884924,0.00124633323707177,0.00114115908028192,0.00104211958937386,0.000949267599427215,0.000866508634450933,0.000788779738332037,0.000716012024587019,0.000651129142586251,0.000590857233933596,0],"text":["density: 5.908572e-04<br />Sepal.Length: 4.300000","density: 6.511291e-04<br />Sepal.Length: 4.307045","density: 7.160120e-04<br />Sepal.Length: 4.314090","density: 7.887797e-04<br />Sepal.Length: 4.321135","density: 8.665086e-04<br />Sepal.Length: 4.328180","density: 9.492676e-04<br />Sepal.Length: 4.335225","density: 1.042120e-03<br />Sepal.Length: 4.342270","density: 1.141159e-03<br />Sepal.Length: 4.349315","density: 1.246333e-03<br />Sepal.Length: 4.356360","density: 1.362613e-03<br />Sepal.Length: 4.363405","density: 1.487281e-03<br />Sepal.Length: 4.370450","density: 1.619309e-03<br />Sepal.Length: 4.377495","density: 1.763292e-03<br />Sepal.Length: 4.384540","density: 1.918310e-03<br />Sepal.Length: 4.391585","density: 2.082008e-03<br />Sepal.Length: 4.398630","density: 2.258273e-03<br />Sepal.Length: 4.405675","density: 2.448660e-03<br />Sepal.Length: 4.412720","density: 2.649105e-03<br />Sepal.Length: 4.419765","density: 2.862413e-03<br />Sepal.Length: 4.426810","density: 3.093333e-03<br />Sepal.Length: 4.433855","density: 3.335692e-03<br />Sepal.Length: 4.440900","density: 3.590828e-03<br />Sepal.Length: 4.447945","density: 3.867383e-03<br />Sepal.Length: 4.454990","density: 4.156696e-03<br />Sepal.Length: 4.462035","density: 4.458853e-03<br />Sepal.Length: 4.469080","density: 4.785240e-03<br />Sepal.Length: 4.476125","density: 5.126148e-03<br />Sepal.Length: 4.483170","density: 5.481053e-03<br />Sepal.Length: 4.490215","density: 5.859891e-03<br />Sepal.Length: 4.497260","density: 6.256322e-03<br />Sepal.Length: 4.504305","density: 6.667651e-03<br />Sepal.Length: 4.511350","density: 7.101947e-03<br />Sepal.Length: 4.518395","density: 7.556771e-03<br />Sepal.Length: 4.525440","density: 8.027041e-03<br />Sepal.Length: 4.532485","density: 8.518633e-03<br />Sepal.Length: 4.539530","density: 9.033297e-03<br />Sepal.Length: 4.546575","density: 9.563500e-03<br />Sepal.Length: 4.553620","density: 1.011275e-02<br />Sepal.Length: 4.560665","density: 1.068692e-02<br />Sepal.Length: 4.567710","density: 1.127617e-02<br />Sepal.Length: 4.574755","density: 1.188165e-02<br />Sepal.Length: 4.581800","density: 1.251291e-02<br />Sepal.Length: 4.588845","density: 1.315814e-02<br />Sepal.Length: 4.595890","density: 1.381724e-02<br />Sepal.Length: 4.602935","density: 1.449992e-02<br />Sepal.Length: 4.609980","density: 1.519563e-02<br />Sepal.Length: 4.617025","density: 1.590335e-02<br />Sepal.Length: 4.624070","density: 1.662944e-02<br />Sepal.Length: 4.631115","density: 1.736754e-02<br />Sepal.Length: 4.638160","density: 1.811508e-02<br />Sepal.Length: 4.645205","density: 1.887542e-02<br />Sepal.Length: 4.652250","density: 1.964523e-02<br />Sepal.Length: 4.659295","density: 2.042119e-02<br />Sepal.Length: 4.666341","density: 2.120433e-02<br />Sepal.Length: 4.673386","density: 2.199269e-02<br />Sepal.Length: 4.680431","density: 2.278326e-02<br />Sepal.Length: 4.687476","density: 2.357568e-02<br />Sepal.Length: 4.694521","density: 2.436720e-02<br />Sepal.Length: 4.701566","density: 2.515648e-02<br />Sepal.Length: 4.708611","density: 2.594288e-02<br />Sepal.Length: 4.715656","density: 2.672041e-02<br />Sepal.Length: 4.722701","density: 2.749085e-02<br />Sepal.Length: 4.729746","density: 2.825383e-02<br />Sepal.Length: 4.736791","density: 2.899984e-02<br />Sepal.Length: 4.743836","density: 2.973289e-02<br />Sepal.Length: 4.750881","density: 3.045345e-02<br />Sepal.Length: 4.757926","density: 3.115076e-02<br />Sepal.Length: 4.764971","density: 3.182763e-02<br />Sepal.Length: 4.772016","density: 3.248702e-02<br />Sepal.Length: 4.779061","density: 3.311845e-02<br />Sepal.Length: 4.786106","density: 3.372090e-02<br />Sepal.Length: 4.793151","density: 3.430119e-02<br />Sepal.Length: 4.800196","density: 3.485059e-02<br />Sepal.Length: 4.807241","density: 3.536181e-02<br />Sepal.Length: 4.814286","density: 3.584674e-02<br />Sepal.Length: 4.821331","density: 3.629972e-02<br />Sepal.Length: 4.828376","density: 3.670521e-02<br />Sepal.Length: 4.835421","density: 3.708104e-02<br />Sepal.Length: 4.842466","density: 3.742570e-02<br />Sepal.Length: 4.849511","density: 3.771400e-02<br />Sepal.Length: 4.856556","density: 3.797030e-02<br />Sepal.Length: 4.863601","density: 3.819447e-02<br />Sepal.Length: 4.870646","density: 3.836131e-02<br />Sepal.Length: 4.877691","density: 3.849154e-02<br />Sepal.Length: 4.884736","density: 3.858856e-02<br />Sepal.Length: 4.891781","density: 3.863209e-02<br />Sepal.Length: 4.898826","density: 3.863415e-02<br />Sepal.Length: 4.905871","density: 3.860319e-02<br />Sepal.Length: 4.912916","density: 3.852439e-02<br />Sepal.Length: 4.919961","density: 3.840087e-02<br />Sepal.Length: 4.927006","density: 3.824583e-02<br />Sepal.Length: 4.934051","density: 3.804998e-02<br />Sepal.Length: 4.941096","density: 3.780825e-02<br />Sepal.Length: 4.948141","density: 3.753781e-02<br />Sepal.Length: 4.955186","density: 3.723441e-02<br />Sepal.Length: 4.962231","density: 3.688652e-02<br />Sepal.Length: 4.969276","density: 3.651397e-02<br />Sepal.Length: 4.976321","density: 3.611656e-02<br />Sepal.Length: 4.983366","density: 3.567890e-02<br />Sepal.Length: 4.990411","density: 3.522176e-02<br />Sepal.Length: 4.997456","density: 3.474554e-02<br />Sepal.Length: 5.004501","density: 3.424048e-02<br />Sepal.Length: 5.011546","density: 3.372002e-02<br />Sepal.Length: 5.018591","density: 3.318670e-02<br />Sepal.Length: 5.025636","density: 3.263661e-02<br />Sepal.Length: 5.032681","density: 3.207726e-02<br />Sepal.Length: 5.039726","density: 3.151202e-02<br />Sepal.Length: 5.046771","density: 3.094118e-02<br />Sepal.Length: 5.053816","density: 3.036982e-02<br />Sepal.Length: 5.060861","density: 2.980011e-02<br />Sepal.Length: 5.067906","density: 2.923461e-02<br />Sepal.Length: 5.074951","density: 2.867992e-02<br />Sepal.Length: 5.081996","density: 2.813479e-02<br />Sepal.Length: 5.089041","density: 2.760193e-02<br />Sepal.Length: 5.096086","density: 2.709370e-02<br />Sepal.Length: 5.103131","density: 2.660315e-02<br />Sepal.Length: 5.110176","density: 2.613089e-02<br />Sepal.Length: 5.117221","density: 2.569943e-02<br />Sepal.Length: 5.124266","density: 2.529378e-02<br />Sepal.Length: 5.131311","density: 2.491449e-02<br />Sepal.Length: 5.138356","density: 2.458582e-02<br />Sepal.Length: 5.145401","density: 2.429522e-02<br />Sepal.Length: 5.152446","density: 2.403899e-02<br />Sepal.Length: 5.159491","density: 2.384065e-02<br />Sepal.Length: 5.166536","density: 2.369462e-02<br />Sepal.Length: 5.173581","density: 2.359072e-02<br />Sepal.Length: 5.180626","density: 2.354950e-02<br />Sepal.Length: 5.187671","density: 2.357656e-02<br />Sepal.Length: 5.194716","density: 2.365319e-02<br />Sepal.Length: 5.201761","density: 2.379480e-02<br />Sepal.Length: 5.208806","density: 2.402216e-02<br />Sepal.Length: 5.215851","density: 2.430612e-02<br />Sepal.Length: 5.222896","density: 2.465492e-02<br />Sepal.Length: 5.229941","density: 2.510819e-02<br />Sepal.Length: 5.236986","density: 2.562458e-02<br />Sepal.Length: 5.244031","density: 2.620450e-02<br />Sepal.Length: 5.251076","density: 2.690627e-02<br />Sepal.Length: 5.258121","density: 2.767822e-02<br />Sepal.Length: 5.265166","density: 2.851961e-02<br />Sepal.Length: 5.272211","density: 2.948203e-02<br />Sepal.Length: 5.279256","density: 3.053043e-02<br />Sepal.Length: 5.286301","density: 3.165346e-02<br />Sepal.Length: 5.293346","density: 3.289422e-02<br />Sepal.Length: 5.300391","density: 3.423731e-02<br />Sepal.Length: 5.307436","density: 3.565942e-02<br />Sepal.Length: 5.314481","density: 3.719362e-02<br />Sepal.Length: 5.321526","density: 3.884658e-02<br />Sepal.Length: 5.328571","density: 4.058207e-02<br />Sepal.Length: 5.335616","density: 4.242179e-02<br />Sepal.Length: 5.342661","density: 4.439633e-02<br />Sepal.Length: 5.349706","density: 4.645583e-02<br />Sepal.Length: 5.356751","density: 4.860979e-02<br />Sepal.Length: 5.363796","density: 5.091360e-02<br />Sepal.Length: 5.370841","density: 5.330360e-02<br />Sepal.Length: 5.377886","density: 5.577983e-02<br />Sepal.Length: 5.384932","density: 5.841290e-02<br />Sepal.Length: 5.391977","density: 6.113523e-02<br />Sepal.Length: 5.399022","density: 6.394352e-02<br />Sepal.Length: 5.406067","density: 6.689479e-02<br />Sepal.Length: 5.413112","density: 6.994613e-02<br />Sepal.Length: 5.420157","density: 7.308169e-02<br />Sepal.Length: 5.427202","density: 7.634453e-02<br />Sepal.Length: 5.434247","density: 7.971603e-02<br />Sepal.Length: 5.441292","density: 8.316844e-02<br />Sepal.Length: 5.448337","density: 8.673110e-02<br />Sepal.Length: 5.455382","density: 9.040811e-02<br />Sepal.Length: 5.462427","density: 9.416107e-02<br />Sepal.Length: 5.469472","density: 9.800654e-02<br />Sepal.Length: 5.476517","density: 1.019685e-01<br />Sepal.Length: 5.483562","density: 1.059999e-01<br />Sepal.Length: 5.490607","density: 1.101059e-01<br />Sepal.Length: 5.497652","density: 1.143265e-01<br />Sepal.Length: 5.504697","density: 1.186084e-01<br />Sepal.Length: 5.511742","density: 1.229510e-01<br />Sepal.Length: 5.518787","density: 1.273954e-01<br />Sepal.Length: 5.525832","density: 1.318948e-01<br />Sepal.Length: 5.532877","density: 1.364453e-01<br />Sepal.Length: 5.539922","density: 1.410740e-01<br />Sepal.Length: 5.546967","density: 1.457535e-01<br />Sepal.Length: 5.554012","density: 1.504734e-01<br />Sepal.Length: 5.561057","density: 1.552493e-01<br />Sepal.Length: 5.568102","density: 1.600680e-01<br />Sepal.Length: 5.575147","density: 1.649159e-01<br />Sepal.Length: 5.582192","density: 1.697997e-01<br />Sepal.Length: 5.589237","density: 1.747149e-01<br />Sepal.Length: 5.596282","density: 1.796475e-01<br />Sepal.Length: 5.603327","density: 1.845990e-01<br />Sepal.Length: 5.610372","density: 1.895672e-01<br />Sepal.Length: 5.617417","density: 1.945414e-01<br />Sepal.Length: 5.624462","density: 1.995209e-01<br />Sepal.Length: 5.631507","density: 2.045001e-01<br />Sepal.Length: 5.638552","density: 2.094745e-01<br />Sepal.Length: 5.645597","density: 2.144435e-01<br />Sepal.Length: 5.652642","density: 2.193954e-01<br />Sepal.Length: 5.659687","density: 2.243322e-01<br />Sepal.Length: 5.666732","density: 2.292543e-01<br />Sepal.Length: 5.673777","density: 2.341473e-01<br />Sepal.Length: 5.680822","density: 2.390142e-01<br />Sepal.Length: 5.687867","density: 2.438588e-01<br />Sepal.Length: 5.694912","density: 2.486672e-01<br />Sepal.Length: 5.701957","density: 2.534391e-01<br />Sepal.Length: 5.709002","density: 2.581832e-01<br />Sepal.Length: 5.716047","density: 2.628888e-01<br />Sepal.Length: 5.723092","density: 2.675490e-01<br />Sepal.Length: 5.730137","density: 2.721787e-01<br />Sepal.Length: 5.737182","density: 2.767716e-01<br />Sepal.Length: 5.744227","density: 2.813132e-01<br />Sepal.Length: 5.751272","density: 2.858244e-01<br />Sepal.Length: 5.758317","density: 2.903038e-01<br />Sepal.Length: 5.765362","density: 2.947305e-01<br />Sepal.Length: 5.772407","density: 2.991299e-01<br />Sepal.Length: 5.779452","density: 3.035021e-01<br />Sepal.Length: 5.786497","density: 3.078306e-01<br />Sepal.Length: 5.793542","density: 3.121353e-01<br />Sepal.Length: 5.800587","density: 3.164190e-01<br />Sepal.Length: 5.807632","density: 3.206735e-01<br />Sepal.Length: 5.814677","density: 3.249108e-01<br />Sepal.Length: 5.821722","density: 3.291359e-01<br />Sepal.Length: 5.828767","density: 3.333476e-01<br />Sepal.Length: 5.835812","density: 3.375538e-01<br />Sepal.Length: 5.842857","density: 3.417589e-01<br />Sepal.Length: 5.849902","density: 3.459661e-01<br />Sepal.Length: 5.856947","density: 3.501848e-01<br />Sepal.Length: 5.863992","density: 3.544152e-01<br />Sepal.Length: 5.871037","density: 3.586616e-01<br />Sepal.Length: 5.878082","density: 3.629417e-01<br />Sepal.Length: 5.885127","density: 3.672470e-01<br />Sepal.Length: 5.892172","density: 3.715797e-01<br />Sepal.Length: 5.899217","density: 3.759723e-01<br />Sepal.Length: 5.906262","density: 3.804041e-01<br />Sepal.Length: 5.913307","density: 3.848758e-01<br />Sepal.Length: 5.920352","density: 3.894266e-01<br />Sepal.Length: 5.927397","density: 3.940346e-01<br />Sepal.Length: 5.934442","density: 3.986955e-01<br />Sepal.Length: 5.941487","density: 4.034470e-01<br />Sepal.Length: 5.948532","density: 4.082767e-01<br />Sepal.Length: 5.955577","density: 4.131701e-01<br />Sepal.Length: 5.962622","density: 4.181597e-01<br />Sepal.Length: 5.969667","density: 4.232483e-01<br />Sepal.Length: 5.976712","density: 4.284088e-01<br />Sepal.Length: 5.983757","density: 4.336648e-01<br />Sepal.Length: 5.990802","density: 4.390383e-01<br />Sepal.Length: 5.997847","density: 4.444884e-01<br />Sepal.Length: 6.004892","density: 4.500276e-01<br />Sepal.Length: 6.011937","density: 4.556982e-01<br />Sepal.Length: 6.018982","density: 4.614458e-01<br />Sepal.Length: 6.026027","density: 4.672710e-01<br />Sepal.Length: 6.033072","density: 4.732347e-01<br />Sepal.Length: 6.040117","density: 4.792709e-01<br />Sepal.Length: 6.047162","density: 4.853788e-01<br />Sepal.Length: 6.054207","density: 4.916044e-01<br />Sepal.Length: 6.061252","density: 4.979019e-01<br />Sepal.Length: 6.068297","density: 5.042611e-01<br />Sepal.Length: 6.075342","density: 5.107101e-01<br />Sepal.Length: 6.082387","density: 5.172231e-01<br />Sepal.Length: 6.089432","density: 5.237823e-01<br />Sepal.Length: 6.096477","density: 5.304007e-01<br />Sepal.Length: 6.103523","density: 5.370647e-01<br />Sepal.Length: 6.110568","density: 5.437548e-01<br />Sepal.Length: 6.117613","density: 5.504724e-01<br />Sepal.Length: 6.124658","density: 5.572060e-01<br />Sepal.Length: 6.131703","density: 5.639409e-01<br />Sepal.Length: 6.138748","density: 5.706736e-01<br />Sepal.Length: 6.145793","density: 5.773805e-01<br />Sepal.Length: 6.152838","density: 5.840607e-01<br />Sepal.Length: 6.159883","density: 5.907123e-01<br />Sepal.Length: 6.166928","density: 5.972852e-01<br />Sepal.Length: 6.173973","density: 6.038005e-01<br />Sepal.Length: 6.181018","density: 6.102565e-01<br />Sepal.Length: 6.188063","density: 6.165905e-01<br />Sepal.Length: 6.195108","density: 6.228247e-01<br />Sepal.Length: 6.202153","density: 6.289678e-01<br />Sepal.Length: 6.209198","density: 6.349537e-01<br />Sepal.Length: 6.216243","density: 6.407890e-01<br />Sepal.Length: 6.223288","density: 6.465021e-01<br />Sepal.Length: 6.230333","density: 6.520324e-01<br />Sepal.Length: 6.237378","density: 6.573549e-01<br />Sepal.Length: 6.244423","density: 6.625261e-01<br />Sepal.Length: 6.251468","density: 6.674997e-01<br />Sepal.Length: 6.258513","density: 6.722044e-01<br />Sepal.Length: 6.265558","density: 6.767319e-01<br />Sepal.Length: 6.272603","density: 6.810583e-01<br />Sepal.Length: 6.279648","density: 6.850536e-01<br />Sepal.Length: 6.286693","density: 6.888505e-01<br />Sepal.Length: 6.293738","density: 6.924476e-01<br />Sepal.Length: 6.300783","density: 6.956662e-01<br />Sepal.Length: 6.307828","density: 6.986644e-01<br />Sepal.Length: 6.314873","density: 7.014477e-01<br />Sepal.Length: 6.321918","density: 7.038631e-01<br />Sepal.Length: 6.328963","density: 7.060164e-01<br />Sepal.Length: 6.336008","density: 7.079462e-01<br />Sepal.Length: 6.343053","density: 7.095306e-01<br />Sepal.Length: 6.350098","density: 7.108166e-01<br />Sepal.Length: 6.357143","density: 7.118769e-01<br />Sepal.Length: 6.364188","density: 7.126245e-01<br />Sepal.Length: 6.371233","density: 7.130446e-01<br />Sepal.Length: 6.378278","density: 7.132437e-01<br />Sepal.Length: 6.385323","density: 7.131698e-01<br />Sepal.Length: 6.392368","density: 7.127493e-01<br />Sepal.Length: 6.399413","density: 7.121184e-01<br />Sepal.Length: 6.406458","density: 7.112584e-01<br />Sepal.Length: 6.413503","density: 7.100436e-01<br />Sepal.Length: 6.420548","density: 7.086344e-01<br />Sepal.Length: 6.427593","density: 7.070321e-01<br />Sepal.Length: 6.434638","density: 7.050967e-01<br />Sepal.Length: 6.441683","density: 7.029781e-01<br />Sepal.Length: 6.448728","density: 7.006869e-01<br />Sepal.Length: 6.455773","density: 6.981233e-01<br />Sepal.Length: 6.462818","density: 6.953767e-01<br />Sepal.Length: 6.469863","density: 6.924808e-01<br />Sepal.Length: 6.476908","density: 6.893703e-01<br />Sepal.Length: 6.483953","density: 6.860851e-01<br />Sepal.Length: 6.490998","density: 6.826753e-01<br />Sepal.Length: 6.498043","density: 6.791032e-01<br />Sepal.Length: 6.505088","density: 6.753717e-01<br />Sepal.Length: 6.512133","density: 6.715402e-01<br />Sepal.Length: 6.519178","density: 6.675915e-01<br />Sepal.Length: 6.526223","density: 6.635041e-01<br />Sepal.Length: 6.533268","density: 6.593399e-01<br />Sepal.Length: 6.540313","density: 6.550957e-01<br />Sepal.Length: 6.547358","density: 6.507367e-01<br />Sepal.Length: 6.554403","density: 6.463218e-01<br />Sepal.Length: 6.561448","density: 6.418523e-01<br />Sepal.Length: 6.568493","density: 6.372997e-01<br />Sepal.Length: 6.575538","density: 6.327057e-01<br />Sepal.Length: 6.582583","density: 6.280745e-01<br />Sepal.Length: 6.589628","density: 6.233914e-01<br />Sepal.Length: 6.596673","density: 6.186775e-01<br />Sepal.Length: 6.603718","density: 6.139401e-01<br />Sepal.Length: 6.610763","density: 6.091730e-01<br />Sepal.Length: 6.617808","density: 6.043850e-01<br />Sepal.Length: 6.624853","density: 5.995838e-01<br />Sepal.Length: 6.631898","density: 5.947672e-01<br />Sepal.Length: 6.638943","density: 5.899382e-01<br />Sepal.Length: 6.645988","density: 5.851025e-01<br />Sepal.Length: 6.653033","density: 5.802600e-01<br />Sepal.Length: 6.660078","density: 5.754111e-01<br />Sepal.Length: 6.667123","density: 5.705594e-01<br />Sepal.Length: 6.674168","density: 5.657051e-01<br />Sepal.Length: 6.681213","density: 5.608479e-01<br />Sepal.Length: 6.688258","density: 5.559897e-01<br />Sepal.Length: 6.695303","density: 5.511304e-01<br />Sepal.Length: 6.702348","density: 5.462699e-01<br />Sepal.Length: 6.709393","density: 5.414087e-01<br />Sepal.Length: 6.716438","density: 5.365468e-01<br />Sepal.Length: 6.723483","density: 5.316840e-01<br />Sepal.Length: 6.730528","density: 5.268204e-01<br />Sepal.Length: 6.737573","density: 5.219563e-01<br />Sepal.Length: 6.744618","density: 5.170915e-01<br />Sepal.Length: 6.751663","density: 5.122264e-01<br />Sepal.Length: 6.758708","density: 5.073612e-01<br />Sepal.Length: 6.765753","density: 5.024965e-01<br />Sepal.Length: 6.772798","density: 4.976332e-01<br />Sepal.Length: 6.779843","density: 4.927716e-01<br />Sepal.Length: 6.786888","density: 4.879125e-01<br />Sepal.Length: 6.793933","density: 4.830591e-01<br />Sepal.Length: 6.800978","density: 4.782108e-01<br />Sepal.Length: 6.808023","density: 4.733678e-01<br />Sepal.Length: 6.815068","density: 4.685385e-01<br />Sepal.Length: 6.822114","density: 4.637189e-01<br />Sepal.Length: 6.829159","density: 4.589097e-01<br />Sepal.Length: 6.836204","density: 4.541230e-01<br />Sepal.Length: 6.843249","density: 4.493542e-01<br />Sepal.Length: 6.850294","density: 4.446024e-01<br />Sepal.Length: 6.857339","density: 4.398821e-01<br />Sepal.Length: 6.864384","density: 4.351911e-01<br />Sepal.Length: 6.871429","density: 4.305252e-01<br />Sepal.Length: 6.878474","density: 4.258991e-01<br />Sepal.Length: 6.885519","density: 4.213170e-01<br />Sepal.Length: 6.892564","density: 4.167688e-01<br />Sepal.Length: 6.899609","density: 4.122671e-01<br />Sepal.Length: 6.906654","density: 4.078269e-01<br />Sepal.Length: 6.913699","density: 4.034292e-01<br />Sepal.Length: 6.920744","density: 3.990825e-01<br />Sepal.Length: 6.927789","density: 3.948165e-01<br />Sepal.Length: 6.934834","density: 3.906012e-01<br />Sepal.Length: 6.941879","density: 3.864382e-01<br />Sepal.Length: 6.948924","density: 3.823757e-01<br />Sepal.Length: 6.955969","density: 3.783709e-01<br />Sepal.Length: 6.963014","density: 3.744242e-01<br />Sepal.Length: 6.970059","density: 3.705816e-01<br />Sepal.Length: 6.977104","density: 3.668095e-01<br />Sepal.Length: 6.984149","density: 3.631002e-01<br />Sepal.Length: 6.991194","density: 3.594923e-01<br />Sepal.Length: 6.998239","density: 3.559675e-01<br />Sepal.Length: 7.005284","density: 3.525081e-01<br />Sepal.Length: 7.012329","density: 3.491431e-01<br />Sepal.Length: 7.019374","density: 3.458715e-01<br />Sepal.Length: 7.026419","density: 3.426655e-01<br />Sepal.Length: 7.033464","density: 3.395436e-01<br />Sepal.Length: 7.040509","density: 3.365223e-01<br />Sepal.Length: 7.047554","density: 3.335641e-01<br />Sepal.Length: 7.054599","density: 3.306777e-01<br />Sepal.Length: 7.061644","density: 3.278951e-01<br />Sepal.Length: 7.068689","density: 3.251712e-01<br />Sepal.Length: 7.075734","density: 3.225055e-01<br />Sepal.Length: 7.082779","density: 3.199427e-01<br />Sepal.Length: 7.089824","density: 3.174326e-01<br />Sepal.Length: 7.096869","density: 3.149744e-01<br />Sepal.Length: 7.103914","density: 3.126001e-01<br />Sepal.Length: 7.110959","density: 3.102784e-01<br />Sepal.Length: 7.118004","density: 3.080014e-01<br />Sepal.Length: 7.125049","density: 3.057904e-01<br />Sepal.Length: 7.132094","density: 3.036295e-01<br />Sepal.Length: 7.139139","density: 3.015059e-01<br />Sepal.Length: 7.146184","density: 2.994323e-01<br />Sepal.Length: 7.153229","density: 2.974049e-01<br />Sepal.Length: 7.160274","density: 2.954078e-01<br />Sepal.Length: 7.167319","density: 2.934476e-01<br />Sepal.Length: 7.174364","density: 2.915292e-01<br />Sepal.Length: 7.181409","density: 2.896354e-01<br />Sepal.Length: 7.188454","density: 2.877686e-01<br />Sepal.Length: 7.195499","density: 2.859398e-01<br />Sepal.Length: 7.202544","density: 2.841317e-01<br />Sepal.Length: 7.209589","density: 2.823440e-01<br />Sepal.Length: 7.216634","density: 2.805920e-01<br />Sepal.Length: 7.223679","density: 2.788593e-01<br />Sepal.Length: 7.230724","density: 2.771455e-01<br />Sepal.Length: 7.237769","density: 2.754635e-01<br />Sepal.Length: 7.244814","density: 2.738041e-01<br />Sepal.Length: 7.251859","density: 2.721643e-01<br />Sepal.Length: 7.258904","density: 2.705557e-01<br />Sepal.Length: 7.265949","density: 2.689756e-01<br />Sepal.Length: 7.272994","density: 2.674181e-01<br />Sepal.Length: 7.280039","density: 2.658934e-01<br />Sepal.Length: 7.287084","density: 2.644061e-01<br />Sepal.Length: 7.294129","density: 2.629462e-01<br />Sepal.Length: 7.301174","density: 2.615215e-01<br />Sepal.Length: 7.308219","density: 2.601465e-01<br />Sepal.Length: 7.315264","density: 2.588047e-01<br />Sepal.Length: 7.322309","density: 2.575004e-01<br />Sepal.Length: 7.329354","density: 2.562606e-01<br />Sepal.Length: 7.336399","density: 2.550605e-01<br />Sepal.Length: 7.343444","density: 2.539005e-01<br />Sepal.Length: 7.350489","density: 2.528181e-01<br />Sepal.Length: 7.357534","density: 2.517832e-01<br />Sepal.Length: 7.364579","density: 2.507944e-01<br />Sepal.Length: 7.371624","density: 2.498857e-01<br />Sepal.Length: 7.378669","density: 2.490367e-01<br />Sepal.Length: 7.385714","density: 2.482384e-01<br />Sepal.Length: 7.392759","density: 2.475191e-01<br />Sepal.Length: 7.399804","density: 2.468705e-01<br />Sepal.Length: 7.406849","density: 2.462753e-01<br />Sepal.Length: 7.413894","density: 2.457540e-01<br />Sepal.Length: 7.420939","density: 2.453120e-01<br />Sepal.Length: 7.427984","density: 2.449233e-01<br />Sepal.Length: 7.435029","density: 2.445996e-01<br />Sepal.Length: 7.442074","density: 2.443596e-01<br />Sepal.Length: 7.449119","density: 2.441694e-01<br />Sepal.Length: 7.456164","density: 2.440326e-01<br />Sepal.Length: 7.463209","density: 2.439778e-01<br />Sepal.Length: 7.470254","density: 2.439660e-01<br />Sepal.Length: 7.477299","density: 2.439966e-01<br />Sepal.Length: 7.484344","density: 2.440955e-01<br />Sepal.Length: 7.491389","density: 2.442294e-01<br />Sepal.Length: 7.498434","density: 2.443950e-01<br />Sepal.Length: 7.505479","density: 2.446051e-01<br />Sepal.Length: 7.512524","density: 2.448401e-01<br />Sepal.Length: 7.519569","density: 2.450931e-01<br />Sepal.Length: 7.526614","density: 2.453663e-01<br />Sepal.Length: 7.533659","density: 2.456474e-01<br />Sepal.Length: 7.540705","density: 2.459304e-01<br />Sepal.Length: 7.547750","density: 2.462106e-01<br />Sepal.Length: 7.554795","density: 2.464750e-01<br />Sepal.Length: 7.561840","density: 2.467235e-01<br />Sepal.Length: 7.568885","density: 2.469491e-01<br />Sepal.Length: 7.575930","density: 2.471292e-01<br />Sepal.Length: 7.582975","density: 2.472748e-01<br />Sepal.Length: 7.590020","density: 2.473817e-01<br />Sepal.Length: 7.597065","density: 2.474082e-01<br />Sepal.Length: 7.604110","density: 2.473820e-01<br />Sepal.Length: 7.611155","density: 2.473019e-01<br />Sepal.Length: 7.618200","density: 2.471129e-01<br />Sepal.Length: 7.625245","density: 2.468493e-01<br />Sepal.Length: 7.632290","density: 2.465150e-01<br />Sepal.Length: 7.639335","density: 2.460567e-01<br />Sepal.Length: 7.646380","density: 2.454968e-01<br />Sepal.Length: 7.653425","density: 2.448519e-01<br />Sepal.Length: 7.660470","density: 2.440756e-01<br />Sepal.Length: 7.667515","density: 2.431704e-01<br />Sepal.Length: 7.674560","density: 2.421690e-01<br />Sepal.Length: 7.681605","density: 2.410364e-01<br />Sepal.Length: 7.688650","density: 2.397493e-01<br />Sepal.Length: 7.695695","density: 2.383586e-01<br />Sepal.Length: 7.702740","density: 2.368438e-01<br />Sepal.Length: 7.709785","density: 2.351523e-01<br />Sepal.Length: 7.716830","density: 2.333541e-01<br />Sepal.Length: 7.723875","density: 2.314444e-01<br />Sepal.Length: 7.730920","density: 2.293414e-01<br />Sepal.Length: 7.737965","density: 2.271327e-01<br />Sepal.Length: 7.745010","density: 2.248185e-01<br />Sepal.Length: 7.752055","density: 2.223226e-01<br />Sepal.Length: 7.759100","density: 2.197156e-01<br />Sepal.Length: 7.766145","density: 2.170086e-01<br />Sepal.Length: 7.773190","density: 2.141454e-01<br />Sepal.Length: 7.780235","density: 2.111665e-01<br />Sepal.Length: 7.787280","density: 2.080968e-01<br />Sepal.Length: 7.794325","density: 2.048990e-01<br />Sepal.Length: 7.801370","density: 2.015872e-01<br />Sepal.Length: 7.808415","density: 1.981969e-01<br />Sepal.Length: 7.815460","density: 1.947072e-01<br />Sepal.Length: 7.822505","density: 1.911117e-01<br />Sepal.Length: 7.829550","density: 1.874526e-01<br />Sepal.Length: 7.836595","density: 1.837215e-01<br />Sepal.Length: 7.843640","density: 1.798997e-01<br />Sepal.Length: 7.850685","density: 1.760304e-01<br />Sepal.Length: 7.857730","density: 1.721140e-01<br />Sepal.Length: 7.864775","density: 1.681279e-01<br />Sepal.Length: 7.871820","density: 1.641115e-01<br />Sepal.Length: 7.878865","density: 1.600662e-01<br />Sepal.Length: 7.885910","density: 1.559828e-01<br />Sepal.Length: 7.892955","density: 1.518838e-01<br />Sepal.Length: 7.900000","density: 1.518838e-01<br />Sepal.Length: 7.900000","density: 1.518838e-01<br />Sepal.Length: 7.900000","density: 1.559828e-01<br />Sepal.Length: 7.892955","density: 1.600662e-01<br />Sepal.Length: 7.885910","density: 1.641115e-01<br />Sepal.Length: 7.878865","density: 1.681279e-01<br />Sepal.Length: 7.871820","density: 1.721140e-01<br />Sepal.Length: 7.864775","density: 1.760304e-01<br />Sepal.Length: 7.857730","density: 1.798997e-01<br />Sepal.Length: 7.850685","density: 1.837215e-01<br />Sepal.Length: 7.843640","density: 1.874526e-01<br />Sepal.Length: 7.836595","density: 1.911117e-01<br />Sepal.Length: 7.829550","density: 1.947072e-01<br />Sepal.Length: 7.822505","density: 1.981969e-01<br />Sepal.Length: 7.815460","density: 2.015872e-01<br />Sepal.Length: 7.808415","density: 2.048990e-01<br />Sepal.Length: 7.801370","density: 2.080968e-01<br />Sepal.Length: 7.794325","density: 2.111665e-01<br />Sepal.Length: 7.787280","density: 2.141454e-01<br />Sepal.Length: 7.780235","density: 2.170086e-01<br />Sepal.Length: 7.773190","density: 2.197156e-01<br />Sepal.Length: 7.766145","density: 2.223226e-01<br />Sepal.Length: 7.759100","density: 2.248185e-01<br />Sepal.Length: 7.752055","density: 2.271327e-01<br />Sepal.Length: 7.745010","density: 2.293414e-01<br />Sepal.Length: 7.737965","density: 2.314444e-01<br />Sepal.Length: 7.730920","density: 2.333541e-01<br />Sepal.Length: 7.723875","density: 2.351523e-01<br />Sepal.Length: 7.716830","density: 2.368438e-01<br />Sepal.Length: 7.709785","density: 2.383586e-01<br />Sepal.Length: 7.702740","density: 2.397493e-01<br />Sepal.Length: 7.695695","density: 2.410364e-01<br />Sepal.Length: 7.688650","density: 2.421690e-01<br />Sepal.Length: 7.681605","density: 2.431704e-01<br />Sepal.Length: 7.674560","density: 2.440756e-01<br />Sepal.Length: 7.667515","density: 2.448519e-01<br />Sepal.Length: 7.660470","density: 2.454968e-01<br />Sepal.Length: 7.653425","density: 2.460567e-01<br />Sepal.Length: 7.646380","density: 2.465150e-01<br />Sepal.Length: 7.639335","density: 2.468493e-01<br />Sepal.Length: 7.632290","density: 2.471129e-01<br />Sepal.Length: 7.625245","density: 2.473019e-01<br />Sepal.Length: 7.618200","density: 2.473820e-01<br />Sepal.Length: 7.611155","density: 2.474082e-01<br />Sepal.Length: 7.604110","density: 2.473817e-01<br />Sepal.Length: 7.597065","density: 2.472748e-01<br />Sepal.Length: 7.590020","density: 2.471292e-01<br />Sepal.Length: 7.582975","density: 2.469491e-01<br />Sepal.Length: 7.575930","density: 2.467235e-01<br />Sepal.Length: 7.568885","density: 2.464750e-01<br />Sepal.Length: 7.561840","density: 2.462106e-01<br />Sepal.Length: 7.554795","density: 2.459304e-01<br />Sepal.Length: 7.547750","density: 2.456474e-01<br />Sepal.Length: 7.540705","density: 2.453663e-01<br />Sepal.Length: 7.533659","density: 2.450931e-01<br />Sepal.Length: 7.526614","density: 2.448401e-01<br />Sepal.Length: 7.519569","density: 2.446051e-01<br />Sepal.Length: 7.512524","density: 2.443950e-01<br />Sepal.Length: 7.505479","density: 2.442294e-01<br />Sepal.Length: 7.498434","density: 2.440955e-01<br />Sepal.Length: 7.491389","density: 2.439966e-01<br />Sepal.Length: 7.484344","density: 2.439660e-01<br />Sepal.Length: 7.477299","density: 2.439778e-01<br />Sepal.Length: 7.470254","density: 2.440326e-01<br />Sepal.Length: 7.463209","density: 2.441694e-01<br />Sepal.Length: 7.456164","density: 2.443596e-01<br />Sepal.Length: 7.449119","density: 2.445996e-01<br />Sepal.Length: 7.442074","density: 2.449233e-01<br />Sepal.Length: 7.435029","density: 2.453120e-01<br />Sepal.Length: 7.427984","density: 2.457540e-01<br />Sepal.Length: 7.420939","density: 2.462753e-01<br />Sepal.Length: 7.413894","density: 2.468705e-01<br />Sepal.Length: 7.406849","density: 2.475191e-01<br />Sepal.Length: 7.399804","density: 2.482384e-01<br />Sepal.Length: 7.392759","density: 2.490367e-01<br />Sepal.Length: 7.385714","density: 2.498857e-01<br />Sepal.Length: 7.378669","density: 2.507944e-01<br />Sepal.Length: 7.371624","density: 2.517832e-01<br />Sepal.Length: 7.364579","density: 2.528181e-01<br />Sepal.Length: 7.357534","density: 2.539005e-01<br />Sepal.Length: 7.350489","density: 2.550605e-01<br />Sepal.Length: 7.343444","density: 2.562606e-01<br />Sepal.Length: 7.336399","density: 2.575004e-01<br />Sepal.Length: 7.329354","density: 2.588047e-01<br />Sepal.Length: 7.322309","density: 2.601465e-01<br />Sepal.Length: 7.315264","density: 2.615215e-01<br />Sepal.Length: 7.308219","density: 2.629462e-01<br />Sepal.Length: 7.301174","density: 2.644061e-01<br />Sepal.Length: 7.294129","density: 2.658934e-01<br />Sepal.Length: 7.287084","density: 2.674181e-01<br />Sepal.Length: 7.280039","density: 2.689756e-01<br />Sepal.Length: 7.272994","density: 2.705557e-01<br />Sepal.Length: 7.265949","density: 2.721643e-01<br />Sepal.Length: 7.258904","density: 2.738041e-01<br />Sepal.Length: 7.251859","density: 2.754635e-01<br />Sepal.Length: 7.244814","density: 2.771455e-01<br />Sepal.Length: 7.237769","density: 2.788593e-01<br />Sepal.Length: 7.230724","density: 2.805920e-01<br />Sepal.Length: 7.223679","density: 2.823440e-01<br />Sepal.Length: 7.216634","density: 2.841317e-01<br />Sepal.Length: 7.209589","density: 2.859398e-01<br />Sepal.Length: 7.202544","density: 2.877686e-01<br />Sepal.Length: 7.195499","density: 2.896354e-01<br />Sepal.Length: 7.188454","density: 2.915292e-01<br />Sepal.Length: 7.181409","density: 2.934476e-01<br />Sepal.Length: 7.174364","density: 2.954078e-01<br />Sepal.Length: 7.167319","density: 2.974049e-01<br />Sepal.Length: 7.160274","density: 2.994323e-01<br />Sepal.Length: 7.153229","density: 3.015059e-01<br />Sepal.Length: 7.146184","density: 3.036295e-01<br />Sepal.Length: 7.139139","density: 3.057904e-01<br />Sepal.Length: 7.132094","density: 3.080014e-01<br />Sepal.Length: 7.125049","density: 3.102784e-01<br />Sepal.Length: 7.118004","density: 3.126001e-01<br />Sepal.Length: 7.110959","density: 3.149744e-01<br />Sepal.Length: 7.103914","density: 3.174326e-01<br />Sepal.Length: 7.096869","density: 3.199427e-01<br />Sepal.Length: 7.089824","density: 3.225055e-01<br />Sepal.Length: 7.082779","density: 3.251712e-01<br />Sepal.Length: 7.075734","density: 3.278951e-01<br />Sepal.Length: 7.068689","density: 3.306777e-01<br />Sepal.Length: 7.061644","density: 3.335641e-01<br />Sepal.Length: 7.054599","density: 3.365223e-01<br />Sepal.Length: 7.047554","density: 3.395436e-01<br />Sepal.Length: 7.040509","density: 3.426655e-01<br />Sepal.Length: 7.033464","density: 3.458715e-01<br />Sepal.Length: 7.026419","density: 3.491431e-01<br />Sepal.Length: 7.019374","density: 3.525081e-01<br />Sepal.Length: 7.012329","density: 3.559675e-01<br />Sepal.Length: 7.005284","density: 3.594923e-01<br />Sepal.Length: 6.998239","density: 3.631002e-01<br />Sepal.Length: 6.991194","density: 3.668095e-01<br />Sepal.Length: 6.984149","density: 3.705816e-01<br />Sepal.Length: 6.977104","density: 3.744242e-01<br />Sepal.Length: 6.970059","density: 3.783709e-01<br />Sepal.Length: 6.963014","density: 3.823757e-01<br />Sepal.Length: 6.955969","density: 3.864382e-01<br />Sepal.Length: 6.948924","density: 3.906012e-01<br />Sepal.Length: 6.941879","density: 3.948165e-01<br />Sepal.Length: 6.934834","density: 3.990825e-01<br />Sepal.Length: 6.927789","density: 4.034292e-01<br />Sepal.Length: 6.920744","density: 4.078269e-01<br />Sepal.Length: 6.913699","density: 4.122671e-01<br />Sepal.Length: 6.906654","density: 4.167688e-01<br />Sepal.Length: 6.899609","density: 4.213170e-01<br />Sepal.Length: 6.892564","density: 4.258991e-01<br />Sepal.Length: 6.885519","density: 4.305252e-01<br />Sepal.Length: 6.878474","density: 4.351911e-01<br />Sepal.Length: 6.871429","density: 4.398821e-01<br />Sepal.Length: 6.864384","density: 4.446024e-01<br />Sepal.Length: 6.857339","density: 4.493542e-01<br />Sepal.Length: 6.850294","density: 4.541230e-01<br />Sepal.Length: 6.843249","density: 4.589097e-01<br />Sepal.Length: 6.836204","density: 4.637189e-01<br />Sepal.Length: 6.829159","density: 4.685385e-01<br />Sepal.Length: 6.822114","density: 4.733678e-01<br />Sepal.Length: 6.815068","density: 4.782108e-01<br />Sepal.Length: 6.808023","density: 4.830591e-01<br />Sepal.Length: 6.800978","density: 4.879125e-01<br />Sepal.Length: 6.793933","density: 4.927716e-01<br />Sepal.Length: 6.786888","density: 4.976332e-01<br />Sepal.Length: 6.779843","density: 5.024965e-01<br />Sepal.Length: 6.772798","density: 5.073612e-01<br />Sepal.Length: 6.765753","density: 5.122264e-01<br />Sepal.Length: 6.758708","density: 5.170915e-01<br />Sepal.Length: 6.751663","density: 5.219563e-01<br />Sepal.Length: 6.744618","density: 5.268204e-01<br />Sepal.Length: 6.737573","density: 5.316840e-01<br />Sepal.Length: 6.730528","density: 5.365468e-01<br />Sepal.Length: 6.723483","density: 5.414087e-01<br />Sepal.Length: 6.716438","density: 5.462699e-01<br />Sepal.Length: 6.709393","density: 5.511304e-01<br />Sepal.Length: 6.702348","density: 5.559897e-01<br />Sepal.Length: 6.695303","density: 5.608479e-01<br />Sepal.Length: 6.688258","density: 5.657051e-01<br />Sepal.Length: 6.681213","density: 5.705594e-01<br />Sepal.Length: 6.674168","density: 5.754111e-01<br />Sepal.Length: 6.667123","density: 5.802600e-01<br />Sepal.Length: 6.660078","density: 5.851025e-01<br />Sepal.Length: 6.653033","density: 5.899382e-01<br />Sepal.Length: 6.645988","density: 5.947672e-01<br />Sepal.Length: 6.638943","density: 5.995838e-01<br />Sepal.Length: 6.631898","density: 6.043850e-01<br />Sepal.Length: 6.624853","density: 6.091730e-01<br />Sepal.Length: 6.617808","density: 6.139401e-01<br />Sepal.Length: 6.610763","density: 6.186775e-01<br />Sepal.Length: 6.603718","density: 6.233914e-01<br />Sepal.Length: 6.596673","density: 6.280745e-01<br />Sepal.Length: 6.589628","density: 6.327057e-01<br />Sepal.Length: 6.582583","density: 6.372997e-01<br />Sepal.Length: 6.575538","density: 6.418523e-01<br />Sepal.Length: 6.568493","density: 6.463218e-01<br />Sepal.Length: 6.561448","density: 6.507367e-01<br />Sepal.Length: 6.554403","density: 6.550957e-01<br />Sepal.Length: 6.547358","density: 6.593399e-01<br />Sepal.Length: 6.540313","density: 6.635041e-01<br />Sepal.Length: 6.533268","density: 6.675915e-01<br />Sepal.Length: 6.526223","density: 6.715402e-01<br />Sepal.Length: 6.519178","density: 6.753717e-01<br />Sepal.Length: 6.512133","density: 6.791032e-01<br />Sepal.Length: 6.505088","density: 6.826753e-01<br />Sepal.Length: 6.498043","density: 6.860851e-01<br />Sepal.Length: 6.490998","density: 6.893703e-01<br />Sepal.Length: 6.483953","density: 6.924808e-01<br />Sepal.Length: 6.476908","density: 6.953767e-01<br />Sepal.Length: 6.469863","density: 6.981233e-01<br />Sepal.Length: 6.462818","density: 7.006869e-01<br />Sepal.Length: 6.455773","density: 7.029781e-01<br />Sepal.Length: 6.448728","density: 7.050967e-01<br />Sepal.Length: 6.441683","density: 7.070321e-01<br />Sepal.Length: 6.434638","density: 7.086344e-01<br />Sepal.Length: 6.427593","density: 7.100436e-01<br />Sepal.Length: 6.420548","density: 7.112584e-01<br />Sepal.Length: 6.413503","density: 7.121184e-01<br />Sepal.Length: 6.406458","density: 7.127493e-01<br />Sepal.Length: 6.399413","density: 7.131698e-01<br />Sepal.Length: 6.392368","density: 7.132437e-01<br />Sepal.Length: 6.385323","density: 7.130446e-01<br />Sepal.Length: 6.378278","density: 7.126245e-01<br />Sepal.Length: 6.371233","density: 7.118769e-01<br />Sepal.Length: 6.364188","density: 7.108166e-01<br />Sepal.Length: 6.357143","density: 7.095306e-01<br />Sepal.Length: 6.350098","density: 7.079462e-01<br />Sepal.Length: 6.343053","density: 7.060164e-01<br />Sepal.Length: 6.336008","density: 7.038631e-01<br />Sepal.Length: 6.328963","density: 7.014477e-01<br />Sepal.Length: 6.321918","density: 6.986644e-01<br />Sepal.Length: 6.314873","density: 6.956662e-01<br />Sepal.Length: 6.307828","density: 6.924476e-01<br />Sepal.Length: 6.300783","density: 6.888505e-01<br />Sepal.Length: 6.293738","density: 6.850536e-01<br />Sepal.Length: 6.286693","density: 6.810583e-01<br />Sepal.Length: 6.279648","density: 6.767319e-01<br />Sepal.Length: 6.272603","density: 6.722044e-01<br />Sepal.Length: 6.265558","density: 6.674997e-01<br />Sepal.Length: 6.258513","density: 6.625261e-01<br />Sepal.Length: 6.251468","density: 6.573549e-01<br />Sepal.Length: 6.244423","density: 6.520324e-01<br />Sepal.Length: 6.237378","density: 6.465021e-01<br />Sepal.Length: 6.230333","density: 6.407890e-01<br />Sepal.Length: 6.223288","density: 6.349537e-01<br />Sepal.Length: 6.216243","density: 6.289678e-01<br />Sepal.Length: 6.209198","density: 6.228247e-01<br />Sepal.Length: 6.202153","density: 6.165905e-01<br />Sepal.Length: 6.195108","density: 6.102565e-01<br />Sepal.Length: 6.188063","density: 6.038005e-01<br />Sepal.Length: 6.181018","density: 5.972852e-01<br />Sepal.Length: 6.173973","density: 5.907123e-01<br />Sepal.Length: 6.166928","density: 5.840607e-01<br />Sepal.Length: 6.159883","density: 5.773805e-01<br />Sepal.Length: 6.152838","density: 5.706736e-01<br />Sepal.Length: 6.145793","density: 5.639409e-01<br />Sepal.Length: 6.138748","density: 5.572060e-01<br />Sepal.Length: 6.131703","density: 5.504724e-01<br />Sepal.Length: 6.124658","density: 5.437548e-01<br />Sepal.Length: 6.117613","density: 5.370647e-01<br />Sepal.Length: 6.110568","density: 5.304007e-01<br />Sepal.Length: 6.103523","density: 5.237823e-01<br />Sepal.Length: 6.096477","density: 5.172231e-01<br />Sepal.Length: 6.089432","density: 5.107101e-01<br />Sepal.Length: 6.082387","density: 5.042611e-01<br />Sepal.Length: 6.075342","density: 4.979019e-01<br />Sepal.Length: 6.068297","density: 4.916044e-01<br />Sepal.Length: 6.061252","density: 4.853788e-01<br />Sepal.Length: 6.054207","density: 4.792709e-01<br />Sepal.Length: 6.047162","density: 4.732347e-01<br />Sepal.Length: 6.040117","density: 4.672710e-01<br />Sepal.Length: 6.033072","density: 4.614458e-01<br />Sepal.Length: 6.026027","density: 4.556982e-01<br />Sepal.Length: 6.018982","density: 4.500276e-01<br />Sepal.Length: 6.011937","density: 4.444884e-01<br />Sepal.Length: 6.004892","density: 4.390383e-01<br />Sepal.Length: 5.997847","density: 4.336648e-01<br />Sepal.Length: 5.990802","density: 4.284088e-01<br />Sepal.Length: 5.983757","density: 4.232483e-01<br />Sepal.Length: 5.976712","density: 4.181597e-01<br />Sepal.Length: 5.969667","density: 4.131701e-01<br />Sepal.Length: 5.962622","density: 4.082767e-01<br />Sepal.Length: 5.955577","density: 4.034470e-01<br />Sepal.Length: 5.948532","density: 3.986955e-01<br />Sepal.Length: 5.941487","density: 3.940346e-01<br />Sepal.Length: 5.934442","density: 3.894266e-01<br />Sepal.Length: 5.927397","density: 3.848758e-01<br />Sepal.Length: 5.920352","density: 3.804041e-01<br />Sepal.Length: 5.913307","density: 3.759723e-01<br />Sepal.Length: 5.906262","density: 3.715797e-01<br />Sepal.Length: 5.899217","density: 3.672470e-01<br />Sepal.Length: 5.892172","density: 3.629417e-01<br />Sepal.Length: 5.885127","density: 3.586616e-01<br />Sepal.Length: 5.878082","density: 3.544152e-01<br />Sepal.Length: 5.871037","density: 3.501848e-01<br />Sepal.Length: 5.863992","density: 3.459661e-01<br />Sepal.Length: 5.856947","density: 3.417589e-01<br />Sepal.Length: 5.849902","density: 3.375538e-01<br />Sepal.Length: 5.842857","density: 3.333476e-01<br />Sepal.Length: 5.835812","density: 3.291359e-01<br />Sepal.Length: 5.828767","density: 3.249108e-01<br />Sepal.Length: 5.821722","density: 3.206735e-01<br />Sepal.Length: 5.814677","density: 3.164190e-01<br />Sepal.Length: 5.807632","density: 3.121353e-01<br />Sepal.Length: 5.800587","density: 3.078306e-01<br />Sepal.Length: 5.793542","density: 3.035021e-01<br />Sepal.Length: 5.786497","density: 2.991299e-01<br />Sepal.Length: 5.779452","density: 2.947305e-01<br />Sepal.Length: 5.772407","density: 2.903038e-01<br />Sepal.Length: 5.765362","density: 2.858244e-01<br />Sepal.Length: 5.758317","density: 2.813132e-01<br />Sepal.Length: 5.751272","density: 2.767716e-01<br />Sepal.Length: 5.744227","density: 2.721787e-01<br />Sepal.Length: 5.737182","density: 2.675490e-01<br />Sepal.Length: 5.730137","density: 2.628888e-01<br />Sepal.Length: 5.723092","density: 2.581832e-01<br />Sepal.Length: 5.716047","density: 2.534391e-01<br />Sepal.Length: 5.709002","density: 2.486672e-01<br />Sepal.Length: 5.701957","density: 2.438588e-01<br />Sepal.Length: 5.694912","density: 2.390142e-01<br />Sepal.Length: 5.687867","density: 2.341473e-01<br />Sepal.Length: 5.680822","density: 2.292543e-01<br />Sepal.Length: 5.673777","density: 2.243322e-01<br />Sepal.Length: 5.666732","density: 2.193954e-01<br />Sepal.Length: 5.659687","density: 2.144435e-01<br />Sepal.Length: 5.652642","density: 2.094745e-01<br />Sepal.Length: 5.645597","density: 2.045001e-01<br />Sepal.Length: 5.638552","density: 1.995209e-01<br />Sepal.Length: 5.631507","density: 1.945414e-01<br />Sepal.Length: 5.624462","density: 1.895672e-01<br />Sepal.Length: 5.617417","density: 1.845990e-01<br />Sepal.Length: 5.610372","density: 1.796475e-01<br />Sepal.Length: 5.603327","density: 1.747149e-01<br />Sepal.Length: 5.596282","density: 1.697997e-01<br />Sepal.Length: 5.589237","density: 1.649159e-01<br />Sepal.Length: 5.582192","density: 1.600680e-01<br />Sepal.Length: 5.575147","density: 1.552493e-01<br />Sepal.Length: 5.568102","density: 1.504734e-01<br />Sepal.Length: 5.561057","density: 1.457535e-01<br />Sepal.Length: 5.554012","density: 1.410740e-01<br />Sepal.Length: 5.546967","density: 1.364453e-01<br />Sepal.Length: 5.539922","density: 1.318948e-01<br />Sepal.Length: 5.532877","density: 1.273954e-01<br />Sepal.Length: 5.525832","density: 1.229510e-01<br />Sepal.Length: 5.518787","density: 1.186084e-01<br />Sepal.Length: 5.511742","density: 1.143265e-01<br />Sepal.Length: 5.504697","density: 1.101059e-01<br />Sepal.Length: 5.497652","density: 1.059999e-01<br />Sepal.Length: 5.490607","density: 1.019685e-01<br />Sepal.Length: 5.483562","density: 9.800654e-02<br />Sepal.Length: 5.476517","density: 9.416107e-02<br />Sepal.Length: 5.469472","density: 9.040811e-02<br />Sepal.Length: 5.462427","density: 8.673110e-02<br />Sepal.Length: 5.455382","density: 8.316844e-02<br />Sepal.Length: 5.448337","density: 7.971603e-02<br />Sepal.Length: 5.441292","density: 7.634453e-02<br />Sepal.Length: 5.434247","density: 7.308169e-02<br />Sepal.Length: 5.427202","density: 6.994613e-02<br />Sepal.Length: 5.420157","density: 6.689479e-02<br />Sepal.Length: 5.413112","density: 6.394352e-02<br />Sepal.Length: 5.406067","density: 6.113523e-02<br />Sepal.Length: 5.399022","density: 5.841290e-02<br />Sepal.Length: 5.391977","density: 5.577983e-02<br />Sepal.Length: 5.384932","density: 5.330360e-02<br />Sepal.Length: 5.377886","density: 5.091360e-02<br />Sepal.Length: 5.370841","density: 4.860979e-02<br />Sepal.Length: 5.363796","density: 4.645583e-02<br />Sepal.Length: 5.356751","density: 4.439633e-02<br />Sepal.Length: 5.349706","density: 4.242179e-02<br />Sepal.Length: 5.342661","density: 4.058207e-02<br />Sepal.Length: 5.335616","density: 3.884658e-02<br />Sepal.Length: 5.328571","density: 3.719362e-02<br />Sepal.Length: 5.321526","density: 3.565942e-02<br />Sepal.Length: 5.314481","density: 3.423731e-02<br />Sepal.Length: 5.307436","density: 3.289422e-02<br />Sepal.Length: 5.300391","density: 3.165346e-02<br />Sepal.Length: 5.293346","density: 3.053043e-02<br />Sepal.Length: 5.286301","density: 2.948203e-02<br />Sepal.Length: 5.279256","density: 2.851961e-02<br />Sepal.Length: 5.272211","density: 2.767822e-02<br />Sepal.Length: 5.265166","density: 2.690627e-02<br />Sepal.Length: 5.258121","density: 2.620450e-02<br />Sepal.Length: 5.251076","density: 2.562458e-02<br />Sepal.Length: 5.244031","density: 2.510819e-02<br />Sepal.Length: 5.236986","density: 2.465492e-02<br />Sepal.Length: 5.229941","density: 2.430612e-02<br />Sepal.Length: 5.222896","density: 2.402216e-02<br />Sepal.Length: 5.215851","density: 2.379480e-02<br />Sepal.Length: 5.208806","density: 2.365319e-02<br />Sepal.Length: 5.201761","density: 2.357656e-02<br />Sepal.Length: 5.194716","density: 2.354950e-02<br />Sepal.Length: 5.187671","density: 2.359072e-02<br />Sepal.Length: 5.180626","density: 2.369462e-02<br />Sepal.Length: 5.173581","density: 2.384065e-02<br />Sepal.Length: 5.166536","density: 2.403899e-02<br />Sepal.Length: 5.159491","density: 2.429522e-02<br />Sepal.Length: 5.152446","density: 2.458582e-02<br />Sepal.Length: 5.145401","density: 2.491449e-02<br />Sepal.Length: 5.138356","density: 2.529378e-02<br />Sepal.Length: 5.131311","density: 2.569943e-02<br />Sepal.Length: 5.124266","density: 2.613089e-02<br />Sepal.Length: 5.117221","density: 2.660315e-02<br />Sepal.Length: 5.110176","density: 2.709370e-02<br />Sepal.Length: 5.103131","density: 2.760193e-02<br />Sepal.Length: 5.096086","density: 2.813479e-02<br />Sepal.Length: 5.089041","density: 2.867992e-02<br />Sepal.Length: 5.081996","density: 2.923461e-02<br />Sepal.Length: 5.074951","density: 2.980011e-02<br />Sepal.Length: 5.067906","density: 3.036982e-02<br />Sepal.Length: 5.060861","density: 3.094118e-02<br />Sepal.Length: 5.053816","density: 3.151202e-02<br />Sepal.Length: 5.046771","density: 3.207726e-02<br />Sepal.Length: 5.039726","density: 3.263661e-02<br />Sepal.Length: 5.032681","density: 3.318670e-02<br />Sepal.Length: 5.025636","density: 3.372002e-02<br />Sepal.Length: 5.018591","density: 3.424048e-02<br />Sepal.Length: 5.011546","density: 3.474554e-02<br />Sepal.Length: 5.004501","density: 3.522176e-02<br />Sepal.Length: 4.997456","density: 3.567890e-02<br />Sepal.Length: 4.990411","density: 3.611656e-02<br />Sepal.Length: 4.983366","density: 3.651397e-02<br />Sepal.Length: 4.976321","density: 3.688652e-02<br />Sepal.Length: 4.969276","density: 3.723441e-02<br />Sepal.Length: 4.962231","density: 3.753781e-02<br />Sepal.Length: 4.955186","density: 3.780825e-02<br />Sepal.Length: 4.948141","density: 3.804998e-02<br />Sepal.Length: 4.941096","density: 3.824583e-02<br />Sepal.Length: 4.934051","density: 3.840087e-02<br />Sepal.Length: 4.927006","density: 3.852439e-02<br />Sepal.Length: 4.919961","density: 3.860319e-02<br />Sepal.Length: 4.912916","density: 3.863415e-02<br />Sepal.Length: 4.905871","density: 3.863209e-02<br />Sepal.Length: 4.898826","density: 3.858856e-02<br />Sepal.Length: 4.891781","density: 3.849154e-02<br />Sepal.Length: 4.884736","density: 3.836131e-02<br />Sepal.Length: 4.877691","density: 3.819447e-02<br />Sepal.Length: 4.870646","density: 3.797030e-02<br />Sepal.Length: 4.863601","density: 3.771400e-02<br />Sepal.Length: 4.856556","density: 3.742570e-02<br />Sepal.Length: 4.849511","density: 3.708104e-02<br />Sepal.Length: 4.842466","density: 3.670521e-02<br />Sepal.Length: 4.835421","density: 3.629972e-02<br />Sepal.Length: 4.828376","density: 3.584674e-02<br />Sepal.Length: 4.821331","density: 3.536181e-02<br />Sepal.Length: 4.814286","density: 3.485059e-02<br />Sepal.Length: 4.807241","density: 3.430119e-02<br />Sepal.Length: 4.800196","density: 3.372090e-02<br />Sepal.Length: 4.793151","density: 3.311845e-02<br />Sepal.Length: 4.786106","density: 3.248702e-02<br />Sepal.Length: 4.779061","density: 3.182763e-02<br />Sepal.Length: 4.772016","density: 3.115076e-02<br />Sepal.Length: 4.764971","density: 3.045345e-02<br />Sepal.Length: 4.757926","density: 2.973289e-02<br />Sepal.Length: 4.750881","density: 2.899984e-02<br />Sepal.Length: 4.743836","density: 2.825383e-02<br />Sepal.Length: 4.736791","density: 2.749085e-02<br />Sepal.Length: 4.729746","density: 2.672041e-02<br />Sepal.Length: 4.722701","density: 2.594288e-02<br />Sepal.Length: 4.715656","density: 2.515648e-02<br />Sepal.Length: 4.708611","density: 2.436720e-02<br />Sepal.Length: 4.701566","density: 2.357568e-02<br />Sepal.Length: 4.694521","density: 2.278326e-02<br />Sepal.Length: 4.687476","density: 2.199269e-02<br />Sepal.Length: 4.680431","density: 2.120433e-02<br />Sepal.Length: 4.673386","density: 2.042119e-02<br />Sepal.Length: 4.666341","density: 1.964523e-02<br />Sepal.Length: 4.659295","density: 1.887542e-02<br />Sepal.Length: 4.652250","density: 1.811508e-02<br />Sepal.Length: 4.645205","density: 1.736754e-02<br />Sepal.Length: 4.638160","density: 1.662944e-02<br />Sepal.Length: 4.631115","density: 1.590335e-02<br />Sepal.Length: 4.624070","density: 1.519563e-02<br />Sepal.Length: 4.617025","density: 1.449992e-02<br />Sepal.Length: 4.609980","density: 1.381724e-02<br />Sepal.Length: 4.602935","density: 1.315814e-02<br />Sepal.Length: 4.595890","density: 1.251291e-02<br />Sepal.Length: 4.588845","density: 1.188165e-02<br />Sepal.Length: 4.581800","density: 1.127617e-02<br />Sepal.Length: 4.574755","density: 1.068692e-02<br />Sepal.Length: 4.567710","density: 1.011275e-02<br />Sepal.Length: 4.560665","density: 9.563500e-03<br />Sepal.Length: 4.553620","density: 9.033297e-03<br />Sepal.Length: 4.546575","density: 8.518633e-03<br />Sepal.Length: 4.539530","density: 8.027041e-03<br />Sepal.Length: 4.532485","density: 7.556771e-03<br />Sepal.Length: 4.525440","density: 7.101947e-03<br />Sepal.Length: 4.518395","density: 6.667651e-03<br />Sepal.Length: 4.511350","density: 6.256322e-03<br />Sepal.Length: 4.504305","density: 5.859891e-03<br />Sepal.Length: 4.497260","density: 5.481053e-03<br />Sepal.Length: 4.490215","density: 5.126148e-03<br />Sepal.Length: 4.483170","density: 4.785240e-03<br />Sepal.Length: 4.476125","density: 4.458853e-03<br />Sepal.Length: 4.469080","density: 4.156696e-03<br />Sepal.Length: 4.462035","density: 3.867383e-03<br />Sepal.Length: 4.454990","density: 3.590828e-03<br />Sepal.Length: 4.447945","density: 3.335692e-03<br />Sepal.Length: 4.440900","density: 3.093333e-03<br />Sepal.Length: 4.433855","density: 2.862413e-03<br />Sepal.Length: 4.426810","density: 2.649105e-03<br />Sepal.Length: 4.419765","density: 2.448660e-03<br />Sepal.Length: 4.412720","density: 2.258273e-03<br />Sepal.Length: 4.405675","density: 2.082008e-03<br />Sepal.Length: 4.398630","density: 1.918310e-03<br />Sepal.Length: 4.391585","density: 1.763292e-03<br />Sepal.Length: 4.384540","density: 1.619309e-03<br />Sepal.Length: 4.377495","density: 1.487281e-03<br />Sepal.Length: 4.370450","density: 1.362613e-03<br />Sepal.Length: 4.363405","density: 1.246333e-03<br />Sepal.Length: 4.356360","density: 1.141159e-03<br />Sepal.Length: 4.349315","density: 1.042120e-03<br />Sepal.Length: 4.342270","density: 9.492676e-04<br />Sepal.Length: 4.335225","density: 8.665086e-04<br />Sepal.Length: 4.328180","density: 7.887797e-04<br />Sepal.Length: 4.321135","density: 7.160120e-04<br />Sepal.Length: 4.314090","density: 6.511291e-04<br />Sepal.Length: 4.307045","density: 5.908572e-04<br />Sepal.Length: 4.300000","density: 5.908572e-04<br />Sepal.Length: 4.300000"],"key":["101","102","103","104","105","106","107","108","109","110","111","112","113","114","115","116","117","118","119","120","121","122","123","124","125","126","127","128","129","130","131","132","133","134","135","136","137","138","139","140","141","142","143","144","145","146","147","148","149","150"],"type":"scatter","mode":"lines","line":{"width":1.88976377952756,"color":"rgba(0,0,0,1)","dash":"solid"},"fill":"toself","fillcolor":"rgba(97,156,255,1)","hoveron":"points","set":"SharedData91833e3a","name":"virginica","legendgroup":"virginica","showlegend":true,"xaxis":"x","yaxis":"y4","hoverinfo":"text","_isSimpleKey":true,"_isNestedKey":false,"frame":null},{"x":[5.1,4.9,4.7,4.6,5,5.4,4.6,5,4.4,4.9,5.4,4.8,4.8,4.3,5.8,5.7,5.4,5.1,5.7,5.1,5.4,5.1,4.6,5.1,4.8,5,5,5.2,5.2,4.7,4.8,5.4,5.2,5.5,4.9,5,5.5,4.9,4.4,5.1,5,4.5,4.4,5,5.1,4.8,5.1,4.6,5.3,5],"y":[3.5,3,3.2,3.1,3.6,3.9,3.4,3.4,2.9,3.1,3.7,3.4,3,3,4,4.4,3.9,3.5,3.8,3.8,3.4,3.7,3.6,3.3,3.4,3,3.4,3.5,3.4,3.2,3.1,3.4,4.1,4.2,3.1,3.2,3.5,3.6,3,3.4,3.5,2.3,3.2,3.5,3.8,3,3.8,3.2,3.7,3.3],"text":["Sepal.Length: 5.1<br />Sepal.Width: 3.5<br />Species: setosa","Sepal.Length: 4.9<br />Sepal.Width: 3.0<br />Species: setosa","Sepal.Length: 4.7<br />Sepal.Width: 3.2<br />Species: setosa","Sepal.Length: 4.6<br />Sepal.Width: 3.1<br />Species: setosa","Sepal.Length: 5.0<br />Sepal.Width: 3.6<br />Species: setosa","Sepal.Length: 5.4<br />Sepal.Width: 3.9<br />Species: setosa","Sepal.Length: 4.6<br />Sepal.Width: 3.4<br />Species: setosa","Sepal.Length: 5.0<br />Sepal.Width: 3.4<br />Species: setosa","Sepal.Length: 4.4<br />Sepal.Width: 2.9<br />Species: setosa","Sepal.Length: 4.9<br />Sepal.Width: 3.1<br />Species: setosa","Sepal.Length: 5.4<br />Sepal.Width: 3.7<br />Species: setosa","Sepal.Length: 4.8<br />Sepal.Width: 3.4<br />Species: setosa","Sepal.Length: 4.8<br />Sepal.Width: 3.0<br />Species: setosa","Sepal.Length: 4.3<br />Sepal.Width: 3.0<br />Species: setosa","Sepal.Length: 5.8<br />Sepal.Width: 4.0<br />Species: setosa","Sepal.Length: 5.7<br />Sepal.Width: 4.4<br />Species: setosa","Sepal.Length: 5.4<br />Sepal.Width: 3.9<br />Species: setosa","Sepal.Length: 5.1<br />Sepal.Width: 3.5<br />Species: setosa","Sepal.Length: 5.7<br />Sepal.Width: 3.8<br />Species: setosa","Sepal.Length: 5.1<br />Sepal.Width: 3.8<br />Species: setosa","Sepal.Length: 5.4<br />Sepal.Width: 3.4<br />Species: setosa","Sepal.Length: 5.1<br />Sepal.Width: 3.7<br />Species: setosa","Sepal.Length: 4.6<br />Sepal.Width: 3.6<br />Species: setosa","Sepal.Length: 5.1<br />Sepal.Width: 3.3<br />Species: setosa","Sepal.Length: 4.8<br />Sepal.Width: 3.4<br />Species: setosa","Sepal.Length: 5.0<br />Sepal.Width: 3.0<br />Species: setosa","Sepal.Length: 5.0<br />Sepal.Width: 3.4<br />Species: setosa","Sepal.Length: 5.2<br />Sepal.Width: 3.5<br />Species: setosa","Sepal.Length: 5.2<br />Sepal.Width: 3.4<br />Species: setosa","Sepal.Length: 4.7<br />Sepal.Width: 3.2<br />Species: setosa","Sepal.Length: 4.8<br />Sepal.Width: 3.1<br />Species: setosa","Sepal.Length: 5.4<br />Sepal.Width: 3.4<br />Species: setosa","Sepal.Length: 5.2<br />Sepal.Width: 4.1<br />Species: setosa","Sepal.Length: 5.5<br />Sepal.Width: 4.2<br />Species: setosa","Sepal.Length: 4.9<br />Sepal.Width: 3.1<br />Species: setosa","Sepal.Length: 5.0<br />Sepal.Width: 3.2<br />Species: setosa","Sepal.Length: 5.5<br />Sepal.Width: 3.5<br />Species: setosa","Sepal.Length: 4.9<br />Sepal.Width: 3.6<br />Species: setosa","Sepal.Length: 4.4<br />Sepal.Width: 3.0<br />Species: setosa","Sepal.Length: 5.1<br />Sepal.Width: 3.4<br />Species: setosa","Sepal.Length: 5.0<br />Sepal.Width: 3.5<br />Species: setosa","Sepal.Length: 4.5<br />Sepal.Width: 2.3<br />Species: setosa","Sepal.Length: 4.4<br />Sepal.Width: 3.2<br />Species: setosa","Sepal.Length: 5.0<br />Sepal.Width: 3.5<br />Species: setosa","Sepal.Length: 5.1<br />Sepal.Width: 3.8<br />Species: setosa","Sepal.Length: 4.8<br />Sepal.Width: 3.0<br />Species: setosa","Sepal.Length: 5.1<br />Sepal.Width: 3.8<br />Species: setosa","Sepal.Length: 4.6<br />Sepal.Width: 3.2<br />Species: setosa","Sepal.Length: 5.3<br />Sepal.Width: 3.7<br />Species: setosa","Sepal.Length: 5.0<br />Sepal.Width: 3.3<br />Species: setosa"],"key":["1","2","3","4","5","6","7","8","9","10","11","12","13","14","15","16","17","18","19","20","21","22","23","24","25","26","27","28","29","30","31","32","33","34","35","36","37","38","39","40","41","42","43","44","45","46","47","48","49","50"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(248,118,109,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(248,118,109,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"setosa","legendgroup":"setosa","showlegend":true,"xaxis":"x","yaxis":"y3","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[7,6.4,6.9,5.5,6.5,5.7,6.3,4.9,6.6,5.2,5,5.9,6,6.1,5.6,6.7,5.6,5.8,6.2,5.6,5.9,6.1,6.3,6.1,6.4,6.6,6.8,6.7,6,5.7,5.5,5.5,5.8,6,5.4,6,6.7,6.3,5.6,5.5,5.5,6.1,5.8,5,5.6,5.7,5.7,6.2,5.1,5.7],"y":[3.2,3.2,3.1,2.3,2.8,2.8,3.3,2.4,2.9,2.7,2,3,2.2,2.9,2.9,3.1,3,2.7,2.2,2.5,3.2,2.8,2.5,2.8,2.9,3,2.8,3,2.9,2.6,2.4,2.4,2.7,2.7,3,3.4,3.1,2.3,3,2.5,2.6,3,2.6,2.3,2.7,3,2.9,2.9,2.5,2.8],"text":["Sepal.Length: 7.0<br />Sepal.Width: 3.2<br />Species: versicolor","Sepal.Length: 6.4<br />Sepal.Width: 3.2<br />Species: versicolor","Sepal.Length: 6.9<br />Sepal.Width: 3.1<br />Species: versicolor","Sepal.Length: 5.5<br />Sepal.Width: 2.3<br />Species: versicolor","Sepal.Length: 6.5<br />Sepal.Width: 2.8<br />Species: versicolor","Sepal.Length: 5.7<br />Sepal.Width: 2.8<br />Species: versicolor","Sepal.Length: 6.3<br />Sepal.Width: 3.3<br />Species: versicolor","Sepal.Length: 4.9<br />Sepal.Width: 2.4<br />Species: versicolor","Sepal.Length: 6.6<br />Sepal.Width: 2.9<br />Species: versicolor","Sepal.Length: 5.2<br />Sepal.Width: 2.7<br />Species: versicolor","Sepal.Length: 5.0<br />Sepal.Width: 2.0<br />Species: versicolor","Sepal.Length: 5.9<br />Sepal.Width: 3.0<br />Species: versicolor","Sepal.Length: 6.0<br />Sepal.Width: 2.2<br />Species: versicolor","Sepal.Length: 6.1<br />Sepal.Width: 2.9<br />Species: versicolor","Sepal.Length: 5.6<br />Sepal.Width: 2.9<br />Species: versicolor","Sepal.Length: 6.7<br />Sepal.Width: 3.1<br />Species: versicolor","Sepal.Length: 5.6<br />Sepal.Width: 3.0<br />Species: versicolor","Sepal.Length: 5.8<br />Sepal.Width: 2.7<br />Species: versicolor","Sepal.Length: 6.2<br />Sepal.Width: 2.2<br />Species: versicolor","Sepal.Length: 5.6<br />Sepal.Width: 2.5<br />Species: versicolor","Sepal.Length: 5.9<br />Sepal.Width: 3.2<br />Species: versicolor","Sepal.Length: 6.1<br />Sepal.Width: 2.8<br />Species: versicolor","Sepal.Length: 6.3<br />Sepal.Width: 2.5<br />Species: versicolor","Sepal.Length: 6.1<br />Sepal.Width: 2.8<br />Species: versicolor","Sepal.Length: 6.4<br />Sepal.Width: 2.9<br />Species: versicolor","Sepal.Length: 6.6<br />Sepal.Width: 3.0<br />Species: versicolor","Sepal.Length: 6.8<br />Sepal.Width: 2.8<br />Species: versicolor","Sepal.Length: 6.7<br />Sepal.Width: 3.0<br />Species: versicolor","Sepal.Length: 6.0<br />Sepal.Width: 2.9<br />Species: versicolor","Sepal.Length: 5.7<br />Sepal.Width: 2.6<br />Species: versicolor","Sepal.Length: 5.5<br />Sepal.Width: 2.4<br />Species: versicolor","Sepal.Length: 5.5<br />Sepal.Width: 2.4<br />Species: versicolor","Sepal.Length: 5.8<br />Sepal.Width: 2.7<br />Species: versicolor","Sepal.Length: 6.0<br />Sepal.Width: 2.7<br />Species: versicolor","Sepal.Length: 5.4<br />Sepal.Width: 3.0<br />Species: versicolor","Sepal.Length: 6.0<br />Sepal.Width: 3.4<br />Species: versicolor","Sepal.Length: 6.7<br />Sepal.Width: 3.1<br />Species: versicolor","Sepal.Length: 6.3<br />Sepal.Width: 2.3<br />Species: versicolor","Sepal.Length: 5.6<br />Sepal.Width: 3.0<br />Species: versicolor","Sepal.Length: 5.5<br />Sepal.Width: 2.5<br />Species: versicolor","Sepal.Length: 5.5<br />Sepal.Width: 2.6<br />Species: versicolor","Sepal.Length: 6.1<br />Sepal.Width: 3.0<br />Species: versicolor","Sepal.Length: 5.8<br />Sepal.Width: 2.6<br />Species: versicolor","Sepal.Length: 5.0<br />Sepal.Width: 2.3<br />Species: versicolor","Sepal.Length: 5.6<br />Sepal.Width: 2.7<br />Species: versicolor","Sepal.Length: 5.7<br />Sepal.Width: 3.0<br />Species: versicolor","Sepal.Length: 5.7<br />Sepal.Width: 2.9<br />Species: versicolor","Sepal.Length: 6.2<br />Sepal.Width: 2.9<br />Species: versicolor","Sepal.Length: 5.1<br />Sepal.Width: 2.5<br />Species: versicolor","Sepal.Length: 5.7<br />Sepal.Width: 2.8<br />Species: versicolor"],"key":["51","52","53","54","55","56","57","58","59","60","61","62","63","64","65","66","67","68","69","70","71","72","73","74","75","76","77","78","79","80","81","82","83","84","85","86","87","88","89","90","91","92","93","94","95","96","97","98","99","100"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(0,186,56,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(0,186,56,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"versicolor","legendgroup":"versicolor","showlegend":true,"xaxis":"x","yaxis":"y3","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[6.3,5.8,7.1,6.3,6.5,7.6,4.9,7.3,6.7,7.2,6.5,6.4,6.8,5.7,5.8,6.4,6.5,7.7,7.7,6,6.9,5.6,7.7,6.3,6.7,7.2,6.2,6.1,6.4,7.2,7.4,7.9,6.4,6.3,6.1,7.7,6.3,6.4,6,6.9,6.7,6.9,5.8,6.8,6.7,6.7,6.3,6.5,6.2,5.9],"y":[3.3,2.7,3,2.9,3,3,2.5,2.9,2.5,3.6,3.2,2.7,3,2.5,2.8,3.2,3,3.8,2.6,2.2,3.2,2.8,2.8,2.7,3.3,3.2,2.8,3,2.8,3,2.8,3.8,2.8,2.8,2.6,3,3.4,3.1,3,3.1,3.1,3.1,2.7,3.2,3.3,3,2.5,3,3.4,3],"text":["Sepal.Length: 6.3<br />Sepal.Width: 3.3<br />Species: virginica","Sepal.Length: 5.8<br />Sepal.Width: 2.7<br />Species: virginica","Sepal.Length: 7.1<br />Sepal.Width: 3.0<br />Species: virginica","Sepal.Length: 6.3<br />Sepal.Width: 2.9<br />Species: virginica","Sepal.Length: 6.5<br />Sepal.Width: 3.0<br />Species: virginica","Sepal.Length: 7.6<br />Sepal.Width: 3.0<br />Species: virginica","Sepal.Length: 4.9<br />Sepal.Width: 2.5<br />Species: virginica","Sepal.Length: 7.3<br />Sepal.Width: 2.9<br />Species: virginica","Sepal.Length: 6.7<br />Sepal.Width: 2.5<br />Species: virginica","Sepal.Length: 7.2<br />Sepal.Width: 3.6<br />Species: virginica","Sepal.Length: 6.5<br />Sepal.Width: 3.2<br />Species: virginica","Sepal.Length: 6.4<br />Sepal.Width: 2.7<br />Species: virginica","Sepal.Length: 6.8<br />Sepal.Width: 3.0<br />Species: virginica","Sepal.Length: 5.7<br />Sepal.Width: 2.5<br />Species: virginica","Sepal.Length: 5.8<br />Sepal.Width: 2.8<br />Species: virginica","Sepal.Length: 6.4<br />Sepal.Width: 3.2<br />Species: virginica","Sepal.Length: 6.5<br />Sepal.Width: 3.0<br />Species: virginica","Sepal.Length: 7.7<br />Sepal.Width: 3.8<br />Species: virginica","Sepal.Length: 7.7<br />Sepal.Width: 2.6<br />Species: virginica","Sepal.Length: 6.0<br />Sepal.Width: 2.2<br />Species: virginica","Sepal.Length: 6.9<br />Sepal.Width: 3.2<br />Species: virginica","Sepal.Length: 5.6<br />Sepal.Width: 2.8<br />Species: virginica","Sepal.Length: 7.7<br />Sepal.Width: 2.8<br />Species: virginica","Sepal.Length: 6.3<br />Sepal.Width: 2.7<br />Species: virginica","Sepal.Length: 6.7<br />Sepal.Width: 3.3<br />Species: virginica","Sepal.Length: 7.2<br />Sepal.Width: 3.2<br />Species: virginica","Sepal.Length: 6.2<br />Sepal.Width: 2.8<br />Species: virginica","Sepal.Length: 6.1<br />Sepal.Width: 3.0<br />Species: virginica","Sepal.Length: 6.4<br />Sepal.Width: 2.8<br />Species: virginica","Sepal.Length: 7.2<br />Sepal.Width: 3.0<br />Species: virginica","Sepal.Length: 7.4<br />Sepal.Width: 2.8<br />Species: virginica","Sepal.Length: 7.9<br />Sepal.Width: 3.8<br />Species: virginica","Sepal.Length: 6.4<br />Sepal.Width: 2.8<br />Species: virginica","Sepal.Length: 6.3<br />Sepal.Width: 2.8<br />Species: virginica","Sepal.Length: 6.1<br />Sepal.Width: 2.6<br />Species: virginica","Sepal.Length: 7.7<br />Sepal.Width: 3.0<br />Species: virginica","Sepal.Length: 6.3<br />Sepal.Width: 3.4<br />Species: virginica","Sepal.Length: 6.4<br />Sepal.Width: 3.1<br />Species: virginica","Sepal.Length: 6.0<br />Sepal.Width: 3.0<br />Species: virginica","Sepal.Length: 6.9<br />Sepal.Width: 3.1<br />Species: virginica","Sepal.Length: 6.7<br />Sepal.Width: 3.1<br />Species: virginica","Sepal.Length: 6.9<br />Sepal.Width: 3.1<br />Species: virginica","Sepal.Length: 5.8<br />Sepal.Width: 2.7<br />Species: virginica","Sepal.Length: 6.8<br />Sepal.Width: 3.2<br />Species: virginica","Sepal.Length: 6.7<br />Sepal.Width: 3.3<br />Species: virginica","Sepal.Length: 6.7<br />Sepal.Width: 3.0<br />Species: virginica","Sepal.Length: 6.3<br />Sepal.Width: 2.5<br />Species: virginica","Sepal.Length: 6.5<br />Sepal.Width: 3.0<br />Species: virginica","Sepal.Length: 6.2<br />Sepal.Width: 3.4<br />Species: virginica","Sepal.Length: 5.9<br />Sepal.Width: 3.0<br />Species: virginica"],"key":["101","102","103","104","105","106","107","108","109","110","111","112","113","114","115","116","117","118","119","120","121","122","123","124","125","126","127","128","129","130","131","132","133","134","135","136","137","138","139","140","141","142","143","144","145","146","147","148","149","150"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(97,156,255,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(97,156,255,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"virginica","legendgroup":"virginica","showlegend":true,"xaxis":"x","yaxis":"y3","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[5.1,4.9,4.7,4.6,5,5.4,4.6,5,4.4,4.9,5.4,4.8,4.8,4.3,5.8,5.7,5.4,5.1,5.7,5.1,5.4,5.1,4.6,5.1,4.8,5,5,5.2,5.2,4.7,4.8,5.4,5.2,5.5,4.9,5,5.5,4.9,4.4,5.1,5,4.5,4.4,5,5.1,4.8,5.1,4.6,5.3,5],"y":[1.4,1.4,1.3,1.5,1.4,1.7,1.4,1.5,1.4,1.5,1.5,1.6,1.4,1.1,1.2,1.5,1.3,1.4,1.7,1.5,1.7,1.5,1,1.7,1.9,1.6,1.6,1.5,1.4,1.6,1.6,1.5,1.5,1.4,1.5,1.2,1.3,1.4,1.3,1.5,1.3,1.3,1.3,1.6,1.9,1.4,1.6,1.4,1.5,1.4],"text":["Sepal.Length: 5.1<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Length: 4.9<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Length: 4.7<br />Petal.Length: 1.3<br />Species: setosa","Sepal.Length: 4.6<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Length: 5.0<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Length: 5.4<br />Petal.Length: 1.7<br />Species: setosa","Sepal.Length: 4.6<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Length: 5.0<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Length: 4.4<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Length: 4.9<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Length: 5.4<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Length: 4.8<br />Petal.Length: 1.6<br />Species: setosa","Sepal.Length: 4.8<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Length: 4.3<br />Petal.Length: 1.1<br />Species: setosa","Sepal.Length: 5.8<br />Petal.Length: 1.2<br />Species: setosa","Sepal.Length: 5.7<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Length: 5.4<br />Petal.Length: 1.3<br />Species: setosa","Sepal.Length: 5.1<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Length: 5.7<br />Petal.Length: 1.7<br />Species: setosa","Sepal.Length: 5.1<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Length: 5.4<br />Petal.Length: 1.7<br />Species: setosa","Sepal.Length: 5.1<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Length: 4.6<br />Petal.Length: 1.0<br />Species: setosa","Sepal.Length: 5.1<br />Petal.Length: 1.7<br />Species: setosa","Sepal.Length: 4.8<br />Petal.Length: 1.9<br />Species: setosa","Sepal.Length: 5.0<br />Petal.Length: 1.6<br />Species: setosa","Sepal.Length: 5.0<br />Petal.Length: 1.6<br />Species: setosa","Sepal.Length: 5.2<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Length: 5.2<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Length: 4.7<br />Petal.Length: 1.6<br />Species: setosa","Sepal.Length: 4.8<br />Petal.Length: 1.6<br />Species: setosa","Sepal.Length: 5.4<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Length: 5.2<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Length: 5.5<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Length: 4.9<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Length: 5.0<br />Petal.Length: 1.2<br />Species: setosa","Sepal.Length: 5.5<br />Petal.Length: 1.3<br />Species: setosa","Sepal.Length: 4.9<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Length: 4.4<br />Petal.Length: 1.3<br />Species: setosa","Sepal.Length: 5.1<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Length: 5.0<br />Petal.Length: 1.3<br />Species: setosa","Sepal.Length: 4.5<br />Petal.Length: 1.3<br />Species: setosa","Sepal.Length: 4.4<br />Petal.Length: 1.3<br />Species: setosa","Sepal.Length: 5.0<br />Petal.Length: 1.6<br />Species: setosa","Sepal.Length: 5.1<br />Petal.Length: 1.9<br />Species: setosa","Sepal.Length: 4.8<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Length: 5.1<br />Petal.Length: 1.6<br />Species: setosa","Sepal.Length: 4.6<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Length: 5.3<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Length: 5.0<br />Petal.Length: 1.4<br />Species: setosa"],"key":["1","2","3","4","5","6","7","8","9","10","11","12","13","14","15","16","17","18","19","20","21","22","23","24","25","26","27","28","29","30","31","32","33","34","35","36","37","38","39","40","41","42","43","44","45","46","47","48","49","50"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(248,118,109,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(248,118,109,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"setosa","legendgroup":"setosa","showlegend":true,"xaxis":"x","yaxis":"y2","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[7,6.4,6.9,5.5,6.5,5.7,6.3,4.9,6.6,5.2,5,5.9,6,6.1,5.6,6.7,5.6,5.8,6.2,5.6,5.9,6.1,6.3,6.1,6.4,6.6,6.8,6.7,6,5.7,5.5,5.5,5.8,6,5.4,6,6.7,6.3,5.6,5.5,5.5,6.1,5.8,5,5.6,5.7,5.7,6.2,5.1,5.7],"y":[4.7,4.5,4.9,4,4.6,4.5,4.7,3.3,4.6,3.9,3.5,4.2,4,4.7,3.6,4.4,4.5,4.1,4.5,3.9,4.8,4,4.9,4.7,4.3,4.4,4.8,5,4.5,3.5,3.8,3.7,3.9,5.1,4.5,4.5,4.7,4.4,4.1,4,4.4,4.6,4,3.3,4.2,4.2,4.2,4.3,3,4.1],"text":["Sepal.Length: 7.0<br />Petal.Length: 4.7<br />Species: versicolor","Sepal.Length: 6.4<br />Petal.Length: 4.5<br />Species: versicolor","Sepal.Length: 6.9<br />Petal.Length: 4.9<br />Species: versicolor","Sepal.Length: 5.5<br />Petal.Length: 4.0<br />Species: versicolor","Sepal.Length: 6.5<br />Petal.Length: 4.6<br />Species: versicolor","Sepal.Length: 5.7<br />Petal.Length: 4.5<br />Species: versicolor","Sepal.Length: 6.3<br />Petal.Length: 4.7<br />Species: versicolor","Sepal.Length: 4.9<br />Petal.Length: 3.3<br />Species: versicolor","Sepal.Length: 6.6<br />Petal.Length: 4.6<br />Species: versicolor","Sepal.Length: 5.2<br />Petal.Length: 3.9<br />Species: versicolor","Sepal.Length: 5.0<br />Petal.Length: 3.5<br />Species: versicolor","Sepal.Length: 5.9<br />Petal.Length: 4.2<br />Species: versicolor","Sepal.Length: 6.0<br />Petal.Length: 4.0<br />Species: versicolor","Sepal.Length: 6.1<br />Petal.Length: 4.7<br />Species: versicolor","Sepal.Length: 5.6<br />Petal.Length: 3.6<br />Species: versicolor","Sepal.Length: 6.7<br />Petal.Length: 4.4<br />Species: versicolor","Sepal.Length: 5.6<br />Petal.Length: 4.5<br />Species: versicolor","Sepal.Length: 5.8<br />Petal.Length: 4.1<br />Species: versicolor","Sepal.Length: 6.2<br />Petal.Length: 4.5<br />Species: versicolor","Sepal.Length: 5.6<br />Petal.Length: 3.9<br />Species: versicolor","Sepal.Length: 5.9<br />Petal.Length: 4.8<br />Species: versicolor","Sepal.Length: 6.1<br />Petal.Length: 4.0<br />Species: versicolor","Sepal.Length: 6.3<br />Petal.Length: 4.9<br />Species: versicolor","Sepal.Length: 6.1<br />Petal.Length: 4.7<br />Species: versicolor","Sepal.Length: 6.4<br />Petal.Length: 4.3<br />Species: versicolor","Sepal.Length: 6.6<br />Petal.Length: 4.4<br />Species: versicolor","Sepal.Length: 6.8<br />Petal.Length: 4.8<br />Species: versicolor","Sepal.Length: 6.7<br />Petal.Length: 5.0<br />Species: versicolor","Sepal.Length: 6.0<br />Petal.Length: 4.5<br />Species: versicolor","Sepal.Length: 5.7<br />Petal.Length: 3.5<br />Species: versicolor","Sepal.Length: 5.5<br />Petal.Length: 3.8<br />Species: versicolor","Sepal.Length: 5.5<br />Petal.Length: 3.7<br />Species: versicolor","Sepal.Length: 5.8<br />Petal.Length: 3.9<br />Species: versicolor","Sepal.Length: 6.0<br />Petal.Length: 5.1<br />Species: versicolor","Sepal.Length: 5.4<br />Petal.Length: 4.5<br />Species: versicolor","Sepal.Length: 6.0<br />Petal.Length: 4.5<br />Species: versicolor","Sepal.Length: 6.7<br />Petal.Length: 4.7<br />Species: versicolor","Sepal.Length: 6.3<br />Petal.Length: 4.4<br />Species: versicolor","Sepal.Length: 5.6<br />Petal.Length: 4.1<br />Species: versicolor","Sepal.Length: 5.5<br />Petal.Length: 4.0<br />Species: versicolor","Sepal.Length: 5.5<br />Petal.Length: 4.4<br />Species: versicolor","Sepal.Length: 6.1<br />Petal.Length: 4.6<br />Species: versicolor","Sepal.Length: 5.8<br />Petal.Length: 4.0<br />Species: versicolor","Sepal.Length: 5.0<br />Petal.Length: 3.3<br />Species: versicolor","Sepal.Length: 5.6<br />Petal.Length: 4.2<br />Species: versicolor","Sepal.Length: 5.7<br />Petal.Length: 4.2<br />Species: versicolor","Sepal.Length: 5.7<br />Petal.Length: 4.2<br />Species: versicolor","Sepal.Length: 6.2<br />Petal.Length: 4.3<br />Species: versicolor","Sepal.Length: 5.1<br />Petal.Length: 3.0<br />Species: versicolor","Sepal.Length: 5.7<br />Petal.Length: 4.1<br />Species: versicolor"],"key":["51","52","53","54","55","56","57","58","59","60","61","62","63","64","65","66","67","68","69","70","71","72","73","74","75","76","77","78","79","80","81","82","83","84","85","86","87","88","89","90","91","92","93","94","95","96","97","98","99","100"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(0,186,56,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(0,186,56,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"versicolor","legendgroup":"versicolor","showlegend":true,"xaxis":"x","yaxis":"y2","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[6.3,5.8,7.1,6.3,6.5,7.6,4.9,7.3,6.7,7.2,6.5,6.4,6.8,5.7,5.8,6.4,6.5,7.7,7.7,6,6.9,5.6,7.7,6.3,6.7,7.2,6.2,6.1,6.4,7.2,7.4,7.9,6.4,6.3,6.1,7.7,6.3,6.4,6,6.9,6.7,6.9,5.8,6.8,6.7,6.7,6.3,6.5,6.2,5.9],"y":[6,5.1,5.9,5.6,5.8,6.6,4.5,6.3,5.8,6.1,5.1,5.3,5.5,5,5.1,5.3,5.5,6.7,6.9,5,5.7,4.9,6.7,4.9,5.7,6,4.8,4.9,5.6,5.8,6.1,6.4,5.6,5.1,5.6,6.1,5.6,5.5,4.8,5.4,5.6,5.1,5.1,5.9,5.7,5.2,5,5.2,5.4,5.1],"text":["Sepal.Length: 6.3<br />Petal.Length: 6.0<br />Species: virginica","Sepal.Length: 5.8<br />Petal.Length: 5.1<br />Species: virginica","Sepal.Length: 7.1<br />Petal.Length: 5.9<br />Species: virginica","Sepal.Length: 6.3<br />Petal.Length: 5.6<br />Species: virginica","Sepal.Length: 6.5<br />Petal.Length: 5.8<br />Species: virginica","Sepal.Length: 7.6<br />Petal.Length: 6.6<br />Species: virginica","Sepal.Length: 4.9<br />Petal.Length: 4.5<br />Species: virginica","Sepal.Length: 7.3<br />Petal.Length: 6.3<br />Species: virginica","Sepal.Length: 6.7<br />Petal.Length: 5.8<br />Species: virginica","Sepal.Length: 7.2<br />Petal.Length: 6.1<br />Species: virginica","Sepal.Length: 6.5<br />Petal.Length: 5.1<br />Species: virginica","Sepal.Length: 6.4<br />Petal.Length: 5.3<br />Species: virginica","Sepal.Length: 6.8<br />Petal.Length: 5.5<br />Species: virginica","Sepal.Length: 5.7<br />Petal.Length: 5.0<br />Species: virginica","Sepal.Length: 5.8<br />Petal.Length: 5.1<br />Species: virginica","Sepal.Length: 6.4<br />Petal.Length: 5.3<br />Species: virginica","Sepal.Length: 6.5<br />Petal.Length: 5.5<br />Species: virginica","Sepal.Length: 7.7<br />Petal.Length: 6.7<br />Species: virginica","Sepal.Length: 7.7<br />Petal.Length: 6.9<br />Species: virginica","Sepal.Length: 6.0<br />Petal.Length: 5.0<br />Species: virginica","Sepal.Length: 6.9<br />Petal.Length: 5.7<br />Species: virginica","Sepal.Length: 5.6<br />Petal.Length: 4.9<br />Species: virginica","Sepal.Length: 7.7<br />Petal.Length: 6.7<br />Species: virginica","Sepal.Length: 6.3<br />Petal.Length: 4.9<br />Species: virginica","Sepal.Length: 6.7<br />Petal.Length: 5.7<br />Species: virginica","Sepal.Length: 7.2<br />Petal.Length: 6.0<br />Species: virginica","Sepal.Length: 6.2<br />Petal.Length: 4.8<br />Species: virginica","Sepal.Length: 6.1<br />Petal.Length: 4.9<br />Species: virginica","Sepal.Length: 6.4<br />Petal.Length: 5.6<br />Species: virginica","Sepal.Length: 7.2<br />Petal.Length: 5.8<br />Species: virginica","Sepal.Length: 7.4<br />Petal.Length: 6.1<br />Species: virginica","Sepal.Length: 7.9<br />Petal.Length: 6.4<br />Species: virginica","Sepal.Length: 6.4<br />Petal.Length: 5.6<br />Species: virginica","Sepal.Length: 6.3<br />Petal.Length: 5.1<br />Species: virginica","Sepal.Length: 6.1<br />Petal.Length: 5.6<br />Species: virginica","Sepal.Length: 7.7<br />Petal.Length: 6.1<br />Species: virginica","Sepal.Length: 6.3<br />Petal.Length: 5.6<br />Species: virginica","Sepal.Length: 6.4<br />Petal.Length: 5.5<br />Species: virginica","Sepal.Length: 6.0<br />Petal.Length: 4.8<br />Species: virginica","Sepal.Length: 6.9<br />Petal.Length: 5.4<br />Species: virginica","Sepal.Length: 6.7<br />Petal.Length: 5.6<br />Species: virginica","Sepal.Length: 6.9<br />Petal.Length: 5.1<br />Species: virginica","Sepal.Length: 5.8<br />Petal.Length: 5.1<br />Species: virginica","Sepal.Length: 6.8<br />Petal.Length: 5.9<br />Species: virginica","Sepal.Length: 6.7<br />Petal.Length: 5.7<br />Species: virginica","Sepal.Length: 6.7<br />Petal.Length: 5.2<br />Species: virginica","Sepal.Length: 6.3<br />Petal.Length: 5.0<br />Species: virginica","Sepal.Length: 6.5<br />Petal.Length: 5.2<br />Species: virginica","Sepal.Length: 6.2<br />Petal.Length: 5.4<br />Species: virginica","Sepal.Length: 5.9<br />Petal.Length: 5.1<br />Species: virginica"],"key":["101","102","103","104","105","106","107","108","109","110","111","112","113","114","115","116","117","118","119","120","121","122","123","124","125","126","127","128","129","130","131","132","133","134","135","136","137","138","139","140","141","142","143","144","145","146","147","148","149","150"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(97,156,255,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(97,156,255,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"virginica","legendgroup":"virginica","showlegend":true,"xaxis":"x","yaxis":"y2","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[5.1,4.9,4.7,4.6,5,5.4,4.6,5,4.4,4.9,5.4,4.8,4.8,4.3,5.8,5.7,5.4,5.1,5.7,5.1,5.4,5.1,4.6,5.1,4.8,5,5,5.2,5.2,4.7,4.8,5.4,5.2,5.5,4.9,5,5.5,4.9,4.4,5.1,5,4.5,4.4,5,5.1,4.8,5.1,4.6,5.3,5],"y":[0.2,0.2,0.2,0.2,0.2,0.4,0.3,0.2,0.2,0.1,0.2,0.2,0.1,0.1,0.2,0.4,0.4,0.3,0.3,0.3,0.2,0.4,0.2,0.5,0.2,0.2,0.4,0.2,0.2,0.2,0.2,0.4,0.1,0.2,0.2,0.2,0.2,0.1,0.2,0.2,0.3,0.3,0.2,0.6,0.4,0.3,0.2,0.2,0.2,0.2],"text":["Sepal.Length: 5.1<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 4.9<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 4.7<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 4.6<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 5.0<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 5.4<br />Petal.Width: 0.4<br />Species: setosa","Sepal.Length: 4.6<br />Petal.Width: 0.3<br />Species: setosa","Sepal.Length: 5.0<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 4.4<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 4.9<br />Petal.Width: 0.1<br />Species: setosa","Sepal.Length: 5.4<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 4.8<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 4.8<br />Petal.Width: 0.1<br />Species: setosa","Sepal.Length: 4.3<br />Petal.Width: 0.1<br />Species: setosa","Sepal.Length: 5.8<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 5.7<br />Petal.Width: 0.4<br />Species: setosa","Sepal.Length: 5.4<br />Petal.Width: 0.4<br />Species: setosa","Sepal.Length: 5.1<br />Petal.Width: 0.3<br />Species: setosa","Sepal.Length: 5.7<br />Petal.Width: 0.3<br />Species: setosa","Sepal.Length: 5.1<br />Petal.Width: 0.3<br />Species: setosa","Sepal.Length: 5.4<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 5.1<br />Petal.Width: 0.4<br />Species: setosa","Sepal.Length: 4.6<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 5.1<br />Petal.Width: 0.5<br />Species: setosa","Sepal.Length: 4.8<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 5.0<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 5.0<br />Petal.Width: 0.4<br />Species: setosa","Sepal.Length: 5.2<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 5.2<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 4.7<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 4.8<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 5.4<br />Petal.Width: 0.4<br />Species: setosa","Sepal.Length: 5.2<br />Petal.Width: 0.1<br />Species: setosa","Sepal.Length: 5.5<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 4.9<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 5.0<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 5.5<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 4.9<br />Petal.Width: 0.1<br />Species: setosa","Sepal.Length: 4.4<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 5.1<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 5.0<br />Petal.Width: 0.3<br />Species: setosa","Sepal.Length: 4.5<br />Petal.Width: 0.3<br />Species: setosa","Sepal.Length: 4.4<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 5.0<br />Petal.Width: 0.6<br />Species: setosa","Sepal.Length: 5.1<br />Petal.Width: 0.4<br />Species: setosa","Sepal.Length: 4.8<br />Petal.Width: 0.3<br />Species: setosa","Sepal.Length: 5.1<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 4.6<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 5.3<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Length: 5.0<br />Petal.Width: 0.2<br />Species: setosa"],"key":["1","2","3","4","5","6","7","8","9","10","11","12","13","14","15","16","17","18","19","20","21","22","23","24","25","26","27","28","29","30","31","32","33","34","35","36","37","38","39","40","41","42","43","44","45","46","47","48","49","50"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(248,118,109,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(248,118,109,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"setosa","legendgroup":"setosa","showlegend":true,"xaxis":"x","yaxis":"y","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[7,6.4,6.9,5.5,6.5,5.7,6.3,4.9,6.6,5.2,5,5.9,6,6.1,5.6,6.7,5.6,5.8,6.2,5.6,5.9,6.1,6.3,6.1,6.4,6.6,6.8,6.7,6,5.7,5.5,5.5,5.8,6,5.4,6,6.7,6.3,5.6,5.5,5.5,6.1,5.8,5,5.6,5.7,5.7,6.2,5.1,5.7],"y":[1.4,1.5,1.5,1.3,1.5,1.3,1.6,1,1.3,1.4,1,1.5,1,1.4,1.3,1.4,1.5,1,1.5,1.1,1.8,1.3,1.5,1.2,1.3,1.4,1.4,1.7,1.5,1,1.1,1,1.2,1.6,1.5,1.6,1.5,1.3,1.3,1.3,1.2,1.4,1.2,1,1.3,1.2,1.3,1.3,1.1,1.3],"text":["Sepal.Length: 7.0<br />Petal.Width: 1.4<br />Species: versicolor","Sepal.Length: 6.4<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Length: 6.9<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Length: 5.5<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Length: 6.5<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Length: 5.7<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Length: 6.3<br />Petal.Width: 1.6<br />Species: versicolor","Sepal.Length: 4.9<br />Petal.Width: 1.0<br />Species: versicolor","Sepal.Length: 6.6<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Length: 5.2<br />Petal.Width: 1.4<br />Species: versicolor","Sepal.Length: 5.0<br />Petal.Width: 1.0<br />Species: versicolor","Sepal.Length: 5.9<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Length: 6.0<br />Petal.Width: 1.0<br />Species: versicolor","Sepal.Length: 6.1<br />Petal.Width: 1.4<br />Species: versicolor","Sepal.Length: 5.6<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Length: 6.7<br />Petal.Width: 1.4<br />Species: versicolor","Sepal.Length: 5.6<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Length: 5.8<br />Petal.Width: 1.0<br />Species: versicolor","Sepal.Length: 6.2<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Length: 5.6<br />Petal.Width: 1.1<br />Species: versicolor","Sepal.Length: 5.9<br />Petal.Width: 1.8<br />Species: versicolor","Sepal.Length: 6.1<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Length: 6.3<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Length: 6.1<br />Petal.Width: 1.2<br />Species: versicolor","Sepal.Length: 6.4<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Length: 6.6<br />Petal.Width: 1.4<br />Species: versicolor","Sepal.Length: 6.8<br />Petal.Width: 1.4<br />Species: versicolor","Sepal.Length: 6.7<br />Petal.Width: 1.7<br />Species: versicolor","Sepal.Length: 6.0<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Length: 5.7<br />Petal.Width: 1.0<br />Species: versicolor","Sepal.Length: 5.5<br />Petal.Width: 1.1<br />Species: versicolor","Sepal.Length: 5.5<br />Petal.Width: 1.0<br />Species: versicolor","Sepal.Length: 5.8<br />Petal.Width: 1.2<br />Species: versicolor","Sepal.Length: 6.0<br />Petal.Width: 1.6<br />Species: versicolor","Sepal.Length: 5.4<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Length: 6.0<br />Petal.Width: 1.6<br />Species: versicolor","Sepal.Length: 6.7<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Length: 6.3<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Length: 5.6<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Length: 5.5<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Length: 5.5<br />Petal.Width: 1.2<br />Species: versicolor","Sepal.Length: 6.1<br />Petal.Width: 1.4<br />Species: versicolor","Sepal.Length: 5.8<br />Petal.Width: 1.2<br />Species: versicolor","Sepal.Length: 5.0<br />Petal.Width: 1.0<br />Species: versicolor","Sepal.Length: 5.6<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Length: 5.7<br />Petal.Width: 1.2<br />Species: versicolor","Sepal.Length: 5.7<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Length: 6.2<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Length: 5.1<br />Petal.Width: 1.1<br />Species: versicolor","Sepal.Length: 5.7<br />Petal.Width: 1.3<br />Species: versicolor"],"key":["51","52","53","54","55","56","57","58","59","60","61","62","63","64","65","66","67","68","69","70","71","72","73","74","75","76","77","78","79","80","81","82","83","84","85","86","87","88","89","90","91","92","93","94","95","96","97","98","99","100"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(0,186,56,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(0,186,56,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"versicolor","legendgroup":"versicolor","showlegend":true,"xaxis":"x","yaxis":"y","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[6.3,5.8,7.1,6.3,6.5,7.6,4.9,7.3,6.7,7.2,6.5,6.4,6.8,5.7,5.8,6.4,6.5,7.7,7.7,6,6.9,5.6,7.7,6.3,6.7,7.2,6.2,6.1,6.4,7.2,7.4,7.9,6.4,6.3,6.1,7.7,6.3,6.4,6,6.9,6.7,6.9,5.8,6.8,6.7,6.7,6.3,6.5,6.2,5.9],"y":[2.5,1.9,2.1,1.8,2.2,2.1,1.7,1.8,1.8,2.5,2,1.9,2.1,2,2.4,2.3,1.8,2.2,2.3,1.5,2.3,2,2,1.8,2.1,1.8,1.8,1.8,2.1,1.6,1.9,2,2.2,1.5,1.4,2.3,2.4,1.8,1.8,2.1,2.4,2.3,1.9,2.3,2.5,2.3,1.9,2,2.3,1.8],"text":["Sepal.Length: 6.3<br />Petal.Width: 2.5<br />Species: virginica","Sepal.Length: 5.8<br />Petal.Width: 1.9<br />Species: virginica","Sepal.Length: 7.1<br />Petal.Width: 2.1<br />Species: virginica","Sepal.Length: 6.3<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Length: 6.5<br />Petal.Width: 2.2<br />Species: virginica","Sepal.Length: 7.6<br />Petal.Width: 2.1<br />Species: virginica","Sepal.Length: 4.9<br />Petal.Width: 1.7<br />Species: virginica","Sepal.Length: 7.3<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Length: 6.7<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Length: 7.2<br />Petal.Width: 2.5<br />Species: virginica","Sepal.Length: 6.5<br />Petal.Width: 2.0<br />Species: virginica","Sepal.Length: 6.4<br />Petal.Width: 1.9<br />Species: virginica","Sepal.Length: 6.8<br />Petal.Width: 2.1<br />Species: virginica","Sepal.Length: 5.7<br />Petal.Width: 2.0<br />Species: virginica","Sepal.Length: 5.8<br />Petal.Width: 2.4<br />Species: virginica","Sepal.Length: 6.4<br />Petal.Width: 2.3<br />Species: virginica","Sepal.Length: 6.5<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Length: 7.7<br />Petal.Width: 2.2<br />Species: virginica","Sepal.Length: 7.7<br />Petal.Width: 2.3<br />Species: virginica","Sepal.Length: 6.0<br />Petal.Width: 1.5<br />Species: virginica","Sepal.Length: 6.9<br />Petal.Width: 2.3<br />Species: virginica","Sepal.Length: 5.6<br />Petal.Width: 2.0<br />Species: virginica","Sepal.Length: 7.7<br />Petal.Width: 2.0<br />Species: virginica","Sepal.Length: 6.3<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Length: 6.7<br />Petal.Width: 2.1<br />Species: virginica","Sepal.Length: 7.2<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Length: 6.2<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Length: 6.1<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Length: 6.4<br />Petal.Width: 2.1<br />Species: virginica","Sepal.Length: 7.2<br />Petal.Width: 1.6<br />Species: virginica","Sepal.Length: 7.4<br />Petal.Width: 1.9<br />Species: virginica","Sepal.Length: 7.9<br />Petal.Width: 2.0<br />Species: virginica","Sepal.Length: 6.4<br />Petal.Width: 2.2<br />Species: virginica","Sepal.Length: 6.3<br />Petal.Width: 1.5<br />Species: virginica","Sepal.Length: 6.1<br />Petal.Width: 1.4<br />Species: virginica","Sepal.Length: 7.7<br />Petal.Width: 2.3<br />Species: virginica","Sepal.Length: 6.3<br />Petal.Width: 2.4<br />Species: virginica","Sepal.Length: 6.4<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Length: 6.0<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Length: 6.9<br />Petal.Width: 2.1<br />Species: virginica","Sepal.Length: 6.7<br />Petal.Width: 2.4<br />Species: virginica","Sepal.Length: 6.9<br />Petal.Width: 2.3<br />Species: virginica","Sepal.Length: 5.8<br />Petal.Width: 1.9<br />Species: virginica","Sepal.Length: 6.8<br />Petal.Width: 2.3<br />Species: virginica","Sepal.Length: 6.7<br />Petal.Width: 2.5<br />Species: virginica","Sepal.Length: 6.7<br />Petal.Width: 2.3<br />Species: virginica","Sepal.Length: 6.3<br />Petal.Width: 1.9<br />Species: virginica","Sepal.Length: 6.5<br />Petal.Width: 2.0<br />Species: virginica","Sepal.Length: 6.2<br />Petal.Width: 2.3<br />Species: virginica","Sepal.Length: 5.9<br />Petal.Width: 1.8<br />Species: virginica"],"key":["101","102","103","104","105","106","107","108","109","110","111","112","113","114","115","116","117","118","119","120","121","122","123","124","125","126","127","128","129","130","131","132","133","134","135","136","137","138","139","140","141","142","143","144","145","146","147","148","149","150"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(97,156,255,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(97,156,255,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"virginica","legendgroup":"virginica","showlegend":true,"xaxis":"x","yaxis":"y","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[3.2],"y":[7.5688],"text":"Corr: -0.118","hovertext":"x: 3.2<br />y: 7.5688","textfont":{"size":14.6645669291339,"color":"rgba(118,118,118,1)"},"type":"scatter","mode":"text","hoveron":"points","showlegend":false,"xaxis":"x2","yaxis":"y8","hoverinfo":"text","frame":null},{"x":[3.2],"y":[6.712],"text":" setosa: 0.743***","hovertext":"xPos: 3.2<br />yPos: 6.7120<br />labelp: setosa: 0.743***","textfont":{"size":14.6645669291339,"color":"rgba(248,118,109,1)"},"type":"scatter","mode":"text","hoveron":"points","name":" setosa: 0.743***","legendgroup":" setosa: 0.743***","showlegend":true,"xaxis":"x2","yaxis":"y8","hoverinfo":"text","frame":null},{"x":[3.2],"y":[5.8552],"text":"versicolor: 0.526***","hovertext":"xPos: 3.2<br />yPos: 5.8552<br />labelp: versicolor: 0.526***","textfont":{"size":14.6645669291339,"color":"rgba(0,186,56,1)"},"type":"scatter","mode":"text","hoveron":"points","name":"versicolor: 0.526***","legendgroup":"versicolor: 0.526***","showlegend":true,"xaxis":"x2","yaxis":"y8","hoverinfo":"text","frame":null},{"x":[3.2],"y":[4.9984],"text":" virginica: 0.457***","hovertext":"xPos: 3.2<br />yPos: 4.9984<br />labelp: virginica: 0.457***","textfont":{"size":14.6645669291339,"color":"rgba(97,156,255,1)"},"type":"scatter","mode":"text","hoveron":"points","name":" virginica: 0.457***","legendgroup":" virginica: 0.457***","showlegend":true,"xaxis":"x2","yaxis":"y8","hoverinfo":"text","frame":null},{"x":[2,2.00469667318982,2.00939334637965,2.01409001956947,2.0187866927593,2.02348336594912,2.02818003913894,2.03287671232877,2.03757338551859,2.04227005870841,2.04696673189824,2.05166340508806,2.05636007827789,2.06105675146771,2.06575342465753,2.07045009784736,2.07514677103718,2.07984344422701,2.08454011741683,2.08923679060665,2.09393346379648,2.0986301369863,2.10332681017613,2.10802348336595,2.11272015655577,2.1174168297456,2.12211350293542,2.12681017612524,2.13150684931507,2.13620352250489,2.14090019569472,2.14559686888454,2.15029354207436,2.15499021526419,2.15968688845401,2.16438356164384,2.16908023483366,2.17377690802348,2.17847358121331,2.18317025440313,2.18786692759295,2.19256360078278,2.1972602739726,2.20195694716243,2.20665362035225,2.21135029354207,2.2160469667319,2.22074363992172,2.22544031311155,2.23013698630137,2.23483365949119,2.23953033268102,2.24422700587084,2.24892367906067,2.25362035225049,2.25831702544031,2.26301369863014,2.26771037181996,2.27240704500978,2.27710371819961,2.28180039138943,2.28649706457926,2.29119373776908,2.2958904109589,2.30058708414873,2.30528375733855,2.30998043052838,2.3146771037182,2.31937377690802,2.32407045009785,2.32876712328767,2.3334637964775,2.33816046966732,2.34285714285714,2.34755381604697,2.35225048923679,2.35694716242661,2.36164383561644,2.36634050880626,2.37103718199609,2.37573385518591,2.38043052837573,2.38512720156556,2.38982387475538,2.39452054794521,2.39921722113503,2.40391389432485,2.40861056751468,2.4133072407045,2.41800391389432,2.42270058708415,2.42739726027397,2.4320939334638,2.43679060665362,2.44148727984344,2.44618395303327,2.45088062622309,2.45557729941292,2.46027397260274,2.46497064579256,2.46966731898239,2.47436399217221,2.47906066536204,2.48375733855186,2.48845401174168,2.49315068493151,2.49784735812133,2.50254403131115,2.50724070450098,2.5119373776908,2.51663405088063,2.52133072407045,2.52602739726027,2.5307240704501,2.53542074363992,2.54011741682975,2.54481409001957,2.54951076320939,2.55420743639922,2.55890410958904,2.56360078277886,2.56829745596869,2.57299412915851,2.57769080234834,2.58238747553816,2.58708414872798,2.59178082191781,2.59647749510763,2.60117416829746,2.60587084148728,2.6105675146771,2.61526418786693,2.61996086105675,2.62465753424658,2.6293542074364,2.63405088062622,2.63874755381605,2.64344422700587,2.64814090019569,2.65283757338552,2.65753424657534,2.66223091976517,2.66692759295499,2.67162426614481,2.67632093933464,2.68101761252446,2.68571428571429,2.69041095890411,2.69510763209393,2.69980430528376,2.70450097847358,2.70919765166341,2.71389432485323,2.71859099804305,2.72328767123288,2.7279843444227,2.73268101761252,2.73737769080235,2.74207436399217,2.746771037182,2.75146771037182,2.75616438356164,2.76086105675147,2.76555772994129,2.77025440313112,2.77495107632094,2.77964774951076,2.78434442270059,2.78904109589041,2.79373776908024,2.79843444227006,2.80313111545988,2.80782778864971,2.81252446183953,2.81722113502935,2.82191780821918,2.826614481409,2.83131115459883,2.83600782778865,2.84070450097847,2.8454011741683,2.85009784735812,2.85479452054795,2.85949119373777,2.86418786692759,2.86888454011742,2.87358121330724,2.87827788649706,2.88297455968689,2.88767123287671,2.89236790606654,2.89706457925636,2.90176125244618,2.90645792563601,2.91115459882583,2.91585127201566,2.92054794520548,2.9252446183953,2.92994129158513,2.93463796477495,2.93933463796477,2.9440313111546,2.94872798434442,2.95342465753425,2.95812133072407,2.96281800391389,2.96751467710372,2.97221135029354,2.97690802348337,2.98160469667319,2.98630136986301,2.99099804305284,2.99569471624266,3.00039138943249,3.00508806262231,3.00978473581213,3.01448140900196,3.01917808219178,3.0238747553816,3.02857142857143,3.03326810176125,3.03796477495108,3.0426614481409,3.04735812133072,3.05205479452055,3.05675146771037,3.0614481409002,3.06614481409002,3.07084148727984,3.07553816046967,3.08023483365949,3.08493150684932,3.08962818003914,3.09432485322896,3.09902152641879,3.10371819960861,3.10841487279843,3.11311154598826,3.11780821917808,3.12250489236791,3.12720156555773,3.13189823874755,3.13659491193738,3.1412915851272,3.14598825831703,3.15068493150685,3.15538160469667,3.1600782778865,3.16477495107632,3.16947162426615,3.17416829745597,3.17886497064579,3.18356164383562,3.18825831702544,3.19295499021526,3.19765166340509,3.20234833659491,3.20704500978474,3.21174168297456,3.21643835616438,3.22113502935421,3.22583170254403,3.23052837573386,3.23522504892368,3.2399217221135,3.24461839530333,3.24931506849315,3.25401174168297,3.2587084148728,3.26340508806262,3.26810176125245,3.27279843444227,3.27749510763209,3.28219178082192,3.28688845401174,3.29158512720157,3.29628180039139,3.30097847358121,3.30567514677104,3.31037181996086,3.31506849315068,3.31976516634051,3.32446183953033,3.32915851272016,3.33385518590998,3.3385518590998,3.34324853228963,3.34794520547945,3.35264187866928,3.3573385518591,3.36203522504892,3.36673189823875,3.37142857142857,3.3761252446184,3.38082191780822,3.38551859099804,3.39021526418787,3.39491193737769,3.39960861056751,3.40430528375734,3.40900195694716,3.41369863013699,3.41839530332681,3.42309197651663,3.42778864970646,3.43248532289628,3.43718199608611,3.44187866927593,3.44657534246575,3.45127201565558,3.4559686888454,3.46066536203523,3.46536203522505,3.47005870841487,3.4747553816047,3.47945205479452,3.48414872798434,3.48884540117417,3.49354207436399,3.49823874755382,3.50293542074364,3.50763209393346,3.51232876712329,3.51702544031311,3.52172211350294,3.52641878669276,3.53111545988258,3.53581213307241,3.54050880626223,3.54520547945206,3.54990215264188,3.5545988258317,3.55929549902153,3.56399217221135,3.56868884540117,3.573385518591,3.57808219178082,3.58277886497065,3.58747553816047,3.59217221135029,3.59686888454012,3.60156555772994,3.60626223091977,3.61095890410959,3.61565557729941,3.62035225048924,3.62504892367906,3.62974559686888,3.63444227005871,3.63913894324853,3.64383561643836,3.64853228962818,3.653228962818,3.65792563600783,3.66262230919765,3.66731898238748,3.6720156555773,3.67671232876712,3.68140900195695,3.68610567514677,3.6908023483366,3.69549902152642,3.70019569471624,3.70489236790607,3.70958904109589,3.71428571428571,3.71898238747554,3.72367906066536,3.72837573385519,3.73307240704501,3.73776908023483,3.74246575342466,3.74716242661448,3.75185909980431,3.75655577299413,3.76125244618395,3.76594911937378,3.7706457925636,3.77534246575342,3.78003913894325,3.78473581213307,3.7894324853229,3.79412915851272,3.79882583170254,3.80352250489237,3.80821917808219,3.81291585127202,3.81761252446184,3.82230919765166,3.82700587084149,3.83170254403131,3.83639921722114,3.84109589041096,3.84579256360078,3.85048923679061,3.85518590998043,3.85988258317026,3.86457925636008,3.8692759295499,3.87397260273973,3.87866927592955,3.88336594911937,3.8880626223092,3.89275929549902,3.89745596868885,3.90215264187867,3.90684931506849,3.91154598825832,3.91624266144814,3.92093933463797,3.92563600782779,3.93033268101761,3.93502935420744,3.93972602739726,3.94442270058708,3.94911937377691,3.95381604696673,3.95851272015656,3.96320939334638,3.9679060665362,3.97260273972603,3.97729941291585,3.98199608610568,3.9866927592955,3.99138943248532,3.99608610567515,4.00078277886497,4.00547945205479,4.01017612524462,4.01487279843444,4.01956947162427,4.02426614481409,4.02896281800391,4.03365949119374,4.03835616438356,4.04305283757339,4.04774951076321,4.05244618395303,4.05714285714286,4.06183953033268,4.0665362035225,4.07123287671233,4.07592954990215,4.08062622309198,4.0853228962818,4.09001956947163,4.09471624266145,4.09941291585127,4.1041095890411,4.10880626223092,4.11350293542074,4.11819960861057,4.12289628180039,4.12759295499022,4.13228962818004,4.13698630136986,4.14168297455969,4.14637964774951,4.15107632093934,4.15577299412916,4.16046966731898,4.16516634050881,4.16986301369863,4.17455968688845,4.17925636007828,4.1839530332681,4.18864970645793,4.19334637964775,4.19804305283757,4.2027397260274,4.20743639921722,4.21213307240705,4.21682974559687,4.22152641878669,4.22622309197652,4.23091976516634,4.23561643835616,4.24031311154599,4.24500978473581,4.24970645792564,4.25440313111546,4.25909980430528,4.26379647749511,4.26849315068493,4.27318982387476,4.27788649706458,4.2825831702544,4.28727984344423,4.29197651663405,4.29667318982388,4.3013698630137,4.30606653620352,4.31076320939335,4.31545988258317,4.32015655577299,4.32485322896282,4.32954990215264,4.33424657534247,4.33894324853229,4.34363992172211,4.34833659491194,4.35303326810176,4.35772994129159,4.36242661448141,4.36712328767123,4.37181996086106,4.37651663405088,4.3812133072407,4.38590998043053,4.39060665362035,4.39530332681018,4.4,4.4,4.39530332681018,4.39060665362035,4.38590998043053,4.3812133072407,4.37651663405088,4.37181996086106,4.36712328767123,4.36242661448141,4.35772994129159,4.35303326810176,4.34833659491194,4.34363992172211,4.33894324853229,4.33424657534247,4.32954990215264,4.32485322896282,4.32015655577299,4.31545988258317,4.31076320939335,4.30606653620352,4.3013698630137,4.29667318982388,4.29197651663405,4.28727984344423,4.2825831702544,4.27788649706458,4.27318982387476,4.26849315068493,4.26379647749511,4.25909980430528,4.25440313111546,4.24970645792564,4.24500978473581,4.24031311154599,4.23561643835616,4.23091976516634,4.22622309197652,4.22152641878669,4.21682974559687,4.21213307240705,4.20743639921722,4.2027397260274,4.19804305283757,4.19334637964775,4.18864970645793,4.1839530332681,4.17925636007828,4.17455968688845,4.16986301369863,4.16516634050881,4.16046966731898,4.15577299412916,4.15107632093934,4.14637964774951,4.14168297455969,4.13698630136986,4.13228962818004,4.12759295499022,4.12289628180039,4.11819960861057,4.11350293542074,4.10880626223092,4.1041095890411,4.09941291585127,4.09471624266145,4.09001956947163,4.0853228962818,4.08062622309198,4.07592954990215,4.07123287671233,4.0665362035225,4.06183953033268,4.05714285714286,4.05244618395303,4.04774951076321,4.04305283757339,4.03835616438356,4.03365949119374,4.02896281800391,4.02426614481409,4.01956947162427,4.01487279843444,4.01017612524462,4.00547945205479,4.00078277886497,3.99608610567515,3.99138943248532,3.9866927592955,3.98199608610568,3.97729941291585,3.97260273972603,3.9679060665362,3.96320939334638,3.95851272015656,3.95381604696673,3.94911937377691,3.94442270058708,3.93972602739726,3.93502935420744,3.93033268101761,3.92563600782779,3.92093933463797,3.91624266144814,3.91154598825832,3.90684931506849,3.90215264187867,3.89745596868885,3.89275929549902,3.8880626223092,3.88336594911937,3.87866927592955,3.87397260273973,3.8692759295499,3.86457925636008,3.85988258317026,3.85518590998043,3.85048923679061,3.84579256360078,3.84109589041096,3.83639921722114,3.83170254403131,3.82700587084149,3.82230919765166,3.81761252446184,3.81291585127202,3.80821917808219,3.80352250489237,3.79882583170254,3.79412915851272,3.7894324853229,3.78473581213307,3.78003913894325,3.77534246575342,3.7706457925636,3.76594911937378,3.76125244618395,3.75655577299413,3.75185909980431,3.74716242661448,3.74246575342466,3.73776908023483,3.73307240704501,3.72837573385519,3.72367906066536,3.71898238747554,3.71428571428571,3.70958904109589,3.70489236790607,3.70019569471624,3.69549902152642,3.6908023483366,3.68610567514677,3.68140900195695,3.67671232876712,3.6720156555773,3.66731898238748,3.66262230919765,3.65792563600783,3.653228962818,3.64853228962818,3.64383561643836,3.63913894324853,3.63444227005871,3.62974559686888,3.62504892367906,3.62035225048924,3.61565557729941,3.61095890410959,3.60626223091977,3.60156555772994,3.59686888454012,3.59217221135029,3.58747553816047,3.58277886497065,3.57808219178082,3.573385518591,3.56868884540117,3.56399217221135,3.55929549902153,3.5545988258317,3.54990215264188,3.54520547945206,3.54050880626223,3.53581213307241,3.53111545988258,3.52641878669276,3.52172211350294,3.51702544031311,3.51232876712329,3.50763209393346,3.50293542074364,3.49823874755382,3.49354207436399,3.48884540117417,3.48414872798434,3.47945205479452,3.4747553816047,3.47005870841487,3.46536203522505,3.46066536203523,3.4559686888454,3.45127201565558,3.44657534246575,3.44187866927593,3.43718199608611,3.43248532289628,3.42778864970646,3.42309197651663,3.41839530332681,3.41369863013699,3.40900195694716,3.40430528375734,3.39960861056751,3.39491193737769,3.39021526418787,3.38551859099804,3.38082191780822,3.3761252446184,3.37142857142857,3.36673189823875,3.36203522504892,3.3573385518591,3.35264187866928,3.34794520547945,3.34324853228963,3.3385518590998,3.33385518590998,3.32915851272016,3.32446183953033,3.31976516634051,3.31506849315068,3.31037181996086,3.30567514677104,3.30097847358121,3.29628180039139,3.29158512720157,3.28688845401174,3.28219178082192,3.27749510763209,3.27279843444227,3.26810176125245,3.26340508806262,3.2587084148728,3.25401174168297,3.24931506849315,3.24461839530333,3.2399217221135,3.23522504892368,3.23052837573386,3.22583170254403,3.22113502935421,3.21643835616438,3.21174168297456,3.20704500978474,3.20234833659491,3.19765166340509,3.19295499021526,3.18825831702544,3.18356164383562,3.17886497064579,3.17416829745597,3.16947162426615,3.16477495107632,3.1600782778865,3.15538160469667,3.15068493150685,3.14598825831703,3.1412915851272,3.13659491193738,3.13189823874755,3.12720156555773,3.12250489236791,3.11780821917808,3.11311154598826,3.10841487279843,3.10371819960861,3.09902152641879,3.09432485322896,3.08962818003914,3.08493150684932,3.08023483365949,3.07553816046967,3.07084148727984,3.06614481409002,3.0614481409002,3.05675146771037,3.05205479452055,3.04735812133072,3.0426614481409,3.03796477495108,3.03326810176125,3.02857142857143,3.0238747553816,3.01917808219178,3.01448140900196,3.00978473581213,3.00508806262231,3.00039138943249,2.99569471624266,2.99099804305284,2.98630136986301,2.98160469667319,2.97690802348337,2.97221135029354,2.96751467710372,2.96281800391389,2.95812133072407,2.95342465753425,2.94872798434442,2.9440313111546,2.93933463796477,2.93463796477495,2.92994129158513,2.9252446183953,2.92054794520548,2.91585127201566,2.91115459882583,2.90645792563601,2.90176125244618,2.89706457925636,2.89236790606654,2.88767123287671,2.88297455968689,2.87827788649706,2.87358121330724,2.86888454011742,2.86418786692759,2.85949119373777,2.85479452054795,2.85009784735812,2.8454011741683,2.84070450097847,2.83600782778865,2.83131115459883,2.826614481409,2.82191780821918,2.81722113502935,2.81252446183953,2.80782778864971,2.80313111545988,2.79843444227006,2.79373776908024,2.78904109589041,2.78434442270059,2.77964774951076,2.77495107632094,2.77025440313112,2.76555772994129,2.76086105675147,2.75616438356164,2.75146771037182,2.746771037182,2.74207436399217,2.73737769080235,2.73268101761252,2.7279843444227,2.72328767123288,2.71859099804305,2.71389432485323,2.70919765166341,2.70450097847358,2.69980430528376,2.69510763209393,2.69041095890411,2.68571428571429,2.68101761252446,2.67632093933464,2.67162426614481,2.66692759295499,2.66223091976517,2.65753424657534,2.65283757338552,2.64814090019569,2.64344422700587,2.63874755381605,2.63405088062622,2.6293542074364,2.62465753424658,2.61996086105675,2.61526418786693,2.6105675146771,2.60587084148728,2.60117416829746,2.59647749510763,2.59178082191781,2.58708414872798,2.58238747553816,2.57769080234834,2.57299412915851,2.56829745596869,2.56360078277886,2.55890410958904,2.55420743639922,2.54951076320939,2.54481409001957,2.54011741682975,2.53542074363992,2.5307240704501,2.52602739726027,2.52133072407045,2.51663405088063,2.5119373776908,2.50724070450098,2.50254403131115,2.49784735812133,2.49315068493151,2.48845401174168,2.48375733855186,2.47906066536204,2.47436399217221,2.46966731898239,2.46497064579256,2.46027397260274,2.45557729941292,2.45088062622309,2.44618395303327,2.44148727984344,2.43679060665362,2.4320939334638,2.42739726027397,2.42270058708415,2.41800391389432,2.4133072407045,2.40861056751468,2.40391389432485,2.39921722113503,2.39452054794521,2.38982387475538,2.38512720156556,2.38043052837573,2.37573385518591,2.37103718199609,2.36634050880626,2.36164383561644,2.35694716242661,2.35225048923679,2.34755381604697,2.34285714285714,2.33816046966732,2.3334637964775,2.32876712328767,2.32407045009785,2.31937377690802,2.3146771037182,2.30998043052838,2.30528375733855,2.30058708414873,2.2958904109589,2.29119373776908,2.28649706457926,2.28180039138943,2.27710371819961,2.27240704500978,2.26771037181996,2.26301369863014,2.25831702544031,2.25362035225049,2.24892367906067,2.24422700587084,2.23953033268102,2.23483365949119,2.23013698630137,2.22544031311155,2.22074363992172,2.2160469667319,2.21135029354207,2.20665362035225,2.20195694716243,2.1972602739726,2.19256360078278,2.18786692759295,2.18317025440313,2.17847358121331,2.17377690802348,2.16908023483366,2.16438356164384,2.15968688845401,2.15499021526419,2.15029354207436,2.14559686888454,2.14090019569472,2.13620352250489,2.13150684931507,2.12681017612524,2.12211350293542,2.1174168297456,2.11272015655577,2.10802348336595,2.10332681017613,2.0986301369863,2.09393346379648,2.08923679060665,2.08454011741683,2.07984344422701,2.07514677103718,2.07045009784736,2.06575342465753,2.06105675146771,2.05636007827789,2.05166340508806,2.04696673189824,2.04227005870841,2.03757338551859,2.03287671232877,2.02818003913894,2.02348336594912,2.0187866927593,2.01409001956947,2.00939334637965,2.00469667318982,2,2],"y":[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0.0843713630760147,0.0859056948154118,0.0874383433064483,0.0889694239205466,0.0905002161659777,0.0920308158206397,0.0935626240893455,0.0950959325204047,0.0966309347099747,0.0981709490492189,0.0997146862989821,0.101262221124735,0.102819467834408,0.104382976632824,0.105952817491745,0.107537034762375,0.109130631843918,0.110733300507608,0.112354949175266,0.113989484948709,0.115635913203803,0.117305703663788,0.118992180304146,0.120693363614851,0.122421940089956,0.124171223912733,0.125937963182227,0.12773570908056,0.129558362383996,0.131401129969583,0.133278094620754,0.135184338638416,0.137113276275373,0.13907922416338,0.1410790296788,0.143104070690827,0.145168640302435,0.14727192733952,0.149403020615956,0.151575966933005,0.153792881274377,0.156040281479711,0.158331765258418,0.160672988751267,0.16304756743214,0.16546844682805,0.167945491638171,0.170459018336922,0.173021094944362,0.175646529865458,0.178311871671359,0.18102804407427,0.18381560400582,0.186646796362418,0.189531079861572,0.192495617493075,0.195507768266255,0.198575149185142,0.201732400194203,0.204941402306614,0.208207508463427,0.211573657206016,0.214995701527856,0.218476286090853,0.222067336608858,0.225718234977734,0.229429080526776,0.23325946359025,0.237153810383925,0.241111605560892,0.245191541657117,0.249341759796754,0.253558224418588,0.257897670393667,0.262313002615659,0.266796410094933,0.271401569491692,0.276087086957984,0.28084128894163,0.285713727219402,0.290669434041242,0.295693002509177,0.300828905484249,0.306049018810298,0.311334584682411,0.31672423145895,0.322196712059999,0.327730573033387,0.333358086649058,0.339064484414577,0.344826542687304,0.350669968609008,0.356585631421415,0.362549704623661,0.368581449647702,0.374676123447136,0.380810650321984,0.386998293216515,0.393237116778034,0.39950625408616,0.405813715593078,0.412158586970544,0.418523667169296,0.424912702114761,0.431323966455393,0.437745258569034,0.444177208334948,0.450615590582354,0.457054281109018,0.463492002192029,0.469920684875266,0.476340973364759,0.482750838783101,0.489137566380832,0.495508753174207,0.501862609552293,0.508181688396648,0.514480121445086,0.520757076972116,0.526991134598871,0.533201879456007,0.539389275360396,0.545531168673374,0.551649271439901,0.557744176202336,0.563797470702958,0.569828698152443,0.575839711307486,0.581817741997448,0.587778706036378,0.593725209491287,0.599651434392356,0.605569038552697,0.611480457217147,0.617387455302277,0.623297637742054,0.629211995927308,0.635139811518405,0.64108560015039,0.647047400978795,0.653041277117206,0.65907021575115,0.665127714270257,0.671235297133722,0.677396344393153,0.683598327429069,0.689866461442361,0.696206504064506,0.702598728426245,0.709069994935314,0.715630151261504,0.722251624131705,0.728960803130274,0.735772718689435,0.742652027523887,0.749622679916931,0.756705032852336,0.763856944828472,0.771098320422641,0.778453758415553,0.785876309870707,0.793380782498287,0.800993503678554,0.808665787120106,0.816407002399415,0.824241170062357,0.832121999982706,0.840053893215275,0.848053038318224,0.856080637768161,0.864137440108889,0.872224957663759,0.880317756731685,0.88841505852942,0.896495846159377,0.904554423427115,0.912590021594397,0.920554528690742,0.92846466045674,0.936320836090773,0.944049742510105,0.951686455861583,0.959236094185738,0.966602940883758,0.973835399866868,0.980946906444839,0.987823335780636,0.994520329553146,1.00106323868225,1.00732445404533,1.01336020656398,1.01921102980778,1.02474135104935,1.03000133772026,1.03504933153342,1.03974754286098,1.04413398407295,1.04828650549711,1.05207069141113,1.05550739824961,1.0586945289741,1.06150611228846,1.06394238681954,1.06612053888879,1.06792727262936,1.06934061312707,1.07049488293586,1.07129260319236,1.0716900310204,1.07183513896042,1.07164815533713,1.07106606073306,1.07024579737183,1.06912587701928,1.0676283702616,1.06591356044004,1.06393754526523,1.0616133912135,1.05909847836069,1.05636465691326,1.0533229579134,1.05012139542524,1.04674482515682,1.04310969327239,1.03934840164637,1.03545543832196,1.0313599571364,1.02717315940089,1.02289549405604,1.01847530866315,1.01399808874049,1.00946468938951,1.0048535047167,1.00021544187175,0.995553153559966,0.990870057975627,0.986189275633306,0.981512470162723,0.97686131396825,0.972239245113067,0.967644357908055,0.963109883567209,0.958627001523746,0.954188549643324,0.949833099014955,0.945545796716576,0.941313583649177,0.937174962686618,0.933113691262621,0.929111289310575,0.925201845132018,0.921370549478895,0.917595077996081,0.913901978554206,0.910278798634266,0.906701949347769,0.903188597774987,0.899727743809904,0.896297946696381,0.892906413654409,0.889541073945319,0.886186651046927,0.882840970712974,0.879487073912939,0.876120195074269,0.87273034420009,0.86929097357712,0.865812207127252,0.862278571346664,0.858648815949703,0.854952057655795,0.851170183602154,0.847242208638506,0.843219772060005,0.839085206217297,0.834753330440708,0.830300989178577,0.825714013011528,0.820879592655355,0.815901378145558,0.810771462447789,0.805347397718801,0.799759973796793,0.794008721440164,0.787924492654871,0.781660949265559,0.775218886531111,0.768430479612716,0.761443412390306,0.754268555699689,0.746745119750407,0.739008889713213,0.731081489880365,0.72281414998565,0.714326248360122,0.705649909750837,0.696652424825482,0.687433901965237,0.678035760171317,0.668344297113803,0.658439251177717,0.648369215179035,0.638041260381211,0.627515419922517,0.616844523470511,0.605956988257623,0.594895461630659,0.583713407104689,0.572360018233899,0.56086431972921,0.54927633386344,0.537564431768319,0.525748927309157,0.513872079584285,0.501918973472191,0.489906915159283,0.477866073948979,0.465795122117295,0.453714458748258,0.441638075477726,0.429574669676408,0.41755382797267,0.405569744159036,0.39363737745949,0.381801205246597,0.370032670729662,0.358349258892496,0.346815306938974,0.335377380177653,0.324051987497008,0.312927282093135,0.30192375631808,0.291053849699313,0.28043227517014,0.269953239472936,0.259622561076261,0.249582932759099,0.239703037489241,0.229983271086588,0.220585015781874,0.211362469871831,0.202312742067912,0.193595157339978,0.185071444379113,0.176728917337856,0.168720690483973,0.160920953259543,0.153306638007358,0.146021367351618,0.138955379430297,0.132075248914371,0.125512707267702,0.119176326612469,0.113022879870063,0.107170650495847,0.101547634972913,0.0961017895001726,0.0909371582113137,0.0860012162769688,0.0812344014182117,0.0767263876905367,0.0724433390356895,0.0683196667612495,0.0644310826637323,0.0607610120847627,0.0572394117170113,0.0539288488535432,0.0508281507019785,0.0478643695044404,0.0450880296465103,0.0425112582932642,0.0400596487410163,0.0377729519561066,0.0356744141760488,0.0336894495842805,0.0318483730481768,0.0301834195917028,0.0286209011488742,0.027183021251439,0.0259090158381438,0.0247269547578795,0.0236521835728403,0.0227291470996856,0.0218883246302163,0.0211393485655871,0.020530293724974,0.0199945184675179,0.0195369614238304,0.0192079476857949,0.01894404364673,0.0187463889656195,0.0186663397111629,0.0186439095133476,0.0186789124798765,0.0188175566057589,0.0190085722159089,0.0192501023521986,0.0195802073483087,0.019958485531526,0.0203807589849391,0.020877807755512,0.0214184292875206,0.0219967712197302,0.0226370559475446,0.0233157864758974,0.0240260606122961,0.0247861648122945,0.0255789312712121,0.0263970385947185,0.0272534035039825,0.0281358733193392,0.0290373649376697,0.0299659782761887,0.0309132794968019,0.031873118996962,0.0328493544578236,0.0338359637807392,0.0348284627241851,0.0358270873063599,0.0368269002300098,0.0378258373804201,0.0388211913999359,0.0398077750894958,0.0407866961918052,0.0417530381597243,0.042700053826722,0.0436327350857352,0.044544731383994,0.0454264999456765,0.0462875456805063,0.0471208739108378,0.047913047317002,0.0486785815028846,0.0494106073287955,0.0500908990938039,0.0507393024866172,0.0513497834459908,0.0518987063395427,0.0524113462960051,0.0528831129820258,0.0532846700364338,0.0536465694427802,0.05396613495343,0.0542084121746194,0.0544088072259965,0.0545668599087581,0.0546424752397739,0.0546752189825422,0.0546650873876843,0.0545733337364906,0.0544371127695135,0.0542583419525879,0.0540018345784583,0.0537001792797922,0.0533575393704444,0.0529430226100734,0.0524841056551176,0.0519869260244613,0.0514253499912342,0.0508215825694425,0.0501832971224602,0.0494893102005921,0.0487567590167979,0.047994280372882,0.0471855755814829,0.0463432353435934,0.0454761953311494,0.0445727486553844,0.0436417134105986,0.0426916150396704,0.0417148595308397,0.0407174413174852,0.0397067714656691,0.0386787532061971,0.0376375977792204,0.0365889500714083,0.0355315109703688,0.0344687579300648,0.0334040119022234,0.0323380400174213,0.0312745689442227,0.0302141650587223,0.0291589462736449,0.0281137421343826,0.0270760833074458,0.0260487794548701,0.0250384404674987,0.024039440381426,0.0230547091351173,0.022093109413177,0.0211458969761441,0.0202156588811607,0.019313767464902,0.0184285462667621,0.0175618949370481,0.0167277430898576,0.0159117953752094,0.0151150388124798,0.0143538193801469,0.0136116365230602,0.0128885064412408,0.0122027278218119,0.011536243918023,0.010888993551353,0.0102779192296185,0.00968682136552681,0.00911460194415665,0.00857653321234728,0.00805862115158893,0.00755879562515243,0.00709048713572472,0.00664205632776599,0],"text":["density: 6.642056e-03<br />Sepal.Width: 2.000000","density: 7.090487e-03<br />Sepal.Width: 2.004697","density: 7.558796e-03<br />Sepal.Width: 2.009393","density: 8.058621e-03<br />Sepal.Width: 2.014090","density: 8.576533e-03<br />Sepal.Width: 2.018787","density: 9.114602e-03<br />Sepal.Width: 2.023483","density: 9.686821e-03<br />Sepal.Width: 2.028180","density: 1.027792e-02<br />Sepal.Width: 2.032877","density: 1.088899e-02<br />Sepal.Width: 2.037573","density: 1.153624e-02<br />Sepal.Width: 2.042270","density: 1.220273e-02<br />Sepal.Width: 2.046967","density: 1.288851e-02<br />Sepal.Width: 2.051663","density: 1.361164e-02<br />Sepal.Width: 2.056360","density: 1.435382e-02<br />Sepal.Width: 2.061057","density: 1.511504e-02<br />Sepal.Width: 2.065753","density: 1.591180e-02<br />Sepal.Width: 2.070450","density: 1.672774e-02<br />Sepal.Width: 2.075147","density: 1.756189e-02<br />Sepal.Width: 2.079843","density: 1.842855e-02<br />Sepal.Width: 2.084540","density: 1.931377e-02<br />Sepal.Width: 2.089237","density: 2.021566e-02<br />Sepal.Width: 2.093933","density: 2.114590e-02<br />Sepal.Width: 2.098630","density: 2.209311e-02<br />Sepal.Width: 2.103327","density: 2.305471e-02<br />Sepal.Width: 2.108023","density: 2.403944e-02<br />Sepal.Width: 2.112720","density: 2.503844e-02<br />Sepal.Width: 2.117417","density: 2.604878e-02<br />Sepal.Width: 2.122114","density: 2.707608e-02<br />Sepal.Width: 2.126810","density: 2.811374e-02<br />Sepal.Width: 2.131507","density: 2.915895e-02<br />Sepal.Width: 2.136204","density: 3.021417e-02<br />Sepal.Width: 2.140900","density: 3.127457e-02<br />Sepal.Width: 2.145597","density: 3.233804e-02<br />Sepal.Width: 2.150294","density: 3.340401e-02<br />Sepal.Width: 2.154990","density: 3.446876e-02<br />Sepal.Width: 2.159687","density: 3.553151e-02<br />Sepal.Width: 2.164384","density: 3.658895e-02<br />Sepal.Width: 2.169080","density: 3.763760e-02<br />Sepal.Width: 2.173777","density: 3.867875e-02<br />Sepal.Width: 2.178474","density: 3.970677e-02<br />Sepal.Width: 2.183170","density: 4.071744e-02<br />Sepal.Width: 2.187867","density: 4.171486e-02<br />Sepal.Width: 2.192564","density: 4.269162e-02<br />Sepal.Width: 2.197260","density: 4.364171e-02<br />Sepal.Width: 2.201957","density: 4.457275e-02<br />Sepal.Width: 2.206654","density: 4.547620e-02<br />Sepal.Width: 2.211350","density: 4.634324e-02<br />Sepal.Width: 2.216047","density: 4.718558e-02<br />Sepal.Width: 2.220744","density: 4.799428e-02<br />Sepal.Width: 2.225440","density: 4.875676e-02<br />Sepal.Width: 2.230137","density: 4.948931e-02<br />Sepal.Width: 2.234834","density: 5.018330e-02<br />Sepal.Width: 2.239530","density: 5.082158e-02<br />Sepal.Width: 2.244227","density: 5.142535e-02<br />Sepal.Width: 2.248924","density: 5.198693e-02<br />Sepal.Width: 2.253620","density: 5.248411e-02<br />Sepal.Width: 2.258317","density: 5.294302e-02<br />Sepal.Width: 2.263014","density: 5.335754e-02<br />Sepal.Width: 2.267710","density: 5.370018e-02<br />Sepal.Width: 2.272407","density: 5.400183e-02<br />Sepal.Width: 2.277104","density: 5.425834e-02<br />Sepal.Width: 2.281800","density: 5.443711e-02<br />Sepal.Width: 2.286497","density: 5.457333e-02<br />Sepal.Width: 2.291194","density: 5.466509e-02<br />Sepal.Width: 2.295890","density: 5.467522e-02<br />Sepal.Width: 2.300587","density: 5.464248e-02<br />Sepal.Width: 2.305284","density: 5.456686e-02<br />Sepal.Width: 2.309980","density: 5.440881e-02<br />Sepal.Width: 2.314677","density: 5.420841e-02<br />Sepal.Width: 2.319374","density: 5.396613e-02<br />Sepal.Width: 2.324070","density: 5.364657e-02<br />Sepal.Width: 2.328767","density: 5.328467e-02<br />Sepal.Width: 2.333464","density: 5.288311e-02<br />Sepal.Width: 2.338160","density: 5.241135e-02<br />Sepal.Width: 2.342857","density: 5.189871e-02<br />Sepal.Width: 2.347554","density: 5.134978e-02<br />Sepal.Width: 2.352250","density: 5.073930e-02<br />Sepal.Width: 2.356947","density: 5.009090e-02<br />Sepal.Width: 2.361644","density: 4.941061e-02<br />Sepal.Width: 2.366341","density: 4.867858e-02<br />Sepal.Width: 2.371037","density: 4.791305e-02<br />Sepal.Width: 2.375734","density: 4.712087e-02<br />Sepal.Width: 2.380431","density: 4.628755e-02<br />Sepal.Width: 2.385127","density: 4.542650e-02<br />Sepal.Width: 2.389824","density: 4.454473e-02<br />Sepal.Width: 2.394521","density: 4.363274e-02<br />Sepal.Width: 2.399217","density: 4.270005e-02<br />Sepal.Width: 2.403914","density: 4.175304e-02<br />Sepal.Width: 2.408611","density: 4.078670e-02<br />Sepal.Width: 2.413307","density: 3.980778e-02<br />Sepal.Width: 2.418004","density: 3.882119e-02<br />Sepal.Width: 2.422701","density: 3.782584e-02<br />Sepal.Width: 2.427397","density: 3.682690e-02<br />Sepal.Width: 2.432094","density: 3.582709e-02<br />Sepal.Width: 2.436791","density: 3.482846e-02<br />Sepal.Width: 2.441487","density: 3.383596e-02<br />Sepal.Width: 2.446184","density: 3.284935e-02<br />Sepal.Width: 2.450881","density: 3.187312e-02<br />Sepal.Width: 2.455577","density: 3.091328e-02<br />Sepal.Width: 2.460274","density: 2.996598e-02<br />Sepal.Width: 2.464971","density: 2.903736e-02<br />Sepal.Width: 2.469667","density: 2.813587e-02<br />Sepal.Width: 2.474364","density: 2.725340e-02<br />Sepal.Width: 2.479061","density: 2.639704e-02<br />Sepal.Width: 2.483757","density: 2.557893e-02<br />Sepal.Width: 2.488454","density: 2.478616e-02<br />Sepal.Width: 2.493151","density: 2.402606e-02<br />Sepal.Width: 2.497847","density: 2.331579e-02<br />Sepal.Width: 2.502544","density: 2.263706e-02<br />Sepal.Width: 2.507241","density: 2.199677e-02<br />Sepal.Width: 2.511937","density: 2.141843e-02<br />Sepal.Width: 2.516634","density: 2.087781e-02<br />Sepal.Width: 2.521331","density: 2.038076e-02<br />Sepal.Width: 2.526027","density: 1.995849e-02<br />Sepal.Width: 2.530724","density: 1.958021e-02<br />Sepal.Width: 2.535421","density: 1.925010e-02<br />Sepal.Width: 2.540117","density: 1.900857e-02<br />Sepal.Width: 2.544814","density: 1.881756e-02<br />Sepal.Width: 2.549511","density: 1.867891e-02<br />Sepal.Width: 2.554207","density: 1.864391e-02<br />Sepal.Width: 2.558904","density: 1.866634e-02<br />Sepal.Width: 2.563601","density: 1.874639e-02<br />Sepal.Width: 2.568297","density: 1.894404e-02<br />Sepal.Width: 2.572994","density: 1.920795e-02<br />Sepal.Width: 2.577691","density: 1.953696e-02<br />Sepal.Width: 2.582387","density: 1.999452e-02<br />Sepal.Width: 2.587084","density: 2.053029e-02<br />Sepal.Width: 2.591781","density: 2.113935e-02<br />Sepal.Width: 2.596477","density: 2.188832e-02<br />Sepal.Width: 2.601174","density: 2.272915e-02<br />Sepal.Width: 2.605871","density: 2.365218e-02<br />Sepal.Width: 2.610568","density: 2.472695e-02<br />Sepal.Width: 2.615264","density: 2.590902e-02<br />Sepal.Width: 2.619961","density: 2.718302e-02<br />Sepal.Width: 2.624658","density: 2.862090e-02<br />Sepal.Width: 2.629354","density: 3.018342e-02<br />Sepal.Width: 2.634051","density: 3.184837e-02<br />Sepal.Width: 2.638748","density: 3.368945e-02<br />Sepal.Width: 2.643444","density: 3.567441e-02<br />Sepal.Width: 2.648141","density: 3.777295e-02<br />Sepal.Width: 2.652838","density: 4.005965e-02<br />Sepal.Width: 2.657534","density: 4.251126e-02<br />Sepal.Width: 2.662231","density: 4.508803e-02<br />Sepal.Width: 2.666928","density: 4.786437e-02<br />Sepal.Width: 2.671624","density: 5.082815e-02<br />Sepal.Width: 2.676321","density: 5.392885e-02<br />Sepal.Width: 2.681018","density: 5.723941e-02<br />Sepal.Width: 2.685714","density: 6.076101e-02<br />Sepal.Width: 2.690411","density: 6.443108e-02<br />Sepal.Width: 2.695108","density: 6.831967e-02<br />Sepal.Width: 2.699804","density: 7.244334e-02<br />Sepal.Width: 2.704501","density: 7.672639e-02<br />Sepal.Width: 2.709198","density: 8.123440e-02<br />Sepal.Width: 2.713894","density: 8.600122e-02<br />Sepal.Width: 2.718591","density: 9.093716e-02<br />Sepal.Width: 2.723288","density: 9.610179e-02<br />Sepal.Width: 2.727984","density: 1.015476e-01<br />Sepal.Width: 2.732681","density: 1.071707e-01<br />Sepal.Width: 2.737378","density: 1.130229e-01<br />Sepal.Width: 2.742074","density: 1.191763e-01<br />Sepal.Width: 2.746771","density: 1.255127e-01<br />Sepal.Width: 2.751468","density: 1.320752e-01<br />Sepal.Width: 2.756164","density: 1.389554e-01<br />Sepal.Width: 2.760861","density: 1.460214e-01<br />Sepal.Width: 2.765558","density: 1.533066e-01<br />Sepal.Width: 2.770254","density: 1.609210e-01<br />Sepal.Width: 2.774951","density: 1.687207e-01<br />Sepal.Width: 2.779648","density: 1.767289e-01<br />Sepal.Width: 2.784344","density: 1.850714e-01<br />Sepal.Width: 2.789041","density: 1.935952e-01<br />Sepal.Width: 2.793738","density: 2.023127e-01<br />Sepal.Width: 2.798434","density: 2.113625e-01<br />Sepal.Width: 2.803131","density: 2.205850e-01<br />Sepal.Width: 2.807828","density: 2.299833e-01<br />Sepal.Width: 2.812524","density: 2.397030e-01<br />Sepal.Width: 2.817221","density: 2.495829e-01<br />Sepal.Width: 2.821918","density: 2.596226e-01<br />Sepal.Width: 2.826614","density: 2.699532e-01<br />Sepal.Width: 2.831311","density: 2.804323e-01<br />Sepal.Width: 2.836008","density: 2.910538e-01<br />Sepal.Width: 2.840705","density: 3.019238e-01<br />Sepal.Width: 2.845401","density: 3.129273e-01<br />Sepal.Width: 2.850098","density: 3.240520e-01<br />Sepal.Width: 2.854795","density: 3.353774e-01<br />Sepal.Width: 2.859491","density: 3.468153e-01<br />Sepal.Width: 2.864188","density: 3.583493e-01<br />Sepal.Width: 2.868885","density: 3.700327e-01<br />Sepal.Width: 2.873581","density: 3.818012e-01<br />Sepal.Width: 2.878278","density: 3.936374e-01<br />Sepal.Width: 2.882975","density: 4.055697e-01<br />Sepal.Width: 2.887671","density: 4.175538e-01<br />Sepal.Width: 2.892368","density: 4.295747e-01<br />Sepal.Width: 2.897065","density: 4.416381e-01<br />Sepal.Width: 2.901761","density: 4.537145e-01<br />Sepal.Width: 2.906458","density: 4.657951e-01<br />Sepal.Width: 2.911155","density: 4.778661e-01<br />Sepal.Width: 2.915851","density: 4.899069e-01<br />Sepal.Width: 2.920548","density: 5.019190e-01<br />Sepal.Width: 2.925245","density: 5.138721e-01<br />Sepal.Width: 2.929941","density: 5.257489e-01<br />Sepal.Width: 2.934638","density: 5.375644e-01<br />Sepal.Width: 2.939335","density: 5.492763e-01<br />Sepal.Width: 2.944031","density: 5.608643e-01<br />Sepal.Width: 2.948728","density: 5.723600e-01<br />Sepal.Width: 2.953425","density: 5.837134e-01<br />Sepal.Width: 2.958121","density: 5.948955e-01<br />Sepal.Width: 2.962818","density: 6.059570e-01<br />Sepal.Width: 2.967515","density: 6.168445e-01<br />Sepal.Width: 2.972211","density: 6.275154e-01<br />Sepal.Width: 2.976908","density: 6.380413e-01<br />Sepal.Width: 2.981605","density: 6.483692e-01<br />Sepal.Width: 2.986301","density: 6.584393e-01<br />Sepal.Width: 2.990998","density: 6.683443e-01<br />Sepal.Width: 2.995695","density: 6.780358e-01<br />Sepal.Width: 3.000391","density: 6.874339e-01<br />Sepal.Width: 3.005088","density: 6.966524e-01<br />Sepal.Width: 3.009785","density: 7.056499e-01<br />Sepal.Width: 3.014481","density: 7.143262e-01<br />Sepal.Width: 3.019178","density: 7.228141e-01<br />Sepal.Width: 3.023875","density: 7.310815e-01<br />Sepal.Width: 3.028571","density: 7.390089e-01<br />Sepal.Width: 3.033268","density: 7.467451e-01<br />Sepal.Width: 3.037965","density: 7.542686e-01<br />Sepal.Width: 3.042661","density: 7.614434e-01<br />Sepal.Width: 3.047358","density: 7.684305e-01<br />Sepal.Width: 3.052055","density: 7.752189e-01<br />Sepal.Width: 3.056751","density: 7.816609e-01<br />Sepal.Width: 3.061448","density: 7.879245e-01<br />Sepal.Width: 3.066145","density: 7.940087e-01<br />Sepal.Width: 3.070841","density: 7.997600e-01<br />Sepal.Width: 3.075538","density: 8.053474e-01<br />Sepal.Width: 3.080235","density: 8.107715e-01<br />Sepal.Width: 3.084932","density: 8.159014e-01<br />Sepal.Width: 3.089628","density: 8.208796e-01<br />Sepal.Width: 3.094325","density: 8.257140e-01<br />Sepal.Width: 3.099022","density: 8.303010e-01<br />Sepal.Width: 3.103718","density: 8.347533e-01<br />Sepal.Width: 3.108415","density: 8.390852e-01<br />Sepal.Width: 3.113112","density: 8.432198e-01<br />Sepal.Width: 3.117808","density: 8.472422e-01<br />Sepal.Width: 3.122505","density: 8.511702e-01<br />Sepal.Width: 3.127202","density: 8.549521e-01<br />Sepal.Width: 3.131898","density: 8.586488e-01<br />Sepal.Width: 3.136595","density: 8.622786e-01<br />Sepal.Width: 3.141292","density: 8.658122e-01<br />Sepal.Width: 3.145988","density: 8.692910e-01<br />Sepal.Width: 3.150685","density: 8.727303e-01<br />Sepal.Width: 3.155382","density: 8.761202e-01<br />Sepal.Width: 3.160078","density: 8.794871e-01<br />Sepal.Width: 3.164775","density: 8.828410e-01<br />Sepal.Width: 3.169472","density: 8.861867e-01<br />Sepal.Width: 3.174168","density: 8.895411e-01<br />Sepal.Width: 3.178865","density: 8.929064e-01<br />Sepal.Width: 3.183562","density: 8.962979e-01<br />Sepal.Width: 3.188258","density: 8.997277e-01<br />Sepal.Width: 3.192955","density: 9.031886e-01<br />Sepal.Width: 3.197652","density: 9.067019e-01<br />Sepal.Width: 3.202348","density: 9.102788e-01<br />Sepal.Width: 3.207045","density: 9.139020e-01<br />Sepal.Width: 3.211742","density: 9.175951e-01<br />Sepal.Width: 3.216438","density: 9.213705e-01<br />Sepal.Width: 3.221135","density: 9.252018e-01<br />Sepal.Width: 3.225832","density: 9.291113e-01<br />Sepal.Width: 3.230528","density: 9.331137e-01<br />Sepal.Width: 3.235225","density: 9.371750e-01<br />Sepal.Width: 3.239922","density: 9.413136e-01<br />Sepal.Width: 3.244618","density: 9.455458e-01<br />Sepal.Width: 3.249315","density: 9.498331e-01<br />Sepal.Width: 3.254012","density: 9.541885e-01<br />Sepal.Width: 3.258708","density: 9.586270e-01<br />Sepal.Width: 3.263405","density: 9.631099e-01<br />Sepal.Width: 3.268102","density: 9.676444e-01<br />Sepal.Width: 3.272798","density: 9.722392e-01<br />Sepal.Width: 3.277495","density: 9.768613e-01<br />Sepal.Width: 3.282192","density: 9.815125e-01<br />Sepal.Width: 3.286888","density: 9.861893e-01<br />Sepal.Width: 3.291585","density: 9.908701e-01<br />Sepal.Width: 3.296282","density: 9.955532e-01<br />Sepal.Width: 3.300978","density: 1.000215e+00<br />Sepal.Width: 3.305675","density: 1.004854e+00<br />Sepal.Width: 3.310372","density: 1.009465e+00<br />Sepal.Width: 3.315068","density: 1.013998e+00<br />Sepal.Width: 3.319765","density: 1.018475e+00<br />Sepal.Width: 3.324462","density: 1.022895e+00<br />Sepal.Width: 3.329159","density: 1.027173e+00<br />Sepal.Width: 3.333855","density: 1.031360e+00<br />Sepal.Width: 3.338552","density: 1.035455e+00<br />Sepal.Width: 3.343249","density: 1.039348e+00<br />Sepal.Width: 3.347945","density: 1.043110e+00<br />Sepal.Width: 3.352642","density: 1.046745e+00<br />Sepal.Width: 3.357339","density: 1.050121e+00<br />Sepal.Width: 3.362035","density: 1.053323e+00<br />Sepal.Width: 3.366732","density: 1.056365e+00<br />Sepal.Width: 3.371429","density: 1.059098e+00<br />Sepal.Width: 3.376125","density: 1.061613e+00<br />Sepal.Width: 3.380822","density: 1.063938e+00<br />Sepal.Width: 3.385519","density: 1.065914e+00<br />Sepal.Width: 3.390215","density: 1.067628e+00<br />Sepal.Width: 3.394912","density: 1.069126e+00<br />Sepal.Width: 3.399609","density: 1.070246e+00<br />Sepal.Width: 3.404305","density: 1.071066e+00<br />Sepal.Width: 3.409002","density: 1.071648e+00<br />Sepal.Width: 3.413699","density: 1.071835e+00<br />Sepal.Width: 3.418395","density: 1.071690e+00<br />Sepal.Width: 3.423092","density: 1.071293e+00<br />Sepal.Width: 3.427789","density: 1.070495e+00<br />Sepal.Width: 3.432485","density: 1.069341e+00<br />Sepal.Width: 3.437182","density: 1.067927e+00<br />Sepal.Width: 3.441879","density: 1.066121e+00<br />Sepal.Width: 3.446575","density: 1.063942e+00<br />Sepal.Width: 3.451272","density: 1.061506e+00<br />Sepal.Width: 3.455969","density: 1.058695e+00<br />Sepal.Width: 3.460665","density: 1.055507e+00<br />Sepal.Width: 3.465362","density: 1.052071e+00<br />Sepal.Width: 3.470059","density: 1.048287e+00<br />Sepal.Width: 3.474755","density: 1.044134e+00<br />Sepal.Width: 3.479452","density: 1.039748e+00<br />Sepal.Width: 3.484149","density: 1.035049e+00<br />Sepal.Width: 3.488845","density: 1.030001e+00<br />Sepal.Width: 3.493542","density: 1.024741e+00<br />Sepal.Width: 3.498239","density: 1.019211e+00<br />Sepal.Width: 3.502935","density: 1.013360e+00<br />Sepal.Width: 3.507632","density: 1.007324e+00<br />Sepal.Width: 3.512329","density: 1.001063e+00<br />Sepal.Width: 3.517025","density: 9.945203e-01<br />Sepal.Width: 3.521722","density: 9.878233e-01<br />Sepal.Width: 3.526419","density: 9.809469e-01<br />Sepal.Width: 3.531115","density: 9.738354e-01<br />Sepal.Width: 3.535812","density: 9.666029e-01<br />Sepal.Width: 3.540509","density: 9.592361e-01<br />Sepal.Width: 3.545205","density: 9.516865e-01<br />Sepal.Width: 3.549902","density: 9.440497e-01<br />Sepal.Width: 3.554599","density: 9.363208e-01<br />Sepal.Width: 3.559295","density: 9.284647e-01<br />Sepal.Width: 3.563992","density: 9.205545e-01<br />Sepal.Width: 3.568689","density: 9.125900e-01<br />Sepal.Width: 3.573386","density: 9.045544e-01<br />Sepal.Width: 3.578082","density: 8.964958e-01<br />Sepal.Width: 3.582779","density: 8.884151e-01<br />Sepal.Width: 3.587476","density: 8.803178e-01<br />Sepal.Width: 3.592172","density: 8.722250e-01<br />Sepal.Width: 3.596869","density: 8.641374e-01<br />Sepal.Width: 3.601566","density: 8.560806e-01<br />Sepal.Width: 3.606262","density: 8.480530e-01<br />Sepal.Width: 3.610959","density: 8.400539e-01<br />Sepal.Width: 3.615656","density: 8.321220e-01<br />Sepal.Width: 3.620352","density: 8.242412e-01<br />Sepal.Width: 3.625049","density: 8.164070e-01<br />Sepal.Width: 3.629746","density: 8.086658e-01<br />Sepal.Width: 3.634442","density: 8.009935e-01<br />Sepal.Width: 3.639139","density: 7.933808e-01<br />Sepal.Width: 3.643836","density: 7.858763e-01<br />Sepal.Width: 3.648532","density: 7.784538e-01<br />Sepal.Width: 3.653229","density: 7.710983e-01<br />Sepal.Width: 3.657926","density: 7.638569e-01<br />Sepal.Width: 3.662622","density: 7.567050e-01<br />Sepal.Width: 3.667319","density: 7.496227e-01<br />Sepal.Width: 3.672016","density: 7.426520e-01<br />Sepal.Width: 3.676712","density: 7.357727e-01<br />Sepal.Width: 3.681409","density: 7.289608e-01<br />Sepal.Width: 3.686106","density: 7.222516e-01<br />Sepal.Width: 3.690802","density: 7.156302e-01<br />Sepal.Width: 3.695499","density: 7.090700e-01<br />Sepal.Width: 3.700196","density: 7.025987e-01<br />Sepal.Width: 3.704892","density: 6.962065e-01<br />Sepal.Width: 3.709589","density: 6.898665e-01<br />Sepal.Width: 3.714286","density: 6.835983e-01<br />Sepal.Width: 3.718982","density: 6.773963e-01<br />Sepal.Width: 3.723679","density: 6.712353e-01<br />Sepal.Width: 3.728376","density: 6.651277e-01<br />Sepal.Width: 3.733072","density: 6.590702e-01<br />Sepal.Width: 3.737769","density: 6.530413e-01<br />Sepal.Width: 3.742466","density: 6.470474e-01<br />Sepal.Width: 3.747162","density: 6.410856e-01<br />Sepal.Width: 3.751859","density: 6.351398e-01<br />Sepal.Width: 3.756556","density: 6.292120e-01<br />Sepal.Width: 3.761252","density: 6.232976e-01<br />Sepal.Width: 3.765949","density: 6.173875e-01<br />Sepal.Width: 3.770646","density: 6.114805e-01<br />Sepal.Width: 3.775342","density: 6.055690e-01<br />Sepal.Width: 3.780039","density: 5.996514e-01<br />Sepal.Width: 3.784736","density: 5.937252e-01<br />Sepal.Width: 3.789432","density: 5.877787e-01<br />Sepal.Width: 3.794129","density: 5.818177e-01<br />Sepal.Width: 3.798826","density: 5.758397e-01<br />Sepal.Width: 3.803523","density: 5.698287e-01<br />Sepal.Width: 3.808219","density: 5.637975e-01<br />Sepal.Width: 3.812916","density: 5.577442e-01<br />Sepal.Width: 3.817613","density: 5.516493e-01<br />Sepal.Width: 3.822309","density: 5.455312e-01<br />Sepal.Width: 3.827006","density: 5.393893e-01<br />Sepal.Width: 3.831703","density: 5.332019e-01<br />Sepal.Width: 3.836399","density: 5.269911e-01<br />Sepal.Width: 3.841096","density: 5.207571e-01<br />Sepal.Width: 3.845793","density: 5.144801e-01<br />Sepal.Width: 3.850489","density: 5.081817e-01<br />Sepal.Width: 3.855186","density: 5.018626e-01<br />Sepal.Width: 3.859883","density: 4.955088e-01<br />Sepal.Width: 3.864579","density: 4.891376e-01<br />Sepal.Width: 3.869276","density: 4.827508e-01<br />Sepal.Width: 3.873973","density: 4.763410e-01<br />Sepal.Width: 3.878669","density: 4.699207e-01<br />Sepal.Width: 3.883366","density: 4.634920e-01<br />Sepal.Width: 3.888063","density: 4.570543e-01<br />Sepal.Width: 3.892759","density: 4.506156e-01<br />Sepal.Width: 3.897456","density: 4.441772e-01<br />Sepal.Width: 3.902153","density: 4.377453e-01<br />Sepal.Width: 3.906849","density: 4.313240e-01<br />Sepal.Width: 3.911546","density: 4.249127e-01<br />Sepal.Width: 3.916243","density: 4.185237e-01<br />Sepal.Width: 3.920939","density: 4.121586e-01<br />Sepal.Width: 3.925636","density: 4.058137e-01<br />Sepal.Width: 3.930333","density: 3.995063e-01<br />Sepal.Width: 3.935029","density: 3.932371e-01<br />Sepal.Width: 3.939726","density: 3.869983e-01<br />Sepal.Width: 3.944423","density: 3.808107e-01<br />Sepal.Width: 3.949119","density: 3.746761e-01<br />Sepal.Width: 3.953816","density: 3.685814e-01<br />Sepal.Width: 3.958513","density: 3.625497e-01<br />Sepal.Width: 3.963209","density: 3.565856e-01<br />Sepal.Width: 3.967906","density: 3.506700e-01<br />Sepal.Width: 3.972603","density: 3.448265e-01<br />Sepal.Width: 3.977299","density: 3.390645e-01<br />Sepal.Width: 3.981996","density: 3.333581e-01<br />Sepal.Width: 3.986693","density: 3.277306e-01<br />Sepal.Width: 3.991389","density: 3.221967e-01<br />Sepal.Width: 3.996086","density: 3.167242e-01<br />Sepal.Width: 4.000783","density: 3.113346e-01<br />Sepal.Width: 4.005479","density: 3.060490e-01<br />Sepal.Width: 4.010176","density: 3.008289e-01<br />Sepal.Width: 4.014873","density: 2.956930e-01<br />Sepal.Width: 4.019569","density: 2.906694e-01<br />Sepal.Width: 4.024266","density: 2.857137e-01<br />Sepal.Width: 4.028963","density: 2.808413e-01<br />Sepal.Width: 4.033659","density: 2.760871e-01<br />Sepal.Width: 4.038356","density: 2.714016e-01<br />Sepal.Width: 4.043053","density: 2.667964e-01<br />Sepal.Width: 4.047750","density: 2.623130e-01<br />Sepal.Width: 4.052446","density: 2.578977e-01<br />Sepal.Width: 4.057143","density: 2.535582e-01<br />Sepal.Width: 4.061840","density: 2.493418e-01<br />Sepal.Width: 4.066536","density: 2.451915e-01<br />Sepal.Width: 4.071233","density: 2.411116e-01<br />Sepal.Width: 4.075930","density: 2.371538e-01<br />Sepal.Width: 4.080626","density: 2.332595e-01<br />Sepal.Width: 4.085323","density: 2.294291e-01<br />Sepal.Width: 4.090020","density: 2.257182e-01<br />Sepal.Width: 4.094716","density: 2.220673e-01<br />Sepal.Width: 4.099413","density: 2.184763e-01<br />Sepal.Width: 4.104110","density: 2.149957e-01<br />Sepal.Width: 4.108806","density: 2.115737e-01<br />Sepal.Width: 4.113503","density: 2.082075e-01<br />Sepal.Width: 4.118200","density: 2.049414e-01<br />Sepal.Width: 4.122896","density: 2.017324e-01<br />Sepal.Width: 4.127593","density: 1.985751e-01<br />Sepal.Width: 4.132290","density: 1.955078e-01<br />Sepal.Width: 4.136986","density: 1.924956e-01<br />Sepal.Width: 4.141683","density: 1.895311e-01<br />Sepal.Width: 4.146380","density: 1.866468e-01<br />Sepal.Width: 4.151076","density: 1.838156e-01<br />Sepal.Width: 4.155773","density: 1.810280e-01<br />Sepal.Width: 4.160470","density: 1.783119e-01<br />Sepal.Width: 4.165166","density: 1.756465e-01<br />Sepal.Width: 4.169863","density: 1.730211e-01<br />Sepal.Width: 4.174560","density: 1.704590e-01<br />Sepal.Width: 4.179256","density: 1.679455e-01<br />Sepal.Width: 4.183953","density: 1.654684e-01<br />Sepal.Width: 4.188650","density: 1.630476e-01<br />Sepal.Width: 4.193346","density: 1.606730e-01<br />Sepal.Width: 4.198043","density: 1.583318e-01<br />Sepal.Width: 4.202740","density: 1.560403e-01<br />Sepal.Width: 4.207436","density: 1.537929e-01<br />Sepal.Width: 4.212133","density: 1.515760e-01<br />Sepal.Width: 4.216830","density: 1.494030e-01<br />Sepal.Width: 4.221526","density: 1.472719e-01<br />Sepal.Width: 4.226223","density: 1.451686e-01<br />Sepal.Width: 4.230920","density: 1.431041e-01<br />Sepal.Width: 4.235616","density: 1.410790e-01<br />Sepal.Width: 4.240313","density: 1.390792e-01<br />Sepal.Width: 4.245010","density: 1.371133e-01<br />Sepal.Width: 4.249706","density: 1.351843e-01<br />Sepal.Width: 4.254403","density: 1.332781e-01<br />Sepal.Width: 4.259100","density: 1.314011e-01<br />Sepal.Width: 4.263796","density: 1.295584e-01<br />Sepal.Width: 4.268493","density: 1.277357e-01<br />Sepal.Width: 4.273190","density: 1.259380e-01<br />Sepal.Width: 4.277886","density: 1.241712e-01<br />Sepal.Width: 4.282583","density: 1.224219e-01<br />Sepal.Width: 4.287280","density: 1.206934e-01<br />Sepal.Width: 4.291977","density: 1.189922e-01<br />Sepal.Width: 4.296673","density: 1.173057e-01<br />Sepal.Width: 4.301370","density: 1.156359e-01<br />Sepal.Width: 4.306067","density: 1.139895e-01<br />Sepal.Width: 4.310763","density: 1.123549e-01<br />Sepal.Width: 4.315460","density: 1.107333e-01<br />Sepal.Width: 4.320157","density: 1.091306e-01<br />Sepal.Width: 4.324853","density: 1.075370e-01<br />Sepal.Width: 4.329550","density: 1.059528e-01<br />Sepal.Width: 4.334247","density: 1.043830e-01<br />Sepal.Width: 4.338943","density: 1.028195e-01<br />Sepal.Width: 4.343640","density: 1.012622e-01<br />Sepal.Width: 4.348337","density: 9.971469e-02<br />Sepal.Width: 4.353033","density: 9.817095e-02<br />Sepal.Width: 4.357730","density: 9.663093e-02<br />Sepal.Width: 4.362427","density: 9.509593e-02<br />Sepal.Width: 4.367123","density: 9.356262e-02<br />Sepal.Width: 4.371820","density: 9.203082e-02<br />Sepal.Width: 4.376517","density: 9.050022e-02<br />Sepal.Width: 4.381213","density: 8.896942e-02<br />Sepal.Width: 4.385910","density: 8.743834e-02<br />Sepal.Width: 4.390607","density: 8.590569e-02<br />Sepal.Width: 4.395303","density: 8.437136e-02<br />Sepal.Width: 4.400000","density: 8.437136e-02<br />Sepal.Width: 4.400000","density: 8.590569e-02<br />Sepal.Width: 4.395303","density: 8.743834e-02<br />Sepal.Width: 4.390607","density: 8.896942e-02<br />Sepal.Width: 4.385910","density: 9.050022e-02<br />Sepal.Width: 4.381213","density: 9.203082e-02<br />Sepal.Width: 4.376517","density: 9.356262e-02<br />Sepal.Width: 4.371820","density: 9.509593e-02<br />Sepal.Width: 4.367123","density: 9.663093e-02<br />Sepal.Width: 4.362427","density: 9.817095e-02<br />Sepal.Width: 4.357730","density: 9.971469e-02<br />Sepal.Width: 4.353033","density: 1.012622e-01<br />Sepal.Width: 4.348337","density: 1.028195e-01<br />Sepal.Width: 4.343640","density: 1.043830e-01<br />Sepal.Width: 4.338943","density: 1.059528e-01<br />Sepal.Width: 4.334247","density: 1.075370e-01<br />Sepal.Width: 4.329550","density: 1.091306e-01<br />Sepal.Width: 4.324853","density: 1.107333e-01<br />Sepal.Width: 4.320157","density: 1.123549e-01<br />Sepal.Width: 4.315460","density: 1.139895e-01<br />Sepal.Width: 4.310763","density: 1.156359e-01<br />Sepal.Width: 4.306067","density: 1.173057e-01<br />Sepal.Width: 4.301370","density: 1.189922e-01<br />Sepal.Width: 4.296673","density: 1.206934e-01<br />Sepal.Width: 4.291977","density: 1.224219e-01<br />Sepal.Width: 4.287280","density: 1.241712e-01<br />Sepal.Width: 4.282583","density: 1.259380e-01<br />Sepal.Width: 4.277886","density: 1.277357e-01<br />Sepal.Width: 4.273190","density: 1.295584e-01<br />Sepal.Width: 4.268493","density: 1.314011e-01<br />Sepal.Width: 4.263796","density: 1.332781e-01<br />Sepal.Width: 4.259100","density: 1.351843e-01<br />Sepal.Width: 4.254403","density: 1.371133e-01<br />Sepal.Width: 4.249706","density: 1.390792e-01<br />Sepal.Width: 4.245010","density: 1.410790e-01<br />Sepal.Width: 4.240313","density: 1.431041e-01<br />Sepal.Width: 4.235616","density: 1.451686e-01<br />Sepal.Width: 4.230920","density: 1.472719e-01<br />Sepal.Width: 4.226223","density: 1.494030e-01<br />Sepal.Width: 4.221526","density: 1.515760e-01<br />Sepal.Width: 4.216830","density: 1.537929e-01<br />Sepal.Width: 4.212133","density: 1.560403e-01<br />Sepal.Width: 4.207436","density: 1.583318e-01<br />Sepal.Width: 4.202740","density: 1.606730e-01<br />Sepal.Width: 4.198043","density: 1.630476e-01<br />Sepal.Width: 4.193346","density: 1.654684e-01<br />Sepal.Width: 4.188650","density: 1.679455e-01<br />Sepal.Width: 4.183953","density: 1.704590e-01<br />Sepal.Width: 4.179256","density: 1.730211e-01<br />Sepal.Width: 4.174560","density: 1.756465e-01<br />Sepal.Width: 4.169863","density: 1.783119e-01<br />Sepal.Width: 4.165166","density: 1.810280e-01<br />Sepal.Width: 4.160470","density: 1.838156e-01<br />Sepal.Width: 4.155773","density: 1.866468e-01<br />Sepal.Width: 4.151076","density: 1.895311e-01<br />Sepal.Width: 4.146380","density: 1.924956e-01<br />Sepal.Width: 4.141683","density: 1.955078e-01<br />Sepal.Width: 4.136986","density: 1.985751e-01<br />Sepal.Width: 4.132290","density: 2.017324e-01<br />Sepal.Width: 4.127593","density: 2.049414e-01<br />Sepal.Width: 4.122896","density: 2.082075e-01<br />Sepal.Width: 4.118200","density: 2.115737e-01<br />Sepal.Width: 4.113503","density: 2.149957e-01<br />Sepal.Width: 4.108806","density: 2.184763e-01<br />Sepal.Width: 4.104110","density: 2.220673e-01<br />Sepal.Width: 4.099413","density: 2.257182e-01<br />Sepal.Width: 4.094716","density: 2.294291e-01<br />Sepal.Width: 4.090020","density: 2.332595e-01<br />Sepal.Width: 4.085323","density: 2.371538e-01<br />Sepal.Width: 4.080626","density: 2.411116e-01<br />Sepal.Width: 4.075930","density: 2.451915e-01<br />Sepal.Width: 4.071233","density: 2.493418e-01<br />Sepal.Width: 4.066536","density: 2.535582e-01<br />Sepal.Width: 4.061840","density: 2.578977e-01<br />Sepal.Width: 4.057143","density: 2.623130e-01<br />Sepal.Width: 4.052446","density: 2.667964e-01<br />Sepal.Width: 4.047750","density: 2.714016e-01<br />Sepal.Width: 4.043053","density: 2.760871e-01<br />Sepal.Width: 4.038356","density: 2.808413e-01<br />Sepal.Width: 4.033659","density: 2.857137e-01<br />Sepal.Width: 4.028963","density: 2.906694e-01<br />Sepal.Width: 4.024266","density: 2.956930e-01<br />Sepal.Width: 4.019569","density: 3.008289e-01<br />Sepal.Width: 4.014873","density: 3.060490e-01<br />Sepal.Width: 4.010176","density: 3.113346e-01<br />Sepal.Width: 4.005479","density: 3.167242e-01<br />Sepal.Width: 4.000783","density: 3.221967e-01<br />Sepal.Width: 3.996086","density: 3.277306e-01<br />Sepal.Width: 3.991389","density: 3.333581e-01<br />Sepal.Width: 3.986693","density: 3.390645e-01<br />Sepal.Width: 3.981996","density: 3.448265e-01<br />Sepal.Width: 3.977299","density: 3.506700e-01<br />Sepal.Width: 3.972603","density: 3.565856e-01<br />Sepal.Width: 3.967906","density: 3.625497e-01<br />Sepal.Width: 3.963209","density: 3.685814e-01<br />Sepal.Width: 3.958513","density: 3.746761e-01<br />Sepal.Width: 3.953816","density: 3.808107e-01<br />Sepal.Width: 3.949119","density: 3.869983e-01<br />Sepal.Width: 3.944423","density: 3.932371e-01<br />Sepal.Width: 3.939726","density: 3.995063e-01<br />Sepal.Width: 3.935029","density: 4.058137e-01<br />Sepal.Width: 3.930333","density: 4.121586e-01<br />Sepal.Width: 3.925636","density: 4.185237e-01<br />Sepal.Width: 3.920939","density: 4.249127e-01<br />Sepal.Width: 3.916243","density: 4.313240e-01<br />Sepal.Width: 3.911546","density: 4.377453e-01<br />Sepal.Width: 3.906849","density: 4.441772e-01<br />Sepal.Width: 3.902153","density: 4.506156e-01<br />Sepal.Width: 3.897456","density: 4.570543e-01<br />Sepal.Width: 3.892759","density: 4.634920e-01<br />Sepal.Width: 3.888063","density: 4.699207e-01<br />Sepal.Width: 3.883366","density: 4.763410e-01<br />Sepal.Width: 3.878669","density: 4.827508e-01<br />Sepal.Width: 3.873973","density: 4.891376e-01<br />Sepal.Width: 3.869276","density: 4.955088e-01<br />Sepal.Width: 3.864579","density: 5.018626e-01<br />Sepal.Width: 3.859883","density: 5.081817e-01<br />Sepal.Width: 3.855186","density: 5.144801e-01<br />Sepal.Width: 3.850489","density: 5.207571e-01<br />Sepal.Width: 3.845793","density: 5.269911e-01<br />Sepal.Width: 3.841096","density: 5.332019e-01<br />Sepal.Width: 3.836399","density: 5.393893e-01<br />Sepal.Width: 3.831703","density: 5.455312e-01<br />Sepal.Width: 3.827006","density: 5.516493e-01<br />Sepal.Width: 3.822309","density: 5.577442e-01<br />Sepal.Width: 3.817613","density: 5.637975e-01<br />Sepal.Width: 3.812916","density: 5.698287e-01<br />Sepal.Width: 3.808219","density: 5.758397e-01<br />Sepal.Width: 3.803523","density: 5.818177e-01<br />Sepal.Width: 3.798826","density: 5.877787e-01<br />Sepal.Width: 3.794129","density: 5.937252e-01<br />Sepal.Width: 3.789432","density: 5.996514e-01<br />Sepal.Width: 3.784736","density: 6.055690e-01<br />Sepal.Width: 3.780039","density: 6.114805e-01<br />Sepal.Width: 3.775342","density: 6.173875e-01<br />Sepal.Width: 3.770646","density: 6.232976e-01<br />Sepal.Width: 3.765949","density: 6.292120e-01<br />Sepal.Width: 3.761252","density: 6.351398e-01<br />Sepal.Width: 3.756556","density: 6.410856e-01<br />Sepal.Width: 3.751859","density: 6.470474e-01<br />Sepal.Width: 3.747162","density: 6.530413e-01<br />Sepal.Width: 3.742466","density: 6.590702e-01<br />Sepal.Width: 3.737769","density: 6.651277e-01<br />Sepal.Width: 3.733072","density: 6.712353e-01<br />Sepal.Width: 3.728376","density: 6.773963e-01<br />Sepal.Width: 3.723679","density: 6.835983e-01<br />Sepal.Width: 3.718982","density: 6.898665e-01<br />Sepal.Width: 3.714286","density: 6.962065e-01<br />Sepal.Width: 3.709589","density: 7.025987e-01<br />Sepal.Width: 3.704892","density: 7.090700e-01<br />Sepal.Width: 3.700196","density: 7.156302e-01<br />Sepal.Width: 3.695499","density: 7.222516e-01<br />Sepal.Width: 3.690802","density: 7.289608e-01<br />Sepal.Width: 3.686106","density: 7.357727e-01<br />Sepal.Width: 3.681409","density: 7.426520e-01<br />Sepal.Width: 3.676712","density: 7.496227e-01<br />Sepal.Width: 3.672016","density: 7.567050e-01<br />Sepal.Width: 3.667319","density: 7.638569e-01<br />Sepal.Width: 3.662622","density: 7.710983e-01<br />Sepal.Width: 3.657926","density: 7.784538e-01<br />Sepal.Width: 3.653229","density: 7.858763e-01<br />Sepal.Width: 3.648532","density: 7.933808e-01<br />Sepal.Width: 3.643836","density: 8.009935e-01<br />Sepal.Width: 3.639139","density: 8.086658e-01<br />Sepal.Width: 3.634442","density: 8.164070e-01<br />Sepal.Width: 3.629746","density: 8.242412e-01<br />Sepal.Width: 3.625049","density: 8.321220e-01<br />Sepal.Width: 3.620352","density: 8.400539e-01<br />Sepal.Width: 3.615656","density: 8.480530e-01<br />Sepal.Width: 3.610959","density: 8.560806e-01<br />Sepal.Width: 3.606262","density: 8.641374e-01<br />Sepal.Width: 3.601566","density: 8.722250e-01<br />Sepal.Width: 3.596869","density: 8.803178e-01<br />Sepal.Width: 3.592172","density: 8.884151e-01<br />Sepal.Width: 3.587476","density: 8.964958e-01<br />Sepal.Width: 3.582779","density: 9.045544e-01<br />Sepal.Width: 3.578082","density: 9.125900e-01<br />Sepal.Width: 3.573386","density: 9.205545e-01<br />Sepal.Width: 3.568689","density: 9.284647e-01<br />Sepal.Width: 3.563992","density: 9.363208e-01<br />Sepal.Width: 3.559295","density: 9.440497e-01<br />Sepal.Width: 3.554599","density: 9.516865e-01<br />Sepal.Width: 3.549902","density: 9.592361e-01<br />Sepal.Width: 3.545205","density: 9.666029e-01<br />Sepal.Width: 3.540509","density: 9.738354e-01<br />Sepal.Width: 3.535812","density: 9.809469e-01<br />Sepal.Width: 3.531115","density: 9.878233e-01<br />Sepal.Width: 3.526419","density: 9.945203e-01<br />Sepal.Width: 3.521722","density: 1.001063e+00<br />Sepal.Width: 3.517025","density: 1.007324e+00<br />Sepal.Width: 3.512329","density: 1.013360e+00<br />Sepal.Width: 3.507632","density: 1.019211e+00<br />Sepal.Width: 3.502935","density: 1.024741e+00<br />Sepal.Width: 3.498239","density: 1.030001e+00<br />Sepal.Width: 3.493542","density: 1.035049e+00<br />Sepal.Width: 3.488845","density: 1.039748e+00<br />Sepal.Width: 3.484149","density: 1.044134e+00<br />Sepal.Width: 3.479452","density: 1.048287e+00<br />Sepal.Width: 3.474755","density: 1.052071e+00<br />Sepal.Width: 3.470059","density: 1.055507e+00<br />Sepal.Width: 3.465362","density: 1.058695e+00<br />Sepal.Width: 3.460665","density: 1.061506e+00<br />Sepal.Width: 3.455969","density: 1.063942e+00<br />Sepal.Width: 3.451272","density: 1.066121e+00<br />Sepal.Width: 3.446575","density: 1.067927e+00<br />Sepal.Width: 3.441879","density: 1.069341e+00<br />Sepal.Width: 3.437182","density: 1.070495e+00<br />Sepal.Width: 3.432485","density: 1.071293e+00<br />Sepal.Width: 3.427789","density: 1.071690e+00<br />Sepal.Width: 3.423092","density: 1.071835e+00<br />Sepal.Width: 3.418395","density: 1.071648e+00<br />Sepal.Width: 3.413699","density: 1.071066e+00<br />Sepal.Width: 3.409002","density: 1.070246e+00<br />Sepal.Width: 3.404305","density: 1.069126e+00<br />Sepal.Width: 3.399609","density: 1.067628e+00<br />Sepal.Width: 3.394912","density: 1.065914e+00<br />Sepal.Width: 3.390215","density: 1.063938e+00<br />Sepal.Width: 3.385519","density: 1.061613e+00<br />Sepal.Width: 3.380822","density: 1.059098e+00<br />Sepal.Width: 3.376125","density: 1.056365e+00<br />Sepal.Width: 3.371429","density: 1.053323e+00<br />Sepal.Width: 3.366732","density: 1.050121e+00<br />Sepal.Width: 3.362035","density: 1.046745e+00<br />Sepal.Width: 3.357339","density: 1.043110e+00<br />Sepal.Width: 3.352642","density: 1.039348e+00<br />Sepal.Width: 3.347945","density: 1.035455e+00<br />Sepal.Width: 3.343249","density: 1.031360e+00<br />Sepal.Width: 3.338552","density: 1.027173e+00<br />Sepal.Width: 3.333855","density: 1.022895e+00<br />Sepal.Width: 3.329159","density: 1.018475e+00<br />Sepal.Width: 3.324462","density: 1.013998e+00<br />Sepal.Width: 3.319765","density: 1.009465e+00<br />Sepal.Width: 3.315068","density: 1.004854e+00<br />Sepal.Width: 3.310372","density: 1.000215e+00<br />Sepal.Width: 3.305675","density: 9.955532e-01<br />Sepal.Width: 3.300978","density: 9.908701e-01<br />Sepal.Width: 3.296282","density: 9.861893e-01<br />Sepal.Width: 3.291585","density: 9.815125e-01<br />Sepal.Width: 3.286888","density: 9.768613e-01<br />Sepal.Width: 3.282192","density: 9.722392e-01<br />Sepal.Width: 3.277495","density: 9.676444e-01<br />Sepal.Width: 3.272798","density: 9.631099e-01<br />Sepal.Width: 3.268102","density: 9.586270e-01<br />Sepal.Width: 3.263405","density: 9.541885e-01<br />Sepal.Width: 3.258708","density: 9.498331e-01<br />Sepal.Width: 3.254012","density: 9.455458e-01<br />Sepal.Width: 3.249315","density: 9.413136e-01<br />Sepal.Width: 3.244618","density: 9.371750e-01<br />Sepal.Width: 3.239922","density: 9.331137e-01<br />Sepal.Width: 3.235225","density: 9.291113e-01<br />Sepal.Width: 3.230528","density: 9.252018e-01<br />Sepal.Width: 3.225832","density: 9.213705e-01<br />Sepal.Width: 3.221135","density: 9.175951e-01<br />Sepal.Width: 3.216438","density: 9.139020e-01<br />Sepal.Width: 3.211742","density: 9.102788e-01<br />Sepal.Width: 3.207045","density: 9.067019e-01<br />Sepal.Width: 3.202348","density: 9.031886e-01<br />Sepal.Width: 3.197652","density: 8.997277e-01<br />Sepal.Width: 3.192955","density: 8.962979e-01<br />Sepal.Width: 3.188258","density: 8.929064e-01<br />Sepal.Width: 3.183562","density: 8.895411e-01<br />Sepal.Width: 3.178865","density: 8.861867e-01<br />Sepal.Width: 3.174168","density: 8.828410e-01<br />Sepal.Width: 3.169472","density: 8.794871e-01<br />Sepal.Width: 3.164775","density: 8.761202e-01<br />Sepal.Width: 3.160078","density: 8.727303e-01<br />Sepal.Width: 3.155382","density: 8.692910e-01<br />Sepal.Width: 3.150685","density: 8.658122e-01<br />Sepal.Width: 3.145988","density: 8.622786e-01<br />Sepal.Width: 3.141292","density: 8.586488e-01<br />Sepal.Width: 3.136595","density: 8.549521e-01<br />Sepal.Width: 3.131898","density: 8.511702e-01<br />Sepal.Width: 3.127202","density: 8.472422e-01<br />Sepal.Width: 3.122505","density: 8.432198e-01<br />Sepal.Width: 3.117808","density: 8.390852e-01<br />Sepal.Width: 3.113112","density: 8.347533e-01<br />Sepal.Width: 3.108415","density: 8.303010e-01<br />Sepal.Width: 3.103718","density: 8.257140e-01<br />Sepal.Width: 3.099022","density: 8.208796e-01<br />Sepal.Width: 3.094325","density: 8.159014e-01<br />Sepal.Width: 3.089628","density: 8.107715e-01<br />Sepal.Width: 3.084932","density: 8.053474e-01<br />Sepal.Width: 3.080235","density: 7.997600e-01<br />Sepal.Width: 3.075538","density: 7.940087e-01<br />Sepal.Width: 3.070841","density: 7.879245e-01<br />Sepal.Width: 3.066145","density: 7.816609e-01<br />Sepal.Width: 3.061448","density: 7.752189e-01<br />Sepal.Width: 3.056751","density: 7.684305e-01<br />Sepal.Width: 3.052055","density: 7.614434e-01<br />Sepal.Width: 3.047358","density: 7.542686e-01<br />Sepal.Width: 3.042661","density: 7.467451e-01<br />Sepal.Width: 3.037965","density: 7.390089e-01<br />Sepal.Width: 3.033268","density: 7.310815e-01<br />Sepal.Width: 3.028571","density: 7.228141e-01<br />Sepal.Width: 3.023875","density: 7.143262e-01<br />Sepal.Width: 3.019178","density: 7.056499e-01<br />Sepal.Width: 3.014481","density: 6.966524e-01<br />Sepal.Width: 3.009785","density: 6.874339e-01<br />Sepal.Width: 3.005088","density: 6.780358e-01<br />Sepal.Width: 3.000391","density: 6.683443e-01<br />Sepal.Width: 2.995695","density: 6.584393e-01<br />Sepal.Width: 2.990998","density: 6.483692e-01<br />Sepal.Width: 2.986301","density: 6.380413e-01<br />Sepal.Width: 2.981605","density: 6.275154e-01<br />Sepal.Width: 2.976908","density: 6.168445e-01<br />Sepal.Width: 2.972211","density: 6.059570e-01<br />Sepal.Width: 2.967515","density: 5.948955e-01<br />Sepal.Width: 2.962818","density: 5.837134e-01<br />Sepal.Width: 2.958121","density: 5.723600e-01<br />Sepal.Width: 2.953425","density: 5.608643e-01<br />Sepal.Width: 2.948728","density: 5.492763e-01<br />Sepal.Width: 2.944031","density: 5.375644e-01<br />Sepal.Width: 2.939335","density: 5.257489e-01<br />Sepal.Width: 2.934638","density: 5.138721e-01<br />Sepal.Width: 2.929941","density: 5.019190e-01<br />Sepal.Width: 2.925245","density: 4.899069e-01<br />Sepal.Width: 2.920548","density: 4.778661e-01<br />Sepal.Width: 2.915851","density: 4.657951e-01<br />Sepal.Width: 2.911155","density: 4.537145e-01<br />Sepal.Width: 2.906458","density: 4.416381e-01<br />Sepal.Width: 2.901761","density: 4.295747e-01<br />Sepal.Width: 2.897065","density: 4.175538e-01<br />Sepal.Width: 2.892368","density: 4.055697e-01<br />Sepal.Width: 2.887671","density: 3.936374e-01<br />Sepal.Width: 2.882975","density: 3.818012e-01<br />Sepal.Width: 2.878278","density: 3.700327e-01<br />Sepal.Width: 2.873581","density: 3.583493e-01<br />Sepal.Width: 2.868885","density: 3.468153e-01<br />Sepal.Width: 2.864188","density: 3.353774e-01<br />Sepal.Width: 2.859491","density: 3.240520e-01<br />Sepal.Width: 2.854795","density: 3.129273e-01<br />Sepal.Width: 2.850098","density: 3.019238e-01<br />Sepal.Width: 2.845401","density: 2.910538e-01<br />Sepal.Width: 2.840705","density: 2.804323e-01<br />Sepal.Width: 2.836008","density: 2.699532e-01<br />Sepal.Width: 2.831311","density: 2.596226e-01<br />Sepal.Width: 2.826614","density: 2.495829e-01<br />Sepal.Width: 2.821918","density: 2.397030e-01<br />Sepal.Width: 2.817221","density: 2.299833e-01<br />Sepal.Width: 2.812524","density: 2.205850e-01<br />Sepal.Width: 2.807828","density: 2.113625e-01<br />Sepal.Width: 2.803131","density: 2.023127e-01<br />Sepal.Width: 2.798434","density: 1.935952e-01<br />Sepal.Width: 2.793738","density: 1.850714e-01<br />Sepal.Width: 2.789041","density: 1.767289e-01<br />Sepal.Width: 2.784344","density: 1.687207e-01<br />Sepal.Width: 2.779648","density: 1.609210e-01<br />Sepal.Width: 2.774951","density: 1.533066e-01<br />Sepal.Width: 2.770254","density: 1.460214e-01<br />Sepal.Width: 2.765558","density: 1.389554e-01<br />Sepal.Width: 2.760861","density: 1.320752e-01<br />Sepal.Width: 2.756164","density: 1.255127e-01<br />Sepal.Width: 2.751468","density: 1.191763e-01<br />Sepal.Width: 2.746771","density: 1.130229e-01<br />Sepal.Width: 2.742074","density: 1.071707e-01<br />Sepal.Width: 2.737378","density: 1.015476e-01<br />Sepal.Width: 2.732681","density: 9.610179e-02<br />Sepal.Width: 2.727984","density: 9.093716e-02<br />Sepal.Width: 2.723288","density: 8.600122e-02<br />Sepal.Width: 2.718591","density: 8.123440e-02<br />Sepal.Width: 2.713894","density: 7.672639e-02<br />Sepal.Width: 2.709198","density: 7.244334e-02<br />Sepal.Width: 2.704501","density: 6.831967e-02<br />Sepal.Width: 2.699804","density: 6.443108e-02<br />Sepal.Width: 2.695108","density: 6.076101e-02<br />Sepal.Width: 2.690411","density: 5.723941e-02<br />Sepal.Width: 2.685714","density: 5.392885e-02<br />Sepal.Width: 2.681018","density: 5.082815e-02<br />Sepal.Width: 2.676321","density: 4.786437e-02<br />Sepal.Width: 2.671624","density: 4.508803e-02<br />Sepal.Width: 2.666928","density: 4.251126e-02<br />Sepal.Width: 2.662231","density: 4.005965e-02<br />Sepal.Width: 2.657534","density: 3.777295e-02<br />Sepal.Width: 2.652838","density: 3.567441e-02<br />Sepal.Width: 2.648141","density: 3.368945e-02<br />Sepal.Width: 2.643444","density: 3.184837e-02<br />Sepal.Width: 2.638748","density: 3.018342e-02<br />Sepal.Width: 2.634051","density: 2.862090e-02<br />Sepal.Width: 2.629354","density: 2.718302e-02<br />Sepal.Width: 2.624658","density: 2.590902e-02<br />Sepal.Width: 2.619961","density: 2.472695e-02<br />Sepal.Width: 2.615264","density: 2.365218e-02<br />Sepal.Width: 2.610568","density: 2.272915e-02<br />Sepal.Width: 2.605871","density: 2.188832e-02<br />Sepal.Width: 2.601174","density: 2.113935e-02<br />Sepal.Width: 2.596477","density: 2.053029e-02<br />Sepal.Width: 2.591781","density: 1.999452e-02<br />Sepal.Width: 2.587084","density: 1.953696e-02<br />Sepal.Width: 2.582387","density: 1.920795e-02<br />Sepal.Width: 2.577691","density: 1.894404e-02<br />Sepal.Width: 2.572994","density: 1.874639e-02<br />Sepal.Width: 2.568297","density: 1.866634e-02<br />Sepal.Width: 2.563601","density: 1.864391e-02<br />Sepal.Width: 2.558904","density: 1.867891e-02<br />Sepal.Width: 2.554207","density: 1.881756e-02<br />Sepal.Width: 2.549511","density: 1.900857e-02<br />Sepal.Width: 2.544814","density: 1.925010e-02<br />Sepal.Width: 2.540117","density: 1.958021e-02<br />Sepal.Width: 2.535421","density: 1.995849e-02<br />Sepal.Width: 2.530724","density: 2.038076e-02<br />Sepal.Width: 2.526027","density: 2.087781e-02<br />Sepal.Width: 2.521331","density: 2.141843e-02<br />Sepal.Width: 2.516634","density: 2.199677e-02<br />Sepal.Width: 2.511937","density: 2.263706e-02<br />Sepal.Width: 2.507241","density: 2.331579e-02<br />Sepal.Width: 2.502544","density: 2.402606e-02<br />Sepal.Width: 2.497847","density: 2.478616e-02<br />Sepal.Width: 2.493151","density: 2.557893e-02<br />Sepal.Width: 2.488454","density: 2.639704e-02<br />Sepal.Width: 2.483757","density: 2.725340e-02<br />Sepal.Width: 2.479061","density: 2.813587e-02<br />Sepal.Width: 2.474364","density: 2.903736e-02<br />Sepal.Width: 2.469667","density: 2.996598e-02<br />Sepal.Width: 2.464971","density: 3.091328e-02<br />Sepal.Width: 2.460274","density: 3.187312e-02<br />Sepal.Width: 2.455577","density: 3.284935e-02<br />Sepal.Width: 2.450881","density: 3.383596e-02<br />Sepal.Width: 2.446184","density: 3.482846e-02<br />Sepal.Width: 2.441487","density: 3.582709e-02<br />Sepal.Width: 2.436791","density: 3.682690e-02<br />Sepal.Width: 2.432094","density: 3.782584e-02<br />Sepal.Width: 2.427397","density: 3.882119e-02<br />Sepal.Width: 2.422701","density: 3.980778e-02<br />Sepal.Width: 2.418004","density: 4.078670e-02<br />Sepal.Width: 2.413307","density: 4.175304e-02<br />Sepal.Width: 2.408611","density: 4.270005e-02<br />Sepal.Width: 2.403914","density: 4.363274e-02<br />Sepal.Width: 2.399217","density: 4.454473e-02<br />Sepal.Width: 2.394521","density: 4.542650e-02<br />Sepal.Width: 2.389824","density: 4.628755e-02<br />Sepal.Width: 2.385127","density: 4.712087e-02<br />Sepal.Width: 2.380431","density: 4.791305e-02<br />Sepal.Width: 2.375734","density: 4.867858e-02<br />Sepal.Width: 2.371037","density: 4.941061e-02<br />Sepal.Width: 2.366341","density: 5.009090e-02<br />Sepal.Width: 2.361644","density: 5.073930e-02<br />Sepal.Width: 2.356947","density: 5.134978e-02<br />Sepal.Width: 2.352250","density: 5.189871e-02<br />Sepal.Width: 2.347554","density: 5.241135e-02<br />Sepal.Width: 2.342857","density: 5.288311e-02<br />Sepal.Width: 2.338160","density: 5.328467e-02<br />Sepal.Width: 2.333464","density: 5.364657e-02<br />Sepal.Width: 2.328767","density: 5.396613e-02<br />Sepal.Width: 2.324070","density: 5.420841e-02<br />Sepal.Width: 2.319374","density: 5.440881e-02<br />Sepal.Width: 2.314677","density: 5.456686e-02<br />Sepal.Width: 2.309980","density: 5.464248e-02<br />Sepal.Width: 2.305284","density: 5.467522e-02<br />Sepal.Width: 2.300587","density: 5.466509e-02<br />Sepal.Width: 2.295890","density: 5.457333e-02<br />Sepal.Width: 2.291194","density: 5.443711e-02<br />Sepal.Width: 2.286497","density: 5.425834e-02<br />Sepal.Width: 2.281800","density: 5.400183e-02<br />Sepal.Width: 2.277104","density: 5.370018e-02<br />Sepal.Width: 2.272407","density: 5.335754e-02<br />Sepal.Width: 2.267710","density: 5.294302e-02<br />Sepal.Width: 2.263014","density: 5.248411e-02<br />Sepal.Width: 2.258317","density: 5.198693e-02<br />Sepal.Width: 2.253620","density: 5.142535e-02<br />Sepal.Width: 2.248924","density: 5.082158e-02<br />Sepal.Width: 2.244227","density: 5.018330e-02<br />Sepal.Width: 2.239530","density: 4.948931e-02<br />Sepal.Width: 2.234834","density: 4.875676e-02<br />Sepal.Width: 2.230137","density: 4.799428e-02<br />Sepal.Width: 2.225440","density: 4.718558e-02<br />Sepal.Width: 2.220744","density: 4.634324e-02<br />Sepal.Width: 2.216047","density: 4.547620e-02<br />Sepal.Width: 2.211350","density: 4.457275e-02<br />Sepal.Width: 2.206654","density: 4.364171e-02<br />Sepal.Width: 2.201957","density: 4.269162e-02<br />Sepal.Width: 2.197260","density: 4.171486e-02<br />Sepal.Width: 2.192564","density: 4.071744e-02<br />Sepal.Width: 2.187867","density: 3.970677e-02<br />Sepal.Width: 2.183170","density: 3.867875e-02<br />Sepal.Width: 2.178474","density: 3.763760e-02<br />Sepal.Width: 2.173777","density: 3.658895e-02<br />Sepal.Width: 2.169080","density: 3.553151e-02<br />Sepal.Width: 2.164384","density: 3.446876e-02<br />Sepal.Width: 2.159687","density: 3.340401e-02<br />Sepal.Width: 2.154990","density: 3.233804e-02<br />Sepal.Width: 2.150294","density: 3.127457e-02<br />Sepal.Width: 2.145597","density: 3.021417e-02<br />Sepal.Width: 2.140900","density: 2.915895e-02<br />Sepal.Width: 2.136204","density: 2.811374e-02<br />Sepal.Width: 2.131507","density: 2.707608e-02<br />Sepal.Width: 2.126810","density: 2.604878e-02<br />Sepal.Width: 2.122114","density: 2.503844e-02<br />Sepal.Width: 2.117417","density: 2.403944e-02<br />Sepal.Width: 2.112720","density: 2.305471e-02<br />Sepal.Width: 2.108023","density: 2.209311e-02<br />Sepal.Width: 2.103327","density: 2.114590e-02<br />Sepal.Width: 2.098630","density: 2.021566e-02<br />Sepal.Width: 2.093933","density: 1.931377e-02<br />Sepal.Width: 2.089237","density: 1.842855e-02<br />Sepal.Width: 2.084540","density: 1.756189e-02<br />Sepal.Width: 2.079843","density: 1.672774e-02<br />Sepal.Width: 2.075147","density: 1.591180e-02<br />Sepal.Width: 2.070450","density: 1.511504e-02<br />Sepal.Width: 2.065753","density: 1.435382e-02<br />Sepal.Width: 2.061057","density: 1.361164e-02<br />Sepal.Width: 2.056360","density: 1.288851e-02<br />Sepal.Width: 2.051663","density: 1.220273e-02<br />Sepal.Width: 2.046967","density: 1.153624e-02<br />Sepal.Width: 2.042270","density: 1.088899e-02<br />Sepal.Width: 2.037573","density: 1.027792e-02<br />Sepal.Width: 2.032877","density: 9.686821e-03<br />Sepal.Width: 2.028180","density: 9.114602e-03<br />Sepal.Width: 2.023483","density: 8.576533e-03<br />Sepal.Width: 2.018787","density: 8.058621e-03<br />Sepal.Width: 2.014090","density: 7.558796e-03<br />Sepal.Width: 2.009393","density: 7.090487e-03<br />Sepal.Width: 2.004697","density: 6.642056e-03<br />Sepal.Width: 2.000000","density: 6.642056e-03<br />Sepal.Width: 2.000000"],"key":["1","2","3","4","5","6","7","8","9","10","11","12","13","14","15","16","17","18","19","20","21","22","23","24","25","26","27","28","29","30","31","32","33","34","35","36","37","38","39","40","41","42","43","44","45","46","47","48","49","50"],"type":"scatter","mode":"lines","line":{"width":1.88976377952756,"color":"rgba(0,0,0,1)","dash":"solid"},"fill":"toself","fillcolor":"rgba(248,118,109,1)","hoveron":"points","set":"SharedData91833e3a","name":"setosa","legendgroup":"setosa","showlegend":true,"xaxis":"x2","yaxis":"y7","hoverinfo":"text","_isSimpleKey":true,"_isNestedKey":false,"frame":null},{"x":[2,2.00469667318982,2.00939334637965,2.01409001956947,2.0187866927593,2.02348336594912,2.02818003913894,2.03287671232877,2.03757338551859,2.04227005870841,2.04696673189824,2.05166340508806,2.05636007827789,2.06105675146771,2.06575342465753,2.07045009784736,2.07514677103718,2.07984344422701,2.08454011741683,2.08923679060665,2.09393346379648,2.0986301369863,2.10332681017613,2.10802348336595,2.11272015655577,2.1174168297456,2.12211350293542,2.12681017612524,2.13150684931507,2.13620352250489,2.14090019569472,2.14559686888454,2.15029354207436,2.15499021526419,2.15968688845401,2.16438356164384,2.16908023483366,2.17377690802348,2.17847358121331,2.18317025440313,2.18786692759295,2.19256360078278,2.1972602739726,2.20195694716243,2.20665362035225,2.21135029354207,2.2160469667319,2.22074363992172,2.22544031311155,2.23013698630137,2.23483365949119,2.23953033268102,2.24422700587084,2.24892367906067,2.25362035225049,2.25831702544031,2.26301369863014,2.26771037181996,2.27240704500978,2.27710371819961,2.28180039138943,2.28649706457926,2.29119373776908,2.2958904109589,2.30058708414873,2.30528375733855,2.30998043052838,2.3146771037182,2.31937377690802,2.32407045009785,2.32876712328767,2.3334637964775,2.33816046966732,2.34285714285714,2.34755381604697,2.35225048923679,2.35694716242661,2.36164383561644,2.36634050880626,2.37103718199609,2.37573385518591,2.38043052837573,2.38512720156556,2.38982387475538,2.39452054794521,2.39921722113503,2.40391389432485,2.40861056751468,2.4133072407045,2.41800391389432,2.42270058708415,2.42739726027397,2.4320939334638,2.43679060665362,2.44148727984344,2.44618395303327,2.45088062622309,2.45557729941292,2.46027397260274,2.46497064579256,2.46966731898239,2.47436399217221,2.47906066536204,2.48375733855186,2.48845401174168,2.49315068493151,2.49784735812133,2.50254403131115,2.50724070450098,2.5119373776908,2.51663405088063,2.52133072407045,2.52602739726027,2.5307240704501,2.53542074363992,2.54011741682975,2.54481409001957,2.54951076320939,2.55420743639922,2.55890410958904,2.56360078277886,2.56829745596869,2.57299412915851,2.57769080234834,2.58238747553816,2.58708414872798,2.59178082191781,2.59647749510763,2.60117416829746,2.60587084148728,2.6105675146771,2.61526418786693,2.61996086105675,2.62465753424658,2.6293542074364,2.63405088062622,2.63874755381605,2.64344422700587,2.64814090019569,2.65283757338552,2.65753424657534,2.66223091976517,2.66692759295499,2.67162426614481,2.67632093933464,2.68101761252446,2.68571428571429,2.69041095890411,2.69510763209393,2.69980430528376,2.70450097847358,2.70919765166341,2.71389432485323,2.71859099804305,2.72328767123288,2.7279843444227,2.73268101761252,2.73737769080235,2.74207436399217,2.746771037182,2.75146771037182,2.75616438356164,2.76086105675147,2.76555772994129,2.77025440313112,2.77495107632094,2.77964774951076,2.78434442270059,2.78904109589041,2.79373776908024,2.79843444227006,2.80313111545988,2.80782778864971,2.81252446183953,2.81722113502935,2.82191780821918,2.826614481409,2.83131115459883,2.83600782778865,2.84070450097847,2.8454011741683,2.85009784735812,2.85479452054795,2.85949119373777,2.86418786692759,2.86888454011742,2.87358121330724,2.87827788649706,2.88297455968689,2.88767123287671,2.89236790606654,2.89706457925636,2.90176125244618,2.90645792563601,2.91115459882583,2.91585127201566,2.92054794520548,2.9252446183953,2.92994129158513,2.93463796477495,2.93933463796477,2.9440313111546,2.94872798434442,2.95342465753425,2.95812133072407,2.96281800391389,2.96751467710372,2.97221135029354,2.97690802348337,2.98160469667319,2.98630136986301,2.99099804305284,2.99569471624266,3.00039138943249,3.00508806262231,3.00978473581213,3.01448140900196,3.01917808219178,3.0238747553816,3.02857142857143,3.03326810176125,3.03796477495108,3.0426614481409,3.04735812133072,3.05205479452055,3.05675146771037,3.0614481409002,3.06614481409002,3.07084148727984,3.07553816046967,3.08023483365949,3.08493150684932,3.08962818003914,3.09432485322896,3.09902152641879,3.10371819960861,3.10841487279843,3.11311154598826,3.11780821917808,3.12250489236791,3.12720156555773,3.13189823874755,3.13659491193738,3.1412915851272,3.14598825831703,3.15068493150685,3.15538160469667,3.1600782778865,3.16477495107632,3.16947162426615,3.17416829745597,3.17886497064579,3.18356164383562,3.18825831702544,3.19295499021526,3.19765166340509,3.20234833659491,3.20704500978474,3.21174168297456,3.21643835616438,3.22113502935421,3.22583170254403,3.23052837573386,3.23522504892368,3.2399217221135,3.24461839530333,3.24931506849315,3.25401174168297,3.2587084148728,3.26340508806262,3.26810176125245,3.27279843444227,3.27749510763209,3.28219178082192,3.28688845401174,3.29158512720157,3.29628180039139,3.30097847358121,3.30567514677104,3.31037181996086,3.31506849315068,3.31976516634051,3.32446183953033,3.32915851272016,3.33385518590998,3.3385518590998,3.34324853228963,3.34794520547945,3.35264187866928,3.3573385518591,3.36203522504892,3.36673189823875,3.37142857142857,3.3761252446184,3.38082191780822,3.38551859099804,3.39021526418787,3.39491193737769,3.39960861056751,3.40430528375734,3.40900195694716,3.41369863013699,3.41839530332681,3.42309197651663,3.42778864970646,3.43248532289628,3.43718199608611,3.44187866927593,3.44657534246575,3.45127201565558,3.4559686888454,3.46066536203523,3.46536203522505,3.47005870841487,3.4747553816047,3.47945205479452,3.48414872798434,3.48884540117417,3.49354207436399,3.49823874755382,3.50293542074364,3.50763209393346,3.51232876712329,3.51702544031311,3.52172211350294,3.52641878669276,3.53111545988258,3.53581213307241,3.54050880626223,3.54520547945206,3.54990215264188,3.5545988258317,3.55929549902153,3.56399217221135,3.56868884540117,3.573385518591,3.57808219178082,3.58277886497065,3.58747553816047,3.59217221135029,3.59686888454012,3.60156555772994,3.60626223091977,3.61095890410959,3.61565557729941,3.62035225048924,3.62504892367906,3.62974559686888,3.63444227005871,3.63913894324853,3.64383561643836,3.64853228962818,3.653228962818,3.65792563600783,3.66262230919765,3.66731898238748,3.6720156555773,3.67671232876712,3.68140900195695,3.68610567514677,3.6908023483366,3.69549902152642,3.70019569471624,3.70489236790607,3.70958904109589,3.71428571428571,3.71898238747554,3.72367906066536,3.72837573385519,3.73307240704501,3.73776908023483,3.74246575342466,3.74716242661448,3.75185909980431,3.75655577299413,3.76125244618395,3.76594911937378,3.7706457925636,3.77534246575342,3.78003913894325,3.78473581213307,3.7894324853229,3.79412915851272,3.79882583170254,3.80352250489237,3.80821917808219,3.81291585127202,3.81761252446184,3.82230919765166,3.82700587084149,3.83170254403131,3.83639921722114,3.84109589041096,3.84579256360078,3.85048923679061,3.85518590998043,3.85988258317026,3.86457925636008,3.8692759295499,3.87397260273973,3.87866927592955,3.88336594911937,3.8880626223092,3.89275929549902,3.89745596868885,3.90215264187867,3.90684931506849,3.91154598825832,3.91624266144814,3.92093933463797,3.92563600782779,3.93033268101761,3.93502935420744,3.93972602739726,3.94442270058708,3.94911937377691,3.95381604696673,3.95851272015656,3.96320939334638,3.9679060665362,3.97260273972603,3.97729941291585,3.98199608610568,3.9866927592955,3.99138943248532,3.99608610567515,4.00078277886497,4.00547945205479,4.01017612524462,4.01487279843444,4.01956947162427,4.02426614481409,4.02896281800391,4.03365949119374,4.03835616438356,4.04305283757339,4.04774951076321,4.05244618395303,4.05714285714286,4.06183953033268,4.0665362035225,4.07123287671233,4.07592954990215,4.08062622309198,4.0853228962818,4.09001956947163,4.09471624266145,4.09941291585127,4.1041095890411,4.10880626223092,4.11350293542074,4.11819960861057,4.12289628180039,4.12759295499022,4.13228962818004,4.13698630136986,4.14168297455969,4.14637964774951,4.15107632093934,4.15577299412916,4.16046966731898,4.16516634050881,4.16986301369863,4.17455968688845,4.17925636007828,4.1839530332681,4.18864970645793,4.19334637964775,4.19804305283757,4.2027397260274,4.20743639921722,4.21213307240705,4.21682974559687,4.22152641878669,4.22622309197652,4.23091976516634,4.23561643835616,4.24031311154599,4.24500978473581,4.24970645792564,4.25440313111546,4.25909980430528,4.26379647749511,4.26849315068493,4.27318982387476,4.27788649706458,4.2825831702544,4.28727984344423,4.29197651663405,4.29667318982388,4.3013698630137,4.30606653620352,4.31076320939335,4.31545988258317,4.32015655577299,4.32485322896282,4.32954990215264,4.33424657534247,4.33894324853229,4.34363992172211,4.34833659491194,4.35303326810176,4.35772994129159,4.36242661448141,4.36712328767123,4.37181996086106,4.37651663405088,4.3812133072407,4.38590998043053,4.39060665362035,4.39530332681018,4.4,4.4,4.39530332681018,4.39060665362035,4.38590998043053,4.3812133072407,4.37651663405088,4.37181996086106,4.36712328767123,4.36242661448141,4.35772994129159,4.35303326810176,4.34833659491194,4.34363992172211,4.33894324853229,4.33424657534247,4.32954990215264,4.32485322896282,4.32015655577299,4.31545988258317,4.31076320939335,4.30606653620352,4.3013698630137,4.29667318982388,4.29197651663405,4.28727984344423,4.2825831702544,4.27788649706458,4.27318982387476,4.26849315068493,4.26379647749511,4.25909980430528,4.25440313111546,4.24970645792564,4.24500978473581,4.24031311154599,4.23561643835616,4.23091976516634,4.22622309197652,4.22152641878669,4.21682974559687,4.21213307240705,4.20743639921722,4.2027397260274,4.19804305283757,4.19334637964775,4.18864970645793,4.1839530332681,4.17925636007828,4.17455968688845,4.16986301369863,4.16516634050881,4.16046966731898,4.15577299412916,4.15107632093934,4.14637964774951,4.14168297455969,4.13698630136986,4.13228962818004,4.12759295499022,4.12289628180039,4.11819960861057,4.11350293542074,4.10880626223092,4.1041095890411,4.09941291585127,4.09471624266145,4.09001956947163,4.0853228962818,4.08062622309198,4.07592954990215,4.07123287671233,4.0665362035225,4.06183953033268,4.05714285714286,4.05244618395303,4.04774951076321,4.04305283757339,4.03835616438356,4.03365949119374,4.02896281800391,4.02426614481409,4.01956947162427,4.01487279843444,4.01017612524462,4.00547945205479,4.00078277886497,3.99608610567515,3.99138943248532,3.9866927592955,3.98199608610568,3.97729941291585,3.97260273972603,3.9679060665362,3.96320939334638,3.95851272015656,3.95381604696673,3.94911937377691,3.94442270058708,3.93972602739726,3.93502935420744,3.93033268101761,3.92563600782779,3.92093933463797,3.91624266144814,3.91154598825832,3.90684931506849,3.90215264187867,3.89745596868885,3.89275929549902,3.8880626223092,3.88336594911937,3.87866927592955,3.87397260273973,3.8692759295499,3.86457925636008,3.85988258317026,3.85518590998043,3.85048923679061,3.84579256360078,3.84109589041096,3.83639921722114,3.83170254403131,3.82700587084149,3.82230919765166,3.81761252446184,3.81291585127202,3.80821917808219,3.80352250489237,3.79882583170254,3.79412915851272,3.7894324853229,3.78473581213307,3.78003913894325,3.77534246575342,3.7706457925636,3.76594911937378,3.76125244618395,3.75655577299413,3.75185909980431,3.74716242661448,3.74246575342466,3.73776908023483,3.73307240704501,3.72837573385519,3.72367906066536,3.71898238747554,3.71428571428571,3.70958904109589,3.70489236790607,3.70019569471624,3.69549902152642,3.6908023483366,3.68610567514677,3.68140900195695,3.67671232876712,3.6720156555773,3.66731898238748,3.66262230919765,3.65792563600783,3.653228962818,3.64853228962818,3.64383561643836,3.63913894324853,3.63444227005871,3.62974559686888,3.62504892367906,3.62035225048924,3.61565557729941,3.61095890410959,3.60626223091977,3.60156555772994,3.59686888454012,3.59217221135029,3.58747553816047,3.58277886497065,3.57808219178082,3.573385518591,3.56868884540117,3.56399217221135,3.55929549902153,3.5545988258317,3.54990215264188,3.54520547945206,3.54050880626223,3.53581213307241,3.53111545988258,3.52641878669276,3.52172211350294,3.51702544031311,3.51232876712329,3.50763209393346,3.50293542074364,3.49823874755382,3.49354207436399,3.48884540117417,3.48414872798434,3.47945205479452,3.4747553816047,3.47005870841487,3.46536203522505,3.46066536203523,3.4559686888454,3.45127201565558,3.44657534246575,3.44187866927593,3.43718199608611,3.43248532289628,3.42778864970646,3.42309197651663,3.41839530332681,3.41369863013699,3.40900195694716,3.40430528375734,3.39960861056751,3.39491193737769,3.39021526418787,3.38551859099804,3.38082191780822,3.3761252446184,3.37142857142857,3.36673189823875,3.36203522504892,3.3573385518591,3.35264187866928,3.34794520547945,3.34324853228963,3.3385518590998,3.33385518590998,3.32915851272016,3.32446183953033,3.31976516634051,3.31506849315068,3.31037181996086,3.30567514677104,3.30097847358121,3.29628180039139,3.29158512720157,3.28688845401174,3.28219178082192,3.27749510763209,3.27279843444227,3.26810176125245,3.26340508806262,3.2587084148728,3.25401174168297,3.24931506849315,3.24461839530333,3.2399217221135,3.23522504892368,3.23052837573386,3.22583170254403,3.22113502935421,3.21643835616438,3.21174168297456,3.20704500978474,3.20234833659491,3.19765166340509,3.19295499021526,3.18825831702544,3.18356164383562,3.17886497064579,3.17416829745597,3.16947162426615,3.16477495107632,3.1600782778865,3.15538160469667,3.15068493150685,3.14598825831703,3.1412915851272,3.13659491193738,3.13189823874755,3.12720156555773,3.12250489236791,3.11780821917808,3.11311154598826,3.10841487279843,3.10371819960861,3.09902152641879,3.09432485322896,3.08962818003914,3.08493150684932,3.08023483365949,3.07553816046967,3.07084148727984,3.06614481409002,3.0614481409002,3.05675146771037,3.05205479452055,3.04735812133072,3.0426614481409,3.03796477495108,3.03326810176125,3.02857142857143,3.0238747553816,3.01917808219178,3.01448140900196,3.00978473581213,3.00508806262231,3.00039138943249,2.99569471624266,2.99099804305284,2.98630136986301,2.98160469667319,2.97690802348337,2.97221135029354,2.96751467710372,2.96281800391389,2.95812133072407,2.95342465753425,2.94872798434442,2.9440313111546,2.93933463796477,2.93463796477495,2.92994129158513,2.9252446183953,2.92054794520548,2.91585127201566,2.91115459882583,2.90645792563601,2.90176125244618,2.89706457925636,2.89236790606654,2.88767123287671,2.88297455968689,2.87827788649706,2.87358121330724,2.86888454011742,2.86418786692759,2.85949119373777,2.85479452054795,2.85009784735812,2.8454011741683,2.84070450097847,2.83600782778865,2.83131115459883,2.826614481409,2.82191780821918,2.81722113502935,2.81252446183953,2.80782778864971,2.80313111545988,2.79843444227006,2.79373776908024,2.78904109589041,2.78434442270059,2.77964774951076,2.77495107632094,2.77025440313112,2.76555772994129,2.76086105675147,2.75616438356164,2.75146771037182,2.746771037182,2.74207436399217,2.73737769080235,2.73268101761252,2.7279843444227,2.72328767123288,2.71859099804305,2.71389432485323,2.70919765166341,2.70450097847358,2.69980430528376,2.69510763209393,2.69041095890411,2.68571428571429,2.68101761252446,2.67632093933464,2.67162426614481,2.66692759295499,2.66223091976517,2.65753424657534,2.65283757338552,2.64814090019569,2.64344422700587,2.63874755381605,2.63405088062622,2.6293542074364,2.62465753424658,2.61996086105675,2.61526418786693,2.6105675146771,2.60587084148728,2.60117416829746,2.59647749510763,2.59178082191781,2.58708414872798,2.58238747553816,2.57769080234834,2.57299412915851,2.56829745596869,2.56360078277886,2.55890410958904,2.55420743639922,2.54951076320939,2.54481409001957,2.54011741682975,2.53542074363992,2.5307240704501,2.52602739726027,2.52133072407045,2.51663405088063,2.5119373776908,2.50724070450098,2.50254403131115,2.49784735812133,2.49315068493151,2.48845401174168,2.48375733855186,2.47906066536204,2.47436399217221,2.46966731898239,2.46497064579256,2.46027397260274,2.45557729941292,2.45088062622309,2.44618395303327,2.44148727984344,2.43679060665362,2.4320939334638,2.42739726027397,2.42270058708415,2.41800391389432,2.4133072407045,2.40861056751468,2.40391389432485,2.39921722113503,2.39452054794521,2.38982387475538,2.38512720156556,2.38043052837573,2.37573385518591,2.37103718199609,2.36634050880626,2.36164383561644,2.35694716242661,2.35225048923679,2.34755381604697,2.34285714285714,2.33816046966732,2.3334637964775,2.32876712328767,2.32407045009785,2.31937377690802,2.3146771037182,2.30998043052838,2.30528375733855,2.30058708414873,2.2958904109589,2.29119373776908,2.28649706457926,2.28180039138943,2.27710371819961,2.27240704500978,2.26771037181996,2.26301369863014,2.25831702544031,2.25362035225049,2.24892367906067,2.24422700587084,2.23953033268102,2.23483365949119,2.23013698630137,2.22544031311155,2.22074363992172,2.2160469667319,2.21135029354207,2.20665362035225,2.20195694716243,2.1972602739726,2.19256360078278,2.18786692759295,2.18317025440313,2.17847358121331,2.17377690802348,2.16908023483366,2.16438356164384,2.15968688845401,2.15499021526419,2.15029354207436,2.14559686888454,2.14090019569472,2.13620352250489,2.13150684931507,2.12681017612524,2.12211350293542,2.1174168297456,2.11272015655577,2.10802348336595,2.10332681017613,2.0986301369863,2.09393346379648,2.08923679060665,2.08454011741683,2.07984344422701,2.07514677103718,2.07045009784736,2.06575342465753,2.06105675146771,2.05636007827789,2.05166340508806,2.04696673189824,2.04227005870841,2.03757338551859,2.03287671232877,2.02818003913894,2.02348336594912,2.0187866927593,2.01409001956947,2.00939334637965,2.00469667318982,2,2],"y":[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,6.46436475374112e-15,8.59429176799396e-15,1.13105975017027e-14,1.47254631720678e-14,1.98014775921978e-14,2.61610393332967e-14,3.39626556240268e-14,4.48184359795307e-14,5.91209403076104e-14,7.68881860479074e-14,1.00103451812635e-13,1.32063511804788e-13,1.71907283202191e-13,2.20879561338554e-13,2.91206298964679e-13,3.79113889333159e-13,4.86392049778136e-13,6.33900664885812e-13,8.25005188203191e-13,1.05943369387932e-12,1.36255152873887e-12,1.77191476389201e-12,2.27606813224737e-12,2.89268421749853e-12,3.7573850443846e-12,4.8251760547996e-12,6.11471865925404e-12,7.86691339842269e-12,1.00954909859191e-11,1.27982513562699e-11,1.62651840267845e-11,2.08501465656908e-11,2.64279402672604e-11,3.32128320934226e-11,4.25146915670782e-11,5.38553241993576e-11,6.7405992106117e-11,8.56022008438869e-11,1.08326851121119e-10,1.35573170386379e-10,1.70217040820171e-10,2.15112582102379e-10,2.69072422139139e-10,3.34307726393575e-10,4.21781163591055e-10,5.27083866206961e-10,6.5158393310629e-10,8.16699440389171e-10,1.01927009463426e-09,1.25941212079356e-09,1.56187216715341e-09,1.94612555035928e-09,2.40246701004313e-09,2.95043221717273e-09,3.6693474388366e-09,4.52403571843491e-09,5.52399181650222e-09,6.83282642567997e-09,8.41104352337773e-09,1.02611195155911e-08,1.25677177157458e-08,1.54417334512241e-08,1.88148070668019e-08,2.28350816795489e-08,2.79978059051794e-08,3.40601893922208e-08,4.10794140543508e-08,5.01402786534122e-08,6.08844179500719e-08,7.33401627599211e-08,8.8701681260119e-08,1.0748274216863e-07,1.29268695913159e-07,1.55025182715641e-07,1.87413055728533e-07,2.24983429119162e-07,2.68034680365425e-07,3.22804014391765e-07,3.86702607968226e-07,4.59968651699584e-07,5.49288767945038e-07,6.56492950857018e-07,7.79417569352479e-07,9.23476681855802e-07,1.10093265694188e-06,1.30431511830874e-06,1.53497032159256e-06,1.82396171242303e-06,2.15587769251694e-06,2.53230881179805e-06,2.98564041088775e-06,3.52004457051652e-06,4.12582394815686e-06,4.82908978655968e-06,5.67810108452289e-06,6.63961335500971e-06,7.71886500321925e-06,9.04968583588579e-06,1.05552395662637e-05,1.22442301952377e-05,1.42521555376431e-05,1.65781902003548e-05,1.91850761410155e-05,2.21810874001383e-05,2.57274396711132e-05,2.96966041957428e-05,3.41174641863835e-05,3.94539113199137e-05,4.54165445854373e-05,5.20336358625242e-05,5.9794158147419e-05,6.8633268328612e-05,7.84285736614251e-05,8.95657331783427e-05,0.000102497911608625,0.000116804073847004,0.000132610310832973,0.000151286230944024,0.000171904613196757,0.000194540929260063,0.000220714365739465,0.000250041953134839,0.000282181075154453,0.000318310072593162,0.000359486260905128,0.000404515592269315,0.000453837155472907,0.000510908336977695,0.000573175333516879,0.000640808601041263,0.000717863296926329,0.000802856361428191,0.000894973999318657,0.000997299321371119,0.00111183228703633,0.00123567045710119,0.00137006758532449,0.00152245699062946,0.00168680493419687,0.00186343386978611,0.00206162342232062,0.00227697015722728,0.00250784298211792,0.00276113770220472,0.00303977678161081,0.00333772734773513,0.00365796341117454,0.00401403862301261,0.00439375040140518,0.00479760217655312,0.00524376557849334,0.00572171509792915,0.00622871190397264,0.00677792420597351,0.00737223184253549,0.00800093403930143,0.00866993676300263,0.00940011800512572,0.0101703853801404,0.0109813731122037,0.0118635238176693,0.0127961385721286,0.0137754021237717,0.0148228025883743,0.0159390086117556,0.0171078330779143,0.0183391692341626,0.019660150711586,0.0210395956422762,0.0224781261360779,0.024019549412153,0.0256298984975433,0.0273048289161262,0.0290745007363613,0.0309347371707268,0.0328645823280899,0.0348782037496354,0.0370055559259892,0.0392069583062165,0.041482860465201,0.0438881826881381,0.046375615250983,0.0489412293340065,0.0516221426927714,0.0544077709738766,0.0572745857900273,0.0602404302128299,0.0633341539947624,0.0665114416434182,0.0697725241856548,0.0731794584742567,0.0766745549932172,0.0802552366393714,0.0839632808271376,0.0877820276824108,0.0916878097682151,0.0957014682817592,0.0998487778363715,0.104084421540671,0.108408539152384,0.112888262504544,0.117458439820563,0.122118463688368,0.126914300381466,0.131824242786671,0.1368256782166,0.1419428368089,0.147198925132912,0.152548587283421,0.157993504095558,0.163603751683728,0.169310207436648,0.175113206742573,0.181065253181426,0.187139114971737,0.193312873721859,0.199615901946746,0.206069961972674,0.212627913229755,0.219294350077664,0.226143558847291,0.233101257990164,0.2401679889504,0.247406555178587,0.254781477316791,0.262270624747714,0.269910767232468,0.277721987440154,0.285653019130117,0.293712195567524,0.301979962194333,0.310373367311841,0.318893051316339,0.327614946374294,0.336491658187149,0.345500507217181,0.354687126518719,0.364067798651189,0.373586221318202,0.383255163022095,0.393159252446227,0.403206133466301,0.413396293955048,0.423818046981216,0.434409489936064,0.445148063671611,0.456086997073034,0.467234749802307,0.478531864157889,0.489995029161303,0.501704566588139,0.51356342493524,0.525571417926174,0.537819847700382,0.550234394392094,0.562794496070324,0.575553027245466,0.588505390235882,0.601595775676508,0.614840845739285,0.628298707078003,0.641882322538463,0.655590039158811,0.669491194269773,0.683513357153341,0.69764107519356,0.71190794589612,0.726294416195544,0.740761906712846,0.755317519870506,0.769973312479943,0.784679427577437,0.799432180934646,0.814238590746357,0.829061420011834,0.843893377791863,0.858721060402848,0.873519315704994,0.888283947343104,0.902998787557226,0.917614492839951,0.932149654276621,0.946598997655111,0.960868114022487,0.975000379390504,0.988996945223558,1.00277419637027,1.01632423121566,1.02968804426924,1.04281529733671,1.05560397535645,1.06815721931223,1.08046980494503,1.09233481801403,1.1039083421991,1.11519594037233,1.12604242390782,1.13648204721256,1.14659592968326,1.15629999711859,1.16547218585077,1.17428581810392,1.18273767923171,1.1905398606758,1.1979532028681,1.20498139998868,1.21142417955213,1.21736663371739,1.22290952813135,1.22794915087359,1.23238102089057,1.23640802591265,1.24002646557937,1.2429389224813,1.24545001755549,1.24756079503531,1.24906791259668,1.25009183564815,1.25072814185265,1.25087452312158,1.25046588378564,1.24968923952341,1.24853547673428,1.24677109577454,1.24466401542583,1.24221734282074,1.2392618755585,1.23592369811229,1.23227614683463,1.22823641145421,1.22377911245172,1.21904560184939,1.21402518003343,1.20856943030443,1.20287245929309,1.19693814838381,1.19065197193316,1.1841182873331,1.17738317419795,1.17039344275568,1.16315128202612,1.15574360889187,1.14816276711162,1.14033968052156,1.13238644757033,1.12430685307738,1.11604665908659,1.10767209990042,1.09920540073712,1.09062644430648,1.08195085526993,1.07321600827101,1.06442126882541,1.05556004467381,1.04667082943388,1.03775690423372,1.02882150183371,1.01988561185711,1.01095406929557,1.0020389773025,0.99315702706686,0.98430620202197,0.975495259645283,0.966759649473693,0.958079286816523,0.94945661293382,0.940942525820949,0.93251246191928,0.924160645682697,0.915925422530416,0.907814197599634,0.899798214276457,0.89189496962393,0.884158478835501,0.87653027663444,0.869011540558741,0.861681335550187,0.854478742971276,0.847393774436497,0.840478097146162,0.833724084968784,0.827091428406265,0.820601728109832,0.814304142185049,0.808127242480366,0.802070724592828,0.796214373377931,0.7904820383258,0.784864637339794,0.779409675430973,0.774097111766788,0.76889031233416,0.763807747238824,0.758878124300812,0.754041939661995,0.749297715612059,0.744700273198009,0.740185347286016,0.735747275790046,0.731414402241624,0.727164355204272,0.722974438021383,0.718853637266409,0.71480749088778,0.71080386018132,0.706840807897457,0.702934624247884,0.699053831370843,0.695195808090716,0.691363122713768,0.687542169915816,0.683726630763417,0.679913213901022,0.676091242271928,0.672258220919116,0.668412362228445,0.664531958521618,0.660625254852361,0.656690706493412,0.652706735353685,0.648674749313723,0.644601324592224,0.640471542746343,0.636267823945426,0.632010656163139,0.62769724952671,0.623280859621321,0.618800683919295,0.614255733271936,0.609606338466391,0.604869771970271,0.600060125677823,0.595153647499073,0.590133517761111,0.585033902094895,0.579850078737105,0.574525865860615,0.569117822546019,0.563625640247209,0.558000589988643,0.552273099088832,0.546459749539177,0.5405331323443,0.534483338417095,0.528348502300992,0.522122668755957,0.515756633241948,0.509309125072777,0.502780734927552,0.496127530434469,0.489385912814276,0.482570337070159,0.475658538954087,0.468651479648953,0.461580361281597,0.454440818573651,0.447207061928012,0.439922016153858,0.43258724761584,0.425181562387849,0.417733484830555,0.410251386371935,0.402729113148725,0.395176916952462,0.387608433517878,0.380025061143818,0.372433844796467,0.36484544320468,0.357261982478833,0.349699136223983,0.342160327689035,0.334646123447601,0.327173753683193,0.319754276107951,0.312378583904742,0.305056729212712,0.297823570276255,0.290651990523099,0.283543806564086,0.276551344948814,0.269642959680937,0.262813132874831,0.256099882503008,0.249504731498929,0.243000258952053,0.236604017854147,0.230361111506884,0.224217488561071,0.218173880377867,0.212302183571035,0.206542351899408,0.200886841248947,0.195382117662151,0.190016223408232,0.18475527700738,0.179619221646057,0.174645190676973,0.169773383041357,0.165003347441208,0.160403061046064,0.1559046141464,0.151502052563571,0.147236211387813,0.143087103307732,0.139026017541551,0.135069835400248,0.131240284009876,0.127489711670663,0.123817102767386,0.120272139416234,0.116798795180152,0.11339400047473,0],"text":["density: 1.133940e-01<br />Sepal.Width: 2.000000","density: 1.167988e-01<br />Sepal.Width: 2.004697","density: 1.202721e-01<br />Sepal.Width: 2.009393","density: 1.238171e-01<br />Sepal.Width: 2.014090","density: 1.274897e-01<br />Sepal.Width: 2.018787","density: 1.312403e-01<br />Sepal.Width: 2.023483","density: 1.350698e-01<br />Sepal.Width: 2.028180","density: 1.390260e-01<br />Sepal.Width: 2.032877","density: 1.430871e-01<br />Sepal.Width: 2.037573","density: 1.472362e-01<br />Sepal.Width: 2.042270","density: 1.515021e-01<br />Sepal.Width: 2.046967","density: 1.559046e-01<br />Sepal.Width: 2.051663","density: 1.604031e-01<br />Sepal.Width: 2.056360","density: 1.650033e-01<br />Sepal.Width: 2.061057","density: 1.697734e-01<br />Sepal.Width: 2.065753","density: 1.746452e-01<br />Sepal.Width: 2.070450","density: 1.796192e-01<br />Sepal.Width: 2.075147","density: 1.847553e-01<br />Sepal.Width: 2.079843","density: 1.900162e-01<br />Sepal.Width: 2.084540","density: 1.953821e-01<br />Sepal.Width: 2.089237","density: 2.008868e-01<br />Sepal.Width: 2.093933","density: 2.065424e-01<br />Sepal.Width: 2.098630","density: 2.123022e-01<br />Sepal.Width: 2.103327","density: 2.181739e-01<br />Sepal.Width: 2.108023","density: 2.242175e-01<br />Sepal.Width: 2.112720","density: 2.303611e-01<br />Sepal.Width: 2.117417","density: 2.366040e-01<br />Sepal.Width: 2.122114","density: 2.430003e-01<br />Sepal.Width: 2.126810","density: 2.495047e-01<br />Sepal.Width: 2.131507","density: 2.560999e-01<br />Sepal.Width: 2.136204","density: 2.628131e-01<br />Sepal.Width: 2.140900","density: 2.696430e-01<br />Sepal.Width: 2.145597","density: 2.765513e-01<br />Sepal.Width: 2.150294","density: 2.835438e-01<br />Sepal.Width: 2.154990","density: 2.906520e-01<br />Sepal.Width: 2.159687","density: 2.978236e-01<br />Sepal.Width: 2.164384","density: 3.050567e-01<br />Sepal.Width: 2.169080","density: 3.123786e-01<br />Sepal.Width: 2.173777","density: 3.197543e-01<br />Sepal.Width: 2.178474","density: 3.271738e-01<br />Sepal.Width: 2.183170","density: 3.346461e-01<br />Sepal.Width: 2.187867","density: 3.421603e-01<br />Sepal.Width: 2.192564","density: 3.496991e-01<br />Sepal.Width: 2.197260","density: 3.572620e-01<br />Sepal.Width: 2.201957","density: 3.648454e-01<br />Sepal.Width: 2.206654","density: 3.724338e-01<br />Sepal.Width: 2.211350","density: 3.800251e-01<br />Sepal.Width: 2.216047","density: 3.876084e-01<br />Sepal.Width: 2.220744","density: 3.951769e-01<br />Sepal.Width: 2.225440","density: 4.027291e-01<br />Sepal.Width: 2.230137","density: 4.102514e-01<br />Sepal.Width: 2.234834","density: 4.177335e-01<br />Sepal.Width: 2.239530","density: 4.251816e-01<br />Sepal.Width: 2.244227","density: 4.325872e-01<br />Sepal.Width: 2.248924","density: 4.399220e-01<br />Sepal.Width: 2.253620","density: 4.472071e-01<br />Sepal.Width: 2.258317","density: 4.544408e-01<br />Sepal.Width: 2.263014","density: 4.615804e-01<br />Sepal.Width: 2.267710","density: 4.686515e-01<br />Sepal.Width: 2.272407","density: 4.756585e-01<br />Sepal.Width: 2.277104","density: 4.825703e-01<br />Sepal.Width: 2.281800","density: 4.893859e-01<br />Sepal.Width: 2.286497","density: 4.961275e-01<br />Sepal.Width: 2.291194","density: 5.027807e-01<br />Sepal.Width: 2.295890","density: 5.093091e-01<br />Sepal.Width: 2.300587","density: 5.157566e-01<br />Sepal.Width: 2.305284","density: 5.221227e-01<br />Sepal.Width: 2.309980","density: 5.283485e-01<br />Sepal.Width: 2.314677","density: 5.344833e-01<br />Sepal.Width: 2.319374","density: 5.405331e-01<br />Sepal.Width: 2.324070","density: 5.464597e-01<br />Sepal.Width: 2.328767","density: 5.522731e-01<br />Sepal.Width: 2.333464","density: 5.580006e-01<br />Sepal.Width: 2.338160","density: 5.636256e-01<br />Sepal.Width: 2.342857","density: 5.691178e-01<br />Sepal.Width: 2.347554","density: 5.745259e-01<br />Sepal.Width: 2.352250","density: 5.798501e-01<br />Sepal.Width: 2.356947","density: 5.850339e-01<br />Sepal.Width: 2.361644","density: 5.901335e-01<br />Sepal.Width: 2.366341","density: 5.951536e-01<br />Sepal.Width: 2.371037","density: 6.000601e-01<br />Sepal.Width: 2.375734","density: 6.048698e-01<br />Sepal.Width: 2.380431","density: 6.096063e-01<br />Sepal.Width: 2.385127","density: 6.142557e-01<br />Sepal.Width: 2.389824","density: 6.188007e-01<br />Sepal.Width: 2.394521","density: 6.232809e-01<br />Sepal.Width: 2.399217","density: 6.276972e-01<br />Sepal.Width: 2.403914","density: 6.320107e-01<br />Sepal.Width: 2.408611","density: 6.362678e-01<br />Sepal.Width: 2.413307","density: 6.404715e-01<br />Sepal.Width: 2.418004","density: 6.446013e-01<br />Sepal.Width: 2.422701","density: 6.486747e-01<br />Sepal.Width: 2.427397","density: 6.527067e-01<br />Sepal.Width: 2.432094","density: 6.566907e-01<br />Sepal.Width: 2.436791","density: 6.606253e-01<br />Sepal.Width: 2.441487","density: 6.645320e-01<br />Sepal.Width: 2.446184","density: 6.684124e-01<br />Sepal.Width: 2.450881","density: 6.722582e-01<br />Sepal.Width: 2.455577","density: 6.760912e-01<br />Sepal.Width: 2.460274","density: 6.799132e-01<br />Sepal.Width: 2.464971","density: 6.837266e-01<br />Sepal.Width: 2.469667","density: 6.875422e-01<br />Sepal.Width: 2.474364","density: 6.913631e-01<br />Sepal.Width: 2.479061","density: 6.951958e-01<br />Sepal.Width: 2.483757","density: 6.990538e-01<br />Sepal.Width: 2.488454","density: 7.029346e-01<br />Sepal.Width: 2.493151","density: 7.068408e-01<br />Sepal.Width: 2.497847","density: 7.108039e-01<br />Sepal.Width: 2.502544","density: 7.148075e-01<br />Sepal.Width: 2.507241","density: 7.188536e-01<br />Sepal.Width: 2.511937","density: 7.229744e-01<br />Sepal.Width: 2.516634","density: 7.271644e-01<br />Sepal.Width: 2.521331","density: 7.314144e-01<br />Sepal.Width: 2.526027","density: 7.357473e-01<br />Sepal.Width: 2.530724","density: 7.401853e-01<br />Sepal.Width: 2.535421","density: 7.447003e-01<br />Sepal.Width: 2.540117","density: 7.492977e-01<br />Sepal.Width: 2.544814","density: 7.540419e-01<br />Sepal.Width: 2.549511","density: 7.588781e-01<br />Sepal.Width: 2.554207","density: 7.638077e-01<br />Sepal.Width: 2.558904","density: 7.688903e-01<br />Sepal.Width: 2.563601","density: 7.740971e-01<br />Sepal.Width: 2.568297","density: 7.794097e-01<br />Sepal.Width: 2.572994","density: 7.848646e-01<br />Sepal.Width: 2.577691","density: 7.904820e-01<br />Sepal.Width: 2.582387","density: 7.962144e-01<br />Sepal.Width: 2.587084","density: 8.020707e-01<br />Sepal.Width: 2.591781","density: 8.081272e-01<br />Sepal.Width: 2.596477","density: 8.143041e-01<br />Sepal.Width: 2.601174","density: 8.206017e-01<br />Sepal.Width: 2.605871","density: 8.270914e-01<br />Sepal.Width: 2.610568","density: 8.337241e-01<br />Sepal.Width: 2.615264","density: 8.404781e-01<br />Sepal.Width: 2.619961","density: 8.473938e-01<br />Sepal.Width: 2.624658","density: 8.544787e-01<br />Sepal.Width: 2.629354","density: 8.616813e-01<br />Sepal.Width: 2.634051","density: 8.690115e-01<br />Sepal.Width: 2.638748","density: 8.765303e-01<br />Sepal.Width: 2.643444","density: 8.841585e-01<br />Sepal.Width: 2.648141","density: 8.918950e-01<br />Sepal.Width: 2.652838","density: 8.997982e-01<br />Sepal.Width: 2.657534","density: 9.078142e-01<br />Sepal.Width: 2.662231","density: 9.159254e-01<br />Sepal.Width: 2.666928","density: 9.241606e-01<br />Sepal.Width: 2.671624","density: 9.325125e-01<br />Sepal.Width: 2.676321","density: 9.409425e-01<br />Sepal.Width: 2.681018","density: 9.494566e-01<br />Sepal.Width: 2.685714","density: 9.580793e-01<br />Sepal.Width: 2.690411","density: 9.667596e-01<br />Sepal.Width: 2.695108","density: 9.754953e-01<br />Sepal.Width: 2.699804","density: 9.843062e-01<br />Sepal.Width: 2.704501","density: 9.931570e-01<br />Sepal.Width: 2.709198","density: 1.002039e+00<br />Sepal.Width: 2.713894","density: 1.010954e+00<br />Sepal.Width: 2.718591","density: 1.019886e+00<br />Sepal.Width: 2.723288","density: 1.028822e+00<br />Sepal.Width: 2.727984","density: 1.037757e+00<br />Sepal.Width: 2.732681","density: 1.046671e+00<br />Sepal.Width: 2.737378","density: 1.055560e+00<br />Sepal.Width: 2.742074","density: 1.064421e+00<br />Sepal.Width: 2.746771","density: 1.073216e+00<br />Sepal.Width: 2.751468","density: 1.081951e+00<br />Sepal.Width: 2.756164","density: 1.090626e+00<br />Sepal.Width: 2.760861","density: 1.099205e+00<br />Sepal.Width: 2.765558","density: 1.107672e+00<br />Sepal.Width: 2.770254","density: 1.116047e+00<br />Sepal.Width: 2.774951","density: 1.124307e+00<br />Sepal.Width: 2.779648","density: 1.132386e+00<br />Sepal.Width: 2.784344","density: 1.140340e+00<br />Sepal.Width: 2.789041","density: 1.148163e+00<br />Sepal.Width: 2.793738","density: 1.155744e+00<br />Sepal.Width: 2.798434","density: 1.163151e+00<br />Sepal.Width: 2.803131","density: 1.170393e+00<br />Sepal.Width: 2.807828","density: 1.177383e+00<br />Sepal.Width: 2.812524","density: 1.184118e+00<br />Sepal.Width: 2.817221","density: 1.190652e+00<br />Sepal.Width: 2.821918","density: 1.196938e+00<br />Sepal.Width: 2.826614","density: 1.202872e+00<br />Sepal.Width: 2.831311","density: 1.208569e+00<br />Sepal.Width: 2.836008","density: 1.214025e+00<br />Sepal.Width: 2.840705","density: 1.219046e+00<br />Sepal.Width: 2.845401","density: 1.223779e+00<br />Sepal.Width: 2.850098","density: 1.228236e+00<br />Sepal.Width: 2.854795","density: 1.232276e+00<br />Sepal.Width: 2.859491","density: 1.235924e+00<br />Sepal.Width: 2.864188","density: 1.239262e+00<br />Sepal.Width: 2.868885","density: 1.242217e+00<br />Sepal.Width: 2.873581","density: 1.244664e+00<br />Sepal.Width: 2.878278","density: 1.246771e+00<br />Sepal.Width: 2.882975","density: 1.248535e+00<br />Sepal.Width: 2.887671","density: 1.249689e+00<br />Sepal.Width: 2.892368","density: 1.250466e+00<br />Sepal.Width: 2.897065","density: 1.250875e+00<br />Sepal.Width: 2.901761","density: 1.250728e+00<br />Sepal.Width: 2.906458","density: 1.250092e+00<br />Sepal.Width: 2.911155","density: 1.249068e+00<br />Sepal.Width: 2.915851","density: 1.247561e+00<br />Sepal.Width: 2.920548","density: 1.245450e+00<br />Sepal.Width: 2.925245","density: 1.242939e+00<br />Sepal.Width: 2.929941","density: 1.240026e+00<br />Sepal.Width: 2.934638","density: 1.236408e+00<br />Sepal.Width: 2.939335","density: 1.232381e+00<br />Sepal.Width: 2.944031","density: 1.227949e+00<br />Sepal.Width: 2.948728","density: 1.222910e+00<br />Sepal.Width: 2.953425","density: 1.217367e+00<br />Sepal.Width: 2.958121","density: 1.211424e+00<br />Sepal.Width: 2.962818","density: 1.204981e+00<br />Sepal.Width: 2.967515","density: 1.197953e+00<br />Sepal.Width: 2.972211","density: 1.190540e+00<br />Sepal.Width: 2.976908","density: 1.182738e+00<br />Sepal.Width: 2.981605","density: 1.174286e+00<br />Sepal.Width: 2.986301","density: 1.165472e+00<br />Sepal.Width: 2.990998","density: 1.156300e+00<br />Sepal.Width: 2.995695","density: 1.146596e+00<br />Sepal.Width: 3.000391","density: 1.136482e+00<br />Sepal.Width: 3.005088","density: 1.126042e+00<br />Sepal.Width: 3.009785","density: 1.115196e+00<br />Sepal.Width: 3.014481","density: 1.103908e+00<br />Sepal.Width: 3.019178","density: 1.092335e+00<br />Sepal.Width: 3.023875","density: 1.080470e+00<br />Sepal.Width: 3.028571","density: 1.068157e+00<br />Sepal.Width: 3.033268","density: 1.055604e+00<br />Sepal.Width: 3.037965","density: 1.042815e+00<br />Sepal.Width: 3.042661","density: 1.029688e+00<br />Sepal.Width: 3.047358","density: 1.016324e+00<br />Sepal.Width: 3.052055","density: 1.002774e+00<br />Sepal.Width: 3.056751","density: 9.889969e-01<br />Sepal.Width: 3.061448","density: 9.750004e-01<br />Sepal.Width: 3.066145","density: 9.608681e-01<br />Sepal.Width: 3.070841","density: 9.465990e-01<br />Sepal.Width: 3.075538","density: 9.321497e-01<br />Sepal.Width: 3.080235","density: 9.176145e-01<br />Sepal.Width: 3.084932","density: 9.029988e-01<br />Sepal.Width: 3.089628","density: 8.882839e-01<br />Sepal.Width: 3.094325","density: 8.735193e-01<br />Sepal.Width: 3.099022","density: 8.587211e-01<br />Sepal.Width: 3.103718","density: 8.438934e-01<br />Sepal.Width: 3.108415","density: 8.290614e-01<br />Sepal.Width: 3.113112","density: 8.142386e-01<br />Sepal.Width: 3.117808","density: 7.994322e-01<br />Sepal.Width: 3.122505","density: 7.846794e-01<br />Sepal.Width: 3.127202","density: 7.699733e-01<br />Sepal.Width: 3.131898","density: 7.553175e-01<br />Sepal.Width: 3.136595","density: 7.407619e-01<br />Sepal.Width: 3.141292","density: 7.262944e-01<br />Sepal.Width: 3.145988","density: 7.119079e-01<br />Sepal.Width: 3.150685","density: 6.976411e-01<br />Sepal.Width: 3.155382","density: 6.835134e-01<br />Sepal.Width: 3.160078","density: 6.694912e-01<br />Sepal.Width: 3.164775","density: 6.555900e-01<br />Sepal.Width: 3.169472","density: 6.418823e-01<br />Sepal.Width: 3.174168","density: 6.282987e-01<br />Sepal.Width: 3.178865","density: 6.148408e-01<br />Sepal.Width: 3.183562","density: 6.015958e-01<br />Sepal.Width: 3.188258","density: 5.885054e-01<br />Sepal.Width: 3.192955","density: 5.755530e-01<br />Sepal.Width: 3.197652","density: 5.627945e-01<br />Sepal.Width: 3.202348","density: 5.502344e-01<br />Sepal.Width: 3.207045","density: 5.378198e-01<br />Sepal.Width: 3.211742","density: 5.255714e-01<br />Sepal.Width: 3.216438","density: 5.135634e-01<br />Sepal.Width: 3.221135","density: 5.017046e-01<br />Sepal.Width: 3.225832","density: 4.899950e-01<br />Sepal.Width: 3.230528","density: 4.785319e-01<br />Sepal.Width: 3.235225","density: 4.672347e-01<br />Sepal.Width: 3.239922","density: 4.560870e-01<br />Sepal.Width: 3.244618","density: 4.451481e-01<br />Sepal.Width: 3.249315","density: 4.344095e-01<br />Sepal.Width: 3.254012","density: 4.238180e-01<br />Sepal.Width: 3.258708","density: 4.133963e-01<br />Sepal.Width: 3.263405","density: 4.032061e-01<br />Sepal.Width: 3.268102","density: 3.931593e-01<br />Sepal.Width: 3.272798","density: 3.832552e-01<br />Sepal.Width: 3.277495","density: 3.735862e-01<br />Sepal.Width: 3.282192","density: 3.640678e-01<br />Sepal.Width: 3.286888","density: 3.546871e-01<br />Sepal.Width: 3.291585","density: 3.455005e-01<br />Sepal.Width: 3.296282","density: 3.364917e-01<br />Sepal.Width: 3.300978","density: 3.276149e-01<br />Sepal.Width: 3.305675","density: 3.188931e-01<br />Sepal.Width: 3.310372","density: 3.103734e-01<br />Sepal.Width: 3.315068","density: 3.019800e-01<br />Sepal.Width: 3.319765","density: 2.937122e-01<br />Sepal.Width: 3.324462","density: 2.856530e-01<br />Sepal.Width: 3.329159","density: 2.777220e-01<br />Sepal.Width: 3.333855","density: 2.699108e-01<br />Sepal.Width: 3.338552","density: 2.622706e-01<br />Sepal.Width: 3.343249","density: 2.547815e-01<br />Sepal.Width: 3.347945","density: 2.474066e-01<br />Sepal.Width: 3.352642","density: 2.401680e-01<br />Sepal.Width: 3.357339","density: 2.331013e-01<br />Sepal.Width: 3.362035","density: 2.261436e-01<br />Sepal.Width: 3.366732","density: 2.192944e-01<br />Sepal.Width: 3.371429","density: 2.126279e-01<br />Sepal.Width: 3.376125","density: 2.060700e-01<br />Sepal.Width: 3.380822","density: 1.996159e-01<br />Sepal.Width: 3.385519","density: 1.933129e-01<br />Sepal.Width: 3.390215","density: 1.871391e-01<br />Sepal.Width: 3.394912","density: 1.810653e-01<br />Sepal.Width: 3.399609","density: 1.751132e-01<br />Sepal.Width: 3.404305","density: 1.693102e-01<br />Sepal.Width: 3.409002","density: 1.636038e-01<br />Sepal.Width: 3.413699","density: 1.579935e-01<br />Sepal.Width: 3.418395","density: 1.525486e-01<br />Sepal.Width: 3.423092","density: 1.471989e-01<br />Sepal.Width: 3.427789","density: 1.419428e-01<br />Sepal.Width: 3.432485","density: 1.368257e-01<br />Sepal.Width: 3.437182","density: 1.318242e-01<br />Sepal.Width: 3.441879","density: 1.269143e-01<br />Sepal.Width: 3.446575","density: 1.221185e-01<br />Sepal.Width: 3.451272","density: 1.174584e-01<br />Sepal.Width: 3.455969","density: 1.128883e-01<br />Sepal.Width: 3.460665","density: 1.084085e-01<br />Sepal.Width: 3.465362","density: 1.040844e-01<br />Sepal.Width: 3.470059","density: 9.984878e-02<br />Sepal.Width: 3.474755","density: 9.570147e-02<br />Sepal.Width: 3.479452","density: 9.168781e-02<br />Sepal.Width: 3.484149","density: 8.778203e-02<br />Sepal.Width: 3.488845","density: 8.396328e-02<br />Sepal.Width: 3.493542","density: 8.025524e-02<br />Sepal.Width: 3.498239","density: 7.667455e-02<br />Sepal.Width: 3.502935","density: 7.317946e-02<br />Sepal.Width: 3.507632","density: 6.977252e-02<br />Sepal.Width: 3.512329","density: 6.651144e-02<br />Sepal.Width: 3.517025","density: 6.333415e-02<br />Sepal.Width: 3.521722","density: 6.024043e-02<br />Sepal.Width: 3.526419","density: 5.727459e-02<br />Sepal.Width: 3.531115","density: 5.440777e-02<br />Sepal.Width: 3.535812","density: 5.162214e-02<br />Sepal.Width: 3.540509","density: 4.894123e-02<br />Sepal.Width: 3.545205","density: 4.637562e-02<br />Sepal.Width: 3.549902","density: 4.388818e-02<br />Sepal.Width: 3.554599","density: 4.148286e-02<br />Sepal.Width: 3.559295","density: 3.920696e-02<br />Sepal.Width: 3.563992","density: 3.700556e-02<br />Sepal.Width: 3.568689","density: 3.487820e-02<br />Sepal.Width: 3.573386","density: 3.286458e-02<br />Sepal.Width: 3.578082","density: 3.093474e-02<br />Sepal.Width: 3.582779","density: 2.907450e-02<br />Sepal.Width: 3.587476","density: 2.730483e-02<br />Sepal.Width: 3.592172","density: 2.562990e-02<br />Sepal.Width: 3.596869","density: 2.401955e-02<br />Sepal.Width: 3.601566","density: 2.247813e-02<br />Sepal.Width: 3.606262","density: 2.103960e-02<br />Sepal.Width: 3.610959","density: 1.966015e-02<br />Sepal.Width: 3.615656","density: 1.833917e-02<br />Sepal.Width: 3.620352","density: 1.710783e-02<br />Sepal.Width: 3.625049","density: 1.593901e-02<br />Sepal.Width: 3.629746","density: 1.482280e-02<br />Sepal.Width: 3.634442","density: 1.377540e-02<br />Sepal.Width: 3.639139","density: 1.279614e-02<br />Sepal.Width: 3.643836","density: 1.186352e-02<br />Sepal.Width: 3.648532","density: 1.098137e-02<br />Sepal.Width: 3.653229","density: 1.017039e-02<br />Sepal.Width: 3.657926","density: 9.400118e-03<br />Sepal.Width: 3.662622","density: 8.669937e-03<br />Sepal.Width: 3.667319","density: 8.000934e-03<br />Sepal.Width: 3.672016","density: 7.372232e-03<br />Sepal.Width: 3.676712","density: 6.777924e-03<br />Sepal.Width: 3.681409","density: 6.228712e-03<br />Sepal.Width: 3.686106","density: 5.721715e-03<br />Sepal.Width: 3.690802","density: 5.243766e-03<br />Sepal.Width: 3.695499","density: 4.797602e-03<br />Sepal.Width: 3.700196","density: 4.393750e-03<br />Sepal.Width: 3.704892","density: 4.014039e-03<br />Sepal.Width: 3.709589","density: 3.657963e-03<br />Sepal.Width: 3.714286","density: 3.337727e-03<br />Sepal.Width: 3.718982","density: 3.039777e-03<br />Sepal.Width: 3.723679","density: 2.761138e-03<br />Sepal.Width: 3.728376","density: 2.507843e-03<br />Sepal.Width: 3.733072","density: 2.276970e-03<br />Sepal.Width: 3.737769","density: 2.061623e-03<br />Sepal.Width: 3.742466","density: 1.863434e-03<br />Sepal.Width: 3.747162","density: 1.686805e-03<br />Sepal.Width: 3.751859","density: 1.522457e-03<br />Sepal.Width: 3.756556","density: 1.370068e-03<br />Sepal.Width: 3.761252","density: 1.235670e-03<br />Sepal.Width: 3.765949","density: 1.111832e-03<br />Sepal.Width: 3.770646","density: 9.972993e-04<br />Sepal.Width: 3.775342","density: 8.949740e-04<br />Sepal.Width: 3.780039","density: 8.028564e-04<br />Sepal.Width: 3.784736","density: 7.178633e-04<br />Sepal.Width: 3.789432","density: 6.408086e-04<br />Sepal.Width: 3.794129","density: 5.731753e-04<br />Sepal.Width: 3.798826","density: 5.109083e-04<br />Sepal.Width: 3.803523","density: 4.538372e-04<br />Sepal.Width: 3.808219","density: 4.045156e-04<br />Sepal.Width: 3.812916","density: 3.594863e-04<br />Sepal.Width: 3.817613","density: 3.183101e-04<br />Sepal.Width: 3.822309","density: 2.821811e-04<br />Sepal.Width: 3.827006","density: 2.500420e-04<br />Sepal.Width: 3.831703","density: 2.207144e-04<br />Sepal.Width: 3.836399","density: 1.945409e-04<br />Sepal.Width: 3.841096","density: 1.719046e-04<br />Sepal.Width: 3.845793","density: 1.512862e-04<br />Sepal.Width: 3.850489","density: 1.326103e-04<br />Sepal.Width: 3.855186","density: 1.168041e-04<br />Sepal.Width: 3.859883","density: 1.024979e-04<br />Sepal.Width: 3.864579","density: 8.956573e-05<br />Sepal.Width: 3.869276","density: 7.842857e-05<br />Sepal.Width: 3.873973","density: 6.863327e-05<br />Sepal.Width: 3.878669","density: 5.979416e-05<br />Sepal.Width: 3.883366","density: 5.203364e-05<br />Sepal.Width: 3.888063","density: 4.541654e-05<br />Sepal.Width: 3.892759","density: 3.945391e-05<br />Sepal.Width: 3.897456","density: 3.411746e-05<br />Sepal.Width: 3.902153","density: 2.969660e-05<br />Sepal.Width: 3.906849","density: 2.572744e-05<br />Sepal.Width: 3.911546","density: 2.218109e-05<br />Sepal.Width: 3.916243","density: 1.918508e-05<br />Sepal.Width: 3.920939","density: 1.657819e-05<br />Sepal.Width: 3.925636","density: 1.425216e-05<br />Sepal.Width: 3.930333","density: 1.224423e-05<br />Sepal.Width: 3.935029","density: 1.055524e-05<br />Sepal.Width: 3.939726","density: 9.049686e-06<br />Sepal.Width: 3.944423","density: 7.718865e-06<br />Sepal.Width: 3.949119","density: 6.639613e-06<br />Sepal.Width: 3.953816","density: 5.678101e-06<br />Sepal.Width: 3.958513","density: 4.829090e-06<br />Sepal.Width: 3.963209","density: 4.125824e-06<br />Sepal.Width: 3.967906","density: 3.520045e-06<br />Sepal.Width: 3.972603","density: 2.985640e-06<br />Sepal.Width: 3.977299","density: 2.532309e-06<br />Sepal.Width: 3.981996","density: 2.155878e-06<br />Sepal.Width: 3.986693","density: 1.823962e-06<br />Sepal.Width: 3.991389","density: 1.534970e-06<br />Sepal.Width: 3.996086","density: 1.304315e-06<br />Sepal.Width: 4.000783","density: 1.100933e-06<br />Sepal.Width: 4.005479","density: 9.234767e-07<br />Sepal.Width: 4.010176","density: 7.794176e-07<br />Sepal.Width: 4.014873","density: 6.564930e-07<br />Sepal.Width: 4.019569","density: 5.492888e-07<br />Sepal.Width: 4.024266","density: 4.599687e-07<br />Sepal.Width: 4.028963","density: 3.867026e-07<br />Sepal.Width: 4.033659","density: 3.228040e-07<br />Sepal.Width: 4.038356","density: 2.680347e-07<br />Sepal.Width: 4.043053","density: 2.249834e-07<br />Sepal.Width: 4.047750","density: 1.874131e-07<br />Sepal.Width: 4.052446","density: 1.550252e-07<br />Sepal.Width: 4.057143","density: 1.292687e-07<br />Sepal.Width: 4.061840","density: 1.074827e-07<br />Sepal.Width: 4.066536","density: 8.870168e-08<br />Sepal.Width: 4.071233","density: 7.334016e-08<br />Sepal.Width: 4.075930","density: 6.088442e-08<br />Sepal.Width: 4.080626","density: 5.014028e-08<br />Sepal.Width: 4.085323","density: 4.107941e-08<br />Sepal.Width: 4.090020","density: 3.406019e-08<br />Sepal.Width: 4.094716","density: 2.799781e-08<br />Sepal.Width: 4.099413","density: 2.283508e-08<br />Sepal.Width: 4.104110","density: 1.881481e-08<br />Sepal.Width: 4.108806","density: 1.544173e-08<br />Sepal.Width: 4.113503","density: 1.256772e-08<br />Sepal.Width: 4.118200","density: 1.026112e-08<br />Sepal.Width: 4.122896","density: 8.411044e-09<br />Sepal.Width: 4.127593","density: 6.832826e-09<br />Sepal.Width: 4.132290","density: 5.523992e-09<br />Sepal.Width: 4.136986","density: 4.524036e-09<br />Sepal.Width: 4.141683","density: 3.669347e-09<br />Sepal.Width: 4.146380","density: 2.950432e-09<br />Sepal.Width: 4.151076","density: 2.402467e-09<br />Sepal.Width: 4.155773","density: 1.946126e-09<br />Sepal.Width: 4.160470","density: 1.561872e-09<br />Sepal.Width: 4.165166","density: 1.259412e-09<br />Sepal.Width: 4.169863","density: 1.019270e-09<br />Sepal.Width: 4.174560","density: 8.166994e-10<br />Sepal.Width: 4.179256","density: 6.515839e-10<br />Sepal.Width: 4.183953","density: 5.270839e-10<br />Sepal.Width: 4.188650","density: 4.217812e-10<br />Sepal.Width: 4.193346","density: 3.343077e-10<br />Sepal.Width: 4.198043","density: 2.690724e-10<br />Sepal.Width: 4.202740","density: 2.151126e-10<br />Sepal.Width: 4.207436","density: 1.702170e-10<br />Sepal.Width: 4.212133","density: 1.355732e-10<br />Sepal.Width: 4.216830","density: 1.083269e-10<br />Sepal.Width: 4.221526","density: 8.560220e-11<br />Sepal.Width: 4.226223","density: 6.740599e-11<br />Sepal.Width: 4.230920","density: 5.385532e-11<br />Sepal.Width: 4.235616","density: 4.251469e-11<br />Sepal.Width: 4.240313","density: 3.321283e-11<br />Sepal.Width: 4.245010","density: 2.642794e-11<br />Sepal.Width: 4.249706","density: 2.085015e-11<br />Sepal.Width: 4.254403","density: 1.626518e-11<br />Sepal.Width: 4.259100","density: 1.279825e-11<br />Sepal.Width: 4.263796","density: 1.009549e-11<br />Sepal.Width: 4.268493","density: 7.866913e-12<br />Sepal.Width: 4.273190","density: 6.114719e-12<br />Sepal.Width: 4.277886","density: 4.825176e-12<br />Sepal.Width: 4.282583","density: 3.757385e-12<br />Sepal.Width: 4.287280","density: 2.892684e-12<br />Sepal.Width: 4.291977","density: 2.276068e-12<br />Sepal.Width: 4.296673","density: 1.771915e-12<br />Sepal.Width: 4.301370","density: 1.362552e-12<br />Sepal.Width: 4.306067","density: 1.059434e-12<br />Sepal.Width: 4.310763","density: 8.250052e-13<br />Sepal.Width: 4.315460","density: 6.339007e-13<br />Sepal.Width: 4.320157","density: 4.863920e-13<br />Sepal.Width: 4.324853","density: 3.791139e-13<br />Sepal.Width: 4.329550","density: 2.912063e-13<br />Sepal.Width: 4.334247","density: 2.208796e-13<br />Sepal.Width: 4.338943","density: 1.719073e-13<br />Sepal.Width: 4.343640","density: 1.320635e-13<br />Sepal.Width: 4.348337","density: 1.001035e-13<br />Sepal.Width: 4.353033","density: 7.688819e-14<br />Sepal.Width: 4.357730","density: 5.912094e-14<br />Sepal.Width: 4.362427","density: 4.481844e-14<br />Sepal.Width: 4.367123","density: 3.396266e-14<br />Sepal.Width: 4.371820","density: 2.616104e-14<br />Sepal.Width: 4.376517","density: 1.980148e-14<br />Sepal.Width: 4.381213","density: 1.472546e-14<br />Sepal.Width: 4.385910","density: 1.131060e-14<br />Sepal.Width: 4.390607","density: 8.594292e-15<br />Sepal.Width: 4.395303","density: 6.464365e-15<br />Sepal.Width: 4.400000","density: 6.464365e-15<br />Sepal.Width: 4.400000","density: 8.594292e-15<br />Sepal.Width: 4.395303","density: 1.131060e-14<br />Sepal.Width: 4.390607","density: 1.472546e-14<br />Sepal.Width: 4.385910","density: 1.980148e-14<br />Sepal.Width: 4.381213","density: 2.616104e-14<br />Sepal.Width: 4.376517","density: 3.396266e-14<br />Sepal.Width: 4.371820","density: 4.481844e-14<br />Sepal.Width: 4.367123","density: 5.912094e-14<br />Sepal.Width: 4.362427","density: 7.688819e-14<br />Sepal.Width: 4.357730","density: 1.001035e-13<br />Sepal.Width: 4.353033","density: 1.320635e-13<br />Sepal.Width: 4.348337","density: 1.719073e-13<br />Sepal.Width: 4.343640","density: 2.208796e-13<br />Sepal.Width: 4.338943","density: 2.912063e-13<br />Sepal.Width: 4.334247","density: 3.791139e-13<br />Sepal.Width: 4.329550","density: 4.863920e-13<br />Sepal.Width: 4.324853","density: 6.339007e-13<br />Sepal.Width: 4.320157","density: 8.250052e-13<br />Sepal.Width: 4.315460","density: 1.059434e-12<br />Sepal.Width: 4.310763","density: 1.362552e-12<br />Sepal.Width: 4.306067","density: 1.771915e-12<br />Sepal.Width: 4.301370","density: 2.276068e-12<br />Sepal.Width: 4.296673","density: 2.892684e-12<br />Sepal.Width: 4.291977","density: 3.757385e-12<br />Sepal.Width: 4.287280","density: 4.825176e-12<br />Sepal.Width: 4.282583","density: 6.114719e-12<br />Sepal.Width: 4.277886","density: 7.866913e-12<br />Sepal.Width: 4.273190","density: 1.009549e-11<br />Sepal.Width: 4.268493","density: 1.279825e-11<br />Sepal.Width: 4.263796","density: 1.626518e-11<br />Sepal.Width: 4.259100","density: 2.085015e-11<br />Sepal.Width: 4.254403","density: 2.642794e-11<br />Sepal.Width: 4.249706","density: 3.321283e-11<br />Sepal.Width: 4.245010","density: 4.251469e-11<br />Sepal.Width: 4.240313","density: 5.385532e-11<br />Sepal.Width: 4.235616","density: 6.740599e-11<br />Sepal.Width: 4.230920","density: 8.560220e-11<br />Sepal.Width: 4.226223","density: 1.083269e-10<br />Sepal.Width: 4.221526","density: 1.355732e-10<br />Sepal.Width: 4.216830","density: 1.702170e-10<br />Sepal.Width: 4.212133","density: 2.151126e-10<br />Sepal.Width: 4.207436","density: 2.690724e-10<br />Sepal.Width: 4.202740","density: 3.343077e-10<br />Sepal.Width: 4.198043","density: 4.217812e-10<br />Sepal.Width: 4.193346","density: 5.270839e-10<br />Sepal.Width: 4.188650","density: 6.515839e-10<br />Sepal.Width: 4.183953","density: 8.166994e-10<br />Sepal.Width: 4.179256","density: 1.019270e-09<br />Sepal.Width: 4.174560","density: 1.259412e-09<br />Sepal.Width: 4.169863","density: 1.561872e-09<br />Sepal.Width: 4.165166","density: 1.946126e-09<br />Sepal.Width: 4.160470","density: 2.402467e-09<br />Sepal.Width: 4.155773","density: 2.950432e-09<br />Sepal.Width: 4.151076","density: 3.669347e-09<br />Sepal.Width: 4.146380","density: 4.524036e-09<br />Sepal.Width: 4.141683","density: 5.523992e-09<br />Sepal.Width: 4.136986","density: 6.832826e-09<br />Sepal.Width: 4.132290","density: 8.411044e-09<br />Sepal.Width: 4.127593","density: 1.026112e-08<br />Sepal.Width: 4.122896","density: 1.256772e-08<br />Sepal.Width: 4.118200","density: 1.544173e-08<br />Sepal.Width: 4.113503","density: 1.881481e-08<br />Sepal.Width: 4.108806","density: 2.283508e-08<br />Sepal.Width: 4.104110","density: 2.799781e-08<br />Sepal.Width: 4.099413","density: 3.406019e-08<br />Sepal.Width: 4.094716","density: 4.107941e-08<br />Sepal.Width: 4.090020","density: 5.014028e-08<br />Sepal.Width: 4.085323","density: 6.088442e-08<br />Sepal.Width: 4.080626","density: 7.334016e-08<br />Sepal.Width: 4.075930","density: 8.870168e-08<br />Sepal.Width: 4.071233","density: 1.074827e-07<br />Sepal.Width: 4.066536","density: 1.292687e-07<br />Sepal.Width: 4.061840","density: 1.550252e-07<br />Sepal.Width: 4.057143","density: 1.874131e-07<br />Sepal.Width: 4.052446","density: 2.249834e-07<br />Sepal.Width: 4.047750","density: 2.680347e-07<br />Sepal.Width: 4.043053","density: 3.228040e-07<br />Sepal.Width: 4.038356","density: 3.867026e-07<br />Sepal.Width: 4.033659","density: 4.599687e-07<br />Sepal.Width: 4.028963","density: 5.492888e-07<br />Sepal.Width: 4.024266","density: 6.564930e-07<br />Sepal.Width: 4.019569","density: 7.794176e-07<br />Sepal.Width: 4.014873","density: 9.234767e-07<br />Sepal.Width: 4.010176","density: 1.100933e-06<br />Sepal.Width: 4.005479","density: 1.304315e-06<br />Sepal.Width: 4.000783","density: 1.534970e-06<br />Sepal.Width: 3.996086","density: 1.823962e-06<br />Sepal.Width: 3.991389","density: 2.155878e-06<br />Sepal.Width: 3.986693","density: 2.532309e-06<br />Sepal.Width: 3.981996","density: 2.985640e-06<br />Sepal.Width: 3.977299","density: 3.520045e-06<br />Sepal.Width: 3.972603","density: 4.125824e-06<br />Sepal.Width: 3.967906","density: 4.829090e-06<br />Sepal.Width: 3.963209","density: 5.678101e-06<br />Sepal.Width: 3.958513","density: 6.639613e-06<br />Sepal.Width: 3.953816","density: 7.718865e-06<br />Sepal.Width: 3.949119","density: 9.049686e-06<br />Sepal.Width: 3.944423","density: 1.055524e-05<br />Sepal.Width: 3.939726","density: 1.224423e-05<br />Sepal.Width: 3.935029","density: 1.425216e-05<br />Sepal.Width: 3.930333","density: 1.657819e-05<br />Sepal.Width: 3.925636","density: 1.918508e-05<br />Sepal.Width: 3.920939","density: 2.218109e-05<br />Sepal.Width: 3.916243","density: 2.572744e-05<br />Sepal.Width: 3.911546","density: 2.969660e-05<br />Sepal.Width: 3.906849","density: 3.411746e-05<br />Sepal.Width: 3.902153","density: 3.945391e-05<br />Sepal.Width: 3.897456","density: 4.541654e-05<br />Sepal.Width: 3.892759","density: 5.203364e-05<br />Sepal.Width: 3.888063","density: 5.979416e-05<br />Sepal.Width: 3.883366","density: 6.863327e-05<br />Sepal.Width: 3.878669","density: 7.842857e-05<br />Sepal.Width: 3.873973","density: 8.956573e-05<br />Sepal.Width: 3.869276","density: 1.024979e-04<br />Sepal.Width: 3.864579","density: 1.168041e-04<br />Sepal.Width: 3.859883","density: 1.326103e-04<br />Sepal.Width: 3.855186","density: 1.512862e-04<br />Sepal.Width: 3.850489","density: 1.719046e-04<br />Sepal.Width: 3.845793","density: 1.945409e-04<br />Sepal.Width: 3.841096","density: 2.207144e-04<br />Sepal.Width: 3.836399","density: 2.500420e-04<br />Sepal.Width: 3.831703","density: 2.821811e-04<br />Sepal.Width: 3.827006","density: 3.183101e-04<br />Sepal.Width: 3.822309","density: 3.594863e-04<br />Sepal.Width: 3.817613","density: 4.045156e-04<br />Sepal.Width: 3.812916","density: 4.538372e-04<br />Sepal.Width: 3.808219","density: 5.109083e-04<br />Sepal.Width: 3.803523","density: 5.731753e-04<br />Sepal.Width: 3.798826","density: 6.408086e-04<br />Sepal.Width: 3.794129","density: 7.178633e-04<br />Sepal.Width: 3.789432","density: 8.028564e-04<br />Sepal.Width: 3.784736","density: 8.949740e-04<br />Sepal.Width: 3.780039","density: 9.972993e-04<br />Sepal.Width: 3.775342","density: 1.111832e-03<br />Sepal.Width: 3.770646","density: 1.235670e-03<br />Sepal.Width: 3.765949","density: 1.370068e-03<br />Sepal.Width: 3.761252","density: 1.522457e-03<br />Sepal.Width: 3.756556","density: 1.686805e-03<br />Sepal.Width: 3.751859","density: 1.863434e-03<br />Sepal.Width: 3.747162","density: 2.061623e-03<br />Sepal.Width: 3.742466","density: 2.276970e-03<br />Sepal.Width: 3.737769","density: 2.507843e-03<br />Sepal.Width: 3.733072","density: 2.761138e-03<br />Sepal.Width: 3.728376","density: 3.039777e-03<br />Sepal.Width: 3.723679","density: 3.337727e-03<br />Sepal.Width: 3.718982","density: 3.657963e-03<br />Sepal.Width: 3.714286","density: 4.014039e-03<br />Sepal.Width: 3.709589","density: 4.393750e-03<br />Sepal.Width: 3.704892","density: 4.797602e-03<br />Sepal.Width: 3.700196","density: 5.243766e-03<br />Sepal.Width: 3.695499","density: 5.721715e-03<br />Sepal.Width: 3.690802","density: 6.228712e-03<br />Sepal.Width: 3.686106","density: 6.777924e-03<br />Sepal.Width: 3.681409","density: 7.372232e-03<br />Sepal.Width: 3.676712","density: 8.000934e-03<br />Sepal.Width: 3.672016","density: 8.669937e-03<br />Sepal.Width: 3.667319","density: 9.400118e-03<br />Sepal.Width: 3.662622","density: 1.017039e-02<br />Sepal.Width: 3.657926","density: 1.098137e-02<br />Sepal.Width: 3.653229","density: 1.186352e-02<br />Sepal.Width: 3.648532","density: 1.279614e-02<br />Sepal.Width: 3.643836","density: 1.377540e-02<br />Sepal.Width: 3.639139","density: 1.482280e-02<br />Sepal.Width: 3.634442","density: 1.593901e-02<br />Sepal.Width: 3.629746","density: 1.710783e-02<br />Sepal.Width: 3.625049","density: 1.833917e-02<br />Sepal.Width: 3.620352","density: 1.966015e-02<br />Sepal.Width: 3.615656","density: 2.103960e-02<br />Sepal.Width: 3.610959","density: 2.247813e-02<br />Sepal.Width: 3.606262","density: 2.401955e-02<br />Sepal.Width: 3.601566","density: 2.562990e-02<br />Sepal.Width: 3.596869","density: 2.730483e-02<br />Sepal.Width: 3.592172","density: 2.907450e-02<br />Sepal.Width: 3.587476","density: 3.093474e-02<br />Sepal.Width: 3.582779","density: 3.286458e-02<br />Sepal.Width: 3.578082","density: 3.487820e-02<br />Sepal.Width: 3.573386","density: 3.700556e-02<br />Sepal.Width: 3.568689","density: 3.920696e-02<br />Sepal.Width: 3.563992","density: 4.148286e-02<br />Sepal.Width: 3.559295","density: 4.388818e-02<br />Sepal.Width: 3.554599","density: 4.637562e-02<br />Sepal.Width: 3.549902","density: 4.894123e-02<br />Sepal.Width: 3.545205","density: 5.162214e-02<br />Sepal.Width: 3.540509","density: 5.440777e-02<br />Sepal.Width: 3.535812","density: 5.727459e-02<br />Sepal.Width: 3.531115","density: 6.024043e-02<br />Sepal.Width: 3.526419","density: 6.333415e-02<br />Sepal.Width: 3.521722","density: 6.651144e-02<br />Sepal.Width: 3.517025","density: 6.977252e-02<br />Sepal.Width: 3.512329","density: 7.317946e-02<br />Sepal.Width: 3.507632","density: 7.667455e-02<br />Sepal.Width: 3.502935","density: 8.025524e-02<br />Sepal.Width: 3.498239","density: 8.396328e-02<br />Sepal.Width: 3.493542","density: 8.778203e-02<br />Sepal.Width: 3.488845","density: 9.168781e-02<br />Sepal.Width: 3.484149","density: 9.570147e-02<br />Sepal.Width: 3.479452","density: 9.984878e-02<br />Sepal.Width: 3.474755","density: 1.040844e-01<br />Sepal.Width: 3.470059","density: 1.084085e-01<br />Sepal.Width: 3.465362","density: 1.128883e-01<br />Sepal.Width: 3.460665","density: 1.174584e-01<br />Sepal.Width: 3.455969","density: 1.221185e-01<br />Sepal.Width: 3.451272","density: 1.269143e-01<br />Sepal.Width: 3.446575","density: 1.318242e-01<br />Sepal.Width: 3.441879","density: 1.368257e-01<br />Sepal.Width: 3.437182","density: 1.419428e-01<br />Sepal.Width: 3.432485","density: 1.471989e-01<br />Sepal.Width: 3.427789","density: 1.525486e-01<br />Sepal.Width: 3.423092","density: 1.579935e-01<br />Sepal.Width: 3.418395","density: 1.636038e-01<br />Sepal.Width: 3.413699","density: 1.693102e-01<br />Sepal.Width: 3.409002","density: 1.751132e-01<br />Sepal.Width: 3.404305","density: 1.810653e-01<br />Sepal.Width: 3.399609","density: 1.871391e-01<br />Sepal.Width: 3.394912","density: 1.933129e-01<br />Sepal.Width: 3.390215","density: 1.996159e-01<br />Sepal.Width: 3.385519","density: 2.060700e-01<br />Sepal.Width: 3.380822","density: 2.126279e-01<br />Sepal.Width: 3.376125","density: 2.192944e-01<br />Sepal.Width: 3.371429","density: 2.261436e-01<br />Sepal.Width: 3.366732","density: 2.331013e-01<br />Sepal.Width: 3.362035","density: 2.401680e-01<br />Sepal.Width: 3.357339","density: 2.474066e-01<br />Sepal.Width: 3.352642","density: 2.547815e-01<br />Sepal.Width: 3.347945","density: 2.622706e-01<br />Sepal.Width: 3.343249","density: 2.699108e-01<br />Sepal.Width: 3.338552","density: 2.777220e-01<br />Sepal.Width: 3.333855","density: 2.856530e-01<br />Sepal.Width: 3.329159","density: 2.937122e-01<br />Sepal.Width: 3.324462","density: 3.019800e-01<br />Sepal.Width: 3.319765","density: 3.103734e-01<br />Sepal.Width: 3.315068","density: 3.188931e-01<br />Sepal.Width: 3.310372","density: 3.276149e-01<br />Sepal.Width: 3.305675","density: 3.364917e-01<br />Sepal.Width: 3.300978","density: 3.455005e-01<br />Sepal.Width: 3.296282","density: 3.546871e-01<br />Sepal.Width: 3.291585","density: 3.640678e-01<br />Sepal.Width: 3.286888","density: 3.735862e-01<br />Sepal.Width: 3.282192","density: 3.832552e-01<br />Sepal.Width: 3.277495","density: 3.931593e-01<br />Sepal.Width: 3.272798","density: 4.032061e-01<br />Sepal.Width: 3.268102","density: 4.133963e-01<br />Sepal.Width: 3.263405","density: 4.238180e-01<br />Sepal.Width: 3.258708","density: 4.344095e-01<br />Sepal.Width: 3.254012","density: 4.451481e-01<br />Sepal.Width: 3.249315","density: 4.560870e-01<br />Sepal.Width: 3.244618","density: 4.672347e-01<br />Sepal.Width: 3.239922","density: 4.785319e-01<br />Sepal.Width: 3.235225","density: 4.899950e-01<br />Sepal.Width: 3.230528","density: 5.017046e-01<br />Sepal.Width: 3.225832","density: 5.135634e-01<br />Sepal.Width: 3.221135","density: 5.255714e-01<br />Sepal.Width: 3.216438","density: 5.378198e-01<br />Sepal.Width: 3.211742","density: 5.502344e-01<br />Sepal.Width: 3.207045","density: 5.627945e-01<br />Sepal.Width: 3.202348","density: 5.755530e-01<br />Sepal.Width: 3.197652","density: 5.885054e-01<br />Sepal.Width: 3.192955","density: 6.015958e-01<br />Sepal.Width: 3.188258","density: 6.148408e-01<br />Sepal.Width: 3.183562","density: 6.282987e-01<br />Sepal.Width: 3.178865","density: 6.418823e-01<br />Sepal.Width: 3.174168","density: 6.555900e-01<br />Sepal.Width: 3.169472","density: 6.694912e-01<br />Sepal.Width: 3.164775","density: 6.835134e-01<br />Sepal.Width: 3.160078","density: 6.976411e-01<br />Sepal.Width: 3.155382","density: 7.119079e-01<br />Sepal.Width: 3.150685","density: 7.262944e-01<br />Sepal.Width: 3.145988","density: 7.407619e-01<br />Sepal.Width: 3.141292","density: 7.553175e-01<br />Sepal.Width: 3.136595","density: 7.699733e-01<br />Sepal.Width: 3.131898","density: 7.846794e-01<br />Sepal.Width: 3.127202","density: 7.994322e-01<br />Sepal.Width: 3.122505","density: 8.142386e-01<br />Sepal.Width: 3.117808","density: 8.290614e-01<br />Sepal.Width: 3.113112","density: 8.438934e-01<br />Sepal.Width: 3.108415","density: 8.587211e-01<br />Sepal.Width: 3.103718","density: 8.735193e-01<br />Sepal.Width: 3.099022","density: 8.882839e-01<br />Sepal.Width: 3.094325","density: 9.029988e-01<br />Sepal.Width: 3.089628","density: 9.176145e-01<br />Sepal.Width: 3.084932","density: 9.321497e-01<br />Sepal.Width: 3.080235","density: 9.465990e-01<br />Sepal.Width: 3.075538","density: 9.608681e-01<br />Sepal.Width: 3.070841","density: 9.750004e-01<br />Sepal.Width: 3.066145","density: 9.889969e-01<br />Sepal.Width: 3.061448","density: 1.002774e+00<br />Sepal.Width: 3.056751","density: 1.016324e+00<br />Sepal.Width: 3.052055","density: 1.029688e+00<br />Sepal.Width: 3.047358","density: 1.042815e+00<br />Sepal.Width: 3.042661","density: 1.055604e+00<br />Sepal.Width: 3.037965","density: 1.068157e+00<br />Sepal.Width: 3.033268","density: 1.080470e+00<br />Sepal.Width: 3.028571","density: 1.092335e+00<br />Sepal.Width: 3.023875","density: 1.103908e+00<br />Sepal.Width: 3.019178","density: 1.115196e+00<br />Sepal.Width: 3.014481","density: 1.126042e+00<br />Sepal.Width: 3.009785","density: 1.136482e+00<br />Sepal.Width: 3.005088","density: 1.146596e+00<br />Sepal.Width: 3.000391","density: 1.156300e+00<br />Sepal.Width: 2.995695","density: 1.165472e+00<br />Sepal.Width: 2.990998","density: 1.174286e+00<br />Sepal.Width: 2.986301","density: 1.182738e+00<br />Sepal.Width: 2.981605","density: 1.190540e+00<br />Sepal.Width: 2.976908","density: 1.197953e+00<br />Sepal.Width: 2.972211","density: 1.204981e+00<br />Sepal.Width: 2.967515","density: 1.211424e+00<br />Sepal.Width: 2.962818","density: 1.217367e+00<br />Sepal.Width: 2.958121","density: 1.222910e+00<br />Sepal.Width: 2.953425","density: 1.227949e+00<br />Sepal.Width: 2.948728","density: 1.232381e+00<br />Sepal.Width: 2.944031","density: 1.236408e+00<br />Sepal.Width: 2.939335","density: 1.240026e+00<br />Sepal.Width: 2.934638","density: 1.242939e+00<br />Sepal.Width: 2.929941","density: 1.245450e+00<br />Sepal.Width: 2.925245","density: 1.247561e+00<br />Sepal.Width: 2.920548","density: 1.249068e+00<br />Sepal.Width: 2.915851","density: 1.250092e+00<br />Sepal.Width: 2.911155","density: 1.250728e+00<br />Sepal.Width: 2.906458","density: 1.250875e+00<br />Sepal.Width: 2.901761","density: 1.250466e+00<br />Sepal.Width: 2.897065","density: 1.249689e+00<br />Sepal.Width: 2.892368","density: 1.248535e+00<br />Sepal.Width: 2.887671","density: 1.246771e+00<br />Sepal.Width: 2.882975","density: 1.244664e+00<br />Sepal.Width: 2.878278","density: 1.242217e+00<br />Sepal.Width: 2.873581","density: 1.239262e+00<br />Sepal.Width: 2.868885","density: 1.235924e+00<br />Sepal.Width: 2.864188","density: 1.232276e+00<br />Sepal.Width: 2.859491","density: 1.228236e+00<br />Sepal.Width: 2.854795","density: 1.223779e+00<br />Sepal.Width: 2.850098","density: 1.219046e+00<br />Sepal.Width: 2.845401","density: 1.214025e+00<br />Sepal.Width: 2.840705","density: 1.208569e+00<br />Sepal.Width: 2.836008","density: 1.202872e+00<br />Sepal.Width: 2.831311","density: 1.196938e+00<br />Sepal.Width: 2.826614","density: 1.190652e+00<br />Sepal.Width: 2.821918","density: 1.184118e+00<br />Sepal.Width: 2.817221","density: 1.177383e+00<br />Sepal.Width: 2.812524","density: 1.170393e+00<br />Sepal.Width: 2.807828","density: 1.163151e+00<br />Sepal.Width: 2.803131","density: 1.155744e+00<br />Sepal.Width: 2.798434","density: 1.148163e+00<br />Sepal.Width: 2.793738","density: 1.140340e+00<br />Sepal.Width: 2.789041","density: 1.132386e+00<br />Sepal.Width: 2.784344","density: 1.124307e+00<br />Sepal.Width: 2.779648","density: 1.116047e+00<br />Sepal.Width: 2.774951","density: 1.107672e+00<br />Sepal.Width: 2.770254","density: 1.099205e+00<br />Sepal.Width: 2.765558","density: 1.090626e+00<br />Sepal.Width: 2.760861","density: 1.081951e+00<br />Sepal.Width: 2.756164","density: 1.073216e+00<br />Sepal.Width: 2.751468","density: 1.064421e+00<br />Sepal.Width: 2.746771","density: 1.055560e+00<br />Sepal.Width: 2.742074","density: 1.046671e+00<br />Sepal.Width: 2.737378","density: 1.037757e+00<br />Sepal.Width: 2.732681","density: 1.028822e+00<br />Sepal.Width: 2.727984","density: 1.019886e+00<br />Sepal.Width: 2.723288","density: 1.010954e+00<br />Sepal.Width: 2.718591","density: 1.002039e+00<br />Sepal.Width: 2.713894","density: 9.931570e-01<br />Sepal.Width: 2.709198","density: 9.843062e-01<br />Sepal.Width: 2.704501","density: 9.754953e-01<br />Sepal.Width: 2.699804","density: 9.667596e-01<br />Sepal.Width: 2.695108","density: 9.580793e-01<br />Sepal.Width: 2.690411","density: 9.494566e-01<br />Sepal.Width: 2.685714","density: 9.409425e-01<br />Sepal.Width: 2.681018","density: 9.325125e-01<br />Sepal.Width: 2.676321","density: 9.241606e-01<br />Sepal.Width: 2.671624","density: 9.159254e-01<br />Sepal.Width: 2.666928","density: 9.078142e-01<br />Sepal.Width: 2.662231","density: 8.997982e-01<br />Sepal.Width: 2.657534","density: 8.918950e-01<br />Sepal.Width: 2.652838","density: 8.841585e-01<br />Sepal.Width: 2.648141","density: 8.765303e-01<br />Sepal.Width: 2.643444","density: 8.690115e-01<br />Sepal.Width: 2.638748","density: 8.616813e-01<br />Sepal.Width: 2.634051","density: 8.544787e-01<br />Sepal.Width: 2.629354","density: 8.473938e-01<br />Sepal.Width: 2.624658","density: 8.404781e-01<br />Sepal.Width: 2.619961","density: 8.337241e-01<br />Sepal.Width: 2.615264","density: 8.270914e-01<br />Sepal.Width: 2.610568","density: 8.206017e-01<br />Sepal.Width: 2.605871","density: 8.143041e-01<br />Sepal.Width: 2.601174","density: 8.081272e-01<br />Sepal.Width: 2.596477","density: 8.020707e-01<br />Sepal.Width: 2.591781","density: 7.962144e-01<br />Sepal.Width: 2.587084","density: 7.904820e-01<br />Sepal.Width: 2.582387","density: 7.848646e-01<br />Sepal.Width: 2.577691","density: 7.794097e-01<br />Sepal.Width: 2.572994","density: 7.740971e-01<br />Sepal.Width: 2.568297","density: 7.688903e-01<br />Sepal.Width: 2.563601","density: 7.638077e-01<br />Sepal.Width: 2.558904","density: 7.588781e-01<br />Sepal.Width: 2.554207","density: 7.540419e-01<br />Sepal.Width: 2.549511","density: 7.492977e-01<br />Sepal.Width: 2.544814","density: 7.447003e-01<br />Sepal.Width: 2.540117","density: 7.401853e-01<br />Sepal.Width: 2.535421","density: 7.357473e-01<br />Sepal.Width: 2.530724","density: 7.314144e-01<br />Sepal.Width: 2.526027","density: 7.271644e-01<br />Sepal.Width: 2.521331","density: 7.229744e-01<br />Sepal.Width: 2.516634","density: 7.188536e-01<br />Sepal.Width: 2.511937","density: 7.148075e-01<br />Sepal.Width: 2.507241","density: 7.108039e-01<br />Sepal.Width: 2.502544","density: 7.068408e-01<br />Sepal.Width: 2.497847","density: 7.029346e-01<br />Sepal.Width: 2.493151","density: 6.990538e-01<br />Sepal.Width: 2.488454","density: 6.951958e-01<br />Sepal.Width: 2.483757","density: 6.913631e-01<br />Sepal.Width: 2.479061","density: 6.875422e-01<br />Sepal.Width: 2.474364","density: 6.837266e-01<br />Sepal.Width: 2.469667","density: 6.799132e-01<br />Sepal.Width: 2.464971","density: 6.760912e-01<br />Sepal.Width: 2.460274","density: 6.722582e-01<br />Sepal.Width: 2.455577","density: 6.684124e-01<br />Sepal.Width: 2.450881","density: 6.645320e-01<br />Sepal.Width: 2.446184","density: 6.606253e-01<br />Sepal.Width: 2.441487","density: 6.566907e-01<br />Sepal.Width: 2.436791","density: 6.527067e-01<br />Sepal.Width: 2.432094","density: 6.486747e-01<br />Sepal.Width: 2.427397","density: 6.446013e-01<br />Sepal.Width: 2.422701","density: 6.404715e-01<br />Sepal.Width: 2.418004","density: 6.362678e-01<br />Sepal.Width: 2.413307","density: 6.320107e-01<br />Sepal.Width: 2.408611","density: 6.276972e-01<br />Sepal.Width: 2.403914","density: 6.232809e-01<br />Sepal.Width: 2.399217","density: 6.188007e-01<br />Sepal.Width: 2.394521","density: 6.142557e-01<br />Sepal.Width: 2.389824","density: 6.096063e-01<br />Sepal.Width: 2.385127","density: 6.048698e-01<br />Sepal.Width: 2.380431","density: 6.000601e-01<br />Sepal.Width: 2.375734","density: 5.951536e-01<br />Sepal.Width: 2.371037","density: 5.901335e-01<br />Sepal.Width: 2.366341","density: 5.850339e-01<br />Sepal.Width: 2.361644","density: 5.798501e-01<br />Sepal.Width: 2.356947","density: 5.745259e-01<br />Sepal.Width: 2.352250","density: 5.691178e-01<br />Sepal.Width: 2.347554","density: 5.636256e-01<br />Sepal.Width: 2.342857","density: 5.580006e-01<br />Sepal.Width: 2.338160","density: 5.522731e-01<br />Sepal.Width: 2.333464","density: 5.464597e-01<br />Sepal.Width: 2.328767","density: 5.405331e-01<br />Sepal.Width: 2.324070","density: 5.344833e-01<br />Sepal.Width: 2.319374","density: 5.283485e-01<br />Sepal.Width: 2.314677","density: 5.221227e-01<br />Sepal.Width: 2.309980","density: 5.157566e-01<br />Sepal.Width: 2.305284","density: 5.093091e-01<br />Sepal.Width: 2.300587","density: 5.027807e-01<br />Sepal.Width: 2.295890","density: 4.961275e-01<br />Sepal.Width: 2.291194","density: 4.893859e-01<br />Sepal.Width: 2.286497","density: 4.825703e-01<br />Sepal.Width: 2.281800","density: 4.756585e-01<br />Sepal.Width: 2.277104","density: 4.686515e-01<br />Sepal.Width: 2.272407","density: 4.615804e-01<br />Sepal.Width: 2.267710","density: 4.544408e-01<br />Sepal.Width: 2.263014","density: 4.472071e-01<br />Sepal.Width: 2.258317","density: 4.399220e-01<br />Sepal.Width: 2.253620","density: 4.325872e-01<br />Sepal.Width: 2.248924","density: 4.251816e-01<br />Sepal.Width: 2.244227","density: 4.177335e-01<br />Sepal.Width: 2.239530","density: 4.102514e-01<br />Sepal.Width: 2.234834","density: 4.027291e-01<br />Sepal.Width: 2.230137","density: 3.951769e-01<br />Sepal.Width: 2.225440","density: 3.876084e-01<br />Sepal.Width: 2.220744","density: 3.800251e-01<br />Sepal.Width: 2.216047","density: 3.724338e-01<br />Sepal.Width: 2.211350","density: 3.648454e-01<br />Sepal.Width: 2.206654","density: 3.572620e-01<br />Sepal.Width: 2.201957","density: 3.496991e-01<br />Sepal.Width: 2.197260","density: 3.421603e-01<br />Sepal.Width: 2.192564","density: 3.346461e-01<br />Sepal.Width: 2.187867","density: 3.271738e-01<br />Sepal.Width: 2.183170","density: 3.197543e-01<br />Sepal.Width: 2.178474","density: 3.123786e-01<br />Sepal.Width: 2.173777","density: 3.050567e-01<br />Sepal.Width: 2.169080","density: 2.978236e-01<br />Sepal.Width: 2.164384","density: 2.906520e-01<br />Sepal.Width: 2.159687","density: 2.835438e-01<br />Sepal.Width: 2.154990","density: 2.765513e-01<br />Sepal.Width: 2.150294","density: 2.696430e-01<br />Sepal.Width: 2.145597","density: 2.628131e-01<br />Sepal.Width: 2.140900","density: 2.560999e-01<br />Sepal.Width: 2.136204","density: 2.495047e-01<br />Sepal.Width: 2.131507","density: 2.430003e-01<br />Sepal.Width: 2.126810","density: 2.366040e-01<br />Sepal.Width: 2.122114","density: 2.303611e-01<br />Sepal.Width: 2.117417","density: 2.242175e-01<br />Sepal.Width: 2.112720","density: 2.181739e-01<br />Sepal.Width: 2.108023","density: 2.123022e-01<br />Sepal.Width: 2.103327","density: 2.065424e-01<br />Sepal.Width: 2.098630","density: 2.008868e-01<br />Sepal.Width: 2.093933","density: 1.953821e-01<br />Sepal.Width: 2.089237","density: 1.900162e-01<br />Sepal.Width: 2.084540","density: 1.847553e-01<br />Sepal.Width: 2.079843","density: 1.796192e-01<br />Sepal.Width: 2.075147","density: 1.746452e-01<br />Sepal.Width: 2.070450","density: 1.697734e-01<br />Sepal.Width: 2.065753","density: 1.650033e-01<br />Sepal.Width: 2.061057","density: 1.604031e-01<br />Sepal.Width: 2.056360","density: 1.559046e-01<br />Sepal.Width: 2.051663","density: 1.515021e-01<br />Sepal.Width: 2.046967","density: 1.472362e-01<br />Sepal.Width: 2.042270","density: 1.430871e-01<br />Sepal.Width: 2.037573","density: 1.390260e-01<br />Sepal.Width: 2.032877","density: 1.350698e-01<br />Sepal.Width: 2.028180","density: 1.312403e-01<br />Sepal.Width: 2.023483","density: 1.274897e-01<br />Sepal.Width: 2.018787","density: 1.238171e-01<br />Sepal.Width: 2.014090","density: 1.202721e-01<br />Sepal.Width: 2.009393","density: 1.167988e-01<br />Sepal.Width: 2.004697","density: 1.133940e-01<br />Sepal.Width: 2.000000","density: 1.133940e-01<br />Sepal.Width: 2.000000"],"key":["51","52","53","54","55","56","57","58","59","60","61","62","63","64","65","66","67","68","69","70","71","72","73","74","75","76","77","78","79","80","81","82","83","84","85","86","87","88","89","90","91","92","93","94","95","96","97","98","99","100"],"type":"scatter","mode":"lines","line":{"width":1.88976377952756,"color":"rgba(0,0,0,1)","dash":"solid"},"fill":"toself","fillcolor":"rgba(0,186,56,1)","hoveron":"points","set":"SharedData91833e3a","name":"versicolor","legendgroup":"versicolor","showlegend":true,"xaxis":"x2","yaxis":"y7","hoverinfo":"text","_isSimpleKey":true,"_isNestedKey":false,"frame":null},{"x":[2,2.00469667318982,2.00939334637965,2.01409001956947,2.0187866927593,2.02348336594912,2.02818003913894,2.03287671232877,2.03757338551859,2.04227005870841,2.04696673189824,2.05166340508806,2.05636007827789,2.06105675146771,2.06575342465753,2.07045009784736,2.07514677103718,2.07984344422701,2.08454011741683,2.08923679060665,2.09393346379648,2.0986301369863,2.10332681017613,2.10802348336595,2.11272015655577,2.1174168297456,2.12211350293542,2.12681017612524,2.13150684931507,2.13620352250489,2.14090019569472,2.14559686888454,2.15029354207436,2.15499021526419,2.15968688845401,2.16438356164384,2.16908023483366,2.17377690802348,2.17847358121331,2.18317025440313,2.18786692759295,2.19256360078278,2.1972602739726,2.20195694716243,2.20665362035225,2.21135029354207,2.2160469667319,2.22074363992172,2.22544031311155,2.23013698630137,2.23483365949119,2.23953033268102,2.24422700587084,2.24892367906067,2.25362035225049,2.25831702544031,2.26301369863014,2.26771037181996,2.27240704500978,2.27710371819961,2.28180039138943,2.28649706457926,2.29119373776908,2.2958904109589,2.30058708414873,2.30528375733855,2.30998043052838,2.3146771037182,2.31937377690802,2.32407045009785,2.32876712328767,2.3334637964775,2.33816046966732,2.34285714285714,2.34755381604697,2.35225048923679,2.35694716242661,2.36164383561644,2.36634050880626,2.37103718199609,2.37573385518591,2.38043052837573,2.38512720156556,2.38982387475538,2.39452054794521,2.39921722113503,2.40391389432485,2.40861056751468,2.4133072407045,2.41800391389432,2.42270058708415,2.42739726027397,2.4320939334638,2.43679060665362,2.44148727984344,2.44618395303327,2.45088062622309,2.45557729941292,2.46027397260274,2.46497064579256,2.46966731898239,2.47436399217221,2.47906066536204,2.48375733855186,2.48845401174168,2.49315068493151,2.49784735812133,2.50254403131115,2.50724070450098,2.5119373776908,2.51663405088063,2.52133072407045,2.52602739726027,2.5307240704501,2.53542074363992,2.54011741682975,2.54481409001957,2.54951076320939,2.55420743639922,2.55890410958904,2.56360078277886,2.56829745596869,2.57299412915851,2.57769080234834,2.58238747553816,2.58708414872798,2.59178082191781,2.59647749510763,2.60117416829746,2.60587084148728,2.6105675146771,2.61526418786693,2.61996086105675,2.62465753424658,2.6293542074364,2.63405088062622,2.63874755381605,2.64344422700587,2.64814090019569,2.65283757338552,2.65753424657534,2.66223091976517,2.66692759295499,2.67162426614481,2.67632093933464,2.68101761252446,2.68571428571429,2.69041095890411,2.69510763209393,2.69980430528376,2.70450097847358,2.70919765166341,2.71389432485323,2.71859099804305,2.72328767123288,2.7279843444227,2.73268101761252,2.73737769080235,2.74207436399217,2.746771037182,2.75146771037182,2.75616438356164,2.76086105675147,2.76555772994129,2.77025440313112,2.77495107632094,2.77964774951076,2.78434442270059,2.78904109589041,2.79373776908024,2.79843444227006,2.80313111545988,2.80782778864971,2.81252446183953,2.81722113502935,2.82191780821918,2.826614481409,2.83131115459883,2.83600782778865,2.84070450097847,2.8454011741683,2.85009784735812,2.85479452054795,2.85949119373777,2.86418786692759,2.86888454011742,2.87358121330724,2.87827788649706,2.88297455968689,2.88767123287671,2.89236790606654,2.89706457925636,2.90176125244618,2.90645792563601,2.91115459882583,2.91585127201566,2.92054794520548,2.9252446183953,2.92994129158513,2.93463796477495,2.93933463796477,2.9440313111546,2.94872798434442,2.95342465753425,2.95812133072407,2.96281800391389,2.96751467710372,2.97221135029354,2.97690802348337,2.98160469667319,2.98630136986301,2.99099804305284,2.99569471624266,3.00039138943249,3.00508806262231,3.00978473581213,3.01448140900196,3.01917808219178,3.0238747553816,3.02857142857143,3.03326810176125,3.03796477495108,3.0426614481409,3.04735812133072,3.05205479452055,3.05675146771037,3.0614481409002,3.06614481409002,3.07084148727984,3.07553816046967,3.08023483365949,3.08493150684932,3.08962818003914,3.09432485322896,3.09902152641879,3.10371819960861,3.10841487279843,3.11311154598826,3.11780821917808,3.12250489236791,3.12720156555773,3.13189823874755,3.13659491193738,3.1412915851272,3.14598825831703,3.15068493150685,3.15538160469667,3.1600782778865,3.16477495107632,3.16947162426615,3.17416829745597,3.17886497064579,3.18356164383562,3.18825831702544,3.19295499021526,3.19765166340509,3.20234833659491,3.20704500978474,3.21174168297456,3.21643835616438,3.22113502935421,3.22583170254403,3.23052837573386,3.23522504892368,3.2399217221135,3.24461839530333,3.24931506849315,3.25401174168297,3.2587084148728,3.26340508806262,3.26810176125245,3.27279843444227,3.27749510763209,3.28219178082192,3.28688845401174,3.29158512720157,3.29628180039139,3.30097847358121,3.30567514677104,3.31037181996086,3.31506849315068,3.31976516634051,3.32446183953033,3.32915851272016,3.33385518590998,3.3385518590998,3.34324853228963,3.34794520547945,3.35264187866928,3.3573385518591,3.36203522504892,3.36673189823875,3.37142857142857,3.3761252446184,3.38082191780822,3.38551859099804,3.39021526418787,3.39491193737769,3.39960861056751,3.40430528375734,3.40900195694716,3.41369863013699,3.41839530332681,3.42309197651663,3.42778864970646,3.43248532289628,3.43718199608611,3.44187866927593,3.44657534246575,3.45127201565558,3.4559686888454,3.46066536203523,3.46536203522505,3.47005870841487,3.4747553816047,3.47945205479452,3.48414872798434,3.48884540117417,3.49354207436399,3.49823874755382,3.50293542074364,3.50763209393346,3.51232876712329,3.51702544031311,3.52172211350294,3.52641878669276,3.53111545988258,3.53581213307241,3.54050880626223,3.54520547945206,3.54990215264188,3.5545988258317,3.55929549902153,3.56399217221135,3.56868884540117,3.573385518591,3.57808219178082,3.58277886497065,3.58747553816047,3.59217221135029,3.59686888454012,3.60156555772994,3.60626223091977,3.61095890410959,3.61565557729941,3.62035225048924,3.62504892367906,3.62974559686888,3.63444227005871,3.63913894324853,3.64383561643836,3.64853228962818,3.653228962818,3.65792563600783,3.66262230919765,3.66731898238748,3.6720156555773,3.67671232876712,3.68140900195695,3.68610567514677,3.6908023483366,3.69549902152642,3.70019569471624,3.70489236790607,3.70958904109589,3.71428571428571,3.71898238747554,3.72367906066536,3.72837573385519,3.73307240704501,3.73776908023483,3.74246575342466,3.74716242661448,3.75185909980431,3.75655577299413,3.76125244618395,3.76594911937378,3.7706457925636,3.77534246575342,3.78003913894325,3.78473581213307,3.7894324853229,3.79412915851272,3.79882583170254,3.80352250489237,3.80821917808219,3.81291585127202,3.81761252446184,3.82230919765166,3.82700587084149,3.83170254403131,3.83639921722114,3.84109589041096,3.84579256360078,3.85048923679061,3.85518590998043,3.85988258317026,3.86457925636008,3.8692759295499,3.87397260273973,3.87866927592955,3.88336594911937,3.8880626223092,3.89275929549902,3.89745596868885,3.90215264187867,3.90684931506849,3.91154598825832,3.91624266144814,3.92093933463797,3.92563600782779,3.93033268101761,3.93502935420744,3.93972602739726,3.94442270058708,3.94911937377691,3.95381604696673,3.95851272015656,3.96320939334638,3.9679060665362,3.97260273972603,3.97729941291585,3.98199608610568,3.9866927592955,3.99138943248532,3.99608610567515,4.00078277886497,4.00547945205479,4.01017612524462,4.01487279843444,4.01956947162427,4.02426614481409,4.02896281800391,4.03365949119374,4.03835616438356,4.04305283757339,4.04774951076321,4.05244618395303,4.05714285714286,4.06183953033268,4.0665362035225,4.07123287671233,4.07592954990215,4.08062622309198,4.0853228962818,4.09001956947163,4.09471624266145,4.09941291585127,4.1041095890411,4.10880626223092,4.11350293542074,4.11819960861057,4.12289628180039,4.12759295499022,4.13228962818004,4.13698630136986,4.14168297455969,4.14637964774951,4.15107632093934,4.15577299412916,4.16046966731898,4.16516634050881,4.16986301369863,4.17455968688845,4.17925636007828,4.1839530332681,4.18864970645793,4.19334637964775,4.19804305283757,4.2027397260274,4.20743639921722,4.21213307240705,4.21682974559687,4.22152641878669,4.22622309197652,4.23091976516634,4.23561643835616,4.24031311154599,4.24500978473581,4.24970645792564,4.25440313111546,4.25909980430528,4.26379647749511,4.26849315068493,4.27318982387476,4.27788649706458,4.2825831702544,4.28727984344423,4.29197651663405,4.29667318982388,4.3013698630137,4.30606653620352,4.31076320939335,4.31545988258317,4.32015655577299,4.32485322896282,4.32954990215264,4.33424657534247,4.33894324853229,4.34363992172211,4.34833659491194,4.35303326810176,4.35772994129159,4.36242661448141,4.36712328767123,4.37181996086106,4.37651663405088,4.3812133072407,4.38590998043053,4.39060665362035,4.39530332681018,4.4,4.4,4.4,4.39530332681018,4.39060665362035,4.38590998043053,4.3812133072407,4.37651663405088,4.37181996086106,4.36712328767123,4.36242661448141,4.35772994129159,4.35303326810176,4.34833659491194,4.34363992172211,4.33894324853229,4.33424657534247,4.32954990215264,4.32485322896282,4.32015655577299,4.31545988258317,4.31076320939335,4.30606653620352,4.3013698630137,4.29667318982388,4.29197651663405,4.28727984344423,4.2825831702544,4.27788649706458,4.27318982387476,4.26849315068493,4.26379647749511,4.25909980430528,4.25440313111546,4.24970645792564,4.24500978473581,4.24031311154599,4.23561643835616,4.23091976516634,4.22622309197652,4.22152641878669,4.21682974559687,4.21213307240705,4.20743639921722,4.2027397260274,4.19804305283757,4.19334637964775,4.18864970645793,4.1839530332681,4.17925636007828,4.17455968688845,4.16986301369863,4.16516634050881,4.16046966731898,4.15577299412916,4.15107632093934,4.14637964774951,4.14168297455969,4.13698630136986,4.13228962818004,4.12759295499022,4.12289628180039,4.11819960861057,4.11350293542074,4.10880626223092,4.1041095890411,4.09941291585127,4.09471624266145,4.09001956947163,4.0853228962818,4.08062622309198,4.07592954990215,4.07123287671233,4.0665362035225,4.06183953033268,4.05714285714286,4.05244618395303,4.04774951076321,4.04305283757339,4.03835616438356,4.03365949119374,4.02896281800391,4.02426614481409,4.01956947162427,4.01487279843444,4.01017612524462,4.00547945205479,4.00078277886497,3.99608610567515,3.99138943248532,3.9866927592955,3.98199608610568,3.97729941291585,3.97260273972603,3.9679060665362,3.96320939334638,3.95851272015656,3.95381604696673,3.94911937377691,3.94442270058708,3.93972602739726,3.93502935420744,3.93033268101761,3.92563600782779,3.92093933463797,3.91624266144814,3.91154598825832,3.90684931506849,3.90215264187867,3.89745596868885,3.89275929549902,3.8880626223092,3.88336594911937,3.87866927592955,3.87397260273973,3.8692759295499,3.86457925636008,3.85988258317026,3.85518590998043,3.85048923679061,3.84579256360078,3.84109589041096,3.83639921722114,3.83170254403131,3.82700587084149,3.82230919765166,3.81761252446184,3.81291585127202,3.80821917808219,3.80352250489237,3.79882583170254,3.79412915851272,3.7894324853229,3.78473581213307,3.78003913894325,3.77534246575342,3.7706457925636,3.76594911937378,3.76125244618395,3.75655577299413,3.75185909980431,3.74716242661448,3.74246575342466,3.73776908023483,3.73307240704501,3.72837573385519,3.72367906066536,3.71898238747554,3.71428571428571,3.70958904109589,3.70489236790607,3.70019569471624,3.69549902152642,3.6908023483366,3.68610567514677,3.68140900195695,3.67671232876712,3.6720156555773,3.66731898238748,3.66262230919765,3.65792563600783,3.653228962818,3.64853228962818,3.64383561643836,3.63913894324853,3.63444227005871,3.62974559686888,3.62504892367906,3.62035225048924,3.61565557729941,3.61095890410959,3.60626223091977,3.60156555772994,3.59686888454012,3.59217221135029,3.58747553816047,3.58277886497065,3.57808219178082,3.573385518591,3.56868884540117,3.56399217221135,3.55929549902153,3.5545988258317,3.54990215264188,3.54520547945206,3.54050880626223,3.53581213307241,3.53111545988258,3.52641878669276,3.52172211350294,3.51702544031311,3.51232876712329,3.50763209393346,3.50293542074364,3.49823874755382,3.49354207436399,3.48884540117417,3.48414872798434,3.47945205479452,3.4747553816047,3.47005870841487,3.46536203522505,3.46066536203523,3.4559686888454,3.45127201565558,3.44657534246575,3.44187866927593,3.43718199608611,3.43248532289628,3.42778864970646,3.42309197651663,3.41839530332681,3.41369863013699,3.40900195694716,3.40430528375734,3.39960861056751,3.39491193737769,3.39021526418787,3.38551859099804,3.38082191780822,3.3761252446184,3.37142857142857,3.36673189823875,3.36203522504892,3.3573385518591,3.35264187866928,3.34794520547945,3.34324853228963,3.3385518590998,3.33385518590998,3.32915851272016,3.32446183953033,3.31976516634051,3.31506849315068,3.31037181996086,3.30567514677104,3.30097847358121,3.29628180039139,3.29158512720157,3.28688845401174,3.28219178082192,3.27749510763209,3.27279843444227,3.26810176125245,3.26340508806262,3.2587084148728,3.25401174168297,3.24931506849315,3.24461839530333,3.2399217221135,3.23522504892368,3.23052837573386,3.22583170254403,3.22113502935421,3.21643835616438,3.21174168297456,3.20704500978474,3.20234833659491,3.19765166340509,3.19295499021526,3.18825831702544,3.18356164383562,3.17886497064579,3.17416829745597,3.16947162426615,3.16477495107632,3.1600782778865,3.15538160469667,3.15068493150685,3.14598825831703,3.1412915851272,3.13659491193738,3.13189823874755,3.12720156555773,3.12250489236791,3.11780821917808,3.11311154598826,3.10841487279843,3.10371819960861,3.09902152641879,3.09432485322896,3.08962818003914,3.08493150684932,3.08023483365949,3.07553816046967,3.07084148727984,3.06614481409002,3.0614481409002,3.05675146771037,3.05205479452055,3.04735812133072,3.0426614481409,3.03796477495108,3.03326810176125,3.02857142857143,3.0238747553816,3.01917808219178,3.01448140900196,3.00978473581213,3.00508806262231,3.00039138943249,2.99569471624266,2.99099804305284,2.98630136986301,2.98160469667319,2.97690802348337,2.97221135029354,2.96751467710372,2.96281800391389,2.95812133072407,2.95342465753425,2.94872798434442,2.9440313111546,2.93933463796477,2.93463796477495,2.92994129158513,2.9252446183953,2.92054794520548,2.91585127201566,2.91115459882583,2.90645792563601,2.90176125244618,2.89706457925636,2.89236790606654,2.88767123287671,2.88297455968689,2.87827788649706,2.87358121330724,2.86888454011742,2.86418786692759,2.85949119373777,2.85479452054795,2.85009784735812,2.8454011741683,2.84070450097847,2.83600782778865,2.83131115459883,2.826614481409,2.82191780821918,2.81722113502935,2.81252446183953,2.80782778864971,2.80313111545988,2.79843444227006,2.79373776908024,2.78904109589041,2.78434442270059,2.77964774951076,2.77495107632094,2.77025440313112,2.76555772994129,2.76086105675147,2.75616438356164,2.75146771037182,2.746771037182,2.74207436399217,2.73737769080235,2.73268101761252,2.7279843444227,2.72328767123288,2.71859099804305,2.71389432485323,2.70919765166341,2.70450097847358,2.69980430528376,2.69510763209393,2.69041095890411,2.68571428571429,2.68101761252446,2.67632093933464,2.67162426614481,2.66692759295499,2.66223091976517,2.65753424657534,2.65283757338552,2.64814090019569,2.64344422700587,2.63874755381605,2.63405088062622,2.6293542074364,2.62465753424658,2.61996086105675,2.61526418786693,2.6105675146771,2.60587084148728,2.60117416829746,2.59647749510763,2.59178082191781,2.58708414872798,2.58238747553816,2.57769080234834,2.57299412915851,2.56829745596869,2.56360078277886,2.55890410958904,2.55420743639922,2.54951076320939,2.54481409001957,2.54011741682975,2.53542074363992,2.5307240704501,2.52602739726027,2.52133072407045,2.51663405088063,2.5119373776908,2.50724070450098,2.50254403131115,2.49784735812133,2.49315068493151,2.48845401174168,2.48375733855186,2.47906066536204,2.47436399217221,2.46966731898239,2.46497064579256,2.46027397260274,2.45557729941292,2.45088062622309,2.44618395303327,2.44148727984344,2.43679060665362,2.4320939334638,2.42739726027397,2.42270058708415,2.41800391389432,2.4133072407045,2.40861056751468,2.40391389432485,2.39921722113503,2.39452054794521,2.38982387475538,2.38512720156556,2.38043052837573,2.37573385518591,2.37103718199609,2.36634050880626,2.36164383561644,2.35694716242661,2.35225048923679,2.34755381604697,2.34285714285714,2.33816046966732,2.3334637964775,2.32876712328767,2.32407045009785,2.31937377690802,2.3146771037182,2.30998043052838,2.30528375733855,2.30058708414873,2.2958904109589,2.29119373776908,2.28649706457926,2.28180039138943,2.27710371819961,2.27240704500978,2.26771037181996,2.26301369863014,2.25831702544031,2.25362035225049,2.24892367906067,2.24422700587084,2.23953033268102,2.23483365949119,2.23013698630137,2.22544031311155,2.22074363992172,2.2160469667319,2.21135029354207,2.20665362035225,2.20195694716243,2.1972602739726,2.19256360078278,2.18786692759295,2.18317025440313,2.17847358121331,2.17377690802348,2.16908023483366,2.16438356164384,2.15968688845401,2.15499021526419,2.15029354207436,2.14559686888454,2.14090019569472,2.13620352250489,2.13150684931507,2.12681017612524,2.12211350293542,2.1174168297456,2.11272015655577,2.10802348336595,2.10332681017613,2.0986301369863,2.09393346379648,2.08923679060665,2.08454011741683,2.07984344422701,2.07514677103718,2.07045009784736,2.06575342465753,2.06105675146771,2.05636007827789,2.05166340508806,2.04696673189824,2.04227005870841,2.03757338551859,2.03287671232877,2.02818003913894,2.02348336594912,2.0187866927593,2.01409001956947,2.00939334637965,2.00469667318982,2,2],"y":[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1.84033790345484e-07,2.2853426383547e-07,2.81638882032015e-07,3.44230725292613e-07,4.2493855936226e-07,5.22896699330664e-07,6.38628765033152e-07,7.77240455257003e-07,9.54442252151286e-07,1.16400623334322e-06,1.40919822496943e-06,1.71353489309245e-06,2.08555738013627e-06,2.52132987117373e-06,3.02699191694683e-06,3.67498630648743e-06,4.43404343946844e-06,5.31497327604508e-06,6.37129181145973e-06,7.66826021147331e-06,9.17363173833267e-06,1.09051843582715e-05,1.30466013104178e-05,1.55693790766072e-05,1.84716564653888e-05,2.1844871265149e-05,2.59940035875599e-05,3.07634816787201e-05,3.62037825143506e-05,4.27069769497038e-05,5.03974479152012e-05,5.91625254496631e-05,6.90789984110278e-05,8.12419632234127e-05,9.50951140316324e-05,0.000110754782685505,0.000128908508209692,0.000150399635065689,0.000174653511942206,0.000201855199821985,0.000234123641718926,0.000270988705012861,0.000312263302160606,0.000358811152005025,0.000413826935735924,0.000475277277157446,0.000543554828415966,0.000622148075101515,0.000711957748821955,0.000811497955961008,0.000921292226625866,0.00104992256970681,0.00119235041104039,0.00134903893951934,0.00152459264478417,0.00172467891711363,0.00194410476699274,0.00218378035300811,0.00245639785697583,0.00275802794865437,0.00308640264911565,0.00344555668380571,0.00385265441131715,0.00429418107637778,0.0047714751475457,0.00530015332525149,0.0058828930936528,0.00651035908334341,0.00718403908317059,0.00793723728418436,0.00874671924406949,0.00961222292556928,0.0105484679832111,0.0115733411636686,0.0126641902592908,0.0138225607281233,0.0150841313295383,0.0164329431349431,0.0178584524919583,0.0193684981222109,0.0210045529679746,0.0227248459062823,0.0245303060777163,0.0264508543036004,0.0284863920734941,0.0306112991116918,0.0328259613376525,0.0351822843908106,0.0376329022066828,0.0401726525119879,0.042818028889187,0.0455864117287604,0.0484383185667221,0.0513723418704148,0.0544197019123379,0.0575536950539074,0.0607574436733334,0.0640324668429251,0.0674002805108313,0.0708197591531191,0.0742874315886669,0.0778098328433085,0.0813735350826111,0.0849610881437274,0.0885682757158961,0.0921889282076234,0.0958056534637868,0.0994130482220084,0.103001030527899,0.106546500845226,0.110051741025885,0.113511816760658,0.116892457888696,0.12019119621,0.123414372717008,0.126552641006552,0.12954524734832,0.132434832944468,0.135217577510298,0.137853046149395,0.140326031428264,0.142671146579888,0.144885615876566,0.146889561739054,0.148740911292008,0.150449478406859,0.15198943449528,0.153316219748773,0.154496578115448,0.155530791353444,0.156360171718604,0.157017648573266,0.157535765413257,0.157911452966482,0.158078251055373,0.158120420524694,0.158041090327031,0.157813085057713,0.15744229762141,0.156972494951185,0.156407665456511,0.155713019841026,0.154938296442138,0.154096178536372,0.153183878894262,0.152200364672804,0.151179224801673,0.150125254000548,0.149039184638345,0.147940505335229,0.146839046960295,0.145739828454172,0.144663708087972,0.143613655441685,0.142594023129079,0.141622379289341,0.140717125559719,0.139868697763689,0.139081087714349,0.138399461168594,0.137806439927341,0.137297940205196,0.136891141676513,0.136630842816111,0.136476793593303,0.136432271430637,0.136548357738717,0.136818668634186,0.137218439174672,0.137751722679454,0.138512208851812,0.139421038272943,0.140481102364407,0.141736773716055,0.143216529611799,0.144865351241612,0.146685980988788,0.148773669413778,0.151069173057493,0.153552966790002,0.156251912995903,0.159247725264458,0.16244655941274,0.165850504339008,0.169540816213884,0.173499410584486,0.177674619859843,0.182068023096942,0.186819584275105,0.19179722966214,0.19700001199996,0.202483170915393,0.208281015354053,0.214306256389574,0.220558796152456,0.227147832752214,0.233992560910435,0.24106054098695,0.24837447567592,0.256018739386273,0.263876854548977,0.271946788226113,0.280298104370714,0.288904552240094,0.297707438720354,0.306703990264357,0.315998054347019,0.325473105527788,0.335121577291887,0.34497559299013,0.355050829337195,0.365276424715737,0.37564855058259,0.386223348780179,0.396944878563692,0.407788192242247,0.418759340815278,0.42989322291478,0.441124247064764,0.452448632688187,0.463888586507222,0.475426841387861,0.487035726777133,0.498711833585232,0.510481565091021,0.522304281967467,0.534175178730826,0.54609930956346,0.558076676458306,0.570087415314591,0.582129630714014,0.594212216861553,0.606323023542415,0.618456646920419,0.630613661555713,0.642801813042923,0.655009708526198,0.667237473149477,0.679492093496584,0.691774954814782,0.704081225517558,0.716411863424543,0.728785678811423,0.741192522730323,0.753632745285754,0.766115990888277,0.778657693117111,0.791244513655684,0.803878406230625,0.816586300073639,0.829360649531249,0.842193736227435,0.855091478984124,0.868089469582949,0.881154197253722,0.894286193312498,0.907507474401836,0.920817351979775,0.934193045424858,0.947633458055248,0.961167724503055,0.97475835723908,0.98839744870286,1.00208713282086,1.01581979496172,1.02956907315642,1.04332828121262,1.05707954639462,1.0707973436766,1.0844725318832,1.09809147486588,1.11158142474358,1.12495966110595,1.13821359816268,1.15127680635001,1.16409752627801,1.17670902896724,1.1890970965508,1.20108538800928,1.21275171529589,1.22410194271992,1.23505850924813,1.24546760900266,1.25546987357484,1.26505173950443,1.27400034580871,1.28236752117901,1.29023715925144,1.2975787491896,1.30407199659917,1.31001079825788,1.31538908255368,1.32003514333135,1.32390220480122,1.32718473767223,1.32988150429806,1.33168398759776,1.33285408756854,1.33345207766078,1.3333984529007,1.33253961017607,1.33115300714956,1.32924845793911,1.32664978270313,1.323482836764,1.31987328989077,1.31582600660466,1.31115462844902,1.30613331174907,1.3007778490564,1.29503294096056,1.28892950812498,1.28259224685106,1.27603688371296,1.269206390738,1.26221773371369,1.25510401552116,1.24786925728267,1.24052557348351,1.23313375151079,1.22570379161091,1.21824736467059,1.21079256431439,1.20334873158424,1.19592201934557,1.18854030138576,1.18119246238334,1.17387908857115,1.16661008760976,1.15938561277917,1.15218855798511,1.14501591966308,1.13786632565762,1.13071906497421,1.12356606435003,1.11639782281801,1.10918318408309,1.10192144378802,1.09460554916649,1.08719618930507,1.07967798167303,1.07206173432156,1.06434076367813,1.05642430642159,1.0483715492245,1.04017881495961,1.03180339596573,1.02321115360917,1.01445743506078,1.00554030853221,0.996376413623348,0.987023833747662,0.977507138664726,0.967810302070069,0.957877595289746,0.947797731977407,0.937575098344726,0.927171298558008,0.916616926929818,0.905954723800686,0.895191118112021,0.88429427796177,0.873332730983204,0.862316015023503,0.851246867420805,0.840146948377293,0.829040950214626,0.817936510709853,0.806859137289537,0.795824495428932,0.784836023586252,0.773906943108009,0.763084120711456,0.75234262414885,0.741686677096001,0.731155647684415,0.720754264646454,0.71045742656401,0.700266996214776,0.690249169769024,0.68034359924745,0.670545146253383,0.660874201084445,0.651344159589282,0.641907622591453,0.632561267779974,0.623334194360553,0.614188925905395,0.605106671295177,0.596085999993577,0.587131446448375,0.578206130947158,0.569304410907068,0.560419974606214,0.55153319371053,0.542635009331947,0.533720101785936,0.524759062685862,0.515754689531189,0.506704672233992,0.497593023552845,0.488395994528346,0.479134230977059,0.46980586711524,0.46037705263464,0.450864647734606,0.441282254801418,0.43162596004948,0.421868057121755,0.412051192893027,0.40217868611922,0.392241529850939,0.382255033030086,0.372240419174163,0.362203028403464,0.352153169464938,0.342113413213904,0.332091478217228,0.322103059559452,0.31218424901256,0.302330609836971,0.292549983113088,0.282897737454828,0.27337797178879,0.263980830434452,0.254721867722149,0.245700725192676,0.236848722973246,0.228172508148987,0.219739691327768,0.211576469217528,0.203626443693732,0.195894712136604,0.188516957608294,0.181397013786684,0.174519316236068,0.167929035201407,0.161704362282975,0.155732838938364,0.150014871641465,0.144655343278543,0.139599588583129,0.134792229647836,0.130240408799968,0.126068628514655,0.122127852146874,0.118414103771874,0.11498260249276,0.11182925547336,0.108873638800827,0.106110461792373,0.103620977580951,0.10130986001479,0.0991541077733019,0.0971674151868268,0.0953715519332741,0.0936910700913721,0.0921195848025772,0.0906822554956272,0.0893419854459238,0.0880713812319322,0.086865459992192,0.0857363148326099,0.0846410631498973,0.0835746573606258,0.0825339299678123,0.0815041589731559,0.080474534459741,0.0794411095553626,0.0783890780883621,0.0773144665295433,0.0762162409152311,0.0750872192029783,0.0739102425980991,0.0726976548993097,0.0714483297732899,0.0701455020508991,0.0687933381463363,0.067401335707756,0.0659691760839394,0.0644744611507541,0.0629431069481601,0.0613762319693335,0.0597671648089618,0.0581171802691691,0.0564405603351044,0.0547390121584238,0.053006461188114,0.0512566000754296,0.0494940519596653,0.0477203281122103,0.0459401004539019,0.0441604564544657,0.0423834906943666,0.0406158570139207,0.0388634551370866,0.0371264132973867,0.0354066456246166,0.0337224420637632,0.0320647683254247,0.0304349605706284,0.0288427101428474,0.0272971417539268,0.0257875786667103,0.0243150710172822,0.0229008068263108,0.0215333233178297,0.0202077448408875,0.0189294557282289,0.0177171787473768,0.0165489907607368,0.0154249798261166,0],"text":["density: 1.542498e-02<br />Sepal.Width: 2.000000","density: 1.654899e-02<br />Sepal.Width: 2.004697","density: 1.771718e-02<br />Sepal.Width: 2.009393","density: 1.892946e-02<br />Sepal.Width: 2.014090","density: 2.020774e-02<br />Sepal.Width: 2.018787","density: 2.153332e-02<br />Sepal.Width: 2.023483","density: 2.290081e-02<br />Sepal.Width: 2.028180","density: 2.431507e-02<br />Sepal.Width: 2.032877","density: 2.578758e-02<br />Sepal.Width: 2.037573","density: 2.729714e-02<br />Sepal.Width: 2.042270","density: 2.884271e-02<br />Sepal.Width: 2.046967","density: 3.043496e-02<br />Sepal.Width: 2.051663","density: 3.206477e-02<br />Sepal.Width: 2.056360","density: 3.372244e-02<br />Sepal.Width: 2.061057","density: 3.540665e-02<br />Sepal.Width: 2.065753","density: 3.712641e-02<br />Sepal.Width: 2.070450","density: 3.886346e-02<br />Sepal.Width: 2.075147","density: 4.061586e-02<br />Sepal.Width: 2.079843","density: 4.238349e-02<br />Sepal.Width: 2.084540","density: 4.416046e-02<br />Sepal.Width: 2.089237","density: 4.594010e-02<br />Sepal.Width: 2.093933","density: 4.772033e-02<br />Sepal.Width: 2.098630","density: 4.949405e-02<br />Sepal.Width: 2.103327","density: 5.125660e-02<br />Sepal.Width: 2.108023","density: 5.300646e-02<br />Sepal.Width: 2.112720","density: 5.473901e-02<br />Sepal.Width: 2.117417","density: 5.644056e-02<br />Sepal.Width: 2.122114","density: 5.811718e-02<br />Sepal.Width: 2.126810","density: 5.976716e-02<br />Sepal.Width: 2.131507","density: 6.137623e-02<br />Sepal.Width: 2.136204","density: 6.294311e-02<br />Sepal.Width: 2.140900","density: 6.447446e-02<br />Sepal.Width: 2.145597","density: 6.596918e-02<br />Sepal.Width: 2.150294","density: 6.740134e-02<br />Sepal.Width: 2.154990","density: 6.879334e-02<br />Sepal.Width: 2.159687","density: 7.014550e-02<br />Sepal.Width: 2.164384","density: 7.144833e-02<br />Sepal.Width: 2.169080","density: 7.269765e-02<br />Sepal.Width: 2.173777","density: 7.391024e-02<br />Sepal.Width: 2.178474","density: 7.508722e-02<br />Sepal.Width: 2.183170","density: 7.621624e-02<br />Sepal.Width: 2.187867","density: 7.731447e-02<br />Sepal.Width: 2.192564","density: 7.838908e-02<br />Sepal.Width: 2.197260","density: 7.944111e-02<br />Sepal.Width: 2.201957","density: 8.047453e-02<br />Sepal.Width: 2.206654","density: 8.150416e-02<br />Sepal.Width: 2.211350","density: 8.253393e-02<br />Sepal.Width: 2.216047","density: 8.357466e-02<br />Sepal.Width: 2.220744","density: 8.464106e-02<br />Sepal.Width: 2.225440","density: 8.573631e-02<br />Sepal.Width: 2.230137","density: 8.686546e-02<br />Sepal.Width: 2.234834","density: 8.807138e-02<br />Sepal.Width: 2.239530","density: 8.934199e-02<br />Sepal.Width: 2.244227","density: 9.068226e-02<br />Sepal.Width: 2.248924","density: 9.211958e-02<br />Sepal.Width: 2.253620","density: 9.369107e-02<br />Sepal.Width: 2.258317","density: 9.537155e-02<br />Sepal.Width: 2.263014","density: 9.716742e-02<br />Sepal.Width: 2.267710","density: 9.915411e-02<br />Sepal.Width: 2.272407","density: 1.013099e-01<br />Sepal.Width: 2.277104","density: 1.036210e-01<br />Sepal.Width: 2.281800","density: 1.061105e-01<br />Sepal.Width: 2.286497","density: 1.088736e-01<br />Sepal.Width: 2.291194","density: 1.118293e-01<br />Sepal.Width: 2.295890","density: 1.149826e-01<br />Sepal.Width: 2.300587","density: 1.184141e-01<br />Sepal.Width: 2.305284","density: 1.221279e-01<br />Sepal.Width: 2.309980","density: 1.260686e-01<br />Sepal.Width: 2.314677","density: 1.302404e-01<br />Sepal.Width: 2.319374","density: 1.347922e-01<br />Sepal.Width: 2.324070","density: 1.395996e-01<br />Sepal.Width: 2.328767","density: 1.446553e-01<br />Sepal.Width: 2.333464","density: 1.500149e-01<br />Sepal.Width: 2.338160","density: 1.557328e-01<br />Sepal.Width: 2.342857","density: 1.617044e-01<br />Sepal.Width: 2.347554","density: 1.679290e-01<br />Sepal.Width: 2.352250","density: 1.745193e-01<br />Sepal.Width: 2.356947","density: 1.813970e-01<br />Sepal.Width: 2.361644","density: 1.885170e-01<br />Sepal.Width: 2.366341","density: 1.958947e-01<br />Sepal.Width: 2.371037","density: 2.036264e-01<br />Sepal.Width: 2.375734","density: 2.115765e-01<br />Sepal.Width: 2.380431","density: 2.197397e-01<br />Sepal.Width: 2.385127","density: 2.281725e-01<br />Sepal.Width: 2.389824","density: 2.368487e-01<br />Sepal.Width: 2.394521","density: 2.457007e-01<br />Sepal.Width: 2.399217","density: 2.547219e-01<br />Sepal.Width: 2.403914","density: 2.639808e-01<br />Sepal.Width: 2.408611","density: 2.733780e-01<br />Sepal.Width: 2.413307","density: 2.828977e-01<br />Sepal.Width: 2.418004","density: 2.925500e-01<br />Sepal.Width: 2.422701","density: 3.023306e-01<br />Sepal.Width: 2.427397","density: 3.121842e-01<br />Sepal.Width: 2.432094","density: 3.221031e-01<br />Sepal.Width: 2.436791","density: 3.320915e-01<br />Sepal.Width: 2.441487","density: 3.421134e-01<br />Sepal.Width: 2.446184","density: 3.521532e-01<br />Sepal.Width: 2.450881","density: 3.622030e-01<br />Sepal.Width: 2.455577","density: 3.722404e-01<br />Sepal.Width: 2.460274","density: 3.822550e-01<br />Sepal.Width: 2.464971","density: 3.922415e-01<br />Sepal.Width: 2.469667","density: 4.021787e-01<br />Sepal.Width: 2.474364","density: 4.120512e-01<br />Sepal.Width: 2.479061","density: 4.218681e-01<br />Sepal.Width: 2.483757","density: 4.316260e-01<br />Sepal.Width: 2.488454","density: 4.412823e-01<br />Sepal.Width: 2.493151","density: 4.508646e-01<br />Sepal.Width: 2.497847","density: 4.603771e-01<br />Sepal.Width: 2.502544","density: 4.698059e-01<br />Sepal.Width: 2.507241","density: 4.791342e-01<br />Sepal.Width: 2.511937","density: 4.883960e-01<br />Sepal.Width: 2.516634","density: 4.975930e-01<br />Sepal.Width: 2.521331","density: 5.067047e-01<br />Sepal.Width: 2.526027","density: 5.157547e-01<br />Sepal.Width: 2.530724","density: 5.247591e-01<br />Sepal.Width: 2.535421","density: 5.337201e-01<br />Sepal.Width: 2.540117","density: 5.426350e-01<br />Sepal.Width: 2.544814","density: 5.515332e-01<br />Sepal.Width: 2.549511","density: 5.604200e-01<br />Sepal.Width: 2.554207","density: 5.693044e-01<br />Sepal.Width: 2.558904","density: 5.782061e-01<br />Sepal.Width: 2.563601","density: 5.871314e-01<br />Sepal.Width: 2.568297","density: 5.960860e-01<br />Sepal.Width: 2.572994","density: 6.051067e-01<br />Sepal.Width: 2.577691","density: 6.141889e-01<br />Sepal.Width: 2.582387","density: 6.233342e-01<br />Sepal.Width: 2.587084","density: 6.325613e-01<br />Sepal.Width: 2.591781","density: 6.419076e-01<br />Sepal.Width: 2.596477","density: 6.513442e-01<br />Sepal.Width: 2.601174","density: 6.608742e-01<br />Sepal.Width: 2.605871","density: 6.705451e-01<br />Sepal.Width: 2.610568","density: 6.803436e-01<br />Sepal.Width: 2.615264","density: 6.902492e-01<br />Sepal.Width: 2.619961","density: 7.002670e-01<br />Sepal.Width: 2.624658","density: 7.104574e-01<br />Sepal.Width: 2.629354","density: 7.207543e-01<br />Sepal.Width: 2.634051","density: 7.311556e-01<br />Sepal.Width: 2.638748","density: 7.416867e-01<br />Sepal.Width: 2.643444","density: 7.523426e-01<br />Sepal.Width: 2.648141","density: 7.630841e-01<br />Sepal.Width: 2.652838","density: 7.739069e-01<br />Sepal.Width: 2.657534","density: 7.848360e-01<br />Sepal.Width: 2.662231","density: 7.958245e-01<br />Sepal.Width: 2.666928","density: 8.068591e-01<br />Sepal.Width: 2.671624","density: 8.179365e-01<br />Sepal.Width: 2.676321","density: 8.290410e-01<br />Sepal.Width: 2.681018","density: 8.401469e-01<br />Sepal.Width: 2.685714","density: 8.512469e-01<br />Sepal.Width: 2.690411","density: 8.623160e-01<br />Sepal.Width: 2.695108","density: 8.733327e-01<br />Sepal.Width: 2.699804","density: 8.842943e-01<br />Sepal.Width: 2.704501","density: 8.951911e-01<br />Sepal.Width: 2.709198","density: 9.059547e-01<br />Sepal.Width: 2.713894","density: 9.166169e-01<br />Sepal.Width: 2.718591","density: 9.271713e-01<br />Sepal.Width: 2.723288","density: 9.375751e-01<br />Sepal.Width: 2.727984","density: 9.477977e-01<br />Sepal.Width: 2.732681","density: 9.578776e-01<br />Sepal.Width: 2.737378","density: 9.678103e-01<br />Sepal.Width: 2.742074","density: 9.775071e-01<br />Sepal.Width: 2.746771","density: 9.870238e-01<br />Sepal.Width: 2.751468","density: 9.963764e-01<br />Sepal.Width: 2.756164","density: 1.005540e+00<br />Sepal.Width: 2.760861","density: 1.014457e+00<br />Sepal.Width: 2.765558","density: 1.023211e+00<br />Sepal.Width: 2.770254","density: 1.031803e+00<br />Sepal.Width: 2.774951","density: 1.040179e+00<br />Sepal.Width: 2.779648","density: 1.048372e+00<br />Sepal.Width: 2.784344","density: 1.056424e+00<br />Sepal.Width: 2.789041","density: 1.064341e+00<br />Sepal.Width: 2.793738","density: 1.072062e+00<br />Sepal.Width: 2.798434","density: 1.079678e+00<br />Sepal.Width: 2.803131","density: 1.087196e+00<br />Sepal.Width: 2.807828","density: 1.094606e+00<br />Sepal.Width: 2.812524","density: 1.101921e+00<br />Sepal.Width: 2.817221","density: 1.109183e+00<br />Sepal.Width: 2.821918","density: 1.116398e+00<br />Sepal.Width: 2.826614","density: 1.123566e+00<br />Sepal.Width: 2.831311","density: 1.130719e+00<br />Sepal.Width: 2.836008","density: 1.137866e+00<br />Sepal.Width: 2.840705","density: 1.145016e+00<br />Sepal.Width: 2.845401","density: 1.152189e+00<br />Sepal.Width: 2.850098","density: 1.159386e+00<br />Sepal.Width: 2.854795","density: 1.166610e+00<br />Sepal.Width: 2.859491","density: 1.173879e+00<br />Sepal.Width: 2.864188","density: 1.181192e+00<br />Sepal.Width: 2.868885","density: 1.188540e+00<br />Sepal.Width: 2.873581","density: 1.195922e+00<br />Sepal.Width: 2.878278","density: 1.203349e+00<br />Sepal.Width: 2.882975","density: 1.210793e+00<br />Sepal.Width: 2.887671","density: 1.218247e+00<br />Sepal.Width: 2.892368","density: 1.225704e+00<br />Sepal.Width: 2.897065","density: 1.233134e+00<br />Sepal.Width: 2.901761","density: 1.240526e+00<br />Sepal.Width: 2.906458","density: 1.247869e+00<br />Sepal.Width: 2.911155","density: 1.255104e+00<br />Sepal.Width: 2.915851","density: 1.262218e+00<br />Sepal.Width: 2.920548","density: 1.269206e+00<br />Sepal.Width: 2.925245","density: 1.276037e+00<br />Sepal.Width: 2.929941","density: 1.282592e+00<br />Sepal.Width: 2.934638","density: 1.288930e+00<br />Sepal.Width: 2.939335","density: 1.295033e+00<br />Sepal.Width: 2.944031","density: 1.300778e+00<br />Sepal.Width: 2.948728","density: 1.306133e+00<br />Sepal.Width: 2.953425","density: 1.311155e+00<br />Sepal.Width: 2.958121","density: 1.315826e+00<br />Sepal.Width: 2.962818","density: 1.319873e+00<br />Sepal.Width: 2.967515","density: 1.323483e+00<br />Sepal.Width: 2.972211","density: 1.326650e+00<br />Sepal.Width: 2.976908","density: 1.329248e+00<br />Sepal.Width: 2.981605","density: 1.331153e+00<br />Sepal.Width: 2.986301","density: 1.332540e+00<br />Sepal.Width: 2.990998","density: 1.333398e+00<br />Sepal.Width: 2.995695","density: 1.333452e+00<br />Sepal.Width: 3.000391","density: 1.332854e+00<br />Sepal.Width: 3.005088","density: 1.331684e+00<br />Sepal.Width: 3.009785","density: 1.329882e+00<br />Sepal.Width: 3.014481","density: 1.327185e+00<br />Sepal.Width: 3.019178","density: 1.323902e+00<br />Sepal.Width: 3.023875","density: 1.320035e+00<br />Sepal.Width: 3.028571","density: 1.315389e+00<br />Sepal.Width: 3.033268","density: 1.310011e+00<br />Sepal.Width: 3.037965","density: 1.304072e+00<br />Sepal.Width: 3.042661","density: 1.297579e+00<br />Sepal.Width: 3.047358","density: 1.290237e+00<br />Sepal.Width: 3.052055","density: 1.282368e+00<br />Sepal.Width: 3.056751","density: 1.274000e+00<br />Sepal.Width: 3.061448","density: 1.265052e+00<br />Sepal.Width: 3.066145","density: 1.255470e+00<br />Sepal.Width: 3.070841","density: 1.245468e+00<br />Sepal.Width: 3.075538","density: 1.235059e+00<br />Sepal.Width: 3.080235","density: 1.224102e+00<br />Sepal.Width: 3.084932","density: 1.212752e+00<br />Sepal.Width: 3.089628","density: 1.201085e+00<br />Sepal.Width: 3.094325","density: 1.189097e+00<br />Sepal.Width: 3.099022","density: 1.176709e+00<br />Sepal.Width: 3.103718","density: 1.164098e+00<br />Sepal.Width: 3.108415","density: 1.151277e+00<br />Sepal.Width: 3.113112","density: 1.138214e+00<br />Sepal.Width: 3.117808","density: 1.124960e+00<br />Sepal.Width: 3.122505","density: 1.111581e+00<br />Sepal.Width: 3.127202","density: 1.098091e+00<br />Sepal.Width: 3.131898","density: 1.084473e+00<br />Sepal.Width: 3.136595","density: 1.070797e+00<br />Sepal.Width: 3.141292","density: 1.057080e+00<br />Sepal.Width: 3.145988","density: 1.043328e+00<br />Sepal.Width: 3.150685","density: 1.029569e+00<br />Sepal.Width: 3.155382","density: 1.015820e+00<br />Sepal.Width: 3.160078","density: 1.002087e+00<br />Sepal.Width: 3.164775","density: 9.883974e-01<br />Sepal.Width: 3.169472","density: 9.747584e-01<br />Sepal.Width: 3.174168","density: 9.611677e-01<br />Sepal.Width: 3.178865","density: 9.476335e-01<br />Sepal.Width: 3.183562","density: 9.341930e-01<br />Sepal.Width: 3.188258","density: 9.208174e-01<br />Sepal.Width: 3.192955","density: 9.075075e-01<br />Sepal.Width: 3.197652","density: 8.942862e-01<br />Sepal.Width: 3.202348","density: 8.811542e-01<br />Sepal.Width: 3.207045","density: 8.680895e-01<br />Sepal.Width: 3.211742","density: 8.550915e-01<br />Sepal.Width: 3.216438","density: 8.421937e-01<br />Sepal.Width: 3.221135","density: 8.293606e-01<br />Sepal.Width: 3.225832","density: 8.165863e-01<br />Sepal.Width: 3.230528","density: 8.038784e-01<br />Sepal.Width: 3.235225","density: 7.912445e-01<br />Sepal.Width: 3.239922","density: 7.786577e-01<br />Sepal.Width: 3.244618","density: 7.661160e-01<br />Sepal.Width: 3.249315","density: 7.536327e-01<br />Sepal.Width: 3.254012","density: 7.411925e-01<br />Sepal.Width: 3.258708","density: 7.287857e-01<br />Sepal.Width: 3.263405","density: 7.164119e-01<br />Sepal.Width: 3.268102","density: 7.040812e-01<br />Sepal.Width: 3.272798","density: 6.917750e-01<br />Sepal.Width: 3.277495","density: 6.794921e-01<br />Sepal.Width: 3.282192","density: 6.672375e-01<br />Sepal.Width: 3.286888","density: 6.550097e-01<br />Sepal.Width: 3.291585","density: 6.428018e-01<br />Sepal.Width: 3.296282","density: 6.306137e-01<br />Sepal.Width: 3.300978","density: 6.184566e-01<br />Sepal.Width: 3.305675","density: 6.063230e-01<br />Sepal.Width: 3.310372","density: 5.942122e-01<br />Sepal.Width: 3.315068","density: 5.821296e-01<br />Sepal.Width: 3.319765","density: 5.700874e-01<br />Sepal.Width: 3.324462","density: 5.580767e-01<br />Sepal.Width: 3.329159","density: 5.460993e-01<br />Sepal.Width: 3.333855","density: 5.341752e-01<br />Sepal.Width: 3.338552","density: 5.223043e-01<br />Sepal.Width: 3.343249","density: 5.104816e-01<br />Sepal.Width: 3.347945","density: 4.987118e-01<br />Sepal.Width: 3.352642","density: 4.870357e-01<br />Sepal.Width: 3.357339","density: 4.754268e-01<br />Sepal.Width: 3.362035","density: 4.638886e-01<br />Sepal.Width: 3.366732","density: 4.524486e-01<br />Sepal.Width: 3.371429","density: 4.411242e-01<br />Sepal.Width: 3.376125","density: 4.298932e-01<br />Sepal.Width: 3.380822","density: 4.187593e-01<br />Sepal.Width: 3.385519","density: 4.077882e-01<br />Sepal.Width: 3.390215","density: 3.969449e-01<br />Sepal.Width: 3.394912","density: 3.862233e-01<br />Sepal.Width: 3.399609","density: 3.756486e-01<br />Sepal.Width: 3.404305","density: 3.652764e-01<br />Sepal.Width: 3.409002","density: 3.550508e-01<br />Sepal.Width: 3.413699","density: 3.449756e-01<br />Sepal.Width: 3.418395","density: 3.351216e-01<br />Sepal.Width: 3.423092","density: 3.254731e-01<br />Sepal.Width: 3.427789","density: 3.159981e-01<br />Sepal.Width: 3.432485","density: 3.067040e-01<br />Sepal.Width: 3.437182","density: 2.977074e-01<br />Sepal.Width: 3.441879","density: 2.889046e-01<br />Sepal.Width: 3.446575","density: 2.802981e-01<br />Sepal.Width: 3.451272","density: 2.719468e-01<br />Sepal.Width: 3.455969","density: 2.638769e-01<br />Sepal.Width: 3.460665","density: 2.560187e-01<br />Sepal.Width: 3.465362","density: 2.483745e-01<br />Sepal.Width: 3.470059","density: 2.410605e-01<br />Sepal.Width: 3.474755","density: 2.339926e-01<br />Sepal.Width: 3.479452","density: 2.271478e-01<br />Sepal.Width: 3.484149","density: 2.205588e-01<br />Sepal.Width: 3.488845","density: 2.143063e-01<br />Sepal.Width: 3.493542","density: 2.082810e-01<br />Sepal.Width: 3.498239","density: 2.024832e-01<br />Sepal.Width: 3.502935","density: 1.970000e-01<br />Sepal.Width: 3.507632","density: 1.917972e-01<br />Sepal.Width: 3.512329","density: 1.868196e-01<br />Sepal.Width: 3.517025","density: 1.820680e-01<br />Sepal.Width: 3.521722","density: 1.776746e-01<br />Sepal.Width: 3.526419","density: 1.734994e-01<br />Sepal.Width: 3.531115","density: 1.695408e-01<br />Sepal.Width: 3.535812","density: 1.658505e-01<br />Sepal.Width: 3.540509","density: 1.624466e-01<br />Sepal.Width: 3.545205","density: 1.592477e-01<br />Sepal.Width: 3.549902","density: 1.562519e-01<br />Sepal.Width: 3.554599","density: 1.535530e-01<br />Sepal.Width: 3.559295","density: 1.510692e-01<br />Sepal.Width: 3.563992","density: 1.487737e-01<br />Sepal.Width: 3.568689","density: 1.466860e-01<br />Sepal.Width: 3.573386","density: 1.448654e-01<br />Sepal.Width: 3.578082","density: 1.432165e-01<br />Sepal.Width: 3.582779","density: 1.417368e-01<br />Sepal.Width: 3.587476","density: 1.404811e-01<br />Sepal.Width: 3.592172","density: 1.394210e-01<br />Sepal.Width: 3.596869","density: 1.385122e-01<br />Sepal.Width: 3.601566","density: 1.377517e-01<br />Sepal.Width: 3.606262","density: 1.372184e-01<br />Sepal.Width: 3.610959","density: 1.368187e-01<br />Sepal.Width: 3.615656","density: 1.365484e-01<br />Sepal.Width: 3.620352","density: 1.364323e-01<br />Sepal.Width: 3.625049","density: 1.364768e-01<br />Sepal.Width: 3.629746","density: 1.366308e-01<br />Sepal.Width: 3.634442","density: 1.368911e-01<br />Sepal.Width: 3.639139","density: 1.372979e-01<br />Sepal.Width: 3.643836","density: 1.378064e-01<br />Sepal.Width: 3.648532","density: 1.383995e-01<br />Sepal.Width: 3.653229","density: 1.390811e-01<br />Sepal.Width: 3.657926","density: 1.398687e-01<br />Sepal.Width: 3.662622","density: 1.407171e-01<br />Sepal.Width: 3.667319","density: 1.416224e-01<br />Sepal.Width: 3.672016","density: 1.425940e-01<br />Sepal.Width: 3.676712","density: 1.436137e-01<br />Sepal.Width: 3.681409","density: 1.446637e-01<br />Sepal.Width: 3.686106","density: 1.457398e-01<br />Sepal.Width: 3.690802","density: 1.468390e-01<br />Sepal.Width: 3.695499","density: 1.479405e-01<br />Sepal.Width: 3.700196","density: 1.490392e-01<br />Sepal.Width: 3.704892","density: 1.501253e-01<br />Sepal.Width: 3.709589","density: 1.511792e-01<br />Sepal.Width: 3.714286","density: 1.522004e-01<br />Sepal.Width: 3.718982","density: 1.531839e-01<br />Sepal.Width: 3.723679","density: 1.540962e-01<br />Sepal.Width: 3.728376","density: 1.549383e-01<br />Sepal.Width: 3.733072","density: 1.557130e-01<br />Sepal.Width: 3.737769","density: 1.564077e-01<br />Sepal.Width: 3.742466","density: 1.569725e-01<br />Sepal.Width: 3.747162","density: 1.574423e-01<br />Sepal.Width: 3.751859","density: 1.578131e-01<br />Sepal.Width: 3.756556","density: 1.580411e-01<br />Sepal.Width: 3.761252","density: 1.581204e-01<br />Sepal.Width: 3.765949","density: 1.580783e-01<br />Sepal.Width: 3.770646","density: 1.579115e-01<br />Sepal.Width: 3.775342","density: 1.575358e-01<br />Sepal.Width: 3.780039","density: 1.570176e-01<br />Sepal.Width: 3.784736","density: 1.563602e-01<br />Sepal.Width: 3.789432","density: 1.555308e-01<br />Sepal.Width: 3.794129","density: 1.544966e-01<br />Sepal.Width: 3.798826","density: 1.533162e-01<br />Sepal.Width: 3.803523","density: 1.519894e-01<br />Sepal.Width: 3.808219","density: 1.504495e-01<br />Sepal.Width: 3.812916","density: 1.487409e-01<br />Sepal.Width: 3.817613","density: 1.468896e-01<br />Sepal.Width: 3.822309","density: 1.448856e-01<br />Sepal.Width: 3.827006","density: 1.426711e-01<br />Sepal.Width: 3.831703","density: 1.403260e-01<br />Sepal.Width: 3.836399","density: 1.378530e-01<br />Sepal.Width: 3.841096","density: 1.352176e-01<br />Sepal.Width: 3.845793","density: 1.324348e-01<br />Sepal.Width: 3.850489","density: 1.295452e-01<br />Sepal.Width: 3.855186","density: 1.265526e-01<br />Sepal.Width: 3.859883","density: 1.234144e-01<br />Sepal.Width: 3.864579","density: 1.201912e-01<br />Sepal.Width: 3.869276","density: 1.168925e-01<br />Sepal.Width: 3.873973","density: 1.135118e-01<br />Sepal.Width: 3.878669","density: 1.100517e-01<br />Sepal.Width: 3.883366","density: 1.065465e-01<br />Sepal.Width: 3.888063","density: 1.030010e-01<br />Sepal.Width: 3.892759","density: 9.941305e-02<br />Sepal.Width: 3.897456","density: 9.580565e-02<br />Sepal.Width: 3.902153","density: 9.218893e-02<br />Sepal.Width: 3.906849","density: 8.856828e-02<br />Sepal.Width: 3.911546","density: 8.496109e-02<br />Sepal.Width: 3.916243","density: 8.137354e-02<br />Sepal.Width: 3.920939","density: 7.780983e-02<br />Sepal.Width: 3.925636","density: 7.428743e-02<br />Sepal.Width: 3.930333","density: 7.081976e-02<br />Sepal.Width: 3.935029","density: 6.740028e-02<br />Sepal.Width: 3.939726","density: 6.403247e-02<br />Sepal.Width: 3.944423","density: 6.075744e-02<br />Sepal.Width: 3.949119","density: 5.755370e-02<br />Sepal.Width: 3.953816","density: 5.441970e-02<br />Sepal.Width: 3.958513","density: 5.137234e-02<br />Sepal.Width: 3.963209","density: 4.843832e-02<br />Sepal.Width: 3.967906","density: 4.558641e-02<br />Sepal.Width: 3.972603","density: 4.281803e-02<br />Sepal.Width: 3.977299","density: 4.017265e-02<br />Sepal.Width: 3.981996","density: 3.763290e-02<br />Sepal.Width: 3.986693","density: 3.518228e-02<br />Sepal.Width: 3.991389","density: 3.282596e-02<br />Sepal.Width: 3.996086","density: 3.061130e-02<br />Sepal.Width: 4.000783","density: 2.848639e-02<br />Sepal.Width: 4.005479","density: 2.645085e-02<br />Sepal.Width: 4.010176","density: 2.453031e-02<br />Sepal.Width: 4.014873","density: 2.272485e-02<br />Sepal.Width: 4.019569","density: 2.100455e-02<br />Sepal.Width: 4.024266","density: 1.936850e-02<br />Sepal.Width: 4.028963","density: 1.785845e-02<br />Sepal.Width: 4.033659","density: 1.643294e-02<br />Sepal.Width: 4.038356","density: 1.508413e-02<br />Sepal.Width: 4.043053","density: 1.382256e-02<br />Sepal.Width: 4.047750","density: 1.266419e-02<br />Sepal.Width: 4.052446","density: 1.157334e-02<br />Sepal.Width: 4.057143","density: 1.054847e-02<br />Sepal.Width: 4.061840","density: 9.612223e-03<br />Sepal.Width: 4.066536","density: 8.746719e-03<br />Sepal.Width: 4.071233","density: 7.937237e-03<br />Sepal.Width: 4.075930","density: 7.184039e-03<br />Sepal.Width: 4.080626","density: 6.510359e-03<br />Sepal.Width: 4.085323","density: 5.882893e-03<br />Sepal.Width: 4.090020","density: 5.300153e-03<br />Sepal.Width: 4.094716","density: 4.771475e-03<br />Sepal.Width: 4.099413","density: 4.294181e-03<br />Sepal.Width: 4.104110","density: 3.852654e-03<br />Sepal.Width: 4.108806","density: 3.445557e-03<br />Sepal.Width: 4.113503","density: 3.086403e-03<br />Sepal.Width: 4.118200","density: 2.758028e-03<br />Sepal.Width: 4.122896","density: 2.456398e-03<br />Sepal.Width: 4.127593","density: 2.183780e-03<br />Sepal.Width: 4.132290","density: 1.944105e-03<br />Sepal.Width: 4.136986","density: 1.724679e-03<br />Sepal.Width: 4.141683","density: 1.524593e-03<br />Sepal.Width: 4.146380","density: 1.349039e-03<br />Sepal.Width: 4.151076","density: 1.192350e-03<br />Sepal.Width: 4.155773","density: 1.049923e-03<br />Sepal.Width: 4.160470","density: 9.212922e-04<br />Sepal.Width: 4.165166","density: 8.114980e-04<br />Sepal.Width: 4.169863","density: 7.119577e-04<br />Sepal.Width: 4.174560","density: 6.221481e-04<br />Sepal.Width: 4.179256","density: 5.435548e-04<br />Sepal.Width: 4.183953","density: 4.752773e-04<br />Sepal.Width: 4.188650","density: 4.138269e-04<br />Sepal.Width: 4.193346","density: 3.588112e-04<br />Sepal.Width: 4.198043","density: 3.122633e-04<br />Sepal.Width: 4.202740","density: 2.709887e-04<br />Sepal.Width: 4.207436","density: 2.341236e-04<br />Sepal.Width: 4.212133","density: 2.018552e-04<br />Sepal.Width: 4.216830","density: 1.746535e-04<br />Sepal.Width: 4.221526","density: 1.503996e-04<br />Sepal.Width: 4.226223","density: 1.289085e-04<br />Sepal.Width: 4.230920","density: 1.107548e-04<br />Sepal.Width: 4.235616","density: 9.509511e-05<br />Sepal.Width: 4.240313","density: 8.124196e-05<br />Sepal.Width: 4.245010","density: 6.907900e-05<br />Sepal.Width: 4.249706","density: 5.916253e-05<br />Sepal.Width: 4.254403","density: 5.039745e-05<br />Sepal.Width: 4.259100","density: 4.270698e-05<br />Sepal.Width: 4.263796","density: 3.620378e-05<br />Sepal.Width: 4.268493","density: 3.076348e-05<br />Sepal.Width: 4.273190","density: 2.599400e-05<br />Sepal.Width: 4.277886","density: 2.184487e-05<br />Sepal.Width: 4.282583","density: 1.847166e-05<br />Sepal.Width: 4.287280","density: 1.556938e-05<br />Sepal.Width: 4.291977","density: 1.304660e-05<br />Sepal.Width: 4.296673","density: 1.090518e-05<br />Sepal.Width: 4.301370","density: 9.173632e-06<br />Sepal.Width: 4.306067","density: 7.668260e-06<br />Sepal.Width: 4.310763","density: 6.371292e-06<br />Sepal.Width: 4.315460","density: 5.314973e-06<br />Sepal.Width: 4.320157","density: 4.434043e-06<br />Sepal.Width: 4.324853","density: 3.674986e-06<br />Sepal.Width: 4.329550","density: 3.026992e-06<br />Sepal.Width: 4.334247","density: 2.521330e-06<br />Sepal.Width: 4.338943","density: 2.085557e-06<br />Sepal.Width: 4.343640","density: 1.713535e-06<br />Sepal.Width: 4.348337","density: 1.409198e-06<br />Sepal.Width: 4.353033","density: 1.164006e-06<br />Sepal.Width: 4.357730","density: 9.544423e-07<br />Sepal.Width: 4.362427","density: 7.772405e-07<br />Sepal.Width: 4.367123","density: 6.386288e-07<br />Sepal.Width: 4.371820","density: 5.228967e-07<br />Sepal.Width: 4.376517","density: 4.249386e-07<br />Sepal.Width: 4.381213","density: 3.442307e-07<br />Sepal.Width: 4.385910","density: 2.816389e-07<br />Sepal.Width: 4.390607","density: 2.285343e-07<br />Sepal.Width: 4.395303","density: 1.840338e-07<br />Sepal.Width: 4.400000","density: 1.840338e-07<br />Sepal.Width: 4.400000","density: 1.840338e-07<br />Sepal.Width: 4.400000","density: 2.285343e-07<br />Sepal.Width: 4.395303","density: 2.816389e-07<br />Sepal.Width: 4.390607","density: 3.442307e-07<br />Sepal.Width: 4.385910","density: 4.249386e-07<br />Sepal.Width: 4.381213","density: 5.228967e-07<br />Sepal.Width: 4.376517","density: 6.386288e-07<br />Sepal.Width: 4.371820","density: 7.772405e-07<br />Sepal.Width: 4.367123","density: 9.544423e-07<br />Sepal.Width: 4.362427","density: 1.164006e-06<br />Sepal.Width: 4.357730","density: 1.409198e-06<br />Sepal.Width: 4.353033","density: 1.713535e-06<br />Sepal.Width: 4.348337","density: 2.085557e-06<br />Sepal.Width: 4.343640","density: 2.521330e-06<br />Sepal.Width: 4.338943","density: 3.026992e-06<br />Sepal.Width: 4.334247","density: 3.674986e-06<br />Sepal.Width: 4.329550","density: 4.434043e-06<br />Sepal.Width: 4.324853","density: 5.314973e-06<br />Sepal.Width: 4.320157","density: 6.371292e-06<br />Sepal.Width: 4.315460","density: 7.668260e-06<br />Sepal.Width: 4.310763","density: 9.173632e-06<br />Sepal.Width: 4.306067","density: 1.090518e-05<br />Sepal.Width: 4.301370","density: 1.304660e-05<br />Sepal.Width: 4.296673","density: 1.556938e-05<br />Sepal.Width: 4.291977","density: 1.847166e-05<br />Sepal.Width: 4.287280","density: 2.184487e-05<br />Sepal.Width: 4.282583","density: 2.599400e-05<br />Sepal.Width: 4.277886","density: 3.076348e-05<br />Sepal.Width: 4.273190","density: 3.620378e-05<br />Sepal.Width: 4.268493","density: 4.270698e-05<br />Sepal.Width: 4.263796","density: 5.039745e-05<br />Sepal.Width: 4.259100","density: 5.916253e-05<br />Sepal.Width: 4.254403","density: 6.907900e-05<br />Sepal.Width: 4.249706","density: 8.124196e-05<br />Sepal.Width: 4.245010","density: 9.509511e-05<br />Sepal.Width: 4.240313","density: 1.107548e-04<br />Sepal.Width: 4.235616","density: 1.289085e-04<br />Sepal.Width: 4.230920","density: 1.503996e-04<br />Sepal.Width: 4.226223","density: 1.746535e-04<br />Sepal.Width: 4.221526","density: 2.018552e-04<br />Sepal.Width: 4.216830","density: 2.341236e-04<br />Sepal.Width: 4.212133","density: 2.709887e-04<br />Sepal.Width: 4.207436","density: 3.122633e-04<br />Sepal.Width: 4.202740","density: 3.588112e-04<br />Sepal.Width: 4.198043","density: 4.138269e-04<br />Sepal.Width: 4.193346","density: 4.752773e-04<br />Sepal.Width: 4.188650","density: 5.435548e-04<br />Sepal.Width: 4.183953","density: 6.221481e-04<br />Sepal.Width: 4.179256","density: 7.119577e-04<br />Sepal.Width: 4.174560","density: 8.114980e-04<br />Sepal.Width: 4.169863","density: 9.212922e-04<br />Sepal.Width: 4.165166","density: 1.049923e-03<br />Sepal.Width: 4.160470","density: 1.192350e-03<br />Sepal.Width: 4.155773","density: 1.349039e-03<br />Sepal.Width: 4.151076","density: 1.524593e-03<br />Sepal.Width: 4.146380","density: 1.724679e-03<br />Sepal.Width: 4.141683","density: 1.944105e-03<br />Sepal.Width: 4.136986","density: 2.183780e-03<br />Sepal.Width: 4.132290","density: 2.456398e-03<br />Sepal.Width: 4.127593","density: 2.758028e-03<br />Sepal.Width: 4.122896","density: 3.086403e-03<br />Sepal.Width: 4.118200","density: 3.445557e-03<br />Sepal.Width: 4.113503","density: 3.852654e-03<br />Sepal.Width: 4.108806","density: 4.294181e-03<br />Sepal.Width: 4.104110","density: 4.771475e-03<br />Sepal.Width: 4.099413","density: 5.300153e-03<br />Sepal.Width: 4.094716","density: 5.882893e-03<br />Sepal.Width: 4.090020","density: 6.510359e-03<br />Sepal.Width: 4.085323","density: 7.184039e-03<br />Sepal.Width: 4.080626","density: 7.937237e-03<br />Sepal.Width: 4.075930","density: 8.746719e-03<br />Sepal.Width: 4.071233","density: 9.612223e-03<br />Sepal.Width: 4.066536","density: 1.054847e-02<br />Sepal.Width: 4.061840","density: 1.157334e-02<br />Sepal.Width: 4.057143","density: 1.266419e-02<br />Sepal.Width: 4.052446","density: 1.382256e-02<br />Sepal.Width: 4.047750","density: 1.508413e-02<br />Sepal.Width: 4.043053","density: 1.643294e-02<br />Sepal.Width: 4.038356","density: 1.785845e-02<br />Sepal.Width: 4.033659","density: 1.936850e-02<br />Sepal.Width: 4.028963","density: 2.100455e-02<br />Sepal.Width: 4.024266","density: 2.272485e-02<br />Sepal.Width: 4.019569","density: 2.453031e-02<br />Sepal.Width: 4.014873","density: 2.645085e-02<br />Sepal.Width: 4.010176","density: 2.848639e-02<br />Sepal.Width: 4.005479","density: 3.061130e-02<br />Sepal.Width: 4.000783","density: 3.282596e-02<br />Sepal.Width: 3.996086","density: 3.518228e-02<br />Sepal.Width: 3.991389","density: 3.763290e-02<br />Sepal.Width: 3.986693","density: 4.017265e-02<br />Sepal.Width: 3.981996","density: 4.281803e-02<br />Sepal.Width: 3.977299","density: 4.558641e-02<br />Sepal.Width: 3.972603","density: 4.843832e-02<br />Sepal.Width: 3.967906","density: 5.137234e-02<br />Sepal.Width: 3.963209","density: 5.441970e-02<br />Sepal.Width: 3.958513","density: 5.755370e-02<br />Sepal.Width: 3.953816","density: 6.075744e-02<br />Sepal.Width: 3.949119","density: 6.403247e-02<br />Sepal.Width: 3.944423","density: 6.740028e-02<br />Sepal.Width: 3.939726","density: 7.081976e-02<br />Sepal.Width: 3.935029","density: 7.428743e-02<br />Sepal.Width: 3.930333","density: 7.780983e-02<br />Sepal.Width: 3.925636","density: 8.137354e-02<br />Sepal.Width: 3.920939","density: 8.496109e-02<br />Sepal.Width: 3.916243","density: 8.856828e-02<br />Sepal.Width: 3.911546","density: 9.218893e-02<br />Sepal.Width: 3.906849","density: 9.580565e-02<br />Sepal.Width: 3.902153","density: 9.941305e-02<br />Sepal.Width: 3.897456","density: 1.030010e-01<br />Sepal.Width: 3.892759","density: 1.065465e-01<br />Sepal.Width: 3.888063","density: 1.100517e-01<br />Sepal.Width: 3.883366","density: 1.135118e-01<br />Sepal.Width: 3.878669","density: 1.168925e-01<br />Sepal.Width: 3.873973","density: 1.201912e-01<br />Sepal.Width: 3.869276","density: 1.234144e-01<br />Sepal.Width: 3.864579","density: 1.265526e-01<br />Sepal.Width: 3.859883","density: 1.295452e-01<br />Sepal.Width: 3.855186","density: 1.324348e-01<br />Sepal.Width: 3.850489","density: 1.352176e-01<br />Sepal.Width: 3.845793","density: 1.378530e-01<br />Sepal.Width: 3.841096","density: 1.403260e-01<br />Sepal.Width: 3.836399","density: 1.426711e-01<br />Sepal.Width: 3.831703","density: 1.448856e-01<br />Sepal.Width: 3.827006","density: 1.468896e-01<br />Sepal.Width: 3.822309","density: 1.487409e-01<br />Sepal.Width: 3.817613","density: 1.504495e-01<br />Sepal.Width: 3.812916","density: 1.519894e-01<br />Sepal.Width: 3.808219","density: 1.533162e-01<br />Sepal.Width: 3.803523","density: 1.544966e-01<br />Sepal.Width: 3.798826","density: 1.555308e-01<br />Sepal.Width: 3.794129","density: 1.563602e-01<br />Sepal.Width: 3.789432","density: 1.570176e-01<br />Sepal.Width: 3.784736","density: 1.575358e-01<br />Sepal.Width: 3.780039","density: 1.579115e-01<br />Sepal.Width: 3.775342","density: 1.580783e-01<br />Sepal.Width: 3.770646","density: 1.581204e-01<br />Sepal.Width: 3.765949","density: 1.580411e-01<br />Sepal.Width: 3.761252","density: 1.578131e-01<br />Sepal.Width: 3.756556","density: 1.574423e-01<br />Sepal.Width: 3.751859","density: 1.569725e-01<br />Sepal.Width: 3.747162","density: 1.564077e-01<br />Sepal.Width: 3.742466","density: 1.557130e-01<br />Sepal.Width: 3.737769","density: 1.549383e-01<br />Sepal.Width: 3.733072","density: 1.540962e-01<br />Sepal.Width: 3.728376","density: 1.531839e-01<br />Sepal.Width: 3.723679","density: 1.522004e-01<br />Sepal.Width: 3.718982","density: 1.511792e-01<br />Sepal.Width: 3.714286","density: 1.501253e-01<br />Sepal.Width: 3.709589","density: 1.490392e-01<br />Sepal.Width: 3.704892","density: 1.479405e-01<br />Sepal.Width: 3.700196","density: 1.468390e-01<br />Sepal.Width: 3.695499","density: 1.457398e-01<br />Sepal.Width: 3.690802","density: 1.446637e-01<br />Sepal.Width: 3.686106","density: 1.436137e-01<br />Sepal.Width: 3.681409","density: 1.425940e-01<br />Sepal.Width: 3.676712","density: 1.416224e-01<br />Sepal.Width: 3.672016","density: 1.407171e-01<br />Sepal.Width: 3.667319","density: 1.398687e-01<br />Sepal.Width: 3.662622","density: 1.390811e-01<br />Sepal.Width: 3.657926","density: 1.383995e-01<br />Sepal.Width: 3.653229","density: 1.378064e-01<br />Sepal.Width: 3.648532","density: 1.372979e-01<br />Sepal.Width: 3.643836","density: 1.368911e-01<br />Sepal.Width: 3.639139","density: 1.366308e-01<br />Sepal.Width: 3.634442","density: 1.364768e-01<br />Sepal.Width: 3.629746","density: 1.364323e-01<br />Sepal.Width: 3.625049","density: 1.365484e-01<br />Sepal.Width: 3.620352","density: 1.368187e-01<br />Sepal.Width: 3.615656","density: 1.372184e-01<br />Sepal.Width: 3.610959","density: 1.377517e-01<br />Sepal.Width: 3.606262","density: 1.385122e-01<br />Sepal.Width: 3.601566","density: 1.394210e-01<br />Sepal.Width: 3.596869","density: 1.404811e-01<br />Sepal.Width: 3.592172","density: 1.417368e-01<br />Sepal.Width: 3.587476","density: 1.432165e-01<br />Sepal.Width: 3.582779","density: 1.448654e-01<br />Sepal.Width: 3.578082","density: 1.466860e-01<br />Sepal.Width: 3.573386","density: 1.487737e-01<br />Sepal.Width: 3.568689","density: 1.510692e-01<br />Sepal.Width: 3.563992","density: 1.535530e-01<br />Sepal.Width: 3.559295","density: 1.562519e-01<br />Sepal.Width: 3.554599","density: 1.592477e-01<br />Sepal.Width: 3.549902","density: 1.624466e-01<br />Sepal.Width: 3.545205","density: 1.658505e-01<br />Sepal.Width: 3.540509","density: 1.695408e-01<br />Sepal.Width: 3.535812","density: 1.734994e-01<br />Sepal.Width: 3.531115","density: 1.776746e-01<br />Sepal.Width: 3.526419","density: 1.820680e-01<br />Sepal.Width: 3.521722","density: 1.868196e-01<br />Sepal.Width: 3.517025","density: 1.917972e-01<br />Sepal.Width: 3.512329","density: 1.970000e-01<br />Sepal.Width: 3.507632","density: 2.024832e-01<br />Sepal.Width: 3.502935","density: 2.082810e-01<br />Sepal.Width: 3.498239","density: 2.143063e-01<br />Sepal.Width: 3.493542","density: 2.205588e-01<br />Sepal.Width: 3.488845","density: 2.271478e-01<br />Sepal.Width: 3.484149","density: 2.339926e-01<br />Sepal.Width: 3.479452","density: 2.410605e-01<br />Sepal.Width: 3.474755","density: 2.483745e-01<br />Sepal.Width: 3.470059","density: 2.560187e-01<br />Sepal.Width: 3.465362","density: 2.638769e-01<br />Sepal.Width: 3.460665","density: 2.719468e-01<br />Sepal.Width: 3.455969","density: 2.802981e-01<br />Sepal.Width: 3.451272","density: 2.889046e-01<br />Sepal.Width: 3.446575","density: 2.977074e-01<br />Sepal.Width: 3.441879","density: 3.067040e-01<br />Sepal.Width: 3.437182","density: 3.159981e-01<br />Sepal.Width: 3.432485","density: 3.254731e-01<br />Sepal.Width: 3.427789","density: 3.351216e-01<br />Sepal.Width: 3.423092","density: 3.449756e-01<br />Sepal.Width: 3.418395","density: 3.550508e-01<br />Sepal.Width: 3.413699","density: 3.652764e-01<br />Sepal.Width: 3.409002","density: 3.756486e-01<br />Sepal.Width: 3.404305","density: 3.862233e-01<br />Sepal.Width: 3.399609","density: 3.969449e-01<br />Sepal.Width: 3.394912","density: 4.077882e-01<br />Sepal.Width: 3.390215","density: 4.187593e-01<br />Sepal.Width: 3.385519","density: 4.298932e-01<br />Sepal.Width: 3.380822","density: 4.411242e-01<br />Sepal.Width: 3.376125","density: 4.524486e-01<br />Sepal.Width: 3.371429","density: 4.638886e-01<br />Sepal.Width: 3.366732","density: 4.754268e-01<br />Sepal.Width: 3.362035","density: 4.870357e-01<br />Sepal.Width: 3.357339","density: 4.987118e-01<br />Sepal.Width: 3.352642","density: 5.104816e-01<br />Sepal.Width: 3.347945","density: 5.223043e-01<br />Sepal.Width: 3.343249","density: 5.341752e-01<br />Sepal.Width: 3.338552","density: 5.460993e-01<br />Sepal.Width: 3.333855","density: 5.580767e-01<br />Sepal.Width: 3.329159","density: 5.700874e-01<br />Sepal.Width: 3.324462","density: 5.821296e-01<br />Sepal.Width: 3.319765","density: 5.942122e-01<br />Sepal.Width: 3.315068","density: 6.063230e-01<br />Sepal.Width: 3.310372","density: 6.184566e-01<br />Sepal.Width: 3.305675","density: 6.306137e-01<br />Sepal.Width: 3.300978","density: 6.428018e-01<br />Sepal.Width: 3.296282","density: 6.550097e-01<br />Sepal.Width: 3.291585","density: 6.672375e-01<br />Sepal.Width: 3.286888","density: 6.794921e-01<br />Sepal.Width: 3.282192","density: 6.917750e-01<br />Sepal.Width: 3.277495","density: 7.040812e-01<br />Sepal.Width: 3.272798","density: 7.164119e-01<br />Sepal.Width: 3.268102","density: 7.287857e-01<br />Sepal.Width: 3.263405","density: 7.411925e-01<br />Sepal.Width: 3.258708","density: 7.536327e-01<br />Sepal.Width: 3.254012","density: 7.661160e-01<br />Sepal.Width: 3.249315","density: 7.786577e-01<br />Sepal.Width: 3.244618","density: 7.912445e-01<br />Sepal.Width: 3.239922","density: 8.038784e-01<br />Sepal.Width: 3.235225","density: 8.165863e-01<br />Sepal.Width: 3.230528","density: 8.293606e-01<br />Sepal.Width: 3.225832","density: 8.421937e-01<br />Sepal.Width: 3.221135","density: 8.550915e-01<br />Sepal.Width: 3.216438","density: 8.680895e-01<br />Sepal.Width: 3.211742","density: 8.811542e-01<br />Sepal.Width: 3.207045","density: 8.942862e-01<br />Sepal.Width: 3.202348","density: 9.075075e-01<br />Sepal.Width: 3.197652","density: 9.208174e-01<br />Sepal.Width: 3.192955","density: 9.341930e-01<br />Sepal.Width: 3.188258","density: 9.476335e-01<br />Sepal.Width: 3.183562","density: 9.611677e-01<br />Sepal.Width: 3.178865","density: 9.747584e-01<br />Sepal.Width: 3.174168","density: 9.883974e-01<br />Sepal.Width: 3.169472","density: 1.002087e+00<br />Sepal.Width: 3.164775","density: 1.015820e+00<br />Sepal.Width: 3.160078","density: 1.029569e+00<br />Sepal.Width: 3.155382","density: 1.043328e+00<br />Sepal.Width: 3.150685","density: 1.057080e+00<br />Sepal.Width: 3.145988","density: 1.070797e+00<br />Sepal.Width: 3.141292","density: 1.084473e+00<br />Sepal.Width: 3.136595","density: 1.098091e+00<br />Sepal.Width: 3.131898","density: 1.111581e+00<br />Sepal.Width: 3.127202","density: 1.124960e+00<br />Sepal.Width: 3.122505","density: 1.138214e+00<br />Sepal.Width: 3.117808","density: 1.151277e+00<br />Sepal.Width: 3.113112","density: 1.164098e+00<br />Sepal.Width: 3.108415","density: 1.176709e+00<br />Sepal.Width: 3.103718","density: 1.189097e+00<br />Sepal.Width: 3.099022","density: 1.201085e+00<br />Sepal.Width: 3.094325","density: 1.212752e+00<br />Sepal.Width: 3.089628","density: 1.224102e+00<br />Sepal.Width: 3.084932","density: 1.235059e+00<br />Sepal.Width: 3.080235","density: 1.245468e+00<br />Sepal.Width: 3.075538","density: 1.255470e+00<br />Sepal.Width: 3.070841","density: 1.265052e+00<br />Sepal.Width: 3.066145","density: 1.274000e+00<br />Sepal.Width: 3.061448","density: 1.282368e+00<br />Sepal.Width: 3.056751","density: 1.290237e+00<br />Sepal.Width: 3.052055","density: 1.297579e+00<br />Sepal.Width: 3.047358","density: 1.304072e+00<br />Sepal.Width: 3.042661","density: 1.310011e+00<br />Sepal.Width: 3.037965","density: 1.315389e+00<br />Sepal.Width: 3.033268","density: 1.320035e+00<br />Sepal.Width: 3.028571","density: 1.323902e+00<br />Sepal.Width: 3.023875","density: 1.327185e+00<br />Sepal.Width: 3.019178","density: 1.329882e+00<br />Sepal.Width: 3.014481","density: 1.331684e+00<br />Sepal.Width: 3.009785","density: 1.332854e+00<br />Sepal.Width: 3.005088","density: 1.333452e+00<br />Sepal.Width: 3.000391","density: 1.333398e+00<br />Sepal.Width: 2.995695","density: 1.332540e+00<br />Sepal.Width: 2.990998","density: 1.331153e+00<br />Sepal.Width: 2.986301","density: 1.329248e+00<br />Sepal.Width: 2.981605","density: 1.326650e+00<br />Sepal.Width: 2.976908","density: 1.323483e+00<br />Sepal.Width: 2.972211","density: 1.319873e+00<br />Sepal.Width: 2.967515","density: 1.315826e+00<br />Sepal.Width: 2.962818","density: 1.311155e+00<br />Sepal.Width: 2.958121","density: 1.306133e+00<br />Sepal.Width: 2.953425","density: 1.300778e+00<br />Sepal.Width: 2.948728","density: 1.295033e+00<br />Sepal.Width: 2.944031","density: 1.288930e+00<br />Sepal.Width: 2.939335","density: 1.282592e+00<br />Sepal.Width: 2.934638","density: 1.276037e+00<br />Sepal.Width: 2.929941","density: 1.269206e+00<br />Sepal.Width: 2.925245","density: 1.262218e+00<br />Sepal.Width: 2.920548","density: 1.255104e+00<br />Sepal.Width: 2.915851","density: 1.247869e+00<br />Sepal.Width: 2.911155","density: 1.240526e+00<br />Sepal.Width: 2.906458","density: 1.233134e+00<br />Sepal.Width: 2.901761","density: 1.225704e+00<br />Sepal.Width: 2.897065","density: 1.218247e+00<br />Sepal.Width: 2.892368","density: 1.210793e+00<br />Sepal.Width: 2.887671","density: 1.203349e+00<br />Sepal.Width: 2.882975","density: 1.195922e+00<br />Sepal.Width: 2.878278","density: 1.188540e+00<br />Sepal.Width: 2.873581","density: 1.181192e+00<br />Sepal.Width: 2.868885","density: 1.173879e+00<br />Sepal.Width: 2.864188","density: 1.166610e+00<br />Sepal.Width: 2.859491","density: 1.159386e+00<br />Sepal.Width: 2.854795","density: 1.152189e+00<br />Sepal.Width: 2.850098","density: 1.145016e+00<br />Sepal.Width: 2.845401","density: 1.137866e+00<br />Sepal.Width: 2.840705","density: 1.130719e+00<br />Sepal.Width: 2.836008","density: 1.123566e+00<br />Sepal.Width: 2.831311","density: 1.116398e+00<br />Sepal.Width: 2.826614","density: 1.109183e+00<br />Sepal.Width: 2.821918","density: 1.101921e+00<br />Sepal.Width: 2.817221","density: 1.094606e+00<br />Sepal.Width: 2.812524","density: 1.087196e+00<br />Sepal.Width: 2.807828","density: 1.079678e+00<br />Sepal.Width: 2.803131","density: 1.072062e+00<br />Sepal.Width: 2.798434","density: 1.064341e+00<br />Sepal.Width: 2.793738","density: 1.056424e+00<br />Sepal.Width: 2.789041","density: 1.048372e+00<br />Sepal.Width: 2.784344","density: 1.040179e+00<br />Sepal.Width: 2.779648","density: 1.031803e+00<br />Sepal.Width: 2.774951","density: 1.023211e+00<br />Sepal.Width: 2.770254","density: 1.014457e+00<br />Sepal.Width: 2.765558","density: 1.005540e+00<br />Sepal.Width: 2.760861","density: 9.963764e-01<br />Sepal.Width: 2.756164","density: 9.870238e-01<br />Sepal.Width: 2.751468","density: 9.775071e-01<br />Sepal.Width: 2.746771","density: 9.678103e-01<br />Sepal.Width: 2.742074","density: 9.578776e-01<br />Sepal.Width: 2.737378","density: 9.477977e-01<br />Sepal.Width: 2.732681","density: 9.375751e-01<br />Sepal.Width: 2.727984","density: 9.271713e-01<br />Sepal.Width: 2.723288","density: 9.166169e-01<br />Sepal.Width: 2.718591","density: 9.059547e-01<br />Sepal.Width: 2.713894","density: 8.951911e-01<br />Sepal.Width: 2.709198","density: 8.842943e-01<br />Sepal.Width: 2.704501","density: 8.733327e-01<br />Sepal.Width: 2.699804","density: 8.623160e-01<br />Sepal.Width: 2.695108","density: 8.512469e-01<br />Sepal.Width: 2.690411","density: 8.401469e-01<br />Sepal.Width: 2.685714","density: 8.290410e-01<br />Sepal.Width: 2.681018","density: 8.179365e-01<br />Sepal.Width: 2.676321","density: 8.068591e-01<br />Sepal.Width: 2.671624","density: 7.958245e-01<br />Sepal.Width: 2.666928","density: 7.848360e-01<br />Sepal.Width: 2.662231","density: 7.739069e-01<br />Sepal.Width: 2.657534","density: 7.630841e-01<br />Sepal.Width: 2.652838","density: 7.523426e-01<br />Sepal.Width: 2.648141","density: 7.416867e-01<br />Sepal.Width: 2.643444","density: 7.311556e-01<br />Sepal.Width: 2.638748","density: 7.207543e-01<br />Sepal.Width: 2.634051","density: 7.104574e-01<br />Sepal.Width: 2.629354","density: 7.002670e-01<br />Sepal.Width: 2.624658","density: 6.902492e-01<br />Sepal.Width: 2.619961","density: 6.803436e-01<br />Sepal.Width: 2.615264","density: 6.705451e-01<br />Sepal.Width: 2.610568","density: 6.608742e-01<br />Sepal.Width: 2.605871","density: 6.513442e-01<br />Sepal.Width: 2.601174","density: 6.419076e-01<br />Sepal.Width: 2.596477","density: 6.325613e-01<br />Sepal.Width: 2.591781","density: 6.233342e-01<br />Sepal.Width: 2.587084","density: 6.141889e-01<br />Sepal.Width: 2.582387","density: 6.051067e-01<br />Sepal.Width: 2.577691","density: 5.960860e-01<br />Sepal.Width: 2.572994","density: 5.871314e-01<br />Sepal.Width: 2.568297","density: 5.782061e-01<br />Sepal.Width: 2.563601","density: 5.693044e-01<br />Sepal.Width: 2.558904","density: 5.604200e-01<br />Sepal.Width: 2.554207","density: 5.515332e-01<br />Sepal.Width: 2.549511","density: 5.426350e-01<br />Sepal.Width: 2.544814","density: 5.337201e-01<br />Sepal.Width: 2.540117","density: 5.247591e-01<br />Sepal.Width: 2.535421","density: 5.157547e-01<br />Sepal.Width: 2.530724","density: 5.067047e-01<br />Sepal.Width: 2.526027","density: 4.975930e-01<br />Sepal.Width: 2.521331","density: 4.883960e-01<br />Sepal.Width: 2.516634","density: 4.791342e-01<br />Sepal.Width: 2.511937","density: 4.698059e-01<br />Sepal.Width: 2.507241","density: 4.603771e-01<br />Sepal.Width: 2.502544","density: 4.508646e-01<br />Sepal.Width: 2.497847","density: 4.412823e-01<br />Sepal.Width: 2.493151","density: 4.316260e-01<br />Sepal.Width: 2.488454","density: 4.218681e-01<br />Sepal.Width: 2.483757","density: 4.120512e-01<br />Sepal.Width: 2.479061","density: 4.021787e-01<br />Sepal.Width: 2.474364","density: 3.922415e-01<br />Sepal.Width: 2.469667","density: 3.822550e-01<br />Sepal.Width: 2.464971","density: 3.722404e-01<br />Sepal.Width: 2.460274","density: 3.622030e-01<br />Sepal.Width: 2.455577","density: 3.521532e-01<br />Sepal.Width: 2.450881","density: 3.421134e-01<br />Sepal.Width: 2.446184","density: 3.320915e-01<br />Sepal.Width: 2.441487","density: 3.221031e-01<br />Sepal.Width: 2.436791","density: 3.121842e-01<br />Sepal.Width: 2.432094","density: 3.023306e-01<br />Sepal.Width: 2.427397","density: 2.925500e-01<br />Sepal.Width: 2.422701","density: 2.828977e-01<br />Sepal.Width: 2.418004","density: 2.733780e-01<br />Sepal.Width: 2.413307","density: 2.639808e-01<br />Sepal.Width: 2.408611","density: 2.547219e-01<br />Sepal.Width: 2.403914","density: 2.457007e-01<br />Sepal.Width: 2.399217","density: 2.368487e-01<br />Sepal.Width: 2.394521","density: 2.281725e-01<br />Sepal.Width: 2.389824","density: 2.197397e-01<br />Sepal.Width: 2.385127","density: 2.115765e-01<br />Sepal.Width: 2.380431","density: 2.036264e-01<br />Sepal.Width: 2.375734","density: 1.958947e-01<br />Sepal.Width: 2.371037","density: 1.885170e-01<br />Sepal.Width: 2.366341","density: 1.813970e-01<br />Sepal.Width: 2.361644","density: 1.745193e-01<br />Sepal.Width: 2.356947","density: 1.679290e-01<br />Sepal.Width: 2.352250","density: 1.617044e-01<br />Sepal.Width: 2.347554","density: 1.557328e-01<br />Sepal.Width: 2.342857","density: 1.500149e-01<br />Sepal.Width: 2.338160","density: 1.446553e-01<br />Sepal.Width: 2.333464","density: 1.395996e-01<br />Sepal.Width: 2.328767","density: 1.347922e-01<br />Sepal.Width: 2.324070","density: 1.302404e-01<br />Sepal.Width: 2.319374","density: 1.260686e-01<br />Sepal.Width: 2.314677","density: 1.221279e-01<br />Sepal.Width: 2.309980","density: 1.184141e-01<br />Sepal.Width: 2.305284","density: 1.149826e-01<br />Sepal.Width: 2.300587","density: 1.118293e-01<br />Sepal.Width: 2.295890","density: 1.088736e-01<br />Sepal.Width: 2.291194","density: 1.061105e-01<br />Sepal.Width: 2.286497","density: 1.036210e-01<br />Sepal.Width: 2.281800","density: 1.013099e-01<br />Sepal.Width: 2.277104","density: 9.915411e-02<br />Sepal.Width: 2.272407","density: 9.716742e-02<br />Sepal.Width: 2.267710","density: 9.537155e-02<br />Sepal.Width: 2.263014","density: 9.369107e-02<br />Sepal.Width: 2.258317","density: 9.211958e-02<br />Sepal.Width: 2.253620","density: 9.068226e-02<br />Sepal.Width: 2.248924","density: 8.934199e-02<br />Sepal.Width: 2.244227","density: 8.807138e-02<br />Sepal.Width: 2.239530","density: 8.686546e-02<br />Sepal.Width: 2.234834","density: 8.573631e-02<br />Sepal.Width: 2.230137","density: 8.464106e-02<br />Sepal.Width: 2.225440","density: 8.357466e-02<br />Sepal.Width: 2.220744","density: 8.253393e-02<br />Sepal.Width: 2.216047","density: 8.150416e-02<br />Sepal.Width: 2.211350","density: 8.047453e-02<br />Sepal.Width: 2.206654","density: 7.944111e-02<br />Sepal.Width: 2.201957","density: 7.838908e-02<br />Sepal.Width: 2.197260","density: 7.731447e-02<br />Sepal.Width: 2.192564","density: 7.621624e-02<br />Sepal.Width: 2.187867","density: 7.508722e-02<br />Sepal.Width: 2.183170","density: 7.391024e-02<br />Sepal.Width: 2.178474","density: 7.269765e-02<br />Sepal.Width: 2.173777","density: 7.144833e-02<br />Sepal.Width: 2.169080","density: 7.014550e-02<br />Sepal.Width: 2.164384","density: 6.879334e-02<br />Sepal.Width: 2.159687","density: 6.740134e-02<br />Sepal.Width: 2.154990","density: 6.596918e-02<br />Sepal.Width: 2.150294","density: 6.447446e-02<br />Sepal.Width: 2.145597","density: 6.294311e-02<br />Sepal.Width: 2.140900","density: 6.137623e-02<br />Sepal.Width: 2.136204","density: 5.976716e-02<br />Sepal.Width: 2.131507","density: 5.811718e-02<br />Sepal.Width: 2.126810","density: 5.644056e-02<br />Sepal.Width: 2.122114","density: 5.473901e-02<br />Sepal.Width: 2.117417","density: 5.300646e-02<br />Sepal.Width: 2.112720","density: 5.125660e-02<br />Sepal.Width: 2.108023","density: 4.949405e-02<br />Sepal.Width: 2.103327","density: 4.772033e-02<br />Sepal.Width: 2.098630","density: 4.594010e-02<br />Sepal.Width: 2.093933","density: 4.416046e-02<br />Sepal.Width: 2.089237","density: 4.238349e-02<br />Sepal.Width: 2.084540","density: 4.061586e-02<br />Sepal.Width: 2.079843","density: 3.886346e-02<br />Sepal.Width: 2.075147","density: 3.712641e-02<br />Sepal.Width: 2.070450","density: 3.540665e-02<br />Sepal.Width: 2.065753","density: 3.372244e-02<br />Sepal.Width: 2.061057","density: 3.206477e-02<br />Sepal.Width: 2.056360","density: 3.043496e-02<br />Sepal.Width: 2.051663","density: 2.884271e-02<br />Sepal.Width: 2.046967","density: 2.729714e-02<br />Sepal.Width: 2.042270","density: 2.578758e-02<br />Sepal.Width: 2.037573","density: 2.431507e-02<br />Sepal.Width: 2.032877","density: 2.290081e-02<br />Sepal.Width: 2.028180","density: 2.153332e-02<br />Sepal.Width: 2.023483","density: 2.020774e-02<br />Sepal.Width: 2.018787","density: 1.892946e-02<br />Sepal.Width: 2.014090","density: 1.771718e-02<br />Sepal.Width: 2.009393","density: 1.654899e-02<br />Sepal.Width: 2.004697","density: 1.542498e-02<br />Sepal.Width: 2.000000","density: 1.542498e-02<br />Sepal.Width: 2.000000"],"key":["101","102","103","104","105","106","107","108","109","110","111","112","113","114","115","116","117","118","119","120","121","122","123","124","125","126","127","128","129","130","131","132","133","134","135","136","137","138","139","140","141","142","143","144","145","146","147","148","149","150"],"type":"scatter","mode":"lines","line":{"width":1.88976377952756,"color":"rgba(0,0,0,1)","dash":"solid"},"fill":"toself","fillcolor":"rgba(97,156,255,1)","hoveron":"points","set":"SharedData91833e3a","name":"virginica","legendgroup":"virginica","showlegend":true,"xaxis":"x2","yaxis":"y7","hoverinfo":"text","_isSimpleKey":true,"_isNestedKey":false,"frame":null},{"x":[3.5,3,3.2,3.1,3.6,3.9,3.4,3.4,2.9,3.1,3.7,3.4,3,3,4,4.4,3.9,3.5,3.8,3.8,3.4,3.7,3.6,3.3,3.4,3,3.4,3.5,3.4,3.2,3.1,3.4,4.1,4.2,3.1,3.2,3.5,3.6,3,3.4,3.5,2.3,3.2,3.5,3.8,3,3.8,3.2,3.7,3.3],"y":[1.4,1.4,1.3,1.5,1.4,1.7,1.4,1.5,1.4,1.5,1.5,1.6,1.4,1.1,1.2,1.5,1.3,1.4,1.7,1.5,1.7,1.5,1,1.7,1.9,1.6,1.6,1.5,1.4,1.6,1.6,1.5,1.5,1.4,1.5,1.2,1.3,1.4,1.3,1.5,1.3,1.3,1.3,1.6,1.9,1.4,1.6,1.4,1.5,1.4],"text":["Sepal.Width: 3.5<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Width: 3.0<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Width: 3.2<br />Petal.Length: 1.3<br />Species: setosa","Sepal.Width: 3.1<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Width: 3.6<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Width: 3.9<br />Petal.Length: 1.7<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Width: 2.9<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Width: 3.1<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Width: 3.7<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Length: 1.6<br />Species: setosa","Sepal.Width: 3.0<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Width: 3.0<br />Petal.Length: 1.1<br />Species: setosa","Sepal.Width: 4.0<br />Petal.Length: 1.2<br />Species: setosa","Sepal.Width: 4.4<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Width: 3.9<br />Petal.Length: 1.3<br />Species: setosa","Sepal.Width: 3.5<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Width: 3.8<br />Petal.Length: 1.7<br />Species: setosa","Sepal.Width: 3.8<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Length: 1.7<br />Species: setosa","Sepal.Width: 3.7<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Width: 3.6<br />Petal.Length: 1.0<br />Species: setosa","Sepal.Width: 3.3<br />Petal.Length: 1.7<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Length: 1.9<br />Species: setosa","Sepal.Width: 3.0<br />Petal.Length: 1.6<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Length: 1.6<br />Species: setosa","Sepal.Width: 3.5<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Width: 3.2<br />Petal.Length: 1.6<br />Species: setosa","Sepal.Width: 3.1<br />Petal.Length: 1.6<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Width: 4.1<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Width: 4.2<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Width: 3.1<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Width: 3.2<br />Petal.Length: 1.2<br />Species: setosa","Sepal.Width: 3.5<br />Petal.Length: 1.3<br />Species: setosa","Sepal.Width: 3.6<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Width: 3.0<br />Petal.Length: 1.3<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Width: 3.5<br />Petal.Length: 1.3<br />Species: setosa","Sepal.Width: 2.3<br />Petal.Length: 1.3<br />Species: setosa","Sepal.Width: 3.2<br />Petal.Length: 1.3<br />Species: setosa","Sepal.Width: 3.5<br />Petal.Length: 1.6<br />Species: setosa","Sepal.Width: 3.8<br />Petal.Length: 1.9<br />Species: setosa","Sepal.Width: 3.0<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Width: 3.8<br />Petal.Length: 1.6<br />Species: setosa","Sepal.Width: 3.2<br />Petal.Length: 1.4<br />Species: setosa","Sepal.Width: 3.7<br />Petal.Length: 1.5<br />Species: setosa","Sepal.Width: 3.3<br />Petal.Length: 1.4<br />Species: setosa"],"key":["1","2","3","4","5","6","7","8","9","10","11","12","13","14","15","16","17","18","19","20","21","22","23","24","25","26","27","28","29","30","31","32","33","34","35","36","37","38","39","40","41","42","43","44","45","46","47","48","49","50"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(248,118,109,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(248,118,109,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"setosa","legendgroup":"setosa","showlegend":true,"xaxis":"x2","yaxis":"y6","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[3.2,3.2,3.1,2.3,2.8,2.8,3.3,2.4,2.9,2.7,2,3,2.2,2.9,2.9,3.1,3,2.7,2.2,2.5,3.2,2.8,2.5,2.8,2.9,3,2.8,3,2.9,2.6,2.4,2.4,2.7,2.7,3,3.4,3.1,2.3,3,2.5,2.6,3,2.6,2.3,2.7,3,2.9,2.9,2.5,2.8],"y":[4.7,4.5,4.9,4,4.6,4.5,4.7,3.3,4.6,3.9,3.5,4.2,4,4.7,3.6,4.4,4.5,4.1,4.5,3.9,4.8,4,4.9,4.7,4.3,4.4,4.8,5,4.5,3.5,3.8,3.7,3.9,5.1,4.5,4.5,4.7,4.4,4.1,4,4.4,4.6,4,3.3,4.2,4.2,4.2,4.3,3,4.1],"text":["Sepal.Width: 3.2<br />Petal.Length: 4.7<br />Species: versicolor","Sepal.Width: 3.2<br />Petal.Length: 4.5<br />Species: versicolor","Sepal.Width: 3.1<br />Petal.Length: 4.9<br />Species: versicolor","Sepal.Width: 2.3<br />Petal.Length: 4.0<br />Species: versicolor","Sepal.Width: 2.8<br />Petal.Length: 4.6<br />Species: versicolor","Sepal.Width: 2.8<br />Petal.Length: 4.5<br />Species: versicolor","Sepal.Width: 3.3<br />Petal.Length: 4.7<br />Species: versicolor","Sepal.Width: 2.4<br />Petal.Length: 3.3<br />Species: versicolor","Sepal.Width: 2.9<br />Petal.Length: 4.6<br />Species: versicolor","Sepal.Width: 2.7<br />Petal.Length: 3.9<br />Species: versicolor","Sepal.Width: 2.0<br />Petal.Length: 3.5<br />Species: versicolor","Sepal.Width: 3.0<br />Petal.Length: 4.2<br />Species: versicolor","Sepal.Width: 2.2<br />Petal.Length: 4.0<br />Species: versicolor","Sepal.Width: 2.9<br />Petal.Length: 4.7<br />Species: versicolor","Sepal.Width: 2.9<br />Petal.Length: 3.6<br />Species: versicolor","Sepal.Width: 3.1<br />Petal.Length: 4.4<br />Species: versicolor","Sepal.Width: 3.0<br />Petal.Length: 4.5<br />Species: versicolor","Sepal.Width: 2.7<br />Petal.Length: 4.1<br />Species: versicolor","Sepal.Width: 2.2<br />Petal.Length: 4.5<br />Species: versicolor","Sepal.Width: 2.5<br />Petal.Length: 3.9<br />Species: versicolor","Sepal.Width: 3.2<br />Petal.Length: 4.8<br />Species: versicolor","Sepal.Width: 2.8<br />Petal.Length: 4.0<br />Species: versicolor","Sepal.Width: 2.5<br />Petal.Length: 4.9<br />Species: versicolor","Sepal.Width: 2.8<br />Petal.Length: 4.7<br />Species: versicolor","Sepal.Width: 2.9<br />Petal.Length: 4.3<br />Species: versicolor","Sepal.Width: 3.0<br />Petal.Length: 4.4<br />Species: versicolor","Sepal.Width: 2.8<br />Petal.Length: 4.8<br />Species: versicolor","Sepal.Width: 3.0<br />Petal.Length: 5.0<br />Species: versicolor","Sepal.Width: 2.9<br />Petal.Length: 4.5<br />Species: versicolor","Sepal.Width: 2.6<br />Petal.Length: 3.5<br />Species: versicolor","Sepal.Width: 2.4<br />Petal.Length: 3.8<br />Species: versicolor","Sepal.Width: 2.4<br />Petal.Length: 3.7<br />Species: versicolor","Sepal.Width: 2.7<br />Petal.Length: 3.9<br />Species: versicolor","Sepal.Width: 2.7<br />Petal.Length: 5.1<br />Species: versicolor","Sepal.Width: 3.0<br />Petal.Length: 4.5<br />Species: versicolor","Sepal.Width: 3.4<br />Petal.Length: 4.5<br />Species: versicolor","Sepal.Width: 3.1<br />Petal.Length: 4.7<br />Species: versicolor","Sepal.Width: 2.3<br />Petal.Length: 4.4<br />Species: versicolor","Sepal.Width: 3.0<br />Petal.Length: 4.1<br />Species: versicolor","Sepal.Width: 2.5<br />Petal.Length: 4.0<br />Species: versicolor","Sepal.Width: 2.6<br />Petal.Length: 4.4<br />Species: versicolor","Sepal.Width: 3.0<br />Petal.Length: 4.6<br />Species: versicolor","Sepal.Width: 2.6<br />Petal.Length: 4.0<br />Species: versicolor","Sepal.Width: 2.3<br />Petal.Length: 3.3<br />Species: versicolor","Sepal.Width: 2.7<br />Petal.Length: 4.2<br />Species: versicolor","Sepal.Width: 3.0<br />Petal.Length: 4.2<br />Species: versicolor","Sepal.Width: 2.9<br />Petal.Length: 4.2<br />Species: versicolor","Sepal.Width: 2.9<br />Petal.Length: 4.3<br />Species: versicolor","Sepal.Width: 2.5<br />Petal.Length: 3.0<br />Species: versicolor","Sepal.Width: 2.8<br />Petal.Length: 4.1<br />Species: versicolor"],"key":["51","52","53","54","55","56","57","58","59","60","61","62","63","64","65","66","67","68","69","70","71","72","73","74","75","76","77","78","79","80","81","82","83","84","85","86","87","88","89","90","91","92","93","94","95","96","97","98","99","100"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(0,186,56,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(0,186,56,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"versicolor","legendgroup":"versicolor","showlegend":true,"xaxis":"x2","yaxis":"y6","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[3.3,2.7,3,2.9,3,3,2.5,2.9,2.5,3.6,3.2,2.7,3,2.5,2.8,3.2,3,3.8,2.6,2.2,3.2,2.8,2.8,2.7,3.3,3.2,2.8,3,2.8,3,2.8,3.8,2.8,2.8,2.6,3,3.4,3.1,3,3.1,3.1,3.1,2.7,3.2,3.3,3,2.5,3,3.4,3],"y":[6,5.1,5.9,5.6,5.8,6.6,4.5,6.3,5.8,6.1,5.1,5.3,5.5,5,5.1,5.3,5.5,6.7,6.9,5,5.7,4.9,6.7,4.9,5.7,6,4.8,4.9,5.6,5.8,6.1,6.4,5.6,5.1,5.6,6.1,5.6,5.5,4.8,5.4,5.6,5.1,5.1,5.9,5.7,5.2,5,5.2,5.4,5.1],"text":["Sepal.Width: 3.3<br />Petal.Length: 6.0<br />Species: virginica","Sepal.Width: 2.7<br />Petal.Length: 5.1<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Length: 5.9<br />Species: virginica","Sepal.Width: 2.9<br />Petal.Length: 5.6<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Length: 5.8<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Length: 6.6<br />Species: virginica","Sepal.Width: 2.5<br />Petal.Length: 4.5<br />Species: virginica","Sepal.Width: 2.9<br />Petal.Length: 6.3<br />Species: virginica","Sepal.Width: 2.5<br />Petal.Length: 5.8<br />Species: virginica","Sepal.Width: 3.6<br />Petal.Length: 6.1<br />Species: virginica","Sepal.Width: 3.2<br />Petal.Length: 5.1<br />Species: virginica","Sepal.Width: 2.7<br />Petal.Length: 5.3<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Length: 5.5<br />Species: virginica","Sepal.Width: 2.5<br />Petal.Length: 5.0<br />Species: virginica","Sepal.Width: 2.8<br />Petal.Length: 5.1<br />Species: virginica","Sepal.Width: 3.2<br />Petal.Length: 5.3<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Length: 5.5<br />Species: virginica","Sepal.Width: 3.8<br />Petal.Length: 6.7<br />Species: virginica","Sepal.Width: 2.6<br />Petal.Length: 6.9<br />Species: virginica","Sepal.Width: 2.2<br />Petal.Length: 5.0<br />Species: virginica","Sepal.Width: 3.2<br />Petal.Length: 5.7<br />Species: virginica","Sepal.Width: 2.8<br />Petal.Length: 4.9<br />Species: virginica","Sepal.Width: 2.8<br />Petal.Length: 6.7<br />Species: virginica","Sepal.Width: 2.7<br />Petal.Length: 4.9<br />Species: virginica","Sepal.Width: 3.3<br />Petal.Length: 5.7<br />Species: virginica","Sepal.Width: 3.2<br />Petal.Length: 6.0<br />Species: virginica","Sepal.Width: 2.8<br />Petal.Length: 4.8<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Length: 4.9<br />Species: virginica","Sepal.Width: 2.8<br />Petal.Length: 5.6<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Length: 5.8<br />Species: virginica","Sepal.Width: 2.8<br />Petal.Length: 6.1<br />Species: virginica","Sepal.Width: 3.8<br />Petal.Length: 6.4<br />Species: virginica","Sepal.Width: 2.8<br />Petal.Length: 5.6<br />Species: virginica","Sepal.Width: 2.8<br />Petal.Length: 5.1<br />Species: virginica","Sepal.Width: 2.6<br />Petal.Length: 5.6<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Length: 6.1<br />Species: virginica","Sepal.Width: 3.4<br />Petal.Length: 5.6<br />Species: virginica","Sepal.Width: 3.1<br />Petal.Length: 5.5<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Length: 4.8<br />Species: virginica","Sepal.Width: 3.1<br />Petal.Length: 5.4<br />Species: virginica","Sepal.Width: 3.1<br />Petal.Length: 5.6<br />Species: virginica","Sepal.Width: 3.1<br />Petal.Length: 5.1<br />Species: virginica","Sepal.Width: 2.7<br />Petal.Length: 5.1<br />Species: virginica","Sepal.Width: 3.2<br />Petal.Length: 5.9<br />Species: virginica","Sepal.Width: 3.3<br />Petal.Length: 5.7<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Length: 5.2<br />Species: virginica","Sepal.Width: 2.5<br />Petal.Length: 5.0<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Length: 5.2<br />Species: virginica","Sepal.Width: 3.4<br />Petal.Length: 5.4<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Length: 5.1<br />Species: virginica"],"key":["101","102","103","104","105","106","107","108","109","110","111","112","113","114","115","116","117","118","119","120","121","122","123","124","125","126","127","128","129","130","131","132","133","134","135","136","137","138","139","140","141","142","143","144","145","146","147","148","149","150"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(97,156,255,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(97,156,255,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"virginica","legendgroup":"virginica","showlegend":true,"xaxis":"x2","yaxis":"y6","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[3.5,3,3.2,3.1,3.6,3.9,3.4,3.4,2.9,3.1,3.7,3.4,3,3,4,4.4,3.9,3.5,3.8,3.8,3.4,3.7,3.6,3.3,3.4,3,3.4,3.5,3.4,3.2,3.1,3.4,4.1,4.2,3.1,3.2,3.5,3.6,3,3.4,3.5,2.3,3.2,3.5,3.8,3,3.8,3.2,3.7,3.3],"y":[0.2,0.2,0.2,0.2,0.2,0.4,0.3,0.2,0.2,0.1,0.2,0.2,0.1,0.1,0.2,0.4,0.4,0.3,0.3,0.3,0.2,0.4,0.2,0.5,0.2,0.2,0.4,0.2,0.2,0.2,0.2,0.4,0.1,0.2,0.2,0.2,0.2,0.1,0.2,0.2,0.3,0.3,0.2,0.6,0.4,0.3,0.2,0.2,0.2,0.2],"text":["Sepal.Width: 3.5<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.0<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.2<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.1<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.6<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.9<br />Petal.Width: 0.4<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Width: 0.3<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 2.9<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.1<br />Petal.Width: 0.1<br />Species: setosa","Sepal.Width: 3.7<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.0<br />Petal.Width: 0.1<br />Species: setosa","Sepal.Width: 3.0<br />Petal.Width: 0.1<br />Species: setosa","Sepal.Width: 4.0<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 4.4<br />Petal.Width: 0.4<br />Species: setosa","Sepal.Width: 3.9<br />Petal.Width: 0.4<br />Species: setosa","Sepal.Width: 3.5<br />Petal.Width: 0.3<br />Species: setosa","Sepal.Width: 3.8<br />Petal.Width: 0.3<br />Species: setosa","Sepal.Width: 3.8<br />Petal.Width: 0.3<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.7<br />Petal.Width: 0.4<br />Species: setosa","Sepal.Width: 3.6<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.3<br />Petal.Width: 0.5<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.0<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Width: 0.4<br />Species: setosa","Sepal.Width: 3.5<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.2<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.1<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Width: 0.4<br />Species: setosa","Sepal.Width: 4.1<br />Petal.Width: 0.1<br />Species: setosa","Sepal.Width: 4.2<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.1<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.2<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.5<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.6<br />Petal.Width: 0.1<br />Species: setosa","Sepal.Width: 3.0<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.4<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.5<br />Petal.Width: 0.3<br />Species: setosa","Sepal.Width: 2.3<br />Petal.Width: 0.3<br />Species: setosa","Sepal.Width: 3.2<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.5<br />Petal.Width: 0.6<br />Species: setosa","Sepal.Width: 3.8<br />Petal.Width: 0.4<br />Species: setosa","Sepal.Width: 3.0<br />Petal.Width: 0.3<br />Species: setosa","Sepal.Width: 3.8<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.2<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.7<br />Petal.Width: 0.2<br />Species: setosa","Sepal.Width: 3.3<br />Petal.Width: 0.2<br />Species: setosa"],"key":["1","2","3","4","5","6","7","8","9","10","11","12","13","14","15","16","17","18","19","20","21","22","23","24","25","26","27","28","29","30","31","32","33","34","35","36","37","38","39","40","41","42","43","44","45","46","47","48","49","50"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(248,118,109,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(248,118,109,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"setosa","legendgroup":"setosa","showlegend":true,"xaxis":"x2","yaxis":"y5","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[3.2,3.2,3.1,2.3,2.8,2.8,3.3,2.4,2.9,2.7,2,3,2.2,2.9,2.9,3.1,3,2.7,2.2,2.5,3.2,2.8,2.5,2.8,2.9,3,2.8,3,2.9,2.6,2.4,2.4,2.7,2.7,3,3.4,3.1,2.3,3,2.5,2.6,3,2.6,2.3,2.7,3,2.9,2.9,2.5,2.8],"y":[1.4,1.5,1.5,1.3,1.5,1.3,1.6,1,1.3,1.4,1,1.5,1,1.4,1.3,1.4,1.5,1,1.5,1.1,1.8,1.3,1.5,1.2,1.3,1.4,1.4,1.7,1.5,1,1.1,1,1.2,1.6,1.5,1.6,1.5,1.3,1.3,1.3,1.2,1.4,1.2,1,1.3,1.2,1.3,1.3,1.1,1.3],"text":["Sepal.Width: 3.2<br />Petal.Width: 1.4<br />Species: versicolor","Sepal.Width: 3.2<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Width: 3.1<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Width: 2.3<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Width: 2.8<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Width: 2.8<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Width: 3.3<br />Petal.Width: 1.6<br />Species: versicolor","Sepal.Width: 2.4<br />Petal.Width: 1.0<br />Species: versicolor","Sepal.Width: 2.9<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Width: 2.7<br />Petal.Width: 1.4<br />Species: versicolor","Sepal.Width: 2.0<br />Petal.Width: 1.0<br />Species: versicolor","Sepal.Width: 3.0<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Width: 2.2<br />Petal.Width: 1.0<br />Species: versicolor","Sepal.Width: 2.9<br />Petal.Width: 1.4<br />Species: versicolor","Sepal.Width: 2.9<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Width: 3.1<br />Petal.Width: 1.4<br />Species: versicolor","Sepal.Width: 3.0<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Width: 2.7<br />Petal.Width: 1.0<br />Species: versicolor","Sepal.Width: 2.2<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Width: 2.5<br />Petal.Width: 1.1<br />Species: versicolor","Sepal.Width: 3.2<br />Petal.Width: 1.8<br />Species: versicolor","Sepal.Width: 2.8<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Width: 2.5<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Width: 2.8<br />Petal.Width: 1.2<br />Species: versicolor","Sepal.Width: 2.9<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Width: 3.0<br />Petal.Width: 1.4<br />Species: versicolor","Sepal.Width: 2.8<br />Petal.Width: 1.4<br />Species: versicolor","Sepal.Width: 3.0<br />Petal.Width: 1.7<br />Species: versicolor","Sepal.Width: 2.9<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Width: 2.6<br />Petal.Width: 1.0<br />Species: versicolor","Sepal.Width: 2.4<br />Petal.Width: 1.1<br />Species: versicolor","Sepal.Width: 2.4<br />Petal.Width: 1.0<br />Species: versicolor","Sepal.Width: 2.7<br />Petal.Width: 1.2<br />Species: versicolor","Sepal.Width: 2.7<br />Petal.Width: 1.6<br />Species: versicolor","Sepal.Width: 3.0<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Width: 3.4<br />Petal.Width: 1.6<br />Species: versicolor","Sepal.Width: 3.1<br />Petal.Width: 1.5<br />Species: versicolor","Sepal.Width: 2.3<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Width: 3.0<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Width: 2.5<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Width: 2.6<br />Petal.Width: 1.2<br />Species: versicolor","Sepal.Width: 3.0<br />Petal.Width: 1.4<br />Species: versicolor","Sepal.Width: 2.6<br />Petal.Width: 1.2<br />Species: versicolor","Sepal.Width: 2.3<br />Petal.Width: 1.0<br />Species: versicolor","Sepal.Width: 2.7<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Width: 3.0<br />Petal.Width: 1.2<br />Species: versicolor","Sepal.Width: 2.9<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Width: 2.9<br />Petal.Width: 1.3<br />Species: versicolor","Sepal.Width: 2.5<br />Petal.Width: 1.1<br />Species: versicolor","Sepal.Width: 2.8<br />Petal.Width: 1.3<br />Species: versicolor"],"key":["51","52","53","54","55","56","57","58","59","60","61","62","63","64","65","66","67","68","69","70","71","72","73","74","75","76","77","78","79","80","81","82","83","84","85","86","87","88","89","90","91","92","93","94","95","96","97","98","99","100"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(0,186,56,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(0,186,56,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"versicolor","legendgroup":"versicolor","showlegend":true,"xaxis":"x2","yaxis":"y5","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[3.3,2.7,3,2.9,3,3,2.5,2.9,2.5,3.6,3.2,2.7,3,2.5,2.8,3.2,3,3.8,2.6,2.2,3.2,2.8,2.8,2.7,3.3,3.2,2.8,3,2.8,3,2.8,3.8,2.8,2.8,2.6,3,3.4,3.1,3,3.1,3.1,3.1,2.7,3.2,3.3,3,2.5,3,3.4,3],"y":[2.5,1.9,2.1,1.8,2.2,2.1,1.7,1.8,1.8,2.5,2,1.9,2.1,2,2.4,2.3,1.8,2.2,2.3,1.5,2.3,2,2,1.8,2.1,1.8,1.8,1.8,2.1,1.6,1.9,2,2.2,1.5,1.4,2.3,2.4,1.8,1.8,2.1,2.4,2.3,1.9,2.3,2.5,2.3,1.9,2,2.3,1.8],"text":["Sepal.Width: 3.3<br />Petal.Width: 2.5<br />Species: virginica","Sepal.Width: 2.7<br />Petal.Width: 1.9<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Width: 2.1<br />Species: virginica","Sepal.Width: 2.9<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Width: 2.2<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Width: 2.1<br />Species: virginica","Sepal.Width: 2.5<br />Petal.Width: 1.7<br />Species: virginica","Sepal.Width: 2.9<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Width: 2.5<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Width: 3.6<br />Petal.Width: 2.5<br />Species: virginica","Sepal.Width: 3.2<br />Petal.Width: 2.0<br />Species: virginica","Sepal.Width: 2.7<br />Petal.Width: 1.9<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Width: 2.1<br />Species: virginica","Sepal.Width: 2.5<br />Petal.Width: 2.0<br />Species: virginica","Sepal.Width: 2.8<br />Petal.Width: 2.4<br />Species: virginica","Sepal.Width: 3.2<br />Petal.Width: 2.3<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Width: 3.8<br />Petal.Width: 2.2<br />Species: virginica","Sepal.Width: 2.6<br />Petal.Width: 2.3<br />Species: virginica","Sepal.Width: 2.2<br />Petal.Width: 1.5<br />Species: virginica","Sepal.Width: 3.2<br />Petal.Width: 2.3<br />Species: virginica","Sepal.Width: 2.8<br />Petal.Width: 2.0<br />Species: virginica","Sepal.Width: 2.8<br />Petal.Width: 2.0<br />Species: virginica","Sepal.Width: 2.7<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Width: 3.3<br />Petal.Width: 2.1<br />Species: virginica","Sepal.Width: 3.2<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Width: 2.8<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Width: 2.8<br />Petal.Width: 2.1<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Width: 1.6<br />Species: virginica","Sepal.Width: 2.8<br />Petal.Width: 1.9<br />Species: virginica","Sepal.Width: 3.8<br />Petal.Width: 2.0<br />Species: virginica","Sepal.Width: 2.8<br />Petal.Width: 2.2<br />Species: virginica","Sepal.Width: 2.8<br />Petal.Width: 1.5<br />Species: virginica","Sepal.Width: 2.6<br />Petal.Width: 1.4<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Width: 2.3<br />Species: virginica","Sepal.Width: 3.4<br />Petal.Width: 2.4<br />Species: virginica","Sepal.Width: 3.1<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Width: 1.8<br />Species: virginica","Sepal.Width: 3.1<br />Petal.Width: 2.1<br />Species: virginica","Sepal.Width: 3.1<br />Petal.Width: 2.4<br />Species: virginica","Sepal.Width: 3.1<br />Petal.Width: 2.3<br />Species: virginica","Sepal.Width: 2.7<br />Petal.Width: 1.9<br />Species: virginica","Sepal.Width: 3.2<br />Petal.Width: 2.3<br />Species: virginica","Sepal.Width: 3.3<br />Petal.Width: 2.5<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Width: 2.3<br />Species: virginica","Sepal.Width: 2.5<br />Petal.Width: 1.9<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Width: 2.0<br />Species: virginica","Sepal.Width: 3.4<br />Petal.Width: 2.3<br />Species: virginica","Sepal.Width: 3.0<br />Petal.Width: 1.8<br />Species: virginica"],"key":["101","102","103","104","105","106","107","108","109","110","111","112","113","114","115","116","117","118","119","120","121","122","123","124","125","126","127","128","129","130","131","132","133","134","135","136","137","138","139","140","141","142","143","144","145","146","147","148","149","150"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(97,156,255,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(97,156,255,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"virginica","legendgroup":"virginica","showlegend":true,"xaxis":"x2","yaxis":"y5","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[3.95],"y":[7.5688],"text":"Corr: 0.872***","hovertext":"x: 3.95<br />y: 7.5688","textfont":{"size":14.6645669291339,"color":"rgba(118,118,118,1)"},"type":"scatter","mode":"text","hoveron":"points","showlegend":false,"xaxis":"x3","yaxis":"y12","hoverinfo":"text","frame":null},{"x":[3.95],"y":[6.712],"text":" setosa: 0.267. ","hovertext":"xPos: 3.95<br />yPos: 6.7120<br />labelp: setosa: 0.267. ","textfont":{"size":14.6645669291339,"color":"rgba(248,118,109,1)"},"type":"scatter","mode":"text","hoveron":"points","name":" setosa: 0.267. ","legendgroup":" setosa: 0.267. ","showlegend":true,"xaxis":"x3","yaxis":"y12","hoverinfo":"text","frame":null},{"x":[3.95],"y":[5.8552],"text":"versicolor: 0.754***","hovertext":"xPos: 3.95<br />yPos: 5.8552<br />labelp: versicolor: 0.754***","textfont":{"size":14.6645669291339,"color":"rgba(0,186,56,1)"},"type":"scatter","mode":"text","hoveron":"points","name":"versicolor: 0.754***","legendgroup":"versicolor: 0.754***","showlegend":true,"xaxis":"x3","yaxis":"y12","hoverinfo":"text","frame":null},{"x":[3.95],"y":[4.9984],"text":" virginica: 0.864***","hovertext":"xPos: 3.95<br />yPos: 4.9984<br />labelp: virginica: 0.864***","textfont":{"size":14.6645669291339,"color":"rgba(97,156,255,1)"},"type":"scatter","mode":"text","hoveron":"points","name":" virginica: 0.864***","legendgroup":" virginica: 0.864***","showlegend":true,"xaxis":"x3","yaxis":"y12","hoverinfo":"text","frame":null},{"x":[3.95],"y":[4.1792],"text":"Corr: -0.428***","hovertext":"x: 3.95<br />y: 4.1792","textfont":{"size":14.6645669291339,"color":"rgba(118,118,118,1)"},"type":"scatter","mode":"text","hoveron":"points","showlegend":false,"xaxis":"x3","yaxis":"y11","hoverinfo":"text","frame":null},{"x":[3.95],"y":[3.608],"text":" setosa: 0.178 ","hovertext":"xPos: 3.95<br />yPos: 3.6080<br />labelp: setosa: 0.178 ","textfont":{"size":14.6645669291339,"color":"rgba(248,118,109,1)"},"type":"scatter","mode":"text","hoveron":"points","name":" setosa: 0.178 ","legendgroup":" setosa: 0.178 ","showlegend":true,"xaxis":"x3","yaxis":"y11","hoverinfo":"text","frame":null},{"x":[3.95],"y":[3.0368],"text":"versicolor: 0.561***","hovertext":"xPos: 3.95<br />yPos: 3.0368<br />labelp: versicolor: 0.561***","textfont":{"size":14.6645669291339,"color":"rgba(0,186,56,1)"},"type":"scatter","mode":"text","hoveron":"points","name":"versicolor: 0.561***","legendgroup":"versicolor: 0.561***","showlegend":true,"xaxis":"x3","yaxis":"y11","hoverinfo":"text","frame":null},{"x":[3.95],"y":[2.4656],"text":" virginica: 0.401** ","hovertext":"xPos: 3.95<br />yPos: 2.4656<br />labelp: virginica: 0.401** ","textfont":{"size":14.6645669291339,"color":"rgba(97,156,255,1)"},"type":"scatter","mode":"text","hoveron":"points","name":" virginica: 0.401** ","legendgroup":" virginica: 0.401** ","showlegend":true,"xaxis":"x3","yaxis":"y11","hoverinfo":"text","frame":null},{"x":[1,1.01154598825832,1.02309197651663,1.03463796477495,1.04618395303327,1.05772994129159,1.0692759295499,1.08082191780822,1.09236790606654,1.10391389432485,1.11545988258317,1.12700587084149,1.1385518590998,1.15009784735812,1.16164383561644,1.17318982387476,1.18473581213307,1.19628180039139,1.20782778864971,1.21937377690802,1.23091976516634,1.24246575342466,1.25401174168297,1.26555772994129,1.27710371819961,1.28864970645793,1.30019569471624,1.31174168297456,1.32328767123288,1.33483365949119,1.34637964774951,1.35792563600783,1.36947162426614,1.38101761252446,1.39256360078278,1.4041095890411,1.41565557729941,1.42720156555773,1.43874755381605,1.45029354207436,1.46183953033268,1.473385518591,1.48493150684931,1.49647749510763,1.50802348336595,1.51956947162427,1.53111545988258,1.5426614481409,1.55420743639922,1.56575342465753,1.57729941291585,1.58884540117417,1.60039138943249,1.6119373776908,1.62348336594912,1.63502935420744,1.64657534246575,1.65812133072407,1.66966731898239,1.6812133072407,1.69275929549902,1.70430528375734,1.71585127201566,1.72739726027397,1.73894324853229,1.75048923679061,1.76203522504892,1.77358121330724,1.78512720156556,1.79667318982387,1.80821917808219,1.81976516634051,1.83131115459883,1.84285714285714,1.85440313111546,1.86594911937378,1.87749510763209,1.88904109589041,1.90058708414873,1.91213307240705,1.92367906066536,1.93522504892368,1.946771037182,1.95831702544031,1.96986301369863,1.98140900195695,1.99295499021526,2.00450097847358,2.0160469667319,2.02759295499022,2.03913894324853,2.05068493150685,2.06223091976517,2.07377690802348,2.0853228962818,2.09686888454012,2.10841487279843,2.11996086105675,2.13150684931507,2.14305283757339,2.1545988258317,2.16614481409002,2.17769080234834,2.18923679060665,2.20078277886497,2.21232876712329,2.2238747553816,2.23542074363992,2.24696673189824,2.25851272015656,2.27005870841487,2.28160469667319,2.29315068493151,2.30469667318982,2.31624266144814,2.32778864970646,2.33933463796478,2.35088062622309,2.36242661448141,2.37397260273973,2.38551859099804,2.39706457925636,2.40861056751468,2.42015655577299,2.43170254403131,2.44324853228963,2.45479452054795,2.46634050880626,2.47788649706458,2.4894324853229,2.50097847358121,2.51252446183953,2.52407045009785,2.53561643835616,2.54716242661448,2.5587084148728,2.57025440313112,2.58180039138943,2.59334637964775,2.60489236790607,2.61643835616438,2.6279843444227,2.63953033268102,2.65107632093933,2.66262230919765,2.67416829745597,2.68571428571429,2.6972602739726,2.70880626223092,2.72035225048924,2.73189823874755,2.74344422700587,2.75499021526419,2.7665362035225,2.77808219178082,2.78962818003914,2.80117416829746,2.81272015655577,2.82426614481409,2.83581213307241,2.84735812133072,2.85890410958904,2.87045009784736,2.88199608610568,2.89354207436399,2.90508806262231,2.91663405088063,2.92818003913894,2.93972602739726,2.95127201565558,2.96281800391389,2.97436399217221,2.98590998043053,2.99745596868885,3.00900195694716,3.02054794520548,3.0320939334638,3.04363992172211,3.05518590998043,3.06673189823875,3.07827788649706,3.08982387475538,3.1013698630137,3.11291585127202,3.12446183953033,3.13600782778865,3.14755381604697,3.15909980430528,3.1706457925636,3.18219178082192,3.19373776908024,3.20528375733855,3.21682974559687,3.22837573385519,3.2399217221135,3.25146771037182,3.26301369863014,3.27455968688845,3.28610567514677,3.29765166340509,3.30919765166341,3.32074363992172,3.33228962818004,3.34383561643836,3.35538160469667,3.36692759295499,3.37847358121331,3.39001956947162,3.40156555772994,3.41311154598826,3.42465753424658,3.43620352250489,3.44774951076321,3.45929549902153,3.47084148727984,3.48238747553816,3.49393346379648,3.50547945205479,3.51702544031311,3.52857142857143,3.54011741682975,3.55166340508806,3.56320939334638,3.5747553816047,3.58630136986301,3.59784735812133,3.60939334637965,3.62093933463796,3.63248532289628,3.6440313111546,3.65557729941292,3.66712328767123,3.67866927592955,3.69021526418787,3.70176125244618,3.7133072407045,3.72485322896282,3.73639921722114,3.74794520547945,3.75949119373777,3.77103718199609,3.7825831702544,3.79412915851272,3.80567514677104,3.81722113502935,3.82876712328767,3.84031311154599,3.85185909980431,3.86340508806262,3.87495107632094,3.88649706457926,3.89804305283757,3.90958904109589,3.92113502935421,3.93268101761252,3.94422700587084,3.95577299412916,3.96731898238748,3.97886497064579,3.99041095890411,4.00195694716243,4.01350293542074,4.02504892367906,4.03659491193738,4.04814090019569,4.05968688845401,4.07123287671233,4.08277886497065,4.09432485322896,4.10587084148728,4.1174168297456,4.12896281800391,4.14050880626223,4.15205479452055,4.16360078277886,4.17514677103718,4.1866927592955,4.19823874755382,4.20978473581213,4.22133072407045,4.23287671232877,4.24442270058708,4.2559686888454,4.26751467710372,4.27906066536204,4.29060665362035,4.30215264187867,4.31369863013699,4.3252446183953,4.33679060665362,4.34833659491194,4.35988258317025,4.37142857142857,4.38297455968689,4.39452054794521,4.40606653620352,4.41761252446184,4.42915851272016,4.44070450097847,4.45225048923679,4.46379647749511,4.47534246575342,4.48688845401174,4.49843444227006,4.50998043052838,4.52152641878669,4.53307240704501,4.54461839530333,4.55616438356164,4.56771037181996,4.57925636007828,4.5908023483366,4.60234833659491,4.61389432485323,4.62544031311155,4.63698630136986,4.64853228962818,4.6600782778865,4.67162426614481,4.68317025440313,4.69471624266145,4.70626223091977,4.71780821917808,4.7293542074364,4.74090019569472,4.75244618395303,4.76399217221135,4.77553816046967,4.78708414872798,4.7986301369863,4.81017612524462,4.82172211350294,4.83326810176125,4.84481409001957,4.85636007827789,4.8679060665362,4.87945205479452,4.89099804305284,4.90254403131116,4.91409001956947,4.92563600782779,4.93718199608611,4.94872798434442,4.96027397260274,4.97181996086106,4.98336594911937,4.99491193737769,5.00645792563601,5.01800391389432,5.02954990215264,5.04109589041096,5.05264187866928,5.06418786692759,5.07573385518591,5.08727984344423,5.09882583170254,5.11037181996086,5.12191780821918,5.1334637964775,5.14500978473581,5.15655577299413,5.16810176125245,5.17964774951076,5.19119373776908,5.2027397260274,5.21428571428571,5.22583170254403,5.23737769080235,5.24892367906067,5.26046966731898,5.2720156555773,5.28356164383562,5.29510763209393,5.30665362035225,5.31819960861057,5.32974559686888,5.3412915851272,5.35283757338552,5.36438356164384,5.37592954990215,5.38747553816047,5.39902152641879,5.4105675146771,5.42211350293542,5.43365949119374,5.44520547945205,5.45675146771037,5.46829745596869,5.47984344422701,5.49138943248532,5.50293542074364,5.51448140900196,5.52602739726027,5.53757338551859,5.54911937377691,5.56066536203523,5.57221135029354,5.58375733855186,5.59530332681018,5.60684931506849,5.61839530332681,5.62994129158513,5.64148727984344,5.65303326810176,5.66457925636008,5.6761252446184,5.68767123287671,5.69921722113503,5.71076320939335,5.72230919765166,5.73385518590998,5.7454011741683,5.75694716242661,5.76849315068493,5.78003913894325,5.79158512720157,5.80313111545988,5.8146771037182,5.82622309197652,5.83776908023483,5.84931506849315,5.86086105675147,5.87240704500979,5.8839530332681,5.89549902152642,5.90704500978474,5.91859099804305,5.93013698630137,5.94168297455969,5.953228962818,5.96477495107632,5.97632093933464,5.98786692759295,5.99941291585127,6.01095890410959,6.02250489236791,6.03405088062622,6.04559686888454,6.05714285714286,6.06868884540117,6.08023483365949,6.09178082191781,6.10332681017613,6.11487279843444,6.12641878669276,6.13796477495108,6.14951076320939,6.16105675146771,6.17260273972603,6.18414872798434,6.19569471624266,6.20724070450098,6.2187866927593,6.23033268101761,6.24187866927593,6.25342465753425,6.26497064579256,6.27651663405088,6.2880626223092,6.29960861056751,6.31115459882583,6.32270058708415,6.33424657534247,6.34579256360078,6.3573385518591,6.36888454011742,6.38043052837573,6.39197651663405,6.40352250489237,6.41506849315069,6.426614481409,6.43816046966732,6.44970645792564,6.46125244618395,6.47279843444227,6.48434442270059,6.4958904109589,6.50743639921722,6.51898238747554,6.53052837573386,6.54207436399217,6.55362035225049,6.56516634050881,6.57671232876712,6.58825831702544,6.59980430528376,6.61135029354207,6.62289628180039,6.63444227005871,6.64598825831703,6.65753424657534,6.66908023483366,6.68062622309198,6.69217221135029,6.70371819960861,6.71526418786693,6.72681017612524,6.73835616438356,6.74990215264188,6.7614481409002,6.77299412915851,6.78454011741683,6.79608610567515,6.80763209393346,6.81917808219178,6.8307240704501,6.84227005870842,6.85381604696673,6.86536203522505,6.87690802348337,6.88845401174168,6.9,6.9,6.88845401174168,6.87690802348337,6.86536203522505,6.85381604696673,6.84227005870842,6.8307240704501,6.81917808219178,6.80763209393346,6.79608610567515,6.78454011741683,6.77299412915851,6.7614481409002,6.74990215264188,6.73835616438356,6.72681017612524,6.71526418786693,6.70371819960861,6.69217221135029,6.68062622309198,6.66908023483366,6.65753424657534,6.64598825831703,6.63444227005871,6.62289628180039,6.61135029354207,6.59980430528376,6.58825831702544,6.57671232876712,6.56516634050881,6.55362035225049,6.54207436399217,6.53052837573386,6.51898238747554,6.50743639921722,6.4958904109589,6.48434442270059,6.47279843444227,6.46125244618395,6.44970645792564,6.43816046966732,6.426614481409,6.41506849315069,6.40352250489237,6.39197651663405,6.38043052837573,6.36888454011742,6.3573385518591,6.34579256360078,6.33424657534247,6.32270058708415,6.31115459882583,6.29960861056751,6.2880626223092,6.27651663405088,6.26497064579256,6.25342465753425,6.24187866927593,6.23033268101761,6.2187866927593,6.20724070450098,6.19569471624266,6.18414872798434,6.17260273972603,6.16105675146771,6.14951076320939,6.13796477495108,6.12641878669276,6.11487279843444,6.10332681017613,6.09178082191781,6.08023483365949,6.06868884540117,6.05714285714286,6.04559686888454,6.03405088062622,6.02250489236791,6.01095890410959,5.99941291585127,5.98786692759295,5.97632093933464,5.96477495107632,5.953228962818,5.94168297455969,5.93013698630137,5.91859099804305,5.90704500978474,5.89549902152642,5.8839530332681,5.87240704500979,5.86086105675147,5.84931506849315,5.83776908023483,5.82622309197652,5.8146771037182,5.80313111545988,5.79158512720157,5.78003913894325,5.76849315068493,5.75694716242661,5.7454011741683,5.73385518590998,5.72230919765166,5.71076320939335,5.69921722113503,5.68767123287671,5.6761252446184,5.66457925636008,5.65303326810176,5.64148727984344,5.62994129158513,5.61839530332681,5.60684931506849,5.59530332681018,5.58375733855186,5.57221135029354,5.56066536203523,5.54911937377691,5.53757338551859,5.52602739726027,5.51448140900196,5.50293542074364,5.49138943248532,5.47984344422701,5.46829745596869,5.45675146771037,5.44520547945205,5.43365949119374,5.42211350293542,5.4105675146771,5.39902152641879,5.38747553816047,5.37592954990215,5.36438356164384,5.35283757338552,5.3412915851272,5.32974559686888,5.31819960861057,5.30665362035225,5.29510763209393,5.28356164383562,5.2720156555773,5.26046966731898,5.24892367906067,5.23737769080235,5.22583170254403,5.21428571428571,5.2027397260274,5.19119373776908,5.17964774951076,5.16810176125245,5.15655577299413,5.14500978473581,5.1334637964775,5.12191780821918,5.11037181996086,5.09882583170254,5.08727984344423,5.07573385518591,5.06418786692759,5.05264187866928,5.04109589041096,5.02954990215264,5.01800391389432,5.00645792563601,4.99491193737769,4.98336594911937,4.97181996086106,4.96027397260274,4.94872798434442,4.93718199608611,4.92563600782779,4.91409001956947,4.90254403131116,4.89099804305284,4.87945205479452,4.8679060665362,4.85636007827789,4.84481409001957,4.83326810176125,4.82172211350294,4.81017612524462,4.7986301369863,4.78708414872798,4.77553816046967,4.76399217221135,4.75244618395303,4.74090019569472,4.7293542074364,4.71780821917808,4.70626223091977,4.69471624266145,4.68317025440313,4.67162426614481,4.6600782778865,4.64853228962818,4.63698630136986,4.62544031311155,4.61389432485323,4.60234833659491,4.5908023483366,4.57925636007828,4.56771037181996,4.55616438356164,4.54461839530333,4.53307240704501,4.52152641878669,4.50998043052838,4.49843444227006,4.48688845401174,4.47534246575342,4.46379647749511,4.45225048923679,4.44070450097847,4.42915851272016,4.41761252446184,4.40606653620352,4.39452054794521,4.38297455968689,4.37142857142857,4.35988258317025,4.34833659491194,4.33679060665362,4.3252446183953,4.31369863013699,4.30215264187867,4.29060665362035,4.27906066536204,4.26751467710372,4.2559686888454,4.24442270058708,4.23287671232877,4.22133072407045,4.20978473581213,4.19823874755382,4.1866927592955,4.17514677103718,4.16360078277886,4.15205479452055,4.14050880626223,4.12896281800391,4.1174168297456,4.10587084148728,4.09432485322896,4.08277886497065,4.07123287671233,4.05968688845401,4.04814090019569,4.03659491193738,4.02504892367906,4.01350293542074,4.00195694716243,3.99041095890411,3.97886497064579,3.96731898238748,3.95577299412916,3.94422700587084,3.93268101761252,3.92113502935421,3.90958904109589,3.89804305283757,3.88649706457926,3.87495107632094,3.86340508806262,3.85185909980431,3.84031311154599,3.82876712328767,3.81722113502935,3.80567514677104,3.79412915851272,3.7825831702544,3.77103718199609,3.75949119373777,3.74794520547945,3.73639921722114,3.72485322896282,3.7133072407045,3.70176125244618,3.69021526418787,3.67866927592955,3.66712328767123,3.65557729941292,3.6440313111546,3.63248532289628,3.62093933463796,3.60939334637965,3.59784735812133,3.58630136986301,3.5747553816047,3.56320939334638,3.55166340508806,3.54011741682975,3.52857142857143,3.51702544031311,3.50547945205479,3.49393346379648,3.48238747553816,3.47084148727984,3.45929549902153,3.44774951076321,3.43620352250489,3.42465753424658,3.41311154598826,3.40156555772994,3.39001956947162,3.37847358121331,3.36692759295499,3.35538160469667,3.34383561643836,3.33228962818004,3.32074363992172,3.30919765166341,3.29765166340509,3.28610567514677,3.27455968688845,3.26301369863014,3.25146771037182,3.2399217221135,3.22837573385519,3.21682974559687,3.20528375733855,3.19373776908024,3.18219178082192,3.1706457925636,3.15909980430528,3.14755381604697,3.13600782778865,3.12446183953033,3.11291585127202,3.1013698630137,3.08982387475538,3.07827788649706,3.06673189823875,3.05518590998043,3.04363992172211,3.0320939334638,3.02054794520548,3.00900195694716,2.99745596868885,2.98590998043053,2.97436399217221,2.96281800391389,2.95127201565558,2.93972602739726,2.92818003913894,2.91663405088063,2.90508806262231,2.89354207436399,2.88199608610568,2.87045009784736,2.85890410958904,2.84735812133072,2.83581213307241,2.82426614481409,2.81272015655577,2.80117416829746,2.78962818003914,2.77808219178082,2.7665362035225,2.75499021526419,2.74344422700587,2.73189823874755,2.72035225048924,2.70880626223092,2.6972602739726,2.68571428571429,2.67416829745597,2.66262230919765,2.65107632093933,2.63953033268102,2.6279843444227,2.61643835616438,2.60489236790607,2.59334637964775,2.58180039138943,2.57025440313112,2.5587084148728,2.54716242661448,2.53561643835616,2.52407045009785,2.51252446183953,2.50097847358121,2.4894324853229,2.47788649706458,2.46634050880626,2.45479452054795,2.44324853228963,2.43170254403131,2.42015655577299,2.40861056751468,2.39706457925636,2.38551859099804,2.37397260273973,2.36242661448141,2.35088062622309,2.33933463796478,2.32778864970646,2.31624266144814,2.30469667318982,2.29315068493151,2.28160469667319,2.27005870841487,2.25851272015656,2.24696673189824,2.23542074363992,2.2238747553816,2.21232876712329,2.20078277886497,2.18923679060665,2.17769080234834,2.16614481409002,2.1545988258317,2.14305283757339,2.13150684931507,2.11996086105675,2.10841487279843,2.09686888454012,2.0853228962818,2.07377690802348,2.06223091976517,2.05068493150685,2.03913894324853,2.02759295499022,2.0160469667319,2.00450097847358,1.99295499021526,1.98140900195695,1.96986301369863,1.95831702544031,1.946771037182,1.93522504892368,1.92367906066536,1.91213307240705,1.90058708414873,1.88904109589041,1.87749510763209,1.86594911937378,1.85440313111546,1.84285714285714,1.83131115459883,1.81976516634051,1.80821917808219,1.79667318982387,1.78512720156556,1.77358121330724,1.76203522504892,1.75048923679061,1.73894324853229,1.72739726027397,1.71585127201566,1.70430528375734,1.69275929549902,1.6812133072407,1.66966731898239,1.65812133072407,1.64657534246575,1.63502935420744,1.62348336594912,1.6119373776908,1.60039138943249,1.58884540117417,1.57729941291585,1.56575342465753,1.55420743639922,1.5426614481409,1.53111545988258,1.51956947162427,1.50802348336595,1.49647749510763,1.48493150684931,1.473385518591,1.46183953033268,1.45029354207436,1.43874755381605,1.42720156555773,1.41565557729941,1.4041095890411,1.39256360078278,1.38101761252446,1.36947162426614,1.35792563600783,1.34637964774951,1.33483365949119,1.32328767123288,1.31174168297456,1.30019569471624,1.28864970645793,1.27710371819961,1.26555772994129,1.25401174168297,1.24246575342466,1.23091976516634,1.21937377690802,1.20782778864971,1.19628180039139,1.18473581213307,1.17318982387476,1.16164383561644,1.15009784735812,1.1385518590998,1.12700587084149,1.11545988258317,1.10391389432485,1.09236790606654,1.08082191780822,1.0692759295499,1.05772994129159,1.04618395303327,1.03463796477495,1.02309197651663,1.01154598825832,1,1],"y":[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,3.69223522167699e-17,7.61769816443942e-18,3.41598331576525e-17,8.23235857872113e-17,1.03333690711882e-16,8.05321742422807e-17,4.19520263126407e-18,4.47506267889994e-17,1.79202359802484e-17,2.64000221266581e-18,2.30211300878288e-17,2.23978043660373e-17,5.0936707582377e-17,4.28999396824839e-17,9.04087430135309e-18,2.52961888359974e-17,3.11294227894161e-17,2.47669790993752e-17,3.6456698662026e-18,2.8110837761008e-17,3.98047223505298e-17,6.87599887741625e-17,8.61895627000471e-17,1.8262747926148e-17,0,0,0,0,0,3.26261990106643e-17,8.65777783981438e-17,6.83127112136979e-17,0,0,0,0,0,0,0,2.7224683666812e-17,1.38952966867543e-17,0,1.81752216023005e-17,2.03929259487705e-17,1.55767963639152e-17,2.97006715261186e-17,7.55992978098532e-18,2.73089894581855e-17,3.78674293944164e-18,3.90872196361779e-17,2.25747031956127e-17,1.33627657902942e-17,8.695932867253e-18,5.76463073856769e-17,1.60629677267823e-16,1.1638179780076e-16,2.91694921512355e-17,6.49964180241908e-18,4.28076866951494e-17,1.16153954305863e-16,1.3531687547756e-16,6.61223244645343e-17,1.8049873171233e-16,7.70593751013183e-17,1.49503861212175e-16,5.23563160885923e-17,1.78838423543306e-16,1.25079959331183e-16,1.32383053228021e-16,4.67536381118143e-17,4.31477474538072e-17,7.79007847264508e-17,7.3188309467764e-17,9.19004564992152e-17,1.19544606307682e-16,7.11037198543e-17,5.6660784538152e-17,1.12299446375594e-16,1.91843104810714e-18,3.2882946836993e-17,2.97484551622997e-17,9.28119080251971e-17,3.21624072377662e-17,4.01463244336301e-17,3.91628891940879e-17,1.02049229867492e-16,9.50706333791682e-17,5.44850887408515e-17,5.49619395967193e-17,4.70669923900061e-17,7.79987339488386e-17,4.55680523026378e-17,1.62310435874796e-17,4.46348860600934e-17,7.42123363560418e-17,9.97490887478725e-18,6.02075487020516e-17,2.4533948247967e-17,7.17291536889067e-17,5.02329325323909e-17,5.00323331034479e-17,3.61333941900623e-17,0,1.60619279074157e-17,5.2528379853121e-17,5.55860765261468e-17,5.46676050649022e-18,3.2316850038789e-17,3.6889310652252e-18,2.56079484371814e-17,3.10294533293848e-18,0,2.25990591633521e-17,6.42233613107825e-17,3.11060348417642e-17,1.3839178802536e-17,3.63101901285393e-17,5.11087425640566e-17,3.6412858197052e-17,0,7.29058249187294e-18,3.07547505622429e-17,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,2.56796985291065e-17,1.13715999386738e-17,1.67728325060905e-17,3.89211026262503e-17,7.13255931579235e-17,5.82369640486425e-17,0,0,8.19008033176868e-18,4.16333634234434e-17,4.24779270421023e-17,5.5412983270618e-17,5.55111512312578e-17,5.55111512312578e-17,3.36083010127005e-17,7.73818855566697e-18,1.31456448284893e-16,2.28643825716731e-16,2.22933711292379e-16,1.83898781705365e-16,2.078208947399e-16,2.80275946190158e-16,2.70460100839796e-16,2.14335727126301e-16,1.75628299393021e-16,1.49758186935988e-16,1.23888074478958e-16,1.82890453128876e-16,1.55645817811587e-16,1.13113411540795e-16,9.38213658903513e-17,1.47423665199428e-16,7.82306631351892e-17,1.3661692998088e-16,2.06872269229172e-16,1.17961196366423e-16,9.54710464175157e-17,4.79261366099066e-17,5.94724914763785e-17,1.95481282502499e-17,1.44125812705374e-16,7.94505315627949e-17,1.10181753330502e-16,2.07075237811235e-17,2.21532383601573e-17,9.34036533745814e-18,0,0,0,0,1.54981575780373e-17,3.26685304300771e-17,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1.80297414540804e-17,1.5316427166923e-17,0,0,1.9620600994149e-17,4.89243011712492e-17,8.83810855592133e-19,8.7394138502992e-18,4.16296447879969e-17,8.91312782026189e-17,4.47766199351888e-17,1.45666524474002e-17,2.24830894345125e-18,4.584420810371e-17,5.02050701833412e-17,5.56156852108815e-17,3.43168340608868e-17,6.2956531096671e-17,1.06172927861431e-16,2.01725491333326e-17,0,0,0,0,0,0,1.51045969619955e-17,7.13779907845653e-18,1.02979072799057e-18,0,0,0,0,0,1.11886649118781e-18,3.34360336588493e-18,0,0,0,1.11699852686923e-17,1.68283595117764e-18,3.87857157576735e-19,1.34967799193072e-19,0,0,0,0,0,0,1.53170526740361e-17,5.04298730211564e-17,0,2.73304766746698e-19,5.93381344177045e-18,1.20567104039909e-17,2.39699982013287e-17,5.91876053427146e-18,1.27667086269615e-17,2.03725580116582e-17,9.3272284174518e-18,0,0,0,1.50727930830902e-17,3.89861316470221e-17,0,0,0,0,0,0,0,3.54223729253157e-17,1.96906248242697e-17,0,1.84784575204013e-17,2.94814416163897e-17,3.06754673480388e-17,5.24252109648893e-17,3.33525118241125e-17,2.90084672674881e-17,1.87725656007325e-16,8.84277918359829e-19,1.17400751532287e-16,6.88620465166031e-17,1.71112414255735e-16,1.41332674945419e-16,4.0199572808478e-17,9.81191849109218e-18,4.33782699736098e-17,6.02937471047154e-17,3.16198827125384e-17,1.05911766995904e-17,2.39173226441906e-17,2.85956929370373e-17,1.37733566780618e-16,5.47842548428707e-17,6.901017303973e-19,3.61191227329637e-17,5.79963630437379e-18,3.37007571144309e-17,1.21864472405707e-17,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1.11761411731152e-15,8.0375171885468e-15,4.42235225908563e-14,2.23182498156696e-13,1.05400600127999e-12,4.66702243191415e-12,1.93600389713714e-11,7.50745190685338e-11,2.85694227156459e-10,1.16021782839763e-09,4.46389296849664e-09,1.62737306088124e-08,5.62272568560506e-08,1.8416661421634e-07,5.72050305423628e-07,1.68584124505439e-06,4.71640097745618e-06,1.25351361156937e-05,3.16774939913891e-05,7.61946992909692e-05,0.000174649423080082,0.000381989942345034,0.000798344940551069,0.0016293615371213,0.00321460954287157,0.00603160705865666,0.0107699246251697,0.0183134501916207,0.0296772387517745,0.0458664140803032,0.0676545261869854,0.0953053420315228,0.128292571064086,0.16509642609934,0.203162910329539,0.239092558792032,0.26907767530551,0.289538176191551,0.296963178700738,0.291197371433341,0.273716091778852,0.247588802074231,0.217170870431105,0.187506543878362,0.163720422412581,0.150487748927325,0.151614485075058,0.169712073966721,0.205939555364443,0.259811752012319,0.329122923389784,0.410081770973002,0.497771099198642,0.586592375892215,0.672019892577224,0.751988733255122,0.826763922134062,0.898562942577085,0.970584311001888,1.04579410387695,1.12593823938228,1.21117754253386,1.30049909275768,1.39272435597267,1.48763856608892,1.58663708058872,1.69241136336144,1.80788454693939,1.93325905992369,2.06381895988349,2.19163358255908,2.30706069455458,2.4015774934155,2.47039059456206,2.51376669166625,2.53651354960027,2.54576267393452,2.54788084602031,2.54571055307302,2.53725723187937,2.51640993864327,2.47547821105795,2.40712236285046,2.31155331897565,2.19469993826932,2.06510764481844,1.93167522172108,1.80102190062914,1.67605319603567,1.55607567067049,1.43819451259637,1.3192987159709,1.19781809828058,1.07462566678766,0.95285570679636,0.836847045283915,0.730732015765482,0.63827555712345,0.559178732467227,0.492380205123583,0.436165371681983,0.38879693243745,0.34889233173202,0.315487226288247,0.287888687147809,0.265472808056132,0.247554724912778,0.233384084597811,0.222236270983436,0.213516185837874,0.206788312898052,0.201713589277792,0.197799958892236,0.194164081737767,0.189754040186259,0.183425591807649,0.174205857547399,0],"text":["density: 1.742059e-01<br />Petal.Length: 1.000000","density: 1.834256e-01<br />Petal.Length: 1.011546","density: 1.897540e-01<br />Petal.Length: 1.023092","density: 1.941641e-01<br />Petal.Length: 1.034638","density: 1.978000e-01<br />Petal.Length: 1.046184","density: 2.017136e-01<br />Petal.Length: 1.057730","density: 2.067883e-01<br />Petal.Length: 1.069276","density: 2.135162e-01<br />Petal.Length: 1.080822","density: 2.222363e-01<br />Petal.Length: 1.092368","density: 2.333841e-01<br />Petal.Length: 1.103914","density: 2.475547e-01<br />Petal.Length: 1.115460","density: 2.654728e-01<br />Petal.Length: 1.127006","density: 2.878887e-01<br />Petal.Length: 1.138552","density: 3.154872e-01<br />Petal.Length: 1.150098","density: 3.488923e-01<br />Petal.Length: 1.161644","density: 3.887969e-01<br />Petal.Length: 1.173190","density: 4.361654e-01<br />Petal.Length: 1.184736","density: 4.923802e-01<br />Petal.Length: 1.196282","density: 5.591787e-01<br />Petal.Length: 1.207828","density: 6.382756e-01<br />Petal.Length: 1.219374","density: 7.307320e-01<br />Petal.Length: 1.230920","density: 8.368470e-01<br />Petal.Length: 1.242466","density: 9.528557e-01<br />Petal.Length: 1.254012","density: 1.074626e+00<br />Petal.Length: 1.265558","density: 1.197818e+00<br />Petal.Length: 1.277104","density: 1.319299e+00<br />Petal.Length: 1.288650","density: 1.438195e+00<br />Petal.Length: 1.300196","density: 1.556076e+00<br />Petal.Length: 1.311742","density: 1.676053e+00<br />Petal.Length: 1.323288","density: 1.801022e+00<br />Petal.Length: 1.334834","density: 1.931675e+00<br />Petal.Length: 1.346380","density: 2.065108e+00<br />Petal.Length: 1.357926","density: 2.194700e+00<br />Petal.Length: 1.369472","density: 2.311553e+00<br />Petal.Length: 1.381018","density: 2.407122e+00<br />Petal.Length: 1.392564","density: 2.475478e+00<br />Petal.Length: 1.404110","density: 2.516410e+00<br />Petal.Length: 1.415656","density: 2.537257e+00<br />Petal.Length: 1.427202","density: 2.545711e+00<br />Petal.Length: 1.438748","density: 2.547881e+00<br />Petal.Length: 1.450294","density: 2.545763e+00<br />Petal.Length: 1.461840","density: 2.536514e+00<br />Petal.Length: 1.473386","density: 2.513767e+00<br />Petal.Length: 1.484932","density: 2.470391e+00<br />Petal.Length: 1.496477","density: 2.401577e+00<br />Petal.Length: 1.508023","density: 2.307061e+00<br />Petal.Length: 1.519569","density: 2.191634e+00<br />Petal.Length: 1.531115","density: 2.063819e+00<br />Petal.Length: 1.542661","density: 1.933259e+00<br />Petal.Length: 1.554207","density: 1.807885e+00<br />Petal.Length: 1.565753","density: 1.692411e+00<br />Petal.Length: 1.577299","density: 1.586637e+00<br />Petal.Length: 1.588845","density: 1.487639e+00<br />Petal.Length: 1.600391","density: 1.392724e+00<br />Petal.Length: 1.611937","density: 1.300499e+00<br />Petal.Length: 1.623483","density: 1.211178e+00<br />Petal.Length: 1.635029","density: 1.125938e+00<br />Petal.Length: 1.646575","density: 1.045794e+00<br />Petal.Length: 1.658121","density: 9.705843e-01<br />Petal.Length: 1.669667","density: 8.985629e-01<br />Petal.Length: 1.681213","density: 8.267639e-01<br />Petal.Length: 1.692759","density: 7.519887e-01<br />Petal.Length: 1.704305","density: 6.720199e-01<br />Petal.Length: 1.715851","density: 5.865924e-01<br />Petal.Length: 1.727397","density: 4.977711e-01<br />Petal.Length: 1.738943","density: 4.100818e-01<br />Petal.Length: 1.750489","density: 3.291229e-01<br />Petal.Length: 1.762035","density: 2.598118e-01<br />Petal.Length: 1.773581","density: 2.059396e-01<br />Petal.Length: 1.785127","density: 1.697121e-01<br />Petal.Length: 1.796673","density: 1.516145e-01<br />Petal.Length: 1.808219","density: 1.504877e-01<br />Petal.Length: 1.819765","density: 1.637204e-01<br />Petal.Length: 1.831311","density: 1.875065e-01<br />Petal.Length: 1.842857","density: 2.171709e-01<br />Petal.Length: 1.854403","density: 2.475888e-01<br />Petal.Length: 1.865949","density: 2.737161e-01<br />Petal.Length: 1.877495","density: 2.911974e-01<br />Petal.Length: 1.889041","density: 2.969632e-01<br />Petal.Length: 1.900587","density: 2.895382e-01<br />Petal.Length: 1.912133","density: 2.690777e-01<br />Petal.Length: 1.923679","density: 2.390926e-01<br />Petal.Length: 1.935225","density: 2.031629e-01<br />Petal.Length: 1.946771","density: 1.650964e-01<br />Petal.Length: 1.958317","density: 1.282926e-01<br />Petal.Length: 1.969863","density: 9.530534e-02<br />Petal.Length: 1.981409","density: 6.765453e-02<br />Petal.Length: 1.992955","density: 4.586641e-02<br />Petal.Length: 2.004501","density: 2.967724e-02<br />Petal.Length: 2.016047","density: 1.831345e-02<br />Petal.Length: 2.027593","density: 1.076992e-02<br />Petal.Length: 2.039139","density: 6.031607e-03<br />Petal.Length: 2.050685","density: 3.214610e-03<br />Petal.Length: 2.062231","density: 1.629362e-03<br />Petal.Length: 2.073777","density: 7.983449e-04<br />Petal.Length: 2.085323","density: 3.819899e-04<br />Petal.Length: 2.096869","density: 1.746494e-04<br />Petal.Length: 2.108415","density: 7.619470e-05<br />Petal.Length: 2.119961","density: 3.167749e-05<br />Petal.Length: 2.131507","density: 1.253514e-05<br />Petal.Length: 2.143053","density: 4.716401e-06<br />Petal.Length: 2.154599","density: 1.685841e-06<br />Petal.Length: 2.166145","density: 5.720503e-07<br />Petal.Length: 2.177691","density: 1.841666e-07<br />Petal.Length: 2.189237","density: 5.622726e-08<br />Petal.Length: 2.200783","density: 1.627373e-08<br />Petal.Length: 2.212329","density: 4.463893e-09<br />Petal.Length: 2.223875","density: 1.160218e-09<br />Petal.Length: 2.235421","density: 2.856942e-10<br />Petal.Length: 2.246967","density: 7.507452e-11<br />Petal.Length: 2.258513","density: 1.936004e-11<br />Petal.Length: 2.270059","density: 4.667022e-12<br />Petal.Length: 2.281605","density: 1.054006e-12<br />Petal.Length: 2.293151","density: 2.231825e-13<br />Petal.Length: 2.304697","density: 4.422352e-14<br />Petal.Length: 2.316243","density: 8.037517e-15<br />Petal.Length: 2.327789","density: 1.117614e-15<br />Petal.Length: 2.339335","density: 0.000000e+00<br />Petal.Length: 2.350881","density: 0.000000e+00<br />Petal.Length: 2.362427","density: 0.000000e+00<br />Petal.Length: 2.373973","density: 0.000000e+00<br />Petal.Length: 2.385519","density: 0.000000e+00<br />Petal.Length: 2.397065","density: 0.000000e+00<br />Petal.Length: 2.408611","density: 0.000000e+00<br />Petal.Length: 2.420157","density: 0.000000e+00<br />Petal.Length: 2.431703","density: 0.000000e+00<br />Petal.Length: 2.443249","density: 0.000000e+00<br />Petal.Length: 2.454795","density: 0.000000e+00<br />Petal.Length: 2.466341","density: 0.000000e+00<br />Petal.Length: 2.477886","density: 0.000000e+00<br />Petal.Length: 2.489432","density: 0.000000e+00<br />Petal.Length: 2.500978","density: 0.000000e+00<br />Petal.Length: 2.512524","density: 0.000000e+00<br />Petal.Length: 2.524070","density: 0.000000e+00<br />Petal.Length: 2.535616","density: 0.000000e+00<br />Petal.Length: 2.547162","density: 0.000000e+00<br />Petal.Length: 2.558708","density: 0.000000e+00<br />Petal.Length: 2.570254","density: 0.000000e+00<br />Petal.Length: 2.581800","density: 0.000000e+00<br />Petal.Length: 2.593346","density: 0.000000e+00<br />Petal.Length: 2.604892","density: 0.000000e+00<br />Petal.Length: 2.616438","density: 0.000000e+00<br />Petal.Length: 2.627984","density: 0.000000e+00<br />Petal.Length: 2.639530","density: 0.000000e+00<br />Petal.Length: 2.651076","density: 0.000000e+00<br />Petal.Length: 2.662622","density: 0.000000e+00<br />Petal.Length: 2.674168","density: 0.000000e+00<br />Petal.Length: 2.685714","density: 0.000000e+00<br />Petal.Length: 2.697260","density: 0.000000e+00<br />Petal.Length: 2.708806","density: 0.000000e+00<br />Petal.Length: 2.720352","density: 0.000000e+00<br />Petal.Length: 2.731898","density: 0.000000e+00<br />Petal.Length: 2.743444","density: 0.000000e+00<br />Petal.Length: 2.754990","density: 0.000000e+00<br />Petal.Length: 2.766536","density: 0.000000e+00<br />Petal.Length: 2.778082","density: 0.000000e+00<br />Petal.Length: 2.789628","density: 0.000000e+00<br />Petal.Length: 2.801174","density: 0.000000e+00<br />Petal.Length: 2.812720","density: 0.000000e+00<br />Petal.Length: 2.824266","density: 0.000000e+00<br />Petal.Length: 2.835812","density: 0.000000e+00<br />Petal.Length: 2.847358","density: 0.000000e+00<br />Petal.Length: 2.858904","density: 0.000000e+00<br />Petal.Length: 2.870450","density: 1.218645e-17<br />Petal.Length: 2.881996","density: 3.370076e-17<br />Petal.Length: 2.893542","density: 5.799636e-18<br />Petal.Length: 2.905088","density: 3.611912e-17<br />Petal.Length: 2.916634","density: 6.901017e-19<br />Petal.Length: 2.928180","density: 5.478425e-17<br />Petal.Length: 2.939726","density: 1.377336e-16<br />Petal.Length: 2.951272","density: 2.859569e-17<br />Petal.Length: 2.962818","density: 2.391732e-17<br />Petal.Length: 2.974364","density: 1.059118e-17<br />Petal.Length: 2.985910","density: 3.161988e-17<br />Petal.Length: 2.997456","density: 6.029375e-17<br />Petal.Length: 3.009002","density: 4.337827e-17<br />Petal.Length: 3.020548","density: 9.811918e-18<br />Petal.Length: 3.032094","density: 4.019957e-17<br />Petal.Length: 3.043640","density: 1.413327e-16<br />Petal.Length: 3.055186","density: 1.711124e-16<br />Petal.Length: 3.066732","density: 6.886205e-17<br />Petal.Length: 3.078278","density: 1.174008e-16<br />Petal.Length: 3.089824","density: 8.842779e-19<br />Petal.Length: 3.101370","density: 1.877257e-16<br />Petal.Length: 3.112916","density: 2.900847e-17<br />Petal.Length: 3.124462","density: 3.335251e-17<br />Petal.Length: 3.136008","density: 5.242521e-17<br />Petal.Length: 3.147554","density: 3.067547e-17<br />Petal.Length: 3.159100","density: 2.948144e-17<br />Petal.Length: 3.170646","density: 1.847846e-17<br />Petal.Length: 3.182192","density: 0.000000e+00<br />Petal.Length: 3.193738","density: 1.969062e-17<br />Petal.Length: 3.205284","density: 3.542237e-17<br />Petal.Length: 3.216830","density: 0.000000e+00<br />Petal.Length: 3.228376","density: 0.000000e+00<br />Petal.Length: 3.239922","density: 0.000000e+00<br />Petal.Length: 3.251468","density: 0.000000e+00<br />Petal.Length: 3.263014","density: 0.000000e+00<br />Petal.Length: 3.274560","density: 0.000000e+00<br />Petal.Length: 3.286106","density: 0.000000e+00<br />Petal.Length: 3.297652","density: 3.898613e-17<br />Petal.Length: 3.309198","density: 1.507279e-17<br />Petal.Length: 3.320744","density: 0.000000e+00<br />Petal.Length: 3.332290","density: 0.000000e+00<br />Petal.Length: 3.343836","density: 0.000000e+00<br />Petal.Length: 3.355382","density: 9.327228e-18<br />Petal.Length: 3.366928","density: 2.037256e-17<br />Petal.Length: 3.378474","density: 1.276671e-17<br />Petal.Length: 3.390020","density: 5.918761e-18<br />Petal.Length: 3.401566","density: 2.397000e-17<br />Petal.Length: 3.413112","density: 1.205671e-17<br />Petal.Length: 3.424658","density: 5.933813e-18<br />Petal.Length: 3.436204","density: 2.733048e-19<br />Petal.Length: 3.447750","density: 0.000000e+00<br />Petal.Length: 3.459295","density: 5.042987e-17<br />Petal.Length: 3.470841","density: 1.531705e-17<br />Petal.Length: 3.482387","density: 0.000000e+00<br />Petal.Length: 3.493933","density: 0.000000e+00<br />Petal.Length: 3.505479","density: 0.000000e+00<br />Petal.Length: 3.517025","density: 0.000000e+00<br />Petal.Length: 3.528571","density: 0.000000e+00<br />Petal.Length: 3.540117","density: 0.000000e+00<br />Petal.Length: 3.551663","density: 1.349678e-19<br />Petal.Length: 3.563209","density: 3.878572e-19<br />Petal.Length: 3.574755","density: 1.682836e-18<br />Petal.Length: 3.586301","density: 1.116999e-17<br />Petal.Length: 3.597847","density: 0.000000e+00<br />Petal.Length: 3.609393","density: 0.000000e+00<br />Petal.Length: 3.620939","density: 0.000000e+00<br />Petal.Length: 3.632485","density: 3.343603e-18<br />Petal.Length: 3.644031","density: 1.118866e-18<br />Petal.Length: 3.655577","density: 0.000000e+00<br />Petal.Length: 3.667123","density: 0.000000e+00<br />Petal.Length: 3.678669","density: 0.000000e+00<br />Petal.Length: 3.690215","density: 0.000000e+00<br />Petal.Length: 3.701761","density: 0.000000e+00<br />Petal.Length: 3.713307","density: 1.029791e-18<br />Petal.Length: 3.724853","density: 7.137799e-18<br />Petal.Length: 3.736399","density: 1.510460e-17<br />Petal.Length: 3.747945","density: 0.000000e+00<br />Petal.Length: 3.759491","density: 0.000000e+00<br />Petal.Length: 3.771037","density: 0.000000e+00<br />Petal.Length: 3.782583","density: 0.000000e+00<br />Petal.Length: 3.794129","density: 0.000000e+00<br />Petal.Length: 3.805675","density: 0.000000e+00<br />Petal.Length: 3.817221","density: 2.017255e-17<br />Petal.Length: 3.828767","density: 1.061729e-16<br />Petal.Length: 3.840313","density: 6.295653e-17<br />Petal.Length: 3.851859","density: 3.431683e-17<br />Petal.Length: 3.863405","density: 5.561569e-17<br />Petal.Length: 3.874951","density: 5.020507e-17<br />Petal.Length: 3.886497","density: 4.584421e-17<br />Petal.Length: 3.898043","density: 2.248309e-18<br />Petal.Length: 3.909589","density: 1.456665e-17<br />Petal.Length: 3.921135","density: 4.477662e-17<br />Petal.Length: 3.932681","density: 8.913128e-17<br />Petal.Length: 3.944227","density: 4.162964e-17<br />Petal.Length: 3.955773","density: 8.739414e-18<br />Petal.Length: 3.967319","density: 8.838109e-19<br />Petal.Length: 3.978865","density: 4.892430e-17<br />Petal.Length: 3.990411","density: 1.962060e-17<br />Petal.Length: 4.001957","density: 0.000000e+00<br />Petal.Length: 4.013503","density: 0.000000e+00<br />Petal.Length: 4.025049","density: 1.531643e-17<br />Petal.Length: 4.036595","density: 1.802974e-17<br />Petal.Length: 4.048141","density: 0.000000e+00<br />Petal.Length: 4.059687","density: 0.000000e+00<br />Petal.Length: 4.071233","density: 0.000000e+00<br />Petal.Length: 4.082779","density: 0.000000e+00<br />Petal.Length: 4.094325","density: 0.000000e+00<br />Petal.Length: 4.105871","density: 0.000000e+00<br />Petal.Length: 4.117417","density: 0.000000e+00<br />Petal.Length: 4.128963","density: 0.000000e+00<br />Petal.Length: 4.140509","density: 0.000000e+00<br />Petal.Length: 4.152055","density: 0.000000e+00<br />Petal.Length: 4.163601","density: 0.000000e+00<br />Petal.Length: 4.175147","density: 0.000000e+00<br />Petal.Length: 4.186693","density: 0.000000e+00<br />Petal.Length: 4.198239","density: 0.000000e+00<br />Petal.Length: 4.209785","density: 0.000000e+00<br />Petal.Length: 4.221331","density: 0.000000e+00<br />Petal.Length: 4.232877","density: 0.000000e+00<br />Petal.Length: 4.244423","density: 0.000000e+00<br />Petal.Length: 4.255969","density: 0.000000e+00<br />Petal.Length: 4.267515","density: 0.000000e+00<br />Petal.Length: 4.279061","density: 0.000000e+00<br />Petal.Length: 4.290607","density: 0.000000e+00<br />Petal.Length: 4.302153","density: 0.000000e+00<br />Petal.Length: 4.313699","density: 0.000000e+00<br />Petal.Length: 4.325245","density: 3.266853e-17<br />Petal.Length: 4.336791","density: 1.549816e-17<br />Petal.Length: 4.348337","density: 0.000000e+00<br />Petal.Length: 4.359883","density: 0.000000e+00<br />Petal.Length: 4.371429","density: 0.000000e+00<br />Petal.Length: 4.382975","density: 0.000000e+00<br />Petal.Length: 4.394521","density: 9.340365e-18<br />Petal.Length: 4.406067","density: 2.215324e-17<br />Petal.Length: 4.417613","density: 2.070752e-17<br />Petal.Length: 4.429159","density: 1.101818e-16<br />Petal.Length: 4.440705","density: 7.945053e-17<br />Petal.Length: 4.452250","density: 1.441258e-16<br />Petal.Length: 4.463796","density: 1.954813e-17<br />Petal.Length: 4.475342","density: 5.947249e-17<br />Petal.Length: 4.486888","density: 4.792614e-17<br />Petal.Length: 4.498434","density: 9.547105e-17<br />Petal.Length: 4.509980","density: 1.179612e-16<br />Petal.Length: 4.521526","density: 2.068723e-16<br />Petal.Length: 4.533072","density: 1.366169e-16<br />Petal.Length: 4.544618","density: 7.823066e-17<br />Petal.Length: 4.556164","density: 1.474237e-16<br />Petal.Length: 4.567710","density: 9.382137e-17<br />Petal.Length: 4.579256","density: 1.131134e-16<br />Petal.Length: 4.590802","density: 1.556458e-16<br />Petal.Length: 4.602348","density: 1.828905e-16<br />Petal.Length: 4.613894","density: 1.238881e-16<br />Petal.Length: 4.625440","density: 1.497582e-16<br />Petal.Length: 4.636986","density: 1.756283e-16<br />Petal.Length: 4.648532","density: 2.143357e-16<br />Petal.Length: 4.660078","density: 2.704601e-16<br />Petal.Length: 4.671624","density: 2.802759e-16<br />Petal.Length: 4.683170","density: 2.078209e-16<br />Petal.Length: 4.694716","density: 1.838988e-16<br />Petal.Length: 4.706262","density: 2.229337e-16<br />Petal.Length: 4.717808","density: 2.286438e-16<br />Petal.Length: 4.729354","density: 1.314564e-16<br />Petal.Length: 4.740900","density: 7.738189e-18<br />Petal.Length: 4.752446","density: 3.360830e-17<br />Petal.Length: 4.763992","density: 5.551115e-17<br />Petal.Length: 4.775538","density: 5.551115e-17<br />Petal.Length: 4.787084","density: 5.541298e-17<br />Petal.Length: 4.798630","density: 4.247793e-17<br />Petal.Length: 4.810176","density: 4.163336e-17<br />Petal.Length: 4.821722","density: 8.190080e-18<br />Petal.Length: 4.833268","density: 0.000000e+00<br />Petal.Length: 4.844814","density: 0.000000e+00<br />Petal.Length: 4.856360","density: 5.823696e-17<br />Petal.Length: 4.867906","density: 7.132559e-17<br />Petal.Length: 4.879452","density: 3.892110e-17<br />Petal.Length: 4.890998","density: 1.677283e-17<br />Petal.Length: 4.902544","density: 1.137160e-17<br />Petal.Length: 4.914090","density: 2.567970e-17<br />Petal.Length: 4.925636","density: 0.000000e+00<br />Petal.Length: 4.937182","density: 0.000000e+00<br />Petal.Length: 4.948728","density: 0.000000e+00<br />Petal.Length: 4.960274","density: 0.000000e+00<br />Petal.Length: 4.971820","density: 0.000000e+00<br />Petal.Length: 4.983366","density: 0.000000e+00<br />Petal.Length: 4.994912","density: 0.000000e+00<br />Petal.Length: 5.006458","density: 0.000000e+00<br />Petal.Length: 5.018004","density: 0.000000e+00<br />Petal.Length: 5.029550","density: 0.000000e+00<br />Petal.Length: 5.041096","density: 0.000000e+00<br />Petal.Length: 5.052642","density: 0.000000e+00<br />Petal.Length: 5.064188","density: 0.000000e+00<br />Petal.Length: 5.075734","density: 0.000000e+00<br />Petal.Length: 5.087280","density: 0.000000e+00<br />Petal.Length: 5.098826","density: 0.000000e+00<br />Petal.Length: 5.110372","density: 0.000000e+00<br />Petal.Length: 5.121918","density: 0.000000e+00<br />Petal.Length: 5.133464","density: 0.000000e+00<br />Petal.Length: 5.145010","density: 0.000000e+00<br />Petal.Length: 5.156556","density: 0.000000e+00<br />Petal.Length: 5.168102","density: 0.000000e+00<br />Petal.Length: 5.179648","density: 0.000000e+00<br />Petal.Length: 5.191194","density: 0.000000e+00<br />Petal.Length: 5.202740","density: 0.000000e+00<br />Petal.Length: 5.214286","density: 0.000000e+00<br />Petal.Length: 5.225832","density: 0.000000e+00<br />Petal.Length: 5.237378","density: 0.000000e+00<br />Petal.Length: 5.248924","density: 0.000000e+00<br />Petal.Length: 5.260470","density: 0.000000e+00<br />Petal.Length: 5.272016","density: 0.000000e+00<br />Petal.Length: 5.283562","density: 0.000000e+00<br />Petal.Length: 5.295108","density: 0.000000e+00<br />Petal.Length: 5.306654","density: 0.000000e+00<br />Petal.Length: 5.318200","density: 0.000000e+00<br />Petal.Length: 5.329746","density: 0.000000e+00<br />Petal.Length: 5.341292","density: 0.000000e+00<br />Petal.Length: 5.352838","density: 0.000000e+00<br />Petal.Length: 5.364384","density: 0.000000e+00<br />Petal.Length: 5.375930","density: 0.000000e+00<br />Petal.Length: 5.387476","density: 0.000000e+00<br />Petal.Length: 5.399022","density: 0.000000e+00<br />Petal.Length: 5.410568","density: 0.000000e+00<br />Petal.Length: 5.422114","density: 0.000000e+00<br />Petal.Length: 5.433659","density: 0.000000e+00<br />Petal.Length: 5.445205","density: 0.000000e+00<br />Petal.Length: 5.456751","density: 0.000000e+00<br />Petal.Length: 5.468297","density: 0.000000e+00<br />Petal.Length: 5.479843","density: 0.000000e+00<br />Petal.Length: 5.491389","density: 3.075475e-17<br />Petal.Length: 5.502935","density: 7.290582e-18<br />Petal.Length: 5.514481","density: 0.000000e+00<br />Petal.Length: 5.526027","density: 3.641286e-17<br />Petal.Length: 5.537573","density: 5.110874e-17<br />Petal.Length: 5.549119","density: 3.631019e-17<br />Petal.Length: 5.560665","density: 1.383918e-17<br />Petal.Length: 5.572211","density: 3.110603e-17<br />Petal.Length: 5.583757","density: 6.422336e-17<br />Petal.Length: 5.595303","density: 2.259906e-17<br />Petal.Length: 5.606849","density: 0.000000e+00<br />Petal.Length: 5.618395","density: 3.102945e-18<br />Petal.Length: 5.629941","density: 2.560795e-17<br />Petal.Length: 5.641487","density: 3.688931e-18<br />Petal.Length: 5.653033","density: 3.231685e-17<br />Petal.Length: 5.664579","density: 5.466761e-18<br />Petal.Length: 5.676125","density: 5.558608e-17<br />Petal.Length: 5.687671","density: 5.252838e-17<br />Petal.Length: 5.699217","density: 1.606193e-17<br />Petal.Length: 5.710763","density: 0.000000e+00<br />Petal.Length: 5.722309","density: 3.613339e-17<br />Petal.Length: 5.733855","density: 5.003233e-17<br />Petal.Length: 5.745401","density: 5.023293e-17<br />Petal.Length: 5.756947","density: 7.172915e-17<br />Petal.Length: 5.768493","density: 2.453395e-17<br />Petal.Length: 5.780039","density: 6.020755e-17<br />Petal.Length: 5.791585","density: 9.974909e-18<br />Petal.Length: 5.803131","density: 7.421234e-17<br />Petal.Length: 5.814677","density: 4.463489e-17<br />Petal.Length: 5.826223","density: 1.623104e-17<br />Petal.Length: 5.837769","density: 4.556805e-17<br />Petal.Length: 5.849315","density: 7.799873e-17<br />Petal.Length: 5.860861","density: 4.706699e-17<br />Petal.Length: 5.872407","density: 5.496194e-17<br />Petal.Length: 5.883953","density: 5.448509e-17<br />Petal.Length: 5.895499","density: 9.507063e-17<br />Petal.Length: 5.907045","density: 1.020492e-16<br />Petal.Length: 5.918591","density: 3.916289e-17<br />Petal.Length: 5.930137","density: 4.014632e-17<br />Petal.Length: 5.941683","density: 3.216241e-17<br />Petal.Length: 5.953229","density: 9.281191e-17<br />Petal.Length: 5.964775","density: 2.974846e-17<br />Petal.Length: 5.976321","density: 3.288295e-17<br />Petal.Length: 5.987867","density: 1.918431e-18<br />Petal.Length: 5.999413","density: 1.122994e-16<br />Petal.Length: 6.010959","density: 5.666078e-17<br />Petal.Length: 6.022505","density: 7.110372e-17<br />Petal.Length: 6.034051","density: 1.195446e-16<br />Petal.Length: 6.045597","density: 9.190046e-17<br />Petal.Length: 6.057143","density: 7.318831e-17<br />Petal.Length: 6.068689","density: 7.790078e-17<br />Petal.Length: 6.080235","density: 4.314775e-17<br />Petal.Length: 6.091781","density: 4.675364e-17<br />Petal.Length: 6.103327","density: 1.323831e-16<br />Petal.Length: 6.114873","density: 1.250800e-16<br />Petal.Length: 6.126419","density: 1.788384e-16<br />Petal.Length: 6.137965","density: 5.235632e-17<br />Petal.Length: 6.149511","density: 1.495039e-16<br />Petal.Length: 6.161057","density: 7.705938e-17<br />Petal.Length: 6.172603","density: 1.804987e-16<br />Petal.Length: 6.184149","density: 6.612232e-17<br />Petal.Length: 6.195695","density: 1.353169e-16<br />Petal.Length: 6.207241","density: 1.161540e-16<br />Petal.Length: 6.218787","density: 4.280769e-17<br />Petal.Length: 6.230333","density: 6.499642e-18<br />Petal.Length: 6.241879","density: 2.916949e-17<br />Petal.Length: 6.253425","density: 1.163818e-16<br />Petal.Length: 6.264971","density: 1.606297e-16<br />Petal.Length: 6.276517","density: 5.764631e-17<br />Petal.Length: 6.288063","density: 8.695933e-18<br />Petal.Length: 6.299609","density: 1.336277e-17<br />Petal.Length: 6.311155","density: 2.257470e-17<br />Petal.Length: 6.322701","density: 3.908722e-17<br />Petal.Length: 6.334247","density: 3.786743e-18<br />Petal.Length: 6.345793","density: 2.730899e-17<br />Petal.Length: 6.357339","density: 7.559930e-18<br />Petal.Length: 6.368885","density: 2.970067e-17<br />Petal.Length: 6.380431","density: 1.557680e-17<br />Petal.Length: 6.391977","density: 2.039293e-17<br />Petal.Length: 6.403523","density: 1.817522e-17<br />Petal.Length: 6.415068","density: 0.000000e+00<br />Petal.Length: 6.426614","density: 1.389530e-17<br />Petal.Length: 6.438160","density: 2.722468e-17<br />Petal.Length: 6.449706","density: 0.000000e+00<br />Petal.Length: 6.461252","density: 0.000000e+00<br />Petal.Length: 6.472798","density: 0.000000e+00<br />Petal.Length: 6.484344","density: 0.000000e+00<br />Petal.Length: 6.495890","density: 0.000000e+00<br />Petal.Length: 6.507436","density: 0.000000e+00<br />Petal.Length: 6.518982","density: 0.000000e+00<br />Petal.Length: 6.530528","density: 6.831271e-17<br />Petal.Length: 6.542074","density: 8.657778e-17<br />Petal.Length: 6.553620","density: 3.262620e-17<br />Petal.Length: 6.565166","density: 0.000000e+00<br />Petal.Length: 6.576712","density: 0.000000e+00<br />Petal.Length: 6.588258","density: 0.000000e+00<br />Petal.Length: 6.599804","density: 0.000000e+00<br />Petal.Length: 6.611350","density: 0.000000e+00<br />Petal.Length: 6.622896","density: 1.826275e-17<br />Petal.Length: 6.634442","density: 8.618956e-17<br />Petal.Length: 6.645988","density: 6.875999e-17<br />Petal.Length: 6.657534","density: 3.980472e-17<br />Petal.Length: 6.669080","density: 2.811084e-17<br />Petal.Length: 6.680626","density: 3.645670e-18<br />Petal.Length: 6.692172","density: 2.476698e-17<br />Petal.Length: 6.703718","density: 3.112942e-17<br />Petal.Length: 6.715264","density: 2.529619e-17<br />Petal.Length: 6.726810","density: 9.040874e-18<br />Petal.Length: 6.738356","density: 4.289994e-17<br />Petal.Length: 6.749902","density: 5.093671e-17<br />Petal.Length: 6.761448","density: 2.239780e-17<br />Petal.Length: 6.772994","density: 2.302113e-17<br />Petal.Length: 6.784540","density: 2.640002e-18<br />Petal.Length: 6.796086","density: 1.792024e-17<br />Petal.Length: 6.807632","density: 4.475063e-17<br />Petal.Length: 6.819178","density: 4.195203e-18<br />Petal.Length: 6.830724","density: 8.053217e-17<br />Petal.Length: 6.842270","density: 1.033337e-16<br />Petal.Length: 6.853816","density: 8.232359e-17<br />Petal.Length: 6.865362","density: 3.415983e-17<br />Petal.Length: 6.876908","density: 7.617698e-18<br />Petal.Length: 6.888454","density: 3.692235e-17<br />Petal.Length: 6.900000","density: 3.692235e-17<br />Petal.Length: 6.900000","density: 7.617698e-18<br />Petal.Length: 6.888454","density: 3.415983e-17<br />Petal.Length: 6.876908","density: 8.232359e-17<br />Petal.Length: 6.865362","density: 1.033337e-16<br />Petal.Length: 6.853816","density: 8.053217e-17<br />Petal.Length: 6.842270","density: 4.195203e-18<br />Petal.Length: 6.830724","density: 4.475063e-17<br />Petal.Length: 6.819178","density: 1.792024e-17<br />Petal.Length: 6.807632","density: 2.640002e-18<br />Petal.Length: 6.796086","density: 2.302113e-17<br />Petal.Length: 6.784540","density: 2.239780e-17<br />Petal.Length: 6.772994","density: 5.093671e-17<br />Petal.Length: 6.761448","density: 4.289994e-17<br />Petal.Length: 6.749902","density: 9.040874e-18<br />Petal.Length: 6.738356","density: 2.529619e-17<br />Petal.Length: 6.726810","density: 3.112942e-17<br />Petal.Length: 6.715264","density: 2.476698e-17<br />Petal.Length: 6.703718","density: 3.645670e-18<br />Petal.Length: 6.692172","density: 2.811084e-17<br />Petal.Length: 6.680626","density: 3.980472e-17<br />Petal.Length: 6.669080","density: 6.875999e-17<br />Petal.Length: 6.657534","density: 8.618956e-17<br />Petal.Length: 6.645988","density: 1.826275e-17<br />Petal.Length: 6.634442","density: 0.000000e+00<br />Petal.Length: 6.622896","density: 0.000000e+00<br />Petal.Length: 6.611350","density: 0.000000e+00<br />Petal.Length: 6.599804","density: 0.000000e+00<br />Petal.Length: 6.588258","density: 0.000000e+00<br />Petal.Length: 6.576712","density: 3.262620e-17<br />Petal.Length: 6.565166","density: 8.657778e-17<br />Petal.Length: 6.553620","density: 6.831271e-17<br />Petal.Length: 6.542074","density: 0.000000e+00<br />Petal.Length: 6.530528","density: 0.000000e+00<br />Petal.Length: 6.518982","density: 0.000000e+00<br />Petal.Length: 6.507436","density: 0.000000e+00<br />Petal.Length: 6.495890","density: 0.000000e+00<br />Petal.Length: 6.484344","density: 0.000000e+00<br />Petal.Length: 6.472798","density: 0.000000e+00<br />Petal.Length: 6.461252","density: 2.722468e-17<br />Petal.Length: 6.449706","density: 1.389530e-17<br />Petal.Length: 6.438160","density: 0.000000e+00<br />Petal.Length: 6.426614","density: 1.817522e-17<br />Petal.Length: 6.415068","density: 2.039293e-17<br />Petal.Length: 6.403523","density: 1.557680e-17<br />Petal.Length: 6.391977","density: 2.970067e-17<br />Petal.Length: 6.380431","density: 7.559930e-18<br />Petal.Length: 6.368885","density: 2.730899e-17<br />Petal.Length: 6.357339","density: 3.786743e-18<br />Petal.Length: 6.345793","density: 3.908722e-17<br />Petal.Length: 6.334247","density: 2.257470e-17<br />Petal.Length: 6.322701","density: 1.336277e-17<br />Petal.Length: 6.311155","density: 8.695933e-18<br />Petal.Length: 6.299609","density: 5.764631e-17<br />Petal.Length: 6.288063","density: 1.606297e-16<br />Petal.Length: 6.276517","density: 1.163818e-16<br />Petal.Length: 6.264971","density: 2.916949e-17<br />Petal.Length: 6.253425","density: 6.499642e-18<br />Petal.Length: 6.241879","density: 4.280769e-17<br />Petal.Length: 6.230333","density: 1.161540e-16<br />Petal.Length: 6.218787","density: 1.353169e-16<br />Petal.Length: 6.207241","density: 6.612232e-17<br />Petal.Length: 6.195695","density: 1.804987e-16<br />Petal.Length: 6.184149","density: 7.705938e-17<br />Petal.Length: 6.172603","density: 1.495039e-16<br />Petal.Length: 6.161057","density: 5.235632e-17<br />Petal.Length: 6.149511","density: 1.788384e-16<br />Petal.Length: 6.137965","density: 1.250800e-16<br />Petal.Length: 6.126419","density: 1.323831e-16<br />Petal.Length: 6.114873","density: 4.675364e-17<br />Petal.Length: 6.103327","density: 4.314775e-17<br />Petal.Length: 6.091781","density: 7.790078e-17<br />Petal.Length: 6.080235","density: 7.318831e-17<br />Petal.Length: 6.068689","density: 9.190046e-17<br />Petal.Length: 6.057143","density: 1.195446e-16<br />Petal.Length: 6.045597","density: 7.110372e-17<br />Petal.Length: 6.034051","density: 5.666078e-17<br />Petal.Length: 6.022505","density: 1.122994e-16<br />Petal.Length: 6.010959","density: 1.918431e-18<br />Petal.Length: 5.999413","density: 3.288295e-17<br />Petal.Length: 5.987867","density: 2.974846e-17<br />Petal.Length: 5.976321","density: 9.281191e-17<br />Petal.Length: 5.964775","density: 3.216241e-17<br />Petal.Length: 5.953229","density: 4.014632e-17<br />Petal.Length: 5.941683","density: 3.916289e-17<br />Petal.Length: 5.930137","density: 1.020492e-16<br />Petal.Length: 5.918591","density: 9.507063e-17<br />Petal.Length: 5.907045","density: 5.448509e-17<br />Petal.Length: 5.895499","density: 5.496194e-17<br />Petal.Length: 5.883953","density: 4.706699e-17<br />Petal.Length: 5.872407","density: 7.799873e-17<br />Petal.Length: 5.860861","density: 4.556805e-17<br />Petal.Length: 5.849315","density: 1.623104e-17<br />Petal.Length: 5.837769","density: 4.463489e-17<br />Petal.Length: 5.826223","density: 7.421234e-17<br />Petal.Length: 5.814677","density: 9.974909e-18<br />Petal.Length: 5.803131","density: 6.020755e-17<br />Petal.Length: 5.791585","density: 2.453395e-17<br />Petal.Length: 5.780039","density: 7.172915e-17<br />Petal.Length: 5.768493","density: 5.023293e-17<br />Petal.Length: 5.756947","density: 5.003233e-17<br />Petal.Length: 5.745401","density: 3.613339e-17<br />Petal.Length: 5.733855","density: 0.000000e+00<br />Petal.Length: 5.722309","density: 1.606193e-17<br />Petal.Length: 5.710763","density: 5.252838e-17<br />Petal.Length: 5.699217","density: 5.558608e-17<br />Petal.Length: 5.687671","density: 5.466761e-18<br />Petal.Length: 5.676125","density: 3.231685e-17<br />Petal.Length: 5.664579","density: 3.688931e-18<br />Petal.Length: 5.653033","density: 2.560795e-17<br />Petal.Length: 5.641487","density: 3.102945e-18<br />Petal.Length: 5.629941","density: 0.000000e+00<br />Petal.Length: 5.618395","density: 2.259906e-17<br />Petal.Length: 5.606849","density: 6.422336e-17<br />Petal.Length: 5.595303","density: 3.110603e-17<br />Petal.Length: 5.583757","density: 1.383918e-17<br />Petal.Length: 5.572211","density: 3.631019e-17<br />Petal.Length: 5.560665","density: 5.110874e-17<br />Petal.Length: 5.549119","density: 3.641286e-17<br />Petal.Length: 5.537573","density: 0.000000e+00<br />Petal.Length: 5.526027","density: 7.290582e-18<br />Petal.Length: 5.514481","density: 3.075475e-17<br />Petal.Length: 5.502935","density: 0.000000e+00<br />Petal.Length: 5.491389","density: 0.000000e+00<br />Petal.Length: 5.479843","density: 0.000000e+00<br />Petal.Length: 5.468297","density: 0.000000e+00<br />Petal.Length: 5.456751","density: 0.000000e+00<br />Petal.Length: 5.445205","density: 0.000000e+00<br />Petal.Length: 5.433659","density: 0.000000e+00<br />Petal.Length: 5.422114","density: 0.000000e+00<br />Petal.Length: 5.410568","density: 0.000000e+00<br />Petal.Length: 5.399022","density: 0.000000e+00<br />Petal.Length: 5.387476","density: 0.000000e+00<br />Petal.Length: 5.375930","density: 0.000000e+00<br />Petal.Length: 5.364384","density: 0.000000e+00<br />Petal.Length: 5.352838","density: 0.000000e+00<br />Petal.Length: 5.341292","density: 0.000000e+00<br />Petal.Length: 5.329746","density: 0.000000e+00<br />Petal.Length: 5.318200","density: 0.000000e+00<br />Petal.Length: 5.306654","density: 0.000000e+00<br />Petal.Length: 5.295108","density: 0.000000e+00<br />Petal.Length: 5.283562","density: 0.000000e+00<br />Petal.Length: 5.272016","density: 0.000000e+00<br />Petal.Length: 5.260470","density: 0.000000e+00<br />Petal.Length: 5.248924","density: 0.000000e+00<br />Petal.Length: 5.237378","density: 0.000000e+00<br />Petal.Length: 5.225832","density: 0.000000e+00<br />Petal.Length: 5.214286","density: 0.000000e+00<br />Petal.Length: 5.202740","density: 0.000000e+00<br />Petal.Length: 5.191194","density: 0.000000e+00<br />Petal.Length: 5.179648","density: 0.000000e+00<br />Petal.Length: 5.168102","density: 0.000000e+00<br />Petal.Length: 5.156556","density: 0.000000e+00<br />Petal.Length: 5.145010","density: 0.000000e+00<br />Petal.Length: 5.133464","density: 0.000000e+00<br />Petal.Length: 5.121918","density: 0.000000e+00<br />Petal.Length: 5.110372","density: 0.000000e+00<br />Petal.Length: 5.098826","density: 0.000000e+00<br />Petal.Length: 5.087280","density: 0.000000e+00<br />Petal.Length: 5.075734","density: 0.000000e+00<br />Petal.Length: 5.064188","density: 0.000000e+00<br />Petal.Length: 5.052642","density: 0.000000e+00<br />Petal.Length: 5.041096","density: 0.000000e+00<br />Petal.Length: 5.029550","density: 0.000000e+00<br />Petal.Length: 5.018004","density: 0.000000e+00<br />Petal.Length: 5.006458","density: 0.000000e+00<br />Petal.Length: 4.994912","density: 0.000000e+00<br />Petal.Length: 4.983366","density: 0.000000e+00<br />Petal.Length: 4.971820","density: 0.000000e+00<br />Petal.Length: 4.960274","density: 0.000000e+00<br />Petal.Length: 4.948728","density: 0.000000e+00<br />Petal.Length: 4.937182","density: 2.567970e-17<br />Petal.Length: 4.925636","density: 1.137160e-17<br />Petal.Length: 4.914090","density: 1.677283e-17<br />Petal.Length: 4.902544","density: 3.892110e-17<br />Petal.Length: 4.890998","density: 7.132559e-17<br />Petal.Length: 4.879452","density: 5.823696e-17<br />Petal.Length: 4.867906","density: 0.000000e+00<br />Petal.Length: 4.856360","density: 0.000000e+00<br />Petal.Length: 4.844814","density: 8.190080e-18<br />Petal.Length: 4.833268","density: 4.163336e-17<br />Petal.Length: 4.821722","density: 4.247793e-17<br />Petal.Length: 4.810176","density: 5.541298e-17<br />Petal.Length: 4.798630","density: 5.551115e-17<br />Petal.Length: 4.787084","density: 5.551115e-17<br />Petal.Length: 4.775538","density: 3.360830e-17<br />Petal.Length: 4.763992","density: 7.738189e-18<br />Petal.Length: 4.752446","density: 1.314564e-16<br />Petal.Length: 4.740900","density: 2.286438e-16<br />Petal.Length: 4.729354","density: 2.229337e-16<br />Petal.Length: 4.717808","density: 1.838988e-16<br />Petal.Length: 4.706262","density: 2.078209e-16<br />Petal.Length: 4.694716","density: 2.802759e-16<br />Petal.Length: 4.683170","density: 2.704601e-16<br />Petal.Length: 4.671624","density: 2.143357e-16<br />Petal.Length: 4.660078","density: 1.756283e-16<br />Petal.Length: 4.648532","density: 1.497582e-16<br />Petal.Length: 4.636986","density: 1.238881e-16<br />Petal.Length: 4.625440","density: 1.828905e-16<br />Petal.Length: 4.613894","density: 1.556458e-16<br />Petal.Length: 4.602348","density: 1.131134e-16<br />Petal.Length: 4.590802","density: 9.382137e-17<br />Petal.Length: 4.579256","density: 1.474237e-16<br />Petal.Length: 4.567710","density: 7.823066e-17<br />Petal.Length: 4.556164","density: 1.366169e-16<br />Petal.Length: 4.544618","density: 2.068723e-16<br />Petal.Length: 4.533072","density: 1.179612e-16<br />Petal.Length: 4.521526","density: 9.547105e-17<br />Petal.Length: 4.509980","density: 4.792614e-17<br />Petal.Length: 4.498434","density: 5.947249e-17<br />Petal.Length: 4.486888","density: 1.954813e-17<br />Petal.Length: 4.475342","density: 1.441258e-16<br />Petal.Length: 4.463796","density: 7.945053e-17<br />Petal.Length: 4.452250","density: 1.101818e-16<br />Petal.Length: 4.440705","density: 2.070752e-17<br />Petal.Length: 4.429159","density: 2.215324e-17<br />Petal.Length: 4.417613","density: 9.340365e-18<br />Petal.Length: 4.406067","density: 0.000000e+00<br />Petal.Length: 4.394521","density: 0.000000e+00<br />Petal.Length: 4.382975","density: 0.000000e+00<br />Petal.Length: 4.371429","density: 0.000000e+00<br />Petal.Length: 4.359883","density: 1.549816e-17<br />Petal.Length: 4.348337","density: 3.266853e-17<br />Petal.Length: 4.336791","density: 0.000000e+00<br />Petal.Length: 4.325245","density: 0.000000e+00<br />Petal.Length: 4.313699","density: 0.000000e+00<br />Petal.Length: 4.302153","density: 0.000000e+00<br />Petal.Length: 4.290607","density: 0.000000e+00<br />Petal.Length: 4.279061","density: 0.000000e+00<br />Petal.Length: 4.267515","density: 0.000000e+00<br />Petal.Length: 4.255969","density: 0.000000e+00<br />Petal.Length: 4.244423","density: 0.000000e+00<br />Petal.Length: 4.232877","density: 0.000000e+00<br />Petal.Length: 4.221331","density: 0.000000e+00<br />Petal.Length: 4.209785","density: 0.000000e+00<br />Petal.Length: 4.198239","density: 0.000000e+00<br />Petal.Length: 4.186693","density: 0.000000e+00<br />Petal.Length: 4.175147","density: 0.000000e+00<br />Petal.Length: 4.163601","density: 0.000000e+00<br />Petal.Length: 4.152055","density: 0.000000e+00<br />Petal.Length: 4.140509","density: 0.000000e+00<br />Petal.Length: 4.128963","density: 0.000000e+00<br />Petal.Length: 4.117417","density: 0.000000e+00<br />Petal.Length: 4.105871","density: 0.000000e+00<br />Petal.Length: 4.094325","density: 0.000000e+00<br />Petal.Length: 4.082779","density: 0.000000e+00<br />Petal.Length: 4.071233","density: 0.000000e+00<br />Petal.Length: 4.059687","density: 1.802974e-17<br />Petal.Length: 4.048141","density: 1.531643e-17<br />Petal.Length: 4.036595","density: 0.000000e+00<br />Petal.Length: 4.025049","density: 0.000000e+00<br />Petal.Length: 4.013503","density: 1.962060e-17<br />Petal.Length: 4.001957","density: 4.892430e-17<br />Petal.Length: 3.990411","density: 8.838109e-19<br />Petal.Length: 3.978865","density: 8.739414e-18<br />Petal.Length: 3.967319","density: 4.162964e-17<br />Petal.Length: 3.955773","density: 8.913128e-17<br />Petal.Length: 3.944227","density: 4.477662e-17<br />Petal.Length: 3.932681","density: 1.456665e-17<br />Petal.Length: 3.921135","density: 2.248309e-18<br />Petal.Length: 3.909589","density: 4.584421e-17<br />Petal.Length: 3.898043","density: 5.020507e-17<br />Petal.Length: 3.886497","density: 5.561569e-17<br />Petal.Length: 3.874951","density: 3.431683e-17<br />Petal.Length: 3.863405","density: 6.295653e-17<br />Petal.Length: 3.851859","density: 1.061729e-16<br />Petal.Length: 3.840313","density: 2.017255e-17<br />Petal.Length: 3.828767","density: 0.000000e+00<br />Petal.Length: 3.817221","density: 0.000000e+00<br />Petal.Length: 3.805675","density: 0.000000e+00<br />Petal.Length: 3.794129","density: 0.000000e+00<br />Petal.Length: 3.782583","density: 0.000000e+00<br />Petal.Length: 3.771037","density: 0.000000e+00<br />Petal.Length: 3.759491","density: 1.510460e-17<br />Petal.Length: 3.747945","density: 7.137799e-18<br />Petal.Length: 3.736399","density: 1.029791e-18<br />Petal.Length: 3.724853","density: 0.000000e+00<br />Petal.Length: 3.713307","density: 0.000000e+00<br />Petal.Length: 3.701761","density: 0.000000e+00<br />Petal.Length: 3.690215","density: 0.000000e+00<br />Petal.Length: 3.678669","density: 0.000000e+00<br />Petal.Length: 3.667123","density: 1.118866e-18<br />Petal.Length: 3.655577","density: 3.343603e-18<br />Petal.Length: 3.644031","density: 0.000000e+00<br />Petal.Length: 3.632485","density: 0.000000e+00<br />Petal.Length: 3.620939","density: 0.000000e+00<br />Petal.Length: 3.609393","density: 1.116999e-17<br />Petal.Length: 3.597847","density: 1.682836e-18<br />Petal.Length: 3.586301","density: 3.878572e-19<br />Petal.Length: 3.574755","density: 1.349678e-19<br />Petal.Length: 3.563209","density: 0.000000e+00<br />Petal.Length: 3.551663","density: 0.000000e+00<br />Petal.Length: 3.540117","density: 0.000000e+00<br />Petal.Length: 3.528571","density: 0.000000e+00<br />Petal.Length: 3.517025","density: 0.000000e+00<br />Petal.Length: 3.505479","density: 0.000000e+00<br />Petal.Length: 3.493933","density: 1.531705e-17<br />Petal.Length: 3.482387","density: 5.042987e-17<br />Petal.Length: 3.470841","density: 0.000000e+00<br />Petal.Length: 3.459295","density: 2.733048e-19<br />Petal.Length: 3.447750","density: 5.933813e-18<br />Petal.Length: 3.436204","density: 1.205671e-17<br />Petal.Length: 3.424658","density: 2.397000e-17<br />Petal.Length: 3.413112","density: 5.918761e-18<br />Petal.Length: 3.401566","density: 1.276671e-17<br />Petal.Length: 3.390020","density: 2.037256e-17<br />Petal.Length: 3.378474","density: 9.327228e-18<br />Petal.Length: 3.366928","density: 0.000000e+00<br />Petal.Length: 3.355382","density: 0.000000e+00<br />Petal.Length: 3.343836","density: 0.000000e+00<br />Petal.Length: 3.332290","density: 1.507279e-17<br />Petal.Length: 3.320744","density: 3.898613e-17<br />Petal.Length: 3.309198","density: 0.000000e+00<br />Petal.Length: 3.297652","density: 0.000000e+00<br />Petal.Length: 3.286106","density: 0.000000e+00<br />Petal.Length: 3.274560","density: 0.000000e+00<br />Petal.Length: 3.263014","density: 0.000000e+00<br />Petal.Length: 3.251468","density: 0.000000e+00<br />Petal.Length: 3.239922","density: 0.000000e+00<br />Petal.Length: 3.228376","density: 3.542237e-17<br />Petal.Length: 3.216830","density: 1.969062e-17<br />Petal.Length: 3.205284","density: 0.000000e+00<br />Petal.Length: 3.193738","density: 1.847846e-17<br />Petal.Length: 3.182192","density: 2.948144e-17<br />Petal.Length: 3.170646","density: 3.067547e-17<br />Petal.Length: 3.159100","density: 5.242521e-17<br />Petal.Length: 3.147554","density: 3.335251e-17<br />Petal.Length: 3.136008","density: 2.900847e-17<br />Petal.Length: 3.124462","density: 1.877257e-16<br />Petal.Length: 3.112916","density: 8.842779e-19<br />Petal.Length: 3.101370","density: 1.174008e-16<br />Petal.Length: 3.089824","density: 6.886205e-17<br />Petal.Length: 3.078278","density: 1.711124e-16<br />Petal.Length: 3.066732","density: 1.413327e-16<br />Petal.Length: 3.055186","density: 4.019957e-17<br />Petal.Length: 3.043640","density: 9.811918e-18<br />Petal.Length: 3.032094","density: 4.337827e-17<br />Petal.Length: 3.020548","density: 6.029375e-17<br />Petal.Length: 3.009002","density: 3.161988e-17<br />Petal.Length: 2.997456","density: 1.059118e-17<br />Petal.Length: 2.985910","density: 2.391732e-17<br />Petal.Length: 2.974364","density: 2.859569e-17<br />Petal.Length: 2.962818","density: 1.377336e-16<br />Petal.Length: 2.951272","density: 5.478425e-17<br />Petal.Length: 2.939726","density: 6.901017e-19<br />Petal.Length: 2.928180","density: 3.611912e-17<br />Petal.Length: 2.916634","density: 5.799636e-18<br />Petal.Length: 2.905088","density: 3.370076e-17<br />Petal.Length: 2.893542","density: 1.218645e-17<br />Petal.Length: 2.881996","density: 0.000000e+00<br />Petal.Length: 2.870450","density: 0.000000e+00<br />Petal.Length: 2.858904","density: 0.000000e+00<br />Petal.Length: 2.847358","density: 0.000000e+00<br />Petal.Length: 2.835812","density: 0.000000e+00<br />Petal.Length: 2.824266","density: 0.000000e+00<br />Petal.Length: 2.812720","density: 0.000000e+00<br />Petal.Length: 2.801174","density: 0.000000e+00<br />Petal.Length: 2.789628","density: 0.000000e+00<br />Petal.Length: 2.778082","density: 0.000000e+00<br />Petal.Length: 2.766536","density: 0.000000e+00<br />Petal.Length: 2.754990","density: 0.000000e+00<br />Petal.Length: 2.743444","density: 0.000000e+00<br />Petal.Length: 2.731898","density: 0.000000e+00<br />Petal.Length: 2.720352","density: 0.000000e+00<br />Petal.Length: 2.708806","density: 0.000000e+00<br />Petal.Length: 2.697260","density: 0.000000e+00<br />Petal.Length: 2.685714","density: 0.000000e+00<br />Petal.Length: 2.674168","density: 0.000000e+00<br />Petal.Length: 2.662622","density: 0.000000e+00<br />Petal.Length: 2.651076","density: 0.000000e+00<br />Petal.Length: 2.639530","density: 0.000000e+00<br />Petal.Length: 2.627984","density: 0.000000e+00<br />Petal.Length: 2.616438","density: 0.000000e+00<br />Petal.Length: 2.604892","density: 0.000000e+00<br />Petal.Length: 2.593346","density: 0.000000e+00<br />Petal.Length: 2.581800","density: 0.000000e+00<br />Petal.Length: 2.570254","density: 0.000000e+00<br />Petal.Length: 2.558708","density: 0.000000e+00<br />Petal.Length: 2.547162","density: 0.000000e+00<br />Petal.Length: 2.535616","density: 0.000000e+00<br />Petal.Length: 2.524070","density: 0.000000e+00<br />Petal.Length: 2.512524","density: 0.000000e+00<br />Petal.Length: 2.500978","density: 0.000000e+00<br />Petal.Length: 2.489432","density: 0.000000e+00<br />Petal.Length: 2.477886","density: 0.000000e+00<br />Petal.Length: 2.466341","density: 0.000000e+00<br />Petal.Length: 2.454795","density: 0.000000e+00<br />Petal.Length: 2.443249","density: 0.000000e+00<br />Petal.Length: 2.431703","density: 0.000000e+00<br />Petal.Length: 2.420157","density: 0.000000e+00<br />Petal.Length: 2.408611","density: 0.000000e+00<br />Petal.Length: 2.397065","density: 0.000000e+00<br />Petal.Length: 2.385519","density: 0.000000e+00<br />Petal.Length: 2.373973","density: 0.000000e+00<br />Petal.Length: 2.362427","density: 0.000000e+00<br />Petal.Length: 2.350881","density: 1.117614e-15<br />Petal.Length: 2.339335","density: 8.037517e-15<br />Petal.Length: 2.327789","density: 4.422352e-14<br />Petal.Length: 2.316243","density: 2.231825e-13<br />Petal.Length: 2.304697","density: 1.054006e-12<br />Petal.Length: 2.293151","density: 4.667022e-12<br />Petal.Length: 2.281605","density: 1.936004e-11<br />Petal.Length: 2.270059","density: 7.507452e-11<br />Petal.Length: 2.258513","density: 2.856942e-10<br />Petal.Length: 2.246967","density: 1.160218e-09<br />Petal.Length: 2.235421","density: 4.463893e-09<br />Petal.Length: 2.223875","density: 1.627373e-08<br />Petal.Length: 2.212329","density: 5.622726e-08<br />Petal.Length: 2.200783","density: 1.841666e-07<br />Petal.Length: 2.189237","density: 5.720503e-07<br />Petal.Length: 2.177691","density: 1.685841e-06<br />Petal.Length: 2.166145","density: 4.716401e-06<br />Petal.Length: 2.154599","density: 1.253514e-05<br />Petal.Length: 2.143053","density: 3.167749e-05<br />Petal.Length: 2.131507","density: 7.619470e-05<br />Petal.Length: 2.119961","density: 1.746494e-04<br />Petal.Length: 2.108415","density: 3.819899e-04<br />Petal.Length: 2.096869","density: 7.983449e-04<br />Petal.Length: 2.085323","density: 1.629362e-03<br />Petal.Length: 2.073777","density: 3.214610e-03<br />Petal.Length: 2.062231","density: 6.031607e-03<br />Petal.Length: 2.050685","density: 1.076992e-02<br />Petal.Length: 2.039139","density: 1.831345e-02<br />Petal.Length: 2.027593","density: 2.967724e-02<br />Petal.Length: 2.016047","density: 4.586641e-02<br />Petal.Length: 2.004501","density: 6.765453e-02<br />Petal.Length: 1.992955","density: 9.530534e-02<br />Petal.Length: 1.981409","density: 1.282926e-01<br />Petal.Length: 1.969863","density: 1.650964e-01<br />Petal.Length: 1.958317","density: 2.031629e-01<br />Petal.Length: 1.946771","density: 2.390926e-01<br />Petal.Length: 1.935225","density: 2.690777e-01<br />Petal.Length: 1.923679","density: 2.895382e-01<br />Petal.Length: 1.912133","density: 2.969632e-01<br />Petal.Length: 1.900587","density: 2.911974e-01<br />Petal.Length: 1.889041","density: 2.737161e-01<br />Petal.Length: 1.877495","density: 2.475888e-01<br />Petal.Length: 1.865949","density: 2.171709e-01<br />Petal.Length: 1.854403","density: 1.875065e-01<br />Petal.Length: 1.842857","density: 1.637204e-01<br />Petal.Length: 1.831311","density: 1.504877e-01<br />Petal.Length: 1.819765","density: 1.516145e-01<br />Petal.Length: 1.808219","density: 1.697121e-01<br />Petal.Length: 1.796673","density: 2.059396e-01<br />Petal.Length: 1.785127","density: 2.598118e-01<br />Petal.Length: 1.773581","density: 3.291229e-01<br />Petal.Length: 1.762035","density: 4.100818e-01<br />Petal.Length: 1.750489","density: 4.977711e-01<br />Petal.Length: 1.738943","density: 5.865924e-01<br />Petal.Length: 1.727397","density: 6.720199e-01<br />Petal.Length: 1.715851","density: 7.519887e-01<br />Petal.Length: 1.704305","density: 8.267639e-01<br />Petal.Length: 1.692759","density: 8.985629e-01<br />Petal.Length: 1.681213","density: 9.705843e-01<br />Petal.Length: 1.669667","density: 1.045794e+00<br />Petal.Length: 1.658121","density: 1.125938e+00<br />Petal.Length: 1.646575","density: 1.211178e+00<br />Petal.Length: 1.635029","density: 1.300499e+00<br />Petal.Length: 1.623483","density: 1.392724e+00<br />Petal.Length: 1.611937","density: 1.487639e+00<br />Petal.Length: 1.600391","density: 1.586637e+00<br />Petal.Length: 1.588845","density: 1.692411e+00<br />Petal.Length: 1.577299","density: 1.807885e+00<br />Petal.Length: 1.565753","density: 1.933259e+00<br />Petal.Length: 1.554207","density: 2.063819e+00<br />Petal.Length: 1.542661","density: 2.191634e+00<br />Petal.Length: 1.531115","density: 2.307061e+00<br />Petal.Length: 1.519569","density: 2.401577e+00<br />Petal.Length: 1.508023","density: 2.470391e+00<br />Petal.Length: 1.496477","density: 2.513767e+00<br />Petal.Length: 1.484932","density: 2.536514e+00<br />Petal.Length: 1.473386","density: 2.545763e+00<br />Petal.Length: 1.461840","density: 2.547881e+00<br />Petal.Length: 1.450294","density: 2.545711e+00<br />Petal.Length: 1.438748","density: 2.537257e+00<br />Petal.Length: 1.427202","density: 2.516410e+00<br />Petal.Length: 1.415656","density: 2.475478e+00<br />Petal.Length: 1.404110","density: 2.407122e+00<br />Petal.Length: 1.392564","density: 2.311553e+00<br />Petal.Length: 1.381018","density: 2.194700e+00<br />Petal.Length: 1.369472","density: 2.065108e+00<br />Petal.Length: 1.357926","density: 1.931675e+00<br />Petal.Length: 1.346380","density: 1.801022e+00<br />Petal.Length: 1.334834","density: 1.676053e+00<br />Petal.Length: 1.323288","density: 1.556076e+00<br />Petal.Length: 1.311742","density: 1.438195e+00<br />Petal.Length: 1.300196","density: 1.319299e+00<br />Petal.Length: 1.288650","density: 1.197818e+00<br />Petal.Length: 1.277104","density: 1.074626e+00<br />Petal.Length: 1.265558","density: 9.528557e-01<br />Petal.Length: 1.254012","density: 8.368470e-01<br />Petal.Length: 1.242466","density: 7.307320e-01<br />Petal.Length: 1.230920","density: 6.382756e-01<br />Petal.Length: 1.219374","density: 5.591787e-01<br />Petal.Length: 1.207828","density: 4.923802e-01<br />Petal.Length: 1.196282","density: 4.361654e-01<br />Petal.Length: 1.184736","density: 3.887969e-01<br />Petal.Length: 1.173190","density: 3.488923e-01<br />Petal.Length: 1.161644","density: 3.154872e-01<br />Petal.Length: 1.150098","density: 2.878887e-01<br />Petal.Length: 1.138552","density: 2.654728e-01<br />Petal.Length: 1.127006","density: 2.475547e-01<br />Petal.Length: 1.115460","density: 2.333841e-01<br />Petal.Length: 1.103914","density: 2.222363e-01<br />Petal.Length: 1.092368","density: 2.135162e-01<br />Petal.Length: 1.080822","density: 2.067883e-01<br />Petal.Length: 1.069276","density: 2.017136e-01<br />Petal.Length: 1.057730","density: 1.978000e-01<br />Petal.Length: 1.046184","density: 1.941641e-01<br />Petal.Length: 1.034638","density: 1.897540e-01<br />Petal.Length: 1.023092","density: 1.834256e-01<br />Petal.Length: 1.011546","density: 1.742059e-01<br />Petal.Length: 1.000000","density: 1.742059e-01<br />Petal.Length: 1.000000"],"key":["1","2","3","4","5","6","7","8","9","10","11","12","13","14","15","16","17","18","19","20","21","22","23","24","25","26","27","28","29","30","31","32","33","34","35","36","37","38","39","40","41","42","43","44","45","46","47","48","49","50"],"type":"scatter","mode":"lines","line":{"width":1.88976377952756,"color":"rgba(0,0,0,1)","dash":"solid"},"fill":"toself","fillcolor":"rgba(248,118,109,1)","hoveron":"points","set":"SharedData91833e3a","name":"setosa","legendgroup":"setosa","showlegend":true,"xaxis":"x3","yaxis":"y10","hoverinfo":"text","_isSimpleKey":true,"_isNestedKey":false,"frame":null},{"x":[1,1.01154598825832,1.02309197651663,1.03463796477495,1.04618395303327,1.05772994129159,1.0692759295499,1.08082191780822,1.09236790606654,1.10391389432485,1.11545988258317,1.12700587084149,1.1385518590998,1.15009784735812,1.16164383561644,1.17318982387476,1.18473581213307,1.19628180039139,1.20782778864971,1.21937377690802,1.23091976516634,1.24246575342466,1.25401174168297,1.26555772994129,1.27710371819961,1.28864970645793,1.30019569471624,1.31174168297456,1.32328767123288,1.33483365949119,1.34637964774951,1.35792563600783,1.36947162426614,1.38101761252446,1.39256360078278,1.4041095890411,1.41565557729941,1.42720156555773,1.43874755381605,1.45029354207436,1.46183953033268,1.473385518591,1.48493150684931,1.49647749510763,1.50802348336595,1.51956947162427,1.53111545988258,1.5426614481409,1.55420743639922,1.56575342465753,1.57729941291585,1.58884540117417,1.60039138943249,1.6119373776908,1.62348336594912,1.63502935420744,1.64657534246575,1.65812133072407,1.66966731898239,1.6812133072407,1.69275929549902,1.70430528375734,1.71585127201566,1.72739726027397,1.73894324853229,1.75048923679061,1.76203522504892,1.77358121330724,1.78512720156556,1.79667318982387,1.80821917808219,1.81976516634051,1.83131115459883,1.84285714285714,1.85440313111546,1.86594911937378,1.87749510763209,1.88904109589041,1.90058708414873,1.91213307240705,1.92367906066536,1.93522504892368,1.946771037182,1.95831702544031,1.96986301369863,1.98140900195695,1.99295499021526,2.00450097847358,2.0160469667319,2.02759295499022,2.03913894324853,2.05068493150685,2.06223091976517,2.07377690802348,2.0853228962818,2.09686888454012,2.10841487279843,2.11996086105675,2.13150684931507,2.14305283757339,2.1545988258317,2.16614481409002,2.17769080234834,2.18923679060665,2.20078277886497,2.21232876712329,2.2238747553816,2.23542074363992,2.24696673189824,2.25851272015656,2.27005870841487,2.28160469667319,2.29315068493151,2.30469667318982,2.31624266144814,2.32778864970646,2.33933463796478,2.35088062622309,2.36242661448141,2.37397260273973,2.38551859099804,2.39706457925636,2.40861056751468,2.42015655577299,2.43170254403131,2.44324853228963,2.45479452054795,2.46634050880626,2.47788649706458,2.4894324853229,2.50097847358121,2.51252446183953,2.52407045009785,2.53561643835616,2.54716242661448,2.5587084148728,2.57025440313112,2.58180039138943,2.59334637964775,2.60489236790607,2.61643835616438,2.6279843444227,2.63953033268102,2.65107632093933,2.66262230919765,2.67416829745597,2.68571428571429,2.6972602739726,2.70880626223092,2.72035225048924,2.73189823874755,2.74344422700587,2.75499021526419,2.7665362035225,2.77808219178082,2.78962818003914,2.80117416829746,2.81272015655577,2.82426614481409,2.83581213307241,2.84735812133072,2.85890410958904,2.87045009784736,2.88199608610568,2.89354207436399,2.90508806262231,2.91663405088063,2.92818003913894,2.93972602739726,2.95127201565558,2.96281800391389,2.97436399217221,2.98590998043053,2.99745596868885,3.00900195694716,3.02054794520548,3.0320939334638,3.04363992172211,3.05518590998043,3.06673189823875,3.07827788649706,3.08982387475538,3.1013698630137,3.11291585127202,3.12446183953033,3.13600782778865,3.14755381604697,3.15909980430528,3.1706457925636,3.18219178082192,3.19373776908024,3.20528375733855,3.21682974559687,3.22837573385519,3.2399217221135,3.25146771037182,3.26301369863014,3.27455968688845,3.28610567514677,3.29765166340509,3.30919765166341,3.32074363992172,3.33228962818004,3.34383561643836,3.35538160469667,3.36692759295499,3.37847358121331,3.39001956947162,3.40156555772994,3.41311154598826,3.42465753424658,3.43620352250489,3.44774951076321,3.45929549902153,3.47084148727984,3.48238747553816,3.49393346379648,3.50547945205479,3.51702544031311,3.52857142857143,3.54011741682975,3.55166340508806,3.56320939334638,3.5747553816047,3.58630136986301,3.59784735812133,3.60939334637965,3.62093933463796,3.63248532289628,3.6440313111546,3.65557729941292,3.66712328767123,3.67866927592955,3.69021526418787,3.70176125244618,3.7133072407045,3.72485322896282,3.73639921722114,3.74794520547945,3.75949119373777,3.77103718199609,3.7825831702544,3.79412915851272,3.80567514677104,3.81722113502935,3.82876712328767,3.84031311154599,3.85185909980431,3.86340508806262,3.87495107632094,3.88649706457926,3.89804305283757,3.90958904109589,3.92113502935421,3.93268101761252,3.94422700587084,3.95577299412916,3.96731898238748,3.97886497064579,3.99041095890411,4.00195694716243,4.01350293542074,4.02504892367906,4.03659491193738,4.04814090019569,4.05968688845401,4.07123287671233,4.08277886497065,4.09432485322896,4.10587084148728,4.1174168297456,4.12896281800391,4.14050880626223,4.15205479452055,4.16360078277886,4.17514677103718,4.1866927592955,4.19823874755382,4.20978473581213,4.22133072407045,4.23287671232877,4.24442270058708,4.2559686888454,4.26751467710372,4.27906066536204,4.29060665362035,4.30215264187867,4.31369863013699,4.3252446183953,4.33679060665362,4.34833659491194,4.35988258317025,4.37142857142857,4.38297455968689,4.39452054794521,4.40606653620352,4.41761252446184,4.42915851272016,4.44070450097847,4.45225048923679,4.46379647749511,4.47534246575342,4.48688845401174,4.49843444227006,4.50998043052838,4.52152641878669,4.53307240704501,4.54461839530333,4.55616438356164,4.56771037181996,4.57925636007828,4.5908023483366,4.60234833659491,4.61389432485323,4.62544031311155,4.63698630136986,4.64853228962818,4.6600782778865,4.67162426614481,4.68317025440313,4.69471624266145,4.70626223091977,4.71780821917808,4.7293542074364,4.74090019569472,4.75244618395303,4.76399217221135,4.77553816046967,4.78708414872798,4.7986301369863,4.81017612524462,4.82172211350294,4.83326810176125,4.84481409001957,4.85636007827789,4.8679060665362,4.87945205479452,4.89099804305284,4.90254403131116,4.91409001956947,4.92563600782779,4.93718199608611,4.94872798434442,4.96027397260274,4.97181996086106,4.98336594911937,4.99491193737769,5.00645792563601,5.01800391389432,5.02954990215264,5.04109589041096,5.05264187866928,5.06418786692759,5.07573385518591,5.08727984344423,5.09882583170254,5.11037181996086,5.12191780821918,5.1334637964775,5.14500978473581,5.15655577299413,5.16810176125245,5.17964774951076,5.19119373776908,5.2027397260274,5.21428571428571,5.22583170254403,5.23737769080235,5.24892367906067,5.26046966731898,5.2720156555773,5.28356164383562,5.29510763209393,5.30665362035225,5.31819960861057,5.32974559686888,5.3412915851272,5.35283757338552,5.36438356164384,5.37592954990215,5.38747553816047,5.39902152641879,5.4105675146771,5.42211350293542,5.43365949119374,5.44520547945205,5.45675146771037,5.46829745596869,5.47984344422701,5.49138943248532,5.50293542074364,5.51448140900196,5.52602739726027,5.53757338551859,5.54911937377691,5.56066536203523,5.57221135029354,5.58375733855186,5.59530332681018,5.60684931506849,5.61839530332681,5.62994129158513,5.64148727984344,5.65303326810176,5.66457925636008,5.6761252446184,5.68767123287671,5.69921722113503,5.71076320939335,5.72230919765166,5.73385518590998,5.7454011741683,5.75694716242661,5.76849315068493,5.78003913894325,5.79158512720157,5.80313111545988,5.8146771037182,5.82622309197652,5.83776908023483,5.84931506849315,5.86086105675147,5.87240704500979,5.8839530332681,5.89549902152642,5.90704500978474,5.91859099804305,5.93013698630137,5.94168297455969,5.953228962818,5.96477495107632,5.97632093933464,5.98786692759295,5.99941291585127,6.01095890410959,6.02250489236791,6.03405088062622,6.04559686888454,6.05714285714286,6.06868884540117,6.08023483365949,6.09178082191781,6.10332681017613,6.11487279843444,6.12641878669276,6.13796477495108,6.14951076320939,6.16105675146771,6.17260273972603,6.18414872798434,6.19569471624266,6.20724070450098,6.2187866927593,6.23033268101761,6.24187866927593,6.25342465753425,6.26497064579256,6.27651663405088,6.2880626223092,6.29960861056751,6.31115459882583,6.32270058708415,6.33424657534247,6.34579256360078,6.3573385518591,6.36888454011742,6.38043052837573,6.39197651663405,6.40352250489237,6.41506849315069,6.426614481409,6.43816046966732,6.44970645792564,6.46125244618395,6.47279843444227,6.48434442270059,6.4958904109589,6.50743639921722,6.51898238747554,6.53052837573386,6.54207436399217,6.55362035225049,6.56516634050881,6.57671232876712,6.58825831702544,6.59980430528376,6.61135029354207,6.62289628180039,6.63444227005871,6.64598825831703,6.65753424657534,6.66908023483366,6.68062622309198,6.69217221135029,6.70371819960861,6.71526418786693,6.72681017612524,6.73835616438356,6.74990215264188,6.7614481409002,6.77299412915851,6.78454011741683,6.79608610567515,6.80763209393346,6.81917808219178,6.8307240704501,6.84227005870842,6.85381604696673,6.86536203522505,6.87690802348337,6.88845401174168,6.9,6.9,6.88845401174168,6.87690802348337,6.86536203522505,6.85381604696673,6.84227005870842,6.8307240704501,6.81917808219178,6.80763209393346,6.79608610567515,6.78454011741683,6.77299412915851,6.7614481409002,6.74990215264188,6.73835616438356,6.72681017612524,6.71526418786693,6.70371819960861,6.69217221135029,6.68062622309198,6.66908023483366,6.65753424657534,6.64598825831703,6.63444227005871,6.62289628180039,6.61135029354207,6.59980430528376,6.58825831702544,6.57671232876712,6.56516634050881,6.55362035225049,6.54207436399217,6.53052837573386,6.51898238747554,6.50743639921722,6.4958904109589,6.48434442270059,6.47279843444227,6.46125244618395,6.44970645792564,6.43816046966732,6.426614481409,6.41506849315069,6.40352250489237,6.39197651663405,6.38043052837573,6.36888454011742,6.3573385518591,6.34579256360078,6.33424657534247,6.32270058708415,6.31115459882583,6.29960861056751,6.2880626223092,6.27651663405088,6.26497064579256,6.25342465753425,6.24187866927593,6.23033268101761,6.2187866927593,6.20724070450098,6.19569471624266,6.18414872798434,6.17260273972603,6.16105675146771,6.14951076320939,6.13796477495108,6.12641878669276,6.11487279843444,6.10332681017613,6.09178082191781,6.08023483365949,6.06868884540117,6.05714285714286,6.04559686888454,6.03405088062622,6.02250489236791,6.01095890410959,5.99941291585127,5.98786692759295,5.97632093933464,5.96477495107632,5.953228962818,5.94168297455969,5.93013698630137,5.91859099804305,5.90704500978474,5.89549902152642,5.8839530332681,5.87240704500979,5.86086105675147,5.84931506849315,5.83776908023483,5.82622309197652,5.8146771037182,5.80313111545988,5.79158512720157,5.78003913894325,5.76849315068493,5.75694716242661,5.7454011741683,5.73385518590998,5.72230919765166,5.71076320939335,5.69921722113503,5.68767123287671,5.6761252446184,5.66457925636008,5.65303326810176,5.64148727984344,5.62994129158513,5.61839530332681,5.60684931506849,5.59530332681018,5.58375733855186,5.57221135029354,5.56066536203523,5.54911937377691,5.53757338551859,5.52602739726027,5.51448140900196,5.50293542074364,5.49138943248532,5.47984344422701,5.46829745596869,5.45675146771037,5.44520547945205,5.43365949119374,5.42211350293542,5.4105675146771,5.39902152641879,5.38747553816047,5.37592954990215,5.36438356164384,5.35283757338552,5.3412915851272,5.32974559686888,5.31819960861057,5.30665362035225,5.29510763209393,5.28356164383562,5.2720156555773,5.26046966731898,5.24892367906067,5.23737769080235,5.22583170254403,5.21428571428571,5.2027397260274,5.19119373776908,5.17964774951076,5.16810176125245,5.15655577299413,5.14500978473581,5.1334637964775,5.12191780821918,5.11037181996086,5.09882583170254,5.08727984344423,5.07573385518591,5.06418786692759,5.05264187866928,5.04109589041096,5.02954990215264,5.01800391389432,5.00645792563601,4.99491193737769,4.98336594911937,4.97181996086106,4.96027397260274,4.94872798434442,4.93718199608611,4.92563600782779,4.91409001956947,4.90254403131116,4.89099804305284,4.87945205479452,4.8679060665362,4.85636007827789,4.84481409001957,4.83326810176125,4.82172211350294,4.81017612524462,4.7986301369863,4.78708414872798,4.77553816046967,4.76399217221135,4.75244618395303,4.74090019569472,4.7293542074364,4.71780821917808,4.70626223091977,4.69471624266145,4.68317025440313,4.67162426614481,4.6600782778865,4.64853228962818,4.63698630136986,4.62544031311155,4.61389432485323,4.60234833659491,4.5908023483366,4.57925636007828,4.56771037181996,4.55616438356164,4.54461839530333,4.53307240704501,4.52152641878669,4.50998043052838,4.49843444227006,4.48688845401174,4.47534246575342,4.46379647749511,4.45225048923679,4.44070450097847,4.42915851272016,4.41761252446184,4.40606653620352,4.39452054794521,4.38297455968689,4.37142857142857,4.35988258317025,4.34833659491194,4.33679060665362,4.3252446183953,4.31369863013699,4.30215264187867,4.29060665362035,4.27906066536204,4.26751467710372,4.2559686888454,4.24442270058708,4.23287671232877,4.22133072407045,4.20978473581213,4.19823874755382,4.1866927592955,4.17514677103718,4.16360078277886,4.15205479452055,4.14050880626223,4.12896281800391,4.1174168297456,4.10587084148728,4.09432485322896,4.08277886497065,4.07123287671233,4.05968688845401,4.04814090019569,4.03659491193738,4.02504892367906,4.01350293542074,4.00195694716243,3.99041095890411,3.97886497064579,3.96731898238748,3.95577299412916,3.94422700587084,3.93268101761252,3.92113502935421,3.90958904109589,3.89804305283757,3.88649706457926,3.87495107632094,3.86340508806262,3.85185909980431,3.84031311154599,3.82876712328767,3.81722113502935,3.80567514677104,3.79412915851272,3.7825831702544,3.77103718199609,3.75949119373777,3.74794520547945,3.73639921722114,3.72485322896282,3.7133072407045,3.70176125244618,3.69021526418787,3.67866927592955,3.66712328767123,3.65557729941292,3.6440313111546,3.63248532289628,3.62093933463796,3.60939334637965,3.59784735812133,3.58630136986301,3.5747553816047,3.56320939334638,3.55166340508806,3.54011741682975,3.52857142857143,3.51702544031311,3.50547945205479,3.49393346379648,3.48238747553816,3.47084148727984,3.45929549902153,3.44774951076321,3.43620352250489,3.42465753424658,3.41311154598826,3.40156555772994,3.39001956947162,3.37847358121331,3.36692759295499,3.35538160469667,3.34383561643836,3.33228962818004,3.32074363992172,3.30919765166341,3.29765166340509,3.28610567514677,3.27455968688845,3.26301369863014,3.25146771037182,3.2399217221135,3.22837573385519,3.21682974559687,3.20528375733855,3.19373776908024,3.18219178082192,3.1706457925636,3.15909980430528,3.14755381604697,3.13600782778865,3.12446183953033,3.11291585127202,3.1013698630137,3.08982387475538,3.07827788649706,3.06673189823875,3.05518590998043,3.04363992172211,3.0320939334638,3.02054794520548,3.00900195694716,2.99745596868885,2.98590998043053,2.97436399217221,2.96281800391389,2.95127201565558,2.93972602739726,2.92818003913894,2.91663405088063,2.90508806262231,2.89354207436399,2.88199608610568,2.87045009784736,2.85890410958904,2.84735812133072,2.83581213307241,2.82426614481409,2.81272015655577,2.80117416829746,2.78962818003914,2.77808219178082,2.7665362035225,2.75499021526419,2.74344422700587,2.73189823874755,2.72035225048924,2.70880626223092,2.6972602739726,2.68571428571429,2.67416829745597,2.66262230919765,2.65107632093933,2.63953033268102,2.6279843444227,2.61643835616438,2.60489236790607,2.59334637964775,2.58180039138943,2.57025440313112,2.5587084148728,2.54716242661448,2.53561643835616,2.52407045009785,2.51252446183953,2.50097847358121,2.4894324853229,2.47788649706458,2.46634050880626,2.45479452054795,2.44324853228963,2.43170254403131,2.42015655577299,2.40861056751468,2.39706457925636,2.38551859099804,2.37397260273973,2.36242661448141,2.35088062622309,2.33933463796478,2.32778864970646,2.31624266144814,2.30469667318982,2.29315068493151,2.28160469667319,2.27005870841487,2.25851272015656,2.24696673189824,2.23542074363992,2.2238747553816,2.21232876712329,2.20078277886497,2.18923679060665,2.17769080234834,2.16614481409002,2.1545988258317,2.14305283757339,2.13150684931507,2.11996086105675,2.10841487279843,2.09686888454012,2.0853228962818,2.07377690802348,2.06223091976517,2.05068493150685,2.03913894324853,2.02759295499022,2.0160469667319,2.00450097847358,1.99295499021526,1.98140900195695,1.96986301369863,1.95831702544031,1.946771037182,1.93522504892368,1.92367906066536,1.91213307240705,1.90058708414873,1.88904109589041,1.87749510763209,1.86594911937378,1.85440313111546,1.84285714285714,1.83131115459883,1.81976516634051,1.80821917808219,1.79667318982387,1.78512720156556,1.77358121330724,1.76203522504892,1.75048923679061,1.73894324853229,1.72739726027397,1.71585127201566,1.70430528375734,1.69275929549902,1.6812133072407,1.66966731898239,1.65812133072407,1.64657534246575,1.63502935420744,1.62348336594912,1.6119373776908,1.60039138943249,1.58884540117417,1.57729941291585,1.56575342465753,1.55420743639922,1.5426614481409,1.53111545988258,1.51956947162427,1.50802348336595,1.49647749510763,1.48493150684931,1.473385518591,1.46183953033268,1.45029354207436,1.43874755381605,1.42720156555773,1.41565557729941,1.4041095890411,1.39256360078278,1.38101761252446,1.36947162426614,1.35792563600783,1.34637964774951,1.33483365949119,1.32328767123288,1.31174168297456,1.30019569471624,1.28864970645793,1.27710371819961,1.26555772994129,1.25401174168297,1.24246575342466,1.23091976516634,1.21937377690802,1.20782778864971,1.19628180039139,1.18473581213307,1.17318982387476,1.16164383561644,1.15009784735812,1.1385518590998,1.12700587084149,1.11545988258317,1.10391389432485,1.09236790606654,1.08082191780822,1.0692759295499,1.05772994129159,1.04618395303327,1.03463796477495,1.02309197651663,1.01154598825832,1,1],"y":[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,8.37986955182749e-19,9.16545356210356e-18,1.51313937313039e-17,1.9015136447843e-17,1.5458158550366e-17,1.59439738158592e-18,2.06486280432448e-19,1.53585226693742e-17,1.42052807287195e-17,9.10657099652711e-18,1.75310935769803e-17,2.26407146429217e-18,0,0,0,0,0,4.73099209776783e-18,2.11497999474243e-17,3.00705904231735e-17,1.27393006080369e-17,1.6360117988092e-18,3.96442224811301e-17,6.02047295739208e-17,7.32253124215921e-17,1.06761779185453e-16,1.66441956535057e-16,3.16111547225033e-16,5.54951747356178e-16,9.19985112139771e-16,1.47215227701544e-15,2.41315600359749e-15,4.0436742372532e-15,6.67707674528396e-15,1.07591654098459e-14,1.69010862849017e-14,2.71345018114815e-14,4.40775461711309e-14,7.05751381486467e-14,1.11159514283227e-13,1.71752969588967e-13,2.6986995697391e-13,4.29656965632408e-13,6.74475876751055e-13,1.04235674243797e-12,1.5825073543338e-12,2.43913267854345e-12,3.80304745391847e-12,5.85137304707869e-12,8.87207261053318e-12,1.32329626485229e-11,2.00056897652761e-11,3.05508378247328e-11,4.60731571844476e-11,6.85386143051137e-11,1.00425905896896e-10,1.48917595736062e-10,2.22755653758003e-10,3.29293295409317e-10,4.80620604917069e-10,6.91782987937315e-10,1.0061956195994e-09,1.4744193740831e-09,2.13666716489574e-09,3.05986895412244e-09,4.32622832949382e-09,6.17230846060489e-09,8.86110151356931e-09,1.25892867884527e-08,1.76901382292463e-08,2.45676020758053e-08,3.43830439027757e-08,4.83655638735267e-08,6.73732332398239e-08,9.28973910279046e-08,1.26721643335296e-07,1.73979908616641e-07,2.39827544935756e-07,3.27592214997018e-07,4.43264618226063e-07,5.93913434905449e-07,7.99960627268944e-07,1.0807869099782e-06,1.44780688692094e-06,1.92259539635326e-06,2.53026385773751e-06,3.3438567175336e-06,4.42856740134286e-06,5.81877450701168e-06,7.58403082261421e-06,9.80417419550394e-06,1.27139238253431e-05,1.65091065935355e-05,2.12795642794922e-05,2.72256318208365e-05,3.45740380810819e-05,4.40017465318515e-05,5.60326589953235e-05,7.08662260237488e-05,8.90167892815914e-05,0.000111058085667777,0.000138740649843585,0.000173308636495373,0.000215121759058643,0.000265353569553743,0.000325293764926376,0.000398996033462695,0.000489071240584654,0.000595983351429843,0.000722102773873294,0.000869994499949943,0.00104805416020144,0.00126109281733935,0.00150928221920549,0.00179683895157645,0.00212825218084037,0.00251905396817989,0.00297695706832977,0.00350075348001719,0.00409700163268332,0.00477256386248322,0.00555308010871369,0.0064491502122407,0.00745611595578952,0.00858280500996849,0.00983828061514402,0.0112602956602542,0.0128610190115837,0.0146294008576888,0.0165755632606596,0.0187096773284264,0.0210813906525496,0.0237017562641817,0.0265503361657839,0.0296367670564369,0.0329704852871535,0.0366101005166382,0.0405623872013967,0.0447958489913513,0.0493177577893414,0.0541349940639166,0.059310455237362,0.0648446087356469,0.0706957211841409,0.0768677213802105,0.0833641242701774,0.0902478383213113,0.0975134001407342,0.10511207492603,0.113044915988179,0.121312707948538,0.129976326090167,0.139026853094422,0.148412539769659,0.158132939960077,0.168187506222306,0.178635125881802,0.189465895575431,0.200627524439962,0.212118897013929,0.223938785840596,0.236143737805265,0.248722736044023,0.261622858540594,0.274841261866057,0.288374703636709,0.3022733827038,0.31652142203543,0.331063964809267,0.345893384942915,0.361001192370851,0.376421990584093,0.39212925264549,0.408066758933111,0.424218421723335,0.440566886766255,0.45711813219175,0.473830409751378,0.490651308450843,0.507553985861265,0.524510230452455,0.541484456977551,0.558419328791318,0.575276565695453,0.59201924250186,0.608609403038226,0.624961772161076,0.641008841707709,0.656740144537198,0.672113106107672,0.687084954297532,0.701522018579955,0.71535823279572,0.728624699476095,0.741281546838976,0.753289996132882,0.764482573144471,0.774811644451536,0.784359177184881,0.793099285193708,0.801008654604942,0.80791482103954,0.813807143746387,0.818817309315317,0.82294424914905,0.82619051327678,0.828415862305178,0.829658572204598,0.830082929082245,0.829717288557581,0.828593471043337,0.82663527301335,0.823925504291516,0.820627922746539,0.816793608844678,0.81247540882083,0.807671374163617,0.802484413120322,0.797038431556185,0.791389437768613,0.7855925016057,0.779700452662392,0.773799663672789,0.767945983334097,0.762178663619268,0.756533569674402,0.751076178582671,0.745844365692719,0.740823939001203,0.736022399391075,0.73144290164602,0.727118151949619,0.723026177455073,0.719109766886533,0.715343891796603,0.711700175104315,0.708150821251161,0.704628816019663,0.701077973688646,0.69745443406555,0.693713440151082,0.689768511199799,0.685541360579701,0.681017726815684,0.676156507338343,0.670918472638656,0.665187847368834,0.65891250155932,0.652139751224769,0.644850990604584,0.637031359641817,0.62857891790135,0.619492713864488,0.609871991793875,0.59973003269104,0.589084056205627,0.577883626957699,0.566181398411147,0.554094562451508,0.541663618229095,0.528931501281531,0.515915480557803,0.502701598466945,0.489382299396535,0.476009322301053,0.462634415451909,0.44933185054561,0.436188455640096,0.423239653006876,0.410529076939681,0.398098190449598,0.386048094059904,0.374436919502412,0.363234607208072,0.352462967079961,0.342140644374417,0.332361463276639,0.323141021722841,0.314400295833499,0.306136387146641,0.298343650892336,0.291085233210027,0.284339162573412,0.278008327041,0.27207214261997,0.266508602392875,0.261343854975358,0.256535632979223,0.251998672966334,0.247706159417971,0.243631276316204,0.239770889804257,0.236081445248006,0.232507304580584,0.229025353530396,0.225613417257051,0.222252982540315,0.218912160994385,0.215567981335425,0.212206368241587,0.20881451639927,0.205371842896479,0.20186313677114,0.198290697735005,0.19465045338048,0.190939492934702,0.187144706445977,0.183266693470073,0.179321223037937,0.175312371061645,0.171245007666077,0.167118363729001,0.162944371346992,0.158740294065185,0.154514786097403,0.150276762064732,0.146037430025351,0.141813114185195,0.137613032595502,0.133445891557921,0.129320087653234,0.125253415174306,0.121258843062835,0.117332874734284,0.113480256105322,0.109705064921938,0.106023895071499,0.102441122708868,0.098942431325741,0.0955271379017672,0.0921939569438205,0.0889527494803037,0.0857996178526983,0.0827165810276548,0.0796994846162687,0.0767440064244806,0.0738533899454609,0.0710209574202162,0.0682327361725264,0.0654849810176511,0.0627742522234669,0.0601021416240858,0.0574656766231069,0.0548580110325809,0.0522787186931034,0.0497278535327777,0.0472108975269504,0.0447314323139674,0.0422868844362454,0.039880198015014,0.0375146024668214,0.0352015529180266,0.0329490563760875,0.0307530588311238,0.028617563708329,0.0265465042732151,0.0245551522246641,0.0226512813655215,0.0208252799894179,0.0190795952906943,0.0174163575151763,0.0158505311976315,0.0143860365633543,0.013007714503305,0.011715260181145,0.0105080328587664,0.0093974555668436,0.00838285071152809,0.00744671672365149,0.0065864994382628,0.00579945425772126,0.00509246813596356,0.00446183677451031,0.00389152972912462,0.00337809395080882,0.0029180600502874,0.00251460944572213,0.00216321537276343,0.0018516985363816,0.00157691087853383,0.00133579711406485,0.00112932645805717,0.000953692407197246,0.000801017392850044,0.000669039351189991,0.000555616880788488,0.0004607628358492,0.000381934022717507,0.000314721646330514,0.000257771828380757,0.000209828968089994,0.000170664509412043,0.000138856797898139,0.000112247627637969,9.01436193486023e-05,7.1913296930391e-05,5.73634576056984e-05,4.58121259838416e-05,3.63282613806524e-05,2.86032547208454e-05,2.2360701551007e-05,1.74922077338682e-05,1.37129067600766e-05,1.06669247353036e-05,8.23371014390246e-06,6.30697568069635e-06,4.83844324320379e-06,3.72353273013715e-06,2.8412343001724e-06,2.14994011299885e-06,1.61351318946906e-06,1.21388959472754e-06,9.17111203288702e-07,6.86460585430452e-07,5.09188108430237e-07,3.74380432702491e-07,2.76211552100925e-07,2.04884402921193e-07,1.50433783147185e-07,1.09378851482551e-07,7.87820305614618e-08,5.70006356620971e-08,4.15148413254578e-08,2.99009240326172e-08,2.13098708356779e-08,1.50350568600021e-08,1.06680385756437e-08,7.62953589967751e-09,5.39046481555925e-09,3.76542482589078e-09,2.60220266281471e-09,1.81072169773647e-09,1.27171231777156e-09,8.8138550308469e-10,6.03432558850525e-10,4.0844336696653e-10,2.78726624954616e-10,1.92253135868893e-10,1.3070773070433e-10,8.77046834216271e-11,5.81400760886807e-11,3.8910225792229e-11,2.63603599164086e-11,1.75805011767183e-11,1.15609715864598e-11,7.50536942685931e-12,4.92616938070824e-12,3.27810160594106e-12,2.14465023398257e-12,1.38212149839274e-12,8.78651929897356e-13,5.65606846143299e-13,3.69738872479223e-13,2.37280873151091e-13,1.49840585658858e-13,9.32903657678029e-14,5.89499753475121e-14,3.78665463854715e-14,2.38559501578006e-14,1.4780303203672e-14,9.01274070016622e-15,5.59084588027224e-15,3.53971123175922e-15,2.19635678791565e-15,1.3422755571173e-15,8.12011236601116e-16,4.90015861133489e-16,3.02188042388852e-16,1.95457092490912e-16,1.25222293774315e-16,5.98638400648311e-17,1.8226507030225e-17,2.25784643639934e-17,1.616329653157e-17,4.74157772753408e-18,1.01258334635009e-17,7.95213885211765e-17,7.92744814252388e-17,9.06766812754114e-17,8.20837827204607e-17,4.52597396183182e-17,1.17431116878302e-16,1.02503104316038e-16,1.00854784683847e-16,8.88306790848817e-17,5.50951332630572e-17,9.95082884999704e-17,8.41643909694709e-17,8.53090951879575e-17,8.15461630099079e-17,5.71471630300679e-17,6.8250451839296e-17,4.44769350154422e-17,4.1884772130402e-17,5.51959627457699e-17,7.59541574938457e-17,1.20367312730759e-16,9.62360688003662e-17,1.03889888315455e-16,1.08167354036888e-16,8.47370721806629e-17,1.18046938608348e-16,1.18936796508572e-16,1.18861307396502e-16,1.09736544047796e-16,8.59930619096271e-17,6.93381286957846e-17,8.76155756712995e-17,9.45090185467849e-17,7.0962697160983e-17,3.07337465382686e-17,8.55562350338333e-17,0],"text":["density: 8.555624e-17<br />Petal.Length: 1.000000","density: 3.073375e-17<br />Petal.Length: 1.011546","density: 7.096270e-17<br />Petal.Length: 1.023092","density: 9.450902e-17<br />Petal.Length: 1.034638","density: 8.761558e-17<br />Petal.Length: 1.046184","density: 6.933813e-17<br />Petal.Length: 1.057730","density: 8.599306e-17<br />Petal.Length: 1.069276","density: 1.097365e-16<br />Petal.Length: 1.080822","density: 1.188613e-16<br />Petal.Length: 1.092368","density: 1.189368e-16<br />Petal.Length: 1.103914","density: 1.180469e-16<br />Petal.Length: 1.115460","density: 8.473707e-17<br />Petal.Length: 1.127006","density: 1.081674e-16<br />Petal.Length: 1.138552","density: 1.038899e-16<br />Petal.Length: 1.150098","density: 9.623607e-17<br />Petal.Length: 1.161644","density: 1.203673e-16<br />Petal.Length: 1.173190","density: 7.595416e-17<br />Petal.Length: 1.184736","density: 5.519596e-17<br />Petal.Length: 1.196282","density: 4.188477e-17<br />Petal.Length: 1.207828","density: 4.447694e-17<br />Petal.Length: 1.219374","density: 6.825045e-17<br />Petal.Length: 1.230920","density: 5.714716e-17<br />Petal.Length: 1.242466","density: 8.154616e-17<br />Petal.Length: 1.254012","density: 8.530910e-17<br />Petal.Length: 1.265558","density: 8.416439e-17<br />Petal.Length: 1.277104","density: 9.950829e-17<br />Petal.Length: 1.288650","density: 5.509513e-17<br />Petal.Length: 1.300196","density: 8.883068e-17<br />Petal.Length: 1.311742","density: 1.008548e-16<br />Petal.Length: 1.323288","density: 1.025031e-16<br />Petal.Length: 1.334834","density: 1.174311e-16<br />Petal.Length: 1.346380","density: 4.525974e-17<br />Petal.Length: 1.357926","density: 8.208378e-17<br />Petal.Length: 1.369472","density: 9.067668e-17<br />Petal.Length: 1.381018","density: 7.927448e-17<br />Petal.Length: 1.392564","density: 7.952139e-17<br />Petal.Length: 1.404110","density: 1.012583e-17<br />Petal.Length: 1.415656","density: 4.741578e-18<br />Petal.Length: 1.427202","density: 1.616330e-17<br />Petal.Length: 1.438748","density: 2.257846e-17<br />Petal.Length: 1.450294","density: 1.822651e-17<br />Petal.Length: 1.461840","density: 5.986384e-17<br />Petal.Length: 1.473386","density: 1.252223e-16<br />Petal.Length: 1.484932","density: 1.954571e-16<br />Petal.Length: 1.496477","density: 3.021880e-16<br />Petal.Length: 1.508023","density: 4.900159e-16<br />Petal.Length: 1.519569","density: 8.120112e-16<br />Petal.Length: 1.531115","density: 1.342276e-15<br />Petal.Length: 1.542661","density: 2.196357e-15<br />Petal.Length: 1.554207","density: 3.539711e-15<br />Petal.Length: 1.565753","density: 5.590846e-15<br />Petal.Length: 1.577299","density: 9.012741e-15<br />Petal.Length: 1.588845","density: 1.478030e-14<br />Petal.Length: 1.600391","density: 2.385595e-14<br />Petal.Length: 1.611937","density: 3.786655e-14<br />Petal.Length: 1.623483","density: 5.894998e-14<br />Petal.Length: 1.635029","density: 9.329037e-14<br />Petal.Length: 1.646575","density: 1.498406e-13<br />Petal.Length: 1.658121","density: 2.372809e-13<br />Petal.Length: 1.669667","density: 3.697389e-13<br />Petal.Length: 1.681213","density: 5.656068e-13<br />Petal.Length: 1.692759","density: 8.786519e-13<br />Petal.Length: 1.704305","density: 1.382121e-12<br />Petal.Length: 1.715851","density: 2.144650e-12<br />Petal.Length: 1.727397","density: 3.278102e-12<br />Petal.Length: 1.738943","density: 4.926169e-12<br />Petal.Length: 1.750489","density: 7.505369e-12<br />Petal.Length: 1.762035","density: 1.156097e-11<br />Petal.Length: 1.773581","density: 1.758050e-11<br />Petal.Length: 1.785127","density: 2.636036e-11<br />Petal.Length: 1.796673","density: 3.891023e-11<br />Petal.Length: 1.808219","density: 5.814008e-11<br />Petal.Length: 1.819765","density: 8.770468e-11<br />Petal.Length: 1.831311","density: 1.307077e-10<br />Petal.Length: 1.842857","density: 1.922531e-10<br />Petal.Length: 1.854403","density: 2.787266e-10<br />Petal.Length: 1.865949","density: 4.084434e-10<br />Petal.Length: 1.877495","density: 6.034326e-10<br />Petal.Length: 1.889041","density: 8.813855e-10<br />Petal.Length: 1.900587","density: 1.271712e-09<br />Petal.Length: 1.912133","density: 1.810722e-09<br />Petal.Length: 1.923679","density: 2.602203e-09<br />Petal.Length: 1.935225","density: 3.765425e-09<br />Petal.Length: 1.946771","density: 5.390465e-09<br />Petal.Length: 1.958317","density: 7.629536e-09<br />Petal.Length: 1.969863","density: 1.066804e-08<br />Petal.Length: 1.981409","density: 1.503506e-08<br />Petal.Length: 1.992955","density: 2.130987e-08<br />Petal.Length: 2.004501","density: 2.990092e-08<br />Petal.Length: 2.016047","density: 4.151484e-08<br />Petal.Length: 2.027593","density: 5.700064e-08<br />Petal.Length: 2.039139","density: 7.878203e-08<br />Petal.Length: 2.050685","density: 1.093789e-07<br />Petal.Length: 2.062231","density: 1.504338e-07<br />Petal.Length: 2.073777","density: 2.048844e-07<br />Petal.Length: 2.085323","density: 2.762116e-07<br />Petal.Length: 2.096869","density: 3.743804e-07<br />Petal.Length: 2.108415","density: 5.091881e-07<br />Petal.Length: 2.119961","density: 6.864606e-07<br />Petal.Length: 2.131507","density: 9.171112e-07<br />Petal.Length: 2.143053","density: 1.213890e-06<br />Petal.Length: 2.154599","density: 1.613513e-06<br />Petal.Length: 2.166145","density: 2.149940e-06<br />Petal.Length: 2.177691","density: 2.841234e-06<br />Petal.Length: 2.189237","density: 3.723533e-06<br />Petal.Length: 2.200783","density: 4.838443e-06<br />Petal.Length: 2.212329","density: 6.306976e-06<br />Petal.Length: 2.223875","density: 8.233710e-06<br />Petal.Length: 2.235421","density: 1.066692e-05<br />Petal.Length: 2.246967","density: 1.371291e-05<br />Petal.Length: 2.258513","density: 1.749221e-05<br />Petal.Length: 2.270059","density: 2.236070e-05<br />Petal.Length: 2.281605","density: 2.860325e-05<br />Petal.Length: 2.293151","density: 3.632826e-05<br />Petal.Length: 2.304697","density: 4.581213e-05<br />Petal.Length: 2.316243","density: 5.736346e-05<br />Petal.Length: 2.327789","density: 7.191330e-05<br />Petal.Length: 2.339335","density: 9.014362e-05<br />Petal.Length: 2.350881","density: 1.122476e-04<br />Petal.Length: 2.362427","density: 1.388568e-04<br />Petal.Length: 2.373973","density: 1.706645e-04<br />Petal.Length: 2.385519","density: 2.098290e-04<br />Petal.Length: 2.397065","density: 2.577718e-04<br />Petal.Length: 2.408611","density: 3.147216e-04<br />Petal.Length: 2.420157","density: 3.819340e-04<br />Petal.Length: 2.431703","density: 4.607628e-04<br />Petal.Length: 2.443249","density: 5.556169e-04<br />Petal.Length: 2.454795","density: 6.690394e-04<br />Petal.Length: 2.466341","density: 8.010174e-04<br />Petal.Length: 2.477886","density: 9.536924e-04<br />Petal.Length: 2.489432","density: 1.129326e-03<br />Petal.Length: 2.500978","density: 1.335797e-03<br />Petal.Length: 2.512524","density: 1.576911e-03<br />Petal.Length: 2.524070","density: 1.851699e-03<br />Petal.Length: 2.535616","density: 2.163215e-03<br />Petal.Length: 2.547162","density: 2.514609e-03<br />Petal.Length: 2.558708","density: 2.918060e-03<br />Petal.Length: 2.570254","density: 3.378094e-03<br />Petal.Length: 2.581800","density: 3.891530e-03<br />Petal.Length: 2.593346","density: 4.461837e-03<br />Petal.Length: 2.604892","density: 5.092468e-03<br />Petal.Length: 2.616438","density: 5.799454e-03<br />Petal.Length: 2.627984","density: 6.586499e-03<br />Petal.Length: 2.639530","density: 7.446717e-03<br />Petal.Length: 2.651076","density: 8.382851e-03<br />Petal.Length: 2.662622","density: 9.397456e-03<br />Petal.Length: 2.674168","density: 1.050803e-02<br />Petal.Length: 2.685714","density: 1.171526e-02<br />Petal.Length: 2.697260","density: 1.300771e-02<br />Petal.Length: 2.708806","density: 1.438604e-02<br />Petal.Length: 2.720352","density: 1.585053e-02<br />Petal.Length: 2.731898","density: 1.741636e-02<br />Petal.Length: 2.743444","density: 1.907960e-02<br />Petal.Length: 2.754990","density: 2.082528e-02<br />Petal.Length: 2.766536","density: 2.265128e-02<br />Petal.Length: 2.778082","density: 2.455515e-02<br />Petal.Length: 2.789628","density: 2.654650e-02<br />Petal.Length: 2.801174","density: 2.861756e-02<br />Petal.Length: 2.812720","density: 3.075306e-02<br />Petal.Length: 2.824266","density: 3.294906e-02<br />Petal.Length: 2.835812","density: 3.520155e-02<br />Petal.Length: 2.847358","density: 3.751460e-02<br />Petal.Length: 2.858904","density: 3.988020e-02<br />Petal.Length: 2.870450","density: 4.228688e-02<br />Petal.Length: 2.881996","density: 4.473143e-02<br />Petal.Length: 2.893542","density: 4.721090e-02<br />Petal.Length: 2.905088","density: 4.972785e-02<br />Petal.Length: 2.916634","density: 5.227872e-02<br />Petal.Length: 2.928180","density: 5.485801e-02<br />Petal.Length: 2.939726","density: 5.746568e-02<br />Petal.Length: 2.951272","density: 6.010214e-02<br />Petal.Length: 2.962818","density: 6.277425e-02<br />Petal.Length: 2.974364","density: 6.548498e-02<br />Petal.Length: 2.985910","density: 6.823274e-02<br />Petal.Length: 2.997456","density: 7.102096e-02<br />Petal.Length: 3.009002","density: 7.385339e-02<br />Petal.Length: 3.020548","density: 7.674401e-02<br />Petal.Length: 3.032094","density: 7.969948e-02<br />Petal.Length: 3.043640","density: 8.271658e-02<br />Petal.Length: 3.055186","density: 8.579962e-02<br />Petal.Length: 3.066732","density: 8.895275e-02<br />Petal.Length: 3.078278","density: 9.219396e-02<br />Petal.Length: 3.089824","density: 9.552714e-02<br />Petal.Length: 3.101370","density: 9.894243e-02<br />Petal.Length: 3.112916","density: 1.024411e-01<br />Petal.Length: 3.124462","density: 1.060239e-01<br />Petal.Length: 3.136008","density: 1.097051e-01<br />Petal.Length: 3.147554","density: 1.134803e-01<br />Petal.Length: 3.159100","density: 1.173329e-01<br />Petal.Length: 3.170646","density: 1.212588e-01<br />Petal.Length: 3.182192","density: 1.252534e-01<br />Petal.Length: 3.193738","density: 1.293201e-01<br />Petal.Length: 3.205284","density: 1.334459e-01<br />Petal.Length: 3.216830","density: 1.376130e-01<br />Petal.Length: 3.228376","density: 1.418131e-01<br />Petal.Length: 3.239922","density: 1.460374e-01<br />Petal.Length: 3.251468","density: 1.502768e-01<br />Petal.Length: 3.263014","density: 1.545148e-01<br />Petal.Length: 3.274560","density: 1.587403e-01<br />Petal.Length: 3.286106","density: 1.629444e-01<br />Petal.Length: 3.297652","density: 1.671184e-01<br />Petal.Length: 3.309198","density: 1.712450e-01<br />Petal.Length: 3.320744","density: 1.753124e-01<br />Petal.Length: 3.332290","density: 1.793212e-01<br />Petal.Length: 3.343836","density: 1.832667e-01<br />Petal.Length: 3.355382","density: 1.871447e-01<br />Petal.Length: 3.366928","density: 1.909395e-01<br />Petal.Length: 3.378474","density: 1.946505e-01<br />Petal.Length: 3.390020","density: 1.982907e-01<br />Petal.Length: 3.401566","density: 2.018631e-01<br />Petal.Length: 3.413112","density: 2.053718e-01<br />Petal.Length: 3.424658","density: 2.088145e-01<br />Petal.Length: 3.436204","density: 2.122064e-01<br />Petal.Length: 3.447750","density: 2.155680e-01<br />Petal.Length: 3.459295","density: 2.189122e-01<br />Petal.Length: 3.470841","density: 2.222530e-01<br />Petal.Length: 3.482387","density: 2.256134e-01<br />Petal.Length: 3.493933","density: 2.290254e-01<br />Petal.Length: 3.505479","density: 2.325073e-01<br />Petal.Length: 3.517025","density: 2.360814e-01<br />Petal.Length: 3.528571","density: 2.397709e-01<br />Petal.Length: 3.540117","density: 2.436313e-01<br />Petal.Length: 3.551663","density: 2.477062e-01<br />Petal.Length: 3.563209","density: 2.519987e-01<br />Petal.Length: 3.574755","density: 2.565356e-01<br />Petal.Length: 3.586301","density: 2.613439e-01<br />Petal.Length: 3.597847","density: 2.665086e-01<br />Petal.Length: 3.609393","density: 2.720721e-01<br />Petal.Length: 3.620939","density: 2.780083e-01<br />Petal.Length: 3.632485","density: 2.843392e-01<br />Petal.Length: 3.644031","density: 2.910852e-01<br />Petal.Length: 3.655577","density: 2.983437e-01<br />Petal.Length: 3.667123","density: 3.061364e-01<br />Petal.Length: 3.678669","density: 3.144003e-01<br />Petal.Length: 3.690215","density: 3.231410e-01<br />Petal.Length: 3.701761","density: 3.323615e-01<br />Petal.Length: 3.713307","density: 3.421406e-01<br />Petal.Length: 3.724853","density: 3.524630e-01<br />Petal.Length: 3.736399","density: 3.632346e-01<br />Petal.Length: 3.747945","density: 3.744369e-01<br />Petal.Length: 3.759491","density: 3.860481e-01<br />Petal.Length: 3.771037","density: 3.980982e-01<br />Petal.Length: 3.782583","density: 4.105291e-01<br />Petal.Length: 3.794129","density: 4.232397e-01<br />Petal.Length: 3.805675","density: 4.361885e-01<br />Petal.Length: 3.817221","density: 4.493319e-01<br />Petal.Length: 3.828767","density: 4.626344e-01<br />Petal.Length: 3.840313","density: 4.760093e-01<br />Petal.Length: 3.851859","density: 4.893823e-01<br />Petal.Length: 3.863405","density: 5.027016e-01<br />Petal.Length: 3.874951","density: 5.159155e-01<br />Petal.Length: 3.886497","density: 5.289315e-01<br />Petal.Length: 3.898043","density: 5.416636e-01<br />Petal.Length: 3.909589","density: 5.540946e-01<br />Petal.Length: 3.921135","density: 5.661814e-01<br />Petal.Length: 3.932681","density: 5.778836e-01<br />Petal.Length: 3.944227","density: 5.890841e-01<br />Petal.Length: 3.955773","density: 5.997300e-01<br />Petal.Length: 3.967319","density: 6.098720e-01<br />Petal.Length: 3.978865","density: 6.194927e-01<br />Petal.Length: 3.990411","density: 6.285789e-01<br />Petal.Length: 4.001957","density: 6.370314e-01<br />Petal.Length: 4.013503","density: 6.448510e-01<br />Petal.Length: 4.025049","density: 6.521398e-01<br />Petal.Length: 4.036595","density: 6.589125e-01<br />Petal.Length: 4.048141","density: 6.651878e-01<br />Petal.Length: 4.059687","density: 6.709185e-01<br />Petal.Length: 4.071233","density: 6.761565e-01<br />Petal.Length: 4.082779","density: 6.810177e-01<br />Petal.Length: 4.094325","density: 6.855414e-01<br />Petal.Length: 4.105871","density: 6.897685e-01<br />Petal.Length: 4.117417","density: 6.937134e-01<br />Petal.Length: 4.128963","density: 6.974544e-01<br />Petal.Length: 4.140509","density: 7.010780e-01<br />Petal.Length: 4.152055","density: 7.046288e-01<br />Petal.Length: 4.163601","density: 7.081508e-01<br />Petal.Length: 4.175147","density: 7.117002e-01<br />Petal.Length: 4.186693","density: 7.153439e-01<br />Petal.Length: 4.198239","density: 7.191098e-01<br />Petal.Length: 4.209785","density: 7.230262e-01<br />Petal.Length: 4.221331","density: 7.271182e-01<br />Petal.Length: 4.232877","density: 7.314429e-01<br />Petal.Length: 4.244423","density: 7.360224e-01<br />Petal.Length: 4.255969","density: 7.408239e-01<br />Petal.Length: 4.267515","density: 7.458444e-01<br />Petal.Length: 4.279061","density: 7.510762e-01<br />Petal.Length: 4.290607","density: 7.565336e-01<br />Petal.Length: 4.302153","density: 7.621787e-01<br />Petal.Length: 4.313699","density: 7.679460e-01<br />Petal.Length: 4.325245","density: 7.737997e-01<br />Petal.Length: 4.336791","density: 7.797005e-01<br />Petal.Length: 4.348337","density: 7.855925e-01<br />Petal.Length: 4.359883","density: 7.913894e-01<br />Petal.Length: 4.371429","density: 7.970384e-01<br />Petal.Length: 4.382975","density: 8.024844e-01<br />Petal.Length: 4.394521","density: 8.076714e-01<br />Petal.Length: 4.406067","density: 8.124754e-01<br />Petal.Length: 4.417613","density: 8.167936e-01<br />Petal.Length: 4.429159","density: 8.206279e-01<br />Petal.Length: 4.440705","density: 8.239255e-01<br />Petal.Length: 4.452250","density: 8.266353e-01<br />Petal.Length: 4.463796","density: 8.285935e-01<br />Petal.Length: 4.475342","density: 8.297173e-01<br />Petal.Length: 4.486888","density: 8.300829e-01<br />Petal.Length: 4.498434","density: 8.296586e-01<br />Petal.Length: 4.509980","density: 8.284159e-01<br />Petal.Length: 4.521526","density: 8.261905e-01<br />Petal.Length: 4.533072","density: 8.229442e-01<br />Petal.Length: 4.544618","density: 8.188173e-01<br />Petal.Length: 4.556164","density: 8.138071e-01<br />Petal.Length: 4.567710","density: 8.079148e-01<br />Petal.Length: 4.579256","density: 8.010087e-01<br />Petal.Length: 4.590802","density: 7.930993e-01<br />Petal.Length: 4.602348","density: 7.843592e-01<br />Petal.Length: 4.613894","density: 7.748116e-01<br />Petal.Length: 4.625440","density: 7.644826e-01<br />Petal.Length: 4.636986","density: 7.532900e-01<br />Petal.Length: 4.648532","density: 7.412815e-01<br />Petal.Length: 4.660078","density: 7.286247e-01<br />Petal.Length: 4.671624","density: 7.153582e-01<br />Petal.Length: 4.683170","density: 7.015220e-01<br />Petal.Length: 4.694716","density: 6.870850e-01<br />Petal.Length: 4.706262","density: 6.721131e-01<br />Petal.Length: 4.717808","density: 6.567401e-01<br />Petal.Length: 4.729354","density: 6.410088e-01<br />Petal.Length: 4.740900","density: 6.249618e-01<br />Petal.Length: 4.752446","density: 6.086094e-01<br />Petal.Length: 4.763992","density: 5.920192e-01<br />Petal.Length: 4.775538","density: 5.752766e-01<br />Petal.Length: 4.787084","density: 5.584193e-01<br />Petal.Length: 4.798630","density: 5.414845e-01<br />Petal.Length: 4.810176","density: 5.245102e-01<br />Petal.Length: 4.821722","density: 5.075540e-01<br />Petal.Length: 4.833268","density: 4.906513e-01<br />Petal.Length: 4.844814","density: 4.738304e-01<br />Petal.Length: 4.856360","density: 4.571181e-01<br />Petal.Length: 4.867906","density: 4.405669e-01<br />Petal.Length: 4.879452","density: 4.242184e-01<br />Petal.Length: 4.890998","density: 4.080668e-01<br />Petal.Length: 4.902544","density: 3.921293e-01<br />Petal.Length: 4.914090","density: 3.764220e-01<br />Petal.Length: 4.925636","density: 3.610012e-01<br />Petal.Length: 4.937182","density: 3.458934e-01<br />Petal.Length: 4.948728","density: 3.310640e-01<br />Petal.Length: 4.960274","density: 3.165214e-01<br />Petal.Length: 4.971820","density: 3.022734e-01<br />Petal.Length: 4.983366","density: 2.883747e-01<br />Petal.Length: 4.994912","density: 2.748413e-01<br />Petal.Length: 5.006458","density: 2.616229e-01<br />Petal.Length: 5.018004","density: 2.487227e-01<br />Petal.Length: 5.029550","density: 2.361437e-01<br />Petal.Length: 5.041096","density: 2.239388e-01<br />Petal.Length: 5.052642","density: 2.121189e-01<br />Petal.Length: 5.064188","density: 2.006275e-01<br />Petal.Length: 5.075734","density: 1.894659e-01<br />Petal.Length: 5.087280","density: 1.786351e-01<br />Petal.Length: 5.098826","density: 1.681875e-01<br />Petal.Length: 5.110372","density: 1.581329e-01<br />Petal.Length: 5.121918","density: 1.484125e-01<br />Petal.Length: 5.133464","density: 1.390269e-01<br />Petal.Length: 5.145010","density: 1.299763e-01<br />Petal.Length: 5.156556","density: 1.213127e-01<br />Petal.Length: 5.168102","density: 1.130449e-01<br />Petal.Length: 5.179648","density: 1.051121e-01<br />Petal.Length: 5.191194","density: 9.751340e-02<br />Petal.Length: 5.202740","density: 9.024784e-02<br />Petal.Length: 5.214286","density: 8.336412e-02<br />Petal.Length: 5.225832","density: 7.686772e-02<br />Petal.Length: 5.237378","density: 7.069572e-02<br />Petal.Length: 5.248924","density: 6.484461e-02<br />Petal.Length: 5.260470","density: 5.931046e-02<br />Petal.Length: 5.272016","density: 5.413499e-02<br />Petal.Length: 5.283562","density: 4.931776e-02<br />Petal.Length: 5.295108","density: 4.479585e-02<br />Petal.Length: 5.306654","density: 4.056239e-02<br />Petal.Length: 5.318200","density: 3.661010e-02<br />Petal.Length: 5.329746","density: 3.297049e-02<br />Petal.Length: 5.341292","density: 2.963677e-02<br />Petal.Length: 5.352838","density: 2.655034e-02<br />Petal.Length: 5.364384","density: 2.370176e-02<br />Petal.Length: 5.375930","density: 2.108139e-02<br />Petal.Length: 5.387476","density: 1.870968e-02<br />Petal.Length: 5.399022","density: 1.657556e-02<br />Petal.Length: 5.410568","density: 1.462940e-02<br />Petal.Length: 5.422114","density: 1.286102e-02<br />Petal.Length: 5.433659","density: 1.126030e-02<br />Petal.Length: 5.445205","density: 9.838281e-03<br />Petal.Length: 5.456751","density: 8.582805e-03<br />Petal.Length: 5.468297","density: 7.456116e-03<br />Petal.Length: 5.479843","density: 6.449150e-03<br />Petal.Length: 5.491389","density: 5.553080e-03<br />Petal.Length: 5.502935","density: 4.772564e-03<br />Petal.Length: 5.514481","density: 4.097002e-03<br />Petal.Length: 5.526027","density: 3.500753e-03<br />Petal.Length: 5.537573","density: 2.976957e-03<br />Petal.Length: 5.549119","density: 2.519054e-03<br />Petal.Length: 5.560665","density: 2.128252e-03<br />Petal.Length: 5.572211","density: 1.796839e-03<br />Petal.Length: 5.583757","density: 1.509282e-03<br />Petal.Length: 5.595303","density: 1.261093e-03<br />Petal.Length: 5.606849","density: 1.048054e-03<br />Petal.Length: 5.618395","density: 8.699945e-04<br />Petal.Length: 5.629941","density: 7.221028e-04<br />Petal.Length: 5.641487","density: 5.959834e-04<br />Petal.Length: 5.653033","density: 4.890712e-04<br />Petal.Length: 5.664579","density: 3.989960e-04<br />Petal.Length: 5.676125","density: 3.252938e-04<br />Petal.Length: 5.687671","density: 2.653536e-04<br />Petal.Length: 5.699217","density: 2.151218e-04<br />Petal.Length: 5.710763","density: 1.733086e-04<br />Petal.Length: 5.722309","density: 1.387406e-04<br />Petal.Length: 5.733855","density: 1.110581e-04<br />Petal.Length: 5.745401","density: 8.901679e-05<br />Petal.Length: 5.756947","density: 7.086623e-05<br />Petal.Length: 5.768493","density: 5.603266e-05<br />Petal.Length: 5.780039","density: 4.400175e-05<br />Petal.Length: 5.791585","density: 3.457404e-05<br />Petal.Length: 5.803131","density: 2.722563e-05<br />Petal.Length: 5.814677","density: 2.127956e-05<br />Petal.Length: 5.826223","density: 1.650911e-05<br />Petal.Length: 5.837769","density: 1.271392e-05<br />Petal.Length: 5.849315","density: 9.804174e-06<br />Petal.Length: 5.860861","density: 7.584031e-06<br />Petal.Length: 5.872407","density: 5.818775e-06<br />Petal.Length: 5.883953","density: 4.428567e-06<br />Petal.Length: 5.895499","density: 3.343857e-06<br />Petal.Length: 5.907045","density: 2.530264e-06<br />Petal.Length: 5.918591","density: 1.922595e-06<br />Petal.Length: 5.930137","density: 1.447807e-06<br />Petal.Length: 5.941683","density: 1.080787e-06<br />Petal.Length: 5.953229","density: 7.999606e-07<br />Petal.Length: 5.964775","density: 5.939134e-07<br />Petal.Length: 5.976321","density: 4.432646e-07<br />Petal.Length: 5.987867","density: 3.275922e-07<br />Petal.Length: 5.999413","density: 2.398275e-07<br />Petal.Length: 6.010959","density: 1.739799e-07<br />Petal.Length: 6.022505","density: 1.267216e-07<br />Petal.Length: 6.034051","density: 9.289739e-08<br />Petal.Length: 6.045597","density: 6.737323e-08<br />Petal.Length: 6.057143","density: 4.836556e-08<br />Petal.Length: 6.068689","density: 3.438304e-08<br />Petal.Length: 6.080235","density: 2.456760e-08<br />Petal.Length: 6.091781","density: 1.769014e-08<br />Petal.Length: 6.103327","density: 1.258929e-08<br />Petal.Length: 6.114873","density: 8.861102e-09<br />Petal.Length: 6.126419","density: 6.172308e-09<br />Petal.Length: 6.137965","density: 4.326228e-09<br />Petal.Length: 6.149511","density: 3.059869e-09<br />Petal.Length: 6.161057","density: 2.136667e-09<br />Petal.Length: 6.172603","density: 1.474419e-09<br />Petal.Length: 6.184149","density: 1.006196e-09<br />Petal.Length: 6.195695","density: 6.917830e-10<br />Petal.Length: 6.207241","density: 4.806206e-10<br />Petal.Length: 6.218787","density: 3.292933e-10<br />Petal.Length: 6.230333","density: 2.227557e-10<br />Petal.Length: 6.241879","density: 1.489176e-10<br />Petal.Length: 6.253425","density: 1.004259e-10<br />Petal.Length: 6.264971","density: 6.853861e-11<br />Petal.Length: 6.276517","density: 4.607316e-11<br />Petal.Length: 6.288063","density: 3.055084e-11<br />Petal.Length: 6.299609","density: 2.000569e-11<br />Petal.Length: 6.311155","density: 1.323296e-11<br />Petal.Length: 6.322701","density: 8.872073e-12<br />Petal.Length: 6.334247","density: 5.851373e-12<br />Petal.Length: 6.345793","density: 3.803047e-12<br />Petal.Length: 6.357339","density: 2.439133e-12<br />Petal.Length: 6.368885","density: 1.582507e-12<br />Petal.Length: 6.380431","density: 1.042357e-12<br />Petal.Length: 6.391977","density: 6.744759e-13<br />Petal.Length: 6.403523","density: 4.296570e-13<br />Petal.Length: 6.415068","density: 2.698700e-13<br />Petal.Length: 6.426614","density: 1.717530e-13<br />Petal.Length: 6.438160","density: 1.111595e-13<br />Petal.Length: 6.449706","density: 7.057514e-14<br />Petal.Length: 6.461252","density: 4.407755e-14<br />Petal.Length: 6.472798","density: 2.713450e-14<br />Petal.Length: 6.484344","density: 1.690109e-14<br />Petal.Length: 6.495890","density: 1.075917e-14<br />Petal.Length: 6.507436","density: 6.677077e-15<br />Petal.Length: 6.518982","density: 4.043674e-15<br />Petal.Length: 6.530528","density: 2.413156e-15<br />Petal.Length: 6.542074","density: 1.472152e-15<br />Petal.Length: 6.553620","density: 9.199851e-16<br />Petal.Length: 6.565166","density: 5.549517e-16<br />Petal.Length: 6.576712","density: 3.161115e-16<br />Petal.Length: 6.588258","density: 1.664420e-16<br />Petal.Length: 6.599804","density: 1.067618e-16<br />Petal.Length: 6.611350","density: 7.322531e-17<br />Petal.Length: 6.622896","density: 6.020473e-17<br />Petal.Length: 6.634442","density: 3.964422e-17<br />Petal.Length: 6.645988","density: 1.636012e-18<br />Petal.Length: 6.657534","density: 1.273930e-17<br />Petal.Length: 6.669080","density: 3.007059e-17<br />Petal.Length: 6.680626","density: 2.114980e-17<br />Petal.Length: 6.692172","density: 4.730992e-18<br />Petal.Length: 6.703718","density: 0.000000e+00<br />Petal.Length: 6.715264","density: 0.000000e+00<br />Petal.Length: 6.726810","density: 0.000000e+00<br />Petal.Length: 6.738356","density: 0.000000e+00<br />Petal.Length: 6.749902","density: 0.000000e+00<br />Petal.Length: 6.761448","density: 2.264071e-18<br />Petal.Length: 6.772994","density: 1.753109e-17<br />Petal.Length: 6.784540","density: 9.106571e-18<br />Petal.Length: 6.796086","density: 1.420528e-17<br />Petal.Length: 6.807632","density: 1.535852e-17<br />Petal.Length: 6.819178","density: 2.064863e-19<br />Petal.Length: 6.830724","density: 1.594397e-18<br />Petal.Length: 6.842270","density: 1.545816e-17<br />Petal.Length: 6.853816","density: 1.901514e-17<br />Petal.Length: 6.865362","density: 1.513139e-17<br />Petal.Length: 6.876908","density: 9.165454e-18<br />Petal.Length: 6.888454","density: 8.379870e-19<br />Petal.Length: 6.900000","density: 8.379870e-19<br />Petal.Length: 6.900000","density: 9.165454e-18<br />Petal.Length: 6.888454","density: 1.513139e-17<br />Petal.Length: 6.876908","density: 1.901514e-17<br />Petal.Length: 6.865362","density: 1.545816e-17<br />Petal.Length: 6.853816","density: 1.594397e-18<br />Petal.Length: 6.842270","density: 2.064863e-19<br />Petal.Length: 6.830724","density: 1.535852e-17<br />Petal.Length: 6.819178","density: 1.420528e-17<br />Petal.Length: 6.807632","density: 9.106571e-18<br />Petal.Length: 6.796086","density: 1.753109e-17<br />Petal.Length: 6.784540","density: 2.264071e-18<br />Petal.Length: 6.772994","density: 0.000000e+00<br />Petal.Length: 6.761448","density: 0.000000e+00<br />Petal.Length: 6.749902","density: 0.000000e+00<br />Petal.Length: 6.738356","density: 0.000000e+00<br />Petal.Length: 6.726810","density: 0.000000e+00<br />Petal.Length: 6.715264","density: 4.730992e-18<br />Petal.Length: 6.703718","density: 2.114980e-17<br />Petal.Length: 6.692172","density: 3.007059e-17<br />Petal.Length: 6.680626","density: 1.273930e-17<br />Petal.Length: 6.669080","density: 1.636012e-18<br />Petal.Length: 6.657534","density: 3.964422e-17<br />Petal.Length: 6.645988","density: 6.020473e-17<br />Petal.Length: 6.634442","density: 7.322531e-17<br />Petal.Length: 6.622896","density: 1.067618e-16<br />Petal.Length: 6.611350","density: 1.664420e-16<br />Petal.Length: 6.599804","density: 3.161115e-16<br />Petal.Length: 6.588258","density: 5.549517e-16<br />Petal.Length: 6.576712","density: 9.199851e-16<br />Petal.Length: 6.565166","density: 1.472152e-15<br />Petal.Length: 6.553620","density: 2.413156e-15<br />Petal.Length: 6.542074","density: 4.043674e-15<br />Petal.Length: 6.530528","density: 6.677077e-15<br />Petal.Length: 6.518982","density: 1.075917e-14<br />Petal.Length: 6.507436","density: 1.690109e-14<br />Petal.Length: 6.495890","density: 2.713450e-14<br />Petal.Length: 6.484344","density: 4.407755e-14<br />Petal.Length: 6.472798","density: 7.057514e-14<br />Petal.Length: 6.461252","density: 1.111595e-13<br />Petal.Length: 6.449706","density: 1.717530e-13<br />Petal.Length: 6.438160","density: 2.698700e-13<br />Petal.Length: 6.426614","density: 4.296570e-13<br />Petal.Length: 6.415068","density: 6.744759e-13<br />Petal.Length: 6.403523","density: 1.042357e-12<br />Petal.Length: 6.391977","density: 1.582507e-12<br />Petal.Length: 6.380431","density: 2.439133e-12<br />Petal.Length: 6.368885","density: 3.803047e-12<br />Petal.Length: 6.357339","density: 5.851373e-12<br />Petal.Length: 6.345793","density: 8.872073e-12<br />Petal.Length: 6.334247","density: 1.323296e-11<br />Petal.Length: 6.322701","density: 2.000569e-11<br />Petal.Length: 6.311155","density: 3.055084e-11<br />Petal.Length: 6.299609","density: 4.607316e-11<br />Petal.Length: 6.288063","density: 6.853861e-11<br />Petal.Length: 6.276517","density: 1.004259e-10<br />Petal.Length: 6.264971","density: 1.489176e-10<br />Petal.Length: 6.253425","density: 2.227557e-10<br />Petal.Length: 6.241879","density: 3.292933e-10<br />Petal.Length: 6.230333","density: 4.806206e-10<br />Petal.Length: 6.218787","density: 6.917830e-10<br />Petal.Length: 6.207241","density: 1.006196e-09<br />Petal.Length: 6.195695","density: 1.474419e-09<br />Petal.Length: 6.184149","density: 2.136667e-09<br />Petal.Length: 6.172603","density: 3.059869e-09<br />Petal.Length: 6.161057","density: 4.326228e-09<br />Petal.Length: 6.149511","density: 6.172308e-09<br />Petal.Length: 6.137965","density: 8.861102e-09<br />Petal.Length: 6.126419","density: 1.258929e-08<br />Petal.Length: 6.114873","density: 1.769014e-08<br />Petal.Length: 6.103327","density: 2.456760e-08<br />Petal.Length: 6.091781","density: 3.438304e-08<br />Petal.Length: 6.080235","density: 4.836556e-08<br />Petal.Length: 6.068689","density: 6.737323e-08<br />Petal.Length: 6.057143","density: 9.289739e-08<br />Petal.Length: 6.045597","density: 1.267216e-07<br />Petal.Length: 6.034051","density: 1.739799e-07<br />Petal.Length: 6.022505","density: 2.398275e-07<br />Petal.Length: 6.010959","density: 3.275922e-07<br />Petal.Length: 5.999413","density: 4.432646e-07<br />Petal.Length: 5.987867","density: 5.939134e-07<br />Petal.Length: 5.976321","density: 7.999606e-07<br />Petal.Length: 5.964775","density: 1.080787e-06<br />Petal.Length: 5.953229","density: 1.447807e-06<br />Petal.Length: 5.941683","density: 1.922595e-06<br />Petal.Length: 5.930137","density: 2.530264e-06<br />Petal.Length: 5.918591","density: 3.343857e-06<br />Petal.Length: 5.907045","density: 4.428567e-06<br />Petal.Length: 5.895499","density: 5.818775e-06<br />Petal.Length: 5.883953","density: 7.584031e-06<br />Petal.Length: 5.872407","density: 9.804174e-06<br />Petal.Length: 5.860861","density: 1.271392e-05<br />Petal.Length: 5.849315","density: 1.650911e-05<br />Petal.Length: 5.837769","density: 2.127956e-05<br />Petal.Length: 5.826223","density: 2.722563e-05<br />Petal.Length: 5.814677","density: 3.457404e-05<br />Petal.Length: 5.803131","density: 4.400175e-05<br />Petal.Length: 5.791585","density: 5.603266e-05<br />Petal.Length: 5.780039","density: 7.086623e-05<br />Petal.Length: 5.768493","density: 8.901679e-05<br />Petal.Length: 5.756947","density: 1.110581e-04<br />Petal.Length: 5.745401","density: 1.387406e-04<br />Petal.Length: 5.733855","density: 1.733086e-04<br />Petal.Length: 5.722309","density: 2.151218e-04<br />Petal.Length: 5.710763","density: 2.653536e-04<br />Petal.Length: 5.699217","density: 3.252938e-04<br />Petal.Length: 5.687671","density: 3.989960e-04<br />Petal.Length: 5.676125","density: 4.890712e-04<br />Petal.Length: 5.664579","density: 5.959834e-04<br />Petal.Length: 5.653033","density: 7.221028e-04<br />Petal.Length: 5.641487","density: 8.699945e-04<br />Petal.Length: 5.629941","density: 1.048054e-03<br />Petal.Length: 5.618395","density: 1.261093e-03<br />Petal.Length: 5.606849","density: 1.509282e-03<br />Petal.Length: 5.595303","density: 1.796839e-03<br />Petal.Length: 5.583757","density: 2.128252e-03<br />Petal.Length: 5.572211","density: 2.519054e-03<br />Petal.Length: 5.560665","density: 2.976957e-03<br />Petal.Length: 5.549119","density: 3.500753e-03<br />Petal.Length: 5.537573","density: 4.097002e-03<br />Petal.Length: 5.526027","density: 4.772564e-03<br />Petal.Length: 5.514481","density: 5.553080e-03<br />Petal.Length: 5.502935","density: 6.449150e-03<br />Petal.Length: 5.491389","density: 7.456116e-03<br />Petal.Length: 5.479843","density: 8.582805e-03<br />Petal.Length: 5.468297","density: 9.838281e-03<br />Petal.Length: 5.456751","density: 1.126030e-02<br />Petal.Length: 5.445205","density: 1.286102e-02<br />Petal.Length: 5.433659","density: 1.462940e-02<br />Petal.Length: 5.422114","density: 1.657556e-02<br />Petal.Length: 5.410568","density: 1.870968e-02<br />Petal.Length: 5.399022","density: 2.108139e-02<br />Petal.Length: 5.387476","density: 2.370176e-02<br />Petal.Length: 5.375930","density: 2.655034e-02<br />Petal.Length: 5.364384","density: 2.963677e-02<br />Petal.Length: 5.352838","density: 3.297049e-02<br />Petal.Length: 5.341292","density: 3.661010e-02<br />Petal.Length: 5.329746","density: 4.056239e-02<br />Petal.Length: 5.318200","density: 4.479585e-02<br />Petal.Length: 5.306654","density: 4.931776e-02<br />Petal.Length: 5.295108","density: 5.413499e-02<br />Petal.Length: 5.283562","density: 5.931046e-02<br />Petal.Length: 5.272016","density: 6.484461e-02<br />Petal.Length: 5.260470","density: 7.069572e-02<br />Petal.Length: 5.248924","density: 7.686772e-02<br />Petal.Length: 5.237378","density: 8.336412e-02<br />Petal.Length: 5.225832","density: 9.024784e-02<br />Petal.Length: 5.214286","density: 9.751340e-02<br />Petal.Length: 5.202740","density: 1.051121e-01<br />Petal.Length: 5.191194","density: 1.130449e-01<br />Petal.Length: 5.179648","density: 1.213127e-01<br />Petal.Length: 5.168102","density: 1.299763e-01<br />Petal.Length: 5.156556","density: 1.390269e-01<br />Petal.Length: 5.145010","density: 1.484125e-01<br />Petal.Length: 5.133464","density: 1.581329e-01<br />Petal.Length: 5.121918","density: 1.681875e-01<br />Petal.Length: 5.110372","density: 1.786351e-01<br />Petal.Length: 5.098826","density: 1.894659e-01<br />Petal.Length: 5.087280","density: 2.006275e-01<br />Petal.Length: 5.075734","density: 2.121189e-01<br />Petal.Length: 5.064188","density: 2.239388e-01<br />Petal.Length: 5.052642","density: 2.361437e-01<br />Petal.Length: 5.041096","density: 2.487227e-01<br />Petal.Length: 5.029550","density: 2.616229e-01<br />Petal.Length: 5.018004","density: 2.748413e-01<br />Petal.Length: 5.006458","density: 2.883747e-01<br />Petal.Length: 4.994912","density: 3.022734e-01<br />Petal.Length: 4.983366","density: 3.165214e-01<br />Petal.Length: 4.971820","density: 3.310640e-01<br />Petal.Length: 4.960274","density: 3.458934e-01<br />Petal.Length: 4.948728","density: 3.610012e-01<br />Petal.Length: 4.937182","density: 3.764220e-01<br />Petal.Length: 4.925636","density: 3.921293e-01<br />Petal.Length: 4.914090","density: 4.080668e-01<br />Petal.Length: 4.902544","density: 4.242184e-01<br />Petal.Length: 4.890998","density: 4.405669e-01<br />Petal.Length: 4.879452","density: 4.571181e-01<br />Petal.Length: 4.867906","density: 4.738304e-01<br />Petal.Length: 4.856360","density: 4.906513e-01<br />Petal.Length: 4.844814","density: 5.075540e-01<br />Petal.Length: 4.833268","density: 5.245102e-01<br />Petal.Length: 4.821722","density: 5.414845e-01<br />Petal.Length: 4.810176","density: 5.584193e-01<br />Petal.Length: 4.798630","density: 5.752766e-01<br />Petal.Length: 4.787084","density: 5.920192e-01<br />Petal.Length: 4.775538","density: 6.086094e-01<br />Petal.Length: 4.763992","density: 6.249618e-01<br />Petal.Length: 4.752446","density: 6.410088e-01<br />Petal.Length: 4.740900","density: 6.567401e-01<br />Petal.Length: 4.729354","density: 6.721131e-01<br />Petal.Length: 4.717808","density: 6.870850e-01<br />Petal.Length: 4.706262","density: 7.015220e-01<br />Petal.Length: 4.694716","density: 7.153582e-01<br />Petal.Length: 4.683170","density: 7.286247e-01<br />Petal.Length: 4.671624","density: 7.412815e-01<br />Petal.Length: 4.660078","density: 7.532900e-01<br />Petal.Length: 4.648532","density: 7.644826e-01<br />Petal.Length: 4.636986","density: 7.748116e-01<br />Petal.Length: 4.625440","density: 7.843592e-01<br />Petal.Length: 4.613894","density: 7.930993e-01<br />Petal.Length: 4.602348","density: 8.010087e-01<br />Petal.Length: 4.590802","density: 8.079148e-01<br />Petal.Length: 4.579256","density: 8.138071e-01<br />Petal.Length: 4.567710","density: 8.188173e-01<br />Petal.Length: 4.556164","density: 8.229442e-01<br />Petal.Length: 4.544618","density: 8.261905e-01<br />Petal.Length: 4.533072","density: 8.284159e-01<br />Petal.Length: 4.521526","density: 8.296586e-01<br />Petal.Length: 4.509980","density: 8.300829e-01<br />Petal.Length: 4.498434","density: 8.297173e-01<br />Petal.Length: 4.486888","density: 8.285935e-01<br />Petal.Length: 4.475342","density: 8.266353e-01<br />Petal.Length: 4.463796","density: 8.239255e-01<br />Petal.Length: 4.452250","density: 8.206279e-01<br />Petal.Length: 4.440705","density: 8.167936e-01<br />Petal.Length: 4.429159","density: 8.124754e-01<br />Petal.Length: 4.417613","density: 8.076714e-01<br />Petal.Length: 4.406067","density: 8.024844e-01<br />Petal.Length: 4.394521","density: 7.970384e-01<br />Petal.Length: 4.382975","density: 7.913894e-01<br />Petal.Length: 4.371429","density: 7.855925e-01<br />Petal.Length: 4.359883","density: 7.797005e-01<br />Petal.Length: 4.348337","density: 7.737997e-01<br />Petal.Length: 4.336791","density: 7.679460e-01<br />Petal.Length: 4.325245","density: 7.621787e-01<br />Petal.Length: 4.313699","density: 7.565336e-01<br />Petal.Length: 4.302153","density: 7.510762e-01<br />Petal.Length: 4.290607","density: 7.458444e-01<br />Petal.Length: 4.279061","density: 7.408239e-01<br />Petal.Length: 4.267515","density: 7.360224e-01<br />Petal.Length: 4.255969","density: 7.314429e-01<br />Petal.Length: 4.244423","density: 7.271182e-01<br />Petal.Length: 4.232877","density: 7.230262e-01<br />Petal.Length: 4.221331","density: 7.191098e-01<br />Petal.Length: 4.209785","density: 7.153439e-01<br />Petal.Length: 4.198239","density: 7.117002e-01<br />Petal.Length: 4.186693","density: 7.081508e-01<br />Petal.Length: 4.175147","density: 7.046288e-01<br />Petal.Length: 4.163601","density: 7.010780e-01<br />Petal.Length: 4.152055","density: 6.974544e-01<br />Petal.Length: 4.140509","density: 6.937134e-01<br />Petal.Length: 4.128963","density: 6.897685e-01<br />Petal.Length: 4.117417","density: 6.855414e-01<br />Petal.Length: 4.105871","density: 6.810177e-01<br />Petal.Length: 4.094325","density: 6.761565e-01<br />Petal.Length: 4.082779","density: 6.709185e-01<br />Petal.Length: 4.071233","density: 6.651878e-01<br />Petal.Length: 4.059687","density: 6.589125e-01<br />Petal.Length: 4.048141","density: 6.521398e-01<br />Petal.Length: 4.036595","density: 6.448510e-01<br />Petal.Length: 4.025049","density: 6.370314e-01<br />Petal.Length: 4.013503","density: 6.285789e-01<br />Petal.Length: 4.001957","density: 6.194927e-01<br />Petal.Length: 3.990411","density: 6.098720e-01<br />Petal.Length: 3.978865","density: 5.997300e-01<br />Petal.Length: 3.967319","density: 5.890841e-01<br />Petal.Length: 3.955773","density: 5.778836e-01<br />Petal.Length: 3.944227","density: 5.661814e-01<br />Petal.Length: 3.932681","density: 5.540946e-01<br />Petal.Length: 3.921135","density: 5.416636e-01<br />Petal.Length: 3.909589","density: 5.289315e-01<br />Petal.Length: 3.898043","density: 5.159155e-01<br />Petal.Length: 3.886497","density: 5.027016e-01<br />Petal.Length: 3.874951","density: 4.893823e-01<br />Petal.Length: 3.863405","density: 4.760093e-01<br />Petal.Length: 3.851859","density: 4.626344e-01<br />Petal.Length: 3.840313","density: 4.493319e-01<br />Petal.Length: 3.828767","density: 4.361885e-01<br />Petal.Length: 3.817221","density: 4.232397e-01<br />Petal.Length: 3.805675","density: 4.105291e-01<br />Petal.Length: 3.794129","density: 3.980982e-01<br />Petal.Length: 3.782583","density: 3.860481e-01<br />Petal.Length: 3.771037","density: 3.744369e-01<br />Petal.Length: 3.759491","density: 3.632346e-01<br />Petal.Length: 3.747945","density: 3.524630e-01<br />Petal.Length: 3.736399","density: 3.421406e-01<br />Petal.Length: 3.724853","density: 3.323615e-01<br />Petal.Length: 3.713307","density: 3.231410e-01<br />Petal.Length: 3.701761","density: 3.144003e-01<br />Petal.Length: 3.690215","density: 3.061364e-01<br />Petal.Length: 3.678669","density: 2.983437e-01<br />Petal.Length: 3.667123","density: 2.910852e-01<br />Petal.Length: 3.655577","density: 2.843392e-01<br />Petal.Length: 3.644031","density: 2.780083e-01<br />Petal.Length: 3.632485","density: 2.720721e-01<br />Petal.Length: 3.620939","density: 2.665086e-01<br />Petal.Length: 3.609393","density: 2.613439e-01<br />Petal.Length: 3.597847","density: 2.565356e-01<br />Petal.Length: 3.586301","density: 2.519987e-01<br />Petal.Length: 3.574755","density: 2.477062e-01<br />Petal.Length: 3.563209","density: 2.436313e-01<br />Petal.Length: 3.551663","density: 2.397709e-01<br />Petal.Length: 3.540117","density: 2.360814e-01<br />Petal.Length: 3.528571","density: 2.325073e-01<br />Petal.Length: 3.517025","density: 2.290254e-01<br />Petal.Length: 3.505479","density: 2.256134e-01<br />Petal.Length: 3.493933","density: 2.222530e-01<br />Petal.Length: 3.482387","density: 2.189122e-01<br />Petal.Length: 3.470841","density: 2.155680e-01<br />Petal.Length: 3.459295","density: 2.122064e-01<br />Petal.Length: 3.447750","density: 2.088145e-01<br />Petal.Length: 3.436204","density: 2.053718e-01<br />Petal.Length: 3.424658","density: 2.018631e-01<br />Petal.Length: 3.413112","density: 1.982907e-01<br />Petal.Length: 3.401566","density: 1.946505e-01<br />Petal.Length: 3.390020","density: 1.909395e-01<br />Petal.Length: 3.378474","density: 1.871447e-01<br />Petal.Length: 3.366928","density: 1.832667e-01<br />Petal.Length: 3.355382","density: 1.793212e-01<br />Petal.Length: 3.343836","density: 1.753124e-01<br />Petal.Length: 3.332290","density: 1.712450e-01<br />Petal.Length: 3.320744","density: 1.671184e-01<br />Petal.Length: 3.309198","density: 1.629444e-01<br />Petal.Length: 3.297652","density: 1.587403e-01<br />Petal.Length: 3.286106","density: 1.545148e-01<br />Petal.Length: 3.274560","density: 1.502768e-01<br />Petal.Length: 3.263014","density: 1.460374e-01<br />Petal.Length: 3.251468","density: 1.418131e-01<br />Petal.Length: 3.239922","density: 1.376130e-01<br />Petal.Length: 3.228376","density: 1.334459e-01<br />Petal.Length: 3.216830","density: 1.293201e-01<br />Petal.Length: 3.205284","density: 1.252534e-01<br />Petal.Length: 3.193738","density: 1.212588e-01<br />Petal.Length: 3.182192","density: 1.173329e-01<br />Petal.Length: 3.170646","density: 1.134803e-01<br />Petal.Length: 3.159100","density: 1.097051e-01<br />Petal.Length: 3.147554","density: 1.060239e-01<br />Petal.Length: 3.136008","density: 1.024411e-01<br />Petal.Length: 3.124462","density: 9.894243e-02<br />Petal.Length: 3.112916","density: 9.552714e-02<br />Petal.Length: 3.101370","density: 9.219396e-02<br />Petal.Length: 3.089824","density: 8.895275e-02<br />Petal.Length: 3.078278","density: 8.579962e-02<br />Petal.Length: 3.066732","density: 8.271658e-02<br />Petal.Length: 3.055186","density: 7.969948e-02<br />Petal.Length: 3.043640","density: 7.674401e-02<br />Petal.Length: 3.032094","density: 7.385339e-02<br />Petal.Length: 3.020548","density: 7.102096e-02<br />Petal.Length: 3.009002","density: 6.823274e-02<br />Petal.Length: 2.997456","density: 6.548498e-02<br />Petal.Length: 2.985910","density: 6.277425e-02<br />Petal.Length: 2.974364","density: 6.010214e-02<br />Petal.Length: 2.962818","density: 5.746568e-02<br />Petal.Length: 2.951272","density: 5.485801e-02<br />Petal.Length: 2.939726","density: 5.227872e-02<br />Petal.Length: 2.928180","density: 4.972785e-02<br />Petal.Length: 2.916634","density: 4.721090e-02<br />Petal.Length: 2.905088","density: 4.473143e-02<br />Petal.Length: 2.893542","density: 4.228688e-02<br />Petal.Length: 2.881996","density: 3.988020e-02<br />Petal.Length: 2.870450","density: 3.751460e-02<br />Petal.Length: 2.858904","density: 3.520155e-02<br />Petal.Length: 2.847358","density: 3.294906e-02<br />Petal.Length: 2.835812","density: 3.075306e-02<br />Petal.Length: 2.824266","density: 2.861756e-02<br />Petal.Length: 2.812720","density: 2.654650e-02<br />Petal.Length: 2.801174","density: 2.455515e-02<br />Petal.Length: 2.789628","density: 2.265128e-02<br />Petal.Length: 2.778082","density: 2.082528e-02<br />Petal.Length: 2.766536","density: 1.907960e-02<br />Petal.Length: 2.754990","density: 1.741636e-02<br />Petal.Length: 2.743444","density: 1.585053e-02<br />Petal.Length: 2.731898","density: 1.438604e-02<br />Petal.Length: 2.720352","density: 1.300771e-02<br />Petal.Length: 2.708806","density: 1.171526e-02<br />Petal.Length: 2.697260","density: 1.050803e-02<br />Petal.Length: 2.685714","density: 9.397456e-03<br />Petal.Length: 2.674168","density: 8.382851e-03<br />Petal.Length: 2.662622","density: 7.446717e-03<br />Petal.Length: 2.651076","density: 6.586499e-03<br />Petal.Length: 2.639530","density: 5.799454e-03<br />Petal.Length: 2.627984","density: 5.092468e-03<br />Petal.Length: 2.616438","density: 4.461837e-03<br />Petal.Length: 2.604892","density: 3.891530e-03<br />Petal.Length: 2.593346","density: 3.378094e-03<br />Petal.Length: 2.581800","density: 2.918060e-03<br />Petal.Length: 2.570254","density: 2.514609e-03<br />Petal.Length: 2.558708","density: 2.163215e-03<br />Petal.Length: 2.547162","density: 1.851699e-03<br />Petal.Length: 2.535616","density: 1.576911e-03<br />Petal.Length: 2.524070","density: 1.335797e-03<br />Petal.Length: 2.512524","density: 1.129326e-03<br />Petal.Length: 2.500978","density: 9.536924e-04<br />Petal.Length: 2.489432","density: 8.010174e-04<br />Petal.Length: 2.477886","density: 6.690394e-04<br />Petal.Length: 2.466341","density: 5.556169e-04<br />Petal.Length: 2.454795","density: 4.607628e-04<br />Petal.Length: 2.443249","density: 3.819340e-04<br />Petal.Length: 2.431703","density: 3.147216e-04<br />Petal.Length: 2.420157","density: 2.577718e-04<br />Petal.Length: 2.408611","density: 2.098290e-04<br />Petal.Length: 2.397065","density: 1.706645e-04<br />Petal.Length: 2.385519","density: 1.388568e-04<br />Petal.Length: 2.373973","density: 1.122476e-04<br />Petal.Length: 2.362427","density: 9.014362e-05<br />Petal.Length: 2.350881","density: 7.191330e-05<br />Petal.Length: 2.339335","density: 5.736346e-05<br />Petal.Length: 2.327789","density: 4.581213e-05<br />Petal.Length: 2.316243","density: 3.632826e-05<br />Petal.Length: 2.304697","density: 2.860325e-05<br />Petal.Length: 2.293151","density: 2.236070e-05<br />Petal.Length: 2.281605","density: 1.749221e-05<br />Petal.Length: 2.270059","density: 1.371291e-05<br />Petal.Length: 2.258513","density: 1.066692e-05<br />Petal.Length: 2.246967","density: 8.233710e-06<br />Petal.Length: 2.235421","density: 6.306976e-06<br />Petal.Length: 2.223875","density: 4.838443e-06<br />Petal.Length: 2.212329","density: 3.723533e-06<br />Petal.Length: 2.200783","density: 2.841234e-06<br />Petal.Length: 2.189237","density: 2.149940e-06<br />Petal.Length: 2.177691","density: 1.613513e-06<br />Petal.Length: 2.166145","density: 1.213890e-06<br />Petal.Length: 2.154599","density: 9.171112e-07<br />Petal.Length: 2.143053","density: 6.864606e-07<br />Petal.Length: 2.131507","density: 5.091881e-07<br />Petal.Length: 2.119961","density: 3.743804e-07<br />Petal.Length: 2.108415","density: 2.762116e-07<br />Petal.Length: 2.096869","density: 2.048844e-07<br />Petal.Length: 2.085323","density: 1.504338e-07<br />Petal.Length: 2.073777","density: 1.093789e-07<br />Petal.Length: 2.062231","density: 7.878203e-08<br />Petal.Length: 2.050685","density: 5.700064e-08<br />Petal.Length: 2.039139","density: 4.151484e-08<br />Petal.Length: 2.027593","density: 2.990092e-08<br />Petal.Length: 2.016047","density: 2.130987e-08<br />Petal.Length: 2.004501","density: 1.503506e-08<br />Petal.Length: 1.992955","density: 1.066804e-08<br />Petal.Length: 1.981409","density: 7.629536e-09<br />Petal.Length: 1.969863","density: 5.390465e-09<br />Petal.Length: 1.958317","density: 3.765425e-09<br />Petal.Length: 1.946771","density: 2.602203e-09<br />Petal.Length: 1.935225","density: 1.810722e-09<br />Petal.Length: 1.923679","density: 1.271712e-09<br />Petal.Length: 1.912133","density: 8.813855e-10<br />Petal.Length: 1.900587","density: 6.034326e-10<br />Petal.Length: 1.889041","density: 4.084434e-10<br />Petal.Length: 1.877495","density: 2.787266e-10<br />Petal.Length: 1.865949","density: 1.922531e-10<br />Petal.Length: 1.854403","density: 1.307077e-10<br />Petal.Length: 1.842857","density: 8.770468e-11<br />Petal.Length: 1.831311","density: 5.814008e-11<br />Petal.Length: 1.819765","density: 3.891023e-11<br />Petal.Length: 1.808219","density: 2.636036e-11<br />Petal.Length: 1.796673","density: 1.758050e-11<br />Petal.Length: 1.785127","density: 1.156097e-11<br />Petal.Length: 1.773581","density: 7.505369e-12<br />Petal.Length: 1.762035","density: 4.926169e-12<br />Petal.Length: 1.750489","density: 3.278102e-12<br />Petal.Length: 1.738943","density: 2.144650e-12<br />Petal.Length: 1.727397","density: 1.382121e-12<br />Petal.Length: 1.715851","density: 8.786519e-13<br />Petal.Length: 1.704305","density: 5.656068e-13<br />Petal.Length: 1.692759","density: 3.697389e-13<br />Petal.Length: 1.681213","density: 2.372809e-13<br />Petal.Length: 1.669667","density: 1.498406e-13<br />Petal.Length: 1.658121","density: 9.329037e-14<br />Petal.Length: 1.646575","density: 5.894998e-14<br />Petal.Length: 1.635029","density: 3.786655e-14<br />Petal.Length: 1.623483","density: 2.385595e-14<br />Petal.Length: 1.611937","density: 1.478030e-14<br />Petal.Length: 1.600391","density: 9.012741e-15<br />Petal.Length: 1.588845","density: 5.590846e-15<br />Petal.Length: 1.577299","density: 3.539711e-15<br />Petal.Length: 1.565753","density: 2.196357e-15<br />Petal.Length: 1.554207","density: 1.342276e-15<br />Petal.Length: 1.542661","density: 8.120112e-16<br />Petal.Length: 1.531115","density: 4.900159e-16<br />Petal.Length: 1.519569","density: 3.021880e-16<br />Petal.Length: 1.508023","density: 1.954571e-16<br />Petal.Length: 1.496477","density: 1.252223e-16<br />Petal.Length: 1.484932","density: 5.986384e-17<br />Petal.Length: 1.473386","density: 1.822651e-17<br />Petal.Length: 1.461840","density: 2.257846e-17<br />Petal.Length: 1.450294","density: 1.616330e-17<br />Petal.Length: 1.438748","density: 4.741578e-18<br />Petal.Length: 1.427202","density: 1.012583e-17<br />Petal.Length: 1.415656","density: 7.952139e-17<br />Petal.Length: 1.404110","density: 7.927448e-17<br />Petal.Length: 1.392564","density: 9.067668e-17<br />Petal.Length: 1.381018","density: 8.208378e-17<br />Petal.Length: 1.369472","density: 4.525974e-17<br />Petal.Length: 1.357926","density: 1.174311e-16<br />Petal.Length: 1.346380","density: 1.025031e-16<br />Petal.Length: 1.334834","density: 1.008548e-16<br />Petal.Length: 1.323288","density: 8.883068e-17<br />Petal.Length: 1.311742","density: 5.509513e-17<br />Petal.Length: 1.300196","density: 9.950829e-17<br />Petal.Length: 1.288650","density: 8.416439e-17<br />Petal.Length: 1.277104","density: 8.530910e-17<br />Petal.Length: 1.265558","density: 8.154616e-17<br />Petal.Length: 1.254012","density: 5.714716e-17<br />Petal.Length: 1.242466","density: 6.825045e-17<br />Petal.Length: 1.230920","density: 4.447694e-17<br />Petal.Length: 1.219374","density: 4.188477e-17<br />Petal.Length: 1.207828","density: 5.519596e-17<br />Petal.Length: 1.196282","density: 7.595416e-17<br />Petal.Length: 1.184736","density: 1.203673e-16<br />Petal.Length: 1.173190","density: 9.623607e-17<br />Petal.Length: 1.161644","density: 1.038899e-16<br />Petal.Length: 1.150098","density: 1.081674e-16<br />Petal.Length: 1.138552","density: 8.473707e-17<br />Petal.Length: 1.127006","density: 1.180469e-16<br />Petal.Length: 1.115460","density: 1.189368e-16<br />Petal.Length: 1.103914","density: 1.188613e-16<br />Petal.Length: 1.092368","density: 1.097365e-16<br />Petal.Length: 1.080822","density: 8.599306e-17<br />Petal.Length: 1.069276","density: 6.933813e-17<br />Petal.Length: 1.057730","density: 8.761558e-17<br />Petal.Length: 1.046184","density: 9.450902e-17<br />Petal.Length: 1.034638","density: 7.096270e-17<br />Petal.Length: 1.023092","density: 3.073375e-17<br />Petal.Length: 1.011546","density: 8.555624e-17<br />Petal.Length: 1.000000","density: 8.555624e-17<br />Petal.Length: 1.000000"],"key":["51","52","53","54","55","56","57","58","59","60","61","62","63","64","65","66","67","68","69","70","71","72","73","74","75","76","77","78","79","80","81","82","83","84","85","86","87","88","89","90","91","92","93","94","95","96","97","98","99","100"],"type":"scatter","mode":"lines","line":{"width":1.88976377952756,"color":"rgba(0,0,0,1)","dash":"solid"},"fill":"toself","fillcolor":"rgba(0,186,56,1)","hoveron":"points","set":"SharedData91833e3a","name":"versicolor","legendgroup":"versicolor","showlegend":true,"xaxis":"x3","yaxis":"y10","hoverinfo":"text","_isSimpleKey":true,"_isNestedKey":false,"frame":null},{"x":[1,1.01154598825832,1.02309197651663,1.03463796477495,1.04618395303327,1.05772994129159,1.0692759295499,1.08082191780822,1.09236790606654,1.10391389432485,1.11545988258317,1.12700587084149,1.1385518590998,1.15009784735812,1.16164383561644,1.17318982387476,1.18473581213307,1.19628180039139,1.20782778864971,1.21937377690802,1.23091976516634,1.24246575342466,1.25401174168297,1.26555772994129,1.27710371819961,1.28864970645793,1.30019569471624,1.31174168297456,1.32328767123288,1.33483365949119,1.34637964774951,1.35792563600783,1.36947162426614,1.38101761252446,1.39256360078278,1.4041095890411,1.41565557729941,1.42720156555773,1.43874755381605,1.45029354207436,1.46183953033268,1.473385518591,1.48493150684931,1.49647749510763,1.50802348336595,1.51956947162427,1.53111545988258,1.5426614481409,1.55420743639922,1.56575342465753,1.57729941291585,1.58884540117417,1.60039138943249,1.6119373776908,1.62348336594912,1.63502935420744,1.64657534246575,1.65812133072407,1.66966731898239,1.6812133072407,1.69275929549902,1.70430528375734,1.71585127201566,1.72739726027397,1.73894324853229,1.75048923679061,1.76203522504892,1.77358121330724,1.78512720156556,1.79667318982387,1.80821917808219,1.81976516634051,1.83131115459883,1.84285714285714,1.85440313111546,1.86594911937378,1.87749510763209,1.88904109589041,1.90058708414873,1.91213307240705,1.92367906066536,1.93522504892368,1.946771037182,1.95831702544031,1.96986301369863,1.98140900195695,1.99295499021526,2.00450097847358,2.0160469667319,2.02759295499022,2.03913894324853,2.05068493150685,2.06223091976517,2.07377690802348,2.0853228962818,2.09686888454012,2.10841487279843,2.11996086105675,2.13150684931507,2.14305283757339,2.1545988258317,2.16614481409002,2.17769080234834,2.18923679060665,2.20078277886497,2.21232876712329,2.2238747553816,2.23542074363992,2.24696673189824,2.25851272015656,2.27005870841487,2.28160469667319,2.29315068493151,2.30469667318982,2.31624266144814,2.32778864970646,2.33933463796478,2.35088062622309,2.36242661448141,2.37397260273973,2.38551859099804,2.39706457925636,2.40861056751468,2.42015655577299,2.43170254403131,2.44324853228963,2.45479452054795,2.46634050880626,2.47788649706458,2.4894324853229,2.50097847358121,2.51252446183953,2.52407045009785,2.53561643835616,2.54716242661448,2.5587084148728,2.57025440313112,2.58180039138943,2.59334637964775,2.60489236790607,2.61643835616438,2.6279843444227,2.63953033268102,2.65107632093933,2.66262230919765,2.67416829745597,2.68571428571429,2.6972602739726,2.70880626223092,2.72035225048924,2.73189823874755,2.74344422700587,2.75499021526419,2.7665362035225,2.77808219178082,2.78962818003914,2.80117416829746,2.81272015655577,2.82426614481409,2.83581213307241,2.84735812133072,2.85890410958904,2.87045009784736,2.88199608610568,2.89354207436399,2.90508806262231,2.91663405088063,2.92818003913894,2.93972602739726,2.95127201565558,2.96281800391389,2.97436399217221,2.98590998043053,2.99745596868885,3.00900195694716,3.02054794520548,3.0320939334638,3.04363992172211,3.05518590998043,3.06673189823875,3.07827788649706,3.08982387475538,3.1013698630137,3.11291585127202,3.12446183953033,3.13600782778865,3.14755381604697,3.15909980430528,3.1706457925636,3.18219178082192,3.19373776908024,3.20528375733855,3.21682974559687,3.22837573385519,3.2399217221135,3.25146771037182,3.26301369863014,3.27455968688845,3.28610567514677,3.29765166340509,3.30919765166341,3.32074363992172,3.33228962818004,3.34383561643836,3.35538160469667,3.36692759295499,3.37847358121331,3.39001956947162,3.40156555772994,3.41311154598826,3.42465753424658,3.43620352250489,3.44774951076321,3.45929549902153,3.47084148727984,3.48238747553816,3.49393346379648,3.50547945205479,3.51702544031311,3.52857142857143,3.54011741682975,3.55166340508806,3.56320939334638,3.5747553816047,3.58630136986301,3.59784735812133,3.60939334637965,3.62093933463796,3.63248532289628,3.6440313111546,3.65557729941292,3.66712328767123,3.67866927592955,3.69021526418787,3.70176125244618,3.7133072407045,3.72485322896282,3.73639921722114,3.74794520547945,3.75949119373777,3.77103718199609,3.7825831702544,3.79412915851272,3.80567514677104,3.81722113502935,3.82876712328767,3.84031311154599,3.85185909980431,3.86340508806262,3.87495107632094,3.88649706457926,3.89804305283757,3.90958904109589,3.92113502935421,3.93268101761252,3.94422700587084,3.95577299412916,3.96731898238748,3.97886497064579,3.99041095890411,4.00195694716243,4.01350293542074,4.02504892367906,4.03659491193738,4.04814090019569,4.05968688845401,4.07123287671233,4.08277886497065,4.09432485322896,4.10587084148728,4.1174168297456,4.12896281800391,4.14050880626223,4.15205479452055,4.16360078277886,4.17514677103718,4.1866927592955,4.19823874755382,4.20978473581213,4.22133072407045,4.23287671232877,4.24442270058708,4.2559686888454,4.26751467710372,4.27906066536204,4.29060665362035,4.30215264187867,4.31369863013699,4.3252446183953,4.33679060665362,4.34833659491194,4.35988258317025,4.37142857142857,4.38297455968689,4.39452054794521,4.40606653620352,4.41761252446184,4.42915851272016,4.44070450097847,4.45225048923679,4.46379647749511,4.47534246575342,4.48688845401174,4.49843444227006,4.50998043052838,4.52152641878669,4.53307240704501,4.54461839530333,4.55616438356164,4.56771037181996,4.57925636007828,4.5908023483366,4.60234833659491,4.61389432485323,4.62544031311155,4.63698630136986,4.64853228962818,4.6600782778865,4.67162426614481,4.68317025440313,4.69471624266145,4.70626223091977,4.71780821917808,4.7293542074364,4.74090019569472,4.75244618395303,4.76399217221135,4.77553816046967,4.78708414872798,4.7986301369863,4.81017612524462,4.82172211350294,4.83326810176125,4.84481409001957,4.85636007827789,4.8679060665362,4.87945205479452,4.89099804305284,4.90254403131116,4.91409001956947,4.92563600782779,4.93718199608611,4.94872798434442,4.96027397260274,4.97181996086106,4.98336594911937,4.99491193737769,5.00645792563601,5.01800391389432,5.02954990215264,5.04109589041096,5.05264187866928,5.06418786692759,5.07573385518591,5.08727984344423,5.09882583170254,5.11037181996086,5.12191780821918,5.1334637964775,5.14500978473581,5.15655577299413,5.16810176125245,5.17964774951076,5.19119373776908,5.2027397260274,5.21428571428571,5.22583170254403,5.23737769080235,5.24892367906067,5.26046966731898,5.2720156555773,5.28356164383562,5.29510763209393,5.30665362035225,5.31819960861057,5.32974559686888,5.3412915851272,5.35283757338552,5.36438356164384,5.37592954990215,5.38747553816047,5.39902152641879,5.4105675146771,5.42211350293542,5.43365949119374,5.44520547945205,5.45675146771037,5.46829745596869,5.47984344422701,5.49138943248532,5.50293542074364,5.51448140900196,5.52602739726027,5.53757338551859,5.54911937377691,5.56066536203523,5.57221135029354,5.58375733855186,5.59530332681018,5.60684931506849,5.61839530332681,5.62994129158513,5.64148727984344,5.65303326810176,5.66457925636008,5.6761252446184,5.68767123287671,5.69921722113503,5.71076320939335,5.72230919765166,5.73385518590998,5.7454011741683,5.75694716242661,5.76849315068493,5.78003913894325,5.79158512720157,5.80313111545988,5.8146771037182,5.82622309197652,5.83776908023483,5.84931506849315,5.86086105675147,5.87240704500979,5.8839530332681,5.89549902152642,5.90704500978474,5.91859099804305,5.93013698630137,5.94168297455969,5.953228962818,5.96477495107632,5.97632093933464,5.98786692759295,5.99941291585127,6.01095890410959,6.02250489236791,6.03405088062622,6.04559686888454,6.05714285714286,6.06868884540117,6.08023483365949,6.09178082191781,6.10332681017613,6.11487279843444,6.12641878669276,6.13796477495108,6.14951076320939,6.16105675146771,6.17260273972603,6.18414872798434,6.19569471624266,6.20724070450098,6.2187866927593,6.23033268101761,6.24187866927593,6.25342465753425,6.26497064579256,6.27651663405088,6.2880626223092,6.29960861056751,6.31115459882583,6.32270058708415,6.33424657534247,6.34579256360078,6.3573385518591,6.36888454011742,6.38043052837573,6.39197651663405,6.40352250489237,6.41506849315069,6.426614481409,6.43816046966732,6.44970645792564,6.46125244618395,6.47279843444227,6.48434442270059,6.4958904109589,6.50743639921722,6.51898238747554,6.53052837573386,6.54207436399217,6.55362035225049,6.56516634050881,6.57671232876712,6.58825831702544,6.59980430528376,6.61135029354207,6.62289628180039,6.63444227005871,6.64598825831703,6.65753424657534,6.66908023483366,6.68062622309198,6.69217221135029,6.70371819960861,6.71526418786693,6.72681017612524,6.73835616438356,6.74990215264188,6.7614481409002,6.77299412915851,6.78454011741683,6.79608610567515,6.80763209393346,6.81917808219178,6.8307240704501,6.84227005870842,6.85381604696673,6.86536203522505,6.87690802348337,6.88845401174168,6.9,6.9,6.9,6.88845401174168,6.87690802348337,6.86536203522505,6.85381604696673,6.84227005870842,6.8307240704501,6.81917808219178,6.80763209393346,6.79608610567515,6.78454011741683,6.77299412915851,6.7614481409002,6.74990215264188,6.73835616438356,6.72681017612524,6.71526418786693,6.70371819960861,6.69217221135029,6.68062622309198,6.66908023483366,6.65753424657534,6.64598825831703,6.63444227005871,6.62289628180039,6.61135029354207,6.59980430528376,6.58825831702544,6.57671232876712,6.56516634050881,6.55362035225049,6.54207436399217,6.53052837573386,6.51898238747554,6.50743639921722,6.4958904109589,6.48434442270059,6.47279843444227,6.46125244618395,6.44970645792564,6.43816046966732,6.426614481409,6.41506849315069,6.40352250489237,6.39197651663405,6.38043052837573,6.36888454011742,6.3573385518591,6.34579256360078,6.33424657534247,6.32270058708415,6.31115459882583,6.29960861056751,6.2880626223092,6.27651663405088,6.26497064579256,6.25342465753425,6.24187866927593,6.23033268101761,6.2187866927593,6.20724070450098,6.19569471624266,6.18414872798434,6.17260273972603,6.16105675146771,6.14951076320939,6.13796477495108,6.12641878669276,6.11487279843444,6.10332681017613,6.09178082191781,6.08023483365949,6.06868884540117,6.05714285714286,6.04559686888454,6.03405088062622,6.02250489236791,6.01095890410959,5.99941291585127,5.98786692759295,5.97632093933464,5.96477495107632,5.953228962818,5.94168297455969,5.93013698630137,5.91859099804305,5.90704500978474,5.89549902152642,5.8839530332681,5.87240704500979,5.86086105675147,5.84931506849315,5.83776908023483,5.82622309197652,5.8146771037182,5.80313111545988,5.79158512720157,5.78003913894325,5.76849315068493,5.75694716242661,5.7454011741683,5.73385518590998,5.72230919765166,5.71076320939335,5.69921722113503,5.68767123287671,5.6761252446184,5.66457925636008,5.65303326810176,5.64148727984344,5.62994129158513,5.61839530332681,5.60684931506849,5.59530332681018,5.58375733855186,5.57221135029354,5.56066536203523,5.54911937377691,5.53757338551859,5.52602739726027,5.51448140900196,5.50293542074364,5.49138943248532,5.47984344422701,5.46829745596869,5.45675146771037,5.44520547945205,5.43365949119374,5.42211350293542,5.4105675146771,5.39902152641879,5.38747553816047,5.37592954990215,5.36438356164384,5.35283757338552,5.3412915851272,5.32974559686888,5.31819960861057,5.30665362035225,5.29510763209393,5.28356164383562,5.2720156555773,5.26046966731898,5.24892367906067,5.23737769080235,5.22583170254403,5.21428571428571,5.2027397260274,5.19119373776908,5.17964774951076,5.16810176125245,5.15655577299413,5.14500978473581,5.1334637964775,5.12191780821918,5.11037181996086,5.09882583170254,5.08727984344423,5.07573385518591,5.06418786692759,5.05264187866928,5.04109589041096,5.02954990215264,5.01800391389432,5.00645792563601,4.99491193737769,4.98336594911937,4.97181996086106,4.96027397260274,4.94872798434442,4.93718199608611,4.92563600782779,4.91409001956947,4.90254403131116,4.89099804305284,4.87945205479452,4.8679060665362,4.85636007827789,4.84481409001957,4.83326810176125,4.82172211350294,4.81017612524462,4.7986301369863,4.78708414872798,4.77553816046967,4.76399217221135,4.75244618395303,4.74090019569472,4.7293542074364,4.71780821917808,4.70626223091977,4.69471624266145,4.68317025440313,4.67162426614481,4.6600782778865,4.64853228962818,4.63698630136986,4.62544031311155,4.61389432485323,4.60234833659491,4.5908023483366,4.57925636007828,4.56771037181996,4.55616438356164,4.54461839530333,4.53307240704501,4.52152641878669,4.50998043052838,4.49843444227006,4.48688845401174,4.47534246575342,4.46379647749511,4.45225048923679,4.44070450097847,4.42915851272016,4.41761252446184,4.40606653620352,4.39452054794521,4.38297455968689,4.37142857142857,4.35988258317025,4.34833659491194,4.33679060665362,4.3252446183953,4.31369863013699,4.30215264187867,4.29060665362035,4.27906066536204,4.26751467710372,4.2559686888454,4.24442270058708,4.23287671232877,4.22133072407045,4.20978473581213,4.19823874755382,4.1866927592955,4.17514677103718,4.16360078277886,4.15205479452055,4.14050880626223,4.12896281800391,4.1174168297456,4.10587084148728,4.09432485322896,4.08277886497065,4.07123287671233,4.05968688845401,4.04814090019569,4.03659491193738,4.02504892367906,4.01350293542074,4.00195694716243,3.99041095890411,3.97886497064579,3.96731898238748,3.95577299412916,3.94422700587084,3.93268101761252,3.92113502935421,3.90958904109589,3.89804305283757,3.88649706457926,3.87495107632094,3.86340508806262,3.85185909980431,3.84031311154599,3.82876712328767,3.81722113502935,3.80567514677104,3.79412915851272,3.7825831702544,3.77103718199609,3.75949119373777,3.74794520547945,3.73639921722114,3.72485322896282,3.7133072407045,3.70176125244618,3.69021526418787,3.67866927592955,3.66712328767123,3.65557729941292,3.6440313111546,3.63248532289628,3.62093933463796,3.60939334637965,3.59784735812133,3.58630136986301,3.5747553816047,3.56320939334638,3.55166340508806,3.54011741682975,3.52857142857143,3.51702544031311,3.50547945205479,3.49393346379648,3.48238747553816,3.47084148727984,3.45929549902153,3.44774951076321,3.43620352250489,3.42465753424658,3.41311154598826,3.40156555772994,3.39001956947162,3.37847358121331,3.36692759295499,3.35538160469667,3.34383561643836,3.33228962818004,3.32074363992172,3.30919765166341,3.29765166340509,3.28610567514677,3.27455968688845,3.26301369863014,3.25146771037182,3.2399217221135,3.22837573385519,3.21682974559687,3.20528375733855,3.19373776908024,3.18219178082192,3.1706457925636,3.15909980430528,3.14755381604697,3.13600782778865,3.12446183953033,3.11291585127202,3.1013698630137,3.08982387475538,3.07827788649706,3.06673189823875,3.05518590998043,3.04363992172211,3.0320939334638,3.02054794520548,3.00900195694716,2.99745596868885,2.98590998043053,2.97436399217221,2.96281800391389,2.95127201565558,2.93972602739726,2.92818003913894,2.91663405088063,2.90508806262231,2.89354207436399,2.88199608610568,2.87045009784736,2.85890410958904,2.84735812133072,2.83581213307241,2.82426614481409,2.81272015655577,2.80117416829746,2.78962818003914,2.77808219178082,2.7665362035225,2.75499021526419,2.74344422700587,2.73189823874755,2.72035225048924,2.70880626223092,2.6972602739726,2.68571428571429,2.67416829745597,2.66262230919765,2.65107632093933,2.63953033268102,2.6279843444227,2.61643835616438,2.60489236790607,2.59334637964775,2.58180039138943,2.57025440313112,2.5587084148728,2.54716242661448,2.53561643835616,2.52407045009785,2.51252446183953,2.50097847358121,2.4894324853229,2.47788649706458,2.46634050880626,2.45479452054795,2.44324853228963,2.43170254403131,2.42015655577299,2.40861056751468,2.39706457925636,2.38551859099804,2.37397260273973,2.36242661448141,2.35088062622309,2.33933463796478,2.32778864970646,2.31624266144814,2.30469667318982,2.29315068493151,2.28160469667319,2.27005870841487,2.25851272015656,2.24696673189824,2.23542074363992,2.2238747553816,2.21232876712329,2.20078277886497,2.18923679060665,2.17769080234834,2.16614481409002,2.1545988258317,2.14305283757339,2.13150684931507,2.11996086105675,2.10841487279843,2.09686888454012,2.0853228962818,2.07377690802348,2.06223091976517,2.05068493150685,2.03913894324853,2.02759295499022,2.0160469667319,2.00450097847358,1.99295499021526,1.98140900195695,1.96986301369863,1.95831702544031,1.946771037182,1.93522504892368,1.92367906066536,1.91213307240705,1.90058708414873,1.88904109589041,1.87749510763209,1.86594911937378,1.85440313111546,1.84285714285714,1.83131115459883,1.81976516634051,1.80821917808219,1.79667318982387,1.78512720156556,1.77358121330724,1.76203522504892,1.75048923679061,1.73894324853229,1.72739726027397,1.71585127201566,1.70430528375734,1.69275929549902,1.6812133072407,1.66966731898239,1.65812133072407,1.64657534246575,1.63502935420744,1.62348336594912,1.6119373776908,1.60039138943249,1.58884540117417,1.57729941291585,1.56575342465753,1.55420743639922,1.5426614481409,1.53111545988258,1.51956947162427,1.50802348336595,1.49647749510763,1.48493150684931,1.473385518591,1.46183953033268,1.45029354207436,1.43874755381605,1.42720156555773,1.41565557729941,1.4041095890411,1.39256360078278,1.38101761252446,1.36947162426614,1.35792563600783,1.34637964774951,1.33483365949119,1.32328767123288,1.31174168297456,1.30019569471624,1.28864970645793,1.27710371819961,1.26555772994129,1.25401174168297,1.24246575342466,1.23091976516634,1.21937377690802,1.20782778864971,1.19628180039139,1.18473581213307,1.17318982387476,1.16164383561644,1.15009784735812,1.1385518590998,1.12700587084149,1.11545988258317,1.10391389432485,1.09236790606654,1.08082191780822,1.0692759295499,1.05772994129159,1.04618395303327,1.03463796477495,1.02309197651663,1.01154598825832,1,1],"y":[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0.101996862441166,0.105634024966893,0.109215424589166,0.112743220349615,0.11621059854995,0.119608627225697,0.122894905816137,0.126092292836774,0.129195694202423,0.132200166580964,0.135060192318402,0.137798918952927,0.140422819257167,0.142929868497606,0.145288082910024,0.147504251326372,0.1496025658843,0.151584643140614,0.153436956576737,0.155149359540491,0.156761320230362,0.158278234386759,0.159703174010678,0.161019688178542,0.162272970731151,0.163471666064559,0.164624583209088,0.165736478007498,0.166835819046194,0.167935665373943,0.169046833548677,0.170192670962244,0.171395314495607,0.172660988788519,0.174000979318604,0.175442712747477,0.177024889633891,0.178731605928383,0.18057301144304,0.182565467766611,0.184785949137251,0.187183308127156,0.189765189676406,0.192539046477643,0.195591843041304,0.198872657473276,0.202371342296019,0.206091915484468,0.210099080456263,0.214378351285116,0.218889328630874,0.223632380220904,0.228643113056214,0.233950255919104,0.239483683398394,0.245240516162785,0.251227181890275,0.257511988313714,0.264001620854678,0.270690831023312,0.277574261126014,0.284712188262788,0.292034680201435,0.299523168273741,0.307171089463508,0.315008839010959,0.323007801598205,0.331134928386568,0.339383360678282,0.347762274567568,0.356269927108321,0.364867535510898,0.373548370892534,0.382308925570656,0.391158517981255,0.400060438609528,0.409007881639393,0.417994015219607,0.427017364660271,0.436055365654781,0.445098806759623,0.454139981678903,0.463162979601972,0.47215135856716,0.481098248776854,0.489994224228255,0.498816091831308,0.507532644570251,0.516150226905375,0.524657321872641,0.533034140336448,0.541208864220896,0.549216454453603,0.557043904181735,0.564678162432848,0.572012423516872,0.579094572544359,0.585923514045739,0.592486675261359,0.598685828815022,0.604530768249182,0.610058292144527,0.615259061054111,0.62006387560996,0.624422567699444,0.628423829316253,0.632064440803927,0.63531920560187,0.638071007967135,0.640463552588198,0.642501769863758,0.644191123712188,0.645412178431763,0.646301818866644,0.646889559615017,0.647189193999976,0.647148236922841,0.646837415051442,0.646318341864944,0.645610952674723,0.644713928917646,0.643661853155078,0.642520038716625,0.641310366653101,0.640054416820304,0.638791389773416,0.637556902708951,0.636369498807985,0.635247346666177,0.634255767586117,0.633386584452355,0.632646030815736,0.632043604625423,0.631633627768132,0.631402252726073,0.631325305588622,0.631400506313823,0.631648847482493,0.63206323487114,0.632591480021146,0.633219169694736,0.633934630032552,0.634715645026951,0.635508025543341,0.636287831583358,0.637030668228843,0.637657348581741,0.638146875178529,0.638473792999921,0.638609605598286,0.638440110196538,0.637942618211776,0.637124686561768,0.63595983813856,0.634341890534245,0.632191718221403,0.629587501148821,0.626510405264725,0.622901324981256,0.618582917740457,0.613727664551171,0.608328229327759,0.602378054358976,0.595642374001121,0.588336170032908,0.580482289056077,0.572087001746926,0.563004202974188,0.553346216706346,0.543204536309163,0.532596819053539,0.52146377269357,0.509830214054113,0.497831971794702,0.485494675629895,0.472822575073239,0.459798300514425,0.446562575414263,0.433144537410634,0.41957340581111,0.405857952969404,0.392091695552864,0.378305361786526,0.364526844268036,0.350813743124877,0.337209699801383,0.323731372686155,0.310401732196737,0.297286176233739,0.284437002435204,0.271826821623025,0.259471485144399,0.247412911891779,0.235753905564928,0.224406332957708,0.213378291809191,0.20267740905252,0.192456394838071,0.182584670864034,0.1730566541663,0.163873085261821,0.155141174797514,0.146792376902949,0.138777994304612,0.131093670851232,0.123798872024353,0.116890982027371,0.110285331473834,0.103974625094998,0.0979774179709013,0.0923470152402705,0.0869751236015474,0.081853512363587,0.0769739400597586,0.0724199212408306,0.0680820561411753,0.0639494301530343,0.0600144301965125,0.0563252194698922,0.052831153539768,0.0495028089435985,0.0463339380062595,0.04334823382544,0.0405369128575336,0.0378597744632279,0.0353120661935617,0.0329009344844679,0.0306463369909195,0.0285020814622748,0.0264646419783394,0.0245305577495023,0.0227367975490825,0.0210362233958028,0.019425281441751,0.0179012815188586,0.0164872346354775,0.015160983096673,0.0139104168388491,0.012733245765648,0.0116415902775909,0.0106314182890446,0.00968457741365953,0.00879895705205037,0.00797834173649686,0.00723133011219629,0.00653606504211874,0.00589054942112726,0.00529279339458479,0.00475886758468263,0.00426588953713499,0.00381188806726599,0.00339503005437656,0.00302386313341143,0.00268742018060747,0.00238021855724387,0.00210065794284578,0.00185223497860043,0.00163173258187025,0.00143215344036197,0.00125218821642621,0.00109234594140847,0.000953781756466341,0.000829465666641791,0.00071839693470986,0.000619623973111173,0.000536225163702669,0.000462050301839955,0.000396384829449577,0.000338530059359107,0.000289761185716105,0.000247393296829401,0.000210222534166027,0.000177783066428223,0.000150413839124369,0.000127256413627336,0.00010711711382397,8.97044555665802e-05,7.4971324870726e-05,6.28626977324571e-05,5.24208685854706e-05,4.3474415423933e-05,3.58673563142811e-05,2.98117271960112e-05,2.46316196521771e-05,2.02321864196175e-05,1.65219646736019e-05,1.35688248028498e-05,1.11102345339835e-05,9.03969917077735e-06,7.30941440855155e-06,5.92586644171759e-06,4.80957529209139e-06,3.87697876127707e-06,3.10447842500538e-06,2.48265777164717e-06,1.99785314586372e-06,1.59586467770312e-06,1.26568736990868e-06,9.97563023027435e-07,7.96192194238555e-07,6.30379166017733e-07,4.95276811330674e-07,3.8626844773254e-07,3.04364884201921e-07,2.38920173390118e-07,1.85997962534255e-07,1.43660198012217e-07,1.11587536916964e-07,8.68741591768211e-08,6.702900092085e-08,5.12814195274221e-08,3.92281851071866e-08,3.03008152264678e-08,2.31774951652482e-08,1.75681578210364e-08,1.32205868959588e-08,1.01362757039383e-08,7.6890352281363e-09,5.77564040083377e-09,4.29893219154891e-09,3.25155295464182e-09,2.44696281984614e-09,1.82198468018241e-09,1.34341645208227e-09,1.00002459278337e-09,7.46923869808501e-10,5.5146717507433e-10,4.02896557270693e-10,2.94812554033432e-10,2.18652935002059e-10,1.60132894363258e-10,1.1595190888172e-10,8.32895492264637e-11,6.13750900966023e-11,4.46043613783196e-11,3.20205405033596e-11,2.27336937219184e-11,1.65159893961675e-11,1.19166389846729e-11,8.48413055962574e-12,5.96893378500528e-12,4.25990941738913e-12,3.05313906758314e-12,2.15658848726133e-12,1.50393146258026e-12,1.0528801794381e-12,7.50056247116917e-13,5.25828166182545e-13,3.63552305597568e-13,2.49266953311956e-13,1.76627043845019e-13,1.22968085387504e-13,8.43170385795513e-14,5.70290461881174e-14,3.98488431812502e-14,2.75550295045735e-14,1.87318874099431e-14,1.25535511464262e-14,8.54563285966142e-15,5.85317908428149e-15,3.97963003445539e-15,2.71115893776356e-15,1.82787846821179e-15,1.23057815154967e-15,8.20652224548616e-16,5.45241345967415e-16,3.38346821567909e-16,2.08996378021915e-16,1.32944310425063e-16,8.25469262764648e-17,3.41586953318161e-17,2.3548719721585e-17,8.99977531116934e-18,0,0,0,0,0,0,0,0,6.22320343774891e-18,1.83108746680628e-17,3.42258380834094e-17,8.95058288750118e-18,0,9.40482564411943e-18,3.48179990345522e-17,3.747049293711e-17,1.66141424215307e-17,0,2.60131415509245e-18,2.38212653755555e-17,1.04699346352402e-17,0,2.10719609502805e-18,1.27171717052595e-17,9.15310805397616e-18,3.84812024886061e-18,0,0,0,3.49404316378752e-17,6.38085278440842e-17,4.76673334625331e-17,2.64473822420703e-17,1.69193690391933e-17,1.8404600955288e-17,2.71260751406836e-17,2.18210873355679e-17,7.91493516791328e-18,5.33335216495573e-18,1.6976097758873e-17,3.28910611742196e-17,1.58790627594951e-17,3.86894831268837e-18,0,0,0,0,0,0,0,1.55059384348028e-17,1.87852615759925e-17,0,0,0,2.6458480186501e-18,5.92695198404868e-18,6.21964178933035e-19,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,5.67016283461767e-19,5.87200408857767e-18,2.70079591412121e-18,0,0,0,2.32068374926343e-17,6.17616221171552e-17,8.20354063443197e-17,2.36805404880476e-17,1.07274076971857e-17,2.05818175378127e-17,4.35785935679601e-17,6.85314267737131e-17,3.67014999430198e-17,1.24995739467598e-17,1.74407545474064e-17,5.27802101330873e-17,2.6255271107509e-17,5.0337571193429e-17,6.42222113847391e-17,6.02584398731334e-17,8.67833788987117e-17,5.78618447079003e-17,5.05932634311659e-17,5.97445906609331e-17,4.91346150507017e-17,7.87151109879912e-17,8.34258150614969e-17,7.98398264346932e-17,8.99614394566634e-17,7.93514638464318e-17,8.39141776497584e-17,6.86129582048154e-17,2.44942028023656e-17,6.16291174381744e-17,7.35853809012026e-17,7.37652971796407e-17,7.3103457697261e-17,9.43234089177236e-17,6.49465350586025e-17,5.74696731435896e-17,6.40613423459195e-17,4.28413911254569e-17,5.66423060622802e-17,7.25572694776269e-17,7.45908834367411e-17,4.81418913485907e-17,2.55956931768496e-17,2.09681303805257e-17,2.36206242830835e-17,2.52640540016845e-17,1.73065722940109e-17,0],"text":["density: 1.730657e-17<br />Petal.Length: 1.000000","density: 2.526405e-17<br />Petal.Length: 1.011546","density: 2.362062e-17<br />Petal.Length: 1.023092","density: 2.096813e-17<br />Petal.Length: 1.034638","density: 2.559569e-17<br />Petal.Length: 1.046184","density: 4.814189e-17<br />Petal.Length: 1.057730","density: 7.459088e-17<br />Petal.Length: 1.069276","density: 7.255727e-17<br />Petal.Length: 1.080822","density: 5.664231e-17<br />Petal.Length: 1.092368","density: 4.284139e-17<br />Petal.Length: 1.103914","density: 6.406134e-17<br />Petal.Length: 1.115460","density: 5.746967e-17<br />Petal.Length: 1.127006","density: 6.494654e-17<br />Petal.Length: 1.138552","density: 9.432341e-17<br />Petal.Length: 1.150098","density: 7.310346e-17<br />Petal.Length: 1.161644","density: 7.376530e-17<br />Petal.Length: 1.173190","density: 7.358538e-17<br />Petal.Length: 1.184736","density: 6.162912e-17<br />Petal.Length: 1.196282","density: 2.449420e-17<br />Petal.Length: 1.207828","density: 6.861296e-17<br />Petal.Length: 1.219374","density: 8.391418e-17<br />Petal.Length: 1.230920","density: 7.935146e-17<br />Petal.Length: 1.242466","density: 8.996144e-17<br />Petal.Length: 1.254012","density: 7.983983e-17<br />Petal.Length: 1.265558","density: 8.342582e-17<br />Petal.Length: 1.277104","density: 7.871511e-17<br />Petal.Length: 1.288650","density: 4.913462e-17<br />Petal.Length: 1.300196","density: 5.974459e-17<br />Petal.Length: 1.311742","density: 5.059326e-17<br />Petal.Length: 1.323288","density: 5.786184e-17<br />Petal.Length: 1.334834","density: 8.678338e-17<br />Petal.Length: 1.346380","density: 6.025844e-17<br />Petal.Length: 1.357926","density: 6.422221e-17<br />Petal.Length: 1.369472","density: 5.033757e-17<br />Petal.Length: 1.381018","density: 2.625527e-17<br />Petal.Length: 1.392564","density: 5.278021e-17<br />Petal.Length: 1.404110","density: 1.744075e-17<br />Petal.Length: 1.415656","density: 1.249957e-17<br />Petal.Length: 1.427202","density: 3.670150e-17<br />Petal.Length: 1.438748","density: 6.853143e-17<br />Petal.Length: 1.450294","density: 4.357859e-17<br />Petal.Length: 1.461840","density: 2.058182e-17<br />Petal.Length: 1.473386","density: 1.072741e-17<br />Petal.Length: 1.484932","density: 2.368054e-17<br />Petal.Length: 1.496477","density: 8.203541e-17<br />Petal.Length: 1.508023","density: 6.176162e-17<br />Petal.Length: 1.519569","density: 2.320684e-17<br />Petal.Length: 1.531115","density: 0.000000e+00<br />Petal.Length: 1.542661","density: 0.000000e+00<br />Petal.Length: 1.554207","density: 0.000000e+00<br />Petal.Length: 1.565753","density: 2.700796e-18<br />Petal.Length: 1.577299","density: 5.872004e-18<br />Petal.Length: 1.588845","density: 5.670163e-19<br />Petal.Length: 1.600391","density: 0.000000e+00<br />Petal.Length: 1.611937","density: 0.000000e+00<br />Petal.Length: 1.623483","density: 0.000000e+00<br />Petal.Length: 1.635029","density: 0.000000e+00<br />Petal.Length: 1.646575","density: 0.000000e+00<br />Petal.Length: 1.658121","density: 0.000000e+00<br />Petal.Length: 1.669667","density: 0.000000e+00<br />Petal.Length: 1.681213","density: 0.000000e+00<br />Petal.Length: 1.692759","density: 0.000000e+00<br />Petal.Length: 1.704305","density: 0.000000e+00<br />Petal.Length: 1.715851","density: 0.000000e+00<br />Petal.Length: 1.727397","density: 0.000000e+00<br />Petal.Length: 1.738943","density: 0.000000e+00<br />Petal.Length: 1.750489","density: 0.000000e+00<br />Petal.Length: 1.762035","density: 0.000000e+00<br />Petal.Length: 1.773581","density: 0.000000e+00<br />Petal.Length: 1.785127","density: 0.000000e+00<br />Petal.Length: 1.796673","density: 0.000000e+00<br />Petal.Length: 1.808219","density: 0.000000e+00<br />Petal.Length: 1.819765","density: 0.000000e+00<br />Petal.Length: 1.831311","density: 0.000000e+00<br />Petal.Length: 1.842857","density: 0.000000e+00<br />Petal.Length: 1.854403","density: 0.000000e+00<br />Petal.Length: 1.865949","density: 0.000000e+00<br />Petal.Length: 1.877495","density: 0.000000e+00<br />Petal.Length: 1.889041","density: 0.000000e+00<br />Petal.Length: 1.900587","density: 0.000000e+00<br />Petal.Length: 1.912133","density: 0.000000e+00<br />Petal.Length: 1.923679","density: 6.219642e-19<br />Petal.Length: 1.935225","density: 5.926952e-18<br />Petal.Length: 1.946771","density: 2.645848e-18<br />Petal.Length: 1.958317","density: 0.000000e+00<br />Petal.Length: 1.969863","density: 0.000000e+00<br />Petal.Length: 1.981409","density: 0.000000e+00<br />Petal.Length: 1.992955","density: 1.878526e-17<br />Petal.Length: 2.004501","density: 1.550594e-17<br />Petal.Length: 2.016047","density: 0.000000e+00<br />Petal.Length: 2.027593","density: 0.000000e+00<br />Petal.Length: 2.039139","density: 0.000000e+00<br />Petal.Length: 2.050685","density: 0.000000e+00<br />Petal.Length: 2.062231","density: 0.000000e+00<br />Petal.Length: 2.073777","density: 0.000000e+00<br />Petal.Length: 2.085323","density: 0.000000e+00<br />Petal.Length: 2.096869","density: 3.868948e-18<br />Petal.Length: 2.108415","density: 1.587906e-17<br />Petal.Length: 2.119961","density: 3.289106e-17<br />Petal.Length: 2.131507","density: 1.697610e-17<br />Petal.Length: 2.143053","density: 5.333352e-18<br />Petal.Length: 2.154599","density: 7.914935e-18<br />Petal.Length: 2.166145","density: 2.182109e-17<br />Petal.Length: 2.177691","density: 2.712608e-17<br />Petal.Length: 2.189237","density: 1.840460e-17<br />Petal.Length: 2.200783","density: 1.691937e-17<br />Petal.Length: 2.212329","density: 2.644738e-17<br />Petal.Length: 2.223875","density: 4.766733e-17<br />Petal.Length: 2.235421","density: 6.380853e-17<br />Petal.Length: 2.246967","density: 3.494043e-17<br />Petal.Length: 2.258513","density: 0.000000e+00<br />Petal.Length: 2.270059","density: 0.000000e+00<br />Petal.Length: 2.281605","density: 0.000000e+00<br />Petal.Length: 2.293151","density: 3.848120e-18<br />Petal.Length: 2.304697","density: 9.153108e-18<br />Petal.Length: 2.316243","density: 1.271717e-17<br />Petal.Length: 2.327789","density: 2.107196e-18<br />Petal.Length: 2.339335","density: 0.000000e+00<br />Petal.Length: 2.350881","density: 1.046993e-17<br />Petal.Length: 2.362427","density: 2.382127e-17<br />Petal.Length: 2.373973","density: 2.601314e-18<br />Petal.Length: 2.385519","density: 0.000000e+00<br />Petal.Length: 2.397065","density: 1.661414e-17<br />Petal.Length: 2.408611","density: 3.747049e-17<br />Petal.Length: 2.420157","density: 3.481800e-17<br />Petal.Length: 2.431703","density: 9.404826e-18<br />Petal.Length: 2.443249","density: 0.000000e+00<br />Petal.Length: 2.454795","density: 8.950583e-18<br />Petal.Length: 2.466341","density: 3.422584e-17<br />Petal.Length: 2.477886","density: 1.831087e-17<br />Petal.Length: 2.489432","density: 6.223203e-18<br />Petal.Length: 2.500978","density: 0.000000e+00<br />Petal.Length: 2.512524","density: 0.000000e+00<br />Petal.Length: 2.524070","density: 0.000000e+00<br />Petal.Length: 2.535616","density: 0.000000e+00<br />Petal.Length: 2.547162","density: 0.000000e+00<br />Petal.Length: 2.558708","density: 0.000000e+00<br />Petal.Length: 2.570254","density: 0.000000e+00<br />Petal.Length: 2.581800","density: 0.000000e+00<br />Petal.Length: 2.593346","density: 8.999775e-18<br />Petal.Length: 2.604892","density: 2.354872e-17<br />Petal.Length: 2.616438","density: 3.415870e-17<br />Petal.Length: 2.627984","density: 8.254693e-17<br />Petal.Length: 2.639530","density: 1.329443e-16<br />Petal.Length: 2.651076","density: 2.089964e-16<br />Petal.Length: 2.662622","density: 3.383468e-16<br />Petal.Length: 2.674168","density: 5.452413e-16<br />Petal.Length: 2.685714","density: 8.206522e-16<br />Petal.Length: 2.697260","density: 1.230578e-15<br />Petal.Length: 2.708806","density: 1.827878e-15<br />Petal.Length: 2.720352","density: 2.711159e-15<br />Petal.Length: 2.731898","density: 3.979630e-15<br />Petal.Length: 2.743444","density: 5.853179e-15<br />Petal.Length: 2.754990","density: 8.545633e-15<br />Petal.Length: 2.766536","density: 1.255355e-14<br />Petal.Length: 2.778082","density: 1.873189e-14<br />Petal.Length: 2.789628","density: 2.755503e-14<br />Petal.Length: 2.801174","density: 3.984884e-14<br />Petal.Length: 2.812720","density: 5.702905e-14<br />Petal.Length: 2.824266","density: 8.431704e-14<br />Petal.Length: 2.835812","density: 1.229681e-13<br />Petal.Length: 2.847358","density: 1.766270e-13<br />Petal.Length: 2.858904","density: 2.492670e-13<br />Petal.Length: 2.870450","density: 3.635523e-13<br />Petal.Length: 2.881996","density: 5.258282e-13<br />Petal.Length: 2.893542","density: 7.500562e-13<br />Petal.Length: 2.905088","density: 1.052880e-12<br />Petal.Length: 2.916634","density: 1.503931e-12<br />Petal.Length: 2.928180","density: 2.156588e-12<br />Petal.Length: 2.939726","density: 3.053139e-12<br />Petal.Length: 2.951272","density: 4.259909e-12<br />Petal.Length: 2.962818","density: 5.968934e-12<br />Petal.Length: 2.974364","density: 8.484131e-12<br />Petal.Length: 2.985910","density: 1.191664e-11<br />Petal.Length: 2.997456","density: 1.651599e-11<br />Petal.Length: 3.009002","density: 2.273369e-11<br />Petal.Length: 3.020548","density: 3.202054e-11<br />Petal.Length: 3.032094","density: 4.460436e-11<br />Petal.Length: 3.043640","density: 6.137509e-11<br />Petal.Length: 3.055186","density: 8.328955e-11<br />Petal.Length: 3.066732","density: 1.159519e-10<br />Petal.Length: 3.078278","density: 1.601329e-10<br />Petal.Length: 3.089824","density: 2.186529e-10<br />Petal.Length: 3.101370","density: 2.948126e-10<br />Petal.Length: 3.112916","density: 4.028966e-10<br />Petal.Length: 3.124462","density: 5.514672e-10<br />Petal.Length: 3.136008","density: 7.469239e-10<br />Petal.Length: 3.147554","density: 1.000025e-09<br />Petal.Length: 3.159100","density: 1.343416e-09<br />Petal.Length: 3.170646","density: 1.821985e-09<br />Petal.Length: 3.182192","density: 2.446963e-09<br />Petal.Length: 3.193738","density: 3.251553e-09<br />Petal.Length: 3.205284","density: 4.298932e-09<br />Petal.Length: 3.216830","density: 5.775640e-09<br />Petal.Length: 3.228376","density: 7.689035e-09<br />Petal.Length: 3.239922","density: 1.013628e-08<br />Petal.Length: 3.251468","density: 1.322059e-08<br />Petal.Length: 3.263014","density: 1.756816e-08<br />Petal.Length: 3.274560","density: 2.317750e-08<br />Petal.Length: 3.286106","density: 3.030082e-08<br />Petal.Length: 3.297652","density: 3.922819e-08<br />Petal.Length: 3.309198","density: 5.128142e-08<br />Petal.Length: 3.320744","density: 6.702900e-08<br />Petal.Length: 3.332290","density: 8.687416e-08<br />Petal.Length: 3.343836","density: 1.115875e-07<br />Petal.Length: 3.355382","density: 1.436602e-07<br />Petal.Length: 3.366928","density: 1.859980e-07<br />Petal.Length: 3.378474","density: 2.389202e-07<br />Petal.Length: 3.390020","density: 3.043649e-07<br />Petal.Length: 3.401566","density: 3.862684e-07<br />Petal.Length: 3.413112","density: 4.952768e-07<br />Petal.Length: 3.424658","density: 6.303792e-07<br />Petal.Length: 3.436204","density: 7.961922e-07<br />Petal.Length: 3.447750","density: 9.975630e-07<br />Petal.Length: 3.459295","density: 1.265687e-06<br />Petal.Length: 3.470841","density: 1.595865e-06<br />Petal.Length: 3.482387","density: 1.997853e-06<br />Petal.Length: 3.493933","density: 2.482658e-06<br />Petal.Length: 3.505479","density: 3.104478e-06<br />Petal.Length: 3.517025","density: 3.876979e-06<br />Petal.Length: 3.528571","density: 4.809575e-06<br />Petal.Length: 3.540117","density: 5.925866e-06<br />Petal.Length: 3.551663","density: 7.309414e-06<br />Petal.Length: 3.563209","density: 9.039699e-06<br />Petal.Length: 3.574755","density: 1.111023e-05<br />Petal.Length: 3.586301","density: 1.356882e-05<br />Petal.Length: 3.597847","density: 1.652196e-05<br />Petal.Length: 3.609393","density: 2.023219e-05<br />Petal.Length: 3.620939","density: 2.463162e-05<br />Petal.Length: 3.632485","density: 2.981173e-05<br />Petal.Length: 3.644031","density: 3.586736e-05<br />Petal.Length: 3.655577","density: 4.347442e-05<br />Petal.Length: 3.667123","density: 5.242087e-05<br />Petal.Length: 3.678669","density: 6.286270e-05<br />Petal.Length: 3.690215","density: 7.497132e-05<br />Petal.Length: 3.701761","density: 8.970446e-05<br />Petal.Length: 3.713307","density: 1.071171e-04<br />Petal.Length: 3.724853","density: 1.272564e-04<br />Petal.Length: 3.736399","density: 1.504138e-04<br />Petal.Length: 3.747945","density: 1.777831e-04<br />Petal.Length: 3.759491","density: 2.102225e-04<br />Petal.Length: 3.771037","density: 2.473933e-04<br />Petal.Length: 3.782583","density: 2.897612e-04<br />Petal.Length: 3.794129","density: 3.385301e-04<br />Petal.Length: 3.805675","density: 3.963848e-04<br />Petal.Length: 3.817221","density: 4.620503e-04<br />Petal.Length: 3.828767","density: 5.362252e-04<br />Petal.Length: 3.840313","density: 6.196240e-04<br />Petal.Length: 3.851859","density: 7.183969e-04<br />Petal.Length: 3.863405","density: 8.294657e-04<br />Petal.Length: 3.874951","density: 9.537818e-04<br />Petal.Length: 3.886497","density: 1.092346e-03<br />Petal.Length: 3.898043","density: 1.252188e-03<br />Petal.Length: 3.909589","density: 1.432153e-03<br />Petal.Length: 3.921135","density: 1.631733e-03<br />Petal.Length: 3.932681","density: 1.852235e-03<br />Petal.Length: 3.944227","density: 2.100658e-03<br />Petal.Length: 3.955773","density: 2.380219e-03<br />Petal.Length: 3.967319","density: 2.687420e-03<br />Petal.Length: 3.978865","density: 3.023863e-03<br />Petal.Length: 3.990411","density: 3.395030e-03<br />Petal.Length: 4.001957","density: 3.811888e-03<br />Petal.Length: 4.013503","density: 4.265890e-03<br />Petal.Length: 4.025049","density: 4.758868e-03<br />Petal.Length: 4.036595","density: 5.292793e-03<br />Petal.Length: 4.048141","density: 5.890549e-03<br />Petal.Length: 4.059687","density: 6.536065e-03<br />Petal.Length: 4.071233","density: 7.231330e-03<br />Petal.Length: 4.082779","density: 7.978342e-03<br />Petal.Length: 4.094325","density: 8.798957e-03<br />Petal.Length: 4.105871","density: 9.684577e-03<br />Petal.Length: 4.117417","density: 1.063142e-02<br />Petal.Length: 4.128963","density: 1.164159e-02<br />Petal.Length: 4.140509","density: 1.273325e-02<br />Petal.Length: 4.152055","density: 1.391042e-02<br />Petal.Length: 4.163601","density: 1.516098e-02<br />Petal.Length: 4.175147","density: 1.648723e-02<br />Petal.Length: 4.186693","density: 1.790128e-02<br />Petal.Length: 4.198239","density: 1.942528e-02<br />Petal.Length: 4.209785","density: 2.103622e-02<br />Petal.Length: 4.221331","density: 2.273680e-02<br />Petal.Length: 4.232877","density: 2.453056e-02<br />Petal.Length: 4.244423","density: 2.646464e-02<br />Petal.Length: 4.255969","density: 2.850208e-02<br />Petal.Length: 4.267515","density: 3.064634e-02<br />Petal.Length: 4.279061","density: 3.290093e-02<br />Petal.Length: 4.290607","density: 3.531207e-02<br />Petal.Length: 4.302153","density: 3.785977e-02<br />Petal.Length: 4.313699","density: 4.053691e-02<br />Petal.Length: 4.325245","density: 4.334823e-02<br />Petal.Length: 4.336791","density: 4.633394e-02<br />Petal.Length: 4.348337","density: 4.950281e-02<br />Petal.Length: 4.359883","density: 5.283115e-02<br />Petal.Length: 4.371429","density: 5.632522e-02<br />Petal.Length: 4.382975","density: 6.001443e-02<br />Petal.Length: 4.394521","density: 6.394943e-02<br />Petal.Length: 4.406067","density: 6.808206e-02<br />Petal.Length: 4.417613","density: 7.241992e-02<br />Petal.Length: 4.429159","density: 7.697394e-02<br />Petal.Length: 4.440705","density: 8.185351e-02<br />Petal.Length: 4.452250","density: 8.697512e-02<br />Petal.Length: 4.463796","density: 9.234702e-02<br />Petal.Length: 4.475342","density: 9.797742e-02<br />Petal.Length: 4.486888","density: 1.039746e-01<br />Petal.Length: 4.498434","density: 1.102853e-01<br />Petal.Length: 4.509980","density: 1.168910e-01<br />Petal.Length: 4.521526","density: 1.237989e-01<br />Petal.Length: 4.533072","density: 1.310937e-01<br />Petal.Length: 4.544618","density: 1.387780e-01<br />Petal.Length: 4.556164","density: 1.467924e-01<br />Petal.Length: 4.567710","density: 1.551412e-01<br />Petal.Length: 4.579256","density: 1.638731e-01<br />Petal.Length: 4.590802","density: 1.730567e-01<br />Petal.Length: 4.602348","density: 1.825847e-01<br />Petal.Length: 4.613894","density: 1.924564e-01<br />Petal.Length: 4.625440","density: 2.026774e-01<br />Petal.Length: 4.636986","density: 2.133783e-01<br />Petal.Length: 4.648532","density: 2.244063e-01<br />Petal.Length: 4.660078","density: 2.357539e-01<br />Petal.Length: 4.671624","density: 2.474129e-01<br />Petal.Length: 4.683170","density: 2.594715e-01<br />Petal.Length: 4.694716","density: 2.718268e-01<br />Petal.Length: 4.706262","density: 2.844370e-01<br />Petal.Length: 4.717808","density: 2.972862e-01<br />Petal.Length: 4.729354","density: 3.104017e-01<br />Petal.Length: 4.740900","density: 3.237314e-01<br />Petal.Length: 4.752446","density: 3.372097e-01<br />Petal.Length: 4.763992","density: 3.508137e-01<br />Petal.Length: 4.775538","density: 3.645268e-01<br />Petal.Length: 4.787084","density: 3.783054e-01<br />Petal.Length: 4.798630","density: 3.920917e-01<br />Petal.Length: 4.810176","density: 4.058580e-01<br />Petal.Length: 4.821722","density: 4.195734e-01<br />Petal.Length: 4.833268","density: 4.331445e-01<br />Petal.Length: 4.844814","density: 4.465626e-01<br />Petal.Length: 4.856360","density: 4.597983e-01<br />Petal.Length: 4.867906","density: 4.728226e-01<br />Petal.Length: 4.879452","density: 4.854947e-01<br />Petal.Length: 4.890998","density: 4.978320e-01<br />Petal.Length: 4.902544","density: 5.098302e-01<br />Petal.Length: 4.914090","density: 5.214638e-01<br />Petal.Length: 4.925636","density: 5.325968e-01<br />Petal.Length: 4.937182","density: 5.432045e-01<br />Petal.Length: 4.948728","density: 5.533462e-01<br />Petal.Length: 4.960274","density: 5.630042e-01<br />Petal.Length: 4.971820","density: 5.720870e-01<br />Petal.Length: 4.983366","density: 5.804823e-01<br />Petal.Length: 4.994912","density: 5.883362e-01<br />Petal.Length: 5.006458","density: 5.956424e-01<br />Petal.Length: 5.018004","density: 6.023781e-01<br />Petal.Length: 5.029550","density: 6.083282e-01<br />Petal.Length: 5.041096","density: 6.137277e-01<br />Petal.Length: 5.052642","density: 6.185829e-01<br />Petal.Length: 5.064188","density: 6.229013e-01<br />Petal.Length: 5.075734","density: 6.265104e-01<br />Petal.Length: 5.087280","density: 6.295875e-01<br />Petal.Length: 5.098826","density: 6.321917e-01<br />Petal.Length: 5.110372","density: 6.343419e-01<br />Petal.Length: 5.121918","density: 6.359598e-01<br />Petal.Length: 5.133464","density: 6.371247e-01<br />Petal.Length: 5.145010","density: 6.379426e-01<br />Petal.Length: 5.156556","density: 6.384401e-01<br />Petal.Length: 5.168102","density: 6.386096e-01<br />Petal.Length: 5.179648","density: 6.384738e-01<br />Petal.Length: 5.191194","density: 6.381469e-01<br />Petal.Length: 5.202740","density: 6.376573e-01<br />Petal.Length: 5.214286","density: 6.370307e-01<br />Petal.Length: 5.225832","density: 6.362878e-01<br />Petal.Length: 5.237378","density: 6.355080e-01<br />Petal.Length: 5.248924","density: 6.347156e-01<br />Petal.Length: 5.260470","density: 6.339346e-01<br />Petal.Length: 5.272016","density: 6.332192e-01<br />Petal.Length: 5.283562","density: 6.325915e-01<br />Petal.Length: 5.295108","density: 6.320632e-01<br />Petal.Length: 5.306654","density: 6.316488e-01<br />Petal.Length: 5.318200","density: 6.314005e-01<br />Petal.Length: 5.329746","density: 6.313253e-01<br />Petal.Length: 5.341292","density: 6.314023e-01<br />Petal.Length: 5.352838","density: 6.316336e-01<br />Petal.Length: 5.364384","density: 6.320436e-01<br />Petal.Length: 5.375930","density: 6.326460e-01<br />Petal.Length: 5.387476","density: 6.333866e-01<br />Petal.Length: 5.399022","density: 6.342558e-01<br />Petal.Length: 5.410568","density: 6.352473e-01<br />Petal.Length: 5.422114","density: 6.363695e-01<br />Petal.Length: 5.433659","density: 6.375569e-01<br />Petal.Length: 5.445205","density: 6.387914e-01<br />Petal.Length: 5.456751","density: 6.400544e-01<br />Petal.Length: 5.468297","density: 6.413104e-01<br />Petal.Length: 5.479843","density: 6.425200e-01<br />Petal.Length: 5.491389","density: 6.436619e-01<br />Petal.Length: 5.502935","density: 6.447139e-01<br />Petal.Length: 5.514481","density: 6.456110e-01<br />Petal.Length: 5.526027","density: 6.463183e-01<br />Petal.Length: 5.537573","density: 6.468374e-01<br />Petal.Length: 5.549119","density: 6.471482e-01<br />Petal.Length: 5.560665","density: 6.471892e-01<br />Petal.Length: 5.572211","density: 6.468896e-01<br />Petal.Length: 5.583757","density: 6.463018e-01<br />Petal.Length: 5.595303","density: 6.454122e-01<br />Petal.Length: 5.606849","density: 6.441911e-01<br />Petal.Length: 5.618395","density: 6.425018e-01<br />Petal.Length: 5.629941","density: 6.404636e-01<br />Petal.Length: 5.641487","density: 6.380710e-01<br />Petal.Length: 5.653033","density: 6.353192e-01<br />Petal.Length: 5.664579","density: 6.320644e-01<br />Petal.Length: 5.676125","density: 6.284238e-01<br />Petal.Length: 5.687671","density: 6.244226e-01<br />Petal.Length: 5.699217","density: 6.200639e-01<br />Petal.Length: 5.710763","density: 6.152591e-01<br />Petal.Length: 5.722309","density: 6.100583e-01<br />Petal.Length: 5.733855","density: 6.045308e-01<br />Petal.Length: 5.745401","density: 5.986858e-01<br />Petal.Length: 5.756947","density: 5.924867e-01<br />Petal.Length: 5.768493","density: 5.859235e-01<br />Petal.Length: 5.780039","density: 5.790946e-01<br />Petal.Length: 5.791585","density: 5.720124e-01<br />Petal.Length: 5.803131","density: 5.646782e-01<br />Petal.Length: 5.814677","density: 5.570439e-01<br />Petal.Length: 5.826223","density: 5.492165e-01<br />Petal.Length: 5.837769","density: 5.412089e-01<br />Petal.Length: 5.849315","density: 5.330341e-01<br />Petal.Length: 5.860861","density: 5.246573e-01<br />Petal.Length: 5.872407","density: 5.161502e-01<br />Petal.Length: 5.883953","density: 5.075326e-01<br />Petal.Length: 5.895499","density: 4.988161e-01<br />Petal.Length: 5.907045","density: 4.899942e-01<br />Petal.Length: 5.918591","density: 4.810982e-01<br />Petal.Length: 5.930137","density: 4.721514e-01<br />Petal.Length: 5.941683","density: 4.631630e-01<br />Petal.Length: 5.953229","density: 4.541400e-01<br />Petal.Length: 5.964775","density: 4.450988e-01<br />Petal.Length: 5.976321","density: 4.360554e-01<br />Petal.Length: 5.987867","density: 4.270174e-01<br />Petal.Length: 5.999413","density: 4.179940e-01<br />Petal.Length: 6.010959","density: 4.090079e-01<br />Petal.Length: 6.022505","density: 4.000604e-01<br />Petal.Length: 6.034051","density: 3.911585e-01<br />Petal.Length: 6.045597","density: 3.823089e-01<br />Petal.Length: 6.057143","density: 3.735484e-01<br />Petal.Length: 6.068689","density: 3.648675e-01<br />Petal.Length: 6.080235","density: 3.562699e-01<br />Petal.Length: 6.091781","density: 3.477623e-01<br />Petal.Length: 6.103327","density: 3.393834e-01<br />Petal.Length: 6.114873","density: 3.311349e-01<br />Petal.Length: 6.126419","density: 3.230078e-01<br />Petal.Length: 6.137965","density: 3.150088e-01<br />Petal.Length: 6.149511","density: 3.071711e-01<br />Petal.Length: 6.161057","density: 2.995232e-01<br />Petal.Length: 6.172603","density: 2.920347e-01<br />Petal.Length: 6.184149","density: 2.847122e-01<br />Petal.Length: 6.195695","density: 2.775743e-01<br />Petal.Length: 6.207241","density: 2.706908e-01<br />Petal.Length: 6.218787","density: 2.640016e-01<br />Petal.Length: 6.230333","density: 2.575120e-01<br />Petal.Length: 6.241879","density: 2.512272e-01<br />Petal.Length: 6.253425","density: 2.452405e-01<br />Petal.Length: 6.264971","density: 2.394837e-01<br />Petal.Length: 6.276517","density: 2.339503e-01<br />Petal.Length: 6.288063","density: 2.286431e-01<br />Petal.Length: 6.299609","density: 2.236324e-01<br />Petal.Length: 6.311155","density: 2.188893e-01<br />Petal.Length: 6.322701","density: 2.143784e-01<br />Petal.Length: 6.334247","density: 2.100991e-01<br />Petal.Length: 6.345793","density: 2.060919e-01<br />Petal.Length: 6.357339","density: 2.023713e-01<br />Petal.Length: 6.368885","density: 1.988727e-01<br />Petal.Length: 6.380431","density: 1.955918e-01<br />Petal.Length: 6.391977","density: 1.925390e-01<br />Petal.Length: 6.403523","density: 1.897652e-01<br />Petal.Length: 6.415068","density: 1.871833e-01<br />Petal.Length: 6.426614","density: 1.847859e-01<br />Petal.Length: 6.438160","density: 1.825655e-01<br />Petal.Length: 6.449706","density: 1.805730e-01<br />Petal.Length: 6.461252","density: 1.787316e-01<br />Petal.Length: 6.472798","density: 1.770249e-01<br />Petal.Length: 6.484344","density: 1.754427e-01<br />Petal.Length: 6.495890","density: 1.740010e-01<br />Petal.Length: 6.507436","density: 1.726610e-01<br />Petal.Length: 6.518982","density: 1.713953e-01<br />Petal.Length: 6.530528","density: 1.701927e-01<br />Petal.Length: 6.542074","density: 1.690468e-01<br />Petal.Length: 6.553620","density: 1.679357e-01<br />Petal.Length: 6.565166","density: 1.668358e-01<br />Petal.Length: 6.576712","density: 1.657365e-01<br />Petal.Length: 6.588258","density: 1.646246e-01<br />Petal.Length: 6.599804","density: 1.634717e-01<br />Petal.Length: 6.611350","density: 1.622730e-01<br />Petal.Length: 6.622896","density: 1.610197e-01<br />Petal.Length: 6.634442","density: 1.597032e-01<br />Petal.Length: 6.645988","density: 1.582782e-01<br />Petal.Length: 6.657534","density: 1.567613e-01<br />Petal.Length: 6.669080","density: 1.551494e-01<br />Petal.Length: 6.680626","density: 1.534370e-01<br />Petal.Length: 6.692172","density: 1.515846e-01<br />Petal.Length: 6.703718","density: 1.496026e-01<br />Petal.Length: 6.715264","density: 1.475043e-01<br />Petal.Length: 6.726810","density: 1.452881e-01<br />Petal.Length: 6.738356","density: 1.429299e-01<br />Petal.Length: 6.749902","density: 1.404228e-01<br />Petal.Length: 6.761448","density: 1.377989e-01<br />Petal.Length: 6.772994","density: 1.350602e-01<br />Petal.Length: 6.784540","density: 1.322002e-01<br />Petal.Length: 6.796086","density: 1.291957e-01<br />Petal.Length: 6.807632","density: 1.260923e-01<br />Petal.Length: 6.819178","density: 1.228949e-01<br />Petal.Length: 6.830724","density: 1.196086e-01<br />Petal.Length: 6.842270","density: 1.162106e-01<br />Petal.Length: 6.853816","density: 1.127432e-01<br />Petal.Length: 6.865362","density: 1.092154e-01<br />Petal.Length: 6.876908","density: 1.056340e-01<br />Petal.Length: 6.888454","density: 1.019969e-01<br />Petal.Length: 6.900000","density: 1.019969e-01<br />Petal.Length: 6.900000","density: 1.019969e-01<br />Petal.Length: 6.900000","density: 1.056340e-01<br />Petal.Length: 6.888454","density: 1.092154e-01<br />Petal.Length: 6.876908","density: 1.127432e-01<br />Petal.Length: 6.865362","density: 1.162106e-01<br />Petal.Length: 6.853816","density: 1.196086e-01<br />Petal.Length: 6.842270","density: 1.228949e-01<br />Petal.Length: 6.830724","density: 1.260923e-01<br />Petal.Length: 6.819178","density: 1.291957e-01<br />Petal.Length: 6.807632","density: 1.322002e-01<br />Petal.Length: 6.796086","density: 1.350602e-01<br />Petal.Length: 6.784540","density: 1.377989e-01<br />Petal.Length: 6.772994","density: 1.404228e-01<br />Petal.Length: 6.761448","density: 1.429299e-01<br />Petal.Length: 6.749902","density: 1.452881e-01<br />Petal.Length: 6.738356","density: 1.475043e-01<br />Petal.Length: 6.726810","density: 1.496026e-01<br />Petal.Length: 6.715264","density: 1.515846e-01<br />Petal.Length: 6.703718","density: 1.534370e-01<br />Petal.Length: 6.692172","density: 1.551494e-01<br />Petal.Length: 6.680626","density: 1.567613e-01<br />Petal.Length: 6.669080","density: 1.582782e-01<br />Petal.Length: 6.657534","density: 1.597032e-01<br />Petal.Length: 6.645988","density: 1.610197e-01<br />Petal.Length: 6.634442","density: 1.622730e-01<br />Petal.Length: 6.622896","density: 1.634717e-01<br />Petal.Length: 6.611350","density: 1.646246e-01<br />Petal.Length: 6.599804","density: 1.657365e-01<br />Petal.Length: 6.588258","density: 1.668358e-01<br />Petal.Length: 6.576712","density: 1.679357e-01<br />Petal.Length: 6.565166","density: 1.690468e-01<br />Petal.Length: 6.553620","density: 1.701927e-01<br />Petal.Length: 6.542074","density: 1.713953e-01<br />Petal.Length: 6.530528","density: 1.726610e-01<br />Petal.Length: 6.518982","density: 1.740010e-01<br />Petal.Length: 6.507436","density: 1.754427e-01<br />Petal.Length: 6.495890","density: 1.770249e-01<br />Petal.Length: 6.484344","density: 1.787316e-01<br />Petal.Length: 6.472798","density: 1.805730e-01<br />Petal.Length: 6.461252","density: 1.825655e-01<br />Petal.Length: 6.449706","density: 1.847859e-01<br />Petal.Length: 6.438160","density: 1.871833e-01<br />Petal.Length: 6.426614","density: 1.897652e-01<br />Petal.Length: 6.415068","density: 1.925390e-01<br />Petal.Length: 6.403523","density: 1.955918e-01<br />Petal.Length: 6.391977","density: 1.988727e-01<br />Petal.Length: 6.380431","density: 2.023713e-01<br />Petal.Length: 6.368885","density: 2.060919e-01<br />Petal.Length: 6.357339","density: 2.100991e-01<br />Petal.Length: 6.345793","density: 2.143784e-01<br />Petal.Length: 6.334247","density: 2.188893e-01<br />Petal.Length: 6.322701","density: 2.236324e-01<br />Petal.Length: 6.311155","density: 2.286431e-01<br />Petal.Length: 6.299609","density: 2.339503e-01<br />Petal.Length: 6.288063","density: 2.394837e-01<br />Petal.Length: 6.276517","density: 2.452405e-01<br />Petal.Length: 6.264971","density: 2.512272e-01<br />Petal.Length: 6.253425","density: 2.575120e-01<br />Petal.Length: 6.241879","density: 2.640016e-01<br />Petal.Length: 6.230333","density: 2.706908e-01<br />Petal.Length: 6.218787","density: 2.775743e-01<br />Petal.Length: 6.207241","density: 2.847122e-01<br />Petal.Length: 6.195695","density: 2.920347e-01<br />Petal.Length: 6.184149","density: 2.995232e-01<br />Petal.Length: 6.172603","density: 3.071711e-01<br />Petal.Length: 6.161057","density: 3.150088e-01<br />Petal.Length: 6.149511","density: 3.230078e-01<br />Petal.Length: 6.137965","density: 3.311349e-01<br />Petal.Length: 6.126419","density: 3.393834e-01<br />Petal.Length: 6.114873","density: 3.477623e-01<br />Petal.Length: 6.103327","density: 3.562699e-01<br />Petal.Length: 6.091781","density: 3.648675e-01<br />Petal.Length: 6.080235","density: 3.735484e-01<br />Petal.Length: 6.068689","density: 3.823089e-01<br />Petal.Length: 6.057143","density: 3.911585e-01<br />Petal.Length: 6.045597","density: 4.000604e-01<br />Petal.Length: 6.034051","density: 4.090079e-01<br />Petal.Length: 6.022505","density: 4.179940e-01<br />Petal.Length: 6.010959","density: 4.270174e-01<br />Petal.Length: 5.999413","density: 4.360554e-01<br />Petal.Length: 5.987867","density: 4.450988e-01<br />Petal.Length: 5.976321","density: 4.541400e-01<br />Petal.Length: 5.964775","density: 4.631630e-01<br />Petal.Length: 5.953229","density: 4.721514e-01<br />Petal.Length: 5.941683","density: 4.810982e-01<br />Petal.Length: 5.930137","density: 4.899942e-01<br />Petal.Length: 5.918591","density: 4.988161e-01<br />Petal.Length: 5.907045","density: 5.075326e-01<br />Petal.Length: 5.895499","density: 5.161502e-01<br />Petal.Length: 5.883953","density: 5.246573e-01<br />Petal.Length: 5.872407","density: 5.330341e-01<br />Petal.Length: 5.860861","density: 5.412089e-01<br />Petal.Length: 5.849315","density: 5.492165e-01<br />Petal.Length: 5.837769","density: 5.570439e-01<br />Petal.Length: 5.826223","density: 5.646782e-01<br />Petal.Length: 5.814677","density: 5.720124e-01<br />Petal.Length: 5.803131","density: 5.790946e-01<br />Petal.Length: 5.791585","density: 5.859235e-01<br />Petal.Length: 5.780039","density: 5.924867e-01<br />Petal.Length: 5.768493","density: 5.986858e-01<br />Petal.Length: 5.756947","density: 6.045308e-01<br />Petal.Length: 5.745401","density: 6.100583e-01<br />Petal.Length: 5.733855","density: 6.152591e-01<br />Petal.Length: 5.722309","density: 6.200639e-01<br />Petal.Length: 5.710763","density: 6.244226e-01<br />Petal.Length: 5.699217","density: 6.284238e-01<br />Petal.Length: 5.687671","density: 6.320644e-01<br />Petal.Length: 5.676125","density: 6.353192e-01<br />Petal.Length: 5.664579","density: 6.380710e-01<br />Petal.Length: 5.653033","density: 6.404636e-01<br />Petal.Length: 5.641487","density: 6.425018e-01<br />Petal.Length: 5.629941","density: 6.441911e-01<br />Petal.Length: 5.618395","density: 6.454122e-01<br />Petal.Length: 5.606849","density: 6.463018e-01<br />Petal.Length: 5.595303","density: 6.468896e-01<br />Petal.Length: 5.583757","density: 6.471892e-01<br />Petal.Length: 5.572211","density: 6.471482e-01<br />Petal.Length: 5.560665","density: 6.468374e-01<br />Petal.Length: 5.549119","density: 6.463183e-01<br />Petal.Length: 5.537573","density: 6.456110e-01<br />Petal.Length: 5.526027","density: 6.447139e-01<br />Petal.Length: 5.514481","density: 6.436619e-01<br />Petal.Length: 5.502935","density: 6.425200e-01<br />Petal.Length: 5.491389","density: 6.413104e-01<br />Petal.Length: 5.479843","density: 6.400544e-01<br />Petal.Length: 5.468297","density: 6.387914e-01<br />Petal.Length: 5.456751","density: 6.375569e-01<br />Petal.Length: 5.445205","density: 6.363695e-01<br />Petal.Length: 5.433659","density: 6.352473e-01<br />Petal.Length: 5.422114","density: 6.342558e-01<br />Petal.Length: 5.410568","density: 6.333866e-01<br />Petal.Length: 5.399022","density: 6.326460e-01<br />Petal.Length: 5.387476","density: 6.320436e-01<br />Petal.Length: 5.375930","density: 6.316336e-01<br />Petal.Length: 5.364384","density: 6.314023e-01<br />Petal.Length: 5.352838","density: 6.313253e-01<br />Petal.Length: 5.341292","density: 6.314005e-01<br />Petal.Length: 5.329746","density: 6.316488e-01<br />Petal.Length: 5.318200","density: 6.320632e-01<br />Petal.Length: 5.306654","density: 6.325915e-01<br />Petal.Length: 5.295108","density: 6.332192e-01<br />Petal.Length: 5.283562","density: 6.339346e-01<br />Petal.Length: 5.272016","density: 6.347156e-01<br />Petal.Length: 5.260470","density: 6.355080e-01<br />Petal.Length: 5.248924","density: 6.362878e-01<br />Petal.Length: 5.237378","density: 6.370307e-01<br />Petal.Length: 5.225832","density: 6.376573e-01<br />Petal.Length: 5.214286","density: 6.381469e-01<br />Petal.Length: 5.202740","density: 6.384738e-01<br />Petal.Length: 5.191194","density: 6.386096e-01<br />Petal.Length: 5.179648","density: 6.384401e-01<br />Petal.Length: 5.168102","density: 6.379426e-01<br />Petal.Length: 5.156556","density: 6.371247e-01<br />Petal.Length: 5.145010","density: 6.359598e-01<br />Petal.Length: 5.133464","density: 6.343419e-01<br />Petal.Length: 5.121918","density: 6.321917e-01<br />Petal.Length: 5.110372","density: 6.295875e-01<br />Petal.Length: 5.098826","density: 6.265104e-01<br />Petal.Length: 5.087280","density: 6.229013e-01<br />Petal.Length: 5.075734","density: 6.185829e-01<br />Petal.Length: 5.064188","density: 6.137277e-01<br />Petal.Length: 5.052642","density: 6.083282e-01<br />Petal.Length: 5.041096","density: 6.023781e-01<br />Petal.Length: 5.029550","density: 5.956424e-01<br />Petal.Length: 5.018004","density: 5.883362e-01<br />Petal.Length: 5.006458","density: 5.804823e-01<br />Petal.Length: 4.994912","density: 5.720870e-01<br />Petal.Length: 4.983366","density: 5.630042e-01<br />Petal.Length: 4.971820","density: 5.533462e-01<br />Petal.Length: 4.960274","density: 5.432045e-01<br />Petal.Length: 4.948728","density: 5.325968e-01<br />Petal.Length: 4.937182","density: 5.214638e-01<br />Petal.Length: 4.925636","density: 5.098302e-01<br />Petal.Length: 4.914090","density: 4.978320e-01<br />Petal.Length: 4.902544","density: 4.854947e-01<br />Petal.Length: 4.890998","density: 4.728226e-01<br />Petal.Length: 4.879452","density: 4.597983e-01<br />Petal.Length: 4.867906","density: 4.465626e-01<br />Petal.Length: 4.856360","density: 4.331445e-01<br />Petal.Length: 4.844814","density: 4.195734e-01<br />Petal.Length: 4.833268","density: 4.058580e-01<br />Petal.Length: 4.821722","density: 3.920917e-01<br />Petal.Length: 4.810176","density: 3.783054e-01<br />Petal.Length: 4.798630","density: 3.645268e-01<br />Petal.Length: 4.787084","density: 3.508137e-01<br />Petal.Length: 4.775538","density: 3.372097e-01<br />Petal.Length: 4.763992","density: 3.237314e-01<br />Petal.Length: 4.752446","density: 3.104017e-01<br />Petal.Length: 4.740900","density: 2.972862e-01<br />Petal.Length: 4.729354","density: 2.844370e-01<br />Petal.Length: 4.717808","density: 2.718268e-01<br />Petal.Length: 4.706262","density: 2.594715e-01<br />Petal.Length: 4.694716","density: 2.474129e-01<br />Petal.Length: 4.683170","density: 2.357539e-01<br />Petal.Length: 4.671624","density: 2.244063e-01<br />Petal.Length: 4.660078","density: 2.133783e-01<br />Petal.Length: 4.648532","density: 2.026774e-01<br />Petal.Length: 4.636986","density: 1.924564e-01<br />Petal.Length: 4.625440","density: 1.825847e-01<br />Petal.Length: 4.613894","density: 1.730567e-01<br />Petal.Length: 4.602348","density: 1.638731e-01<br />Petal.Length: 4.590802","density: 1.551412e-01<br />Petal.Length: 4.579256","density: 1.467924e-01<br />Petal.Length: 4.567710","density: 1.387780e-01<br />Petal.Length: 4.556164","density: 1.310937e-01<br />Petal.Length: 4.544618","density: 1.237989e-01<br />Petal.Length: 4.533072","density: 1.168910e-01<br />Petal.Length: 4.521526","density: 1.102853e-01<br />Petal.Length: 4.509980","density: 1.039746e-01<br />Petal.Length: 4.498434","density: 9.797742e-02<br />Petal.Length: 4.486888","density: 9.234702e-02<br />Petal.Length: 4.475342","density: 8.697512e-02<br />Petal.Length: 4.463796","density: 8.185351e-02<br />Petal.Length: 4.452250","density: 7.697394e-02<br />Petal.Length: 4.440705","density: 7.241992e-02<br />Petal.Length: 4.429159","density: 6.808206e-02<br />Petal.Length: 4.417613","density: 6.394943e-02<br />Petal.Length: 4.406067","density: 6.001443e-02<br />Petal.Length: 4.394521","density: 5.632522e-02<br />Petal.Length: 4.382975","density: 5.283115e-02<br />Petal.Length: 4.371429","density: 4.950281e-02<br />Petal.Length: 4.359883","density: 4.633394e-02<br />Petal.Length: 4.348337","density: 4.334823e-02<br />Petal.Length: 4.336791","density: 4.053691e-02<br />Petal.Length: 4.325245","density: 3.785977e-02<br />Petal.Length: 4.313699","density: 3.531207e-02<br />Petal.Length: 4.302153","density: 3.290093e-02<br />Petal.Length: 4.290607","density: 3.064634e-02<br />Petal.Length: 4.279061","density: 2.850208e-02<br />Petal.Length: 4.267515","density: 2.646464e-02<br />Petal.Length: 4.255969","density: 2.453056e-02<br />Petal.Length: 4.244423","density: 2.273680e-02<br />Petal.Length: 4.232877","density: 2.103622e-02<br />Petal.Length: 4.221331","density: 1.942528e-02<br />Petal.Length: 4.209785","density: 1.790128e-02<br />Petal.Length: 4.198239","density: 1.648723e-02<br />Petal.Length: 4.186693","density: 1.516098e-02<br />Petal.Length: 4.175147","density: 1.391042e-02<br />Petal.Length: 4.163601","density: 1.273325e-02<br />Petal.Length: 4.152055","density: 1.164159e-02<br />Petal.Length: 4.140509","density: 1.063142e-02<br />Petal.Length: 4.128963","density: 9.684577e-03<br />Petal.Length: 4.117417","density: 8.798957e-03<br />Petal.Length: 4.105871","density: 7.978342e-03<br />Petal.Length: 4.094325","density: 7.231330e-03<br />Petal.Length: 4.082779","density: 6.536065e-03<br />Petal.Length: 4.071233","density: 5.890549e-03<br />Petal.Length: 4.059687","density: 5.292793e-03<br />Petal.Length: 4.048141","density: 4.758868e-03<br />Petal.Length: 4.036595","density: 4.265890e-03<br />Petal.Length: 4.025049","density: 3.811888e-03<br />Petal.Length: 4.013503","density: 3.395030e-03<br />Petal.Length: 4.001957","density: 3.023863e-03<br />Petal.Length: 3.990411","density: 2.687420e-03<br />Petal.Length: 3.978865","density: 2.380219e-03<br />Petal.Length: 3.967319","density: 2.100658e-03<br />Petal.Length: 3.955773","density: 1.852235e-03<br />Petal.Length: 3.944227","density: 1.631733e-03<br />Petal.Length: 3.932681","density: 1.432153e-03<br />Petal.Length: 3.921135","density: 1.252188e-03<br />Petal.Length: 3.909589","density: 1.092346e-03<br />Petal.Length: 3.898043","density: 9.537818e-04<br />Petal.Length: 3.886497","density: 8.294657e-04<br />Petal.Length: 3.874951","density: 7.183969e-04<br />Petal.Length: 3.863405","density: 6.196240e-04<br />Petal.Length: 3.851859","density: 5.362252e-04<br />Petal.Length: 3.840313","density: 4.620503e-04<br />Petal.Length: 3.828767","density: 3.963848e-04<br />Petal.Length: 3.817221","density: 3.385301e-04<br />Petal.Length: 3.805675","density: 2.897612e-04<br />Petal.Length: 3.794129","density: 2.473933e-04<br />Petal.Length: 3.782583","density: 2.102225e-04<br />Petal.Length: 3.771037","density: 1.777831e-04<br />Petal.Length: 3.759491","density: 1.504138e-04<br />Petal.Length: 3.747945","density: 1.272564e-04<br />Petal.Length: 3.736399","density: 1.071171e-04<br />Petal.Length: 3.724853","density: 8.970446e-05<br />Petal.Length: 3.713307","density: 7.497132e-05<br />Petal.Length: 3.701761","density: 6.286270e-05<br />Petal.Length: 3.690215","density: 5.242087e-05<br />Petal.Length: 3.678669","density: 4.347442e-05<br />Petal.Length: 3.667123","density: 3.586736e-05<br />Petal.Length: 3.655577","density: 2.981173e-05<br />Petal.Length: 3.644031","density: 2.463162e-05<br />Petal.Length: 3.632485","density: 2.023219e-05<br />Petal.Length: 3.620939","density: 1.652196e-05<br />Petal.Length: 3.609393","density: 1.356882e-05<br />Petal.Length: 3.597847","density: 1.111023e-05<br />Petal.Length: 3.586301","density: 9.039699e-06<br />Petal.Length: 3.574755","density: 7.309414e-06<br />Petal.Length: 3.563209","density: 5.925866e-06<br />Petal.Length: 3.551663","density: 4.809575e-06<br />Petal.Length: 3.540117","density: 3.876979e-06<br />Petal.Length: 3.528571","density: 3.104478e-06<br />Petal.Length: 3.517025","density: 2.482658e-06<br />Petal.Length: 3.505479","density: 1.997853e-06<br />Petal.Length: 3.493933","density: 1.595865e-06<br />Petal.Length: 3.482387","density: 1.265687e-06<br />Petal.Length: 3.470841","density: 9.975630e-07<br />Petal.Length: 3.459295","density: 7.961922e-07<br />Petal.Length: 3.447750","density: 6.303792e-07<br />Petal.Length: 3.436204","density: 4.952768e-07<br />Petal.Length: 3.424658","density: 3.862684e-07<br />Petal.Length: 3.413112","density: 3.043649e-07<br />Petal.Length: 3.401566","density: 2.389202e-07<br />Petal.Length: 3.390020","density: 1.859980e-07<br />Petal.Length: 3.378474","density: 1.436602e-07<br />Petal.Length: 3.366928","density: 1.115875e-07<br />Petal.Length: 3.355382","density: 8.687416e-08<br />Petal.Length: 3.343836","density: 6.702900e-08<br />Petal.Length: 3.332290","density: 5.128142e-08<br />Petal.Length: 3.320744","density: 3.922819e-08<br />Petal.Length: 3.309198","density: 3.030082e-08<br />Petal.Length: 3.297652","density: 2.317750e-08<br />Petal.Length: 3.286106","density: 1.756816e-08<br />Petal.Length: 3.274560","density: 1.322059e-08<br />Petal.Length: 3.263014","density: 1.013628e-08<br />Petal.Length: 3.251468","density: 7.689035e-09<br />Petal.Length: 3.239922","density: 5.775640e-09<br />Petal.Length: 3.228376","density: 4.298932e-09<br />Petal.Length: 3.216830","density: 3.251553e-09<br />Petal.Length: 3.205284","density: 2.446963e-09<br />Petal.Length: 3.193738","density: 1.821985e-09<br />Petal.Length: 3.182192","density: 1.343416e-09<br />Petal.Length: 3.170646","density: 1.000025e-09<br />Petal.Length: 3.159100","density: 7.469239e-10<br />Petal.Length: 3.147554","density: 5.514672e-10<br />Petal.Length: 3.136008","density: 4.028966e-10<br />Petal.Length: 3.124462","density: 2.948126e-10<br />Petal.Length: 3.112916","density: 2.186529e-10<br />Petal.Length: 3.101370","density: 1.601329e-10<br />Petal.Length: 3.089824","density: 1.159519e-10<br />Petal.Length: 3.078278","density: 8.328955e-11<br />Petal.Length: 3.066732","density: 6.137509e-11<br />Petal.Length: 3.055186","density: 4.460436e-11<br />Petal.Length: 3.043640","density: 3.202054e-11<br />Petal.Length: 3.032094","density: 2.273369e-11<br />Petal.Length: 3.020548","density: 1.651599e-11<br />Petal.Length: 3.009002","density: 1.191664e-11<br />Petal.Length: 2.997456","density: 8.484131e-12<br />Petal.Length: 2.985910","density: 5.968934e-12<br />Petal.Length: 2.974364","density: 4.259909e-12<br />Petal.Length: 2.962818","density: 3.053139e-12<br />Petal.Length: 2.951272","density: 2.156588e-12<br />Petal.Length: 2.939726","density: 1.503931e-12<br />Petal.Length: 2.928180","density: 1.052880e-12<br />Petal.Length: 2.916634","density: 7.500562e-13<br />Petal.Length: 2.905088","density: 5.258282e-13<br />Petal.Length: 2.893542","density: 3.635523e-13<br />Petal.Length: 2.881996","density: 2.492670e-13<br />Petal.Length: 2.870450","density: 1.766270e-13<br />Petal.Length: 2.858904","density: 1.229681e-13<br />Petal.Length: 2.847358","density: 8.431704e-14<br />Petal.Length: 2.835812","density: 5.702905e-14<br />Petal.Length: 2.824266","density: 3.984884e-14<br />Petal.Length: 2.812720","density: 2.755503e-14<br />Petal.Length: 2.801174","density: 1.873189e-14<br />Petal.Length: 2.789628","density: 1.255355e-14<br />Petal.Length: 2.778082","density: 8.545633e-15<br />Petal.Length: 2.766536","density: 5.853179e-15<br />Petal.Length: 2.754990","density: 3.979630e-15<br />Petal.Length: 2.743444","density: 2.711159e-15<br />Petal.Length: 2.731898","density: 1.827878e-15<br />Petal.Length: 2.720352","density: 1.230578e-15<br />Petal.Length: 2.708806","density: 8.206522e-16<br />Petal.Length: 2.697260","density: 5.452413e-16<br />Petal.Length: 2.685714","density: 3.383468e-16<br />Petal.Length: 2.674168","density: 2.089964e-16<br />Petal.Length: 2.662622","density: 1.329443e-16<br />Petal.Length: 2.651076","density: 8.254693e-17<br />Petal.Length: 2.639530","density: 3.415870e-17<br />Petal.Length: 2.627984","density: 2.354872e-17<br />Petal.Length: 2.616438","density: 8.999775e-18<br />Petal.Length: 2.604892","density: 0.000000e+00<br />Petal.Length: 2.593346","density: 0.000000e+00<br />Petal.Length: 2.581800","density: 0.000000e+00<br />Petal.Length: 2.570254","density: 0.000000e+00<br />Petal.Length: 2.558708","density: 0.000000e+00<br />Petal.Length: 2.547162","density: 0.000000e+00<br />Petal.Length: 2.535616","density: 0.000000e+00<br />Petal.Length: 2.524070","density: 0.000000e+00<br />Petal.Length: 2.512524","density: 6.223203e-18<br />Petal.Length: 2.500978","density: 1.831087e-17<br />Petal.Length: 2.489432","density: 3.422584e-17<br />Petal.Length: 2.477886","density: 8.950583e-18<br />Petal.Length: 2.466341","density: 0.000000e+00<br />Petal.Length: 2.454795","density: 9.404826e-18<br />Petal.Length: 2.443249","density: 3.481800e-17<br />Petal.Length: 2.431703","density: 3.747049e-17<br />Petal.Length: 2.420157","density: 1.661414e-17<br />Petal.Length: 2.408611","density: 0.000000e+00<br />Petal.Length: 2.397065","density: 2.601314e-18<br />Petal.Length: 2.385519","density: 2.382127e-17<br />Petal.Length: 2.373973","density: 1.046993e-17<br />Petal.Length: 2.362427","density: 0.000000e+00<br />Petal.Length: 2.350881","density: 2.107196e-18<br />Petal.Length: 2.339335","density: 1.271717e-17<br />Petal.Length: 2.327789","density: 9.153108e-18<br />Petal.Length: 2.316243","density: 3.848120e-18<br />Petal.Length: 2.304697","density: 0.000000e+00<br />Petal.Length: 2.293151","density: 0.000000e+00<br />Petal.Length: 2.281605","density: 0.000000e+00<br />Petal.Length: 2.270059","density: 3.494043e-17<br />Petal.Length: 2.258513","density: 6.380853e-17<br />Petal.Length: 2.246967","density: 4.766733e-17<br />Petal.Length: 2.235421","density: 2.644738e-17<br />Petal.Length: 2.223875","density: 1.691937e-17<br />Petal.Length: 2.212329","density: 1.840460e-17<br />Petal.Length: 2.200783","density: 2.712608e-17<br />Petal.Length: 2.189237","density: 2.182109e-17<br />Petal.Length: 2.177691","density: 7.914935e-18<br />Petal.Length: 2.166145","density: 5.333352e-18<br />Petal.Length: 2.154599","density: 1.697610e-17<br />Petal.Length: 2.143053","density: 3.289106e-17<br />Petal.Length: 2.131507","density: 1.587906e-17<br />Petal.Length: 2.119961","density: 3.868948e-18<br />Petal.Length: 2.108415","density: 0.000000e+00<br />Petal.Length: 2.096869","density: 0.000000e+00<br />Petal.Length: 2.085323","density: 0.000000e+00<br />Petal.Length: 2.073777","density: 0.000000e+00<br />Petal.Length: 2.062231","density: 0.000000e+00<br />Petal.Length: 2.050685","density: 0.000000e+00<br />Petal.Length: 2.039139","density: 0.000000e+00<br />Petal.Length: 2.027593","density: 1.550594e-17<br />Petal.Length: 2.016047","density: 1.878526e-17<br />Petal.Length: 2.004501","density: 0.000000e+00<br />Petal.Length: 1.992955","density: 0.000000e+00<br />Petal.Length: 1.981409","density: 0.000000e+00<br />Petal.Length: 1.969863","density: 2.645848e-18<br />Petal.Length: 1.958317","density: 5.926952e-18<br />Petal.Length: 1.946771","density: 6.219642e-19<br />Petal.Length: 1.935225","density: 0.000000e+00<br />Petal.Length: 1.923679","density: 0.000000e+00<br />Petal.Length: 1.912133","density: 0.000000e+00<br />Petal.Length: 1.900587","density: 0.000000e+00<br />Petal.Length: 1.889041","density: 0.000000e+00<br />Petal.Length: 1.877495","density: 0.000000e+00<br />Petal.Length: 1.865949","density: 0.000000e+00<br />Petal.Length: 1.854403","density: 0.000000e+00<br />Petal.Length: 1.842857","density: 0.000000e+00<br />Petal.Length: 1.831311","density: 0.000000e+00<br />Petal.Length: 1.819765","density: 0.000000e+00<br />Petal.Length: 1.808219","density: 0.000000e+00<br />Petal.Length: 1.796673","density: 0.000000e+00<br />Petal.Length: 1.785127","density: 0.000000e+00<br />Petal.Length: 1.773581","density: 0.000000e+00<br />Petal.Length: 1.762035","density: 0.000000e+00<br />Petal.Length: 1.750489","density: 0.000000e+00<br />Petal.Length: 1.738943","density: 0.000000e+00<br />Petal.Length: 1.727397","density: 0.000000e+00<br />Petal.Length: 1.715851","density: 0.000000e+00<br />Petal.Length: 1.704305","density: 0.000000e+00<br />Petal.Length: 1.692759","density: 0.000000e+00<br />Petal.Length: 1.681213","density: 0.000000e+00<br />Petal.Length: 1.669667","density: 0.000000e+00<br />Petal.Length: 1.658121","density: 0.000000e+00<br />Petal.Length: 1.646575","density: 0.000000e+00<br />Petal.Length: 1.635029","density: 0.000000e+00<br />Petal.Length: 1.623483","density: 0.000000e+00<br />Petal.Length: 1.611937","density: 5.670163e-19<br />Petal.Length: 1.600391","density: 5.872004e-18<br />Petal.Length: 1.588845","density: 2.700796e-18<br />Petal.Length: 1.577299","density: 0.000000e+00<br />Petal.Length: 1.565753","density: 0.000000e+00<br />Petal.Length: 1.554207","density: 0.000000e+00<br />Petal.Length: 1.542661","density: 2.320684e-17<br />Petal.Length: 1.531115","density: 6.176162e-17<br />Petal.Length: 1.519569","density: 8.203541e-17<br />Petal.Length: 1.508023","density: 2.368054e-17<br />Petal.Length: 1.496477","density: 1.072741e-17<br />Petal.Length: 1.484932","density: 2.058182e-17<br />Petal.Length: 1.473386","density: 4.357859e-17<br />Petal.Length: 1.461840","density: 6.853143e-17<br />Petal.Length: 1.450294","density: 3.670150e-17<br />Petal.Length: 1.438748","density: 1.249957e-17<br />Petal.Length: 1.427202","density: 1.744075e-17<br />Petal.Length: 1.415656","density: 5.278021e-17<br />Petal.Length: 1.404110","density: 2.625527e-17<br />Petal.Length: 1.392564","density: 5.033757e-17<br />Petal.Length: 1.381018","density: 6.422221e-17<br />Petal.Length: 1.369472","density: 6.025844e-17<br />Petal.Length: 1.357926","density: 8.678338e-17<br />Petal.Length: 1.346380","density: 5.786184e-17<br />Petal.Length: 1.334834","density: 5.059326e-17<br />Petal.Length: 1.323288","density: 5.974459e-17<br />Petal.Length: 1.311742","density: 4.913462e-17<br />Petal.Length: 1.300196","density: 7.871511e-17<br />Petal.Length: 1.288650","density: 8.342582e-17<br />Petal.Length: 1.277104","density: 7.983983e-17<br />Petal.Length: 1.265558","density: 8.996144e-17<br />Petal.Length: 1.254012","density: 7.935146e-17<br />Petal.Length: 1.242466","density: 8.391418e-17<br />Petal.Length: 1.230920","density: 6.861296e-17<br />Petal.Length: 1.219374","density: 2.449420e-17<br />Petal.Length: 1.207828","density: 6.162912e-17<br />Petal.Length: 1.196282","density: 7.358538e-17<br />Petal.Length: 1.184736","density: 7.376530e-17<br />Petal.Length: 1.173190","density: 7.310346e-17<br />Petal.Length: 1.161644","density: 9.432341e-17<br />Petal.Length: 1.150098","density: 6.494654e-17<br />Petal.Length: 1.138552","density: 5.746967e-17<br />Petal.Length: 1.127006","density: 6.406134e-17<br />Petal.Length: 1.115460","density: 4.284139e-17<br />Petal.Length: 1.103914","density: 5.664231e-17<br />Petal.Length: 1.092368","density: 7.255727e-17<br />Petal.Length: 1.080822","density: 7.459088e-17<br />Petal.Length: 1.069276","density: 4.814189e-17<br />Petal.Length: 1.057730","density: 2.559569e-17<br />Petal.Length: 1.046184","density: 2.096813e-17<br />Petal.Length: 1.034638","density: 2.362062e-17<br />Petal.Length: 1.023092","density: 2.526405e-17<br />Petal.Length: 1.011546","density: 1.730657e-17<br />Petal.Length: 1.000000","density: 1.730657e-17<br />Petal.Length: 1.000000"],"key":["101","102","103","104","105","106","107","108","109","110","111","112","113","114","115","116","117","118","119","120","121","122","123","124","125","126","127","128","129","130","131","132","133","134","135","136","137","138","139","140","141","142","143","144","145","146","147","148","149","150"],"type":"scatter","mode":"lines","line":{"width":1.88976377952756,"color":"rgba(0,0,0,1)","dash":"solid"},"fill":"toself","fillcolor":"rgba(97,156,255,1)","hoveron":"points","set":"SharedData91833e3a","name":"virginica","legendgroup":"virginica","showlegend":true,"xaxis":"x3","yaxis":"y10","hoverinfo":"text","_isSimpleKey":true,"_isNestedKey":false,"frame":null},{"x":[1.4,1.4,1.3,1.5,1.4,1.7,1.4,1.5,1.4,1.5,1.5,1.6,1.4,1.1,1.2,1.5,1.3,1.4,1.7,1.5,1.7,1.5,1,1.7,1.9,1.6,1.6,1.5,1.4,1.6,1.6,1.5,1.5,1.4,1.5,1.2,1.3,1.4,1.3,1.5,1.3,1.3,1.3,1.6,1.9,1.4,1.6,1.4,1.5,1.4],"y":[0.2,0.2,0.2,0.2,0.2,0.4,0.3,0.2,0.2,0.1,0.2,0.2,0.1,0.1,0.2,0.4,0.4,0.3,0.3,0.3,0.2,0.4,0.2,0.5,0.2,0.2,0.4,0.2,0.2,0.2,0.2,0.4,0.1,0.2,0.2,0.2,0.2,0.1,0.2,0.2,0.3,0.3,0.2,0.6,0.4,0.3,0.2,0.2,0.2,0.2],"text":["Petal.Length: 1.4<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.4<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.3<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.5<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.4<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.7<br />Petal.Width: 0.4<br />Species: setosa","Petal.Length: 1.4<br />Petal.Width: 0.3<br />Species: setosa","Petal.Length: 1.5<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.4<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.5<br />Petal.Width: 0.1<br />Species: setosa","Petal.Length: 1.5<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.6<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.4<br />Petal.Width: 0.1<br />Species: setosa","Petal.Length: 1.1<br />Petal.Width: 0.1<br />Species: setosa","Petal.Length: 1.2<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.5<br />Petal.Width: 0.4<br />Species: setosa","Petal.Length: 1.3<br />Petal.Width: 0.4<br />Species: setosa","Petal.Length: 1.4<br />Petal.Width: 0.3<br />Species: setosa","Petal.Length: 1.7<br />Petal.Width: 0.3<br />Species: setosa","Petal.Length: 1.5<br />Petal.Width: 0.3<br />Species: setosa","Petal.Length: 1.7<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.5<br />Petal.Width: 0.4<br />Species: setosa","Petal.Length: 1.0<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.7<br />Petal.Width: 0.5<br />Species: setosa","Petal.Length: 1.9<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.6<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.6<br />Petal.Width: 0.4<br />Species: setosa","Petal.Length: 1.5<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.4<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.6<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.6<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.5<br />Petal.Width: 0.4<br />Species: setosa","Petal.Length: 1.5<br />Petal.Width: 0.1<br />Species: setosa","Petal.Length: 1.4<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.5<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.2<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.3<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.4<br />Petal.Width: 0.1<br />Species: setosa","Petal.Length: 1.3<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.5<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.3<br />Petal.Width: 0.3<br />Species: setosa","Petal.Length: 1.3<br />Petal.Width: 0.3<br />Species: setosa","Petal.Length: 1.3<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.6<br />Petal.Width: 0.6<br />Species: setosa","Petal.Length: 1.9<br />Petal.Width: 0.4<br />Species: setosa","Petal.Length: 1.4<br />Petal.Width: 0.3<br />Species: setosa","Petal.Length: 1.6<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.4<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.5<br />Petal.Width: 0.2<br />Species: setosa","Petal.Length: 1.4<br />Petal.Width: 0.2<br />Species: setosa"],"key":["1","2","3","4","5","6","7","8","9","10","11","12","13","14","15","16","17","18","19","20","21","22","23","24","25","26","27","28","29","30","31","32","33","34","35","36","37","38","39","40","41","42","43","44","45","46","47","48","49","50"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(248,118,109,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(248,118,109,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"setosa","legendgroup":"setosa","showlegend":true,"xaxis":"x3","yaxis":"y9","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[4.7,4.5,4.9,4,4.6,4.5,4.7,3.3,4.6,3.9,3.5,4.2,4,4.7,3.6,4.4,4.5,4.1,4.5,3.9,4.8,4,4.9,4.7,4.3,4.4,4.8,5,4.5,3.5,3.8,3.7,3.9,5.1,4.5,4.5,4.7,4.4,4.1,4,4.4,4.6,4,3.3,4.2,4.2,4.2,4.3,3,4.1],"y":[1.4,1.5,1.5,1.3,1.5,1.3,1.6,1,1.3,1.4,1,1.5,1,1.4,1.3,1.4,1.5,1,1.5,1.1,1.8,1.3,1.5,1.2,1.3,1.4,1.4,1.7,1.5,1,1.1,1,1.2,1.6,1.5,1.6,1.5,1.3,1.3,1.3,1.2,1.4,1.2,1,1.3,1.2,1.3,1.3,1.1,1.3],"text":["Petal.Length: 4.7<br />Petal.Width: 1.4<br />Species: versicolor","Petal.Length: 4.5<br />Petal.Width: 1.5<br />Species: versicolor","Petal.Length: 4.9<br />Petal.Width: 1.5<br />Species: versicolor","Petal.Length: 4.0<br />Petal.Width: 1.3<br />Species: versicolor","Petal.Length: 4.6<br />Petal.Width: 1.5<br />Species: versicolor","Petal.Length: 4.5<br />Petal.Width: 1.3<br />Species: versicolor","Petal.Length: 4.7<br />Petal.Width: 1.6<br />Species: versicolor","Petal.Length: 3.3<br />Petal.Width: 1.0<br />Species: versicolor","Petal.Length: 4.6<br />Petal.Width: 1.3<br />Species: versicolor","Petal.Length: 3.9<br />Petal.Width: 1.4<br />Species: versicolor","Petal.Length: 3.5<br />Petal.Width: 1.0<br />Species: versicolor","Petal.Length: 4.2<br />Petal.Width: 1.5<br />Species: versicolor","Petal.Length: 4.0<br />Petal.Width: 1.0<br />Species: versicolor","Petal.Length: 4.7<br />Petal.Width: 1.4<br />Species: versicolor","Petal.Length: 3.6<br />Petal.Width: 1.3<br />Species: versicolor","Petal.Length: 4.4<br />Petal.Width: 1.4<br />Species: versicolor","Petal.Length: 4.5<br />Petal.Width: 1.5<br />Species: versicolor","Petal.Length: 4.1<br />Petal.Width: 1.0<br />Species: versicolor","Petal.Length: 4.5<br />Petal.Width: 1.5<br />Species: versicolor","Petal.Length: 3.9<br />Petal.Width: 1.1<br />Species: versicolor","Petal.Length: 4.8<br />Petal.Width: 1.8<br />Species: versicolor","Petal.Length: 4.0<br />Petal.Width: 1.3<br />Species: versicolor","Petal.Length: 4.9<br />Petal.Width: 1.5<br />Species: versicolor","Petal.Length: 4.7<br />Petal.Width: 1.2<br />Species: versicolor","Petal.Length: 4.3<br />Petal.Width: 1.3<br />Species: versicolor","Petal.Length: 4.4<br />Petal.Width: 1.4<br />Species: versicolor","Petal.Length: 4.8<br />Petal.Width: 1.4<br />Species: versicolor","Petal.Length: 5.0<br />Petal.Width: 1.7<br />Species: versicolor","Petal.Length: 4.5<br />Petal.Width: 1.5<br />Species: versicolor","Petal.Length: 3.5<br />Petal.Width: 1.0<br />Species: versicolor","Petal.Length: 3.8<br />Petal.Width: 1.1<br />Species: versicolor","Petal.Length: 3.7<br />Petal.Width: 1.0<br />Species: versicolor","Petal.Length: 3.9<br />Petal.Width: 1.2<br />Species: versicolor","Petal.Length: 5.1<br />Petal.Width: 1.6<br />Species: versicolor","Petal.Length: 4.5<br />Petal.Width: 1.5<br />Species: versicolor","Petal.Length: 4.5<br />Petal.Width: 1.6<br />Species: versicolor","Petal.Length: 4.7<br />Petal.Width: 1.5<br />Species: versicolor","Petal.Length: 4.4<br />Petal.Width: 1.3<br />Species: versicolor","Petal.Length: 4.1<br />Petal.Width: 1.3<br />Species: versicolor","Petal.Length: 4.0<br />Petal.Width: 1.3<br />Species: versicolor","Petal.Length: 4.4<br />Petal.Width: 1.2<br />Species: versicolor","Petal.Length: 4.6<br />Petal.Width: 1.4<br />Species: versicolor","Petal.Length: 4.0<br />Petal.Width: 1.2<br />Species: versicolor","Petal.Length: 3.3<br />Petal.Width: 1.0<br />Species: versicolor","Petal.Length: 4.2<br />Petal.Width: 1.3<br />Species: versicolor","Petal.Length: 4.2<br />Petal.Width: 1.2<br />Species: versicolor","Petal.Length: 4.2<br />Petal.Width: 1.3<br />Species: versicolor","Petal.Length: 4.3<br />Petal.Width: 1.3<br />Species: versicolor","Petal.Length: 3.0<br />Petal.Width: 1.1<br />Species: versicolor","Petal.Length: 4.1<br />Petal.Width: 1.3<br />Species: versicolor"],"key":["51","52","53","54","55","56","57","58","59","60","61","62","63","64","65","66","67","68","69","70","71","72","73","74","75","76","77","78","79","80","81","82","83","84","85","86","87","88","89","90","91","92","93","94","95","96","97","98","99","100"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(0,186,56,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(0,186,56,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"versicolor","legendgroup":"versicolor","showlegend":true,"xaxis":"x3","yaxis":"y9","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[6,5.1,5.9,5.6,5.8,6.6,4.5,6.3,5.8,6.1,5.1,5.3,5.5,5,5.1,5.3,5.5,6.7,6.9,5,5.7,4.9,6.7,4.9,5.7,6,4.8,4.9,5.6,5.8,6.1,6.4,5.6,5.1,5.6,6.1,5.6,5.5,4.8,5.4,5.6,5.1,5.1,5.9,5.7,5.2,5,5.2,5.4,5.1],"y":[2.5,1.9,2.1,1.8,2.2,2.1,1.7,1.8,1.8,2.5,2,1.9,2.1,2,2.4,2.3,1.8,2.2,2.3,1.5,2.3,2,2,1.8,2.1,1.8,1.8,1.8,2.1,1.6,1.9,2,2.2,1.5,1.4,2.3,2.4,1.8,1.8,2.1,2.4,2.3,1.9,2.3,2.5,2.3,1.9,2,2.3,1.8],"text":["Petal.Length: 6.0<br />Petal.Width: 2.5<br />Species: virginica","Petal.Length: 5.1<br />Petal.Width: 1.9<br />Species: virginica","Petal.Length: 5.9<br />Petal.Width: 2.1<br />Species: virginica","Petal.Length: 5.6<br />Petal.Width: 1.8<br />Species: virginica","Petal.Length: 5.8<br />Petal.Width: 2.2<br />Species: virginica","Petal.Length: 6.6<br />Petal.Width: 2.1<br />Species: virginica","Petal.Length: 4.5<br />Petal.Width: 1.7<br />Species: virginica","Petal.Length: 6.3<br />Petal.Width: 1.8<br />Species: virginica","Petal.Length: 5.8<br />Petal.Width: 1.8<br />Species: virginica","Petal.Length: 6.1<br />Petal.Width: 2.5<br />Species: virginica","Petal.Length: 5.1<br />Petal.Width: 2.0<br />Species: virginica","Petal.Length: 5.3<br />Petal.Width: 1.9<br />Species: virginica","Petal.Length: 5.5<br />Petal.Width: 2.1<br />Species: virginica","Petal.Length: 5.0<br />Petal.Width: 2.0<br />Species: virginica","Petal.Length: 5.1<br />Petal.Width: 2.4<br />Species: virginica","Petal.Length: 5.3<br />Petal.Width: 2.3<br />Species: virginica","Petal.Length: 5.5<br />Petal.Width: 1.8<br />Species: virginica","Petal.Length: 6.7<br />Petal.Width: 2.2<br />Species: virginica","Petal.Length: 6.9<br />Petal.Width: 2.3<br />Species: virginica","Petal.Length: 5.0<br />Petal.Width: 1.5<br />Species: virginica","Petal.Length: 5.7<br />Petal.Width: 2.3<br />Species: virginica","Petal.Length: 4.9<br />Petal.Width: 2.0<br />Species: virginica","Petal.Length: 6.7<br />Petal.Width: 2.0<br />Species: virginica","Petal.Length: 4.9<br />Petal.Width: 1.8<br />Species: virginica","Petal.Length: 5.7<br />Petal.Width: 2.1<br />Species: virginica","Petal.Length: 6.0<br />Petal.Width: 1.8<br />Species: virginica","Petal.Length: 4.8<br />Petal.Width: 1.8<br />Species: virginica","Petal.Length: 4.9<br />Petal.Width: 1.8<br />Species: virginica","Petal.Length: 5.6<br />Petal.Width: 2.1<br />Species: virginica","Petal.Length: 5.8<br />Petal.Width: 1.6<br />Species: virginica","Petal.Length: 6.1<br />Petal.Width: 1.9<br />Species: virginica","Petal.Length: 6.4<br />Petal.Width: 2.0<br />Species: virginica","Petal.Length: 5.6<br />Petal.Width: 2.2<br />Species: virginica","Petal.Length: 5.1<br />Petal.Width: 1.5<br />Species: virginica","Petal.Length: 5.6<br />Petal.Width: 1.4<br />Species: virginica","Petal.Length: 6.1<br />Petal.Width: 2.3<br />Species: virginica","Petal.Length: 5.6<br />Petal.Width: 2.4<br />Species: virginica","Petal.Length: 5.5<br />Petal.Width: 1.8<br />Species: virginica","Petal.Length: 4.8<br />Petal.Width: 1.8<br />Species: virginica","Petal.Length: 5.4<br />Petal.Width: 2.1<br />Species: virginica","Petal.Length: 5.6<br />Petal.Width: 2.4<br />Species: virginica","Petal.Length: 5.1<br />Petal.Width: 2.3<br />Species: virginica","Petal.Length: 5.1<br />Petal.Width: 1.9<br />Species: virginica","Petal.Length: 5.9<br />Petal.Width: 2.3<br />Species: virginica","Petal.Length: 5.7<br />Petal.Width: 2.5<br />Species: virginica","Petal.Length: 5.2<br />Petal.Width: 2.3<br />Species: virginica","Petal.Length: 5.0<br />Petal.Width: 1.9<br />Species: virginica","Petal.Length: 5.2<br />Petal.Width: 2.0<br />Species: virginica","Petal.Length: 5.4<br />Petal.Width: 2.3<br />Species: virginica","Petal.Length: 5.1<br />Petal.Width: 1.8<br />Species: virginica"],"key":["101","102","103","104","105","106","107","108","109","110","111","112","113","114","115","116","117","118","119","120","121","122","123","124","125","126","127","128","129","130","131","132","133","134","135","136","137","138","139","140","141","142","143","144","145","146","147","148","149","150"],"type":"scatter","mode":"markers","marker":{"autocolorscale":false,"color":"rgba(97,156,255,1)","opacity":1,"size":5.66929133858268,"symbol":"circle","line":{"width":1.88976377952756,"color":"rgba(97,156,255,1)"}},"hoveron":"points","set":"SharedData91833e3a","name":"virginica","legendgroup":"virginica","showlegend":true,"xaxis":"x3","yaxis":"y9","hoverinfo":"text","_isNestedKey":false,"frame":null},{"x":[1.3],"y":[7.5688],"text":"Corr: 0.818***","hovertext":"x: 1.3<br />y: 7.5688","textfont":{"size":14.6645669291339,"color":"rgba(118,118,118,1)"},"type":"scatter","mode":"text","hoveron":"points","showlegend":false,"xaxis":"x4","yaxis":"y16","hoverinfo":"text","frame":null},{"x":[1.3],"y":[6.712],"text":" setosa: 0.278. ","hovertext":"xPos: 1.3<br />yPos: 6.7120<br />labelp: setosa: 0.278. ","textfont":{"size":14.6645669291339,"color":"rgba(248,118,109,1)"},"type":"scatter","mode":"text","hoveron":"points","name":" setosa: 0.278. ","legendgroup":" setosa: 0.278. ","showlegend":true,"xaxis":"x4","yaxis":"y16","hoverinfo":"text","frame":null},{"x":[1.3],"y":[5.8552],"text":"versicolor: 0.546***","hovertext":"xPos: 1.3<br />yPos: 5.8552<br />labelp: versicolor: 0.546***","textfont":{"size":14.6645669291339,"color":"rgba(0,186,56,1)"},"type":"scatter","mode":"text","hoveron":"points","name":"versicolor: 0.546***","legendgroup":"versicolor: 0.546***","showlegend":true,"xaxis":"x4","yaxis":"y16","hoverinfo":"text","frame":null},{"x":[1.3],"y":[4.9984],"text":" virginica: 0.281* ","hovertext":"xPos: 1.3<br />yPos: 4.9984<br />labelp: virginica: 0.281* ","textfont":{"size":14.6645669291339,"color":"rgba(97,156,255,1)"},"type":"scatter","mode":"text","hoveron":"points","name":" virginica: 0.281* ","legendgroup":" virginica: 0.281* ","showlegend":true,"xaxis":"x4","yaxis":"y16","hoverinfo":"text","frame":null},{"x":[1.3],"y":[4.1792],"text":"Corr: -0.366***","hovertext":"x: 1.3<br />y: 4.1792","textfont":{"size":14.6645669291339,"color":"rgba(118,118,118,1)"},"type":"scatter","mode":"text","hoveron":"points","showlegend":false,"xaxis":"x4","yaxis":"y15","hoverinfo":"text","frame":null},{"x":[1.3],"y":[3.608],"text":" setosa: 0.233 ","hovertext":"xPos: 1.3<br />yPos: 3.6080<br />labelp: setosa: 0.233 ","textfont":{"size":14.6645669291339,"color":"rgba(248,118,109,1)"},"type":"scatter","mode":"text","hoveron":"points","name":" setosa: 0.233 ","legendgroup":" setosa: 0.233 ","showlegend":true,"xaxis":"x4","yaxis":"y15","hoverinfo":"text","frame":null},{"x":[1.3],"y":[3.0368],"text":"versicolor: 0.664***","hovertext":"xPos: 1.3<br />yPos: 3.0368<br />labelp: versicolor: 0.664***","textfont":{"size":14.6645669291339,"color":"rgba(0,186,56,1)"},"type":"scatter","mode":"text","hoveron":"points","name":"versicolor: 0.664***","legendgroup":"versicolor: 0.664***","showlegend":true,"xaxis":"x4","yaxis":"y15","hoverinfo":"text","frame":null},{"x":[1.3],"y":[2.4656],"text":" virginica: 0.538***","hovertext":"xPos: 1.3<br />yPos: 2.4656<br />labelp: virginica: 0.538***","textfont":{"size":14.6645669291339,"color":"rgba(97,156,255,1)"},"type":"scatter","mode":"text","hoveron":"points","name":" virginica: 0.538***","legendgroup":" virginica: 0.538***","showlegend":true,"xaxis":"x4","yaxis":"y15","hoverinfo":"text","frame":null},{"x":[1.3],"y":[6.3572],"text":"Corr: 0.963***","hovertext":"x: 1.3<br />y: 6.3572","textfont":{"size":14.6645669291339,"color":"rgba(118,118,118,1)"},"type":"scatter","mode":"text","hoveron":"points","showlegend":false,"xaxis":"x4","yaxis":"y14","hoverinfo":"text","frame":null},{"x":[1.3],"y":[4.953],"text":" setosa: 0.332* ","hovertext":"xPos: 1.3<br />yPos: 4.9530<br />labelp: setosa: 0.332* ","textfont":{"size":14.6645669291339,"color":"rgba(248,118,109,1)"},"type":"scatter","mode":"text","hoveron":"points","name":" setosa: 0.332* ","legendgroup":" setosa: 0.332* ","showlegend":true,"xaxis":"x4","yaxis":"y14","hoverinfo":"text","frame":null},{"x":[1.3],"y":[3.5488],"text":"versicolor: 0.787***","hovertext":"xPos: 1.3<br />yPos: 3.5488<br />labelp: versicolor: 0.787***","textfont":{"size":14.6645669291339,"color":"rgba(0,186,56,1)"},"type":"scatter","mode":"text","hoveron":"points","name":"versicolor: 0.787***","legendgroup":"versicolor: 0.787***","showlegend":true,"xaxis":"x4","yaxis":"y14","hoverinfo":"text","frame":null},{"x":[1.3],"y":[2.1446],"text":" virginica: 0.322* ","hovertext":"xPos: 1.3<br />yPos: 2.1446<br />labelp: virginica: 0.322* ","textfont":{"size":14.6645669291339,"color":"rgba(97,156,255,1)"},"type":"scatter","mode":"text","hoveron":"points","name":" virginica: 0.322* ","legendgroup":" virginica: 0.322* ","showlegend":true,"xaxis":"x4","yaxis":"y14","hoverinfo":"text","frame":null},{"x":[0.1,0.104696673189824,0.109393346379648,0.114090019569472,0.118786692759296,0.123483365949119,0.128180039138943,0.132876712328767,0.137573385518591,0.142270058708415,0.146966731898239,0.151663405088063,0.156360078277887,0.16105675146771,0.165753424657534,0.170450097847358,0.175146771037182,0.179843444227006,0.18454011741683,0.189236790606654,0.193933463796477,0.198630136986301,0.203326810176125,0.208023483365949,0.212720156555773,0.217416829745597,0.222113502935421,0.226810176125245,0.231506849315068,0.236203522504892,0.240900195694716,0.24559686888454,0.250293542074364,0.254990215264188,0.259686888454012,0.264383561643836,0.26908023483366,0.273776908023483,0.278473581213307,0.283170254403131,0.287866927592955,0.292563600782779,0.297260273972603,0.301956947162427,0.306653620352251,0.311350293542074,0.316046966731898,0.320743639921722,0.325440313111546,0.33013698630137,0.334833659491194,0.339530332681018,0.344227005870841,0.348923679060665,0.353620352250489,0.358317025440313,0.363013698630137,0.367710371819961,0.372407045009785,0.377103718199609,0.381800391389432,0.386497064579256,0.39119373776908,0.395890410958904,0.400587084148728,0.405283757338552,0.409980430528376,0.4146771037182,0.419373776908023,0.424070450097847,0.428767123287671,0.433463796477495,0.438160469667319,0.442857142857143,0.447553816046967,0.452250489236791,0.456947162426614,0.461643835616438,0.466340508806262,0.471037181996086,0.47573385518591,0.480430528375734,0.485127201565558,0.489823874755382,0.494520547945206,0.499217221135029,0.503913894324853,0.508610567514677,0.513307240704501,0.518003913894325,0.522700587084149,0.527397260273973,0.532093933463797,0.53679060665362,0.541487279843444,0.546183953033268,0.550880626223092,0.555577299412916,0.56027397260274,0.564970645792564,0.569667318982387,0.574363992172211,0.579060665362035,0.583757338551859,0.588454011741683,0.593150684931507,0.597847358121331,0.602544031311155,0.607240704500978,0.611937377690802,0.616634050880626,0.62133072407045,0.626027397260274,0.630724070450098,0.635420743639922,0.640117416829746,0.644814090019569,0.649510763209393,0.654207436399217,0.658904109589041,0.663600782778865,0.668297455968689,0.672994129158513,0.677690802348337,0.68238747553816,0.687084148727984,0.691780821917808,0.696477495107632,0.701174168297456,0.70587084148728,0.710567514677104,0.715264187866928,0.719960861056751,0.724657534246575,0.729354207436399,0.734050880626223,0.738747553816047,0.743444227005871,0.748140900195695,0.752837573385519,0.757534246575342,0.762230919765166,0.76692759295499,0.771624266144814,0.776320939334638,0.781017612524462,0.785714285714286,0.79041095890411,0.795107632093933,0.799804305283757,0.804500978473581,0.809197651663405,0.813894324853229,0.818590998043053,0.823287671232877,0.827984344422701,0.832681017612524,0.837377690802348,0.842074363992172,0.846771037181996,0.85146771037182,0.856164383561644,0.860861056751468,0.865557729941292,0.870254403131115,0.874951076320939,0.879647749510763,0.884344422700587,0.889041095890411,0.893737769080235,0.898434442270059,0.903131115459883,0.907827788649706,0.91252446183953,0.917221135029354,0.921917808219178,0.926614481409002,0.931311154598826,0.93600782778865,0.940704500978474,0.945401174168297,0.950097847358121,0.954794520547945,0.959491193737769,0.964187866927593,0.968884540117417,0.973581213307241,0.978277886497064,0.982974559686888,0.987671232876712,0.992367906066536,0.99706457925636,1.00176125244618,1.00645792563601,1.01115459882583,1.01585127201566,1.02054794520548,1.0252446183953,1.02994129158513,1.03463796477495,1.03933463796477,1.0440313111546,1.04872798434442,1.05342465753425,1.05812133072407,1.06281800391389,1.06751467710372,1.07221135029354,1.07690802348337,1.08160469667319,1.08630136986301,1.09099804305284,1.09569471624266,1.10039138943249,1.10508806262231,1.10978473581213,1.11448140900196,1.11917808219178,1.1238747553816,1.12857142857143,1.13326810176125,1.13796477495108,1.1426614481409,1.14735812133072,1.15205479452055,1.15675146771037,1.1614481409002,1.16614481409002,1.17084148727984,1.17553816046967,1.18023483365949,1.18493150684932,1.18962818003914,1.19432485322896,1.19902152641879,1.20371819960861,1.20841487279843,1.21311154598826,1.21780821917808,1.22250489236791,1.22720156555773,1.23189823874755,1.23659491193738,1.2412915851272,1.24598825831703,1.25068493150685,1.25538160469667,1.2600782778865,1.26477495107632,1.26947162426614,1.27416829745597,1.27886497064579,1.28356164383562,1.28825831702544,1.29295499021526,1.29765166340509,1.30234833659491,1.30704500978474,1.31174168297456,1.31643835616438,1.32113502935421,1.32583170254403,1.33052837573386,1.33522504892368,1.3399217221135,1.34461839530333,1.34931506849315,1.35401174168297,1.3587084148728,1.36340508806262,1.36810176125245,1.37279843444227,1.37749510763209,1.38219178082192,1.38688845401174,1.39158512720157,1.39628180039139,1.40097847358121,1.40567514677104,1.41037181996086,1.41506849315068,1.41976516634051,1.42446183953033,1.42915851272016,1.43385518590998,1.4385518590998,1.44324853228963,1.44794520547945,1.45264187866928,1.4573385518591,1.46203522504892,1.46673189823875,1.47142857142857,1.4761252446184,1.48082191780822,1.48551859099804,1.49021526418787,1.49491193737769,1.49960861056751,1.50430528375734,1.50900195694716,1.51369863013699,1.51839530332681,1.52309197651663,1.52778864970646,1.53248532289628,1.53718199608611,1.54187866927593,1.54657534246575,1.55127201565558,1.5559686888454,1.56066536203523,1.56536203522505,1.57005870841487,1.5747553816047,1.57945205479452,1.58414872798434,1.58884540117417,1.59354207436399,1.59823874755382,1.60293542074364,1.60763209393346,1.61232876712329,1.61702544031311,1.62172211350294,1.62641878669276,1.63111545988258,1.63581213307241,1.64050880626223,1.64520547945205,1.64990215264188,1.6545988258317,1.65929549902153,1.66399217221135,1.66868884540117,1.673385518591,1.67808219178082,1.68277886497065,1.68747553816047,1.69217221135029,1.69686888454012,1.70156555772994,1.70626223091977,1.71095890410959,1.71565557729941,1.72035225048924,1.72504892367906,1.72974559686888,1.73444227005871,1.73913894324853,1.74383561643836,1.74853228962818,1.753228962818,1.75792563600783,1.76262230919765,1.76731898238748,1.7720156555773,1.77671232876712,1.78140900195695,1.78610567514677,1.79080234833659,1.79549902152642,1.80019569471624,1.80489236790607,1.80958904109589,1.81428571428571,1.81898238747554,1.82367906066536,1.82837573385519,1.83307240704501,1.83776908023483,1.84246575342466,1.84716242661448,1.85185909980431,1.85655577299413,1.86125244618395,1.86594911937378,1.8706457925636,1.87534246575342,1.88003913894325,1.88473581213307,1.8894324853229,1.89412915851272,1.89882583170254,1.90352250489237,1.90821917808219,1.91291585127202,1.91761252446184,1.92230919765166,1.92700587084149,1.93170254403131,1.93639921722114,1.94109589041096,1.94579256360078,1.95048923679061,1.95518590998043,1.95988258317025,1.96457925636008,1.9692759295499,1.97397260273973,1.97866927592955,1.98336594911937,1.9880626223092,1.99275929549902,1.99745596868885,2.00215264187867,2.00684931506849,2.01154598825832,2.01624266144814,2.02093933463796,2.02563600782779,2.03033268101761,2.03502935420744,2.03972602739726,2.04442270058708,2.04911937377691,2.05381604696673,2.05851272015656,2.06320939334638,2.0679060665362,2.07260273972603,2.07729941291585,2.08199608610568,2.0866927592955,2.09138943248532,2.09608610567515,2.10078277886497,2.10547945205479,2.11017612524462,2.11487279843444,2.11956947162427,2.12426614481409,2.12896281800391,2.13365949119374,2.13835616438356,2.14305283757339,2.14774951076321,2.15244618395303,2.15714285714286,2.16183953033268,2.1665362035225,2.17123287671233,2.17592954990215,2.18062622309198,2.1853228962818,2.19001956947162,2.19471624266145,2.19941291585127,2.2041095890411,2.20880626223092,2.21350293542074,2.21819960861057,2.22289628180039,2.22759295499022,2.23228962818004,2.23698630136986,2.24168297455969,2.24637964774951,2.25107632093933,2.25577299412916,2.26046966731898,2.26516634050881,2.26986301369863,2.27455968688845,2.27925636007828,2.2839530332681,2.28864970645793,2.29334637964775,2.29804305283757,2.3027397260274,2.30743639921722,2.31213307240705,2.31682974559687,2.32152641878669,2.32622309197652,2.33091976516634,2.33561643835616,2.34031311154599,2.34500978473581,2.34970645792564,2.35440313111546,2.35909980430528,2.36379647749511,2.36849315068493,2.37318982387476,2.37788649706458,2.3825831702544,2.38727984344423,2.39197651663405,2.39667318982387,2.4013698630137,2.40606653620352,2.41076320939335,2.41545988258317,2.42015655577299,2.42485322896282,2.42954990215264,2.43424657534247,2.43894324853229,2.44363992172211,2.44833659491194,2.45303326810176,2.45772994129159,2.46242661448141,2.46712328767123,2.47181996086106,2.47651663405088,2.4812133072407,2.48590998043053,2.49060665362035,2.49530332681018,2.5,2.5,2.49530332681018,2.49060665362035,2.48590998043053,2.4812133072407,2.47651663405088,2.47181996086106,2.46712328767123,2.46242661448141,2.45772994129159,2.45303326810176,2.44833659491194,2.44363992172211,2.43894324853229,2.43424657534247,2.42954990215264,2.42485322896282,2.42015655577299,2.41545988258317,2.41076320939335,2.40606653620352,2.4013698630137,2.39667318982387,2.39197651663405,2.38727984344423,2.3825831702544,2.37788649706458,2.37318982387476,2.36849315068493,2.36379647749511,2.35909980430528,2.35440313111546,2.34970645792564,2.34500978473581,2.34031311154599,2.33561643835616,2.33091976516634,2.32622309197652,2.32152641878669,2.31682974559687,2.31213307240705,2.30743639921722,2.3027397260274,2.29804305283757,2.29334637964775,2.28864970645793,2.2839530332681,2.27925636007828,2.27455968688845,2.26986301369863,2.26516634050881,2.26046966731898,2.25577299412916,2.25107632093933,2.24637964774951,2.24168297455969,2.23698630136986,2.23228962818004,2.22759295499022,2.22289628180039,2.21819960861057,2.21350293542074,2.20880626223092,2.2041095890411,2.19941291585127,2.19471624266145,2.19001956947162,2.1853228962818,2.18062622309198,2.17592954990215,2.17123287671233,2.1665362035225,2.16183953033268,2.15714285714286,2.15244618395303,2.14774951076321,2.14305283757339,2.13835616438356,2.13365949119374,2.12896281800391,2.12426614481409,2.11956947162427,2.11487279843444,2.11017612524462,2.10547945205479,2.10078277886497,2.09608610567515,2.09138943248532,2.0866927592955,2.08199608610568,2.07729941291585,2.07260273972603,2.0679060665362,2.06320939334638,2.05851272015656,2.05381604696673,2.04911937377691,2.04442270058708,2.03972602739726,2.03502935420744,2.03033268101761,2.02563600782779,2.02093933463796,2.01624266144814,2.01154598825832,2.00684931506849,2.00215264187867,1.99745596868885,1.99275929549902,1.9880626223092,1.98336594911937,1.97866927592955,1.97397260273973,1.9692759295499,1.96457925636008,1.95988258317025,1.95518590998043,1.95048923679061,1.94579256360078,1.94109589041096,1.93639921722114,1.93170254403131,1.92700587084149,1.92230919765166,1.91761252446184,1.91291585127202,1.90821917808219,1.90352250489237,1.89882583170254,1.89412915851272,1.8894324853229,1.88473581213307,1.88003913894325,1.87534246575342,1.8706457925636,1.86594911937378,1.86125244618395,1.85655577299413,1.85185909980431,1.84716242661448,1.84246575342466,1.83776908023483,1.83307240704501,1.82837573385519,1.82367906066536,1.81898238747554,1.81428571428571,1.80958904109589,1.80489236790607,1.80019569471624,1.79549902152642,1.79080234833659,1.78610567514677,1.78140900195695,1.77671232876712,1.7720156555773,1.76731898238748,1.76262230919765,1.75792563600783,1.753228962818,1.74853228962818,1.74383561643836,1.73913894324853,1.73444227005871,1.72974559686888,1.72504892367906,1.72035225048924,1.71565557729941,1.71095890410959,1.70626223091977,1.70156555772994,1.69686888454012,1.69217221135029,1.68747553816047,1.68277886497065,1.67808219178082,1.673385518591,1.66868884540117,1.66399217221135,1.65929549902153,1.6545988258317,1.64990215264188,1.64520547945205,1.64050880626223,1.63581213307241,1.63111545988258,1.62641878669276,1.62172211350294,1.61702544031311,1.61232876712329,1.60763209393346,1.60293542074364,1.59823874755382,1.59354207436399,1.58884540117417,1.58414872798434,1.57945205479452,1.5747553816047,1.57005870841487,1.56536203522505,1.56066536203523,1.5559686888454,1.55127201565558,1.54657534246575,1.54187866927593,1.53718199608611,1.53248532289628,1.52778864970646,1.52309197651663,1.51839530332681,1.51369863013699,1.50900195694716,1.50430528375734,1.49960861056751,1.49491193737769,1.49021526418787,1.48551859099804,1.48082191780822,1.4761252446184,1.47142857142857,1.46673189823875,1.46203522504892,1.4573385518591,1.45264187866928,1.44794520547945,1.44324853228963,1.4385518590998,1.43385518590998,1.42915851272016,1.42446183953033,1.41976516634051,1.41506849315068,1.41037181996086,1.40567514677104,1.40097847358121,1.39628180039139,1.39158512720157,1.38688845401174,1.38219178082192,1.37749510763209,1.37279843444227,1.36810176125245,1.36340508806262,1.3587084148728,1.35401174168297,1.34931506849315,1.34461839530333,1.3399217221135,1.33522504892368,1.33052837573386,1.32583170254403,1.32113502935421,1.31643835616438,1.31174168297456,1.30704500978474,1.30234833659491,1.29765166340509,1.29295499021526,1.28825831702544,1.28356164383562,1.27886497064579,1.27416829745597,1.26947162426614,1.26477495107632,1.2600782778865,1.25538160469667,1.25068493150685,1.24598825831703,1.2412915851272,1.23659491193738,1.23189823874755,1.22720156555773,1.22250489236791,1.21780821917808,1.21311154598826,1.20841487279843,1.20371819960861,1.19902152641879,1.19432485322896,1.18962818003914,1.18493150684932,1.18023483365949,1.17553816046967,1.17084148727984,1.16614481409002,1.1614481409002,1.15675146771037,1.15205479452055,1.14735812133072,1.1426614481409,1.13796477495108,1.13326810176125,1.12857142857143,1.1238747553816,1.11917808219178,1.11448140900196,1.10978473581213,1.10508806262231,1.10039138943249,1.09569471624266,1.09099804305284,1.08630136986301,1.08160469667319,1.07690802348337,1.07221135029354,1.06751467710372,1.06281800391389,1.05812133072407,1.05342465753425,1.04872798434442,1.0440313111546,1.03933463796477,1.03463796477495,1.02994129158513,1.0252446183953,1.02054794520548,1.01585127201566,1.01115459882583,1.00645792563601,1.00176125244618,0.99706457925636,0.992367906066536,0.987671232876712,0.982974559686888,0.978277886497064,0.973581213307241,0.968884540117417,0.964187866927593,0.959491193737769,0.954794520547945,0.950097847358121,0.945401174168297,0.940704500978474,0.93600782778865,0.931311154598826,0.926614481409002,0.921917808219178,0.917221135029354,0.91252446183953,0.907827788649706,0.903131115459883,0.898434442270059,0.893737769080235,0.889041095890411,0.884344422700587,0.879647749510763,0.874951076320939,0.870254403131115,0.865557729941292,0.860861056751468,0.856164383561644,0.85146771037182,0.846771037181996,0.842074363992172,0.837377690802348,0.832681017612524,0.827984344422701,0.823287671232877,0.818590998043053,0.813894324853229,0.809197651663405,0.804500978473581,0.799804305283757,0.795107632093933,0.79041095890411,0.785714285714286,0.781017612524462,0.776320939334638,0.771624266144814,0.76692759295499,0.762230919765166,0.757534246575342,0.752837573385519,0.748140900195695,0.743444227005871,0.738747553816047,0.734050880626223,0.729354207436399,0.724657534246575,0.719960861056751,0.715264187866928,0.710567514677104,0.70587084148728,0.701174168297456,0.696477495107632,0.691780821917808,0.687084148727984,0.68238747553816,0.677690802348337,0.672994129158513,0.668297455968689,0.663600782778865,0.658904109589041,0.654207436399217,0.649510763209393,0.644814090019569,0.640117416829746,0.635420743639922,0.630724070450098,0.626027397260274,0.62133072407045,0.616634050880626,0.611937377690802,0.607240704500978,0.602544031311155,0.597847358121331,0.593150684931507,0.588454011741683,0.583757338551859,0.579060665362035,0.574363992172211,0.569667318982387,0.564970645792564,0.56027397260274,0.555577299412916,0.550880626223092,0.546183953033268,0.541487279843444,0.53679060665362,0.532093933463797,0.527397260273973,0.522700587084149,0.518003913894325,0.513307240704501,0.508610567514677,0.503913894324853,0.499217221135029,0.494520547945206,0.489823874755382,0.485127201565558,0.480430528375734,0.47573385518591,0.471037181996086,0.466340508806262,0.461643835616438,0.456947162426614,0.452250489236791,0.447553816046967,0.442857142857143,0.438160469667319,0.433463796477495,0.428767123287671,0.424070450097847,0.419373776908023,0.4146771037182,0.409980430528376,0.405283757338552,0.400587084148728,0.395890410958904,0.39119373776908,0.386497064579256,0.381800391389432,0.377103718199609,0.372407045009785,0.367710371819961,0.363013698630137,0.358317025440313,0.353620352250489,0.348923679060665,0.344227005870841,0.339530332681018,0.334833659491194,0.33013698630137,0.325440313111546,0.320743639921722,0.316046966731898,0.311350293542074,0.306653620352251,0.301956947162427,0.297260273972603,0.292563600782779,0.287866927592955,0.283170254403131,0.278473581213307,0.273776908023483,0.26908023483366,0.264383561643836,0.259686888454012,0.254990215264188,0.250293542074364,0.24559686888454,0.240900195694716,0.236203522504892,0.231506849315068,0.226810176125245,0.222113502935421,0.217416829745597,0.212720156555773,0.208023483365949,0.203326810176125,0.198630136986301,0.193933463796477,0.189236790606654,0.18454011741683,0.179843444227006,0.175146771037182,0.170450097847358,0.165753424657534,0.16105675146771,0.156360078277887,0.151663405088063,0.146966731898239,0.142270058708415,0.137573385518591,0.132876712328767,0.128180039138943,0.123483365949119,0.118786692759296,0.114090019569472,0.109393346379648,0.104696673189824,0.1,0.1],"y":[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1.88415463161855e-17,1.02629576629897e-17,0,0,3.49871786190468e-17,7.88098298478689e-17,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,9.93669342272487e-17,4.40573882310217e-17,0,0,7.09112344929855e-18,1.30030340346012e-17,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,3.27161027824412e-18,1.10539644401533e-17,0,0,0,0,0,0,0,0,6.34187064375916e-17,2.15553842671771e-16,2.38849698515583e-16,0,0,1.00857323220436e-16,1.25250339309831e-16,5.89748769197831e-17,4.19530194241353e-17,2.78702245360349e-17,6.19202273357496e-17,2.00688689394646e-16,2.90809193187918e-16,2.44624967293684e-16,1.88792790352593e-16,3.18604035485706e-16,1.47767938957661e-16,1.64414106745456e-16,1.19506523103649e-16,1.30251330590773e-16,9.6450537346528e-17,7.09056270187452e-17,1.53075646586248e-16,1.55106641493728e-16,2.74104972429594e-16,7.2419965303132e-17,2.32129484259539e-16,1.7287652263723e-16,8.25536090209945e-17,1.76182792300934e-16,1.36719275241051e-16,2.19845834300325e-17,0,7.07870855946203e-18,2.32776493296329e-17,8.06393941735445e-19,1.95868712591819e-20,0,0,0,0,8.01002752503199e-18,9.05246777397808e-18,0,8.9468511718057e-18,3.27614706702182e-17,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,3.86669525333532e-17,2.57913630406838e-17,0,0,0,0,0,0,0,0,1.51307212264593e-17,7.61927250555375e-18,0,0,0,0,0,2.66256180736465e-19,3.41348446264137e-18,1.2093754707048e-16,7.99274886753061e-17,5.12385213721446e-17,1.44959470150412e-16,1.00023350373312e-16,0,9.26363017841025e-18,3.10985160246398e-17,3.54711662369935e-17,1.53038209724356e-18,2.6708208352484e-17,3.62511662353344e-18,2.1552709631707e-17,8.78061534431044e-18,0,0,4.49668487606071e-17,7.63664511434625e-17,0,0,2.79214424578977e-18,7.8325623011511e-17,1.53859101777232e-16,4.08151999954135e-17,0,9.47852613736909e-17,8.72146884824136e-17,0,0,8.22981918108993e-18,3.72701682829362e-17,9.93142199848859e-19,7.65266209655701e-17,1.44733539624821e-17,2.03533749344118e-17,9.97995004160557e-18,4.36938941141983e-17,5.3768268260115e-17,1.43919734747525e-17,1.07304029404213e-17,9.76248954426434e-17,3.37165412202191e-16,3.55921432025568e-18,1.44784761656775e-16,5.94420109490237e-17,1.51982108310008e-16,7.60409999924487e-17,0,2.77973126043833e-17,5.35617196989073e-17,2.48076051127731e-17,3.26899871036651e-17,2.03302636511836e-16,8.05607335175389e-17,9.47660080877181e-18,0,0,8.2881454383641e-17,6.8785170496446e-17,6.85247529357001e-17,1.13475196920404e-16,1.19040575268287e-16,4.57292599276044e-16,1.22077418583562e-17,2.89163830665988e-16,1.80834505823373e-16,3.80848568589314e-17,0,0,0,0,4.533856495803e-17,1.39553905560868e-16,7.86282902102537e-17,0,0,0,0,0,0,0,4.08858298346243e-17,8.04474700694467e-17,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,3.85671204047871e-18,7.91965625036999e-17,1.15701361214362e-17,0,0,9.31068062225851e-17,8.17069375242413e-17,2.14710532259661e-17,7.23580606615026e-17,1.12268337958211e-16,1.00855079922352e-16,6.50513673867948e-17,7.44747639521454e-18,5.03135742654711e-19,0,0,0,0,1.83587823373918e-17,8.45918589712589e-17,1.53756015249345e-16,5.80950465490228e-17,2.78343779246182e-17,2.50334963375408e-18,0,4.83596256172459e-17,2.93938066460861e-16,1.96622235278595e-16,0,0,3.56444002622788e-17,1.14043417417842e-16,0,0,0,5.58653843425262e-17,2.08474599280955e-17,5.08249353653316e-17,7.07904813803315e-17,4.08121465771699e-17,1.06882508755058e-17,0,1.5553407445586e-17,1.10950304589896e-16,1.83777244186312e-16,2.48427208255176e-17,0,0,7.88640387507201e-17,6.75847049532839e-17,0,0,7.83634787328563e-18,3.16242979647842e-17,0,0,0,0,0,0,0,8.61241534967738e-18,1.60806509488137e-17,0,0,0,0,0,0,0,0,0,0,0,2.40287174936201e-20,1.27049788230747e-19,1.64881847102477e-21,0,0,0,0,0,0,0,0,0,0,0,0,2.66604681905826e-16,3.53826905664536e-16,1.82440125646378e-16,2.78606927763169e-16,3.89840186174752e-16,3.47773891387485e-16,6.32334308325748e-16,1.1137371037938e-15,3.44597631611059e-15,1.15336063801298e-14,3.89775242633707e-14,1.27830056181721e-13,4.09588051313733e-13,1.26988797725907e-12,3.80563718123044e-12,1.10191818176116e-11,3.08019089059473e-11,8.30281880046672e-11,2.15514731220885e-10,5.3769854764537e-10,1.4146519900486e-09,3.68011919360915e-09,9.28905643369015e-09,2.27494429247301e-08,5.40582095089095e-08,1.24640920601122e-07,2.78870863382236e-07,6.05549828147965e-07,1.27642405457049e-06,2.6126175803161e-06,5.21227493760986e-06,1.06571792080748e-05,2.11848619911066e-05,4.09536968985397e-05,7.70164209423286e-05,0.000140948480020502,0.000251140015999656,0.000435883381009247,0.000737348565663843,0.00121646572470881,0.00195862286636598,0.00309880360143773,0.00485225402827561,0.00740315773695306,0.0110103999254251,0.0159698720898893,0.0226005289276121,0.0312221409358595,0.0421246801237608,0.0555303710131992,0.0715506801693475,0.0901418355925688,0.111165995500253,0.133992175418253,0.1576959060216,0.181282815858343,0.203626861539729,0.223555417661778,0.239949125005615,0.251846811685822,0.258544643753919,0.259678553496893,0.255279990849884,0.245456189830148,0.231572105206615,0.215040522137844,0.197310193810768,0.179889164890233,0.164230141364444,0.151624496624553,0.143111184828105,0.139405083100439,0.140847714478065,0.147382044077532,0.158946593314407,0.174009411098318,0.191344102881046,0.209571569831663,0.227275726548061,0.243137934629283,0.256076609792353,0.265382626328573,0.270840530419451,0.272825624478387,0.272359151016034,0.271523214014114,0.273119942067924,0.280209020475047,0.296141523905579,0.324333223849685,0.367994823215236,0.429828622570042,0.511706487401982,0.614350043688605,0.737040360499222,0.878027465467719,1.03205688557414,1.19150293203765,1.34859544660312,1.49488945032964,1.62194217568461,1.72205389722904,1.78899992629608,1.81867789783184,1.80959869358318,1.76316101405687,1.68178380828739,1.57550530209343,1.45502269636812,1.33074403388604,1.21312328531306,1.11187614381141,1.03528200548418,0.989609072142387,0.978687657026668,1.00364680914331,1.06282191220959,1.1540126362315,1.26612883052272,1.38968704463061,1.51465478968806,1.63103156735843,1.7297436173282,1.80352538638802,1.84772067222869,1.86093746188688,1.84549683513764,1.80744901763462,1.75812900982342,1.71424260816279,1.69239051465649,1.71008842775939,1.78447654459103,1.93091547622472,2.16153019382963,2.48378032611707,2.89915412664383,3.40210282222611,3.98144811178498,4.61188433921272,5.25748863334051,5.88305109947038,6.45156083795999,6.92720502366101,7.27845747490034,7.48093807747981,7.51973299470714,7.39090636632204,7.10200553871505,6.66394027739023,6.1144902212476,5.49490358112012,4.84172087031402,4.18988240857721,3.57024571921293,3.00774780834923,2.52028730960513,2.1183230735899,1.80511948667901,1.5775182775571,1.43302920973281,1.35084577165542,1.31467331439784,1.30918863836918,1.31982040197982,1.33362198802909,1.33992854951638,1.33078799914153,0],"text":["density: 1.330788e+00<br />Petal.Width: 0.1000000","density: 1.339929e+00<br />Petal.Width: 0.1046967","density: 1.333622e+00<br />Petal.Width: 0.1093933","density: 1.319820e+00<br />Petal.Width: 0.1140900","density: 1.309189e+00<br />Petal.Width: 0.1187867","density: 1.314673e+00<br />Petal.Width: 0.1234834","density: 1.350846e+00<br />Petal.Width: 0.1281800","density: 1.433029e+00<br />Petal.Width: 0.1328767","density: 1.577518e+00<br />Petal.Width: 0.1375734","density: 1.805119e+00<br />Petal.Width: 0.1422701","density: 2.118323e+00<br />Petal.Width: 0.1469667","density: 2.520287e+00<br />Petal.Width: 0.1516634","density: 3.007748e+00<br />Petal.Width: 0.1563601","density: 3.570246e+00<br />Petal.Width: 0.1610568","density: 4.189882e+00<br />Petal.Width: 0.1657534","density: 4.841721e+00<br />Petal.Width: 0.1704501","density: 5.494904e+00<br />Petal.Width: 0.1751468","density: 6.114490e+00<br />Petal.Width: 0.1798434","density: 6.663940e+00<br />Petal.Width: 0.1845401","density: 7.102006e+00<br />Petal.Width: 0.1892368","density: 7.390906e+00<br />Petal.Width: 0.1939335","density: 7.519733e+00<br />Petal.Width: 0.1986301","density: 7.480938e+00<br />Petal.Width: 0.2033268","density: 7.278457e+00<br />Petal.Width: 0.2080235","density: 6.927205e+00<br />Petal.Width: 0.2127202","density: 6.451561e+00<br />Petal.Width: 0.2174168","density: 5.883051e+00<br />Petal.Width: 0.2221135","density: 5.257489e+00<br />Petal.Width: 0.2268102","density: 4.611884e+00<br />Petal.Width: 0.2315068","density: 3.981448e+00<br />Petal.Width: 0.2362035","density: 3.402103e+00<br />Petal.Width: 0.2409002","density: 2.899154e+00<br />Petal.Width: 0.2455969","density: 2.483780e+00<br />Petal.Width: 0.2502935","density: 2.161530e+00<br />Petal.Width: 0.2549902","density: 1.930915e+00<br />Petal.Width: 0.2596869","density: 1.784477e+00<br />Petal.Width: 0.2643836","density: 1.710088e+00<br />Petal.Width: 0.2690802","density: 1.692391e+00<br />Petal.Width: 0.2737769","density: 1.714243e+00<br />Petal.Width: 0.2784736","density: 1.758129e+00<br />Petal.Width: 0.2831703","density: 1.807449e+00<br />Petal.Width: 0.2878669","density: 1.845497e+00<br />Petal.Width: 0.2925636","density: 1.860937e+00<br />Petal.Width: 0.2972603","density: 1.847721e+00<br />Petal.Width: 0.3019569","density: 1.803525e+00<br />Petal.Width: 0.3066536","density: 1.729744e+00<br />Petal.Width: 0.3113503","density: 1.631032e+00<br />Petal.Width: 0.3160470","density: 1.514655e+00<br />Petal.Width: 0.3207436","density: 1.389687e+00<br />Petal.Width: 0.3254403","density: 1.266129e+00<br />Petal.Width: 0.3301370","density: 1.154013e+00<br />Petal.Width: 0.3348337","density: 1.062822e+00<br />Petal.Width: 0.3395303","density: 1.003647e+00<br />Petal.Width: 0.3442270","density: 9.786877e-01<br />Petal.Width: 0.3489237","density: 9.896091e-01<br />Petal.Width: 0.3536204","density: 1.035282e+00<br />Petal.Width: 0.3583170","density: 1.111876e+00<br />Petal.Width: 0.3630137","density: 1.213123e+00<br />Petal.Width: 0.3677104","density: 1.330744e+00<br />Petal.Width: 0.3724070","density: 1.455023e+00<br />Petal.Width: 0.3771037","density: 1.575505e+00<br />Petal.Width: 0.3818004","density: 1.681784e+00<br />Petal.Width: 0.3864971","density: 1.763161e+00<br />Petal.Width: 0.3911937","density: 1.809599e+00<br />Petal.Width: 0.3958904","density: 1.818678e+00<br />Petal.Width: 0.4005871","density: 1.789000e+00<br />Petal.Width: 0.4052838","density: 1.722054e+00<br />Petal.Width: 0.4099804","density: 1.621942e+00<br />Petal.Width: 0.4146771","density: 1.494889e+00<br />Petal.Width: 0.4193738","density: 1.348595e+00<br />Petal.Width: 0.4240705","density: 1.191503e+00<br />Petal.Width: 0.4287671","density: 1.032057e+00<br />Petal.Width: 0.4334638","density: 8.780275e-01<br />Petal.Width: 0.4381605","density: 7.370404e-01<br />Petal.Width: 0.4428571","density: 6.143500e-01<br />Petal.Width: 0.4475538","density: 5.117065e-01<br />Petal.Width: 0.4522505","density: 4.298286e-01<br />Petal.Width: 0.4569472","density: 3.679948e-01<br />Petal.Width: 0.4616438","density: 3.243332e-01<br />Petal.Width: 0.4663405","density: 2.961415e-01<br />Petal.Width: 0.4710372","density: 2.802090e-01<br />Petal.Width: 0.4757339","density: 2.731199e-01<br />Petal.Width: 0.4804305","density: 2.715232e-01<br />Petal.Width: 0.4851272","density: 2.723592e-01<br />Petal.Width: 0.4898239","density: 2.728256e-01<br />Petal.Width: 0.4945205","density: 2.708405e-01<br />Petal.Width: 0.4992172","density: 2.653826e-01<br />Petal.Width: 0.5039139","density: 2.560766e-01<br />Petal.Width: 0.5086106","density: 2.431379e-01<br />Petal.Width: 0.5133072","density: 2.272757e-01<br />Petal.Width: 0.5180039","density: 2.095716e-01<br />Petal.Width: 0.5227006","density: 1.913441e-01<br />Petal.Width: 0.5273973","density: 1.740094e-01<br />Petal.Width: 0.5320939","density: 1.589466e-01<br />Petal.Width: 0.5367906","density: 1.473820e-01<br />Petal.Width: 0.5414873","density: 1.408477e-01<br />Petal.Width: 0.5461840","density: 1.394051e-01<br />Petal.Width: 0.5508806","density: 1.431112e-01<br />Petal.Width: 0.5555773","density: 1.516245e-01<br />Petal.Width: 0.5602740","density: 1.642301e-01<br />Petal.Width: 0.5649706","density: 1.798892e-01<br />Petal.Width: 0.5696673","density: 1.973102e-01<br />Petal.Width: 0.5743640","density: 2.150405e-01<br />Petal.Width: 0.5790607","density: 2.315721e-01<br />Petal.Width: 0.5837573","density: 2.454562e-01<br />Petal.Width: 0.5884540","density: 2.552800e-01<br />Petal.Width: 0.5931507","density: 2.596786e-01<br />Petal.Width: 0.5978474","density: 2.585446e-01<br />Petal.Width: 0.6025440","density: 2.518468e-01<br />Petal.Width: 0.6072407","density: 2.399491e-01<br />Petal.Width: 0.6119374","density: 2.235554e-01<br />Petal.Width: 0.6166341","density: 2.036269e-01<br />Petal.Width: 0.6213307","density: 1.812828e-01<br />Petal.Width: 0.6260274","density: 1.576959e-01<br />Petal.Width: 0.6307241","density: 1.339922e-01<br />Petal.Width: 0.6354207","density: 1.111660e-01<br />Petal.Width: 0.6401174","density: 9.014184e-02<br />Petal.Width: 0.6448141","density: 7.155068e-02<br />Petal.Width: 0.6495108","density: 5.553037e-02<br />Petal.Width: 0.6542074","density: 4.212468e-02<br />Petal.Width: 0.6589041","density: 3.122214e-02<br />Petal.Width: 0.6636008","density: 2.260053e-02<br />Petal.Width: 0.6682975","density: 1.596987e-02<br />Petal.Width: 0.6729941","density: 1.101040e-02<br />Petal.Width: 0.6776908","density: 7.403158e-03<br />Petal.Width: 0.6823875","density: 4.852254e-03<br />Petal.Width: 0.6870841","density: 3.098804e-03<br />Petal.Width: 0.6917808","density: 1.958623e-03<br />Petal.Width: 0.6964775","density: 1.216466e-03<br />Petal.Width: 0.7011742","density: 7.373486e-04<br />Petal.Width: 0.7058708","density: 4.358834e-04<br />Petal.Width: 0.7105675","density: 2.511400e-04<br />Petal.Width: 0.7152642","density: 1.409485e-04<br />Petal.Width: 0.7199609","density: 7.701642e-05<br />Petal.Width: 0.7246575","density: 4.095370e-05<br />Petal.Width: 0.7293542","density: 2.118486e-05<br />Petal.Width: 0.7340509","density: 1.065718e-05<br />Petal.Width: 0.7387476","density: 5.212275e-06<br />Petal.Width: 0.7434442","density: 2.612618e-06<br />Petal.Width: 0.7481409","density: 1.276424e-06<br />Petal.Width: 0.7528376","density: 6.055498e-07<br />Petal.Width: 0.7575342","density: 2.788709e-07<br />Petal.Width: 0.7622309","density: 1.246409e-07<br />Petal.Width: 0.7669276","density: 5.405821e-08<br />Petal.Width: 0.7716243","density: 2.274944e-08<br />Petal.Width: 0.7763209","density: 9.289056e-09<br />Petal.Width: 0.7810176","density: 3.680119e-09<br />Petal.Width: 0.7857143","density: 1.414652e-09<br />Petal.Width: 0.7904110","density: 5.376985e-10<br />Petal.Width: 0.7951076","density: 2.155147e-10<br />Petal.Width: 0.7998043","density: 8.302819e-11<br />Petal.Width: 0.8045010","density: 3.080191e-11<br />Petal.Width: 0.8091977","density: 1.101918e-11<br />Petal.Width: 0.8138943","density: 3.805637e-12<br />Petal.Width: 0.8185910","density: 1.269888e-12<br />Petal.Width: 0.8232877","density: 4.095881e-13<br />Petal.Width: 0.8279843","density: 1.278301e-13<br />Petal.Width: 0.8326810","density: 3.897752e-14<br />Petal.Width: 0.8373777","density: 1.153361e-14<br />Petal.Width: 0.8420744","density: 3.445976e-15<br />Petal.Width: 0.8467710","density: 1.113737e-15<br />Petal.Width: 0.8514677","density: 6.323343e-16<br />Petal.Width: 0.8561644","density: 3.477739e-16<br />Petal.Width: 0.8608611","density: 3.898402e-16<br />Petal.Width: 0.8655577","density: 2.786069e-16<br />Petal.Width: 0.8702544","density: 1.824401e-16<br />Petal.Width: 0.8749511","density: 3.538269e-16<br />Petal.Width: 0.8796477","density: 2.666047e-16<br />Petal.Width: 0.8843444","density: 0.000000e+00<br />Petal.Width: 0.8890411","density: 0.000000e+00<br />Petal.Width: 0.8937378","density: 0.000000e+00<br />Petal.Width: 0.8984344","density: 0.000000e+00<br />Petal.Width: 0.9031311","density: 0.000000e+00<br />Petal.Width: 0.9078278","density: 0.000000e+00<br />Petal.Width: 0.9125245","density: 0.000000e+00<br />Petal.Width: 0.9172211","density: 0.000000e+00<br />Petal.Width: 0.9219178","density: 0.000000e+00<br />Petal.Width: 0.9266145","density: 0.000000e+00<br />Petal.Width: 0.9313112","density: 0.000000e+00<br />Petal.Width: 0.9360078","density: 0.000000e+00<br />Petal.Width: 0.9407045","density: 1.648818e-21<br />Petal.Width: 0.9454012","density: 1.270498e-19<br />Petal.Width: 0.9500978","density: 2.402872e-20<br />Petal.Width: 0.9547945","density: 0.000000e+00<br />Petal.Width: 0.9594912","density: 0.000000e+00<br />Petal.Width: 0.9641879","density: 0.000000e+00<br />Petal.Width: 0.9688845","density: 0.000000e+00<br />Petal.Width: 0.9735812","density: 0.000000e+00<br />Petal.Width: 0.9782779","density: 0.000000e+00<br />Petal.Width: 0.9829746","density: 0.000000e+00<br />Petal.Width: 0.9876712","density: 0.000000e+00<br />Petal.Width: 0.9923679","density: 0.000000e+00<br />Petal.Width: 0.9970646","density: 0.000000e+00<br />Petal.Width: 1.0017613","density: 0.000000e+00<br />Petal.Width: 1.0064579","density: 1.608065e-17<br />Petal.Width: 1.0111546","density: 8.612415e-18<br />Petal.Width: 1.0158513","density: 0.000000e+00<br />Petal.Width: 1.0205479","density: 0.000000e+00<br />Petal.Width: 1.0252446","density: 0.000000e+00<br />Petal.Width: 1.0299413","density: 0.000000e+00<br />Petal.Width: 1.0346380","density: 0.000000e+00<br />Petal.Width: 1.0393346","density: 0.000000e+00<br />Petal.Width: 1.0440313","density: 0.000000e+00<br />Petal.Width: 1.0487280","density: 3.162430e-17<br />Petal.Width: 1.0534247","density: 7.836348e-18<br />Petal.Width: 1.0581213","density: 0.000000e+00<br />Petal.Width: 1.0628180","density: 0.000000e+00<br />Petal.Width: 1.0675147","density: 6.758470e-17<br />Petal.Width: 1.0722114","density: 7.886404e-17<br />Petal.Width: 1.0769080","density: 0.000000e+00<br />Petal.Width: 1.0816047","density: 0.000000e+00<br />Petal.Width: 1.0863014","density: 2.484272e-17<br />Petal.Width: 1.0909980","density: 1.837772e-16<br />Petal.Width: 1.0956947","density: 1.109503e-16<br />Petal.Width: 1.1003914","density: 1.555341e-17<br />Petal.Width: 1.1050881","density: 0.000000e+00<br />Petal.Width: 1.1097847","density: 1.068825e-17<br />Petal.Width: 1.1144814","density: 4.081215e-17<br />Petal.Width: 1.1191781","density: 7.079048e-17<br />Petal.Width: 1.1238748","density: 5.082494e-17<br />Petal.Width: 1.1285714","density: 2.084746e-17<br />Petal.Width: 1.1332681","density: 5.586538e-17<br />Petal.Width: 1.1379648","density: 0.000000e+00<br />Petal.Width: 1.1426614","density: 0.000000e+00<br />Petal.Width: 1.1473581","density: 0.000000e+00<br />Petal.Width: 1.1520548","density: 1.140434e-16<br />Petal.Width: 1.1567515","density: 3.564440e-17<br />Petal.Width: 1.1614481","density: 0.000000e+00<br />Petal.Width: 1.1661448","density: 0.000000e+00<br />Petal.Width: 1.1708415","density: 1.966222e-16<br />Petal.Width: 1.1755382","density: 2.939381e-16<br />Petal.Width: 1.1802348","density: 4.835963e-17<br />Petal.Width: 1.1849315","density: 0.000000e+00<br />Petal.Width: 1.1896282","density: 2.503350e-18<br />Petal.Width: 1.1943249","density: 2.783438e-17<br />Petal.Width: 1.1990215","density: 5.809505e-17<br />Petal.Width: 1.2037182","density: 1.537560e-16<br />Petal.Width: 1.2084149","density: 8.459186e-17<br />Petal.Width: 1.2131115","density: 1.835878e-17<br />Petal.Width: 1.2178082","density: 0.000000e+00<br />Petal.Width: 1.2225049","density: 0.000000e+00<br />Petal.Width: 1.2272016","density: 0.000000e+00<br />Petal.Width: 1.2318982","density: 0.000000e+00<br />Petal.Width: 1.2365949","density: 5.031357e-19<br />Petal.Width: 1.2412916","density: 7.447476e-18<br />Petal.Width: 1.2459883","density: 6.505137e-17<br />Petal.Width: 1.2506849","density: 1.008551e-16<br />Petal.Width: 1.2553816","density: 1.122683e-16<br />Petal.Width: 1.2600783","density: 7.235806e-17<br />Petal.Width: 1.2647750","density: 2.147105e-17<br />Petal.Width: 1.2694716","density: 8.170694e-17<br />Petal.Width: 1.2741683","density: 9.310681e-17<br />Petal.Width: 1.2788650","density: 0.000000e+00<br />Petal.Width: 1.2835616","density: 0.000000e+00<br />Petal.Width: 1.2882583","density: 1.157014e-17<br />Petal.Width: 1.2929550","density: 7.919656e-17<br />Petal.Width: 1.2976517","density: 3.856712e-18<br />Petal.Width: 1.3023483","density: 0.000000e+00<br />Petal.Width: 1.3070450","density: 0.000000e+00<br />Petal.Width: 1.3117417","density: 0.000000e+00<br />Petal.Width: 1.3164384","density: 0.000000e+00<br />Petal.Width: 1.3211350","density: 0.000000e+00<br />Petal.Width: 1.3258317","density: 0.000000e+00<br />Petal.Width: 1.3305284","density: 0.000000e+00<br />Petal.Width: 1.3352250","density: 0.000000e+00<br />Petal.Width: 1.3399217","density: 0.000000e+00<br />Petal.Width: 1.3446184","density: 0.000000e+00<br />Petal.Width: 1.3493151","density: 0.000000e+00<br />Petal.Width: 1.3540117","density: 0.000000e+00<br />Petal.Width: 1.3587084","density: 0.000000e+00<br />Petal.Width: 1.3634051","density: 0.000000e+00<br />Petal.Width: 1.3681018","density: 0.000000e+00<br />Petal.Width: 1.3727984","density: 0.000000e+00<br />Petal.Width: 1.3774951","density: 0.000000e+00<br />Petal.Width: 1.3821918","density: 0.000000e+00<br />Petal.Width: 1.3868885","density: 0.000000e+00<br />Petal.Width: 1.3915851","density: 0.000000e+00<br />Petal.Width: 1.3962818","density: 0.000000e+00<br />Petal.Width: 1.4009785","density: 0.000000e+00<br />Petal.Width: 1.4056751","density: 0.000000e+00<br />Petal.Width: 1.4103718","density: 8.044747e-17<br />Petal.Width: 1.4150685","density: 4.088583e-17<br />Petal.Width: 1.4197652","density: 0.000000e+00<br />Petal.Width: 1.4244618","density: 0.000000e+00<br />Petal.Width: 1.4291585","density: 0.000000e+00<br />Petal.Width: 1.4338552","density: 0.000000e+00<br />Petal.Width: 1.4385519","density: 0.000000e+00<br />Petal.Width: 1.4432485","density: 0.000000e+00<br />Petal.Width: 1.4479452","density: 0.000000e+00<br />Petal.Width: 1.4526419","density: 7.862829e-17<br />Petal.Width: 1.4573386","density: 1.395539e-16<br />Petal.Width: 1.4620352","density: 4.533856e-17<br />Petal.Width: 1.4667319","density: 0.000000e+00<br />Petal.Width: 1.4714286","density: 0.000000e+00<br />Petal.Width: 1.4761252","density: 0.000000e+00<br />Petal.Width: 1.4808219","density: 0.000000e+00<br />Petal.Width: 1.4855186","density: 3.808486e-17<br />Petal.Width: 1.4902153","density: 1.808345e-16<br />Petal.Width: 1.4949119","density: 2.891638e-16<br />Petal.Width: 1.4996086","density: 1.220774e-17<br />Petal.Width: 1.5043053","density: 4.572926e-16<br />Petal.Width: 1.5090020","density: 1.190406e-16<br />Petal.Width: 1.5136986","density: 1.134752e-16<br />Petal.Width: 1.5183953","density: 6.852475e-17<br />Petal.Width: 1.5230920","density: 6.878517e-17<br />Petal.Width: 1.5277886","density: 8.288145e-17<br />Petal.Width: 1.5324853","density: 0.000000e+00<br />Petal.Width: 1.5371820","density: 0.000000e+00<br />Petal.Width: 1.5418787","density: 9.476601e-18<br />Petal.Width: 1.5465753","density: 8.056073e-17<br />Petal.Width: 1.5512720","density: 2.033026e-16<br />Petal.Width: 1.5559687","density: 3.268999e-17<br />Petal.Width: 1.5606654","density: 2.480761e-17<br />Petal.Width: 1.5653620","density: 5.356172e-17<br />Petal.Width: 1.5700587","density: 2.779731e-17<br />Petal.Width: 1.5747554","density: 0.000000e+00<br />Petal.Width: 1.5794521","density: 7.604100e-17<br />Petal.Width: 1.5841487","density: 1.519821e-16<br />Petal.Width: 1.5888454","density: 5.944201e-17<br />Petal.Width: 1.5935421","density: 1.447848e-16<br />Petal.Width: 1.5982387","density: 3.559214e-18<br />Petal.Width: 1.6029354","density: 3.371654e-16<br />Petal.Width: 1.6076321","density: 9.762490e-17<br />Petal.Width: 1.6123288","density: 1.073040e-17<br />Petal.Width: 1.6170254","density: 1.439197e-17<br />Petal.Width: 1.6217221","density: 5.376827e-17<br />Petal.Width: 1.6264188","density: 4.369389e-17<br />Petal.Width: 1.6311155","density: 9.979950e-18<br />Petal.Width: 1.6358121","density: 2.035337e-17<br />Petal.Width: 1.6405088","density: 1.447335e-17<br />Petal.Width: 1.6452055","density: 7.652662e-17<br />Petal.Width: 1.6499022","density: 9.931422e-19<br />Petal.Width: 1.6545988","density: 3.727017e-17<br />Petal.Width: 1.6592955","density: 8.229819e-18<br />Petal.Width: 1.6639922","density: 0.000000e+00<br />Petal.Width: 1.6686888","density: 0.000000e+00<br />Petal.Width: 1.6733855","density: 8.721469e-17<br />Petal.Width: 1.6780822","density: 9.478526e-17<br />Petal.Width: 1.6827789","density: 0.000000e+00<br />Petal.Width: 1.6874755","density: 4.081520e-17<br />Petal.Width: 1.6921722","density: 1.538591e-16<br />Petal.Width: 1.6968689","density: 7.832562e-17<br />Petal.Width: 1.7015656","density: 2.792144e-18<br />Petal.Width: 1.7062622","density: 0.000000e+00<br />Petal.Width: 1.7109589","density: 0.000000e+00<br />Petal.Width: 1.7156556","density: 7.636645e-17<br />Petal.Width: 1.7203523","density: 4.496685e-17<br />Petal.Width: 1.7250489","density: 0.000000e+00<br />Petal.Width: 1.7297456","density: 0.000000e+00<br />Petal.Width: 1.7344423","density: 8.780615e-18<br />Petal.Width: 1.7391389","density: 2.155271e-17<br />Petal.Width: 1.7438356","density: 3.625117e-18<br />Petal.Width: 1.7485323","density: 2.670821e-17<br />Petal.Width: 1.7532290","density: 1.530382e-18<br />Petal.Width: 1.7579256","density: 3.547117e-17<br />Petal.Width: 1.7626223","density: 3.109852e-17<br />Petal.Width: 1.7673190","density: 9.263630e-18<br />Petal.Width: 1.7720157","density: 0.000000e+00<br />Petal.Width: 1.7767123","density: 1.000234e-16<br />Petal.Width: 1.7814090","density: 1.449595e-16<br />Petal.Width: 1.7861057","density: 5.123852e-17<br />Petal.Width: 1.7908023","density: 7.992749e-17<br />Petal.Width: 1.7954990","density: 1.209375e-16<br />Petal.Width: 1.8001957","density: 3.413484e-18<br />Petal.Width: 1.8048924","density: 2.662562e-19<br />Petal.Width: 1.8095890","density: 0.000000e+00<br />Petal.Width: 1.8142857","density: 0.000000e+00<br />Petal.Width: 1.8189824","density: 0.000000e+00<br />Petal.Width: 1.8236791","density: 0.000000e+00<br />Petal.Width: 1.8283757","density: 0.000000e+00<br />Petal.Width: 1.8330724","density: 7.619273e-18<br />Petal.Width: 1.8377691","density: 1.513072e-17<br />Petal.Width: 1.8424658","density: 0.000000e+00<br />Petal.Width: 1.8471624","density: 0.000000e+00<br />Petal.Width: 1.8518591","density: 0.000000e+00<br />Petal.Width: 1.8565558","density: 0.000000e+00<br />Petal.Width: 1.8612524","density: 0.000000e+00<br />Petal.Width: 1.8659491","density: 0.000000e+00<br />Petal.Width: 1.8706458","density: 0.000000e+00<br />Petal.Width: 1.8753425","density: 0.000000e+00<br />Petal.Width: 1.8800391","density: 2.579136e-17<br />Petal.Width: 1.8847358","density: 3.866695e-17<br />Petal.Width: 1.8894325","density: 0.000000e+00<br />Petal.Width: 1.8941292","density: 0.000000e+00<br />Petal.Width: 1.8988258","density: 0.000000e+00<br />Petal.Width: 1.9035225","density: 0.000000e+00<br />Petal.Width: 1.9082192","density: 0.000000e+00<br />Petal.Width: 1.9129159","density: 0.000000e+00<br />Petal.Width: 1.9176125","density: 0.000000e+00<br />Petal.Width: 1.9223092","density: 0.000000e+00<br />Petal.Width: 1.9270059","density: 0.000000e+00<br />Petal.Width: 1.9317025","density: 0.000000e+00<br />Petal.Width: 1.9363992","density: 0.000000e+00<br />Petal.Width: 1.9410959","density: 0.000000e+00<br />Petal.Width: 1.9457926","density: 0.000000e+00<br />Petal.Width: 1.9504892","density: 0.000000e+00<br />Petal.Width: 1.9551859","density: 0.000000e+00<br />Petal.Width: 1.9598826","density: 3.276147e-17<br />Petal.Width: 1.9645793","density: 8.946851e-18<br />Petal.Width: 1.9692759","density: 0.000000e+00<br />Petal.Width: 1.9739726","density: 9.052468e-18<br />Petal.Width: 1.9786693","density: 8.010028e-18<br />Petal.Width: 1.9833659","density: 0.000000e+00<br />Petal.Width: 1.9880626","density: 0.000000e+00<br />Petal.Width: 1.9927593","density: 0.000000e+00<br />Petal.Width: 1.9974560","density: 0.000000e+00<br />Petal.Width: 2.0021526","density: 1.958687e-20<br />Petal.Width: 2.0068493","density: 8.063939e-19<br />Petal.Width: 2.0115460","density: 2.327765e-17<br />Petal.Width: 2.0162427","density: 7.078709e-18<br />Petal.Width: 2.0209393","density: 0.000000e+00<br />Petal.Width: 2.0256360","density: 2.198458e-17<br />Petal.Width: 2.0303327","density: 1.367193e-16<br />Petal.Width: 2.0350294","density: 1.761828e-16<br />Petal.Width: 2.0397260","density: 8.255361e-17<br />Petal.Width: 2.0444227","density: 1.728765e-16<br />Petal.Width: 2.0491194","density: 2.321295e-16<br />Petal.Width: 2.0538160","density: 7.241997e-17<br />Petal.Width: 2.0585127","density: 2.741050e-16<br />Petal.Width: 2.0632094","density: 1.551066e-16<br />Petal.Width: 2.0679061","density: 1.530756e-16<br />Petal.Width: 2.0726027","density: 7.090563e-17<br />Petal.Width: 2.0772994","density: 9.645054e-17<br />Petal.Width: 2.0819961","density: 1.302513e-16<br />Petal.Width: 2.0866928","density: 1.195065e-16<br />Petal.Width: 2.0913894","density: 1.644141e-16<br />Petal.Width: 2.0960861","density: 1.477679e-16<br />Petal.Width: 2.1007828","density: 3.186040e-16<br />Petal.Width: 2.1054795","density: 1.887928e-16<br />Petal.Width: 2.1101761","density: 2.446250e-16<br />Petal.Width: 2.1148728","density: 2.908092e-16<br />Petal.Width: 2.1195695","density: 2.006887e-16<br />Petal.Width: 2.1242661","density: 6.192023e-17<br />Petal.Width: 2.1289628","density: 2.787022e-17<br />Petal.Width: 2.1336595","density: 4.195302e-17<br />Petal.Width: 2.1383562","density: 5.897488e-17<br />Petal.Width: 2.1430528","density: 1.252503e-16<br />Petal.Width: 2.1477495","density: 1.008573e-16<br />Petal.Width: 2.1524462","density: 0.000000e+00<br />Petal.Width: 2.1571429","density: 0.000000e+00<br />Petal.Width: 2.1618395","density: 2.388497e-16<br />Petal.Width: 2.1665362","density: 2.155538e-16<br />Petal.Width: 2.1712329","density: 6.341871e-17<br />Petal.Width: 2.1759295","density: 0.000000e+00<br />Petal.Width: 2.1806262","density: 0.000000e+00<br />Petal.Width: 2.1853229","density: 0.000000e+00<br />Petal.Width: 2.1900196","density: 0.000000e+00<br />Petal.Width: 2.1947162","density: 0.000000e+00<br />Petal.Width: 2.1994129","density: 0.000000e+00<br />Petal.Width: 2.2041096","density: 0.000000e+00<br />Petal.Width: 2.2088063","density: 0.000000e+00<br />Petal.Width: 2.2135029","density: 1.105396e-17<br />Petal.Width: 2.2181996","density: 3.271610e-18<br />Petal.Width: 2.2228963","density: 0.000000e+00<br />Petal.Width: 2.2275930","density: 0.000000e+00<br />Petal.Width: 2.2322896","density: 0.000000e+00<br />Petal.Width: 2.2369863","density: 0.000000e+00<br />Petal.Width: 2.2416830","density: 0.000000e+00<br />Petal.Width: 2.2463796","density: 0.000000e+00<br />Petal.Width: 2.2510763","density: 0.000000e+00<br />Petal.Width: 2.2557730","density: 0.000000e+00<br />Petal.Width: 2.2604697","density: 0.000000e+00<br />Petal.Width: 2.2651663","density: 0.000000e+00<br />Petal.Width: 2.2698630","density: 0.000000e+00<br />Petal.Width: 2.2745597","density: 0.000000e+00<br />Petal.Width: 2.2792564","density: 0.000000e+00<br />Petal.Width: 2.2839530","density: 0.000000e+00<br />Petal.Width: 2.2886497","density: 0.000000e+00<br />Petal.Width: 2.2933464","density: 0.000000e+00<br />Petal.Width: 2.2980431","density: 0.000000e+00<br />Petal.Width: 2.3027397","density: 0.000000e+00<br />Petal.Width: 2.3074364","density: 0.000000e+00<br />Petal.Width: 2.3121331","density: 0.000000e+00<br />Petal.Width: 2.3168297","density: 0.000000e+00<br />Petal.Width: 2.3215264","density: 1.300303e-17<br />Petal.Width: 2.3262231","density: 7.091123e-18<br />Petal.Width: 2.3309198","density: 0.000000e+00<br />Petal.Width: 2.3356164","density: 0.000000e+00<br />Petal.Width: 2.3403131","density: 4.405739e-17<br />Petal.Width: 2.3450098","density: 9.936693e-17<br />Petal.Width: 2.3497065","density: 0.000000e+00<br />Petal.Width: 2.3544031","density: 0.000000e+00<br />Petal.Width: 2.3590998","density: 0.000000e+00<br />Petal.Width: 2.3637965","density: 0.000000e+00<br />Petal.Width: 2.3684932","density: 0.000000e+00<br />Petal.Width: 2.3731898","density: 0.000000e+00<br />Petal.Width: 2.3778865","density: 0.000000e+00<br />Petal.Width: 2.3825832","density: 0.000000e+00<br />Petal.Width: 2.3872798","density: 0.000000e+00<br />Petal.Width: 2.3919765","density: 0.000000e+00<br />Petal.Width: 2.3966732","density: 0.000000e+00<br />Petal.Width: 2.4013699","density: 0.000000e+00<br />Petal.Width: 2.4060665","density: 0.000000e+00<br />Petal.Width: 2.4107632","density: 0.000000e+00<br />Petal.Width: 2.4154599","density: 0.000000e+00<br />Petal.Width: 2.4201566","density: 7.880983e-17<br />Petal.Width: 2.4248532","density: 3.498718e-17<br />Petal.Width: 2.4295499","density: 0.000000e+00<br />Petal.Width: 2.4342466","density: 0.000000e+00<br />Petal.Width: 2.4389432","density: 1.026296e-17<br />Petal.Width: 2.4436399","density: 1.884155e-17<br />Petal.Width: 2.4483366","density: 0.000000e+00<br />Petal.Width: 2.4530333","density: 0.000000e+00<br />Petal.Width: 2.4577299","density: 0.000000e+00<br />Petal.Width: 2.4624266","density: 0.000000e+00<br />Petal.Width: 2.4671233","density: 0.000000e+00<br />Petal.Width: 2.4718200","density: 0.000000e+00<br />Petal.Width: 2.4765166","density: 0.000000e+00<br />Petal.Width: 2.4812133","density: 0.000000e+00<br />Petal.Width: 2.4859100","density: 0.000000e+00<br />Petal.Width: 2.4906067","density: 0.000000e+00<br />Petal.Width: 2.4953033","density: 0.000000e+00<br />Petal.Width: 2.5000000","density: 0.000000e+00<br />Petal.Width: 2.5000000","density: 0.000000e+00<br />Petal.Width: 2.4953033","density: 0.000000e+00<br />Petal.Width: 2.4906067","density: 0.000000e+00<br />Petal.Width: 2.4859100","density: 0.000000e+00<br />Petal.Width: 2.4812133","density: 0.000000e+00<br />Petal.Width: 2.4765166","density: 0.000000e+00<br />Petal.Width: 2.4718200","density: 0.000000e+00<br />Petal.Width: 2.4671233","density: 0.000000e+00<br />Petal.Width: 2.4624266","density: 0.000000e+00<br />Petal.Width: 2.4577299","density: 0.000000e+00<br />Petal.Width: 2.4530333","density: 1.884155e-17<br />Petal.Width: 2.4483366","density: 1.026296e-17<br />Petal.Width: 2.4436399","density: 0.000000e+00<br />Petal.Width: 2.4389432","density: 0.000000e+00<br />Petal.Width: 2.4342466","density: 3.498718e-17<br />Petal.Width: 2.4295499","density: 7.880983e-17<br />Petal.Width: 2.4248532","density: 0.000000e+00<br />Petal.Width: 2.4201566","density: 0.000000e+00<br />Petal.Width: 2.4154599","density: 0.000000e+00<br />Petal.Width: 2.4107632","density: 0.000000e+00<br />Petal.Width: 2.4060665","density: 0.000000e+00<br />Petal.Width: 2.4013699","density: 0.000000e+00<br />Petal.Width: 2.3966732","density: 0.000000e+00<br />Petal.Width: 2.3919765","density: 0.000000e+00<br />Petal.Width: 2.3872798","density: 0.000000e+00<br />Petal.Width: 2.3825832","density: 0.000000e+00<br />Petal.Width: 2.3778865","density: 0.000000e+00<br />Petal.Width: 2.3731898","density: 0.000000e+00<br />Petal.Width: 2.3684932","density: 0.000000e+00<br />Petal.Width: 2.3637965","density: 0.000000e+00<br />Petal.Width: 2.3590998","density: 0.000000e+00<br />Petal.Width: 2.3544031","density: 9.936693e-17<br />Petal.Width: 2.3497065","density: 4.405739e-17<br />Petal.Width: 2.3450098","density: 0.000000e+00<br />Petal.Width: 2.3403131","density: 0.000000e+00<br />Petal.Width: 2.3356164","density: 7.091123e-18<br />Petal.Width: 2.3309198","density: 1.300303e-17<br />Petal.Width: 2.3262231","density: 0.000000e+00<br />Petal.Width: 2.3215264","density: 0.000000e+00<br />Petal.Width: 2.3168297","density: 0.000000e+00<br />Petal.Width: 2.3121331","density: 0.000000e+00<br />Petal.Width: 2.3074364","density: 0.000000e+00<br />Petal.Width: 2.3027397","density: 0.000000e+00<br />Petal.Width: 2.2980431","density: 0.000000e+00<br />Petal.Width: 2.2933464","density: 0.000000e+00<br />Petal.Width: 2.2886497","density: 0.000000e+00<br />Petal.Width: 2.2839530","density: 0.000000e+00<br />Petal.Width: 2.2792564","density: 0.000000e+00<br />Petal.Width: 2.2745597","density: 0.000000e+00<br />Petal.Width: 2.2698630","density: 0.000000e+00<br />Petal.Width: 2.2651663","density: 0.000000e+00<br />Petal.Width: 2.2604697","density: 0.000000e+00<br />Petal.Width: 2.2557730","density: 0.000000e+00<br />Petal.Width: 2.2510763","density: 0.000000e+00<br />Petal.Width: 2.2463796","density: 0.000000e+00<br />Petal.Width: 2.2416830","density: 0.000000e+00<br />Petal.Width: 2.2369863","density: 0.000000e+00<br />Petal.Width: 2.2322896","density: 0.000000e+00<br />Petal.Width: 2.2275930","density: 3.271610e-18<br />Petal.Width: 2.2228963","density: 1.105396e-17<br />Petal.Width: 2.2181996","density: 0.000000e+00<br />Petal.Width: 2.2135029","density: 0.000000e+00<br />Petal.Width: 2.2088063","density: 0.000000e+00<br />Petal.Width: 2.2041096","density: 0.000000e+00<br />Petal.Width: 2.1994129","density: 0.000000e+00<br />Petal.Width: 2.1947162","density: 0.000000e+00<br />Petal.Width: 2.1900196","density: 0.000000e+00<br />Petal.Width: 2.1853229","density: 0.000000e+00<br />Petal.Width: 2.1806262","density: 6.341871e-17<br />Petal.Width: 2.1759295","density: 2.155538e-16<br />Petal.Width: 2.1712329","density: 2.388497e-16<br />Petal.Width: 2.1665362","density: 0.000000e+00<br />Petal.Width: 2.1618395","density: 0.000000e+00<br />Petal.Width: 2.1571429","density: 1.008573e-16<br />Petal.Width: 2.1524462","density: 1.252503e-16<br />Petal.Width: 2.1477495","density: 5.897488e-17<br />Petal.Width: 2.1430528","density: 4.195302e-17<br />Petal.Width: 2.1383562","density: 2.787022e-17<br />Petal.Width: 2.1336595","density: 6.192023e-17<br />Petal.Width: 2.1289628","density: 2.006887e-16<br />Petal.Width: 2.1242661","density: 2.908092e-16<br />Petal.Width: 2.1195695","density: 2.446250e-16<br />Petal.Width: 2.1148728","density: 1.887928e-16<br />Petal.Width: 2.1101761","density: 3.186040e-16<br />Petal.Width: 2.1054795","density: 1.477679e-16<br />Petal.Width: 2.1007828","density: 1.644141e-16<br />Petal.Width: 2.0960861","density: 1.195065e-16<br />Petal.Width: 2.0913894","density: 1.302513e-16<br />Petal.Width: 2.0866928","density: 9.645054e-17<br />Petal.Width: 2.0819961","density: 7.090563e-17<br />Petal.Width: 2.0772994","density: 1.530756e-16<br />Petal.Width: 2.0726027","density: 1.551066e-16<br />Petal.Width: 2.0679061","density: 2.741050e-16<br />Petal.Width: 2.0632094","density: 7.241997e-17<br />Petal.Width: 2.0585127","density: 2.321295e-16<br />Petal.Width: 2.0538160","density: 1.728765e-16<br />Petal.Width: 2.0491194","density: 8.255361e-17<br />Petal.Width: 2.0444227","density: 1.761828e-16<br />Petal.Width: 2.0397260","density: 1.367193e-16<br />Petal.Width: 2.0350294","density: 2.198458e-17<br />Petal.Width: 2.0303327","density: 0.000000e+00<br />Petal.Width: 2.0256360","density: 7.078709e-18<br />Petal.Width: 2.0209393","density: 2.327765e-17<br />Petal.Width: 2.0162427","density: 8.063939e-19<br />Petal.Width: 2.0115460","density: 1.958687e-20<br />Petal.Width: 2.0068493","density: 0.000000e+00<br />Petal.Width: 2.0021526","density: 0.000000e+00<br />Petal.Width: 1.9974560","density: 0.000000e+00<br />Petal.Width: 1.9927593","density: 0.000000e+00<br />Petal.Width: 1.9880626","density: 8.010028e-18<br />Petal.Width: 1.9833659","density: 9.052468e-18<br />Petal.Width: 1.9786693","density: 0.000000e+00<br />Petal.Width: 1.9739726","density: 8.946851e-18<br />Petal.Width: 1.9692759","density: 3.276147e-17<br />Petal.Width: 1.9645793","density: 0.000000e+00<br />Petal.Width: 1.9598826","density: 0.000000e+00<br />Petal.Width: 1.9551859","density: 0.000000e+00<br />Petal.Width: 1.9504892","density: 0.000000e+00<br />Petal.Width: 1.9457926","density: 0.000000e+00<br />Petal.Width: 1.9410959","density: 0.000000e+00<br />Petal.Width: 1.9363992","density: 0.000000e+00<br />Petal.Width: 1.9317025","density: 0.000000e+00<br />Petal.Width: 1.9270059","density: 0.000000e+00<br />Petal.Width: 1.9223092","density: 0.000000e+00<br />Petal.Width: 1.9176125","density: 0.000000e+00<br />Petal.Width: 1.9129159","density: 0.000000e+00<br />Petal.Width: 1.9082192","density: 0.000000e+00<br />Petal.Width: 1.9035225","density: 0.000000e+00<br />Petal.Width: 1.8988258","density: 0.000000e+00<br />Petal.Width: 1.8941292","density: 3.866695e-17<br />Petal.Width: 1.8894325","density: 2.579136e-17<br />Petal.Width: 1.8847358","density: 0.000000e+00<br />Petal.Width: 1.8800391","density: 0.000000e+00<br />Petal.Width: 1.8753425","density: 0.000000e+00<br />Petal.Width: 1.8706458","density: 0.000000e+00<br />Petal.Width: 1.8659491","density: 0.000000e+00<br />Petal.Width: 1.8612524","density: 0.000000e+00<br />Petal.Width: 1.8565558","density: 0.000000e+00<br />Petal.Width: 1.8518591","density: 0.000000e+00<br />Petal.Width: 1.8471624","density: 1.513072e-17<br />Petal.Width: 1.8424658","density: 7.619273e-18<br />Petal.Width: 1.8377691","density: 0.000000e+00<br />Petal.Width: 1.8330724","density: 0.000000e+00<br />Petal.Width: 1.8283757","density: 0.000000e+00<br />Petal.Width: 1.8236791","density: 0.000000e+00<br />Petal.Width: 1.8189824","density: 0.000000e+00<br />Petal.Width: 1.8142857","density: 2.662562e-19<br />Petal.Width: 1.8095890","density: 3.413484e-18<br />Petal.Width: 1.8048924","density: 1.209375e-16<br />Petal.Width: 1.8001957","density: 7.992749e-17<br />Petal.Width: 1.7954990","density: 5.123852e-17<br />Petal.Width: 1.7908023","density: 1.449595e-16<br />Petal.Width: 1.7861057","density: 1.000234e-16<br />Petal.Width: 1.7814090","density: 0.000000e+00<br />Petal.Width: 1.7767123","density: 9.263630e-18<br />Petal.Width: 1.7720157","density: 3.109852e-17<br />Petal.Width: 1.7673190","density: 3.547117e-17<br />Petal.Width: 1.7626223","density: 1.530382e-18<br />Petal.Width: 1.7579256","density: 2.670821e-17<br />Petal.Width: 1.7532290","density: 3.625117e-18<br />Petal.Width: 1.7485323","density: 2.155271e-17<br />Petal.Width: 1.7438356","density: 8.780615e-18<br />Petal.Width: 1.7391389","density: 0.000000e+00<br />Petal.Width: 1.7344423","density: 0.000000e+00<br />Petal.Width: 1.7297456","density: 4.496685e-17<br />Petal.Width: 1.7250489","density: 7.636645e-17<br />Petal.Width: 1.7203523","density: 0.000000e+00<br />Petal.Width: 1.7156556","density: 0.000000e+00<br />Petal.Width: 1.7109589","density: 2.792144e-18<br />Petal.Width: 1.7062622","density: 7.832562e-17<br />Petal.Width: 1.7015656","density: 1.538591e-16<br />Petal.Width: 1.6968689","density: 4.081520e-17<br />Petal.Width: 1.6921722","density: 0.000000e+00<br />Petal.Width: 1.6874755","density: 9.478526e-17<br />Petal.Width: 1.6827789","density: 8.721469e-17<br />Petal.Width: 1.6780822","density: 0.000000e+00<br />Petal.Width: 1.6733855","density: 0.000000e+00<br />Petal.Width: 1.6686888","density: 8.229819e-18<br />Petal.Width: 1.6639922","density: 3.727017e-17<br />Petal.Width: 1.6592955","density: 9.931422e-19<br />Petal.Width: 1.6545988","density: 7.652662e-17<br />Petal.Width: 1.6499022","density: 1.447335e-17<br />Petal.Width: 1.6452055","density: 2.035337e-17<br />Petal.Width: 1.6405088","density: 9.979950e-18<br />Petal.Width: 1.6358121","density: 4.369389e-17<br />Petal.Width: 1.6311155","density: 5.376827e-17<br />Petal.Width: 1.6264188","density: 1.439197e-17<br />Petal.Width: 1.6217221","density: 1.073040e-17<br />Petal.Width: 1.6170254","density: 9.762490e-17<br />Petal.Width: 1.6123288","density: 3.371654e-16<br />Petal.Width: 1.6076321","density: 3.559214e-18<br />Petal.Width: 1.6029354","density: 1.447848e-16<br />Petal.Width: 1.5982387","density: 5.944201e-17<br />Petal.Width: 1.5935421","density: 1.519821e-16<br />Petal.Width: 1.5888454","density: 7.604100e-17<br />Petal.Width: 1.5841487","density: 0.000000e+00<br />Petal.Width: 1.5794521","density: 2.779731e-17<br />Petal.Width: 1.5747554","density: 5.356172e-17<br />Petal.Width: 1.5700587","density: 2.480761e-17<br />Petal.Width: 1.5653620","density: 3.268999e-17<br />Petal.Width: 1.5606654","density: 2.033026e-16<br />Petal.Width: 1.5559687","density: 8.056073e-17<br />Petal.Width: 1.5512720","density: 9.476601e-18<br />Petal.Width: 1.5465753","density: 0.000000e+00<br />Petal.Width: 1.5418787","density: 0.000000e+00<br />Petal.Width: 1.5371820","density: 8.288145e-17<br />Petal.Width: 1.5324853","density: 6.878517e-17<br />Petal.Width: 1.5277886","density: 6.852475e-17<br />Petal.Width: 1.5230920","density: 1.134752e-16<br />Petal.Width: 1.5183953","density: 1.190406e-16<br />Petal.Width: 1.5136986","density: 4.572926e-16<br />Petal.Width: 1.5090020","density: 1.220774e-17<br />Petal.Width: 1.5043053","density: 2.891638e-16<br />Petal.Width: 1.4996086","density: 1.808345e-16<br />Petal.Width: 1.4949119","density: 3.808486e-17<br />Petal.Width: 1.4902153","density: 0.000000e+00<br />Petal.Width: 1.4855186","density: 0.000000e+00<br />Petal.Width: 1.4808219","density: 0.000000e+00<br />Petal.Width: 1.4761252","density: 0.000000e+00<br />Petal.Width: 1.4714286","density: 4.533856e-17<br />Petal.Width: 1.4667319","density: 1.395539e-16<br />Petal.Width: 1.4620352","density: 7.862829e-17<br />Petal.Width: 1.4573386","density: 0.000000e+00<br />Petal.Width: 1.4526419","density: 0.000000e+00<br />Petal.Width: 1.4479452","density: 0.000000e+00<br />Petal.Width: 1.4432485","density: 0.000000e+00<br />Petal.Width: 1.4385519","density: 0.000000e+00<br />Petal.Width: 1.4338552","density: 0.000000e+00<br />Petal.Width: 1.4291585","density: 0.000000e+00<br />Petal.Width: 1.4244618","density: 4.088583e-17<br />Petal.Width: 1.4197652","density: 8.044747e-17<br />Petal.Width: 1.4150685","density: 0.000000e+00<br />Petal.Width: 1.4103718","density: 0.000000e+00<br />Petal.Width: 1.4056751","density: 0.000000e+00<br />Petal.Width: 1.4009785","density: 0.000000e+00<br />Petal.Width: 1.3962818","density: 0.000000e+00<br />Petal.Width: 1.3915851","density: 0.000000e+00<br />Petal.Width: 1.3868885","density: 0.000000e+00<br />Petal.Width: 1.3821918","density: 0.000000e+00<br />Petal.Width: 1.3774951","density: 0.000000e+00<br />Petal.Width: 1.3727984","density: 0.000000e+00<br />Petal.Width: 1.3681018","density: 0.000000e+00<br />Petal.Width: 1.3634051","density: 0.000000e+00<br />Petal.Width: 1.3587084","density: 0.000000e+00<br />Petal.Width: 1.3540117","density: 0.000000e+00<br />Petal.Width: 1.3493151","density: 0.000000e+00<br />Petal.Width: 1.3446184","density: 0.000000e+00<br />Petal.Width: 1.3399217","density: 0.000000e+00<br />Petal.Width: 1.3352250","density: 0.000000e+00<br />Petal.Width: 1.3305284","density: 0.000000e+00<br />Petal.Width: 1.3258317","density: 0.000000e+00<br />Petal.Width: 1.3211350","density: 0.000000e+00<br />Petal.Width: 1.3164384","density: 0.000000e+00<br />Petal.Width: 1.3117417","density: 0.000000e+00<br />Petal.Width: 1.3070450","density: 3.856712e-18<br />Petal.Width: 1.3023483","density: 7.919656e-17<br />Petal.Width: 1.2976517","density: 1.157014e-17<br />Petal.Width: 1.2929550","density: 0.000000e+00<br />Petal.Width: 1.2882583","density: 0.000000e+00<br />Petal.Width: 1.2835616","density: 9.310681e-17<br />Petal.Width: 1.2788650","density: 8.170694e-17<br />Petal.Width: 1.2741683","density: 2.147105e-17<br />Petal.Width: 1.2694716","density: 7.235806e-17<br />Petal.Width: 1.2647750","density: 1.122683e-16<br />Petal.Width: 1.2600783","density: 1.008551e-16<br />Petal.Width: 1.2553816","density: 6.505137e-17<br />Petal.Width: 1.2506849","density: 7.447476e-18<br />Petal.Width: 1.2459883","density: 5.031357e-19<br />Petal.Width: 1.2412916","density: 0.000000e+00<br />Petal.Width: 1.2365949","density: 0.000000e+00<br />Petal.Width: 1.2318982","density: 0.000000e+00<br />Petal.Width: 1.2272016","density: 0.000000e+00<br />Petal.Width: 1.2225049","density: 1.835878e-17<br />Petal.Width: 1.2178082","density: 8.459186e-17<br />Petal.Width: 1.2131115","density: 1.537560e-16<br />Petal.Width: 1.2084149","density: 5.809505e-17<br />Petal.Width: 1.2037182","density: 2.783438e-17<br />Petal.Width: 1.1990215","density: 2.503350e-18<br />Petal.Width: 1.1943249","density: 0.000000e+00<br />Petal.Width: 1.1896282","density: 4.835963e-17<br />Petal.Width: 1.1849315","density: 2.939381e-16<br />Petal.Width: 1.1802348","density: 1.966222e-16<br />Petal.Width: 1.1755382","density: 0.000000e+00<br />Petal.Width: 1.1708415","density: 0.000000e+00<br />Petal.Width: 1.1661448","density: 3.564440e-17<br />Petal.Width: 1.1614481","density: 1.140434e-16<br />Petal.Width: 1.1567515","density: 0.000000e+00<br />Petal.Width: 1.1520548","density: 0.000000e+00<br />Petal.Width: 1.1473581","density: 0.000000e+00<br />Petal.Width: 1.1426614","density: 5.586538e-17<br />Petal.Width: 1.1379648","density: 2.084746e-17<br />Petal.Width: 1.1332681","density: 5.082494e-17<br />Petal.Width: 1.1285714","density: 7.079048e-17<br />Petal.Width: 1.1238748","density: 4.081215e-17<br />Petal.Width: 1.1191781","density: 1.068825e-17<br />Petal.Width: 1.1144814","density: 0.000000e+00<br />Petal.Width: 1.1097847","density: 1.555341e-17<br />Petal.Width: 1.1050881","density: 1.109503e-16<br />Petal.Width: 1.1003914","density: 1.837772e-16<br />Petal.Width: 1.0956947","density: 2.484272e-17<br />Petal.Width: 1.0909980","density: 0.000000e+00<br />Petal.Width: 1.0863014","density: 0.000000e+00<br />Petal.Width: 1.0816047","density: 7.886404e-17<br />Petal.Width: 1.0769080","density: 6.758470e-17<br />Petal.Width: 1.0722114","density: 0.000000e+00<br />Petal.Width: 1.0675147","density: 0.000000e+00<br />Petal.Width: 1.0628180","density: 7.836348e-18<br />Petal.Width: 1.0581213","density: 3.162430e-17<br />Petal.Width: 1.0534247","density: 0.000000e+00<br />Petal.Width: 1.0487280","density: 0.000000e+00<br />Petal.Width: 1.0440313","density: 0.000000e+00<br />Petal.Width: 1.0393346","density: 0.000000e+00<br />Petal.Width: 1.0346380","density: 0.000000e+00<br />Petal.Width: 1.0299413","density: 0.000000e+00<br />Petal.Width: 1.0252446","density: 0.000000e+00<br />Petal.Width: 1.0205479","density: 8.612415e-18<br />Petal.Width: 1.0158513","density: 1.608065e-17<br />Petal.Width: 1.0111546","density: 0.000000e+00<br />Petal.Width: 1.0064579","density: 0.000000e+00<br />Petal.Width: 1.0017613","density: 0.000000e+00<br />Petal.Width: 0.9970646","density: 0.000000e+00<br />Petal.Width: 0.9923679","density: 0.000000e+00<br />Petal.Width: 0.9876712","density: 0.000000e+00<br />Petal.Width: 0.9829746","density: 0.000000e+00<br />Petal.Width: 0.9782779","density: 0.000000e+00<br />Petal.Width: 0.9735812","density: 0.000000e+00<br />Petal.Width: 0.9688845","density: 0.000000e+00<br />Petal.Width: 0.9641879","density: 0.000000e+00<br />Petal.Width: 0.9594912","density: 2.402872e-20<br />Petal.Width: 0.9547945","density: 1.270498e-19<br />Petal.Width: 0.9500978","density: 1.648818e-21<br />Petal.Width: 0.9454012","density: 0.000000e+00<br />Petal.Width: 0.9407045","density: 0.000000e+00<br />Petal.Width: 0.9360078","density: 0.000000e+00<br />Petal.Width: 0.9313112","density: 0.000000e+00<br />Petal.Width: 0.9266145","density: 0.000000e+00<br />Petal.Width: 0.9219178","density: 0.000000e+00<br />Petal.Width: 0.9172211","density: 0.000000e+00<br />Petal.Width: 0.9125245","density: 0.000000e+00<br />Petal.Width: 0.9078278","density: 0.000000e+00<br />Petal.Width: 0.9031311","density: 0.000000e+00<br />Petal.Width: 0.8984344","density: 0.000000e+00<br />Petal.Width: 0.8937378","density: 0.000000e+00<br />Petal.Width: 0.8890411","density: 2.666047e-16<br />Petal.Width: 0.8843444","density: 3.538269e-16<br />Petal.Width: 0.8796477","density: 1.824401e-16<br />Petal.Width: 0.8749511","density: 2.786069e-16<br />Petal.Width: 0.8702544","density: 3.898402e-16<br />Petal.Width: 0.8655577","density: 3.477739e-16<br />Petal.Width: 0.8608611","density: 6.323343e-16<br />Petal.Width: 0.8561644","density: 1.113737e-15<br />Petal.Width: 0.8514677","density: 3.445976e-15<br />Petal.Width: 0.8467710","density: 1.153361e-14<br />Petal.Width: 0.8420744","density: 3.897752e-14<br />Petal.Width: 0.8373777","density: 1.278301e-13<br />Petal.Width: 0.8326810","density: 4.095881e-13<br />Petal.Width: 0.8279843","density: 1.269888e-12<br />Petal.Width: 0.8232877","density: 3.805637e-12<br />Petal.Width: 0.8185910","density: 1.101918e-11<br />Petal.Width: 0.8138943","density: 3.080191e-11<br />Petal.Width: 0.8091977","density: 8.302819e-11<br />Petal.Width: 0.8045010","density: 2.155147e-10<br />Petal.Width: 0.7998043","density: 5.376985e-10<br />Petal.Width: 0.7951076","density: 1.414652e-09<br />Petal.Width: 0.7904110","density: 3.680119e-09<br />Petal.Width: 0.7857143","density: 9.289056e-09<br />Petal.Width: 0.7810176","density: 2.274944e-08<br />Petal.Width: 0.7763209","density: 5.405821e-08<br />Petal.Width: 0.7716243","density: 1.246409e-07<br />Petal.Width: 0.7669276","density: 2.788709e-07<br />Petal.Width: 0.7622309","density: 6.055498e-07<br />Petal.Width: 0.7575342","density: 1.276424e-06<br />Petal.Width: 0.7528376","density: 2.612618e-06<br />Petal.Width: 0.7481409","density: 5.212275e-06<br />Petal.Width: 0.7434442","density: 1.065718e-05<br />Petal.Width: 0.7387476","density: 2.118486e-05<br />Petal.Width: 0.7340509","density: 4.095370e-05<br />Petal.Width: 0.7293542","density: 7.701642e-05<br />Petal.Width: 0.7246575","density: 1.409485e-04<br />Petal.Width: 0.7199609","density: 2.511400e-04<br />Petal.Width: 0.7152642","density: 4.358834e-04<br />Petal.Width: 0.7105675","density: 7.373486e-04<br />Petal.Width: 0.7058708","density: 1.216466e-03<br />Petal.Width: 0.7011742","density: 1.958623e-03<br />Petal.Width: 0.6964775","density: 3.098804e-03<br />Petal.Width: 0.6917808","density: 4.852254e-03<br />Petal.Width: 0.6870841","density: 7.403158e-03<br />Petal.Width: 0.6823875","density: 1.101040e-02<br />Petal.Width: 0.6776908","density: 1.596987e-02<br />Petal.Width: 0.6729941","density: 2.260053e-02<br />Petal.Width: 0.6682975","density: 3.122214e-02<br />Petal.Width: 0.6636008","density: 4.212468e-02<br />Petal.Width: 0.6589041","density: 5.553037e-02<br />Petal.Width: 0.6542074","density: 7.155068e-02<br />Petal.Width: 0.6495108","density: 9.014184e-02<br />Petal.Width: 0.6448141","density: 1.111660e-01<br />Petal.Width: 0.6401174","density: 1.339922e-01<br />Petal.Width: 0.6354207","density: 1.576959e-01<br />Petal.Width: 0.6307241","density: 1.812828e-01<br />Petal.Width: 0.6260274","density: 2.036269e-01<br />Petal.Width: 0.6213307","density: 2.235554e-01<br />Petal.Width: 0.6166341","density: 2.399491e-01<br />Petal.Width: 0.6119374","density: 2.518468e-01<br />Petal.Width: 0.6072407","density: 2.585446e-01<br />Petal.Width: 0.6025440","density: 2.596786e-01<br />Petal.Width: 0.5978474","density: 2.552800e-01<br />Petal.Width: 0.5931507","density: 2.454562e-01<br />Petal.Width: 0.5884540","density: 2.315721e-01<br />Petal.Width: 0.5837573","density: 2.150405e-01<br />Petal.Width: 0.5790607","density: 1.973102e-01<br />Petal.Width: 0.5743640","density: 1.798892e-01<br />Petal.Width: 0.5696673","density: 1.642301e-01<br />Petal.Width: 0.5649706","density: 1.516245e-01<br />Petal.Width: 0.5602740","density: 1.431112e-01<br />Petal.Width: 0.5555773","density: 1.394051e-01<br />Petal.Width: 0.5508806","density: 1.408477e-01<br />Petal.Width: 0.5461840","density: 1.473820e-01<br />Petal.Width: 0.5414873","density: 1.589466e-01<br />Petal.Width: 0.5367906","density: 1.740094e-01<br />Petal.Width: 0.5320939","density: 1.913441e-01<br />Petal.Width: 0.5273973","density: 2.095716e-01<br />Petal.Width: 0.5227006","density: 2.272757e-01<br />Petal.Width: 0.5180039","density: 2.431379e-01<br />Petal.Width: 0.5133072","density: 2.560766e-01<br />Petal.Width: 0.5086106","density: 2.653826e-01<br />Petal.Width: 0.5039139","density: 2.708405e-01<br />Petal.Width: 0.4992172","density: 2.728256e-01<br />Petal.Width: 0.4945205","density: 2.723592e-01<br />Petal.Width: 0.4898239","density: 2.715232e-01<br />Petal.Width: 0.4851272","density: 2.731199e-01<br />Petal.Width: 0.4804305","density: 2.802090e-01<br />Petal.Width: 0.4757339","density: 2.961415e-01<br />Petal.Width: 0.4710372","density: 3.243332e-01<br />Petal.Width: 0.4663405","density: 3.679948e-01<br />Petal.Width: 0.4616438","density: 4.298286e-01<br />Petal.Width: 0.4569472","density: 5.117065e-01<br />Petal.Width: 0.4522505","density: 6.143500e-01<br />Petal.Width: 0.4475538","density: 7.370404e-01<br />Petal.Width: 0.4428571","density: 8.780275e-01<br />Petal.Width: 0.4381605","density: 1.032057e+00<br />Petal.Width: 0.4334638","density: 1.191503e+00<br />Petal.Width: 0.4287671","density: 1.348595e+00<br />Petal.Width: 0.4240705","density: 1.494889e+00<br />Petal.Width: 0.4193738","density: 1.621942e+00<br />Petal.Width: 0.4146771","density: 1.722054e+00<br />Petal.Width: 0.4099804","density: 1.789000e+00<br />Petal.Width: 0.4052838","density: 1.818678e+00<br />Petal.Width: 0.4005871","density: 1.809599e+00<br />Petal.Width: 0.3958904","density: 1.763161e+00<br />Petal.Width: 0.3911937","density: 1.681784e+00<br />Petal.Width: 0.3864971","density: 1.575505e+00<br />Petal.Width: 0.3818004","density: 1.455023e+00<br />Petal.Width: 0.3771037","density: 1.330744e+00<br />Petal.Width: 0.3724070","density: 1.213123e+00<br />Petal.Width: 0.3677104","density: 1.111876e+00<br />Petal.Width: 0.3630137","density: 1.035282e+00<br />Petal.Width: 0.3583170","density: 9.896091e-01<br />Petal.Width: 0.3536204","density: 9.786877e-01<br />Petal.Width: 0.3489237","density: 1.003647e+00<br />Petal.Width: 0.3442270","density: 1.062822e+00<br />Petal.Width: 0.3395303","density: 1.154013e+00<br />Petal.Width: 0.3348337","density: 1.266129e+00<br />Petal.Width: 0.3301370","density: 1.389687e+00<br />Petal.Width: 0.3254403","density: 1.514655e+00<br />Petal.Width: 0.3207436","density: 1.631032e+00<br />Petal.Width: 0.3160470","density: 1.729744e+00<br />Petal.Width: 0.3113503","density: 1.803525e+00<br />Petal.Width: 0.3066536","density: 1.847721e+00<br />Petal.Width: 0.3019569","density: 1.860937e+00<br />Petal.Width: 0.2972603","density: 1.845497e+00<br />Petal.Width: 0.2925636","density: 1.807449e+00<br />Petal.Width: 0.2878669","density: 1.758129e+00<br />Petal.Width: 0.2831703","density: 1.714243e+00<br />Petal.Width: 0.2784736","density: 1.692391e+00<br />Petal.Width: 0.2737769","density: 1.710088e+00<br />Petal.Width: 0.2690802","density: 1.784477e+00<br />Petal.Width: 0.2643836","density: 1.930915e+00<br />Petal.Width: 0.2596869","density: 2.161530e+00<br />Petal.Width: 0.2549902","density: 2.483780e+00<br />Petal.Width: 0.2502935","density: 2.899154e+00<br />Petal.Width: 0.2455969","density: 3.402103e+00<br />Petal.Width: 0.2409002","density: 3.981448e+00<br />Petal.Width: 0.2362035","density: 4.611884e+00<br />Petal.Width: 0.2315068","density: 5.257489e+00<br />Petal.Width: 0.2268102","density: 5.883051e+00<br />Petal.Width: 0.2221135","density: 6.451561e+00<br />Petal.Width: 0.2174168","density: 6.927205e+00<br />Petal.Width: 0.2127202","density: 7.278457e+00<br />Petal.Width: 0.2080235","density: 7.480938e+00<br />Petal.Width: 0.2033268","density: 7.519733e+00<br />Petal.Width: 0.1986301","density: 7.390906e+00<br />Petal.Width: 0.1939335","density: 7.102006e+00<br />Petal.Width: 0.1892368","density: 6.663940e+00<br />Petal.Width: 0.1845401","density: 6.114490e+00<br />Petal.Width: 0.1798434","density: 5.494904e+00<br />Petal.Width: 0.1751468","density: 4.841721e+00<br />Petal.Width: 0.1704501","density: 4.189882e+00<br />Petal.Width: 0.1657534","density: 3.570246e+00<br />Petal.Width: 0.1610568","density: 3.007748e+00<br />Petal.Width: 0.1563601","density: 2.520287e+00<br />Petal.Width: 0.1516634","density: 2.118323e+00<br />Petal.Width: 0.1469667","density: 1.805119e+00<br />Petal.Width: 0.1422701","density: 1.577518e+00<br />Petal.Width: 0.1375734","density: 1.433029e+00<br />Petal.Width: 0.1328767","density: 1.350846e+00<br />Petal.Width: 0.1281800","density: 1.314673e+00<br />Petal.Width: 0.1234834","density: 1.309189e+00<br />Petal.Width: 0.1187867","density: 1.319820e+00<br />Petal.Width: 0.1140900","density: 1.333622e+00<br />Petal.Width: 0.1093933","density: 1.339929e+00<br />Petal.Width: 0.1046967","density: 1.330788e+00<br />Petal.Width: 0.1000000","density: 1.330788e+00<br />Petal.Width: 0.1000000"],"key":["1","2","3","4","5","6","7","8","9","10","11","12","13","14","15","16","17","18","19","20","21","22","23","24","25","26","27","28","29","30","31","32","33","34","35","36","37","38","39","40","41","42","43","44","45","46","47","48","49","50"],"type":"scatter","mode":"lines","line":{"width":1.88976377952756,"color":"rgba(0,0,0,1)","dash":"solid"},"fill":"toself","fillcolor":"rgba(248,118,109,1)","hoveron":"points","set":"SharedData91833e3a","name":"setosa","legendgroup":"setosa","showlegend":true,"xaxis":"x4","yaxis":"y13","hoverinfo":"text","_isSimpleKey":true,"_isNestedKey":false,"frame":null},{"x":[0.1,0.104696673189824,0.109393346379648,0.114090019569472,0.118786692759296,0.123483365949119,0.128180039138943,0.132876712328767,0.137573385518591,0.142270058708415,0.146966731898239,0.151663405088063,0.156360078277887,0.16105675146771,0.165753424657534,0.170450097847358,0.175146771037182,0.179843444227006,0.18454011741683,0.189236790606654,0.193933463796477,0.198630136986301,0.203326810176125,0.208023483365949,0.212720156555773,0.217416829745597,0.222113502935421,0.226810176125245,0.231506849315068,0.236203522504892,0.240900195694716,0.24559686888454,0.250293542074364,0.254990215264188,0.259686888454012,0.264383561643836,0.26908023483366,0.273776908023483,0.278473581213307,0.283170254403131,0.287866927592955,0.292563600782779,0.297260273972603,0.301956947162427,0.306653620352251,0.311350293542074,0.316046966731898,0.320743639921722,0.325440313111546,0.33013698630137,0.334833659491194,0.339530332681018,0.344227005870841,0.348923679060665,0.353620352250489,0.358317025440313,0.363013698630137,0.367710371819961,0.372407045009785,0.377103718199609,0.381800391389432,0.386497064579256,0.39119373776908,0.395890410958904,0.400587084148728,0.405283757338552,0.409980430528376,0.4146771037182,0.419373776908023,0.424070450097847,0.428767123287671,0.433463796477495,0.438160469667319,0.442857142857143,0.447553816046967,0.452250489236791,0.456947162426614,0.461643835616438,0.466340508806262,0.471037181996086,0.47573385518591,0.480430528375734,0.485127201565558,0.489823874755382,0.494520547945206,0.499217221135029,0.503913894324853,0.508610567514677,0.513307240704501,0.518003913894325,0.522700587084149,0.527397260273973,0.532093933463797,0.53679060665362,0.541487279843444,0.546183953033268,0.550880626223092,0.555577299412916,0.56027397260274,0.564970645792564,0.569667318982387,0.574363992172211,0.579060665362035,0.583757338551859,0.588454011741683,0.593150684931507,0.597847358121331,0.602544031311155,0.607240704500978,0.611937377690802,0.616634050880626,0.62133072407045,0.626027397260274,0.630724070450098,0.635420743639922,0.640117416829746,0.644814090019569,0.649510763209393,0.654207436399217,0.658904109589041,0.663600782778865,0.668297455968689,0.672994129158513,0.677690802348337,0.68238747553816,0.687084148727984,0.691780821917808,0.696477495107632,0.701174168297456,0.70587084148728,0.710567514677104,0.715264187866928,0.719960861056751,0.724657534246575,0.729354207436399,0.734050880626223,0.738747553816047,0.743444227005871,0.748140900195695,0.752837573385519,0.757534246575342,0.762230919765166,0.76692759295499,0.771624266144814,0.776320939334638,0.781017612524462,0.785714285714286,0.79041095890411,0.795107632093933,0.799804305283757,0.804500978473581,0.809197651663405,0.813894324853229,0.818590998043053,0.823287671232877,0.827984344422701,0.832681017612524,0.837377690802348,0.842074363992172,0.846771037181996,0.85146771037182,0.856164383561644,0.860861056751468,0.865557729941292,0.870254403131115,0.874951076320939,0.879647749510763,0.884344422700587,0.889041095890411,0.893737769080235,0.898434442270059,0.903131115459883,0.907827788649706,0.91252446183953,0.917221135029354,0.921917808219178,0.926614481409002,0.931311154598826,0.93600782778865,0.940704500978474,0.945401174168297,0.950097847358121,0.954794520547945,0.959491193737769,0.964187866927593,0.968884540117417,0.973581213307241,0.978277886497064,0.982974559686888,0.987671232876712,0.992367906066536,0.99706457925636,1.00176125244618,1.00645792563601,1.01115459882583,1.01585127201566,1.02054794520548,1.0252446183953,1.02994129158513,1.03463796477495,1.03933463796477,1.0440313111546,1.04872798434442,1.05342465753425,1.05812133072407,1.06281800391389,1.06751467710372,1.07221135029354,1.07690802348337,1.08160469667319,1.08630136986301,1.09099804305284,1.09569471624266,1.10039138943249,1.10508806262231,1.10978473581213,1.11448140900196,1.11917808219178,1.1238747553816,1.12857142857143,1.13326810176125,1.13796477495108,1.1426614481409,1.14735812133072,1.15205479452055,1.15675146771037,1.1614481409002,1.16614481409002,1.17084148727984,1.17553816046967,1.18023483365949,1.18493150684932,1.18962818003914,1.19432485322896,1.19902152641879,1.20371819960861,1.20841487279843,1.21311154598826,1.21780821917808,1.22250489236791,1.22720156555773,1.23189823874755,1.23659491193738,1.2412915851272,1.24598825831703,1.25068493150685,1.25538160469667,1.2600782778865,1.26477495107632,1.26947162426614,1.27416829745597,1.27886497064579,1.28356164383562,1.28825831702544,1.29295499021526,1.29765166340509,1.30234833659491,1.30704500978474,1.31174168297456,1.31643835616438,1.32113502935421,1.32583170254403,1.33052837573386,1.33522504892368,1.3399217221135,1.34461839530333,1.34931506849315,1.35401174168297,1.3587084148728,1.36340508806262,1.36810176125245,1.37279843444227,1.37749510763209,1.38219178082192,1.38688845401174,1.39158512720157,1.39628180039139,1.40097847358121,1.40567514677104,1.41037181996086,1.41506849315068,1.41976516634051,1.42446183953033,1.42915851272016,1.43385518590998,1.4385518590998,1.44324853228963,1.44794520547945,1.45264187866928,1.4573385518591,1.46203522504892,1.46673189823875,1.47142857142857,1.4761252446184,1.48082191780822,1.48551859099804,1.49021526418787,1.49491193737769,1.49960861056751,1.50430528375734,1.50900195694716,1.51369863013699,1.51839530332681,1.52309197651663,1.52778864970646,1.53248532289628,1.53718199608611,1.54187866927593,1.54657534246575,1.55127201565558,1.5559686888454,1.56066536203523,1.56536203522505,1.57005870841487,1.5747553816047,1.57945205479452,1.58414872798434,1.58884540117417,1.59354207436399,1.59823874755382,1.60293542074364,1.60763209393346,1.61232876712329,1.61702544031311,1.62172211350294,1.62641878669276,1.63111545988258,1.63581213307241,1.64050880626223,1.64520547945205,1.64990215264188,1.6545988258317,1.65929549902153,1.66399217221135,1.66868884540117,1.673385518591,1.67808219178082,1.68277886497065,1.68747553816047,1.69217221135029,1.69686888454012,1.70156555772994,1.70626223091977,1.71095890410959,1.71565557729941,1.72035225048924,1.72504892367906,1.72974559686888,1.73444227005871,1.73913894324853,1.74383561643836,1.74853228962818,1.753228962818,1.75792563600783,1.76262230919765,1.76731898238748,1.7720156555773,1.77671232876712,1.78140900195695,1.78610567514677,1.79080234833659,1.79549902152642,1.80019569471624,1.80489236790607,1.80958904109589,1.81428571428571,1.81898238747554,1.82367906066536,1.82837573385519,1.83307240704501,1.83776908023483,1.84246575342466,1.84716242661448,1.85185909980431,1.85655577299413,1.86125244618395,1.86594911937378,1.8706457925636,1.87534246575342,1.88003913894325,1.88473581213307,1.8894324853229,1.89412915851272,1.89882583170254,1.90352250489237,1.90821917808219,1.91291585127202,1.91761252446184,1.92230919765166,1.92700587084149,1.93170254403131,1.93639921722114,1.94109589041096,1.94579256360078,1.95048923679061,1.95518590998043,1.95988258317025,1.96457925636008,1.9692759295499,1.97397260273973,1.97866927592955,1.98336594911937,1.9880626223092,1.99275929549902,1.99745596868885,2.00215264187867,2.00684931506849,2.01154598825832,2.01624266144814,2.02093933463796,2.02563600782779,2.03033268101761,2.03502935420744,2.03972602739726,2.04442270058708,2.04911937377691,2.05381604696673,2.05851272015656,2.06320939334638,2.0679060665362,2.07260273972603,2.07729941291585,2.08199608610568,2.0866927592955,2.09138943248532,2.09608610567515,2.10078277886497,2.10547945205479,2.11017612524462,2.11487279843444,2.11956947162427,2.12426614481409,2.12896281800391,2.13365949119374,2.13835616438356,2.14305283757339,2.14774951076321,2.15244618395303,2.15714285714286,2.16183953033268,2.1665362035225,2.17123287671233,2.17592954990215,2.18062622309198,2.1853228962818,2.19001956947162,2.19471624266145,2.19941291585127,2.2041095890411,2.20880626223092,2.21350293542074,2.21819960861057,2.22289628180039,2.22759295499022,2.23228962818004,2.23698630136986,2.24168297455969,2.24637964774951,2.25107632093933,2.25577299412916,2.26046966731898,2.26516634050881,2.26986301369863,2.27455968688845,2.27925636007828,2.2839530332681,2.28864970645793,2.29334637964775,2.29804305283757,2.3027397260274,2.30743639921722,2.31213307240705,2.31682974559687,2.32152641878669,2.32622309197652,2.33091976516634,2.33561643835616,2.34031311154599,2.34500978473581,2.34970645792564,2.35440313111546,2.35909980430528,2.36379647749511,2.36849315068493,2.37318982387476,2.37788649706458,2.3825831702544,2.38727984344423,2.39197651663405,2.39667318982387,2.4013698630137,2.40606653620352,2.41076320939335,2.41545988258317,2.42015655577299,2.42485322896282,2.42954990215264,2.43424657534247,2.43894324853229,2.44363992172211,2.44833659491194,2.45303326810176,2.45772994129159,2.46242661448141,2.46712328767123,2.47181996086106,2.47651663405088,2.4812133072407,2.48590998043053,2.49060665362035,2.49530332681018,2.5,2.5,2.49530332681018,2.49060665362035,2.48590998043053,2.4812133072407,2.47651663405088,2.47181996086106,2.46712328767123,2.46242661448141,2.45772994129159,2.45303326810176,2.44833659491194,2.44363992172211,2.43894324853229,2.43424657534247,2.42954990215264,2.42485322896282,2.42015655577299,2.41545988258317,2.41076320939335,2.40606653620352,2.4013698630137,2.39667318982387,2.39197651663405,2.38727984344423,2.3825831702544,2.37788649706458,2.37318982387476,2.36849315068493,2.36379647749511,2.35909980430528,2.35440313111546,2.34970645792564,2.34500978473581,2.34031311154599,2.33561643835616,2.33091976516634,2.32622309197652,2.32152641878669,2.31682974559687,2.31213307240705,2.30743639921722,2.3027397260274,2.29804305283757,2.29334637964775,2.28864970645793,2.2839530332681,2.27925636007828,2.27455968688845,2.26986301369863,2.26516634050881,2.26046966731898,2.25577299412916,2.25107632093933,2.24637964774951,2.24168297455969,2.23698630136986,2.23228962818004,2.22759295499022,2.22289628180039,2.21819960861057,2.21350293542074,2.20880626223092,2.2041095890411,2.19941291585127,2.19471624266145,2.19001956947162,2.1853228962818,2.18062622309198,2.17592954990215,2.17123287671233,2.1665362035225,2.16183953033268,2.15714285714286,2.15244618395303,2.14774951076321,2.14305283757339,2.13835616438356,2.13365949119374,2.12896281800391,2.12426614481409,2.11956947162427,2.11487279843444,2.11017612524462,2.10547945205479,2.10078277886497,2.09608610567515,2.09138943248532,2.0866927592955,2.08199608610568,2.07729941291585,2.07260273972603,2.0679060665362,2.06320939334638,2.05851272015656,2.05381604696673,2.04911937377691,2.04442270058708,2.03972602739726,2.03502935420744,2.03033268101761,2.02563600782779,2.02093933463796,2.01624266144814,2.01154598825832,2.00684931506849,2.00215264187867,1.99745596868885,1.99275929549902,1.9880626223092,1.98336594911937,1.97866927592955,1.97397260273973,1.9692759295499,1.96457925636008,1.95988258317025,1.95518590998043,1.95048923679061,1.94579256360078,1.94109589041096,1.93639921722114,1.93170254403131,1.92700587084149,1.92230919765166,1.91761252446184,1.91291585127202,1.90821917808219,1.90352250489237,1.89882583170254,1.89412915851272,1.8894324853229,1.88473581213307,1.88003913894325,1.87534246575342,1.8706457925636,1.86594911937378,1.86125244618395,1.85655577299413,1.85185909980431,1.84716242661448,1.84246575342466,1.83776908023483,1.83307240704501,1.82837573385519,1.82367906066536,1.81898238747554,1.81428571428571,1.80958904109589,1.80489236790607,1.80019569471624,1.79549902152642,1.79080234833659,1.78610567514677,1.78140900195695,1.77671232876712,1.7720156555773,1.76731898238748,1.76262230919765,1.75792563600783,1.753228962818,1.74853228962818,1.74383561643836,1.73913894324853,1.73444227005871,1.72974559686888,1.72504892367906,1.72035225048924,1.71565557729941,1.71095890410959,1.70626223091977,1.70156555772994,1.69686888454012,1.69217221135029,1.68747553816047,1.68277886497065,1.67808219178082,1.673385518591,1.66868884540117,1.66399217221135,1.65929549902153,1.6545988258317,1.64990215264188,1.64520547945205,1.64050880626223,1.63581213307241,1.63111545988258,1.62641878669276,1.62172211350294,1.61702544031311,1.61232876712329,1.60763209393346,1.60293542074364,1.59823874755382,1.59354207436399,1.58884540117417,1.58414872798434,1.57945205479452,1.5747553816047,1.57005870841487,1.56536203522505,1.56066536203523,1.5559686888454,1.55127201565558,1.54657534246575,1.54187866927593,1.53718199608611,1.53248532289628,1.52778864970646,1.52309197651663,1.51839530332681,1.51369863013699,1.50900195694716,1.50430528375734,1.49960861056751,1.49491193737769,1.49021526418787,1.48551859099804,1.48082191780822,1.4761252446184,1.47142857142857,1.46673189823875,1.46203522504892,1.4573385518591,1.45264187866928,1.44794520547945,1.44324853228963,1.4385518590998,1.43385518590998,1.42915851272016,1.42446183953033,1.41976516634051,1.41506849315068,1.41037181996086,1.40567514677104,1.40097847358121,1.39628180039139,1.39158512720157,1.38688845401174,1.38219178082192,1.37749510763209,1.37279843444227,1.36810176125245,1.36340508806262,1.3587084148728,1.35401174168297,1.34931506849315,1.34461839530333,1.3399217221135,1.33522504892368,1.33052837573386,1.32583170254403,1.32113502935421,1.31643835616438,1.31174168297456,1.30704500978474,1.30234833659491,1.29765166340509,1.29295499021526,1.28825831702544,1.28356164383562,1.27886497064579,1.27416829745597,1.26947162426614,1.26477495107632,1.2600782778865,1.25538160469667,1.25068493150685,1.24598825831703,1.2412915851272,1.23659491193738,1.23189823874755,1.22720156555773,1.22250489236791,1.21780821917808,1.21311154598826,1.20841487279843,1.20371819960861,1.19902152641879,1.19432485322896,1.18962818003914,1.18493150684932,1.18023483365949,1.17553816046967,1.17084148727984,1.16614481409002,1.1614481409002,1.15675146771037,1.15205479452055,1.14735812133072,1.1426614481409,1.13796477495108,1.13326810176125,1.12857142857143,1.1238747553816,1.11917808219178,1.11448140900196,1.10978473581213,1.10508806262231,1.10039138943249,1.09569471624266,1.09099804305284,1.08630136986301,1.08160469667319,1.07690802348337,1.07221135029354,1.06751467710372,1.06281800391389,1.05812133072407,1.05342465753425,1.04872798434442,1.0440313111546,1.03933463796477,1.03463796477495,1.02994129158513,1.0252446183953,1.02054794520548,1.01585127201566,1.01115459882583,1.00645792563601,1.00176125244618,0.99706457925636,0.992367906066536,0.987671232876712,0.982974559686888,0.978277886497064,0.973581213307241,0.968884540117417,0.964187866927593,0.959491193737769,0.954794520547945,0.950097847358121,0.945401174168297,0.940704500978474,0.93600782778865,0.931311154598826,0.926614481409002,0.921917808219178,0.917221135029354,0.91252446183953,0.907827788649706,0.903131115459883,0.898434442270059,0.893737769080235,0.889041095890411,0.884344422700587,0.879647749510763,0.874951076320939,0.870254403131115,0.865557729941292,0.860861056751468,0.856164383561644,0.85146771037182,0.846771037181996,0.842074363992172,0.837377690802348,0.832681017612524,0.827984344422701,0.823287671232877,0.818590998043053,0.813894324853229,0.809197651663405,0.804500978473581,0.799804305283757,0.795107632093933,0.79041095890411,0.785714285714286,0.781017612524462,0.776320939334638,0.771624266144814,0.76692759295499,0.762230919765166,0.757534246575342,0.752837573385519,0.748140900195695,0.743444227005871,0.738747553816047,0.734050880626223,0.729354207436399,0.724657534246575,0.719960861056751,0.715264187866928,0.710567514677104,0.70587084148728,0.701174168297456,0.696477495107632,0.691780821917808,0.687084148727984,0.68238747553816,0.677690802348337,0.672994129158513,0.668297455968689,0.663600782778865,0.658904109589041,0.654207436399217,0.649510763209393,0.644814090019569,0.640117416829746,0.635420743639922,0.630724070450098,0.626027397260274,0.62133072407045,0.616634050880626,0.611937377690802,0.607240704500978,0.602544031311155,0.597847358121331,0.593150684931507,0.588454011741683,0.583757338551859,0.579060665362035,0.574363992172211,0.569667318982387,0.564970645792564,0.56027397260274,0.555577299412916,0.550880626223092,0.546183953033268,0.541487279843444,0.53679060665362,0.532093933463797,0.527397260273973,0.522700587084149,0.518003913894325,0.513307240704501,0.508610567514677,0.503913894324853,0.499217221135029,0.494520547945206,0.489823874755382,0.485127201565558,0.480430528375734,0.47573385518591,0.471037181996086,0.466340508806262,0.461643835616438,0.456947162426614,0.452250489236791,0.447553816046967,0.442857142857143,0.438160469667319,0.433463796477495,0.428767123287671,0.424070450097847,0.419373776908023,0.4146771037182,0.409980430528376,0.405283757338552,0.400587084148728,0.395890410958904,0.39119373776908,0.386497064579256,0.381800391389432,0.377103718199609,0.372407045009785,0.367710371819961,0.363013698630137,0.358317025440313,0.353620352250489,0.348923679060665,0.344227005870841,0.339530332681018,0.334833659491194,0.33013698630137,0.325440313111546,0.320743639921722,0.316046966731898,0.311350293542074,0.306653620352251,0.301956947162427,0.297260273972603,0.292563600782779,0.287866927592955,0.283170254403131,0.278473581213307,0.273776908023483,0.26908023483366,0.264383561643836,0.259686888454012,0.254990215264188,0.250293542074364,0.24559686888454,0.240900195694716,0.236203522504892,0.231506849315068,0.226810176125245,0.222113502935421,0.217416829745597,0.212720156555773,0.208023483365949,0.203326810176125,0.198630136986301,0.193933463796477,0.189236790606654,0.18454011741683,0.179843444227006,0.175146771037182,0.170450097847358,0.165753424657534,0.16105675146771,0.156360078277887,0.151663405088063,0.146966731898239,0.142270058708415,0.137573385518591,0.132876712328767,0.128180039138943,0.123483365949119,0.118786692759296,0.114090019569472,0.109393346379648,0.104696673189824,0.1,0.1],"y":[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,7.30283588187437e-17,1.46039667686887e-16,1.14548681075808e-16,3.81996040753735e-16,6.74588811523286e-16,1.13197163238927e-15,1.83233262307725e-15,2.82025042107122e-15,4.52148031870572e-15,7.2035308791984e-15,1.12550361323241e-14,1.71977311867499e-14,2.65856793223832e-14,4.20097211137738e-14,6.5237637192693e-14,9.97028146411535e-14,1.49612428416275e-13,2.30910967648977e-13,3.56271319176782e-13,5.41804721543134e-13,8.10525416778713e-13,1.19732798334961e-12,1.83248361943286e-12,2.76916676237732e-12,4.12563273364903e-12,6.0483177546524e-12,8.91017643365151e-12,1.33589658768696e-11,1.97795814240622e-11,2.88825318883888e-11,4.15199097606907e-11,6.09232607347267e-11,8.95020844435528e-11,1.29890807439163e-10,1.85983855064832e-10,2.6294738370939e-10,3.828079438508e-10,5.512380537625e-10,7.84430408704742e-10,1.10187546291138e-09,1.5493271044198e-09,2.21088038558516e-09,3.12164036207405e-09,4.35759342036904e-09,6.00783025951159e-09,8.38980079229333e-09,1.17387624306938e-08,1.62577205814472e-08,2.22719636803858e-08,3.01536248652063e-08,4.17616261826183e-08,5.7311660832003e-08,7.78874621519481e-08,1.04759425577856e-07,1.40682744656459e-07,1.91121053302049e-07,2.57348824357634e-07,3.43325934546056e-07,4.5357889807497e-07,6.03468583513494e-07,8.04338482696509e-07,1.06306273666063e-06,1.3927673514051e-06,1.80816610406258e-06,2.38023666708873e-06,3.11364040477279e-06,4.04069938995484e-06,5.20100117919189e-06,6.67400290029809e-06,8.63457201610509e-06,1.10893207029709e-05,1.4135916805497e-05,1.78828944556226e-05,2.26882193018551e-05,2.88158451714911e-05,3.63467443165431e-05,4.55278908727892e-05,5.66293938552484e-05,7.09520212226521e-05,8.84954780440584e-05,0.000109668153301987,0.000135034477504379,0.000165645022373802,0.000204185860488711,0.000250183899241269,0.00030471877765856,0.000368952237284267,0.000446748498698175,0.000540951195497899,0.000651364412018569,0.000780007200161915,0.000929021473934995,0.00110937001977357,0.0013200113577179,0.00156255030688787,0.00184033640788232,0.00215918301900827,0.00253795620292602,0.00296864509412632,0.00345595637506209,0.00400475183281711,0.00463291904573955,0.00535343645185252,0.00615826070920895,0.00705332554115353,0.0080446498738815,0.00917113207942877,0.0104228099072001,0.0117966303303401,0.0132986697416349,0.0149391494624267,0.0167724341850574,0.0187573832550335,0.0208986613002034,0.0232006343903818,0.025692293030545,0.0283926916896648,0.0312660128393083,0.0343133450252801,0.0375352965536495,0.0409770846754017,0.0446088981055729,0.0484084891099251,0.0523720785731754,0.0564965269218211,0.06082666954665,0.0652969429092693,0.0698996555408657,0.0746267905005703,0.0794824059345057,0.0844625526681162,0.089529505086395,0.0946739602420506,0.0998866980359366,0.105168861117749,0.110499451859373,0.115864241816294,0.12125696153718,0.126671926907005,0.132108645369844,0.137558524042751,0.143021831004154,0.148500106630134,0.15399876033764,0.15953251555955,0.165106286072176,0.170731191157763,0.176419555820972,0.182209359366943,0.188127082750918,0.194181072302764,0.200393381632446,0.206787139331509,0.213475802366098,0.22043850103632,0.227697191156509,0.235282272113587,0.243258607723858,0.251768867156222,0.260741888802336,0.270211744667157,0.280212642729514,0.290904209354365,0.302336113363881,0.314441626256144,0.327253639768406,0.340804591607475,0.355389053788266,0.370853013004306,0.387178326052505,0.404390289334506,0.422569807026351,0.441989575672101,0.462378805616476,0.483749516137645,0.506111628884447,0.529650048931755,0.554383783521223,0.580109353636142,0.606815423701884,0.634487687930827,0.663370173805068,0.693211295674752,0.723899070341068,0.755391329300044,0.787668113879967,0.820835974970929,0.85457499403118,0.888814411480417,0.92348046118735,0.958526243696487,0.993807454370867,1.02916110391915,1.06449452594444,1.09971385713951,1.1346138696862,1.1690705490183,1.20301591197611,1.23635893278147,1.26899623004653,1.30051404954374,1.33108934996762,1.36065783058733,1.38916008506823,1.41638200995832,1.44213002759015,1.46665537925072,1.48994659663713,1.51199851578109,1.53251333168522,1.55169063097011,1.56971996398288,1.5866467537856,1.60252185102822,1.61709395942179,1.63082424530675,1.64379622644746,1.65609079159811,1.66774126078032,1.67883241887049,1.68959731186706,1.70011527306565,1.71046305509827,1.7207205528212,1.73100420918872,1.74136922514668,1.75185374731377,1.76248978160166,1.77336348585058,1.78441585196169,1.79562173811246,1.80695320898792,1.81837424755265,1.82978705486843,1.8410891904501,1.85219615923203,1.86301810051169,1.87333669264229,1.88296824438581,1.89186430523186,1.89990734210734,1.9069787384165,1.9125250067831,1.91667342779024,1.91937181883103,1.92051758125488,1.91988781773631,1.91692026010509,1.91205813378738,1.90524981720922,1.89645155735581,1.88524702653674,1.87161918147022,1.85596422217436,1.83830926783739,1.81868995723488,1.79661491666843,1.7726491719332,1.74701560216226,1.71981264287536,1.69108479686519,1.66067982718769,1.62920483195138,1.59679613674834,1.56359227091803,1.52967080528218,1.49528225957207,1.46069811305897,1.42605994534006,1.39150789464356,1.35731850541746,1.32366209901204,1.29063236483241,1.25834448851843,1.22693627046536,1.19689621720992,1.16800917952897,1.14035136235397,1.11399375869476,1.0892105667494,1.06621777882226,1.04473110591773,1.02477652547311,1.00637455389358,0.989945753858467,0.975252033486043,0.962097999005491,0.950458886829352,0.940311124343226,0.932100175607942,0.925217507514474,0.919595899212361,0.915163527292496,0.911959671669211,0.909948382622835,0.90875252565642,0.908266789216106,0.908382785664203,0.909049512618189,0.910005416441799,0.911067785194359,0.912113316465286,0.913018209532699,0.913471384693142,0.913391368079597,0.912669508978028,0.911194738565602,0.908755784950904,0.905005720579183,0.90010408829534,0.893978244368005,0.886561657211635,0.877484726015672,0.866761641769834,0.854588774147974,0.840957939617987,0.825868352988945,0.808866290998281,0.790431300418822,0.770670252805223,0.749640979538832,0.727324091007024,0.703626420055426,0.678995421093943,0.6535303220867,0.627333403141851,0.600429599289963,0.573038412475213,0.545377760804771,0.517559805433621,0.489696087129284,0.461985560906453,0.434572747767676,0.407545537520233,0.380995021628153,0.35504815269105,0.330012696761952,0.305768169265758,0.282369240214241,0.259865778189879,0.238476014723645,0.218280800940064,0.199106983903211,0.180965650597239,0.16386367929146,0.148100194361737,0.133440538754242,0.119781194778241,0.107100813359279,0.0954073926266508,0.0849256867116414,0.0752988458223735,0.0664907129596545,0.0584643449341737,0.051284209613408,0.0449244407645576,0.0391845109908732,0.0340260609845307,0.0294111279136157,0.0254191792417428,0.021907113717568,0.0187917541099279,0.016041565357356,0.0136299718611704,0.0116115880024225,0.00984279100513082,0.00830075299351425,0.00696367479554153,0.00583068537557631,0.00488652412148299,0.00407288068181294,0.00337584024864846,0.0027823030054433,0.00229776873029309,0.00189384613473456,0.00155162432382908,0.00126359770362332,0.00102280646150171,0.000833956296420932,0.000675776815955166,0.000544057448753174,0.000435179200570037,0.00034747621916973,0.000278695158754927,0.000221955450214969,0.000175534236036842,0.000137862042592913,0.000108739799476055,8.57371370613266e-05,6.70856776764767e-05,5.20994194185859e-05,4.0163773039089e-05,3.1330987275186e-05,2.42755219660981e-05,1.86551045709058e-05,1.42219751511615e-05,1.08191864629753e-05,8.30989185181558e-06,6.3246389574884e-06,4.77172408749301e-06,3.5698536677735e-06,2.68793463935678e-06,2.02844525538899e-06,1.51592744298072e-06,1.12246166932989e-06,8.23790206032684e-07,6.14733280201064e-07,4.55602719734688e-07,3.34199021145153e-07,2.42771638759364e-07,1.75775525777796e-07,1.29391647247826e-07,9.4139371084165e-08,6.77525648112057e-08,4.82691222176986e-08,3.4673019075497e-08,2.50599920226804e-08,1.78905874943936e-08,1.2628498293663e-08,8.82072171663461e-09,6.29496041373241e-09,4.46490443264234e-09,3.12646252616981e-09,2.1637011483498e-09,1.4891794118988e-09,1.05161542336997e-09,7.31645547684285e-10,5.02303242750525e-10,3.40706138479818e-10,2.33298881244013e-10,1.61612800014366e-10,1.10242267145428e-10,7.41778559295119e-11,4.92970661219376e-11,3.36320213392677e-11,2.28424684679154e-11,1.52707185784011e-11,1.00668703498321e-11,6.57650423521668e-12,4.46008813242016e-12,2.96856576822035e-12,1.94415784757395e-12,1.2552145064292e-12,8.18550063870164e-13,5.43851019805359e-13,3.54509548033722e-13,2.27349457465399e-13,1.43716105838671e-13,9.37491612449678e-14,6.09472934666549e-14,3.88295818140556e-14,2.43252708444702e-14,1.49898570116313e-14,9.8194455834704e-15,6.35372115241131e-15,4.03438483701535e-15,2.4667788205722e-15,1.53412438006276e-15,9.92360330912955e-16,6.01451425500808e-16,3.1549516000994e-16,1.15298452344132e-16,5.38947218058364e-17,5.91965166515612e-17,3.79983817787026e-17,0,0,0,1.92725540067398e-17,2.07209185197645e-17,1.06841547680604e-17,6.39007212345846e-17,1.81911434250834e-17,3.23311597305614e-17,1.01514026466457e-16,1.85773326071666e-16,1.4210845102221e-16,1.4452464491455e-16,1.0227638336219e-16,3.44003143586582e-17,2.40342322705475e-18,8.15460092541936e-17,4.67539452086688e-17,2.90443964880072e-17,4.8937770501627e-17,2.54384076158037e-18,0,0,2.19726495201607e-17,6.25823749608101e-17,1.07303358395815e-17,6.21420130487971e-17,5.16493715705274e-17,2.59934925691732e-17,8.50268888838882e-17,1.23233654552162e-16,9.37799949719824e-17,4.1909018486042e-17,0,0,0,1.88187393415459e-17,2.32796527916166e-17,1.96533811600361e-17,1.53377060998994e-16,5.52948122941593e-17,1.82008737265859e-18,0],"text":["density: 1.820087e-18<br />Petal.Width: 0.1000000","density: 5.529481e-17<br />Petal.Width: 0.1046967","density: 1.533771e-16<br />Petal.Width: 0.1093933","density: 1.965338e-17<br />Petal.Width: 0.1140900","density: 2.327965e-17<br />Petal.Width: 0.1187867","density: 1.881874e-17<br />Petal.Width: 0.1234834","density: 0.000000e+00<br />Petal.Width: 0.1281800","density: 0.000000e+00<br />Petal.Width: 0.1328767","density: 0.000000e+00<br />Petal.Width: 0.1375734","density: 4.190902e-17<br />Petal.Width: 0.1422701","density: 9.377999e-17<br />Petal.Width: 0.1469667","density: 1.232337e-16<br />Petal.Width: 0.1516634","density: 8.502689e-17<br />Petal.Width: 0.1563601","density: 2.599349e-17<br />Petal.Width: 0.1610568","density: 5.164937e-17<br />Petal.Width: 0.1657534","density: 6.214201e-17<br />Petal.Width: 0.1704501","density: 1.073034e-17<br />Petal.Width: 0.1751468","density: 6.258237e-17<br />Petal.Width: 0.1798434","density: 2.197265e-17<br />Petal.Width: 0.1845401","density: 0.000000e+00<br />Petal.Width: 0.1892368","density: 0.000000e+00<br />Petal.Width: 0.1939335","density: 2.543841e-18<br />Petal.Width: 0.1986301","density: 4.893777e-17<br />Petal.Width: 0.2033268","density: 2.904440e-17<br />Petal.Width: 0.2080235","density: 4.675395e-17<br />Petal.Width: 0.2127202","density: 8.154601e-17<br />Petal.Width: 0.2174168","density: 2.403423e-18<br />Petal.Width: 0.2221135","density: 3.440031e-17<br />Petal.Width: 0.2268102","density: 1.022764e-16<br />Petal.Width: 0.2315068","density: 1.445246e-16<br />Petal.Width: 0.2362035","density: 1.421085e-16<br />Petal.Width: 0.2409002","density: 1.857733e-16<br />Petal.Width: 0.2455969","density: 1.015140e-16<br />Petal.Width: 0.2502935","density: 3.233116e-17<br />Petal.Width: 0.2549902","density: 1.819114e-17<br />Petal.Width: 0.2596869","density: 6.390072e-17<br />Petal.Width: 0.2643836","density: 1.068415e-17<br />Petal.Width: 0.2690802","density: 2.072092e-17<br />Petal.Width: 0.2737769","density: 1.927255e-17<br />Petal.Width: 0.2784736","density: 0.000000e+00<br />Petal.Width: 0.2831703","density: 0.000000e+00<br />Petal.Width: 0.2878669","density: 0.000000e+00<br />Petal.Width: 0.2925636","density: 3.799838e-17<br />Petal.Width: 0.2972603","density: 5.919652e-17<br />Petal.Width: 0.3019569","density: 5.389472e-17<br />Petal.Width: 0.3066536","density: 1.152985e-16<br />Petal.Width: 0.3113503","density: 3.154952e-16<br />Petal.Width: 0.3160470","density: 6.014514e-16<br />Petal.Width: 0.3207436","density: 9.923603e-16<br />Petal.Width: 0.3254403","density: 1.534124e-15<br />Petal.Width: 0.3301370","density: 2.466779e-15<br />Petal.Width: 0.3348337","density: 4.034385e-15<br />Petal.Width: 0.3395303","density: 6.353721e-15<br />Petal.Width: 0.3442270","density: 9.819446e-15<br />Petal.Width: 0.3489237","density: 1.498986e-14<br />Petal.Width: 0.3536204","density: 2.432527e-14<br />Petal.Width: 0.3583170","density: 3.882958e-14<br />Petal.Width: 0.3630137","density: 6.094729e-14<br />Petal.Width: 0.3677104","density: 9.374916e-14<br />Petal.Width: 0.3724070","density: 1.437161e-13<br />Petal.Width: 0.3771037","density: 2.273495e-13<br />Petal.Width: 0.3818004","density: 3.545095e-13<br />Petal.Width: 0.3864971","density: 5.438510e-13<br />Petal.Width: 0.3911937","density: 8.185501e-13<br />Petal.Width: 0.3958904","density: 1.255215e-12<br />Petal.Width: 0.4005871","density: 1.944158e-12<br />Petal.Width: 0.4052838","density: 2.968566e-12<br />Petal.Width: 0.4099804","density: 4.460088e-12<br />Petal.Width: 0.4146771","density: 6.576504e-12<br />Petal.Width: 0.4193738","density: 1.006687e-11<br />Petal.Width: 0.4240705","density: 1.527072e-11<br />Petal.Width: 0.4287671","density: 2.284247e-11<br />Petal.Width: 0.4334638","density: 3.363202e-11<br />Petal.Width: 0.4381605","density: 4.929707e-11<br />Petal.Width: 0.4428571","density: 7.417786e-11<br />Petal.Width: 0.4475538","density: 1.102423e-10<br />Petal.Width: 0.4522505","density: 1.616128e-10<br />Petal.Width: 0.4569472","density: 2.332989e-10<br />Petal.Width: 0.4616438","density: 3.407061e-10<br />Petal.Width: 0.4663405","density: 5.023032e-10<br />Petal.Width: 0.4710372","density: 7.316455e-10<br />Petal.Width: 0.4757339","density: 1.051615e-09<br />Petal.Width: 0.4804305","density: 1.489179e-09<br />Petal.Width: 0.4851272","density: 2.163701e-09<br />Petal.Width: 0.4898239","density: 3.126463e-09<br />Petal.Width: 0.4945205","density: 4.464904e-09<br />Petal.Width: 0.4992172","density: 6.294960e-09<br />Petal.Width: 0.5039139","density: 8.820722e-09<br />Petal.Width: 0.5086106","density: 1.262850e-08<br />Petal.Width: 0.5133072","density: 1.789059e-08<br />Petal.Width: 0.5180039","density: 2.505999e-08<br />Petal.Width: 0.5227006","density: 3.467302e-08<br />Petal.Width: 0.5273973","density: 4.826912e-08<br />Petal.Width: 0.5320939","density: 6.775256e-08<br />Petal.Width: 0.5367906","density: 9.413937e-08<br />Petal.Width: 0.5414873","density: 1.293916e-07<br />Petal.Width: 0.5461840","density: 1.757755e-07<br />Petal.Width: 0.5508806","density: 2.427716e-07<br />Petal.Width: 0.5555773","density: 3.341990e-07<br />Petal.Width: 0.5602740","density: 4.556027e-07<br />Petal.Width: 0.5649706","density: 6.147333e-07<br />Petal.Width: 0.5696673","density: 8.237902e-07<br />Petal.Width: 0.5743640","density: 1.122462e-06<br />Petal.Width: 0.5790607","density: 1.515927e-06<br />Petal.Width: 0.5837573","density: 2.028445e-06<br />Petal.Width: 0.5884540","density: 2.687935e-06<br />Petal.Width: 0.5931507","density: 3.569854e-06<br />Petal.Width: 0.5978474","density: 4.771724e-06<br />Petal.Width: 0.6025440","density: 6.324639e-06<br />Petal.Width: 0.6072407","density: 8.309892e-06<br />Petal.Width: 0.6119374","density: 1.081919e-05<br />Petal.Width: 0.6166341","density: 1.422198e-05<br />Petal.Width: 0.6213307","density: 1.865510e-05<br />Petal.Width: 0.6260274","density: 2.427552e-05<br />Petal.Width: 0.6307241","density: 3.133099e-05<br />Petal.Width: 0.6354207","density: 4.016377e-05<br />Petal.Width: 0.6401174","density: 5.209942e-05<br />Petal.Width: 0.6448141","density: 6.708568e-05<br />Petal.Width: 0.6495108","density: 8.573714e-05<br />Petal.Width: 0.6542074","density: 1.087398e-04<br />Petal.Width: 0.6589041","density: 1.378620e-04<br />Petal.Width: 0.6636008","density: 1.755342e-04<br />Petal.Width: 0.6682975","density: 2.219555e-04<br />Petal.Width: 0.6729941","density: 2.786952e-04<br />Petal.Width: 0.6776908","density: 3.474762e-04<br />Petal.Width: 0.6823875","density: 4.351792e-04<br />Petal.Width: 0.6870841","density: 5.440574e-04<br />Petal.Width: 0.6917808","density: 6.757768e-04<br />Petal.Width: 0.6964775","density: 8.339563e-04<br />Petal.Width: 0.7011742","density: 1.022806e-03<br />Petal.Width: 0.7058708","density: 1.263598e-03<br />Petal.Width: 0.7105675","density: 1.551624e-03<br />Petal.Width: 0.7152642","density: 1.893846e-03<br />Petal.Width: 0.7199609","density: 2.297769e-03<br />Petal.Width: 0.7246575","density: 2.782303e-03<br />Petal.Width: 0.7293542","density: 3.375840e-03<br />Petal.Width: 0.7340509","density: 4.072881e-03<br />Petal.Width: 0.7387476","density: 4.886524e-03<br />Petal.Width: 0.7434442","density: 5.830685e-03<br />Petal.Width: 0.7481409","density: 6.963675e-03<br />Petal.Width: 0.7528376","density: 8.300753e-03<br />Petal.Width: 0.7575342","density: 9.842791e-03<br />Petal.Width: 0.7622309","density: 1.161159e-02<br />Petal.Width: 0.7669276","density: 1.362997e-02<br />Petal.Width: 0.7716243","density: 1.604157e-02<br />Petal.Width: 0.7763209","density: 1.879175e-02<br />Petal.Width: 0.7810176","density: 2.190711e-02<br />Petal.Width: 0.7857143","density: 2.541918e-02<br />Petal.Width: 0.7904110","density: 2.941113e-02<br />Petal.Width: 0.7951076","density: 3.402606e-02<br />Petal.Width: 0.7998043","density: 3.918451e-02<br />Petal.Width: 0.8045010","density: 4.492444e-02<br />Petal.Width: 0.8091977","density: 5.128421e-02<br />Petal.Width: 0.8138943","density: 5.846434e-02<br />Petal.Width: 0.8185910","density: 6.649071e-02<br />Petal.Width: 0.8232877","density: 7.529885e-02<br />Petal.Width: 0.8279843","density: 8.492569e-02<br />Petal.Width: 0.8326810","density: 9.540739e-02<br />Petal.Width: 0.8373777","density: 1.071008e-01<br />Petal.Width: 0.8420744","density: 1.197812e-01<br />Petal.Width: 0.8467710","density: 1.334405e-01<br />Petal.Width: 0.8514677","density: 1.481002e-01<br />Petal.Width: 0.8561644","density: 1.638637e-01<br />Petal.Width: 0.8608611","density: 1.809657e-01<br />Petal.Width: 0.8655577","density: 1.991070e-01<br />Petal.Width: 0.8702544","density: 2.182808e-01<br />Petal.Width: 0.8749511","density: 2.384760e-01<br />Petal.Width: 0.8796477","density: 2.598658e-01<br />Petal.Width: 0.8843444","density: 2.823692e-01<br />Petal.Width: 0.8890411","density: 3.057682e-01<br />Petal.Width: 0.8937378","density: 3.300127e-01<br />Petal.Width: 0.8984344","density: 3.550482e-01<br />Petal.Width: 0.9031311","density: 3.809950e-01<br />Petal.Width: 0.9078278","density: 4.075455e-01<br />Petal.Width: 0.9125245","density: 4.345727e-01<br />Petal.Width: 0.9172211","density: 4.619856e-01<br />Petal.Width: 0.9219178","density: 4.896961e-01<br />Petal.Width: 0.9266145","density: 5.175598e-01<br />Petal.Width: 0.9313112","density: 5.453778e-01<br />Petal.Width: 0.9360078","density: 5.730384e-01<br />Petal.Width: 0.9407045","density: 6.004296e-01<br />Petal.Width: 0.9454012","density: 6.273334e-01<br />Petal.Width: 0.9500978","density: 6.535303e-01<br />Petal.Width: 0.9547945","density: 6.789954e-01<br />Petal.Width: 0.9594912","density: 7.036264e-01<br />Petal.Width: 0.9641879","density: 7.273241e-01<br />Petal.Width: 0.9688845","density: 7.496410e-01<br />Petal.Width: 0.9735812","density: 7.706703e-01<br />Petal.Width: 0.9782779","density: 7.904313e-01<br />Petal.Width: 0.9829746","density: 8.088663e-01<br />Petal.Width: 0.9876712","density: 8.258684e-01<br />Petal.Width: 0.9923679","density: 8.409579e-01<br />Petal.Width: 0.9970646","density: 8.545888e-01<br />Petal.Width: 1.0017613","density: 8.667616e-01<br />Petal.Width: 1.0064579","density: 8.774847e-01<br />Petal.Width: 1.0111546","density: 8.865617e-01<br />Petal.Width: 1.0158513","density: 8.939782e-01<br />Petal.Width: 1.0205479","density: 9.001041e-01<br />Petal.Width: 1.0252446","density: 9.050057e-01<br />Petal.Width: 1.0299413","density: 9.087558e-01<br />Petal.Width: 1.0346380","density: 9.111947e-01<br />Petal.Width: 1.0393346","density: 9.126695e-01<br />Petal.Width: 1.0440313","density: 9.133914e-01<br />Petal.Width: 1.0487280","density: 9.134714e-01<br />Petal.Width: 1.0534247","density: 9.130182e-01<br />Petal.Width: 1.0581213","density: 9.121133e-01<br />Petal.Width: 1.0628180","density: 9.110678e-01<br />Petal.Width: 1.0675147","density: 9.100054e-01<br />Petal.Width: 1.0722114","density: 9.090495e-01<br />Petal.Width: 1.0769080","density: 9.083828e-01<br />Petal.Width: 1.0816047","density: 9.082668e-01<br />Petal.Width: 1.0863014","density: 9.087525e-01<br />Petal.Width: 1.0909980","density: 9.099484e-01<br />Petal.Width: 1.0956947","density: 9.119597e-01<br />Petal.Width: 1.1003914","density: 9.151635e-01<br />Petal.Width: 1.1050881","density: 9.195959e-01<br />Petal.Width: 1.1097847","density: 9.252175e-01<br />Petal.Width: 1.1144814","density: 9.321002e-01<br />Petal.Width: 1.1191781","density: 9.403111e-01<br />Petal.Width: 1.1238748","density: 9.504589e-01<br />Petal.Width: 1.1285714","density: 9.620980e-01<br />Petal.Width: 1.1332681","density: 9.752520e-01<br />Petal.Width: 1.1379648","density: 9.899458e-01<br />Petal.Width: 1.1426614","density: 1.006375e+00<br />Petal.Width: 1.1473581","density: 1.024777e+00<br />Petal.Width: 1.1520548","density: 1.044731e+00<br />Petal.Width: 1.1567515","density: 1.066218e+00<br />Petal.Width: 1.1614481","density: 1.089211e+00<br />Petal.Width: 1.1661448","density: 1.113994e+00<br />Petal.Width: 1.1708415","density: 1.140351e+00<br />Petal.Width: 1.1755382","density: 1.168009e+00<br />Petal.Width: 1.1802348","density: 1.196896e+00<br />Petal.Width: 1.1849315","density: 1.226936e+00<br />Petal.Width: 1.1896282","density: 1.258344e+00<br />Petal.Width: 1.1943249","density: 1.290632e+00<br />Petal.Width: 1.1990215","density: 1.323662e+00<br />Petal.Width: 1.2037182","density: 1.357319e+00<br />Petal.Width: 1.2084149","density: 1.391508e+00<br />Petal.Width: 1.2131115","density: 1.426060e+00<br />Petal.Width: 1.2178082","density: 1.460698e+00<br />Petal.Width: 1.2225049","density: 1.495282e+00<br />Petal.Width: 1.2272016","density: 1.529671e+00<br />Petal.Width: 1.2318982","density: 1.563592e+00<br />Petal.Width: 1.2365949","density: 1.596796e+00<br />Petal.Width: 1.2412916","density: 1.629205e+00<br />Petal.Width: 1.2459883","density: 1.660680e+00<br />Petal.Width: 1.2506849","density: 1.691085e+00<br />Petal.Width: 1.2553816","density: 1.719813e+00<br />Petal.Width: 1.2600783","density: 1.747016e+00<br />Petal.Width: 1.2647750","density: 1.772649e+00<br />Petal.Width: 1.2694716","density: 1.796615e+00<br />Petal.Width: 1.2741683","density: 1.818690e+00<br />Petal.Width: 1.2788650","density: 1.838309e+00<br />Petal.Width: 1.2835616","density: 1.855964e+00<br />Petal.Width: 1.2882583","density: 1.871619e+00<br />Petal.Width: 1.2929550","density: 1.885247e+00<br />Petal.Width: 1.2976517","density: 1.896452e+00<br />Petal.Width: 1.3023483","density: 1.905250e+00<br />Petal.Width: 1.3070450","density: 1.912058e+00<br />Petal.Width: 1.3117417","density: 1.916920e+00<br />Petal.Width: 1.3164384","density: 1.919888e+00<br />Petal.Width: 1.3211350","density: 1.920518e+00<br />Petal.Width: 1.3258317","density: 1.919372e+00<br />Petal.Width: 1.3305284","density: 1.916673e+00<br />Petal.Width: 1.3352250","density: 1.912525e+00<br />Petal.Width: 1.3399217","density: 1.906979e+00<br />Petal.Width: 1.3446184","density: 1.899907e+00<br />Petal.Width: 1.3493151","density: 1.891864e+00<br />Petal.Width: 1.3540117","density: 1.882968e+00<br />Petal.Width: 1.3587084","density: 1.873337e+00<br />Petal.Width: 1.3634051","density: 1.863018e+00<br />Petal.Width: 1.3681018","density: 1.852196e+00<br />Petal.Width: 1.3727984","density: 1.841089e+00<br />Petal.Width: 1.3774951","density: 1.829787e+00<br />Petal.Width: 1.3821918","density: 1.818374e+00<br />Petal.Width: 1.3868885","density: 1.806953e+00<br />Petal.Width: 1.3915851","density: 1.795622e+00<br />Petal.Width: 1.3962818","density: 1.784416e+00<br />Petal.Width: 1.4009785","density: 1.773363e+00<br />Petal.Width: 1.4056751","density: 1.762490e+00<br />Petal.Width: 1.4103718","density: 1.751854e+00<br />Petal.Width: 1.4150685","density: 1.741369e+00<br />Petal.Width: 1.4197652","density: 1.731004e+00<br />Petal.Width: 1.4244618","density: 1.720721e+00<br />Petal.Width: 1.4291585","density: 1.710463e+00<br />Petal.Width: 1.4338552","density: 1.700115e+00<br />Petal.Width: 1.4385519","density: 1.689597e+00<br />Petal.Width: 1.4432485","density: 1.678832e+00<br />Petal.Width: 1.4479452","density: 1.667741e+00<br />Petal.Width: 1.4526419","density: 1.656091e+00<br />Petal.Width: 1.4573386","density: 1.643796e+00<br />Petal.Width: 1.4620352","density: 1.630824e+00<br />Petal.Width: 1.4667319","density: 1.617094e+00<br />Petal.Width: 1.4714286","density: 1.602522e+00<br />Petal.Width: 1.4761252","density: 1.586647e+00<br />Petal.Width: 1.4808219","density: 1.569720e+00<br />Petal.Width: 1.4855186","density: 1.551691e+00<br />Petal.Width: 1.4902153","density: 1.532513e+00<br />Petal.Width: 1.4949119","density: 1.511999e+00<br />Petal.Width: 1.4996086","density: 1.489947e+00<br />Petal.Width: 1.5043053","density: 1.466655e+00<br />Petal.Width: 1.5090020","density: 1.442130e+00<br />Petal.Width: 1.5136986","density: 1.416382e+00<br />Petal.Width: 1.5183953","density: 1.389160e+00<br />Petal.Width: 1.5230920","density: 1.360658e+00<br />Petal.Width: 1.5277886","density: 1.331089e+00<br />Petal.Width: 1.5324853","density: 1.300514e+00<br />Petal.Width: 1.5371820","density: 1.268996e+00<br />Petal.Width: 1.5418787","density: 1.236359e+00<br />Petal.Width: 1.5465753","density: 1.203016e+00<br />Petal.Width: 1.5512720","density: 1.169071e+00<br />Petal.Width: 1.5559687","density: 1.134614e+00<br />Petal.Width: 1.5606654","density: 1.099714e+00<br />Petal.Width: 1.5653620","density: 1.064495e+00<br />Petal.Width: 1.5700587","density: 1.029161e+00<br />Petal.Width: 1.5747554","density: 9.938075e-01<br />Petal.Width: 1.5794521","density: 9.585262e-01<br />Petal.Width: 1.5841487","density: 9.234805e-01<br />Petal.Width: 1.5888454","density: 8.888144e-01<br />Petal.Width: 1.5935421","density: 8.545750e-01<br />Petal.Width: 1.5982387","density: 8.208360e-01<br />Petal.Width: 1.6029354","density: 7.876681e-01<br />Petal.Width: 1.6076321","density: 7.553913e-01<br />Petal.Width: 1.6123288","density: 7.238991e-01<br />Petal.Width: 1.6170254","density: 6.932113e-01<br />Petal.Width: 1.6217221","density: 6.633702e-01<br />Petal.Width: 1.6264188","density: 6.344877e-01<br />Petal.Width: 1.6311155","density: 6.068154e-01<br />Petal.Width: 1.6358121","density: 5.801094e-01<br />Petal.Width: 1.6405088","density: 5.543838e-01<br />Petal.Width: 1.6452055","density: 5.296500e-01<br />Petal.Width: 1.6499022","density: 5.061116e-01<br />Petal.Width: 1.6545988","density: 4.837495e-01<br />Petal.Width: 1.6592955","density: 4.623788e-01<br />Petal.Width: 1.6639922","density: 4.419896e-01<br />Petal.Width: 1.6686888","density: 4.225698e-01<br />Petal.Width: 1.6733855","density: 4.043903e-01<br />Petal.Width: 1.6780822","density: 3.871783e-01<br />Petal.Width: 1.6827789","density: 3.708530e-01<br />Petal.Width: 1.6874755","density: 3.553891e-01<br />Petal.Width: 1.6921722","density: 3.408046e-01<br />Petal.Width: 1.6968689","density: 3.272536e-01<br />Petal.Width: 1.7015656","density: 3.144416e-01<br />Petal.Width: 1.7062622","density: 3.023361e-01<br />Petal.Width: 1.7109589","density: 2.909042e-01<br />Petal.Width: 1.7156556","density: 2.802126e-01<br />Petal.Width: 1.7203523","density: 2.702117e-01<br />Petal.Width: 1.7250489","density: 2.607419e-01<br />Petal.Width: 1.7297456","density: 2.517689e-01<br />Petal.Width: 1.7344423","density: 2.432586e-01<br />Petal.Width: 1.7391389","density: 2.352823e-01<br />Petal.Width: 1.7438356","density: 2.276972e-01<br />Petal.Width: 1.7485323","density: 2.204385e-01<br />Petal.Width: 1.7532290","density: 2.134758e-01<br />Petal.Width: 1.7579256","density: 2.067871e-01<br />Petal.Width: 1.7626223","density: 2.003934e-01<br />Petal.Width: 1.7673190","density: 1.941811e-01<br />Petal.Width: 1.7720157","density: 1.881271e-01<br />Petal.Width: 1.7767123","density: 1.822094e-01<br />Petal.Width: 1.7814090","density: 1.764196e-01<br />Petal.Width: 1.7861057","density: 1.707312e-01<br />Petal.Width: 1.7908023","density: 1.651063e-01<br />Petal.Width: 1.7954990","density: 1.595325e-01<br />Petal.Width: 1.8001957","density: 1.539988e-01<br />Petal.Width: 1.8048924","density: 1.485001e-01<br />Petal.Width: 1.8095890","density: 1.430218e-01<br />Petal.Width: 1.8142857","density: 1.375585e-01<br />Petal.Width: 1.8189824","density: 1.321086e-01<br />Petal.Width: 1.8236791","density: 1.266719e-01<br />Petal.Width: 1.8283757","density: 1.212570e-01<br />Petal.Width: 1.8330724","density: 1.158642e-01<br />Petal.Width: 1.8377691","density: 1.104995e-01<br />Petal.Width: 1.8424658","density: 1.051689e-01<br />Petal.Width: 1.8471624","density: 9.988670e-02<br />Petal.Width: 1.8518591","density: 9.467396e-02<br />Petal.Width: 1.8565558","density: 8.952951e-02<br />Petal.Width: 1.8612524","density: 8.446255e-02<br />Petal.Width: 1.8659491","density: 7.948241e-02<br />Petal.Width: 1.8706458","density: 7.462679e-02<br />Petal.Width: 1.8753425","density: 6.989966e-02<br />Petal.Width: 1.8800391","density: 6.529694e-02<br />Petal.Width: 1.8847358","density: 6.082667e-02<br />Petal.Width: 1.8894325","density: 5.649653e-02<br />Petal.Width: 1.8941292","density: 5.237208e-02<br />Petal.Width: 1.8988258","density: 4.840849e-02<br />Petal.Width: 1.9035225","density: 4.460890e-02<br />Petal.Width: 1.9082192","density: 4.097708e-02<br />Petal.Width: 1.9129159","density: 3.753530e-02<br />Petal.Width: 1.9176125","density: 3.431335e-02<br />Petal.Width: 1.9223092","density: 3.126601e-02<br />Petal.Width: 1.9270059","density: 2.839269e-02<br />Petal.Width: 1.9317025","density: 2.569229e-02<br />Petal.Width: 1.9363992","density: 2.320063e-02<br />Petal.Width: 1.9410959","density: 2.089866e-02<br />Petal.Width: 1.9457926","density: 1.875738e-02<br />Petal.Width: 1.9504892","density: 1.677243e-02<br />Petal.Width: 1.9551859","density: 1.493915e-02<br />Petal.Width: 1.9598826","density: 1.329867e-02<br />Petal.Width: 1.9645793","density: 1.179663e-02<br />Petal.Width: 1.9692759","density: 1.042281e-02<br />Petal.Width: 1.9739726","density: 9.171132e-03<br />Petal.Width: 1.9786693","density: 8.044650e-03<br />Petal.Width: 1.9833659","density: 7.053326e-03<br />Petal.Width: 1.9880626","density: 6.158261e-03<br />Petal.Width: 1.9927593","density: 5.353436e-03<br />Petal.Width: 1.9974560","density: 4.632919e-03<br />Petal.Width: 2.0021526","density: 4.004752e-03<br />Petal.Width: 2.0068493","density: 3.455956e-03<br />Petal.Width: 2.0115460","density: 2.968645e-03<br />Petal.Width: 2.0162427","density: 2.537956e-03<br />Petal.Width: 2.0209393","density: 2.159183e-03<br />Petal.Width: 2.0256360","density: 1.840336e-03<br />Petal.Width: 2.0303327","density: 1.562550e-03<br />Petal.Width: 2.0350294","density: 1.320011e-03<br />Petal.Width: 2.0397260","density: 1.109370e-03<br />Petal.Width: 2.0444227","density: 9.290215e-04<br />Petal.Width: 2.0491194","density: 7.800072e-04<br />Petal.Width: 2.0538160","density: 6.513644e-04<br />Petal.Width: 2.0585127","density: 5.409512e-04<br />Petal.Width: 2.0632094","density: 4.467485e-04<br />Petal.Width: 2.0679061","density: 3.689522e-04<br />Petal.Width: 2.0726027","density: 3.047188e-04<br />Petal.Width: 2.0772994","density: 2.501839e-04<br />Petal.Width: 2.0819961","density: 2.041859e-04<br />Petal.Width: 2.0866928","density: 1.656450e-04<br />Petal.Width: 2.0913894","density: 1.350345e-04<br />Petal.Width: 2.0960861","density: 1.096682e-04<br />Petal.Width: 2.1007828","density: 8.849548e-05<br />Petal.Width: 2.1054795","density: 7.095202e-05<br />Petal.Width: 2.1101761","density: 5.662939e-05<br />Petal.Width: 2.1148728","density: 4.552789e-05<br />Petal.Width: 2.1195695","density: 3.634674e-05<br />Petal.Width: 2.1242661","density: 2.881585e-05<br />Petal.Width: 2.1289628","density: 2.268822e-05<br />Petal.Width: 2.1336595","density: 1.788289e-05<br />Petal.Width: 2.1383562","density: 1.413592e-05<br />Petal.Width: 2.1430528","density: 1.108932e-05<br />Petal.Width: 2.1477495","density: 8.634572e-06<br />Petal.Width: 2.1524462","density: 6.674003e-06<br />Petal.Width: 2.1571429","density: 5.201001e-06<br />Petal.Width: 2.1618395","density: 4.040699e-06<br />Petal.Width: 2.1665362","density: 3.113640e-06<br />Petal.Width: 2.1712329","density: 2.380237e-06<br />Petal.Width: 2.1759295","density: 1.808166e-06<br />Petal.Width: 2.1806262","density: 1.392767e-06<br />Petal.Width: 2.1853229","density: 1.063063e-06<br />Petal.Width: 2.1900196","density: 8.043385e-07<br />Petal.Width: 2.1947162","density: 6.034686e-07<br />Petal.Width: 2.1994129","density: 4.535789e-07<br />Petal.Width: 2.2041096","density: 3.433259e-07<br />Petal.Width: 2.2088063","density: 2.573488e-07<br />Petal.Width: 2.2135029","density: 1.911211e-07<br />Petal.Width: 2.2181996","density: 1.406827e-07<br />Petal.Width: 2.2228963","density: 1.047594e-07<br />Petal.Width: 2.2275930","density: 7.788746e-08<br />Petal.Width: 2.2322896","density: 5.731166e-08<br />Petal.Width: 2.2369863","density: 4.176163e-08<br />Petal.Width: 2.2416830","density: 3.015362e-08<br />Petal.Width: 2.2463796","density: 2.227196e-08<br />Petal.Width: 2.2510763","density: 1.625772e-08<br />Petal.Width: 2.2557730","density: 1.173876e-08<br />Petal.Width: 2.2604697","density: 8.389801e-09<br />Petal.Width: 2.2651663","density: 6.007830e-09<br />Petal.Width: 2.2698630","density: 4.357593e-09<br />Petal.Width: 2.2745597","density: 3.121640e-09<br />Petal.Width: 2.2792564","density: 2.210880e-09<br />Petal.Width: 2.2839530","density: 1.549327e-09<br />Petal.Width: 2.2886497","density: 1.101875e-09<br />Petal.Width: 2.2933464","density: 7.844304e-10<br />Petal.Width: 2.2980431","density: 5.512381e-10<br />Petal.Width: 2.3027397","density: 3.828079e-10<br />Petal.Width: 2.3074364","density: 2.629474e-10<br />Petal.Width: 2.3121331","density: 1.859839e-10<br />Petal.Width: 2.3168297","density: 1.298908e-10<br />Petal.Width: 2.3215264","density: 8.950208e-11<br />Petal.Width: 2.3262231","density: 6.092326e-11<br />Petal.Width: 2.3309198","density: 4.151991e-11<br />Petal.Width: 2.3356164","density: 2.888253e-11<br />Petal.Width: 2.3403131","density: 1.977958e-11<br />Petal.Width: 2.3450098","density: 1.335897e-11<br />Petal.Width: 2.3497065","density: 8.910176e-12<br />Petal.Width: 2.3544031","density: 6.048318e-12<br />Petal.Width: 2.3590998","density: 4.125633e-12<br />Petal.Width: 2.3637965","density: 2.769167e-12<br />Petal.Width: 2.3684932","density: 1.832484e-12<br />Petal.Width: 2.3731898","density: 1.197328e-12<br />Petal.Width: 2.3778865","density: 8.105254e-13<br />Petal.Width: 2.3825832","density: 5.418047e-13<br />Petal.Width: 2.3872798","density: 3.562713e-13<br />Petal.Width: 2.3919765","density: 2.309110e-13<br />Petal.Width: 2.3966732","density: 1.496124e-13<br />Petal.Width: 2.4013699","density: 9.970281e-14<br />Petal.Width: 2.4060665","density: 6.523764e-14<br />Petal.Width: 2.4107632","density: 4.200972e-14<br />Petal.Width: 2.4154599","density: 2.658568e-14<br />Petal.Width: 2.4201566","density: 1.719773e-14<br />Petal.Width: 2.4248532","density: 1.125504e-14<br />Petal.Width: 2.4295499","density: 7.203531e-15<br />Petal.Width: 2.4342466","density: 4.521480e-15<br />Petal.Width: 2.4389432","density: 2.820250e-15<br />Petal.Width: 2.4436399","density: 1.832333e-15<br />Petal.Width: 2.4483366","density: 1.131972e-15<br />Petal.Width: 2.4530333","density: 6.745888e-16<br />Petal.Width: 2.4577299","density: 3.819960e-16<br />Petal.Width: 2.4624266","density: 1.145487e-16<br />Petal.Width: 2.4671233","density: 1.460397e-16<br />Petal.Width: 2.4718200","density: 7.302836e-17<br />Petal.Width: 2.4765166","density: 0.000000e+00<br />Petal.Width: 2.4812133","density: 0.000000e+00<br />Petal.Width: 2.4859100","density: 0.000000e+00<br />Petal.Width: 2.4906067","density: 0.000000e+00<br />Petal.Width: 2.4953033","density: 0.000000e+00<br />Petal.Width: 2.5000000","density: 0.000000e+00<br />Petal.Width: 2.5000000","density: 0.000000e+00<br />Petal.Width: 2.4953033","density: 0.000000e+00<br />Petal.Width: 2.4906067","density: 0.000000e+00<br />Petal.Width: 2.4859100","density: 0.000000e+00<br />Petal.Width: 2.4812133","density: 7.302836e-17<br />Petal.Width: 2.4765166","density: 1.460397e-16<br />Petal.Width: 2.4718200","density: 1.145487e-16<br />Petal.Width: 2.4671233","density: 3.819960e-16<br />Petal.Width: 2.4624266","density: 6.745888e-16<br />Petal.Width: 2.4577299","density: 1.131972e-15<br />Petal.Width: 2.4530333","density: 1.832333e-15<br />Petal.Width: 2.4483366","density: 2.820250e-15<br />Petal.Width: 2.4436399","density: 4.521480e-15<br />Petal.Width: 2.4389432","density: 7.203531e-15<br />Petal.Width: 2.4342466","density: 1.125504e-14<br />Petal.Width: 2.4295499","density: 1.719773e-14<br />Petal.Width: 2.4248532","density: 2.658568e-14<br />Petal.Width: 2.4201566","density: 4.200972e-14<br />Petal.Width: 2.4154599","density: 6.523764e-14<br />Petal.Width: 2.4107632","density: 9.970281e-14<br />Petal.Width: 2.4060665","density: 1.496124e-13<br />Petal.Width: 2.4013699","density: 2.309110e-13<br />Petal.Width: 2.3966732","density: 3.562713e-13<br />Petal.Width: 2.3919765","density: 5.418047e-13<br />Petal.Width: 2.3872798","density: 8.105254e-13<br />Petal.Width: 2.3825832","density: 1.197328e-12<br />Petal.Width: 2.3778865","density: 1.832484e-12<br />Petal.Width: 2.3731898","density: 2.769167e-12<br />Petal.Width: 2.3684932","density: 4.125633e-12<br />Petal.Width: 2.3637965","density: 6.048318e-12<br />Petal.Width: 2.3590998","density: 8.910176e-12<br />Petal.Width: 2.3544031","density: 1.335897e-11<br />Petal.Width: 2.3497065","density: 1.977958e-11<br />Petal.Width: 2.3450098","density: 2.888253e-11<br />Petal.Width: 2.3403131","density: 4.151991e-11<br />Petal.Width: 2.3356164","density: 6.092326e-11<br />Petal.Width: 2.3309198","density: 8.950208e-11<br />Petal.Width: 2.3262231","density: 1.298908e-10<br />Petal.Width: 2.3215264","density: 1.859839e-10<br />Petal.Width: 2.3168297","density: 2.629474e-10<br />Petal.Width: 2.3121331","density: 3.828079e-10<br />Petal.Width: 2.3074364","density: 5.512381e-10<br />Petal.Width: 2.3027397","density: 7.844304e-10<br />Petal.Width: 2.2980431","density: 1.101875e-09<br />Petal.Width: 2.2933464","density: 1.549327e-09<br />Petal.Width: 2.2886497","density: 2.210880e-09<br />Petal.Width: 2.2839530","density: 3.121640e-09<br />Petal.Width: 2.2792564","density: 4.357593e-09<br />Petal.Width: 2.2745597","density: 6.007830e-09<br />Petal.Width: 2.2698630","density: 8.389801e-09<br />Petal.Width: 2.2651663","density: 1.173876e-08<br />Petal.Width: 2.2604697","density: 1.625772e-08<br />Petal.Width: 2.2557730","density: 2.227196e-08<br />Petal.Width: 2.2510763","density: 3.015362e-08<br />Petal.Width: 2.2463796","density: 4.176163e-08<br />Petal.Width: 2.2416830","density: 5.731166e-08<br />Petal.Width: 2.2369863","density: 7.788746e-08<br />Petal.Width: 2.2322896","density: 1.047594e-07<br />Petal.Width: 2.2275930","density: 1.406827e-07<br />Petal.Width: 2.2228963","density: 1.911211e-07<br />Petal.Width: 2.2181996","density: 2.573488e-07<br />Petal.Width: 2.2135029","density: 3.433259e-07<br />Petal.Width: 2.2088063","density: 4.535789e-07<br />Petal.Width: 2.2041096","density: 6.034686e-07<br />Petal.Width: 2.1994129","density: 8.043385e-07<br />Petal.Width: 2.1947162","density: 1.063063e-06<br />Petal.Width: 2.1900196","density: 1.392767e-06<br />Petal.Width: 2.1853229","density: 1.808166e-06<br />Petal.Width: 2.1806262","density: 2.380237e-06<br />Petal.Width: 2.1759295","density: 3.113640e-06<br />Petal.Width: 2.1712329","density: 4.040699e-06<br />Petal.Width: 2.1665362","density: 5.201001e-06<br />Petal.Width: 2.1618395","density: 6.674003e-06<br />Petal.Width: 2.1571429","density: 8.634572e-06<br />Petal.Width: 2.1524462","density: 1.108932e-05<br />Petal.Width: 2.1477495","density: 1.413592e-05<br />Petal.Width: 2.1430528","density: 1.788289e-05<br />Petal.Width: 2.1383562","density: 2.268822e-05<br />Petal.Width: 2.1336595","density: 2.881585e-05<br />Petal.Width: 2.1289628","density: 3.634674e-05<br />Petal.Width: 2.1242661","density: 4.552789e-05<br />Petal.Width: 2.1195695","density: 5.662939e-05<br />Petal.Width: 2.1148728","density: 7.095202e-05<br />Petal.Width: 2.1101761","density: 8.849548e-05<br />Petal.Width: 2.1054795","density: 1.096682e-04<br />Petal.Width: 2.1007828","density: 1.350345e-04<br />Petal.Width: 2.0960861","density: 1.656450e-04<br />Petal.Width: 2.0913894","density: 2.041859e-04<br />Petal.Width: 2.0866928","density: 2.501839e-04<br />Petal.Width: 2.0819961","density: 3.047188e-04<br />Petal.Width: 2.0772994","density: 3.689522e-04<br />Petal.Width: 2.0726027","density: 4.467485e-04<br />Petal.Width: 2.0679061","density: 5.409512e-04<br />Petal.Width: 2.0632094","density: 6.513644e-04<br />Petal.Width: 2.0585127","density: 7.800072e-04<br />Petal.Width: 2.0538160","density: 9.290215e-04<br />Petal.Width: 2.0491194","density: 1.109370e-03<br />Petal.Width: 2.0444227","density: 1.320011e-03<br />Petal.Width: 2.0397260","density: 1.562550e-03<br />Petal.Width: 2.0350294","density: 1.840336e-03<br />Petal.Width: 2.0303327","density: 2.159183e-03<br />Petal.Width: 2.0256360","density: 2.537956e-03<br />Petal.Width: 2.0209393","density: 2.968645e-03<br />Petal.Width: 2.0162427","density: 3.455956e-03<br />Petal.Width: 2.0115460","density: 4.004752e-03<br />Petal.Width: 2.0068493","density: 4.632919e-03<br />Petal.Width: 2.0021526","density: 5.353436e-03<br />Petal.Width: 1.9974560","density: 6.158261e-03<br />Petal.Width: 1.9927593","density: 7.053326e-03<br />Petal.Width: 1.9880626","density: 8.044650e-03<br />Petal.Width: 1.9833659","density: 9.171132e-03<br />Petal.Width: 1.9786693","density: 1.042281e-02<br />Petal.Width: 1.9739726","density: 1.179663e-02<br />Petal.Width: 1.9692759","density: 1.329867e-02<br />Petal.Width: 1.9645793","density: 1.493915e-02<br />Petal.Width: 1.9598826","density: 1.677243e-02<br />Petal.Width: 1.9551859","density: 1.875738e-02<br />Petal.Width: 1.9504892","density: 2.089866e-02<br />Petal.Width: 1.9457926","density: 2.320063e-02<br />Petal.Width: 1.9410959","density: 2.569229e-02<br />Petal.Width: 1.9363992","density: 2.839269e-02<br />Petal.Width: 1.9317025","density: 3.126601e-02<br />Petal.Width: 1.9270059","density: 3.431335e-02<br />Petal.Width: 1.9223092","density: 3.753530e-02<br />Petal.Width: 1.9176125","density: 4.097708e-02<br />Petal.Width: 1.9129159","density: 4.460890e-02<br />Petal.Width: 1.9082192","density: 4.840849e-02<br />Petal.Width: 1.9035225","density: 5.237208e-02<br />Petal.Width: 1.8988258","density: 5.649653e-02<br />Petal.Width: 1.8941292","density: 6.082667e-02<br />Petal.Width: 1.8894325","density: 6.529694e-02<br />Petal.Width: 1.8847358","density: 6.989966e-02<br />Petal.Width: 1.8800391","density: 7.462679e-02<br />Petal.Width: 1.8753425","density: 7.948241e-02<br />Petal.Width: 1.8706458","density: 8.446255e-02<br />Petal.Width: 1.8659491","density: 8.952951e-02<br />Petal.Width: 1.8612524","density: 9.467396e-02<br />Petal.Width: 1.8565558","density: 9.988670e-02<br />Petal.Width: 1.8518591","density: 1.051689e-01<br />Petal.Width: 1.8471624","density: 1.104995e-01<br />Petal.Width: 1.8424658","density: 1.158642e-01<br />Petal.Width: 1.8377691","density: 1.212570e-01<br />Petal.Width: 1.8330724","density: 1.266719e-01<br />Petal.Width: 1.8283757","density: 1.321086e-01<br />Petal.Width: 1.8236791","density: 1.375585e-01<br />Petal.Width: 1.8189824","density: 1.430218e-01<br />Petal.Width: 1.8142857","density: 1.485001e-01<br />Petal.Width: 1.8095890","density: 1.539988e-01<br />Petal.Width: 1.8048924","density: 1.595325e-01<br />Petal.Width: 1.8001957","density: 1.651063e-01<br />Petal.Width: 1.7954990","density: 1.707312e-01<br />Petal.Width: 1.7908023","density: 1.764196e-01<br />Petal.Width: 1.7861057","density: 1.822094e-01<br />Petal.Width: 1.7814090","density: 1.881271e-01<br />Petal.Width: 1.7767123","density: 1.941811e-01<br />Petal.Width: 1.7720157","density: 2.003934e-01<br />Petal.Width: 1.7673190","density: 2.067871e-01<br />Petal.Width: 1.7626223","density: 2.134758e-01<br />Petal.Width: 1.7579256","density: 2.204385e-01<br />Petal.Width: 1.7532290","density: 2.276972e-01<br />Petal.Width: 1.7485323","density: 2.352823e-01<br />Petal.Width: 1.7438356","density: 2.432586e-01<br />Petal.Width: 1.7391389","density: 2.517689e-01<br />Petal.Width: 1.7344423","density: 2.607419e-01<br />Petal.Width: 1.7297456","density: 2.702117e-01<br />Petal.Width: 1.7250489","density: 2.802126e-01<br />Petal.Width: 1.7203523","density: 2.909042e-01<br />Petal.Width: 1.7156556","density: 3.023361e-01<br />Petal.Width: 1.7109589","density: 3.144416e-01<br />Petal.Width: 1.7062622","density: 3.272536e-01<br />Petal.Width: 1.7015656","density: 3.408046e-01<br />Petal.Width: 1.6968689","density: 3.553891e-01<br />Petal.Width: 1.6921722","density: 3.708530e-01<br />Petal.Width: 1.6874755","density: 3.871783e-01<br />Petal.Width: 1.6827789","density: 4.043903e-01<br />Petal.Width: 1.6780822","density: 4.225698e-01<br />Petal.Width: 1.6733855","density: 4.419896e-01<br />Petal.Width: 1.6686888","density: 4.623788e-01<br />Petal.Width: 1.6639922","density: 4.837495e-01<br />Petal.Width: 1.6592955","density: 5.061116e-01<br />Petal.Width: 1.6545988","density: 5.296500e-01<br />Petal.Width: 1.6499022","density: 5.543838e-01<br />Petal.Width: 1.6452055","density: 5.801094e-01<br />Petal.Width: 1.6405088","density: 6.068154e-01<br />Petal.Width: 1.6358121","density: 6.344877e-01<br />Petal.Width: 1.6311155","density: 6.633702e-01<br />Petal.Width: 1.6264188","density: 6.932113e-01<br />Petal.Width: 1.6217221","density: 7.238991e-01<br />Petal.Width: 1.6170254","density: 7.553913e-01<br />Petal.Width: 1.6123288","density: 7.876681e-01<br />Petal.Width: 1.6076321","density: 8.208360e-01<br />Petal.Width: 1.6029354","density: 8.545750e-01<br />Petal.Width: 1.5982387","density: 8.888144e-01<br />Petal.Width: 1.5935421","density: 9.234805e-01<br />Petal.Width: 1.5888454","density: 9.585262e-01<br />Petal.Width: 1.5841487","density: 9.938075e-01<br />Petal.Width: 1.5794521","density: 1.029161e+00<br />Petal.Width: 1.5747554","density: 1.064495e+00<br />Petal.Width: 1.5700587","density: 1.099714e+00<br />Petal.Width: 1.5653620","density: 1.134614e+00<br />Petal.Width: 1.5606654","density: 1.169071e+00<br />Petal.Width: 1.5559687","density: 1.203016e+00<br />Petal.Width: 1.5512720","density: 1.236359e+00<br />Petal.Width: 1.5465753","density: 1.268996e+00<br />Petal.Width: 1.5418787","density: 1.300514e+00<br />Petal.Width: 1.5371820","density: 1.331089e+00<br />Petal.Width: 1.5324853","density: 1.360658e+00<br />Petal.Width: 1.5277886","density: 1.389160e+00<br />Petal.Width: 1.5230920","density: 1.416382e+00<br />Petal.Width: 1.5183953","density: 1.442130e+00<br />Petal.Width: 1.5136986","density: 1.466655e+00<br />Petal.Width: 1.5090020","density: 1.489947e+00<br />Petal.Width: 1.5043053","density: 1.511999e+00<br />Petal.Width: 1.4996086","density: 1.532513e+00<br />Petal.Width: 1.4949119","density: 1.551691e+00<br />Petal.Width: 1.4902153","density: 1.569720e+00<br />Petal.Width: 1.4855186","density: 1.586647e+00<br />Petal.Width: 1.4808219","density: 1.602522e+00<br />Petal.Width: 1.4761252","density: 1.617094e+00<br />Petal.Width: 1.4714286","density: 1.630824e+00<br />Petal.Width: 1.4667319","density: 1.643796e+00<br />Petal.Width: 1.4620352","density: 1.656091e+00<br />Petal.Width: 1.4573386","density: 1.667741e+00<br />Petal.Width: 1.4526419","density: 1.678832e+00<br />Petal.Width: 1.4479452","density: 1.689597e+00<br />Petal.Width: 1.4432485","density: 1.700115e+00<br />Petal.Width: 1.4385519","density: 1.710463e+00<br />Petal.Width: 1.4338552","density: 1.720721e+00<br />Petal.Width: 1.4291585","density: 1.731004e+00<br />Petal.Width: 1.4244618","density: 1.741369e+00<br />Petal.Width: 1.4197652","density: 1.751854e+00<br />Petal.Width: 1.4150685","density: 1.762490e+00<br />Petal.Width: 1.4103718","density: 1.773363e+00<br />Petal.Width: 1.4056751","density: 1.784416e+00<br />Petal.Width: 1.4009785","density: 1.795622e+00<br />Petal.Width: 1.3962818","density: 1.806953e+00<br />Petal.Width: 1.3915851","density: 1.818374e+00<br />Petal.Width: 1.3868885","density: 1.829787e+00<br />Petal.Width: 1.3821918","density: 1.841089e+00<br />Petal.Width: 1.3774951","density: 1.852196e+00<br />Petal.Width: 1.3727984","density: 1.863018e+00<br />Petal.Width: 1.3681018","density: 1.873337e+00<br />Petal.Width: 1.3634051","density: 1.882968e+00<br />Petal.Width: 1.3587084","density: 1.891864e+00<br />Petal.Width: 1.3540117","density: 1.899907e+00<br />Petal.Width: 1.3493151","density: 1.906979e+00<br />Petal.Width: 1.3446184","density: 1.912525e+00<br />Petal.Width: 1.3399217","density: 1.916673e+00<br />Petal.Width: 1.3352250","density: 1.919372e+00<br />Petal.Width: 1.3305284","density: 1.920518e+00<br />Petal.Width: 1.3258317","density: 1.919888e+00<br />Petal.Width: 1.3211350","density: 1.916920e+00<br />Petal.Width: 1.3164384","density: 1.912058e+00<br />Petal.Width: 1.3117417","density: 1.905250e+00<br />Petal.Width: 1.3070450","density: 1.896452e+00<br />Petal.Width: 1.3023483","density: 1.885247e+00<br />Petal.Width: 1.2976517","density: 1.871619e+00<br />Petal.Width: 1.2929550","density: 1.855964e+00<br />Petal.Width: 1.2882583","density: 1.838309e+00<br />Petal.Width: 1.2835616","density: 1.818690e+00<br />Petal.Width: 1.2788650","density: 1.796615e+00<br />Petal.Width: 1.2741683","density: 1.772649e+00<br />Petal.Width: 1.2694716","density: 1.747016e+00<br />Petal.Width: 1.2647750","density: 1.719813e+00<br />Petal.Width: 1.2600783","density: 1.691085e+00<br />Petal.Width: 1.2553816","density: 1.660680e+00<br />Petal.Width: 1.2506849","density: 1.629205e+00<br />Petal.Width: 1.2459883","density: 1.596796e+00<br />Petal.Width: 1.2412916","density: 1.563592e+00<br />Petal.Width: 1.2365949","density: 1.529671e+00<br />Petal.Width: 1.2318982","density: 1.495282e+00<br />Petal.Width: 1.2272016","density: 1.460698e+00<br />Petal.Width: 1.2225049","density: 1.426060e+00<br />Petal.Width: 1.2178082","density: 1.391508e+00<br />Petal.Width: 1.2131115","density: 1.357319e+00<br />Petal.Width: 1.2084149","density: 1.323662e+00<br />Petal.Width: 1.2037182","density: 1.290632e+00<br />Petal.Width: 1.1990215","density: 1.258344e+00<br />Petal.Width: 1.1943249","density: 1.226936e+00<br />Petal.Width: 1.1896282","density: 1.196896e+00<br />Petal.Width: 1.1849315","density: 1.168009e+00<br />Petal.Width: 1.1802348","density: 1.140351e+00<br />Petal.Width: 1.1755382","density: 1.113994e+00<br />Petal.Width: 1.1708415","density: 1.089211e+00<br />Petal.Width: 1.1661448","density: 1.066218e+00<br />Petal.Width: 1.1614481","density: 1.044731e+00<br />Petal.Width: 1.1567515","density: 1.024777e+00<br />Petal.Width: 1.1520548","density: 1.006375e+00<br />Petal.Width: 1.1473581","density: 9.899458e-01<br />Petal.Width: 1.1426614","density: 9.752520e-01<br />Petal.Width: 1.1379648","density: 9.620980e-01<br />Petal.Width: 1.1332681","density: 9.504589e-01<br />Petal.Width: 1.1285714","density: 9.403111e-01<br />Petal.Width: 1.1238748","density: 9.321002e-01<br />Petal.Width: 1.1191781","density: 9.252175e-01<br />Petal.Width: 1.1144814","density: 9.195959e-01<br />Petal.Width: 1.1097847","density: 9.151635e-01<br />Petal.Width: 1.1050881","density: 9.119597e-01<br />Petal.Width: 1.1003914","density: 9.099484e-01<br />Petal.Width: 1.0956947","density: 9.087525e-01<br />Petal.Width: 1.0909980","density: 9.082668e-01<br />Petal.Width: 1.0863014","density: 9.083828e-01<br />Petal.Width: 1.0816047","density: 9.090495e-01<br />Petal.Width: 1.0769080","density: 9.100054e-01<br />Petal.Width: 1.0722114","density: 9.110678e-01<br />Petal.Width: 1.0675147","density: 9.121133e-01<br />Petal.Width: 1.0628180","density: 9.130182e-01<br />Petal.Width: 1.0581213","density: 9.134714e-01<br />Petal.Width: 1.0534247","density: 9.133914e-01<br />Petal.Width: 1.0487280","density: 9.126695e-01<br />Petal.Width: 1.0440313","density: 9.111947e-01<br />Petal.Width: 1.0393346","density: 9.087558e-01<br />Petal.Width: 1.0346380","density: 9.050057e-01<br />Petal.Width: 1.0299413","density: 9.001041e-01<br />Petal.Width: 1.0252446","density: 8.939782e-01<br />Petal.Width: 1.0205479","density: 8.865617e-01<br />Petal.Width: 1.0158513","density: 8.774847e-01<br />Petal.Width: 1.0111546","density: 8.667616e-01<br />Petal.Width: 1.0064579","density: 8.545888e-01<br />Petal.Width: 1.0017613","density: 8.409579e-01<br />Petal.Width: 0.9970646","density: 8.258684e-01<br />Petal.Width: 0.9923679","density: 8.088663e-01<br />Petal.Width: 0.9876712","density: 7.904313e-01<br />Petal.Width: 0.9829746","density: 7.706703e-01<br />Petal.Width: 0.9782779","density: 7.496410e-01<br />Petal.Width: 0.9735812","density: 7.273241e-01<br />Petal.Width: 0.9688845","density: 7.036264e-01<br />Petal.Width: 0.9641879","density: 6.789954e-01<br />Petal.Width: 0.9594912","density: 6.535303e-01<br />Petal.Width: 0.9547945","density: 6.273334e-01<br />Petal.Width: 0.9500978","density: 6.004296e-01<br />Petal.Width: 0.9454012","density: 5.730384e-01<br />Petal.Width: 0.9407045","density: 5.453778e-01<br />Petal.Width: 0.9360078","density: 5.175598e-01<br />Petal.Width: 0.9313112","density: 4.896961e-01<br />Petal.Width: 0.9266145","density: 4.619856e-01<br />Petal.Width: 0.9219178","density: 4.345727e-01<br />Petal.Width: 0.9172211","density: 4.075455e-01<br />Petal.Width: 0.9125245","density: 3.809950e-01<br />Petal.Width: 0.9078278","density: 3.550482e-01<br />Petal.Width: 0.9031311","density: 3.300127e-01<br />Petal.Width: 0.8984344","density: 3.057682e-01<br />Petal.Width: 0.8937378","density: 2.823692e-01<br />Petal.Width: 0.8890411","density: 2.598658e-01<br />Petal.Width: 0.8843444","density: 2.384760e-01<br />Petal.Width: 0.8796477","density: 2.182808e-01<br />Petal.Width: 0.8749511","density: 1.991070e-01<br />Petal.Width: 0.8702544","density: 1.809657e-01<br />Petal.Width: 0.8655577","density: 1.638637e-01<br />Petal.Width: 0.8608611","density: 1.481002e-01<br />Petal.Width: 0.8561644","density: 1.334405e-01<br />Petal.Width: 0.8514677","density: 1.197812e-01<br />Petal.Width: 0.8467710","density: 1.071008e-01<br />Petal.Width: 0.8420744","density: 9.540739e-02<br />Petal.Width: 0.8373777","density: 8.492569e-02<br />Petal.Width: 0.8326810","density: 7.529885e-02<br />Petal.Width: 0.8279843","density: 6.649071e-02<br />Petal.Width: 0.8232877","density: 5.846434e-02<br />Petal.Width: 0.8185910","density: 5.128421e-02<br />Petal.Width: 0.8138943","density: 4.492444e-02<br />Petal.Width: 0.8091977","density: 3.918451e-02<br />Petal.Width: 0.8045010","density: 3.402606e-02<br />Petal.Width: 0.7998043","density: 2.941113e-02<br />Petal.Width: 0.7951076","density: 2.541918e-02<br />Petal.Width: 0.7904110","density: 2.190711e-02<br />Petal.Width: 0.7857143","density: 1.879175e-02<br />Petal.Width: 0.7810176","density: 1.604157e-02<br />Petal.Width: 0.7763209","density: 1.362997e-02<br />Petal.Width: 0.7716243","density: 1.161159e-02<br />Petal.Width: 0.7669276","density: 9.842791e-03<br />Petal.Width: 0.7622309","density: 8.300753e-03<br />Petal.Width: 0.7575342","density: 6.963675e-03<br />Petal.Width: 0.7528376","density: 5.830685e-03<br />Petal.Width: 0.7481409","density: 4.886524e-03<br />Petal.Width: 0.7434442","density: 4.072881e-03<br />Petal.Width: 0.7387476","density: 3.375840e-03<br />Petal.Width: 0.7340509","density: 2.782303e-03<br />Petal.Width: 0.7293542","density: 2.297769e-03<br />Petal.Width: 0.7246575","density: 1.893846e-03<br />Petal.Width: 0.7199609","density: 1.551624e-03<br />Petal.Width: 0.7152642","density: 1.263598e-03<br />Petal.Width: 0.7105675","density: 1.022806e-03<br />Petal.Width: 0.7058708","density: 8.339563e-04<br />Petal.Width: 0.7011742","density: 6.757768e-04<br />Petal.Width: 0.6964775","density: 5.440574e-04<br />Petal.Width: 0.6917808","density: 4.351792e-04<br />Petal.Width: 0.6870841","density: 3.474762e-04<br />Petal.Width: 0.6823875","density: 2.786952e-04<br />Petal.Width: 0.6776908","density: 2.219555e-04<br />Petal.Width: 0.6729941","density: 1.755342e-04<br />Petal.Width: 0.6682975","density: 1.378620e-04<br />Petal.Width: 0.6636008","density: 1.087398e-04<br />Petal.Width: 0.6589041","density: 8.573714e-05<br />Petal.Width: 0.6542074","density: 6.708568e-05<br />Petal.Width: 0.6495108","density: 5.209942e-05<br />Petal.Width: 0.6448141","density: 4.016377e-05<br />Petal.Width: 0.6401174","density: 3.133099e-05<br />Petal.Width: 0.6354207","density: 2.427552e-05<br />Petal.Width: 0.6307241","density: 1.865510e-05<br />Petal.Width: 0.6260274","density: 1.422198e-05<br />Petal.Width: 0.6213307","density: 1.081919e-05<br />Petal.Width: 0.6166341","density: 8.309892e-06<br />Petal.Width: 0.6119374","density: 6.324639e-06<br />Petal.Width: 0.6072407","density: 4.771724e-06<br />Petal.Width: 0.6025440","density: 3.569854e-06<br />Petal.Width: 0.5978474","density: 2.687935e-06<br />Petal.Width: 0.5931507","density: 2.028445e-06<br />Petal.Width: 0.5884540","density: 1.515927e-06<br />Petal.Width: 0.5837573","density: 1.122462e-06<br />Petal.Width: 0.5790607","density: 8.237902e-07<br />Petal.Width: 0.5743640","density: 6.147333e-07<br />Petal.Width: 0.5696673","density: 4.556027e-07<br />Petal.Width: 0.5649706","density: 3.341990e-07<br />Petal.Width: 0.5602740","density: 2.427716e-07<br />Petal.Width: 0.5555773","density: 1.757755e-07<br />Petal.Width: 0.5508806","density: 1.293916e-07<br />Petal.Width: 0.5461840","density: 9.413937e-08<br />Petal.Width: 0.5414873","density: 6.775256e-08<br />Petal.Width: 0.5367906","density: 4.826912e-08<br />Petal.Width: 0.5320939","density: 3.467302e-08<br />Petal.Width: 0.5273973","density: 2.505999e-08<br />Petal.Width: 0.5227006","density: 1.789059e-08<br />Petal.Width: 0.5180039","density: 1.262850e-08<br />Petal.Width: 0.5133072","density: 8.820722e-09<br />Petal.Width: 0.5086106","density: 6.294960e-09<br />Petal.Width: 0.5039139","density: 4.464904e-09<br />Petal.Width: 0.4992172","density: 3.126463e-09<br />Petal.Width: 0.4945205","density: 2.163701e-09<br />Petal.Width: 0.4898239","density: 1.489179e-09<br />Petal.Width: 0.4851272","density: 1.051615e-09<br />Petal.Width: 0.4804305","density: 7.316455e-10<br />Petal.Width: 0.4757339","density: 5.023032e-10<br />Petal.Width: 0.4710372","density: 3.407061e-10<br />Petal.Width: 0.4663405","density: 2.332989e-10<br />Petal.Width: 0.4616438","density: 1.616128e-10<br />Petal.Width: 0.4569472","density: 1.102423e-10<br />Petal.Width: 0.4522505","density: 7.417786e-11<br />Petal.Width: 0.4475538","density: 4.929707e-11<br />Petal.Width: 0.4428571","density: 3.363202e-11<br />Petal.Width: 0.4381605","density: 2.284247e-11<br />Petal.Width: 0.4334638","density: 1.527072e-11<br />Petal.Width: 0.4287671","density: 1.006687e-11<br />Petal.Width: 0.4240705","density: 6.576504e-12<br />Petal.Width: 0.4193738","density: 4.460088e-12<br />Petal.Width: 0.4146771","density: 2.968566e-12<br />Petal.Width: 0.4099804","density: 1.944158e-12<br />Petal.Width: 0.4052838","density: 1.255215e-12<br />Petal.Width: 0.4005871","density: 8.185501e-13<br />Petal.Width: 0.3958904","density: 5.438510e-13<br />Petal.Width: 0.3911937","density: 3.545095e-13<br />Petal.Width: 0.3864971","density: 2.273495e-13<br />Petal.Width: 0.3818004","density: 1.437161e-13<br />Petal.Width: 0.3771037","density: 9.374916e-14<br />Petal.Width: 0.3724070","density: 6.094729e-14<br />Petal.Width: 0.3677104","density: 3.882958e-14<br />Petal.Width: 0.3630137","density: 2.432527e-14<br />Petal.Width: 0.3583170","density: 1.498986e-14<br />Petal.Width: 0.3536204","density: 9.819446e-15<br />Petal.Width: 0.3489237","density: 6.353721e-15<br />Petal.Width: 0.3442270","density: 4.034385e-15<br />Petal.Width: 0.3395303","density: 2.466779e-15<br />Petal.Width: 0.3348337","density: 1.534124e-15<br />Petal.Width: 0.3301370","density: 9.923603e-16<br />Petal.Width: 0.3254403","density: 6.014514e-16<br />Petal.Width: 0.3207436","density: 3.154952e-16<br />Petal.Width: 0.3160470","density: 1.152985e-16<br />Petal.Width: 0.3113503","density: 5.389472e-17<br />Petal.Width: 0.3066536","density: 5.919652e-17<br />Petal.Width: 0.3019569","density: 3.799838e-17<br />Petal.Width: 0.2972603","density: 0.000000e+00<br />Petal.Width: 0.2925636","density: 0.000000e+00<br />Petal.Width: 0.2878669","density: 0.000000e+00<br />Petal.Width: 0.2831703","density: 1.927255e-17<br />Petal.Width: 0.2784736","density: 2.072092e-17<br />Petal.Width: 0.2737769","density: 1.068415e-17<br />Petal.Width: 0.2690802","density: 6.390072e-17<br />Petal.Width: 0.2643836","density: 1.819114e-17<br />Petal.Width: 0.2596869","density: 3.233116e-17<br />Petal.Width: 0.2549902","density: 1.015140e-16<br />Petal.Width: 0.2502935","density: 1.857733e-16<br />Petal.Width: 0.2455969","density: 1.421085e-16<br />Petal.Width: 0.2409002","density: 1.445246e-16<br />Petal.Width: 0.2362035","density: 1.022764e-16<br />Petal.Width: 0.2315068","density: 3.440031e-17<br />Petal.Width: 0.2268102","density: 2.403423e-18<br />Petal.Width: 0.2221135","density: 8.154601e-17<br />Petal.Width: 0.2174168","density: 4.675395e-17<br />Petal.Width: 0.2127202","density: 2.904440e-17<br />Petal.Width: 0.2080235","density: 4.893777e-17<br />Petal.Width: 0.2033268","density: 2.543841e-18<br />Petal.Width: 0.1986301","density: 0.000000e+00<br />Petal.Width: 0.1939335","density: 0.000000e+00<br />Petal.Width: 0.1892368","density: 2.197265e-17<br />Petal.Width: 0.1845401","density: 6.258237e-17<br />Petal.Width: 0.1798434","density: 1.073034e-17<br />Petal.Width: 0.1751468","density: 6.214201e-17<br />Petal.Width: 0.1704501","density: 5.164937e-17<br />Petal.Width: 0.1657534","density: 2.599349e-17<br />Petal.Width: 0.1610568","density: 8.502689e-17<br />Petal.Width: 0.1563601","density: 1.232337e-16<br />Petal.Width: 0.1516634","density: 9.377999e-17<br />Petal.Width: 0.1469667","density: 4.190902e-17<br />Petal.Width: 0.1422701","density: 0.000000e+00<br />Petal.Width: 0.1375734","density: 0.000000e+00<br />Petal.Width: 0.1328767","density: 0.000000e+00<br />Petal.Width: 0.1281800","density: 1.881874e-17<br />Petal.Width: 0.1234834","density: 2.327965e-17<br />Petal.Width: 0.1187867","density: 1.965338e-17<br />Petal.Width: 0.1140900","density: 1.533771e-16<br />Petal.Width: 0.1093933","density: 5.529481e-17<br />Petal.Width: 0.1046967","density: 1.820087e-18<br />Petal.Width: 0.1000000","density: 1.820087e-18<br />Petal.Width: 0.1000000"],"key":["51","52","53","54","55","56","57","58","59","60","61","62","63","64","65","66","67","68","69","70","71","72","73","74","75","76","77","78","79","80","81","82","83","84","85","86","87","88","89","90","91","92","93","94","95","96","97","98","99","100"],"type":"scatter","mode":"lines","line":{"width":1.88976377952756,"color":"rgba(0,0,0,1)","dash":"solid"},"fill":"toself","fillcolor":"rgba(0,186,56,1)","hoveron":"points","set":"SharedData91833e3a","name":"versicolor","legendgroup":"versicolor","showlegend":true,"xaxis":"x4","yaxis":"y13","hoverinfo":"text","_isSimpleKey":true,"_isNestedKey":false,"frame":null},{"x":[0.1,0.104696673189824,0.109393346379648,0.114090019569472,0.118786692759296,0.123483365949119,0.128180039138943,0.132876712328767,0.137573385518591,0.142270058708415,0.146966731898239,0.151663405088063,0.156360078277887,0.16105675146771,0.165753424657534,0.170450097847358,0.175146771037182,0.179843444227006,0.18454011741683,0.189236790606654,0.193933463796477,0.198630136986301,0.203326810176125,0.208023483365949,0.212720156555773,0.217416829745597,0.222113502935421,0.226810176125245,0.231506849315068,0.236203522504892,0.240900195694716,0.24559686888454,0.250293542074364,0.254990215264188,0.259686888454012,0.264383561643836,0.26908023483366,0.273776908023483,0.278473581213307,0.283170254403131,0.287866927592955,0.292563600782779,0.297260273972603,0.301956947162427,0.306653620352251,0.311350293542074,0.316046966731898,0.320743639921722,0.325440313111546,0.33013698630137,0.334833659491194,0.339530332681018,0.344227005870841,0.348923679060665,0.353620352250489,0.358317025440313,0.363013698630137,0.367710371819961,0.372407045009785,0.377103718199609,0.381800391389432,0.386497064579256,0.39119373776908,0.395890410958904,0.400587084148728,0.405283757338552,0.409980430528376,0.4146771037182,0.419373776908023,0.424070450097847,0.428767123287671,0.433463796477495,0.438160469667319,0.442857142857143,0.447553816046967,0.452250489236791,0.456947162426614,0.461643835616438,0.466340508806262,0.471037181996086,0.47573385518591,0.480430528375734,0.485127201565558,0.489823874755382,0.494520547945206,0.499217221135029,0.503913894324853,0.508610567514677,0.513307240704501,0.518003913894325,0.522700587084149,0.527397260273973,0.532093933463797,0.53679060665362,0.541487279843444,0.546183953033268,0.550880626223092,0.555577299412916,0.56027397260274,0.564970645792564,0.569667318982387,0.574363992172211,0.579060665362035,0.583757338551859,0.588454011741683,0.593150684931507,0.597847358121331,0.602544031311155,0.607240704500978,0.611937377690802,0.616634050880626,0.62133072407045,0.626027397260274,0.630724070450098,0.635420743639922,0.640117416829746,0.644814090019569,0.649510763209393,0.654207436399217,0.658904109589041,0.663600782778865,0.668297455968689,0.672994129158513,0.677690802348337,0.68238747553816,0.687084148727984,0.691780821917808,0.696477495107632,0.701174168297456,0.70587084148728,0.710567514677104,0.715264187866928,0.719960861056751,0.724657534246575,0.729354207436399,0.734050880626223,0.738747553816047,0.743444227005871,0.748140900195695,0.752837573385519,0.757534246575342,0.762230919765166,0.76692759295499,0.771624266144814,0.776320939334638,0.781017612524462,0.785714285714286,0.79041095890411,0.795107632093933,0.799804305283757,0.804500978473581,0.809197651663405,0.813894324853229,0.818590998043053,0.823287671232877,0.827984344422701,0.832681017612524,0.837377690802348,0.842074363992172,0.846771037181996,0.85146771037182,0.856164383561644,0.860861056751468,0.865557729941292,0.870254403131115,0.874951076320939,0.879647749510763,0.884344422700587,0.889041095890411,0.893737769080235,0.898434442270059,0.903131115459883,0.907827788649706,0.91252446183953,0.917221135029354,0.921917808219178,0.926614481409002,0.931311154598826,0.93600782778865,0.940704500978474,0.945401174168297,0.950097847358121,0.954794520547945,0.959491193737769,0.964187866927593,0.968884540117417,0.973581213307241,0.978277886497064,0.982974559686888,0.987671232876712,0.992367906066536,0.99706457925636,1.00176125244618,1.00645792563601,1.01115459882583,1.01585127201566,1.02054794520548,1.0252446183953,1.02994129158513,1.03463796477495,1.03933463796477,1.0440313111546,1.04872798434442,1.05342465753425,1.05812133072407,1.06281800391389,1.06751467710372,1.07221135029354,1.07690802348337,1.08160469667319,1.08630136986301,1.09099804305284,1.09569471624266,1.10039138943249,1.10508806262231,1.10978473581213,1.11448140900196,1.11917808219178,1.1238747553816,1.12857142857143,1.13326810176125,1.13796477495108,1.1426614481409,1.14735812133072,1.15205479452055,1.15675146771037,1.1614481409002,1.16614481409002,1.17084148727984,1.17553816046967,1.18023483365949,1.18493150684932,1.18962818003914,1.19432485322896,1.19902152641879,1.20371819960861,1.20841487279843,1.21311154598826,1.21780821917808,1.22250489236791,1.22720156555773,1.23189823874755,1.23659491193738,1.2412915851272,1.24598825831703,1.25068493150685,1.25538160469667,1.2600782778865,1.26477495107632,1.26947162426614,1.27416829745597,1.27886497064579,1.28356164383562,1.28825831702544,1.29295499021526,1.29765166340509,1.30234833659491,1.30704500978474,1.31174168297456,1.31643835616438,1.32113502935421,1.32583170254403,1.33052837573386,1.33522504892368,1.3399217221135,1.34461839530333,1.34931506849315,1.35401174168297,1.3587084148728,1.36340508806262,1.36810176125245,1.37279843444227,1.37749510763209,1.38219178082192,1.38688845401174,1.39158512720157,1.39628180039139,1.40097847358121,1.40567514677104,1.41037181996086,1.41506849315068,1.41976516634051,1.42446183953033,1.42915851272016,1.43385518590998,1.4385518590998,1.44324853228963,1.44794520547945,1.45264187866928,1.4573385518591,1.46203522504892,1.46673189823875,1.47142857142857,1.4761252446184,1.48082191780822,1.48551859099804,1.49021526418787,1.49491193737769,1.49960861056751,1.50430528375734,1.50900195694716,1.51369863013699,1.51839530332681,1.52309197651663,1.52778864970646,1.53248532289628,1.53718199608611,1.54187866927593,1.54657534246575,1.55127201565558,1.5559686888454,1.56066536203523,1.56536203522505,1.57005870841487,1.5747553816047,1.57945205479452,1.58414872798434,1.58884540117417,1.59354207436399,1.59823874755382,1.60293542074364,1.60763209393346,1.61232876712329,1.61702544031311,1.62172211350294,1.62641878669276,1.63111545988258,1.63581213307241,1.64050880626223,1.64520547945205,1.64990215264188,1.6545988258317,1.65929549902153,1.66399217221135,1.66868884540117,1.673385518591,1.67808219178082,1.68277886497065,1.68747553816047,1.69217221135029,1.69686888454012,1.70156555772994,1.70626223091977,1.71095890410959,1.71565557729941,1.72035225048924,1.72504892367906,1.72974559686888,1.73444227005871,1.73913894324853,1.74383561643836,1.74853228962818,1.753228962818,1.75792563600783,1.76262230919765,1.76731898238748,1.7720156555773,1.77671232876712,1.78140900195695,1.78610567514677,1.79080234833659,1.79549902152642,1.80019569471624,1.80489236790607,1.80958904109589,1.81428571428571,1.81898238747554,1.82367906066536,1.82837573385519,1.83307240704501,1.83776908023483,1.84246575342466,1.84716242661448,1.85185909980431,1.85655577299413,1.86125244618395,1.86594911937378,1.8706457925636,1.87534246575342,1.88003913894325,1.88473581213307,1.8894324853229,1.89412915851272,1.89882583170254,1.90352250489237,1.90821917808219,1.91291585127202,1.91761252446184,1.92230919765166,1.92700587084149,1.93170254403131,1.93639921722114,1.94109589041096,1.94579256360078,1.95048923679061,1.95518590998043,1.95988258317025,1.96457925636008,1.9692759295499,1.97397260273973,1.97866927592955,1.98336594911937,1.9880626223092,1.99275929549902,1.99745596868885,2.00215264187867,2.00684931506849,2.01154598825832,2.01624266144814,2.02093933463796,2.02563600782779,2.03033268101761,2.03502935420744,2.03972602739726,2.04442270058708,2.04911937377691,2.05381604696673,2.05851272015656,2.06320939334638,2.0679060665362,2.07260273972603,2.07729941291585,2.08199608610568,2.0866927592955,2.09138943248532,2.09608610567515,2.10078277886497,2.10547945205479,2.11017612524462,2.11487279843444,2.11956947162427,2.12426614481409,2.12896281800391,2.13365949119374,2.13835616438356,2.14305283757339,2.14774951076321,2.15244618395303,2.15714285714286,2.16183953033268,2.1665362035225,2.17123287671233,2.17592954990215,2.18062622309198,2.1853228962818,2.19001956947162,2.19471624266145,2.19941291585127,2.2041095890411,2.20880626223092,2.21350293542074,2.21819960861057,2.22289628180039,2.22759295499022,2.23228962818004,2.23698630136986,2.24168297455969,2.24637964774951,2.25107632093933,2.25577299412916,2.26046966731898,2.26516634050881,2.26986301369863,2.27455968688845,2.27925636007828,2.2839530332681,2.28864970645793,2.29334637964775,2.29804305283757,2.3027397260274,2.30743639921722,2.31213307240705,2.31682974559687,2.32152641878669,2.32622309197652,2.33091976516634,2.33561643835616,2.34031311154599,2.34500978473581,2.34970645792564,2.35440313111546,2.35909980430528,2.36379647749511,2.36849315068493,2.37318982387476,2.37788649706458,2.3825831702544,2.38727984344423,2.39197651663405,2.39667318982387,2.4013698630137,2.40606653620352,2.41076320939335,2.41545988258317,2.42015655577299,2.42485322896282,2.42954990215264,2.43424657534247,2.43894324853229,2.44363992172211,2.44833659491194,2.45303326810176,2.45772994129159,2.46242661448141,2.46712328767123,2.47181996086106,2.47651663405088,2.4812133072407,2.48590998043053,2.49060665362035,2.49530332681018,2.5,2.5,2.5,2.49530332681018,2.49060665362035,2.48590998043053,2.4812133072407,2.47651663405088,2.47181996086106,2.46712328767123,2.46242661448141,2.45772994129159,2.45303326810176,2.44833659491194,2.44363992172211,2.43894324853229,2.43424657534247,2.42954990215264,2.42485322896282,2.42015655577299,2.41545988258317,2.41076320939335,2.40606653620352,2.4013698630137,2.39667318982387,2.39197651663405,2.38727984344423,2.3825831702544,2.37788649706458,2.37318982387476,2.36849315068493,2.36379647749511,2.35909980430528,2.35440313111546,2.34970645792564,2.34500978473581,2.34031311154599,2.33561643835616,2.33091976516634,2.32622309197652,2.32152641878669,2.31682974559687,2.31213307240705,2.30743639921722,2.3027397260274,2.29804305283757,2.29334637964775,2.28864970645793,2.2839530332681,2.27925636007828,2.27455968688845,2.26986301369863,2.26516634050881,2.26046966731898,2.25577299412916,2.25107632093933,2.24637964774951,2.24168297455969,2.23698630136986,2.23228962818004,2.22759295499022,2.22289628180039,2.21819960861057,2.21350293542074,2.20880626223092,2.2041095890411,2.19941291585127,2.19471624266145,2.19001956947162,2.1853228962818,2.18062622309198,2.17592954990215,2.17123287671233,2.1665362035225,2.16183953033268,2.15714285714286,2.15244618395303,2.14774951076321,2.14305283757339,2.13835616438356,2.13365949119374,2.12896281800391,2.12426614481409,2.11956947162427,2.11487279843444,2.11017612524462,2.10547945205479,2.10078277886497,2.09608610567515,2.09138943248532,2.0866927592955,2.08199608610568,2.07729941291585,2.07260273972603,2.0679060665362,2.06320939334638,2.05851272015656,2.05381604696673,2.04911937377691,2.04442270058708,2.03972602739726,2.03502935420744,2.03033268101761,2.02563600782779,2.02093933463796,2.01624266144814,2.01154598825832,2.00684931506849,2.00215264187867,1.99745596868885,1.99275929549902,1.9880626223092,1.98336594911937,1.97866927592955,1.97397260273973,1.9692759295499,1.96457925636008,1.95988258317025,1.95518590998043,1.95048923679061,1.94579256360078,1.94109589041096,1.93639921722114,1.93170254403131,1.92700587084149,1.92230919765166,1.91761252446184,1.91291585127202,1.90821917808219,1.90352250489237,1.89882583170254,1.89412915851272,1.8894324853229,1.88473581213307,1.88003913894325,1.87534246575342,1.8706457925636,1.86594911937378,1.86125244618395,1.85655577299413,1.85185909980431,1.84716242661448,1.84246575342466,1.83776908023483,1.83307240704501,1.82837573385519,1.82367906066536,1.81898238747554,1.81428571428571,1.80958904109589,1.80489236790607,1.80019569471624,1.79549902152642,1.79080234833659,1.78610567514677,1.78140900195695,1.77671232876712,1.7720156555773,1.76731898238748,1.76262230919765,1.75792563600783,1.753228962818,1.74853228962818,1.74383561643836,1.73913894324853,1.73444227005871,1.72974559686888,1.72504892367906,1.72035225048924,1.71565557729941,1.71095890410959,1.70626223091977,1.70156555772994,1.69686888454012,1.69217221135029,1.68747553816047,1.68277886497065,1.67808219178082,1.673385518591,1.66868884540117,1.66399217221135,1.65929549902153,1.6545988258317,1.64990215264188,1.64520547945205,1.64050880626223,1.63581213307241,1.63111545988258,1.62641878669276,1.62172211350294,1.61702544031311,1.61232876712329,1.60763209393346,1.60293542074364,1.59823874755382,1.59354207436399,1.58884540117417,1.58414872798434,1.57945205479452,1.5747553816047,1.57005870841487,1.56536203522505,1.56066536203523,1.5559686888454,1.55127201565558,1.54657534246575,1.54187866927593,1.53718199608611,1.53248532289628,1.52778864970646,1.52309197651663,1.51839530332681,1.51369863013699,1.50900195694716,1.50430528375734,1.49960861056751,1.49491193737769,1.49021526418787,1.48551859099804,1.48082191780822,1.4761252446184,1.47142857142857,1.46673189823875,1.46203522504892,1.4573385518591,1.45264187866928,1.44794520547945,1.44324853228963,1.4385518590998,1.43385518590998,1.42915851272016,1.42446183953033,1.41976516634051,1.41506849315068,1.41037181996086,1.40567514677104,1.40097847358121,1.39628180039139,1.39158512720157,1.38688845401174,1.38219178082192,1.37749510763209,1.37279843444227,1.36810176125245,1.36340508806262,1.3587084148728,1.35401174168297,1.34931506849315,1.34461839530333,1.3399217221135,1.33522504892368,1.33052837573386,1.32583170254403,1.32113502935421,1.31643835616438,1.31174168297456,1.30704500978474,1.30234833659491,1.29765166340509,1.29295499021526,1.28825831702544,1.28356164383562,1.27886497064579,1.27416829745597,1.26947162426614,1.26477495107632,1.2600782778865,1.25538160469667,1.25068493150685,1.24598825831703,1.2412915851272,1.23659491193738,1.23189823874755,1.22720156555773,1.22250489236791,1.21780821917808,1.21311154598826,1.20841487279843,1.20371819960861,1.19902152641879,1.19432485322896,1.18962818003914,1.18493150684932,1.18023483365949,1.17553816046967,1.17084148727984,1.16614481409002,1.1614481409002,1.15675146771037,1.15205479452055,1.14735812133072,1.1426614481409,1.13796477495108,1.13326810176125,1.12857142857143,1.1238747553816,1.11917808219178,1.11448140900196,1.10978473581213,1.10508806262231,1.10039138943249,1.09569471624266,1.09099804305284,1.08630136986301,1.08160469667319,1.07690802348337,1.07221135029354,1.06751467710372,1.06281800391389,1.05812133072407,1.05342465753425,1.04872798434442,1.0440313111546,1.03933463796477,1.03463796477495,1.02994129158513,1.0252446183953,1.02054794520548,1.01585127201566,1.01115459882583,1.00645792563601,1.00176125244618,0.99706457925636,0.992367906066536,0.987671232876712,0.982974559686888,0.978277886497064,0.973581213307241,0.968884540117417,0.964187866927593,0.959491193737769,0.954794520547945,0.950097847358121,0.945401174168297,0.940704500978474,0.93600782778865,0.931311154598826,0.926614481409002,0.921917808219178,0.917221135029354,0.91252446183953,0.907827788649706,0.903131115459883,0.898434442270059,0.893737769080235,0.889041095890411,0.884344422700587,0.879647749510763,0.874951076320939,0.870254403131115,0.865557729941292,0.860861056751468,0.856164383561644,0.85146771037182,0.846771037181996,0.842074363992172,0.837377690802348,0.832681017612524,0.827984344422701,0.823287671232877,0.818590998043053,0.813894324853229,0.809197651663405,0.804500978473581,0.799804305283757,0.795107632093933,0.79041095890411,0.785714285714286,0.781017612524462,0.776320939334638,0.771624266144814,0.76692759295499,0.762230919765166,0.757534246575342,0.752837573385519,0.748140900195695,0.743444227005871,0.738747553816047,0.734050880626223,0.729354207436399,0.724657534246575,0.719960861056751,0.715264187866928,0.710567514677104,0.70587084148728,0.701174168297456,0.696477495107632,0.691780821917808,0.687084148727984,0.68238747553816,0.677690802348337,0.672994129158513,0.668297455968689,0.663600782778865,0.658904109589041,0.654207436399217,0.649510763209393,0.644814090019569,0.640117416829746,0.635420743639922,0.630724070450098,0.626027397260274,0.62133072407045,0.616634050880626,0.611937377690802,0.607240704500978,0.602544031311155,0.597847358121331,0.593150684931507,0.588454011741683,0.583757338551859,0.579060665362035,0.574363992172211,0.569667318982387,0.564970645792564,0.56027397260274,0.555577299412916,0.550880626223092,0.546183953033268,0.541487279843444,0.53679060665362,0.532093933463797,0.527397260273973,0.522700587084149,0.518003913894325,0.513307240704501,0.508610567514677,0.503913894324853,0.499217221135029,0.494520547945206,0.489823874755382,0.485127201565558,0.480430528375734,0.47573385518591,0.471037181996086,0.466340508806262,0.461643835616438,0.456947162426614,0.452250489236791,0.447553816046967,0.442857142857143,0.438160469667319,0.433463796477495,0.428767123287671,0.424070450097847,0.419373776908023,0.4146771037182,0.409980430528376,0.405283757338552,0.400587084148728,0.395890410958904,0.39119373776908,0.386497064579256,0.381800391389432,0.377103718199609,0.372407045009785,0.367710371819961,0.363013698630137,0.358317025440313,0.353620352250489,0.348923679060665,0.344227005870841,0.339530332681018,0.334833659491194,0.33013698630137,0.325440313111546,0.320743639921722,0.316046966731898,0.311350293542074,0.306653620352251,0.301956947162427,0.297260273972603,0.292563600782779,0.287866927592955,0.283170254403131,0.278473581213307,0.273776908023483,0.26908023483366,0.264383561643836,0.259686888454012,0.254990215264188,0.250293542074364,0.24559686888454,0.240900195694716,0.236203522504892,0.231506849315068,0.226810176125245,0.222113502935421,0.217416829745597,0.212720156555773,0.208023483365949,0.203326810176125,0.198630136986301,0.193933463796477,0.189236790606654,0.18454011741683,0.179843444227006,0.175146771037182,0.170450097847358,0.165753424657534,0.16105675146771,0.156360078277887,0.151663405088063,0.146966731898239,0.142270058708415,0.137573385518591,0.132876712328767,0.128180039138943,0.123483365949119,0.118786692759296,0.114090019569472,0.109393346379648,0.104696673189824,0.1,0.1],"y":[0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0.480633396788584,0.495450801241174,0.510338836582338,0.525291991602453,0.54031973956631,0.555395892875511,0.570501798108408,0.585631312978989,0.600772890023597,0.615904679110501,0.631019327699958,0.646100894997604,0.661115993386698,0.676067700681322,0.690947801177591,0.705704017533822,0.720326754029583,0.734824836563145,0.749188466999408,0.763299592789572,0.777228610551957,0.79096572478504,0.804451683839735,0.817615103127103,0.830527319665818,0.843178703455939,0.855431039609261,0.867324494443396,0.878901739147373,0.890144701243622,0.900842632653073,0.911177516027879,0.92114313759179,0.930640042390467,0.939600885227423,0.94816259911188,0.956321937602609,0.963892076529064,0.970986952473915,0.977672459281634,0.983928615917491,0.989557447799341,0.994791353979835,0.999635286062064,1.0039987803845,1.00787175451958,1.01139511867919,1.01457736114351,1.0172885720119,1.01966568244021,1.02176412929502,1.02357812412549,1.02503410642303,1.02628669437137,1.02734920902145,1.02819741049922,1.02887021685683,1.02943504500743,1.0299054410215,1.03027978819964,1.03061898994256,1.03094249144127,1.03126563008365,1.03163239854632,1.03205238330492,1.03253540313651,1.03312284901232,1.03384174044213,1.03467436494669,1.0356275716986,1.03678385767064,1.03809878246259,1.0395632901575,1.04120001542549,1.04307283019134,1.04510652358938,1.04730120268332,1.04970925181705,1.05231838434258,1.05508169523058,1.0579970529627,1.06113641875225,1.06442238414655,1.06784152483276,1.07140715858907,1.07514588397798,1.07899435092193,1.08294859410037,1.08703572290133,1.09123396946813,1.09551508993108,1.09987557260023,1.1043477979582,1.10888727439299,1.11348852251861,1.11815779784777,1.12290387379928,1.12770001685698,1.13254487275515,1.13745348016834,1.14241628313364,1.14742202918096,1.15247001068318,1.15758242926905,1.16273461564509,1.16792485081598,1.17315878158437,1.17844038160921,1.18375208325547,1.18909179620878,1.19446389869666,1.19985405143726,1.20525271985042,1.21065565745403,1.2160437459102,1.2214059608208,1.2267344035844,1.23200721257717,1.23718332213352,1.24226864289525,1.24725179960623,1.25205418701789,1.25666553606502,1.26109379385125,1.26531810609882,1.26916450757422,1.27272961408778,1.27599561267913,1.27885711568596,1.28120585965196,1.28314382663625,1.28465215126805,1.2854821033696,1.28571199613267,1.28539937726834,1.28449652543061,1.28264998468104,1.2801583686589,1.27700698558916,1.2729955789317,1.26802023513295,1.26230940294149,1.25585339705567,1.24827818624159,1.2398193710196,1.2305801960453,1.22050134794824,1.20922260130782,1.19716900908195,1.18434648326597,1.17055321329711,1.15578882818377,1.14031375571578,1.12414145925248,1.10698207061285,1.08913600093097,1.07070065694359,1.05164848551249,1.03183386624926,1.01157173601744,0.990888448457262,0.969717679766927,0.948152915067644,0.926336199731329,0.904296611479702,0.882022291792392,0.859660422969958,0.837254491343161,0.814841908872492,0.792528527944699,0.770343770085133,0.748314449010155,0.726555454278426,0.705136812611996,0.684023534283772,0.663238140154078,0.643031598473396,0.623277416323946,0.60396537296382,0.585182451645025,0.567127957131479,0.549594470065567,0.532591033464653,0.516323827657586,0.500758081688821,0.48575624883298,0.471320945840927,0.457770660111782,0.444808593391478,0.432404692421347,0.420643714017846,0.409656300684086,0.399189117848239,0.389233330445788,0.379954427341291,0.371248731575068,0.362989485826752,0.355164641334528,0.347976691950223,0.341165071119553,0.334706771491375,0.328643325262166,0.323000273910287,0.317624348306592,0.312501318969529,0.307691813149795,0.303111382327586,0.298700136310018,0.294444827363943,0.290387768550912,0.286424026987704,0.282542452097821,0.278739645671635,0.274991545660318,0.271264207848352,0.267548871654657,0.263826548904204,0.260076933671754,0.25629534863505,0.252474417554157,0.248572647630338,0.24460990992077,0.240583177889462,0.23647172104365,0.232259125739426,0.227970716093853,0.223605778941991,0.219129840279418,0.214566202014515,0.209928972039563,0.205216242601065,0.200398959319298,0.195520693189604,0.190584427157967,0.185581599276495,0.180519109360937,0.175419507952104,0.170286678204398,0.165118077987288,0.15993386905116,0.15474183465134,0.149546900507579,0.144364846269685,0.139201465765125,0.134061066210446,0.128959436401481,0.123911929363785,0.118912523271159,0.113965139018587,0.109107718224036,0.104327668031684,0.0996211107068536,0.0949992431447468,0.0905056364266452,0.0861028241905928,0.0817931359637162,0.0776098931066752,0.0735593607619959,0.0696134719989325,0.0657736974420323,0.0620984956635401,0.0585446958389909,0.0551026196954778,0.0517860143234695,0.0486375541896714,0.0456018771294179,0.0426787041276214,0.039903767886583,0.0372699061662129,0.0347448669843724,0.0323277591281744,0.0300724404405137,0.0279265503444607,0.0258812694534565,0.023948432668562,0.0221545167962684,0.0204518029426561,0.0188385831807814,0.017341631484946,0.0159472754155158,0.0146315599801336,0.0133926429634207,0.0122663697386331,0.0112108822317697,0.0102209423708537,0.00930416321272601,0.00847156084375336,0.00769357270458074,0.0069684633585458,0.0063113215834196,0.00571015337663224,0.00515187541331951,0.00463491873536554,0.00417744726921246,0.00375485315441919,0.00336491811420448,0.00301138930850271,0.00269701454783357,0.00240786179402093,0.00214282963938348,0.00190848064490681,0.00169777294302337,0.00150523788285349,0.00132998440741033,0.00117915954706536,0.00104171958871559,0.000916989672552431,0.00080639626648402,0.000710037677223577,0.000622811202058309,0.000544213607313441,0.000476471822847811,0.000416564356241856,0.000362713989481135,0.000314657318994017,0.000274273918485284,0.000238040173612782,0.000205707810171217,0.000177699135357673,0.000153770522026198,0.00013245516169425,0.000113579574088353,9.77365488495713e-05,8.39444972137608e-05,7.1751056074703e-05,6.10939792728885e-05,5.23417057391679e-05,4.46105507234927e-05,3.7829084191453e-05,3.2094815101666e-05,2.72871588542734e-05,2.30733665007923e-05,1.94073214211614e-05,1.64131789926329e-05,1.38450336568994e-05,1.16123516660703e-05,9.70301734135186e-06,8.16922334272695e-06,6.83542955618748e-06,5.68563324794867e-06,4.73741154025697e-06,3.95651015491241e-06,3.28314178499577e-06,2.7077351603023e-06,2.25078455557944e-06,1.86425284053315e-06,1.53386243908932e-06,1.25764762899141e-06,1.04040802578632e-06,8.5443745167914e-07,6.96918413160294e-07,5.70322403767904e-07,4.67813327346041e-07,3.80859640813138e-07,3.07897952553767e-07,2.51596226270279e-07,2.04581630550998e-07,1.65078105123526e-07,1.32818268713985e-07,1.07953062510168e-07,8.69993064784373e-08,6.95646786548079e-08,5.59193318306327e-08,4.50444654164798e-08,3.59711249272068e-08,2.84971926969399e-08,2.28964531255011e-08,1.82749305218525e-08,1.44583696062388e-08,1.14163249429739e-08,9.11603583016715e-09,7.20798551775399e-09,5.64876769093156e-09,4.46116803433148e-09,3.52865042366648e-09,2.76347750205393e-09,2.14488097789146e-09,1.69496669487953e-09,1.32774028337308e-09,1.0297356144137e-09,7.98071184687988e-10,6.26033189835905e-10,4.85579813051762e-10,3.72881681651022e-10,2.89399494185352e-10,2.24747239693913e-10,1.72582297092912e-10,1.31203048796566e-10,1.02008406584454e-10,7.84142122674244e-11,5.96033618945398e-11,4.53511871087477e-11,3.49457124054863e-11,2.6585434295915e-11,2.00002189365386e-11,1.52589185895003e-11,1.16336019212638e-11,8.75781343601169e-12,6.5200971602546e-12,4.98926288164161e-12,3.76305975594844e-12,2.80282395718325e-12,2.09431717881921e-12,1.58515980662818e-12,1.18258267166295e-12,8.71332841215772e-13,6.53690525688143e-13,4.89227048675243e-13,3.61005174004076e-13,2.63594311774595e-13,1.98383474876079e-13,1.46867923637104e-13,1.07227249837814e-13,7.87418300674787e-14,5.85817708551686e-14,4.28613400321948e-14,3.09007660991656e-14,2.29175911316833e-14,1.68674157710922e-14,1.22007532259175e-14,8.75347312632278e-15,6.42404188184415e-15,4.65453013314065e-15,3.3582645806972e-15,2.47124513986585e-15,1.82198472488667e-15,1.28290595534151e-15,8.42515825865004e-16,6.40920498403324e-16,4.82387746037618e-16,3.61545463302027e-16,2.80907332317355e-16,1.90608381979735e-16,1.58518463804179e-16,1.58274347759817e-16,1.17955282267481e-16,7.76362167751458e-17,7.37051511797045e-17,1.08770866805372e-16,7.85315676861213e-17,5.53244920576037e-17,3.19307572554321e-17,1.69145813618137e-18,9.51594699435686e-18,1.38777878078145e-17,1.1957883682919e-17,1.878117309835e-18,0,0,0,0,0,0,0,5.6226447814578e-17,6.65475672437894e-17,4.34083951897452e-17,5.3488161562829e-17,2.32840444126379e-17,0,5.06807218713089e-18,1.15945502291053e-16,1.18943272018225e-16,6.55999828139891e-17,5.12138457548693e-18,0,5.42818084243881e-18,1.55079472155224e-17,2.55877135886064e-17,1.19317669235064e-17,0,0,0,4.61832449046721e-17,7.75474197931435e-17,6.98905518155762e-18,5.44879795114545e-17,1.67799662588407e-16,2.84517313084314e-16,1.43400583861142e-16,1.03255270475048e-16,8.52374829964901e-17,5.72210424113208e-17,1.5801870614216e-16,1.42667156858526e-16,8.59214394685156e-17,9.60012058415995e-17,9.71445146547012e-17,9.20060785294498e-17,8.19263121563659e-17,7.18465457832822e-17,6.17667794101983e-17,5.16870130371145e-17,4.1659480182856e-17,5.17392465559399e-17,3.65875661379605e-17,3.14466696490604e-17,1.02005034260648e-16,1.28605472865292e-16,1.68739260678091e-16,2.17262435597323e-16,1.76943370104988e-16,1.44101591882933e-16,1.2857935610588e-16,1.38659122478964e-16,1.88582931947658e-16,1.5886290240384e-16,1.01874450463584e-16,1.22033983209752e-16,1.42193515955919e-16,1.28412208845639e-16,8.11653869002199e-17,0],"text":["density: 8.116539e-17<br />Petal.Width: 0.1000000","density: 1.284122e-16<br />Petal.Width: 0.1046967","density: 1.421935e-16<br />Petal.Width: 0.1093933","density: 1.220340e-16<br />Petal.Width: 0.1140900","density: 1.018745e-16<br />Petal.Width: 0.1187867","density: 1.588629e-16<br />Petal.Width: 0.1234834","density: 1.885829e-16<br />Petal.Width: 0.1281800","density: 1.386591e-16<br />Petal.Width: 0.1328767","density: 1.285794e-16<br />Petal.Width: 0.1375734","density: 1.441016e-16<br />Petal.Width: 0.1422701","density: 1.769434e-16<br />Petal.Width: 0.1469667","density: 2.172624e-16<br />Petal.Width: 0.1516634","density: 1.687393e-16<br />Petal.Width: 0.1563601","density: 1.286055e-16<br />Petal.Width: 0.1610568","density: 1.020050e-16<br />Petal.Width: 0.1657534","density: 3.144667e-17<br />Petal.Width: 0.1704501","density: 3.658757e-17<br />Petal.Width: 0.1751468","density: 5.173925e-17<br />Petal.Width: 0.1798434","density: 4.165948e-17<br />Petal.Width: 0.1845401","density: 5.168701e-17<br />Petal.Width: 0.1892368","density: 6.176678e-17<br />Petal.Width: 0.1939335","density: 7.184655e-17<br />Petal.Width: 0.1986301","density: 8.192631e-17<br />Petal.Width: 0.2033268","density: 9.200608e-17<br />Petal.Width: 0.2080235","density: 9.714451e-17<br />Petal.Width: 0.2127202","density: 9.600121e-17<br />Petal.Width: 0.2174168","density: 8.592144e-17<br />Petal.Width: 0.2221135","density: 1.426672e-16<br />Petal.Width: 0.2268102","density: 1.580187e-16<br />Petal.Width: 0.2315068","density: 5.722104e-17<br />Petal.Width: 0.2362035","density: 8.523748e-17<br />Petal.Width: 0.2409002","density: 1.032553e-16<br />Petal.Width: 0.2455969","density: 1.434006e-16<br />Petal.Width: 0.2502935","density: 2.845173e-16<br />Petal.Width: 0.2549902","density: 1.677997e-16<br />Petal.Width: 0.2596869","density: 5.448798e-17<br />Petal.Width: 0.2643836","density: 6.989055e-18<br />Petal.Width: 0.2690802","density: 7.754742e-17<br />Petal.Width: 0.2737769","density: 4.618324e-17<br />Petal.Width: 0.2784736","density: 0.000000e+00<br />Petal.Width: 0.2831703","density: 0.000000e+00<br />Petal.Width: 0.2878669","density: 0.000000e+00<br />Petal.Width: 0.2925636","density: 1.193177e-17<br />Petal.Width: 0.2972603","density: 2.558771e-17<br />Petal.Width: 0.3019569","density: 1.550795e-17<br />Petal.Width: 0.3066536","density: 5.428181e-18<br />Petal.Width: 0.3113503","density: 0.000000e+00<br />Petal.Width: 0.3160470","density: 5.121385e-18<br />Petal.Width: 0.3207436","density: 6.559998e-17<br />Petal.Width: 0.3254403","density: 1.189433e-16<br />Petal.Width: 0.3301370","density: 1.159455e-16<br />Petal.Width: 0.3348337","density: 5.068072e-18<br />Petal.Width: 0.3395303","density: 0.000000e+00<br />Petal.Width: 0.3442270","density: 2.328404e-17<br />Petal.Width: 0.3489237","density: 5.348816e-17<br />Petal.Width: 0.3536204","density: 4.340840e-17<br />Petal.Width: 0.3583170","density: 6.654757e-17<br />Petal.Width: 0.3630137","density: 5.622645e-17<br />Petal.Width: 0.3677104","density: 0.000000e+00<br />Petal.Width: 0.3724070","density: 0.000000e+00<br />Petal.Width: 0.3771037","density: 0.000000e+00<br />Petal.Width: 0.3818004","density: 0.000000e+00<br />Petal.Width: 0.3864971","density: 0.000000e+00<br />Petal.Width: 0.3911937","density: 0.000000e+00<br />Petal.Width: 0.3958904","density: 0.000000e+00<br />Petal.Width: 0.4005871","density: 1.878117e-18<br />Petal.Width: 0.4052838","density: 1.195788e-17<br />Petal.Width: 0.4099804","density: 1.387779e-17<br />Petal.Width: 0.4146771","density: 9.515947e-18<br />Petal.Width: 0.4193738","density: 1.691458e-18<br />Petal.Width: 0.4240705","density: 3.193076e-17<br />Petal.Width: 0.4287671","density: 5.532449e-17<br />Petal.Width: 0.4334638","density: 7.853157e-17<br />Petal.Width: 0.4381605","density: 1.087709e-16<br />Petal.Width: 0.4428571","density: 7.370515e-17<br />Petal.Width: 0.4475538","density: 7.763622e-17<br />Petal.Width: 0.4522505","density: 1.179553e-16<br />Petal.Width: 0.4569472","density: 1.582743e-16<br />Petal.Width: 0.4616438","density: 1.585185e-16<br />Petal.Width: 0.4663405","density: 1.906084e-16<br />Petal.Width: 0.4710372","density: 2.809073e-16<br />Petal.Width: 0.4757339","density: 3.615455e-16<br />Petal.Width: 0.4804305","density: 4.823877e-16<br />Petal.Width: 0.4851272","density: 6.409205e-16<br />Petal.Width: 0.4898239","density: 8.425158e-16<br />Petal.Width: 0.4945205","density: 1.282906e-15<br />Petal.Width: 0.4992172","density: 1.821985e-15<br />Petal.Width: 0.5039139","density: 2.471245e-15<br />Petal.Width: 0.5086106","density: 3.358265e-15<br />Petal.Width: 0.5133072","density: 4.654530e-15<br />Petal.Width: 0.5180039","density: 6.424042e-15<br />Petal.Width: 0.5227006","density: 8.753473e-15<br />Petal.Width: 0.5273973","density: 1.220075e-14<br />Petal.Width: 0.5320939","density: 1.686742e-14<br />Petal.Width: 0.5367906","density: 2.291759e-14<br />Petal.Width: 0.5414873","density: 3.090077e-14<br />Petal.Width: 0.5461840","density: 4.286134e-14<br />Petal.Width: 0.5508806","density: 5.858177e-14<br />Petal.Width: 0.5555773","density: 7.874183e-14<br />Petal.Width: 0.5602740","density: 1.072272e-13<br />Petal.Width: 0.5649706","density: 1.468679e-13<br />Petal.Width: 0.5696673","density: 1.983835e-13<br />Petal.Width: 0.5743640","density: 2.635943e-13<br />Petal.Width: 0.5790607","density: 3.610052e-13<br />Petal.Width: 0.5837573","density: 4.892270e-13<br />Petal.Width: 0.5884540","density: 6.536905e-13<br />Petal.Width: 0.5931507","density: 8.713328e-13<br />Petal.Width: 0.5978474","density: 1.182583e-12<br />Petal.Width: 0.6025440","density: 1.585160e-12<br />Petal.Width: 0.6072407","density: 2.094317e-12<br />Petal.Width: 0.6119374","density: 2.802824e-12<br />Petal.Width: 0.6166341","density: 3.763060e-12<br />Petal.Width: 0.6213307","density: 4.989263e-12<br />Petal.Width: 0.6260274","density: 6.520097e-12<br />Petal.Width: 0.6307241","density: 8.757813e-12<br />Petal.Width: 0.6354207","density: 1.163360e-11<br />Petal.Width: 0.6401174","density: 1.525892e-11<br />Petal.Width: 0.6448141","density: 2.000022e-11<br />Petal.Width: 0.6495108","density: 2.658543e-11<br />Petal.Width: 0.6542074","density: 3.494571e-11<br />Petal.Width: 0.6589041","density: 4.535119e-11<br />Petal.Width: 0.6636008","density: 5.960336e-11<br />Petal.Width: 0.6682975","density: 7.841421e-11<br />Petal.Width: 0.6729941","density: 1.020084e-10<br />Petal.Width: 0.6776908","density: 1.312030e-10<br />Petal.Width: 0.6823875","density: 1.725823e-10<br />Petal.Width: 0.6870841","density: 2.247472e-10<br />Petal.Width: 0.6917808","density: 2.893995e-10<br />Petal.Width: 0.6964775","density: 3.728817e-10<br />Petal.Width: 0.7011742","density: 4.855798e-10<br />Petal.Width: 0.7058708","density: 6.260332e-10<br />Petal.Width: 0.7105675","density: 7.980712e-10<br />Petal.Width: 0.7152642","density: 1.029736e-09<br />Petal.Width: 0.7199609","density: 1.327740e-09<br />Petal.Width: 0.7246575","density: 1.694967e-09<br />Petal.Width: 0.7293542","density: 2.144881e-09<br />Petal.Width: 0.7340509","density: 2.763478e-09<br />Petal.Width: 0.7387476","density: 3.528650e-09<br />Petal.Width: 0.7434442","density: 4.461168e-09<br />Petal.Width: 0.7481409","density: 5.648768e-09<br />Petal.Width: 0.7528376","density: 7.207986e-09<br />Petal.Width: 0.7575342","density: 9.116036e-09<br />Petal.Width: 0.7622309","density: 1.141632e-08<br />Petal.Width: 0.7669276","density: 1.445837e-08<br />Petal.Width: 0.7716243","density: 1.827493e-08<br />Petal.Width: 0.7763209","density: 2.289645e-08<br />Petal.Width: 0.7810176","density: 2.849719e-08<br />Petal.Width: 0.7857143","density: 3.597112e-08<br />Petal.Width: 0.7904110","density: 4.504447e-08<br />Petal.Width: 0.7951076","density: 5.591933e-08<br />Petal.Width: 0.7998043","density: 6.956468e-08<br />Petal.Width: 0.8045010","density: 8.699931e-08<br />Petal.Width: 0.8091977","density: 1.079531e-07<br />Petal.Width: 0.8138943","density: 1.328183e-07<br />Petal.Width: 0.8185910","density: 1.650781e-07<br />Petal.Width: 0.8232877","density: 2.045816e-07<br />Petal.Width: 0.8279843","density: 2.515962e-07<br />Petal.Width: 0.8326810","density: 3.078980e-07<br />Petal.Width: 0.8373777","density: 3.808596e-07<br />Petal.Width: 0.8420744","density: 4.678133e-07<br />Petal.Width: 0.8467710","density: 5.703224e-07<br />Petal.Width: 0.8514677","density: 6.969184e-07<br />Petal.Width: 0.8561644","density: 8.544375e-07<br />Petal.Width: 0.8608611","density: 1.040408e-06<br />Petal.Width: 0.8655577","density: 1.257648e-06<br />Petal.Width: 0.8702544","density: 1.533862e-06<br />Petal.Width: 0.8749511","density: 1.864253e-06<br />Petal.Width: 0.8796477","density: 2.250785e-06<br />Petal.Width: 0.8843444","density: 2.707735e-06<br />Petal.Width: 0.8890411","density: 3.283142e-06<br />Petal.Width: 0.8937378","density: 3.956510e-06<br />Petal.Width: 0.8984344","density: 4.737412e-06<br />Petal.Width: 0.9031311","density: 5.685633e-06<br />Petal.Width: 0.9078278","density: 6.835430e-06<br />Petal.Width: 0.9125245","density: 8.169223e-06<br />Petal.Width: 0.9172211","density: 9.703017e-06<br />Petal.Width: 0.9219178","density: 1.161235e-05<br />Petal.Width: 0.9266145","density: 1.384503e-05<br />Petal.Width: 0.9313112","density: 1.641318e-05<br />Petal.Width: 0.9360078","density: 1.940732e-05<br />Petal.Width: 0.9407045","density: 2.307337e-05<br />Petal.Width: 0.9454012","density: 2.728716e-05<br />Petal.Width: 0.9500978","density: 3.209482e-05<br />Petal.Width: 0.9547945","density: 3.782908e-05<br />Petal.Width: 0.9594912","density: 4.461055e-05<br />Petal.Width: 0.9641879","density: 5.234171e-05<br />Petal.Width: 0.9688845","density: 6.109398e-05<br />Petal.Width: 0.9735812","density: 7.175106e-05<br />Petal.Width: 0.9782779","density: 8.394450e-05<br />Petal.Width: 0.9829746","density: 9.773655e-05<br />Petal.Width: 0.9876712","density: 1.135796e-04<br />Petal.Width: 0.9923679","density: 1.324552e-04<br />Petal.Width: 0.9970646","density: 1.537705e-04<br />Petal.Width: 1.0017613","density: 1.776991e-04<br />Petal.Width: 1.0064579","density: 2.057078e-04<br />Petal.Width: 1.0111546","density: 2.380402e-04<br />Petal.Width: 1.0158513","density: 2.742739e-04<br />Petal.Width: 1.0205479","density: 3.146573e-04<br />Petal.Width: 1.0252446","density: 3.627140e-04<br />Petal.Width: 1.0299413","density: 4.165644e-04<br />Petal.Width: 1.0346380","density: 4.764718e-04<br />Petal.Width: 1.0393346","density: 5.442136e-04<br />Petal.Width: 1.0440313","density: 6.228112e-04<br />Petal.Width: 1.0487280","density: 7.100377e-04<br />Petal.Width: 1.0534247","density: 8.063963e-04<br />Petal.Width: 1.0581213","density: 9.169897e-04<br />Petal.Width: 1.0628180","density: 1.041720e-03<br />Petal.Width: 1.0675147","density: 1.179160e-03<br />Petal.Width: 1.0722114","density: 1.329984e-03<br />Petal.Width: 1.0769080","density: 1.505238e-03<br />Petal.Width: 1.0816047","density: 1.697773e-03<br />Petal.Width: 1.0863014","density: 1.908481e-03<br />Petal.Width: 1.0909980","density: 2.142830e-03<br />Petal.Width: 1.0956947","density: 2.407862e-03<br />Petal.Width: 1.1003914","density: 2.697015e-03<br />Petal.Width: 1.1050881","density: 3.011389e-03<br />Petal.Width: 1.1097847","density: 3.364918e-03<br />Petal.Width: 1.1144814","density: 3.754853e-03<br />Petal.Width: 1.1191781","density: 4.177447e-03<br />Petal.Width: 1.1238748","density: 4.634919e-03<br />Petal.Width: 1.1285714","density: 5.151875e-03<br />Petal.Width: 1.1332681","density: 5.710153e-03<br />Petal.Width: 1.1379648","density: 6.311322e-03<br />Petal.Width: 1.1426614","density: 6.968463e-03<br />Petal.Width: 1.1473581","density: 7.693573e-03<br />Petal.Width: 1.1520548","density: 8.471561e-03<br />Petal.Width: 1.1567515","density: 9.304163e-03<br />Petal.Width: 1.1614481","density: 1.022094e-02<br />Petal.Width: 1.1661448","density: 1.121088e-02<br />Petal.Width: 1.1708415","density: 1.226637e-02<br />Petal.Width: 1.1755382","density: 1.339264e-02<br />Petal.Width: 1.1802348","density: 1.463156e-02<br />Petal.Width: 1.1849315","density: 1.594728e-02<br />Petal.Width: 1.1896282","density: 1.734163e-02<br />Petal.Width: 1.1943249","density: 1.883858e-02<br />Petal.Width: 1.1990215","density: 2.045180e-02<br />Petal.Width: 1.2037182","density: 2.215452e-02<br />Petal.Width: 1.2084149","density: 2.394843e-02<br />Petal.Width: 1.2131115","density: 2.588127e-02<br />Petal.Width: 1.2178082","density: 2.792655e-02<br />Petal.Width: 1.2225049","density: 3.007244e-02<br />Petal.Width: 1.2272016","density: 3.232776e-02<br />Petal.Width: 1.2318982","density: 3.474487e-02<br />Petal.Width: 1.2365949","density: 3.726991e-02<br />Petal.Width: 1.2412916","density: 3.990377e-02<br />Petal.Width: 1.2459883","density: 4.267870e-02<br />Petal.Width: 1.2506849","density: 4.560188e-02<br />Petal.Width: 1.2553816","density: 4.863755e-02<br />Petal.Width: 1.2600783","density: 5.178601e-02<br />Petal.Width: 1.2647750","density: 5.510262e-02<br />Petal.Width: 1.2694716","density: 5.854470e-02<br />Petal.Width: 1.2741683","density: 6.209850e-02<br />Petal.Width: 1.2788650","density: 6.577370e-02<br />Petal.Width: 1.2835616","density: 6.961347e-02<br />Petal.Width: 1.2882583","density: 7.355936e-02<br />Petal.Width: 1.2929550","density: 7.760989e-02<br />Petal.Width: 1.2976517","density: 8.179314e-02<br />Petal.Width: 1.3023483","density: 8.610282e-02<br />Petal.Width: 1.3070450","density: 9.050564e-02<br />Petal.Width: 1.3117417","density: 9.499924e-02<br />Petal.Width: 1.3164384","density: 9.962111e-02<br />Petal.Width: 1.3211350","density: 1.043277e-01<br />Petal.Width: 1.3258317","density: 1.091077e-01<br />Petal.Width: 1.3305284","density: 1.139651e-01<br />Petal.Width: 1.3352250","density: 1.189125e-01<br />Petal.Width: 1.3399217","density: 1.239119e-01<br />Petal.Width: 1.3446184","density: 1.289594e-01<br />Petal.Width: 1.3493151","density: 1.340611e-01<br />Petal.Width: 1.3540117","density: 1.392015e-01<br />Petal.Width: 1.3587084","density: 1.443648e-01<br />Petal.Width: 1.3634051","density: 1.495469e-01<br />Petal.Width: 1.3681018","density: 1.547418e-01<br />Petal.Width: 1.3727984","density: 1.599339e-01<br />Petal.Width: 1.3774951","density: 1.651181e-01<br />Petal.Width: 1.3821918","density: 1.702867e-01<br />Petal.Width: 1.3868885","density: 1.754195e-01<br />Petal.Width: 1.3915851","density: 1.805191e-01<br />Petal.Width: 1.3962818","density: 1.855816e-01<br />Petal.Width: 1.4009785","density: 1.905844e-01<br />Petal.Width: 1.4056751","density: 1.955207e-01<br />Petal.Width: 1.4103718","density: 2.003990e-01<br />Petal.Width: 1.4150685","density: 2.052162e-01<br />Petal.Width: 1.4197652","density: 2.099290e-01<br />Petal.Width: 1.4244618","density: 2.145662e-01<br />Petal.Width: 1.4291585","density: 2.191298e-01<br />Petal.Width: 1.4338552","density: 2.236058e-01<br />Petal.Width: 1.4385519","density: 2.279707e-01<br />Petal.Width: 1.4432485","density: 2.322591e-01<br />Petal.Width: 1.4479452","density: 2.364717e-01<br />Petal.Width: 1.4526419","density: 2.405832e-01<br />Petal.Width: 1.4573386","density: 2.446099e-01<br />Petal.Width: 1.4620352","density: 2.485726e-01<br />Petal.Width: 1.4667319","density: 2.524744e-01<br />Petal.Width: 1.4714286","density: 2.562953e-01<br />Petal.Width: 1.4761252","density: 2.600769e-01<br />Petal.Width: 1.4808219","density: 2.638265e-01<br />Petal.Width: 1.4855186","density: 2.675489e-01<br />Petal.Width: 1.4902153","density: 2.712642e-01<br />Petal.Width: 1.4949119","density: 2.749915e-01<br />Petal.Width: 1.4996086","density: 2.787396e-01<br />Petal.Width: 1.5043053","density: 2.825425e-01<br />Petal.Width: 1.5090020","density: 2.864240e-01<br />Petal.Width: 1.5136986","density: 2.903878e-01<br />Petal.Width: 1.5183953","density: 2.944448e-01<br />Petal.Width: 1.5230920","density: 2.987001e-01<br />Petal.Width: 1.5277886","density: 3.031114e-01<br />Petal.Width: 1.5324853","density: 3.076918e-01<br />Petal.Width: 1.5371820","density: 3.125013e-01<br />Petal.Width: 1.5418787","density: 3.176243e-01<br />Petal.Width: 1.5465753","density: 3.230003e-01<br />Petal.Width: 1.5512720","density: 3.286433e-01<br />Petal.Width: 1.5559687","density: 3.347068e-01<br />Petal.Width: 1.5606654","density: 3.411651e-01<br />Petal.Width: 1.5653620","density: 3.479767e-01<br />Petal.Width: 1.5700587","density: 3.551646e-01<br />Petal.Width: 1.5747554","density: 3.629895e-01<br />Petal.Width: 1.5794521","density: 3.712487e-01<br />Petal.Width: 1.5841487","density: 3.799544e-01<br />Petal.Width: 1.5888454","density: 3.892333e-01<br />Petal.Width: 1.5935421","density: 3.991891e-01<br />Petal.Width: 1.5982387","density: 4.096563e-01<br />Petal.Width: 1.6029354","density: 4.206437e-01<br />Petal.Width: 1.6076321","density: 4.324047e-01<br />Petal.Width: 1.6123288","density: 4.448086e-01<br />Petal.Width: 1.6170254","density: 4.577707e-01<br />Petal.Width: 1.6217221","density: 4.713209e-01<br />Petal.Width: 1.6264188","density: 4.857562e-01<br />Petal.Width: 1.6311155","density: 5.007581e-01<br />Petal.Width: 1.6358121","density: 5.163238e-01<br />Petal.Width: 1.6405088","density: 5.325910e-01<br />Petal.Width: 1.6452055","density: 5.495945e-01<br />Petal.Width: 1.6499022","density: 5.671280e-01<br />Petal.Width: 1.6545988","density: 5.851825e-01<br />Petal.Width: 1.6592955","density: 6.039654e-01<br />Petal.Width: 1.6639922","density: 6.232774e-01<br />Petal.Width: 1.6686888","density: 6.430316e-01<br />Petal.Width: 1.6733855","density: 6.632381e-01<br />Petal.Width: 1.6780822","density: 6.840235e-01<br />Petal.Width: 1.6827789","density: 7.051368e-01<br />Petal.Width: 1.6874755","density: 7.265555e-01<br />Petal.Width: 1.6921722","density: 7.483144e-01<br />Petal.Width: 1.6968689","density: 7.703438e-01<br />Petal.Width: 1.7015656","density: 7.925285e-01<br />Petal.Width: 1.7062622","density: 8.148419e-01<br />Petal.Width: 1.7109589","density: 8.372545e-01<br />Petal.Width: 1.7156556","density: 8.596604e-01<br />Petal.Width: 1.7203523","density: 8.820223e-01<br />Petal.Width: 1.7250489","density: 9.042966e-01<br />Petal.Width: 1.7297456","density: 9.263362e-01<br />Petal.Width: 1.7344423","density: 9.481529e-01<br />Petal.Width: 1.7391389","density: 9.697177e-01<br />Petal.Width: 1.7438356","density: 9.908884e-01<br />Petal.Width: 1.7485323","density: 1.011572e+00<br />Petal.Width: 1.7532290","density: 1.031834e+00<br />Petal.Width: 1.7579256","density: 1.051648e+00<br />Petal.Width: 1.7626223","density: 1.070701e+00<br />Petal.Width: 1.7673190","density: 1.089136e+00<br />Petal.Width: 1.7720157","density: 1.106982e+00<br />Petal.Width: 1.7767123","density: 1.124141e+00<br />Petal.Width: 1.7814090","density: 1.140314e+00<br />Petal.Width: 1.7861057","density: 1.155789e+00<br />Petal.Width: 1.7908023","density: 1.170553e+00<br />Petal.Width: 1.7954990","density: 1.184346e+00<br />Petal.Width: 1.8001957","density: 1.197169e+00<br />Petal.Width: 1.8048924","density: 1.209223e+00<br />Petal.Width: 1.8095890","density: 1.220501e+00<br />Petal.Width: 1.8142857","density: 1.230580e+00<br />Petal.Width: 1.8189824","density: 1.239819e+00<br />Petal.Width: 1.8236791","density: 1.248278e+00<br />Petal.Width: 1.8283757","density: 1.255853e+00<br />Petal.Width: 1.8330724","density: 1.262309e+00<br />Petal.Width: 1.8377691","density: 1.268020e+00<br />Petal.Width: 1.8424658","density: 1.272996e+00<br />Petal.Width: 1.8471624","density: 1.277007e+00<br />Petal.Width: 1.8518591","density: 1.280158e+00<br />Petal.Width: 1.8565558","density: 1.282650e+00<br />Petal.Width: 1.8612524","density: 1.284497e+00<br />Petal.Width: 1.8659491","density: 1.285399e+00<br />Petal.Width: 1.8706458","density: 1.285712e+00<br />Petal.Width: 1.8753425","density: 1.285482e+00<br />Petal.Width: 1.8800391","density: 1.284652e+00<br />Petal.Width: 1.8847358","density: 1.283144e+00<br />Petal.Width: 1.8894325","density: 1.281206e+00<br />Petal.Width: 1.8941292","density: 1.278857e+00<br />Petal.Width: 1.8988258","density: 1.275996e+00<br />Petal.Width: 1.9035225","density: 1.272730e+00<br />Petal.Width: 1.9082192","density: 1.269165e+00<br />Petal.Width: 1.9129159","density: 1.265318e+00<br />Petal.Width: 1.9176125","density: 1.261094e+00<br />Petal.Width: 1.9223092","density: 1.256666e+00<br />Petal.Width: 1.9270059","density: 1.252054e+00<br />Petal.Width: 1.9317025","density: 1.247252e+00<br />Petal.Width: 1.9363992","density: 1.242269e+00<br />Petal.Width: 1.9410959","density: 1.237183e+00<br />Petal.Width: 1.9457926","density: 1.232007e+00<br />Petal.Width: 1.9504892","density: 1.226734e+00<br />Petal.Width: 1.9551859","density: 1.221406e+00<br />Petal.Width: 1.9598826","density: 1.216044e+00<br />Petal.Width: 1.9645793","density: 1.210656e+00<br />Petal.Width: 1.9692759","density: 1.205253e+00<br />Petal.Width: 1.9739726","density: 1.199854e+00<br />Petal.Width: 1.9786693","density: 1.194464e+00<br />Petal.Width: 1.9833659","density: 1.189092e+00<br />Petal.Width: 1.9880626","density: 1.183752e+00<br />Petal.Width: 1.9927593","density: 1.178440e+00<br />Petal.Width: 1.9974560","density: 1.173159e+00<br />Petal.Width: 2.0021526","density: 1.167925e+00<br />Petal.Width: 2.0068493","density: 1.162735e+00<br />Petal.Width: 2.0115460","density: 1.157582e+00<br />Petal.Width: 2.0162427","density: 1.152470e+00<br />Petal.Width: 2.0209393","density: 1.147422e+00<br />Petal.Width: 2.0256360","density: 1.142416e+00<br />Petal.Width: 2.0303327","density: 1.137453e+00<br />Petal.Width: 2.0350294","density: 1.132545e+00<br />Petal.Width: 2.0397260","density: 1.127700e+00<br />Petal.Width: 2.0444227","density: 1.122904e+00<br />Petal.Width: 2.0491194","density: 1.118158e+00<br />Petal.Width: 2.0538160","density: 1.113489e+00<br />Petal.Width: 2.0585127","density: 1.108887e+00<br />Petal.Width: 2.0632094","density: 1.104348e+00<br />Petal.Width: 2.0679061","density: 1.099876e+00<br />Petal.Width: 2.0726027","density: 1.095515e+00<br />Petal.Width: 2.0772994","density: 1.091234e+00<br />Petal.Width: 2.0819961","density: 1.087036e+00<br />Petal.Width: 2.0866928","density: 1.082949e+00<br />Petal.Width: 2.0913894","density: 1.078994e+00<br />Petal.Width: 2.0960861","density: 1.075146e+00<br />Petal.Width: 2.1007828","density: 1.071407e+00<br />Petal.Width: 2.1054795","density: 1.067842e+00<br />Petal.Width: 2.1101761","density: 1.064422e+00<br />Petal.Width: 2.1148728","density: 1.061136e+00<br />Petal.Width: 2.1195695","density: 1.057997e+00<br />Petal.Width: 2.1242661","density: 1.055082e+00<br />Petal.Width: 2.1289628","density: 1.052318e+00<br />Petal.Width: 2.1336595","density: 1.049709e+00<br />Petal.Width: 2.1383562","density: 1.047301e+00<br />Petal.Width: 2.1430528","density: 1.045107e+00<br />Petal.Width: 2.1477495","density: 1.043073e+00<br />Petal.Width: 2.1524462","density: 1.041200e+00<br />Petal.Width: 2.1571429","density: 1.039563e+00<br />Petal.Width: 2.1618395","density: 1.038099e+00<br />Petal.Width: 2.1665362","density: 1.036784e+00<br />Petal.Width: 2.1712329","density: 1.035628e+00<br />Petal.Width: 2.1759295","density: 1.034674e+00<br />Petal.Width: 2.1806262","density: 1.033842e+00<br />Petal.Width: 2.1853229","density: 1.033123e+00<br />Petal.Width: 2.1900196","density: 1.032535e+00<br />Petal.Width: 2.1947162","density: 1.032052e+00<br />Petal.Width: 2.1994129","density: 1.031632e+00<br />Petal.Width: 2.2041096","density: 1.031266e+00<br />Petal.Width: 2.2088063","density: 1.030942e+00<br />Petal.Width: 2.2135029","density: 1.030619e+00<br />Petal.Width: 2.2181996","density: 1.030280e+00<br />Petal.Width: 2.2228963","density: 1.029905e+00<br />Petal.Width: 2.2275930","density: 1.029435e+00<br />Petal.Width: 2.2322896","density: 1.028870e+00<br />Petal.Width: 2.2369863","density: 1.028197e+00<br />Petal.Width: 2.2416830","density: 1.027349e+00<br />Petal.Width: 2.2463796","density: 1.026287e+00<br />Petal.Width: 2.2510763","density: 1.025034e+00<br />Petal.Width: 2.2557730","density: 1.023578e+00<br />Petal.Width: 2.2604697","density: 1.021764e+00<br />Petal.Width: 2.2651663","density: 1.019666e+00<br />Petal.Width: 2.2698630","density: 1.017289e+00<br />Petal.Width: 2.2745597","density: 1.014577e+00<br />Petal.Width: 2.2792564","density: 1.011395e+00<br />Petal.Width: 2.2839530","density: 1.007872e+00<br />Petal.Width: 2.2886497","density: 1.003999e+00<br />Petal.Width: 2.2933464","density: 9.996353e-01<br />Petal.Width: 2.2980431","density: 9.947914e-01<br />Petal.Width: 2.3027397","density: 9.895574e-01<br />Petal.Width: 2.3074364","density: 9.839286e-01<br />Petal.Width: 2.3121331","density: 9.776725e-01<br />Petal.Width: 2.3168297","density: 9.709870e-01<br />Petal.Width: 2.3215264","density: 9.638921e-01<br />Petal.Width: 2.3262231","density: 9.563219e-01<br />Petal.Width: 2.3309198","density: 9.481626e-01<br />Petal.Width: 2.3356164","density: 9.396009e-01<br />Petal.Width: 2.3403131","density: 9.306400e-01<br />Petal.Width: 2.3450098","density: 9.211431e-01<br />Petal.Width: 2.3497065","density: 9.111775e-01<br />Petal.Width: 2.3544031","density: 9.008426e-01<br />Petal.Width: 2.3590998","density: 8.901447e-01<br />Petal.Width: 2.3637965","density: 8.789017e-01<br />Petal.Width: 2.3684932","density: 8.673245e-01<br />Petal.Width: 2.3731898","density: 8.554310e-01<br />Petal.Width: 2.3778865","density: 8.431787e-01<br />Petal.Width: 2.3825832","density: 8.305273e-01<br />Petal.Width: 2.3872798","density: 8.176151e-01<br />Petal.Width: 2.3919765","density: 8.044517e-01<br />Petal.Width: 2.3966732","density: 7.909657e-01<br />Petal.Width: 2.4013699","density: 7.772286e-01<br />Petal.Width: 2.4060665","density: 7.632996e-01<br />Petal.Width: 2.4107632","density: 7.491885e-01<br />Petal.Width: 2.4154599","density: 7.348248e-01<br />Petal.Width: 2.4201566","density: 7.203268e-01<br />Petal.Width: 2.4248532","density: 7.057040e-01<br />Petal.Width: 2.4295499","density: 6.909478e-01<br />Petal.Width: 2.4342466","density: 6.760677e-01<br />Petal.Width: 2.4389432","density: 6.611160e-01<br />Petal.Width: 2.4436399","density: 6.461009e-01<br />Petal.Width: 2.4483366","density: 6.310193e-01<br />Petal.Width: 2.4530333","density: 6.159047e-01<br />Petal.Width: 2.4577299","density: 6.007729e-01<br />Petal.Width: 2.4624266","density: 5.856313e-01<br />Petal.Width: 2.4671233","density: 5.705018e-01<br />Petal.Width: 2.4718200","density: 5.553959e-01<br />Petal.Width: 2.4765166","density: 5.403197e-01<br />Petal.Width: 2.4812133","density: 5.252920e-01<br />Petal.Width: 2.4859100","density: 5.103388e-01<br />Petal.Width: 2.4906067","density: 4.954508e-01<br />Petal.Width: 2.4953033","density: 4.806334e-01<br />Petal.Width: 2.5000000","density: 4.806334e-01<br />Petal.Width: 2.5000000","density: 4.806334e-01<br />Petal.Width: 2.5000000","density: 4.954508e-01<br />Petal.Width: 2.4953033","density: 5.103388e-01<br />Petal.Width: 2.4906067","density: 5.252920e-01<br />Petal.Width: 2.4859100","density: 5.403197e-01<br />Petal.Width: 2.4812133","density: 5.553959e-01<br />Petal.Width: 2.4765166","density: 5.705018e-01<br />Petal.Width: 2.4718200","density: 5.856313e-01<br />Petal.Width: 2.4671233","density: 6.007729e-01<br />Petal.Width: 2.4624266","density: 6.159047e-01<br />Petal.Width: 2.4577299","density: 6.310193e-01<br />Petal.Width: 2.4530333","density: 6.461009e-01<br />Petal.Width: 2.4483366","density: 6.611160e-01<br />Petal.Width: 2.4436399","density: 6.760677e-01<br />Petal.Width: 2.4389432","density: 6.909478e-01<br />Petal.Width: 2.4342466","density: 7.057040e-01<br />Petal.Width: 2.4295499","density: 7.203268e-01<br />Petal.Width: 2.4248532","density: 7.348248e-01<br />Petal.Width: 2.4201566","density: 7.491885e-01<br />Petal.Width: 2.4154599","density: 7.632996e-01<br />Petal.Width: 2.4107632","density: 7.772286e-01<br />Petal.Width: 2.4060665","density: 7.909657e-01<br />Petal.Width: 2.4013699","density: 8.044517e-01<br />Petal.Width: 2.3966732","density: 8.176151e-01<br />Petal.Width: 2.3919765","density: 8.305273e-01<br />Petal.Width: 2.3872798","density: 8.431787e-01<br />Petal.Width: 2.3825832","density: 8.554310e-01<br />Petal.Width: 2.3778865","density: 8.673245e-01<br />Petal.Width: 2.3731898","density: 8.789017e-01<br />Petal.Width: 2.3684932","density: 8.901447e-01<br />Petal.Width: 2.3637965","density: 9.008426e-01<br />Petal.Width: 2.3590998","density: 9.111775e-01<br />Petal.Width: 2.3544031","density: 9.211431e-01<br />Petal.Width: 2.3497065","density: 9.306400e-01<br />Petal.Width: 2.3450098","density: 9.396009e-01<br />Petal.Width: 2.3403131","density: 9.481626e-01<br />Petal.Width: 2.3356164","density: 9.563219e-01<br />Petal.Width: 2.3309198","density: 9.638921e-01<br />Petal.Width: 2.3262231","density: 9.709870e-01<br />Petal.Width: 2.3215264","density: 9.776725e-01<br />Petal.Width: 2.3168297","density: 9.839286e-01<br />Petal.Width: 2.3121331","density: 9.895574e-01<br />Petal.Width: 2.3074364","density: 9.947914e-01<br />Petal.Width: 2.3027397","density: 9.996353e-01<br />Petal.Width: 2.2980431","density: 1.003999e+00<br />Petal.Width: 2.2933464","density: 1.007872e+00<br />Petal.Width: 2.2886497","density: 1.011395e+00<br />Petal.Width: 2.2839530","density: 1.014577e+00<br />Petal.Width: 2.2792564","density: 1.017289e+00<br />Petal.Width: 2.2745597","density: 1.019666e+00<br />Petal.Width: 2.2698630","density: 1.021764e+00<br />Petal.Width: 2.2651663","density: 1.023578e+00<br />Petal.Width: 2.2604697","density: 1.025034e+00<br />Petal.Width: 2.2557730","density: 1.026287e+00<br />Petal.Width: 2.2510763","density: 1.027349e+00<br />Petal.Width: 2.2463796","density: 1.028197e+00<br />Petal.Width: 2.2416830","density: 1.028870e+00<br />Petal.Width: 2.2369863","density: 1.029435e+00<br />Petal.Width: 2.2322896","density: 1.029905e+00<br />Petal.Width: 2.2275930","density: 1.030280e+00<br />Petal.Width: 2.2228963","density: 1.030619e+00<br />Petal.Width: 2.2181996","density: 1.030942e+00<br />Petal.Width: 2.2135029","density: 1.031266e+00<br />Petal.Width: 2.2088063","density: 1.031632e+00<br />Petal.Width: 2.2041096","density: 1.032052e+00<br />Petal.Width: 2.1994129","density: 1.032535e+00<br />Petal.Width: 2.1947162","density: 1.033123e+00<br />Petal.Width: 2.1900196","density: 1.033842e+00<br />Petal.Width: 2.1853229","density: 1.034674e+00<br />Petal.Width: 2.1806262","density: 1.035628e+00<br />Petal.Width: 2.1759295","density: 1.036784e+00<br />Petal.Width: 2.1712329","density: 1.038099e+00<br />Petal.Width: 2.1665362","density: 1.039563e+00<br />Petal.Width: 2.1618395","density: 1.041200e+00<br />Petal.Width: 2.1571429","density: 1.043073e+00<br />Petal.Width: 2.1524462","density: 1.045107e+00<br />Petal.Width: 2.1477495","density: 1.047301e+00<br />Petal.Width: 2.1430528","density: 1.049709e+00<br />Petal.Width: 2.1383562","density: 1.052318e+00<br />Petal.Width: 2.1336595","density: 1.055082e+00<br />Petal.Width: 2.1289628","density: 1.057997e+00<br />Petal.Width: 2.1242661","density: 1.061136e+00<br />Petal.Width: 2.1195695","density: 1.064422e+00<br />Petal.Width: 2.1148728","density: 1.067842e+00<br />Petal.Width: 2.1101761","density: 1.071407e+00<br />Petal.Width: 2.1054795","density: 1.075146e+00<br />Petal.Width: 2.1007828","density: 1.078994e+00<br />Petal.Width: 2.0960861","density: 1.082949e+00<br />Petal.Width: 2.0913894","density: 1.087036e+00<br />Petal.Width: 2.0866928","density: 1.091234e+00<br />Petal.Width: 2.0819961","density: 1.095515e+00<br />Petal.Width: 2.0772994","density: 1.099876e+00<br />Petal.Width: 2.0726027","density: 1.104348e+00<br />Petal.Width: 2.0679061","density: 1.108887e+00<br />Petal.Width: 2.0632094","density: 1.113489e+00<br />Petal.Width: 2.0585127","density: 1.118158e+00<br />Petal.Width: 2.0538160","density: 1.122904e+00<br />Petal.Width: 2.0491194","density: 1.127700e+00<br />Petal.Width: 2.0444227","density: 1.132545e+00<br />Petal.Width: 2.0397260","density: 1.137453e+00<br />Petal.Width: 2.0350294","density: 1.142416e+00<br />Petal.Width: 2.0303327","density: 1.147422e+00<br />Petal.Width: 2.0256360","density: 1.152470e+00<br />Petal.Width: 2.0209393","density: 1.157582e+00<br />Petal.Width: 2.0162427","density: 1.162735e+00<br />Petal.Width: 2.0115460","density: 1.167925e+00<br />Petal.Width: 2.0068493","density: 1.173159e+00<br />Petal.Width: 2.0021526","density: 1.178440e+00<br />Petal.Width: 1.9974560","density: 1.183752e+00<br />Petal.Width: 1.9927593","density: 1.189092e+00<br />Petal.Width: 1.9880626","density: 1.194464e+00<br />Petal.Width: 1.9833659","density: 1.199854e+00<br />Petal.Width: 1.9786693","density: 1.205253e+00<br />Petal.Width: 1.9739726","density: 1.210656e+00<br />Petal.Width: 1.9692759","density: 1.216044e+00<br />Petal.Width: 1.9645793","density: 1.221406e+00<br />Petal.Width: 1.9598826","density: 1.226734e+00<br />Petal.Width: 1.9551859","density: 1.232007e+00<br />Petal.Width: 1.9504892","density: 1.237183e+00<br />Petal.Width: 1.9457926","density: 1.242269e+00<br />Petal.Width: 1.9410959","density: 1.247252e+00<br />Petal.Width: 1.9363992","density: 1.252054e+00<br />Petal.Width: 1.9317025","density: 1.256666e+00<br />Petal.Width: 1.9270059","density: 1.261094e+00<br />Petal.Width: 1.9223092","density: 1.265318e+00<br />Petal.Width: 1.9176125","density: 1.269165e+00<br />Petal.Width: 1.9129159","density: 1.272730e+00<br />Petal.Width: 1.9082192","density: 1.275996e+00<br />Petal.Width: 1.9035225","density: 1.278857e+00<br />Petal.Width: 1.8988258","density: 1.281206e+00<br />Petal.Width: 1.8941292","density: 1.283144e+00<br />Petal.Width: 1.8894325","density: 1.284652e+00<br />Petal.Width: 1.8847358","density: 1.285482e+00<br />Petal.Width: 1.8800391","density: 1.285712e+00<br />Petal.Width: 1.8753425","density: 1.285399e+00<br />Petal.Width: 1.8706458","density: 1.284497e+00<br />Petal.Width: 1.8659491","density: 1.282650e+00<br />Petal.Width: 1.8612524","density: 1.280158e+00<br />Petal.Width: 1.8565558","density: 1.277007e+00<br />Petal.Width: 1.8518591","density: 1.272996e+00<br />Petal.Width: 1.8471624","density: 1.268020e+00<br />Petal.Width: 1.8424658","density: 1.262309e+00<br />Petal.Width: 1.8377691","density: 1.255853e+00<br />Petal.Width: 1.8330724","density: 1.248278e+00<br />Petal.Width: 1.8283757","density: 1.239819e+00<br />Petal.Width: 1.8236791","density: 1.230580e+00<br />Petal.Width: 1.8189824","density: 1.220501e+00<br />Petal.Width: 1.8142857","density: 1.209223e+00<br />Petal.Width: 1.8095890","density: 1.197169e+00<br />Petal.Width: 1.8048924","density: 1.184346e+00<br />Petal.Width: 1.8001957","density: 1.170553e+00<br />Petal.Width: 1.7954990","density: 1.155789e+00<br />Petal.Width: 1.7908023","density: 1.140314e+00<br />Petal.Width: 1.7861057","density: 1.124141e+00<br />Petal.Width: 1.7814090","density: 1.106982e+00<br />Petal.Width: 1.7767123","density: 1.089136e+00<br />Petal.Width: 1.7720157","density: 1.070701e+00<br />Petal.Width: 1.7673190","density: 1.051648e+00<br />Petal.Width: 1.7626223","density: 1.031834e+00<br />Petal.Width: 1.7579256","density: 1.011572e+00<br />Petal.Width: 1.7532290","density: 9.908884e-01<br />Petal.Width: 1.7485323","density: 9.697177e-01<br />Petal.Width: 1.7438356","density: 9.481529e-01<br />Petal.Width: 1.7391389","density: 9.263362e-01<br />Petal.Width: 1.7344423","density: 9.042966e-01<br />Petal.Width: 1.7297456","density: 8.820223e-01<br />Petal.Width: 1.7250489","density: 8.596604e-01<br />Petal.Width: 1.7203523","density: 8.372545e-01<br />Petal.Width: 1.7156556","density: 8.148419e-01<br />Petal.Width: 1.7109589","density: 7.925285e-01<br />Petal.Width: 1.7062622","density: 7.703438e-01<br />Petal.Width: 1.7015656","density: 7.483144e-01<br />Petal.Width: 1.6968689","density: 7.265555e-01<br />Petal.Width: 1.6921722","density: 7.051368e-01<br />Petal.Width: 1.6874755","density: 6.840235e-01<br />Petal.Width: 1.6827789","density: 6.632381e-01<br />Petal.Width: 1.6780822","density: 6.430316e-01<br />Petal.Width: 1.6733855","density: 6.232774e-01<br />Petal.Width: 1.6686888","density: 6.039654e-01<br />Petal.Width: 1.6639922","density: 5.851825e-01<br />Petal.Width: 1.6592955","density: 5.671280e-01<br />Petal.Width: 1.6545988","density: 5.495945e-01<br />Petal.Width: 1.6499022","density: 5.325910e-01<br />Petal.Width: 1.6452055","density: 5.163238e-01<br />Petal.Width: 1.6405088","density: 5.007581e-01<br />Petal.Width: 1.6358121","density: 4.857562e-01<br />Petal.Width: 1.6311155","density: 4.713209e-01<br />Petal.Width: 1.6264188","density: 4.577707e-01<br />Petal.Width: 1.6217221","density: 4.448086e-01<br />Petal.Width: 1.6170254","density: 4.324047e-01<br />Petal.Width: 1.6123288","density: 4.206437e-01<br />Petal.Width: 1.6076321","density: 4.096563e-01<br />Petal.Width: 1.6029354","density: 3.991891e-01<br />Petal.Width: 1.5982387","density: 3.892333e-01<br />Petal.Width: 1.5935421","density: 3.799544e-01<br />Petal.Width: 1.5888454","density: 3.712487e-01<br />Petal.Width: 1.5841487","density: 3.629895e-01<br />Petal.Width: 1.5794521","density: 3.551646e-01<br />Petal.Width: 1.5747554","density: 3.479767e-01<br />Petal.Width: 1.5700587","density: 3.411651e-01<br />Petal.Width: 1.5653620","density: 3.347068e-01<br />Petal.Width: 1.5606654","density: 3.286433e-01<br />Petal.Width: 1.5559687","density: 3.230003e-01<br />Petal.Width: 1.5512720","density: 3.176243e-01<br />Petal.Width: 1.5465753","density: 3.125013e-01<br />Petal.Width: 1.5418787","density: 3.076918e-01<br />Petal.Width: 1.5371820","density: 3.031114e-01<br />Petal.Width: 1.5324853","density: 2.987001e-01<br />Petal.Width: 1.5277886","density: 2.944448e-01<br />Petal.Width: 1.5230920","density: 2.903878e-01<br />Petal.Width: 1.5183953","density: 2.864240e-01<br />Petal.Width: 1.5136986","density: 2.825425e-01<br />Petal.Width: 1.5090020","density: 2.787396e-01<br />Petal.Width: 1.5043053","density: 2.749915e-01<br />Petal.Width: 1.4996086","density: 2.712642e-01<br />Petal.Width: 1.4949119","density: 2.675489e-01<br />Petal.Width: 1.4902153","density: 2.638265e-01<br />Petal.Width: 1.4855186","density: 2.600769e-01<br />Petal.Width: 1.4808219","density: 2.562953e-01<br />Petal.Width: 1.4761252","density: 2.524744e-01<br />Petal.Width: 1.4714286","density: 2.485726e-01<br />Petal.Width: 1.4667319","density: 2.446099e-01<br />Petal.Width: 1.4620352","density: 2.405832e-01<br />Petal.Width: 1.4573386","density: 2.364717e-01<br />Petal.Width: 1.4526419","density: 2.322591e-01<br />Petal.Width: 1.4479452","density: 2.279707e-01<br />Petal.Width: 1.4432485","density: 2.236058e-01<br />Petal.Width: 1.4385519","density: 2.191298e-01<br />Petal.Width: 1.4338552","density: 2.145662e-01<br />Petal.Width: 1.4291585","density: 2.099290e-01<br />Petal.Width: 1.4244618","density: 2.052162e-01<br />Petal.Width: 1.4197652","density: 2.003990e-01<br />Petal.Width: 1.4150685","density: 1.955207e-01<br />Petal.Width: 1.4103718","density: 1.905844e-01<br />Petal.Width: 1.4056751","density: 1.855816e-01<br />Petal.Width: 1.4009785","density: 1.805191e-01<br />Petal.Width: 1.3962818","density: 1.754195e-01<br />Petal.Width: 1.3915851","density: 1.702867e-01<br />Petal.Width: 1.3868885","density: 1.651181e-01<br />Petal.Width: 1.3821918","density: 1.599339e-01<br />Petal.Width: 1.3774951","density: 1.547418e-01<br />Petal.Width: 1.3727984","density: 1.495469e-01<br />Petal.Width: 1.3681018","density: 1.443648e-01<br />Petal.Width: 1.3634051","density: 1.392015e-01<br />Petal.Width: 1.3587084","density: 1.340611e-01<br />Petal.Width: 1.3540117","density: 1.289594e-01<br />Petal.Width: 1.3493151","density: 1.239119e-01<br />Petal.Width: 1.3446184","density: 1.189125e-01<br />Petal.Width: 1.3399217","density: 1.139651e-01<br />Petal.Width: 1.3352250","density: 1.091077e-01<br />Petal.Width: 1.3305284","density: 1.043277e-01<br />Petal.Width: 1.3258317","density: 9.962111e-02<br />Petal.Width: 1.3211350","density: 9.499924e-02<br />Petal.Width: 1.3164384","density: 9.050564e-02<br />Petal.Width: 1.3117417","density: 8.610282e-02<br />Petal.Width: 1.3070450","density: 8.179314e-02<br />Petal.Width: 1.3023483","density: 7.760989e-02<br />Petal.Width: 1.2976517","density: 7.355936e-02<br />Petal.Width: 1.2929550","density: 6.961347e-02<br />Petal.Width: 1.2882583","density: 6.577370e-02<br />Petal.Width: 1.2835616","density: 6.209850e-02<br />Petal.Width: 1.2788650","density: 5.854470e-02<br />Petal.Width: 1.2741683","density: 5.510262e-02<br />Petal.Width: 1.2694716","density: 5.178601e-02<br />Petal.Width: 1.2647750","density: 4.863755e-02<br />Petal.Width: 1.2600783","density: 4.560188e-02<br />Petal.Width: 1.2553816","density: 4.267870e-02<br />Petal.Width: 1.2506849","density: 3.990377e-02<br />Petal.Width: 1.2459883","density: 3.726991e-02<br />Petal.Width: 1.2412916","density: 3.474487e-02<br />Petal.Width: 1.2365949","density: 3.232776e-02<br />Petal.Width: 1.2318982","density: 3.007244e-02<br />Petal.Width: 1.2272016","density: 2.792655e-02<br />Petal.Width: 1.2225049","density: 2.588127e-02<br />Petal.Width: 1.2178082","density: 2.394843e-02<br />Petal.Width: 1.2131115","density: 2.215452e-02<br />Petal.Width: 1.2084149","density: 2.045180e-02<br />Petal.Width: 1.2037182","density: 1.883858e-02<br />Petal.Width: 1.1990215","density: 1.734163e-02<br />Petal.Width: 1.1943249","density: 1.594728e-02<br />Petal.Width: 1.1896282","density: 1.463156e-02<br />Petal.Width: 1.1849315","density: 1.339264e-02<br />Petal.Width: 1.1802348","density: 1.226637e-02<br />Petal.Width: 1.1755382","density: 1.121088e-02<br />Petal.Width: 1.1708415","density: 1.022094e-02<br />Petal.Width: 1.1661448","density: 9.304163e-03<br />Petal.Width: 1.1614481","density: 8.471561e-03<br />Petal.Width: 1.1567515","density: 7.693573e-03<br />Petal.Width: 1.1520548","density: 6.968463e-03<br />Petal.Width: 1.1473581","density: 6.311322e-03<br />Petal.Width: 1.1426614","density: 5.710153e-03<br />Petal.Width: 1.1379648","density: 5.151875e-03<br />Petal.Width: 1.1332681","density: 4.634919e-03<br />Petal.Width: 1.1285714","density: 4.177447e-03<br />Petal.Width: 1.1238748","density: 3.754853e-03<br />Petal.Width: 1.1191781","density: 3.364918e-03<br />Petal.Width: 1.1144814","density: 3.011389e-03<br />Petal.Width: 1.1097847","density: 2.697015e-03<br />Petal.Width: 1.1050881","density: 2.407862e-03<br />Petal.Width: 1.1003914","density: 2.142830e-03<br />Petal.Width: 1.0956947","density: 1.908481e-03<br />Petal.Width: 1.0909980","density: 1.697773e-03<br />Petal.Width: 1.0863014","density: 1.505238e-03<br />Petal.Width: 1.0816047","density: 1.329984e-03<br />Petal.Width: 1.0769080","density: 1.179160e-03<br />Petal.Width: 1.0722114","density: 1.041720e-03<br />Petal.Width: 1.0675147","density: 9.169897e-04<br />Petal.Width: 1.0628180","density: 8.063963e-04<br />Petal.Width: 1.0581213","density: 7.100377e-04<br />Petal.Width: 1.0534247","density: 6.228112e-04<br />Petal.Width: 1.0487280","density: 5.442136e-04<br />Petal.Width: 1.0440313","density: 4.764718e-04<br />Petal.Width: 1.0393346","density: 4.165644e-04<br />Petal.Width: 1.0346380","density: 3.627140e-04<br />Petal.Width: 1.0299413","density: 3.146573e-04<br />Petal.Width: 1.0252446","density: 2.742739e-04<br />Petal.Width: 1.0205479","density: 2.380402e-04<br />Petal.Width: 1.0158513","density: 2.057078e-04<br />Petal.Width: 1.0111546","density: 1.776991e-04<br />Petal.Width: 1.0064579","density: 1.537705e-04<br />Petal.Width: 1.0017613","density: 1.324552e-04<br />Petal.Width: 0.9970646","density: 1.135796e-04<br />Petal.Width: 0.9923679","density: 9.773655e-05<br />Petal.Width: 0.9876712","density: 8.394450e-05<br />Petal.Width: 0.9829746","density: 7.175106e-05<br />Petal.Width: 0.9782779","density: 6.109398e-05<br />Petal.Width: 0.9735812","density: 5.234171e-05<br />Petal.Width: 0.9688845","density: 4.461055e-05<br />Petal.Width: 0.9641879","density: 3.782908e-05<br />Petal.Width: 0.9594912","density: 3.209482e-05<br />Petal.Width: 0.9547945","density: 2.728716e-05<br />Petal.Width: 0.9500978","density: 2.307337e-05<br />Petal.Width: 0.9454012","density: 1.940732e-05<br />Petal.Width: 0.9407045","density: 1.641318e-05<br />Petal.Width: 0.9360078","density: 1.384503e-05<br />Petal.Width: 0.9313112","density: 1.161235e-05<br />Petal.Width: 0.9266145","density: 9.703017e-06<br />Petal.Width: 0.9219178","density: 8.169223e-06<br />Petal.Width: 0.9172211","density: 6.835430e-06<br />Petal.Width: 0.9125245","density: 5.685633e-06<br />Petal.Width: 0.9078278","density: 4.737412e-06<br />Petal.Width: 0.9031311","density: 3.956510e-06<br />Petal.Width: 0.8984344","density: 3.283142e-06<br />Petal.Width: 0.8937378","density: 2.707735e-06<br />Petal.Width: 0.8890411","density: 2.250785e-06<br />Petal.Width: 0.8843444","density: 1.864253e-06<br />Petal.Width: 0.8796477","density: 1.533862e-06<br />Petal.Width: 0.8749511","density: 1.257648e-06<br />Petal.Width: 0.8702544","density: 1.040408e-06<br />Petal.Width: 0.8655577","density: 8.544375e-07<br />Petal.Width: 0.8608611","density: 6.969184e-07<br />Petal.Width: 0.8561644","density: 5.703224e-07<br />Petal.Width: 0.8514677","density: 4.678133e-07<br />Petal.Width: 0.8467710","density: 3.808596e-07<br />Petal.Width: 0.8420744","density: 3.078980e-07<br />Petal.Width: 0.8373777","density: 2.515962e-07<br />Petal.Width: 0.8326810","density: 2.045816e-07<br />Petal.Width: 0.8279843","density: 1.650781e-07<br />Petal.Width: 0.8232877","density: 1.328183e-07<br />Petal.Width: 0.8185910","density: 1.079531e-07<br />Petal.Width: 0.8138943","density: 8.699931e-08<br />Petal.Width: 0.8091977","density: 6.956468e-08<br />Petal.Width: 0.8045010","density: 5.591933e-08<br />Petal.Width: 0.7998043","density: 4.504447e-08<br />Petal.Width: 0.7951076","density: 3.597112e-08<br />Petal.Width: 0.7904110","density: 2.849719e-08<br />Petal.Width: 0.7857143","density: 2.289645e-08<br />Petal.Width: 0.7810176","density: 1.827493e-08<br />Petal.Width: 0.7763209","density: 1.445837e-08<br />Petal.Width: 0.7716243","density: 1.141632e-08<br />Petal.Width: 0.7669276","density: 9.116036e-09<br />Petal.Width: 0.7622309","density: 7.207986e-09<br />Petal.Width: 0.7575342","density: 5.648768e-09<br />Petal.Width: 0.7528376","density: 4.461168e-09<br />Petal.Width: 0.7481409","density: 3.528650e-09<br />Petal.Width: 0.7434442","density: 2.763478e-09<br />Petal.Width: 0.7387476","density: 2.144881e-09<br />Petal.Width: 0.7340509","density: 1.694967e-09<br />Petal.Width: 0.7293542","density: 1.327740e-09<br />Petal.Width: 0.7246575","density: 1.029736e-09<br />Petal.Width: 0.7199609","density: 7.980712e-10<br />Petal.Width: 0.7152642","density: 6.260332e-10<br />Petal.Width: 0.7105675","density: 4.855798e-10<br />Petal.Width: 0.7058708","density: 3.728817e-10<br />Petal.Width: 0.7011742","density: 2.893995e-10<br />Petal.Width: 0.6964775","density: 2.247472e-10<br />Petal.Width: 0.6917808","density: 1.725823e-10<br />Petal.Width: 0.6870841","density: 1.312030e-10<br />Petal.Width: 0.6823875","density: 1.020084e-10<br />Petal.Width: 0.6776908","density: 7.841421e-11<br />Petal.Width: 0.6729941","density: 5.960336e-11<br />Petal.Width: 0.6682975","density: 4.535119e-11<br />Petal.Width: 0.6636008","density: 3.494571e-11<br />Petal.Width: 0.6589041","density: 2.658543e-11<br />Petal.Width: 0.6542074","density: 2.000022e-11<br />Petal.Width: 0.6495108","density: 1.525892e-11<br />Petal.Width: 0.6448141","density: 1.163360e-11<br />Petal.Width: 0.6401174","density: 8.757813e-12<br />Petal.Width: 0.6354207","density: 6.520097e-12<br />Petal.Width: 0.6307241","density: 4.989263e-12<br />Petal.Width: 0.6260274","density: 3.763060e-12<br />Petal.Width: 0.6213307","density: 2.802824e-12<br />Petal.Width: 0.6166341","density: 2.094317e-12<br />Petal.Width: 0.6119374","density: 1.585160e-12<br />Petal.Width: 0.6072407","density: 1.182583e-12<br />Petal.Width: 0.6025440","density: 8.713328e-13<br />Petal.Width: 0.5978474","density: 6.536905e-13<br />Petal.Width: 0.5931507","density: 4.892270e-13<br />Petal.Width: 0.5884540","density: 3.610052e-13<br />Petal.Width: 0.5837573","density: 2.635943e-13<br />Petal.Width: 0.5790607","density: 1.983835e-13<br />Petal.Width: 0.5743640","density: 1.468679e-13<br />Petal.Width: 0.5696673","density: 1.072272e-13<br />Petal.Width: 0.5649706","density: 7.874183e-14<br />Petal.Width: 0.5602740","density: 5.858177e-14<br />Petal.Width: 0.5555773","density: 4.286134e-14<br />Petal.Width: 0.5508806","density: 3.090077e-14<br />Petal.Width: 0.5461840","density: 2.291759e-14<br />Petal.Width: 0.5414873","density: 1.686742e-14<br />Petal.Width: 0.5367906","density: 1.220075e-14<br />Petal.Width: 0.5320939","density: 8.753473e-15<br />Petal.Width: 0.5273973","density: 6.424042e-15<br />Petal.Width: 0.5227006","density: 4.654530e-15<br />Petal.Width: 0.5180039","density: 3.358265e-15<br />Petal.Width: 0.5133072","density: 2.471245e-15<br />Petal.Width: 0.5086106","density: 1.821985e-15<br />Petal.Width: 0.5039139","density: 1.282906e-15<br />Petal.Width: 0.4992172","density: 8.425158e-16<br />Petal.Width: 0.4945205","density: 6.409205e-16<br />Petal.Width: 0.4898239","density: 4.823877e-16<br />Petal.Width: 0.4851272","density: 3.615455e-16<br />Petal.Width: 0.4804305","density: 2.809073e-16<br />Petal.Width: 0.4757339","density: 1.906084e-16<br />Petal.Width: 0.4710372","density: 1.585185e-16<br />Petal.Width: 0.4663405","density: 1.582743e-16<br />Petal.Width: 0.4616438","density: 1.179553e-16<br />Petal.Width: 0.4569472","density: 7.763622e-17<br />Petal.Width: 0.4522505","density: 7.370515e-17<br />Petal.Width: 0.4475538","density: 1.087709e-16<br />Petal.Width: 0.4428571","density: 7.853157e-17<br />Petal.Width: 0.4381605","density: 5.532449e-17<br />Petal.Width: 0.4334638","density: 3.193076e-17<br />Petal.Width: 0.4287671","density: 1.691458e-18<br />Petal.Width: 0.4240705","density: 9.515947e-18<br />Petal.Width: 0.4193738","density: 1.387779e-17<br />Petal.Width: 0.4146771","density: 1.195788e-17<br />Petal.Width: 0.4099804","density: 1.878117e-18<br />Petal.Width: 0.4052838","density: 0.000000e+00<br />Petal.Width: 0.4005871","density: 0.000000e+00<br />Petal.Width: 0.3958904","density: 0.000000e+00<br />Petal.Width: 0.3911937","density: 0.000000e+00<br />Petal.Width: 0.3864971","density: 0.000000e+00<br />Petal.Width: 0.3818004","density: 0.000000e+00<br />Petal.Width: 0.3771037","density: 0.000000e+00<br />Petal.Width: 0.3724070","density: 5.622645e-17<br />Petal.Width: 0.3677104","density: 6.654757e-17<br />Petal.Width: 0.3630137","density: 4.340840e-17<br />Petal.Width: 0.3583170","density: 5.348816e-17<br />Petal.Width: 0.3536204","density: 2.328404e-17<br />Petal.Width: 0.3489237","density: 0.000000e+00<br />Petal.Width: 0.3442270","density: 5.068072e-18<br />Petal.Width: 0.3395303","density: 1.159455e-16<br />Petal.Width: 0.3348337","density: 1.189433e-16<br />Petal.Width: 0.3301370","density: 6.559998e-17<br />Petal.Width: 0.3254403","density: 5.121385e-18<br />Petal.Width: 0.3207436","density: 0.000000e+00<br />Petal.Width: 0.3160470","density: 5.428181e-18<br />Petal.Width: 0.3113503","density: 1.550795e-17<br />Petal.Width: 0.3066536","density: 2.558771e-17<br />Petal.Width: 0.3019569","density: 1.193177e-17<br />Petal.Width: 0.2972603","density: 0.000000e+00<br />Petal.Width: 0.2925636","density: 0.000000e+00<br />Petal.Width: 0.2878669","density: 0.000000e+00<br />Petal.Width: 0.2831703","density: 4.618324e-17<br />Petal.Width: 0.2784736","density: 7.754742e-17<br />Petal.Width: 0.2737769","density: 6.989055e-18<br />Petal.Width: 0.2690802","density: 5.448798e-17<br />Petal.Width: 0.2643836","density: 1.677997e-16<br />Petal.Width: 0.2596869","density: 2.845173e-16<br />Petal.Width: 0.2549902","density: 1.434006e-16<br />Petal.Width: 0.2502935","density: 1.032553e-16<br />Petal.Width: 0.2455969","density: 8.523748e-17<br />Petal.Width: 0.2409002","density: 5.722104e-17<br />Petal.Width: 0.2362035","density: 1.580187e-16<br />Petal.Width: 0.2315068","density: 1.426672e-16<br />Petal.Width: 0.2268102","density: 8.592144e-17<br />Petal.Width: 0.2221135","density: 9.600121e-17<br />Petal.Width: 0.2174168","density: 9.714451e-17<br />Petal.Width: 0.2127202","density: 9.200608e-17<br />Petal.Width: 0.2080235","density: 8.192631e-17<br />Petal.Width: 0.2033268","density: 7.184655e-17<br />Petal.Width: 0.1986301","density: 6.176678e-17<br />Petal.Width: 0.1939335","density: 5.168701e-17<br />Petal.Width: 0.1892368","density: 4.165948e-17<br />Petal.Width: 0.1845401","density: 5.173925e-17<br />Petal.Width: 0.1798434","density: 3.658757e-17<br />Petal.Width: 0.1751468","density: 3.144667e-17<br />Petal.Width: 0.1704501","density: 1.020050e-16<br />Petal.Width: 0.1657534","density: 1.286055e-16<br />Petal.Width: 0.1610568","density: 1.687393e-16<br />Petal.Width: 0.1563601","density: 2.172624e-16<br />Petal.Width: 0.1516634","density: 1.769434e-16<br />Petal.Width: 0.1469667","density: 1.441016e-16<br />Petal.Width: 0.1422701","density: 1.285794e-16<br />Petal.Width: 0.1375734","density: 1.386591e-16<br />Petal.Width: 0.1328767","density: 1.885829e-16<br />Petal.Width: 0.1281800","density: 1.588629e-16<br />Petal.Width: 0.1234834","density: 1.018745e-16<br />Petal.Width: 0.1187867","density: 1.220340e-16<br />Petal.Width: 0.1140900","density: 1.421935e-16<br />Petal.Width: 0.1093933","density: 1.284122e-16<br />Petal.Width: 0.1046967","density: 8.116539e-17<br />Petal.Width: 0.1000000","density: 8.116539e-17<br />Petal.Width: 0.1000000"],"key":["101","102","103","104","105","106","107","108","109","110","111","112","113","114","115","116","117","118","119","120","121","122","123","124","125","126","127","128","129","130","131","132","133","134","135","136","137","138","139","140","141","142","143","144","145","146","147","148","149","150"],"type":"scatter","mode":"lines","line":{"width":1.88976377952756,"color":"rgba(0,0,0,1)","dash":"solid"},"fill":"toself","fillcolor":"rgba(97,156,255,1)","hoveron":"points","set":"SharedData91833e3a","name":"virginica","legendgroup":"virginica","showlegend":true,"xaxis":"x4","yaxis":"y13","hoverinfo":"text","_isSimpleKey":true,"_isNestedKey":false,"frame":null}],"layout":{"xaxis":{"domain":[0,0.24],"automargin":true,"type":"linear","autorange":false,"range":[4.12,8.08],"tickmode":"array","ticktext":["5","6","7","8"],"tickvals":[5,6,7,8],"categoryorder":"array","categoryarray":["5","6","7","8"],"nticks":null,"ticks":"outside","tickcolor":"rgba(51,51,51,1)","ticklen":3.65296803652968,"tickwidth":0.66417600664176,"showticklabels":true,"tickfont":{"color":"rgba(77,77,77,1)","family":"","size":18.5969281859693},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"y","title":"Sepal.Length","hoverformat":".2f"},"xaxis2":{"domain":[0.26,0.49],"automargin":true,"type":"linear","autorange":false,"range":[1.88,4.52],"tickmode":"array","ticktext":["2.0","2.5","3.0","3.5","4.0","4.5"],"tickvals":[2,2.5,3,3.5,4,4.5],"categoryorder":"array","categoryarray":["2.0","2.5","3.0","3.5","4.0","4.5"],"nticks":null,"ticks":"outside","tickcolor":"rgba(51,51,51,1)","ticklen":3.65296803652968,"tickwidth":0.66417600664176,"showticklabels":true,"tickfont":{"color":"rgba(77,77,77,1)","family":"","size":18.5969281859693},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"y5","title":"Sepal.Width","hoverformat":".2f"},"xaxis3":{"domain":[0.51,0.74],"automargin":true,"type":"linear","autorange":false,"range":[0.705,7.195],"tickmode":"array","ticktext":["2","4","6"],"tickvals":[2,4,6],"categoryorder":"array","categoryarray":["2","4","6"],"nticks":null,"ticks":"outside","tickcolor":"rgba(51,51,51,1)","ticklen":3.65296803652968,"tickwidth":0.66417600664176,"showticklabels":true,"tickfont":{"color":"rgba(77,77,77,1)","family":"","size":18.5969281859693},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"y9","title":"Petal.Length","hoverformat":".2f"},"xaxis4":{"domain":[0.76,1],"automargin":true,"type":"linear","autorange":false,"range":[-0.02,2.62],"tickmode":"array","ticktext":["0.0","0.5","1.0","1.5","2.0","2.5"],"tickvals":[0,0.5,1,1.5,2,2.5],"categoryorder":"array","categoryarray":["0.0","0.5","1.0","1.5","2.0","2.5"],"nticks":null,"ticks":"outside","tickcolor":"rgba(51,51,51,1)","ticklen":3.65296803652968,"tickwidth":0.66417600664176,"showticklabels":true,"tickfont":{"color":"rgba(77,77,77,1)","family":"","size":18.5969281859693},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"y13","title":"Petal.Width","hoverformat":".2f"},"yaxis13":{"domain":[0,0.24],"automargin":true,"type":"linear","autorange":false,"range":[-0.375986649735357,7.8957196444425],"tickmode":"array","ticktext":["0","2","4","6"],"tickvals":[0,2,4,6],"categoryorder":"array","categoryarray":["0","2","4","6"],"nticks":null,"ticks":"","tickcolor":null,"ticklen":3.65296803652968,"tickwidth":0,"showticklabels":false,"tickfont":{"color":null,"family":null,"size":0},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x4","title":{"text":"","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis14":{"domain":[0.26,0.49],"automargin":true,"type":"linear","autorange":false,"range":[0.6401,7.2599],"tickmode":"array","ticktext":["2","4","6"],"tickvals":[2,4,6],"categoryorder":"array","categoryarray":["2","4","6"],"nticks":null,"ticks":"","tickcolor":null,"ticklen":3.65296803652968,"tickwidth":0,"showticklabels":false,"tickfont":{"color":null,"family":null,"size":0},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x4","title":{"text":"","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis15":{"domain":[0.51,0.74],"automargin":true,"type":"linear","autorange":false,"range":[1.8536,4.5464],"tickmode":"array","ticktext":["2.0","2.5","3.0","3.5","4.0","4.5"],"tickvals":[2,2.5,3,3.5,4,4.5],"categoryorder":"array","categoryarray":["2.0","2.5","3.0","3.5","4.0","4.5"],"nticks":null,"ticks":"","tickcolor":null,"ticklen":3.65296803652968,"tickwidth":0,"showticklabels":false,"tickfont":{"color":null,"family":null,"size":0},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x4","title":{"text":"","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis16":{"domain":[0.76,1],"automargin":true,"type":"linear","autorange":false,"range":[4.0804,8.1196],"tickmode":"array","ticktext":["5","6","7","8"],"tickvals":[5,6,7,8],"categoryorder":"array","categoryarray":["5","6","7","8"],"nticks":null,"ticks":"","tickcolor":null,"ticklen":3.65296803652968,"tickwidth":0,"showticklabels":false,"tickfont":{"color":null,"family":null,"size":0},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x4","title":{"text":"","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis9":{"domain":[0,0.24],"automargin":true,"type":"linear","autorange":false,"range":[-0.02,2.62],"tickmode":"array","ticktext":["0.0","0.5","1.0","1.5","2.0","2.5"],"tickvals":[0,0.5,1,1.5,2,2.5],"categoryorder":"array","categoryarray":["0.0","0.5","1.0","1.5","2.0","2.5"],"nticks":null,"ticks":"","tickcolor":null,"ticklen":3.65296803652968,"tickwidth":0,"showticklabels":false,"tickfont":{"color":null,"family":null,"size":0},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x3","title":{"text":"","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis10":{"domain":[0.26,0.49],"automargin":true,"type":"linear","autorange":false,"range":[-0.127394042301015,2.67527488832132],"tickmode":"array","ticktext":["0","1","2"],"tickvals":[0,1,2],"categoryorder":"array","categoryarray":["0","1","2"],"nticks":null,"ticks":"","tickcolor":null,"ticklen":3.65296803652968,"tickwidth":0,"showticklabels":false,"tickfont":{"color":null,"family":null,"size":0},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x3","title":{"text":"","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis11":{"domain":[0.51,0.74],"automargin":true,"type":"linear","autorange":false,"range":[1.8536,4.5464],"tickmode":"array","ticktext":["2.0","2.5","3.0","3.5","4.0","4.5"],"tickvals":[2,2.5,3,3.5,4,4.5],"categoryorder":"array","categoryarray":["2.0","2.5","3.0","3.5","4.0","4.5"],"nticks":null,"ticks":"","tickcolor":null,"ticklen":3.65296803652968,"tickwidth":0,"showticklabels":false,"tickfont":{"color":null,"family":null,"size":0},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x3","title":{"text":"","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis12":{"domain":[0.76,1],"automargin":true,"type":"linear","autorange":false,"range":[4.0804,8.1196],"tickmode":"array","ticktext":["5","6","7","8"],"tickvals":[5,6,7,8],"categoryorder":"array","categoryarray":["5","6","7","8"],"nticks":null,"ticks":"","tickcolor":null,"ticklen":3.65296803652968,"tickwidth":0,"showticklabels":false,"tickfont":{"color":null,"family":null,"size":0},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x3","title":{"text":"","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis5":{"domain":[0,0.24],"automargin":true,"type":"linear","autorange":false,"range":[-0.02,2.62],"tickmode":"array","ticktext":["0.0","0.5","1.0","1.5","2.0","2.5"],"tickvals":[0,0.5,1,1.5,2,2.5],"categoryorder":"array","categoryarray":["0.0","0.5","1.0","1.5","2.0","2.5"],"nticks":null,"ticks":"","tickcolor":null,"ticklen":3.65296803652968,"tickwidth":0,"showticklabels":false,"tickfont":{"color":null,"family":null,"size":0},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x2","title":{"text":"","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis6":{"domain":[0.26,0.49],"automargin":true,"type":"linear","autorange":false,"range":[0.705,7.195],"tickmode":"array","ticktext":["2","4","6"],"tickvals":[2,4,6],"categoryorder":"array","categoryarray":["2","4","6"],"nticks":null,"ticks":"","tickcolor":null,"ticklen":3.65296803652968,"tickwidth":0,"showticklabels":false,"tickfont":{"color":null,"family":null,"size":0},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x2","title":{"text":"","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis7":{"domain":[0.51,0.74],"automargin":true,"type":"linear","autorange":false,"range":[-0.0666726038830392,1.40012468154382],"tickmode":"array","ticktext":["0.0","0.5","1.0"],"tickvals":[0,0.5,1],"categoryorder":"array","categoryarray":["0.0","0.5","1.0"],"nticks":null,"ticks":"","tickcolor":null,"ticklen":3.65296803652968,"tickwidth":0,"showticklabels":false,"tickfont":{"color":null,"family":null,"size":0},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x2","title":{"text":"","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis8":{"domain":[0.76,1],"automargin":true,"type":"linear","autorange":false,"range":[4.0804,8.1196],"tickmode":"array","ticktext":["5","6","7","8"],"tickvals":[5,6,7,8],"categoryorder":"array","categoryarray":["5","6","7","8"],"nticks":null,"ticks":"","tickcolor":null,"ticklen":3.65296803652968,"tickwidth":0,"showticklabels":false,"tickfont":{"color":null,"family":null,"size":0},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x2","title":{"text":"","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis":{"domain":[0,0.24],"automargin":true,"type":"linear","autorange":false,"range":[-0.02,2.62],"tickmode":"array","ticktext":["0.0","0.5","1.0","1.5","2.0","2.5"],"tickvals":[0,0.5,1,1.5,2,2.5],"categoryorder":"array","categoryarray":["0.0","0.5","1.0","1.5","2.0","2.5"],"nticks":null,"ticks":"outside","tickcolor":"rgba(51,51,51,1)","ticklen":3.65296803652968,"tickwidth":0.66417600664176,"showticklabels":true,"tickfont":{"color":"rgba(77,77,77,1)","family":"","size":18.5969281859693},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x","title":{"text":"Petal.Width","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis2":{"domain":[0.26,0.49],"automargin":true,"type":"linear","autorange":false,"range":[0.705,7.195],"tickmode":"array","ticktext":["2","4","6"],"tickvals":[2,4,6],"categoryorder":"array","categoryarray":["2","4","6"],"nticks":null,"ticks":"outside","tickcolor":"rgba(51,51,51,1)","ticklen":3.65296803652968,"tickwidth":0.66417600664176,"showticklabels":true,"tickfont":{"color":"rgba(77,77,77,1)","family":"","size":18.5969281859693},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x","title":{"text":"Petal.Length","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis3":{"domain":[0.51,0.74],"automargin":true,"type":"linear","autorange":false,"range":[1.88,4.52],"tickmode":"array","ticktext":["2.0","2.5","3.0","3.5","4.0","4.5"],"tickvals":[2,2.5,3,3.5,4,4.5],"categoryorder":"array","categoryarray":["2.0","2.5","3.0","3.5","4.0","4.5"],"nticks":null,"ticks":"outside","tickcolor":"rgba(51,51,51,1)","ticklen":3.65296803652968,"tickwidth":0.66417600664176,"showticklabels":true,"tickfont":{"color":"rgba(77,77,77,1)","family":"","size":18.5969281859693},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x","title":{"text":"Sepal.Width","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"yaxis4":{"domain":[0.76,1],"automargin":true,"type":"linear","autorange":false,"range":[-0.0619923272297817,1.30183887182541],"tickmode":"array","ticktext":["0.0","0.4","0.8","1.2"],"tickvals":[0,0.4,0.8,1.2],"categoryorder":"array","categoryarray":["0.0","0.4","0.8","1.2"],"nticks":null,"ticks":"outside","tickcolor":"rgba(51,51,51,1)","ticklen":3.65296803652968,"tickwidth":0.66417600664176,"showticklabels":true,"tickfont":{"color":"rgba(77,77,77,1)","family":"","size":18.5969281859693},"tickangle":-0,"showline":true,"linecolor":"rgba(0,0,0,1)","linewidth":0.66417600664176,"showgrid":false,"gridcolor":null,"gridwidth":0,"zeroline":false,"anchor":"x","title":{"text":"Sepal.Length","font":{"color":"rgba(0,0,0,1)","family":"","size":21.2536322125363}},"hoverformat":".2f"},"annotations":[],"shapes":[{"type":"rect","fillcolor":null,"line":{"color":null,"width":0,"linetype":[]},"yref":"paper","xref":"paper","x0":0,"x1":0.24,"y0":0.76,"y1":1},{"type":"rect","fillcolor":null,"line":{"color":null,"width":0,"linetype":[]},"yref":"paper","xref":"paper","x0":0,"x1":0.24,"y0":0.51,"y1":0.74},{"type":"rect","fillcolor":null,"line":{"color":null,"width":0,"linetype":[]},"yref":"paper","xref":"paper","x0":0,"x1":0.24,"y0":0.26,"y1":0.49},{"type":"rect","fillcolor":null,"line":{"color":null,"width":0,"linetype":[]},"yref":"paper","xref":"paper","x0":0,"x1":0.24,"y0":0,"y1":0.24},{"type":"rect","fillcolor":"transparent","line":{"color":"rgba(255,255,255,1)","width":0.66417600664176,"linetype":"solid"},"yref":"paper","xref":"paper","x0":0.26,"x1":0.49,"y0":0.76,"y1":1},{"type":"rect","fillcolor":null,"line":{"color":null,"width":0,"linetype":[]},"yref":"paper","xref":"paper","x0":0.26,"x1":0.49,"y0":0.51,"y1":0.74},{"type":"rect","fillcolor":null,"line":{"color":null,"width":0,"linetype":[]},"yref":"paper","xref":"paper","x0":0.26,"x1":0.49,"y0":0.26,"y1":0.49},{"type":"rect","fillcolor":null,"line":{"color":null,"width":0,"linetype":[]},"yref":"paper","xref":"paper","x0":0.26,"x1":0.49,"y0":0,"y1":0.24},{"type":"rect","fillcolor":"transparent","line":{"color":"rgba(255,255,255,1)","width":0.66417600664176,"linetype":"solid"},"yref":"paper","xref":"paper","x0":0.51,"x1":0.74,"y0":0.76,"y1":1},{"type":"rect","fillcolor":"transparent","line":{"color":"rgba(255,255,255,1)","width":0.66417600664176,"linetype":"solid"},"yref":"paper","xref":"paper","x0":0.51,"x1":0.74,"y0":0.51,"y1":0.74},{"type":"rect","fillcolor":null,"line":{"color":null,"width":0,"linetype":[]},"yref":"paper","xref":"paper","x0":0.51,"x1":0.74,"y0":0.26,"y1":0.49},{"type":"rect","fillcolor":null,"line":{"color":null,"width":0,"linetype":[]},"yref":"paper","xref":"paper","x0":0.51,"x1":0.74,"y0":0,"y1":0.24},{"type":"rect","fillcolor":"transparent","line":{"color":"rgba(255,255,255,1)","width":0.66417600664176,"linetype":"solid"},"yref":"paper","xref":"paper","x0":0.76,"x1":1,"y0":0.76,"y1":1},{"type":"rect","fillcolor":"transparent","line":{"color":"rgba(255,255,255,1)","width":0.66417600664176,"linetype":"solid"},"yref":"paper","xref":"paper","x0":0.76,"x1":1,"y0":0.51,"y1":0.74},{"type":"rect","fillcolor":"transparent","line":{"color":"rgba(255,255,255,1)","width":0.66417600664176,"linetype":"solid"},"yref":"paper","xref":"paper","x0":0.76,"x1":1,"y0":0.26,"y1":0.49},{"type":"rect","fillcolor":null,"line":{"color":null,"width":0,"linetype":[]},"yref":"paper","xref":"paper","x0":0.76,"x1":1,"y0":0,"y1":0.24}],"images":[],"margin":{"t":26.2283105022831,"r":7.30593607305936,"b":53.7318389373184,"l":60.958904109589},"plot_bgcolor":"rgba(255,255,255,1)","paper_bgcolor":"rgba(255,255,255,1)","font":{"color":"rgba(0,0,0,1)","family":"","size":14.6118721461187},"showlegend":false,"legend":{"bgcolor":"rgba(255,255,255,1)","bordercolor":"transparent","borderwidth":1.88976377952756,"font":{"color":"rgba(0,0,0,1)","family":"","size":18.5969281859693},"y":0.874015748031496},"hovermode":"closest","barmode":"relative","dragmode":"select"},"attrs":{"1841049cbab10":{"x":{},"fill":{},"type":"scatter"},"1841064c565b1":{"x":{},"y":{},"colour":{},"type":"scatter"},"184106951c599":{"x":{},"y":{},"colour":{},"type":"scatter"},"184102706c005":{"x":{},"y":{},"colour":{},"type":"scatter"},"184102f8aff15":{"x":{},"y":{},"type":"scatter"},"184104e70d415":{"x":{},"y":{},"label":{},"colour":{}},"1841026a1019":{"x":{},"fill":{},"type":"scatter"},"18410326a7429":{"x":{},"y":{},"colour":{},"type":"scatter"},"1841051af3226":{"x":{},"y":{},"colour":{},"type":"scatter"},"18410536cee75":{"x":{},"y":{},"type":"scatter"},"18410149b6e1d":{"x":{},"y":{},"label":{},"colour":{}},"18410209e0066":{"x":{},"y":{},"type":"scatter"},"18410652c3944":{"x":{},"y":{},"label":{},"colour":{}},"1841035049370":{"x":{},"fill":{},"type":"scatter"},"18410110fbca7":{"x":{},"y":{},"colour":{},"type":"scatter"},"1841049530ac1":{"x":{},"y":{},"type":"scatter"},"184106ae72a82":{"x":{},"y":{},"label":{},"colour":{}},"1841012dd05a2":{"x":{},"y":{},"type":"scatter"},"184106c9cd85a":{"x":{},"y":{},"label":{},"colour":{}},"18410395268e9":{"x":{},"y":{},"type":"scatter"},"184105165b665":{"x":{},"y":{},"label":{},"colour":{}},"1841055778dbe":{"x":{},"fill":{},"type":"scatter"}},"source":"A","config":{"doubleClick":"reset","showSendToCloud":false},"highlight":{"on":"plotly_selected","off":"plotly_deselect","persistent":false,"dynamic":false,"color":null,"selectize":false,"defaultValues":null,"opacityDim":0.2,"selected":{"opacity":1},"debounce":0,"ctGroups":["SharedData91833e3a"]},"subplot":true,".hideLegend":true,"shinyEvents":["plotly_hover","plotly_click","plotly_selected","plotly_relayout","plotly_brushed","plotly_brushing","plotly_clickannotation","plotly_doubleclick","plotly_deselect","plotly_afterplot","plotly_sunburstclick"],"base_url":"https://plot.ly"},"evals":[],"jsHooks":[]}</script> ] .right-column70[ **GGally** is a **ggplot2** extension that provides additional composite plot types like the pairs plot we use here. ] --- class: middle, inverse # Dygraphs https://rstudio.github.io/dygraphs --- ## Dygraphs The *dygraphs* package wraps the `dygraphs` Javascript pacman::p_load, which is build to chart time-series data. ```r pacman::p_load(dygraphs) dygraph(nhtemp, main = "New Haven temperatures") %>% dyRangeSelector(dateWindow = c("1920-01-01","1960-01-01")) ``` <div id="htmlwidget-5f88b4faa0d69ea0b10d" style="width:504px;height:432px;" class="dygraphs html-widget"></div> <script type="application/json" data-for="htmlwidget-5f88b4faa0d69ea0b10d">{"x":{"attrs":{"title":"New Haven temperatures","labels":["year","V1"],"legend":"auto","retainDateWindow":false,"axes":{"x":{"pixelsPerLabel":60}},"showRangeSelector":true,"dateWindow":["1920-01-01T00:00:00.000Z","1960-01-01T00:00:00.000Z"],"rangeSelectorHeight":40,"rangeSelectorPlotFillColor":" #A7B1C4","rangeSelectorPlotStrokeColor":"#808FAB","interactionModel":"Dygraph.Interaction.defaultModel"},"scale":"yearly","annotations":[],"shadings":[],"events":[],"format":"date","data":[["1912-01-01T00:00:00.000Z","1913-01-01T00:00:00.000Z","1914-01-01T00:00:00.000Z","1915-01-01T00:00:00.000Z","1916-01-01T00:00:00.000Z","1917-01-01T00:00:00.000Z","1918-01-01T00:00:00.000Z","1919-01-01T00:00:00.000Z","1920-01-01T00:00:00.000Z","1921-01-01T00:00:00.000Z","1922-01-01T00:00:00.000Z","1923-01-01T00:00:00.000Z","1924-01-01T00:00:00.000Z","1925-01-01T00:00:00.000Z","1926-01-01T00:00:00.000Z","1927-01-01T00:00:00.000Z","1928-01-01T00:00:00.000Z","1929-01-01T00:00:00.000Z","1930-01-01T00:00:00.000Z","1931-01-01T00:00:00.000Z","1932-01-01T00:00:00.000Z","1933-01-01T00:00:00.000Z","1934-01-01T00:00:00.000Z","1935-01-01T00:00:00.000Z","1936-01-01T00:00:00.000Z","1937-01-01T00:00:00.000Z","1938-01-01T00:00:00.000Z","1939-01-01T00:00:00.000Z","1940-01-01T00:00:00.000Z","1941-01-01T00:00:00.000Z","1942-01-01T00:00:00.000Z","1943-01-01T00:00:00.000Z","1944-01-01T00:00:00.000Z","1945-01-01T00:00:00.000Z","1946-01-01T00:00:00.000Z","1947-01-01T00:00:00.000Z","1948-01-01T00:00:00.000Z","1949-01-01T00:00:00.000Z","1950-01-01T00:00:00.000Z","1951-01-01T00:00:00.000Z","1952-01-01T00:00:00.000Z","1953-01-01T00:00:00.000Z","1954-01-01T00:00:00.000Z","1955-01-01T00:00:00.000Z","1956-01-01T00:00:00.000Z","1957-01-01T00:00:00.000Z","1958-01-01T00:00:00.000Z","1959-01-01T00:00:00.000Z","1960-01-01T00:00:00.000Z","1961-01-01T00:00:00.000Z","1962-01-01T00:00:00.000Z","1963-01-01T00:00:00.000Z","1964-01-01T00:00:00.000Z","1965-01-01T00:00:00.000Z","1966-01-01T00:00:00.000Z","1967-01-01T00:00:00.000Z","1968-01-01T00:00:00.000Z","1969-01-01T00:00:00.000Z","1970-01-01T00:00:00.000Z","1971-01-01T00:00:00.000Z"],[49.9,52.3,49.4,51.1,49.4,47.9,49.8,50.9,49.3,51.9,50.8,49.6,49.3,50.6,48.4,50.7,50.9,50.6,51.5,52.8,51.8,51.1,49.8,50.2,50.4,51.6,51.8,50.9,48.8,51.7,51,50.6,51.7,51.5,52.1,51.3,51,54,51.4,52.7,53.1,54.6,52,52,50.9,52.6,50.2,52.6,51.6,51.9,50.5,50.9,51.7,51.4,51.7,50.8,51.9,51.8,51.9,53]]},"evals":["attrs.interactionModel"],"jsHooks":[]}</script> ------------------------------------------------------------------------------- This package is built for linked charts, in the form of line charts + annotations + controls --- ## Dygraphs We can also create multiple linked time series .left-column30[ ```r dygraph(ldeaths, main = "All", * group = "lung-deaths") dygraph(mdeaths, main = "Male", * group = "lung-deaths") dygraph(fdeaths, main = "Female", * group = "lung-deaths") ``` You could name the group anything you like, as long as it's the same across the plots. ] .right-column30[ <div id="htmlwidget-02821e85d8fbaed04314" style="width:504px;height:144px;" class="dygraphs html-widget"></div> <script type="application/json" data-for="htmlwidget-02821e85d8fbaed04314">{"x":{"attrs":{"title":"All","labels":["month","V1"],"legend":"auto","retainDateWindow":false,"axes":{"x":{"pixelsPerLabel":60}}},"scale":"monthly","group":"lung-deaths","annotations":[],"shadings":[],"events":[],"format":"date","data":[["1974-01-01T00:00:00.000Z","1974-02-01T00:00:00.000Z","1974-03-01T00:00:00.000Z","1974-04-01T00:00:00.000Z","1974-05-01T00:00:00.000Z","1974-06-01T00:00:00.000Z","1974-07-01T00:00:00.000Z","1974-08-01T00:00:00.000Z","1974-09-01T00:00:00.000Z","1974-10-01T00:00:00.000Z","1974-11-01T00:00:00.000Z","1974-12-01T00:00:00.000Z","1975-01-01T00:00:00.000Z","1975-02-01T00:00:00.000Z","1975-03-01T00:00:00.000Z","1975-04-01T00:00:00.000Z","1975-05-01T00:00:00.000Z","1975-06-01T00:00:00.000Z","1975-07-01T00:00:00.000Z","1975-08-01T00:00:00.000Z","1975-09-01T00:00:00.000Z","1975-10-01T00:00:00.000Z","1975-11-01T00:00:00.000Z","1975-12-01T00:00:00.000Z","1976-01-01T00:00:00.000Z","1976-02-01T00:00:00.000Z","1976-03-01T00:00:00.000Z","1976-04-01T00:00:00.000Z","1976-05-01T00:00:00.000Z","1976-06-01T00:00:00.000Z","1976-07-01T00:00:00.000Z","1976-08-01T00:00:00.000Z","1976-09-01T00:00:00.000Z","1976-10-01T00:00:00.000Z","1976-11-01T00:00:00.000Z","1976-12-01T00:00:00.000Z","1977-01-01T00:00:00.000Z","1977-02-01T00:00:00.000Z","1977-03-01T00:00:00.000Z","1977-04-01T00:00:00.000Z","1977-05-01T00:00:00.000Z","1977-06-01T00:00:00.000Z","1977-07-01T00:00:00.000Z","1977-08-01T00:00:00.000Z","1977-09-01T00:00:00.000Z","1977-10-01T00:00:00.000Z","1977-11-01T00:00:00.000Z","1977-12-01T00:00:00.000Z","1978-01-01T00:00:00.000Z","1978-02-01T00:00:00.000Z","1978-03-01T00:00:00.000Z","1978-04-01T00:00:00.000Z","1978-05-01T00:00:00.000Z","1978-06-01T00:00:00.000Z","1978-07-01T00:00:00.000Z","1978-08-01T00:00:00.000Z","1978-09-01T00:00:00.000Z","1978-10-01T00:00:00.000Z","1978-11-01T00:00:00.000Z","1978-12-01T00:00:00.000Z","1979-01-01T00:00:00.000Z","1979-02-01T00:00:00.000Z","1979-03-01T00:00:00.000Z","1979-04-01T00:00:00.000Z","1979-05-01T00:00:00.000Z","1979-06-01T00:00:00.000Z","1979-07-01T00:00:00.000Z","1979-08-01T00:00:00.000Z","1979-09-01T00:00:00.000Z","1979-10-01T00:00:00.000Z","1979-11-01T00:00:00.000Z","1979-12-01T00:00:00.000Z"],[3035,2552,2704,2554,2014,1655,1721,1524,1596,2074,2199,2512,2933,2889,2938,2497,1870,1726,1607,1545,1396,1787,2076,2837,2787,3891,3179,2011,1636,1580,1489,1300,1356,1653,2013,2823,3102,2294,2385,2444,1748,1554,1498,1361,1346,1564,1640,2293,2815,3137,2679,1969,1870,1633,1529,1366,1357,1570,1535,2491,3084,2605,2573,2143,1693,1504,1461,1354,1333,1492,1781,1915]]},"evals":[],"jsHooks":[]}</script><div id="htmlwidget-c0e9f7fe5fa8f99f21ea" style="width:504px;height:144px;" class="dygraphs html-widget"></div> <script type="application/json" data-for="htmlwidget-c0e9f7fe5fa8f99f21ea">{"x":{"attrs":{"title":"Male","labels":["month","V1"],"legend":"auto","retainDateWindow":false,"axes":{"x":{"pixelsPerLabel":60}}},"scale":"monthly","group":"lung-deaths","annotations":[],"shadings":[],"events":[],"format":"date","data":[["1974-01-01T00:00:00.000Z","1974-02-01T00:00:00.000Z","1974-03-01T00:00:00.000Z","1974-04-01T00:00:00.000Z","1974-05-01T00:00:00.000Z","1974-06-01T00:00:00.000Z","1974-07-01T00:00:00.000Z","1974-08-01T00:00:00.000Z","1974-09-01T00:00:00.000Z","1974-10-01T00:00:00.000Z","1974-11-01T00:00:00.000Z","1974-12-01T00:00:00.000Z","1975-01-01T00:00:00.000Z","1975-02-01T00:00:00.000Z","1975-03-01T00:00:00.000Z","1975-04-01T00:00:00.000Z","1975-05-01T00:00:00.000Z","1975-06-01T00:00:00.000Z","1975-07-01T00:00:00.000Z","1975-08-01T00:00:00.000Z","1975-09-01T00:00:00.000Z","1975-10-01T00:00:00.000Z","1975-11-01T00:00:00.000Z","1975-12-01T00:00:00.000Z","1976-01-01T00:00:00.000Z","1976-02-01T00:00:00.000Z","1976-03-01T00:00:00.000Z","1976-04-01T00:00:00.000Z","1976-05-01T00:00:00.000Z","1976-06-01T00:00:00.000Z","1976-07-01T00:00:00.000Z","1976-08-01T00:00:00.000Z","1976-09-01T00:00:00.000Z","1976-10-01T00:00:00.000Z","1976-11-01T00:00:00.000Z","1976-12-01T00:00:00.000Z","1977-01-01T00:00:00.000Z","1977-02-01T00:00:00.000Z","1977-03-01T00:00:00.000Z","1977-04-01T00:00:00.000Z","1977-05-01T00:00:00.000Z","1977-06-01T00:00:00.000Z","1977-07-01T00:00:00.000Z","1977-08-01T00:00:00.000Z","1977-09-01T00:00:00.000Z","1977-10-01T00:00:00.000Z","1977-11-01T00:00:00.000Z","1977-12-01T00:00:00.000Z","1978-01-01T00:00:00.000Z","1978-02-01T00:00:00.000Z","1978-03-01T00:00:00.000Z","1978-04-01T00:00:00.000Z","1978-05-01T00:00:00.000Z","1978-06-01T00:00:00.000Z","1978-07-01T00:00:00.000Z","1978-08-01T00:00:00.000Z","1978-09-01T00:00:00.000Z","1978-10-01T00:00:00.000Z","1978-11-01T00:00:00.000Z","1978-12-01T00:00:00.000Z","1979-01-01T00:00:00.000Z","1979-02-01T00:00:00.000Z","1979-03-01T00:00:00.000Z","1979-04-01T00:00:00.000Z","1979-05-01T00:00:00.000Z","1979-06-01T00:00:00.000Z","1979-07-01T00:00:00.000Z","1979-08-01T00:00:00.000Z","1979-09-01T00:00:00.000Z","1979-10-01T00:00:00.000Z","1979-11-01T00:00:00.000Z","1979-12-01T00:00:00.000Z"],[2134,1863,1877,1877,1492,1249,1280,1131,1209,1492,1621,1846,2103,2137,2153,1833,1403,1288,1186,1133,1053,1347,1545,2066,2020,2750,2283,1479,1189,1160,1113,970,999,1208,1467,2059,2240,1634,1722,1801,1246,1162,1087,1013,959,1179,1229,1655,2019,2284,1942,1423,1340,1187,1098,1004,970,1140,1110,1812,2263,1820,1846,1531,1215,1075,1056,975,940,1081,1294,1341]]},"evals":[],"jsHooks":[]}</script><div id="htmlwidget-1c9631a34451a862e997" style="width:504px;height:144px;" class="dygraphs html-widget"></div> <script type="application/json" data-for="htmlwidget-1c9631a34451a862e997">{"x":{"attrs":{"title":"Female","labels":["month","V1"],"legend":"auto","retainDateWindow":false,"axes":{"x":{"pixelsPerLabel":60}}},"scale":"monthly","group":"lung-deaths","annotations":[],"shadings":[],"events":[],"format":"date","data":[["1974-01-01T00:00:00.000Z","1974-02-01T00:00:00.000Z","1974-03-01T00:00:00.000Z","1974-04-01T00:00:00.000Z","1974-05-01T00:00:00.000Z","1974-06-01T00:00:00.000Z","1974-07-01T00:00:00.000Z","1974-08-01T00:00:00.000Z","1974-09-01T00:00:00.000Z","1974-10-01T00:00:00.000Z","1974-11-01T00:00:00.000Z","1974-12-01T00:00:00.000Z","1975-01-01T00:00:00.000Z","1975-02-01T00:00:00.000Z","1975-03-01T00:00:00.000Z","1975-04-01T00:00:00.000Z","1975-05-01T00:00:00.000Z","1975-06-01T00:00:00.000Z","1975-07-01T00:00:00.000Z","1975-08-01T00:00:00.000Z","1975-09-01T00:00:00.000Z","1975-10-01T00:00:00.000Z","1975-11-01T00:00:00.000Z","1975-12-01T00:00:00.000Z","1976-01-01T00:00:00.000Z","1976-02-01T00:00:00.000Z","1976-03-01T00:00:00.000Z","1976-04-01T00:00:00.000Z","1976-05-01T00:00:00.000Z","1976-06-01T00:00:00.000Z","1976-07-01T00:00:00.000Z","1976-08-01T00:00:00.000Z","1976-09-01T00:00:00.000Z","1976-10-01T00:00:00.000Z","1976-11-01T00:00:00.000Z","1976-12-01T00:00:00.000Z","1977-01-01T00:00:00.000Z","1977-02-01T00:00:00.000Z","1977-03-01T00:00:00.000Z","1977-04-01T00:00:00.000Z","1977-05-01T00:00:00.000Z","1977-06-01T00:00:00.000Z","1977-07-01T00:00:00.000Z","1977-08-01T00:00:00.000Z","1977-09-01T00:00:00.000Z","1977-10-01T00:00:00.000Z","1977-11-01T00:00:00.000Z","1977-12-01T00:00:00.000Z","1978-01-01T00:00:00.000Z","1978-02-01T00:00:00.000Z","1978-03-01T00:00:00.000Z","1978-04-01T00:00:00.000Z","1978-05-01T00:00:00.000Z","1978-06-01T00:00:00.000Z","1978-07-01T00:00:00.000Z","1978-08-01T00:00:00.000Z","1978-09-01T00:00:00.000Z","1978-10-01T00:00:00.000Z","1978-11-01T00:00:00.000Z","1978-12-01T00:00:00.000Z","1979-01-01T00:00:00.000Z","1979-02-01T00:00:00.000Z","1979-03-01T00:00:00.000Z","1979-04-01T00:00:00.000Z","1979-05-01T00:00:00.000Z","1979-06-01T00:00:00.000Z","1979-07-01T00:00:00.000Z","1979-08-01T00:00:00.000Z","1979-09-01T00:00:00.000Z","1979-10-01T00:00:00.000Z","1979-11-01T00:00:00.000Z","1979-12-01T00:00:00.000Z"],[901,689,827,677,522,406,441,393,387,582,578,666,830,752,785,664,467,438,421,412,343,440,531,771,767,1141,896,532,447,420,376,330,357,445,546,764,862,660,663,643,502,392,411,348,387,385,411,638,796,853,737,546,530,446,431,362,387,430,425,679,821,785,727,612,478,429,405,379,393,411,487,574]]},"evals":[],"jsHooks":[]}</script> ] --- ## Dygraphs You could also annotate dygraphs ```r dygraph(presidents, main = "Quarterly Presidential Approval Ratings") %>% dyAxis("y", valueRange = c(0, 100)) %>% dyEvent("1950-6-30", "Korea", labelLoc = "bottom") %>% dyEvent("1965-2-09", "Vietnam", labelLoc = "bottom") ``` <div id="htmlwidget-0f38e7772d45d116e52d" style="width:504px;height:360px;" class="dygraphs html-widget"></div> <script type="application/json" data-for="htmlwidget-0f38e7772d45d116e52d">{"x":{"attrs":{"axes":{"x":{"pixelsPerLabel":60},"y":{"valueRange":[0,100]}},"title":"Quarterly Presidential Approval Ratings","labels":["quarter","V1"],"legend":"auto","retainDateWindow":false},"scale":"quarterly","annotations":[],"shadings":[],"events":[{"pos":"1950-06-30T00:00:00.000Z","label":"Korea","labelLoc":"bottom","color":"black","strokePattern":[7,3],"axis":"x"},{"pos":"1965-02-09T00:00:00.000Z","label":"Vietnam","labelLoc":"bottom","color":"black","strokePattern":[7,3],"axis":"x"}],"format":"date","data":[["1945-01-01T00:00:00.000Z","1945-04-01T00:00:00.000Z","1945-07-01T00:00:00.000Z","1945-10-01T00:00:00.000Z","1946-01-01T00:00:00.000Z","1946-04-01T00:00:00.000Z","1946-07-01T00:00:00.000Z","1946-10-01T00:00:00.000Z","1947-01-01T00:00:00.000Z","1947-04-01T00:00:00.000Z","1947-07-01T00:00:00.000Z","1947-10-01T00:00:00.000Z","1948-01-01T00:00:00.000Z","1948-04-01T00:00:00.000Z","1948-07-01T00:00:00.000Z","1948-10-01T00:00:00.000Z","1949-01-01T00:00:00.000Z","1949-04-01T00:00:00.000Z","1949-07-01T00:00:00.000Z","1949-10-01T00:00:00.000Z","1950-01-01T00:00:00.000Z","1950-04-01T00:00:00.000Z","1950-07-01T00:00:00.000Z","1950-10-01T00:00:00.000Z","1951-01-01T00:00:00.000Z","1951-04-01T00:00:00.000Z","1951-07-01T00:00:00.000Z","1951-10-01T00:00:00.000Z","1952-01-01T00:00:00.000Z","1952-04-01T00:00:00.000Z","1952-07-01T00:00:00.000Z","1952-10-01T00:00:00.000Z","1953-01-01T00:00:00.000Z","1953-04-01T00:00:00.000Z","1953-07-01T00:00:00.000Z","1953-10-01T00:00:00.000Z","1954-01-01T00:00:00.000Z","1954-04-01T00:00:00.000Z","1954-07-01T00:00:00.000Z","1954-10-01T00:00:00.000Z","1955-01-01T00:00:00.000Z","1955-04-01T00:00:00.000Z","1955-07-01T00:00:00.000Z","1955-10-01T00:00:00.000Z","1956-01-01T00:00:00.000Z","1956-04-01T00:00:00.000Z","1956-07-01T00:00:00.000Z","1956-10-01T00:00:00.000Z","1957-01-01T00:00:00.000Z","1957-04-01T00:00:00.000Z","1957-07-01T00:00:00.000Z","1957-10-01T00:00:00.000Z","1958-01-01T00:00:00.000Z","1958-04-01T00:00:00.000Z","1958-07-01T00:00:00.000Z","1958-10-01T00:00:00.000Z","1959-01-01T00:00:00.000Z","1959-04-01T00:00:00.000Z","1959-07-01T00:00:00.000Z","1959-10-01T00:00:00.000Z","1960-01-01T00:00:00.000Z","1960-04-01T00:00:00.000Z","1960-07-01T00:00:00.000Z","1960-10-01T00:00:00.000Z","1961-01-01T00:00:00.000Z","1961-04-01T00:00:00.000Z","1961-07-01T00:00:00.000Z","1961-10-01T00:00:00.000Z","1962-01-01T00:00:00.000Z","1962-04-01T00:00:00.000Z","1962-07-01T00:00:00.000Z","1962-10-01T00:00:00.000Z","1963-01-01T00:00:00.000Z","1963-04-01T00:00:00.000Z","1963-07-01T00:00:00.000Z","1963-10-01T00:00:00.000Z","1964-01-01T00:00:00.000Z","1964-04-01T00:00:00.000Z","1964-07-01T00:00:00.000Z","1964-10-01T00:00:00.000Z","1965-01-01T00:00:00.000Z","1965-04-01T00:00:00.000Z","1965-07-01T00:00:00.000Z","1965-10-01T00:00:00.000Z","1966-01-01T00:00:00.000Z","1966-04-01T00:00:00.000Z","1966-07-01T00:00:00.000Z","1966-10-01T00:00:00.000Z","1967-01-01T00:00:00.000Z","1967-04-01T00:00:00.000Z","1967-07-01T00:00:00.000Z","1967-10-01T00:00:00.000Z","1968-01-01T00:00:00.000Z","1968-04-01T00:00:00.000Z","1968-07-01T00:00:00.000Z","1968-10-01T00:00:00.000Z","1969-01-01T00:00:00.000Z","1969-04-01T00:00:00.000Z","1969-07-01T00:00:00.000Z","1969-10-01T00:00:00.000Z","1970-01-01T00:00:00.000Z","1970-04-01T00:00:00.000Z","1970-07-01T00:00:00.000Z","1970-10-01T00:00:00.000Z","1971-01-01T00:00:00.000Z","1971-04-01T00:00:00.000Z","1971-07-01T00:00:00.000Z","1971-10-01T00:00:00.000Z","1972-01-01T00:00:00.000Z","1972-04-01T00:00:00.000Z","1972-07-01T00:00:00.000Z","1972-10-01T00:00:00.000Z","1973-01-01T00:00:00.000Z","1973-04-01T00:00:00.000Z","1973-07-01T00:00:00.000Z","1973-10-01T00:00:00.000Z","1974-01-01T00:00:00.000Z","1974-04-01T00:00:00.000Z","1974-07-01T00:00:00.000Z","1974-10-01T00:00:00.000Z"],[null,87,82,75,63,50,43,32,35,60,54,55,36,39,null,null,69,57,57,51,45,37,46,39,36,24,32,23,25,32,null,32,59,74,75,60,71,61,71,57,71,68,79,73,76,71,67,75,79,62,63,57,60,49,48,52,57,62,61,66,71,62,61,57,72,83,71,78,79,71,62,74,76,64,62,57,80,73,69,69,71,64,69,62,63,46,56,44,44,52,38,46,36,49,35,44,59,65,65,56,66,53,61,52,51,48,54,49,49,61,null,null,68,44,40,27,28,25,24,24]]},"evals":[],"jsHooks":[]}</script> --- class: middle, inverse # Highcharter https://jkunst.com/highcharter/index.html --- ## Highcharter + The **highcharter** package provides a rich set of plotting functions for dynamic graphics + The R package is, in spirit, similar to **ggplot2** + It uses *mappings*, *aesthetics* and *geometries* (called *types*) + Customization can be added using additional functions in a pipe + Very rich set of chart types --- ## Highcharter .pull-left[ ```r pacman::p_load(highcharter) hchart(object=penguins, type="scatter", mapping = hcaes(x = body_mass_g, y = flipper_length_mm , group = species)) %>% hc_legend(align='right', verticalAlign='top') ``` See how the elements are similar to what you'd put in a **ggplot2** pipe, except they are in a single function Options can be added with pipes ] .pull-right[ <div id="htmlwidget-b989035eb423fb915da9" style="width:100%;height:500px;" class="highchart html-widget"></div> <script type="application/json" data-for="htmlwidget-b989035eb423fb915da9">{"x":{"hc_opts":{"chart":{"reflow":true},"title":{"text":null},"yAxis":{"title":{"text":"flipper_length_mm"},"type":"linear"},"credits":{"enabled":false},"exporting":{"enabled":false},"boost":{"enabled":false},"plotOptions":{"series":{"label":{"enabled":false},"turboThreshold":0,"showInLegend":true},"treemap":{"layoutAlgorithm":"squarified"},"scatter":{"marker":{"symbol":"circle"}}},"series":[{"name":"Adelie","data":[{"species":"Adelie","island":"Torgersen","bill_length_mm":39.1,"bill_depth_mm":18.7,"flipper_length_mm":181,"body_mass_g":3750,"sex":"male","year":2007,"x":3750,"y":181},{"species":"Adelie","island":"Torgersen","bill_length_mm":39.5,"bill_depth_mm":17.4,"flipper_length_mm":186,"body_mass_g":3800,"sex":"female","year":2007,"x":3800,"y":186},{"species":"Adelie","island":"Torgersen","bill_length_mm":40.3,"bill_depth_mm":18,"flipper_length_mm":195,"body_mass_g":3250,"sex":"female","year":2007,"x":3250,"y":195},{"species":"Adelie","island":"Torgersen","bill_length_mm":null,"bill_depth_mm":null,"flipper_length_mm":null,"body_mass_g":null,"sex":null,"year":2007,"x":null,"y":null},{"species":"Adelie","island":"Torgersen","bill_length_mm":36.7,"bill_depth_mm":19.3,"flipper_length_mm":193,"body_mass_g":3450,"sex":"female","year":2007,"x":3450,"y":193},{"species":"Adelie","island":"Torgersen","bill_length_mm":39.3,"bill_depth_mm":20.6,"flipper_length_mm":190,"body_mass_g":3650,"sex":"male","year":2007,"x":3650,"y":190},{"species":"Adelie","island":"Torgersen","bill_length_mm":38.9,"bill_depth_mm":17.8,"flipper_length_mm":181,"body_mass_g":3625,"sex":"female","year":2007,"x":3625,"y":181},{"species":"Adelie","island":"Torgersen","bill_length_mm":39.2,"bill_depth_mm":19.6,"flipper_length_mm":195,"body_mass_g":4675,"sex":"male","year":2007,"x":4675,"y":195},{"species":"Adelie","island":"Torgersen","bill_length_mm":34.1,"bill_depth_mm":18.1,"flipper_length_mm":193,"body_mass_g":3475,"sex":null,"year":2007,"x":3475,"y":193},{"species":"Adelie","island":"Torgersen","bill_length_mm":42,"bill_depth_mm":20.2,"flipper_length_mm":190,"body_mass_g":4250,"sex":null,"year":2007,"x":4250,"y":190},{"species":"Adelie","island":"Torgersen","bill_length_mm":37.8,"bill_depth_mm":17.1,"flipper_length_mm":186,"body_mass_g":3300,"sex":null,"year":2007,"x":3300,"y":186},{"species":"Adelie","island":"Torgersen","bill_length_mm":37.8,"bill_depth_mm":17.3,"flipper_length_mm":180,"body_mass_g":3700,"sex":null,"year":2007,"x":3700,"y":180},{"species":"Adelie","island":"Torgersen","bill_length_mm":41.1,"bill_depth_mm":17.6,"flipper_length_mm":182,"body_mass_g":3200,"sex":"female","year":2007,"x":3200,"y":182},{"species":"Adelie","island":"Torgersen","bill_length_mm":38.6,"bill_depth_mm":21.2,"flipper_length_mm":191,"body_mass_g":3800,"sex":"male","year":2007,"x":3800,"y":191},{"species":"Adelie","island":"Torgersen","bill_length_mm":34.6,"bill_depth_mm":21.1,"flipper_length_mm":198,"body_mass_g":4400,"sex":"male","year":2007,"x":4400,"y":198},{"species":"Adelie","island":"Torgersen","bill_length_mm":36.6,"bill_depth_mm":17.8,"flipper_length_mm":185,"body_mass_g":3700,"sex":"female","year":2007,"x":3700,"y":185},{"species":"Adelie","island":"Torgersen","bill_length_mm":38.7,"bill_depth_mm":19,"flipper_length_mm":195,"body_mass_g":3450,"sex":"female","year":2007,"x":3450,"y":195},{"species":"Adelie","island":"Torgersen","bill_length_mm":42.5,"bill_depth_mm":20.7,"flipper_length_mm":197,"body_mass_g":4500,"sex":"male","year":2007,"x":4500,"y":197},{"species":"Adelie","island":"Torgersen","bill_length_mm":34.4,"bill_depth_mm":18.4,"flipper_length_mm":184,"body_mass_g":3325,"sex":"female","year":2007,"x":3325,"y":184},{"species":"Adelie","island":"Torgersen","bill_length_mm":46,"bill_depth_mm":21.5,"flipper_length_mm":194,"body_mass_g":4200,"sex":"male","year":2007,"x":4200,"y":194},{"species":"Adelie","island":"Biscoe","bill_length_mm":37.8,"bill_depth_mm":18.3,"flipper_length_mm":174,"body_mass_g":3400,"sex":"female","year":2007,"x":3400,"y":174},{"species":"Adelie","island":"Biscoe","bill_length_mm":37.7,"bill_depth_mm":18.7,"flipper_length_mm":180,"body_mass_g":3600,"sex":"male","year":2007,"x":3600,"y":180},{"species":"Adelie","island":"Biscoe","bill_length_mm":35.9,"bill_depth_mm":19.2,"flipper_length_mm":189,"body_mass_g":3800,"sex":"female","year":2007,"x":3800,"y":189},{"species":"Adelie","island":"Biscoe","bill_length_mm":38.2,"bill_depth_mm":18.1,"flipper_length_mm":185,"body_mass_g":3950,"sex":"male","year":2007,"x":3950,"y":185},{"species":"Adelie","island":"Biscoe","bill_length_mm":38.8,"bill_depth_mm":17.2,"flipper_length_mm":180,"body_mass_g":3800,"sex":"male","year":2007,"x":3800,"y":180},{"species":"Adelie","island":"Biscoe","bill_length_mm":35.3,"bill_depth_mm":18.9,"flipper_length_mm":187,"body_mass_g":3800,"sex":"female","year":2007,"x":3800,"y":187},{"species":"Adelie","island":"Biscoe","bill_length_mm":40.6,"bill_depth_mm":18.6,"flipper_length_mm":183,"body_mass_g":3550,"sex":"male","year":2007,"x":3550,"y":183},{"species":"Adelie","island":"Biscoe","bill_length_mm":40.5,"bill_depth_mm":17.9,"flipper_length_mm":187,"body_mass_g":3200,"sex":"female","year":2007,"x":3200,"y":187},{"species":"Adelie","island":"Biscoe","bill_length_mm":37.9,"bill_depth_mm":18.6,"flipper_length_mm":172,"body_mass_g":3150,"sex":"female","year":2007,"x":3150,"y":172},{"species":"Adelie","island":"Biscoe","bill_length_mm":40.5,"bill_depth_mm":18.9,"flipper_length_mm":180,"body_mass_g":3950,"sex":"male","year":2007,"x":3950,"y":180},{"species":"Adelie","island":"Dream","bill_length_mm":39.5,"bill_depth_mm":16.7,"flipper_length_mm":178,"body_mass_g":3250,"sex":"female","year":2007,"x":3250,"y":178},{"species":"Adelie","island":"Dream","bill_length_mm":37.2,"bill_depth_mm":18.1,"flipper_length_mm":178,"body_mass_g":3900,"sex":"male","year":2007,"x":3900,"y":178},{"species":"Adelie","island":"Dream","bill_length_mm":39.5,"bill_depth_mm":17.8,"flipper_length_mm":188,"body_mass_g":3300,"sex":"female","year":2007,"x":3300,"y":188},{"species":"Adelie","island":"Dream","bill_length_mm":40.9,"bill_depth_mm":18.9,"flipper_length_mm":184,"body_mass_g":3900,"sex":"male","year":2007,"x":3900,"y":184},{"species":"Adelie","island":"Dream","bill_length_mm":36.4,"bill_depth_mm":17,"flipper_length_mm":195,"body_mass_g":3325,"sex":"female","year":2007,"x":3325,"y":195},{"species":"Adelie","island":"Dream","bill_length_mm":39.2,"bill_depth_mm":21.1,"flipper_length_mm":196,"body_mass_g":4150,"sex":"male","year":2007,"x":4150,"y":196},{"species":"Adelie","island":"Dream","bill_length_mm":38.8,"bill_depth_mm":20,"flipper_length_mm":190,"body_mass_g":3950,"sex":"male","year":2007,"x":3950,"y":190},{"species":"Adelie","island":"Dream","bill_length_mm":42.2,"bill_depth_mm":18.5,"flipper_length_mm":180,"body_mass_g":3550,"sex":"female","year":2007,"x":3550,"y":180},{"species":"Adelie","island":"Dream","bill_length_mm":37.6,"bill_depth_mm":19.3,"flipper_length_mm":181,"body_mass_g":3300,"sex":"female","year":2007,"x":3300,"y":181},{"species":"Adelie","island":"Dream","bill_length_mm":39.8,"bill_depth_mm":19.1,"flipper_length_mm":184,"body_mass_g":4650,"sex":"male","year":2007,"x":4650,"y":184},{"species":"Adelie","island":"Dream","bill_length_mm":36.5,"bill_depth_mm":18,"flipper_length_mm":182,"body_mass_g":3150,"sex":"female","year":2007,"x":3150,"y":182},{"species":"Adelie","island":"Dream","bill_length_mm":40.8,"bill_depth_mm":18.4,"flipper_length_mm":195,"body_mass_g":3900,"sex":"male","year":2007,"x":3900,"y":195},{"species":"Adelie","island":"Dream","bill_length_mm":36,"bill_depth_mm":18.5,"flipper_length_mm":186,"body_mass_g":3100,"sex":"female","year":2007,"x":3100,"y":186},{"species":"Adelie","island":"Dream","bill_length_mm":44.1,"bill_depth_mm":19.7,"flipper_length_mm":196,"body_mass_g":4400,"sex":"male","year":2007,"x":4400,"y":196},{"species":"Adelie","island":"Dream","bill_length_mm":37,"bill_depth_mm":16.9,"flipper_length_mm":185,"body_mass_g":3000,"sex":"female","year":2007,"x":3000,"y":185},{"species":"Adelie","island":"Dream","bill_length_mm":39.6,"bill_depth_mm":18.8,"flipper_length_mm":190,"body_mass_g":4600,"sex":"male","year":2007,"x":4600,"y":190},{"species":"Adelie","island":"Dream","bill_length_mm":41.1,"bill_depth_mm":19,"flipper_length_mm":182,"body_mass_g":3425,"sex":"male","year":2007,"x":3425,"y":182},{"species":"Adelie","island":"Dream","bill_length_mm":37.5,"bill_depth_mm":18.9,"flipper_length_mm":179,"body_mass_g":2975,"sex":null,"year":2007,"x":2975,"y":179},{"species":"Adelie","island":"Dream","bill_length_mm":36,"bill_depth_mm":17.9,"flipper_length_mm":190,"body_mass_g":3450,"sex":"female","year":2007,"x":3450,"y":190},{"species":"Adelie","island":"Dream","bill_length_mm":42.3,"bill_depth_mm":21.2,"flipper_length_mm":191,"body_mass_g":4150,"sex":"male","year":2007,"x":4150,"y":191},{"species":"Adelie","island":"Biscoe","bill_length_mm":39.6,"bill_depth_mm":17.7,"flipper_length_mm":186,"body_mass_g":3500,"sex":"female","year":2008,"x":3500,"y":186},{"species":"Adelie","island":"Biscoe","bill_length_mm":40.1,"bill_depth_mm":18.9,"flipper_length_mm":188,"body_mass_g":4300,"sex":"male","year":2008,"x":4300,"y":188},{"species":"Adelie","island":"Biscoe","bill_length_mm":35,"bill_depth_mm":17.9,"flipper_length_mm":190,"body_mass_g":3450,"sex":"female","year":2008,"x":3450,"y":190},{"species":"Adelie","island":"Biscoe","bill_length_mm":42,"bill_depth_mm":19.5,"flipper_length_mm":200,"body_mass_g":4050,"sex":"male","year":2008,"x":4050,"y":200},{"species":"Adelie","island":"Biscoe","bill_length_mm":34.5,"bill_depth_mm":18.1,"flipper_length_mm":187,"body_mass_g":2900,"sex":"female","year":2008,"x":2900,"y":187},{"species":"Adelie","island":"Biscoe","bill_length_mm":41.4,"bill_depth_mm":18.6,"flipper_length_mm":191,"body_mass_g":3700,"sex":"male","year":2008,"x":3700,"y":191},{"species":"Adelie","island":"Biscoe","bill_length_mm":39,"bill_depth_mm":17.5,"flipper_length_mm":186,"body_mass_g":3550,"sex":"female","year":2008,"x":3550,"y":186},{"species":"Adelie","island":"Biscoe","bill_length_mm":40.6,"bill_depth_mm":18.8,"flipper_length_mm":193,"body_mass_g":3800,"sex":"male","year":2008,"x":3800,"y":193},{"species":"Adelie","island":"Biscoe","bill_length_mm":36.5,"bill_depth_mm":16.6,"flipper_length_mm":181,"body_mass_g":2850,"sex":"female","year":2008,"x":2850,"y":181},{"species":"Adelie","island":"Biscoe","bill_length_mm":37.6,"bill_depth_mm":19.1,"flipper_length_mm":194,"body_mass_g":3750,"sex":"male","year":2008,"x":3750,"y":194},{"species":"Adelie","island":"Biscoe","bill_length_mm":35.7,"bill_depth_mm":16.9,"flipper_length_mm":185,"body_mass_g":3150,"sex":"female","year":2008,"x":3150,"y":185},{"species":"Adelie","island":"Biscoe","bill_length_mm":41.3,"bill_depth_mm":21.1,"flipper_length_mm":195,"body_mass_g":4400,"sex":"male","year":2008,"x":4400,"y":195},{"species":"Adelie","island":"Biscoe","bill_length_mm":37.6,"bill_depth_mm":17,"flipper_length_mm":185,"body_mass_g":3600,"sex":"female","year":2008,"x":3600,"y":185},{"species":"Adelie","island":"Biscoe","bill_length_mm":41.1,"bill_depth_mm":18.2,"flipper_length_mm":192,"body_mass_g":4050,"sex":"male","year":2008,"x":4050,"y":192},{"species":"Adelie","island":"Biscoe","bill_length_mm":36.4,"bill_depth_mm":17.1,"flipper_length_mm":184,"body_mass_g":2850,"sex":"female","year":2008,"x":2850,"y":184},{"species":"Adelie","island":"Biscoe","bill_length_mm":41.6,"bill_depth_mm":18,"flipper_length_mm":192,"body_mass_g":3950,"sex":"male","year":2008,"x":3950,"y":192},{"species":"Adelie","island":"Biscoe","bill_length_mm":35.5,"bill_depth_mm":16.2,"flipper_length_mm":195,"body_mass_g":3350,"sex":"female","year":2008,"x":3350,"y":195},{"species":"Adelie","island":"Biscoe","bill_length_mm":41.1,"bill_depth_mm":19.1,"flipper_length_mm":188,"body_mass_g":4100,"sex":"male","year":2008,"x":4100,"y":188},{"species":"Adelie","island":"Torgersen","bill_length_mm":35.9,"bill_depth_mm":16.6,"flipper_length_mm":190,"body_mass_g":3050,"sex":"female","year":2008,"x":3050,"y":190},{"species":"Adelie","island":"Torgersen","bill_length_mm":41.8,"bill_depth_mm":19.4,"flipper_length_mm":198,"body_mass_g":4450,"sex":"male","year":2008,"x":4450,"y":198},{"species":"Adelie","island":"Torgersen","bill_length_mm":33.5,"bill_depth_mm":19,"flipper_length_mm":190,"body_mass_g":3600,"sex":"female","year":2008,"x":3600,"y":190},{"species":"Adelie","island":"Torgersen","bill_length_mm":39.7,"bill_depth_mm":18.4,"flipper_length_mm":190,"body_mass_g":3900,"sex":"male","year":2008,"x":3900,"y":190},{"species":"Adelie","island":"Torgersen","bill_length_mm":39.6,"bill_depth_mm":17.2,"flipper_length_mm":196,"body_mass_g":3550,"sex":"female","year":2008,"x":3550,"y":196},{"species":"Adelie","island":"Torgersen","bill_length_mm":45.8,"bill_depth_mm":18.9,"flipper_length_mm":197,"body_mass_g":4150,"sex":"male","year":2008,"x":4150,"y":197},{"species":"Adelie","island":"Torgersen","bill_length_mm":35.5,"bill_depth_mm":17.5,"flipper_length_mm":190,"body_mass_g":3700,"sex":"female","year":2008,"x":3700,"y":190},{"species":"Adelie","island":"Torgersen","bill_length_mm":42.8,"bill_depth_mm":18.5,"flipper_length_mm":195,"body_mass_g":4250,"sex":"male","year":2008,"x":4250,"y":195},{"species":"Adelie","island":"Torgersen","bill_length_mm":40.9,"bill_depth_mm":16.8,"flipper_length_mm":191,"body_mass_g":3700,"sex":"female","year":2008,"x":3700,"y":191},{"species":"Adelie","island":"Torgersen","bill_length_mm":37.2,"bill_depth_mm":19.4,"flipper_length_mm":184,"body_mass_g":3900,"sex":"male","year":2008,"x":3900,"y":184},{"species":"Adelie","island":"Torgersen","bill_length_mm":36.2,"bill_depth_mm":16.1,"flipper_length_mm":187,"body_mass_g":3550,"sex":"female","year":2008,"x":3550,"y":187},{"species":"Adelie","island":"Torgersen","bill_length_mm":42.1,"bill_depth_mm":19.1,"flipper_length_mm":195,"body_mass_g":4000,"sex":"male","year":2008,"x":4000,"y":195},{"species":"Adelie","island":"Torgersen","bill_length_mm":34.6,"bill_depth_mm":17.2,"flipper_length_mm":189,"body_mass_g":3200,"sex":"female","year":2008,"x":3200,"y":189},{"species":"Adelie","island":"Torgersen","bill_length_mm":42.9,"bill_depth_mm":17.6,"flipper_length_mm":196,"body_mass_g":4700,"sex":"male","year":2008,"x":4700,"y":196},{"species":"Adelie","island":"Torgersen","bill_length_mm":36.7,"bill_depth_mm":18.8,"flipper_length_mm":187,"body_mass_g":3800,"sex":"female","year":2008,"x":3800,"y":187},{"species":"Adelie","island":"Torgersen","bill_length_mm":35.1,"bill_depth_mm":19.4,"flipper_length_mm":193,"body_mass_g":4200,"sex":"male","year":2008,"x":4200,"y":193},{"species":"Adelie","island":"Dream","bill_length_mm":37.3,"bill_depth_mm":17.8,"flipper_length_mm":191,"body_mass_g":3350,"sex":"female","year":2008,"x":3350,"y":191},{"species":"Adelie","island":"Dream","bill_length_mm":41.3,"bill_depth_mm":20.3,"flipper_length_mm":194,"body_mass_g":3550,"sex":"male","year":2008,"x":3550,"y":194},{"species":"Adelie","island":"Dream","bill_length_mm":36.3,"bill_depth_mm":19.5,"flipper_length_mm":190,"body_mass_g":3800,"sex":"male","year":2008,"x":3800,"y":190},{"species":"Adelie","island":"Dream","bill_length_mm":36.9,"bill_depth_mm":18.6,"flipper_length_mm":189,"body_mass_g":3500,"sex":"female","year":2008,"x":3500,"y":189},{"species":"Adelie","island":"Dream","bill_length_mm":38.3,"bill_depth_mm":19.2,"flipper_length_mm":189,"body_mass_g":3950,"sex":"male","year":2008,"x":3950,"y":189},{"species":"Adelie","island":"Dream","bill_length_mm":38.9,"bill_depth_mm":18.8,"flipper_length_mm":190,"body_mass_g":3600,"sex":"female","year":2008,"x":3600,"y":190},{"species":"Adelie","island":"Dream","bill_length_mm":35.7,"bill_depth_mm":18,"flipper_length_mm":202,"body_mass_g":3550,"sex":"female","year":2008,"x":3550,"y":202},{"species":"Adelie","island":"Dream","bill_length_mm":41.1,"bill_depth_mm":18.1,"flipper_length_mm":205,"body_mass_g":4300,"sex":"male","year":2008,"x":4300,"y":205},{"species":"Adelie","island":"Dream","bill_length_mm":34,"bill_depth_mm":17.1,"flipper_length_mm":185,"body_mass_g":3400,"sex":"female","year":2008,"x":3400,"y":185},{"species":"Adelie","island":"Dream","bill_length_mm":39.6,"bill_depth_mm":18.1,"flipper_length_mm":186,"body_mass_g":4450,"sex":"male","year":2008,"x":4450,"y":186},{"species":"Adelie","island":"Dream","bill_length_mm":36.2,"bill_depth_mm":17.3,"flipper_length_mm":187,"body_mass_g":3300,"sex":"female","year":2008,"x":3300,"y":187},{"species":"Adelie","island":"Dream","bill_length_mm":40.8,"bill_depth_mm":18.9,"flipper_length_mm":208,"body_mass_g":4300,"sex":"male","year":2008,"x":4300,"y":208},{"species":"Adelie","island":"Dream","bill_length_mm":38.1,"bill_depth_mm":18.6,"flipper_length_mm":190,"body_mass_g":3700,"sex":"female","year":2008,"x":3700,"y":190},{"species":"Adelie","island":"Dream","bill_length_mm":40.3,"bill_depth_mm":18.5,"flipper_length_mm":196,"body_mass_g":4350,"sex":"male","year":2008,"x":4350,"y":196},{"species":"Adelie","island":"Dream","bill_length_mm":33.1,"bill_depth_mm":16.1,"flipper_length_mm":178,"body_mass_g":2900,"sex":"female","year":2008,"x":2900,"y":178},{"species":"Adelie","island":"Dream","bill_length_mm":43.2,"bill_depth_mm":18.5,"flipper_length_mm":192,"body_mass_g":4100,"sex":"male","year":2008,"x":4100,"y":192},{"species":"Adelie","island":"Biscoe","bill_length_mm":35,"bill_depth_mm":17.9,"flipper_length_mm":192,"body_mass_g":3725,"sex":"female","year":2009,"x":3725,"y":192},{"species":"Adelie","island":"Biscoe","bill_length_mm":41,"bill_depth_mm":20,"flipper_length_mm":203,"body_mass_g":4725,"sex":"male","year":2009,"x":4725,"y":203},{"species":"Adelie","island":"Biscoe","bill_length_mm":37.7,"bill_depth_mm":16,"flipper_length_mm":183,"body_mass_g":3075,"sex":"female","year":2009,"x":3075,"y":183},{"species":"Adelie","island":"Biscoe","bill_length_mm":37.8,"bill_depth_mm":20,"flipper_length_mm":190,"body_mass_g":4250,"sex":"male","year":2009,"x":4250,"y":190},{"species":"Adelie","island":"Biscoe","bill_length_mm":37.9,"bill_depth_mm":18.6,"flipper_length_mm":193,"body_mass_g":2925,"sex":"female","year":2009,"x":2925,"y":193},{"species":"Adelie","island":"Biscoe","bill_length_mm":39.7,"bill_depth_mm":18.9,"flipper_length_mm":184,"body_mass_g":3550,"sex":"male","year":2009,"x":3550,"y":184},{"species":"Adelie","island":"Biscoe","bill_length_mm":38.6,"bill_depth_mm":17.2,"flipper_length_mm":199,"body_mass_g":3750,"sex":"female","year":2009,"x":3750,"y":199},{"species":"Adelie","island":"Biscoe","bill_length_mm":38.2,"bill_depth_mm":20,"flipper_length_mm":190,"body_mass_g":3900,"sex":"male","year":2009,"x":3900,"y":190},{"species":"Adelie","island":"Biscoe","bill_length_mm":38.1,"bill_depth_mm":17,"flipper_length_mm":181,"body_mass_g":3175,"sex":"female","year":2009,"x":3175,"y":181},{"species":"Adelie","island":"Biscoe","bill_length_mm":43.2,"bill_depth_mm":19,"flipper_length_mm":197,"body_mass_g":4775,"sex":"male","year":2009,"x":4775,"y":197},{"species":"Adelie","island":"Biscoe","bill_length_mm":38.1,"bill_depth_mm":16.5,"flipper_length_mm":198,"body_mass_g":3825,"sex":"female","year":2009,"x":3825,"y":198},{"species":"Adelie","island":"Biscoe","bill_length_mm":45.6,"bill_depth_mm":20.3,"flipper_length_mm":191,"body_mass_g":4600,"sex":"male","year":2009,"x":4600,"y":191},{"species":"Adelie","island":"Biscoe","bill_length_mm":39.7,"bill_depth_mm":17.7,"flipper_length_mm":193,"body_mass_g":3200,"sex":"female","year":2009,"x":3200,"y":193},{"species":"Adelie","island":"Biscoe","bill_length_mm":42.2,"bill_depth_mm":19.5,"flipper_length_mm":197,"body_mass_g":4275,"sex":"male","year":2009,"x":4275,"y":197},{"species":"Adelie","island":"Biscoe","bill_length_mm":39.6,"bill_depth_mm":20.7,"flipper_length_mm":191,"body_mass_g":3900,"sex":"female","year":2009,"x":3900,"y":191},{"species":"Adelie","island":"Biscoe","bill_length_mm":42.7,"bill_depth_mm":18.3,"flipper_length_mm":196,"body_mass_g":4075,"sex":"male","year":2009,"x":4075,"y":196},{"species":"Adelie","island":"Torgersen","bill_length_mm":38.6,"bill_depth_mm":17,"flipper_length_mm":188,"body_mass_g":2900,"sex":"female","year":2009,"x":2900,"y":188},{"species":"Adelie","island":"Torgersen","bill_length_mm":37.3,"bill_depth_mm":20.5,"flipper_length_mm":199,"body_mass_g":3775,"sex":"male","year":2009,"x":3775,"y":199},{"species":"Adelie","island":"Torgersen","bill_length_mm":35.7,"bill_depth_mm":17,"flipper_length_mm":189,"body_mass_g":3350,"sex":"female","year":2009,"x":3350,"y":189},{"species":"Adelie","island":"Torgersen","bill_length_mm":41.1,"bill_depth_mm":18.6,"flipper_length_mm":189,"body_mass_g":3325,"sex":"male","year":2009,"x":3325,"y":189},{"species":"Adelie","island":"Torgersen","bill_length_mm":36.2,"bill_depth_mm":17.2,"flipper_length_mm":187,"body_mass_g":3150,"sex":"female","year":2009,"x":3150,"y":187},{"species":"Adelie","island":"Torgersen","bill_length_mm":37.7,"bill_depth_mm":19.8,"flipper_length_mm":198,"body_mass_g":3500,"sex":"male","year":2009,"x":3500,"y":198},{"species":"Adelie","island":"Torgersen","bill_length_mm":40.2,"bill_depth_mm":17,"flipper_length_mm":176,"body_mass_g":3450,"sex":"female","year":2009,"x":3450,"y":176},{"species":"Adelie","island":"Torgersen","bill_length_mm":41.4,"bill_depth_mm":18.5,"flipper_length_mm":202,"body_mass_g":3875,"sex":"male","year":2009,"x":3875,"y":202},{"species":"Adelie","island":"Torgersen","bill_length_mm":35.2,"bill_depth_mm":15.9,"flipper_length_mm":186,"body_mass_g":3050,"sex":"female","year":2009,"x":3050,"y":186},{"species":"Adelie","island":"Torgersen","bill_length_mm":40.6,"bill_depth_mm":19,"flipper_length_mm":199,"body_mass_g":4000,"sex":"male","year":2009,"x":4000,"y":199},{"species":"Adelie","island":"Torgersen","bill_length_mm":38.8,"bill_depth_mm":17.6,"flipper_length_mm":191,"body_mass_g":3275,"sex":"female","year":2009,"x":3275,"y":191},{"species":"Adelie","island":"Torgersen","bill_length_mm":41.5,"bill_depth_mm":18.3,"flipper_length_mm":195,"body_mass_g":4300,"sex":"male","year":2009,"x":4300,"y":195},{"species":"Adelie","island":"Torgersen","bill_length_mm":39,"bill_depth_mm":17.1,"flipper_length_mm":191,"body_mass_g":3050,"sex":"female","year":2009,"x":3050,"y":191},{"species":"Adelie","island":"Torgersen","bill_length_mm":44.1,"bill_depth_mm":18,"flipper_length_mm":210,"body_mass_g":4000,"sex":"male","year":2009,"x":4000,"y":210},{"species":"Adelie","island":"Torgersen","bill_length_mm":38.5,"bill_depth_mm":17.9,"flipper_length_mm":190,"body_mass_g":3325,"sex":"female","year":2009,"x":3325,"y":190},{"species":"Adelie","island":"Torgersen","bill_length_mm":43.1,"bill_depth_mm":19.2,"flipper_length_mm":197,"body_mass_g":3500,"sex":"male","year":2009,"x":3500,"y":197},{"species":"Adelie","island":"Dream","bill_length_mm":36.8,"bill_depth_mm":18.5,"flipper_length_mm":193,"body_mass_g":3500,"sex":"female","year":2009,"x":3500,"y":193},{"species":"Adelie","island":"Dream","bill_length_mm":37.5,"bill_depth_mm":18.5,"flipper_length_mm":199,"body_mass_g":4475,"sex":"male","year":2009,"x":4475,"y":199},{"species":"Adelie","island":"Dream","bill_length_mm":38.1,"bill_depth_mm":17.6,"flipper_length_mm":187,"body_mass_g":3425,"sex":"female","year":2009,"x":3425,"y":187},{"species":"Adelie","island":"Dream","bill_length_mm":41.1,"bill_depth_mm":17.5,"flipper_length_mm":190,"body_mass_g":3900,"sex":"male","year":2009,"x":3900,"y":190},{"species":"Adelie","island":"Dream","bill_length_mm":35.6,"bill_depth_mm":17.5,"flipper_length_mm":191,"body_mass_g":3175,"sex":"female","year":2009,"x":3175,"y":191},{"species":"Adelie","island":"Dream","bill_length_mm":40.2,"bill_depth_mm":20.1,"flipper_length_mm":200,"body_mass_g":3975,"sex":"male","year":2009,"x":3975,"y":200},{"species":"Adelie","island":"Dream","bill_length_mm":37,"bill_depth_mm":16.5,"flipper_length_mm":185,"body_mass_g":3400,"sex":"female","year":2009,"x":3400,"y":185},{"species":"Adelie","island":"Dream","bill_length_mm":39.7,"bill_depth_mm":17.9,"flipper_length_mm":193,"body_mass_g":4250,"sex":"male","year":2009,"x":4250,"y":193},{"species":"Adelie","island":"Dream","bill_length_mm":40.2,"bill_depth_mm":17.1,"flipper_length_mm":193,"body_mass_g":3400,"sex":"female","year":2009,"x":3400,"y":193},{"species":"Adelie","island":"Dream","bill_length_mm":40.6,"bill_depth_mm":17.2,"flipper_length_mm":187,"body_mass_g":3475,"sex":"male","year":2009,"x":3475,"y":187},{"species":"Adelie","island":"Dream","bill_length_mm":32.1,"bill_depth_mm":15.5,"flipper_length_mm":188,"body_mass_g":3050,"sex":"female","year":2009,"x":3050,"y":188},{"species":"Adelie","island":"Dream","bill_length_mm":40.7,"bill_depth_mm":17,"flipper_length_mm":190,"body_mass_g":3725,"sex":"male","year":2009,"x":3725,"y":190},{"species":"Adelie","island":"Dream","bill_length_mm":37.3,"bill_depth_mm":16.8,"flipper_length_mm":192,"body_mass_g":3000,"sex":"female","year":2009,"x":3000,"y":192},{"species":"Adelie","island":"Dream","bill_length_mm":39,"bill_depth_mm":18.7,"flipper_length_mm":185,"body_mass_g":3650,"sex":"male","year":2009,"x":3650,"y":185},{"species":"Adelie","island":"Dream","bill_length_mm":39.2,"bill_depth_mm":18.6,"flipper_length_mm":190,"body_mass_g":4250,"sex":"male","year":2009,"x":4250,"y":190},{"species":"Adelie","island":"Dream","bill_length_mm":36.6,"bill_depth_mm":18.4,"flipper_length_mm":184,"body_mass_g":3475,"sex":"female","year":2009,"x":3475,"y":184},{"species":"Adelie","island":"Dream","bill_length_mm":36,"bill_depth_mm":17.8,"flipper_length_mm":195,"body_mass_g":3450,"sex":"female","year":2009,"x":3450,"y":195},{"species":"Adelie","island":"Dream","bill_length_mm":37.8,"bill_depth_mm":18.1,"flipper_length_mm":193,"body_mass_g":3750,"sex":"male","year":2009,"x":3750,"y":193},{"species":"Adelie","island":"Dream","bill_length_mm":36,"bill_depth_mm":17.1,"flipper_length_mm":187,"body_mass_g":3700,"sex":"female","year":2009,"x":3700,"y":187},{"species":"Adelie","island":"Dream","bill_length_mm":41.5,"bill_depth_mm":18.5,"flipper_length_mm":201,"body_mass_g":4000,"sex":"male","year":2009,"x":4000,"y":201}],"type":"scatter"},{"name":"Chinstrap","data":[{"species":"Chinstrap","island":"Dream","bill_length_mm":46.5,"bill_depth_mm":17.9,"flipper_length_mm":192,"body_mass_g":3500,"sex":"female","year":2007,"x":3500,"y":192},{"species":"Chinstrap","island":"Dream","bill_length_mm":50,"bill_depth_mm":19.5,"flipper_length_mm":196,"body_mass_g":3900,"sex":"male","year":2007,"x":3900,"y":196},{"species":"Chinstrap","island":"Dream","bill_length_mm":51.3,"bill_depth_mm":19.2,"flipper_length_mm":193,"body_mass_g":3650,"sex":"male","year":2007,"x":3650,"y":193},{"species":"Chinstrap","island":"Dream","bill_length_mm":45.4,"bill_depth_mm":18.7,"flipper_length_mm":188,"body_mass_g":3525,"sex":"female","year":2007,"x":3525,"y":188},{"species":"Chinstrap","island":"Dream","bill_length_mm":52.7,"bill_depth_mm":19.8,"flipper_length_mm":197,"body_mass_g":3725,"sex":"male","year":2007,"x":3725,"y":197},{"species":"Chinstrap","island":"Dream","bill_length_mm":45.2,"bill_depth_mm":17.8,"flipper_length_mm":198,"body_mass_g":3950,"sex":"female","year":2007,"x":3950,"y":198},{"species":"Chinstrap","island":"Dream","bill_length_mm":46.1,"bill_depth_mm":18.2,"flipper_length_mm":178,"body_mass_g":3250,"sex":"female","year":2007,"x":3250,"y":178},{"species":"Chinstrap","island":"Dream","bill_length_mm":51.3,"bill_depth_mm":18.2,"flipper_length_mm":197,"body_mass_g":3750,"sex":"male","year":2007,"x":3750,"y":197},{"species":"Chinstrap","island":"Dream","bill_length_mm":46,"bill_depth_mm":18.9,"flipper_length_mm":195,"body_mass_g":4150,"sex":"female","year":2007,"x":4150,"y":195},{"species":"Chinstrap","island":"Dream","bill_length_mm":51.3,"bill_depth_mm":19.9,"flipper_length_mm":198,"body_mass_g":3700,"sex":"male","year":2007,"x":3700,"y":198},{"species":"Chinstrap","island":"Dream","bill_length_mm":46.6,"bill_depth_mm":17.8,"flipper_length_mm":193,"body_mass_g":3800,"sex":"female","year":2007,"x":3800,"y":193},{"species":"Chinstrap","island":"Dream","bill_length_mm":51.7,"bill_depth_mm":20.3,"flipper_length_mm":194,"body_mass_g":3775,"sex":"male","year":2007,"x":3775,"y":194},{"species":"Chinstrap","island":"Dream","bill_length_mm":47,"bill_depth_mm":17.3,"flipper_length_mm":185,"body_mass_g":3700,"sex":"female","year":2007,"x":3700,"y":185},{"species":"Chinstrap","island":"Dream","bill_length_mm":52,"bill_depth_mm":18.1,"flipper_length_mm":201,"body_mass_g":4050,"sex":"male","year":2007,"x":4050,"y":201},{"species":"Chinstrap","island":"Dream","bill_length_mm":45.9,"bill_depth_mm":17.1,"flipper_length_mm":190,"body_mass_g":3575,"sex":"female","year":2007,"x":3575,"y":190},{"species":"Chinstrap","island":"Dream","bill_length_mm":50.5,"bill_depth_mm":19.6,"flipper_length_mm":201,"body_mass_g":4050,"sex":"male","year":2007,"x":4050,"y":201},{"species":"Chinstrap","island":"Dream","bill_length_mm":50.3,"bill_depth_mm":20,"flipper_length_mm":197,"body_mass_g":3300,"sex":"male","year":2007,"x":3300,"y":197},{"species":"Chinstrap","island":"Dream","bill_length_mm":58,"bill_depth_mm":17.8,"flipper_length_mm":181,"body_mass_g":3700,"sex":"female","year":2007,"x":3700,"y":181},{"species":"Chinstrap","island":"Dream","bill_length_mm":46.4,"bill_depth_mm":18.6,"flipper_length_mm":190,"body_mass_g":3450,"sex":"female","year":2007,"x":3450,"y":190},{"species":"Chinstrap","island":"Dream","bill_length_mm":49.2,"bill_depth_mm":18.2,"flipper_length_mm":195,"body_mass_g":4400,"sex":"male","year":2007,"x":4400,"y":195},{"species":"Chinstrap","island":"Dream","bill_length_mm":42.4,"bill_depth_mm":17.3,"flipper_length_mm":181,"body_mass_g":3600,"sex":"female","year":2007,"x":3600,"y":181},{"species":"Chinstrap","island":"Dream","bill_length_mm":48.5,"bill_depth_mm":17.5,"flipper_length_mm":191,"body_mass_g":3400,"sex":"male","year":2007,"x":3400,"y":191},{"species":"Chinstrap","island":"Dream","bill_length_mm":43.2,"bill_depth_mm":16.6,"flipper_length_mm":187,"body_mass_g":2900,"sex":"female","year":2007,"x":2900,"y":187},{"species":"Chinstrap","island":"Dream","bill_length_mm":50.6,"bill_depth_mm":19.4,"flipper_length_mm":193,"body_mass_g":3800,"sex":"male","year":2007,"x":3800,"y":193},{"species":"Chinstrap","island":"Dream","bill_length_mm":46.7,"bill_depth_mm":17.9,"flipper_length_mm":195,"body_mass_g":3300,"sex":"female","year":2007,"x":3300,"y":195},{"species":"Chinstrap","island":"Dream","bill_length_mm":52,"bill_depth_mm":19,"flipper_length_mm":197,"body_mass_g":4150,"sex":"male","year":2007,"x":4150,"y":197},{"species":"Chinstrap","island":"Dream","bill_length_mm":50.5,"bill_depth_mm":18.4,"flipper_length_mm":200,"body_mass_g":3400,"sex":"female","year":2008,"x":3400,"y":200},{"species":"Chinstrap","island":"Dream","bill_length_mm":49.5,"bill_depth_mm":19,"flipper_length_mm":200,"body_mass_g":3800,"sex":"male","year":2008,"x":3800,"y":200},{"species":"Chinstrap","island":"Dream","bill_length_mm":46.4,"bill_depth_mm":17.8,"flipper_length_mm":191,"body_mass_g":3700,"sex":"female","year":2008,"x":3700,"y":191},{"species":"Chinstrap","island":"Dream","bill_length_mm":52.8,"bill_depth_mm":20,"flipper_length_mm":205,"body_mass_g":4550,"sex":"male","year":2008,"x":4550,"y":205},{"species":"Chinstrap","island":"Dream","bill_length_mm":40.9,"bill_depth_mm":16.6,"flipper_length_mm":187,"body_mass_g":3200,"sex":"female","year":2008,"x":3200,"y":187},{"species":"Chinstrap","island":"Dream","bill_length_mm":54.2,"bill_depth_mm":20.8,"flipper_length_mm":201,"body_mass_g":4300,"sex":"male","year":2008,"x":4300,"y":201},{"species":"Chinstrap","island":"Dream","bill_length_mm":42.5,"bill_depth_mm":16.7,"flipper_length_mm":187,"body_mass_g":3350,"sex":"female","year":2008,"x":3350,"y":187},{"species":"Chinstrap","island":"Dream","bill_length_mm":51,"bill_depth_mm":18.8,"flipper_length_mm":203,"body_mass_g":4100,"sex":"male","year":2008,"x":4100,"y":203},{"species":"Chinstrap","island":"Dream","bill_length_mm":49.7,"bill_depth_mm":18.6,"flipper_length_mm":195,"body_mass_g":3600,"sex":"male","year":2008,"x":3600,"y":195},{"species":"Chinstrap","island":"Dream","bill_length_mm":47.5,"bill_depth_mm":16.8,"flipper_length_mm":199,"body_mass_g":3900,"sex":"female","year":2008,"x":3900,"y":199},{"species":"Chinstrap","island":"Dream","bill_length_mm":47.6,"bill_depth_mm":18.3,"flipper_length_mm":195,"body_mass_g":3850,"sex":"female","year":2008,"x":3850,"y":195},{"species":"Chinstrap","island":"Dream","bill_length_mm":52,"bill_depth_mm":20.7,"flipper_length_mm":210,"body_mass_g":4800,"sex":"male","year":2008,"x":4800,"y":210},{"species":"Chinstrap","island":"Dream","bill_length_mm":46.9,"bill_depth_mm":16.6,"flipper_length_mm":192,"body_mass_g":2700,"sex":"female","year":2008,"x":2700,"y":192},{"species":"Chinstrap","island":"Dream","bill_length_mm":53.5,"bill_depth_mm":19.9,"flipper_length_mm":205,"body_mass_g":4500,"sex":"male","year":2008,"x":4500,"y":205},{"species":"Chinstrap","island":"Dream","bill_length_mm":49,"bill_depth_mm":19.5,"flipper_length_mm":210,"body_mass_g":3950,"sex":"male","year":2008,"x":3950,"y":210},{"species":"Chinstrap","island":"Dream","bill_length_mm":46.2,"bill_depth_mm":17.5,"flipper_length_mm":187,"body_mass_g":3650,"sex":"female","year":2008,"x":3650,"y":187},{"species":"Chinstrap","island":"Dream","bill_length_mm":50.9,"bill_depth_mm":19.1,"flipper_length_mm":196,"body_mass_g":3550,"sex":"male","year":2008,"x":3550,"y":196},{"species":"Chinstrap","island":"Dream","bill_length_mm":45.5,"bill_depth_mm":17,"flipper_length_mm":196,"body_mass_g":3500,"sex":"female","year":2008,"x":3500,"y":196},{"species":"Chinstrap","island":"Dream","bill_length_mm":50.9,"bill_depth_mm":17.9,"flipper_length_mm":196,"body_mass_g":3675,"sex":"female","year":2009,"x":3675,"y":196},{"species":"Chinstrap","island":"Dream","bill_length_mm":50.8,"bill_depth_mm":18.5,"flipper_length_mm":201,"body_mass_g":4450,"sex":"male","year":2009,"x":4450,"y":201},{"species":"Chinstrap","island":"Dream","bill_length_mm":50.1,"bill_depth_mm":17.9,"flipper_length_mm":190,"body_mass_g":3400,"sex":"female","year":2009,"x":3400,"y":190},{"species":"Chinstrap","island":"Dream","bill_length_mm":49,"bill_depth_mm":19.6,"flipper_length_mm":212,"body_mass_g":4300,"sex":"male","year":2009,"x":4300,"y":212},{"species":"Chinstrap","island":"Dream","bill_length_mm":51.5,"bill_depth_mm":18.7,"flipper_length_mm":187,"body_mass_g":3250,"sex":"male","year":2009,"x":3250,"y":187},{"species":"Chinstrap","island":"Dream","bill_length_mm":49.8,"bill_depth_mm":17.3,"flipper_length_mm":198,"body_mass_g":3675,"sex":"female","year":2009,"x":3675,"y":198},{"species":"Chinstrap","island":"Dream","bill_length_mm":48.1,"bill_depth_mm":16.4,"flipper_length_mm":199,"body_mass_g":3325,"sex":"female","year":2009,"x":3325,"y":199},{"species":"Chinstrap","island":"Dream","bill_length_mm":51.4,"bill_depth_mm":19,"flipper_length_mm":201,"body_mass_g":3950,"sex":"male","year":2009,"x":3950,"y":201},{"species":"Chinstrap","island":"Dream","bill_length_mm":45.7,"bill_depth_mm":17.3,"flipper_length_mm":193,"body_mass_g":3600,"sex":"female","year":2009,"x":3600,"y":193},{"species":"Chinstrap","island":"Dream","bill_length_mm":50.7,"bill_depth_mm":19.7,"flipper_length_mm":203,"body_mass_g":4050,"sex":"male","year":2009,"x":4050,"y":203},{"species":"Chinstrap","island":"Dream","bill_length_mm":42.5,"bill_depth_mm":17.3,"flipper_length_mm":187,"body_mass_g":3350,"sex":"female","year":2009,"x":3350,"y":187},{"species":"Chinstrap","island":"Dream","bill_length_mm":52.2,"bill_depth_mm":18.8,"flipper_length_mm":197,"body_mass_g":3450,"sex":"male","year":2009,"x":3450,"y":197},{"species":"Chinstrap","island":"Dream","bill_length_mm":45.2,"bill_depth_mm":16.6,"flipper_length_mm":191,"body_mass_g":3250,"sex":"female","year":2009,"x":3250,"y":191},{"species":"Chinstrap","island":"Dream","bill_length_mm":49.3,"bill_depth_mm":19.9,"flipper_length_mm":203,"body_mass_g":4050,"sex":"male","year":2009,"x":4050,"y":203},{"species":"Chinstrap","island":"Dream","bill_length_mm":50.2,"bill_depth_mm":18.8,"flipper_length_mm":202,"body_mass_g":3800,"sex":"male","year":2009,"x":3800,"y":202},{"species":"Chinstrap","island":"Dream","bill_length_mm":45.6,"bill_depth_mm":19.4,"flipper_length_mm":194,"body_mass_g":3525,"sex":"female","year":2009,"x":3525,"y":194},{"species":"Chinstrap","island":"Dream","bill_length_mm":51.9,"bill_depth_mm":19.5,"flipper_length_mm":206,"body_mass_g":3950,"sex":"male","year":2009,"x":3950,"y":206},{"species":"Chinstrap","island":"Dream","bill_length_mm":46.8,"bill_depth_mm":16.5,"flipper_length_mm":189,"body_mass_g":3650,"sex":"female","year":2009,"x":3650,"y":189},{"species":"Chinstrap","island":"Dream","bill_length_mm":45.7,"bill_depth_mm":17,"flipper_length_mm":195,"body_mass_g":3650,"sex":"female","year":2009,"x":3650,"y":195},{"species":"Chinstrap","island":"Dream","bill_length_mm":55.8,"bill_depth_mm":19.8,"flipper_length_mm":207,"body_mass_g":4000,"sex":"male","year":2009,"x":4000,"y":207},{"species":"Chinstrap","island":"Dream","bill_length_mm":43.5,"bill_depth_mm":18.1,"flipper_length_mm":202,"body_mass_g":3400,"sex":"female","year":2009,"x":3400,"y":202},{"species":"Chinstrap","island":"Dream","bill_length_mm":49.6,"bill_depth_mm":18.2,"flipper_length_mm":193,"body_mass_g":3775,"sex":"male","year":2009,"x":3775,"y":193},{"species":"Chinstrap","island":"Dream","bill_length_mm":50.8,"bill_depth_mm":19,"flipper_length_mm":210,"body_mass_g":4100,"sex":"male","year":2009,"x":4100,"y":210},{"species":"Chinstrap","island":"Dream","bill_length_mm":50.2,"bill_depth_mm":18.7,"flipper_length_mm":198,"body_mass_g":3775,"sex":"female","year":2009,"x":3775,"y":198}],"type":"scatter"},{"name":"Gentoo","data":[{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.1,"bill_depth_mm":13.2,"flipper_length_mm":211,"body_mass_g":4500,"sex":"female","year":2007,"x":4500,"y":211},{"species":"Gentoo","island":"Biscoe","bill_length_mm":50,"bill_depth_mm":16.3,"flipper_length_mm":230,"body_mass_g":5700,"sex":"male","year":2007,"x":5700,"y":230},{"species":"Gentoo","island":"Biscoe","bill_length_mm":48.7,"bill_depth_mm":14.1,"flipper_length_mm":210,"body_mass_g":4450,"sex":"female","year":2007,"x":4450,"y":210},{"species":"Gentoo","island":"Biscoe","bill_length_mm":50,"bill_depth_mm":15.2,"flipper_length_mm":218,"body_mass_g":5700,"sex":"male","year":2007,"x":5700,"y":218},{"species":"Gentoo","island":"Biscoe","bill_length_mm":47.6,"bill_depth_mm":14.5,"flipper_length_mm":215,"body_mass_g":5400,"sex":"male","year":2007,"x":5400,"y":215},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.5,"bill_depth_mm":13.5,"flipper_length_mm":210,"body_mass_g":4550,"sex":"female","year":2007,"x":4550,"y":210},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.4,"bill_depth_mm":14.6,"flipper_length_mm":211,"body_mass_g":4800,"sex":"female","year":2007,"x":4800,"y":211},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.7,"bill_depth_mm":15.3,"flipper_length_mm":219,"body_mass_g":5200,"sex":"male","year":2007,"x":5200,"y":219},{"species":"Gentoo","island":"Biscoe","bill_length_mm":43.3,"bill_depth_mm":13.4,"flipper_length_mm":209,"body_mass_g":4400,"sex":"female","year":2007,"x":4400,"y":209},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.8,"bill_depth_mm":15.4,"flipper_length_mm":215,"body_mass_g":5150,"sex":"male","year":2007,"x":5150,"y":215},{"species":"Gentoo","island":"Biscoe","bill_length_mm":40.9,"bill_depth_mm":13.7,"flipper_length_mm":214,"body_mass_g":4650,"sex":"female","year":2007,"x":4650,"y":214},{"species":"Gentoo","island":"Biscoe","bill_length_mm":49,"bill_depth_mm":16.1,"flipper_length_mm":216,"body_mass_g":5550,"sex":"male","year":2007,"x":5550,"y":216},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.5,"bill_depth_mm":13.7,"flipper_length_mm":214,"body_mass_g":4650,"sex":"female","year":2007,"x":4650,"y":214},{"species":"Gentoo","island":"Biscoe","bill_length_mm":48.4,"bill_depth_mm":14.6,"flipper_length_mm":213,"body_mass_g":5850,"sex":"male","year":2007,"x":5850,"y":213},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.8,"bill_depth_mm":14.6,"flipper_length_mm":210,"body_mass_g":4200,"sex":"female","year":2007,"x":4200,"y":210},{"species":"Gentoo","island":"Biscoe","bill_length_mm":49.3,"bill_depth_mm":15.7,"flipper_length_mm":217,"body_mass_g":5850,"sex":"male","year":2007,"x":5850,"y":217},{"species":"Gentoo","island":"Biscoe","bill_length_mm":42,"bill_depth_mm":13.5,"flipper_length_mm":210,"body_mass_g":4150,"sex":"female","year":2007,"x":4150,"y":210},{"species":"Gentoo","island":"Biscoe","bill_length_mm":49.2,"bill_depth_mm":15.2,"flipper_length_mm":221,"body_mass_g":6300,"sex":"male","year":2007,"x":6300,"y":221},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.2,"bill_depth_mm":14.5,"flipper_length_mm":209,"body_mass_g":4800,"sex":"female","year":2007,"x":4800,"y":209},{"species":"Gentoo","island":"Biscoe","bill_length_mm":48.7,"bill_depth_mm":15.1,"flipper_length_mm":222,"body_mass_g":5350,"sex":"male","year":2007,"x":5350,"y":222},{"species":"Gentoo","island":"Biscoe","bill_length_mm":50.2,"bill_depth_mm":14.3,"flipper_length_mm":218,"body_mass_g":5700,"sex":"male","year":2007,"x":5700,"y":218},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.1,"bill_depth_mm":14.5,"flipper_length_mm":215,"body_mass_g":5000,"sex":"female","year":2007,"x":5000,"y":215},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.5,"bill_depth_mm":14.5,"flipper_length_mm":213,"body_mass_g":4400,"sex":"female","year":2007,"x":4400,"y":213},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.3,"bill_depth_mm":15.8,"flipper_length_mm":215,"body_mass_g":5050,"sex":"male","year":2007,"x":5050,"y":215},{"species":"Gentoo","island":"Biscoe","bill_length_mm":42.9,"bill_depth_mm":13.1,"flipper_length_mm":215,"body_mass_g":5000,"sex":"female","year":2007,"x":5000,"y":215},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.1,"bill_depth_mm":15.1,"flipper_length_mm":215,"body_mass_g":5100,"sex":"male","year":2007,"x":5100,"y":215},{"species":"Gentoo","island":"Biscoe","bill_length_mm":44.5,"bill_depth_mm":14.3,"flipper_length_mm":216,"body_mass_g":4100,"sex":null,"year":2007,"x":4100,"y":216},{"species":"Gentoo","island":"Biscoe","bill_length_mm":47.8,"bill_depth_mm":15,"flipper_length_mm":215,"body_mass_g":5650,"sex":"male","year":2007,"x":5650,"y":215},{"species":"Gentoo","island":"Biscoe","bill_length_mm":48.2,"bill_depth_mm":14.3,"flipper_length_mm":210,"body_mass_g":4600,"sex":"female","year":2007,"x":4600,"y":210},{"species":"Gentoo","island":"Biscoe","bill_length_mm":50,"bill_depth_mm":15.3,"flipper_length_mm":220,"body_mass_g":5550,"sex":"male","year":2007,"x":5550,"y":220},{"species":"Gentoo","island":"Biscoe","bill_length_mm":47.3,"bill_depth_mm":15.3,"flipper_length_mm":222,"body_mass_g":5250,"sex":"male","year":2007,"x":5250,"y":222},{"species":"Gentoo","island":"Biscoe","bill_length_mm":42.8,"bill_depth_mm":14.2,"flipper_length_mm":209,"body_mass_g":4700,"sex":"female","year":2007,"x":4700,"y":209},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.1,"bill_depth_mm":14.5,"flipper_length_mm":207,"body_mass_g":5050,"sex":"female","year":2007,"x":5050,"y":207},{"species":"Gentoo","island":"Biscoe","bill_length_mm":59.6,"bill_depth_mm":17,"flipper_length_mm":230,"body_mass_g":6050,"sex":"male","year":2007,"x":6050,"y":230},{"species":"Gentoo","island":"Biscoe","bill_length_mm":49.1,"bill_depth_mm":14.8,"flipper_length_mm":220,"body_mass_g":5150,"sex":"female","year":2008,"x":5150,"y":220},{"species":"Gentoo","island":"Biscoe","bill_length_mm":48.4,"bill_depth_mm":16.3,"flipper_length_mm":220,"body_mass_g":5400,"sex":"male","year":2008,"x":5400,"y":220},{"species":"Gentoo","island":"Biscoe","bill_length_mm":42.6,"bill_depth_mm":13.7,"flipper_length_mm":213,"body_mass_g":4950,"sex":"female","year":2008,"x":4950,"y":213},{"species":"Gentoo","island":"Biscoe","bill_length_mm":44.4,"bill_depth_mm":17.3,"flipper_length_mm":219,"body_mass_g":5250,"sex":"male","year":2008,"x":5250,"y":219},{"species":"Gentoo","island":"Biscoe","bill_length_mm":44,"bill_depth_mm":13.6,"flipper_length_mm":208,"body_mass_g":4350,"sex":"female","year":2008,"x":4350,"y":208},{"species":"Gentoo","island":"Biscoe","bill_length_mm":48.7,"bill_depth_mm":15.7,"flipper_length_mm":208,"body_mass_g":5350,"sex":"male","year":2008,"x":5350,"y":208},{"species":"Gentoo","island":"Biscoe","bill_length_mm":42.7,"bill_depth_mm":13.7,"flipper_length_mm":208,"body_mass_g":3950,"sex":"female","year":2008,"x":3950,"y":208},{"species":"Gentoo","island":"Biscoe","bill_length_mm":49.6,"bill_depth_mm":16,"flipper_length_mm":225,"body_mass_g":5700,"sex":"male","year":2008,"x":5700,"y":225},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.3,"bill_depth_mm":13.7,"flipper_length_mm":210,"body_mass_g":4300,"sex":"female","year":2008,"x":4300,"y":210},{"species":"Gentoo","island":"Biscoe","bill_length_mm":49.6,"bill_depth_mm":15,"flipper_length_mm":216,"body_mass_g":4750,"sex":"male","year":2008,"x":4750,"y":216},{"species":"Gentoo","island":"Biscoe","bill_length_mm":50.5,"bill_depth_mm":15.9,"flipper_length_mm":222,"body_mass_g":5550,"sex":"male","year":2008,"x":5550,"y":222},{"species":"Gentoo","island":"Biscoe","bill_length_mm":43.6,"bill_depth_mm":13.9,"flipper_length_mm":217,"body_mass_g":4900,"sex":"female","year":2008,"x":4900,"y":217},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.5,"bill_depth_mm":13.9,"flipper_length_mm":210,"body_mass_g":4200,"sex":"female","year":2008,"x":4200,"y":210},{"species":"Gentoo","island":"Biscoe","bill_length_mm":50.5,"bill_depth_mm":15.9,"flipper_length_mm":225,"body_mass_g":5400,"sex":"male","year":2008,"x":5400,"y":225},{"species":"Gentoo","island":"Biscoe","bill_length_mm":44.9,"bill_depth_mm":13.3,"flipper_length_mm":213,"body_mass_g":5100,"sex":"female","year":2008,"x":5100,"y":213},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.2,"bill_depth_mm":15.8,"flipper_length_mm":215,"body_mass_g":5300,"sex":"male","year":2008,"x":5300,"y":215},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.6,"bill_depth_mm":14.2,"flipper_length_mm":210,"body_mass_g":4850,"sex":"female","year":2008,"x":4850,"y":210},{"species":"Gentoo","island":"Biscoe","bill_length_mm":48.5,"bill_depth_mm":14.1,"flipper_length_mm":220,"body_mass_g":5300,"sex":"male","year":2008,"x":5300,"y":220},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.1,"bill_depth_mm":14.4,"flipper_length_mm":210,"body_mass_g":4400,"sex":"female","year":2008,"x":4400,"y":210},{"species":"Gentoo","island":"Biscoe","bill_length_mm":50.1,"bill_depth_mm":15,"flipper_length_mm":225,"body_mass_g":5000,"sex":"male","year":2008,"x":5000,"y":225},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.5,"bill_depth_mm":14.4,"flipper_length_mm":217,"body_mass_g":4900,"sex":"female","year":2008,"x":4900,"y":217},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45,"bill_depth_mm":15.4,"flipper_length_mm":220,"body_mass_g":5050,"sex":"male","year":2008,"x":5050,"y":220},{"species":"Gentoo","island":"Biscoe","bill_length_mm":43.8,"bill_depth_mm":13.9,"flipper_length_mm":208,"body_mass_g":4300,"sex":"female","year":2008,"x":4300,"y":208},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.5,"bill_depth_mm":15,"flipper_length_mm":220,"body_mass_g":5000,"sex":"male","year":2008,"x":5000,"y":220},{"species":"Gentoo","island":"Biscoe","bill_length_mm":43.2,"bill_depth_mm":14.5,"flipper_length_mm":208,"body_mass_g":4450,"sex":"female","year":2008,"x":4450,"y":208},{"species":"Gentoo","island":"Biscoe","bill_length_mm":50.4,"bill_depth_mm":15.3,"flipper_length_mm":224,"body_mass_g":5550,"sex":"male","year":2008,"x":5550,"y":224},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.3,"bill_depth_mm":13.8,"flipper_length_mm":208,"body_mass_g":4200,"sex":"female","year":2008,"x":4200,"y":208},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.2,"bill_depth_mm":14.9,"flipper_length_mm":221,"body_mass_g":5300,"sex":"male","year":2008,"x":5300,"y":221},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.7,"bill_depth_mm":13.9,"flipper_length_mm":214,"body_mass_g":4400,"sex":"female","year":2008,"x":4400,"y":214},{"species":"Gentoo","island":"Biscoe","bill_length_mm":54.3,"bill_depth_mm":15.7,"flipper_length_mm":231,"body_mass_g":5650,"sex":"male","year":2008,"x":5650,"y":231},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.8,"bill_depth_mm":14.2,"flipper_length_mm":219,"body_mass_g":4700,"sex":"female","year":2008,"x":4700,"y":219},{"species":"Gentoo","island":"Biscoe","bill_length_mm":49.8,"bill_depth_mm":16.8,"flipper_length_mm":230,"body_mass_g":5700,"sex":"male","year":2008,"x":5700,"y":230},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.2,"bill_depth_mm":14.4,"flipper_length_mm":214,"body_mass_g":4650,"sex":null,"year":2008,"x":4650,"y":214},{"species":"Gentoo","island":"Biscoe","bill_length_mm":49.5,"bill_depth_mm":16.2,"flipper_length_mm":229,"body_mass_g":5800,"sex":"male","year":2008,"x":5800,"y":229},{"species":"Gentoo","island":"Biscoe","bill_length_mm":43.5,"bill_depth_mm":14.2,"flipper_length_mm":220,"body_mass_g":4700,"sex":"female","year":2008,"x":4700,"y":220},{"species":"Gentoo","island":"Biscoe","bill_length_mm":50.7,"bill_depth_mm":15,"flipper_length_mm":223,"body_mass_g":5550,"sex":"male","year":2008,"x":5550,"y":223},{"species":"Gentoo","island":"Biscoe","bill_length_mm":47.7,"bill_depth_mm":15,"flipper_length_mm":216,"body_mass_g":4750,"sex":"female","year":2008,"x":4750,"y":216},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.4,"bill_depth_mm":15.6,"flipper_length_mm":221,"body_mass_g":5000,"sex":"male","year":2008,"x":5000,"y":221},{"species":"Gentoo","island":"Biscoe","bill_length_mm":48.2,"bill_depth_mm":15.6,"flipper_length_mm":221,"body_mass_g":5100,"sex":"male","year":2008,"x":5100,"y":221},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.5,"bill_depth_mm":14.8,"flipper_length_mm":217,"body_mass_g":5200,"sex":"female","year":2008,"x":5200,"y":217},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.4,"bill_depth_mm":15,"flipper_length_mm":216,"body_mass_g":4700,"sex":"female","year":2008,"x":4700,"y":216},{"species":"Gentoo","island":"Biscoe","bill_length_mm":48.6,"bill_depth_mm":16,"flipper_length_mm":230,"body_mass_g":5800,"sex":"male","year":2008,"x":5800,"y":230},{"species":"Gentoo","island":"Biscoe","bill_length_mm":47.5,"bill_depth_mm":14.2,"flipper_length_mm":209,"body_mass_g":4600,"sex":"female","year":2008,"x":4600,"y":209},{"species":"Gentoo","island":"Biscoe","bill_length_mm":51.1,"bill_depth_mm":16.3,"flipper_length_mm":220,"body_mass_g":6000,"sex":"male","year":2008,"x":6000,"y":220},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.2,"bill_depth_mm":13.8,"flipper_length_mm":215,"body_mass_g":4750,"sex":"female","year":2008,"x":4750,"y":215},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.2,"bill_depth_mm":16.4,"flipper_length_mm":223,"body_mass_g":5950,"sex":"male","year":2008,"x":5950,"y":223},{"species":"Gentoo","island":"Biscoe","bill_length_mm":49.1,"bill_depth_mm":14.5,"flipper_length_mm":212,"body_mass_g":4625,"sex":"female","year":2009,"x":4625,"y":212},{"species":"Gentoo","island":"Biscoe","bill_length_mm":52.5,"bill_depth_mm":15.6,"flipper_length_mm":221,"body_mass_g":5450,"sex":"male","year":2009,"x":5450,"y":221},{"species":"Gentoo","island":"Biscoe","bill_length_mm":47.4,"bill_depth_mm":14.6,"flipper_length_mm":212,"body_mass_g":4725,"sex":"female","year":2009,"x":4725,"y":212},{"species":"Gentoo","island":"Biscoe","bill_length_mm":50,"bill_depth_mm":15.9,"flipper_length_mm":224,"body_mass_g":5350,"sex":"male","year":2009,"x":5350,"y":224},{"species":"Gentoo","island":"Biscoe","bill_length_mm":44.9,"bill_depth_mm":13.8,"flipper_length_mm":212,"body_mass_g":4750,"sex":"female","year":2009,"x":4750,"y":212},{"species":"Gentoo","island":"Biscoe","bill_length_mm":50.8,"bill_depth_mm":17.3,"flipper_length_mm":228,"body_mass_g":5600,"sex":"male","year":2009,"x":5600,"y":228},{"species":"Gentoo","island":"Biscoe","bill_length_mm":43.4,"bill_depth_mm":14.4,"flipper_length_mm":218,"body_mass_g":4600,"sex":"female","year":2009,"x":4600,"y":218},{"species":"Gentoo","island":"Biscoe","bill_length_mm":51.3,"bill_depth_mm":14.2,"flipper_length_mm":218,"body_mass_g":5300,"sex":"male","year":2009,"x":5300,"y":218},{"species":"Gentoo","island":"Biscoe","bill_length_mm":47.5,"bill_depth_mm":14,"flipper_length_mm":212,"body_mass_g":4875,"sex":"female","year":2009,"x":4875,"y":212},{"species":"Gentoo","island":"Biscoe","bill_length_mm":52.1,"bill_depth_mm":17,"flipper_length_mm":230,"body_mass_g":5550,"sex":"male","year":2009,"x":5550,"y":230},{"species":"Gentoo","island":"Biscoe","bill_length_mm":47.5,"bill_depth_mm":15,"flipper_length_mm":218,"body_mass_g":4950,"sex":"female","year":2009,"x":4950,"y":218},{"species":"Gentoo","island":"Biscoe","bill_length_mm":52.2,"bill_depth_mm":17.1,"flipper_length_mm":228,"body_mass_g":5400,"sex":"male","year":2009,"x":5400,"y":228},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.5,"bill_depth_mm":14.5,"flipper_length_mm":212,"body_mass_g":4750,"sex":"female","year":2009,"x":4750,"y":212},{"species":"Gentoo","island":"Biscoe","bill_length_mm":49.5,"bill_depth_mm":16.1,"flipper_length_mm":224,"body_mass_g":5650,"sex":"male","year":2009,"x":5650,"y":224},{"species":"Gentoo","island":"Biscoe","bill_length_mm":44.5,"bill_depth_mm":14.7,"flipper_length_mm":214,"body_mass_g":4850,"sex":"female","year":2009,"x":4850,"y":214},{"species":"Gentoo","island":"Biscoe","bill_length_mm":50.8,"bill_depth_mm":15.7,"flipper_length_mm":226,"body_mass_g":5200,"sex":"male","year":2009,"x":5200,"y":226},{"species":"Gentoo","island":"Biscoe","bill_length_mm":49.4,"bill_depth_mm":15.8,"flipper_length_mm":216,"body_mass_g":4925,"sex":"male","year":2009,"x":4925,"y":216},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.9,"bill_depth_mm":14.6,"flipper_length_mm":222,"body_mass_g":4875,"sex":"female","year":2009,"x":4875,"y":222},{"species":"Gentoo","island":"Biscoe","bill_length_mm":48.4,"bill_depth_mm":14.4,"flipper_length_mm":203,"body_mass_g":4625,"sex":"female","year":2009,"x":4625,"y":203},{"species":"Gentoo","island":"Biscoe","bill_length_mm":51.1,"bill_depth_mm":16.5,"flipper_length_mm":225,"body_mass_g":5250,"sex":"male","year":2009,"x":5250,"y":225},{"species":"Gentoo","island":"Biscoe","bill_length_mm":48.5,"bill_depth_mm":15,"flipper_length_mm":219,"body_mass_g":4850,"sex":"female","year":2009,"x":4850,"y":219},{"species":"Gentoo","island":"Biscoe","bill_length_mm":55.9,"bill_depth_mm":17,"flipper_length_mm":228,"body_mass_g":5600,"sex":"male","year":2009,"x":5600,"y":228},{"species":"Gentoo","island":"Biscoe","bill_length_mm":47.2,"bill_depth_mm":15.5,"flipper_length_mm":215,"body_mass_g":4975,"sex":"female","year":2009,"x":4975,"y":215},{"species":"Gentoo","island":"Biscoe","bill_length_mm":49.1,"bill_depth_mm":15,"flipper_length_mm":228,"body_mass_g":5500,"sex":"male","year":2009,"x":5500,"y":228},{"species":"Gentoo","island":"Biscoe","bill_length_mm":47.3,"bill_depth_mm":13.8,"flipper_length_mm":216,"body_mass_g":4725,"sex":null,"year":2009,"x":4725,"y":216},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.8,"bill_depth_mm":16.1,"flipper_length_mm":215,"body_mass_g":5500,"sex":"male","year":2009,"x":5500,"y":215},{"species":"Gentoo","island":"Biscoe","bill_length_mm":41.7,"bill_depth_mm":14.7,"flipper_length_mm":210,"body_mass_g":4700,"sex":"female","year":2009,"x":4700,"y":210},{"species":"Gentoo","island":"Biscoe","bill_length_mm":53.4,"bill_depth_mm":15.8,"flipper_length_mm":219,"body_mass_g":5500,"sex":"male","year":2009,"x":5500,"y":219},{"species":"Gentoo","island":"Biscoe","bill_length_mm":43.3,"bill_depth_mm":14,"flipper_length_mm":208,"body_mass_g":4575,"sex":"female","year":2009,"x":4575,"y":208},{"species":"Gentoo","island":"Biscoe","bill_length_mm":48.1,"bill_depth_mm":15.1,"flipper_length_mm":209,"body_mass_g":5500,"sex":"male","year":2009,"x":5500,"y":209},{"species":"Gentoo","island":"Biscoe","bill_length_mm":50.5,"bill_depth_mm":15.2,"flipper_length_mm":216,"body_mass_g":5000,"sex":"female","year":2009,"x":5000,"y":216},{"species":"Gentoo","island":"Biscoe","bill_length_mm":49.8,"bill_depth_mm":15.9,"flipper_length_mm":229,"body_mass_g":5950,"sex":"male","year":2009,"x":5950,"y":229},{"species":"Gentoo","island":"Biscoe","bill_length_mm":43.5,"bill_depth_mm":15.2,"flipper_length_mm":213,"body_mass_g":4650,"sex":"female","year":2009,"x":4650,"y":213},{"species":"Gentoo","island":"Biscoe","bill_length_mm":51.5,"bill_depth_mm":16.3,"flipper_length_mm":230,"body_mass_g":5500,"sex":"male","year":2009,"x":5500,"y":230},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.2,"bill_depth_mm":14.1,"flipper_length_mm":217,"body_mass_g":4375,"sex":"female","year":2009,"x":4375,"y":217},{"species":"Gentoo","island":"Biscoe","bill_length_mm":55.1,"bill_depth_mm":16,"flipper_length_mm":230,"body_mass_g":5850,"sex":"male","year":2009,"x":5850,"y":230},{"species":"Gentoo","island":"Biscoe","bill_length_mm":44.5,"bill_depth_mm":15.7,"flipper_length_mm":217,"body_mass_g":4875,"sex":null,"year":2009,"x":4875,"y":217},{"species":"Gentoo","island":"Biscoe","bill_length_mm":48.8,"bill_depth_mm":16.2,"flipper_length_mm":222,"body_mass_g":6000,"sex":"male","year":2009,"x":6000,"y":222},{"species":"Gentoo","island":"Biscoe","bill_length_mm":47.2,"bill_depth_mm":13.7,"flipper_length_mm":214,"body_mass_g":4925,"sex":"female","year":2009,"x":4925,"y":214},{"species":"Gentoo","island":"Biscoe","bill_length_mm":null,"bill_depth_mm":null,"flipper_length_mm":null,"body_mass_g":null,"sex":null,"year":2009,"x":null,"y":null},{"species":"Gentoo","island":"Biscoe","bill_length_mm":46.8,"bill_depth_mm":14.3,"flipper_length_mm":215,"body_mass_g":4850,"sex":"female","year":2009,"x":4850,"y":215},{"species":"Gentoo","island":"Biscoe","bill_length_mm":50.4,"bill_depth_mm":15.7,"flipper_length_mm":222,"body_mass_g":5750,"sex":"male","year":2009,"x":5750,"y":222},{"species":"Gentoo","island":"Biscoe","bill_length_mm":45.2,"bill_depth_mm":14.8,"flipper_length_mm":212,"body_mass_g":5200,"sex":"female","year":2009,"x":5200,"y":212},{"species":"Gentoo","island":"Biscoe","bill_length_mm":49.9,"bill_depth_mm":16.1,"flipper_length_mm":213,"body_mass_g":5400,"sex":"male","year":2009,"x":5400,"y":213}],"type":"scatter"}],"xAxis":{"type":"linear","title":{"text":"body_mass_g"},"categories":null},"legend":{"align":"right","verticalAlign":"top"}},"theme":{"chart":{"backgroundColor":"transparent"},"colors":["#7cb5ec","#434348","#90ed7d","#f7a35c","#8085e9","#f15c80","#e4d354","#2b908f","#f45b5b","#91e8e1"]},"conf_opts":{"global":{"Date":null,"VMLRadialGradientURL":"http =//code.highcharts.com/list(version)/gfx/vml-radial-gradient.png","canvasToolsURL":"http =//code.highcharts.com/list(version)/modules/canvas-tools.js","getTimezoneOffset":null,"timezoneOffset":0,"useUTC":true},"lang":{"contextButtonTitle":"Chart context menu","decimalPoint":".","downloadJPEG":"Download JPEG image","downloadPDF":"Download PDF document","downloadPNG":"Download PNG image","downloadSVG":"Download SVG vector image","drillUpText":"Back to {series.name}","invalidDate":null,"loading":"Loading...","months":["January","February","March","April","May","June","July","August","September","October","November","December"],"noData":"No data to display","numericSymbols":["k","M","G","T","P","E"],"printChart":"Print chart","resetZoom":"Reset zoom","resetZoomTitle":"Reset zoom level 1:1","shortMonths":["Jan","Feb","Mar","Apr","May","Jun","Jul","Aug","Sep","Oct","Nov","Dec"],"thousandsSep":" ","weekdays":["Sunday","Monday","Tuesday","Wednesday","Thursday","Friday","Saturday"]}},"type":"chart","fonts":[],"debug":false},"evals":[],"jsHooks":[]}</script> ] --- ## Highcharter Unlike **ggplot2**, the object to be plotted doesn't have to be a data frame .pull-left[ ```r hc1=hchart(diamonds$cut, type='column') hc1 ``` ] .pull-right[ ```r hc1=hchart(diamonds$cut, type='column') hc1 ``` <div id="htmlwidget-f27116ca46fd434b4d48" style="width:100%;height:500px;" class="highchart html-widget"></div> <script type="application/json" data-for="htmlwidget-f27116ca46fd434b4d48">{"x":{"hc_opts":{"chart":{"reflow":true},"title":{"text":null},"yAxis":{"title":{"text":null}},"credits":{"enabled":false},"exporting":{"enabled":false},"boost":{"enabled":false},"plotOptions":{"series":{"label":{"enabled":false},"turboThreshold":0},"treemap":{"layoutAlgorithm":"squarified"}},"xAxis":{"type":"category"},"series":[{"data":[{"name":"Fair","y":1610},{"name":"Good","y":4906},{"name":"Very Good","y":12082},{"name":"Premium","y":13791},{"name":"Ideal","y":21551}],"type":"column"}]},"theme":{"chart":{"backgroundColor":"transparent"},"colors":["#7cb5ec","#434348","#90ed7d","#f7a35c","#8085e9","#f15c80","#e4d354","#2b908f","#f45b5b","#91e8e1"]},"conf_opts":{"global":{"Date":null,"VMLRadialGradientURL":"http =//code.highcharts.com/list(version)/gfx/vml-radial-gradient.png","canvasToolsURL":"http =//code.highcharts.com/list(version)/modules/canvas-tools.js","getTimezoneOffset":null,"timezoneOffset":0,"useUTC":true},"lang":{"contextButtonTitle":"Chart context menu","decimalPoint":".","downloadJPEG":"Download JPEG image","downloadPDF":"Download PDF document","downloadPNG":"Download PNG image","downloadSVG":"Download SVG vector image","drillUpText":"Back to {series.name}","invalidDate":null,"loading":"Loading...","months":["January","February","March","April","May","June","July","August","September","October","November","December"],"noData":"No data to display","numericSymbols":["k","M","G","T","P","E"],"printChart":"Print chart","resetZoom":"Reset zoom","resetZoomTitle":"Reset zoom level 1:1","shortMonths":["Jan","Feb","Mar","Apr","May","Jun","Jul","Aug","Sep","Oct","Nov","Dec"],"thousandsSep":" ","weekdays":["Sunday","Monday","Tuesday","Wednesday","Thursday","Friday","Saturday"]}},"type":"chart","fonts":[],"debug":false},"evals":[],"jsHooks":[]}</script> ] --- ## Highcharter We can make some more complex plots using **highcharter** .pull-left[ ```r pacman::p_load(survival) pacman::p_load(widgetframe) data(lung) lung <- dplyr::mutate(lung, sex = ifelse(sex == 1, "Male", "Female")) fit <- survfit(Surv(time, status) ~ sex, data = lung) hchart(fit, ranges=TRUE) ``` ] .pull-right[ <div id="htmlwidget-ebfc3ffd5490f1b4e6ad" style="width:100%;height:504px;" class="widgetframe html-widget"></div> <script type="application/json" data-for="htmlwidget-ebfc3ffd5490f1b4e6ad">{"x":{"url":"09-animation_files/figure-html//widgets/widget_unnamed-chunk-10.html","options":{"xdomain":"*","allowfullscreen":false,"lazyload":false}},"evals":[],"jsHooks":[]}</script> ] --- ## Highcharter .pull-left[ ```r pacman::p_load(highcharter) mtcars2 <- mtcars[1:20, ] x <- dist(mtcars2) hchart(x) ``` ] .pull-right[ <div id="htmlwidget-2887d9cc796538c18091" style="width:100%;height:504px;" class="widgetframe html-widget"></div> <script type="application/json" data-for="htmlwidget-2887d9cc796538c18091">{"x":{"url":"09-animation_files/figure-html//widgets/widget_unnamed-chunk-12.html","options":{"xdomain":"*","allowfullscreen":false,"lazyload":false}},"evals":[],"jsHooks":[]}</script> ] --- ## Highcharter ### Licensing issues ### <quote> Highcharter has a dependency on Highcharts, a commercial JavaScript charting pacman::p_load. Highcharts offers both a commercial license as well as a free non-commercial license. Please review the licensing options and terms before using this software, as the highcharter license neither provides nor implies a license for Highcharts. Highcharts (http://highcharts.com) is a Highsoft product which is not free for commercial and Governmental use. </quote> --- class: middle, inverse # rbokeh http://hafen.github.io/rbokeh/ --- ## rbokeh **rbokeh** wraps the **bokeh** plotting pacman::p_load in Python It uses a layering concept similar to **ggplot2** --- .pull-left[ ```r pacman::p_load(rbokeh) p <- figure(width=400,height=400) %>% ly_points(bill_length_mm, body_mass_g, data=penguins, color=species, glyph = species, hover=list(bill_length_mm,body_mass_g)) p ``` <div id="htmlwidget-68a8fd85caeb1e4e42f7" style="width:400px;height:400px;" class="rbokeh html-widget"></div> <script type="application/json" data-for="htmlwidget-68a8fd85caeb1e4e42f7">{"x":{"elementid":"fc4964edffd5d409b7be7c3d37d1696a","modeltype":"Plot","modelid":"1621a675938f639006197165d3c9ce03","docid":"aa8d0ef8604b1fe14104707a0c04511e","docs_json":{"aa8d0ef8604b1fe14104707a0c04511e":{"version":"0.12.5","title":"Bokeh Figure","roots":{"root_ids":["1621a675938f639006197165d3c9ce03"],"references":[{"type":"Plot","id":"1621a675938f639006197165d3c9ce03","attributes":{"id":"1621a675938f639006197165d3c9ce03","plot_width":400,"plot_height":400,"sizing_mode":"scale_both","x_range":{"type":"Range1d","id":"6d9676613b83d2a44a506e406addde02"},"y_range":{"type":"Range1d","id":"923c71391bc396b4e778c438f3bf2ba9"},"left":[{"type":"LinearAxis","id":"9a9bf02e31f070fd8c8ec7c1a129cac3"}],"below":[{"type":"LinearAxis","id":"5c653efddc2cb95adba9bf1dc6fd37c6"}],"right":[],"above":[],"renderers":[{"type":"BoxAnnotation","id":"edf8b40e4a44e517ac1e6363633c6d79"},{"type":"GlyphRenderer","id":"9ccaa5363ce5a13556d89a8174cf026d"},{"type":"GlyphRenderer","id":"3afab1fd590be5674b6cf3da93c9c7ec"},{"type":"GlyphRenderer","id":"567d44e97486a704c40726341abdca53"},{"type":"GlyphRenderer","id":"0c485508601ef5c54531fd5d9ae5b904"},{"type":"GlyphRenderer","id":"30af114dfeb8c7265ba10235a1ad2a3d"},{"type":"GlyphRenderer","id":"ca4589f86c1d81bc66400bbdf50a84fd"},{"type":"Legend","id":"03167715f40addfa0c1b2343554ad4da"},{"type":"LinearAxis","id":"5c653efddc2cb95adba9bf1dc6fd37c6"},{"type":"Grid","id":"06ecd62185e8c52a1f49fdc7fc43f60c"},{"type":"LinearAxis","id":"9a9bf02e31f070fd8c8ec7c1a129cac3"},{"type":"Grid","id":"54e35646d8cb8d5c963f0a34aac20cfa"}],"extra_y_ranges":{},"extra_x_ranges":{},"tags":[],"min_border_left":4,"min_border_right":4,"min_border_top":4,"min_border_bottom":4,"lod_threshold":null,"toolbar":{"type":"Toolbar","id":"6a1d08217a6ed1c5eb899ed930c9f2fc"},"tool_events":{"type":"ToolEvents","id":"975cb14ef0ac6fae089ec5844db0e150"}},"subtype":"Figure"},{"type":"Toolbar","id":"6a1d08217a6ed1c5eb899ed930c9f2fc","attributes":{"id":"6a1d08217a6ed1c5eb899ed930c9f2fc","tags":[],"active_drag":"auto","active_scroll":"auto","active_tap":"auto","tools":[{"type":"PanTool","id":"0a2e3f9d18a54812bb3c3e32a046b04c"},{"type":"WheelZoomTool","id":"0b5b45b0be5e90b8015d6bcded32739b"},{"type":"BoxZoomTool","id":"16290db1eac04336f6fd3f4534ee4aaf"},{"type":"ResetTool","id":"8422bf8cd74a79569ded6912fb84a14a"},{"type":"SaveTool","id":"ef4ee8d28b8ebd9f58543e9c56aae5be"},{"type":"HelpTool","id":"ffbf947bd8003e8949f480bb526a4464"},{"type":"HoverTool","id":"6f878b87d5b48d299c82a067cde0e39f"},{"type":"HoverTool","id":"90fa60c5873174759d3971c25416c852"},{"type":"HoverTool","id":"dcf97fcd5084178d7657e55f41768256"}],"logo":null}},{"type":"PanTool","id":"0a2e3f9d18a54812bb3c3e32a046b04c","attributes":{"id":"0a2e3f9d18a54812bb3c3e32a046b04c","tags":[],"plot":{"type":"Plot","id":"1621a675938f639006197165d3c9ce03","subtype":"Figure"},"dimensions":"both"}},{"type":"ToolEvents","id":"975cb14ef0ac6fae089ec5844db0e150","attributes":{"id":"975cb14ef0ac6fae089ec5844db0e150","tags":[]},"geometries":[]},{"type":"WheelZoomTool","id":"0b5b45b0be5e90b8015d6bcded32739b","attributes":{"id":"0b5b45b0be5e90b8015d6bcded32739b","tags":[],"plot":{"type":"Plot","id":"1621a675938f639006197165d3c9ce03","subtype":"Figure"},"dimensions":"both"}},{"type":"BoxAnnotation","id":"edf8b40e4a44e517ac1e6363633c6d79","attributes":{"id":"edf8b40e4a44e517ac1e6363633c6d79","tags":[],"line_color":{"units":"data","value":"black"},"line_alpha":{"units":"data","value":1},"fill_color":{"units":"data","value":"lightgrey"},"fill_alpha":{"units":"data","value":0.5},"line_dash":[4,4],"line_width":{"units":"data","value":2},"level":"overlay","top_units":"screen","bottom_units":"screen","left_units":"screen","right_units":"screen","render_mode":"css"}},{"type":"BoxZoomTool","id":"16290db1eac04336f6fd3f4534ee4aaf","attributes":{"id":"16290db1eac04336f6fd3f4534ee4aaf","tags":[],"plot":{"type":"Plot","id":"1621a675938f639006197165d3c9ce03","subtype":"Figure"},"overlay":{"type":"BoxAnnotation","id":"edf8b40e4a44e517ac1e6363633c6d79"}}},{"type":"ResetTool","id":"8422bf8cd74a79569ded6912fb84a14a","attributes":{"id":"8422bf8cd74a79569ded6912fb84a14a","tags":[],"plot":{"type":"Plot","id":"1621a675938f639006197165d3c9ce03","subtype":"Figure"}}},{"type":"SaveTool","id":"ef4ee8d28b8ebd9f58543e9c56aae5be","attributes":{"id":"ef4ee8d28b8ebd9f58543e9c56aae5be","tags":[],"plot":{"type":"Plot","id":"1621a675938f639006197165d3c9ce03","subtype":"Figure"}}},{"type":"HelpTool","id":"ffbf947bd8003e8949f480bb526a4464","attributes":{"id":"ffbf947bd8003e8949f480bb526a4464","tags":[],"plot":{"type":"Plot","id":"1621a675938f639006197165d3c9ce03","subtype":"Figure"},"redirect":"https://hafen.github.io/rbokeh","help_tooltip":"Click to learn more about rbokeh."}},{"type":"HoverTool","id":"6f878b87d5b48d299c82a067cde0e39f","attributes":{"id":"6f878b87d5b48d299c82a067cde0e39f","tags":[],"plot":{"type":"Plot","id":"1621a675938f639006197165d3c9ce03","subtype":"Figure"},"renderers":[{"type":"GlyphRenderer","id":"9ccaa5363ce5a13556d89a8174cf026d"}],"names":[],"anchor":"center","attachment":"horizontal","line_policy":"prev","mode":"mouse","point_policy":"snap_to_data","tooltips":[["bill_length_mm","@hover_col_1"],["body_mass_g","@hover_col_2"]]}},{"type":"ColumnDataSource","id":"879646d753e6386e2e009e21cff9e8a9","attributes":{"id":"879646d753e6386e2e009e21cff9e8a9","tags":[],"column_names":["x","y","line_color","fill_color","hover_col_1","hover_col_2"],"selected":[],"data":{"x":[39.1,39.5,40.3,null,36.7,39.3,38.9,39.2,34.1,42,37.8,37.8,41.1,38.6,34.6,36.6,38.7,42.5,34.4,46,37.8,37.7,35.9,38.2,38.8,35.3,40.6,40.5,37.9,40.5,39.5,37.2,39.5,40.9,36.4,39.2,38.8,42.2,37.6,39.8,36.5,40.8,36,44.1,37,39.6,41.1,37.5,36,42.3,39.6,40.1,35,42,34.5,41.4,39,40.6,36.5,37.6,35.7,41.3,37.6,41.1,36.4,41.6,35.5,41.1,35.9,41.8,33.5,39.7,39.6,45.8,35.5,42.8,40.9,37.2,36.2,42.1,34.6,42.9,36.7,35.1,37.3,41.3,36.3,36.9,38.3,38.9,35.7,41.1,34,39.6,36.2,40.8,38.1,40.3,33.1,43.2,35,41,37.7,37.8,37.9,39.7,38.6,38.2,38.1,43.2,38.1,45.6,39.7,42.2,39.6,42.7,38.6,37.3,35.7,41.1,36.2,37.7,40.2,41.4,35.2,40.6,38.8,41.5,39,44.1,38.5,43.1,36.8,37.5,38.1,41.1,35.6,40.2,37,39.7,40.2,40.6,32.1,40.7,37.3,39,39.2,36.6,36,37.8,36,41.5],"y":[3750,3800,3250,null,3450,3650,3625,4675,3475,4250,3300,3700,3200,3800,4400,3700,3450,4500,3325,4200,3400,3600,3800,3950,3800,3800,3550,3200,3150,3950,3250,3900,3300,3900,3325,4150,3950,3550,3300,4650,3150,3900,3100,4400,3000,4600,3425,2975,3450,4150,3500,4300,3450,4050,2900,3700,3550,3800,2850,3750,3150,4400,3600,4050,2850,3950,3350,4100,3050,4450,3600,3900,3550,4150,3700,4250,3700,3900,3550,4000,3200,4700,3800,4200,3350,3550,3800,3500,3950,3600,3550,4300,3400,4450,3300,4300,3700,4350,2900,4100,3725,4725,3075,4250,2925,3550,3750,3900,3175,4775,3825,4600,3200,4275,3900,4075,2900,3775,3350,3325,3150,3500,3450,3875,3050,4000,3275,4300,3050,4000,3325,3500,3500,4475,3425,3900,3175,3975,3400,4250,3400,3475,3050,3725,3000,3650,4250,3475,3450,3750,3700,4000],"line_color":["#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4"],"fill_color":["#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4","#1F77B4"],"hover_col_1":["39.1","39.5","40.3"," NA","36.7","39.3","38.9","39.2","34.1","42.0","37.8","37.8","41.1","38.6","34.6","36.6","38.7","42.5","34.4","46.0","37.8","37.7","35.9","38.2","38.8","35.3","40.6","40.5","37.9","40.5","39.5","37.2","39.5","40.9","36.4","39.2","38.8","42.2","37.6","39.8","36.5","40.8","36.0","44.1","37.0","39.6","41.1","37.5","36.0","42.3","39.6","40.1","35.0","42.0","34.5","41.4","39.0","40.6","36.5","37.6","35.7","41.3","37.6","41.1","36.4","41.6","35.5","41.1","35.9","41.8","33.5","39.7","39.6","45.8","35.5","42.8","40.9","37.2","36.2","42.1","34.6","42.9","36.7","35.1","37.3","41.3","36.3","36.9","38.3","38.9","35.7","41.1","34.0","39.6","36.2","40.8","38.1","40.3","33.1","43.2","35.0","41.0","37.7","37.8","37.9","39.7","38.6","38.2","38.1","43.2","38.1","45.6","39.7","42.2","39.6","42.7","38.6","37.3","35.7","41.1","36.2","37.7","40.2","41.4","35.2","40.6","38.8","41.5","39.0","44.1","38.5","43.1","36.8","37.5","38.1","41.1","35.6","40.2","37.0","39.7","40.2","40.6","32.1","40.7","37.3","39.0","39.2","36.6","36.0","37.8","36.0","41.5"],"hover_col_2":["3750","3800","3250"," NA","3450","3650","3625","4675","3475","4250","3300","3700","3200","3800","4400","3700","3450","4500","3325","4200","3400","3600","3800","3950","3800","3800","3550","3200","3150","3950","3250","3900","3300","3900","3325","4150","3950","3550","3300","4650","3150","3900","3100","4400","3000","4600","3425","2975","3450","4150","3500","4300","3450","4050","2900","3700","3550","3800","2850","3750","3150","4400","3600","4050","2850","3950","3350","4100","3050","4450","3600","3900","3550","4150","3700","4250","3700","3900","3550","4000","3200","4700","3800","4200","3350","3550","3800","3500","3950","3600","3550","4300","3400","4450","3300","4300","3700","4350","2900","4100","3725","4725","3075","4250","2925","3550","3750","3900","3175","4775","3825","4600","3200","4275","3900","4075","2900","3775","3350","3325","3150","3500","3450","3875","3050","4000","3275","4300","3050","4000","3325","3500","3500","4475","3425","3900","3175","3975","3400","4250","3400","3475","3050","3725","3000","3650","4250","3475","3450","3750","3700","4000"]}}},{"type":"Circle","id":"26239098e2ced5af29e8f9beca8e6eb5","attributes":{"id":"26239098e2ced5af29e8f9beca8e6eb5","tags":[],"size":{"units":"screen","value":10},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"},"line_color":{"units":"data","field":"line_color"},"fill_color":{"units":"data","field":"fill_color"}}},{"type":"Circle","id":"d5680a039a043c8aab1bcc505a10faac","attributes":{"id":"d5680a039a043c8aab1bcc505a10faac","tags":[],"size":{"units":"screen","value":10},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"},"line_color":{"units":"data","value":"#e1e1e1"},"fill_color":{"units":"data","value":"#e1e1e1"}}},{"type":"Circle","id":"56c01c5afe228aa8f373ef287dc68d2c","attributes":{"id":"56c01c5afe228aa8f373ef287dc68d2c","tags":[],"size":{"units":"screen","value":10},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":1},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"},"line_color":{"units":"data","field":"line_color"},"fill_color":{"units":"data","field":"fill_color"}}},{"type":"GlyphRenderer","id":"9ccaa5363ce5a13556d89a8174cf026d","attributes":{"id":"9ccaa5363ce5a13556d89a8174cf026d","tags":[],"selection_glyph":null,"nonselection_glyph":{"type":"Circle","id":"d5680a039a043c8aab1bcc505a10faac"},"hover_glyph":{"type":"Circle","id":"56c01c5afe228aa8f373ef287dc68d2c"},"name":null,"data_source":{"type":"ColumnDataSource","id":"879646d753e6386e2e009e21cff9e8a9"},"glyph":{"type":"Circle","id":"26239098e2ced5af29e8f9beca8e6eb5"}}},{"type":"HoverTool","id":"90fa60c5873174759d3971c25416c852","attributes":{"id":"90fa60c5873174759d3971c25416c852","tags":[],"plot":{"type":"Plot","id":"1621a675938f639006197165d3c9ce03","subtype":"Figure"},"renderers":[{"type":"GlyphRenderer","id":"3afab1fd590be5674b6cf3da93c9c7ec"}],"names":[],"anchor":"center","attachment":"horizontal","line_policy":"prev","mode":"mouse","point_policy":"snap_to_data","tooltips":[["bill_length_mm","@hover_col_1"],["body_mass_g","@hover_col_2"]]}},{"type":"ColumnDataSource","id":"cc0273f2df9e48f2ed884dadabf27f35","attributes":{"id":"cc0273f2df9e48f2ed884dadabf27f35","tags":[],"column_names":["x","y","line_color","fill_color","hover_col_1","hover_col_2"],"selected":[],"data":{"x":[46.5,50,51.3,45.4,52.7,45.2,46.1,51.3,46,51.3,46.6,51.7,47,52,45.9,50.5,50.3,58,46.4,49.2,42.4,48.5,43.2,50.6,46.7,52,50.5,49.5,46.4,52.8,40.9,54.2,42.5,51,49.7,47.5,47.6,52,46.9,53.5,49,46.2,50.9,45.5,50.9,50.8,50.1,49,51.5,49.8,48.1,51.4,45.7,50.7,42.5,52.2,45.2,49.3,50.2,45.6,51.9,46.8,45.7,55.8,43.5,49.6,50.8,50.2],"y":[3500,3900,3650,3525,3725,3950,3250,3750,4150,3700,3800,3775,3700,4050,3575,4050,3300,3700,3450,4400,3600,3400,2900,3800,3300,4150,3400,3800,3700,4550,3200,4300,3350,4100,3600,3900,3850,4800,2700,4500,3950,3650,3550,3500,3675,4450,3400,4300,3250,3675,3325,3950,3600,4050,3350,3450,3250,4050,3800,3525,3950,3650,3650,4000,3400,3775,4100,3775],"line_color":["#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E"],"fill_color":["#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E","#FF7F0E"],"hover_col_1":["46.5","50.0","51.3","45.4","52.7","45.2","46.1","51.3","46.0","51.3","46.6","51.7","47.0","52.0","45.9","50.5","50.3","58.0","46.4","49.2","42.4","48.5","43.2","50.6","46.7","52.0","50.5","49.5","46.4","52.8","40.9","54.2","42.5","51.0","49.7","47.5","47.6","52.0","46.9","53.5","49.0","46.2","50.9","45.5","50.9","50.8","50.1","49.0","51.5","49.8","48.1","51.4","45.7","50.7","42.5","52.2","45.2","49.3","50.2","45.6","51.9","46.8","45.7","55.8","43.5","49.6","50.8","50.2"],"hover_col_2":["3500","3900","3650","3525","3725","3950","3250","3750","4150","3700","3800","3775","3700","4050","3575","4050","3300","3700","3450","4400","3600","3400","2900","3800","3300","4150","3400","3800","3700","4550","3200","4300","3350","4100","3600","3900","3850","4800","2700","4500","3950","3650","3550","3500","3675","4450","3400","4300","3250","3675","3325","3950","3600","4050","3350","3450","3250","4050","3800","3525","3950","3650","3650","4000","3400","3775","4100","3775"]}}},{"type":"Square","id":"109fd49771c84c9f92650904309e8549","attributes":{"id":"109fd49771c84c9f92650904309e8549","tags":[],"size":{"units":"screen","value":10},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"},"line_color":{"units":"data","field":"line_color"},"fill_color":{"units":"data","field":"fill_color"}}},{"type":"Square","id":"4c0e0d6281719f3e203b82160ef1dd7a","attributes":{"id":"4c0e0d6281719f3e203b82160ef1dd7a","tags":[],"size":{"units":"screen","value":10},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"},"line_color":{"units":"data","value":"#e1e1e1"},"fill_color":{"units":"data","value":"#e1e1e1"}}},{"type":"Square","id":"523a75474c74793ab92c83ce887628bb","attributes":{"id":"523a75474c74793ab92c83ce887628bb","tags":[],"size":{"units":"screen","value":10},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":1},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"},"line_color":{"units":"data","field":"line_color"},"fill_color":{"units":"data","field":"fill_color"}}},{"type":"GlyphRenderer","id":"3afab1fd590be5674b6cf3da93c9c7ec","attributes":{"id":"3afab1fd590be5674b6cf3da93c9c7ec","tags":[],"selection_glyph":null,"nonselection_glyph":{"type":"Square","id":"4c0e0d6281719f3e203b82160ef1dd7a"},"hover_glyph":{"type":"Square","id":"523a75474c74793ab92c83ce887628bb"},"name":null,"data_source":{"type":"ColumnDataSource","id":"cc0273f2df9e48f2ed884dadabf27f35"},"glyph":{"type":"Square","id":"109fd49771c84c9f92650904309e8549"}}},{"type":"HoverTool","id":"dcf97fcd5084178d7657e55f41768256","attributes":{"id":"dcf97fcd5084178d7657e55f41768256","tags":[],"plot":{"type":"Plot","id":"1621a675938f639006197165d3c9ce03","subtype":"Figure"},"renderers":[{"type":"GlyphRenderer","id":"567d44e97486a704c40726341abdca53"}],"names":[],"anchor":"center","attachment":"horizontal","line_policy":"prev","mode":"mouse","point_policy":"snap_to_data","tooltips":[["bill_length_mm","@hover_col_1"],["body_mass_g","@hover_col_2"]]}},{"type":"ColumnDataSource","id":"2db403d25f10a1bff7065e662ef3b9ea","attributes":{"id":"2db403d25f10a1bff7065e662ef3b9ea","tags":[],"column_names":["x","y","line_color","fill_color","hover_col_1","hover_col_2"],"selected":[],"data":{"x":[46.1,50,48.7,50,47.6,46.5,45.4,46.7,43.3,46.8,40.9,49,45.5,48.4,45.8,49.3,42,49.2,46.2,48.7,50.2,45.1,46.5,46.3,42.9,46.1,44.5,47.8,48.2,50,47.3,42.8,45.1,59.6,49.1,48.4,42.6,44.4,44,48.7,42.7,49.6,45.3,49.6,50.5,43.6,45.5,50.5,44.9,45.2,46.6,48.5,45.1,50.1,46.5,45,43.8,45.5,43.2,50.4,45.3,46.2,45.7,54.3,45.8,49.8,46.2,49.5,43.5,50.7,47.7,46.4,48.2,46.5,46.4,48.6,47.5,51.1,45.2,45.2,49.1,52.5,47.4,50,44.9,50.8,43.4,51.3,47.5,52.1,47.5,52.2,45.5,49.5,44.5,50.8,49.4,46.9,48.4,51.1,48.5,55.9,47.2,49.1,47.3,46.8,41.7,53.4,43.3,48.1,50.5,49.8,43.5,51.5,46.2,55.1,44.5,48.8,47.2,null,46.8,50.4,45.2,49.9],"y":[4500,5700,4450,5700,5400,4550,4800,5200,4400,5150,4650,5550,4650,5850,4200,5850,4150,6300,4800,5350,5700,5000,4400,5050,5000,5100,4100,5650,4600,5550,5250,4700,5050,6050,5150,5400,4950,5250,4350,5350,3950,5700,4300,4750,5550,4900,4200,5400,5100,5300,4850,5300,4400,5000,4900,5050,4300,5000,4450,5550,4200,5300,4400,5650,4700,5700,4650,5800,4700,5550,4750,5000,5100,5200,4700,5800,4600,6000,4750,5950,4625,5450,4725,5350,4750,5600,4600,5300,4875,5550,4950,5400,4750,5650,4850,5200,4925,4875,4625,5250,4850,5600,4975,5500,4725,5500,4700,5500,4575,5500,5000,5950,4650,5500,4375,5850,4875,6000,4925,null,4850,5750,5200,5400],"line_color":["#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C"],"fill_color":["#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C","#2CA02C"],"hover_col_1":["46.1","50.0","48.7","50.0","47.6","46.5","45.4","46.7","43.3","46.8","40.9","49.0","45.5","48.4","45.8","49.3","42.0","49.2","46.2","48.7","50.2","45.1","46.5","46.3","42.9","46.1","44.5","47.8","48.2","50.0","47.3","42.8","45.1","59.6","49.1","48.4","42.6","44.4","44.0","48.7","42.7","49.6","45.3","49.6","50.5","43.6","45.5","50.5","44.9","45.2","46.6","48.5","45.1","50.1","46.5","45.0","43.8","45.5","43.2","50.4","45.3","46.2","45.7","54.3","45.8","49.8","46.2","49.5","43.5","50.7","47.7","46.4","48.2","46.5","46.4","48.6","47.5","51.1","45.2","45.2","49.1","52.5","47.4","50.0","44.9","50.8","43.4","51.3","47.5","52.1","47.5","52.2","45.5","49.5","44.5","50.8","49.4","46.9","48.4","51.1","48.5","55.9","47.2","49.1","47.3","46.8","41.7","53.4","43.3","48.1","50.5","49.8","43.5","51.5","46.2","55.1","44.5","48.8","47.2"," NA","46.8","50.4","45.2","49.9"],"hover_col_2":["4500","5700","4450","5700","5400","4550","4800","5200","4400","5150","4650","5550","4650","5850","4200","5850","4150","6300","4800","5350","5700","5000","4400","5050","5000","5100","4100","5650","4600","5550","5250","4700","5050","6050","5150","5400","4950","5250","4350","5350","3950","5700","4300","4750","5550","4900","4200","5400","5100","5300","4850","5300","4400","5000","4900","5050","4300","5000","4450","5550","4200","5300","4400","5650","4700","5700","4650","5800","4700","5550","4750","5000","5100","5200","4700","5800","4600","6000","4750","5950","4625","5450","4725","5350","4750","5600","4600","5300","4875","5550","4950","5400","4750","5650","4850","5200","4925","4875","4625","5250","4850","5600","4975","5500","4725","5500","4700","5500","4575","5500","5000","5950","4650","5500","4375","5850","4875","6000","4925"," NA","4850","5750","5200","5400"]}}},{"type":"Triangle","id":"e0c9b42e6a55ce8700df841b322cfa85","attributes":{"id":"e0c9b42e6a55ce8700df841b322cfa85","tags":[],"size":{"units":"screen","value":10},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"},"line_color":{"units":"data","field":"line_color"},"fill_color":{"units":"data","field":"fill_color"}}},{"type":"Triangle","id":"5732054e668aedac1b4e7eb2f080a7b4","attributes":{"id":"5732054e668aedac1b4e7eb2f080a7b4","tags":[],"size":{"units":"screen","value":10},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"},"line_color":{"units":"data","value":"#e1e1e1"},"fill_color":{"units":"data","value":"#e1e1e1"}}},{"type":"Triangle","id":"8e02415d845f4cf16058f217d2295a39","attributes":{"id":"8e02415d845f4cf16058f217d2295a39","tags":[],"size":{"units":"screen","value":10},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":1},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"},"line_color":{"units":"data","field":"line_color"},"fill_color":{"units":"data","field":"fill_color"}}},{"type":"GlyphRenderer","id":"567d44e97486a704c40726341abdca53","attributes":{"id":"567d44e97486a704c40726341abdca53","tags":[],"selection_glyph":null,"nonselection_glyph":{"type":"Triangle","id":"5732054e668aedac1b4e7eb2f080a7b4"},"hover_glyph":{"type":"Triangle","id":"8e02415d845f4cf16058f217d2295a39"},"name":null,"data_source":{"type":"ColumnDataSource","id":"2db403d25f10a1bff7065e662ef3b9ea"},"glyph":{"type":"Triangle","id":"e0c9b42e6a55ce8700df841b322cfa85"}}},{"type":"ColumnDataSource","id":"9b0a1983ed172b5d35d17cd0a61ce312","attributes":{"id":"9b0a1983ed172b5d35d17cd0a61ce312","tags":[],"column_names":["x","y"],"selected":[],"data":{"x":[null,null],"y":[null,null]}}},{"type":"Circle","id":"22be7a87b0cccd7b708cd4193650b462","attributes":{"id":"22be7a87b0cccd7b708cd4193650b462","tags":[],"size":{"units":"screen","value":0},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"line_color":{"units":"data","value":"#1F77B4"},"fill_color":{"units":"data","value":"#1F77B4"},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"Circle","id":"0df7f9f1f7cb5cb61622ccb69788d973","attributes":{"id":"0df7f9f1f7cb5cb61622ccb69788d973","tags":[],"size":{"units":"screen","value":0},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"line_color":{"units":"data","value":"#e1e1e1"},"fill_color":{"units":"data","value":"#e1e1e1"},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"Circle","id":"7c4a3a57a09b3640896e764296ee788f","attributes":{"id":"7c4a3a57a09b3640896e764296ee788f","tags":[],"size":{"units":"screen","value":0},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":1},"line_color":{"units":"data","value":"#1F77B4"},"fill_color":{"units":"data","value":"#1F77B4"},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"GlyphRenderer","id":"0c485508601ef5c54531fd5d9ae5b904","attributes":{"id":"0c485508601ef5c54531fd5d9ae5b904","tags":[],"selection_glyph":null,"nonselection_glyph":{"type":"Circle","id":"0df7f9f1f7cb5cb61622ccb69788d973"},"hover_glyph":{"type":"Circle","id":"7c4a3a57a09b3640896e764296ee788f"},"name":null,"data_source":{"type":"ColumnDataSource","id":"9b0a1983ed172b5d35d17cd0a61ce312"},"glyph":{"type":"Circle","id":"22be7a87b0cccd7b708cd4193650b462"}}},{"type":"Square","id":"9e06bfa514e5f49aba25d94180b249ca","attributes":{"id":"9e06bfa514e5f49aba25d94180b249ca","tags":[],"size":{"units":"screen","value":0},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"line_color":{"units":"data","value":"#FF7F0E"},"fill_color":{"units":"data","value":"#FF7F0E"},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"Square","id":"a78846046bfa8edafc31990bc353d118","attributes":{"id":"a78846046bfa8edafc31990bc353d118","tags":[],"size":{"units":"screen","value":0},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"line_color":{"units":"data","value":"#e1e1e1"},"fill_color":{"units":"data","value":"#e1e1e1"},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"Square","id":"0387d7e017b290c1a81ce8cf984f8533","attributes":{"id":"0387d7e017b290c1a81ce8cf984f8533","tags":[],"size":{"units":"screen","value":0},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":1},"line_color":{"units":"data","value":"#FF7F0E"},"fill_color":{"units":"data","value":"#FF7F0E"},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"GlyphRenderer","id":"30af114dfeb8c7265ba10235a1ad2a3d","attributes":{"id":"30af114dfeb8c7265ba10235a1ad2a3d","tags":[],"selection_glyph":null,"nonselection_glyph":{"type":"Square","id":"a78846046bfa8edafc31990bc353d118"},"hover_glyph":{"type":"Square","id":"0387d7e017b290c1a81ce8cf984f8533"},"name":null,"data_source":{"type":"ColumnDataSource","id":"9b0a1983ed172b5d35d17cd0a61ce312"},"glyph":{"type":"Square","id":"9e06bfa514e5f49aba25d94180b249ca"}}},{"type":"Triangle","id":"2364cc6df4edb048ff42eaf0010cfa11","attributes":{"id":"2364cc6df4edb048ff42eaf0010cfa11","tags":[],"size":{"units":"screen","value":0},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"line_color":{"units":"data","value":"#2CA02C"},"fill_color":{"units":"data","value":"#2CA02C"},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"Triangle","id":"b0aab7776f36e978541f9c80d949eb65","attributes":{"id":"b0aab7776f36e978541f9c80d949eb65","tags":[],"size":{"units":"screen","value":0},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"line_color":{"units":"data","value":"#e1e1e1"},"fill_color":{"units":"data","value":"#e1e1e1"},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"Triangle","id":"271298e03ead6cabbb96ccd070692341","attributes":{"id":"271298e03ead6cabbb96ccd070692341","tags":[],"size":{"units":"screen","value":0},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":1},"line_color":{"units":"data","value":"#2CA02C"},"fill_color":{"units":"data","value":"#2CA02C"},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"GlyphRenderer","id":"ca4589f86c1d81bc66400bbdf50a84fd","attributes":{"id":"ca4589f86c1d81bc66400bbdf50a84fd","tags":[],"selection_glyph":null,"nonselection_glyph":{"type":"Triangle","id":"b0aab7776f36e978541f9c80d949eb65"},"hover_glyph":{"type":"Triangle","id":"271298e03ead6cabbb96ccd070692341"},"name":null,"data_source":{"type":"ColumnDataSource","id":"9b0a1983ed172b5d35d17cd0a61ce312"},"glyph":{"type":"Triangle","id":"2364cc6df4edb048ff42eaf0010cfa11"}}},{"type":"LegendItem","id":"7a26e0cd5af05ade84fb31fa5e84d7ca","attributes":{"id":"7a26e0cd5af05ade84fb31fa5e84d7ca","tags":[],"label":{"value":"species"},"renderers":[]}},{"type":"LegendItem","id":"7ace73026bc2c54951ca5c95d6485f85","attributes":{"id":"7ace73026bc2c54951ca5c95d6485f85","tags":[],"label":{"value":" Adelie"},"renderers":[{"type":"GlyphRenderer","id":"0c485508601ef5c54531fd5d9ae5b904"}]}},{"type":"LegendItem","id":"4ff99bfa921ed31294c847efd92e0d0f","attributes":{"id":"4ff99bfa921ed31294c847efd92e0d0f","tags":[],"label":{"value":" Chinstrap"},"renderers":[{"type":"GlyphRenderer","id":"30af114dfeb8c7265ba10235a1ad2a3d"}]}},{"type":"LegendItem","id":"6117101beb162a2ac4eef3f36d812f73","attributes":{"id":"6117101beb162a2ac4eef3f36d812f73","tags":[],"label":{"value":" Gentoo"},"renderers":[{"type":"GlyphRenderer","id":"ca4589f86c1d81bc66400bbdf50a84fd"}]}},{"type":"Legend","id":"03167715f40addfa0c1b2343554ad4da","attributes":{"id":"03167715f40addfa0c1b2343554ad4da","tags":[],"plot":{"type":"Plot","id":"1621a675938f639006197165d3c9ce03","subtype":"Figure"},"items":[{"type":"LegendItem","id":"7a26e0cd5af05ade84fb31fa5e84d7ca"},{"type":"LegendItem","id":"7ace73026bc2c54951ca5c95d6485f85"},{"type":"LegendItem","id":"4ff99bfa921ed31294c847efd92e0d0f"},{"type":"LegendItem","id":"6117101beb162a2ac4eef3f36d812f73"}],"location":"top_right"}},{"type":"Range1d","id":"6d9676613b83d2a44a506e406addde02","attributes":{"id":"6d9676613b83d2a44a506e406addde02","tags":[],"start":30.175,"end":61.525}},{"type":"Range1d","id":"923c71391bc396b4e778c438f3bf2ba9","attributes":{"id":"923c71391bc396b4e778c438f3bf2ba9","tags":[],"start":2448,"end":6552}},{"type":"LinearAxis","id":"5c653efddc2cb95adba9bf1dc6fd37c6","attributes":{"id":"5c653efddc2cb95adba9bf1dc6fd37c6","tags":[],"plot":{"type":"Plot","id":"1621a675938f639006197165d3c9ce03","subtype":"Figure"},"axis_label":"bill_length_mm","formatter":{"type":"BasicTickFormatter","id":"5b76b978466021a607f2fdab6a1b9479"},"ticker":{"type":"BasicTicker","id":"d7a93166a99d9a42762ef06807d6ec5f"},"visible":true,"axis_label_text_font_size":"12pt"}},{"type":"BasicTickFormatter","id":"5b76b978466021a607f2fdab6a1b9479","attributes":{"id":"5b76b978466021a607f2fdab6a1b9479","tags":[]}},{"type":"BasicTicker","id":"d7a93166a99d9a42762ef06807d6ec5f","attributes":{"id":"d7a93166a99d9a42762ef06807d6ec5f","tags":[],"num_minor_ticks":5}},{"type":"Grid","id":"06ecd62185e8c52a1f49fdc7fc43f60c","attributes":{"id":"06ecd62185e8c52a1f49fdc7fc43f60c","tags":[],"dimension":0,"plot":{"type":"Plot","id":"1621a675938f639006197165d3c9ce03","subtype":"Figure"},"ticker":{"type":"BasicTicker","id":"d7a93166a99d9a42762ef06807d6ec5f"}}},{"type":"LinearAxis","id":"9a9bf02e31f070fd8c8ec7c1a129cac3","attributes":{"id":"9a9bf02e31f070fd8c8ec7c1a129cac3","tags":[],"plot":{"type":"Plot","id":"1621a675938f639006197165d3c9ce03","subtype":"Figure"},"axis_label":"body_mass_g","formatter":{"type":"BasicTickFormatter","id":"e035b7705c14422d50df26f8322a8f17"},"ticker":{"type":"BasicTicker","id":"d575ae06ad1853d38b99074e2e6f8fdd"},"visible":true,"axis_label_text_font_size":"12pt"}},{"type":"BasicTickFormatter","id":"e035b7705c14422d50df26f8322a8f17","attributes":{"id":"e035b7705c14422d50df26f8322a8f17","tags":[]}},{"type":"BasicTicker","id":"d575ae06ad1853d38b99074e2e6f8fdd","attributes":{"id":"d575ae06ad1853d38b99074e2e6f8fdd","tags":[],"num_minor_ticks":5}},{"type":"Grid","id":"54e35646d8cb8d5c963f0a34aac20cfa","attributes":{"id":"54e35646d8cb8d5c963f0a34aac20cfa","tags":[],"dimension":1,"plot":{"type":"Plot","id":"1621a675938f639006197165d3c9ce03","subtype":"Figure"},"ticker":{"type":"BasicTicker","id":"d575ae06ad1853d38b99074e2e6f8fdd"}}}]}}},"debug":false},"evals":[],"jsHooks":[]}</script> ] .pull-right[ ```r z <- lm(dist ~ speed, data = cars) p <- figure(width = 400, height = 400) %>% ly_points(cars, hover = cars) %>% ly_lines(lowess(cars), legend = "lowess") %>% ly_abline(z, type = 2, legend = "lm") p ``` <div id="htmlwidget-ee96b862b5e7e446cfd1" style="width:400px;height:400px;" class="rbokeh html-widget"></div> <script type="application/json" data-for="htmlwidget-ee96b862b5e7e446cfd1">{"x":{"elementid":"a9c06d31f21dbe7a1238eaafb23d317a","modeltype":"Plot","modelid":"c4f50f3bf319a2cfcf494303502a97a3","docid":"aa8d0ef8604b1fe14104707a0c04511e","docs_json":{"aa8d0ef8604b1fe14104707a0c04511e":{"version":"0.12.5","title":"Bokeh Figure","roots":{"root_ids":["c4f50f3bf319a2cfcf494303502a97a3"],"references":[{"type":"Plot","id":"c4f50f3bf319a2cfcf494303502a97a3","attributes":{"id":"c4f50f3bf319a2cfcf494303502a97a3","plot_width":400,"plot_height":400,"sizing_mode":"scale_both","x_range":{"type":"Range1d","id":"4ac0d983e79745cdfa11441d4f0d0a96"},"y_range":{"type":"Range1d","id":"1d4709903946a4c8916bf96a2380393f"},"left":[{"type":"LinearAxis","id":"770849ee389f715466cb8cfcbb419bee"}],"below":[{"type":"LinearAxis","id":"e24d0b8869860a70f7db4ffad0a6e09b"}],"right":[],"above":[],"renderers":[{"type":"BoxAnnotation","id":"b6309ce64697cd557204b8b0af3c86f7"},{"type":"GlyphRenderer","id":"a67d4650b55a4aaf41b3c9d203420cbf"},{"type":"GlyphRenderer","id":"63e50343b9bd739a6be7af0fef18bc86"},{"type":"GlyphRenderer","id":"66333ea16171b28ab8afd6f5569743c1"},{"type":"GlyphRenderer","id":"e76dfd34e85fb4cbc960c72d00e52f97"},{"type":"GlyphRenderer","id":"4b0a545eaea5ba4ac8b9f0cdb67e6dd4"},{"type":"Legend","id":"3e3eed12fe26c83629aaf792789544aa"},{"type":"LinearAxis","id":"e24d0b8869860a70f7db4ffad0a6e09b"},{"type":"Grid","id":"c4c5c4937f331b303b2f83043ec9ec57"},{"type":"LinearAxis","id":"770849ee389f715466cb8cfcbb419bee"},{"type":"Grid","id":"74b79f7a98e0d98a6b86977b073c0ad9"}],"extra_y_ranges":{},"extra_x_ranges":{},"tags":[],"min_border_left":4,"min_border_right":4,"min_border_top":4,"min_border_bottom":4,"lod_threshold":null,"toolbar":{"type":"Toolbar","id":"764e66bb9e598c4952231dddc83ccdb9"},"tool_events":{"type":"ToolEvents","id":"b6d30b68e4ffe7fa0bfd7f076a21551e"}},"subtype":"Figure"},{"type":"Toolbar","id":"764e66bb9e598c4952231dddc83ccdb9","attributes":{"id":"764e66bb9e598c4952231dddc83ccdb9","tags":[],"active_drag":"auto","active_scroll":"auto","active_tap":"auto","tools":[{"type":"PanTool","id":"f53ff27756d2e8061edef59bac95dedd"},{"type":"WheelZoomTool","id":"2b4e52e3c2baac0617c3e61d8dddf737"},{"type":"BoxZoomTool","id":"0c1358420fc1d1e1e2e7b7e22121e2e2"},{"type":"ResetTool","id":"fc72264d2010d69d5d5dcfa33b964dae"},{"type":"SaveTool","id":"47db294fd35d67e3b39fb68585f85576"},{"type":"HelpTool","id":"c2b1db9f0fd075e4cba9c7eb468f745d"},{"type":"HoverTool","id":"c1685f99cd992fdc7205b9a02771a28e"}],"logo":null}},{"type":"PanTool","id":"f53ff27756d2e8061edef59bac95dedd","attributes":{"id":"f53ff27756d2e8061edef59bac95dedd","tags":[],"plot":{"type":"Plot","id":"c4f50f3bf319a2cfcf494303502a97a3","subtype":"Figure"},"dimensions":"both"}},{"type":"ToolEvents","id":"b6d30b68e4ffe7fa0bfd7f076a21551e","attributes":{"id":"b6d30b68e4ffe7fa0bfd7f076a21551e","tags":[]},"geometries":[]},{"type":"WheelZoomTool","id":"2b4e52e3c2baac0617c3e61d8dddf737","attributes":{"id":"2b4e52e3c2baac0617c3e61d8dddf737","tags":[],"plot":{"type":"Plot","id":"c4f50f3bf319a2cfcf494303502a97a3","subtype":"Figure"},"dimensions":"both"}},{"type":"BoxAnnotation","id":"b6309ce64697cd557204b8b0af3c86f7","attributes":{"id":"b6309ce64697cd557204b8b0af3c86f7","tags":[],"line_color":{"units":"data","value":"black"},"line_alpha":{"units":"data","value":1},"fill_color":{"units":"data","value":"lightgrey"},"fill_alpha":{"units":"data","value":0.5},"line_dash":[4,4],"line_width":{"units":"data","value":2},"level":"overlay","top_units":"screen","bottom_units":"screen","left_units":"screen","right_units":"screen","render_mode":"css"}},{"type":"BoxZoomTool","id":"0c1358420fc1d1e1e2e7b7e22121e2e2","attributes":{"id":"0c1358420fc1d1e1e2e7b7e22121e2e2","tags":[],"plot":{"type":"Plot","id":"c4f50f3bf319a2cfcf494303502a97a3","subtype":"Figure"},"overlay":{"type":"BoxAnnotation","id":"b6309ce64697cd557204b8b0af3c86f7"}}},{"type":"ResetTool","id":"fc72264d2010d69d5d5dcfa33b964dae","attributes":{"id":"fc72264d2010d69d5d5dcfa33b964dae","tags":[],"plot":{"type":"Plot","id":"c4f50f3bf319a2cfcf494303502a97a3","subtype":"Figure"}}},{"type":"SaveTool","id":"47db294fd35d67e3b39fb68585f85576","attributes":{"id":"47db294fd35d67e3b39fb68585f85576","tags":[],"plot":{"type":"Plot","id":"c4f50f3bf319a2cfcf494303502a97a3","subtype":"Figure"}}},{"type":"HelpTool","id":"c2b1db9f0fd075e4cba9c7eb468f745d","attributes":{"id":"c2b1db9f0fd075e4cba9c7eb468f745d","tags":[],"plot":{"type":"Plot","id":"c4f50f3bf319a2cfcf494303502a97a3","subtype":"Figure"},"redirect":"https://hafen.github.io/rbokeh","help_tooltip":"Click to learn more about rbokeh."}},{"type":"HoverTool","id":"c1685f99cd992fdc7205b9a02771a28e","attributes":{"id":"c1685f99cd992fdc7205b9a02771a28e","tags":[],"plot":{"type":"Plot","id":"c4f50f3bf319a2cfcf494303502a97a3","subtype":"Figure"},"renderers":[{"type":"GlyphRenderer","id":"a67d4650b55a4aaf41b3c9d203420cbf"}],"names":[],"anchor":"center","attachment":"horizontal","line_policy":"prev","mode":"mouse","point_policy":"snap_to_data","tooltips":[["speed","@hover_col_1"],["dist","@hover_col_2"]]}},{"type":"ColumnDataSource","id":"0594ead4fca1a163dd7363e1b3d730c1","attributes":{"id":"0594ead4fca1a163dd7363e1b3d730c1","tags":[],"column_names":["x","y","hover_col_1","hover_col_2"],"selected":[],"data":{"x":[4,4,7,7,8,9,10,10,10,11,11,12,12,12,12,13,13,13,13,14,14,14,14,15,15,15,16,16,17,17,17,18,18,18,18,19,19,19,20,20,20,20,20,22,23,24,24,24,24,25],"y":[2,10,4,22,16,10,18,26,34,17,28,14,20,24,28,26,34,34,46,26,36,60,80,20,26,54,32,40,32,40,50,42,56,76,84,36,46,68,32,48,52,56,64,66,54,70,92,93,120,85],"hover_col_1":[" 4"," 4"," 7"," 7"," 8"," 9","10","10","10","11","11","12","12","12","12","13","13","13","13","14","14","14","14","15","15","15","16","16","17","17","17","18","18","18","18","19","19","19","20","20","20","20","20","22","23","24","24","24","24","25"],"hover_col_2":[" 2"," 10"," 4"," 22"," 16"," 10"," 18"," 26"," 34"," 17"," 28"," 14"," 20"," 24"," 28"," 26"," 34"," 34"," 46"," 26"," 36"," 60"," 80"," 20"," 26"," 54"," 32"," 40"," 32"," 40"," 50"," 42"," 56"," 76"," 84"," 36"," 46"," 68"," 32"," 48"," 52"," 56"," 64"," 66"," 54"," 70"," 92"," 93","120"," 85"]}}},{"type":"Circle","id":"ef08bccba98458360b79692646152fd2","attributes":{"id":"ef08bccba98458360b79692646152fd2","tags":[],"size":{"units":"screen","value":10},"line_color":{"units":"data","value":"#1F77B4"},"fill_color":{"units":"data","value":"#1F77B4"},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"Circle","id":"b572332bf5e33ec722caea30cbee1f0c","attributes":{"id":"b572332bf5e33ec722caea30cbee1f0c","tags":[],"size":{"units":"screen","value":10},"line_color":{"units":"data","value":"#e1e1e1"},"fill_color":{"units":"data","value":"#e1e1e1"},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"Circle","id":"53a2be13f5bf78e8c3e17b8908282fca","attributes":{"id":"53a2be13f5bf78e8c3e17b8908282fca","tags":[],"size":{"units":"screen","value":10},"line_color":{"units":"data","value":"#1F77B4"},"fill_color":{"units":"data","value":"#1F77B4"},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":1},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"GlyphRenderer","id":"a67d4650b55a4aaf41b3c9d203420cbf","attributes":{"id":"a67d4650b55a4aaf41b3c9d203420cbf","tags":[],"selection_glyph":null,"nonselection_glyph":{"type":"Circle","id":"b572332bf5e33ec722caea30cbee1f0c"},"hover_glyph":{"type":"Circle","id":"53a2be13f5bf78e8c3e17b8908282fca"},"name":null,"data_source":{"type":"ColumnDataSource","id":"0594ead4fca1a163dd7363e1b3d730c1"},"glyph":{"type":"Circle","id":"ef08bccba98458360b79692646152fd2"}}},{"type":"ColumnDataSource","id":"a236e6099d13b4ea89aaa8d124b2522d","attributes":{"id":"a236e6099d13b4ea89aaa8d124b2522d","tags":[],"column_names":["x","y"],"selected":[],"data":{"x":[4,4,7,7,8,9,10,10,10,11,11,12,12,12,12,13,13,13,13,14,14,14,14,15,15,15,16,16,17,17,17,18,18,18,18,19,19,19,20,20,20,20,20,22,23,24,24,24,24,25],"y":[4.96545927718688,4.96545927718688,13.1244950396664,13.1244950396664,15.8586333820983,18.5796905142177,21.2803125785285,21.2803125785285,21.2803125785285,24.1292771489265,24.1292771489265,27.1195485506035,27.1195485506035,27.1195485506035,27.1195485506035,30.027276331154,30.027276331154,30.027276331154,30.027276331154,32.9625061361576,32.9625061361576,32.9625061361576,32.9625061361576,36.7577283416497,36.7577283416497,36.7577283416497,40.4350745619887,40.4350745619887,43.4634917818176,43.4634917818176,43.4634917818176,46.885478946024,46.885478946024,46.885478946024,46.885478946024,50.7931517254206,50.7931517254206,50.7931517254206,56.4912240928772,56.4912240928772,56.4912240928772,56.4912240928772,56.4912240928772,67.5858242314313,73.07969526937,78.6431635543999,78.6431635543999,78.6431635543999,78.6431635543999,84.3286980968342]}}},{"type":"Line","id":"038b58293ecca9b2bf0f8b8c20a92924","attributes":{"id":"038b58293ecca9b2bf0f8b8c20a92924","tags":[],"line_color":{"units":"data","value":"black"},"line_alpha":{"units":"data","value":1},"line_width":{"units":"data","value":1},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"Line","id":"44dd49f0062c145ae4405a92146f7077","attributes":{"id":"44dd49f0062c145ae4405a92146f7077","tags":[],"line_color":{"units":"data","value":"#e1e1e1"},"line_alpha":{"units":"data","value":1},"line_width":{"units":"data","value":1},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"Line","id":"a839a690fda25c4ee3cdce25dbd1b575","attributes":{"id":"a839a690fda25c4ee3cdce25dbd1b575","tags":[],"line_color":{"units":"data","value":"black"},"line_alpha":{"units":"data","value":1},"line_width":{"units":"data","value":1},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"GlyphRenderer","id":"63e50343b9bd739a6be7af0fef18bc86","attributes":{"id":"63e50343b9bd739a6be7af0fef18bc86","tags":[],"selection_glyph":null,"nonselection_glyph":{"type":"Line","id":"44dd49f0062c145ae4405a92146f7077"},"hover_glyph":{"type":"Line","id":"a839a690fda25c4ee3cdce25dbd1b575"},"name":null,"data_source":{"type":"ColumnDataSource","id":"a236e6099d13b4ea89aaa8d124b2522d"},"glyph":{"type":"Line","id":"038b58293ecca9b2bf0f8b8c20a92924"}}},{"type":"ColumnDataSource","id":"cdd342fd763c2e52bffe451c2a4dce97","attributes":{"id":"cdd342fd763c2e52bffe451c2a4dce97","tags":[],"column_names":["x0","y0","x1","y1"],"selected":[],"data":{"x0":[0],"y0":[-17.5790948905109],"x1":[1],"y1":[-13.6466861313868]}}},{"type":"Segment","id":"f124a696c38a2497397ea07d63383164","attributes":{"id":"f124a696c38a2497397ea07d63383164","tags":[],"line_color":{"units":"data","value":"black"},"line_dash":[8,8],"line_width":{"units":"data","value":1},"x0":{"units":"data","field":"x0"},"y0":{"units":"data","field":"y0"},"x1":{"units":"data","field":"x1"},"y1":{"units":"data","field":"y1"}}},{"type":"Segment","id":"992cf047c8a0c2a1159bc914dc4ec3fd","attributes":{"id":"992cf047c8a0c2a1159bc914dc4ec3fd","tags":[],"line_color":{"units":"data","value":"#e1e1e1"},"line_dash":[8,8],"line_width":{"units":"data","value":1},"x0":{"units":"data","field":"x0"},"y0":{"units":"data","field":"y0"},"x1":{"units":"data","field":"x1"},"y1":{"units":"data","field":"y1"}}},{"type":"Segment","id":"ec9f160c7edaf44ba84d89f4ba116e20","attributes":{"id":"ec9f160c7edaf44ba84d89f4ba116e20","tags":[],"line_color":{"units":"data","value":"black"},"line_dash":[8,8],"line_width":{"units":"data","value":1},"x0":{"units":"data","field":"x0"},"y0":{"units":"data","field":"y0"},"x1":{"units":"data","field":"x1"},"y1":{"units":"data","field":"y1"}}},{"type":"GlyphRenderer","id":"66333ea16171b28ab8afd6f5569743c1","attributes":{"id":"66333ea16171b28ab8afd6f5569743c1","tags":[],"selection_glyph":null,"nonselection_glyph":{"type":"Segment","id":"992cf047c8a0c2a1159bc914dc4ec3fd"},"hover_glyph":{"type":"Segment","id":"ec9f160c7edaf44ba84d89f4ba116e20"},"name":null,"data_source":{"type":"ColumnDataSource","id":"842d5a9c215353e5248445230a0b8ebf"},"glyph":{"type":"Segment","id":"f124a696c38a2497397ea07d63383164"}}},{"type":"ColumnDataSource","id":"968e7e49d4e9186bc1159a9d8a7bc9b7","attributes":{"id":"968e7e49d4e9186bc1159a9d8a7bc9b7","tags":[],"column_names":["x","y"],"selected":[],"data":{"x":[null,null],"y":[null,null]}}},{"type":"Line","id":"b6fba13c3d30e8817c4381538487eca4","attributes":{"id":"b6fba13c3d30e8817c4381538487eca4","tags":[],"line_color":{"units":"data","value":"black"},"line_alpha":{"units":"data","value":1},"line_width":{"units":"data","value":1}}},{"type":"Line","id":"2c6e20a921b95dfe5a01663547122230","attributes":{"id":"2c6e20a921b95dfe5a01663547122230","tags":[],"line_color":{"units":"data","value":"#e1e1e1"},"line_alpha":{"units":"data","value":1},"line_width":{"units":"data","value":1}}},{"type":"Line","id":"18bfaa0f60717036be08b2c48a012b5c","attributes":{"id":"18bfaa0f60717036be08b2c48a012b5c","tags":[],"line_color":{"units":"data","value":"black"},"line_alpha":{"units":"data","value":1},"line_width":{"units":"data","value":1}}},{"type":"GlyphRenderer","id":"e76dfd34e85fb4cbc960c72d00e52f97","attributes":{"id":"e76dfd34e85fb4cbc960c72d00e52f97","tags":[],"selection_glyph":null,"nonselection_glyph":{"type":"Line","id":"2c6e20a921b95dfe5a01663547122230"},"hover_glyph":{"type":"Line","id":"18bfaa0f60717036be08b2c48a012b5c"},"name":null,"data_source":{"type":"ColumnDataSource","id":"968e7e49d4e9186bc1159a9d8a7bc9b7"},"glyph":{"type":"Line","id":"b6fba13c3d30e8817c4381538487eca4"}}},{"type":"Segment","id":"ba105f3ea6633acd6bde2104da8fcfbd","attributes":{"id":"ba105f3ea6633acd6bde2104da8fcfbd","tags":[],"line_color":{"units":"data","value":"black"},"line_dash":[8,8],"line_width":{"units":"data","value":1}}},{"type":"Segment","id":"400a9b9b2e253f4b8979806b35d12b7b","attributes":{"id":"400a9b9b2e253f4b8979806b35d12b7b","tags":[],"line_color":{"units":"data","value":"#e1e1e1"},"line_dash":[8,8],"line_width":{"units":"data","value":1}}},{"type":"Segment","id":"0caeebb003d4a983f44d2422f6070d04","attributes":{"id":"0caeebb003d4a983f44d2422f6070d04","tags":[],"line_color":{"units":"data","value":"black"},"line_dash":[8,8],"line_width":{"units":"data","value":1}}},{"type":"GlyphRenderer","id":"4b0a545eaea5ba4ac8b9f0cdb67e6dd4","attributes":{"id":"4b0a545eaea5ba4ac8b9f0cdb67e6dd4","tags":[],"selection_glyph":null,"nonselection_glyph":{"type":"Segment","id":"400a9b9b2e253f4b8979806b35d12b7b"},"hover_glyph":{"type":"Segment","id":"0caeebb003d4a983f44d2422f6070d04"},"name":null,"data_source":{"type":"ColumnDataSource","id":"968e7e49d4e9186bc1159a9d8a7bc9b7"},"glyph":{"type":"Segment","id":"ba105f3ea6633acd6bde2104da8fcfbd"}}},{"type":"LegendItem","id":"071b3807960dc3aa37517e15c03de6cc","attributes":{"id":"071b3807960dc3aa37517e15c03de6cc","tags":[],"label":{"value":"lowess"},"renderers":[{"type":"GlyphRenderer","id":"e76dfd34e85fb4cbc960c72d00e52f97"}]}},{"type":"LegendItem","id":"6745adc8e81a4442d6987219c55501be","attributes":{"id":"6745adc8e81a4442d6987219c55501be","tags":[],"label":{"value":"lm"},"renderers":[{"type":"GlyphRenderer","id":"4b0a545eaea5ba4ac8b9f0cdb67e6dd4"}]}},{"type":"Legend","id":"3e3eed12fe26c83629aaf792789544aa","attributes":{"id":"3e3eed12fe26c83629aaf792789544aa","tags":[],"plot":{"type":"Plot","id":"c4f50f3bf319a2cfcf494303502a97a3","subtype":"Figure"},"items":[{"type":"LegendItem","id":"071b3807960dc3aa37517e15c03de6cc"},{"type":"LegendItem","id":"6745adc8e81a4442d6987219c55501be"}],"location":"top_right"}},{"type":"Range1d","id":"4ac0d983e79745cdfa11441d4f0d0a96","attributes":{"id":"4ac0d983e79745cdfa11441d4f0d0a96","tags":[],"start":2.53,"end":26.47}},{"type":"Range1d","id":"1d4709903946a4c8916bf96a2380393f","attributes":{"id":"1d4709903946a4c8916bf96a2380393f","tags":[],"start":-6.26,"end":128.26}},{"type":"LinearAxis","id":"e24d0b8869860a70f7db4ffad0a6e09b","attributes":{"id":"e24d0b8869860a70f7db4ffad0a6e09b","tags":[],"plot":{"type":"Plot","id":"c4f50f3bf319a2cfcf494303502a97a3","subtype":"Figure"},"axis_label":"speed","formatter":{"type":"BasicTickFormatter","id":"e08c3ad653cf48c176e9186726e491da"},"ticker":{"type":"BasicTicker","id":"f862ebab6aa2d84a741782d45f05a38b"},"visible":true,"axis_label_text_font_size":"12pt"}},{"type":"BasicTickFormatter","id":"e08c3ad653cf48c176e9186726e491da","attributes":{"id":"e08c3ad653cf48c176e9186726e491da","tags":[]}},{"type":"BasicTicker","id":"f862ebab6aa2d84a741782d45f05a38b","attributes":{"id":"f862ebab6aa2d84a741782d45f05a38b","tags":[],"num_minor_ticks":5}},{"type":"Grid","id":"c4c5c4937f331b303b2f83043ec9ec57","attributes":{"id":"c4c5c4937f331b303b2f83043ec9ec57","tags":[],"dimension":0,"plot":{"type":"Plot","id":"c4f50f3bf319a2cfcf494303502a97a3","subtype":"Figure"},"ticker":{"type":"BasicTicker","id":"f862ebab6aa2d84a741782d45f05a38b"}}},{"type":"LinearAxis","id":"770849ee389f715466cb8cfcbb419bee","attributes":{"id":"770849ee389f715466cb8cfcbb419bee","tags":[],"plot":{"type":"Plot","id":"c4f50f3bf319a2cfcf494303502a97a3","subtype":"Figure"},"axis_label":"dist","formatter":{"type":"BasicTickFormatter","id":"8c809a30ec41446f76b603dd95fc723d"},"ticker":{"type":"BasicTicker","id":"a995a494bd492a5f7f386dff89bf8470"},"visible":true,"axis_label_text_font_size":"12pt"}},{"type":"BasicTickFormatter","id":"8c809a30ec41446f76b603dd95fc723d","attributes":{"id":"8c809a30ec41446f76b603dd95fc723d","tags":[]}},{"type":"BasicTicker","id":"a995a494bd492a5f7f386dff89bf8470","attributes":{"id":"a995a494bd492a5f7f386dff89bf8470","tags":[],"num_minor_ticks":5}},{"type":"Grid","id":"74b79f7a98e0d98a6b86977b073c0ad9","attributes":{"id":"74b79f7a98e0d98a6b86977b073c0ad9","tags":[],"dimension":1,"plot":{"type":"Plot","id":"c4f50f3bf319a2cfcf494303502a97a3","subtype":"Figure"},"ticker":{"type":"BasicTicker","id":"a995a494bd492a5f7f386dff89bf8470"}}},{"type":"ColumnDataSource","id":"842d5a9c215353e5248445230a0b8ebf","attributes":{"id":"842d5a9c215353e5248445230a0b8ebf","tags":[],"column_names":["x0","y0","x1","y1"],"selected":[],"data":{"x0":[2.53],"y0":[-7.63010072992694],"x1":[26.47],"y1":[86.5117649635037]}}}]}}},"debug":false},"evals":[],"jsHooks":[]}</script> ] --- class: middle, inverse # Maps --- ## Maps We've already seen maps with **leaflet** that allow data to be overlayed over cartographic maps Using the packages introduced here, we also have mapping capabilities. See the respective websites for details .pull-left[ **rbokeh** <div id="htmlwidget-62e928774f326605bd27" style="width:400px;height:250px;" class="rbokeh html-widget"></div> <script type="application/json" data-for="htmlwidget-62e928774f326605bd27">{"x":{"elementid":"fce09c848f7fe3f334dd824f2dd5a675","modeltype":"Plot","modelid":"f77af90710cde2b4e2ddd44c50c3e2ba","docid":"ac46c2aed6e6831b41b2f613a1bfd311","docs_json":{"ac46c2aed6e6831b41b2f613a1bfd311":{"version":"0.12.5","title":"Bokeh Figure","roots":{"root_ids":["f77af90710cde2b4e2ddd44c50c3e2ba"],"references":[{"type":"Plot","id":"f77af90710cde2b4e2ddd44c50c3e2ba","attributes":{"id":"f77af90710cde2b4e2ddd44c50c3e2ba","plot_width":400,"plot_height":250,"sizing_mode":"scale_both","x_range":{"type":"Range1d","id":"fd4371919ebaacd5d415338d5ce9c7fb"},"y_range":{"type":"Range1d","id":"19e287b0f3e3f36a6926be14219eb799"},"left":[{"type":"LinearAxis","id":"54bc5a7168a4b9e88594538b8e5c725b"}],"below":[{"type":"LinearAxis","id":"eae859109634ea0b96d0a5268118226b"}],"right":[],"above":[],"renderers":[{"type":"BoxAnnotation","id":"ff520bb2f7b76139f0e4c613c9df196f"},{"type":"GlyphRenderer","id":"d74047225abb80aa550c08706d1e8ee5"},{"type":"GlyphRenderer","id":"d0d5df621e402ab18060410f73ca9d28"},{"type":"LinearAxis","id":"eae859109634ea0b96d0a5268118226b"},{"type":"Grid","id":"810e26aab35b527dac544cc889fca40e"},{"type":"LinearAxis","id":"54bc5a7168a4b9e88594538b8e5c725b"},{"type":"Grid","id":"2c01b0f5c3bec758e9da037873c4f0b2"}],"extra_y_ranges":{},"extra_x_ranges":{},"tags":[],"min_border_left":4,"min_border_right":4,"min_border_top":4,"min_border_bottom":4,"lod_threshold":null,"toolbar":{"type":"Toolbar","id":"0e166e359102383885985a0dcf2587ea"},"tool_events":{"type":"ToolEvents","id":"8a9b48b0b5ff199e2f31b244b2368633"}},"subtype":"Figure"},{"type":"Toolbar","id":"0e166e359102383885985a0dcf2587ea","attributes":{"id":"0e166e359102383885985a0dcf2587ea","tags":[],"active_drag":"auto","active_scroll":"auto","active_tap":"auto","tools":[{"type":"PanTool","id":"6000f114c2c7a7ccfe72190c55e85593"},{"type":"WheelZoomTool","id":"01938119b7f24f54ea43ee32c968bd8a"},{"type":"BoxZoomTool","id":"e7aa0a07a060ab5b9456531b9727ac96"},{"type":"ResetTool","id":"5a43dd21fe7e5692b6726ffa57e04f06"},{"type":"SaveTool","id":"8513d2a2263b701e03caf8b784cbdfcb"},{"type":"HelpTool","id":"d2bcbd681e86b75c4863f62cf7b17f8b"},{"type":"HoverTool","id":"33b3db0d39b742e31fada3b78ac8a341"}],"logo":null}},{"type":"PanTool","id":"6000f114c2c7a7ccfe72190c55e85593","attributes":{"id":"6000f114c2c7a7ccfe72190c55e85593","tags":[],"plot":{"type":"Plot","id":"f77af90710cde2b4e2ddd44c50c3e2ba","subtype":"Figure"},"dimensions":"both"}},{"type":"ToolEvents","id":"8a9b48b0b5ff199e2f31b244b2368633","attributes":{"id":"8a9b48b0b5ff199e2f31b244b2368633","tags":[]},"geometries":[]},{"type":"WheelZoomTool","id":"01938119b7f24f54ea43ee32c968bd8a","attributes":{"id":"01938119b7f24f54ea43ee32c968bd8a","tags":[],"plot":{"type":"Plot","id":"f77af90710cde2b4e2ddd44c50c3e2ba","subtype":"Figure"},"dimensions":"both"}},{"type":"BoxAnnotation","id":"ff520bb2f7b76139f0e4c613c9df196f","attributes":{"id":"ff520bb2f7b76139f0e4c613c9df196f","tags":[],"line_color":{"units":"data","value":"black"},"line_alpha":{"units":"data","value":1},"fill_color":{"units":"data","value":"lightgrey"},"fill_alpha":{"units":"data","value":0.5},"line_dash":[4,4],"line_width":{"units":"data","value":2},"level":"overlay","top_units":"screen","bottom_units":"screen","left_units":"screen","right_units":"screen","render_mode":"css"}},{"type":"BoxZoomTool","id":"e7aa0a07a060ab5b9456531b9727ac96","attributes":{"id":"e7aa0a07a060ab5b9456531b9727ac96","tags":[],"plot":{"type":"Plot","id":"f77af90710cde2b4e2ddd44c50c3e2ba","subtype":"Figure"},"overlay":{"type":"BoxAnnotation","id":"ff520bb2f7b76139f0e4c613c9df196f"}}},{"type":"ResetTool","id":"5a43dd21fe7e5692b6726ffa57e04f06","attributes":{"id":"5a43dd21fe7e5692b6726ffa57e04f06","tags":[],"plot":{"type":"Plot","id":"f77af90710cde2b4e2ddd44c50c3e2ba","subtype":"Figure"}}},{"type":"SaveTool","id":"8513d2a2263b701e03caf8b784cbdfcb","attributes":{"id":"8513d2a2263b701e03caf8b784cbdfcb","tags":[],"plot":{"type":"Plot","id":"f77af90710cde2b4e2ddd44c50c3e2ba","subtype":"Figure"}}},{"type":"HelpTool","id":"d2bcbd681e86b75c4863f62cf7b17f8b","attributes":{"id":"d2bcbd681e86b75c4863f62cf7b17f8b","tags":[],"plot":{"type":"Plot","id":"f77af90710cde2b4e2ddd44c50c3e2ba","subtype":"Figure"},"redirect":"https://hafen.github.io/rbokeh","help_tooltip":"Click to learn more about rbokeh."}},{"type":"ColumnDataSource","id":"c28bc5d1124875495f93d9a639613f8a","attributes":{"id":"c28bc5d1124875495f93d9a639613f8a","tags":[],"column_names":["xs","ys"],"selected":[],"data":{"xs":[[-69.8991241455078,-69.8957061767578,-69.9421920776367,-70.004150390625,-70.0661163330078,-70.0508804321289,-70.0351104736328,-69.97314453125,-69.9118118286133,-69.8991241455078],[74.8913116455078,74.8402328491211,74.7673797607422,74.7389602661133,74.7266616821289,74.6689453125,74.5589904785156,74.3721694946289,74.3761672973633,74.4979553222656,74.5264587402344,74.5414047241211,74.4310607910156,74.1947250366211,74.0388717651367,74.0018539428711,73.9078140258789,73.7691421508789,73.7318344116211,73.4111328125,73.1167984008789,72.9937438964844,72.7662124633789,72.6228561401367,72.5313491821289,72.43115234375,72.3269577026367,72.2498016357422,72.15673828125,72.0956039428711,71.9207000732422,71.822265625,71.7726593017578,71.7164077758789,71.6205062866211,71.5458984375,71.4632797241211,71.3125991821289,71.23291015625,71.18505859375,71.22021484375,71.3428726196289,71.3975601196289,71.4275360107422,71.4835891723633,71.51904296875,71.5719757080078,71.58740234375,71.6005859375,71.5719757080078,71.5455093383789,71.5455093383789,71.5772476196289,71.6052703857422,71.6205062866211,71.6016616821289,71.5455093383789,71.51708984375,71.455078125,71.3581085205078,71.2941436767578,71.2257766723633,71.11328125,71.0656280517578,71.0163116455078,70.9656219482422,70.9789123535156,71.02294921875,71.095703125,71.0923843383789,71.0890579223633,71.09130859375,71.0515594482422,70.8484420776367,70.6540069580078,70.4157257080078,70.32568359375,70.2536087036133,69.9947280883789,69.8896484375,69.8680648803711,70.056640625,70.1341781616211,70.2197265625,70.2841796875,70.2611312866211,70.0902328491211,69.9201202392578,69.7037124633789,69.5677719116211,69.5015563964844,69.453125,69.4045944213867,69.4053726196289,69.3594741821289,69.2899398803711,69.2414016723633,69.2565383911133,69.279296875,69.1869125366211,69.0831069946289,68.9734344482422,68.8689498901367,68.7823257446289,68.7136688232422,68.6732406616211,68.59765625,68.5207061767578,68.4432601928711,68.31982421875,68.2139663696289,68.1610336303711,68.1301727294922,68.0171890258789,67.7398452758789,67.6267547607422,67.5782165527344,67.5975570678711,67.6470642089844,67.7334976196289,67.7378921508789,67.6615219116211,67.5963897705078,67.4528274536133,67.2873001098633,67.1159210205078,67.0277328491211,66.92431640625,66.8292922973633,66.7313461303711,66.6242218017578,66.5958023071289,66.5667953491211,66.4973602294922,66.3971710205078,66.3468704223633,66.2869110107422,66.3009719848633,66.3054656982422,66.2818374633789,66.2384796142578,66.2471694946289,66.3133773803711,66.2869110107422,66.2312545776367,66.1770553588867,65.9616241455078,65.6662139892578,65.4709930419922,65.1804656982422,65.0955047607422,64.9189453125,64.8273391723633,64.7035140991211,64.5210952758789,64.3937530517578,64.26611328125,64.1721649169922,64.1179733276367,64.0987319946289,63.9709968566895,63.5675811767578,62.4765586853027,62.3734359741211,62.0009727478027,61.5214805603027,61.2244148254395,60.8433609008789,61.1041030883789,61.3316421508789,61.5594711303711,61.7841758728027,61.8108406066895,61.8142585754395,61.7550811767578,61.6601524353027,61.3464851379395,61.1107406616211,60.8541030883789,60.8207054138184,60.7915992736816,60.8042984008789,60.7875022888184,60.7899398803711,60.8272476196289,60.8292961120605,60.71044921875,60.64453125,60.5765647888184,60.5619163513184,60.560546875,60.7180671691895,60.7668952941895,60.8592796325684,60.9169883728027,60.9069328308105,60.8064422607422,60.6545867919922,60.5738258361816,60.5108375549316,60.4859352111816,60.5270538330078,60.4857444763184,60.5702133178711,60.6426734924316,60.8894577026367,60.8039093017578,60.7625999450684,60.7361335754395,60.7262687683105,60.7394523620605,60.8023452758789,60.8453140258789,60.9147491455078,60.9511756896973,60.9578132629395,60.9908218383789,61.0404319763184,61.080078125,61.0702171325684,61.1066436767578,61.1231460571289,61.1496124267578,61.12646484375,61.1066436767578,61.0999984741211,61.1396446228027,61.1892623901367,61.19921875,61.2256851196289,61.2455062866211,61.2785186767578,61.2818374633789,61.2620086669922,61.3447303771973,61.3777313232422,61.4217796325684,61.5427703857422,61.6209983825684,61.7197265625,61.8410186767578,61.9380874633789,61.9838829040527,62.0896492004395,62.2130851745605,62.2528305053711,62.2711906433105,62.3078117370605,62.3866195678711,62.4628944396973,62.5331077575684,62.6105461120605,62.6880836486816,62.7226600646973,62.8580055236816,62.9802742004395,63.056640625,63.0841789245605,63.1193351745605,63.1697273254395,63.1507797241211,63.1299819946289,63.1085929870605,63.1299819946289,63.1789093017578,63.3016586303711,63.5169906616211,63.6965827941895,63.8624992370605,63.9380836486816,64.0096740722656,64.0423812866211,64.0513687133789,64.0921859741211,64.1843719482422,64.3580093383789,64.5110321044922,64.5658187866211,64.6025390625,64.67431640625,64.7531204223633,64.7824249267578,64.8163070678711,64.9515609741211,65.0896453857422,65.3036193847656,65.5549774169922,65.6080093383789,65.6412124633789,65.6830062866211,65.7438507080078,65.7650375366211,65.9006805419922,66.1083984375,66.3502960205078,66.4718704223633,66.5222625732422,66.8277282714844,67.06884765625,67.1955108642578,67.3197250366211,67.4416961669922,67.5172882080078,67.5464859008789,67.607421875,67.7000045776367,67.7529296875,67.7589874267578,67.7660140991211,67.83447265625,67.9580078125,68.0677719116211,68.2121124267578,68.2609405517578,68.2847671508789,68.2995147705078,68.3869171142578,68.5464859008789,68.6370162963867,68.6691360473633,68.7232360839844,68.7820358276367,68.82373046875,68.8384780883789,68.8553695678711,68.88525390625,68.9118118286133,68.96044921875,69.0500030517578,69.18017578125,69.2648468017578,69.3039093017578,69.3538055419922,69.4144515991211,69.4296875,69.3992156982422,69.4201202392578,69.4920959472656,69.6257781982422,69.8208999633789,69.9406280517578,69.9849624633789,70.0447235107422,70.1198196411133,70.1886672973633,70.25146484375,70.2549743652344,70.1994171142578,70.2146453857422,70.23876953125,70.3132781982422,70.4177780151367,70.5185546875,70.6158218383789,70.7359390258789,70.87890625,71.0521469116211,71.255859375,71.3327102661133,71.2828140258789,71.2785186767578,71.3199234008789,71.3896484375,71.48779296875,71.5519561767578,71.5822296142578,71.5803680419922,71.5461883544922,71.5050811767578,71.4796829223633,71.4547805786133,71.4329147338867,71.4718780517578,71.5308532714844,71.5974578857422,71.6656265258789,71.7337875366211,71.8020477294922,71.9419937133789,72.1535186767578,72.3587951660156,72.6574249267578,72.7570343017578,72.8955078125,73.2111282348633,73.3829116821289,73.4813461303711,73.6046829223633,73.6326141357422,73.6571273803711,73.7206115722656,73.7337875366211,73.71728515625,73.6488265991211,73.6275405883789,73.6535186767578,73.7496109008789,73.9488220214844,74.0777282714844,74.1670913696289,74.2035140991211,74.2596664428711,74.3490219116211,74.4449234008789,74.5242233276367,74.6593780517578,74.7305679321289,74.8304672241211,74.8753890991211,74.8913116455078],[23.9665031433105,23.98828125,24.0100574493408,24.0255870819092,24.0414066314697,24.0466804504395,24.029296875,24.0146484375,23.98681640625,23.9709968566895,23.98388671875,23.9734382629395,23.9623050689697,23.9588871002197,23.9964847564697,23.9913101196289,23.9447269439697,23.9093761444092,23.8865242004395,23.8824214935303,23.9680652618408,23.9629878997803,23.8974609375,23.8431644439697,23.6358413696289,23.3386726379395,23.04150390625,22.7443351745605,22.4709968566895,22.2095699310303,21.9789066314697,21.97900390625,21.9791011810303,21.9791011810303,21.9792976379395,21.9793949127197,21.9794921875,21.9795894622803,21.9796867370605,21.9797840118408,21.9797840118408,22.0402355194092,22.1506843566895,22.1939449310303,22.3050765991211,22.4594745635986,22.5459957122803,22.7219734191895,22.9558601379395,23.181640625,23.3806648254395,23.0682621002197,22.6240234375,22.32421875,21.9608383178711,21.7184562683105,21.4168949127197,21.3687496185303,21.2878913879395,21.1134757995605,20.9083003997803,20.7455081939697,20.6250991821289,20.5076179504395,20.3929691314697,20.1943359375,19.9118156433105,19.6393547058105,19.3771476745605,19.189453125,19.0764636993408,18.9552726745605,18.8259773254395,18.7180671691895,18.5881843566895,18.4866199493408,18.4603519439697,18.42822265625,18.3963871002197,18.1087875366211,17.8353519439697,17.6788082122803,17.2962894439697,16.9136714935303,16.5310535430908,16.1484375,15.7658195495605,15.3832035064697,15.0005865097046,14.6179695129395,14.4147453308105,14.2258787155151,14.0174798965454,13.9874019622803,13.9379892349243,13.9041986465454,13.7919921875,13.6943359375,13.5617189407349,13.4759769439697,13.4037113189697,13.2756834030151,13.1794919967651,13.1011714935303,12.9631843566895,12.8592777252197,12.78515625,12.6565427780151,12.5481443405151,12.3592767715454,12.3184566497803,12.21337890625,12.1143560409546,12.0139646530151,11.9025392532349,11.7430667877197,11.7800788879395,11.8189458847046,11.8199214935303,11.7969722747803,11.7694339752197,11.7508792877197,11.8497076034546,11.89990234375,11.9678707122803,12.01611328125,12.0732421875,12.2804679870605,12.37890625,12.5037107467651,12.5504875183105,12.8976564407349,12.9832029342651,13.1626949310303,13.4169921875,13.5979499816895,13.685546875,13.7853507995605,13.7842769622803,13.8474617004395,13.8335933685303,13.7389650344849,13.7213869094849,13.6334962844849,13.5394535064697,13.4954109191895,13.3322257995605,13.2874994277954,13.2093753814697,13.1968746185303,13.1556644439697,13.0759763717651,12.99853515625,12.99853515625,13.0467767715454,13.0927734375,13.0772457122803,13.0465812683105,13.0538091659546,13.3589839935303,13.3783206939697,13.3680658340454,13.3664064407349,13.3785161972046,13.0908212661743,12.8623046875,12.8234376907349,12.5212888717651,12.4021482467651,12.3342771530151,12.2833003997803,12.3025388717651,12.38037109375,12.5535154342651,12.7906246185303,13.0097665786743,13.0681638717651,13.1843748092651,13.3026371002197,13.3464841842651,13.3714847564697,13.6490230560303,13.7645502090454,13.978515625,14.11376953125,14.1908206939697,14.3986330032349,14.6579103469849,14.7494144439697,15.0893545150757,15.4250001907349,15.7269525527954,16.0601558685303,16.3152351379395,16.4314460754395,16.537109375,16.5851554870605,16.6080074310303,16.6395511627197,16.697265625,16.7177734375,16.7009773254395,16.7093753814697,16.7429695129395,16.8130855560303,16.91943359375,16.9658203125,16.9520511627197,16.9847660064697,17.0637702941895,17.12158203125,17.1550788879395,17.2450180053711,17.4113292694092,17.5360336303711,17.57958984375,17.6433582305908,17.77880859375,17.9130859375,18.0087890625,18.0471687316895,18.1915035247803,18.3348617553711,18.4846668243408,18.5626945495605,18.6534175872803,18.8983402252197,18.9444332122803,19.1426753997803,19.3408203125,19.3699207305908,19.3716793060303,19.4193363189697,19.4798812866211,19.4874019622803,19.4837894439697,19.5276374816895,19.6603507995605,19.8751945495605,19.9974613189697,20.1900386810303,20.4822254180908,20.5900382995605,20.5987300872803,20.5369129180908,20.5358390808105,20.5583972930908,20.6078128814697,20.9109382629395,21.1903324127197,21.5108413696289,21.7510738372803,21.7816410064697,21.8060550689697,21.8416004180908,21.8335952758789,21.7800788879395,21.8008804321289,21.8958969116211,21.9053707122803,21.8718738555908,21.8294906616211,21.8131847381592,21.8566417694092,21.9486331939697,22.0891590118408,22.19775390625,22.2745113372803,22.3024406433105,22.2816410064697,22.283203125,22.3070316314697,22.2804679870605,22.2035160064697,22.1779308319092,22.2166996002197,22.2261714935303,22.2566413879395,22.27880859375,22.31494140625,22.3929672241211,22.4861316680908,22.56103515625,22.66650390625,22.8147468566895,23.0762691497803,23.15673828125,23.4001960754395,23.4639644622803,23.5599613189697,23.6963863372803,23.8338871002197,23.9011707305908,23.9073238372803,23.9287109375,23.9665031433105],[12.2136716842651,12.1990232467651,12.1554679870605,12.1800785064697,12.2065439224243,12.1771478652954,12.1105470657349,12.0399417877197,12.0183591842651,12.0775394439697,12.1670894622803,12.2042961120605,12.3079099655151,12.3466796875,12.3740234375,12.3845710754395,12.5014638900757,12.6416988372803,12.7194337844849,12.7982416152954,12.84814453125,12.8810548782349,12.9713869094849,13.0480470657349,13.07275390625,13.0573234558105,12.9474611282349,12.8296871185303,12.6748046875,12.59619140625,12.5735359191895,12.5027351379395,12.4514646530151,12.4532222747803,12.4874029159546,12.5223627090454,12.5189456939697,12.5037107467651,12.4845705032349,12.3860349655151,12.2552738189697,12.2136716842651],[-63.001220703125,-63.1600074768066,-63.1533241271973,-63.0260276794434,-62.9795913696289,-63.001220703125],[20.06396484375,20.103515625,20.1857414245605,20.2405261993408,20.3482418060303,20.4083003997803,20.4854507446289,20.5228519439697,20.5753898620605,20.5814456939697,20.5662117004395,20.5531253814697,20.5051746368408,20.5166015625,20.5162105560303,20.4755859375,20.4486331939697,20.4923820495605,20.4870128631592,20.4889659881592,20.56787109375,20.6144542694092,20.6560535430908,20.7092761993408,20.7408199310303,20.8702144622803,20.9334964752197,20.9585933685303,20.9642581939697,20.9557609558105,20.9878902435303,21.0310535430908,21.0308589935303,21.001953125,20.9501953125,20.8816394805908,20.8060531616211,20.77001953125,20.7516593933105,20.7178707122803,20.6969718933105,20.6649417877197,20.6574230194092,20.6062507629395,20.5270500183105,20.4560546875,20.4080066680908,20.3836917877197,20.3384761810303,20.3111324310303,20.3113288879395,20.34423828125,20.3816413879395,20.3824214935303,20.3640613555908,20.30615234375,20.2938480377197,20.28759765625,20.2720699310303,20.2482433319092,20.2068367004395,20.1310558319092,20.0597648620605,20.0225582122803,20.0012702941895,19.99560546875,19.96484375,19.8518562316895,19.4845714569092,19.3981437683105,19.3601551055908,19.322265625,19.3585929870605,19.39453125,19.4405269622803,19.4591808319092,19.4392585754395,19.3446292877197,19.3374996185303,19.3838882446289,19.4612312316895,19.4560546875,19.4800777435303,19.4534168243408,19.4406242370605,19.4973621368408,19.5458011627197,19.57568359375,19.5775394439697,19.46826171875,19.3423843383789,19.3455085754395,19.3611316680908,19.3521499633789,19.3614253997803,19.3308601379395,19.2806644439697,19.3290042877197,19.3996086120605,19.4651374816895,19.5445308685303,19.5974597930908,19.6544933319092,19.7034187316895,19.7278308868408,19.74072265625,19.7377910614014,19.7544918060303,19.7882804870605,19.8597660064697,19.9390621185303,20.0457038879395,20.06396484375],[20.611328125,20.6034183502197,20.5217761993408,20.4875011444092,20.4112319946289,20.39794921875,20.4295902252197,20.4901371002197,20.5691394805908,20.611328125],[19.6623039245605,19.66748046875,19.6291980743408,19.5998058319092,19.5798835754395,19.5365238189697,19.51904296875,19.5513668060303,19.6288089752197,19.6623039245605],[19.9895515441895,20.0202159881592,20.0338878631592,20.08740234375,20.1678695678711,20.18408203125,20.2395515441895,20.2588882446289,20.1947269439697,20.1550769805908,20.12548828125,20.0732421875,20.0425796508789,20.0323238372803,20.0339851379395,19.7998046875,19.7459964752197,19.6722660064697,19.6869144439697,19.7365226745605,19.7790031433105,19.7852535247803,19.84765625,19.8671855926514,19.87158203125,19.8546867370605,19.8123035430908,19.7877922058105,19.8230457305908,19.8882808685303,19.9445323944092,19.9895515441895],[1.70605456829071,1.67851531505585,1.58642566204071,1.53408193588257,1.48623073101044,1.44882810115814,1.42812478542328,1.43027341365814,1.42197239398956,1.41484367847443,1.42832016944885,1.45888686180115,1.50136709213257,1.56816422939301,1.70986306667328,1.73945295810699,1.74023449420929,1.71396493911743,1.70605456829071],[53.9278335571289,53.9281234741211,53.8263664245605,53.7991218566895,53.7158203125,53.6344718933105,53.6896476745605,53.8337898254395,53.8937492370605,53.9278335571289],[52.6168937683105,52.6000022888184,52.5822257995605,52.5835952758789,52.62939453125,52.6576156616211,52.6168937683105],[53.3322257995605,53.25830078125,53.19091796875,53.33251953125,53.3708953857422,53.4124031066895,53.4453125,53.4089851379395,53.3826179504395,53.3322257995605],[54.4654312133789,54.4566421508789,54.4284172058105,54.3577156066895,54.3347663879395,54.37890625,54.3983421325684,54.4265632629395,54.4654312133789],[56.2978515625,56.3634757995605,56.3728485107422,56.3879890441895,56.3529281616211,56.3135757446289,56.2678718566895,56.2046890258789,56.1544914245605,56.1065444946289,56.0638656616211,56.0083999633789,55.9703140258789,55.9796867370605,56.00634765625,56.0167007446289,56.0005836486816,55.9630851745605,55.9158210754395,55.8707008361816,55.8228530883789,55.7957038879395,55.7915992736816,55.80419921875,55.8039054870605,55.7775382995605,55.7681617736816,55.7868156433105,55.8040046691895,55.8056640625,55.7608413696289,55.7997093200684,55.9286155700684,55.9663124084473,55.9921875,55.9851531982422,55.8941383361816,55.7791023254395,55.6965827941895,55.5478515625,55.4684562683105,55.4917984008789,55.5193367004395,55.5316390991211,55.5084953308105,55.46630859375,55.4138679504395,55.3532218933105,55.2702140808105,55.1999015808105,55.1921882629395,55.1940422058105,55.1858367919922,55.1194343566895,55.1042976379395,55.0250015258789,54.9982452392578,54.9224586486816,54.8048820495605,54.6522483825684,54.4716796875,54.2701187133789,54.0545883178711,53.8321304321289,53.6095695495605,53.39404296875,53.1923828125,53.0119132995605,52.8592796325684,52.7416000366211,52.6659202575684,52.63916015625,52.5550765991211,52.5095672607422,52.45458984375,52.3996086120605,52.3445320129395,52.28955078125,52.2345733642578,52.1794929504395,52.12451171875,52.0694351196289,52.0144500732422,51.95947265625,51.9043922424316,51.8494148254395,51.7943344116211,51.7393569946289,51.6843757629395,51.6292991638184,51.5925788879395,51.5721664428711,51.568359375,51.568359375,51.60546875,51.6231422424316,51.66455078125,51.7347640991211,51.767578125,51.7916984558105,51.8431625366211,51.9060516357422,52.1185531616211,52.2508773803711,52.5114250183105,52.6482429504395,53.0263671875,53.32958984375,53.8017578125,53.8933601379395,54.14794921875,54.3042984008789,54.3970718383789,54.4584007263184,54.4988288879395,54.53466796875,54.5804672241211,54.6241188049316,54.6589851379395,54.7467765808105,55.09814453125,55.3035163879395,55.3216819763184,55.4333992004395,55.5228500366211,55.9412078857422,56.0251922607422,56.0746116638184,56.0804710388184,56.1165046691895,56.16748046875,56.1725578308105,56.1541023254395,56.1519508361816,56.1446304321289,56.18359375,56.24951171875,56.2785148620605,56.2978515625],[56.2105484008789,56.240234375,56.2818336486816,56.2877922058105,56.27734375,56.2342758178711,56.2165031433105,56.2105484008789],[-64.5491638183594,-64.4388198852539,-64.2205047607422,-64.1053237915039,-64.054931640625,-64.0324249267578,-63.8819313049316,-63.8154258728027,-63.8325729370117,-63.9712371826172,-64.0283203125,-64.3229064941406,-64.4532699584961,-64.5086898803711,-64.6373519897461,-64.7314453125,-64.75732421875,-64.689208984375,-64.6250991821289,-64.5813446044922,-64.5491638183594],[-68.6532211303711,-68.6475143432617,-68.6397933959961,-68.6382293701172,-68.6366729736328,-68.6350631713867,-68.6334533691406,-68.6316909790039,-68.6299285888672,-68.5711898803711,-68.3387680053711,-68.2782287597656,-68.2401428222656,-68.3330078125,-68.43115234375,-68.4794921875,-68.5207977294922,-68.5205078125,-68.488525390625,-68.3931121826172,-68.1611328125,-68.1440963745117,-68.0084915161133,-67.9402847290039,-67.861083984375,-67.6781234741211,-67.5025863647461,-67.2942428588867,-67.0694808959961,-66.8651351928711,-66.6700744628906,-66.4620132446289,-66.2356414794922,-65.9925765991211,-65.7470703125,-65.3692855834961,-65.251953125,-65.1790084838867,-65.2523422241211,-65.3460006713867,-65.4711380004883,-65.6033172607422,-65.7227478027344,-65.8419952392578,-65.9537582397461,-66.0606460571289,-66.1720199584961,-66.2867660522461,-66.398681640625,-66.5111312866211,-66.627685546875,-66.9304733276367,-67.1270980834961,-67.7932586669922,-68.0071258544922,-68.2201156616211,-68.3316955566406,-68.4910125732422,-68.61865234375,-68.6532211303711],[-61.8757820129395,-61.865966796875,-61.9180145263672,-62.0415992736816,-62.0832977294922,-62.093017578125,-61.9666519165039,-61.9071273803711,-61.8757820129395],[-62.6509742736816,-62.6256828308105,-62.6259765625,-62.5415496826172,-62.3725090026855,-62.2141571044922,-62.0665969848633,-61.9280281066895,-61.7985343933105,-61.6794891357422,-61.5709915161133,-61.5130386352539,-61.5055198669434,-61.4039535522461,-61.2083969116211,-61.084716796875,-61.0329093933105,-60.8398399353027,-60.5053749084473,-60.26220703125,-60.1103019714355,-59.8924789428711,-59.6085891723633,-59.4353981018066,-59.3729476928711,-59.187255859375,-58.7240257263184,-58.5196266174316,-58.4228019714355,-58.3653793334961,-58.3086891174316,-58.2527847290039,-58.136474609375,-57.9598159790039,-57.8216781616211,-57.6438941955566,-57.5871543884277,-57.5631332397461,-57.5716781616211,-57.6258277893066,-57.7254867553711,-57.7547836303711,-57.7570762634277,-57.782470703125,-57.865234375,-57.88623046875,-57.8906288146973,-57.9431190490723,-58.0824203491211,-58.1111335754395,-58.1180648803711,-58.1356468200684,-58.1546897888184,-58.1814918518066,-58.2030258178711,-58.2051773071289,-58.1879386901855,-58.1913070678711,-58.2220687866211,-58.2393569946289,-58.2455596923828,-58.2716751098633,-58.3176765441895,-58.3346633911133,-58.3225631713867,-58.3564453125,-58.4363250732422,-58.4852561950684,-58.5032196044922,-58.5477066040039,-58.6186027526855,-58.6417503356934,-58.6048316955566,-58.1682624816895,-57.8122100830078,-57.3912582397461,-57.11181640625,-56.9739761352539,-56.8717269897461,-56.80517578125,-56.7157211303711,-56.6033706665039,-56.5105476379395,-56.4371566772461,-56.3705062866211,-56.310546875,-56.24169921875,-56.1640625,-56.0673332214355,-55.9514656066895,-55.8590316772461,-55.7899894714355,-55.7146492004395,-55.6329078674316,-55.5937995910645,-55.5972633361816,-55.5648918151855,-55.4967269897461,-55.4506340026855,-55.4266624450684,-55.3458023071289,-55.2080078125,-55.1359367370605,-55.129638671875,-55.0888671875,-55.0136222839355,-54.962158203125,-54.9344749450684,-54.888916015625,-54.8254890441895,-54.7550773620605,-54.677734375,-54.6319351196289,-54.6158676147461,-54.537841796875,-54.5015144348145,-54.4439468383789,-54.3833503723145,-54.3318824768066,-54.2500991821289,-54.2061538696289,-54.1545906066895,-54.1192398071289,-54.0850105285645,-54.0123062133789,-53.9547843933105,-53.8911628723145,-53.8642120361328,-53.8232421875,-53.7469215393066,-53.6712875366211,-53.6685523986816,-53.7109375,-53.7181625366211,-53.7445793151855,-53.7533187866211,-53.7271499633789,-53.71728515625,-53.7584953308105,-53.8381843566895,-53.9156265258789,-53.9353485107422,-54.0401344299316,-54.1138191223145,-54.1564445495605,-54.2052230834961,-54.2601547241211,-54.3270034790039,-54.4481430053711,-54.4843254089355,-54.554931640625,-54.6154289245605,-54.6658706665039,-54.7197265625,-54.777099609375,-54.8291015625,-54.875732421875,-54.9027862548828,-54.9102058410645,-54.9559059143066,-55.0399436950684,-55.0689926147461,-55.0638694763184,-55.1015129089355,-55.2437515258789,-55.3464851379395,-55.4098129272461,-55.4766616821289,-55.5823707580566,-55.7254867553711,-55.7459945678711,-55.6915054321289,-55.6719741821289,-55.687255859375,-55.7319831848145,-55.8060531616211,-55.85888671875,-55.8905258178711,-55.9054183959961,-55.9036598205566,-55.93017578125,-55.9849128723145,-56.0196304321289,-56.0342292785645,-56.1028785705566,-56.2255363464355,-56.3223648071289,-56.3932609558105,-56.4759750366211,-56.5707054138184,-56.6358413696289,-56.6715354919434,-56.7724609375,-56.9386215209961,-57.08935546875,-57.2246551513672,-57.3006820678711,-57.3174781799316,-57.4052276611328,-57.5638656616211,-57.6088905334473,-57.645751953125,-57.6508827209473,-57.7126960754395,-57.8311996459961,-57.8725051879883,-57.8185539245605,-57.8105964660645,-57.8340797424316,-57.8863296508789,-57.8982925415039,-57.8700675964355,-57.868408203125,-57.8933601379395,-57.9483413696289,-58.0333976745605,-58.0538558959961,-58.0096702575684,-57.9879837036133,-57.9888648986816,-58.0069808959961,-58.0423316955566,-58.0958480834961,-58.1674842834473,-58.1890106201172,-58.160400390625,-58.1563491821289,-58.177001953125,-58.1647987365723,-58.1197242736816,-58.123046875,-58.2011680603027,-58.219970703125,-58.1709938049316,-58.2007789611816,-58.2503929138184,-58.3088874816895,-58.3759727478027,-58.4244613647461,-58.454833984375,-58.5472145080566,-58.5305671691895,-58.4565925598145,-58.4294967651367,-58.4090309143066,-58.3924827575684,-58.4354972839355,-58.4752426147461,-58.5254859924316,-58.4662094116211,-58.4189414978027,-58.2833480834961,-57.7635726928711,-57.5478553771973,-57.3036613464355,-57.170654296875,-57.1588859558105,-57.3539047241211,-57.37548828125,-57.33544921875,-57.2649879455566,-57.076171875,-56.9371566772461,-56.7494621276855,-56.7173805236816,-56.6980972290039,-56.6682624816895,-56.6720237731934,-56.7271499633789,-57.0876960754395,-57.395751953125,-57.5072746276855,-57.5469741821289,-57.6456069946289,-58.1791954040527,-59.0072250366211,-59.67626953125,-59.8283195495605,-60.9039573669434,-61.1122093200684,-61.3828620910645,-61.6025428771973,-61.8479042053223,-62.0668907165527,-62.1892547607422,-62.3347663879395,-62.3744621276855,-62.3036117553711,-62.3380851745605,-62.2950668334961,-62.2090835571289,-62.12646484375,-62.0536613464355,-62.1793441772461,-62.1305618286133,-62.0768089294434,-62.082763671875,-62.1315383911133,-62.2539558410645,-62.2869148254395,-62.323974609375,-62.4018592834473,-62.427001953125,-62.3936004638672,-62.2463340759277,-62.3018569946289,-62.39501953125,-62.7979965209961,-62.9590301513672,-63.2128410339355,-63.6217765808105,-63.7729988098145,-64.1231918334961,-64.3834457397461,-64.6214828491211,-64.8529815673828,-64.8198776245117,-64.8043899536133,-64.8694839477539,-64.9169006347656,-65.0694351196289,-65.1333999633789,-65.15185546875,-65.1549758911133,-65.1278839111328,-65.0182647705078,-65.0070266723633,-65.05908203125,-64.9863739013672,-64.8980484008789,-64.6995162963867,-64.6224594116211,-64.5377502441406,-64.51171875,-64.5242156982422,-64.5741195678711,-64.5709991455078,-64.42041015625,-64.2646026611328,-64.1008834838867,-64.0622100830078,-64.0611877441406,-64.2529296875,-64.228515625,-64.083251953125,-63.8928680419922,-63.7955589294434,-63.7294883728027,-63.6847648620605,-63.6298866271973,-63.5958976745605,-63.5944366455078,-63.6173324584961,-63.6444816589355,-63.6924781799316,-64.0347671508789,-64.1306610107422,-64.2199249267578,-64.2479476928711,-64.3242721557617,-64.4878387451172,-64.6504898071289,-64.8119659423828,-64.970703125,-65.0269012451172,-64.6291961669922,-64.4415512084961,-64.38037109375,-64.3191375732422,-64.3756790161133,-64.4322280883789,-64.7152328491211,-64.8399429321289,-64.9855499267578,-65.1897430419922,-65.2523422241211,-65.2835998535156,-65.3046875,-65.2385711669922,-65.3083953857422,-65.2655258178711,-65.2898483276367,-65.3612747192383,-65.6476058959961,-65.6983413696289,-65.59912109375,-65.605712890625,-65.6387634277344,-65.7577133178711,-66.1901321411133,-66.3477554321289,-66.4936065673828,-66.533447265625,-66.5850601196289,-66.8824691772461,-66.94140625,-67.2576141357422,-67.3930206298828,-67.556640625,-67.5995559692383,-67.60888671875,-67.5860824584961,-67.5633773803711,-67.5064468383789,-67.3866195678711,-66.77685546875,-66.650390625,-65.99853515625,-65.8536682128906,-65.7690963745117,-65.7380828857422,-65.775390625,-65.8143081665039,-65.8863296508789,-66.0406265258789,-66.2252426147461,-66.17236328125,-66.0973663330078,-65.9342269897461,-65.8636703491211,-65.81005859375,-65.9121551513672,-65.9434051513672,-66.0171890258789,-66.3933563232422,-66.5962905883789,-66.7828063964844,-67.0331115722656,-67.1309585571289,-67.2633285522461,-67.46630859375,-67.6848678588867,-67.6937026977539,-67.6619644165039,-67.7834930419922,-67.8259811401367,-67.9139633178711,-68.1456527709961,-68.2572326660156,-68.4046401977539,-68.4878921508789,-68.5692901611328,-68.6675796508789,-68.6726531982422,-68.6384735107422,-68.66162109375,-68.9129867553711,-68.9795913696289,-68.752685546875,-68.5979461669922,-68.5325698852539,-68.4737319946289,-68.421875,-68.4654235839844,-68.58935546875,-68.7498550415039,-68.939453125,-69.0447769165039,-69.0901870727539,-69.1414031982422,-69.1549835205078,-69.2351531982422,-69.3585968017578,-69.3517608642578,-69.2679672241211,-69.2010269165039,-69.1435089111328,-69.0657272338867,-69.0295867919922,-69.0353012084961,-69.0583038330078,-69.2180633544922,-69.3605422973633,-69.4654312133789,-69.4090805053711,-69.3130416870117,-69.1801223754883,-69.0325164794922,-68.96533203125,-68.9168014526367,-68.6908264160156,-68.4935073852539,-68.3937530517578,-68.443359375,-68.4609909057617,-68.5897979736328,-68.7151870727539,-68.924560546875,-69.2061996459961,-69.4884262084961,-69.7126007080078,-69.9602584838867,-70.4828567504883,-70.9431686401367,-71.4147415161133,-71.7166061401367,-71.9186553955078,-71.9710998535156,-71.9534683227539,-72.0284118652344,-72.136962890625,-72.2689971923828,-72.3345260620117,-72.4076690673828,-72.3664093017578,-72.30322265625,-72.3018569946289,-72.3591842651367,-72.3768081665039,-72.3591842651367,-72.307373046875,-72.2763137817383,-72.3006362915039,-72.3402328491211,-72.392578125,-72.4601593017578,-72.5098114013672,-72.6204071044922,-72.8036117553711,-72.8659210205078,-72.95556640625,-73.0823745727539,-73.1529312133789,-73.1745147705078,-73.2216262817383,-73.2516174316406,-73.274169921875,-73.3117218017578,-73.3866195678711,-73.5012741088867,-73.5077133178711,-73.5289077758789,-73.4704132080078,-73.5045394897461,-73.5762634277344,-73.55419921875,-73.483642578125,-73.4615707397461,-73.13525390625,-73.1488723754883,-73.0945816040039,-73.0336456298828,-72.9817428588867,-72.8654251098633,-72.7284698486328,-72.6512680053711,-72.6144027709961,-72.5917510986328,-72.5859375,-72.6083984375,-72.5828628540039,-72.4981384277344,-72.354736328125,-72.2930145263672,-72.3283233642578,-72.4079055786133,-72.5090866088867,-72.5179214477539,-72.4722137451172,-72.41259765625,-72.3415069580078,-72.345947265625,-72.2829055786133,-72.1034164428711,-72.0417022705078,-71.978515625,-71.9049835205078,-71.9005355834961,-71.9542465209961,-71.9629898071289,-71.9566421508789,-71.9402313232422,-71.8564453125,-71.7327194213867,-71.6996612548828,-71.6952133178711,-71.7312927246094,-71.7621078491211,-71.7776412963867,-71.8341293334961,-71.8756866455078,-71.8092727661133,-71.6844711303711,-71.6315383911133,-71.6800765991211,-71.7506332397461,-71.7726593017578,-71.7461929321289,-71.6933135986328,-71.5081024169922,-71.4904251098633,-71.3493194580078,-71.353759765625,-71.4434585571289,-71.5313034057617,-71.5962905883789,-71.8123550415039,-72.0417022705078,-72.072509765625,-72.063720703125,-71.95703125,-71.7828140258789,-71.6516647338867,-71.5603942871094,-71.4551773071289,-71.358154296875,-71.2611312866211,-71.2214813232422,-71.1597213745117,-71.15087890625,-71.2126007080078,-71.3257293701172,-71.8200225830078,-71.8350601196289,-71.8307647705078,-71.8121109008789,-71.8123550415039,-71.7671890258789,-71.7161636352539,-71.6800765991211,-71.7159652709961,-71.7947311401367,-71.7374038696289,-71.7327651977539,-71.7506332397461,-71.8324203491211,-71.9049835205078,-71.9049835205078,-71.8202209472656,-71.7638702392578,-71.7506332397461,-71.781494140625,-71.8985900878906,-72.0546875,-72.1023941040039,-72.1464309692383,-72.1136245727539,-72.1300277709961,-72.1436996459961,-72.1054153442383,-72.053466796875,-72.078125,-72.1246109008789,-72.1082077026367,-72.0644073486328,-72.026123046875,-71.9933090209961,-71.944091796875,-71.8607864379883,-71.7609405517578,-71.75,-71.77001953125,-71.844482421875,-71.9112777709961,-71.8976058959961,-71.8711471557617,-71.8921890258789,-71.8855972290039,-71.8807144165039,-71.873046875,-71.9413604736328,-71.93212890625,-71.8837890625,-71.8385238647461,-71.8046417236328,-71.7691421508789,-71.708984375,-71.6953125,-71.72265625,-71.8005905151367,-71.8183135986328,-71.8019561767578,-71.763671875,-71.7043914794922,-71.6597595214844,-71.6471176147461,-71.6378936767578,-71.6720733642578,-71.6968231201172,-71.7199249267578,-71.6925735473633,-71.654296875,-71.5870132446289,-71.5394592285156,-71.53125,-71.5257797241211,-71.5077667236328,-71.4653854370117,-71.4200134277344,-71.4093780517578,-71.4255828857422,-71.4015579223633,-71.3531799316406,-71.2857437133789,-71.197265625,-71.0871047973633,-70.9516067504883,-70.8969268798828,-70.858642578125,-70.84765625,-70.899658203125,-70.9679641723633,-71.00048828125,-71.0181655883789,-71.0281753540039,-71.09619140625,-71.1675796508789,-71.1867141723633,-71.162841796875,-71.1348114013672,-71.1648941040039,-71.2003936767578,-71.1634750366211,-71.118408203125,-71.1238250732422,-71.1593780517578,-71.1921920776367,-71.1593780517578,-71.107421875,-71.06640625,-71.0732421875,-71.0555191040039,-70.9779357910156,-70.9051284790039,-70.8531723022461,-70.790283203125,-70.749267578125,-70.7328567504883,-70.721923828125,-70.6218719482422,-70.5633773803711,-70.4567413330078,-70.40478515625,-70.4036636352539,-70.4157257080078,-70.3801727294922,-70.4197311401367,-70.4157257080078,-70.4567413330078,-70.4485397338867,-70.4704055786133,-70.5323257446289,-70.55517578125,-70.5251007080078,-70.4666061401367,-70.3931655883789,-70.338134765625,-70.3121109008789,-70.2867660522461,-70.2899398803711,-70.2546920776367,-70.210693359375,-70.1412582397461,-70.1014633178711,-70.06298828125,-70.0520553588867,-70.0028305053711,-69.9463348388672,-69.8797912597656,-69.8524398803711,-69.8573684692383,-69.8615264892578,-69.8814926147461,-69.8943328857422,-69.882568359375,-69.8387680053711,-69.7977523803711,-69.8086929321289,-69.8196258544922,-69.8961944580078,-69.9690475463867,-70.0198211669922,-70.0848617553711,-70.10400390625,-70.0930709838867,-70.0421371459961,-70.02197265625,-70.0520553588867,-70.1161575317383,-70.1769561767578,-70.1696243286133,-70.2297897338867,-70.2578125,-70.3200225830078,-70.3446273803711,-70.36376953125,-70.3555679321289,-70.2909164428711,-70.25439453125,-70.28173828125,-70.3309555053711,-70.3938522338867,-70.4501419067383,-70.525634765625,-70.585205078125,-70.56640625,-70.5546875,-70.529052734375,-70.5195770263672,-70.4730911254883,-70.4293899536133,-70.3883743286133,-70.3505859375,-70.30908203125,-70.3118209838867,-70.33642578125,-70.34814453125,-70.3192367553711,-70.2693786621094,-70.1939392089844,-70.1614227294922,-70.1696243286133,-70.1532211303711,-70.1020050048828,-69.9563446044922,-69.9071273803711,-69.8880386352539,-69.8442840576172,-69.8633804321289,-69.9235382080078,-69.9599609375,-69.9454574584961,-69.9241256713867,-69.9276351928711,-69.9826202392578,-70.0268096923828,-69.99560546875,-69.9003372192383,-69.827880859375,-69.8148498535156,-69.7431640625,-69.7349166870117,-69.6878890991211,-69.6569366455078,-69.5271453857422,-69.4888687133789,-69.4369125366211,-69.4095687866211,-69.3407211303711,-69.251220703125,-69.1744155883789,-69.1552734375,-69.1185073852539,-69.0421905517578,-68.9994125366211,-68.9419860839844,-68.8750915527344,-68.8463363647461,-68.769775390625,-68.7096176147461,-68.6521987915039,-68.5920944213867,-68.537353515625,-68.4053726196289,-68.3459930419922,-68.3186492919922,-68.3186492919922,-68.3733444213867,-68.485107421875,-68.5811538696289,-68.5915985107422,-68.5921859741211,-68.5757827758789,-68.5298385620117,-68.4144973754883,-68.4267578125,-68.5108413696289,-68.5419006347656,-68.6002960205078,-68.5920944213867,-68.5408172607422,-68.496337890625,-68.4307174682617,-68.3952102661133,-68.3842315673828,-68.4280242919922,-68.4471206665039,-68.46630859375,-68.5270462036133,-68.56201171875,-68.56201171875,-68.5072784423828,-68.4471206665039,-68.4225616455078,-68.3581008911133,-68.2995147705078,-68.2502975463867,-68.04736328125,-67.88623046875,-67.57177734375,-67.356201171875,-67.3355941772461,-67.3191452026367,-67.2191390991211,-67.0897445678711,-67.0087890625,-67.1948776245117,-67.1619110107422,-67.055419921875,-67.0335464477539,-66.9911117553711,-66.80029296875,-66.7674865722656,-66.7506332397461,-66.7117233276367,-66.6390151977539,-66.5069808959961,-66.3651809692383,-66.3224639892578,-66.2821273803711,-66.2476119995117,-66.2201690673828,-66.1746597290039,-66.0985794067383,-66.05859375,-65.8601531982422,-65.7710494995117,-65.6861801147461,-65.518798828125,-65.48486328125,-65.0578079223633,-64.9926223754883,-64.8430633544922,-64.7586441040039,-64.7001037597656,-64.6055145263672,-64.5236358642578,-64.4777374267578,-64.4455032348633,-64.3739700317383,-64.3252944946289,-64.3079147338867,-64.2664031982422,-64.2090835571289,-64.1318359375,-63.9761199951172,-63.9216766357422,-63.8610305786133,-63.8186492919922,-63.7755851745605,-63.7169418334961,-63.6753387451172,-63.2671890258789,-62.8433609008789,-62.8342781066895,-62.8150863647461,-62.7445831298828,-62.665283203125,-62.6509742736816],[45.5523452758789,45.5143547058105,45.4788093566895,45.4788093566895,45.5044937133789,45.5341796875,45.5623016357422,45.5523452758789],[46.4906234741211,46.3177719116211,46.1701202392578,46.1144523620605,46.0774383544922,46.0458984375,45.9518547058105,45.9774436950684,45.9249992370605,45.7986297607422,45.7663078308105,45.7841796875,45.7964820861816,45.7844734191895,45.75048828125,45.6874008178711,45.6107444763184,45.4568367004395,45.3499031066895,45.2882804870605,45.2525405883789,45.1725578308105,45.15283203125,45.1486320495605,45.1246109008789,45.0764656066895,45.0316429138184,44.8671875,44.7682647705078,44.7337875366211,44.5604476928711,44.3996124267578,44.2892570495605,44.1780242919922,44.00537109375,43.9419898986816,43.7916984558105,43.6662139892578,43.6833000183105,43.7098655700684,43.6781234741211,43.6083984375,43.6158180236816,43.59375,43.5693359375,43.6678733825684,43.712890625,43.72265625,43.6964836120605,43.6316413879395,43.5916976928711,43.5174827575684,43.4552726745605,43.439453125,43.4919929504395,43.64501953125,43.7931632995605,43.9091796875,44.0772476196289,44.146484375,44.2273445129395,44.4730491638184,44.5648422241211,44.8414077758789,44.8485336303711,44.8109397888184,44.8113288879395,44.9758796691895,45.0013656616211,45.02294921875,45.0847625732422,45.15234375,45.1885719299316,45.1902351379395,45.0707054138184,45.0625953674316,45.0705108642578,45.1060523986816,45.2734375,45.3689422607422,45.4191398620605,45.4442367553711,45.5240249633789,45.5875015258789,45.5914077758789,45.5793952941895,45.4013671875,45.37890625,45.3761711120605,45.4543914794922,45.5695304870605,45.7357406616211,45.9646492004395,45.9675788879395,45.9312515258789,45.9000968933105,45.8859367370605,45.8581047058105,45.6301765441895,45.5959968566895,45.5809555053711,45.5797843933105,45.6618156433105,45.7896499633789,45.8631820678711,45.93994140625,46.02587890625,46.0948219299316,46.2020492553711,46.3216819763184,46.4814453125,46.4880828857422,46.4781265258789,46.3776359558105,46.3651390075684,46.365234375,46.37841796875,46.4373054504395,46.5066413879395,46.5847663879395,46.5499992370605,46.4771499633789,46.4203109741211,46.4002914428711,46.4014625549316,46.4753913879395,46.4898414611816,46.4867172241211,46.4906234741211],[44.9690437316895,45.0020484924316,45.0236320495605,45.0287132263184,45.0210914611816,44.9943351745605,44.96142578125,44.9588851928711,44.9690437316895],[-170.726272583008,-170.769241333008,-170.820510864258,-170.720855712891,-170.689163208008,-170.568115234375,-170.640472412109,-170.726272583008],[-161.993789672852,-162.304931640625,-163.046539306641,-163.242141723633,-163.348388671875,-163.552108764648,-163.601760864258,-163.602203369141,-163.634323120117,-163.703918457031,-163.735305786133,-163.795944213867,-162.798492431641,-162.410598754883,-162.339797973633,-161.635162353516,-161.828216552734,-161.993789672852],[-157.987487792969,-158.076217651367,-158.154098510742,-158.545318603516,-158.773193359375,-158.926315307617,-158.988723754883,-158.913726806641,-158.346740722656,-158.260833740234,-157.834564208984,-157.987487792969],[-160.467132568359,-160.570999145508,-163.253082275391,-163.766067504883,-163.89013671875,-163.939254760742,-163.951217651367,-163.930038452148,-163.869003295898,-163.200637817383,-162.456497192383,-161.55859375,-160.937896728516,-160.616897583008,-160.485443115234,-160.467132568359],[-153.930465698242,-154.114059448242,-154.348831176758,-154.529434204102,-154.941650390625,-155.044769287109,-155.525344848633,-155.751159667969,-155.674270629883,-155.162216186523,-154.535110473633,-154.025390625,-153.930465698242],[-59.7339401245117,-59.7737312316895,-59.7713394165039,-59.8254890441895,-59.9263648986816,-60.1246604919434,-60.2682113647461,-60.5828132629395,-62.0233421325684,-62.6705093383789,-62.9403762817383,-62.986083984375,-63.0675277709961,-63.1439933776855,-63.7149429321289,-64.0650329589844,-64.12646484375,-64.2199249267578,-64.2682647705078,-65.2028274536133,-66.1837387084961,-66.5914077758789,-66.7335891723633,-66.7711410522461,-66.6810989379883,-66.5884323120117,-66.482666015625,-66.376953125,-66.2958068847656,-66.2173767089844,-66.1676254272461,-66.115478515625,-65.9800796508789,-62.518798828125,-62.2318344116211,-61.6332969665527,-61.3128395080566,-61.1939926147461,-61.4847679138184,-61.5974617004395,-61.6943321228027,-61.7168464660645,-61.6841812133789,-61.3023414611816,-61.2462387084961,-61.3462371826172,-61.3431167602539,-61.114990234375,-61.0263214111328,-60.5788040161133,-59.8733444213867,-59.7063446044922,-59.75244140625,-59.78564453125,-59.7877960205078,-59.4981460571289,-59.4078102111816,-59.3216819763184,-59.4266128540039,-59.5300788879395,-59.6124038696289,-59.6833000183105,-59.7339401245117],[-31.1188468933105,-30.9850578308105,-30.8408203125,-30.8614749908447,-30.7799339294434,-30.66015625,-29.8708992004395,-29.6144542694092,-29.7207527160645,-29.8000984191895,-30.0290050506592,-30.4221210479736,-30.8444347381592,-31.59423828125,-31.8241214752197,-32.0002937316895,-31.6804180145264,-31.6048831939697,-31.1188468933105],[-32.342529296875,-32.514892578125,-32.583251953125,-32.5008316040039,-32.3766136169434,-32.1500015258789,-31.9334468841553,-31.9567375183105,-32.0011711120605,-32.342529296875],[-66.1736297607422,-66.2671813964844,-66.3194808959961,-66.3668899536133,-66.410400390625,-66.9041976928711,-66.9623565673828,-66.99365234375,-67.0772476196289,-67.7191390991211,-67.770751953125,-67.8087387084961,-67.6879425048828,-67.438232421875,-66.9789047241211,-66.8813018798828,-66.7852096557617,-66.2737731933594,-66.01416015625,-65.8703079223633,-65.5792465209961,-65.53955078125,-65.5044479370117,-65.8990249633789,-65.9894027709961,-66.1736297607422],[-67.2619171142578,-67.434326171875,-68.1614761352539,-68.4081115722656,-68.5489273071289,-68.4222640991211,-68.3243637084961,-68.2330093383789,-68.0325698852539,-67.7141571044922,-67.4745101928711,-67.0686569213867,-67.1729507446289,-67.239501953125,-67.3048858642578,-67.2619171142578],[-33.9341812133789,-34.0499496459961,-36.4813003540039,-36.6007804870605,-36.5659675598145,-36.23779296875,-36.0479965209961,-35.7902374267578,-35.5973129272461,-35.53466796875,-34.3915061950684,-33.9947242736816,-33.9471664428711,-33.9341812133789],[-159.052932739258,-160.302093505859,-160.806701660156,-161.865524291992,-163.317245483398,-163.71240234375,-163.970901489258,-164.225784301758,-164.281631469727,-164.244873046875,-164.199508666992,-164.125534057617,-163.814636230469,-163.660247802734,-163.345306396484,-163.256103515625,-163.124130249023,-162.872756958008,-162.62158203125,-162.390045166016,-162.160705566406,-161.642974853516,-161.283447265625,-160.764404296875,-160.249618530273,-159.963516235352,-159.68408203125,-159.418807983398,-159.366394042969,-159.256103515625,-159.189697265625,-159.118743896484,-159.051361083984,-158.99658203125,-159.009567260742,-159.052932739258],[-70.3340835571289,-70.5525436401367,-70.9837417602539,-71.4140090942383,-71.5257797241211,-71.6866683959961,-71.7351989746094,-71.7777328491211,-71.7835464477539,-71.66748046875,-71.4540023803711,-71.2544937133789,-70.6258773803711,-70.5439987182617,-69.9718704223633,-69.7476577758789,-69.3982391357422,-67.478515625,-67.0381393432617,-66.8402328491211,-66.7280731201172,-66.7870101928711,-67.046142578125,-67.1664047241211,-67.48095703125,-68.1570281982422,-68.637939453125,-69.2508773803711,-69.3944320678711,-69.6864700317383,-69.6344757080078,-69.7317352294922,-70.1163101196289,-70.3340835571289],[167.642776489258,167.517486572266,167.376953125,166.936233520508,166.6259765625,166.284866333008,166.121887207031,166.050582885742,166.012496948242,166.012878417969,166.111129760742,166.567092895508,166.759857177734,166.863662719727,167.137878417969,167.364059448242,167.42236328125,167.497848510742,167.593856811523,167.639846801758,167.642776489258],[-45.22265625,-45.0916023254395,-44.5663070678711,-44.0411605834961,-43.7220687866211,-43.6273460388184,-43.5443344116211,-43.4506340026855,-43.3636741638184,-43.2672386169434,-43.2105445861816,-43.1187019348145,-42.9653816223145,-42.9446792602539,-42.989990234375,-43.06591796875,-43.2672843933105,-43.4958000183105,-43.6002922058105,-43.7036628723145,-43.7421875,-43.7582054138184,-43.489990234375,-43.4588356018066,-43.4537582397461,-43.4893531799316,-43.5279273986816,-49.1875,-49.4104499816895,-49.6296882629395,-49.7013206481934,-49.7730484008789,-54.1624984741211,-54.2020988464355,-54.2413101196289,-54.3507804870605,-54.37158203125,-54.34716796875,-54.1295890808105,-54.044921875,-53.6760711669922,-53.4823226928711,-53.3938980102539,-53.3464851379395,-53.1764183044434,-53.053466796875,-52.8072242736816,-52.5667953491211,-52.4609870910645,-52.3571281433105,-52.3380851745605,-52.2971687316895,-51.7113265991211,-51.1838340759277,-50.6643562316895,-50.4015617370605,-50.3392601013184,-50.294921875,-50.3313446044922,-50.378662109375,-50.4195327758789,-50.4638710021973,-50.7330589294434,-50.6490211486816,-50.5732917785645,-50.5203132629395,-50.5023918151855,-50.5137176513672,-50.5024909973145,-50.3797874450684,-50.2977523803711,-50.241943359375,-50.33544921875,-50.3773918151855,-50.294189453125,-50.2196273803711,-50.1416511535645,-49.9397468566895,-49.3541526794434,-49.1436500549316,-49.0812492370605,-47.6920890808105,-47.4634780883789,-47.0299301147461,-46.82568359375,-46.2578620910645,-45.9931640625,-45.5302734375,-44.8519515991211,-44.594482421875,-44.33984375,-44.093994140625,-43.8544921875,-43.80859375,-43.7845687866211,-43.7767066955566,-43.7882804870605,-43.852294921875,-43.9472160339355,-45.0678215026855,-45.2130851745605,-45.2947769165039,-45.3523445129395,-45.22265625],[-149.218124389648,-148.928863525391,-149.438659667969,-149.662353515625,-149.518646240234,-149.375442504883,-149.249130249023,-149.218124389648],[-150.397064208984,-150.474853515625,-151.344482421875,-151.511627197266,-151.218017578125,-151.021530151367,-150.499114990234,-150.356246948242,-150.397064208984],[167.084075927734,167.4609375,168.450790405273,169.275588989258,169.352737426758,169.117294311523,168.754592895508,168.518859863281,168.322555541992,167.917572021484,167.392288208008,167.279495239258,167.025100708008,166.72900390625,166.650390625,166.532531738281,166.23681640625,166.216796875,166.37841796875,166.4580078125,166.626373291016,166.607116699219,166.469039916992,166.4130859375,166.50634765625,166.716400146484,166.9873046875,167.106842041016,167.0849609375,167.084075927734],[-148.595840454102,-149.0146484375,-149.244827270508,-149.302536010742,-149.238082885742,-148.7041015625,-148.508941650391,-148.439743041992,-148.474304199219,-148.595840454102],[-149.230651855469,-149.29345703125,-149.728622436523,-149.81689453125,-149.856338500977,-150.461807250977,-150.735778808594,-150.788528442383,-150.680236816406,-150.475769042969,-150.393310546875,-149.87060546875,-149.789840698242,-149.742584228516,-149.505767822266,-149.441757202148,-149.416412353516,-149.288330078125,-149.230651855469],[-146.606735229492,-146.9814453125,-147.078903198242,-147.044143676758,-147.101470947266,-147.115615844727,-147.086669921875,-146.866500854492,-146.244384765625,-146.163955688477,-146.606735229492],[-149.333251953125,-148.927932739258,-148.66259765625,-148.384185791016,-148.32080078125,-148.370941162109,-148.669525146484,-148.814605712891,-148.98388671875,-149.238479614258,-149.468948364258,-149.333251953125],[-150.232528686523,-150.655181884766,-150.830459594727,-150.87353515625,-150.837646484375,-150.177383422852,-150.103576660156,-150.084762573242,-150.232528686523],[-147.588287353516,-147.578857421875,-147.729644775391,-147.954299926758,-148.001068115234,-147.899703979492,-147.769683837891,-147.649108886719,-147.588287353516],[-146.7900390625,-146.907806396484,-147.221343994141,-147.355316162109,-147.278610229492,-147.135299682617,-146.947463989258,-146.877883911133,-146.7900390625],[-146.690139770508,-146.894485473633,-147.150939941406,-147.345397949219,-147.407760620117,-147.4208984375,-147.418060302734,-147.360641479492,-146.949020385742,-146.690139770508],[162.968063354492,162.788284301758,162.661911010742,162.591201782227,162.720016479492,162.842376708984,162.9169921875,162.968063354492],[-145.238082885742,-145.348388671875,-145.541168212891,-146.042770385742,-146.150833129883,-146.075729370117,-145.8955078125,-145.760787963867,-145.417373657227,-145.3154296875,-145.252258300781,-145.238082885742],[163.975967407227,163.844619750977,163.763381958008,163.737197875977,163.74169921875,164.002349853516,164.20849609375,164.098251342773,164.059173583984,163.975967407227],[-132.391265869141,-132.546295166016,-132.857177734375,-132.831695556641,-132.552490234375,-132.3623046875,-132.391265869141],[-131.066696166992,-131.178909301758,-131.597946166992,-131.840881347656,-131.952499389648,-132.025100708008,-132.04931640625,-132.162643432617,-131.937789916992,-131.7626953125,-131.594085693359,-131.559616088867,-131.23388671875,-130.981109619141,-130.956787109375,-130.96728515625,-131.066696166992],[-127.365814208984,-127.517723083496,-127.817085266113,-127.915237426758,-128.000061035156,-128.070465087891,-128.096237182617,-128.133453369141,-128.042861938477,-127.852973937988,-127.486419677734,-127.22972869873,-127.145156860352,-127.23193359375,-127.365814208984],[-116.738632202148,-117.230323791504,-117.362991333008,-117.398284912109,-117.376457214355,-116.381301879883,-116.202690124512,-116.154975891113,-116.451416015625,-116.584564208984,-116.60863494873,-116.534172058105,-116.514106750488,-116.570892333984,-116.738632202148],[-119.548927307129,-119.758354187012,-119.820999145508,-119.88720703125,-119.905128479004,-119.796684265137,-119.693748474121,-119.661567687988,-119.802581787109,-119.669036865234,-119.516357421875,-119.216209411621,-118.959274291992,-118.909812927246,-118.877342224121,-118.989791870117,-119.058601379395,-119.448532104492,-119.548927307129],[-120.556251525879,-120.378120422363,-120.312309265137,-120.2724609375,-120.98950958252,-121.01904296875,-121.054107666016,-121.036430358887,-121.004684448242,-121.00244140625,-121.062400817871,-122.286628723145,-122.859077453613,-122.938430786133,-122.9560546875,-122.890625,-122.764739990234,-122.794189453125,-122.87523651123,-122.880760192871,-122.710006713867,-122.624710083008,-122.951179504395,-122.991554260254,-123.03466796875,-123.190818786621,-123.34619140625,-123.291801452637,-123.249069213867,-123.112152099609,-123.012496948242,-122.910446166992,-122.435691833496,-121.966705322266,-121.497314453125,-120.722213745117,-120.556251525879],[-20.607421875,-20.6542472839355,-20.6412601470947,-20.600341796875,-20.42333984375,-20.4114265441895,-20.4166488647461,-20.489013671875,-20.7370109558105,-20.8176746368408,-20.8456535339355,-20.9767570495605,-21.0512199401855,-21.1665534973145,-21.60986328125,-22.0353507995605,-21.9303722381592,-21.2882804870605,-21.1263675689697,-21.0245113372803,-20.9791984558105,-20.8670406341553,-20.6901378631592,-20.5802230834961,-20.5207023620605,-20.5207023620605,-20.607421875],[169.843643188477,169.709182739258,169.522354125977,169.4794921875,169.659378051758,169.645416259766,169.671875,169.740036010742,169.783203125,169.886535644531,169.960357666016,169.858779907227,169.843643188477],[-126.329879760742,-126.065277099609,-125.975875854492,-125.856643676758,-125.735794067383,-125.626808166504,-125.561332702637,-125.50390625,-125.326171875,-125.263961791992,-125.276077270508,-125.612396240234,-125.723197937012,-125.828422546387,-125.859909057617,-125.857025146484,-125.79859161377,-125.674415588379,-125.552383422852,-125.326850891113,-125.224418640137,-125.108795166016,-124.99340057373,-124.694381713867,-124.617485046387,-124.539985656738,-124.128509521484,-124.042037963867,-124.100532531738,-124.151802062988,-124.129341125488,-123.932319641113,-123.851181030273,-123.800445556641,-123.811027526855,-123.838775634766,-123.836723327637,-123.937400817871,-123.982467651367,-124.199409484863,-124.873001098633,-125.089546203613,-125.420799255371,-125.54931640625,-125.682723999023,-125.886863708496,-126.244094848633,-126.4716796875,-126.46556854248,-126.496101379395,-126.538375854492,-126.582664489746,-126.7109375,-126.838241577148,-126.901664733887,-127.006401062012,-127.1220703125,-127.211616516113,-127.231643676758,-127.232917785645,-127.33203125,-127.414352416992,-127.429046630859,-127.394340515137,-127.267623901367,-127.12353515625,-126.977836608887,-126.829971313477,-126.596870422363,-126.329879760742],[-73.87841796875,-73.9748077392578,-74.0382843017578,-74.1466293334961,-74.1344223022461,-74.08447265625,-74.0487289428711,-73.8321304321289,-73.6742172241211,-73.5424270629883,-73.6822738647461,-73.7213897705078,-73.87841796875],[-104.537940979004,-104.659568786621,-104.880958557129,-105.052925109863,-105.123435974121,-105.131790161133,-105.084571838379,-104.972412109375,-104.537940979004],[-74.3544464111328,-74.4989242553711,-74.5224609375,-74.6676788330078,-74.6153869628906,-74.5754852294922,-74.5506362915039,-74.4671401977539,-74.3661193847656,-74.4521026611328,-74.5746536254883,-75.9008331298828,-76.0032272338867,-76.0531234741211,-76.0904312133789,-76.0963897705078,-76.0624008178711,-76.0176773071289,-75.8976593017578,-75.774658203125,-75.505859375,-75.4675827026367,-75.417236328125,-75.2762145996094,-75.2438507080078,-75.439453125,-75.6002883911133,-75.7017593383789,-75.7310562133789,-75.3768539428711,-74.473876953125,-74.3355407714844,-74.2757797241211,-74.223876953125,-74.3544464111328],[-93.7956085205078,-93.9655303955078,-94.078125,-94.1131820678711,-94.0469741821289,-94.0042495727539,-93.799560546875,-93.7558059692383,-93.7956085205078],[-91.1606903076172,-91.3441925048828,-91.5108413696289,-91.4503936767578,-91.3568878173828,-91.3821334838867,-91.5514678955078,-91.6700210571289,-91.6124038696289,-91.3035202026367,-90.9474105834961,-90.80712890625,-90.7633285522461,-90.7801742553711,-90.8953552246094,-90.7762222290039,-90.7509765625,-90.7755813598633,-90.89306640625,-90.9984359741211,-91.1606903076172],[-95.0270462036133,-95.2194366455078,-95.27294921875,-95.2156219482422,-94.7530288696289,-94.5661163330078,-94.5383758544922,-94.5138702392578,-94.4339370727539,-94.426025390625,-95.0270462036133],[-16.1044921875,-16.1748046875,-16.3175773620605,-16.4530277252197,-16.509765625,-16.5165538787842,-16.4553718566895,-16.3558597564697,-16.3028812408447,-16.1725101470947,-16.1044921875],[68.4617156982422,68.4088897705078,68.4361343383789,68.5663070678711,68.6670913696289,68.7292938232422,68.8404312133789,68.8171920776367,68.6695327758789,68.4617156982422],[-12.5088872909546,-12.5884275436401,-12.7201662063599,-12.888427734375,-12.9437007904053,-12.9632816314697,-12.914794921875,-12.87548828125,-12.7888679504395,-12.6366205215454,-12.5347661972046,-12.5088872909546],[69.9183578491211,69.7919921875,69.7435531616211,69.6925811767578,69.7371139526367,69.7960891723633,69.8952178955078,69.9183578491211],[-98.0911102294922,-98.1759338378906,-98.16796875,-97.9231414794922,-97.8160171508789,-97.5847625732422,-97.4734878540039,-97.5819854736328,-97.525634765625,-97.4602584838867,-97.34521484375,-97.2419891357422,-97.1955108642578,-97.15478515625,-97.0887222290039,-96.8694305419922,-96.3833465576172,-96.125,-96.2981948852539,-96.7149353027344,-96.9789047241211,-96.89013671875,-96.7987289428711,-96.7175750732422,-96.4823226928711,-95.9063491821289,-95.6853942871094,-95.609375,-95.6095199584961,-95.5310592651367,-95.5753936767578,-95.82568359375,-96.0781707763672,-96.0143051147461,-96.0298843383789,-96.0517578125,-96.6926727294922,-96.8039093017578,-96.914794921875,-97.0276412963867,-97.2502975463867,-97.3655319213867,-97.5956039428711,-97.8283233642578,-98.1634292602539,-98.4078140258789,-98.6406784057617,-98.8815460205078,-99.1488265991211,-99.434326171875,-99.67236328125,-100.014251708984,-100.104049682617,-100.195213317871,-100.357421875,-101.601951599121,-101.784759521484,-101.9033203125,-102.022117614746,-102.264793395996,-102.313621520996,-102.288276672363,-102.236526489258,-102.128120422363,-100.400924682617,-100.218658447266,-100.084617614746,-99.9851608276367,-99.8331985473633,-99.7839813232422,-99.7349166870117,-99.5630874633789,-99.2541046142578,-99.0821228027344,-98.9645538330078,-98.6154327392578,-98.394287109375,-98.1891632080078,-98.0911102294922],[-2.95498061180115,-3.06059551239014,-3.20146465301514,-3.30937504768372,-3.38564419746399,-3.40385746955872,-3.39169931411743,-3.39897465705872,-3.26303696632385,-3.21279311180115,-3.19135713577271,-2.95498061180115],[-60.5522422790527,-60.6521492004395,-60.7897491455078,-60.9063491821289,-60.9464797973633,-60.8890609741211,-60.7828102111816,-60.6131324768066,-60.5333023071289,-60.5163078308105,-60.5359382629395,-60.5522422790527],[-2.53281259536743,-2.42343735694885,-2.25556635856628,-2.09228491783142,-2.11904311180115,-2.21269559860229,-2.29316377639771,-2.36894559860229,-2.60673856735229,-2.78349614143372,-2.82514667510986,-2.82187485694885,-2.80517554283142,-2.800537109375,-2.96313500404358,-2.97500038146973,-3.00698232650757,-3.48896455764771,-3.57465839385986,-3.53706049919128,-3.0400390625,-2.74980497360229,-2.53281259536743],[-73.7066421508789,-73.5503387451172,-73.6945343017578,-74.2050247192383,-74.5047378540039,-74.8058090209961,-76.1763229370117,-76.2712860107422,-76.3639678955078,-76.4214859008789,-76.5115280151367,-76.500244140625,-76.3776321411133,-76.2488784790039,-76.0345687866211,-75.2100067138672,-75.126953125,-75.0599136352539,-75.0375518798828,-75.0074691772461,-74.95361328125,-74.8984909057617,-74.7904739379883,-74.5896987915039,-74.5271530151367,-74.4686508178711,-74.4561538696289,-74.4009780883789,-74.2249984741211,-74.1145553588867,-74.1126480102539,-74.038330078125,-73.9578170776367,-73.8794937133789,-73.7066421508789],[-60.7406272888184,-60.8260726928711,-60.896484375,-60.9580116271973,-60.9755363464355,-60.9417953491211,-60.8838882446289,-60.5536651611328,-60.4524879455566,-60.4489784240723,-60.4876976013184,-60.7406272888184],[-5.89409208297729,-6.15610361099243,-6.18085956573486,-6.26611328125,-6.43798828125,-6.24365234375,-6.06826210021973,-5.97163105010986,-5.94951200485229,-5.89409208297729],[3.03691411018372,2.69775390625,2.62275385856628,2.58466815948486,2.63144493103027,3.07216835021973,3.19277334213257,3.23056626319885,3.25986337661743,3.22128891944885,3.17109394073486,3.03691411018372],[-3.28022456169128,-3.44189429283142,-3.490234375,-3.496826171875,-3.28745126724243,-3.17324185371399,-2.94990229606628,-2.80512690544128,-2.71352529525757,-2.68437480926514,-2.68271493911743,-2.73803687095642,-3.28022456169128],[-71.6950225830078,-71.6477584838867,-71.4315948486328,-71.3548812866211,-71.3402786254883,-71.437744140625,-71.5512161254883,-71.6846694946289,-71.781982421875,-71.7952651977539,-71.6950225830078],[72.0022506713867,71.9290008544922,71.8412170410156,71.7257843017578,71.6593704223633,71.6371078491211,71.6465835571289,71.705078125,71.7965850830078,71.8379898071289,71.8511734008789,71.8799819946289,72,72.0556640625,72.0734405517578,72.0973663330078,72.078125,72.0022506713867],[4.52587890625,4.36523389816284,4.1796875,4.12959003448486,4.076171875,4.06972694396973,4.11171865463257,4.25605487823486,4.49501943588257,4.58623027801514,4.61757802963257,4.58994150161743,4.52587890625],[26.8572254180908,26.79296875,26.6087894439697,26.47021484375,26.3576164245605,26.00537109375,25.9642562866211,25.9541015625,25.9825191497803,26.3010749816895,26.4255847930908,26.6047859191895,26.6862297058105,26.7374019622803,26.8748054504395,26.8572254180908],[1.29931652545929,1.21152329444885,1.15634739398956,1.10458981990814,0.990331768989563,0.952538907527924,0.949609339237213,1.02675759792328,1.31484365463257,1.41220688819885,1.4609375,1.29931652545929],[-61.1584510803223,-61.3086471557617,-61.3784713745117,-61.4043426513672,-61.386474609375,-61.3271484375,-61.1518592834473,-61.1079139709473,-61.1584510803223],[-74.9871063232422,-74.8101501464844,-74.5497055053711,-74.4654235839844,-74.43798828125,-74.4600601196289,-74.5784225463867,-74.6717758178711,-74.8488235473633,-75.2684097290039,-75.7267608642578,-75.7644577026367,-75.8041534423828,-75.8129425048828,-75.759521484375,-75.6813507080078,-75.3399429321289,-75.3139114379883,-75.2645492553711,-75.178955078125,-75.1363754272461,-74.9871063232422],[16.22265625,16.1592769622803,15.8449220657349,15.6634759902954,15.6138668060303,15.5709953308105,15.5625982284546,15.5968751907349,15.6990242004395,15.9095697402954,16.2468757629395,16.5734367370605,16.62548828125,16.3154296875,16.22265625],[-71.9853515625,-72.2020492553711,-72.3445816040039,-72.7767562866211,-72.9573211669922,-72.936767578125,-72.8573303222656,-72.7261734008789,-72.4643020629883,-72.3311538696289,-71.9853515625],[-61.9976081848145,-62.0851554870605,-62.1719284057617,-62.2160148620605,-62.4962387084961,-62.5676765441895,-62.515869140625,-62.4421348571777,-62.2389640808105,-62.117919921875,-61.9783668518066,-61.8159637451172,-61.78369140625,-61.8071823120117,-61.9113273620605,-61.9078102111816,-61.9701156616211,-61.9976081848145],[-70.0511245727539,-70.079345703125,-69.9132843017578,-69.85498046875,-69.7075653076172,-69.4169387817383,-69.3529739379883,-69.2337875366211,-69.0912551879883,-68.7305679321289,-68.5535125732422,-68.45947265625,-68.4507827758789,-68.3359909057617,-68.3140182495117,-68.2779312133789,-68.25244140625,-68.2278289794922,-68.2410125732422,-68.3938980102539,-68.4607391357422,-68.5425796508789,-68.6400375366211,-69.1492691040039,-69.2093200683594,-70.0630798339844,-70.54345703125,-70.7313461303711,-70.9229507446289,-71.1585922241211,-71.8461456298828,-72.3674774169922,-72.4333038330078,-72.4798812866211,-72.5308074951172,-72.6702117919922,-72.7803726196289,-72.8876495361328,-73.0070343017578,-73.0574264526367,-73.0863800048828,-72.8548355102539,-72.7374954223633,-72.6182174682617,-72.3760757446289,-72.1349563598633,-71.6053161621094,-70.8726043701172,-70.671142578125,-70.4276885986328,-70.2060012817383,-70.314208984375,-70.4241714477539,-70.533203125,-70.6414108276367,-70.9451675415039,-71.177734375,-71.4125518798828,-71.6614761352539,-71.8921890258789,-71.8975524902344,-71.1066436767578,-71.0343780517578,-70.89111328125,-70.8446273803711,-70.8209915161133,-71.35498046875,-71.4646453857422,-71.5744094848633,-71.816162109375,-72.0459518432617,-72.2590866088867,-72.3366241455078,-72.4122009277344,-72.9276351928711,-72.9720687866211,-73.1667022705078,-73.4099578857422,-73.6329116821289,-73.7759704589844,-73.8298873901367,-73.6905746459961,-73.5723114013672,-73.537109375,-73.8992691040039,-73.99560546875,-74.1529769897461,-74.2089309692383,-74.32177734375,-74.4292984008789,-74.6632308959961,-74.7858352661133,-74.9082489013672,-75.0241241455078,-75.1297378540039,-75.2588806152344,-75.3530807495117,-75.3827133178711,-75.3732376098633,-75.330810546875,-75.3249053955078,-75.3531265258789,-75.33544921875,-75.2925796508789,-75.0996551513672,-74.8631362915039,-74.6363830566406,-74.4874496459961,-74.4186553955078,-74.3917999267578,-74.3733444213867,-74.3800735473633,-74.4203109741211,-74.4253921508789,-74.375,-74.3079071044922,-74.236083984375,-74.1872024536133,-74.0409622192383,-73.9371109008789,-73.7242660522461,-73.5453567504883,-73.47900390625,-73.4271469116211,-73.3801803588867,-73.5921859741211,-73.6170883178711,-73.6044921875,-73.4739761352539,-73.3974075317383,-73.0197296142578,-72.821044921875,-72.62158203125,-72.211669921875,-72.4300765991211,-72.9054641723633,-72.9945297241211,-73.0604019165039,-72.71044921875,-72.3563461303711,-71.718505859375,-71.5044937133789,-71.3075180053711,-71.1940460205078,-70.7411193847656,-70.3806686401367,-70.3229064941406,-70.2676773071289,-69.9164505004883,-69.8697738647461,-69.8351058959961,-69.8228530883789,-69.822509765625,-69.8304214477539,-69.8757781982422,-69.9331970214844,-69.9930191040039,-70.093994140625,-70.1962356567383,-70.2986831665039,-70.66064453125,-70.9169464111328,-71.0494155883789,-71.1726531982422,-71.1900405883789,-71.0610809326172,-70.5621109008789,-70.3280715942383,-70.0904312133789,-69.9752960205078,-69.7020568847656,-69.6598663330078,-69.6183547973633,-69.8830032348633,-70.1178665161133,-70.2340850830078,-70.32763671875,-70.7195358276367,-70.9262237548828,-71.0237274169922,-71.1205062866211,-71.6960906982422,-71.728515625,-71.8099594116211,-71.8536071777344,-71.8792495727539,-71.86767578125,-71.8520050048828,-71.7665023803711,-71.7182159423828,-71.742919921875,-71.8335876464844,-71.9632339477539,-72.0806655883789,-72.1143493652344,-72.1351547241211,-72.1378936767578,-72.108642578125,-72.057861328125,-71.9901428222656,-71.8689956665039,-71.3915557861328,-70.4169921875,-70.3119125366211,-70.154296875,-70.10546875,-70.052734375,-70.0511245727539],[-90.5361785888672,-90.5471725463867,-90.566650390625,-90.5808639526367,-90.59521484375,-90.609130859375,-90.6173858642578,-90.620361328125,-90.6236343383789,-90.6283645629883,-90.6383819580078,-90.6470642089844,-90.6520538330078,-90.6480407714844,-90.63720703125,-90.63671875,-90.63671875,-90.6314468383789,-90.6283645629883,-90.629150390625,-90.6256790161133,-90.6167449951172,-90.60009765625,-90.5896453857422,-90.5706024169922,-90.5616683959961,-90.5484848022461,-90.5354461669922,-90.525146484375,-90.525146484375,-90.5229034423828,-90.5211944580078,-90.5191955566406,-90.5147018432617,-90.5181579589844,-90.5259246826172,-90.532958984375,-90.5361785888672],[-60.6559104919434,-60.6933631896973,-60.820068359375,-60.8940467834473,-61.0149421691895,-60.9471702575684,-60.8135719299316,-60.7048377990723,-60.6559104919434],[-67.3489227294922,-67.5445327758789,-67.693359375,-67.689697265625,-67.7306594848633,-67.7432632446289,-67.5567398071289,-67.4176788330078,-67.2467269897461,-67.1749038696289,-67.1494140625,-67.2796859741211,-67.2997055053711,-67.3489227294922],[164.833602905273,164.746292114258,164.692092895508,164.638961791992,164.6962890625,164.675186157227,164.683990478516,164.825012207031,164.85009765625,164.9072265625,164.918640136719,164.860458374023,164.833602905273],[-67.3623962402344,-67.4092254638672,-67.5207977294922,-67.59326171875,-67.49951171875,-67.5108413696289,-67.4259719848633,-67.3316879272461,-67.2687530517578,-67.2569808959961,-67.3623962402344],[85.8223648071289,85.6501007080078,85.6222686767578,85.6173858642578,85.3587875366211,85.314453125,85.34033203125,85.5528335571289,85.8062515258789,85.9376907348633,85.8223648071289],[48.5459976196289,48.3779296875,48.3044929504395,48.2959976196289,48.2939453125,48.30078125,48.3577156066895,48.6377944946289,48.7510719299316,48.7824249267578,48.7855491638184,48.7747077941895,48.5459976196289],[163.301956176758,163.283599853516,163.234573364258,163.163864135742,163.089645385742,163.156158447266,163.237884521484,163.271102905273,163.299133300781,163.301956176758],[86.5417938232422,86.4266586303711,86.3370132446289,86.2322235107422,86.2777328491211,86.38330078125,86.5206069946289,86.5567398071289,86.6519546508789,86.5417938232422],[-67.9884719848633,-68.092529296875,-68.1750946044922,-68.2503890991211,-68.3251037597656,-68.3813018798828,-68.4394073486328,-68.5067367553711,-68.5804214477539,-68.6223678588867,-68.6640625,-68.7336883544922,-68.8183059692383,-68.9013671875,-68.9823226928711,-69.0975646972656,-69.120361328125,-69.1380386352539,-69.1326675415039,-69.0824279785156,-68.8199234008789,-68.7335968017578,-68.6563415527344,-68.5746078491211,-68.4168472290039,-68.3359375,-67.9376449584961,-67.8305206298828,-67.7111358642578,-67.68115234375,-67.7408218383789,-67.932373046875,-67.9691848754883,-67.9684066772461,-67.9489288330078,-67.8760757446289,-67.8277816772461,-67.7611770629883,-67.6878433227539,-67.8480453491211,-67.9563446044922,-68.0300750732422,-68.1751403808594,-68.235107421875,-68.1444320678711,-68.0069808959961,-67.969482421875,-67.9884719848633],[85.3285140991211,85.2224655151367,85.1361389160156,85.07958984375,85.0687484741211,85.12109375,85.1647491455078,85.1939468383789,85.3285140991211],[98.8460922241211,98.7517547607422,98.6550827026367,98.6051712036133,98.5964889526367,98.7486343383789,98.9498062133789,98.8460922241211],[162.611419677734,162.55712890625,162.511337280273,162.302734375,162.326278686523,162.297256469727,162.302062988281,162.310546875,162.563278198242,162.611419677734],[100.264747619629,100.133201599121,100.08203125,100.076263427734,100.174415588379,100.29052734375,100.281539916992,100.264747619629],[-66.5953140258789,-66.8186569213867,-66.8499984741211,-66.8675308227539,-66.8669891357422,-66.79150390625,-66.7790069580078,-66.63134765625,-66.5751953125,-66.6228561401367,-66.5926284790039,-66.5953140258789],[96.6127014160156,96.7273406982422,96.931640625,97.0055618286133,97.0188446044922,97.015625,96.9339828491211,96.39453125,96.3070297241211,96.3988342285156,96.4998016357422,96.6127014160156],[92.6013717651367,92.4705123901367,92.3330078125,92.2627868652344,92.2481460571289,92.3014602661133,92.4963912963867,92.6337890625,92.66455078125,92.6696319580078,92.6013717651367],[-65.8452606201172,-66.0639190673828,-66.17529296875,-66.1814422607422,-66.1534729003906,-66.0496063232422,-66.0669479370117,-65.9997100830078,-65.9683074951172,-65.8335876464844,-65.6369171142578,-65.66796875,-65.669677734375,-65.7837371826172,-65.8138656616211,-65.8408203125,-65.8357391357422,-65.8452606201172],[100.9814453125,100.546875,100.512306213379,100.3505859375,100.292579650879,100.270309448242,100.324119567871,100.409378051758,100.545120239258,100.60693359375,100.883399963379,101.078712463379,101.220603942871,101.258979797363,101.238380432129,100.9814453125],[103.397270202637,103.337203979492,103.175979614258,103.138481140137,103.124214172363,103.11279296875,103.054389953613,102.788764953613,102.759567260742,102.796096801758,102.892875671387,103.136817932129,103.190818786621,103.181732177734,103.186134338379,103.261032104492,103.37890625,103.397270202637],[-63.3162078857422,-63.4744148254395,-63.5583534240723,-63.4592742919922,-63.3668937683105,-63.2193832397461,-63.1772499084473,-63.2569351196289,-63.3162078857422],[-57.240478515625,-57.3262710571289,-57.4333992004395,-57.4479026794434,-57.4458999633789,-57.3656234741211,-57.3145484924316,-57.0225601196289,-56.8947296142578,-56.9517097473145,-56.9452629089355,-56.9910163879395,-57.240478515625],[-63.1805686950684,-63.2769546508789,-63.1305160522461,-63.0320816040039,-62.9282188415527,-62.8365249633789,-63.0255851745605,-63.2025871276855,-63.2754402160645,-63.3548812866211,-63.4578132629395,-63.5584487915039,-63.6468734741211,-63.739501953125,-63.7699241638184,-63.8043975830078,-64.007080078125,-64.0991668701172,-64.1837387084961,-64.2720718383789,-64.2262191772461,-64.1710968017578,-63.8671379089355,-63.8969268798828,-63.9161605834961,-63.6744117736816,-63.6683120727539,-63.68310546875,-63.6055679321289,-63.5341796875,-63.485595703125,-63.3335914611816,-63.2296371459961,-63.2707061767578,-63.1805686950684],[-62.3257827758789,-62.3958969116211,-62.4551734924316,-62.5081100463867,-62.5796890258789,-62.7270011901855,-62.7817878723145,-62.7468223571777,-62.7208976745605,-62.6430168151855,-62.5040054321289,-62.479736328125,-62.5902824401855,-62.6107406616211,-62.585693359375,-62.5449714660645,-62.4514198303223,-62.3287620544434,-62.2676239013672,-62.2687530517578,-62.0584983825684,-62.0938491821289,-62.1742706298828,-62.1856956481934,-62.3037071228027,-62.3257827758789],[-61.9524383544922,-62.0438957214355,-62.0207557678223,-61.936279296875,-61.7982406616211,-61.8862342834473,-61.9111289978027,-61.9524383544922],[-57.8460006713867,-57.8085441589355,-57.773681640625,-57.7411613464355,-57.7100563049316,-57.5929183959961,-57.479736328125,-57.51708984375,-57.2494621276855,-57.2728042602539,-57.2222671508789,-57.32763671875,-57.4139633178711,-57.33837890625,-57.294677734375,-57.3878898620605,-57.5807609558105,-57.6832008361816,-57.6707496643066,-57.7033233642578,-57.8228530883789,-57.8714828491211,-57.9097671508789,-57.9522438049316,-57.9207000732422,-57.9710922241211,-58.0215835571289,-58.1694831848145,-58.2140655517578,-58.3044395446777,-58.0199699401855,-58.1376953125,-58.1622085571289,-58.1470756530762,-58.2504425048828,-58.3520011901855,-58.3976058959961,-58.4380874633789,-58.4249534606934,-58.3418922424316,-58.2748069763184,-58.1456527709961,-58.0703620910645,-57.970703125,-57.9256858825684,-57.8313484191895,-57.7794952392578,-57.7806625366211,-57.8269577026367,-57.8460006713867],[-57.3741683959961,-57.3602027893066,-57.1608390808105,-57.10400390625,-57.218017578125,-57.2477531433105,-57.3437461853027,-57.6163558959961,-57.6832504272461,-57.4393577575684,-57.3741683959961],[-60.6531257629395,-60.7776870727539,-60.8524436950684,-60.97216796875,-60.8100547790527,-60.7966804504395,-60.71484375,-60.5623512268066,-60.6559104919434,-60.6888656616211,-60.6549835205078,-60.6531257629395],[-55.87255859375,-55.9567375183105,-56.1783714294434,-56.2352066040039,-56.2098655700684,-55.85791015625,-55.7618141174316,-55.7191925048828,-55.87255859375],[-180,-178.389495849609,-178.069046020508,-177.730407714844,-176.985534667969,-176.289016723633,-176.107376098633,-175.874618530273,-175.381011962891,-174.986709594727,-174.663177490234,-171.703659057617,-168.667785644531,-168.048583984375,-167.4921875,-166.911087036133,-163.463714599609,-162.933395385742,-160.820907592773,-157.127487182617,-156.810302734375,-156.459136962891,-156.642776489258,-156.98828125,-157.453903198242,-157.149612426758,-156.489654541016,-156.620986938477,-156.986328125,-158.303421020508,-163.568511962891,-163.685455322266,-163.758987426758,-163.897033691406,-164.114166259766,-164.916839599609,-165.135360717773,-165.240432739258,-165.184814453125,-165.125152587891,-163.899169921875,-163.765197753906,-163.757629394531,-163.821395874023,-164.032135009766,-164.5283203125,-164.68505859375,-164.602828979492,-164.502532958984,-164.123931884766,-164.011322021484,-164.082901000977,-164.950881958008,-165.536331176758,-165.921783447266,-166.649459838867,-167.552886962891,-167.801223754883,-168.052734375,-168.347366333008,-168.497268676758,-168.785018920898,-169.167678833008,-171.187850952148,-171.539413452148,-174.065963745117,-174.172027587891,-174.235946655273,-173.071151733398,-172.851516723633,-172.592926025391,-172.392028808594,-172.124359130859,-171.8212890625,-171.031295776367,-169.440765380859,-169.016067504883,-168.790130615234,-168.603759765625,-168.417663574219,-168.191024780273,-168.054748535156,-167.724273681641,-166.216888427734,-165.619186401367,-164.915634155273,-164.644332885742,-164.445556640625,-164.058395385742,-163.7333984375,-163.111083984375,-162.912063598633,-162.574172973633,-162.197265625,-160.5947265625,-159.923538208008,-159.444381713867,-157.699279785156,-157.428466796875,-157.027786254883,-157.355804443359,-157.589218139648,-157.679489135742,-157.521881103516,-157.018264770508,-156.037017822266,-155.459426879883,-155.150253295898,-153.822265625,-153.398635864258,-153.009857177734,-153.882614135742,-154.717437744141,-154.451461791992,-154.1884765625,-154.061370849609,-153.956649780273,-154.232070922852,-154.485153198242,-154.907821655273,-156.492584228516,-157.032516479492,-156.815093994141,-156.528228759766,-155.921142578125,-152.034759521484,-148.122741699219,-148.0234375,-148.54296875,-148.984191894531,-149.147171020508,-149.207427978516,-149.214065551758,-149.264404296875,-149.428604125977,-150.132766723633,-150.281692504883,-150.51611328125,-150.575393676758,-150.435455322266,-150.220703125,-149.845367431641,-149.57763671875,-148.766067504883,-148.447998046875,-148.317138671875,-148.339797973633,-148.430267333984,-148.433486938477,-148.296432495117,-148.129287719727,-148.082962036133,-148.176513671875,-148.41748046875,-149.051422119141,-150.490631103516,-151.048431396484,-151.368255615234,-151.636123657227,-151.903564453125,-152.091400146484,-152.053405761719,-152.1376953125,-152.243515014648,-152.701370239258,-153.517578125,-154.517730712891,-155.209915161133,-156.114547729492,-156.469329833984,-156.207916259766,-155.919784545898,-154.716400146484,-154.537643432617,-154.293014526367,-154.695068359375,-155.03662109375,-155.341506958008,-156.569229125977,-157.266799926758,-157.848052978516,-158.285888671875,-158.406936645508,-158.500381469727,-158.351425170898,-158.22998046875,-158.246475219727,-158.213577270508,-158.003082275391,-157.842041015625,-157.465377807617,-157.139312744141,-156.667663574219,-156.368209838867,-156.211227416992,-155.919586181641,-155.358840942383,-155.101745605469,-154.81494140625,-153.909973144531,-153.712600708008,-153.606109619141,-153.573043823242,-153.460601806641,-153.076950073242,-151.998382568359,-151.718994140625,-150.956405639648,-150.305572509766,-150.084320068359,-149.717712402344,-149.588485717773,-149.474014282227,-149.1259765625,-148.339950561523,-148.155715942383,-148.259811401367,-148.559280395508,-148.744384765625,-148.843612670898,-148.839019775391,-148.777496337891,-148.572402954102,-148.196334838867,-147.730224609375,-147.56640625,-147.442276000977,-147.207229614258,-146.927581787109,-146.390625,-146.073638916016,-145.677139282227,-145.600631713867,-145.649658203125,-145.713821411133,-145.794281005859,-145.807952880859,-145.634567260742,-145.515716552734,-145.563186645508,-145.753128051758,-145.864303588867,-145.966995239258,-145.933929443359,-145.806350708008,-145.629257202148,-145.685699462891,-145.675674438477,-145.75048828125,-146.166458129883,-146.776657104492,-147.340423583984,-148.60107421875,-149.045852661133,-149.339660644531,-149.654251098633,-149.284957885742,-148.894821166992,-148.780364990234,-148.631790161133,-148.458984375,-148.3203125,-147.860214233398,-146.817337036133,-146.597412109375,-145.8857421875,-145.686874389648,-145.442092895508,-145.642333984375,-145.860397338867,-146.382995605469,-146.323486328125,-145.987731933594,-145.105514526367,-144.721282958984,-144.220611572266,-143.574264526367,-143.022171020508,-142.329833984375,-142.094192504883,-141.505722045898,-141.134613037109,-141.008987426758,-140.874313354492,-141.223342895508,-140.998733520508,-140.709274291992,-140.470993041992,-140.293792724609,-139.691162109375,-139.148834228516,-137.6181640625,-137.090133666992,-136.649856567383,-136.549514770508,-136.4619140625,-136.227828979492,-136.030075073242,-135.362060546875,-134.840377807617,-134.465087890625,-134.117141723633,-133.796340942383,-133.474853515625,-132.991653442383,-132.351272583008,-132.049362182617,-131.70654296875,-130.857482910156,-130.195602416992,-129.790817260742,-129.23828125,-128.940628051758,-127.863380432129,-127.020210266113,-126.383987426758,-125.353424072266,-124.312446594238,-123.889457702637,-121.543937683105,-119.677001953125,-119.422210693359,-119.022415161133,-118.802932739258,-118.655769348145,-118.342041015625,-117.806198120117,-117.068305969238,-116.433006286621,-115.222610473633,-115.105178833008,-114.991012573242,-114.791015625,-114.623725891113,-114.345947265625,-113.508499145508,-113.489402770996,-113.574661254883,-113.713569641113,-113.753273010254,-113.640815734863,-113.454254150391,-113.332962036133,-113.597312927246,-113.783103942871,-113.903465270996,-113.984764099121,-114.097213745117,-114.110450744629,-113.931838989258,-113.752540588379,-113.593406677246,-113.091506958008,-112.170021057129,-111.868209838867,-111.696243286133,-111.584426879883,-111.738716125488,-111.788719177246,-111.695899963379,-111.722267150879,-111.806343078613,-111.629981994629,-111.466796875,-111.18017578125,-111.019828796387,-110.770408630371,-110.533935546875,-110.307075500488,-110.229789733887,-110.300437927246,-110.531936645508,-110.967575073242,-111.463134765625,-111.358787536621,-111.104202270508,-109.989990234375,-109.272163391113,-108.822265625,-108.254486083984,-107.804733276367,-107.266792297363,-106.93212890625,-106.618850708008,-105.399368286133,-104.90185546875,-104.617820739746,-104.15966796875,-103.901313781738,-103.424903869629,-103.121040344238,-102.771339416504,-101.708106994629,-101.627830505371,-101.3037109375,-101.039360046387,-100.706344604492,-100.463432312012,-100.082809448242,-99.5313491821289,-98.980224609375,-98.7523422241211,-98.6457061767578,-98.557861328125,-98.7272491455078,-99.2081527709961,-99.6519088745117,-99.8485794067383,-100.1640625,-100.31298828125,-100.473342895508,-100.264892578125,-100.0126953125,-100.118598937988,-100.238136291504,-100.530853271484,-100.8818359375,-101.023139953613,-101.251708984375,-101.3427734375,-101.586723327637,-101.715431213379,-102.105125427246,-102.440826416016,-102.766448974609,-102.862747192383,-102.799507141113,-102.41064453125,-102.03662109375,-101.828369140625,-101.58740234375,-101.310737609863,-101.130226135254,-100.985443115234,-100.7177734375,-99.7811050415039,-99.6561584472656,-99.541015625,-99.3433609008789,-99.1619186401367,-98.8961410522461,-99.2003402709961,-99.5279312133789,-100.020797729492,-100.436370849609,-101.189453125,-101.57373046875,-101.815963745117,-102.675048828125,-102.908790588379,-103.076171875,-103.307716369629,-103.375,-103.216606140137,-103.110107421875,-102.855857849121,-102.484771728516,-102.362892150879,-102.272018432617,-102.362937927246,-102.482032775879,-102.409271240234,-102.028861999512,-101.841499328613,-101.681205749512,-101.331832885742,-100.820503234863,-100.563575744629,-100.2587890625,-99.8107452392578,-98.2085952758789,-98.0124053955078,-97.8185043334961,-97.6510238647461,-97.4764633178711,-96.9557647705078,-96.6758270263672,-96.3942413330078,-96.1521453857422,-95.8805618286133,-95.5292434692383,-95.2366180419922,-95.0295867919922,-94.5864715576172,-94.2461929321289,-93.9846649169922,-93.7059555053711,-92.828369140625,-92.2410125732422,-91.1686553955078,-90.9209442138672,-90.430908203125,-90.2737731933594,-90.29541015625,-90.1524353027344,-90.0352020263672,-89.8176803588867,-89.5223617553711,-89.3412551879883,-89.2293930053711,-89.1271438598633,-88.7799835205078,-88.5269012451172,-88.194091796875,-88.1945343017578,-88.3317337036133,-88.5607452392578,-88.4193878173828,-88.2049789428711,-87.9363327026367,-87.6084442138672,-87.4010238647461,-87.0379409790039,-86.791015625,-86.6021499633789,-85.9807662963867,-85.8014144897461,-85.5821762084961,-85.2605972290039,-84.981201171875,-84.5712890625,-84.2141647338867,-83.7962951660156,-83.5648422241211,-83.0419006347656,-82.8152389526367,-82.1834945678711,-81.6061019897461,-81.3087387084961,-81.1631851196289,-81.2359924316406,-81.2623977661133,-81.1764144897461,-81.0243148803711,-80.3363800048828,-80.3798294067383,-80.4387664794922,-80.6142120361328,-80.5877456665039,-80.4422378540039,-80.1517562866211,-79.8080062866211,-79.521728515625,-78.9637222290039,-78.7862319946289,-78.4078598022461,-78.1441421508789,-77.8456039428711,-77.4440460205078,-77.1355438232422,-76.8504867553711,-76.7645034790039,-77.0330047607422,-77.1349105834961,-77.0489273071289,-76.8875045776367,-76.7549819946289,-76.2912139892578,-75.9162139892578,-75.5950164794922,-75.2930679321289,-75.0435562133789,-74.85546875,-74.5940399169922,-74.3452606201172,-74.1973114013672,-73.9960403442383,-72.92919921875,-72.6874084472656,-72.3808135986328,-71.9941864013672,-71.6975631713867,-71.4527359008789,-71.0171890258789,-70.3226547241211,-69.9686050415039,-69.2822265625,-68.8209457397461,-68.0003356933594,-67.6670837402344,-67.3067398071289,-67.0795440673828,-66.8277282714844,-66.95166015625,-67.0841293334961,-67.1957473754883,-67.4603500366211,-67.5299301147461,-67.5045928955078,-67.598388671875,-67.6921844482422,-67.8884735107422,-68.1256866455078,-68.4033203125,-68.4036636352539,-68.4698257446289,-68.6375503540039,-68.7079620361328,-68.5800323486328,-68.4615173339844,-68.140869140625,-67.3717803955078,-67.3043441772461,-67.1104507446289,-66.9748992919922,-67.021240234375,-67.1875991821289,-67.3905258178711,-67.2990188598633,-67.1336898803711,-67.0542984008789,-67.1168975830078,-67.041015625,-66.8935089111328,-66.7933654785156,-66.9775848388672,-67.1498489379883,-67.106689453125,-67.0212860107422,-66.9153823852539,-66.7698745727539,-66.67724609375,-66.7049789428711,-66.9231414794922,-67.1243209838867,-67.4869155883789,-67.5445327758789,-67.5647430419922,-67.5857925415039,-67.5503921508789,-67.4933624267578,-67.4404830932617,-67.2990188598633,-67.16015625,-67.0344772338867,-66.955078125,-66.9286117553711,-66.8861312866211,-66.9021453857422,-66.8360366821289,-66.75732421875,-66.6100158691406,-66.5515594482422,-66.4986801147461,-66.4722137451172,-66.490966796875,-66.51513671875,-66.5333023071289,-66.5021057128906,-66.4646911621094,-66.5268096923828,-66.5036087036133,-66.3709030151367,-66.3065414428711,-66.181884765625,-65.9537582397461,-65.8474655151367,-65.7664108276367,-65.7179718017578,-65.678466796875,-65.7757797241211,-65.7745132446289,-65.7174758911133,-65.6172866821289,-65.465087890625,-65.3163528442383,-65.1720199584961,-65.2220687866211,-65.2674789428711,-65.1050796508789,-64.9987335205078,-64.7216796875,-64.613525390625,-64.5143051147461,-64.54736328125,-64.6532287597656,-64.6730499267578,-64.6465759277344,-64.474609375,-64.4353485107422,-64.3900375366211,-64.4168014526367,-64.4389114379883,-64.2134780883789,-64.1799850463867,-64.1322250366211,-64.06591796875,-63.8622055053711,-63.8181190490723,-63.7979507446289,-63.9080123901367,-64.05126953125,-64.0710983276367,-64.0380401611328,-63.9124031066895,-63.76025390625,-63.4821281433105,-63.2641639709473,-63.1781234741211,-63.0590782165527,-63.0326194763184,-63.0856971740723,-63.1198768615723,-62.774658203125,-62.6645011901855,-62.5275421142578,-62.5762176513672,-62.5034637451172,-62.4042472839355,-62.3380851745605,-62.2433090209961,-62.1396484375,-61.8825187683105,-61.7563972473145,-61.6317901611328,-61.5004844665527,-61.4700241088867,-61.3959503173828,-61.173583984375,-61.0821304321289,-60.8866233825684,-60.9220695495605,-60.8641586303711,-60.2772445678711,-59.9898452758789,-59.5101547241211,-59.2175750732422,-59.0364265441895,-58.8720703125,-58.6735382080078,-58.2155723571777,-57.8680686950684,-57.3896484375,-57.1682586669922,-57.0767097473145,-57.0206527709961,-56.9273414611816,-56.7818336486816,-56.8347663879395,-56.9736824035645,-57.1191902160645,-57.1522483825684,-57.0977058410645,-57.1522483825684,-57.2838897705078,-57.4606475830078,-57.5814933776855,-57.7369155883789,-57.8567352294922,-58.2629890441895,-58.5318870544434,-58.7228507995605,-58.8389663696289,-59.0053215026855,-59.0473136901855,-58.9772491455078,-58.9229965209961,-58.79931640625,-58.8191452026367,-58.9051284790039,-58.8954582214355,-58.805908203125,-58.7860870361328,-58.8918914794922,-59.0506820678711,-59.2293930053711,-59.3696784973145,-59.4607429504395,-59.5467758178711,-59.6121559143066,-59.5731887817383,-59.6459922790527,-59.734375,-59.7650375366211,-59.8501968383789,-59.9630851745605,-60.2424812316895,-60.3405265808105,-60.3936538696289,-60.5559577941895,-60.6599617004395,-60.9153823852539,-61.0598640441895,-61.3318328857422,-61.4394493103027,-61.5030250549316,-61.6032257080078,-61.7031211853027,-61.7361793518066,-61.5774421691895,-61.6634254455566,-61.8556137084961,-61.9478492736816,-62.0245094299316,-62.0846710205078,-62.1453094482422,-62.0536613464355,-61.9033699035645,-61.7957077026367,-61.7560081481934,-61.9914054870605,-62.1505889892578,-62.2224159240723,-62.3050270080566,-62.2932662963867,-62.1691398620605,-62.0050315856934,-61.8390655517578,-61.6248016357422,-61.57470703125,-61.3591346740723,-61.26611328125,-61.1981468200684,-61.1375961303711,-61.0392570495605,-60.9881324768066,-60.9127960205078,-60.8129844665527,-60.6183128356934,-60.5653839111328,-60.6249008178711,-60.7439918518066,-60.8564453125,-60.9556617736816,-61.00927734375,-60.9027328491211,-60.9424324035645,-61.0284156799316,-61.13427734375,-61.1497535705566,-61.29296875,-61.4319381713867,-61.5261268615723,-61.6756324768066,-61.6964836120605,-61.7560081481934,-61.8419952392578,-61.8754425048828,-62.1165046691895,-62.2412605285645,-62.494140625,-62.5828094482422,-62.6820335388184,-62.7548370361328,-62.6502914428711,-62.6153831481934,-62.6179161071777,-62.6375465393066,-62.6550788879395,-62.5431594848633,-62.5365219116211,-62.62890625,-62.7047843933105,-62.9967269897461,-63.1795425415039,-63.2575187683105,-63.4485359191895,-63.5866241455078,-63.7525405883789,-63.6875457763672,-63.6543922424316,-63.7556648254395,-63.8804244995117,-63.9643516540527,-64.0150375366211,-64.0777359008789,-63.8087882995605,-63.76904296875,-63.7547340393066,-63.839599609375,-64.0425796508789,-64.4009780883789,-64.5540008544922,-64.60693359375,-64.686279296875,-64.7354507446289,-64.7933578491211,-64.8781204223633,-64.8541030883789,-64.7854995727539,-64.8387222290039,-64.9508743286133,-65.0269012451172,-64.8582534790039,-64.8264694213867,-64.8192901611328,-65.07958984375,-65.24853515625,-65.35009765625,-65.443115234375,-65.5031280517578,-65.5234375,-65.50390625,-65.4708023071289,-65.44677734375,-65.4180679321289,-65.5740203857422,-65.5892562866211,-65.6000442504883,-65.52783203125,-65.4694290161133,-65.5511703491211,-65.6395034790039,-65.5462417602539,-65.3874969482422,-65.218017578125,-64.9588317871094,-64.8847122192383,-64.8534622192383,-64.8294906616211,-64.8959503173828,-65.3651809692383,-65.4520034790039,-65.3313980102539,-65.0897521972656,-64.9964828491211,-65.0545425415039,-65.1400909423828,-65.2416076660156,-65.1583480834961,-64.8982925415039,-64.4289093017578,-64.0784683227539,-64.1568374633789,-64.1692352294922,-63.9244651794434,-63.7964859008789,-63.2166023254395,-63.0565452575684,-62.9333000183105,-62.9796867370605,-63.11474609375,-63.3475112915039,-63.7073249816895,-63.7734336853027,-63.7470245361328,-63.4427261352539,-63.3435096740723,-63.4782218933105,-63.4559555053711,-63.3014640808105,-63.0943832397461,-62.9940910339355,-62.8397483825684,-62.5868110656738,-62.4505386352539,-62.4071311950684,-62.2024421691895,-61.9610862731934,-61.9346199035645,-62.0139656066895,-62.2178726196289,-62.3314933776855,-62.3777809143066,-62.2322769165039,-62.0007820129395,-61.5046844482422,-61.4914093017578,-61.6053199768066,-61.6964836120605,-61.8089370727539,-61.9941444396973,-62.0404319763184,-61.9610862731934,-61.7021446228027,-61.5133819580078,-61.3128395080566,-61.2516593933105,-61.0172348022461,-60.9622535705566,-61.0030746459961,-61.1484375,-61.2373046875,-61.3692855834961,-61.5159225463867,-61.78955078125,-61.9095726013184,-61.9587898254395,-61.7254371643066,-61.5627899169922,-61.2135772705078,-61.0813522338867,-60.9953117370605,-60.9490242004395,-61.0350570678711,-61.6445350646973,-61.9389152526855,-62.2566413879395,-61.8940391540527,-61.6280250549316,-61.4926795959473,-61.3102569580078,-61.1074676513672,-60.9517593383789,-60.8332023620605,-60.7194328308105,-60.7042961120605,-60.6910667419434,-60.6646003723145,-60.7303199768066,-61.0475082397461,-61.2797889709473,-61.2861328125,-60.9391593933105,-60.7241172790527,-60.5323257446289,-60.5323257446289,-60.3846702575684,-60.2544937133789,-60.148681640625,-60.0097618103027,-59.9568367004395,-60.0164070129395,-60.1222152709961,-60.4037628173828,-60.5606918334961,-60.6866188049316,-60.8958511352539,-61.0813522338867,-61.2420921325684,-61.4284210205078,-61.7264137268066,-62.00830078125,-61.9147415161133,-61.78759765625,-61.7376976013184,-61.6369667053223,-61.4054718017578,-61.0797843933105,-60.8788566589355,-60.790283203125,-60.9027328491211,-61.0882835388184,-61.2034187316895,-61.4040489196777,-61.5454063415527,-61.6916961669922,-61.7416038513184,-61.8382301330566,-61.3194313049316,-61.1606903076172,-61.0416526794434,-61.2268524169922,-61.57080078125,-61.7182579040527,-61.8427734375,-61.331787109375,-61.1206016540527,-60.78369140625,-60.7042961120605,-60.8384742736816,-61.0107879638672,-61.3701667785645,-61.6399917602539,-61.9945335388184,-62.0888633728027,-62.2353057861328,-62.2256889343262,-62.1327133178711,-61.8944358825684,-61.855224609375,-61.9280281066895,-62.1377944946289,-62.3724594116211,-62.5667953491211,-62.7084999084473,-62.8871078491211,-63.0723152160645,-63.1781234741211,-63.1679191589355,-63.125244140625,-63.197998046875,-63.3570327758789,-63.5587882995605,-63.7508811950684,-63.9247093200684,-63.5709953308105,-63.3369102478027,-63.1731910705566,-63.2310562133789,-63.5514144897461,-63.8575210571289,-64.279541015625,-63.9724617004395,-63.6784210205078,-63.474853515625,-63.3038101196289,-63.2575187683105,-63.3633766174316,-64.0526428222656,-64.7782745361328,-65.0443801879883,-65.3217315673828,-65.9656753540039,-66.3704071044922,-67.5182189941406,-69.3043899536133,-69.915283203125,-70.0955047607422,-70.2101593017578,-70.55078125,-70.89501953125,-71.7986831665039,-72.7223129272461,-73.4717788696289,-73.8797836303711,-75.2683563232422,-75.4435043334961,-75.6592788696289,-75.8313522338867,-75.9628448486328,-76.2441940307617,-77.1900405883789,-77.2870635986328,-77.16796875,-76.8235855102539,-76.2485809326172,-75.937255859375,-75.7481460571289,-75.3869171142578,-74.5806198120117,-73.4782257080078,-72.8519515991211,-72.8751449584961,-73.2515640258789,-73.4850082397461,-74.0421371459961,-74.8120574951172,-75.3984375,-76.4378433227539,-77.7421417236328,-79.6790084838867,-80.1041030883789,-80.6015625,-80.8885269165039,-81.1031265258789,-81.5809555053711,-81.4410171508789,-79.5096664428711,-77.8581085205078,-77.664794921875,-77.4325714111328,-77.4524383544922,-77.5450210571289,-77.869140625,-78.7116165161133,-79.7665557861328,-80.2922821044922,-80.8163070678711,-81.9292907714844,-82.6084442138672,-83.08251953125,-83.4120559692383,-83.7790069580078,-83.7520980834961,-83.687744140625,-83.5082473754883,-83.2459030151367,-83.2269515991211,-83.35498046875,-83.54443359375,-83.7059097290039,-83.7627410888672,-83.6966323852539,-83.5951614379883,-83.260009765625,-82.9707565307617,-82.5892028808594,-81.660888671875,-81.5029296875,-81.22216796875,-81.1631851196289,-80.8919906616211,-80.7047882080078,-80.5348129272461,-80.478759765625,-80.488525390625,-80.415771484375,-80.1511688232422,-79.4556655883789,-76.4991226196289,-76.2176742553711,-76.1051330566406,-76.0315856933594,-76.3439483642578,-76.557861328125,-76.9039993286133,-77.2222595214844,-77.7018585205078,-78.6922378540039,-79.660400390625,-78.9071273803711,-78.1760711669922,-77.1604461669922,-76.7571258544922,-76.4073257446289,-76.2596206665039,-75.9856414794922,-75.8224182128906,-75.7090301513672,-75.5550231933594,-75.4945297241211,-75.3445816040039,-75.2365188598633,-75.0755920410156,-74.8065948486328,-74.5111312866211,-73.9378433227539,-73.3833465576172,-73.0294876098633,-72.55322265625,-72.173583984375,-71.3800354003906,-71.2306594848633,-71.0176773071289,-70.6878890991211,-70.56005859375,-70.3924331665039,-70.2391128540039,-70.012451171875,-69.7722702026367,-69.6339797973633,-69.1815948486328,-68.58984375,-68.3265609741211,-68.2846145629883,-68.1437530517578,-67.9654312133789,-65.5736846923828,-64.7501449584961,-63.4777374267578,-62.490234375,-62.3537139892578,-62.1653823852539,-62.5418434143066,-62.9458961486816,-63.553955078125,-63.7686538696289,-64.232666015625,-64.4756774902344,-64.6960906982422,-65.0215835571289,-65.6197204589844,-65.4866180419922,-65.2637710571289,-64.810546875,-64.190185546875,-64.1371078491211,-64.7061538696289,-65.9162063598633,-66.0422897338867,-66.1338424682617,-65.9530334472656,-65.8438491821289,-65.78662109375,-65.9131851196289,-65.7139663696289,-65.5719299316406,-65.4244155883789,-65.1701202392578,-64.9195785522461,-64.3965835571289,-63.7728538513184,-63.46630859375,-62.6453132629395,-61.9016647338867,-60.8590812683105,-60.6871109008789,-60.5277366638184,-60.8171844482422,-62.0945320129395,-62.5530242919922,-62.7356414794922,-62.6309089660645,-62.465576171875,-62.1286163330078,-61.9169883728027,-61.7089347839355,-61.3128395080566,-61.2184104919434,-61.2003936767578,-61.30322265625,-61.4363288879395,-61.5305633544922,-61.58984375,-61.42529296875,-60.9832000732422,-60.3970184326172,-59.8538093566895,-59.5160140991211,-58.2899398803711,-57.7977561950684,-57.55712890625,-57.3536109924316,-56.31787109375,-56.0750465393066,-55.8006820678711,-55.294677734375,-54.6011238098145,-53.986083984375,-53.7396011352539,-53.5575675964355,-53.3390617370605,-52.7988739013672,-52.4149398803711,-51.7306632995605,-51.2096672058105,-50.6530265808105,-50.0292472839355,-48.3607940673828,-47.8868141174316,-47.3602561950684,-47.0198707580566,-46.5666999816895,-46.258056640625,-46.119140625,-46.04638671875,-46.1985359191895,-46.4483375549316,-46.5167465209961,-46.17529296875,-45.78857421875,-45.0437469482422,-44.4548835754395,-44.2917976379395,-44.064208984375,-43.6693382263184,-43.1803703308105,-42.5645523071289,-42.0462875366211,-41.7115707397461,-41.433837890625,-41.1258773803711,-40.91455078125,-40.4408187866211,-39.7623062133789,-38.771728515625,-38.0109367370605,-37.2092742919922,-36.8124046325684,-36.4995079040527,-36.2339820861816,-35.9657707214355,-35.77587890625,-35.5205574035645,-35.3270034790039,-34.3499488830566,-33.3287124633789,-33.1913070678711,-33.0572280883789,-32.7061996459961,-32.2557106018066,-31.6342296600342,-31.3121070861816,-31.0154304504395,-30.42529296875,-29.79736328125,-29.531494140625,-29.3291015625,-24.2402839660645,-24.0198230743408,-23.574462890625,-23.4068355560303,-24.0882816314697,-24.2998523712158,-24.5338859558105,-24.67041015625,-25.2586421966553,-29.9493179321289,-30.049072265625,-30.2112312316895,-30.1779308319092,-30.31591796875,-30.6452617645264,-30.9851570129395,-31.4127922058105,-32.5418472290039,-32.9942398071289,-34.1973648071289,-34.9949226379395,-35.5159645080566,-35.8900871276855,-36.2391586303711,-36.265625,-36.1808586120605,-35.50927734375,-35.0875968933105,-34.808349609375,-34.5514640808105,-34.2901878356934,-34.0757827758789,-33.5911598205566,-33.3767585754395,-32.6140594482422,-32.4052734375,-32.0633811950684,-31.67578125,-30.4892101287842,-30.2219715118408,-29.8915519714355,-28.9336452484131,-28.0793933868408,-27.653076171875,-27.134521484375,-26.56005859375,-26.059326171875,-24.2695789337158,-23.197265625,-22.4654788970947,-21.9480953216553,-21.4337902069092,-20.989013671875,-20.7833003997803,-20.4875011444092,-19.4930191040039,-18.8509273529053,-18.5851573944092,-18.3045902252197,-18.4151363372803,-18.5169429779053,-18.6172847747803,-18.7492198944092,-18.6172847747803,-18.5169429779053,-18.22119140625,-18.0682621002197,-17.9227542877197,-17.4358386993408,-17.2990226745605,-16.9892597198486,-16.7270984649658,-16.4295425415039,-15.6725091934204,-15.5312509536743,-15.2897453308105,-15.0891599655151,-14.6589345932007,-14.5738286972046,-14.6114253997803,-15.2596197128296,-15.7488279342651,-16.2201156616211,-16.2818851470947,-16.1808605194092,-16.0031242370605,-15.9356441497803,-16.0974597930908,-16.3877449035645,-16.5188484191895,-16.5070323944092,-16.4352035522461,-16.2791023254395,-16.1490230560303,-15.8028326034546,-15.5959959030151,-15.0070323944092,-14.3209962844849,-14.1646976470947,-14.0000972747803,-14.1683101654053,-14.2982416152954,-14.2977533340454,-13.93896484375,-13.6028318405151,-13.2085933685303,-12.7469244003296,-12.0947265625,-11.77734375,-11.4969730377197,-11.3464841842651,-11.1213865280151,-10.9581050872803,-10.9610347747803,-11.0092287063599,-11.1793451309204,-11.3330564498901,-11.6968746185303,-12.148193359375,-12.2845211029053,-12.351318359375,-12.2078123092651,-12.0736818313599,-11.9261226654053,-11.6630373001099,-11.3280763626099,-11.16015625,-10.9698238372803,-10.825439453125,-10.6594724655151,-10.5200681686401,-10.4066400527954,-10.2305660247803,-10.0334959030151,-10.122314453125,-10.3310060501099,-10.3599605560303,-10.2706050872803,-10.0987310409546,-9.88798809051514,-9.599365234375,-9.40234375,-9.23066425323486,-8.96591758728027,-8.646484375,-8.49770450592041,-8.21645450592041,-7.91582059860229,-7.71372032165527,-7.66899442672729,-7.59013652801514,-7.61796855926514,-7.75688457489014,-7.87348604202271,-7.85493183135986,-7.75273418426514,-7.61977529525757,-7.38813447952271,-7.03159189224243,-6.83818340301514,-6.54750919342041,-6.24521446228027,-5.93632793426514,-5.69472646713257,-5.587890625,-5.70869112014771,-5.90380859375,-6.08027315139771,-6.12675809860229,-6.11748075485229,-5.95004892349243,-4.45014667510986,-4.25322294235229,-3.99482440948486,-3.71318364143372,-3.23964834213257,-2.81201148033142,-2.61025381088257,-2.26132822036743,-2.01459980010986,-1.50063490867615,-1.35424816608429,-1.21635735034943,-1.06777334213257,-0.895849823951721,-0.840087950229645,-0.759863495826721,-0.543163955211639,-0.326953202486038,-0.184570327401161,0,0.154198989272118,0.538476645946503,0.83496105670929,1.55224645137787,1.90869140625,2.60947275161743,3.50693368911743,5.11308574676514,5.64394521713257,6.50800752639771,6.95097684860229,7.40117168426514,7.6767578125,8.30673789978027,8.52304649353027,8.81748008728027,9.1416015625,9.61347675323486,9.88554668426514,10.2176752090454,10.9688472747803,11.2035160064697,11.7012691497803,11.8335943222046,12.0679683685303,12.3087892532349,12.4616212844849,12.6819343566895,12.9293947219849,12.8645515441895,12.7234373092651,12.5951175689697,12.6262693405151,13.0656251907349,13.2979497909546,13.5326166152954,13.8226556777954,14.4917964935303,15.0638675689697,15.5628900527954,15.8069343566895,16.0251941680908,16.3810539245605,16.5848636627197,16.7091808319092,17.1666984558105,18.1246089935303,18.2320308685303,18.3513679504395,18.4326171875,18.6273422241211,18.8772468566895,19.0093746185303,19.1963863372803,19.13232421875,19.0265617370605,19.1529293060303,19.26513671875,19.4092769622803,19.65185546875,19.9442386627197,20.1281261444092,21.07080078125,21.18603515625,21.3373050689697,21.7049808502197,21.8489246368408,21.9623050689697,22.2158203125,22.3660163879395,22.396484375,22.2336921691895,22.27783203125,22.4454097747803,22.97900390625,23.14990234375,23.4068355560303,23.6648426055908,23.8036136627197,24.0241222381592,24.2357425689697,24.3857421875,24.3857421875,24.58837890625,24.7567386627197,25.1874008178711,25.6501941680908,25.97412109375,26.4988288879395,26.75439453125,26.91796875,27.2068367004395,27.5085945129395,27.69775390625,28.3864250183105,28.9115257263184,29.4638690948486,30.0033206939697,30.8340835571289,31.0628910064697,31.3788070678711,32.1595687866211,32.45654296875,32.6212882995605,32.8097648620605,32.9115219116211,32.9893569946289,32.9759750366211,32.9031257629395,32.7379875183105,32.5675773620605,32.6416015625,32.7761726379395,33.1214866638184,33.4656257629395,33.853515625,34.19287109375,34.2193374633789,34.07421875,33.8848609924316,33.7720718383789,33.8136711120605,34.05859375,34.5958976745605,34.74951171875,35.13134765625,35.2248039245605,35.3570327758789,35.5676765441895,36.0177726745605,36.3311538696289,36.5859375,36.71875,36.8557624816895,37.1148452758789,37.37451171875,37.5597686767578,37.787109375,38.1443367004395,38.4994125366211,38.8855476379395,38.9117164611816,38.8592796325684,39.0187492370605,39.2113265991211,39.4870109558105,39.705078125,39.7623062133789,39.8638648986816,40.0417022705078,40.2156257629395,40.48388671875,40.8170890808105,41.1327133178711,41.3563499450684,41.8246078491211,42.4087867736816,42.8195304870605,42.9609375,43.1708984375,43.5541000366211,44.1775398254395,44.3728523254395,44.6998023986816,44.9895515441895,45.1969718933105,45.5693359375,45.8876953125,46.1539077758789,46.3990211486816,46.4365234375,46.3197288513184,46.3172836303711,46.4541015625,46.5596694946289,46.8838882446289,47.1543922424316,47.3515625,47.4029312133789,47.2313499450684,47.1171875,47.3141632080078,47.4898452758789,47.7035179138184,47.9585914611816,48.2099609375,48.3216781616211,48.3216781616211,48.37451171875,48.5509757995605,48.6480445861816,48.6200180053711,48.63037109375,49.0529289245605,49.2193374633789,48.9230461120605,48.7136688232422,48.5984382629395,48.4652328491211,48.8302726745605,49.2470703125,49.4886703491211,50.0061492919922,50.29296875,50.5530281066895,50.60595703125,50.5088882446289,50.5208969116211,50.3060531616211,50.2443351745605,50.3324203491211,50.5882797241211,50.9369125366211,51.6875,51.8845672607422,52.3782196044922,52.9552726745605,53.6717758178711,54.9478492736816,55.2904281616211,55.5044898986816,55.7103538513184,55.9740219116211,56.3615226745605,56.859375,57.0002937316895,57.1854476928711,56.9865226745605,56.8236312866211,56.5100593566895,56.2945327758789,56.1459007263184,56.2918968200684,56.4532203674316,56.4796867370605,56.3914070129395,55.802734375,56.1548805236816,56.3659172058105,56.5621070861816,56.7600593566895,56.8915977478027,57.3611297607422,57.62744140625,57.828125,58.0267601013184,58.3174819946289,58.7374000549316,59.2507858276367,59.6501922607422,59.8675765991211,60.4820289611816,61.0121040344238,61.30908203125,62.1739273071289,62.6878929138184,63.0176734924316,63.2375984191895,63.6990203857422,63.9312477111816,64.5736312866211,65.70751953125,66.4883804321289,67.1748046875,67.50244140625,68.0985336303711,68.3279266357422,68.8995132446289,69.1671905517578,69.4164047241211,69.5591812133789,69.6559524536133,69.60302734375,69.7044906616211,69.7886734008789,69.9074249267578,69.9822311401367,69.9279251098633,69.7619094848633,69.5341796875,69.546875,69.6456069946289,69.53076171875,69.6146469116211,69.6294860839844,69.5494155883789,69.3718719482422,69.06494140625,68.90625,68.8797836303711,68.9591827392578,69.1355438232422,69.1884765625,69.1620101928711,69.0826187133789,68.9205093383789,68.7437515258789,68.4152297973633,68.1781234741211,68.0271453857422,67.9169921875,67.5753936767578,67.4166030883789,67.2679748535156,67.6589889526367,67.9408187866211,68.5593795776367,68.7574234008789,69.0208969116211,69.1620101928711,69.2501983642578,69.1966781616211,69.1883850097656,68.8727493286133,68.7679672241211,68.6237335205078,68.4475555419922,68.310546875,68.0373992919922,67.8733367919922,67.6935577392578,67.4322204589844,67.2810516357422,67.21484375,67.1133804321289,66.89208984375,66.7464904785156,66.4976577758789,66.5691375732422,66.7647476196289,67.0031280517578,67.3220748901367,67.7486343383789,67.9713821411133,68.0155258178711,67.9713821411133,68.1068344116211,68.4198226928711,69.1570281982422,69.3094711303711,69.5546875,69.7697296142578,69.9621124267578,70.2943344116211,70.5728530883789,70.6162109375,70.7316436767578,71.0788040161133,71.1678695678711,71.2767639160156,71.3492202758789,71.3788070678711,71.46484375,71.6338806152344,71.7713851928711,71.9048843383789,72.2625961303711,72.41796875,72.6223602294922,72.7603530883789,72.7443389892578,72.8220672607422,73.0414047241211,73.3248062133789,73.6760787963867,73.9421844482422,74.2267608642578,74.5710983276367,75.1478500366211,75.423828125,75.6355514526367,75.8207015991211,75.8912048339844,76.1117248535156,76.3597640991211,76.7701187133789,77.1919937133789,77.5409164428711,77.8174819946289,78.01513671875,78.228515625,78.4889602661133,78.5634765625,78.7262725830078,79.03515625,79.2877960205078,80.3630828857422,81.1874084472656,82.0169906616211,82.2732391357422,82.6070327758789,83.1578140258789,83.3042984008789,83.49365234375,83.9037094116211,84.1607437133789,84.4851608276367,84.7483367919922,85.1167984008789,85.4290008544922,85.7107391357422,86.1183547973633,86.7501907348633,86.9465789794922,87.0848617553711,87.9802780151367,88.3141632080078,88.7894515991211,89.0765609741211,89.3517532348633,89.6984329223633,90.29296875,90.5472640991211,91.0216827392578,91.5460891723633,91.7770538330078,92.0734329223633,92.3123016357422,92.48583984375,92.5919876098633,92.7305679321289,93.0749053955078,93.3580093383789,93.7216796875,93.9642562866211,94.0887756347656,94.3134765625,94.5868148803711,94.83984375,95.083984375,95.2479476928711,95.541015625,95.9914016723633,96.4237289428711,96.7888717651367,97.1005859375,97.3884811401367,97.7198257446289,98.2576217651367,98.4617156982422,98.6031265258789,98.7201156616211,98.85888671875,99.3701171875,99.8243179321289,100.211715698242,100.591209411621,100.889068603516,101.3271484375,101.320899963379,101.38134765625,101.474411010742,102.174118041992,102.392189025879,102.674217224121,103.166595458984,103.638771057129,103.763473510742,103.951171875,104.2890625,104.6669921875,105.000389099121,106.386917114258,107.1708984375,107.565528869629,107.667282104492,107.785057067871,107.99169921875,108.157913208008,108.376167297363,108.910057067871,109.462791442871,109.82373046875,110.43701171875,110.622261047363,110.587013244629,110.90673828125,111.453125,112.130271911621,112.5478515625,113.099418640137,113.367973327637,113.502151489258,113.709770202637,113.954490661621,114.3369140625,114.618553161621,114.869537353516,115.082420349121,115.310356140137,115.635345458984,115.441886901855,115.273727416992,114.570602416992,114.259765625,113.9912109375,113.912498474121,114.026565551758,114.319038391113,114.657905578613,114.925788879395,115.171882629395,115.384178161621,115.88525390625,116.214645385742,116.509078979492,116.713478088379,116.923629760742,117.131935119629,117.297859191895,117.744728088379,117.951957702637,118.138664245605,118.325790405273,118.518943786621,118.7138671875,118.964645385742,119.318252563477,119.767967224121,120.187301635742,120.289260864258,120.374801635742,119.953704833984,119.280662536621,118.921676635742,119.133003234863,120.400390625,120.978713989258,121.487594604492,121.613182067871,122.033103942871,122.182807922363,122.633010864258,123.2216796875,123.666595458984,123.969337463379,124.196182250977,124.370506286621,124.597854614258,124.821586608887,125.095123291016,125.286323547363,125.397941589355,125.60302734375,125.865615844727,126.0771484375,126.423728942871,126.664749145508,126.873634338379,127.365432739258,127.541213989258,127.968063354492,128.430557250977,128.627822875977,128.81640625,128.982421875,129.236923217773,129.5,129.7412109375,129.975784301758,130.120513916016,130.300582885742,130.578521728516,130.951766967773,131.232025146484,131.830856323242,132.3203125,132.874328613281,133.148239135742,133.444534301758,133.8427734375,133.95751953125,134.178909301758,134.231842041016,134.289459228516,134.398132324219,134.769729614258,134.971496582031,135.351943969727,135.554794311523,136.009384155273,136.193939208984,136.553329467773,136.739654541016,136.88916015625,137.336227416992,137.753814697266,137.92578125,138.139953613281,138.27099609375,138.37646484375,139.241607666016,139.613174438477,139.900100708008,140.901550292969,141.285934448242,141.517196655273,141.972839355469,142.158981323242,142.32666015625,142.6875,142.888381958008,143.1689453125,143.4482421875,143.730377197266,143.862686157227,143.9111328125,144.11767578125,144.347854614258,144.550582885742,144.621200561523,144.515335083008,144.259674072266,144.153717041016,143.941986083984,143.977340698242,144.189056396484,144.404296875,144.879104614258,145.1279296875,145.556442260742,145.975204467773,146.2763671875,146.827819824219,146.852447509766,146.896575927734,146.878219604492,146.797668457031,147.093658447266,147.353805541992,147.568557739258,148.456253051758,148.880569458008,149.2626953125,149.716903686523,150.065521240234,150.342193603516,150.671981811523,150.935943603516,151.068267822266,151.12109375,151.138870239258,151.288772583008,151.447555541992,151.562103271484,152.265228271484,152.545516967773,152.814163208008,153.081832885742,153.339950561523,153.495697021484,153.705276489258,153.766998291016,153.792373657227,153.78466796875,153.908020019531,154.031539916992,154.19970703125,154.576354980469,154.866012573242,154.987503051758,155.163955688477,155.520309448242,156.010925292969,156.488677978516,157.04638671875,157.481353759766,157.775787353516,157.932525634766,158.157806396484,158.4326171875,158.647171020508,159.386337280273,159.783981323242,159.930953979492,160.125778198242,160.125778198242,160.209671020508,160.65185546875,160.826766967773,161.037002563477,161.424514770508,161.625106811523,161.916107177734,162.189453125,162.277740478516,162.286529541016,162.039657592773,162.02197265625,162.216003417969,162.6748046875,163.026458740234,163.348724365234,163.566497802734,163.998443603516,164.403228759766,164.716003417969,165.209381103516,165.853897094727,166.132034301758,166.626953125,167.228805541992,167.569427490234,167.640045166016,167.798828125,167.878128051758,167.96630859375,168.172653198242,168.3828125,168.7978515625,169.663772583008,169.976943969727,170.162307739258,170.25048828125,170.27685546875,170.435745239258,170.603317260742,170.779693603516,170.859085083008,170.675384521484,170.409271240234,170.224029541016,170.030075073242,169.953521728516,170.127151489258,170.259384155273,170.285842895508,170.206436157227,170.047744750977,169.77490234375,169.440322875977,169.072357177734,168.718841552734,168.576171875,168.428421020508,168.621673583984,168.820022583008,169.26953125,169.82861328125,169.844818115234,169.712112426758,169.545013427734,169.033401489258,168.735946655273,168.38134765625,168.204483032227,167.85302734375,167.155670166016,166.8828125,166.452835083008,166.466995239258,166.833984375,167.2255859375,167.615814208984,167.709075927734,167.5341796875,167.296478271484,166.99609375,166.428817749023,166.159378051758,166.001068115234,165.860168457031,165.970611572266,166.106048583984,165.913284301758,165.733688354492,165.548919677734,165.346969604492,165.249893188477,165.244522094727,165.129501342773,165.0048828125,164.81298828125,164.749618530273,164.8876953125,164.979888916016,164.905960083008,164.775680541992,165.037200927734,165.263076782227,165.399810791016,165.408584594727,165.302825927734,165.001174926758,164.85302734375,164.68896484375,164.410736083984,164.174026489258,163.935836791992,163.735244750977,163.556549072266,163.397857666016,163.265518188477,163.167388916016,162.961227416992,162.752059936523,162.604095458984,162.533599853516,162.410049438477,162.2255859375,162.087799072266,161.910354614258,161.679595947266,160.910751342773,161.032913208008,161.227340698242,161.903518676758,162.189651489258,162.239074707031,162.351959228516,162.57763671875,162.754013061523,162.815719604492,162.7451171875,162.648239135742,162.4365234375,162.498245239258,162.7275390625,162.824600219727,162.674621582031,162.745208740234,162.762786865234,162.609558105469,162.489456176758,162.450302124023,162.679092407227,162.850189208984,163.0869140625,163.249908447266,163.45849609375,163.607620239258,163.619140625,163.766220092773,164.045211791992,164.036437988281,164.232040405273,164.4208984375,164.491500854492,164.4296875,164.107803344727,163.977630615234,164.29736328125,164.628128051758,165.050598144531,165.274017333984,165.417572021484,165.524017333984,165.5341796875,165.662994384766,166.208602905273,166.510360717773,166.801162719727,167.058013916016,167.130279541016,167.049026489258,166.850006103516,166.524612426758,166.286529541016,166.116806030273,164.634765625,164.300582885742,163.901748657227,163.503128051758,162.895126342773,162.639450073242,161.974502563477,161.757415771484,161.669235229492,161.510452270508,161.501770019531,161.813385009766,161.951461791992,161.951461791992,161.8642578125,161.7373046875,161.546096801758,161.190521240234,160.87353515625,160.482711791992,160.5673828125,160.670211791992,160.646087646484,160.209182739258,160.067962646484,159.975891113281,160.083602905273,160.32275390625,160.346282958984,160.264846801758,159.871871948242,159.896484375,160.111419677734,160.387313842773,160.558013916016,160.558792114258,160.381729125977,160.179397583008,158.767471313477,158.560348510742,158.573638916016,159.065139770508,160.542190551758,160.637313842773,160.603317260742,160.521286010742,160.601364135742,160.8232421875,160.830276489258,160.720016479492,160.502532958984,160.260162353516,160.275787353516,160.607223510742,160.716796875,160.727935791016,160.716705322266,160.5400390625,160.478607177734,160.469833374023,160.907806396484,161.582122802734,161.730178833008,161.99609375,162.42529296875,162.57666015625,162.821197509766,162.854400634766,163.004287719727,163.602340698242,163.680068969727,162.426559448242,161.16650390625,161.283203125,162.643753051758,163.011825561523,163.174713134766,163.2685546875,164.001373291016,164.747161865234,164.980072021484,165.981643676758,166.445907592773,166.7421875,166.956741333008,167.116317749023,167.271286010742,167.232620239258,167.297470092773,167.404495239258,167.601959228516,167.827545166016,168.091796875,168.275970458984,168.607315063477,168.481643676758,168.408401489258,168.320022583008,168.240234375,167.973052978516,167.825378417969,167.674407958984,167.657623291016,167.8427734375,168.110046386719,169.837890625,170.33203125,170.817489624023,171.035736083984,171.220703125,171.289260864258,171.537109375,171.917190551758,172.450088500977,172.873931884766,173.397262573242,173.661819458008,173.822357177734,175.011032104492,175.187393188477,175.322860717773,175.605758666992,175.911239624023,177.5810546875,178.208602905273,178.352630615234,178.496002197266,178.944427490234,179.403030395508,179.620315551758,-180],[-55.5280265808105,-55.4662590026855,-55.2155265808105,-55.15625,-55.1064453125,-55.0751953125,-55.1567878723145,-55.5936508178711,-55.7501983642578,-55.8304672241211,-56.0091285705566,-56.0830078125,-56.3785171508789,-56.4628410339355,-56.4990234375,-56.5053215026855,-56.475341796875,-56.4605445861816,-56.4659652709961,-56.3851051330566,-56.0421867370605,-55.5896492004395,-55.5287094116211,-55.5280265808105],[-56.0599632263184,-56.2589836120605,-56.3541030883789,-56.5458488464355,-56.6005363464355,-56.6141624450684,-56.4885749816895,-56.140380859375,-56.061767578125,-56.0584449768066,-56.051025390625,-56.0599632263184],[-60.5048828125,-60.5546875,-60.6197242736816,-60.6177253723145,-60.5636711120605,-60.6216773986816,-60.6929168701172,-60.7404289245605,-60.7058601379395,-60.6374015808105,-60.5048828125],[-62.6150894165039,-62.6552734375,-62.6388664245605,-62.5270500183105,-62.3174285888672,-62.3440399169922,-62.4114761352539,-62.6150894165039],[-60.625,-60.5763206481934,-60.139892578125,-60.0027351379395,-59.8495597839355,-60.2209434509277,-60.3215789794922,-60.3538055419922,-60.3780250549316,-60.61962890625,-60.6969261169434,-60.7958946228027,-60.9950714111328,-61.0633316040039,-61.1498031616211,-61.1523895263672,-60.9747047424316,-60.8377456665039,-60.7992706298828,-60.7318344116211,-60.625],[-59.3890151977539,-59.5252418518066,-59.6194305419922,-59.6606903076172,-59.478515625,-59.3958511352539,-59.3533668518066,-59.3890151977539],[-58.8379402160645,-59.0594215393066,-59.174560546875,-59.2024421691895,-59.0638198852539,-58.9906234741211,-58.9621047973633,-58.878662109375,-58.8379402160645],[-57.9784164428711,-57.8493194580078,-57.7379875183105,-57.6765632629395,-57.6365242004395,-57.6395492553711,-57.8066902160645,-57.9627456665039,-58.1475601196289,-58.1722145080566,-58.1331062316895,-58.1830062866211,-58.3414535522461,-58.4666519165039,-58.50732421875,-58.5619583129883,-58.5940399169922,-58.6439895629883,-58.7457046508789,-58.7553215026855,-58.8190422058105,-59.0037078857422,-58.9552230834961,-58.7095184326172,-58.68359375,-58.3994636535645,-58.2651901245117,-57.9784164428711],[-54.0707054138184,-54.1154289245605,-54.1838874816895,-54.1922378540039,-54.1219711303711,-54.0499038696289,-54.0243186950684,-54.0413055419922,-54.0707054138184],[-55.1654281616211,-55.2970199584961,-55.346923828125,-55.369140625,-55.4402351379395,-55.3870124816895,-54.6709976196289,-54.7099609375,-55.0576171875,-55.1654281616211],[-45.7177734375,-45.4997062683105,-45.3862800598145,-45.357421875,-45.2281265258789,-45.2109870910645,-45.1863784790039,-45.1728515625,-45.1736831665039,-45.3980484008789,-45.7091789245605,-45.780029296875,-45.9373054504395,-45.9547882080078,-45.956298828125,-45.934814453125,-45.8341789245605,-45.7177734375],[123.594528198242,123.595207214355,123.573150634766,123.572463989258,123.594528198242],[69.2824172973633,69.2206039428711,69.2015609741211,69.2039031982422,69.1695327758789,69.1501007080078,69.1671905517578,69.2664031982422,69.3687515258789,69.3947296142578,69.3211898803711,69.2824172973633],[69.1848678588867,69.26513671875,69.3142547607422,69.5349578857422,69.5927734375,69.5873031616211,69.64404296875,69.572265625,69.4362258911133,69.4050750732422,69.5423812866211,69.6107406616211,69.6666030883789,69.7707061767578,69.8543930053711,69.9839859008789,70.0613250732422,70.2083969116211,70.2849655151367,70.3202133178711,70.40625,70.4842834472656,70.5308609008789,70.5554733276367,70.5368194580078,70.4850616455078,70.3898468017578,70.4114227294922,70.3861312866211,70.33837890625,70.2976608276367,70.23779296875,70.1658248901367,69.9931640625,69.9156265258789,69.9021530151367,69.8611373901367,69.818359375,69.7599563598633,69.7492141723633,69.7802734375,69.85595703125,69.9864273071289,70.0628890991211,70.0734405517578,70.1658248901367,70.2477493286133,70.30712890625,70.2587890625,70.2162094116211,70.2074203491211,70.1243133544922,70.0750961303711,69.9189453125,69.8260726928711,69.8039016723633,69.7466812133789,69.6820373535156,69.6128921508789,69.4776306152344,69.3527374267578,69.2746047973633,69.1531219482422,69.0859375,68.9929733276367,68.8726577758789,68.8147430419922,68.7828140258789,68.7912139892578,68.810546875,68.8483428955078,68.8720703125,68.8619155883789,68.8184585571289,68.8414077758789,68.798828125,68.8135757446289,68.8833999633789,68.8538055419922,68.8166961669922,68.7901306152344,68.76953125,68.7965850830078,68.8369140625,68.83203125,68.9002914428711,68.9586868286133,69.00244140625,69.0572280883789,69.0812530517578,69.0930633544922,69.0715789794922,69.1227493286133,69.1361312866211,69.1041030883789,69.0994110107422,69.03271484375,69.0521469116211,69.1848678588867],[51.8345718383789,51.76171875,51.6965827941895,51.6592788696289,51.7418937683105,51.7841796875,51.8154296875,51.8345718383789],[-61.716064453125,-61.7481460571289,-61.8596725463867,-61.8820343017578,-61.8871078491211,-61.8172836303711,-61.7385711669922,-61.7082061767578,-61.68603515625,-61.6864776611328,-61.6949691772461,-61.716064453125],[-61.7471160888672,-61.7620124816895,-61.8437957763672,-61.8687477111816,-61.8661613464355,-61.8524436950684,-61.8199195861816,-61.7767601013184,-61.7496109008789,-61.7471160888672],[158.878799438477,158.84521484375,158.8359375,158.89697265625,158.958877563477,158.945602416992,158.878799438477],[147.356063842773,147.308883666992,147.2314453125,147.15380859375,147.10498046875,147.104690551758,147.1630859375,147.184661865234,147.198425292969,147.2197265625,147.233993530273,147.283889770508,147.3125,147.342483520508,147.356063842773],[147.4345703125,147.371871948242,147.348831176758,147.337600708008,147.296096801758,147.319137573242,147.327346801758,147.3525390625,147.397277832031,147.4345703125],[148.104293823242,148.048141479492,148.029693603516,148.030853271484,148.022750854492,148.072555541992,148.142776489258,148.169540405273,148.1005859375,148.104293823242],[145.04296875,145.15869140625,145.224319458008,145.283020019531,145.349411010742,145.429397583008,145.485153198242,145.533493041992,145.576461791992,145.68603515625,145.733795166016,145.775390625,145.821472167969,146.111129760742,146.317489624023,146.574417114258,146.650588989258,146.723434448242,146.786041259766,146.84814453125,146.836029052734,146.856628417969,146.91943359375,146.989837646484,147.105758666992,147.218856811523,147.268951416016,147.320510864258,147.3876953125,147.454788208008,147.500778198242,147.579299926758,147.621688842773,147.817672729492,147.872955322266,147.96875,148.032806396484,148.215225219727,148.292877197266,148.285461425781,148.291610717773,148.306243896484,148.312210083008,148.289840698242,148.286911010742,148.296569824219,148.28759765625,148.315734863281,148.301651000977,148.301467895508,148.328018188477,148.3408203125,148.3310546875,148.342590332031,148.331253051758,148.290328979492,148.276962280273,148.284561157227,148.277145385742,148.255752563477,148.18310546875,148.204391479492,148.241607666016,148.213668823242,148.167190551758,148.141220092773,148.15625,148.127548217773,148.066604614258,148.022750854492,148.0048828125,148.009384155273,147.973541259766,147.924407958984,147.912109375,147.9150390625,147.957717895508,147.980850219727,147.945404052734,147.838577270508,147.785842895508,147.698913574219,147.64794921875,147.687301635742,147.77392578125,147.800384521484,147.807418823242,147.799987792969,147.693450927734,147.573837280273,147.535842895508,147.549026489258,147.536712646484,147.452346801758,147.408004760742,147.297943115234,147.301956176758,147.34765625,147.342666625977,147.324996948242,147.28076171875,147.259765625,147.259765625,147.245010375977,147.1728515625,146.996963500977,146.98486328125,146.987503051758,147.077346801758,147.035934448242,147.004684448242,146.954696655273,146.873931884766,146.834274291992,146.69921875,146.548538208008,146.413177490234,146.186721801758,146.043151855469,146.013076782227,145.981735229492,145.994430541992,146.108779907227,146.226379394531,146.2080078125,146.176467895508,146.125091552734,145.975296020508,145.873245239258,145.802734375,145.681549072266,145.609954833984,145.5673828125,145.517578125,145.487594604492,145.268157958984,145.237121582031,145.198822021484,145.372955322266,145.434860229492,145.46826171875,145.527252197266,145.5166015625,145.3603515625,145.339645385742,145.3310546875,145.29443359375,145.23486328125,145.258987426758,145.238174438477,145.055374145508,144.91552734375,144.777938842773,144.76611328125,144.764358520508,144.69775390625,144.662399291992,144.646102905273,144.709671020508,144.718551635742,144.818557739258,145.04296875],[148.236923217773,148.187805175781,148.126953125,148.117279052734,148.193161010742,148.218353271484,148.236923217773],[144.784362792969,144.748046875,144.710159301758,144.751174926758,144.783401489258,144.790817260742,144.784362792969],[148.326278686523,148.420700073242,148.474212646484,148.404006958008,148.352737426758,148.319427490234,148.214065551758,148.1025390625,148.020111083984,148.010452270508,148.058792114258,148.198135375977,148.326278686523],[148.000396728516,148.177932739258,148.27001953125,148.29736328125,148.289840698242,148.250793457031,148.3232421875,148.313568115234,148.299423217773,148.210357666016,148.105651855469,148.073638916016,148.046875,148.024795532227,147.890518188477,147.905960083008,147.876266479492,147.812301635742,147.767181396484,147.839157104492,147.932998657227,148.000396728516],[143.927932739258,143.898727416992,143.875778198242,143.887588500977,143.838577270508,143.865234375,143.86181640625,143.87939453125,143.939346313477,143.948837280273,144.000778198242,144.09130859375,144.120895385742,144.106048583984,144.141021728516,144.111907958984,144.035049438477,143.927932739258],[145.314453125,145.349212646484,145.355072021484,145.270904541016,145.12841796875,145.2177734375,145.287887573242,145.314453125],[145.486526489258,145.335845947266,145.2802734375,145.285842895508,145.295303344727,145.426559448242,145.486526489258],[137.596481323242,137.8359375,137.928909301758,138.046585083008,138.123428344727,138.066513061523,138.011901855469,137.835556030273,137.6708984375,137.622253417969,137.590240478516,137.448425292969,137.382217407227,137.209564208984,137.147750854492,137.02587890625,136.912704467773,136.755065917969,136.589263916016,136.540634155273,136.5791015625,136.638671875,137.091796875,137.334075927734,137.530456542969,137.5849609375,137.635452270508,137.59814453125,137.596481323242],[153.538772583008,153.452743530273,153.426559448242,153.395797729492,153.40087890625,153.435440063477,153.521881103516,153.538772583008],[153.442474365234,153.4208984375,153.376556396484,153.365036010742,153.3798828125,153.432312011719,153.466796875,153.426361083984,153.442474365234],[113.183006286621,113.156448364258,112.964256286621,112.908203125,112.947067260742,112.982421875,113.096290588379,113.131546020508,113.1318359375,113.148338317871,113.183006286621],[153.077438354492,153.051956176758,153.006927490234,152.976669311523,152.9990234375,153.051559448242,153.060745239258,153.0380859375,153.189254760742,153.2275390625,153.241989135742,153.186325073242,153.143753051758,153.180953979492,153.22314453125,153.256927490234,153.282135009766,153.297958374023,153.359268188477,153.350189208984,153.141403198242,153.083786010742,153.077438354492],[151.146575927734,151.180755615234,151.212020874023,151.240142822266,151.228805541992,151.274322509766,151.295806884766,151.261520385742,151.23828125,151.184173583984,151.033294677734,151.059967041016,151.146575927734],[150.516708374023,150.488479614258,150.46240234375,150.484664916992,150.488479614258,150.521484375,150.548828125,150.516708374023],[149.928314208984,149.893646240234,149.869522094727,149.875381469727,149.912307739258,149.927932739258,149.928314208984],[115.446189880371,115.388084411621,115.318069458008,115.30859375,115.354301452637,115.4345703125,115.457611083984,115.446189880371],[149.043746948242,149.019912719727,148.987411499023,148.938873291016,148.981048583984,149.00439453125,149.045318603516,149.043746948242],[148.935546875,148.913482666016,148.886901855469,148.906448364258,148.931640625,148.967880249023,148.956253051758,148.935546875],[146.2783203125,146.298828125,146.341995239258,146.327056884766,146.298828125,146.235641479492,146.191299438477,146.1162109375,146.098831176758,146.186721801758,146.230850219727,146.249114990234,146.2783203125],[139.459182739258,139.421676635742,139.408203125,139.459182739258,139.492782592773,139.56005859375,139.570892333984,139.459182739258],[139.5078125,139.430572509766,139.391494750977,139.354293823242,139.283004760742,139.239059448242,139.159576416016,139.147659301758,139.162704467773,139.228713989258,139.29296875,139.458877563477,139.587890625,139.6044921875,139.69775390625,139.559677124023,139.5078125],[137.093658447266,137.050872802734,136.996475219727,136.985061645508,136.942672729492,136.96337890625,136.985748291016,137.009567260742,137.064544677734,137.071090698242,137.093658447266],[136.591003417969,136.531158447266,136.514251708984,136.502731323242,136.522552490234,136.586044311523,136.6123046875,136.591003417969],[136.862701416016,136.846771240234,136.845596313477,136.876861572266,136.890228271484,136.862701416016],[124.597267150879,124.559562683105,124.524223327637,124.523735046387,124.482803344727,124.504104614258,124.519332885742,124.550872802734,124.564552307129,124.605079650879,124.597267150879],[125.198829650879,125.134765625,125.091209411621,125.117385864258,125.159965515137,125.198143005371,125.193557739258,125.198829650879],[136.714645385742,136.758010864258,136.804489135742,136.845306396484,136.870712280273,136.890808105469,136.905563354492,136.842956542969,136.81494140625,136.788177490234,136.745315551758,136.749908447266,136.787002563477,136.885452270508,136.933883666992,136.950790405273,136.931350708008,136.894332885742,136.76318359375,136.649703979492,136.460540771484,136.36328125,136.33544921875,136.392181396484,136.427734375,136.4111328125,136.424713134766,136.533782958984,136.582809448242,136.655670166016,136.701950073242,136.695983886719,136.714645385742],[136.237396240234,136.213668823242,136.122650146484,136.122268676758,136.134368896484,136.159576416016,136.215438842773,136.257431030273,136.275390625,136.237396240234],[136.338668823242,136.180267333984,136.267379760742,136.44921875,136.479293823242,136.470520019531,136.37939453125,136.338668823242],[130.459274291992,130.541793823242,130.579879760742,130.602737426758,130.606246948242,130.502532958984,130.317489624023,130.131256103516,130.076568603516,130.043258666992,130.072067260742,130.139068603516,130.197555541992,130.187103271484,130.15283203125,130.251174926758,130.294921875,130.339263916016,130.376754760742,130.385650634766,130.432815551758,130.459274291992],[130.618850708008,130.752258300781,130.912780761719,130.987396240234,131.023040771484,131.140625,131.217178344727,131.268264770508,131.320510864258,131.436904907227,131.473342895508,131.522262573242,131.53857421875,131.467880249023,131.458587646484,131.3828125,131.292098999023,130.950988769531,130.644927978516,130.511917114258,130.422744750977,130.40478515625,130.368560791016,130.384567260742,130.402923583984,130.426651000977,130.519134521484,130.559768676758,130.618850708008],[136.598541259766,136.526565551758,136.521682739258,136.559768676758,136.649032592773,136.68798828125,136.710556030273,136.727340698242,136.731735229492,136.7802734375,136.741409301758,136.598541259766],[132.593353271484,132.573623657227,132.493743896484,132.516311645508,132.483779907227,132.537796020508,132.578811645508,132.59326171875,132.596878051758,132.629104614258,132.593353271484],[143.178909301758,143.152923583984,143.104690551758,143.099014282227,143.110244750977,143.190628051758,143.254104614258,143.289657592773,143.401550292969,143.397552490234,143.457717895508,143.512008666992,143.529479980469,143.586624145508,143.548431396484,143.589263916016,143.643356323242,143.707214355469,143.75634765625,143.822357177734,143.961807250977,144.105865478516,144.209869384766,144.321685791016,144.473052978516,144.58642578125,144.648056030273,144.915725708008,145.064453125,145.179977416992,145.287704467773,145.276947021484,145.251663208008,145.276168823242,145.293075561523,145.271575927734,145.349517822266,145.375381469727,145.4580078125,145.451858520508,145.436416625977,145.426086425781,145.490432739258,145.549896240234,145.638275146484,145.754791259766,145.837890625,145.912109375,145.901947021484,146.0498046875,146.125869750977,146.074020385742,146.022842407227,146.0322265625,146.223052978516,146.333190917969,146.311721801758,146.296875,146.383392333984,146.481155395508,146.587310791016,146.691986083984,146.828994750977,147.002639770508,147.0927734375,147.138778686523,147.278121948242,147.341506958008,147.418548583984,147.470901489258,147.509765625,147.586029052734,147.742385864258,147.853225708008,147.915634155273,148.004486083984,148.0810546875,148.189636230469,148.366897583008,148.526748657227,148.600494384766,148.759368896484,148.820999145508,148.884765625,148.805084228516,148.72998046875,148.683700561523,148.789459228516,148.912399291992,149.060546875,149.204879760742,149.241409301758,149.2802734375,149.329299926758,149.4541015625,149.460052490234,149.524032592773,149.595703125,149.645309448242,149.703903198242,149.771575927734,149.822463989258,149.920318603516,149.974411010742,150.005569458008,149.94189453125,149.981262207031,150.020614624023,150.076171875,150.142959594727,150.23486328125,150.405075073242,150.541305541992,150.57958984375,150.564361572266,150.568557739258,150.622848510742,150.672454833984,150.763870239258,150.782821655273,150.783004760742,150.843170166016,150.931060791016,150.98876953125,151.087692260742,151.15380859375,151.236328125,151.500778198242,151.575393676758,151.69091796875,151.831634521484,151.902740478516,152.055358886719,152.1298828125,152.282028198242,152.353118896484,152.456344604492,152.4931640625,152.502044677734,152.563278198242,152.654296875,152.789154052734,152.913482666016,152.920501708984,152.984954833984,153.028213500977,153.12548828125,153.164947509766,153.084182739258,153.162109375,153.116806030273,153.197952270508,153.3857421875,153.428421020508,153.454879760742,153.57568359375,153.569137573242,153.616897583008,153.604583740234,153.462203979492,153.348052978516,153.346969604492,153.272369384766,153.223831176758,153.188186645508,153.030563354492,153.023742675781,153.0478515625,153.021575927734,152.982223510742,152.943939208984,152.785842895508,152.559280395508,152.545303344727,152.5166015625,152.470413208008,152.331253051758,152.247451782227,152.215728759766,152.136520385742,152.134567260742,152.188095092773,152.164245605469,151.954284667969,151.812896728516,151.668350219727,151.607711791992,151.530075073242,151.483795166016,151.46337890625,151.432022094727,151.357528686523,151.292098999023,151.32275390625,151.288375854492,151.2802734375,151.24462890625,151.20166015625,151.167861938477,151.124801635742,151.191207885742,151.231552124023,151.089950561523,150.960342407227,150.927444458008,150.871276855469,150.821868896484,150.781051635742,150.809188842773,150.804595947266,150.774612426758,150.756057739258,150.697357177734,150.680953979492,150.705657958984,150.72216796875,150.714660644531,150.690338134766,150.634460449219,150.567474365234,150.374114990234,150.292190551758,150.1953125,150.158493041992,150.12890625,150.095306396484,150.062789916992,149.988174438477,149.960266113281,149.950592041016,149.986328125,149.962890625,149.962310791016,149.932723999023,149.809387207031,149.708984375,149.5654296875,149.480865478516,149.298446655273,148.943939208984,148.262512207031,148.130676269531,147.876754760742,147.631454467773,147.395599365234,146.856842041016,146.435745239258,146.356262207031,146.292587280273,146.217483520508,146.216201782227,146.285552978516,146.336624145508,146.426940917969,146.466598510742,146.481643676758,146.483795166016,146.456649780273,146.399993896484,146.340042114258,146.33203125,146.254287719727,146.158401489258,146.069915771484,146.018157958984,145.935348510742,145.865539550781,145.790832519531,145.69189453125,145.606338500977,145.535354614258,145.397262573242,145.424224853516,145.462799072266,145.542190551758,145.518356323242,145.475769042969,145.366409301758,145.292770385742,145.248931884766,145.191223144531,144.959564208984,144.847259521484,144.7177734375,144.7802734375,144.911422729492,145.020126342773,145.066986083984,145.119918823242,145.049606323242,144.98486328125,144.891296386719,144.538482666016,144.46533203125,144.3955078125,144.517776489258,144.589447021484,144.665237426758,144.543640136719,144.447845458984,144.328720092773,144.1015625,143.811721801758,143.686721801758,143.538970947266,143.338470458984,143.226470947266,143.082611083984,142.840240478516,142.612106323242,142.455856323242,142.344528198242,142.187698364258,141.924713134766,141.725006103516,141.593948364258,141.491790771484,141.424209594727,141.2138671875,141.010940551758,140.627243041992,140.390426635742,140.212097167969,139.874801635742,139.784271240234,139.742279052734,139.738464355469,139.783889770508,139.846572875977,139.857330322266,139.72900390625,139.548721313477,139.465927124023,139.244918823242,139.037689208984,138.985061645508,138.968948364258,139.06689453125,139.112503051758,139.178024291992,139.230560302734,139.289459228516,139.292083740234,139.325088500977,139.302536010742,139.282516479492,139.192764282227,139.09375,139.017684936523,138.915222167969,138.875305175781,138.77099609375,138.729690551758,138.521881103516,138.389266967773,138.184371948242,138.252151489258,138.332916259766,138.399810791016,138.511123657227,138.489944458008,138.436233520508,138.264358520508,138.186233520508,138.089248657227,138.041305541992,138.012298583984,137.919235229492,137.874114990234,137.691696166992,137.56640625,137.459564208984,137.272369384766,137.144439697266,137.029876708984,136.966598510742,136.883590698242,137.014266967773,137.12841796875,137.252044677734,137.308395385742,137.391021728516,137.454284667969,137.492965698242,137.468551635742,137.458984375,137.483596801758,137.493850708008,137.650390625,137.780853271484,137.931838989258,137.913970947266,137.866012573242,137.852340698242,137.92431640625,137.992584228516,137.913192749023,137.863098144531,137.783203125,137.781845092773,137.790908813477,137.68017578125,137.536224365234,137.442291259766,137.354202270508,137.2373046875,137.130264282227,137.034469604492,136.9365234375,136.783493041992,136.635543823242,136.52587890625,136.4306640625,136.12109375,135.979675292969,135.950592041016,135.891006469727,135.902633666992,135.950592041016,135.99853515625,135.9697265625,135.919143676758,135.792388916016,135.712493896484,135.647552490234,135.480865478516,135.411727905273,135.32421875,135.230667114258,135.190826416016,135.123046875,135.129592895508,135.175979614258,135.216796875,135.29248046875,135.378707885742,135.427337646484,135.449996948242,135.367965698242,135.31201171875,135.286315917969,135.218948364258,135.185455322266,135.042083740234,134.888763427734,134.8466796875,134.791015625,134.719039916992,134.607711791992,134.30126953125,134.173538208008,134.100402832031,134.158401489258,134.227157592773,134.249206542969,134.234191894531,133.93017578125,133.786712646484,133.665328979492,133.551376342773,133.400588989258,133.212097167969,132.757415771484,132.648635864258,132.323638916016,132.214645385742,131.72119140625,131.393173217773,131.284957885742,131.143646240234,131.029296875,130.948135375977,130.783020019531,130.129791259766,129.56884765625,129.187698364258,128.946182250977,128.546096801758,128.067672729492,127.678024291992,127.31982421875,127.084083557129,126.779296875,126.136520385742,125.917190551758,125.567481994629,125.463676452637,125.2666015625,124.7587890625,124.524604797363,124.373237609863,124.243751525879,124.126083374023,123.967178344727,123.868362426758,123.650382995605,123.506843566895,123.365425109863,123.207611083984,123.067581176758,122.955657958984,122.777542114258,122.150978088379,122.061134338379,121.946388244629,121.729682922363,121.405082702637,120.814544677734,120.530570983887,120.418365478516,120.209373474121,119.854095458984,119.729103088379,119.635162353516,119.450584411621,119.247657775879,119.081344604492,118.895317077637,118.520118713379,118.135551452637,118.006446838379,117.863082885742,117.675392150879,117.581939697266,117.143943786621,116.865425109863,116.517196655273,116.217086791992,115.986717224121,115.726272583008,115.565040588379,115.277641296387,115.194831848145,115.127937316895,115.0087890625,115.005661010742,114.973434448242,114.975685119629,114.993843078613,115.098922729492,115.181648254395,115.358795166016,115.515327453613,115.6044921875,115.683006286621,115.670890808105,115.618560791016,115.654296875,115.707916259766,115.725387573242,115.738090515137,115.698432922363,115.454597473145,115.294334411621,115.176849365234,115.077926635742,114.994529724121,114.968841552734,114.942085266113,114.971382141113,114.958984375,114.856826782227,114.62841796875,114.590621948242,114.591789245605,114.537399291992,114.353515625,114.165138244629,114.133499145508,114.098442077637,114.028121948242,113.709373474121,113.3330078125,113.231056213379,113.184768676758,113.210739135742,113.253120422363,113.300102233887,113.3232421875,113.345306396484,113.342872619629,113.356056213379,113.388969421387,113.427444458008,113.546577453613,113.581642150879,113.733695983887,113.780372619629,113.83642578125,113.852836608887,113.775779724121,113.706451416016,113.589065551758,113.513381958008,113.395309448242,113.397361755371,113.451370239258,113.539451599121,113.621192932129,113.713088989258,113.697845458984,113.68359375,113.691703796387,113.723731994629,113.765823364258,113.811820983887,113.853904724121,113.8798828125,113.9423828125,113.991989135742,114.09033203125,114.175979614258,114.215721130371,114.203315734863,114.228515625,114.214256286621,113.992774963379,113.792388916016,113.670799255371,113.569236755371,113.503517150879,113.417671203613,113.412986755371,113.421287536621,113.489845275879,113.552932739258,113.757034301758,113.766990661621,113.764846801758,113.794921875,113.795120239258,113.767875671387,113.682807922363,113.795021057129,113.958396911621,114.022850036621,114.123924255371,114.142578125,114.0927734375,114.163864135742,114.1416015625,114.205177307129,114.303512573242,114.377731323242,114.4169921875,114.602828979492,114.709274291992,114.859085083008,115.161712646484,115.456153869629,115.596092224121,115.771484375,115.8935546875,116.010940551758,116.605865478516,116.706733703613,116.836326599121,116.995307922363,117.139053344727,117.292778015137,117.40625,117.683883666992,117.832321166992,118.087303161621,118.199226379395,118.458305358887,118.751457214355,119.104499816895,119.358787536621,119.585945129395,119.767776489258,120.1962890625,120.433692932129,120.87841796875,120.997947692871,121.179779052734,121.337692260742,121.493560791016,121.589454650879,121.630668640137,121.721977233887,121.784866333008,121.833786010742,122.006248474121,122.262107849121,122.345405578613,122.360939025879,122.305763244629,122.237396240234,122.191307067871,122.147453308105,122.143165588379,122.160255432129,122.260932922363,122.332717895508,122.432029724121,122.522552490234,122.597946166992,122.720413208008,122.772079467773,122.848045349121,122.916793823242,122.970695495605,123.074409484863,123.142097473145,123.265914916992,123.383201599121,123.478813171387,123.525199890137,123.563079833984,123.571479797363,123.561813354492,123.607917785645,123.586334228516,123.593559265137,123.617683410645,123.664054870605,123.753807067871,123.799026489258,123.8310546875,123.829483032227,123.874412536621,123.856338500977,123.778121948242,123.745025634766,123.680465698242,123.607131958008,123.517974853516,123.490432739258,123.525093078613,123.581344604492,123.6259765625,123.646492004395,123.607025146484,123.6474609375,123.728904724121,123.859184265137,123.915229797363,123.961318969727,124.04443359375,124.129783630371,124.18603515625,124.300384521484,124.452743530273,124.529975891113,124.6923828125,124.77197265625,124.757026672363,124.669242858887,124.5703125,124.454490661621,124.404884338379,124.388275146484,124.416404724121,124.4345703125,124.509963989258,124.576858520508,124.585052490234,124.608589172363,124.648536682129,124.648345947266,124.606636047363,124.504295349121,124.455268859863,124.381637573242,124.396583557129,124.439544677734,124.505661010742,124.561630249023,124.644340515137,124.690925598145,124.680168151855,124.692573547363,124.750480651855,124.972068786621,125.016410827637,125.06298828125,125.077934265137,125.072952270508,125.024017333984,124.909172058105,124.882713317871,124.892669677734,124.839073181152,124.914161682129,124.978706359863,125.023330688477,125.024017333984,125.038185119629,125.072952270508,125.188667297363,125.302345275879,125.355667114258,125.375579833984,125.3837890625,125.243263244629,125.239456176758,125.180374145508,125.1787109375,125.266502380371,125.284576416016,125.33544921875,125.435935974121,125.503715515137,125.579788208008,125.598342895508,125.597068786621,125.627738952637,125.70458984375,125.681243896484,125.680961608887,125.661613464355,125.690521240234,125.708404541016,125.738479614258,125.819526672363,125.839553833008,125.85009765625,125.890617370605,125.946090698242,126.020706176758,126.016609191895,126.044830322266,126.053611755371,126.100875854492,126.111328125,126.073432922363,126.053901672363,126.119049072266,126.184272766113,126.228218078613,126.258491516113,126.298828125,126.323051452637,126.403129577637,126.482429504395,126.569732666016,126.679092407227,126.780662536621,126.764457702637,126.775588989258,126.903221130371,127.006057739258,127.099220275879,127.293060302734,127.457611083984,127.531051635742,127.672843933105,127.763481140137,127.887603759766,128.180465698242,128.199417114258,128.159866333008,128.124420166016,128.080474853516,128.069427490234,128.111709594727,128.155471801758,128.201766967773,128.254684448242,128.258987426758,128.227249145508,128.172958374023,128.175003051758,128.218353271484,128.28515625,128.358200073242,128.403228759766,128.409866333008,128.477447509766,128.575790405273,128.635543823242,129.058197021484,129.165130615234,129.175201416016,129.2158203125,129.237884521484,129.233596801758,129.267578125,129.381256103516,129.458984375,129.567092895508,129.587692260742,129.634765625,129.650299072266,129.628219604492,129.612701416016,129.637115478516,129.763473510742,129.848724365234,129.808395385742,129.753509521484,129.662979125977,129.604690551758,129.698638916016,129.697937011719,129.60791015625,129.48388671875,129.378707885742,129.459167480469,129.61962890625,129.709869384766,129.718368530273,129.76171875,129.789260864258,129.797164916992,129.8388671875,129.937881469727,130.072662353516,130.135940551758,130.199325561523,130.259765625,130.134963989258,130.145309448242,130.168167114258,130.317977905273,130.39990234375,130.454193115234,130.571884155273,130.617477416992,130.609573364258,130.622650146484,130.67236328125,130.736129760742,130.776565551758,130.867385864258,130.898239135742,130.882919311523,130.873825073242,130.956634521484,131.0234375,131.030075073242,131.01953125,131.045700073242,131.219924926758,131.265426635742,131.291610717773,131.313781738281,131.342086791992,131.438278198242,131.726257324219,131.887985229492,131.956741333008,132.064056396484,132.182327270508,132.253219604492,132.3720703125,132.411026000977,132.441604614258,132.510543823242,132.583786010742,132.676376342773,132.712783813477,132.630477905273,132.63525390625,132.6298828125,132.644729614258,132.674224853516,132.475189208984,132.277923583984,132.133590698242,132.072860717773,131.944625854492,131.822463989258,131.811813354492,131.961517333984,132.0185546875,132.105758666992,132.155471801758,132.19775390625,132.225006103516,132.2626953125,132.333984375,132.557327270508,132.682815551758,132.7470703125,132.857116699219,132.961044311523,133.02490234375,133.114349365234,133.185256958008,133.356155395508,133.443161010742,133.533203125,133.654495239258,133.904205322266,134.139450073242,134.237106323242,134.35107421875,134.417388916016,134.5380859375,134.730270385742,134.81640625,134.854675292969,135.029678344727,135.217971801758,135.352340698242,135.548721313477,135.685546875,135.788482666016,135.88525390625,135.922454833984,135.843551635742,135.833984375,135.895812988281,135.889450073242,135.804290771484,135.702545166016,135.704391479492,135.743942260742,135.790832519531,135.857421875,135.937805175781,136.008483886719,136.031448364258,136.081832885742,136.192672729492,136.260650634766,136.328506469727,136.291885375977,136.249893188477,136.270111083984,136.443359375,136.540237426758,136.609771728516,136.719436645508,136.83642578125,136.8974609375,136.947463989258,136.537017822266,136.517776489258,136.573043823242,136.594329833984,136.461029052734,136.411911010742,136.364547729492,136.294143676758,136.232330322266,136.166122436523,135.927352905273,135.92919921875,135.989547729492,135.954498291016,135.883407592773,135.806350708008,135.744537353516,135.538864135742,135.473236083984,135.405181884766,135.428024291992,135.453323364258,135.53076171875,135.832611083984,135.969528198242,136.205368041992,136.25927734375,136.291412353516,136.4619140625,136.583587646484,136.618759155273,136.644134521484,136.674606323242,136.704879760742,136.700103759766,136.686721801758,136.698150634766,136.78466796875,136.922653198242,137.002151489258,137.08984375,137.1689453125,137.29931640625,137.5263671875,137.703720092773,137.912887573242,138.071578979492,138.245025634766,138.505661010742,138.625686645508,138.8203125,139.009872436523,139.1103515625,139.14453125,139.154113769531,139.248428344727,139.440521240234,139.689651489258,139.89453125,139.945999145508,140.035842895508,140.209671020508,140.511123657227,140.6484375,140.830474853516,140.915817260742,140.966018676758,141.219146728516,141.291412353516,141.355667114258,141.411911010742,141.393173217773,141.451568603516,141.581451416016,141.62548828125,141.603515625,141.52294921875,141.558990478516,141.594329833984,141.535446166992,141.480667114258,141.472564697266,141.5341796875,141.588775634766,141.645401000977,141.613571166992,141.734573364258,141.7822265625,141.875793457031,141.920303344727,141.929794311523,141.892883300781,141.878311157227,141.852142333984,141.794540405273,141.746688842773,141.677734375,141.688568115234,141.805770874023,141.870498657227,141.912979125977,141.961135864258,141.9677734375,141.951568603516,142.04052734375,142.138961791992,142.168365478516,142.326477050781,142.406829833984,142.456451416016,142.544815063477,142.605072021484,142.5654296875,142.552734375,142.723052978516,142.779678344727,142.803314208984,142.836822509766,142.852935791016,142.8505859375,142.87255859375,142.933990478516,142.988479614258,143.06640625,143.178909301758],[142.274810791016,142.19140625,142.13720703125,142.12548828125,142.131057739258,142.197952270508,142.274810791016],[142.338973999023,142.279403686523,142.216201782227,142.195114135742,142.21875,142.298721313477,142.338973999023],[142.167572021484,142.141983032227,142.09765625,142.148834228516,142.192001342773,142.167572021484],[16.953125,16.9488277435303,16.943359375,16.9044933319092,16.8626956939697,16.8654289245605,16.97265625,17.06787109375,17.0859394073486,17.1473636627197,17.0890636444092,17.0777339935303,17.0399398803711,17.0300769805908,17.0458984375,17.0456066131592,17.0666027069092,16.9734363555908,16.8626956939697,16.8230457305908,16.7859382629395,16.74755859375,16.6474609375,16.5909175872803,16.5509777069092,16.5210933685303,16.4696273803711,16.4212894439697,16.43212890625,16.6397457122803,16.6765632629395,16.6366214752197,16.623046875,16.5744132995605,16.5147457122803,16.44287109375,16.4343738555908,16.4625988006592,16.4397468566895,16.4168949127197,16.4383792877197,16.4828128814697,16.49267578125,16.4847660064697,16.4769535064697,16.4612312316895,16.4534187316895,16.4239253997803,16.3318367004395,16.2525386810303,16.0930671691895,16.0372066497803,15.9768562316895,15.98046875,15.9722661972046,15.9576168060303,15.7668943405151,15.7602529525757,15.6326179504395,15.5453128814697,15.4392585754395,15.2169923782349,15.0006837844849,14.9494142532349,14.8932619094849,14.8406248092651,14.8105459213257,14.7567377090454,14.6801748275757,14.5969734191895,14.5771484375,14.5498056411743,14.5035152435303,14.4659185409546,14.4199209213257,14.2672853469849,14.0995121002197,14.0196285247803,13.9288082122803,13.8313484191895,13.7439460754395,13.6999998092651,13.4900398254395,13.3515625,13.1687498092651,12.8055667877197,12.6998052597046,12.5986318588257,12.4791984558105,12.3882808685303,12.330078125,12.2679691314697,12.1541013717651,12.1307621002197,12.16552734375,12.2012701034546,12.1971683502197,12.16943359375,11.9695310592651,11.7756834030151,11.6994142532349,11.62548828125,11.5275392532349,11.4332027435303,11.2444334030151,11.1338863372803,11.0634765625,11.0250978469849,10.9932613372803,10.9273443222046,10.8289060592651,10.759765625,10.6892576217651,10.5797853469849,10.4793949127197,10.4528331756592,10.45458984375,10.4149417877197,10.3494138717651,10.1797857284546,10.1334953308105,9.99687480926514,9.87773418426514,9.86464881896973,9.84531211853027,9.74501895904541,9.61992168426514,9.58027362823486,9.595703125,9.61054706573486,9.60117244720459,9.57187461853027,9.55576133728027,9.55107402801514,9.54218769073486,9.53681659698486,9.52753925323486,9.60908222198486,9.62587833404541,9.55439472198486,9.52402305603027,9.54892539978027,9.65058612823486,9.71513652801514,9.74892520904541,9.83915996551514,9.97158145904541,10.0340824127197,10.0598640441895,10.07421875,10.0663089752197,10.0964851379395,10.1587886810303,10.2002935409546,10.1857423782349,10.1830081939697,10.2406244277954,10.3127927780151,10.369140625,10.4039068222046,10.4303712844849,10.439453125,10.4828119277954,10.65869140625,10.7416009902954,10.873046875,10.87060546875,10.8939447402954,10.9521484375,10.9808588027954,11.0419921875,11.1360349655151,11.1912107467651,11.2119140625,11.2979488372803,11.3741207122803,11.3929691314697,11.4699220657349,11.5739250183105,11.716796875,12.1856451034546,12.2038087844849,12.1968755722046,12.2092771530151,12.2683591842651,12.3631839752197,12.4357423782349,12.48291015625,12.5265626907349,12.59423828125,12.6858396530151,12.7713861465454,12.7961912155151,12.7811517715454,12.7828121185303,12.8093748092651,12.87890625,12.9680671691895,13.0143547058105,13.0315437316895,13.0479488372803,13.0541019439697,13.0335931777954,12.9855470657349,12.9281244277954,12.8976564407349,12.9083013534546,12.9541988372803,12.9535160064697,12.8499021530151,12.7600584030151,12.7603511810303,12.8142576217651,12.8974609375,13.0821285247803,13.1404294967651,13.2152347564697,13.3228521347046,13.3746089935303,13.4093751907349,13.4598636627197,13.4716796875,13.4866209030151,13.6751947402954,13.6921873092651,13.7239255905151,13.7853507995605,13.798828125,13.7974615097046,13.8029298782349,13.8147459030151,13.8431634902954,13.92431640625,13.98876953125,14.0491209030151,14.1898431777954,14.3675775527954,14.4310541152954,14.4886722564697,14.5539054870605,14.6913080215454,14.7066402435303,14.7859373092651,14.8218746185303,14.9225578308105,14.9473638534546,14.9721670150757,14.9934568405151,15.0667963027954,15.1397457122803,15.1617183685303,15.1996097564697,15.2527341842651,15.3109378814697,15.4029302597046,15.5994148254395,15.7007808685303,15.7650394439697,15.8251953125,16.0572261810303,16.2193355560303,16.3672847747803,16.4148426055908,16.4779300689697,16.5435543060303,16.6009769439697,16.7126960754395,16.7644538879395,16.8332023620605,16.8836917877197,16.9283199310303,16.953125],[46.1144523620605,45.921875,45.5750007629395,45.4796905517578,45.3892555236816,45.3355484008789,45.2559547424316,45.1906242370605,45.1412124633789,45.1130867004395,45.0716819763184,45.0001945495605,44.8381843566895,44.8171882629395,44.7833976745605,44.7682647705078,44.8671875,45.0316429138184,45.0764656066895,45.1246109008789,45.1486320495605,45.15283203125,45.1725578308105,45.2525405883789,45.2882804870605,45.3499031066895,45.4568367004395,45.6107444763184,45.6874008178711,45.75048828125,45.7844734191895,45.7964820861816,45.7841796875,45.7663078308105,45.7986297607422,45.9249992370605,45.9774436950684,45.9518547058105,46.0458984375,46.0774383544922,46.1144523620605],[44.9690437316895,44.9588851928711,44.96142578125,44.9943351745605,45.0210914611816,45.0287132263184,45.0236320495605,45.0020484924316,44.9690437316895],[48.5728530883789,48.6646461486816,48.8239288330078,49.0508804321289,49.1066398620605,49.1432609558105,49.1747093200684,49.2264633178711,49.4567375183105,49.55615234375,49.7183609008789,49.7759780883789,49.8517570495605,49.9906272888184,50.119140625,50.1825218200684,50.248046875,50.3068351745605,50.3659172058105,50.1431655883789,49.9188461303711,49.7919883728027,49.6690444946289,49.5511703491211,49.4773445129395,49.4151382446289,49.3244132995605,49.3275413513184,49.3673820495605,49.36279296875,49.3211936950684,49.2693367004395,49.1998062133789,49.1653289794922,49.1209945678711,49.1086921691895,49.111328125,49.0134735107422,48.9617195129395,48.9261703491211,48.8544921875,48.8508796691895,48.8687515258789,48.84033203125,48.6355476379395,48.5926742553711,48.4173851013184,48.3812484741211,48.3055686950684,48.2613296508789,48.2251968383789,48.2046890258789,48.0232429504395,47.9964828491211,47.99267578125,48.0193367004395,48.0500984191895,48.1385765075684,48.2419929504395,48.2750968933105,48.2920913696289,48.291015625,48.2741203308105,48.12548828125,48.1091766357422,48.1043930053711,48.1128921508789,48.1360359191895,48.2572250366211,48.3221664428711,48.28173828125,48.1510734558105,47.9958992004395,47.8922843933105,47.7728500366211,47.5818367004395,47.4761734008789,47.3384780883789,47.1883811950684,47.0654296875,46.9888687133789,46.8525390625,46.783203125,46.5547828674316,46.4906234741211,46.4867172241211,46.4898414611816,46.4753913879395,46.4014625549316,46.4002914428711,46.4203109741211,46.4771499633789,46.5499992370605,46.5847663879395,46.5066413879395,46.4373054504395,46.37841796875,46.365234375,46.3651390075684,46.3776359558105,46.4781265258789,46.4880828857422,46.4814453125,46.3216819763184,46.2020492553711,46.0948219299316,46.02587890625,45.93994140625,45.8631820678711,45.7896499633789,45.6618156433105,45.5797843933105,45.5809555053711,45.5959968566895,45.6301765441895,45.8581047058105,45.8859367370605,45.9000968933105,45.9312515258789,45.9675788879395,45.9646492004395,45.7357406616211,45.5695304870605,45.4543914794922,45.3761711120605,45.37890625,45.4013671875,45.5793952941895,45.5914077758789,45.5875015258789,45.5240249633789,45.4442367553711,45.4191398620605,45.3689422607422,45.2734375,45.1060523986816,45.0705108642578,45.0625953674316,45.0707054138184,45.1902351379395,45.1885719299316,45.15234375,45.0847625732422,45.02294921875,45.0013656616211,45.2171859741211,45.2809562683105,45.4222640991211,45.7156257629395,45.6957054138184,45.7254867553711,45.7927742004395,45.9219703674316,46.03125,46.0865249633789,46.1707038879395,46.2799835205078,46.3807601928711,46.4309577941895,46.4579124450684,46.5343742370605,46.6263694763184,46.6623992919922,46.6725578308105,46.6189422607422,46.5087890625,46.3849639892578,46.3054695129395,46.2546882629395,46.2035140991211,46.1905288696289,46.18212890625,46.1842765808105,46.2018547058105,46.2518539428711,46.3025398254395,46.3482437133789,46.4054679870605,46.4298858642578,46.5376968383789,46.5521469116211,46.5712890625,46.6160163879395,46.6903343200684,46.7493133544922,46.8255844116211,46.9308586120605,46.98779296875,47.0101585388184,47.06396484375,47.142578125,47.2052764892578,47.2611312866211,47.3176765441895,47.5206031799316,47.591796875,47.791015625,47.8611335754395,47.9636726379395,48.0560531616211,48.1422843933105,48.2981452941895,48.3914070129395,48.4306640625,48.5186500549316,48.5728530883789],[45.5523452758789,45.5623016357422,45.5341796875,45.5044937133789,45.4788093566895,45.4788093566895,45.5143547058105,45.5523452758789],[30.5536117553711,30.53369140625,30.4419918060303,30.4242210388184,30.4343738555908,30.4733390808105,30.4504890441895,30.4413089752197,30.4240226745605,30.4334964752197,30.4555683135986,30.5150375366211,30.6042976379395,30.70947265625,30.7802753448486,30.796875,30.7935562133789,30.8111343383789,30.8114280700684,30.7902336120605,30.6818351745605,30.6260738372803,30.6109371185303,30.6246070861816,30.6319332122803,30.5298843383789,30.4249992370605,30.4000015258789,30.3791007995605,30.3484363555908,30.2685527801514,30.1871109008789,30.1471672058105,29.947265625,29.7695293426514,29.7177734375,29.4032230377197,29.3791980743408,29.3313484191895,29.2232418060303,29.2118167877197,29.216796875,29.2171859741211,29.2100582122803,29.2123050689697,29.2260761260986,29.2244148254395,29.1532211303711,29.0647468566895,29.0165996551514,29.0141582489014,29.0143566131592,29.0286140441895,29.0631828308105,29.1020488739014,29.1975593566895,29.2970714569092,29.3498039245605,29.3902359008789,29.4636707305908,29.6513671875,29.6980476379395,29.7833976745605,29.8681640625,29.892578125,29.9124031066895,29.93017578125,29.9734382629395,30.0918941497803,30.1172847747803,30.1422843933105,30.1833000183105,30.2337875366211,30.27099609375,30.4084968566895,30.4822254180908,30.5289058685303,30.5536117553711],[5.99394512176514,6.00595712661743,6.11943340301514,6.15449237823486,6.23593759536743,6.16845655441284,6.1787109375,6.20302724838257,6.294921875,6.34091854095459,6.34365224838257,6.36445331573486,6.17509794235229,6.12128877639771,6.11650419235229,6.11005830764771,6.08906221389771,6.05478572845459,5.97626972198486,5.86689472198486,5.8173828125,5.78808641433716,5.74404287338257,5.73525333404541,5.74082040786743,5.72578191757202,5.72499990463257,5.78798818588257,5.8037109375,5.88037109375,5.85654306411743,5.83759737014771,5.81543016433716,5.78974580764771,5.71044921875,5.61005830764771,5.54238319396973,5.50732421875,5.43466806411743,5.353515625,5.30195331573486,5.27880859375,5.21503877639771,5.12412118911743,5.06103515625,5.00693368911743,4.93056678771973,4.86757802963257,4.84912109375,4.84150362014771,4.7900390625,4.86054706573486,4.81865215301514,4.77285146713257,4.70664072036743,4.67509794235229,4.65615224838257,4.54501962661743,4.36875009536743,4.17607402801514,4.14931631088257,4.13701200485229,4.13681697845459,4.15029335021973,4.18388652801514,4.19218730926514,4.15771436691284,4.13525390625,4.14414072036743,4.16962862014771,4.17460966110229,4.04414033889771,3.94970679283142,3.85810542106628,3.78857398033142,3.74804711341858,3.71884799003601,3.68935537338257,3.66728520393372,3.62675833702087,3.59541034698486,3.47695302963257,3.31621098518372,3.27333998680115,3.24980473518372,3.23496103286743,3.18203091621399,3.15488266944885,3.10683608055115,3.02285122871399,2.92197251319885,2.86240243911743,2.83974623680115,2.75937509536743,2.66914057731628,2.59677720069885,2.57929682731628,2.60146498680115,2.57480478286743,2.53603482246399,2.52490234375,2.96015620231628,3.22519540786743,3.35009741783142,3.38007783889771,3.40283179283142,3.43251967430115,3.47197222709656,3.51709008216858,3.58027362823486,3.68183612823486,3.75566411018372,3.78193330764771,3.83076143264771,3.90205025672913,4.0400390625,4.17255878448486,4.21142578125,4.22617149353027,4.30449199676514,4.37373018264771,4.40400409698486,4.38476610183716,4.44091796875,4.50341796875,4.53164052963257,4.58876943588257,4.63398408889771,4.75566434860229,4.78418016433716,4.81054639816284,4.81601524353027,4.82070302963257,4.84804677963257,4.94394493103027,4.99257850646973,5.03095674514771,5.05947303771973,5.07343769073486,5.09990262985229,5.21415996551514,5.31083965301514,5.42978525161743,5.47685527801514,5.5087890625,5.54042959213257,5.60878896713257,5.75234413146973,5.79648447036743,5.8271484375,5.81826162338257,5.74980449676514,5.74082040786743,5.75,5.73662137985229,5.64755868911743,5.63945341110229,5.66914081573486,5.69355487823486,5.7470703125,5.79736328125,5.89246129989624,5.99394512176514],[3.59541034698486,3.55390644073486,3.49052691459656,3.48779296875,3.63886690139771,3.65624976158142,3.6953125,3.71640658378601,3.73417973518372,3.74492192268372,3.75683617591858,3.82968783378601,3.83447241783142,3.78378915786743,3.77177739143372,3.75849604606628,3.68027377128601,3.64658212661743,3.60410165786743,3.57792949676514,3.57656240463257,3.64589810371399,3.60205101966858,3.55722665786743,3.47675776481628,3.40478515625,3.35449194908142,3.3251953125,3.32949209213257,3.22343754768372,3.16464829444885,3.13613247871399,3.14804673194885,3.11044907569885,3.044921875,2.89804697036743,2.77480483055115,2.73291015625,2.73466801643372,2.7236328125,2.70312476158142,2.71152353286743,2.70234417915344,2.68603539466858,2.70771503448486,2.72040987014771,2.71933603286743,2.75097680091858,2.78515648841858,2.78398418426514,2.76582026481628,2.75048851966858,2.75058579444885,2.75673818588257,2.74775362014771,2.72138690948486,2.73173832893372,2.75292992591858,2.77460932731628,2.75371074676514,2.73564434051514,2.70800757408142,2.70644521713257,2.28691411018372,1.81816411018372,1.62265598773956,1.61093723773956,1.77792978286743,1.74316394329071,1.63925802707672,1.59853506088257,1.57753896713257,1.60292959213257,1.59082019329071,1.58203101158142,1.53095698356628,1.62470722198486,1.62470722198486,1.62460923194885,1.62460923194885,1.62460923194885,1.60664069652557,1.60380852222443,1.60019552707672,1.56630849838257,1.42431628704071,1.38574194908142,1.37890601158142,1.34707021713257,1.34511697292328,1.34287095069885,1.33007800579071,1.17617177963257,0.958300530910492,0.792187511920929,0.779980599880219,0.763379216194153,0.787500083446503,0.821874737739563,0.874804556369781,0.900488495826721,0.924609541893005,0.958007752895355,0.985058844089508,1.01386713981628,1.06230449676514,1.08457016944885,1.08154320716858,1.09755861759186,1.13554680347443,1.14550769329071,1.14580094814301,1.17871117591858,1.23466789722443,1.28046905994415,1.31738293170929,1.36484396457672,1.39150369167328,1.39970719814301,1.42675769329071,1.50136709213257,1.56142580509186,1.60000014305115,1.85761737823486,1.98037135601044,2.23085927963257,2.28720688819885,2.36328125,2.38916015625,2.41269540786743,2.36328125,2.36601543426514,2.46933579444885,2.59843754768372,2.64843726158142,2.68134737014771,2.728515625,2.80527353286743,2.85019516944885,2.87812519073486,3.14960908889771,3.26738309860229,3.29912137985229,3.35996079444885,3.44980478286743,3.53173828125,3.59541034698486],[0.217480450868607,0.203808531165123,0.202734231948853,0.185058489441872,0.163867071270943,0.250585943460464,0.354589819908142,0.38251981139183,0.354882627725601,0.374023407697678,0.429199248552322,0.522363305091858,0.61816418170929,0.684570550918579,0.747754037380219,0.786035060882568,0.842285215854645,0.897949159145355,0.946581840515137,0.977734088897705,1.01787102222443,1.12597668170929,1.20117163658142,1.17089831829071,1.07685530185699,0.988476514816284,0.976757824420929,0.973046779632568,0.98730480670929,1.00791013240814,1.09677755832672,1.30869114398956,1.50048816204071,1.56494140625,1.67109382152557,1.78984391689301,1.84091806411743,1.95615231990814,2.01738262176514,2.07382822036743,2.10458970069885,2.15976548194885,2.21152377128601,2.22626948356628,2.22138667106628,2.20380878448486,2.109375,2.06855463981628,2.05839848518372,2.07294917106628,2.09140658378601,2.19443345069885,2.34335947036743,2.38916015625,2.36328125,2.28720688819885,2.23085927963257,1.98037135601044,1.85761737823486,1.60000014305115,1.56142580509186,1.50136709213257,1.42675769329071,1.39970719814301,1.39150369167328,1.36484396457672,1.31738293170929,1.28046905994415,1.23466789722443,1.17871117591858,1.14580094814301,1.14550769329071,1.13554680347443,1.09755861759186,1.08154320716858,1.08457016944885,1.06230449676514,1.01386713981628,0.985058844089508,0.958007752895355,0.924609541893005,0.900488495826721,0.642968654632568,0.549121022224426,0.492675513029099,0.490722894668579,0.484179675579071,0.159277111291885,0,-0.0686036720871925,-0.299462705850601,-0.31254905462265,-0.345751971006393,-0.395605593919754,-0.430322259664536,-0.453515589237213,-0.491699188947678,-0.545214593410492,-0.597656309604645,-0.627148687839508,-0.648534953594208,-0.701416075229645,-0.771582186222076,-0.902929902076721,-0.961816251277924,-1.04248070716858,-1.23261725902557,-1.53676736354828,-1.58647477626801,-1.59965801239014,-1.90063464641571,-2.23193359375,-2.50917983055115,-2.75166010856628,-2.75209975242615,-2.82993125915527,-2.83857464790344,-2.90732431411743,-2.91489243507385,-2.87841796875,-2.83720684051514,-2.79116225242615,-2.78662085533142,-2.82343745231628,-2.8203125,-2.77709937095642,-2.76650381088257,-2.78847670555115,-2.78320336341858,-2.75073266029358,-2.74980497360229,-2.78051781654358,-2.76596689224243,-2.70620131492615,-2.69584941864014,-2.71718764305115,-2.76660180091858,-2.81674814224243,-2.87514638900757,-2.90087914466858,-2.94814467430115,-2.98828148841858,-3.04262709617615,-3.09580087661743,-3.16069340705872,-3.22353506088257,-3.28969740867615,-3.38627910614014,-3.58115220069885,-3.79062533378601,-3.87763667106628,-3.96347689628601,-4.18115186691284,-4.26718759536743,-4.33222675323486,-4.40620136260986,-4.48027324676514,-4.526611328125,-4.62583017349243,-4.72177743911743,-4.814453125,-4.88271474838257,-4.96992206573486,-4.99404287338257,-5.04931688308716,-5.09985399246216,-5.17529296875,-5.26230430603027,-5.38227558135986,-5.46127986907959,-5.52353477478027,-5.50703144073486,-5.47900390625,-5.47568368911743,-5.45707988739014,-5.46855401992798,-5.490478515625,-5.42421913146973,-5.347412109375,-5.29985332489014,-5.250244140625,-5.22939443588257,-5.24477529525757,-5.27031278610229,-5.29052686691284,-5.302001953125,-5.28813457489014,-5.23017597198486,-5.15751981735229,-5.10590839385986,-4.968994140625,-4.79794931411743,-4.69931602478027,-4.62724590301514,-4.5869140625,-4.54604482650757,-4.47988319396973,-4.4287109375,-4.42158174514771,-4.42192363739014,-4.45986318588257,-4.48061513900757,-4.22709989547729,-4.22524404525757,-4.26064491271973,-4.31025362014771,-4.32871103286743,-4.25869178771973,-4.19619178771973,-4.15102529525757,-4.05117225646973,-3.94731402397156,-3.85346674919128,-3.57578134536743,-3.52763700485229,-3.46992182731628,-3.396728515625,-3.30175757408142,-3.26674795150757,-3.27016592025757,-3.24863290786743,-3.19843769073486,-3.03867197036743,-2.99721670150757,-2.95082998275757,-2.91708970069885,-2.91850614547729,-2.92587876319885,-2.87392592430115,-2.77885723114014,-2.58671879768372,-2.52690434455872,-2.45722627639771,-2.11323213577271,-2.05712866783142,-1.97304654121399,-1.87978529930115,-1.76777362823486,-1.69506812095642,-1.65732395648956,-1.49365222454071,-1.20498073101044,-1.04956066608429,-1.01918935775757,-0.907958805561066,-0.760449111461639,-0.666455328464508,-0.536523401737213,-0.454492479562759,-0.432275474071503,-0.405419766902924,-0.235888451337814,0,0.0073241195641458,0.217480450868607],[91.94921875,91.8888626098633,91.8594741821289,91.8732452392578,91.8570251464844,91.90771484375,91.9339828491211,91.9486312866211,91.9619140625,91.94921875],[91.8738327026367,91.8375930786133,91.8197250366211,91.8351593017578,91.8506851196289,91.861328125,91.8825149536133,91.8738327026367],[91.1507797241211,91.0447311401367,91.0794906616211,91.1583023071289,91.17822265625,91.1507797241211],[91.5567398071289,91.5104522705078,91.4668960571289,91.4113311767578,91.4388732910156,91.4560546875,91.4839859008789,91.5230484008789,91.54833984375,91.5567398071289],[90.7776412963867,90.6036148071289,90.5150375366211,90.6804656982422,90.6749038696289,90.6492156982422,90.56494140625,90.5603485107422,90.5225601196289,90.5029296875,90.5964813232422,90.6722640991211,90.6830062866211,90.69921875,90.7369079589844,90.8681640625,90.8658142089844,90.8298873901367,90.7776412963867],[90.6417922973633,90.6595764160156,90.6039047241211,90.5623016357422,90.5363311767578,90.5798873901367,90.6417922973633],[92.5749053955078,92.5828094482422,92.5842819213867,92.5934600830078,92.6252975463867,92.6316375732422,92.5998077392578,92.5685577392578,92.5391616821289,92.4718780517578,92.3726501464844,92.33056640625,92.2796859741211,92.2082061767578,92.1795883178711,92.1919937133789,92.2147445678711,92.2644577026367,92.2684555053711,92.2862319946289,92.3119125366211,92.3241195678711,92.3078155517578,92.2481460571289,92.1946334838867,92.0560531616211,92.0109405517578,92.0080108642578,91.9131851196289,91.8499984741211,91.8248062133789,91.8578109741211,91.8633804321289,91.8454132080078,91.7970733642578,91.7340850830078,91.6929702758789,91.5296859741211,91.4821243286133,91.4800796508789,91.4095687866211,91.3137664794922,91.2162170410156,91.1513671875,90.9456100463867,90.8267593383789,90.65625,90.6335906982422,90.6560592651367,90.6156311035156,90.6161117553711,90.60400390625,90.5734405517578,90.5616226196289,90.5680694580078,90.5556640625,90.4080123901367,90.2691421508789,90.3915023803711,90.5227508544922,90.5909194946289,90.5992202758789,90.5951156616211,90.5277404785156,90.4660110473633,90.4775390625,90.55224609375,90.4616241455078,90.4369125366211,90.43505859375,90.4806671142578,90.4984359741211,90.4874038696289,90.53173828125,90.5955047607422,90.6161117553711,90.5894546508789,90.5528335571289,90.494140625,90.3557586669922,90.2881851196289,90.2305679321289,90.1587905883789,90.1307601928711,90.0711898803711,90.0700225830078,90.087890625,90.2095718383789,90.1434555053711,90.0685577392578,89.9542007446289,89.9180679321289,89.89404296875,89.8938446044922,89.9851531982422,89.8818359375,89.8532180786133,89.8658218383789,89.8525390625,89.8119125366211,89.7568359375,89.6677780151367,89.6281280517578,89.5685501098633,89.5665969848633,89.5474624633789,89.4832000732422,89.4693374633789,89.5025405883789,89.5005874633789,89.4519577026367,89.3537063598633,89.2786102294922,89.2342758178711,89.1670913696289,89.0939407348633,89.0816421508789,89.0514678955078,89.0558624267578,89.0499954223633,88.9714889526367,88.9207000732422,88.9269561767578,88.9234313964844,88.8997039794922,88.8669891357422,88.8505859375,88.9281234741211,88.8970718383789,88.8076171875,88.7244186401367,88.7040023803711,88.7408218383789,88.6976547241211,88.6357421875,88.6164016723633,88.5959930419922,88.5673828125,88.62255859375,88.6998062133789,88.7137680053711,88.7265625,88.7335968017578,88.7235336303711,88.6422805786133,88.49853515625,88.39697265625,88.3375015258789,88.287109375,88.2249984741211,88.1455078125,88.0791015625,88.0234375,88.0302734375,88.0451202392578,88.1498031616211,88.1888656616211,88.2794952392578,88.3133773803711,88.3729476928711,88.4562530517578,88.5738220214844,88.6775436401367,88.74755859375,88.8172836303711,88.89013671875,88.9297866821289,88.95166015625,88.9441375732422,88.8547821044922,88.8203125,88.79541015625,88.7691421508789,88.5934524536133,88.50244140625,88.4523468017578,88.3630828857422,88.2529296875,88.1474609375,88.1066436767578,88.0845718383789,88.0973663330078,88.1290054321289,88.1507797241211,88.2351531982422,88.333984375,88.3780212402344,88.4404296875,88.4478454589844,88.4367141723633,88.38623046875,88.3514709472656,88.3459014892578,88.3699264526367,88.4181671142578,88.5182647705078,88.6201171875,88.6806640625,88.6828155517578,88.72216796875,88.7619171142578,88.8280334472656,88.896484375,88.9407196044922,88.9704132080078,88.9815444946289,88.9482421875,88.9241180419922,88.9519500732422,88.9833984375,89.0186538696289,89.0667953491211,89.1019592285156,89.1082992553711,89.1864242553711,89.2892608642578,89.3697280883789,89.4668960571289,89.5498962402344,89.5914077758789,89.57275390625,89.5857467651367,89.6190414428711,89.6708984375,89.7098617553711,89.8229522705078,89.7996139526367,89.8249053955078,89.7962951660156,89.8008804321289,89.8140640258789,89.8332977294922,89.8663101196289,90.0038070678711,90.11962890625,90.2503890991211,90.4393539428711,90.5552749633789,90.6130905151367,90.7301788330078,91.0382766723633,91.2931594848633,91.3966827392578,91.4796905517578,91.7634735107422,92.0497055053711,92.2046813964844,92.3734359741211,92.4683609008789,92.4854431152344,92.4750061035156,92.4431610107422,92.3849639892578,92.2512664794922,92.2283172607422,92.2305679321289,92.2266540527344,92.1980438232422,92.1174774169922,92.1019515991211,92.0850601196289,92.0641632080078,92.0010757446289,91.95166015625,91.9310531616211,91.8990173339844,91.876953125,91.84619140625,91.7724609375,91.7265625,91.6687469482422,91.6111373901367,91.5713882446289,91.5263671875,91.3926773071289,91.3670959472656,91.3501968383789,91.33642578125,91.2320327758789,91.1924819946289,91.1604537963867,91.16552734375,91.2538070678711,91.3152389526367,91.3388671875,91.359375,91.36865234375,91.3667984008789,91.37060546875,91.3994140625,91.4362258911133,91.4713897705078,91.51123046875,91.5535125732422,91.6195297241211,91.6949234008789,91.7509765625,91.7738265991211,91.7579116821289,91.7542037963867,91.7900390625,91.9191436767578,91.9378890991211,91.9294891357422,91.9295959472656,91.978515625,92.0440444946289,92.1270523071289,92.15234375,92.1871109008789,92.24609375,92.2893600463867,92.3341751098633,92.3337860107422,92.3412170410156,92.3616256713867,92.3931655883789,92.4304733276367,92.4644546508789,92.4914093017578,92.5095748901367,92.5318374633789,92.5612258911133,92.5749053955078],[28.5853500366211,28.5618171691895,28.4654293060303,28.3196296691895,28.1336917877197,28.03515625,27.9792957305908,27.9289073944092,27.896484375,27.8888683319092,27.818359375,27.7537097930908,27.4847640991211,27.6395511627197,27.7082023620605,27.8213882446289,27.9827156066895,28.0144519805908,27.8791999816895,27.8319339752197,27.8016605377197,27.7388668060303,27.6611328125,27.5798816680908,27.5348625183105,27.4748039245605,27.3628902435303,27.294921875,27.2443351745605,27.193359375,27.01171875,26.96875,26.8848628997803,26.8003902435303,26.67919921875,26.6153335571289,26.5796871185303,26.5497074127197,26.529296875,26.5114250183105,26.3603515625,26.3272457122803,26.3179683685303,26.3208980560303,26.2005844116211,26.107421875,26.0855484008789,26.0660152435303,26.0769538879395,26.1112308502197,26.1435546875,26.1551761627197,26.1353511810303,26.06640625,25.92333984375,25.7849617004395,25.7239246368408,25.6214847564697,25.5270500183105,25.3819332122803,25.2511711120605,25.1333980560303,24.9935550689697,24.8468761444092,24.7958011627197,24.7737293243408,24.6510753631592,24.5959968566895,24.5693359375,24.5182609558105,24.4878921508789,24.38671875,24.2894535064697,24.2303714752197,24.0560550689697,24.0329093933105,24.0113277435303,23.9735355377197,23.880859375,23.7623043060303,23.6351566314697,23.5358390808105,23.4333992004395,23.3720703125,23.2398433685303,23.1559562683105,23.0255851745605,22.916015625,22.9296875,22.9514636993408,23.0056629180908,23.0036125183105,22.9919929504395,22.9439449310303,22.9091796875,22.8368167877197,22.7960948944092,22.6823234558105,22.5827140808105,22.4982414245605,22.3440437316895,22.4220695495605,22.4457015991211,22.5235366821289,22.5324211120605,22.5242195129395,22.4720706939697,22.4362297058105,22.4632816314697,22.4656238555908,22.4392585754395,22.466796875,22.5227527618408,22.55810546875,22.7061519622803,22.7999019622803,22.8568344116211,22.9152355194092,22.9422836303711,22.9679698944092,22.9768562316895,22.8595714569092,22.8197269439697,22.767578125,22.6969738006592,22.5545883178711,22.4991207122803,22.47412109375,22.4363269805908,22.3948249816895,22.3869152069092,22.36962890625,22.3654308319092,22.3990230560303,22.4208011627197,22.4690437316895,22.5974597930908,22.6034183502197,22.6265621185303,22.66748046875,22.705078125,22.7751960754395,22.9454116821289,23.0285148620605,23.0244140625,22.9853496551514,22.9113273620605,22.8682613372803,22.8564453125,22.86767578125,22.9190425872803,23.224609375,23.5345706939697,23.9507808685303,24.2267570495605,24.4305648803711,24.8082027435303,25.15966796875,25.4970703125,25.6861343383789,25.81884765625,25.9333992004395,26.2158203125,26.4892578125,26.8477535247803,27.0869140625,27.1207027435303,27.4253902435303,27.56103515625,27.6708984375,27.7107410430908,27.7385749816895,27.88427734375,27.9489250183105,28.0500011444092,28.2219734191895,28.3751945495605,28.4234371185303,28.5853500366211],[50.6072273254395,50.5749015808105,50.5440406799316,50.4659194946289,50.4894523620605,50.4524421691895,50.4699211120605,50.5640602111816,50.5859375,50.5578117370605,50.6097640991211,50.6174812316895,50.6072273254395],[-73.02685546875,-73.0587463378906,-73.16455078125,-73.4007797241211,-73.6610336303711,-73.68115234375,-73.6868209838867,-73.6678237915039,-73.6695785522461,-73.6803741455078,-73.5850601196289,-73.5230941772461,-73.4245147705078,-73.3015594482422,-73.2353515625,-73.1373062133789,-73.0584945678711,-73.0116653442383,-73.02685546875],[-72.9161148071289,-73.04931640625,-73.0626983642578,-72.9947814941406,-72.9161148071289],[-73.041015625,-72.9789505004883,-72.9452133178711,-72.8307647705078,-72.7625961303711,-72.7472686767578,-72.7838897705078,-72.88916015625,-72.9810562133789,-73.1102066040039,-73.1619110107422,-73.1273956298828,-73.041015625],[-74.2067413330078,-74.2769088745117,-74.2613296508789,-74.1267623901367,-74.0523452758789,-74.0100555419922,-73.9949722290039,-73.9359817504883,-73.9063949584961,-73.91455078125,-73.9763641357422,-73.9754943847656,-73.9542007446289,-73.8499526977539,-73.8774948120117,-73.8365249633789,-73.974609375,-74.0929260253906,-74.2067413330078],[-74.0575180053711,-74.0347595214844,-74.0985870361328,-74.2422332763672,-74.2746124267578,-74.3031311035156,-74.31396484375,-74.3070297241211,-74.2214889526367,-74.1753921508789,-74.0575180053711],[-74.8404769897461,-74.8468704223633,-74.9733428955078,-75.1321258544922,-75.2233352661133,-75.2043914794922,-75.14111328125,-75.1305618286133,-75.1575698852539,-75.2412643432617,-75.2882308959961,-75.309814453125,-75.3159713745117,-75.2166061401367,-75.17529296875,-75.1087875366211,-75.064208984375,-74.9371032714844,-74.8456115722656,-74.8404769897461],[-75.66455078125,-75.7063522338867,-75.781005859375,-75.9559555053711,-76.037109375,-76.0104522705078,-75.9486312866211,-75.8075180053711,-75.7542495727539,-75.66455078125],[-74.429443359375,-74.5086898803711,-74.5509262084961,-74.5269088745117,-74.4720230102539,-74.4504928588867,-74.429443359375],[-77.65771484375,-77.6561508178711,-77.7552795410156,-77.6832504272461,-77.6153793334961,-77.5615234375,-77.5320281982422,-77.5368194580078,-77.5318832397461,-77.5213394165039,-77.5187454223633,-77.57373046875,-77.7712860107422,-77.7757797241211,-77.8062973022461,-77.8522415161133,-77.9140625,-77.9999008178711,-77.9500427246094,-77.8835983276367,-77.8495635986328,-77.7574234008789,-77.7014694213867,-77.65771484375],[-75.3083953857422,-75.3017578125,-75.3687438964844,-75.4676284790039,-75.5032196044922,-75.4810562133789,-75.4124069213867,-75.408935546875,-75.493896484375,-75.5927734375,-75.6390686035156,-75.6610412597656,-75.7439956665039,-75.7266616821289,-75.7096176147461,-75.6535186767578,-75.5264663696289,-75.5181655883789,-75.3083953857422],[-77.3475646972656,-77.4605026245117,-77.5412139892578,-77.5619125366211,-77.52734375,-77.4512710571289,-77.3291015625,-77.2755889892578,-77.2691345214844,-77.3475646972656],[-77.7438507080078,-77.7460403442383,-77.735107421875,-77.7452163696289,-77.8534164428711,-77.8812026977539,-77.9832077026367,-78.044921875,-78.0758285522461,-78.1357421875,-78.1457977294922,-78.1916046142578,-78.2576141357422,-78.3665008544922,-78.435302734375,-78.3389129638672,-78.3189926147461,-78.2427291870117,-78.2600631713867,-78.2738342285156,-78.298828125,-78.18408203125,-78.1593246459961,-78.2113723754883,-78.1627960205078,-78.0333023071289,-77.9752883911133,-77.973388671875,-77.9189453125,-77.8401336669922,-77.7438507080078],[-76.6488265991211,-76.4842224121094,-76.3437957763672,-76.1919860839844,-76.1266098022461,-76.1149368286133,-76.1405258178711,-76.1746597290039,-76.1695327758789,-76.2051773071289,-76.2412109375,-76.30029296875,-76.3199768066406,-76.2137680053711,-76.204345703125,-76.1525421142578,-76.160400390625,-76.2843246459961,-76.3692855834961,-76.4999008178711,-76.6206970214844,-76.6927719116211,-76.7806625366211,-76.7489242553711,-76.7269515991211,-76.7108383178711,-76.6488265991211],[-78.4928665161133,-78.3717269897461,-78.3068313598633,-78.2679214477539,-78.0886688232422,-77.9439468383789,-77.9224624633789,-77.9261169433594,-78.23388671875,-78.5162124633789,-78.6709442138672,-78.74365234375,-78.7992172241211,-78.9856491088867,-78.935791015625,-78.7980499267578,-78.7125015258789,-78.6332473754883,-78.6211395263672,-78.6329574584961,-78.5971145629883,-78.4928665161133],[-77.2256393432617,-77.2464370727539,-77.333251953125,-77.4031753540039,-77.2939453125,-77.2467727661133,-77.2477493286133,-77.2210922241211,-77.2301254272461,-77.2060546875,-77.2386169433594,-77.3299331665039,-77.5105972290039,-77.7959976196289,-77.9437484741211,-77.862548828125,-77.7875518798828,-77.672119140625,-77.5338897705078,-77.4494171142578,-77.3687515258789,-77.2958984375,-77.2659225463867,-77.269287109375,-77.2571792602539,-77.162109375,-77.0663528442383,-77.0382843017578,-77.1672897338867,-77.1910171508789,-77.2256393432617],[19.0071296691895,19.0420894622803,19.1315422058105,19.23681640625,19.3126945495605,19.3486328125,19.3568363189697,19.33447265625,19.2918949127197,19.22314453125,19.1513671875,19.1324214935303,19.1273441314697,19.1184577941895,19.1283206939697,19.15185546875,19.2315425872803,19.3388671875,19.43017578125,19.5471687316895,19.5837898254395,19.5836906433105,19.5495128631592,19.4494152069092,19.34521484375,19.3052730560303,19.26806640625,19.2450199127197,19.2572269439697,19.3640632629395,19.4881839752197,19.4951171875,19.47998046875,19.4512691497803,19.3996086120605,19.3603515625,19.30078125,19.2544918060303,19.1943359375,19.1643543243408,19.11279296875,19.0800762176514,19.0283203125,18.9742202758789,18.9506816864014,18.9402332305908,18.9738292694092,19.0367183685303,19.0266590118408,18.9787101745605,18.9346675872803,18.8956050872803,18.85107421875,18.7492198944092,18.6742191314697,18.6568355560303,18.6299819946289,18.6218757629395,18.6236324310303,18.4884777069092,18.4601573944092,18.44384765625,18.455078125,18.4660148620605,18.5432624816895,18.5458984375,18.5349617004395,18.4800777435303,18.4539051055908,18.4363288879395,18.3465805053711,18.3040027618408,18.1239261627197,18.0445308685303,17.9188480377197,17.84130859375,17.8019542694092,17.740234375,17.6675777435303,17.5851573944092,17.6434574127197,17.6578140258789,17.6504878997803,17.6248054504395,17.4022464752197,17.2930679321289,17.2752933502197,17.2738285064697,17.248046875,17.0845699310303,16.90185546875,16.7134761810303,16.6876964569092,16.5905284881592,16.4720706939697,16.3775386810303,16.3000984191895,16.2142581939697,16.1698246002197,16.1302738189697,16.1034183502197,16.0490226745605,15.8800792694092,15.7366209030151,15.7379884719849,15.7615242004395,15.7880868911743,15.8228521347046,15.8882818222046,15.9631834030151,16.0283203125,16.1573238372803,16.2310543060303,16.2933597564697,16.3650398254395,16.4535160064697,16.5306644439697,16.7908191680908,16.9186515808105,17.1253890991211,17.2106437683105,17.2586917877197,17.3241214752197,17.4691410064697,17.5026359558105,17.5462894439697,17.6535148620605,17.6901378631592,17.8127937316895,17.8744144439697,17.9486331939697,17.9962902069092,18.13720703125,18.2179679870605,18.2849617004395,18.3576164245605,18.4239273071289,18.48828125,18.66259765625,18.74609375,18.7793941497803,18.7801761627197,18.7883777618408,18.83642578125,18.9413089752197,19.0071296691895],[-62.8319358825684,-62.8469276428223,-62.8589363098145,-62.869384765625,-62.8754425048828,-62.8742218017578,-62.8654289245605,-62.7997093200684,-62.8070335388184,-62.8181648254395,-62.8319358825684],[28.14794921875,28.2842769622803,28.3163070678711,28.39208984375,28.4070320129395,28.5639629364014,28.6369132995605,28.6908206939697,28.7408218383789,28.7947254180908,28.9474620819092,29.0317401885986,29.08740234375,29.2830085754395,29.3750019073486,29.3960933685303,29.3979511260986,29.3731422424316,29.3534183502197,29.4129867553711,29.4822254180908,29.6300773620605,29.6845684051514,29.7441425323486,29.8239250183105,29.8816413879395,29.9370136260986,30.0426750183105,30.2335929870605,30.4562511444092,30.4753913879395,30.5867176055908,30.6255855560303,30.6623039245605,30.7216777801514,30.8007831573486,30.85595703125,30.8822250366211,30.9068355560303,30.9087886810303,30.9005851745605,30.86181640625,30.8209972381592,30.810546875,30.814453125,30.87744140625,30.9588871002197,30.9777355194092,30.9777355194092,30.8667964935303,30.8298816680908,30.8044910430908,30.791015625,30.798828125,30.9841804504395,31.1212882995605,31.1521472930908,31.1548824310303,31.0819339752197,31.0748062133789,31.1847648620605,31.2455062866211,31.2991199493408,31.4036140441895,31.6284160614014,31.7919940948486,31.8259773254395,31.8377933502197,31.8252944946289,31.7830085754395,31.7541980743408,31.8207988739014,31.9921855926514,32.2003898620605,32.4501953125,32.4509773254395,32.4251976013184,32.4423828125,32.4696311950684,32.6857452392578,32.7064476013184,32.7102546691895,32.7042999267578,32.6444358825684,32.5780296325684,32.4693374633789,32.42626953125,32.2506866455078,32.1419906616211,32.0554695129395,31.8497085571289,31.7774410247803,31.7474594116211,31.6682605743408,31.5629863739014,31.4178695678711,31.3883800506592,31.3645496368408,31.3029308319092,31.2587871551514,31.2951183319092,31.35302734375,31.4427719116211,31.5351581573486,31.5648441314697,31.5634746551514,31.5194339752197,31.5261726379395,31.6159172058105,31.5855464935303,31.5765628814697,31.5773448944092,31.6015605926514,31.6499004364014,31.6906261444092,31.7585926055908,31.7633781433105,31.57373046875,31.3459949493408,31.2179698944092,31.1684589385986,31.0792980194092,30.9806613922119,30.8457050323486,30.7552738189697,30.6672859191895,30.6394538879395,30.5838871002197,30.5330085754395,30.5607414245605,30.6023445129395,30.6117191314697,30.6325206756592,30.5769538879395,30.5445308685303,30.4495124816895,30.3333988189697,30.3089847564697,30.2195301055908,30.1607418060303,30.0637702941895,29.9087905883789,29.7060527801514,29.5531253814697,29.4696292877197,29.3464851379395,29.298828125,29.2304668426514,29.1742172241211,29.1356430053711,29.1020488739014,29.0607414245605,29.0130863189697,28.9777336120605,28.9275379180908,28.8495140075684,28.7932605743408,28.7312488555908,28.6902332305908,28.6477527618408,28.5990238189697,28.5320301055908,28.4246082305908,28.2916011810303,28.1837882995605,28.1444339752197,28.0802745819092,28.0107421875,27.8585948944092,27.8288078308105,27.7888679504395,27.7413082122803,27.6999988555908,27.6767578125,27.6897449493408,27.6013679504395,27.4523429870605,27.34765625,27.2962894439697,27.2701168060303,27.1419925689697,27.0741214752197,26.9528312683105,26.7734375,26.56689453125,26.4534187316895,26.3943347930908,26.26708984375,25.92529296875,25.7857418060303,25.5802745819092,25.2671871185303,25.0666980743408,24.9738292694092,24.8664073944092,24.6851558685303,24.611328125,24.4952144622803,24.3619136810303,24.32373046875,24.2800769805908,24.1268558502197,23.9783191680908,23.951171875,23.8642578125,23.7916984558105,23.7068347930908,23.6466808319092,23.6085929870605,23.6137714385986,23.6052742004395,23.5396480560303,23.5448226928711,23.5813484191895,23.6256847381592,23.607421875,23.6510734558105,23.6524410247803,23.63330078125,23.5979480743408,23.5011711120605,23.4583988189697,23.3271484375,23.1969718933105,23.1750984191895,23.1812496185303,23.2041015625,23.3033218383789,23.4109382629395,23.4795894622803,23.8447265625,23.9012699127197,23.9154300689697,23.9163093566895,23.9093761444092,23.8871097564697,23.8591785430908,23.7892589569092,23.5989246368408,23.4846687316895,23.55908203125,23.7336921691895,23.87255859375,23.9444332122803,24.0084953308105,24.1039066314697,24.1913089752197,24.2366199493408,24.3179683685303,24.478515625,24.6207027435303,24.7681655883789,24.7892589569092,24.82568359375,24.8695316314697,25.0460929870605,25.1114253997803,25.1794929504395,25.28369140625,25.37060546875,25.4611339569092,25.5056648254395,25.52734375,25.4973621368408,25.5103511810303,25.5730457305908,25.6805667877197,25.7492198944092,25.7652339935303,25.7650394439697,25.7481441497803,25.7025394439697,25.6168937683105,25.5575199127197,25.54736328125,25.5675792694092,25.6203136444092,25.6851577758789,25.7248039245605,25.7316398620605,25.7239246368408,25.7224617004395,25.7808589935303,25.8592777252197,25.9644527435303,26.0929679870605,26.1751956939697,26.2158203125,26.2312488555908,26.2507801055908,26.2917976379395,26.6011714935303,26.6484375,26.6749992370605,26.734375,26.7756843566895,26.7601547241211,26.6812496185303,26.4953136444092,26.4576168060303,26.4695301055908,26.5192375183105,26.5666007995605,26.5908203125,26.5935554504395,26.6202144622803,26.7718753814697,26.8224620819092,26.9530277252197,27.0525379180908,27.3091793060303,27.4271488189697,27.4591808319092,27.5767574310303,27.5894527435303,27.6422843933105,27.6942386627197,27.8962898254395,28.0320301055908,28.1178703308105,28.14794921875],[-87.8529357910156,-87.9299774169922,-87.9348602294922,-87.90283203125,-87.8594207763672,-87.83251953125,-87.826416015625,-87.7886276245117,-87.7981491088867,-87.8529357910156],[-87.9505844116211,-87.9980926513672,-87.9590301513672,-87.9533233642578,-87.8983383178711,-87.8589324951172,-87.8485336303711,-87.9505844116211],[-88.89404296875,-88.9371566772461,-89.1135711669922,-89.2328186035156,-89.2375030517578,-89.2276382446289,-89.2124557495117,-89.2012634277344,-89.1903762817383,-89.1821746826172,-89.1710891723633,-89.1614761352539,-89.162353515625,-89.133544921875,-89.0504455566406,-88.942626953125,-88.8978042602539,-88.8573760986328,-88.8063430786133,-88.7436065673828,-88.586181640625,-88.5230026245117,-88.4612731933594,-88.3724060058594,-88.295654296875,-88.3492660522461,-88.295654296875,-88.2472686767578,-88.1302719116211,-88.0852508544922,-88.0972137451172,-88.2074737548828,-88.221435546875,-88.271728515625,-88.2034683227539,-88.2671890258789,-88.288818359375,-88.2939910888672,-88.2618179321289,-88.3134307861328,-88.404541015625,-88.4611358642578,-88.5623016357422,-88.6951599121094,-88.8790969848633,-88.9117202758789,-88.89404296875],[-64.7302780151367,-64.8201141357422,-64.8450698852539,-64.8628463745117,-64.7711868286133,-64.6946334838867,-64.6683120727539,-64.7302780151367],[-58.1597633361816,-58.1399421691895,-58.1600570678711,-58.1801719665527,-58.4742202758789,-58.7411155700684,-59.0905265808105,-59.5408706665039,-60.0073699951172,-60.4516143798828,-60.8887672424316,-61.0959930419922,-61.5118179321289,-61.7568359375,-61.8208961486816,-61.9169425964355,-62.0118141174316,-62.1216278076172,-62.2763175964355,-62.2765121459961,-62.2766609191895,-62.3854522705078,-62.4778289794922,-62.5669441223145,-62.6285171508789,-62.6509742736816,-62.665283203125,-62.7445831298828,-62.8150863647461,-62.8342781066895,-62.8433609008789,-63.2671890258789,-63.6753387451172,-63.7169418334961,-63.7755851745605,-63.8186492919922,-63.8610305786133,-63.9216766357422,-63.9761199951172,-64.1318359375,-64.2090835571289,-64.2664031982422,-64.3079147338867,-64.3252944946289,-64.3739700317383,-64.4455032348633,-64.4777374267578,-64.5236358642578,-64.6055145263672,-64.7001037597656,-64.7586441040039,-64.8430633544922,-64.9926223754883,-65.0578079223633,-65.48486328125,-65.518798828125,-65.6861801147461,-65.7710494995117,-65.8601531982422,-66.05859375,-66.0985794067383,-66.1746597290039,-66.2201690673828,-66.2476119995117,-66.2821273803711,-66.3224639892578,-66.3651809692383,-66.5069808959961,-66.6390151977539,-66.7117233276367,-66.7506332397461,-66.7674865722656,-66.80029296875,-66.9911117553711,-67.0335464477539,-67.055419921875,-67.1619110107422,-67.1948776245117,-67.3622589111328,-67.5799331665039,-67.7073211669922,-67.79443359375,-67.8205032348633,-67.8794479370117,-67.88916015625,-67.8899917602539,-67.8737258911133,-67.8817367553711,-67.9503860473633,-67.9449157714844,-67.9539108276367,-67.9883804321289,-68.0767517089844,-68.101806640625,-68.1121520996094,-68.1864242553711,-68.1985397338867,-68.197021484375,-68.3138656616211,-68.4354934692383,-68.5338363647461,-68.5582504272461,-68.5689468383789,-68.571044921875,-68.5631790161133,-68.4873962402344,-68.4843292236328,-68.4998474121094,-68.69580078125,-68.7451629638672,-68.7605438232422,-68.7592239379883,-68.7123031616211,-68.6885757446289,-68.7345733642578,-68.7300262451172,-68.7593231201172,-68.7558135986328,-68.7274856567383,-68.6001968383789,-68.5606918334961,-68.5593795776367,-68.5782699584961,-68.6961898803711,-68.6982879638672,-68.5752868652344,-68.4870071411133,-68.462890625,-68.4701690673828,-68.4919891357422,-68.5478515625,-68.6205520629883,-68.6806640625,-68.7590866088867,-68.8579559326172,-68.9310073852539,-68.9683151245117,-68.9690933227539,-68.9788589477539,-69.0268096923828,-69.0393981933594,-69.0601577758789,-69.0809555053711,-69.09228515625,-69.1263732910156,-69.1454620361328,-69.1180648803711,-69.0904312133789,-69.0939483642578,-69.2823257446289,-69.3133850097656,-69.3580093383789,-69.4950180053711,-69.5109405517578,-69.5110778808594,-69.510986328125,-69.5219192504883,-69.5638198852539,-69.6258773803711,-69.6457061767578,-69.6248550415039,-69.5033187866211,-69.4383316040039,-69.4210968017578,-69.3815383911133,-69.2672348022461,-69.1997985839844,-69.1325149536133,-69.0545349121094,-69.0206985473633,-69.0383758544922,-69.0329132080078,-69.0062484741211,-68.9280319213867,-68.8578109741211,-68.8427734375,-68.8488235473633,-68.9134750366211,-69.0462417602539,-69.1341781616211,-69.18798828125,-69.2175827026367,-69.3918914794922,-69.4208984375,-69.4185028076172,-69.3019027709961,-69.2543029785156,-69.1724624633789,-69.1871109008789,-69.3307113647461,-69.3747100830078,-69.3737335205078,-69.3594741821289,-69.2760238647461,-69.2523498535156,-69.2349090576172,-69.1992721557617,-69.1626968383789,-69.1197280883789,-69.0527801513672,-69.0131301879883,-69.0044937133789,-68.9717788696289,-68.8803176879883,-68.8709030151367,-68.8917007446289,-68.9374542236328,-68.9742660522461,-69.0230484008789,-69.0741195678711,-69.0528335571289,-69.0175323486328,-68.9834518432617,-68.9722671508789,-68.9805221557617,-68.9786148071289,-68.9337387084961,-68.86767578125,-68.8118133544922,-68.7590866088867,-68.7628860473633,-68.7281265258789,-68.6852569580078,-68.8186950683594,-68.93603515625,-69.0461883544922,-69.1737289428711,-69.2577133178711,-69.3620147705078,-69.45361328125,-69.57861328125,-69.4625473022461,-69.228515625,-69.0016555786133,-68.8483428955078,-68.7840805053711,-68.7699203491211,-68.7274856567383,-68.6783752441406,-68.6226577758789,-68.4983444213867,-68.3980026245117,-68.3111343383789,-68.2666015625,-68.1586456298828,-68.0716781616211,-67.99169921875,-67.8350067138672,-67.7856903076172,-67.7217788696289,-67.6666488647461,-67.5824203491211,-67.4169464111328,-67.3327178955078,-67.2804718017578,-67.1904754638672,-67.1115264892578,-66.72998046875,-66.5753402709961,-66.4789047241211,-66.3992233276367,-66.2635726928711,-65.9247055053711,-65.706787109375,-65.6371078491211,-65.5586929321289,-65.4919891357422,-65.436767578125,-65.3961410522461,-65.337890625,-65.309326171875,-65.328125,-65.3245544433594,-65.298583984375,-65.3130798339844,-65.3954620361328,-65.4369125366211,-65.4471206665039,-65.4399871826172,-65.4022903442383,-65.3340301513672,-65.3237762451172,-65.37158203125,-65.3936004638672,-65.389892578125,-65.3728561401367,-65.3423843383789,-65.3254852294922,-65.322021484375,-65.2822799682617,-65.2062072753906,-65.1753921508789,-65.1897430419922,-65.1857376098633,-65.1633758544922,-65.1426773071289,-65.1151351928711,-65.0902786254883,-65.037109375,-65.0302734375,-65.001220703125,-64.9925308227539,-64.9143600463867,-64.8298797607422,-64.783447265625,-64.6900405883789,-64.6116714477539,-64.513427734375,-64.4807662963867,-64.4205093383789,-64.2550354003906,-64.0616226196289,-63.9385757446289,-63.7880859375,-63.6885757446289,-63.5856475830078,-63.5418930053711,-63.4652366638184,-63.3466796875,-63.2497596740723,-63.1806640625,-63.1167945861816,-63.0674819946289,-63.0413551330566,-63.0151901245117,-62.9579124450684,-62.8351593017578,-62.7654762268066,-62.6870613098145,-62.5255393981934,-62.3528327941895,-62.263916015625,-62.1760711669922,-62.1180152893066,-62.0947723388672,-61.9447288513184,-61.8741226196289,-61.7899398803711,-61.57568359375,-61.5115699768066,-61.4160652160645,-61.1291465759277,-61.0769996643066,-60.9145011901855,-60.7223625183105,-60.5953140258789,-60.506591796875,-60.4601593017578,-60.4223594665527,-60.4049758911133,-60.4280776977539,-60.4629859924316,-60.4746589660645,-60.4601593017578,-60.396240234375,-60.3727035522461,-60.3380355834961,-60.2988777160645,-60.2733383178711,-60.4020004272461,-60.5832023620605,-60.5304679870605,-60.3804702758789,-60.2423362731934,-60.2204132080078,-60.2066421508789,-60.1872100830078,-60.1755867004395,-59.8311538696289,-59.4342765808105,-58.9572792053223,-58.5379371643066,-58.49658203125,-58.4236831665039,-58.3753929138184,-58.3456077575684,-58.3405723571777,-58.3507843017578,-58.3503875732422,-58.4706077575684,-58.4781265258789,-58.4598159790039,-58.4173851013184,-58.3959922790527,-58.3477516174316,-58.20556640625,-57.9909133911133,-57.9050331115723,-57.8324699401855,-57.7888641357422,-57.7801780700684,-57.6616744995117,-57.5864715576172,-57.5520477294922,-57.4956512451172,-57.5061492919922,-57.5531272888184,-57.5740203857422,-57.63916015625,-57.7250022888184,-57.7831077575684,-57.7308616638184,-57.7286148071289,-57.7168006896973,-57.7814483642578,-57.8003921508789,-57.8745079040527,-57.9716796875,-58.0720252990723,-58.1314926147461,-58.0299301147461,-57.8607444763184,-57.8875961303711,-57.9601593017578,-58.0211448669434,-58.0676231384277,-58.09375,-58.1597633361816],[-48.4858856201172,-48.5545883178711,-48.5421867370605,-48.5051765441895,-48.4647445678711,-48.4148902893066,-48.3779296875,-48.4095687866211,-48.4967765808105,-48.4858856201172],[-48.5844230651855,-48.6030769348145,-48.665771484375,-48.5397453308105,-48.4976081848145,-48.5311050415039,-48.5680656433105,-48.5844230651855],[-45.26025390625,-45.2608871459961,-45.3025360107422,-45.412841796875,-45.451416015625,-45.3023414611816,-45.2722663879395,-45.2490692138672,-45.2331047058105,-45.2502937316895,-45.26025390625],[-44.1292953491211,-44.0980491638184,-44.15576171875,-44.2205085754395,-44.320068359375,-44.3601570129395,-44.2741241455078,-44.2428703308105,-44.2204093933105,-44.1916007995605,-44.1292953491211],[-38.903564453125,-38.9378890991211,-38.9775886535645,-38.9932136535645,-39.0221672058105,-39.006591796875,-38.9801292419434,-38.9071273803711,-38.903564453125],[-38.7438468933105,-38.7830085754395,-38.7869644165039,-38.6848640441895,-38.6681137084961,-38.6145477294922,-38.6002922058105,-38.6011734008789,-38.7438468933105],[-44.4993171691895,-44.5977554321289,-44.5653305053711,-44.5818862915039,-44.569091796875,-44.501953125,-44.4814453125,-44.4873046875,-44.4825706481934,-44.4993171691895],[-44.8831062316895,-44.9471168518066,-44.9678726196289,-45.0208511352539,-45.01123046875,-44.99560546875,-44.9786643981934,-44.8882789611816,-44.8831062316895],[-51.83251953125,-51.9383773803711,-51.8020515441895,-51.6800270080566,-51.6782684326172,-51.5460433959961,-51.4244651794434,-51.2540016174316,-51.1607437133789,-51.2763175964355,-51.3101043701172,-51.4651374816895,-51.6376953125,-51.83251953125],[-49.628662109375,-49.5352058410645,-49.4028816223145,-49.3142585754395,-49.215087890625,-49.1169929504395,-48.7865715026855,-48.5880355834961,-48.5154304504395,-48.4444808959961,-48.3926773071289,-48.3796882629395,-48.4280281066895,-48.4639663696289,-48.4974594116211,-48.5233421325684,-48.566650390625,-48.5396995544434,-48.5495109558105,-48.5709457397461,-48.6240730285645,-48.70458984375,-48.728515625,-48.7898445129395,-48.8396987915039,-48.8290061950684,-48.8040504455566,-48.8335952758789,-48.9289054870605,-48.9859390258789,-49.0384788513184,-49.0868644714355,-49.1727027893066,-49.1816902160645,-49.2047843933105,-49.2339859008789,-49.3448257446289,-49.4065933227539,-49.5066413879395,-49.5256843566895,-49.587890625,-49.6505851745605,-49.748779296875,-49.8051300048828,-49.9113273620605,-50.0099639892578,-50.0657234191895,-50.1092758178711,-50.3384246826172,-50.4434585571289,-50.5076179504395,-50.60205078125,-50.6171875,-50.6733894348145,-50.7238311767578,-50.759765625,-50.7294921875,-50.6683120727539,-50.5959014892578,-50.5805168151855,-50.5769538879395,-50.5929183959961,-50.7096176147461,-50.7832984924316,-50.7960929870605,-50.7809562683105,-50.7713890075684,-50.7199211120605,-50.7030754089355,-50.7158203125,-50.6937026977539,-50.6455078125,-50.4615707397461,-50.2482452392578,-49.628662109375],[-50.6528816223145,-50.9263687133789,-51.0189933776855,-51.0380859375,-51.0223617553711,-51.0257301330566,-50.9950714111328,-50.8421859741211,-50.7652816772461,-50.6669921875,-50.6505851745605,-50.6528816223145],[-49.4438972473145,-49.7088394165039,-49.830078125,-49.8026847839355,-49.7123069763184,-49.6021957397461,-49.5034675598145,-49.4004859924316,-49.372314453125,-49.380859375,-49.4438972473145],[-49.7382316589355,-49.697265625,-49.8389663696289,-49.9170875549316,-50.0024909973145,-50.1131362915039,-50.2855949401855,-50.3394546508789,-50.3451156616211,-50.2726593017578,-50.1279792785645,-49.8790016174316,-49.7382316589355],[-50.4261207580566,-50.4439468383789,-50.6239242553711,-50.6104469299316,-50.5262222290039,-50.4515609741211,-50.4260749816895,-50.424560546875,-50.3968734741211,-50.3727531433105,-50.3509750366211,-50.342529296875,-50.332275390625,-50.4261207580566],[-50.1529273986816,-50.2613258361816,-50.2815437316895,-50.2816886901855,-50.2511711120605,-50.11279296875,-50.0986328125,-50.058837890625,-50.0368156433105,-50.0400390625,-50.1529273986816],[-50.2989730834961,-50.3987808227539,-50.4561042785645,-50.5089874267578,-50.4910163879395,-50.4187507629395,-50.3626441955566,-50.3419914245605,-50.2920913696289,-50.2989730834961],[-56.4828109741211,-56.4528312683105,-56.3858413696289,-56.2271499633789,-56.0199203491211,-55.9633293151855,-55.9296379089355,-55.921630859375,-55.9153327941895,-55.9619636535645,-56.0200691223145,-56.0736351013184,-56.1376953125,-56.12939453125,-56.0877914428711,-56.0451164245605,-56.0203628540039,-55.9935035705566,-55.9755859375,-55.9574699401855,-55.9359359741211,-55.8937492370605,-55.7305679321289,-55.658935546875,-55.3853530883789,-55.343994140625,-55.2860336303711,-55.1876945495605,-55.1488304138184,-55.1141128540039,-55.0703125,-55.0058135986328,-54.9786605834961,-54.9684104919434,-54.9265594482422,-54.8760757446289,-54.8516616821289,-54.766845703125,-54.7222137451172,-54.7029304504395,-54.6974105834961,-54.661865234375,-54.6162605285645,-54.5919418334961,-54.5504875183105,-54.5150871276855,-54.43310546875,-54.2930679321289,-54.2279777526855,-54.1673812866211,-54.1300773620605,-54.0897483825684,-53.9464340209961,-53.8766098022461,-53.8295402526855,-53.7942390441895,-53.7677764892578,-53.7501449584961,-53.7347183227539,-53.6836929321289,-53.56396484375,-53.5089874267578,-53.4318351745605,-53.3660163879395,-53.3344230651855,-53.2854995727539,-53.252197265625,-53.2297859191895,-53.1800765991211,-53.082275390625,-53.0097160339355,-52.96484375,-52.9034652709961,-52.8704109191895,-52.7833976745605,-52.7006340026855,-52.6531753540039,-52.5830078125,-52.5594749450684,-52.5546875,-52.4558601379395,-52.4184112548828,-52.3963851928711,-52.3566398620605,-52.3566398620605,-52.327880859375,-52.271240234375,-52.2294425964355,-52.16259765625,-52.1161117553711,-51.99951171875,-51.9906234741211,-51.9443359375,-51.9289054870605,-51.8794898986816,-51.8274879455566,-51.8052749633789,-51.76708984375,-51.6834487915039,-51.6525382995605,-51.5578117370605,-51.5470695495605,-51.4615249633789,-51.3270988464355,-51.2199249267578,-51.0762710571289,-51.0523910522461,-50.994140625,-50.8271980285645,-50.8165054321289,-50.7896957397461,-50.7369651794434,-50.6787605285645,-50.6765594482422,-50.7144050598145,-50.658935546875,-50.6086921691895,-50.5758781433105,-50.534423828125,-50.4588851928711,-50.3042945861816,-50.1875991821289,-50.0546875,-49.9571266174316,-49.881591796875,-49.90625,-49.8988761901855,-49.9379425048828,-50.0472183227539,-50.0709953308105,-50.29443359375,-50.34326171875,-50.4629898071289,-50.58154296875,-50.7550773620605,-50.8163566589355,-50.91015625,-50.9670906066895,-51.1019515991211,-51.2829093933105,-51.299560546875,-51.4041519165039,-51.4962882995605,-51.5550308227539,-51.70263671875,-51.7215309143066,-51.7206077575684,-51.8191375732422,-51.921630859375,-51.9344749450684,-51.9808082580566,-52.0204582214355,-52.229248046875,-52.5534172058105,-52.6641616821289,-52.310302734375,-52.1966819763184,-51.9475593566895,-51.6462860107422,-51.5312004089355,-51.29736328125,-51.2023429870605,-51.0289535522461,-50.9920425415039,-50.8949241638184,-50.84228515625,-50.8381843566895,-50.9178733825684,-50.8971710205078,-50.8445816040039,-50.8255386352539,-50.8186531066895,-50.7861328125,-50.678955078125,-50.67529296875,-50.6900405883789,-50.6387710571289,-50.5855979919434,-50.4032249450684,-50.2604484558105,-50.1727066040039,-50.1166000366211,-49.9992179870605,-49.9029769897461,-49.7195320129395,-49.5853538513184,-49.3136711120605,-49.3986320495605,-49.4601554870605,-49.5069847106934,-49.5533714294434,-49.5993156433105,-49.6365242004395,-49.5758781433105,-49.52392578125,-49.45751953125,-49.4076652526855,-49.2110366821289,-49.15478515625,-48.9913063049316,-48.7100105285645,-48.5999984741211,-48.5295906066895,-48.4629364013672,-48.4458503723145,-48.3498077392578,-48.4514656066895,-48.4680671691895,-48.4777336120605,-48.4085922241211,-48.4498023986816,-48.3064918518066,-48.3175773620605,-48.2664527893066,-48.2017593383789,-48.1284675598145,-48.1150856018066,-48.06884765625,-48.0325660705566,-47.9609375,-47.8833999633789,-47.8076667785645,-47.7737312316895,-47.7314910888672,-47.6871109008789,-47.6510734558105,-47.5573234558105,-47.470703125,-47.4186515808105,-47.4329109191895,-47.4603500366211,-47.4390602111816,-47.3980941772461,-47.2686042785645,-47.2005386352539,-47.1269035339355,-47.0246086120605,-46.9443359375,-46.8936500549316,-46.8112297058105,-46.7699203491211,-46.6444358825684,-46.6172370910645,-46.5163116455078,-46.4217300415039,-46.3208503723145,-46.2191429138184,-46.2149887084961,-46.140380859375,-46.0446281433105,-45.9722633361816,-45.77880859375,-45.644775390625,-45.5569343566895,-45.4585952758789,-45.35302734375,-45.3291511535645,-45.2821273803711,-45.2385711669922,-45.1820793151855,-45.0763664245605,-45.0257835388184,-44.9197731018066,-44.828369140625,-44.7898445129395,-44.7211418151855,-44.7785148620605,-44.720947265625,-44.6512680053711,-44.5916519165039,-44.5467758178711,-44.5377922058105,-44.5800285339355,-44.6172866821289,-44.6586418151855,-44.70751953125,-44.75634765625,-44.7006340026855,-44.6624031066895,-44.5790061950684,-44.5203628540039,-44.4354476928711,-44.3913078308105,-44.3818359375,-44.5201187133789,-44.5206565856934,-44.56201171875,-44.5890159606934,-44.6107902526855,-44.6389656066895,-44.7213859558105,-44.7230491638184,-44.6226577758789,-44.4375495910645,-44.3811531066895,-44.3081550598145,-44.2286148071289,-44.1793937683105,-44.1055679321289,-44.1013679504395,-44.1126480102539,-44.1916007995605,-44.2251930236816,-44.1926765441895,-44.0132331848145,-43.9329109191895,-43.8644523620605,-43.7286109924316,-43.4551239013672,-43.4346160888672,-43.3800773620605,-43.2296867370605,-42.9367179870605,-42.832275390625,-42.6758804321289,-42.5935554504395,-42.2496109008789,-41.9998550415039,-41.8761749267578,-41.7218742370605,-41.64013671875,-41.4799308776855,-41.3182106018066,-41.1945304870605,-40.8755836486816,-40.4745597839355,-40.2353515625,-39.9646987915039,-39.7718276977539,-39.6094245910645,-39.5111808776855,-39.3526840209961,-39.0143547058105,-38.89599609375,-38.6862297058105,-38.4757804870605,-38.3619155883789,-38.2718772888184,-38.048828125,-37.795654296875,-37.6263160705566,-37.3014678955078,-37.1746559143066,-36.9548835754395,-36.8611335754395,-36.7473640441895,-36.5907249450684,-36.38671875,-36.1617660522461,-35.9798812866211,-35.5494117736816,-35.481689453125,-35.392578125,-35.2354507446289,-35.1417465209961,-35.095458984375,-34.9881820678711,-34.9295921325684,-34.8798828125,-34.8759765625,-34.8338851928711,-34.8054695129395,-34.8166007995605,-34.8577651977539,-34.86083984375,-34.8547859191895,-34.8730010986328,-34.8786125183105,-34.8369140625,-34.8346672058105,-34.8905296325684,-34.9666519165039,-35.1577644348145,-35.3408699035645,-35.5970726013184,-35.7639617919922,-35.8301277160645,-35.8908195495605,-35.8477516174316,-35.8854484558105,-36.0549812316895,-36.2235374450684,-36.3983421325684,-36.41162109375,-36.6357421875,-36.768310546875,-36.9377937316895,-37.0933609008789,-37.12548828125,-37.1828117370605,-37.1812019348145,-37.3151359558105,-37.3560066223145,-37.3548812866211,-37.3316421508789,-37.32080078125,-37.32177734375,-37.3592300415039,-37.4384765625,-37.4118194580078,-37.4693336486816,-37.688720703125,-37.9573249816895,-38.0192375183105,-38.23974609375,-38.4017601013184,-38.4473152160645,-38.4989242553711,-38.52490234375,-38.6540031433105,-38.6909675598145,-38.743896484375,-38.7879867553711,-38.8517570495605,-38.7835922241211,-38.7637214660645,-38.8331527709961,-38.8353004455566,-38.9591789245605,-39.0309066772461,-39.0673828125,-39.08935546875,-39.034912109375,-39.0090827941895,-38.9886207580566,-39.001220703125,-39.0411109924316,-39.034912109375,-39.0481452941895,-39.0084953308105,-38.9665031433105,-38.9423332214355,-39.0595703125,-39.0133781433105,-38.9961929321289,-38.9432106018066,-38.88525390625,-38.880615234375,-38.9607925415039,-39.063232421875,-39.1250495910645,-39.1639633178711,-39.202880859375,-39.2152328491211,-39.1706047058105,-39.1540031433105,-39.2783699035645,-39.41259765625,-39.4867668151855,-39.6507797241211,-39.7397956848145,-39.741943359375,-39.6998519897461,-39.7314453125,-39.7833023071289,-39.8447265625,-40.0013694763184,-40.1416969299316,-40.2027320861816,-40.2988777160645,-40.3185539245605,-40.3959465026855,-40.5965805053711,-40.72705078125,-40.7892570495605,-40.8287620544434,-40.9545402526855,-41.0472640991211,-41.0231437683105,-41.0215835571289,-40.9878425598145,-41.0002937316895,-41.1225090026855,-41.5829086303711,-41.7055168151855,-41.9804191589355,-41.99755859375,-41.9861335754395,-41.94091796875,-41.9874992370605,-42.0423812866211,-42.1224594116211,-42.5810546875,-42.8292961120605,-42.9583015441895,-43.0162086486816,-43.0811500549316,-43.1006813049316,-43.0654296875,-43.0862770080566,-43.154296875,-43.22900390625,-43.241943359375,-43.2366218566895,-43.2088394165039,-43.193603515625,-43.2241706848145,-43.3694801330566,-43.5328102111816,-43.7365226745605,-43.8988265991211,-43.9738273620605,-43.8988265991211,-43.7914047241211,-43.6759757995605,-43.7029304504395,-43.8661613464355,-44.0474586486816,-44.1479949951172,-44.367919921875,-44.6372566223145,-44.68115234375,-44.673828125,-44.62109375,-44.5696792602539,-44.6190910339355,-44.6671867370605,-44.95166015625,-45.2154273986816,-45.3253898620605,-45.4232902526855,-45.4333953857422,-45.4643058776855,-45.527099609375,-45.6646461486816,-45.8431625366211,-45.9720687866211,-46.6307601928711,-46.8672866821289,-47.13720703125,-47.5921859741211,-47.8311538696289,-47.8765640258789,-47.914306640625,-47.9891624450684,-47.9593734741211,-47.9083518981934,-47.9293937683105,-48.0243644714355,-48.2027359008789,-48.242431640625,-48.1859359741211,-48.2734870910645,-48.4024925231934,-48.45849609375,-48.4276351928711,-48.4761238098145,-48.5641632080078,-48.6439933776855,-48.7317352294922,-48.6921882629395,-48.5070304870605,-48.4298820495605,-48.4011726379395,-48.545166015625,-48.665771484375,-48.6790046691895,-48.6128425598145,-48.5763206481934,-48.6194343566895,-48.6790046691895,-48.7137680053711,-48.748291015625,-48.70068359375,-48.651611328125,-48.6581535339355,-48.676513671875,-48.677734375,-48.6156730651855,-48.5934066772461,-48.568359375,-48.5541534423828,-48.5955085754395,-48.5719718933105,-48.642578125,-48.6056632995605,-48.6208000183105,-48.6484375,-48.6932144165039,-48.7972640991211,-48.7996597290039,-49.0235824584961,-49.2712860107422,-49.4999046325684,-49.7459945678711,-50.0333480834961,-50.2995109558105,-50.6199684143066,-50.7481460571289,-50.92138671875,-51.1517562866211,-51.4603996276855,-51.7981414794922,-51.9202156066895,-52.0392074584961,-52.0689468383789,-52.0431671142578,-52.0595703125,-52.063232421875,-51.9951171875,-51.8931655883789,-51.8412094116211,-51.8034210205078,-51.6806640625,-51.4461898803711,-51.2721672058105,-51.17431640625,-51.1575202941895,-51.1614265441895,-51.10595703125,-50.9800796508789,-50.9543914794922,-50.96533203125,-50.9408226013184,-50.7701644897461,-50.6893081665039,-50.71630859375,-50.68505859375,-50.6148452758789,-50.5819320678711,-50.5465354919434,-50.5635261535645,-50.6461906433105,-50.931884765625,-51.0249519348145,-51.0403823852539,-51.1792984008789,-51.2335929870605,-51.2498550415039,-51.2980461120605,-51.2950210571289,-51.2817878723145,-51.1572761535645,-51.1875495910645,-51.24658203125,-51.2876968383789,-51.2830581665039,-51.31640625,-51.3590812683105,-51.37646484375,-51.4591293334961,-51.4852561950684,-51.4636726379395,-51.5062980651855,-51.7168960571289,-51.9268074035645,-51.9724617004395,-51.994873046875,-52.0269546508789,-52.1198234558105,-52.1935577392578,-52.1915512084961,-52.1670913696289,-52.1273918151855,-52.190185546875,-52.2746086120605,-52.3416519165039,-52.5084953308105,-52.6522445678711,-52.7628898620605,-52.9208488464355,-53.37060546875,-53.3975601196289,-53.4635772705078,-53.5188484191895,-53.5313491821289,-53.5376472473145,-53.5303726196289,-53.5313491821289,-53.5118637084961,-53.4828605651855,-53.3952140808105,-53.3101081848145,-53.2140617370605,-53.1255874633789,-53.1572761535645,-53.2312507629395,-53.3627433776855,-53.4894027709961,-53.6017074584961,-53.6536102294922,-53.7011222839355,-53.74658203125,-53.76171875,-53.8061027526855,-53.8765144348145,-53.9206047058105,-53.9851570129395,-54.1004409790039,-54.2205581665039,-54.3699226379395,-54.4776840209961,-54.5309104919434,-54.587646484375,-54.89599609375,-55.0360336303711,-55.0911636352539,-55.1735343933105,-55.254638671875,-55.2789535522461,-55.3132820129395,-55.3455085754395,-55.3660621643066,-55.4495582580566,-55.5573272705078,-55.60302734375,-55.6271476745605,-55.6504898071289,-55.6652374267578,-55.7059555053711,-55.75634765625,-55.8077659606934,-55.8736839294434,-55.9519996643066,-56.0046844482422,-56.0155258178711,-56.0184555053711,-55.9989776611328,-56.0448226928711,-56.1058578491211,-56.1761703491211,-56.4072265625,-56.7216796875,-56.8327178955078,-56.937255859375,-57.03271484375,-57.1205062866211,-57.1869125366211,-57.2144508361816,-57.3838424682617,-57.5522956848145,-57.6088905334473,-57.5638656616211,-57.4052276611328,-57.3174781799316,-57.3006820678711,-57.2246551513672,-57.08935546875,-56.9386215209961,-56.7724609375,-56.6715354919434,-56.6358413696289,-56.5707054138184,-56.4759750366211,-56.3932609558105,-56.3223648071289,-56.2255363464355,-56.1028785705566,-56.0342292785645,-56.0196304321289,-55.9849128723145,-55.93017578125,-55.9036598205566,-55.9054183959961,-55.8905258178711,-55.85888671875,-55.8060531616211,-55.7319831848145,-55.687255859375,-55.6719741821289,-55.6915054321289,-55.7459945678711,-55.7254867553711,-55.5823707580566,-55.4766616821289,-55.4098129272461,-55.3464851379395,-55.2437515258789,-55.1015129089355,-55.0638694763184,-55.0689926147461,-55.0399436950684,-54.9559059143066,-54.9102058410645,-54.9027862548828,-54.875732421875,-54.8291015625,-54.777099609375,-54.7197265625,-54.6658706665039,-54.6154289245605,-54.554931640625,-54.4843254089355,-54.4481430053711,-54.3270034790039,-54.2601547241211,-54.2052230834961,-54.1564445495605,-54.1138191223145,-54.0401344299316,-53.9353485107422,-53.9156265258789,-53.8381843566895,-53.7584953308105,-53.71728515625,-53.7271499633789,-53.7533187866211,-53.7445793151855,-53.7181625366211,-53.7109375,-53.6685523986816,-53.6712875366211,-53.7469215393066,-53.8232421875,-53.8642120361328,-53.8911628723145,-53.9547843933105,-54.0123062133789,-54.0850105285645,-54.1192398071289,-54.1545906066895,-54.2061538696289,-54.2500991821289,-54.3318824768066,-54.3833503723145,-54.4439468383789,-54.5015144348145,-54.537841796875,-54.6158676147461,-54.6105461120605,-54.47314453125,-54.4362297058105,-54.4541015625,-54.4129905700684,-54.3129386901855,-54.281005859375,-54.3172836303711,-54.3182601928711,-54.2668952941895,-54.2417984008789,-54.3708038330078,-54.4402351379395,-54.5295906066895,-54.62548828125,-54.6717796325684,-54.7213859558105,-54.8172836303711,-54.9264640808105,-54.982666015625,-55.0818862915039,-55.1943359375,-55.2869148254395,-55.3663101196289,-55.4159164428711,-55.4423828125,-55.4423828125,-55.4588851928711,-55.5184593200684,-55.5382804870605,-55.5416030883789,-55.5349617004395,-55.5184593200684,-55.5283699035645,-55.5548324584961,-55.5481948852539,-55.5614280700684,-55.6011238098145,-55.6209983825684,-55.6209983825684,-55.6507301330566,-55.654052734375,-55.6275863647461,-55.61767578125,-55.6474113464355,-55.7036628723145,-55.7466316223145,-55.7532691955566,-55.799560546875,-55.8491706848145,-55.9053726196289,-55.9914054870605,-56.0674781799316,-56.1898422241211,-56.2460441589355,-56.2757797241211,-56.3518524169922,-56.3948745727539,-56.4477996826172,-56.5238265991211,-56.55029296875,-56.580078125,-56.6330070495605,-56.7024421691895,-56.7751960754395,-56.8446769714355,-56.937255859375,-57.0298843383789,-57.142333984375,-57.2382316589355,-57.3308601379395,-57.3936500549316,-57.4763679504395,-57.5689468383789,-57.6417007446289,-57.7210960388184,-57.7640647888184,-57.8203125,-57.8798370361328,-57.9559097290039,-57.9856910705566,-57.9790496826172,-57.9625015258789,-57.9327621459961,-57.9493179321289,-57.9426727294922,-57.929443359375,-57.9162139892578,-57.9261703491211,-57.929443359375,-57.9360809326172,-57.9459915161133,-57.9062957763672,-57.8732452392578,-57.89306640625,-57.886474609375,-57.8600082397461,-57.8269577026367,-57.8302268981934,-57.8600082397461,-57.8922386169434,-57.9004898071289,-57.8848152160645,-57.9019050598145,-57.9084968566895,-57.8914070129395,-57.9151344299316,-57.9625015258789,-57.9790496826172,-57.9956092834473,-58.0088386535645,-58.0022468566895,-58.0253944396973,-58.0584449768066,-58.0915031433105,-58.1246109008789,-58.1377906799316,-58.1597633361816,-58.09375,-58.0676231384277,-58.0211448669434,-57.9601593017578,-57.8875961303711,-57.8607444763184,-58.0299301147461,-58.1314926147461,-58.0720252990723,-57.9716796875,-57.8745079040527,-57.8003921508789,-57.7814483642578,-57.7168006896973,-57.7286148071289,-57.7308616638184,-57.7831077575684,-57.7250022888184,-57.63916015625,-57.5740203857422,-57.5531272888184,-57.5061492919922,-57.4956512451172,-57.5520477294922,-57.5864715576172,-57.6616744995117,-57.7801780700684,-57.7888641357422,-57.8324699401855,-57.9050331115723,-57.9909133911133,-58.20556640625,-58.3477516174316,-58.3959922790527,-58.4173851013184,-58.4598159790039,-58.4781265258789,-58.4706077575684,-58.3503875732422,-58.3507843017578,-58.3405723571777,-58.3456077575684,-58.3753929138184,-58.4236831665039,-58.49658203125,-58.5379371643066,-58.9572792053223,-59.4342765808105,-59.8311538696289,-60.1755867004395,-60.1872100830078,-60.2066421508789,-60.2204132080078,-60.2423362731934,-60.3804702758789,-60.5304679870605,-60.5832023620605,-60.4020004272461,-60.2733383178711,-60.2988777160645,-60.3380355834961,-60.3727035522461,-60.396240234375,-60.4601593017578,-60.4746589660645,-60.4629859924316,-60.4280776977539,-60.4049758911133,-60.4223594665527,-60.4601593017578,-60.506591796875,-60.5953140258789,-60.7223625183105,-60.9145011901855,-61.0769996643066,-61.1291465759277,-61.4160652160645,-61.5115699768066,-61.57568359375,-61.7899398803711,-61.8741226196289,-61.9447288513184,-62.0947723388672,-62.1180152893066,-62.1760711669922,-62.263916015625,-62.3528327941895,-62.5255393981934,-62.6870613098145,-62.7654762268066,-62.8351593017578,-62.9579124450684,-63.0151901245117,-63.0413551330566,-63.0674819946289,-63.1167945861816,-63.1806640625,-63.2497596740723,-63.3466796875,-63.4652366638184,-63.5418930053711,-63.5856475830078,-63.6885757446289,-63.7880859375,-63.9385757446289,-64.0616226196289,-64.2550354003906,-64.4205093383789,-64.4807662963867,-64.513427734375,-64.6116714477539,-64.6900405883789,-64.783447265625,-64.8298797607422,-64.9143600463867,-64.9925308227539,-65.001220703125,-65.0302734375,-65.037109375,-65.0902786254883,-65.1151351928711,-65.1426773071289,-65.1633758544922,-65.1857376098633,-65.1897430419922,-65.1753921508789,-65.2062072753906,-65.2822799682617,-65.322021484375,-65.3254852294922,-65.3423843383789,-65.3728561401367,-65.389892578125,-65.3936004638672,-65.37158203125,-65.3237762451172,-65.3340301513672,-65.4022903442383,-65.4399871826172,-65.4471206665039,-65.4369125366211,-65.3954620361328,-65.3130798339844,-65.298583984375,-65.3245544433594,-65.328125,-65.309326171875,-65.337890625,-65.3961410522461,-65.436767578125,-65.4919891357422,-65.5586929321289,-65.6371078491211,-65.706787109375,-65.9247055053711,-66.2635726928711,-66.3992233276367,-66.4789047241211,-66.5753402709961,-66.72998046875,-67.1115264892578,-67.1904754638672,-67.2804718017578,-67.3327178955078,-67.4169464111328,-67.5824203491211,-67.6666488647461,-67.7217788696289,-67.7856903076172,-67.8350067138672,-67.99169921875,-68.0716781616211,-68.1586456298828,-68.2666015625,-68.3111343383789,-68.3980026245117,-68.4983444213867,-68.6226577758789,-68.6783752441406,-68.7274856567383,-68.7699203491211,-68.7840805053711,-68.8483428955078,-69.0016555786133,-69.228515625,-69.4625473022461,-69.57861328125,-69.6740188598633,-69.8397979736328,-69.9603500366211,-70.0663070678711,-70.2200698852539,-70.2903747558594,-70.3419952392578,-70.3922882080078,-70.4508819580078,-70.5332565307617,-70.5965347290039,-70.642333984375,-70.6415557861328,-70.6403274536133,-70.6393508911133,-70.6385269165039,-70.6375961303711,-70.6369171142578,-70.5937957763672,-70.5672302246094,-70.5991744995117,-70.5922393798828,-70.5701599121094,-70.5411148071289,-70.60791015625,-70.6369171142578,-70.6724624633789,-70.7584991455078,-70.8162612915039,-70.884521484375,-70.9707565307617,-71.041748046875,-71.1152801513672,-71.2379379272461,-71.3394088745117,-71.6080093383789,-71.887451171875,-72.1429748535156,-72.1815872192383,-72.1791000366211,-72.1728515625,-72.2599563598633,-72.2658157348633,-72.2890167236328,-72.3180618286133,-72.3790512084961,-72.4647521972656,-72.60546875,-72.8142547607422,-73.0137710571289,-73.2094268798828,-73.08984375,-72.9703598022461,-72.9740219116211,-73.0705108642578,-73.12255859375,-73.203125,-73.3024444580078,-73.3567352294922,-73.3517074584961,-73.3603973388672,-73.3981399536133,-73.4358901977539,-73.4881362915039,-73.5491256713867,-73.5491256713867,-73.5723648071289,-73.610107421875,-73.610107421875,-73.6449203491211,-73.6826705932617,-73.7204132080078,-73.7755813598633,-73.772705078125,-73.7320251464844,-73.714599609375,-73.7204132080078,-73.7668991088867,-73.8220748901367,-73.8946304321289,-73.9468688964844,-73.9817428588867,-74.0020523071289,-73.9817428588867,-73.95849609375,-73.9526824951172,-73.9643096923828,-73.9643096923828,-73.929443359375,-73.8917465209961,-73.85400390625,-73.8046417236328,-73.7494659423828,-73.7204132080078,-73.7233428955078,-73.7582015991211,-73.7930145263672,-73.8046417236328,-73.7762680053711,-73.7581100463867,-73.6945343017578,-73.4999084472656,-73.3254852294922,-73.2403335571289,-73.1774444580078,-73.1373519897461,-73.1263198852539,-73.1353530883789,-73.167724609375,-73.2064971923828,-73.2355499267578,-73.2093734741211,-73.1628875732422,-73.0680694580078,-72.9798812866211,-72.97021484375,-72.9589309692383,-72.9182586669922,-72.8958053588867,-72.907470703125,-72.8870620727539,-72.8319320678711,-72.69873046875,-72.6083526611328,-72.468994140625,-72.3528289794922,-72.2567901611328,-72.08251953125,-71.982421875,-71.9431610107422,-71.8447265625,-71.6683654785156,-71.5213317871094,-71.4382781982422,-71.3167953491211,-71.2350082397461,-71.1442413330078,-70.9736862182617,-70.9156265258789,-70.8660125732422,-70.7995147705078,-70.7215805053711,-70.6345748901367,-70.5306625366211,-70.4046401977539,-70.3436508178711,-70.31689453125,-70.2391586303711,-70.1839828491211,-70.1288070678711,-70.0533218383789,-70.0039520263672,-69.9720230102539,-69.9659194946289,-69.9481964111328,-69.9110336303711,-69.8497543334961,-69.7941360473633,-69.7326202392578,-69.6690444946289,-69.6046905517578,-69.5518569946289,-69.5064468383789,-69.4786071777344,-69.4349136352539,-69.4178695678711,-69.4002380371094,-69.4114303588867,-69.4491271972656,-69.44873046875,-69.4443359375,-69.44873046875,-69.4884262084961,-69.519287109375,-69.5435485839844,-69.5545883178711,-69.5744171142578,-69.583251953125,-69.6119155883789,-69.6207046508789,-69.6008758544922,-69.592041015625,-69.6008758544922,-69.6119155883789,-69.6339797973633,-69.66748046875,-69.7474594116211,-69.8279266357422,-69.9227600097656,-70.0440444946289,-70.0705108642578,-70.0709457397461,-70.0657196044922,-70.0579147338867,-70.0539016723633,-69.9854507446289,-69.9251022338867,-69.862060546875,-69.80712890625,-69.7567367553711,-69.7188949584961,-69.673828125,-69.6387252807617,-69.6036148071289,-69.5648422241211,-69.5270538330078,-69.4721221923828,-69.4207992553711,-69.3919906616211,-69.358642578125,-69.3271484375,-69.3055191040039,-69.2830047607422,-69.2541961669922,-69.2127914428711,-69.174072265625,-69.1560516357422,-69.1533203125,-69.1632308959961,-69.1767578125,-69.1659622192383,-69.1650390625,-69.1632308959961,-69.19384765625,-69.2244644165039,-69.2586898803711,-69.3118133544922,-69.3613739013672,-69.4027786254883,-69.4415054321289,-69.4703140258789,-69.5171432495117,-69.5675811767578,-69.6208953857422,-69.7169876098633,-69.7513198852539,-69.7981491088867,-69.8521499633789,-69.8507843017578,-69.8494644165039,-69.8485794067383,-69.7999496459961,-69.7396011352539,-69.6500473022461,-69.5812454223633,-69.5429153442383,-69.4701690673828,-69.3943328857422,-69.3197250366211,-69.124267578125,-68.9131851196289,-68.678466796875,-68.4434509277344,-68.2395553588867,-68.1765670776367,-68.2132873535156,-68.2559585571289,-68.2394485473633,-68.2183609008789,-68.1937942504883,-68.1302795410156,-68.0770492553711,-68.0328674316406,-67.98974609375,-67.9362335205078,-67.8755340576172,-67.8150939941406,-67.7118682861328,-67.6092300415039,-67.5560531616211,-67.4996566772461,-67.457763671875,-67.4004364013672,-67.3519592285156,-67.3206024169922,-67.205810546875,-67.1192398071289,-67.0901336669922,-67.0882797241211,-67.0936508178711,-67.082275390625,-67.0652313232422,-66.8760299682617,-66.6190414428711,-66.4292449951172,-66.3471221923828,-66.3016586303711,-66.1912155151367,-66.06005859375,-65.996337890625,-65.9258804321289,-65.8113250732422,-65.7181167602539,-65.6814498901367,-65.6446762084961,-65.5660171508789,-65.5229949951172,-65.5626907348633,-65.5560531616211,-65.473388671875,-65.4072265625,-65.36083984375,-65.2639694213867,-65.1696319580078,-65.103759765625,-65.0265655517578,-64.9101104736328,-64.8179702758789,-64.7315368652344,-64.6674346923828,-64.5843734741211,-64.5262756347656,-64.4860382080078,-64.4051284790039,-64.30419921875,-64.2050247192383,-64.1148452758789,-64.0670394897461,-64.0354461669922,-64.0084991455078,-63.9757804870605,-63.9371604919434,-63.844482421875,-63.68212890625,-63.5702629089355,-63.4639167785645,-63.4325218200684,-63.3939437866211,-63.3748512268066,-63.3892593383789,-63.5846214294434,-63.7125511169434,-63.9241714477539,-64.02490234375,-64.0465850830078,-64.048828125,-64.0287170410156,-64.009033203125,-64.0377883911133,-64.1435546875,-64.2188491821289,-64.228759765625,-64.22705078125,-64.2210922241211,-64.2752914428711,-64.5679244995117,-64.6689987182617,-64.7025833129883,-64.81787109375,-64.7886734008789,-64.7222671508789,-64.66552734375,-64.6136703491211,-64.5763626098633,-64.5255355834961,-64.2556686401367,-64.1924819946289,-64.154296875,-64.1217269897461,-64.0733871459961,-64.021484375,-63.9146499633789,-63.7469749450684,-63.6529312133789,-63.5966300964355,-63.5268058776855,-63.3797836303711,-63.3386726379395,-63.2947273254395,-63.1362342834473,-63.0453109741211,-62.9686546325684,-62.8569793701172,-62.7646026611328,-62.7399406433105,-62.7121124267578,-62.6653327941895,-62.6097640991211,-62.5439453125,-62.4725570678711,-62.4106407165527,-62.1531219482422,-62.0815887451172,-61.8208465576172,-61.5542449951172,-61.4793930053711,-61.3675270080566,-61.2800827026367,-61.2094230651855,-61.1024436950684,-61.0362777709961,-61.0028305053711,-60.9664039611816,-60.90625,-60.8334007263184,-60.7417449951172,-60.6791496276855,-60.6275863647461,-60.6038589477539,-60.6044921875,-60.635009765625,-60.6719779968262,-60.7119598388672,-60.7421340942383,-60.6513710021973,-60.576416015625,-60.4595222473145,-60.4087867736816,-60.335205078125,-60.2416534423828,-60.1817398071289,-60.1420440673828,-60.10595703125,-60.0780792236328,-59.9906730651855,-59.9993667602539,-60.0154800415039,-60.0267562866211,-60.0317878723145,-60.0689468383789,-60.1245574951172,-60.1409149169922,-60.1486320495605,-60.1111373901367,-60.0450210571289,-59.9623527526855,-59.9061012268066,-59.8333511352539,-59.7458038330078,-59.7032699584961,-59.69970703125,-59.7274894714355,-59.7385749816895,-59.7168960571289,-59.6912078857422,-59.6202163696289,-59.58642578125,-59.5577621459961,-59.5511207580566,-59.5753898620605,-59.6044464111328,-59.6702117919922,-59.6790046691895,-59.7316436767578,-59.8543968200684,-59.8330574035645,-59.8288116455078,-59.8311538696289,-59.8730506896973,-59.9456520080566,-59.9723129272461,-59.9958953857422,-59.9943313598633,-59.9607887268066,-59.8896446228027,-59.84912109375,-59.7552223205566,-59.7435073852539,-59.7517585754395,-59.7561988830566,-59.74072265625,-59.6985359191895,-59.6685066223145,-59.6637725830078,-59.6665992736816,-59.5966300964355,-59.5356941223145,-59.4794464111328,-59.3776817321777,-59.3372535705566,-59.3169898986816,-59.231201171875,-59.1003913879395,-58.968505859375,-58.9166030883789,-58.8625030517578,-58.8217811584473,-58.7872047424316,-58.7303237915039,-58.6846160888672,-58.6050796508789,-58.5118675231934,-58.4957008361816,-58.4868621826172,-58.5060539245605,-58.4729499816895,-58.3958015441895,-58.38037109375,-58.3626976013184,-58.3406753540039,-58.314208984375,-58.2811508178711,-58.2304229736328,-58.173095703125,-58.1422386169434,-58.0913047790527,-58.03466796875,-58.0117683410645,-57.9951171875,-57.9828147888184,-57.9463424682617,-57.8734359741211,-57.795654296875,-57.6917457580566,-57.5944366455078,-57.5457496643066,-57.5004386901855,-57.4126968383789,-57.3667984008789,-57.3174781799316,-57.2755851745605,-57.1896018981934,-57.118896484375,-57.0926780700684,-57.03759765625,-57.0100593566895,-56.9695320129395,-56.8367195129395,-56.7662582397461,-56.6898422241211,-56.616455078125,-56.5635719299316,-56.5254859924316,-56.4828109741211],[-59.4933090209961,-59.5218734741211,-59.611328125,-59.6427764892578,-59.6466827392578,-59.5916023254395,-59.4878883361816,-59.4276390075684,-59.4933090209961],[115.026748657227,115.140037536621,115.168449401855,115.227928161621,115.266693115234,115.279289245605,115.326759338379,115.319236755371,115.290626525879,115.246681213379,115.170608520508,115.107032775879,115.051559448242,115.026748657227,115.028816223145,115.026748657227],[115.026748657227,114.944717407227,114.864547729492,114.784187316895,114.746688842773,114.759963989258,114.779296875,114.790138244629,114.818267822266,114.840232849121,114.8310546875,114.783500671387,114.810455322266,114.776168823242,114.724990844727,114.654098510742,114.608306884766,114.57177734375,114.51220703125,114.447074890137,114.416603088379,114.322944641113,114.289649963379,114.28759765625,114.261032104492,114.22412109375,114.168846130371,114.095123291016,114.063865661621,114.177932739258,114.299407958984,114.424415588379,114.544723510742,114.645896911621,114.740829467773,114.840629577637,114.995414733887,115.047653198242,115.047065734863,115.026748657227],[91.6319351196289,91.6258850097656,91.59765625,91.5792999267578,91.5947265625,91.6581039428711,91.7430648803711,91.8512725830078,91.9509811401367,91.9908218383789,92.044921875,92.0834045410156,92.0311508178711,92.0025405883789,91.9922866821289,91.9986343383789,92.0308532714844,92.0681610107422,92.0734329223633,92.0497055053711,91.9983367919922,91.9437484741211,91.8986282348633,91.8420944213867,91.7537078857422,91.6715850830078,91.517578125,91.4558563232422,91.4267578125,91.2865219116211,91.1338882446289,90.8557662963867,90.7396469116211,90.6203155517578,90.5598678588867,90.4476547241211,90.3458938598633,90.2423858642578,90.2060546875,90.1229553222656,89.9431610107422,89.7638702392578,89.7109375,89.6099548339844,89.6061553955078,89.6091766357422,89.5861358642578,89.5451202392578,89.474609375,89.3841781616211,89.3321228027344,89.1482467651367,89.0409240722656,88.9191360473633,88.8576202392578,88.8351516723633,88.8135681152344,88.765625,88.73876953125,88.7603530883789,88.8816375732422,88.8914108276367,88.9475555419922,89.0254898071289,89.1023483276367,89.1604461669922,89.2726593017578,89.3958969116211,89.4806594848633,89.5369186401367,89.6527328491211,89.7498092651367,89.81689453125,89.8978500366211,89.9810562133789,90.1044921875,90.2208023071289,90.3482360839844,90.3629837036133,90.3521499633789,90.3337860107422,90.3331069946289,90.3527374267578,90.4773406982422,90.6300811767578,90.7157211303711,90.9066390991211,90.9625015258789,91.0208053588867,91.0777282714844,91.14990234375,91.2258758544922,91.2730484008789,91.3068389892578,91.3675765991211,91.4933624267578,91.6055679321289,91.6418914794922,91.62939453125,91.6319351196289],[25.2587890625,25.2390613555908,25.2240238189697,25.2422847747803,25.2824211120605,25.3402347564697,25.3843746185303,25.4367198944092,25.4892559051514,25.5583019256592,25.76123046875,25.78369140625,25.8119144439697,25.9393558502197,25.9591808319092,25.95068359375,26.0819339752197,26.1680660247803,26.2410144805908,26.474609375,26.67822265625,26.9167003631592,27.091796875,27.17822265625,27.2214851379395,27.2567386627197,27.2746105194092,27.28076171875,27.4689464569092,27.6246089935303,27.6792964935303,27.6996097564697,27.69482421875,27.6969738006592,27.7042980194092,27.6880855560303,27.6769523620605,27.66943359375,27.6934585571289,27.8441410064697,27.9074230194092,27.974609375,28.0140609741211,28.0456047058105,28.181640625,28.5320301055908,28.7477531433105,28.9193344116211,28.9907207489014,29.0255851745605,29.0373058319092,29.0158214569092,29.0233383178711,29.0423812866211,29.0714836120605,29.1068382263184,29.2372093200684,29.3152332305908,29.3648452758789,29.1298828125,29.0134754180908,28.9458026885986,28.8398456573486,28.6955070495605,28.5428695678711,28.3817386627197,28.2101573944092,28.0279312133789,27.93505859375,27.9313488006592,27.8905277252197,27.8125972747803,27.7685546875,27.75830078125,27.716796875,27.6438484191895,27.5926742553711,27.5631847381592,27.4987316131592,27.3992176055908,27.3133792877197,27.2412109375,27.185546875,27.1463871002197,27.0855464935303,26.9870109558105,26.9706058502197,26.8350582122803,26.7611331939697,26.6177730560303,26.5015621185303,26.4517593383789,26.3971691131592,26.130859375,26.0318374633789,25.912109375,25.8818359375,25.8524417877197,25.7699203491211,25.70263671875,25.6591796875,25.5837879180908,25.5181636810303,25.4436531066895,25.34619140625,25.21337890625,25.0924816131592,24.9989261627197,24.8692378997803,24.7481460571289,24.5558605194092,24.4001941680908,24.33056640625,24.1929683685303,24.1044921875,23.9695320129395,23.8937511444092,23.8234367370605,23.6707038879395,23.521484375,23.3892574310303,23.2660160064697,23.1487312316895,23.0575199127197,23.0220718383789,22.9512691497803,22.8788089752197,22.8189449310303,22.7960948944092,22.72900390625,22.6402339935303,22.5976543426514,22.5486335754395,22.4708995819092,22.2175769805908,22.0909175872803,22.0109367370605,21.91455078125,21.8332042694092,21.7882804870605,21.7380867004395,21.6947269439697,21.6462879180908,21.5013675689697,21.4549808502197,21.0709972381592,20.9539070129395,20.8708972930908,20.7398433685303,20.68505859375,20.6414051055908,20.6199226379395,20.6267585754395,20.6978511810303,20.7570323944092,20.8150386810303,20.8226566314697,20.81103515625,20.7994155883789,20.7931632995605,20.7107429504395,20.6092777252197,20.47314453125,20.4306659698486,20.34521484375,20.0286121368408,19.98046875,19.98046875,19.9801750183105,19.9798831939697,19.9795913696289,19.9792976379395,19.9789066314697,19.978515625,19.9782238006592,19.9779300689697,19.9776382446289,19.9773445129395,20.2053718566895,20.4875011444092,20.82275390625,20.9709968566895,20.9794921875,20.9792957305908,20.9787101745605,20.9781246185303,20.9774417877197,20.9768543243408,20.9761714935303,20.9755859375,20.9750003814697,20.9743156433105,20.97412109375,21.2325191497803,21.5296859741211,22.0114250183105,22.4600582122803,22.7527351379395,23.0999031066895,23.2193355560303,23.2515621185303,23.2986316680908,23.4597644805908,23.5601577758789,23.58056640625,23.5997066497803,23.6471691131592,23.7004890441895,23.8642578125,23.8983402252197,24.0026378631592,24.1292972564697,24.2439460754395,24.3589859008789,24.4122085571289,24.4749011993408,24.5305671691895,24.7921886444092,24.9090824127197,25.2160148620605,25.2587890625],[22.8600597381592,22.9307613372803,22.9643535614014,23.255859375,23.3123054504395,23.4566402435303,23.5450191497803,23.6462898254395,23.65625,23.6427726745605,23.6226558685303,23.5960941314697,23.46826171875,23.4627933502197,23.4890632629395,23.5280284881592,23.5518550872803,23.5373039245605,23.5832042694092,23.6792984008789,23.9219722747803,24.0481452941895,24.1473636627197,24.19482421875,24.2208995819092,24.1799793243408,24.2083988189697,24.2914047241211,24.37548828125,24.4560546875,24.7367191314697,24.8533191680908,25.0072269439697,25.2003917694092,25.2473640441895,25.2386722564697,25.1813488006592,25.1901378631592,25.2789058685303,25.3806629180908,25.5666027069092,25.8889636993408,26.0365238189697,26.0869140625,26.1693363189697,26.2845706939697,26.36181640625,26.30859375,26.3246097564697,26.3533210754395,26.4205074310303,26.4474601745605,26.5142574310303,26.5936527252197,26.7263660430908,26.7964859008789,26.9422855377197,27.0833988189697,27.1439456939697,27.1812496185303,27.21337890625,27.2291011810303,27.2325191497803,27.2567386627197,27.3324203491211,27.4033203125,27.1149406433105,27.0718746185303,27.0206050872803,26.8701171875,26.8220710754395,26.767578125,26.6326160430908,26.1735363006592,25.8199214935303,25.7138671875,25.5250988006592,25.4001941680908,25.2831058502197,25.2493171691895,25.0652351379395,24.9784183502197,24.7655258178711,24.4371089935303,24.31982421875,24.2277336120605,23.99169921875,23.8484363555908,23.6818351745605,23.5236320495605,23.4171867370605,23.3128910064697,23.2188472747803,23.1159172058105,22.9928722381592,22.8645496368408,22.7557621002197,22.7117195129395,22.6171875,22.5056648254395,22.4618167877197,22.44970703125,22.4221687316895,21.908203125,21.68701171875,21.53759765625,21.3501949310303,21.2683582305908,21.2297859191895,21.1255855560303,20.9557609558105,20.79296875,20.6474609375,20.55810546875,20.4865245819092,20.3935546875,20.2263679504395,20.0023441314697,19.8624992370605,19.8065433502197,19.68603515625,19.5009765625,19.3234386444092,19.0685539245605,18.8317375183105,18.6999015808105,18.5941410064697,18.5674800872803,18.6199226379395,18.6336917877197,18.5966796875,18.6103515625,18.5538101196289,18.4998054504395,18.4744129180908,18.3181648254395,18.2371082305908,18.1939449310303,18.1609363555908,18.111328125,18.072265625,18.0107421875,17.9479484558105,17.9071292877197,17.88037109375,17.806640625,17.5376949310303,17.4916019439697,17.43798828125,17.2984371185303,17.2247066497803,17.0025386810303,16.7643566131592,16.67333984375,16.6107425689697,16.5704116821289,16.5430660247803,16.4962902069092,16.4767570495605,16.4800777435303,16.4662113189697,16.4595699310303,16.4685535430908,16.4012699127197,16.3196296691895,16.2517566680908,16.1833992004395,16.1361331939697,16.1067390441895,16.0955085754395,16.1018543243408,16.08349609375,16.0821304321289,16.0592765808105,16.0824222564697,16.0634765625,16.0082035064697,15.9580078125,15.9287118911743,15.9048824310303,15.8493175506592,15.7749996185303,15.6765632629395,15.5808601379395,15.4583978652954,15.3601560592651,15.2398443222046,15.1287117004395,15.0621089935303,15.0348634719849,15.0673837661743,15.1154308319092,15.1358394622803,15.1369152069092,15.0875005722046,15.0635738372803,15.0227537155151,14.8935556411743,14.7704105377197,14.7312498092651,14.708984375,14.6617183685303,14.640625,14.6017580032349,14.5735349655151,14.56298828125,14.5680656433105,14.5843744277954,14.5835943222046,14.6168947219849,14.6168947219849,14.5988273620605,14.5772457122803,14.54248046875,14.5031242370605,14.4638662338257,14.4311513900757,14.4407224655151,14.4749994277954,14.5121097564697,14.5593748092651,14.6995115280151,14.7392578125,14.7640619277954,14.7803707122803,14.8619146347046,14.9827146530151,15.0345697402954,15.0863285064697,15.1571292877197,15.1858396530151,15.2067384719849,15.2458992004395,15.3791017532349,15.4800777435303,15.5892581939697,15.7012691497803,15.8450193405151,15.9576168060303,16.0306644439697,16.1911144256592,16.37890625,16.4043941497803,16.4593753814697,16.5232410430908,16.5453128814697,16.5501956939697,16.5889644622803,16.6683597564697,16.7847652435303,16.8181648254395,16.8903312683105,17.0719718933105,17.1179695129395,17.2469730377197,17.4021492004395,17.4364261627197,17.49267578125,17.6494140625,17.7608394622803,17.9401378631592,18.2388668060303,18.455078125,18.5641613006592,18.5916004180908,18.6335945129395,18.6662101745605,18.7474613189697,18.9064445495605,19.0108394622803,19.0398445129395,19.0423831939697,19.0638675689697,19.1086921691895,19.0641593933105,18.8860340118408,18.8885746002197,18.8783206939697,18.8882827758789,18.9562511444092,19.0478515625,19.1455078125,19.4002933502197,19.6174812316895,19.6683597564697,19.8376960754395,19.9535160064697,20.0726547241211,20.3420886993408,20.56689453125,20.6314449310303,20.65966796875,20.6681652069092,20.7732410430908,20.8910160064697,20.9841804504395,21.0094718933105,21.2638664245605,21.3524417877197,21.39599609375,21.4968738555908,21.5280265808105,21.5757808685303,21.6327152252197,21.6827144622803,21.7257823944092,21.7261714935303,21.70654296875,21.70654296875,21.7306632995605,21.771484375,21.96484375,22.0137691497803,22.0431632995605,22.09716796875,22.15625,22.1936531066895,22.2359371185303,22.3698234558105,22.4938488006592,22.6240234375,22.7301769256592,22.8173828125,22.8600597381592],[-59.78759765625,-59.9222679138184,-60.0377464294434,-60.1142578125,-60.1174812316895,-59.93603515625,-59.8663558959961,-59.7271499633789,-59.78759765625],[-66.2737731933594,-66.3241195678711,-66.3119125366211,-66.25048828125,-66.2103500366211,-66.2737731933594],[-66.7624969482422,-66.8970718383789,-66.8447265625,-66.8021469116211,-66.7454071044922,-66.7533721923828,-66.7624969482422],[-60.9615707397461,-61.0028800964355,-61.0125007629395,-61.076171875,-61.0817413330078,-61.0259780883789,-60.9124526977539,-60.9530258178711,-60.9615707397461],[-73.6953125,-73.81591796875,-73.8577194213867,-73.7246551513672,-73.5723648071289,-73.6953125],[-73.5665054321289,-73.6435546875,-73.7753372192383,-73.9202117919922,-73.9605484008789,-73.8529281616211,-73.6874465942383,-73.5224609375,-73.47607421875,-73.5388641357422,-73.5516586303711,-73.5665054321289],[-71.0257339477539,-71.1166534423828,-71.094970703125,-70.9708480834961,-70.879638671875,-70.8257827758789,-70.9134750366211,-71.0257339477539],[-61.1051750183105,-61.0713386535645,-60.9365692138672,-60.8652381896973,-60.8684043884277,-60.9842796325684,-61.0375480651855,-60.9706039428711,-60.9715347290039,-61.0519523620605,-61.0920867919922,-61.0590362548828,-60.9303703308105,-60.8775863647461,-60.8060989379883,-60.7378883361816,-60.6990699768066,-60.4723625183105,-60.4605941772461,-60.7048835754395,-60.7332992553711,-60.5731925964355,-60.5857467651367,-60.5049285888672,-60.4308586120605,-60.3765144348145,-60.2979469299316,-60.2438468933105,-60.2264633178711,-60.0924835205078,-59.96142578125,-59.8650360107422,-59.8499984741211,-59.8487815856934,-59.8809089660645,-59.9340362548828,-59.8280258178711,-59.8421897888184,-60.0158195495605,-60.1144561767578,-60.205078125,-60.3860855102539,-60.6729469299316,-60.7637176513672,-60.87158203125,-60.9786109924316,-61.0836944580078,-61.1864242553711,-61.236328125,-61.2836875915527,-61.3234405517578,-61.4083518981934,-61.4498062133789,-61.4953155517578,-61.4806175231934,-61.4086380004883,-61.3021965026855,-61.2405281066895,-60.9825172424316,-60.9319801330566,-60.8701667785645,-60.7596702575684,-60.6166534423828,-60.571044921875,-60.4890632629395,-60.4081993103027,-60.4313468933105,-60.4254417419434,-60.3317375183105,-60.3329124450684,-60.3840827941895,-60.4824256896973,-60.5077171325684,-60.4945297241211,-60.534423828125,-60.5768585205078,-60.7448272705078,-60.83056640625,-60.9122085571289,-61.1051750183105],[-63.8112831115723,-63.7842254638672,-63.7370109558105,-63.6814460754395,-63.5343704223633,-63.4564933776855,-63.4131355285645,-63.36865234375,-63.2860794067383,-63.12939453125,-62.9640159606934,-62.7120094299316,-62.6819305419922,-62.4230918884277,-62.16357421875,-62.0742683410645,-62.0408706665039,-62.0237312316895,-62.1717758178711,-62.3199691772461,-62.5260734558105,-62.5519981384277,-62.5392074584961,-62.5432624816895,-62.5025901794434,-62.4780769348145,-62.5313453674316,-62.7431640625,-62.8048820495605,-62.8783645629883,-62.9035148620605,-62.99462890625,-63.0220718383789,-62.8945350646973,-62.95263671875,-63.0150375366211,-63.0563468933105,-63.0529327392578,-62.9951133728027,-62.9784698486328,-63.0568885803223,-63.1169929504395,-63.1947288513184,-63.2708015441895,-63.1527824401855,-63.2134742736816,-63.2766151428223,-63.5688972473145,-63.6410179138184,-63.7317848205566,-63.800537109375,-63.7632293701172,-63.7505340576172,-63.7586402893066,-63.8605461120605,-64.0197219848633,-64.11083984375,-64.1065444946289,-64.1360397338867,-64.2356414794922,-64.3880386352539,-64.4031219482422,-64.3545913696289,-64.2799835205078,-64.2232437133789,-64.1569366455078,-63.9935531616211,-63.9972686767578,-63.9814949035645,-64.087890625,-63.9030227661133,-63.8792953491211,-63.8637237548828,-63.8756370544434,-63.9055671691895,-63.8335952758789,-63.8112831115723],[-61.9141159057617,-61.8787117004395,-61.8154792785645,-61.7725601196289,-61.8337440490723,-61.9508323669434,-62.00830078125,-61.9247093200684,-61.8272972106934,-61.6278343200684,-61.5480499267578,-61.4740715026855,-61.3955039978027,-61.4755325317383,-61.5822296142578,-61.68408203125,-61.7508811950684,-61.8312492370605,-61.8866233825684,-61.9141159057617],[-54.2271499633789,-54.2760734558105,-54.3259773254395,-54.3201141357422,-54.2586898803711,-54.2273902893066,-54.2262687683105,-54.2149391174316,-54.1683578491211,-54.128173828125,-54.1475563049316,-54.2271499633789],[-64.5085983276367,-64.5338897705078,-64.6212844848633,-64.6646423339844,-64.6845703125,-64.6604995727539,-64.6632766723633,-64.5911102294922,-64.5648422241211,-64.5085983276367],[-64.47607421875,-64.59130859375,-64.5407257080078,-64.5195770263672,-64.5001983642578,-64.4812469482422,-64.47607421875],[-123.435394287109,-123.477249145508,-123.499610900879,-123.517524719238,-123.582328796387,-123.554695129395,-123.467864990234,-123.487548828125,-123.422744750977,-123.406791687012,-123.435394287109],[-123.372367858887,-123.384811401367,-123.541015625,-123.645599365234,-123.689262390137,-123.482322692871,-123.3779296875,-123.372367858887],[-74.7088928222656,-74.5663070678711,-74.26904296875,-74.0498046875,-73.7646484375,-73.55810546875,-73.518798828125,-73.4841842651367,-73.4652862548828,-73.3688507080078,-73.2530288696289,-73.1595687866211,-72.9899444580078,-72.7334518432617,-72.4961929321289,-72.3661651611328,-72.2401351928711,-72.1872024536133,-72.1092758178711,-71.9009323120117,-71.6712875366211,-71.439208984375,-71.2611770629883,-71.1520004272461,-70.9932632446289,-70.5194854736328,-70.3880844116211,-70.2177734375,-70.069580078125,-70.0171356201172,-69.80224609375,-69.5810546875,-69.4710388183594,-69.3063430786133,-68.987060546875,-68.815673828125,-68.7460479736328,-68.552001953125,-68.4314956665039,-68.2381896972656,-67.8890151977539,-67.5608901977539,-67.1174850463867,-66.5980911254883,-66.1781768798828,-65.8828125,-65.5233917236328,-65.3961410522461,-64.8363265991211,-64.5677261352539,-64.2618179321289,-64.2162094116211,-64.2087936401367,-64.3707504272461,-64.5137176513672,-64.41455078125,-64.24609375,-64.2537612915039,-64.3488311767578,-64.6331481933594,-64.7057647705078,-64.7645034790039,-64.8220672607422,-64.9599151611328,-65.0360870361328,-65.2594223022461,-65.3600158691406,-65.4758834838867,-65.7546920776367,-65.9267044067383,-66.0125503540039,-66.0831069946289,-66.2486343383789,-66.3242645263672,-66.448974609375,-66.7043914794922,-66.6315383911133,-66.4288101196289,-66.359619140625,-66.210205078125,-65.8494186401367,-65.7557144165039,-65.6664581298828,-65.6072235107422,-65.4834976196289,-65.3439483642578,-65.2281799316406,-65.0016632080078,-65.04638671875,-64.8739776611328,-64.7032241821289,-64.7663040161133,-64.8521499633789,-64.9122085571289,-65.0861282348633,-65.3188934326172,-65.2602005004883,-65.1920928955078,-65.0423812866211,-64.9424285888672,-64.8313980102539,-64.8658676147461,-64.90576171875,-64.8825225830078,-64.8166961669922,-64.7258758544922,-64.6894989013672,-64.641357421875,-64.6478500366211,-64.5568389892578,-64.54150390625,-64.2118148803711,-64.14501953125,-63.9159202575684,-63.8726577758789,-63.8319358825684,-64.056396484375,-63.8747062683105,-63.702880859375,-63.5676765441895,-63.5092315673828,-63.3580093383789,-63.3159141540527,-63.2927742004395,-63.2168922424316,-63.10791015625,-62.9107894897461,-62.7006874084473,-62.7183609008789,-62.7500953674316,-62.5856437683105,-62.4830551147461,-62.447265625,-62.4218788146973,-62.2177238464355,-61.9555206298828,-61.9235801696777,-61.91162109375,-61.8772468566895,-61.7765121459961,-61.6568832397461,-61.4922828674316,-61.4276351928711,-61.3504867553711,-61.2770500183105,-61.2819786071777,-61.3761215209961,-61.4609870910645,-61.1067390441895,-61.0707969665527,-61.0315437316895,-61.0676765441895,-61.10107421875,-61.1653327941895,-61.2837905883789,-61.3872528076172,-61.4979019165039,-61.5687522888184,-61.6474075317383,-61.7192344665527,-61.7938957214355,-62.0268058776855,-62.2649879455566,-62.5140151977539,-62.76806640625,-63.0318336486816,-63.0892066955566,-63.1557121276855,-63.3062973022461,-63.3808097839355,-63.4568367004395,-63.5443382263184,-63.60400390625,-63.5582504272461,-63.5448226928711,-63.5676765441895,-63.6097640991211,-63.7611351013184,-63.8206520080566,-63.8913116455078,-63.9236793518066,-63.9997062683105,-64.044921875,-64.0446243286133,-64.1008834838867,-64.1669921875,-64.2860794067383,-64.3385238647461,-64.312255859375,-64.2756881713867,-64.3345718383789,-64.3782196044922,-64.4687957763672,-64.5784683227539,-64.6916046142578,-64.8256301879883,-64.8623504638672,-65.0868225097656,-65.1720733642578,-65.2349090576172,-65.32958984375,-65.3442916870117,-65.3860855102539,-65.4285125732422,-65.450439453125,-65.481689453125,-65.564453125,-65.6619110107422,-65.7381286621094,-65.8353042602539,-65.8869171142578,-65.9784164428711,-66.0021438598633,-66.0376434326172,-66.125732421875,-66.1925277709961,-66.1930618286133,-66.0995559692383,-65.8680114746094,-65.9419479370117,-66.1463851928711,-66.1252975463867,-66.0906219482422,-66.0216827392578,-65.9170455932617,-65.7776870727539,-65.6818389892578,-65.6157684326172,-65.52001953125,-65.5022430419922,-65.587158203125,-65.7282180786133,-65.6920394897461,-65.65673828125,-64.9029312133789,-64.7512741088867,-64.4488220214844,-64.4068908691406,-64.4481430053711,-64.3307571411133,-64.3404312133789,-64.3588409423828,-64.36572265625,-64.354248046875,-64.2350158691406,-64.135498046875,-64.1827163696289,-64.0931625366211,-63.7483406066895,-63.4602508544922,-63.3680152893066,-63.6144561767578,-63.9064445495605,-64.087158203125,-64.33642578125,-64.6001968383789,-64.6811065673828,-64.7466812133789,-64.8319320678711,-64.8731460571289,-64.9128875732422,-64.827392578125,-64.56005859375,-64.3970718383789,-64.3511276245117,-64.3146438598633,-64.404052734375,-64.4822235107422,-64.5363235473633,-64.6327133178711,-64.6420440673828,-64.5936508178711,-64.7785186767578,-64.8979034423828,-65.0572814941406,-65.2823257446289,-65.5450210571289,-65.8844680786133,-65.9556121826172,-66.1097640991211,-66.066650390625,-66.0265655517578,-66.0648956298828,-66.0897445678711,-66.1827163696289,-66.1073226928711,-66.1437530517578,-66.2515640258789,-66.3519515991211,-66.4398422241211,-66.5109405517578,-66.7071762084961,-66.8724594116211,-66.908203125,-66.918701171875,-66.9765625,-67.0840835571289,-67.1248550415039,-67.1709976196289,-67.2132339477539,-67.2496109008789,-67.2707061767578,-67.2906723022461,-67.3152847290039,-67.366943359375,-67.3998107910156,-67.4525833129883,-67.4725570678711,-67.4619598388672,-67.4385299682617,-67.4279251098633,-67.4537582397461,-67.4772491455078,-67.49365234375,-67.48779296875,-67.4549331665039,-67.4244155883789,-67.4138641357422,-67.4326705932617,-67.4866180419922,-67.5312042236328,-67.5957489013672,-67.6579055786133,-67.698974609375,-67.7306594848633,-67.7553176879883,-67.78466796875,-67.80224609375,-67.7999038696289,-67.7916946411133,-67.7752914428711,-67.7741241455078,-67.7811508178711,-67.7822799682617,-67.7776336669922,-67.7670364379883,-67.78466796875,-67.7864685058594,-67.7899398803711,-67.7925338745117,-67.7957992553711,-67.7977066040039,-67.8003463745117,-67.8028335571289,-67.8067855834961,-67.9348678588867,-68.0967712402344,-68.2354965209961,-68.3108901977539,-68.3580093383789,-68.3768997192383,-68.4803695678711,-68.6685562133789,-68.8287124633789,-68.8874053955078,-68.9372100830078,-69.0031280517578,-69.048583984375,-69.0642547607422,-69.0501937866211,-69.1462860107422,-69.2428741455078,-69.3021469116211,-69.35888671875,-69.4714889526367,-69.6297836303711,-69.717529296875,-69.8717269897461,-70.0077209472656,-70.0382385253906,-70.0671844482422,-70.1796875,-70.248291015625,-70.2789001464844,-70.3044891357422,-70.3064422607422,-70.2871627807617,-70.2962417602539,-70.3334503173828,-70.4078674316406,-70.4210968017578,-70.4666061401367,-70.5963821411133,-70.7022476196289,-70.7074203491211,-70.692138671875,-70.6897964477539,-70.7109375,-70.7533187866211,-70.7991714477539,-70.8377914428711,-70.8368148803711,-70.8650360107422,-70.8979949951172,-70.9262237548828,-70.9601516723633,-70.9999084472656,-71.0602569580078,-71.1346664428711,-71.2016143798828,-71.3272933959961,-71.4190368652344,-71.5175323486328,-71.9336395263672,-72.3497543334961,-72.7659225463867,-73.1820373535156,-73.59814453125,-74.0142593383789,-74.4303741455078,-74.6632308959961,-74.7088928222656],[-126.092094421387,-126.064010620117,-126.186813354492,-126.229644775391,-126.2314453125,-126.208549499512,-126.115280151367,-126.092094421387],[-54.5543937683105,-54.7086906433105,-54.7438468933105,-54.7865257263184,-54.8185081481934,-54.8635711669922,-54.8554229736328,-54.8130836486816,-54.7887687683105,-54.7826194763184,-54.7640647888184,-54.7331047058105,-54.6187477111816,-54.5591773986816,-54.5376930236816,-54.5543937683105],[-54.0937004089355,-54.0199203491211,-53.9806671142578,-54.2383766174316,-54.2692375183105,-54.2861328125,-54.2887687683105,-54.2776336669922,-54.2589836120605,-54.1993637084961,-54.1376953125,-54.0937004089355],[-124.153663635254,-124.139793395996,-124.362312316895,-124.457221984863,-124.493942260742,-124.51781463623,-124.630950927734,-124.649856567383,-124.623291015625,-124.547172546387,-124.421485900879,-124.309127807617,-124.153663635254],[-126.641212463379,-126.680419921875,-126.743408203125,-126.814208984375,-126.938575744629,-126.951263427734,-126.940032958984,-126.904884338379,-126.896865844727,-126.925827026367,-126.826080322266,-126.738136291504,-126.698150634766,-126.64990234375,-126.62816619873,-126.625778198242,-126.641212463379],[-61.8011207580566,-62.2195320129395,-62.5526351928711,-62.7996063232422,-63.04150390625,-63.5658645629883,-63.6258773803711,-63.6762199401855,-63.776611328125,-63.8849105834961,-64.4400405883789,-64.4851989746094,-64.3729476928711,-64.2437515258789,-64.1314468383789,-63.7601585388184,-63.2919883728027,-63.0888175964355,-62.8585433959961,-62.6334495544434,-62.1330070495605,-62.0430679321289,-61.817138671875,-61.73583984375,-61.6961441040039,-61.7455101013184,-61.8011207580566],[-124.977737426758,-125.001556396484,-125.025970458984,-124.995658874512,-124.987014770508,-124.990821838379,-124.937850952148,-124.916397094727,-124.907470703125,-124.908447265625,-124.977737426758],[-125.184127807617,-125.195114135742,-125.259574890137,-125.358451843262,-125.345306396484,-125.301170349121,-125.260940551758,-125.195999145508,-125.139503479004,-125.126472473145,-125.091400146484,-125.074028015137,-125.112983703613,-125.184127807617],[-55.5361328125,-55.5696792602539,-55.6007843017578,-55.6293449401855,-55.6338844299316,-55.6044921875,-55.5271987915039,-55.4692878723145,-55.4727554321289,-55.5038070678711,-55.5361328125],[-127.197311401367,-126.700927734375,-126.20386505127,-125.839157104492,-125.615242004395,-125.534324645996,-125.482078552246,-125.420455932617,-125.313957214355,-125.233200073242,-125.06640625,-124.934661865234,-124.904640197754,-124.932426452637,-124.9306640625,-124.830612182617,-124.642868041992,-124.495948791504,-124.185882568359,-123.995803833008,-123.937156677246,-123.854499816895,-123.820022583008,-123.752288818359,-123.626564025879,-123.497016906738,-123.472854614258,-123.457962036133,-123.443061828613,-123.415481567383,-123.389892578125,-123.366302490234,-123.283790588379,-123.310653686523,-123.334518432617,-123.445899963379,-123.484573364258,-123.536468505859,-123.573150634766,-123.594619750977,-123.916938781738,-124.115234375,-124.37621307373,-124.689407348633,-124.868263244629,-125.017242431641,-125.120704650879,-125.140281677246,-125.135688781738,-124.934761047363,-124.849655151367,-124.817031860352,-124.800247192383,-124.812652587891,-124.820747375488,-124.838714599609,-124.868316650391,-124.904449462891,-124.927345275879,-125.168212890625,-125.362739562988,-125.460304260254,-125.489456176758,-125.543113708496,-125.660491943359,-125.828521728516,-125.811958312988,-125.702293395996,-125.644241333008,-125.654632568359,-125.693702697754,-125.728034973145,-125.79638671875,-125.835441589355,-125.918365478516,-125.951667785645,-125.983833312988,-125.937690734863,-125.935394287109,-126.020309448242,-126.04833984375,-126.074897766113,-126.099853515625,-126.168853759766,-126.243598937988,-126.269721984863,-126.279640197754,-126.304489135742,-126.418601989746,-126.444534301758,-126.499855041504,-126.519142150879,-126.548530578613,-126.563720703125,-126.557479858398,-126.541893005371,-126.442771911621,-126.157814025879,-126.134086608887,-126.347557067871,-126.403167724609,-126.462799072266,-126.525238037109,-126.558258056641,-126.592872619629,-126.683113098145,-126.744621276855,-126.84937286377,-126.9033203125,-126.926071166992,-126.947944641113,-126.977096557617,-127.048728942871,-127.114303588867,-127.16552734375,-127.195892333984,-127.20751953125,-127.179107666016,-127.179641723633,-127.192329406738,-127.215675354004,-127.249801635742,-127.268402099609,-127.271530151367,-127.2900390625,-127.349411010742,-127.397903442383,-127.429786682129,-127.467132568359,-127.674858093262,-127.770462036133,-127.816307067871,-127.863914489746,-127.873001098633,-127.828178405762,-127.839164733887,-127.850830078125,-127.946685791016,-127.962936401367,-127.905868530273,-127.8740234375,-127.83154296875,-127.641403198242,-127.578125,-127.486518859863,-127.489356994629,-127.524017333984,-127.528999328613,-127.465919494629,-127.526214599609,-127.75146484375,-127.749710083008,-127.731147766113,-127.864692687988,-127.963676452637,-128.058349609375,-128.135650634766,-128.267425537109,-128.349899291992,-128.346038818359,-128.300827026367,-128.241561889648,-128.101318359375,-127.918060302734,-127.713035583496,-127.197311401367],[-55.458740234375,-55.5324211120605,-55.5834007263184,-55.6307601928711,-55.730712890625,-55.941162109375,-56.0311050415039,-56.0439453125,-56.0306663513184,-55.9999008178711,-55.9608383178711,-55.87353515625,-55.8411140441895,-55.8150863647461,-55.7955093383789,-55.7853507995605,-55.7847175598145,-55.7999992370605,-55.8713874816895,-55.9620094299316,-56.078125,-56.1065444946289,-56.1211929321289,-56.1356430053711,-56.1957550048828,-56.3824234008789,-56.454345703125,-56.4547843933105,-56.4839363098145,-56.5393562316895,-56.6939964294434,-56.7323226928711,-56.7495574951172,-56.7471694946289,-56.7541007995605,-56.7895011901855,-56.8388671875,-56.8486328125,-56.8292007446289,-56.8092269897461,-56.806884765625,-56.8221664428711,-56.7567901611328,-56.6106414794922,-56.5009269714355,-56.4275856018066,-56.3764152526855,-56.3218269348145,-56.2470703125,-56.1793937683105,-56.1483879089355,-56.1221694946289,-56.12744140625,-56.1641616821289,-56.1612777709961,-56.0750007629395,-55.927001953125,-55.8733367919922,-55.7647438049316,-55.6744651794434,-55.530029296875,-55.5029296875,-55.5270042419434,-55.5836906433105,-55.7176284790039,-56.0399894714355,-56.1401863098145,-56.1211929321289,-56.0516128540039,-55.978515625,-55.90185546875,-55.8698234558105,-55.88232421875,-55.8920440673828,-56.0873031616211,-56.0412101745605,-55.8152351379395,-55.6781234741211,-55.48974609375,-55.3759307861328,-55.379150390625,-55.3544921875,-55.3553695678711,-55.3438491821289,-55.2899398803711,-55.2801780700684,-55.2830085754395,-55.266357421875,-55.2295417785645,-55.20703125,-55.2002944946289,-55.2249984741211,-55.2593231201172,-55.3424835205078,-55.3319320678711,-55.3531761169434,-55.3347625732422,-55.2523422241211,-55.2473640441895,-55.2538108825684,-55.2445297241211,-55.1761207580566,-55.0631828308105,-55.0261726379395,-55.0104026794434,-55.0159187316895,-54.9826164245605,-54.9105491638184,-54.8436546325684,-54.7818832397461,-54.7176246643066,-54.65087890625,-54.5790519714355,-54.502197265625,-54.4691925048828,-54.4806137084961,-54.4654312133789,-54.4634780883789,-54.4482421875,-54.3890609741211,-54.3561515808105,-54.3167495727539,-54.2708015441895,-53.9577140808105,-53.8624534606934,-53.7549819946289,-53.6194343566895,-53.569580078125,-53.56005859375,-53.5734405517578,-53.671142578125,-53.758056640625,-53.809326171875,-53.8249015808105,-53.84521484375,-53.9032211303711,-54.1612777709961,-54.0995101928711,-53.95068359375,-53.8528785705566,-53.8477516174316,-53.8868141174316,-53.9615249633789,-53.9695816040039,-53.9660148620605,-53.8861312866211,-53.7840805053711,-53.6980438232422,-53.7063484191895,-53.7746086120605,-53.7946281433105,-53.8855476379395,-54.0677757263184,-54.1144523620605,-54.104248046875,-53.93701171875,-53.8527374267578,-53.79931640625,-53.7388648986816,-53.6444320678711,-53.5520515441895,-53.4113273620605,-53.361083984375,-53.2754364013672,-53.2202644348145,-53.1273422241211,-53.0572738647461,-53.0426750183105,-53.027587890625,-53.020751953125,-53.0373039245605,-53.0602035522461,-53.1357421875,-53.18212890625,-53.22509765625,-53.3011703491211,-53.3343276977539,-53.405517578125,-53.5312004089355,-53.6097640991211,-53.5602073669434,-53.5418472290039,-53.5694313049316,-53.7042961120605,-53.7101554870605,-53.7582015991211,-53.8695793151855,-53.7935562133789,-53.6530265808105,-53.6382331848145,-53.6576194763184,-53.6950187683105,-53.8616714477539,-53.8636703491211,-53.8377418518066,-53.8053703308105,-53.76513671875,-53.67236328125,-53.603759765625,-53.5037612915039,-53.28271484375,-53.08544921875,-52.9209938049316,-52.88330078125,-52.8660163879395,-52.8720245361328,-52.9549789428711,-52.9982414245605,-53.11083984375,-53.1538581848145,-53.175537109375,-53.1698265075684,-53.1576652526855,-53.1224632263184,-53.0568389892578,-52.9450225830078,-52.8731956481934,-52.8169441223145,-52.7824211120605,-52.7449226379395,-52.71142578125,-52.7032699584961,-52.6721687316895,-52.6536636352539,-52.6685066223145,-52.6836433410645,-52.9124031066895,-52.8881340026855,-52.882080078125,-52.8892059326172,-52.9617156982422,-53.0319366455078,-53.0697746276855,-53.1148414611816,-53.1669921875,-53.2136726379395,-53.2548828125,-53.2913093566895,-53.3230476379395,-53.3817405700684,-53.5361328125,-53.5677719116211,-53.5897941589355,-53.6163558959961,-53.5951652526855,-53.5813484191895,-53.6121597290039,-53.5796356201172,-53.5784645080566,-53.5973625183105,-53.6363754272461,-53.6953620910645,-53.7743148803711,-53.8600082397461,-54.0095710754395,-54.0760231018066,-54.1023941040039,-54.1328125,-54.1737289428711,-54.1732902526855,-54.1552238464355,-54.0926780700684,-53.9705085754395,-53.8690910339355,-53.8495101928711,-53.8778800964355,-53.9008293151855,-53.9397468566895,-53.989013671875,-54.0472679138184,-54.1918449401855,-54.2184066772461,-54.23388671875,-54.4046859741211,-54.4344749450684,-54.4559097290039,-54.4881362915039,-54.5625495910645,-54.5423851013184,-54.4632301330566,-54.4739227294922,-54.5745124816895,-54.6511726379395,-54.7446746826172,-54.801513671875,-54.8566398620605,-55.0904312133789,-55.0992164611816,-55.1396484375,-55.2549324035645,-55.3157196044922,-55.4012680053711,-55.4792976379395,-55.5307121276855,-55.65234375,-55.7885246276855,-55.8447265625,-55.880615234375,-55.9499015808105,-55.9582023620605,-55.9544906616211,-55.9192352294922,-55.83837890625,-55.7718276977539,-55.6100578308105,-55.4915046691895,-55.4012222290039,-55.3608894348145,-55.1908226013184,-54.9756355285645,-54.8695335388184,-54.7953605651855,-54.7846183776855,-54.8910179138184,-54.9459495544434,-55.0350112915039,-55.0745620727539,-55.1965827941895,-55.3663101196289,-55.3907699584961,-55.4126968383789,-55.4346694946289,-55.4606437683105,-55.4986343383789,-55.5761222839355,-55.7747077941895,-55.8113784790039,-55.862060546875,-56.0813484191895,-56.1272468566895,-56.0836906433105,-55.8670883178711,-55.8443832397461,-55.85791015625,-55.91845703125,-56.0201187133789,-56.0896453857422,-56.1214332580566,-56.1505851745605,-56.2212867736816,-56.2629852294922,-56.3257789611816,-56.4595680236816,-56.7223129272461,-56.7741203308105,-56.9524917602539,-57.4734382629395,-57.6598167419434,-57.8840827941895,-57.925537109375,-58.2393074035645,-58.3332023620605,-58.3269538879395,-58.3368644714355,-58.4280281066895,-58.5088882446289,-58.6131324768066,-58.9411659240723,-59.1169471740723,-59.2192840576172,-59.259765625,-59.3206558227539,-59.3624038696289,-59.362060546875,-59.3408660888672,-59.2720680236816,-58.9608421325684,-58.7105979919434,-58.6049842834473,-58.5026397705078,-58.3355445861816,-58.3302268981934,-58.4922370910645,-58.6061515808105,-58.7225570678711,-58.9438018798828,-59.1667938232422,-59.1676750183105,-59.0634269714355,-58.8418006896973,-58.8191909790039,-58.8871116638184,-58.9064483642578,-58.8772468566895,-58.8434104919434,-58.7164535522461,-58.6873550415039,-58.6416015625,-58.5456085205078,-58.4937477111816,-58.4036636352539,-58.3586921691895,-58.3187522888184,-58.1861343383789,-58.0496635437012,-58.0055656433105,-57.9905242919922,-58.0405769348145,-58.0818405151367,-58.0989265441895,-58.0490760803223,-57.9906768798828,-57.9800758361816,-58.0968704223633,-58.19091796875,-58.2188949584961,-58.2133827209473,-58.1827125549316,-58.1074256896973,-58.0158195495605,-57.9612312316895,-57.8560523986816,-57.7913093566895,-57.798828125,-57.8974571228027,-57.9290542602539,-57.9261703491211,-57.7125015258789,-57.6079597473145,-57.4655303955078,-57.4326171875,-57.3604469299316,-57.33056640625,-57.2374000549316,-57.1795921325684,-57.26416015625,-57.29443359375,-57.2980003356934,-57.27490234375,-57.2421379089355,-57.1316413879395,-57.0532684326172,-57.0056648254395,-57.0127449035645,-57.0373039245605,-57.0359382629395,-56.9763641357422,-56.8251457214355,-56.8054656982422,-56.7501945495605,-56.6824226379395,-56.6190452575684,-56.5179672241211,-56.2073707580566,-56.0255851745605,-55.902099609375,-55.8658218383789,-55.6904296875,-55.6595687866211,-55.7006340026855,-55.6664085388184,-55.5216293334961,-55.4964332580566,-55.4532241821289,-55.458740234375],[-127.924659729004,-127.941261291504,-127.981246948242,-128.044525146484,-128.091796875,-128.148773193359,-128.142379760742,-128.122756958008,-128.03173828125,-127.998687744141,-127.98681640625,-127.932518005371,-127.916358947754,-127.916305541992,-127.924659729004],[-55.3612327575684,-55.4088897705078,-55.4196281433105,-55.3998031616211,-55.3464851379395,-55.2740707397461,-55.2935523986816,-55.3612327575684],[-79.38427734375,-79.4255828857422,-79.5206069946289,-79.5968704223633,-79.6437454223633,-79.3348617553711,-79.2712936401367,-79.2702102661133,-79.3166046142578,-79.3289489746094,-79.3515090942383,-79.38427734375],[-131.029296875,-131.047256469727,-131.080520629883,-131.103424072266,-131.117340087891,-131.107131958008,-131.098083496094,-131.010650634766,-131.029296875],[-128.368743896484,-128.445404052734,-128.419876098633,-128.412506103516,-128.42626953125,-128.435943603516,-128.439788818359,-128.364898681641,-128.247268676758,-128.248443603516,-128.298156738281,-128.323776245117,-128.343551635742,-128.368743896484],[-128.936874389648,-128.968704223633,-129.102340698242,-129.151016235352,-129.25048828125,-129.267776489258,-129.263534545898,-129.245941162109,-129.215042114258,-129.186187744141,-128.994003295898,-128.940338134766,-128.936874389648],[-129.313720703125,-129.328720092773,-129.370025634766,-129.409729003906,-129.477783203125,-129.500137329102,-129.514739990234,-129.501068115234,-129.471435546875,-129.450729370117,-129.343505859375,-129.313720703125],[-80.731689453125,-80.8023452758789,-81.0098648071289,-81.0965881347656,-81.3522491455078,-81.8390579223633,-82.0050277709961,-82.0392608642578,-81.9511260986328,-81.9013671875,-81.8473129272461,-81.3353576660156,-81.1355972290039,-80.900390625,-80.7653350830078,-80.71044921875,-80.7095260620117,-80.731689453125],[-131.753707885742,-131.65234375,-131.622177124023,-131.634674072266,-131.795257568359,-131.879684448242,-131.916366577148,-131.971786499023,-131.904388427734,-131.81005859375,-131.727294921875,-131.610595703125,-131.455230712891,-131.572799682617,-131.590576171875,-131.443908691406,-131.429977416992,-131.383010864258,-131.273620605469,-131.259719848633,-131.259963989258,-131.327056884766,-131.319915771484,-131.259170532227,-131.142623901367,-131.116165161133,-131.221542358398,-131.421875,-131.511123657227,-131.562072753906,-131.623687744141,-131.809677124023,-132.092239379883,-132.165084838867,-132.238571166992,-132.259963989258,-132.258102416992,-132.229537963867,-132.144927978516,-132.143753051758,-132.468704223633,-132.504837036133,-132.546768188477,-132.524215698242,-132.345413208008,-132.153900146484,-132.035949707031,-131.989501953125,-131.893112182617,-131.853469848633,-131.753707885742],[-128.552444458008,-128.506546020508,-128.509918212891,-128.576812744141,-128.6240234375,-128.678955078125,-128.730911254883,-128.735534667969,-128.749420166016,-128.766448974609,-128.746337890625,-128.769622802734,-128.831207275391,-128.899810791016,-129.022842407227,-129.084716796875,-129.094879150391,-129.175918579102,-129.184326171875,-129.177688598633,-129.111083984375,-129.084091186523,-129.060348510742,-129.033248901367,-128.97021484375,-128.857727050781,-128.740371704102,-128.632659912109,-128.552444458008],[-129.167724609375,-129.173248291016,-129.27685546875,-129.305709838867,-129.323883056641,-129.331237792969,-129.314361572266,-129.253082275391,-129.251159667969,-129.238189697266,-129.195220947266,-129.177001953125,-129.167724609375],[-79.938232421875,-79.9393081665039,-80.0041046142578,-80.0393524169922,-80.06787109375,-80.0740203857422,-80.0497055053711,-79.9745635986328,-79.938232421875],[-129.848587036133,-129.868560791016,-129.934371948242,-130.151412963867,-130.3056640625,-130.410736083984,-130.517578125,-130.451995849609,-130.394821166992,-130.195022583008,-130.035400390625,-129.944732666016,-129.754837036133,-129.768951416016,-129.848587036133],[-130.236282348633,-130.267242431641,-130.337554931641,-130.384231567383,-130.4072265625,-130.470260620117,-130.537490844727,-130.58984375,-130.624603271484,-130.641845703125,-130.646286010742,-130.637893676758,-130.643707275391,-130.66357421875,-130.683441162109,-130.703170776367,-130.707275390625,-130.695693969727,-130.646926879883,-130.49462890625,-130.447998046875,-130.397308349609,-130.315872192383,-130.298477172852,-130.236282348633],[-132.655517578125,-132.564071655273,-132.344436645508,-132.303375244141,-132.261627197266,-132.215927124023,-132.166107177734,-132.155136108398,-132.175109863281,-132.214492797852,-132.564880371094,-132.574127197266,-132.567138671875,-132.53466796875,-132.464401245117,-132.186965942383,-132.171676635742,-132.152252197266,-132.114013671875,-132.110595703125,-132.135879516602,-132.134414672852,-131.940826416016,-131.819625854492,-131.695953369141,-131.667633056641,-131.685409545898,-131.702529907227,-131.821151733398,-131.88916015625,-131.922317504883,-131.928085327148,-131.957427978516,-132.011322021484,-132.347259521484,-132.520462036133,-132.6748046875,-132.747512817383,-132.692581176758,-132.65478515625,-132.546234130859,-132.46240234375,-132.425003051758,-132.431350708008,-132.670166015625,-132.845016479492,-132.897994995117,-132.899566650391,-132.913375854492,-133.05224609375,-133.079483032227,-133.09765625,-133.097946166992,-133.063873291016,-133.048385620117,-132.991455078125,-132.89306640625,-132.655517578125],[-130.927139282227,-130.950302124023,-130.959030151367,-130.953475952148,-130.921783447266,-130.906829833984,-130.777038574219,-130.757995605469,-130.75341796875,-130.763381958008,-130.805130004883,-130.927139282227],[-130.575332641602,-130.493270874023,-130.312545776367,-130.214065551758,-130.203903198242,-130.349426269531,-130.535507202148,-130.575332641602],[-60.9944839477539,-60.9827156066895,-61.1370086669922,-61.1913108825684,-61.1958503723145,-61.1881828308105,-61.1575698852539,-61.0869178771973,-61.0485382080078,-60.9664039611816,-60.9553680419922,-60.9944839477539],[-78.8265151977539,-78.8772888183594,-78.913818359375,-78.9070281982422,-78.8568801879883,-78.8284149169922,-78.8215866088867,-78.7994155883789,-78.7618637084961,-78.7245178222656,-78.6687545776367,-78.6571731567383,-78.6728057861328,-78.7101593017578,-78.7613754272461,-78.8265151977539],[-79.9775848388672,-80.0286178588867,-80.0888671875,-80.0574722290039,-80.0050811767578,-79.8744659423828,-79.8521499633789,-79.8104019165039,-79.7491760253906,-79.6810073852539,-79.6058578491211,-79.5797348022461,-79.632568359375,-79.6879425048828,-79.9775848388672],[-78.9355926513672,-79.0179672241211,-79.0838851928711,-79.1754913330078,-79.2278289794922,-79.2736358642578,-79.1422805786133,-79.1360778808594,-79.1422805786133,-79.18212890625,-79.2218246459961,-79.4074249267578,-79.455322265625,-79.4951171875,-79.5267639160156,-79.605712890625,-79.7647476196289,-79.4974594116211,-79.4946823120117,-79.5447235107422,-79.5645523071289,-79.7811050415039,-79.9045867919922,-79.9874954223633,-80.0082550048828,-80.0007858276367,-79.7900390625,-79.5963439941406,-79.5152893066406,-79.4823684692383,-79.4679183959961,-79.4689407348633,-79.4588928222656,-79.4476547241211,-79.435302734375,-79.4320297241211,-79.4762725830078,-79.5118179321289,-79.55419921875,-79.5363311767578,-79.4583053588867,-79.3926315307617,-79.33935546875,-79.3053207397461,-79.2724151611328,-79.2611389160156,-79.2457504272461,-79.21044921875,-79.1551818847656,-79.12353515625,-79.1002426147461,-79.0777359008789,-78.9949722290039,-78.9631805419922,-78.9403381347656,-78.9424285888672,-78.9312057495117,-78.9066390991211,-78.9355926513672],[-79.5181655883789,-79.553466796875,-79.577392578125,-79.5507354736328,-79.5817337036133,-79.5835494995117,-79.5701217651367,-79.5528793334961,-79.51123046875,-79.4910659790039,-79.482177734375,-79.4845733642578,-79.4965286254883,-79.5181655883789],[-79.8669967651367,-79.8944854736328,-79.9436569213867,-79.9457015991211,-79.8981475830078,-79.8605422973633,-79.82666015625,-79.8350067138672,-79.8669967651367],[-61.7436065673828,-61.6595191955566,-61.6374969482422,-61.7952613830566,-61.9754905700684,-62.0112342834473,-62.0072250366211,-61.9833030700684,-61.9375,-61.8930625915527,-61.8483390808105,-61.7436065673828],[-79.7165069580078,-79.7322235107422,-79.7751922607422,-79.7920379638672,-79.8084487915039,-79.8382339477539,-79.81591796875,-79.8191375732422,-79.8108444213867,-79.7678756713867,-79.7425765991211,-79.7267074584961,-79.7134780883789,-79.7165069580078],[-69.1600570678711,-69.2208557128906,-69.3017120361328,-69.330810546875,-69.3528289794922,-69.3163146972656,-69.3115234375,-69.3299789428711,-69.30322265625,-69.1951675415039,-69.1938018798828,-69.1806640625,-69.1551742553711,-69.1600570678711],[-80.2852554321289,-80.3172378540039,-80.3246536254883,-80.2989730834961,-80.2566452026367,-80.2099609375,-80.167236328125,-80.1830520629883,-80.2405319213867,-80.2852554321289],[-80.064208984375,-80.1670913696289,-80.1222152709961,-80.0836410522461,-80.0411605834961,-79.9558639526367,-79.8986282348633,-79.9496078491211,-80.064208984375],[-64.4070281982422,-64.4419403076172,-64.5582046508789,-64.7379379272461,-64.8089904785156,-64.8337860107422,-64.83642578125,-64.7825698852539,-64.6462860107422,-64.5325241088867,-64.4998016357422,-64.4070281982422],[-68.2337875366211,-68.3241271972656,-68.365234375,-68.3678741455078,-68.3382797241211,-68.2347640991211,-68.1418914794922,-68.0875930786133,-67.97802734375,-67.9142074584961,-67.8475646972656,-67.81884765625,-67.84423828125,-67.9223175048828,-68.0123062133789,-68.2337875366211],[-78.5316467285156,-78.6688995361328,-78.6690979003906,-78.6120147705078,-78.3995590209961,-78.24169921875,-78.2788619995117,-78.3724594116211,-78.5316467285156],[-64.8326187133789,-64.8568344116211,-64.8797836303711,-64.9542465209961,-65.0543975830078,-65.0915069580078,-65.3938980102539,-65.4268035888672,-65.43212890625,-65.3316421508789,-65.1297912597656,-64.9544448852539,-64.7896499633789,-64.75634765625,-64.6695861816406,-64.6909713745117,-64.6963882446289,-64.7323226928711,-64.78759765625,-64.8326187133789],[-93.0439453125,-93.0848159790039,-93.1765594482422,-93.1966781616211,-93.0757827758789,-92.9930191040039,-92.9999542236328,-93.0439453125],[-65.0305633544922,-65.008056640625,-64.9810562133789,-64.9605484008789,-64.9466781616211,-64.9235382080078,-64.8651351928711,-64.8455123901367,-64.8470764160156,-64.8965835571289,-64.927734375,-65.1659164428711,-65.2302703857422,-65.2353515625,-65.2105484008789,-65.1739273071289,-65.1256332397461,-65.068359375,-65.0305633544922],[-79.5453109741211,-79.4662094116211,-79.3360366821289,-79.2864761352539,-79.2720260620117,-79.3064498901367,-79.3239288330078,-79.3722610473633,-79.462158203125,-79.5418472290039,-79.611328125,-79.6687469482422,-79.7142562866211,-79.7633285522461,-79.8161163330078,-79.8963394165039,-80.004150390625,-80.0919952392578,-80.2049331665039,-80.2651901245117,-80.2761688232422,-80.2798385620117,-80.2751007080078,-80.2600555419922,-80.2346725463867,-80.1785659790039,-80.0215835571289,-79.9267578125,-79.8680648803711,-79.7125473022461,-79.6495590209961,-79.59765625,-79.5453109741211],[-64.8238296508789,-64.6318359375,-64.5153274536133,-64.4650421142578,-64.4180679321289,-64.4783172607422,-64.5464859008789,-64.6574172973633,-64.8373031616211,-64.9012222290039,-64.9564895629883,-64.9307632446289,-64.8419418334961,-64.8271026611328,-64.849853515625,-64.8487777709961,-64.8238296508789],[-74.0004425048828,-74.0535659790039,-74.2535171508789,-74.49951171875,-74.62646484375,-74.6199722290039,-74.564208984375,-74.5009307861328,-74.394775390625,-74.1089324951172,-74.0167999267578,-73.9881820678711,-74.0004425048828],[-70.3370590209961,-70.4063491821289,-70.54150390625,-70.6865692138672,-70.7660675048828,-70.8375549316406,-70.8512649536133,-70.9861373901367,-71.1369094848633,-71.2201156616211,-71.1348648071289,-71.013671875,-70.8346252441406,-70.67431640625,-70.442626953125,-70.3667907714844,-70.29150390625,-70.2688522338867,-70.28857421875,-70.3370590209961],[-82.00048828125,-81.9605484008789,-81.9485855102539,-81.9644012451172,-81.9901885986328,-82.0258331298828,-82.1137237548828,-82.3880386352539,-82.490966796875,-82.5682601928711,-83.0158233642578,-83.0713806152344,-83.1296844482422,-83.2523880004883,-83.3768081665039,-83.6988830566406,-83.7144012451172,-83.7286148071289,-83.7609405517578,-83.9031219482422,-83.9124069213867,-83.9104995727539,-83.8992691040039,-83.7390594482422,-83.3764114379883,-83.2894515991211,-83.1109390258789,-83.0262680053711,-82.9657669067383,-82.7064437866211,-82.459716796875,-82.2347640991211,-82.1292495727539,-82.047607421875,-82.00048828125],[-77.876708984375,-77.7920913696289,-77.7037048339844,-77.65478515625,-77.5384826660156,-77.5272979736328,-77.53271484375,-77.5939025878906,-77.6576690673828,-77.7914581298828,-77.9424285888672,-78.0244140625,-78.2559585571289,-78.46875,-78.5367660522461,-78.50732421875,-78.4172821044922,-78.2349090576172,-77.9339370727539,-77.876708984375],[-76.6775894165039,-76.7831573486328,-76.921875,-77.0572280883789,-77.36474609375,-77.1336898803711,-76.7636184692383,-76.6524429321289,-76.6775894165039],[-77.64208984375,-77.7140655517578,-77.9288101196289,-77.9579086303711,-77.9659652709961,-77.9313430786133,-77.7107925415039,-77.6172866821289,-77.5693893432617,-77.5636215209961,-77.64208984375],[-84.9196243286133,-84.8851089477539,-84.8420867919922,-84.7712860107422,-84.6125030517578,-84.5679168701172,-84.5011291503906,-84.2664108276367,-84.1799774169922,-84.1334991455078,-84.0848617553711,-83.9001007080078,-83.7225570678711,-83.4907684326172,-83.4071273803711,-83.2222671508789,-83.2009811401367,-82.9905776977539,-82.6676330566406,-82.5857925415039,-82.2716751098633,-82.1588821411133,-82.0499954223633,-81.9289016723633,-81.7872085571289,-81.6761245727539,-81.6673812866211,-81.680908203125,-81.720947265625,-81.9026336669922,-81.8871078491211,-81.7161102294922,-81.3356475830078,-81.1040496826172,-81.0235824584961,-81.0050277709961,-80.921142578125,-80.8289566040039,-80.6942901611328,-80.6075668334961,-80.56884765625,-80.5792007446289,-80.6682662963867,-80.4505844116211,-80.2613296508789,-80.3020477294922,-80.5040512084961,-80.7117614746094,-80.9535140991211,-81.0138626098633,-81.04638671875,-81.1796875,-81.3717346191406,-81.9633255004883,-82.14599609375,-82.3781280517578,-82.4117202758789,-82.4670867919922,-82.5714797973633,-82.9296875,-83.0338821411133,-83.0386734008789,-83.0161590576172,-83.0651321411133,-83.185546875,-83.303955078125,-83.4943389892578,-83.5835952758789,-83.6170883178711,-83.6379852294922,-83.66162109375,-83.728271484375,-84.0221176147461,-84.1416015625,-84.2604522705078,-84.3076171875,-84.3874969482422,-84.5062026977539,-84.5545959472656,-84.6329116821289,-84.7955551147461,-84.9615249633789,-85.2381362915039,-85.3926315307617,-85.4955062866211,-85.5661087036133,-85.7141647338867,-85.7387237548828,-85.7689437866211,-85.8046875,-86.3015670776367,-86.57568359375,-86.8468704223633,-86.9152374267578,-87.0529327392578,-87.1519012451172,-87.1771469116211,-87.19384765625,-87.1889114379883,-87.1543960571289,-87.0319366455078,-86.9320297241211,-86.8860397338867,-86.4217300415039,-86.30859375,-86.2520980834961,-86.252197265625,-86.274169921875,-86.3544921875,-86.3749008178711,-86.374267578125,-86.3438491821289,-86.2276306152344,-86.1882781982422,-86.1142120361328,-86.0746078491211,-86.01708984375,-85.961669921875,-85.81396484375,-85.6990737915039,-85.5546875,-85.5230484008789,-85.4955062866211,-85.4424285888672,-85.2411193847656,-85.1762237548828,-85.13037109375,-85.1053695678711,-85.1303253173828,-85.226318359375,-85.2427749633789,-85.2399444580078,-85.0560531616211,-84.9196243286133],[-84.6747589111328,-84.7269973754883,-84.7829132080078,-84.8302764892578,-84.8689422607422,-84.93115234375,-85.0719757080078,-85.0963363647461,-85.1362762451172,-85.14404296875,-85.1741638183594,-85.1756820678711,-85.1496124267578,-85.0313949584961,-84.9385757446289,-84.9198226928711,-84.8894500732422,-84.8695297241211,-84.7573776245117,-84.6917495727539,-84.6026382446289,-84.6022491455078,-84.6262664794922,-84.6747589111328],[-83.7259750366211,-83.5975112915039,-83.4694366455078,-83.26318359375,-83.2337341308594,-83.2339324951172,-83.263671875,-83.3324203491211,-83.3814468383789,-83.4954071044922,-83.537109375,-83.5832061767578,-83.6065444946289,-83.6363754272461,-83.6443786621094,-83.6306686401367,-83.6495132446289,-83.7875518798828,-83.8092269897461,-83.7981872558594,-83.701904296875,-83.7865219116211,-83.8135757446289,-83.93896484375,-84.0084991455078,-84.1182632446289,-84.1299362182617,-84.1432113647461,-84.1932144165039,-84.2229461669922,-84.2709045410156,-84.3701171875,-84.4505844116211,-84.4673843383789,-84.4563446044922,-84.4071807861328,-84.1222610473633,-83.9503860473633,-83.7869644165039,-83.7013626098633,-83.6936569213867,-83.7148895263672,-83.76513671875,-83.7259750366211],[-83.1234817504883,-83.0238800048828,-82.9481430053711,-82.9313507080078,-83.0108413696289,-83.0598602294922,-83.1479034423828,-83.2139129638672,-83.2325668334961,-83.2378387451172,-83.2222671508789,-83.1234817504883],[-108.092720031738,-107.966453552246,-107.805519104004,-107.833343505859,-107.895164489746,-107.943946838379,-107.965133666992,-108.059722900391,-108.092720031738],[-62.6815414428711,-62.805419921875,-62.8716316223145,-62.8250961303711,-62.7569808959961,-62.6644020080566,-62.6252937316895,-62.4697303771973,-62.4167976379395,-62.3963394165039,-62.484619140625,-62.6815414428711],[-107.899848937988,-107.950248718262,-107.969535827637,-108.003959655762,-108.073341369629,-108.152244567871,-108.151123046875,-108.120849609375,-108.127532958984,-108.048973083496,-107.990867614746,-107.974899291992,-107.989356994629,-107.931793212891,-107.905174255371,-107.890968322754,-107.899848937988],[-109.166404724121,-109.053909301758,-108.970504760742,-108.909622192383,-108.886032104492,-108.893852233887,-108.920166015625,-109.096244812012,-109.161521911621,-109.18359375,-109.166404724121],[-73.6217269897461,-74.1090850830078,-74.3740768432617,-74.480712890625,-74.5733947753906,-74.6786117553711,-74.7459945678711,-74.749267578125,-74.7314453125,-74.70654296875,-74.37939453125,-74.1113739013672,-73.8807144165039,-73.5840301513672,-73.4937515258789,-73.459228515625,-73.4352569580078,-73.4015655517578,-73.398193359375,-73.4071807861328,-73.6217269897461],[-109.323150634766,-109.36083984375,-109.497955322266,-109.469139099121,-109.341697692871,-109.323539733887,-109.323150634766],[-86.5955581665039,-86.63818359375,-86.7059631347656,-86.861083984375,-86.8925247192383,-86.9083023071289,-86.908447265625,-86.8945846557617,-86.8470687866211,-86.937744140625,-86.9598159790039,-86.9491653442383,-86.898681640625,-86.8848648071289,-86.833984375,-86.7021026611328,-86.5699234008789,-86.4519500732422,-86.421142578125,-86.4303207397461,-86.4200210571289,-86.3903274536133,-86.3824234008789,-86.3964385986328,-86.4469223022461,-86.4896469116211,-86.5460433959961,-86.5955581665039],[-75.6758804321289,-75.15380859375,-75.1031265258789,-75.078125,-75.0634765625,-75.0623550415039,-75.0728530883789,-75.1238784790039,-75.1273422241211,-75.0863800048828,-75.0905227661133,-75.1272964477539,-75.2019500732422,-75.3144989013672,-75.4001007080078,-75.7800827026367,-76.0489730834961,-76.332763671875,-76.6939468383789,-76.8588333129883,-76.9441909790039,-77.0048828125,-77.075927734375,-77.1570816040039,-77.2242126464844,-77.3043975830078,-77.305908203125,-77.2285690307617,-77.1258850097656,-76.9447250366211,-76.740234375,-76.688232421875,-76.5958023071289,-76.3644485473633,-76.1728057861328,-76.0882797241211,-75.9827575683594,-75.8665008544922,-75.6758804321289],[-78.9827194213867,-79.0640640258789,-79.1740264892578,-79.1747512817383,-79.1534652709961,-78.9525909423828,-78.8686981201172,-78.8285140991211,-78.9827194213867],[-104.540672302246,-104.595993041992,-104.699462890625,-104.851119995117,-104.965232849121,-105.041748046875,-105.051368713379,-104.993995666504,-104.907272338867,-104.700393676758,-104.602005004883,-104.472122192383,-104.444534301758,-104.440475463867,-104.457130432129,-104.540672302246],[-74.880859375,-74.9593276977539,-75.072509765625,-75.3101577758789,-75.4002380371094,-75.4034194946289,-75.3961868286133,-75.3701705932617,-75.2873992919922,-75.1997528076172,-75.07470703125,-74.983642578125,-74.884765625,-74.8189468383789,-74.7982406616211,-74.8309555053711,-74.8279342651367,-74.8128890991211,-74.8185577392578,-74.844970703125,-74.880859375],[-101.845901489258,-101.88720703125,-101.944633483887,-102.266357421875,-102.308151245117,-102.2705078125,-102.1533203125,-102.074363708496,-102.013374328613,-101.828369140625,-101.759330749512,-101.732955932617,-101.721633911133,-101.732032775879,-101.794281005859,-101.845901489258],[-100.217231750488,-100.248779296875,-100.287940979004,-100.36572265625,-100.397315979004,-100.442573547363,-100.480659484863,-100.496925354004,-100.521041870117,-100.573387145996,-100.596534729004,-100.615966796875,-100.625396728516,-100.624656677246,-100.599899291992,-100.598335266113,-100.611572265625,-100.600631713867,-100.565475463867,-100.520309448242,-100.413963317871,-100.329933166504,-100.288963317871,-100.206886291504,-100.178466796875,-100.217231750488],[-99.9946823120117,-100.018020629883,-100.141311645508,-100.195709228516,-100.241996765137,-100.247360229492,-100.237060546875,-100.186965942383,-100.153129577637,-100.072799682617,-100.035354614258,-100.005615234375,-99.9946823120117],[-90.4925765991211,-90.5744171142578,-90.6257858276367,-90.6674346923828,-90.6858901977539,-90.7715835571289,-90.7656707763672,-90.7423858642578,-90.6627883911133,-90.5997085571289,-90.5398483276367,-90.5106506347656,-90.4853515625,-90.4925765991211],[-79.2106475830078,-79.2797317504883,-79.3613739013672,-79.3904800415039,-79.40576171875,-79.3911590576172,-79.354736328125,-79.3052215576172,-79.24267578125,-79.1449737548828,-78.9304733276367,-78.9000015258789,-78.8041000366211,-78.7718276977539,-78.6620101928711,-78.6501922607422,-78.6890640258789,-78.6890640258789,-78.6501922607422,-78.5966796875,-78.4579086303711,-78.3325729370117,-78.3004913330078,-78.2724609375,-78.2340774536133,-78.2289581298828,-78.2870101928711,-78.43896484375,-78.5329055786133,-78.5517578125,-78.560302734375,-78.5956573486328,-78.7053680419922,-78.7791976928711,-78.8526840209961,-79.0536117553711,-79.2106475830078],[-90.1998062133789,-90.1773910522461,-90.2672882080078,-90.2954559326172,-90.3302764892578,-90.3640670776367,-90.4646911621094,-90.4920425415039,-90.4551239013672,-90.3772506713867,-90.3220748901367,-90.2528305053711,-90.2285614013672,-90.1998062133789],[-76.995361328125,-77.1216354370117,-77.2150344848633,-77.2755889892578,-77.3219223022461,-77.37939453125,-77.3580551147461,-77.3515167236328,-77.3409194946289,-77.3187026977539,-77.1875534057617,-77.1091842651367,-76.9940948486328,-76.7457046508789,-76.68408203125,-76.6688461303711,-76.6700210571289,-76.6874542236328,-76.810302734375,-76.8693389892578,-76.9112319946289,-76.995361328125],[-101.171730041504,-101.253517150879,-101.268501281738,-101.261520385742,-101.267631530762,-101.289497375488,-101.2177734375,-101.207328796387,-101.230125427246,-101.328468322754,-101.356498718262,-101.351318359375,-101.312889099121,-101.244873046875,-101.098342895508,-101.031150817871,-101.000633239746,-101.049171447754,-101.086868286133,-101.126953125,-101.171730041504],[-95.513671875,-95.3809051513672,-95.382080078125,-95.3994140625,-95.4374465942383,-95.4962387084961,-95.5785217285156,-95.6843719482422,-95.7301254272461,-95.6958923339844,-95.670166015625,-95.6658248901367,-95.6828155517578,-95.7041015625,-95.7636260986328,-95.8062057495117,-95.8177719116211,-95.79833984375,-95.8118209838867,-95.8582000732422,-95.8934555053711,-95.9560546875,-95.9859390258789,-95.9779281616211,-95.9947738647461,-95.9788589477539,-95.9362258911133,-95.8758316040039,-95.7977523803711,-95.7066421508789,-95.6024932861328,-95.513671875],[-139.043121337891,-139.125732421875,-139.256988525391,-139.291412353516,-139.139602661133,-139.072647094727,-138.931533813477,-138.878845214844,-139.043121337891],[-67.9146957397461,-67.9402847290039,-68.2023391723633,-68.2213821411133,-68.09326171875,-67.9891128540039,-67.9088363647461,-67.8291015625,-67.7545852661133,-67.8449249267578,-67.9146957397461],[-78.0290985107422,-77.9778289794922,-77.9691467285156,-78.0399932861328,-78.3072204589844,-78.4700698852539,-78.5523910522461,-78.6620559692383,-78.7953109741211,-78.8481979370117,-78.789306640625,-78.5785675048828,-78.40185546875,-78.3441925048828,-78.2955093383789,-78.267333984375,-78.262451171875,-78.2007369995117,-78.1452102661133,-78.0290985107422],[-79.4306640625,-79.3902816772461,-79.364990234375,-79.4024429321289,-79.5528335571289,-79.8816909790039,-80.0475082397461,-79.9711380004883,-79.9544982910156,-79.9778366088867,-80.046875,-80.1614761352539,-80.2273406982422,-80.2444839477539,-80.2686462402344,-80.2997055053711,-80.32958984375,-80.3978500366211,-80.4480438232422,-80.7782211303711,-80.7947692871094,-80.7775421142578,-80.7266159057617,-80.6525421142578,-80.4659194946289,-80.45068359375,-80.4383316040039,-80.4242172241211,-80.294921875,-80.2136764526367,-80.1688461303711,-80.1246109008789,-80.061767578125,-79.9708480834961,-79.8695831298828,-79.71484375,-79.593994140625,-79.4306640625],[-97.439453125,-97.4086380004883,-97.3506851196289,-97.3057632446289,-97.2784652709961,-97.236083984375,-97.0963363647461,-96.9890594482422,-96.8751983642578,-96.6945266723633,-96.2999496459961,-96.1837387084961,-96.0609893798828,-95.9513626098633,-95.8548812866211,-95.7513732910156,-95.5855026245117,-95.4375534057617,-95.3741760253906,-95.3195343017578,-95.2677688598633,-95.295166015625,-95.3594741821289,-95.465576171875,-95.6142044067383,-95.6856460571289,-95.8021469116211,-95.8946228027344,-96.0240249633789,-96.2676315307617,-96.4015655517578,-96.5988235473633,-97.0083999633789,-97.263671875,-97.4720230102539,-97.7047805786133,-97.8853530883789,-98.2350616455078,-98.2579574584961,-98.2730484008789,-98.2801818847656,-98.2960433959961,-98.320556640625,-98.3755798339844,-98.4318389892578,-98.5396499633789,-98.7038116455078,-98.7752456665039,-98.8296432495117,-98.859130859375,-98.8637237548828,-98.8788604736328,-98.9044952392578,-98.9640121459961,-99.057373046875,-99.0938491821289,-99.0733871459961,-99.0906219482422,-99.2539978027344,-99.3179702758789,-99.4408645629883,-99.4946746826172,-99.5640640258789,-99.557373046875,-99.5132827758789,-99.4557113647461,-99.08544921875,-98.9122009277344,-98.7236328125,-98.5035095214844,-98.4559555053711,-98.4503860473633,-98.4666061401367,-98.5353469848633,-98.5585403442383,-98.5367202758789,-98.4483871459961,-98.494873046875,-98.5343704223633,-98.5482406616211,-98.5459976196289,-98.475830078125,-98.3893508911133,-98.2223129272461,-98.15576171875,-98.0413513183594,-98.1629867553711,-98.288818359375,-98.3044967651367,-98.3012237548828,-98.2682113647461,-98.2386703491211,-98.2004928588867,-98.0807571411133,-97.8889694213867,-97.7907180786133,-97.6912155151367,-97.6043472290039,-97.411376953125,-97.382568359375,-97.3856887817383,-97.4601516723633,-97.4694366455078,-97.439453125],[-86.9130401611328,-86.7987823486328,-86.6912155151367,-86.6127395629883,-86.5633773803711,-86.5309066772461,-86.5152359008789,-86.5576705932617,-86.7343215942383,-86.8549346923828,-86.9839859008789,-87.0438003540039,-87.1908187866211,-87.263916015625,-87.3232421875,-87.3231430053711,-87.1681137084961,-87.1072769165039,-86.9130401611328],[-100.308349609375,-100.321235656738,-100.537254333496,-100.620651245117,-100.647758483887,-100.666938781738,-100.678321838379,-100.635307312012,-100.537940979004,-100.433937072754,-100.276123046875,-100.321098327637,-100.3232421875,-100.305511474609,-100.308349609375],[-74.7088928222656,-74.762451171875,-74.8566360473633,-74.9961395263672,-75.1793899536133,-75.4012756347656,-75.7919464111328,-75.8193359375,-75.8759307861328,-76.0202178955078,-76.1511688232422,-76.185791015625,-76.24853515625,-76.464599609375,-76.5861358642578,-76.6964874267578,-76.8199691772461,-77.0733337402344,-77.2667007446289,-77.5965347290039,-77.8792495727539,-78.2147979736328,-78.458251953125,-78.7204132080078,-78.8455581665039,-79.0024871826172,-79.171875,-79.0830612182617,-79.0592269897461,-79.0660629272461,-79.0479965209961,-79.029052734375,-79.0261688232422,-79.0116729736328,-78.9807662963867,-78.9459991455078,-78.9208450317383,-78.9150924682617,-78.9392547607422,-79.0367126464844,-79.1737289428711,-79.4462432861328,-79.7620162963867,-80.0357437133789,-80.24755859375,-80.6826171875,-81.0282211303711,-81.2776412963867,-81.50732421875,-81.7609329223633,-81.9741668701172,-82.2133255004883,-82.4390640258789,-82.6900405883789,-82.8662109375,-83.0299758911133,-83.1419448852539,-83.149658203125,-83.1095199584961,-83.0731430053711,-83.0037078857422,-82.8677749633789,-82.7441864013672,-82.6451187133789,-82.5453186035156,-82.4883346557617,-82.417236328125,-82.408203125,-82.3047866821289,-82.1903762817383,-82.1378402709961,-82.1965789794922,-82.2407684326172,-82.28125,-82.3268051147461,-82.3682632446289,-82.4073715209961,-82.4465866088867,-82.4850616455078,-82.5152359008789,-82.5510711669922,-82.7603988647461,-82.9193344116211,-83.1792907714844,-83.3973159790039,-83.5926742553711,-83.469482421875,-83.4801330566406,-83.5247573852539,-83.615966796875,-83.6692810058594,-83.76318359375,-83.9130325317383,-83.977783203125,-84.0291976928711,-84.08837890625,-84.1077651977539,-84.1151885986328,-84.1504821777344,-84.1281204223633,-84.1231918334961,-84.1251907348633,-84.1494598388672,-84.1921920776367,-84.3367156982422,-84.4017028808594,-84.4404830932617,-84.5015640258789,-84.561767578125,-84.665771484375,-84.7793960571289,-84.8270492553711,-84.8759765625,-85.070068359375,-85.2641143798828,-85.4581985473633,-85.6522445678711,-85.8463363647461,-86.0403747558594,-86.2344741821289,-86.4285659790039,-86.4955520629883,-86.6721725463867,-86.9218292236328,-87.2080078125,-87.4942398071289,-87.743896484375,-87.9205093383789,-87.9874496459961,-88.16064453125,-88.378173828125,-88.6117630004883,-88.898681640625,-89.0625991821289,-89.1856460571289,-89.273193359375,-89.4556655883789,-89.5505828857422,-89.775390625,-89.9010238647461,-89.99365234375,-90.0399398803711,-90.091796875,-90.3201217651367,-90.6070785522461,-90.744384765625,-90.7973175048828,-90.84033203125,-90.9160690307617,-91.04345703125,-91.2206497192383,-91.38720703125,-91.518310546875,-91.6473159790039,-91.8583984375,-92.0051727294922,-92.1717758178711,-92.2986831665039,-92.3484420776367,-92.4145965576172,-92.4608840942383,-92.5005798339844,-92.583251953125,-92.732666015625,-92.8367156982422,-92.9962463378906,-93.0517120361328,-93.1552200317383,-93.2579650878906,-93.3778762817383,-93.463623046875,-93.5642547607422,-93.7077178955078,-93.8035659790039,-93.8516159057617,-94.05517578125,-94.4141616821289,-94.6208953857422,-94.6753387451172,-94.705078125,-94.7125473022461,-94.7127914428711,-94.803466796875,-94.8425827026367,-94.8603973388672,-94.8543472290039,-94.8748016357422,-94.9393539428711,-95.1552734375,-95.1582489013672,-95.1620635986328,-95.3979034423828,-95.8243179321289,-96.2506866455078,-96.6770553588867,-97.1034698486328,-97.5298385620117,-97.9561996459961,-98.3826217651367,-98.8089828491211,-99.2353515625,-99.6617202758789,-100.088134765625,-100.514503479004,-100.940872192383,-101.367286682129,-101.793647766113,-102.220016479492,-102.646430969238,-103.072799682617,-103.499168395996,-103.925582885742,-104.351951599121,-104.7783203125,-105.204689025879,-105.631103515625,-106.057472229004,-106.483840942383,-106.910255432129,-107.336624145508,-107.762985229492,-108.189407348633,-108.615768432617,-109.042137145996,-109.468551635742,-109.894920349121,-110.3212890625,-110.747657775879,-111.174072265625,-111.600440979004,-112.026809692383,-112.453224182129,-112.879592895508,-113.305953979492,-113.732376098633,-114.158744812012,-114.585105895996,-115.011520385742,-115.437889099121,-115.8642578125,-116.290626525879,-116.71704864502,-117.143409729004,-117.569778442383,-117.996192932129,-118.422554016113,-118.848922729492,-119.275344848633,-119.701713562012,-120.128067016602,-120.554489135742,-120.980857849121,-121.4072265625,-121.833595275879,-122.26000213623,-122.686370849609,-122.788772583008,-122.826705932617,-122.924171447754,-122.962692260742,-123.002296447754,-123.027252197266,-123.049217224121,-123.063278198242,-123.077293395996,-123.08642578125,-123.117630004883,-123.109321594238,-123.077293395996,-123.079544067383,-123.15013885498,-123.181884765625,-123.196342468262,-123.191070556641,-123.229438781738,-123.183937072754,-123.067291259766,-122.947654724121,-122.912986755371,-122.879104614258,-122.964454650879,-123.015533447266,-123.174263000488,-123.276756286621,-123.29052734375,-123.286277770996,-123.264068603516,-123.247703552246,-123.222991943359,-123.190673828125,-123.179595947266,-123.1875,-123.324996948242,-123.336669921875,-123.322410583496,-123.335647583008,-123.398979187012,-123.436965942383,-123.508209228516,-123.530570983887,-123.858940124512,-123.891845703125,-123.948387145996,-124.028610229492,-124.053802490234,-124.024024963379,-123.992630004883,-123.959518432617,-123.922760009766,-123.847122192383,-123.817192077637,-123.739067077637,-123.612747192383,-123.582466125488,-123.708305358887,-123.762702941895,-123.818008422852,-123.874412536621,-123.90380859375,-123.904251098633,-123.884963989258,-123.823829650879,-123.78466796875,-123.787696838379,-123.825439453125,-123.880126953125,-123.93359375,-123.945899963379,-123.863037109375,-123.865730285645,-123.957412719727,-123.971389770508,-123.972114562988,-123.984916687012,-124.058792114258,-124.1416015625,-124.28125,-124.412590026855,-124.483261108398,-124.702285766602,-124.782379150391,-124.784278869629,-124.934173583984,-124.933349609375,-124.98558807373,-125.043609619141,-125.056694030762,-124.936805725098,-124.862648010254,-124.854248046875,-124.857521057129,-124.875434875488,-124.85986328125,-124.933601379395,-124.949264526367,-124.931060791016,-124.942535400391,-124.985397338867,-125.058784484863,-125.209869384766,-125.47631072998,-125.507171630859,-125.525978088379,-125.539352416992,-125.555572509766,-125.585838317871,-125.610153198242,-125.641304016113,-125.697555541992,-125.7412109375,-125.772415161133,-125.839599609375,-125.965042114258,-126.024124145508,-126.094337463379,-126.23656463623,-126.404495239258,-126.44995880127,-126.447456359863,-126.416122436523,-126.238914489746,-126.067230224609,-125.897613525391,-125.904098510742,-125.98070526123,-126.370315551758,-126.492973327637,-126.514350891113,-126.51732635498,-126.472221374512,-126.397117614746,-126.374603271484,-126.418212890625,-126.488174438477,-126.521781921387,-126.48462677002,-126.51732635498,-126.562889099121,-126.631790161133,-126.960403442383,-127.014060974121,-127.057571411133,-127.267471313477,-127.35693359375,-127.441207885742,-127.590873718262,-127.708106994629,-127.714302062988,-127.689155578613,-127.632713317871,-127.419677734375,-127.346580505371,-127.280654907227,-126.968124389648,-126.735404968262,-126.691459655762,-127.034088134766,-127.338729858398,-127.442726135254,-127.575736999512,-127.609573364258,-127.644866943359,-127.66870880127,-127.714057922363,-127.728713989258,-127.747512817383,-127.818946838379,-127.850540161133,-127.869148254395,-127.863227844238,-127.829978942871,-127.727638244629,-127.858795166016,-127.843315124512,-127.795364379883,-127.67333984375,-127.549713134766,-127.437942504883,-127.242233276367,-127.175735473633,-127.007957458496,-126.95947265625,-126.899993896484,-126.826309204102,-126.738571166992,-126.713966369629,-126.752639770508,-126.895217895508,-126.901420593262,-126.938179016113,-127.127044677734,-127.160598754883,-127.193992614746,-127.20825958252,-127.187110900879,-126.995208740234,-126.951316833496,-126.951370239258,-126.966400146484,-127.008255004883,-127.019332885742,-127.006401062012,-127.013229370117,-127.034866333008,-127.066215515137,-127.107070922852,-127.519233703613,-127.56029510498,-127.71337890625,-127.791893005371,-127.834327697754,-127.902191162109,-127.995407104492,-128.102249145508,-128.193557739258,-128.357620239258,-128.037490844727,-128.029144287109,-128.060302734375,-128.051559448242,-128.021301269531,-127.940231323242,-127.943359375,-128.038238525391,-128.183990478516,-128.240966796875,-128.271530151367,-128.275146484375,-128.19677734375,-128.132369995117,-128.108795166016,-128.053268432617,-128.10595703125,-128.365051269531,-128.451950073242,-128.524703979492,-128.65234375,-128.868560791016,-129.080902099609,-129.129531860352,-129.171585083008,-129.114456176758,-129.021438598633,-128.935592651367,-128.854598999023,-128.850433349609,-128.905609130859,-128.833053588867,-128.542144775391,-128.478622436523,-128.358047485352,-128.291061401367,-128.132705688477,-128.079208374023,-127.927833557129,-127.950042724609,-128.115142822266,-128.207229614258,-128.369140625,-128.469635009766,-128.511764526367,-128.600341796875,-128.675537109375,-128.750793457031,-128.767883300781,-128.763671875,-128.745956420898,-128.714752197266,-128.652877807617,-128.560440063477,-128.532119750977,-128.65087890625,-128.704772949219,-128.890197753906,-128.927825927734,-128.944000244141,-128.959381103516,-129.013961791992,-129.056396484375,-129.208114624023,-129.231735229492,-129.240341186523,-129.257858276367,-129.284286499023,-129.46240234375,-129.563720703125,-129.686721801758,-129.82177734375,-129.911865234375,-130.074371337891,-130.263275146484,-130.335250854492,-130.232849121094,-130.0859375,-130.063522338867,-130.043304443359,-129.790771484375,-129.626022338867,-129.794967651367,-129.8984375,-130.084228515625,-130.290328979492,-130.396774291992,-130.430282592773,-130.393463134766,-130.388626098633,-130.370025634766,-130.350479125977,-130.307220458984,-130.218948364258,-130.140869140625,-130.108642578125,-129.948532104492,-129.89013671875,-129.78076171875,-129.560638427734,-129.630126953125,-129.666641235352,-129.701324462891,-129.734176635742,-129.765472412109,-129.795166015625,-129.811904907227,-129.815628051758,-129.837738037109,-129.877151489258,-129.985214233398,-130.048477172852,-130.091796875,-130.058395385742,-129.995849609375,-129.985153198242,-130.044036865234,-130.079986572266,-130.092971801758,-130.094680786133,-130.085098266602,-130.060348510742,-130.020309448242,-130.025100708008,-130.014053344727,-130.022903442383,-130.055953979492,-130.097854614258,-130.214706420898,-130.413131713867,-130.477096557617,-130.649078369141,-130.74169921875,-130.930221557617,-131.082916259766,-131.199417114258,-131.335800170898,-131.471878051758,-131.575103759766,-131.651519775391,-131.824264526367,-131.833099365234,-131.885986328125,-131.866165161133,-131.962509155273,-132.104293823242,-132.062896728516,-132.031539916992,-132.157028198242,-132.337982177734,-132.279403686523,-132.232177734375,-132.301666259766,-132.442474365234,-132.550491333008,-132.691513061523,-132.815521240234,-132.916854858398,-133.001419067383,-133.120407104492,-133.275299072266,-133.422561645508,-133.401123046875,-133.54638671875,-133.673919677734,-133.820755004883,-133.965713500977,-134.069198608398,-134.218505859375,-134.296966552734,-134.329650878906,-134.363525390625,-134.39306640625,-134.410202026367,-134.440765380859,-134.621978759766,-134.67724609375,-134.802383422852,-134.9072265625,-134.943740844727,-135.0712890625,-135.050827026367,-135.036666870117,-135.051025390625,-135.260787963867,-135.367874145508,-135.475921630859,-135.702590942383,-135.934677124023,-136.09716796875,-136.321823120117,-136.247131347656,-136.277984619141,-136.347854614258,-136.466354370117,-136.466735839844,-136.578750610352,-136.813278198242,-136.939315795898,-137.126220703125,-137.277542114258,-137.438568115234,-137.520904541016,-137.484176635742,-137.543701171875,-137.593307495117,-137.696624755859,-137.870544433594,-138.001113891602,-138.187438964844,-138.317626953125,-138.45361328125,-138.632278442383,-138.705474853516,-138.868759155273,-139.04345703125,-139.185150146484,-139.136962890625,-139.079254150391,-139.079254150391,-139.234756469727,-139.467971801758,-139.676315307617,-139.830657958984,-139.973297119141,-140.196929931641,-140.452819824219,-140.525451660156,-140.762741088867,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-140.860015869141,-140.405120849609,-139.976608276367,-139.181533813477,-138.689880371094,-138.291015625,-138.128356933594,-137.869430541992,-137.259963989258,-137.070404052734,-136.717330932617,-136.498687744141,-136.122360229492,-135.866653442383,-135.362167358398,-135.258834838867,-135.231201171875,-135.406936645508,-135.4345703125,-135.638000488281,-135.876312255859,-135.894729614258,-135.939010620117,-135.924758911133,-135.872863769531,-135.695220947266,-135.589996337891,-135.575546264648,-135.651275634766,-135.742630004883,-135.849700927734,-135.910202026367,-135.691467285156,-135.614929199219,-135.499557495117,-135.292831420898,-135.254989624023,-135.229782104492,-135.199020385742,-135.140823364258,-134.852874755859,-134.493835449219,-134.456832885742,-134.4912109375,-134.495361328125,-134.473678588867,-134.451461791992,-134.408935546875,-134.242034912109,-134.189895629883,-134.134033203125,-134.077484130859,-133.899963378906,-133.879776000977,-133.947448730469,-134.018402099609,-134.1650390625,-134.17431640625,-133.948043823242,-133.694046020508,-133.475921630859,-133.293655395508,-133.163131713867,-133.084411621094,-133.028274536133,-132.915328979492,-132.84033203125,-132.526809692383,-132.452346801758,-132.403900146484,-132.412734985352,-132.478942871094,-132.568359375,-132.570617675781,-132.537551879883,-132.488464355469,-132.333984375,-132.232421875,-132.163421630859,-131.934127807617,-131.581832885742,-131.44091796875,-131.318954467773,-131.215866088867,-131.136383056641,-131.031845092773,-130.990631103516,-130.926177978516,-130.665481567383,-130.498443603516,-130.396377563477,-130.274963378906,-130.174957275391,-130.043304443359,-129.944976806641,-129.898040771484,-129.730072021484,-129.675628662109,-129.623001098633,-129.538436889648,-129.538177490234,-129.648284912109,-130.458847045898,-130.708557128906,-130.832077026367,-130.960113525391,-131.207962036133,-131.306350708008,-131.472946166992,-131.86279296875,-131.937789916992,-131.98876953125,-132.128753662109,-132.196823120117,-132.330764770508,-132.481201171875,-132.686721801758,-132.817474365234,-132.967971801758,-133.089462280273,-133.228225708008,-133.378952026367,-133.418304443359,-133.373397827148,-133.348388671875,-133.196823120117,-133.138031005859,-133.192184448242,-133.319534301758,-133.336669921875,-133.304016113281,-132.705993652344,-132.57763671875,-132.532669067383,-132.542236328125,-132.704345703125,-132.739105224609,-132.764694213867,-132.770126342773,-132.755462646484,-132.718948364258,-132.545166015625,-132.358062744141,-132.213958740234,-132.134384155273,-131.919631958008,-131.833404541016,-131.786911010742,-131.781051635742,-131.820159912109,-131.788375854492,-131.631790161133,-131.562942504883,-131.342926025391,-131.303024291992,-131.32470703125,-131.293899536133,-131.209030151367,-131.161727905273,-131.112838745117,-131.063415527344,-131.013427734375,-130.986282348633,-130.981979370117,-130.970657348633,-130.914306640625,-130.875045776367,-130.660705566406,-130.515960693359,-130.353607177734,-130.117630004883,-129.572113037109,-129.264846801758,-129.109130859375,-129.032913208008,-128.984329223633,-128.89892578125,-128.883682250977,-128.916809082031,-128.938613891602,-129.138336181641,-129.157913208008,-129.13623046875,-129.101715087891,-129.054351806641,-128.971435546875,-128.85302734375,-128.705520629883,-128.38671875,-128.359191894531,-128.278610229492,-128.095855712891,-127.764945983887,-127.683792114258,-127.974014282227,-128.0341796875,-128.043655395508,-127.988922119141,-128.121475219727,-128.170104980469,-128.168060302734,-128.127304077148,-128.040481567383,-127.991012573242,-127.861618041992,-127.752830505371,-127.376853942871,-127.225975036621,-127.138481140137,-126.926811218262,-126.83349609375,-126.758697509766,-126.684913635254,-126.612159729004,-126.250442504883,-126.063819885254,-125.907417297363,-125.727783203125,-125.524955749512,-125.386772155762,-125.171875,-125.166847229004,-125.261573791504,-125.35693359375,-125.345512390137,-125.219390869141,-125.227882385254,-125.201179504395,-125.114013671875,-125.07958984375,-125.03101348877,-124.968254089355,-124.88916015625,-124.793647766113,-124.767929077148,-124.862594604492,-124.920021057129,-124.962600708008,-124.990386962891,-124.952438354492,-124.745109558105,-124.706344604492,-124.639938354492,-124.555030822754,-124.502586364746,-124.444473266602,-124.441497802734,-124.467193603516,-124.47191619873,-124.406936645508,-124.34937286377,-124.124603271484,-124.138465881348,-124.398384094238,-124.453903198242,-124.481346130371,-124.472068786621,-124.426170349121,-124.338088989258,-124.111724853516,-124.049659729004,-123.609130859375,-123.528427124023,-123.46044921875,-123.361480712891,-123.248977661133,-123.213676452637,-123.144485473633,-123.110404968262,-123.076606750488,-123.025779724121,-122.956680297852,-122.78539276123,-122.704879760742,-122.387496948242,-122.070068359375,-121.741851806641,-121.531097412109,-121.336235046387,-120.962448120117,-120.814651489258,-120.292526245117,-120.139991760254,-119.852828979492,-118.868705749512,-118.744873046875,-118.48560333252,-118.306976318359,-118.09521484375,-117.830322265625,-117.31128692627,-117.226943969727,-117.131736755371,-117.025726318359,-116.549949645996,-116.424560546875,-116.334075927734,-116.222702026367,-116.059478759766,-116.065238952637,-116.251609802246,-116.24341583252,-116.166748046875,-115.936080932617,-115.883255004883,-115.806350708008,-115.631156921387,-115.442283630371,-115.239837646484,-114.993751525879,-114.620162963867,-114.413864135742,-114.218162536621,-114.11083984375,-114.092041015625,-114.051116943359,-113.988182067871,-113.964401245117,-114.020797729492,-114.05322265625,-114.095947265625,-114.274757385254,-114.765281677246,-114.852195739746,-115.127044677734,-115.175926208496,-115.186767578125,-115.167083740234,-115.201850891113,-115.426849365234,-115.434471130371,-115.288467407227,-115.133209228516,-115.011177062988,-114.856735229492,-114.662895202637,-114.429397583008,-114.267044067383,-114.175727844238,-114.051071166992,-113.893211364746,-113.681938171387,-113.214988708496,-113.074951171875,-112.879440307617,-112.503028869629,-112.435157775879,-112.314552307129,-112.236724853516,-112.101318359375,-111.710884094238,-111.575729370117,-111.45068359375,-111.290824890137,-111.192184448242,-111.15478515625,-111.08740234375,-110.990043640137,-110.804878234863,-110.371971130371,-110.216262817383,-110.101951599121,-110.073928833008,-110.04248046875,-109.9365234375,-109.904243469238,-109.831344604492,-109.760154724121,-109.68603515625,-109.63037109375,-109.224311828613,-109.081253051758,-109.038032531738,-108.994483947754,-108.967674255371,-108.949905395508,-108.890968322754,-108.851997375488,-108.815185546875,-108.715133666992,-108.680221557617,-108.613327026367,-108.592918395996,-108.491500854492,-108.346969604492,-107.988716125488,-107.930519104004,-107.9091796875,-107.929443359375,-107.991310119629,-108.088424682617,-108.220802307129,-108.344329833984,-108.459083557129,-108.496047973633,-108.455276489258,-108.218162536621,-108.157661437988,-108.101463317871,-108.049613952637,-108.001754760742,-107.957954406738,-107.760940551758,-107.704887390137,-107.480323791504,-107.373680114746,-107.291358947754,-107.259475708008,-107.278076171875,-107.564453125,-107.710350036621,-107.730857849121,-107.740234375,-107.745994567871,-107.72509765625,-107.626174926758,-107.499221801758,-107.451271057129,-107.418846130371,-107.402099609375,-107.329193115234,-107.2001953125,-107.156494140625,-107.253753662109,-107.323341369629,-107.347846984863,-107.283157348633,-107.318458557129,-107.482368469238,-107.567481994629,-107.64404296875,-107.650924682617,-107.638381958008,-107.64990234375,-107.753028869629,-107.865089416504,-107.954048156738,-107.972122192383,-107.958396911621,-107.890914916992,-107.763084411621,-107.728614807129,-107.787452697754,-107.798294067383,-107.761032104492,-107.509376525879,-107.446189880371,-107.351119995117,-107.22412109375,-107.124801635742,-106.99365234375,-106.922561645508,-106.835647583008,-106.790725708008,-106.710990905762,-106.668411254883,-106.534866333008,-106.45947265625,-106.424263000488,-106.429489135742,-106.404396057129,-106.271240234375,-106.132133483887,-106.039306640625,-105.933059692383,-105.85693359375,-105.781204223633,-105.750190734863,-105.774314880371,-105.932228088379,-106.027153015137,-106.2373046875,-106.458053588867,-106.543304443359,-106.566650390625,-106.608497619629,-106.780418395996,-106.853713989258,-106.94580078125,-107.043312072754,-107.146186828613,-107.298149108887,-107.499114990234,-107.619338989258,-107.741500854492,-107.734230041504,-107.677635192871,-107.734176635742,-108.027191162109,-108.104583740234,-108.261032104492,-108.322807312012,-108.367919921875,-108.686576843262,-108.718116760254,-108.640922546387,-108.345756530762,-108.3134765625,-107.766357421875,-107.435935974121,-106.830657958984,-106.713478088379,-106.324272155762,-106.164451599121,-106.015678405762,-105.797950744629,-105.685592651367,-105.606056213379,-105.539840698242,-105.456932067871,-105.428611755371,-105.37744140625,-105.194976806641,-105.101318359375,-105.043601989746,-104.993797302246,-104.959815979004,-104.936714172363,-104.911964416504,-104.879440307617,-104.769630432129,-104.653175354004,-104.636375427246,-104.6611328125,-104.628173828125,-104.48681640625,-104.350738525391,-104.193550109863,-103.901557922363,-103.6572265625,-103.47412109375,-103.3232421875,-103.021781921387,-102.841552734375,-102.691986083984,-102.389114379883,-102.320358276367,-102.209762573242,-102.057228088379,-101.883644104004,-101.688865661621,-101.554985046387,-101.096389770508,-101.026412963867,-100.855621337891,-100.74560546875,-100.616111755371,-100.519630432129,-100.456108093262,-100.212936401367,-99.77294921875,-99.4722671508789,-99.2935562133789,-99.1468734741211,-99.0321731567383,-98.9204635620117,-98.8117218017578,-98.697265625,-98.4527816772461,-98.412109375,-98.4171371459961,-98.4678268432617,-98.6064910888672,-98.7035598754883,-98.7222137451172,-98.7200698852539,-98.6898422241211,-98.6315460205078,-98.5398406982422,-98.414794921875,-98.0625457763672,-97.9776382446289,-97.9307632446289,-97.607421875,-97.4549255371094,-97.2742691040039,-97.1944351196289,-97.1554183959961,-97.1571807861328,-97.1398468017578,-97.1580581665039,-97.20654296875,-97.3361282348633,-97.546630859375,-97.7391128540039,-97.913330078125,-98.1104965209961,-98.1925277709961,-98.4383773803711,-98.5002975463867,-98.500244140625,-98.3860778808594,-98.380859375,-98.4491729736328,-98.4912567138672,-98.6330032348633,-98.6504898071289,-98.562255859375,-98.522216796875,-98.4685592651367,-98.2185592651367,-98.0905227661133,-97.7942352294922,-97.9110336303711,-97.9388656616211,-97.9250946044922,-97.8285675048828,-97.6395492553711,-97.5480422973633,-97.4811019897461,-97.4103546142578,-97.3357925415039,-97.2659149169922,-97.135986328125,-97.07177734375,-96.9995574951172,-96.9767074584961,-96.628173828125,-96.4306640625,-96.4349594116211,-96.480224609375,-96.7251434326172,-96.7220687866211,-96.5921859741211,-96.5312957763672,-96.4937057495117,-96.461181640625,-96.4393539428711,-96.0755844116211,-95.9703140258789,-96.0360336303711,-96.1713409423828,-96.1988296508789,-96.2284622192383,-96.3713836669922,-96.369140625,-96.2128448486328,-96.1850128173828,-96.1692428588867,-96.1414566040039,-96.0125961303711,-95.8791046142578,-95.7199249267578,-95.6951675415039,-95.7825241088867,-95.7776870727539,-95.6264190673828,-95.5570297241211,-95.5287094116211,-95.4159164428711,-95.4045867919922,-95.406982421875,-95.4188995361328,-95.4569854736328,-95.5022430419922,-95.5593719482422,-95.6106491088867,-95.7686538696289,-95.86181640625,-95.9540557861328,-96.01953125,-96.095458984375,-96.215576171875,-96.3504409790039,-96.4042510986328,-96.4225616455078,-96.4202651977539,-96.3595199584961,-95.8852996826172,-95.8132781982422,-95.79736328125,-95.7875518798828,-95.7431640625,-95.7721176147461,-96.01611328125,-96.0453643798828,-96.036865234375,-95.9723587036133,-95.6250457763672,-95.4903793334961,-95.399658203125,-95.3540954589844,-95.3210983276367,-95.2587432861328,-95.2956085205078,-95.3895492553711,-95.46337890625,-95.6336898803711,-95.6504898071289,-95.460693359375,-95.426513671875,-95.3840866088867,-95.2347106933594,-95.1258850097656,-94.9552230834961,-94.8610382080078,-94.7444381713867,-94.4853057861328,-94.3838348388672,-94.2547836303711,-94.09814453125,-93.927734375,-93.6517105102539,-93.4830093383789,-93.4489288330078,-93.6058120727539,-93.6439437866211,-93.6761779785156,-93.6598663330078,-93.6627960205078,-93.6814498901367,-93.7157669067383,-93.7657241821289,-93.8113327026367,-93.8524398803711,-93.8807144165039,-93.8960952758789,-93.9380874633789,-93.9915542602539,-94.0648956298828,-94.2169418334961,-94.4783172607422,-94.5867691040039,-94.6004409790039,-94.5625457763672,-94.4756317138672,-94.2366180419922,-94.0836410522461,-94.0811462402344,-94.2218246459961,-94.25537109375,-94.2849655151367,-94.2767562866211,-94.2547378540039,-94.1563491821289,-93.8543930053711,-93.6194839477539,-93.6126480102539,-93.8009719848633,-93.8204650878906,-93.74853515625,-93.5674819946289,-93.4505844116211,-93.4309539794922,-93.5370635986328,-93.5428695678711,-93.5224151611328,-93.5322723388672,-93.6498031616211,-93.7943878173828,-93.9150848388672,-94.0152893066406,-94.1631774902344,-94.2708053588867,-94.338134765625,-94.419189453125,-94.513916015625,-94.6338348388672,-94.67626953125,-94.7126998901367,-94.7892532348633,-94.822509765625,-95.2920913696289,-95.4912643432617,-95.5875930786133,-95.7074279785156,-95.8506774902344,-95.9649429321289,-96.0501403808594,-96.1190948486328,-96.1717758178711,-96.2693862915039,-96.4923858642578,-96.5513687133789,-96.5595703125,-96.5456085205078,-96.3365707397461,-96.2977066040039,-96.2264175415039,-96.1227569580078,-96.0481414794922,-95.8786087036133,-95.9801712036133,-95.9881896972656,-95.8863296508789,-95.9063949584961,-96.1864242553711,-96.2580032348633,-96.35888671875,-96.5489273071289,-96.5510787963867,-96.4913024902344,-96.4704132080078,-96.5247573852539,-96.5044479370117,-96.4454574584961,-96.4207458496094,-96.4465789794922,-96.4056625366211,-96.2713394165039,-96.1396484375,-96.06201171875,-95.9944381713867,-95.924072265625,-95.8508758544922,-95.7253875732422,-95.632568359375,-95.5642547607422,-95.447509765625,-95.40625,-95.4454116821289,-95.6742172241211,-95.7733917236328,-95.8303680419922,-95.872314453125,-95.8377456665039,-95.6159210205078,-95.5116729736328,-95.2012176513672,-94.886962890625,-94.73486328125,-94.6111297607422,-94.5570755004883,-94.4910659790039,-94.4788131713867,-94.308349609375,-94.1712417602539,-94.0859832763672,-93.8102035522461,-93.7462844848633,-93.7508773803711,-93.7816390991211,-93.7628402709961,-93.5758819580078,-93.407470703125,-93.25634765625,-93.0313034057617,-92.9825668334961,-92.9486846923828,-92.8901824951172,-92.8827209472656,-92.9041976928711,-92.9219741821289,-92.9814453125,-92.9608917236328,-92.7830123901367,-92.6417007446289,-92.5674743652344,-92.3884811401367,-92.3558120727539,-92.3153839111328,-92.2144546508789,-92.0491180419922,-92.0372085571289,-92.0726089477539,-92.04736328125,-91.9835433959961,-91.9262161254883,-91.8755416870117,-91.8204116821289,-91.7608413696289,-91.7156219482422,-91.654052734375,-91.5640640258789,-91.5716323852539,-91.6161117553711,-91.8585891723633,-91.9949722290039,-92.1210403442383,-92.2086410522461,-92.2903289794922,-92.3205032348633,-92.36328125,-92.4546890258789,-92.5118637084961,-92.4457473754883,-92.1269989013672,-92.0577163696289,-91.9767074584961,-92.0690460205078,-92.2847595214844,-92.7509307861328,-92.8877944946289,-92.8545455932617,-92.8026351928711,-92.642822265625,-92.4934616088867,-92.3116760253906,-92.2307662963867,-92.25830078125,-92.2090835571289,-91.9119644165039,-91.72412109375,-91.5323791503906,-91.3842315673828,-91.2018051147461,-91.15087890625,-91.1703109741211,-91.3050231933594,-91.4268569946289,-91.43994140625,-91.2881317138672,-90.9501953125,-90.7857437133789,-90.6666488647461,-90.554931640625,-90.4505386352539,-90.4155807495117,-90.5132827758789,-90.6055679321289,-90.6839828491211,-90.74853515625,-90.7945785522461,-90.8221206665039,-90.8922882080078,-91.0049819946289,-91.0492248535156,-91.0248489379883,-91.0577697753906,-91.1478500366211,-91.2179718017578,-91.2372055053711,-90.7447738647461,-90.5873031616211,-90.47900390625,-90.4683609008789,-90.5389633178711,-90.5425338745117,-90.5101547241211,-90.5252456665039,-90.5736389160156,-90.5283203125,-90.4230422973633,-90.3600540161133,-90.3172378540039,-90.2852554321289,-90.2477569580078,-90.2047882080078,-90.1744079589844,-90.1568374633789,-90.1164093017578,-90.0053253173828,-89.897705078125,-89.8794937133789,-89.8965835571289,-89.8842315673828,-89.7831039428711,-89.7508316040039,-89.7201614379883,-89.6666030883789,-89.552001953125,-89.3519058227539,-89.279541015625,-89.198486328125,-89.0567398071289,-88.9535140991211,-88.8145446777344,-88.6377410888672,-88.3155288696289,-88.2235412597656,-88.0413589477539,-87.96435546875,-87.9115219116211,-87.865966796875,-87.8277359008789,-87.810302734375,-87.8135757446289,-87.8279266357422,-87.853271484375,-87.8926773071289,-87.990966796875,-88.1111373901367,-88.1458053588867,-88.2090759277344,-88.2352523803711,-88.3469772338867,-88.3606948852539,-88.3196258544922,-88.3250961303711,-88.3138198852539,-88.1958999633789,-87.9971618652344,-87.4991226196289,-87.4708023071289,-87.4179153442383,-87.3919448852539,-87.359375,-87.3202667236328,-87.2662582397461,-87.0832061767578,-86.9239273071289,-86.8127899169922,-86.7498474121094,-86.6820297241211,-86.609375,-86.560791015625,-86.5364303588867,-86.5035095214844,-86.4755325317383,-86.3980484008789,-86.3696746826172,-85.9844741821289,-85.9525909423828,-85.7888717651367,-85.7311096191406,-85.7228012084961,-85.744873046875,-85.7338409423828,-85.6897964477539,-85.6431579589844,-85.5624465942383,-85.5177764892578,-85.4910659790039,-85.425048828125,-85.3380813598633,-85.2750930786133,-84.8675765991211,-84.8674774169922,-85.1066436767578,-85.1043472290039,-85.0833969116211,-85.00830078125,-84.9160690307617,-84.8953094482422,-84.8927230834961,-84.8622055053711,-84.8900375366211,-85.113525390625,-85.2426223754883,-85.2754821777344,-85.3867721557617,-85.4275360107422,-85.4319305419922,-85.4164047241211,-85.4022445678711,-85.4091796875,-85.4372024536133,-85.4394989013672,-85.4159698486328,-85.4301300048828,-85.4820327758789,-85.50244140625,-85.4478988647461,-85.4460983276367,-85.4974594116211,-85.5348129272461,-85.5073699951172,-85.4151382446289,-85.3049774169922,-85.1768112182617,-85.0198211669922,-84.833984375,-84.6451187133789,-84.3188018798828,-84.2416458129883,-83.9171829223633,-83.6653289794922,-83.5517120361328,-82.9913558959961,-82.74560546875,-82.6183547973633,-82.3741683959961,-82.3902359008789,-82.4957046508789,-82.63330078125,-82.7548294067383,-82.6420440673828,-82.3098678588867,-82.2318344116211,-82.2081604003906,-82.2467727661133,-82.2275390625,-82.1505355834961,-81.9518051147461,-81.732177734375,-81.4123001098633,-81.3778305053711,-81.3215866088867,-81.3286590576172,-81.6114273071289,-81.7583465576172,-81.95166015625,-81.9579086303711,-81.6869125366211,-81.4760208129883,-81.3809127807617,-81.331298828125,-81.2635269165039,-81.2524871826172,-81.2591323852539,-81.2815475463867,-81.52685546875,-81.6395034790039,-81.8313980102539,-81.9148406982422,-82.0064926147461,-82.1063461303711,-82.2101593017578,-82.397216796875,-82.4987335205078,-82.5486373901367,-82.5526885986328,-82.4641571044922,-82.4129867553711,-82.3928756713867,-82.4306640625,-82.4227066040039,-82.3925323486328,-82.222412109375,-82.1865768432617,-82.1513137817383,-82.07763671875,-82.0338821411133,-82.0124969482422,-82.0133819580078,-82.0918960571289,-82.1021423339844,-82.1004867553711,-82.0625534057617,-81.9764633178711,-81.8693389892578,-81.7085952758789,-81.4927749633789,-81.4123001098633,-81.2943420410156,-81.2701187133789,-81.3010711669922,-81.38720703125,-81.4427261352539,-81.4675827026367,-81.6300735473633,-81.7223663330078,-81.8744583129883,-81.925537109375,-82.0050811767578,-82.1131820678711,-82.1983413696289,-82.2605438232422,-82.374755859375,-82.5536651611328,-82.6415023803711,-82.9488754272461,-83.1987838745117,-83.2983932495117,-83.4064483642578,-83.5231018066406,-83.5902786254883,-83.6283721923828,-83.6510772705078,-83.7392120361328,-83.9202117919922,-83.998046875,-84.0500030517578,-84.1544494628906,-84.2080078125,-84.3243713378906,-84.3662567138672,-84.361083984375,-84.2725601196289,-84.3103561401367,-84.466064453125,-84.5306701660156,-84.5384750366211,-84.6925735473633,-84.8457489013672,-85.0400390625,-85.1137161254883,-85.1112823486328,-85.0182571411133,-84.9779739379883,-84.8990249633789,-84.8573760986328,-84.7377471923828,-84.6385803222656,-84.6025390625,-84.5895004272461,-84.3189468383789,-84.2230453491211,-84.18310546875,-84.1527328491211,-84.09423828125,-83.9642105102539,-83.8258361816406,-83.7975540161133,-83.8690414428711,-83.9050750732422,-84.0116653442383,-84.2930679321289,-84.3242645263672,-84.3984375,-84.4593734741211,-84.4784164428711,-84.6280822753906,-84.9083938598633,-85.09619140625,-85.1914978027344,-85.3068389892578,-85.4422378540039,-85.6038589477539,-85.791748046875,-86.063232421875,-86.6332015991211,-86.7081527709961,-86.7371139526367,-86.6886215209961,-86.7001495361328,-86.7383804321289,-86.7469711303711,-86.6851043701172,-86.5847625732422,-86.301025390625,-86.1130828857422,-86.0008316040039,-85.9642562866211,-85.958740234375,-86.0122604370117,-86.0428695678711,-86.7019500732422,-86.9531707763672,-87.0811004638672,-87.1938018798828,-87.2914581298828,-87.452880859375,-87.6781311035156,-87.969970703125,-88.1209945678711,-88.3948745727539,-88.5867233276367,-88.6724624633789,-88.7439422607422,-88.8084945678711,-88.9461441040039,-89.0877456665039,-89.4203567504883,-89.5926742553711,-89.7494201660156,-89.8904800415039,-89.9439926147461,-89.8477478027344,-89.8896942138672,-90.0038070678711,-90.1166000366211,-90.3162612915039,-90.5132827758789,-90.6554718017578,-90.8257293701172,-91.009521484375,-91.3054733276367,-91.4115219116211,-91.42724609375,-91.28515625,-91.0411148071289,-91.0736312866211,-91.06494140625,-90.9834442138672,-90.5968246459961,-90.1586380004883,-90.0475616455078,-89.924072265625,-89.7879867553711,-89.6003875732422,-89.2417526245117,-89.1265640258789,-88.9740295410156,-88.1977996826172,-87.9295425415039,-87.3919448852539,-87.1080017089844,-87.0275421142578,-87.002685546875,-87.0285186767578,-87.1829147338867,-87.280517578125,-87.885009765625,-87.9635772705078,-87.99755859375,-88.1056137084961,-88.3789520263672,-88.6530303955078,-88.8177261352539,-88.9644012451172,-89.0596237182617,-89.2006301879883,-89.2094192504883,-89.107666015625,-89.1315460205078,-89.2145538330078,-89.4035186767578,-89.4647445678711,-89.5009307861328,-89.5513229370117,-89.6158218383789,-89.7327194213867,-89.7638244628906,-89.7920913696289,-89.8113250732422,-90.0416564941406,-90.0800323486328,-89.985595703125,-89.9535675048828,-89.860595703125,-89.855712890625,-89.921875,-90.1418914794922,-90.1681594848633,-90.0596160888672,-90.0179672241211,-90.013427734375,-90.1547317504883,-90.2453155517578,-90.3688430786133,-90.4462356567383,-90.5334930419922,-90.5963897705078,-90.635009765625,-90.7068405151367,-90.8119125366211,-90.9456558227539,-91.1080551147461,-91.538818359375,-91.6746520996094,-91.926025390625,-91.9560012817383,-91.9538040161133,-91.91943359375,-91.9289093017578,-91.9822235107422,-92.03759765625,-92.0948791503906,-92.1952133178711,-92.3384246826172,-92.5500946044922,-92.97021484375,-93.4296875,-93.6963348388672,-93.5967254638672,-93.6048812866211,-93.6558151245117,-93.6641616821289,-93.559814453125,-93.4155731201172,-93.2702178955078,-93.2662124633789,-93.3268585205078,-93.3802795410156,-93.4058609008789,-93.3785171508789,-93.2504425048828,-93.1659164428711,-92.5292510986328,-92.3392105102539,-92.1964797973633,-92.1568908691406,-92.2050247192383,-92.4610366821289,-92.465087890625,-92.28955078125,-92.0766143798828,-91.9568405151367,-91.8418426513672,-91.68603515625,-91.4893035888672,-91.330078125,-91.1030807495117,-90.9700698852539,-90.74658203125,-90.7112808227539,-90.6907272338867,-90.6985855102539,-90.7276382446289,-90.7778778076172,-90.8711929321289,-91.0077133178711,-91.1148910522461,-91.3494644165039,-91.448974609375,-91.86962890625,-92.0342330932617,-92.1100616455078,-92.152099609375,-92.1961441040039,-92.3612823486328,-92.3881378173828,-92.3777313232422,-92.3452606201172,-92.30517578125,-92.2431640625,-92.1491241455078,-91.9558639526367,-91.9358444213867,-91.9444351196289,-92.0078582763672,-92.0811538696289,-92.2072296142578,-92.26953125,-92.3240737915039,-92.4000015258789,-92.4973678588867,-92.5514144897461,-92.5622024536133,-92.594970703125,-92.7074203491211,-92.7679672241211,-92.7659683227539,-92.7014617919922,-92.62744140625,-92.5440444946289,-92.5279769897461,-92.5792007446289,-92.6480941772461,-92.7347640991211,-92.8658218383789,-93.1544418334961,-93.2053680419922,-93.1792526245117,-92.98779296875,-92.9144515991211,-92.905517578125,-93.0658721923828,-93.0702590942383,-93.0277328491211,-93.0162582397461,-93.0733871459961,-93.16748046875,-93.3497619628906,-93.3663558959961,-93.296875,-93.2734375,-93.3330612182617,-93.3720169067383,-93.581787109375,-93.5267105102539,-93.4942398071289,-93.429931640625,-93.3144073486328,-93.31201171875,-93.3523406982422,-93.4206008911133,-93.7096710205078,-93.9127426147461,-93.9408721923828,-93.8892593383789,-93.8888168334961,-93.9419937133789,-94.0607376098633,-94.0834426879883,-94.0552215576172,-94.0499496459961,-94.0677795410156,-94.154052734375,-94.3086929321289,-94.4271926879883,-94.5093765258789,-94.5688934326172,-94.6787567138672,-94.76171875,-94.7052688598633,-94.670654296875,-94.6467819213867,-94.67041015625,-94.7416000366211,-94.7857894897461,-94.7766647338867,-94.7882766723633,-94.8195343017578,-94.8702621459961,-94.9573287963867,-94.8465805053711,-94.7761688232422,-94.7437438964844,-94.71337890625,-94.6733856201172,-94.6237335205078,-94.5792007446289,-94.5396957397461,-94.4192352294922,-94.2870559692383,-94.2808074951172,-94.3326187133789,-94.3322296142578,-94.2721710205078,-94.2089385986328,-94.1231994628906,-94.0557632446289,-93.780029296875,-93.4861755371094,-93.3750534057617,-93.2781219482422,-93.1787567138672,-93.1545867919922,-93.1265182495117,-93.1001968383789,-92.9251480102539,-92.8417510986328,-92.7398452758789,-92.70166015625,-92.4896469116211,-92.4494171142578,-92.4328155517578,-92.4397964477539,-92.4783706665039,-92.5484848022461,-92.6141128540039,-92.6752471923828,-92.7379837036133,-92.8023986816406,-92.7981491088867,-92.7252960205078,-92.6509780883789,-92.5102996826172,-92.456298828125,-92.3033676147461,-92.2982940673828,-92.3721237182617,-92.355712890625,-92.2490234375,-92.0180130004883,-91.1112823486328,-90.8974609375,-90.5921859741211,-90.3448257446289,-90.0751953125,-89.7908248901367,-89.3423385620117,-89.2115707397461,-88.948486328125,-88.8264694213867,-88.6798858642578,-88.4470672607422,-88.2713394165039,-88.0750961303711,-87.8781204223633,-87.5608901977539,-87.482421875,-87.286865234375,-86.9193801879883,-86.376953125,-86.138671875,-85.9844741821289,-85.8305206298828,-85.6766586303711,-85.559326171875,-85.4784622192383,-85.4072723388672,-85.28271484375,-85.218017578125,-85.2120132446289,-85.3620071411133,-85.3652801513672,-85.2135772705078,-85.1288528442383,-85.0609359741211,-84.919921875,-84.7057571411133,-84.5179672241211,-84.3564987182617,-84.2189407348633,-84.1053695678711,-84.0230026245117,-83.9717788696289,-83.9105987548828,-83.6676712036133,-83.5694808959961,-83.21435546875,-82.9862823486328,-82.947021484375,-82.8677749633789,-82.8006820678711,-82.6875,-82.5774459838867,-82.3932647705078,-82.3082504272461,-82.2266082763672,-82.2193832397461,-82.37060546875,-82.4180679321289,-82.4241714477539,-82.3941421508789,-82.2635803222656,-82.2398910522461,-82.1626510620117,-82.1414566040039,-82.1500015258789,-82.1906204223633,-82.1803741455078,-82.1461944580078,-82.1591796875,-82.21923828125,-82.2599105834961,-82.2916030883789,-82.2915496826172,-82.2604522705078,-82.2026824951172,-82.1079559326172,-82.02001953125,-81.8592758178711,-81.7423324584961,-81.5994186401367,-81.5716781616211,-81.6115264892578,-81.6612319946289,-81.7763671875,-81.827880859375,-81.8145523071289,-81.6480026245117,-81.5495147705078,-81.4662170410156,-81.3980941772461,-81.2850570678711,-81.127197265625,-80.968505859375,-80.7055130004883,-80.657958984375,-80.5880355834961,-80.495849609375,-80.4476089477539,-80.4433135986328,-80.4955062866211,-80.6727066040039,-80.8512268066406,-80.7949676513672,-80.67724609375,-80.4783248901367,-80.3679656982422,-80.2656707763672,-80.1035614013672,-79.9604034423828,-79.8362274169922,-79.6515121459961,-79.4561538696289,-79.347900390625,-79.3807144165039,-79.45263671875,-79.6361770629883,-79.7144546508789,-79.7313919067383,-79.7374496459961,-79.7232437133789,-79.6888198852539,-79.6429672241211,-79.5857391357422,-79.5474624633789,-79.5280227661133,-79.49755859375,-79.3386688232422,-79.2969741821289,-79.2642593383789,-79.2261199951172,-79.1527328491211,-79.0908737182617,-79.04052734375,-79.0050277709961,-78.9843292236328,-78.9369125366211,-78.9031753540039,-78.8975067138672,-78.8580093383789,-78.8274383544922,-78.7313461303711,-78.7364273071289,-78.7763214111328,-78.9777374267578,-78.9816436767578,-78.9278869628906,-78.89111328125,-78.8712387084961,-78.8282241821289,-78.7020034790039,-78.5933151245117,-78.537353515625,-78.4916534423828,-78.4480972290039,-78.5130844116211,-78.5260772705078,-78.5291061401367,-78.5570755004883,-78.6005859375,-78.7237777709961,-78.744140625,-78.7657699584961,-78.7536087036133,-78.7216796875,-78.7398376464844,-78.8541030883789,-78.8982391357422,-78.9471206665039,-78.9920425415039,-79.0431137084961,-79.100341796875,-79.1131286621094,-79.0814437866211,-79.0403366088867,-79.003173828125,-78.9457015991211,-78.9443817138672,-79.0320281982422,-79.0751495361328,-79.0732879638672,-78.9960403442383,-79.0099105834961,-79.067138671875,-79.2417984008789,-79.1788101196289,-79.1388168334961,-79.146728515625,-79.2159652709961,-79.295654296875,-79.3561477661133,-79.4305648803711,-79.4759750366211,-79.5206527709961,-79.5979537963867,-79.6317443847656,-79.67041015625,-79.7139663696289,-79.7123489379883,-79.66552734375,-78.9092330932617,-78.8462448120117,-78.4750518798828,-78.3036117553711,-78.1288604736328,-77.89111328125,-77.7752990722656,-77.7021484375,-77.324951171875,-77.1650848388672,-77.0725555419922,-76.9380874633789,-76.7618179321289,-76.6504898071289,-76.6040573120117,-76.54638671875,-76.5298309326172,-76.5196304321289,-76.5255813598633,-76.5728530883789,-76.6014175415039,-76.6554183959961,-76.7862777709961,-76.809814453125,-76.8909149169922,-77.1567916870117,-77.4891586303711,-77.5524368286133,-77.68408203125,-77.8841247558594,-78.0135726928711,-78.3517074584961,-78.4629898071289,-78.5059051513672,-78.5150909423828,-78.5022964477539,-78.4826202392578,-78.4586868286133,-78.4305191040039,-78.2444305419922,-78.1402359008789,-78.0676727294922,-77.9876480102539,-77.8428192138672,-77.7606964111328,-77.7794418334961,-77.8446731567383,-77.8590316772461,-77.7490234375,-77.7334976196289,-77.7475128173828,-77.7261734008789,-77.5904312133789,-77.3966751098633,-77.3490676879883,-77.4110336303711,-77.4852981567383,-77.4745635986328,-77.3316421508789,-77.32763671875,-77.368408203125,-77.3729476928711,-77.2892150878906,-77.3118209838867,-77.5471649169922,-77.5858840942383,-77.5721664428711,-77.4613800048828,-77.452880859375,-77.6486282348633,-77.6814422607422,-77.59814453125,-77.5035629272461,-77.5155792236328,-77.6394500732422,-77.7149887084961,-77.7908172607422,-77.76123046875,-77.7342224121094,-77.66064453125,-77.5895462036133,-77.60302734375,-77.8715362548828,-77.9981460571289,-78.1224594116211,-78.1813507080078,-78.15966796875,-77.9341812133789,-77.8301315307617,-77.7650375366211,-77.7306137084961,-77.726806640625,-77.7496109008789,-77.7361755371094,-77.6488723754883,-77.5143585205078,-77.6984329223633,-77.8137664794922,-77.889892578125,-77.9475631713867,-78.0213851928711,-78.0774917602539,-78.1371612548828,-78.14697265625,-78.1333923339844,-78.1085968017578,-78.068115234375,-77.89990234375,-77.6039505004883,-77.3724136352539,-77.2052764892578,-76.87939453125,-76.6163558959961,-75.81689453125,-75.675537109375,-75.8092269897461,-75.7898406982422,-75.4888229370117,-75.4092330932617,-75.3412094116211,-75.114013671875,-75.0227584838867,-74.9075622558594,-74.632568359375,-74.6128921508789,-74.6898956298828,-74.6457977294922,-74.42919921875,-74.2054672241211,-74.0464859008789,-73.8778305053711,-73.7639694213867,-73.705078125,-73.6299819946289,-73.4283752441406,-73.2989730834961,-73.1951675415039,-73.0493621826172,-72.9923782348633,-72.8818359375,-72.7349548339844,-72.6869583129883,-72.6707992553711,-72.64599609375,-72.6331024169922,-72.6321258544922,-72.666015625,-72.7716293334961,-72.7273864746094,-72.66064453125,-72.5738754272461,-72.5055694580078,-72.3607482910156,-72.2261199951172,-72.178466796875,-72.1261215209961,-72.0814437866211,-72.0400390625,-72.04296875,-72.08203125,-72.2470703125,-72.2158737182617,-72.0230941772461,-71.9644012451172,-71.9222640991211,-71.8661193847656,-71.6382827758789,-71.6047897338867,-71.6194305419922,-71.6562042236328,-71.7557601928711,-71.8410186767578,-71.8543930053711,-71.7936553955078,-71.6453094482422,-71.6464385986328,-71.7321319580078,-71.7434616088867,-71.551513671875,-71.4227066040039,-71.3484344482422,-71.1751480102539,-71.0349578857422,-70.7232437133789,-70.540771484375,-70.3836441040039,-70.279296875,-70.1872100830078,-70.157958984375,-70.1441345214844,-70.1458969116211,-70.0953140258789,-69.992431640625,-69.9092254638672,-69.8004455566406,-69.7083969116211,-69.6775894165039,-69.6504364013672,-69.6236343383789,-69.5569763183594,-69.5033187866211,-69.471923828125,-69.4143600463867,-69.3983383178711,-69.4047317504883,-69.4334487915039,-69.4899444580078,-69.57421875,-69.6404800415039,-69.7213897705078,-69.7512741088867,-69.7594757080078,-69.7559127807617,-69.7405776977539,-69.70849609375,-69.6331024169922,-69.6287612915039,-69.6231460571289,-69.6297836303711,-69.6737289428711,-69.795654296875,-69.9628448486328,-70.5093231201172,-70.6548385620117,-70.6197280883789,-70.4666519165039,-70.3268585205078,-69.8056640625,-69.7339401245117,-69.6734313964844,-69.6302261352539,-69.58740234375,-69.5793991088867,-69.6023483276367,-69.6562042236328,-69.6923828125,-69.7108917236328,-69.681884765625,-69.4000015258789,-69.3440399169922,-69.3504409790039,-69.4504928588867,-69.4597625732422,-69.4141159057617,-69.4203109741211,-69.4480895996094,-69.4746551513672,-69.5000991821289,-69.5316390991211,-69.6082077026367,-69.6483917236328,-69.6773452758789,-69.7530288696289,-69.7841796875,-69.8136749267578,-69.8285140991211,-69.82861328125,-69.8415985107422,-69.8675765991211,-69.9791488647461,-70.1599655151367,-70.1543502807617,-70.0330123901367,-69.8785629272461,-69.7898864746094,-69.6505432128906,-69.3818359375,-69.2710952758789,-69.1734848022461,-69.0636215209961,-68.9415054321289,-68.6981964111328,-68.6372985839844,-68.5628967285156,-68.4748992919922,-68.414306640625,-68.3811492919922,-68.3264694213867,-68.2529830932617,-68.2351531982422,-68.2293930053711,-68.23388671875,-68.314697265625,-68.3565368652344,-68.4681625366211,-68.5968704223633,-68.8257827758789,-68.9453125,-69.0354461669922,-69.0408248901367,-68.7809524536133,-68.4950714111328,-68.41357421875,-68.3518524169922,-68.2891159057617,-68.175537109375,-68.1110382080078,-68.0210494995117,-67.9811553955078,-67.8877944946289,-67.8882827758789,-67.9114227294922,-68.0638656616211,-68.0089874267578,-67.8558578491211,-67.8233413696289,-67.8052291870117,-67.7559509277344,-67.737060546875,-67.689697265625,-67.6881866455078,-67.6805648803711,-67.6976623535156,-67.6782684326172,-67.6322250366211,-67.6171875,-67.5963363647461,-67.5696258544922,-67.3819808959961,-67.162841796875,-67.0194320678711,-66.900390625,-66.72216796875,-66.60791015625,-66.5577163696289,-66.5150375366211,-66.47998046875,-66.3624038696289,-66.2985382080078,-66.2374038696289,-66.168212890625,-66.0909194946289,-66.0446319580078,-66.029541015625,-66.0170440673828,-66.0023956298828,-65.9312515258789,-65.9229049682617,-65.9279327392578,-65.9496536254883,-66.0212936401367,-66.0493621826172,-66.0430679321289,-65.967041015625,-65.8548355102539,-65.8359375,-65.9184112548828,-65.9207000732422,-65.8414077758789,-65.7948226928711,-65.7035675048828,-65.721923828125,-65.7209930419922,-65.6952667236328,-65.5439910888672,-65.5263214111328,-65.396240234375,-65.383544921875,-65.4959945678711,-65.6062469482422,-65.6398468017578,-65.6656265258789,-65.6999969482422,-65.6917037963867,-65.6607360839844,-65.6071319580078,-65.5780258178711,-65.5453109741211,-65.5127944946289,-65.4117202758789,-65.4072799682617,-65.4893569946289,-65.47509765625,-65.3497085571289,-65.2737808227539,-65.0743103027344,-65.0382385253906,-65.06884765625,-65.1709442138672,-65.26318359375,-65.3455047607422,-65.4074249267578,-65.4751968383789,-65.4864730834961,-65.4808578491211,-65.4333953857422,-65.4061508178711,-65.35791015625,-65.2887649536133,-65.2122573852539,-65.0544891357422,-65.0281753540039,-65.11328125,-65.1592254638672,-65.181396484375,-65.1717224121094,-65.1048812866211,-65.0733947753906,-64.9312515258789,-64.8895492553711,-64.8450164794922,-64.8173370361328,-64.7058563232422,-64.4994125366211,-64.4363250732422,-64.4195785522461,-64.5277328491211,-64.7132797241211,-64.7684555053711,-64.7325744628906,-64.5591812133789,-64.40771484375,-64.2834930419922,-64.182861328125,-64.1688003540039,-64.226318359375,-64.1506881713867,-64.0560531616211,-63.9787101745605,-63.969482421875,-63.9288101196289,-63.8412590026855,-63.7501907348633,-63.8503913879395,-63.970703125,-63.9454612731934,-63.7808609008789,-63.7585945129395,-63.77587890625,-63.7520027160645,-63.6375007629395,-63.5398902893066,-63.4151382446289,-63.5062026977539,-63.6455039978027,-63.7563934326172,-63.9104957580566,-63.9711418151855,-63.9410133361816,-63.7936515808105,-63.5678749084473,-63.3989753723145,-63.3255386352539,-63.2484397888184,-63.2225112915039,-63.3037109375,-63.3098640441895,-63.2794418334961,-63.2164077758789,-63.221923828125,-63.2821273803711,-63.1853485107422,-63.0502967834473,-63.0083503723145,-62.9260749816895,-62.8738784790039,-63.1023406982422,-63.2186546325684,-63.3899421691895,-63.4379348754883,-63.5370635986328,-63.4736328125,-63.2964820861816,-63.2099571228027,-63.1455078125,-63.1195335388184,-63.1321296691895,-63.07568359375,-62.8373985290527,-62.7373008728027,-62.6078643798828,-62.5938491821289,-62.6743125915527,-62.8120574951172,-63.0627937316895,-63.1516609191895,-63.2615280151367,-63.2200202941895,-62.9809074401855,-62.8175315856934,-62.5880851745605,-62.4862327575684,-62.3056602478027,-62.2015151977539,-62.1174278259277,-61.9586410522461,-61.8990745544434,-61.9140586853027,-61.9677734375,-61.9949226379395,-61.9312515258789,-61.9679718017578,-62.0839805603027,-62.1668930053711,-62.2536163330078,-62.3385772705078,-62.3771476745605,-62.4955558776855,-62.4549827575684,-62.396484375,-62.30322265625,-62.1942405700684,-62.0880851745605,-61.921142578125,-61.8510704040527,-61.8498001098633,-61.8858375549316,-61.9388656616211,-61.9774436950684,-61.9445304870605,-61.86083984375,-61.7983436584473,-61.7163124084473,-61.6285171508789,-61.333740234375,-61.3457527160645,-61.3904762268066,-61.372802734375,-61.3716316223145,-61.5316925048828,-62.0625,-62.3661155700684,-62.3817367553711,-62.2957992553711,-62.3720245361328,-62.4602088928223,-62.4972686767578,-62.3955078125,-62.1165046691895,-61.9916038513184,-61.8549308776855,-61.8133773803711,-61.7377433776855,-61.7600593566895,-61.8994140625,-62.0096702575684,-61.9404296875,-61.6924819946289,-61.5146026611328,-61.42529296875,-61.4986801147461,-61.7071266174316,-61.7130889892578,-61.5585975646973,-61.4210891723633,-61.3646926879883,-61.3244171142578,-61.3011245727539,-61.4489250183105,-61.4495124816895,-61.3512725830078,-61.1878890991211,-61.1338882446289,-61.1230010986328,-61.08935546875,-60.9957542419434,-60.8926734924316,-60.8318367004395,-60.7432632446289,-60.7366256713867,-60.6309547424316,-60.5925750732422,-60.5621109008789,-60.4758262634277,-60.4126014709473,-60.3410110473633,-60.3654289245605,-60.4083023071289,-60.3519554138184,-60.3083038330078,-60.1923789978027,-60.2240257263184,-60.3609352111816,-60.43310546875,-60.4500961303711,-60.5205116271973,-60.6171379089355,-60.5565452575684,-60.3407745361328,-60.2125511169434,-59.9303207397461,-59.8621063232422,-59.7587852478027,-59.6955032348633,-59.6890640258789,-59.60546875,-59.5176734924316,-59.4378890991211,-59.48583984375,-59.74169921875,-59.81640625,-59.8377914428711,-59.7499008178711,-59.4285659790039,-59.3941917419434,-59.3241691589355,-59.2595672607422,-59.0863265991211,-58.9971199035645,-58.955810546875,-58.8857917785645,-58.7801780700684,-58.4999008178711,-58.3981437683105,-58.2228546142578,-58.1952590942383,-58.0584945678711,-57.9624519348145,-57.9292984008789,-57.8268547058105,-57.7248992919922,-57.6266136169434,-57.4830093383789,-57.4044876098633,-57.4044418334961,-57.4853515625,-57.5632286071777,-57.6992645263672,-57.8891105651855,-58.1513710021973,-58.1619148254395,-58.2197265625,-58.359130859375,-58.4352035522461,-58.5584030151367,-58.6332054138184,-58.7194366455078,-58.8408241271973,-58.9202156066895,-58.9784660339355,-59.0126419067383,-59.038818359375,-59.201416015625,-59.4965362548828,-59.6526870727539,-59.7494621276855,-59.8230438232422,-59.8733444213867,-60.01416015625,-60.0565414428711,-60.0813484191895,-60.1004867553711,-60.1172828674316,-60.1449241638184,-60.2633323669434,-60.3954086303711,-60.3695297241211,-60.1602516174316,-60.1002960205078,-60.1571311950684,-60.290283203125,-60.3057632446289,-60.2511749267578,-60.272705078125,-60.345703125,-60.3375015258789,-60.3294906616211,-60.1483383178711,-59.9871063232422,-59.8817367553711,-59.8290557861328,-59.6210975646973,-59.4981460571289,-59.3222618103027,-59.1293907165527,-58.9195823669434,-58.6520500183105,-58.3267059326172,-58.0880889892578,-57.9359855651855,-57.9282684326172,-58.0648460388184,-58.1774406433105,-58.3174819946289,-58.3607902526855,-58.3561515808105,-58.3099594116211,-58.1920928955078,-57.6149406433105,-57.4160652160645,-57.1988754272461,-57.1489753723145,-57.1349639892578,-57.1569328308105,-57.2439918518066,-57.4894561767578,-57.5240707397461,-57.52734375,-57.4202117919922,-57.3861274719238,-57.3317413330078,-57.2213859558105,-57.0121574401855,-56.8408699035645,-56.6965789794922,-56.5243148803711,-56.4649887084961,-56.4443359375,-56.35400390625,-56.2702178955078,-56.1101570129395,-55.9661140441895,-55.9112319946289,-55.859375,-55.8633804321289,-55.8547821044922,-55.81689453125,-55.7979507446289,-55.8082008361816,-55.892333984375,-55.8298835754395,-55.85791015625,-55.8725128173828,-55.8186531066895,-55.8028335571289,-55.8484382629395,-56.1669921875,-56.2923851013184,-56.3249015808105,-56.2284202575684,-56.0525856018066,-55.8401832580566,-55.7464828491211,-55.7059555053711,-55.7162094116211,-55.7771492004395,-55.8966789245605,-56.01171875,-56.004638671875,-55.8336448669434,-55.7834968566895,-55.6910629272461,-55.6728019714355,-55.6952133178711,-56.0174789428711,-56.2825698852539,-56.548583984375,-56.9759750366211,-57.0182609558105,-57.0956039428711,-57.2992210388184,-57.461669921875,-57.7695846557617,-57.853759765625,-58.0226554870605,-58.0894050598145,-58.2704620361328,-58.4422874450684,-58.5103492736816,-58.59326171875,-58.6147422790527,-58.6375961303711,-59.0549278259277,-59.1653785705566,-59.3780250549316,-59.6119155883789,-59.8153343200684,-59.8863258361816,-60.0801734924316,-60.4380836486816,-60.6082038879395,-60.8072280883789,-60.9562950134277,-61.1807174682617,-61.2897453308105,-61.7248573303223,-61.8353500366211,-61.9195289611816,-62.1652336120605,-62.3616676330566,-62.5409202575684,-62.7154312133789,-62.8302268981934,-62.9497566223145,-63.1356468200684,-63.2386245727539,-63.586669921875,-63.7335929870605,-63.8539581298828,-64.0158157348633,-64.17041015625,-64.5089340209961,-64.8678741455078,-65.180908203125,-65.2686080932617,-65.762451171875,-65.9553756713867,-66.1255340576172,-66.2421875,-66.3688430786133,-66.4110870361328,-66.4955062866211,-66.550048828125,-66.6217269897461,-66.7408676147461,-66.941162109375,-67.234375,-67.2619171142578,-67.3720245361328,-67.46923828125,-67.5492630004883,-68.0562515258789,-68.2819366455078,-68.2206039428711,-68.2945251464844,-68.4144058227539,-68.5438461303711,-68.6278839111328,-68.6690444946289,-68.9290542602539,-69.2307662963867,-69.3749465942383,-69.5500946044922,-69.6738815307617,-69.7619094848633,-69.8517074584961,-70.0010223388672,-70.1106414794922,-70.3836898803711,-71.0182647705078,-70.922607421875,-70.8387680053711,-70.6710968017578,-70.5006332397461,-70.1453094482422,-69.97119140625,-69.8655242919922,-69.7750015258789,-69.83984375,-69.9055633544922,-69.9944305419922,-70.3000946044922,-70.4480438232422,-70.7058563232422,-70.9727096557617,-71.1156234741211,-71.2677688598633,-71.624755859375,-71.7572784423828,-71.8795928955078,-72.0284652709961,-72.2046432495117,-72.2566375732422,-72.6801223754883,-72.8426742553711,-72.9810104370117,-73.0219192504883,-73.1454086303711,-73.1796875,-73.2835464477539,-73.4766159057617,-73.7118682861328,-73.7978515625,-73.8974151611328,-74.037841796875,-74.3150863647461,-74.2476501464844,-73.9996109008789,-73.9738311767578,-74.0980911254883,-74.3582992553711,-74.7088928222656],[-96.7823257446289,-96.9438018798828,-97.0927734375,-97.09765625,-97.0876998901367,-97.0692443847656,-97.0149917602539,-96.8624038696289,-96.7931671142578,-96.7677688598633,-96.7444305419922,-96.64599609375,-96.5984878540039,-96.603515625,-96.6353988647461,-96.6706008911133,-96.709228515625,-96.7823257446289],[-114.521537780762,-114.458106994629,-114.342437744141,-114.174461364746,-113.957809448242,-113.692436218262,-113.622169494629,-113.578071594238,-113.500045776367,-113.486137390137,-113.495903015137,-113.491409301758,-113.449851989746,-113.292381286621,-113.2080078125,-113.073532104492,-112.753616333008,-112.453758239746,-112.048095703125,-111.455421447754,-111.269721984863,-111.250389099121,-111.355514526367,-111.610893249512,-111.816017150879,-111.895164489746,-111.761619567871,-111.675094604492,-111.543556213379,-111.447364807129,-111.311180114746,-111.264793395996,-111.25341796875,-111.277145385742,-111.287254333496,-111.283737182617,-111.26806640625,-111.184616088867,-111.139892578125,-110.958984375,-110.781539916992,-110.512550354004,-110.439308166504,-110.205131530762,-110.207954406738,-110.197166442871,-110.279098510742,-110.553611755371,-110.689407348633,-110.660842895508,-110.50927734375,-110.094627380371,-110.008445739746,-109.609962463379,-109.469093322754,-109.357124328613,-109.121925354004,-109.043022155762,-108.987403869629,-108.968162536621,-108.985404968262,-108.994430541992,-108.950782775879,-108.7978515625,-108.754974365234,-108.698295593262,-108.627731323242,-108.566352844238,-108.469581604004,-108.276412963867,-108.210403442383,-108.188232421875,-108.144683837891,-108.020797729492,-107.925338745117,-107.812843322754,-107.785446166992,-107.757469177246,-107.687255859375,-107.346923828125,-107.329299926758,-107.369430541992,-107.381790161133,-107.376853942871,-107.306007385254,-107.542625427246,-107.695846557617,-107.794044494629,-107.809028625488,-107.82373046875,-107.855613708496,-107.909820556641,-107.932518005371,-107.923683166504,-107.934371948242,-107.997169494629,-108.238235473633,-108.237396240234,-108.204147338867,-108.118309020996,-107.979927062988,-107.936180114746,-107.987060546875,-108.077491760254,-108.089401245117,-108.029052734375,-107.720016479492,-107.496292114258,-107.113479614258,-107.074417114258,-107.032524108887,-106.950782775879,-106.828369140625,-106.482131958008,-106.081642150879,-105.812698364258,-105.624168395996,-105.495948791504,-105.415138244629,-105.411666870117,-105.430076599121,-105.411079406738,-105.354537963867,-105.323188781738,-105.297554016113,-105.246925354004,-105.234085083008,-104.878318786621,-104.810302734375,-104.766990661621,-104.518310546875,-104.385940551758,-104.373146057129,-104.355369567871,-104.349563598633,-104.355812072754,-104.384864807129,-104.436813354492,-104.487060546875,-104.563087463379,-104.569580078125,-104.514793395996,-104.166847229004,-103.953460693359,-103.853469848633,-103.584571838379,-103.294677734375,-103.197174072266,-103.10498046875,-103.077201843262,-103.021194458008,-103.005172729492,-103.001220703125,-103.082809448242,-103.088569641113,-103.049560546875,-102.75048828125,-102.589164733887,-102.368751525879,-101.989845275879,-101.937202453613,-101.732223510742,-101.676322937012,-101.641166687012,-101.626808166504,-101.618453979492,-101.562400817871,-101.239158630371,-101.148536682129,-101.090774536133,-101.042678833008,-100.973335266113,-100.909088134766,-100.905708312988,-100.935104370117,-100.982376098633,-101.043701171875,-101.216209411621,-101.337257385254,-101.400100708008,-101.456733703613,-101.483840942383,-101.508399963379,-101.565093994141,-101.602493286133,-101.647651672363,-101.733589172363,-101.860260009766,-102.097946166992,-102.18212890625,-102.234329223633,-102.348098754883,-102.523490905762,-102.595893859863,-102.565238952637,-102.544921875,-102.534858703613,-102.540916442871,-102.563133239746,-102.62109375,-102.743606567383,-102.919776916504,-103.059181213379,-103.30322265625,-103.359275817871,-103.434768676758,-103.464897155762,-103.418022155762,-103.294044494629,-103.142433166504,-103.101852416992,-103.062744140625,-103.048927307129,-103.031829833984,-103.039794921875,-103.112693786621,-103.120216369629,-103.09033203125,-102.884078979492,-102.777442932129,-102.546485900879,-102.44677734375,-102.151412963867,-102.045951843262,-101.978218078613,-101.975540161133,-102.052879333496,-102.06689453125,-102.070899963379,-102.064018249512,-102.046089172363,-101.992965698242,-101.899124145508,-101.872848510742,-101.822509765625,-101.789260864258,-101.787796020508,-101.857131958008,-101.980567932129,-102.358787536621,-102.488426208496,-102.738334655762,-102.834861755371,-102.895072937012,-103.162254333496,-103.468215942383,-103.820365905762,-104.067337036133,-104.352691650391,-104.460151672363,-104.571434020996,-105.105857849121,-105.169288635254,-105.148338317871,-105.021629333496,-105.013572692871,-105.019584655762,-105.262351989746,-105.533004760742,-105.804977416992,-106.008399963379,-106.140869140625,-106.270172119141,-106.341163635254,-106.353958129883,-106.355712890625,-106.34423828125,-106.361381530762,-106.419967651367,-106.539794921875,-106.659080505371,-106.759963989258,-106.855812072754,-107.033447265625,-107.122505187988,-107.353370666504,-107.439895629883,-107.863380432129,-108.364990234375,-108.552536010742,-108.730415344238,-108.945892333984,-109.472122192383,-109.958549499512,-110.467628479004,-110.848098754883,-110.957229614258,-111.127586364746,-111.310935974121,-111.51806640625,-112.304931640625,-112.666213989258,-112.8642578125,-113.01953125,-113.127731323242,-113.231391906738,-113.338081359863,-113.554832458496,-113.616844177246,-113.592529296875,-113.608543395996,-113.6806640625,-113.694137573242,-114.073432922363,-114.322944641113,-114.699066162109,-115.159034729004,-115.618110656738,-115.860748291016,-116.1015625,-116.513481140137,-116.536811828613,-116.568801879883,-116.609466552734,-116.712005615234,-116.992782592773,-117.10400390625,-117.121963500977,-117.148628234863,-117.184036254883,-117.195404052734,-117.162742614746,-117.135452270508,-116.553802490234,-115.529098510742,-114.592338562012,-114.1669921875,-113.916603088379,-113.66552734375,-113.210739135742,-112.637893676758,-112.522758483887,-112.265968322754,-112.189651489258,-111.78369140625,-111.704879760742,-111.632568359375,-111.725830078125,-112.114158630371,-113.1455078125,-113.397018432617,-113.757270812988,-113.966064453125,-114.232177734375,-114.331398010254,-114.592620849609,-114.840721130371,-115.311225891113,-115.990913391113,-116.086090087891,-116.225875854492,-116.327301025391,-116.992530822754,-117.587059020996,-118.264053344727,-118.376510620117,-118.352531433105,-118.269096374512,-117.93384552002,-117.814056396484,-117.31396484375,-116.815277099609,-116.421531677246,-116.228225708008,-116.042083740234,-115.891647338867,-115.922271728516,-116.045310974121,-116.0439453125,-115.980270385742,-115.733741760254,-115.471878051758,-115.341018676758,-115.303413391113,-115.33814239502,-115.58667755127,-116.780281066895,-117.337104797363,-117.723335266113,-117.935646057129,-118.188186645508,-118.221870422363,-118.226463317871,-118.148338317871,-117.878425598145,-117.742340087891,-117.887603759766,-118.371536254883,-118.583015441895,-118.868408203125,-118.952102661133,-118.987701416016,-118.993751525879,-118.984176635742,-118.959823608398,-118.944633483887,-118.58984375,-118.36865234375,-118.213470458984,-118.207473754883,-118.245903015137,-118.390480041504,-118.448638916016,-118.481300354004,-118.456588745117,-118.37451171875,-118.133110046387,-117.551712036133,-117.256439208984,-116.971672058105,-116.5732421875,-115.552200317383,-114.638236999512,-114.301902770996,-114.206398010254,-114.163963317871,-114.127052307129,-114.095458984375,-114.051704406738,-114.046142578125,-114.053756713867,-114.074752807617,-114.109176635742,-114.177680969238,-114.280319213867,-114.497848510742,-114.521537780762],[-105.288917541504,-105.33935546875,-105.43408203125,-105.572998046875,-105.800148010254,-106.071044921875,-106.112640380859,-106.180030822754,-106.525733947754,-106.750389099121,-106.921539306641,-106.949653625488,-106.831008911133,-106.69482421875,-106.613967895508,-106.362106323242,-105.512306213379,-105.317970275879,-105.114456176758,-104.834671020508,-104.71826171875,-104.648780822754,-104.587493896484,-104.555084228516,-104.552345275879,-104.582862854004,-104.621726989746,-104.791015625,-104.968650817871,-105.002586364746,-105.074615478516,-105.200637817383,-105.288917541504],[-79.5373077392578,-79.3667984008789,-78.2865219116211,-78.0629425048828,-77.3821258544922,-77.20654296875,-77.1197738647461,-77.04150390625,-77.0053253173828,-76.7586898803711,-76.6572723388672,-76.62158203125,-76.5697708129883,-76.4584503173828,-76.3311538696289,-76.28955078125,-76.3095703125,-76.2552719116211,-76.1350631713867,-76.0899887084961,-76.1833953857422,-76.4005355834961,-77.0135726928711,-77.8359375,-78.314208984375,-78.5540542602539,-79.1341857910156,-79.3193359375,-79.5005416870117,-79.8207015991211,-79.9368667602539,-79.9752960205078,-80.0516128540039,-80.1144485473633,-80.1464385986328,-80.1832962036133,-80.292724609375,-80.617919921875,-80.7268524169922,-80.7764205932617,-80.8241729736328,-80.8229446411133,-80.7979965209961,-80.7769546508789,-80.73583984375,-80.8270034790039,-80.8584976196289,-80.8607482910156,-80.848876953125,-80.8228530883789,-80.7627487182617,-80.6213912963867,-80.4123001098633,-80.1202621459961,-79.8893585205078,-79.5373077392578],[-86.58935546875,-86.5496597290039,-86.3212890625,-86.1271514892578,-85.8245544433594,-85.6438522338867,-85.0948715209961,-85.0015640258789,-85.0427703857422,-85.0657730102539,-85.0476608276367,-84.9884719848633,-84.8703155517578,-84.82373046875,-84.7895965576172,-84.7085952758789,-84.67431640625,-84.6581039428711,-84.6599655151367,-84.6994171142578,-84.8401336669922,-85.0322265625,-85.1309127807617,-85.25048828125,-85.3390579223633,-85.3966827392578,-85.5115203857422,-85.59619140625,-85.8133316040039,-85.91162109375,-85.8621063232422,-85.664794921875,-85.5457992553711,-85.4059066772461,-85.3218765258789,-85.0187530517578,-84.6084976196289,-84.3516616821289,-84.2837448120117,-84.2823181152344,-84.3474578857422,-84.6429672241211,-84.7775344848633,-84.8419952392578,-84.81103515625,-84.6446762084961,-84.623046875,-84.8494110107422,-84.9641571044922,-85.0568389892578,-85.1563949584961,-85.3411102294922,-85.3913116455078,-85.4977569580078,-85.5537109375,-85.6155776977539,-85.6378860473633,-85.64990234375,-85.64453125,-85.6194305419922,-85.5746078491211,-85.4547882080078,-85.3875961303711,-85.2621154785156,-84.9895553588867,-84.2566375732422,-84.2742691040039,-85.0940399169922,-85.3838882446289,-85.4547348022461,-85.0184097290039,-84.6160659790039,-84.4160614013672,-84.0889663696289,-83.7818908691406,-83.7765121459961,-83.9149856567383,-83.904052734375,-83.7298355102539,-83.4103546142578,-83.0204544067383,-82.9432144165039,-82.8433074951172,-82.6596221923828,-82.2027816772461,-81.9461441040039,-81.6053695678711,-81.4061508178711,-81.3440933227539,-81.2383346557617,-81.1517562866211,-81.025146484375,-80.8217315673828,-80.68115234375,-80.6034622192383,-80.582763671875,-80.619140625,-80.5918960571289,-80.5009307861328,-80.4308090209961,-80.2772521972656,-80.2747116088867,-80.3226547241211,-80.42431640625,-80.6751022338867,-80.9987258911133,-81.2293395996094,-81.2405776977539,-80.7607955932617,-80.6114807128906,-80.6046905517578,-80.7024383544922,-80.8214874267578,-80.9412078857422,-80.9193344116211,-80.69140625,-80.7332534790039,-80.84326171875,-80.8883819580078,-80.9210968017578,-80.94140625,-80.9426727294922,-80.925048828125,-80.9268035888672,-80.9478988647461,-80.9251480102539,-80.8584518432617,-80.80224609375,-80.7054214477539,-80.3861389160156,-80.1819305419922,-80.1161117553711,-79.9283218383789,-79.8843765258789,-80.0909194946289,-80.1089401245117,-80.0669937133789,-80.0418014526367,-79.9267120361328,-79.831298828125,-79.7778778076172,-79.6933059692383,-79.6538543701172,-79.5836944580078,-79.4274444580078,-79.3233413696289,-79.1943817138672,-79.000244140625,-79.0179672241211,-79.0177764892578,-79.0078125,-78.7759246826172,-78.6144561767578,-78.5851135253906,-78.5888671875,-78.62255859375,-78.7111358642578,-78.7908248901367,-78.8627471923828,-78.8201141357422,-78.6992721557617,-78.5824737548828,-78.4288101196289,-78.3074722290039,-78.1163558959961,-77.7260208129883,-77.5165100097656,-77.5357437133789,-77.6944808959961,-77.9261779785156,-78.2872085571289,-78.4530792236328,-78.4842758178711,-78.4794921875,-78.4588394165039,-78.4224166870117,-78.3502502441406,-78.0010299682617,-77.7532196044922,-77.5667953491211,-77.25537109375,-76.8935012817383,-76.6979522705078,-76.4732437133789,-76.1887741088867,-76.0873031616211,-75.96875,-75.8332061767578,-75.7042922973633,-75.2942352294922,-75.185791015625,-75.1200637817383,-75.0714797973633,-75.0398483276367,-75.0526885986328,-75.3941421508789,-75.5427703857422,-75.6409606933594,-75.7874069213867,-75.9112854003906,-75.9228057861328,-75.8968200683594,-75.8220748901367,-75.693359375,-75.5998992919922,-75.4283676147461,-75.1476516723633,-74.9031753540039,-74.6949234008789,-74.5196762084961,-74.37744140625,-74.29296875,-74.266357421875,-74.2093276977539,-74.2126007080078,-74.2482376098633,-74.3157196044922,-74.6214828491211,-74.7890625,-74.8929672241211,-75.2047882080078,-75.1910629272461,-74.95947265625,-74.7007827758789,-74.7073745727539,-74.8289566040039,-74.8683090209961,-74.83447265625,-74.8407287597656,-74.9312973022461,-75.0353546142578,-74.9961929321289,-74.7589340209961,-74.6953125,-74.5995559692383,-74.4880828857422,-74.4040985107422,-74.1390609741211,-73.9920883178711,-73.8666076660156,-73.8140563964844,-73.7072296142578,-73.7135772705078,-73.8686065673828,-74.197265625,-74.0633239746094,-73.9725112915039,-73.8508834838867,-73.7128448486328,-73.6216812133789,-73.4815902709961,-73.3977966308594,-73.2624053955078,-73.1806106567383,-73.1921844482422,-73.3104476928711,-73.2782211303711,-73.1868133544922,-72.9019546508789,-72.7030258178711,-72.5806121826172,-72.519287109375,-72.3646011352539,-72.1165008544922,-71.8751907348633,-71.6406707763672,-71.4599151611328,-71.3329620361328,-71.2560577392578,-71.2293930053711,-71.1865234375,-71.2193832397461,-71.3965835571289,-71.4950180053711,-71.5930633544922,-71.8561477661133,-71.9379348754883,-72.0238723754883,-72.2976989746094,-72.4491195678711,-72.5980453491211,-72.6327133178711,-72.3125534057617,-72.2239227294922,-72.1500015258789,-72.0091781616211,-71.7425308227539,-71.370849609375,-71.1862335205078,-71.0453567504883,-70.8880386352539,-70.8260803222656,-70.79248046875,-70.6726608276367,-70.636474609375,-70.6391143798828,-70.6552276611328,-70.76171875,-71.0217742919922,-71.1918029785156,-71.3804702758789,-71.5859375,-71.658447265625,-71.7293930053711,-71.8001403808594,-71.8901824951172,-71.7723617553711,-71.7272415161133,-71.6836853027344,-71.5649871826172,-71.4766616821289,-71.4266586303711,-71.3750991821289,-71.3248519897461,-71.27587890625,-71.2795867919922,-71.429443359375,-71.4051284790039,-71.3130798339844,-71.0452651977539,-70.9797821044922,-70.8505325317383,-70.5609359741211,-70.3372573852539,-70.084716796875,-69.9498062133789,-69.7957000732422,-69.6955108642578,-69.5601043701172,-69.3953628540039,-69.2890625,-69.168701171875,-69.0657272338867,-68.8907241821289,-68.4957504272461,-68.4466781616211,-68.4008255004883,-68.3582534790039,-68.363525390625,-68.4166488647461,-68.4825744628906,-68.5613327026367,-68.642822265625,-68.7936553955078,-68.8429183959961,-69.0794372558594,-69.2987289428711,-69.4356994628906,-69.698974609375,-70.0614242553711,-70.0577163696289,-69.9130859375,-69.7958526611328,-69.6345672607422,-69.4830093383789,-69.2461929321289,-68.9185562133789,-68.7782211303711,-68.7529296875,-68.734619140625,-68.7232971191406,-68.7769546508789,-68.839111328125,-69.00830078125,-68.8970184326172,-68.7440414428711,-68.65673828125,-68.5778350830078,-68.4893493652344,-68.3912582397461,-68.3050842285156,-68.2306671142578,-68.21044921875,-68.318603515625,-68.3271942138672,-68.2831039428711,-68.2035140991211,-68.1206512451172,-68.05908203125,-67.8553237915039,-67.7160186767578,-67.3636703491211,-67.3184051513672,-67.1958999633789,-67.1726531982422,-67.1927719116211,-67.2216262817383,-67.25927734375,-67.3367156982422,-67.8061981201172,-68.0204162597656,-68.1139678955078,-68.189453125,-68.2481002807617,-68.289794921875,-68.3720703125,-68.669921875,-68.8371124267578,-69.12451171875,-69.2276840209961,-69.2507858276367,-69.0749053955078,-68.7852554321289,-68.5130386352539,-68.0581588745117,-67.9082565307617,-67.8248519897461,-67.7245101928711,-67.3609390258789,-67.2369613647461,-67.0526809692383,-66.7708511352539,-66.7167510986328,-66.6852569580078,-66.67626953125,-66.6792984008789,-66.7074279785156,-66.8028869628906,-67.2080078125,-67.3316421508789,-67.4837951660156,-67.6072235107422,-67.7650375366211,-67.9384765625,-68.1981964111328,-68.4062957763672,-68.618896484375,-69.0406265258789,-68.9934539794922,-68.41552734375,-68.303955078125,-68.1212844848633,-67.8326187133789,-67.751708984375,-67.7510223388672,-67.7951202392578,-67.8832015991211,-68.015625,-68.32421875,-68.4503936767578,-68.5427703857422,-68.6667022705078,-68.7252960205078,-69.2188415527344,-69.3297882080078,-69.3426742553711,-69.319091796875,-68.8714370727539,-68.5406265258789,-68.3331985473633,-68.2104034423828,-68.1524887084961,-68.1483459472656,-68.0379409790039,-67.9384765625,-67.8750457763672,-67.7660140991211,-67.6559524536133,-67.5669403076172,-67.4555130004883,-67.3207015991211,-67.2024917602539,-67.1111755371094,-66.8542022705078,-66.7427215576172,-66.7139129638672,-66.7623977661133,-66.9972610473633,-67.032958984375,-66.900390625,-66.8309555053711,-66.8343200683594,-66.9051284790039,-66.923095703125,-66.8998031616211,-66.72900390625,-66.7023468017578,-66.6845703125,-66.6626968383789,-66.60546875,-66.6309509277344,-66.53076171875,-66.21240234375,-66.2663116455078,-66.2747039794922,-66.4138717651367,-66.5299835205078,-66.5264663696289,-66.4439468383789,-66.3923797607422,-66.3429641723633,-66.2251968383789,-65.98583984375,-65.9423828125,-65.9439926147461,-65.9748992919922,-65.8643569946289,-65.7589340209961,-65.7017059326172,-65.5693359375,-65.5090789794922,-65.4911117553711,-65.552001953125,-65.5408706665039,-65.4012680053711,-65.3871078491211,-65.4134826660156,-65.4422378540039,-65.4153289794922,-65.3003463745117,-65.0643997192383,-64.9769058227539,-64.9223175048828,-64.83544921875,-64.862548828125,-64.9563980102539,-65.0260238647461,-65.0714340209961,-65.0210952758789,-64.8298797607422,-64.6377944946289,-64.5275344848633,-64.3964385986328,-64.15625,-64.0194320678711,-63.8501930236816,-64.0774917602539,-64.0079574584961,-64.3032760620117,-64.4692916870117,-64.5804672241211,-64.699951171875,-64.5892639160156,-64.3759231567383,-64.3564453125,-64.18896484375,-64.063232421875,-63.8362312316895,-63.8241233825684,-63.6764678955078,-63.5916023254395,-63.5210952758789,-63.3158187866211,-63.0401382446289,-63.16162109375,-63.1946792602539,-63.2355461120605,-63.2583961486816,-63.3067893981934,-63.7015609741211,-63.63623046875,-63.4691925048828,-63.1436996459961,-62.9623031616211,-62.8333473205566,-62.7681617736816,-62.7104454040527,-62.6028785705566,-62.3797340393066,-62.1235809326172,-61.9685592651367,-61.8241195678711,-61.5146980285645,-61.3534164428711,-61.2997055053711,-61.3072242736816,-61.4530754089355,-61.52783203125,-61.7241172790527,-61.9044914245605,-62.0142593383789,-62.1233367919922,-62.0893096923828,-61.6526336669922,-61.576416015625,-61.57080078125,-61.8626976013184,-61.9563446044922,-62.1584434509277,-62.2769050598145,-62.37451171875,-62.5098152160645,-62.5531272888184,-62.4056625366211,-62.4198226928711,-62.4959983825684,-62.5335922241211,-62.2420845031738,-62.0239219665527,-61.9916038513184,-62.138671875,-62.2443351745605,-62.4677772521973,-62.59033203125,-62.6241188049316,-62.4973640441895,-62.4483871459961,-62.4103012084961,-62.38818359375,-62.3819808959961,-62.4856452941895,-62.6102523803711,-62.6588859558105,-62.7717247009277,-62.8172836303711,-62.9688949584961,-63.1689414978027,-63.2406730651855,-63.458740234375,-63.46435546875,-63.4097671508789,-63.4208946228027,-63.6519546508789,-63.6510734558105,-63.5092315673828,-63.3374519348145,-63.3642578125,-63.3633766174316,-63.4018058776855,-63.4858360290527,-63.6065940856934,-63.7371597290039,-63.7893562316895,-63.8332061767578,-63.8956069946289,-63.9762725830078,-64.0614242553711,-64.15185546875,-64.2504425048828,-64.345703125,-64.3097610473633,-64.2696762084961,-64.2857437133789,-64.3399429321289,-64.4698181152344,-64.5550765991211,-64.6653366088867,-64.7647933959961,-64.846923828125,-64.9796371459961,-65.1084976196289,-65.1756896972656,-65.2069854736328,-65.2820358276367,-65.3114776611328,-65.33740234375,-65.401611328125,-65.3781280517578,-65.2769546508789,-65.1848678588867,-65.0322265625,-64.8537139892578,-64.7725067138672,-64.6729965209961,-64.56396484375,-64.4453582763672,-64.50439453125,-64.6551818847656,-64.7611389160156,-64.8872528076172,-65.0044937133789,-65.3053741455078,-65.4155731201172,-65.543701171875,-65.8257293701172,-65.8910675048828,-65.857177734375,-65.6563491821289,-65.6883850097656,-65.7589874267578,-65.85595703125,-65.9400405883789,-66.063720703125,-66.2085952758789,-66.2773895263672,-66.419189453125,-66.4769515991211,-66.7123031616211,-66.759765625,-66.7873992919922,-66.8628921508789,-66.986328125,-67.0147933959961,-66.9704055786133,-66.968994140625,-67.0768585205078,-67.1896514892578,-67.3073272705078,-67.3176727294922,-67.1917495727539,-67.1897430419922,-67.2253952026367,-67.3112258911133,-67.3688430786133,-67.5597610473633,-67.7407684326172,-67.8684539794922,-67.8833999633789,-67.8005828857422,-67.7044906616211,-67.5472183227539,-67.2967300415039,-67.1832046508789,-67.2726593017578,-67.3504409790039,-67.3987731933594,-67.55078125,-67.8280258178711,-67.9582061767578,-68.1472625732422,-68.4599151611328,-68.5277862548828,-68.7489242553711,-68.7142105102539,-68.5716781616211,-68.4670867919922,-68.2171936035156,-68.1983413696289,-68.2606887817383,-68.2568359375,-68.1867218017578,-68.1150360107422,-67.9680633544922,-67.8941879272461,-67.866455078125,-67.954345703125,-67.9618148803711,-67.936767578125,-67.9060516357422,-67.7171401977539,-67.6380844116211,-67.5696258544922,-67.4901351928711,-67.3997039794922,-67.3463897705078,-67.330322265625,-67.3034210205078,-67.1179656982422,-67.1349639892578,-67.3260726928711,-67.3365249633789,-67.29833984375,-67.1775894165039,-67.0665054321289,-66.9985809326172,-66.9849166870117,-66.9856491088867,-66.9703598022461,-66.9115219116211,-66.8875045776367,-66.8606491088867,-66.8309097290039,-66.7996139526367,-66.7327651977539,-66.6974105834961,-66.6771469116211,-66.6666946411133,-66.635498046875,-66.5177688598633,-66.34521484375,-66.2237319946289,-66.209716796875,-66.301513671875,-66.2821273803711,-66.2146453857422,-66.1524963378906,-66.1075210571289,-66.0301742553711,-65.9385223388672,-65.76806640625,-65.6267623901367,-65.6052703857422,-65.51318359375,-65.4319381713867,-65.2748031616211,-65.3493194580078,-65.5127944946289,-65.5293502807617,-65.489990234375,-65.1786117553711,-65.0945281982422,-65.0746078491211,-65.2129898071289,-65.3398895263672,-65.5074691772461,-65.5936508178711,-65.580322265625,-65.3478012084961,-65.281982421875,-65.1927795410156,-65.1496124267578,-65.150634765625,-65.1873016357422,-65.1698760986328,-65.0105972290039,-64.9118194580078,-64.7877960205078,-64.678466796875,-64.6697235107422,-64.6861801147461,-64.7981414794922,-64.7681655883789,-64.63671875,-64.5763168334961,-64.4984893798828,-64.4109344482422,-64.4822235107422,-64.5615692138672,-64.55029296875,-64.4986343383789,-64.4981002807617,-64.5143508911133,-64.5869140625,-64.6646423339844,-64.6956100463867,-64.8862762451172,-64.9333038330078,-64.9897003173828,-65.1918487548828,-65.1839370727539,-65.1338348388672,-65.0894012451172,-65.0313415527344,-65.0047836303711,-65.0167007446289,-65.0580596923828,-65.0689468383789,-65.04931640625,-64.8948211669922,-64.8201675415039,-64.7673873901367,-64.7181167602539,-64.67236328125,-64.6836395263672,-64.7518539428711,-64.8687057495117,-64.9232406616211,-65.1329574584961,-65.1627883911133,-65.0465850830078,-65.0501937866211,-65.1084976196289,-65.1803207397461,-65.2658157348633,-65.3965377807617,-65.5724105834961,-65.7403793334961,-65.7798843383789,-65.8056640625,-65.8336944580078,-65.8640594482422,-65.9202651977539,-65.9788589477539,-66.2240219116211,-66.2492218017578,-66.22607421875,-66.2010726928711,-66.2286605834961,-66.2927703857422,-66.4144515991211,-66.4963836669922,-66.6004867553711,-66.6549835205078,-66.6598129272461,-66.630859375,-66.6364288330078,-66.6974563598633,-66.7232437133789,-66.74853515625,-66.7732391357422,-66.8314437866211,-66.9232940673828,-66.9747009277344,-67.0001449584961,-67.0179214477539,-67.1797866821289,-67.2609329223633,-67.4950180053711,-67.709228515625,-67.84423828125,-67.8932647705078,-67.8214340209961,-67.7425231933594,-67.7225646972656,-67.7587890625,-67.837890625,-68.2435531616211,-68.4937515258789,-68.6328659057617,-68.8589324951172,-68.9110794067383,-68.7892608642578,-68.6705551147461,-68.5551300048828,-68.3739242553711,-68.2080612182617,-68.1412582397461,-67.9153289794922,-67.7974624633789,-67.6759796142578,-67.6648941040039,-67.7237777709961,-67.7369613647461,-67.4682159423828,-67.3666458129883,-67.2685012817383,-67.2126922607422,-66.9795379638672,-66.9215316772461,-66.7140197753906,-66.6448745727539,-66.530517578125,-66.458740234375,-66.3572769165039,-66.28125,-66.0950164794922,-66.015625,-65.9801712036133,-66.0043487548828,-66.0269546508789,-66.1331558227539,-66.1164093017578,-66.0564498901367,-66.0588836669922,-66.1238708496094,-66.2566909790039,-66.32373046875,-66.4245147705078,-66.5513229370117,-66.8031234741211,-67.1810607910156,-67.322021484375,-67.3689956665039,-67.4401321411133,-68.3786163330078,-68.535888671875,-68.6336441040039,-68.724365234375,-69.0823211669922,-69.1255874633789,-69.3660125732422,-69.545166015625,-69.604736328125,-69.7995147705078,-69.9621124267578,-70.0709457397461,-70.2361297607422,-70.3440399169922,-70.5713348388672,-70.8014144897461,-71.0021438598633,-71.1057586669922,-71.09619140625,-70.946044921875,-70.99267578125,-71.2537078857422,-71.3472671508789,-71.5012664794922,-71.6171340942383,-71.85546875,-71.9922332763672,-71.9730529785156,-71.8191833496094,-71.696533203125,-71.6142578125,-71.4558639526367,-71.3874053955078,-71.380859375,-71.5134735107422,-71.5418930053711,-71.5656204223633,-71.6267623901367,-71.7252883911133,-71.8375473022461,-72.2229461669922,-72.2901382446289,-72.2887649536133,-72.2134780883789,-72.1724548339844,-72.1593780517578,-72.1742706298828,-72.2264633178711,-72.4500045776367,-72.4984359741211,-72.5861358642578,-72.6393051147461,-72.6780776977539,-72.7295913696289,-72.9131851196289,-73.17431640625,-73.2703170776367,-73.3770980834961,-73.4545364379883,-73.4436492919922,-73.2781753540039,-73.2712936401367,-73.4130859375,-73.626953125,-73.7284164428711,-73.7927780151367,-73.8678741455078,-73.9103546142578,-73.9503860473633,-73.9811019897461,-74.0255813598633,-74.0647964477539,-74.0987243652344,-74.097900390625,-74.1304702758789,-74.205078125,-74.4158706665039,-74.4612350463867,-74.512451171875,-74.5562515258789,-74.5925750732422,-74.63427734375,-74.681396484375,-74.7191925048828,-74.7477493286133,-74.8134231567383,-74.916259765625,-74.91943359375,-74.8230514526367,-74.7298278808594,-74.6400375366211,-74.6947250366211,-74.8939437866211,-75.0673828125,-75.2150421142578,-75.3284149169922,-75.48779296875,-75.7150344848633,-75.7667007446289,-75.8152389526367,-76.0318374633789,-76.1180648803711,-76.4068298339844,-76.4947204589844,-76.5615234375,-76.6265106201172,-76.7238311767578,-76.8561477661133,-77.0235366821289,-77.1656723022461,-77.2825164794922,-77.4028778076172,-77.5267562866211,-77.6277847290039,-77.760498046875,-77.7911605834961,-77.98486328125,-78.0452117919922,-78.174560546875,-78.1975555419922,-78.2008743286133,-78.189697265625,-78.1446304321289,-78.0956039428711,-78.0552749633789,-77.9945755004883,-77.8761672973633,-77.4474639892578,-77.3608856201172,-77.3638687133789,-77.4614791870117,-77.4604034423828,-77.4276885986328,-77.3580017089844,-77.3267059326172,-77.2511749267578,-77.0941390991211,-76.9585952758789,-76.7789077758789,-76.481689453125,-76.0669937133789,-75.8283233642578,-75.6481399536133,-75.5199203491211,-75.5015563964844,-75.5609359741211,-75.5908737182617,-75.589111328125,-75.5557632446289,-75.4521026611328,-75.4271545410156,-75.4351577758789,-75.4136657714844,-75.36279296875,-75.3571319580078,-75.3966827392578,-75.44580078125,-75.5046844482422,-75.77294921875,-75.7986831665039,-75.7085952758789,-75.316650390625,-75.1663055419922,-75.1092758178711,-75.0477523803711,-74.9817428588867,-74.849853515625,-74.6654739379883,-74.5748977661133,-74.4947738647461,-74.3907241821289,-74.2368621826172,-74.1384811401367,-73.9896011352539,-73.8779296875,-73.6753921508789,-73.55078125,-73.5607376098633,-73.6434097290039,-73.74609375,-73.8260726928711,-74.0331039428711,-74.2761688232422,-74.4010696411133,-74.4339294433594,-74.4164047241211,-74.3749008178711,-73.9336929321289,-73.584228515625,-73.4309539794922,-73.3573760986328,-73.2808151245117,-73.2011260986328,-73.0332489013672,-72.9853515625,-72.974853515625,-72.94677734375,-72.788818359375,-72.667724609375,-72.4851608276367,-72.3649368286133,-72.2200241088867,-72.234130859375,-72.3010787963867,-72.3528823852539,-72.5764617919922,-72.7252960205078,-72.9039535522461,-73.0634307861328,-73.3282241821289,-73.3314437866211,-73.2844772338867,-73.306884765625,-73.5801696777344,-73.6444778442383,-73.7494659423828,-73.8205032348633,-73.8793411254883,-73.8733367919922,-73.8344268798828,-73.7825164794922,-73.7806091308594,-73.7984390258789,-73.8221206665039,-73.8515625,-73.9351577758789,-74.072998046875,-74.1179656982422,-73.966064453125,-73.9892578125,-74.1828155517578,-74.2701187133789,-74.3499984741211,-74.4224090576172,-74.64794921875,-74.69580078125,-74.6805191040039,-74.699951171875,-74.7459945678711,-74.808349609375,-74.8929672241211,-74.910400390625,-74.7523880004883,-74.7432556152344,-74.8161163330078,-74.9251022338867,-74.9540023803711,-74.9172821044922,-74.7693328857422,-74.7166976928711,-74.8054656982422,-74.8548812866211,-74.9544448852539,-75.104248046875,-75.2132797241211,-75.3627395629883,-75.4569778442383,-75.5226593017578,-75.623046875,-75.8422393798828,-76.2347183227539,-76.4034194946289,-76.5850601196289,-76.6194381713867,-76.6162567138672,-76.6036605834961,-76.5817413330078,-76.5745620727539,-76.5876998901367,-76.5572280883789,-76.4951705932617,-76.3809051513672,-76.0892105102539,-75.9537124633789,-75.8585968017578,-75.7633819580078,-75.66796875,-75.6477508544922,-75.7490692138672,-75.7871551513672,-76.0464859008789,-76.1897964477539,-76.3162078857422,-76.407958984375,-76.4649429321289,-76.5203628540039,-76.5249557495117,-76.51611328125,-76.4632797241211,-76.2311096191406,-76.2340850830078,-76.423828125,-76.5132827758789,-76.5900421142578,-76.6865234375,-76.7423324584961,-76.9155731201172,-77.0196304321289,-77.0899429321289,-77.1288146972656,-77.1050796508789,-77.0186996459961,-76.8685989379883,-76.8585968017578,-76.9622573852539,-77.0159683227539,-77.2324752807617,-77.4942855834961,-77.5916519165039,-77.6353073120117,-77.6629867553711,-77.6747589111328,-77.721923828125,-77.7740249633789,-77.842529296875,-78.1567916870117,-78.2314453125,-78.2828140258789,-78.49072265625,-78.5748062133789,-78.6214370727539,-78.7726516723633,-78.8308639526367,-78.89990234375,-78.9797897338867,-79.06640625,-79.1597671508789,-79.253173828125,-79.3466262817383,-79.3973159790039,-79.4052276611328,-79.347412109375,-79.0175323486328,-78.933837890625,-78.8628387451172,-78.809814453125,-78.7748565673828,-78.77783203125,-78.8188018798828,-78.8896484375,-79.0928726196289,-79.3033218383789,-79.5154266357422,-79.6159133911133,-80.162109375,-80.2603988647461,-80.3868179321289,-80.6703109741211,-80.8257827758789,-81.0982894897461,-81.5595703125,-81.6519546508789,-81.5292510986328,-81.4217300415039,-81.3294906616211,-81.1968231201172,-81.0237350463867,-80.9248046875,-80.8428726196289,-80.8402786254883,-80.9217300415039,-81.564697265625,-81.9577178955078,-82.1387176513672,-82.2938461303711,-82.4877471923828,-82.9253845214844,-83.0911636352539,-83.1499481201172,-83.53076171875,-83.8590850830078,-84.5218734741211,-84.76513671875,-84.8292007446289,-84.9090805053711,-85.0526351928711,-85.432373046875,-85.780029296875,-86.1981964111328,-86.322021484375,-86.3614273071289,-86.4831008911133,-86.4998016357422,-86.4653778076172,-86.4309997558594,-86.3968734741211,-86.6243209838867,-86.7041473388672,-86.8092803955078,-87.1224670410156,-87.1719665527344,-87.1558074951172,-87.0740280151367,-87.0632781982422,-87.2378845214844,-87.50244140625,-87.6177749633789,-87.6702117919922,-87.7894515991211,-87.838134765625,-87.9006805419922,-88.1783218383789,-88.402099609375,-88.6629867553711,-88.78271484375,-88.8484420776367,-89.2083053588867,-89.2575225830078,-89.3715362548828,-89.4097671508789,-89.4559097290039,-89.3655242919922,-89.025146484375,-88.6956558227539,-88.5166473388672,-88.30908203125,-88.03857421875,-87.8449172973633,-87.534423828125,-87.1815948486328,-87.1400833129883,-87.3686065673828,-87.5723114013672,-87.76025390625,-87.8724594116211,-88.0606460571289,-88.5895004272461,-89.079345703125,-89.4176712036133,-89.6933135986328,-89.8053665161133,-89.8457489013672,-89.8885269165039,-89.9336853027344,-89.9773483276367,-90.01953125,-90.0251922607422,-89.9314956665039,-89.663818359375,-89.6572799682617,-89.7105484008789,-89.8228988647461,-89.8586959838867,-89.8731002807617,-89.8740234375,-89.8615188598633,-89.8168411254883,-89.7015151977539,-89.5364227294922,-89.3577117919922,-89.3271026611328,-89.3113784790039,-89.2876968383789,-89.2632293701172,-89.225341796875,-89.11474609375,-88.976806640625,-88.7609405517578,-88.7425308227539,-88.7396011352539,-88.7271499633789,-88.7051773071289,-88.1700134277344,-87.9264221191406,-87.7197799682617,-87.4723587036133,-86.7687530517578,-86.4063949584961,-85.9507827758789,-85.1104965209961,-85.0093231201172,-84.9835891723633,-84.94677734375,-84.9745101928711,-85.2042922973633,-85.4935989379883,-85.681884765625,-86.0005340576172,-86.0864715576172,-86.4813995361328,-86.5746536254883,-86.6293487548828,-86.6677703857422,-86.6562957763672,-86.5946273803711,-86.3803253173828,-86.3225631713867,-86.3240203857422,-86.3480453491211,-86.3509750366211,-86.3413543701172,-86.2971649169922,-86.2184524536133,-86.0361328125,-85.7500915527344,-85.5371551513672,-85.3271942138672,-85.0787048339844,-85.0233917236328,-85.1375961303711,-85.4053649902344,-85.7572708129883,-85.9454116821289,-86.179443359375,-86.4732437133789,-86.58935546875],[-100.001907348633,-99.157958984375,-99.0396499633789,-98.7845230102539,-98.5193405151367,-98.15185546875,-97.927734375,-97.8321838378906,-97.6699676513672,-97.5818328857422,-97.3270568847656,-97.2247543334961,-97.1705093383789,-97.1117172241211,-97.0112838745117,-96.99658203125,-97.001708984375,-97.0945816040039,-97.1563949584961,-97.2841796875,-97.394775390625,-97.4897994995117,-97.5969696044922,-97.6258850097656,-97.6146011352539,-97.5858459472656,-97.5318374633789,-97.4701156616211,-97.3502883911133,-97.287109375,-97.2303695678711,-97.2725067138672,-97.4840774536133,-97.7958984375,-98.1758270263672,-98.3755798339844,-98.4168472290039,-98.4369659423828,-98.430908203125,-98.4217758178711,-98.3666458129883,-98.1808166503906,-98.06103515625,-97.9394073486328,-97.7248001098633,-97.6363296508789,-97.4756851196289,-97.3287582397461,-97.2958450317383,-97.3099136352539,-97.3710021972656,-97.377685546875,-97.2376022338867,-97.0830078125,-97.0728988647461,-97.1404800415039,-97.158935546875,-97.1281280517578,-97.0518035888672,-96.8690414428711,-96.6712875366211,-96.5920944213867,-96.5420837402344,-96.4892044067383,-96.4456024169922,-96.4401397705078,-96.4728469848633,-96.5198745727539,-96.6382827758789,-96.7455139160156,-96.8014602661133,-96.7958984375,-96.6687545776367,-96.6155700683594,-96.5928726196289,-96.6005859375,-96.6181106567383,-96.7663116455078,-96.75830078125,-96.71728515625,-96.6243667602539,-96.6134262084961,-96.9464797973633,-97.024658203125,-97.11669921875,-97.2222137451172,-97.4612350463867,-97.582275390625,-98.1813430786133,-98.241943359375,-98.2838897705078,-98.3070831298828,-98.3133773803711,-98.3026885986328,-98.3058166503906,-98.3227005004883,-98.3893051147461,-98.4588394165039,-98.4207992553711,-98.2314453125,-98.1953125,-98.1901397705078,-98.1986312866211,-98.4123001098633,-98.5359344482422,-98.6628875732422,-98.7838363647461,-98.8987731933594,-98.9862289428711,-99.1671371459961,-99.2236328125,-99.2761688232422,-99.4036636352539,-99.5814437866211,-99.7347183227539,-100.124114990234,-100.32568359375,-100.594482421875,-100.706832885742,-100.800201416016,-100.983642578125,-101.026222229004,-101.093116760254,-101.208549499512,-101.250686645508,-101.318695068359,-101.498336791992,-101.723930358887,-101.774513244629,-101.804443359375,-101.832908630371,-101.909324645996,-101.973678588867,-102.402244567871,-102.657081604004,-102.708740234375,-102.713668823242,-102.6875,-102.628471374512,-102.551078796387,-102.503814697266,-102.336128234863,-102.204010009766,-102.019622802734,-101.922462463379,-101.835403442383,-101.798049926758,-101.754539489746,-101.617774963379,-101.543601989746,-101.434623718262,-101.3505859375,-101.273193359375,-101.087600708008,-100.896049499512,-100.484764099121,-100.468017578125,-100.442573547363,-100.395698547363,-100.367523193359,-100.227928161621,-100.188331604004,-100.128128051758,-100.092384338379,-100.096733093262,-100.184471130371,-100.236183166504,-100.282669067383,-100.334373474121,-100.446189880371,-100.531394958496,-100.550193786621,-100.536376953125,-100.489311218262,-100.438819885254,-100.340721130371,-100.225883483887,-100.066993713379,-99.9664077758789,-99.8251495361328,-100.005905151367,-100.257957458496,-100.366111755371,-100.497993469238,-100.587013244629,-100.755325317383,-100.889350891113,-101.450874328613,-101.482086181641,-101.523193359375,-101.518455505371,-101.463043212891,-101.323143005371,-101.114936828613,-100.975784301758,-100.854103088379,-100.676803588867,-100.521675109863,-100.508941650391,-100.536331176758,-100.607131958008,-100.657913208008,-100.78271484375,-100.898246765137,-100.952583312988,-100.981536865234,-100.985107421875,-100.962989807129,-100.915138244629,-100.483642578125,-100.182327270508,-99.9911117553711,-99.911865234375,-99.9394989013672,-100.040084838867,-100.15380859375,-100.224800109863,-100.22705078125,-100.138473510742,-100.001907348633],[-98.2703628540039,-98.5582046508789,-98.6910629272461,-98.7613830566406,-98.8166046142578,-98.9739303588867,-99.2980499267578,-99.3851547241211,-99.4169921875,-99.40380859375,-99.3456039428711,-99.0968704223633,-99.0046844482422,-98.9667053222656,-98.9044952392578,-98.8181686401367,-98.5849609375,-98.06103515625,-97.8004379272461,-97.6982421875,-97.6674270629883,-97.6591339111328,-97.67333984375,-97.7547378540039,-97.861083984375,-98.14697265625,-98.2703628540039],[-93.1708526611328,-92.7780227661133,-92.5868148803711,-92.4928207397461,-92.3138656616211,-92.2227020263672,-91.8741760253906,-91.6304168701172,-91.0879898071289,-90.62744140625,-90.4580078125,-90.3545913696289,-90.3813934326172,-90.4661636352539,-90.5655746459961,-90.7645568847656,-90.9336929321289,-90.9754867553711,-91.001953125,-91.067626953125,-91.2493209838867,-91.2977981567383,-91.5537109375,-91.4660110473633,-91.4259262084961,-91.4596176147461,-91.6209945678711,-91.788330078125,-91.9053268432617,-92.117919921875,-92.2349090576172,-92.3919448852539,-93.3406295776367,-93.5786590576172,-94.2113265991211,-94.1517105102539,-93.9200210571289,-93.7705535888672,-93.572265625,-93.5464859008789,-93.533935546875,-93.5415954589844,-93.55517578125,-93.87060546875,-93.9725570678711,-94.0375442504883,-94.1437454223633,-94.4971694946289,-94.6112289428711,-95.0078659057617,-95.1929702758789,-95.1667938232422,-95.1926727294922,-95.2510223388672,-95.547607421875,-95.580322265625,-95.6021499633789,-95.6131820678711,-95.6122055053711,-95.5915985107422,-95.5892562866211,-95.6041030883789,-95.6442413330078,-95.6479949951172,-95.645263671875,-95.6329040527344,-95.5694351196289,-95.4474105834961,-95.385986328125,-94.9961395263672,-94.8168411254883,-94.6976089477539,-94.6910171508789,-94.7971649169922,-94.8969268798828,-95.0594711303711,-95.1341323852539,-95.1490249633789,-95.152587890625,-95.144775390625,-95.1211929321289,-95.0398406982422,-94.9735336303711,-94.7289505004883,-94.4825668334961,-93.9388122558594,-93.7846145629883,-93.5492248535156,-93.4103012084961,-93.1708526611328],[-119.736328125,-119.728561401367,-119.471084594727,-119.314842224121,-119.205955505371,-119.171432495117,-119.149604797363,-119.138771057129,-119.131889343262,-119.117965698242,-119.08251953125,-119.025680541992,-118.744132995605,-118.625289916992,-118.543998718262,-118.199661254883,-117.965866088867,-117.707466125488,-117.514839172363,-117.198837280273,-116.950393676758,-116.722358703613,-115.957717895508,-115.634323120117,-115.510696411133,-115.455657958984,-115.407524108887,-115.392822265625,-115.411575317383,-115.446876525879,-115.524467468262,-115.992279052734,-116.238616943359,-116.482521057129,-117.0654296875,-117.464462280273,-117.983200073242,-118.961570739746,-119.077980041504,-119.131546020508,-119.407760620117,-119.512847900391,-119.767478942871,-120.089752197266,-120.179885864258,-120.194435119629,-120.310012817383,-120.366264343262,-120.443161010742,-120.460929870605,-120.519668579102,-120.619338989258,-120.930313110352,-121.159812927246,-121.47216796875,-121.546821594238,-121.622169494629,-121.700675964355,-121.749366760254,-122.156639099121,-122.549507141113,-122.719772338867,-122.839942932129,-122.9365234375,-123.095649719238,-123.210601806641,-123.314750671387,-123.393363952637,-123.595169067383,-123.681831359863,-123.755569458008,-123.953277587891,-124.007766723633,-124.759963989258,-125.126129150391,-125.214645385742,-125.296676635742,-125.766899108887,-125.8291015625,-125.845306396484,-125.789657592773,-125.767730712891,-125.76050567627,-125.768600463867,-125.762596130371,-125.583793640137,-125.61279296875,-125.6337890625,-125.646781921387,-125.627304077148,-125.575485229492,-125.512405395508,-125.438087463379,-125.382766723633,-125.306007385254,-125.168312072754,-125.070213317871,-124.987106323242,-124.984664916992,-125.0185546875,-125.030220031738,-125.014739990234,-125.015426635742,-125.000381469727,-124.969673156738,-124.930854797363,-124.582565307617,-124.564949035645,-124.560836791992,-124.570220947266,-124.588279724121,-124.64331817627,-124.736427307129,-124.817092895508,-124.836418151855,-124.804054260254,-124.646919250488,-124.59398651123,-124.424217224121,-124.114151000977,-124.030166625977,-123.797264099121,-123.797805786133,-123.873046875,-124.088035583496,-124.191497802734,-124.2607421875,-124.575340270996,-124.629096984863,-124.645027160645,-124.709320068359,-124.696235656738,-123.468307495117,-122.623153686523,-121.747901916504,-121.504150390625,-121.315238952637,-121.128707885742,-120.881645202637,-120.554489135742,-119.94359588623,-119.562652587891,-119.715377807617,-119.736915588379,-119.736328125],[-97.3555145263672,-97.6561050415039,-97.7215805053711,-97.75,-97.5163116455078,-97.41650390625,-97.3182144165039,-97.2913131713867,-97.3038558959961,-97.3555145263672],[-95.306640625,-95.3524398803711,-95.4415054321289,-95.7771987915039,-95.8343811035156,-95.8507385253906,-95.7744140625,-95.74560546875,-95.6604461669922,-95.5102005004883,-95.3525390625,-95.2783737182617,-95.2744674682617,-95.306640625],[-104.119926452637,-104.308685302734,-104.634323120117,-104.828125,-104.887405395508,-104.881637573242,-104.848098754883,-104.801315307617,-104.690383911133,-104.648826599121,-104.474166870117,-104.34619140625,-104.074661254883,-103.9169921875,-103.851165771484,-103.804100036621,-103.757911682129,-103.746482849121,-103.667236328125,-103.643501281738,-103.642135620117,-103.664260864258,-103.709716796875,-103.813919067383,-104.119926452637],[-93.5425720214844,-93.478271484375,-93.4666061401367,-93.4634780883789,-93.4908676147461,-93.5091857910156,-93.53564453125,-93.5482940673828,-93.5471649169922,-93.5730895996094,-93.6261672973633,-93.9845733642578,-94.2060546875,-94.5345230102539,-94.697265625,-94.8038558959961,-94.958740234375,-95.2860870361328,-95.4512252807617,-95.8654251098633,-96.09423828125,-96.1817398071289,-96.2701187133789,-96.294189453125,-96.3185577392578,-96.3431625366211,-96.3863220214844,-96.5598678588867,-96.5911636352539,-96.599609375,-96.596923828125,-96.5657730102539,-96.3828659057617,-96.2923812866211,-96.1803741455078,-96.118408203125,-96.1249008178711,-95.9546356201172,-95.8531723022461,-95.6707992553711,-95.0495147705078,-94.878173828125,-94.6486358642578,-94.42724609375,-94.2566909790039,-93.9090805053711,-93.7508316040039,-93.6668472290039,-93.5912170410156,-93.49755859375,-93.53173828125,-93.5518112182617,-93.5425720214844],[-96.0785598754883,-96.1563949584961,-96.2366180419922,-96.344482421875,-96.4616241455078,-96.6219711303711,-96.6790084838867,-96.7228469848633,-96.8571319580078,-96.9151382446289,-96.9696273803711,-97.0206527709961,-96.9828109741211,-96.8561553955078,-96.5229034423828,-96.4276885986328,-96.417236328125,-96.3972702026367,-96.3678207397461,-96.1454086303711,-96.0398406982422,-95.9598617553711,-95.9686050415039,-96.0785598754883],[-121.076225280762,-121.154289245605,-121.240913391113,-121.221092224121,-121.026321411133,-121.015434265137,-121.018074035645,-121.042289733887,-120.993019104004,-120.913963317871,-120.887794494629,-120.878707885742,-120.896873474121,-120.921249389648,-120.954933166504,-121.006637573242,-121.076225280762],[-94.5265579223633,-94.6243667602539,-94.75146484375,-94.787353515625,-94.8147430419922,-94.8336486816406,-94.860107421875,-94.8940963745117,-94.9012222290039,-94.88134765625,-94.8397903442383,-94.7448272705078,-94.5378952026367,-94.4986801147461,-94.4712905883789,-94.443359375,-94.4137649536133,-94.3322296142578,-94.2962875366211,-94.3040084838867,-94.3295440673828,-94.5265579223633],[-118.328125,-118.613868713379,-118.817138671875,-119.086669921875,-119.306053161621,-119.383247375488,-119.394577026367,-119.320167541504,-119.22681427002,-119.003463745117,-118.626068115234,-118.379005432129,-118.136672973633,-117.889350891113,-117.752487182617,-117.633697509766,-117.512596130371,-117.499114990234,-117.626365661621,-117.715957641602,-117.890815734863,-118.226516723633,-118.328125],[-79.0630874633789,-79.0517578125,-79.1244125366211,-79.3556671142578,-79.5445327758789,-79.6387710571289,-79.69873046875,-79.55126953125,-79.3817901611328,-79.1783218383789,-79.0093231201172,-78.9258804321289,-78.8451614379883,-78.9464340209961,-79.056640625,-79.0630874633789],[-102.227348327637,-102.017868041992,-102.008010864258,-102.047462463379,-102.318115234375,-102.423439025879,-102.511375427246,-102.57958984375,-102.943550109863,-103.314743041992,-103.244728088379,-103.04150390625,-103.201560974121,-103.769775390625,-103.985252380371,-103.80078125,-103.984519958496,-104.242477416992,-104.406051635742,-104.350631713867,-104.012062072754,-103.571441650391,-103.098243713379,-102.72802734375,-102.584075927734,-102.5361328125,-102.490043640137,-102.425682067871,-102.227348327637],[-104.022850036621,-103.973480224609,-103.821098327637,-103.722755432129,-103.613136291504,-103.584617614746,-103.190139770508,-103.051322937012,-103.033546447754,-103.082954406738,-103.199516296387,-103.311378479004,-103.472213745117,-104.270652770996,-104.357521057129,-104.407661437988,-104.506446838379,-104.576606750488,-104.60302734375,-104.585693359375,-104.500389099121,-104.205131530762,-104.074508666992,-103.992477416992,-103.959083557129,-103.969184875488,-104.022850036621],[-97.700927734375,-97.6897430419922,-97.7018508911133,-97.7371139526367,-97.73876953125,-97.7068328857422,-97.5731430053711,-97.5306625366211,-97.5242691040039,-97.5310592651367,-97.6134796142578,-97.6500015258789,-97.6521530151367,-97.60302734375,-97.6016616821289,-97.6942367553711,-97.8905334472656,-97.86279296875,-97.4395523071289,-97.4075164794922,-97.4096145629883,-97.3360366821289,-97.3634796142578,-97.4652328491211,-97.6533203125,-97.8782196044922,-97.8527297973633,-97.7048873901367,-97.659912109375,-97.67431640625,-97.7993698120117,-97.8427276611328,-97.9708480834961,-98.0453109741211,-98.0687484741211,-98.0916976928711,-98.0767593383789,-97.989990234375,-97.9533233642578,-97.9917984008789,-98.1209487915039,-98.295166015625,-98.5686492919922,-98.7035140991211,-98.8348083496094,-99.0100631713867,-99.1558074951172,-99.2449264526367,-99.3261184692383,-99.4206008911133,-99.6268997192383,-99.9466323852539,-100.234375,-100.292282104492,-100.356636047363,-100.483497619629,-100.45947265625,-100.152053833008,-100.145698547363,-100.364112854004,-100.614898681641,-100.731155395508,-100.704246520996,-100.7119140625,-100.279640197754,-99.9652862548828,-99.7702102661133,-99.7560043334961,-99.5911560058594,-99.2094268798828,-99.194580078125,-99.9151382446289,-100.901756286621,-101.206832885742,-101.461326599121,-102.541404724121,-102.58740234375,-102.700393676758,-102.797508239746,-102.727836608887,-102.410697937012,-102.252052307129,-102.270652770996,-102.144729614258,-101.942817687988,-101.599655151367,-101.421241760254,-101.261428833008,-101.119384765625,-100.972801208496,-101.009910583496,-101.258834838867,-101.288040161133,-101.414993286133,-101.470314025879,-101.505905151367,-101.507858276367,-101.431350708008,-101.716796875,-101.823387145996,-101.872123718262,-101.861373901367,-101.771392822266,-101.528953552246,-101.557029724121,-101.909820556641,-101.987449645996,-102.137748718262,-102.104690551758,-101.964202880859,-101.858497619629,-101.787551879883,-101.67724609375,-101.415191650391,-101.339752197266,-101.139060974121,-101.087890625,-101.094192504883,-101.055809020996,-100.900093078613,-100.230659484863,-100.105712890625,-100.020118713379,-99.865478515625,-99.7748489379883,-99.7012710571289,-99.6889114379883,-99.9783248901367,-100.050971984863,-100.112838745117,-100.085792541504,-100.001754760742,-99.7901840209961,-99.5410690307617,-99.8172378540039,-99.9976119995117,-100.182762145996,-100.414207458496,-100.414352416992,-100.357612609863,-100.042724609375,-99.9831008911133,-99.9777374267578,-100.081886291504,-100.174652099609,-100.650688171387,-100.819877624512,-100.873626708984,-100.890769958496,-100.829742431641,-100.57373046875,-100.387939453125,-100.068695068359,-99.8140640258789,-99.6690444946289,-99.3294982910156,-99.1696243286133,-98.8903350830078,-98.9710006713867,-99.0236282348633,-98.9408721923828,-98.7108383178711,-98.527587890625,-98.2886657714844,-98.2361755371094,-97.9673385620117,-97.8083953857422,-97.7258834838867,-97.700927734375],[-101.226119995117,-101.485206604004,-101.60498046875,-101.613082885742,-101.509475708008,-101.1650390625,-100.962158203125,-100.886474609375,-100.62158203125,-100.467231750488,-100.269134521484,-100.74658203125,-101.226119995117],[-108.292381286621,-108.166313171387,-108.018753051758,-107.852294921875,-107.77685546875,-107.723487854004,-107.721290588379,-107.731834411621,-107.755172729492,-107.970405578613,-108.020706176758,-107.951072692871,-107.917526245117,-107.702590942383,-107.540916442871,-107.418258666992,-107.216209411621,-107.135688781738,-107.080421447754,-107.050384521484,-106.970947265625,-106.913528442383,-106.904205322266,-106.902778625488,-106.891700744629,-106.693115234375,-106.688087463379,-106.759376525879,-106.820068359375,-106.862060546875,-106.845649719238,-106.804100036621,-106.677001953125,-106.528617858887,-106.396583557129,-105.904838562012,-105.711326599121,-105.632667541504,-105.604446411133,-105.563278198242,-105.480903625488,-105.4814453125,-105.519485473633,-105.678413391113,-105.70263671875,-105.862594604492,-105.971969604492,-106.092628479004,-106.588233947754,-106.961128234863,-107.055610656738,-107.153411865234,-107.4619140625,-107.820068359375,-108.023635864258,-108.226608276367,-108.354438781738,-108.474754333496,-108.594184875488,-108.751319885254,-108.670265197754,-108.63330078125,-108.666015625,-108.831298828125,-109.002540588379,-109.503120422363,-110.17578125,-110.38671875,-110.543411254883,-110.624755859375,-110.749320983887,-110.940872192383,-111.287551879883,-111.728713989258,-112.519332885742,-113.016891479492,-113.514060974121,-113.671577453613,-113.836814880371,-114.174751281738,-114.268257141113,-114.376953125,-114.312698364258,-114.132469177246,-113.862892150879,-113.324310302734,-112.835990905762,-112.663032531738,-112.192825317383,-111.955764770508,-111.784225463867,-111.671096801758,-111.503265380859,-111.257957458496,-111.078910827637,-111.033493041992,-111.093460083008,-111.181205749512,-111.473922729492,-111.620849609375,-111.780860900879,-112.00048828125,-112.214164733887,-112.255516052246,-112.478073120117,-112.597068786621,-112.652389526367,-112.703125,-112.799613952637,-112.855323791504,-112.951362609863,-113.339645385742,-113.711769104004,-113.794624328613,-113.844970703125,-113.855377197266,-113.860939025879,-113.886032104492,-113.853317260742,-113.810890197754,-113.7587890625,-113.503021240234,-113.467094421387,-113.588958740234,-113.878517150879,-113.916358947754,-113.984130859375,-114.016502380371,-114.053413391113,-114.074905395508,-114.124664306641,-114.16845703125,-114.284957885742,-114.429008483887,-114.482818603516,-114.513816833496,-114.503952026367,-114.357765197754,-114.356101989746,-114.451759338379,-114.859176635742,-115.020111083984,-115.077049255371,-115.128318786621,-115.173835754395,-115.279632568359,-115.342620849609,-115.413192749023,-115.478073120117,-115.537307739258,-115.574066162109,-115.608985900879,-115.683158874512,-115.728851318359,-116.142623901367,-116.47607421875,-116.841011047363,-117.004829406738,-117.501960754395,-117.565231323242,-117.600090026855,-117.596733093262,-117.576080322266,-117.513130187988,-117.387794494629,-117.335548400879,-117.257614135742,-117.154205322266,-116.890777587891,-116.212745666504,-116.077156066895,-115.335350036621,-115.250686645508,-115.14183807373,-115.117240905762,-115.121871948242,-116.034324645996,-116.425628662109,-117.025199890137,-117.13793182373,-117.163619995117,-117.038566589355,-116.97265625,-116.802146911621,-116.3896484375,-115.838081359863,-115.476860046387,-115.173736572266,-114.991508483887,-115.602249145508,-116.337890625,-116.444236755371,-116.654304504395,-116.66455078125,-116.580467224121,-116.549667358398,-116.609771728516,-116.591300964355,-116.454246520996,-116.209861755371,-116.059135437012,-115.768264770508,-114.939407348633,-114.778610229492,-114.880172729492,-115.024566650391,-115.664459228516,-115.796867370605,-115.822166442871,-115.831741333008,-115.825584411621,-115.779304504395,-115.580665588379,-114.998489379883,-114.766845703125,-114.534767150879,-114.298973083496,-114.193946838379,-114.141258239746,-114.115768432617,-114.11279296875,-114.101463317871,-114.058837890625,-113.923294067383,-113.823486328125,-113.362983703613,-113.171287536621,-112.978469848633,-112.697608947754,-112.333885192871,-111.865234375,-111.867622375488,-112.047172546387,-112.080909729004,-112.056694030762,-111.877395629883,-111.709373474121,-111.549118041992,-111.513236999512,-111.454444885254,-111.372749328613,-111.275688171387,-111.163284301758,-111.052688598633,-110.889595031738,-110.7255859375,-110.459373474121,-109.086380004883,-109.005027770996,-108.947166442871,-108.91259765625,-108.899513244629,-108.918212890625,-108.944778442383,-109.796043395996,-109.870506286621,-109.454643249512,-109.424751281738,-109.416603088379,-109.430374145508,-109.48681640625,-109.710159301758,-109.907814025879,-110.200775146484,-110.247016906738,-110.284866333008,-110.314453125,-110.309471130371,-110.27001953125,-109.981590270996,-109.864845275879,-109.505027770996,-109.338531494141,-109.219535827637,-109.098243713379,-108.831642150879,-108.553901672363,-108.492385864258,-108.466987609863,-108.477783203125,-108.512451171875,-108.61181640625,-108.635154724121,-108.627632141113,-108.559516906738,-108.538627624512,-108.523536682129,-108.512496948242,-108.345458984375,-108.193557739258,-108.123191833496,-108.177932739258,-108.305809020996,-108.381889343262,-108.406150817871,-108.386817932129,-108.292381286621],[-89.7264633178711,-89.7732849121094,-89.9241180419922,-89.97412109375,-90.0542984008789,-90.16455078125,-90.2935028076172,-90.4409637451172,-90.5562515258789,-90.5625,-90.5248031616211,-90.4095230102539,-90.1363296508789,-89.9487762451172,-89.7745590209961,-89.7252960205078,-89.6954116821289,-89.6944351196289,-89.7086410522461,-89.7875518798828,-89.8221206665039,-89.8219223022461,-89.8047790527344,-89.77294921875,-89.7263641357422,-89.7264633178711],[-113.560691833496,-113.712455749512,-114.75146484375,-114.808296203613,-114.835258483887,-114.647071838379,-114.419876098633,-113.891647338867,-113.70751953125,-113.585403442383,-113.516502380371,-113.487594604492,-113.560691833496],[-94.2949676513672,-94.1079559326172,-93.9480514526367,-93.8109359741211,-93.6083984375,-93.4207534790039,-93.2765579223633,-93.2300262451172,-93.2118682861328,-93.1892547607422,-93.1899871826172,-93.2005844116211,-93.263671875,-93.3167495727539,-93.42626953125,-93.4845657348633,-93.5345687866211,-93.421875,-92.995361328125,-92.7162628173828,-92.2970199584961,-91.7894515991211,-91.5484390258789,-91.3050231933594,-91.124267578125,-90.7384262084961,-90.6047897338867,-90.5546417236328,-90.5426254272461,-90.6216354370117,-90.8640594482422,-91.2630310058594,-91.3359909057617,-91.3980941772461,-91.4432601928711,-91.4150848388672,-91.3338851928711,-90.8547897338867,-89.2845230102539,-89.2190933227539,-89.2365264892578,-89.2920837402344,-89.4065856933594,-90.3120574951172,-90.8273468017578,-91.2603988647461,-91.4073257446289,-91.2794418334961,-91.019775390625,-90.8023910522461,-90.7121124267578,-90.2513656616211,-90.176025390625,-90.0327682495117,-89.9125518798828,-89.7934112548828,-89.6953125,-89.6500473022461,-89.51123046875,-89.277587890625,-89.2048797607422,-89.2045364379883,-89.2566375732422,-89.3612289428711,-89.6254425048828,-89.6460494995117,-89.3373031616211,-89.2804183959961,-88.9167022705078,-88.8688507080078,-88.8389205932617,-88.8041076660156,-88.8196334838867,-88.8640594482422,-88.8521499633789,-88.783935546875,-88.7148971557617,-88.6449661254883,-88.5690460205078,-88.2013168334961,-87.729736328125,-87.6436538696289,-87.5724105834961,-87.5391159057617,-87.3646469116211,-87.2569351196289,-86.814453125,-86.5447235107422,-86.4365234375,-86.236328125,-85.9514617919922,-85.904541015625,-86.0687484741211,-85.9729995727539,-85.5812530517578,-85.372314453125,-84.9867630004883,-84.7501983642578,-84.6048812866211,-84.1276397705078,-84.0142593383789,-83.9319839477539,-83.7445831298828,-83.2371139526367,-83.0934143066406,-82.553466796875,-82.3538589477539,-82.1536636352539,-81.6473617553711,-81.2685546875,-81.1507797241211,-81.1926803588867,-81.1735305786133,-81.1244125366211,-81.0007858276367,-80.5277328491211,-80.3219757080078,-80.1583480834961,-80.1191940307617,-80.125732421875,-80.28662109375,-80.2604446411133,-80.099609375,-79.7377014160156,-79.6602020263672,-79.5857391357422,-79.5078125,-79.5090866088867,-79.6344223022461,-79.9771499633789,-80.3575744628906,-80.3819808959961,-80.2606430053711,-80.13525390625,-80.0364227294922,-79.7330093383789,-79.6640625,-79.5248107910156,-79.4603958129883,-79.4014129638672,-79.5079574584961,-79.9444808959961,-80.2024459838867,-80.2892074584961,-80.3477554321289,-80.3145446777344,-80.1897506713867,-80.14892578125,-80.1922302246094,-80.2126998901367,-80.2102508544922,-80.2206039428711,-80.2627410888672,-80.2777328491211,-81.2262191772461,-81.3404769897461,-81.607177734375,-81.808837890625,-81.940185546875,-82.0685043334961,-82.4147415161133,-82.6454086303711,-82.7357864379883,-82.9310607910156,-82.9784164428711,-83.0576171875,-83.1169967651367,-83.1123046875,-83.0873031616211,-83.1026382446289,-83.1582946777344,-83.2203140258789,-83.4070358276367,-83.5220718383789,-83.5435562133789,-83.509765625,-83.4873046875,-83.3642044067383,-83.34130859375,-83.3936996459961,-83.4122085571289,-83.5318832397461,-83.6218795776367,-83.8680648803711,-84.2451629638672,-84.425537109375,-84.6670913696289,-84.8182601928711,-84.9163055419922,-85.0115280151367,-85.0614242553711,-85.0867691040039,-85.1334457397461,-85.2143096923828,-85.3392562866211,-85.4423370361328,-85.4744110107422,-85.4886703491211,-85.51171875,-85.5435104370117,-85.8080062866211,-85.9556121826172,-86.10986328125,-86.2109375,-86.340576171875,-86.6554260253906,-86.7307586669922,-86.6661148071289,-86.7701644897461,-86.9947280883789,-87.36376953125,-87.5925827026367,-88.005859375,-88.4230499267578,-88.500732421875,-88.5556640625,-88.557861328125,-88.5376434326172,-88.4766082763672,-88.3747024536133,-88.3395462036133,-88.4314422607422,-88.4881820678711,-88.5349578857422,-88.6820297241211,-88.77783203125,-88.85107421875,-88.8833999633789,-88.9078140258789,-88.9401321411133,-88.9803695678711,-89.0196304321289,-89.057861328125,-89.1152801513672,-89.1919403076172,-89.2191390991211,-89.1968231201172,-89.1890640258789,-89.19580078125,-89.2618637084961,-89.4500045776367,-89.5586929321289,-89.8443832397461,-90.0153274536133,-90.3616180419922,-90.5532684326172,-90.7840805053711,-90.966796875,-90.95751953125,-90.8776321411133,-90.8802261352539,-91.1298828125,-91.1637268066406,-91.1345748901367,-91.167724609375,-91.3394470214844,-91.5083465576172,-91.5491256713867,-91.665771484375,-91.8710403442383,-91.9615707397461,-92.1025390625,-92.1741714477539,-92.1652374267578,-92.0605010986328,-92.0763168334961,-92.2068328857422,-92.3474655151367,-92.3892593383789,-92.4083480834961,-92.4270935058594,-92.427978515625,-92.4110870361328,-92.3306579589844,-92.1104507446289,-92.0807113647461,-92.06884765625,-92.0991744995117,-92.141845703125,-92.1851119995117,-92.3065948486328,-92.4737319946289,-92.7088928222656,-92.88330078125,-93.0917510986328,-93.1922836303711,-93.30859375,-93.5599594116211,-93.6444320678711,-93.6651840209961,-93.8523483276367,-94.382568359375,-94.5853576660156,-94.7367172241211,-94.9966354370117,-95.2738800048828,-95.4474105834961,-95.8416519165039,-95.9592742919922,-96.0397033691406,-96.0130844116211,-95.7888641357422,-95.6957015991211,-95.6509704589844,-95.8731918334961,-95.9713363647461,-96.6396942138672,-96.8456573486328,-96.8807144165039,-96.8979949951172,-96.8975601196289,-96.8780288696289,-96.6793975830078,-96.5902786254883,-96.451171875,-96.4015655517578,-96.4331970214844,-96.6610336303711,-96.7711410522461,-96.8135223388672,-96.7698287963867,-96.75830078125,-96.6851043701172,-96.5501022338867,-96.3772964477539,-96.0612258911133,-95.8495101928711,-95.6382369995117,-95.1264114379883,-94.6161117553711,-94.2949676513672],[-115.55126953125,-115.475593566895,-115.470207214355,-115.506637573242,-115.623931884766,-116.213714599609,-116.329200744629,-116.285743713379,-116.073097229004,-115.856834411621,-115.810066223145,-115.912887573242,-116.109756469727,-116.183250427246,-116.252731323242,-116.233985900879,-116.016212463379,-115.944580078125,-115.9462890625,-115.984809875488,-116.076225280762,-116.22046661377,-116.467620849609,-116.999214172363,-117.016792297363,-117.013282775879,-117.026168823242,-117.044486999512,-117.107757568359,-117.153907775879,-117.233596801758,-117.346923828125,-117.492378234863,-117.732421875,-117.841407775879,-117.992965698242,-118.020217895508,-118.005424499512,-117.965423583984,-117.899513244629,-117.807624816895,-117.780464172363,-117.817924499512,-117.880813598633,-118.076416015625,-118.202789306641,-118.300582885742,-118.369873046875,-118.409126281738,-118.430999755859,-118.468170166016,-118.573684692383,-118.731544494629,-118.791404724121,-118.820755004883,-118.79955291748,-118.643898010254,-118.62451171875,-118.643402099609,-118.811576843262,-118.851127624512,-118.955459594727,-118.993942260742,-119.080711364746,-119.168212890625,-119.249267578125,-119.36792755127,-119.447799682617,-119.488815307617,-119.523735046387,-119.580368041992,-119.648933410645,-119.650772094727,-119.635841369629,-119.639640808105,-119.739646911621,-119.725196838379,-119.549705505371,-119.527153015137,-119.526123046875,-119.537750244141,-119.607963562012,-119.667137145996,-119.734817504883,-119.912887573242,-120.160537719727,-120.365379333496,-120.408882141113,-120.458297729492,-120.513824462891,-120.563278198242,-120.637161254883,-120.728660583496,-120.771583557129,-120.813385009766,-120.84839630127,-120.900100708008,-121.019287109375,-121.213485717773,-121.320167541504,-121.427978515625,-121.694534301758,-121.908210754395,-122.057418823242,-122.302734375,-122.400489807129,-122.533058166504,-122.591117858887,-122.640472412109,-122.645950317383,-122.607376098633,-122.546241760254,-122.548194885254,-122.608642578125,-122.609474182129,-122.587890625,-122.592727661133,-122.623931884766,-122.684661865234,-122.902778625488,-122.878273010254,-122.774017333984,-122.519386291504,-122.423042297363,-122.365379333496,-121.61376953125,-121.561141967773,-121.203903198242,-121.101997375488,-120.997604370117,-120.485359191895,-120.437599182129,-120.35765838623,-120.310943603516,-120.200347900391,-119.831146240234,-119.494827270508,-119.32398223877,-119.090179443359,-118.820022583008,-118.005180358887,-117.418601989746,-117.279098510742,-117.210784912109,-117.148971557617,-117.061386108398,-116.843551635742,-116.795303344727,-116.703620910645,-116.766265869141,-117.02978515625,-117.045753479004,-117.039741516113,-116.947166442871,-116.835304260254,-116.511337280273,-116.362602233887,-116.208892822266,-116.008003234863,-115.55126953125],[-89.833251953125,-90.0947265625,-90.228271484375,-90.9932174682617,-91.1472625732422,-91.1766128540039,-91.18505859375,-91.1826629638672,-91.1494598388672,-91.109130859375,-91.01904296875,-90.8425827026367,-90.6748504638672,-90.4227600097656,-90.1719284057617,-89.8389663696289,-89.719482421875,-89.6941909790039,-89.694580078125,-89.7120132446289,-89.7466812133789,-89.833251953125],[-104.558151245117,-104.711380004883,-105.015579223633,-105.215087890625,-105.379928588867,-105.556343078613,-105.695114135742,-105.747215270996,-105.84814453125,-105.883155822754,-106.066108703613,-106.035591125488,-105.862983703613,-105.587890625,-105.456108093262,-105.289649963379,-105.073875427246,-105.007225036621,-104.994293212891,-104.955322265625,-104.770217895508,-104.542236328125,-104.500778198242,-104.453712463379,-104.456985473633,-104.493354797363,-104.558151245117],[-95.484375,-95.2330551147461,-94.9599151611328,-94.6667938232422,-94.0147476196289,-93.5828628540039,-93.4710922241211,-93.3009796142578,-93.2107467651367,-93.1287155151367,-93.3391571044922,-93.5195770263672,-93.5439453125,-93.7401885986328,-93.836181640625,-94.4089889526367,-95.987060546875,-96.0561065673828,-96.2638702392578,-96.276611328125,-96.2391586303711,-96.194580078125,-96.1429748535156,-95.6839370727539,-95.484375],[-101.693550109863,-101.8310546875,-102.079833984375,-102.377838134766,-102.458206176758,-102.475051879883,-102.471534729004,-102.447708129883,-102.26318359375,-101.917869567871,-101.639404296875,-101.322021484375,-101.193214416504,-101.127586364746,-101.046241760254,-101.019577026367,-101.002052307129,-101.397651672363,-101.584571838379,-101.693550109863],[-113.832473754883,-114.105903625488,-114.287208557129,-114.608345031738,-114.980415344238,-115.029350280762,-114.890434265137,-114.789497375488,-114.726470947266,-114.606887817383,-114.330375671387,-114.296875,-114.302879333496,-114.279838562012,-114.180953979492,-114.087203979492,-113.897758483887,-113.768013000488,-113.721389770508,-113.696678161621,-113.617919921875,-113.619384765625,-113.725830078125,-113.832473754883],[-110.458061218262,-109.656883239746,-109.622268676758,-109.619041442871,-109.679397583008,-109.771774291992,-110.199508666992,-110.751121520996,-110.865623474121,-110.856544494629,-110.811622619629,-110.719390869141,-110.292190551758,-110.189208984375,-110.152732849121,-110.130859375,-110.117576599121,-110.116897583008,-110.139450073242,-110.198486328125,-110.371536254883,-110.682861328125,-110.893997192383,-111.060447692871,-111.226219177246,-111.951950073242,-112.176513671875,-112.372657775879,-112.643798828125,-112.925636291504,-113.046043395996,-113.164360046387,-113.197120666504,-113.208549499512,-113.188621520996,-113.137405395508,-113.120658874512,-113.167770385742,-113.189498901367,-113.271293640137,-113.283447265625,-113.282958984375,-113.269340515137,-113.215187072754,-113.187057495117,-113.021636962891,-112.804542541504,-112.304595947266,-111.206588745117,-110.873245239258,-110.727348327637,-110.458061218262],[-103.003372192383,-103.118217468262,-103.252243041992,-103.27099609375,-103.273590087891,-103.260055541992,-103.110450744629,-102.973289489746,-102.891799926758,-102.825538635254,-102.788284301758,-103.003372192383],[-109.815963745117,-109.640968322754,-109.580856323242,-109.504150390625,-109.46728515625,-109.362205505371,-109.342140197754,-109.336036682129,-109.352104187012,-109.390525817871,-109.484466552734,-109.708786010742,-110.021873474121,-110.29345703125,-110.418357849121,-110.755073547363,-110.840034484863,-111.026763916016,-111.169189453125,-111.22900390625,-111.300483703613,-111.43505859375,-111.517478942871,-111.759712219238,-112.131248474121,-112.557815551758,-112.999900817871,-113.172515869141,-113.223045349121,-113.292579650879,-113.281692504883,-113.149955749512,-112.855857849121,-112.640823364258,-112.214012145996,-111.708786010742,-111.519874572754,-111.400344848633,-111.071487426758,-110.877586364746,-110.618072509766,-110.407814025879,-110.140480041504,-109.940872192383,-109.815963745117],[-96.2044906616211,-95.9684524536133,-95.5611343383789,-95.4129409790039,-95.0312042236328,-94.9153823852539,-94.8877487182617,-94.8871612548828,-95.0138702392578,-95.2678680419922,-95.3292541503906,-95.1027374267578,-94.98779296875,-94.93603515625,-94.9342727661133,-95.0870132446289,-95.1991195678711,-95.3705062866211,-95.4515686035156,-95.6707992553711,-96.0115737915039,-96.4768524169922,-96.60302734375,-96.833984375,-96.9896469116211,-97.0404815673828,-97.0638198852539,-97.0519561767578,-97.0190963745117,-97.0933074951172,-97.4266586303711,-97.620849609375,-97.6483840942383,-97.6581573486328,-97.2266082763672,-97.0409164428711,-96.9583435058594,-96.9446792602539,-97.02734375,-97.3230438232422,-97.8190460205078,-97.8427276611328,-98.0495071411133,-98.0692901611328,-98.1143112182617,-98.2549285888672,-98.2756881713867,-98.3173370361328,-98.32373046875,-98.3156280517578,-98.0603485107422,-98.0959930419922,-98.2898941040039,-98.3408203125,-98.3326110839844,-98.2121124267578,-98.0428695678711,-97.5958938598633,-97.38232421875,-97.1693344116211,-96.935791015625,-96.5870590209961,-96.475341796875,-96.2652816772461,-96.2426223754883,-96.2565002441406,-96.2044906616211],[-103.426025390625,-103.191650390625,-102.914352416992,-102.652290344238,-102.638969421387,-102.637603759766,-102.648193359375,-102.682861328125,-102.730522155762,-102.595603942871,-102.580757141113,-102.5927734375,-102.576171875,-102.494575500488,-102.4248046875,-102.407318115234,-102.393165588379,-102.188575744629,-101.9736328125,-101.872604370117,-101.703659057617,-101.299018859863,-101.144584655762,-101.088470458984,-101.037155151367,-101.03369140625,-101.115913391113,-101.1474609375,-101.128120422363,-100.9169921875,-100.435501098633,-100.014747619629,-99.7818374633789,-99.6094207763672,-99.5821304321289,-99.631103515625,-99.68017578125,-99.8183135986328,-99.8478012084961,-99.7741165161133,-99.7686538696289,-99.7782211303711,-99.7513656616211,-99.5624542236328,-99.1315383911133,-99.0531234741211,-99.0045928955078,-98.9996109008789,-99.0611877441406,-99.1283645629883,-99.1664047241211,-99.34130859375,-99.6591339111328,-99.9559020996094,-100.274658203125,-100.586036682129,-100.680274963379,-100.757911682129,-100.778228759766,-100.8095703125,-100.826171875,-100.957611083984,-101.074119567871,-101.297996520996,-101.829490661621,-102.056983947754,-102.284469604492,-102.606979370117,-102.667671203613,-102.722702026367,-102.772071838379,-102.784324645996,-102.731346130371,-103.67724609375,-103.946578979492,-104.32421875,-104.512641906738,-104.763572692871,-104.879341125488,-104.985404968262,-104.995552062988,-104.909622192383,-104.820121765137,-104.72705078125,-104.213966369629,-103.764358520508,-103.570510864258,-103.482566833496,-103.587989807129,-104.020851135254,-103.928512573242,-103.562698364258,-103.37158203125,-103.408401489258,-103.518363952637,-104.008743286133,-104.185005187988,-104.194580078125,-104.154983520508,-103.875633239746,-103.887153625488,-104.007225036621,-104.112747192383,-104.151947021484,-104.394821166992,-104.736030578613,-104.817428588867,-104.8955078125,-104.97021484375,-104.969528198242,-104.893409729004,-104.735252380371,-104.746780395508,-104.9013671875,-105.308792114258,-105.53564453125,-105.570747375488,-105.580177307129,-105.571044921875,-105.514549255371,-105.435691833496,-105.3876953125,-104.847366333008,-103.964599609375,-103.706398010254,-103.426025390625],[-98.7916030883789,-98.7689514160156,-98.789794921875,-98.8406219482422,-98.8852081298828,-98.9452133178711,-99.2184524536133,-99.3017578125,-99.3062515258789,-99.3330078125,-99.515625,-99.8574676513672,-99.9999084472656,-100.056831359863,-100.092430114746,-100.126022338867,-100.120361328125,-100.078514099121,-100.053268432617,-99.8027877807617,-99.731201171875,-99.4248504638672,-99.1532211303711,-99.0166015625,-98.8946762084961,-98.8231964111328,-98.7916030883789],[-91.8855438232422,-91.7549819946289,-91.2724609375,-91.0539093017578,-90.6829147338867,-90.63671875,-90.6324691772461,-90.6430206298828,-90.5372619628906,-90.2176284790039,-89.8618621826172,-89.7978515625,-89.673828125,-89.5248031616211,-89.3290557861328,-89.235595703125,-89.1669006347656,-89.1382751464844,-89.1341781616211,-89.2043914794922,-89.1965789794922,-89.1546859741211,-89.1472702026367,-89.2176742553711,-89.1983413696289,-89.0192337036133,-88.8573226928711,-88.5375518798828,-88.3292465209961,-88.1998977661133,-88.1968231201172,-88.25537109375,-88.3807601928711,-88.6124954223633,-88.646240234375,-88.6634292602539,-88.6436462402344,-88.5248031616211,-88.4243698120117,-88.125244140625,-87.9600067138672,-87.6750030517578,-87.6455078125,-87.6302719116211,-87.6183547973633,-87.62548828125,-87.869140625,-87.9223098754883,-87.8606948852539,-87.6513671875,-87.3285140991211,-87.2020492553711,-87.076171875,-86.9772033691406,-87.0495147705078,-87.1442413330078,-87.2202682495117,-87.295166015625,-87.2428741455078,-86.9252395629883,-86.8610382080078,-86.6488265991211,-86.3369674682617,-86.2322235107422,-86.1804733276367,-86.0855484008789,-86.0070343017578,-85.948974609375,-85.8038558959961,-85.7509307861328,-85.6786117553711,-85.6478500366211,-85.5013656616211,-85.1755905151367,-85.0637664794922,-85.0421371459961,-85.1813430786133,-85.2898406982422,-86.0916519165039,-86.4505386352539,-86.6294403076172,-86.7208023071289,-86.9134750366211,-86.9571838378906,-87.0164566040039,-87.0803680419922,-87.2463836669922,-87.478759765625,-87.6173782348633,-87.8614807128906,-87.9223098754883,-87.9562530517578,-87.9607391357422,-87.9531707763672,-87.922607421875,-87.8167495727539,-87.8293991088867,-87.8783645629883,-88.0401840209961,-88.1040496826172,-88.163818359375,-88.1902313232422,-88.1666030883789,-88.189697265625,-88.2537612915039,-88.2278823852539,-88.0370101928711,-88.0031204223633,-87.9819793701172,-87.9736328125,-87.9828643798828,-88.0402374267578,-88.1475601196289,-88.2845764160156,-88.5806655883789,-88.7092742919922,-88.7416000366211,-88.7139663696289,-88.623046875,-88.6064453125,-88.6483917236328,-88.7329635620117,-88.791015625,-88.8224105834961,-88.9696273803711,-89.095703125,-89.4700241088867,-89.6552734375,-89.9262237548828,-89.995361328125,-90.037109375,-90.0763168334961,-90.0010299682617,-89.7572784423828,-89.6116714477539,-89.5068359375,-89.4898452758789,-89.5256805419922,-89.5794982910156,-89.651123046875,-89.873046875,-89.9651794433594,-90.0254364013672,-90.1360855102539,-90.2972183227539,-90.4590301513672,-90.62158203125,-90.6523971557617,-90.4691848754883,-90.4054183959961,-90.3579559326172,-90.3267593383789,-90.386962890625,-90.6144027709961,-90.9181137084961,-91.4096145629883,-91.899169921875,-92.3512649536133,-92.6782684326172,-92.8076171875,-92.8482360839844,-92.7255859375,-92.2967300415039,-91.8668899536133,-91.9349594116211,-92.7155303955078,-92.9725112915039,-93.109375,-93.2666015625,-93.3894500732422,-93.5520477294922,-93.6344223022461,-93.6233444213867,-93.5614242553711,-93.2083435058594,-93.1598587036133,-93.3364715576172,-93.9022445678711,-94.1145935058594,-94.1536102294922,-94.169677734375,-94.1627883911133,-93.9501953125,-93.2938919067383,-93.0684585571289,-92.8415985107422,-92.6836471557617,-92.5472183227539,-91.8675842285156,-91.3436508178711,-91.2999038696289,-91.692626953125,-92.2479476928711,-92.4845733642578,-92.6447296142578,-92.8219223022461,-93.0281295776367,-93.380859375,-93.5504379272461,-93.9331512451172,-94.0398406982422,-94.0933609008789,-94.109375,-94.040283203125,-93.939697265625,-93.9602584838867,-94.1102981567383,-94.2841262817383,-94.4048767089844,-94.8460388183594,-95.043701171875,-95.1031723022461,-95.3166046142578,-95.6570281982422,-95.7330093383789,-95.6628952026367,-95.5634765625,-95.3023452758789,-94.5196762084961,-94.4755401611328,-94.40185546875,-94.5808563232422,-94.9730529785156,-95.2969741821289,-95.552490234375,-95.7393569946289,-95.8575210571289,-95.9996643066406,-96.4627380371094,-96.5890655517578,-96.6067352294922,-96.6392059326172,-96.7732391357422,-95.781982421875,-95.3938522338867,-94.6458969116211,-94.61083984375,-94.5998077392578,-94.6126937866211,-94.6069869995117,-94.5826187133789,-94.304443359375,-94.2626037597656,-94.5901336669922,-95.1923828125,-95.4050827026367,-95.646240234375,-95.9040069580078,-96.0256805419922,-96.215087890625,-96.3083038330078,-96.368408203125,-96.3940887451172,-96.3853530883789,-96.3343734741211,-96.1121597290039,-96.0118637084961,-95.7473602294922,-95.549072265625,-95.6144485473633,-95.9010772705078,-96.1518020629883,-96.1328125,-95.9269561767578,-95.713623046875,-95.5052719116211,-95.225830078125,-95.0257339477539,-94.892578125,-94.7344741821289,-94.4854049682617,-93.9279251098633,-94.0287170410156,-94.2021484375,-94.5962905883789,-94.7884750366211,-95.1958465576172,-95.5147476196289,-95.50927734375,-95.269775390625,-94.9805145263672,-94.5194396972656,-94.21630859375,-93.8259735107422,-93.4437484741211,-93.3451156616211,-93.2867202758789,-93.2356491088867,-93.2354049682617,-93.2859344482422,-93.4065475463867,-93.8944320678711,-94.1103515625,-94.1944351196289,-94.2186584472656,-94.2314987182617,-94.2329559326172,-94.2201156616211,-94.1793441772461,-94.0597152709961,-93.6048812866211,-93.332763671875,-93.03466796875,-92.41259765625,-92.2117691040039,-91.9978561401367,-91.8855438232422],[-69.4888687133789,-68.6732406616211,-68.4090270996094,-68.1068801879883,-67.9246139526367,-67.6244583129883,-67.4056625366211,-66.5916519165039,-66.4225616455078,-66.4248046875,-66.6003875732422,-66.8363342285156,-68.3575744628906,-68.4693374633789,-68.1728515625,-67.7358856201172,-67.3970718383789,-66.9977111816406,-66.86572265625,-66.6118621826172,-66.1204605102539,-65.7274398803711,-65.5496139526367,-65.3999938964844,-65.2990264892578,-65.24658203125,-65.1623992919922,-65.1131820678711,-64.98388671875,-64.9048843383789,-64.7767562866211,-64.634765625,-64.5040054321289,-64.4333953857422,-64.1342239379883,-63.9835929870605,-63.6410179138184,-63.4987335205078,-63.4730453491211,-63.5640640258789,-63.6206016540527,-63.6425285339355,-63.5926742553711,-63.3853988647461,-63.0853500366211,-63.0870628356934,-63.2508316040039,-63.2467803955078,-62.4751930236816,-61.6971664428711,-61.47705078125,-61.3924789428711,-61.302490234375,-61.2072257995605,-61.2735366821289,-61.6153831481934,-61.9686546325684,-62.1767082214355,-62.4964828491211,-63.59228515625,-64.1279296875,-64.435791015625,-64.5740203857422,-65.2261657714844,-65.399169921875,-65.4954071044922,-65.7010726928711,-66.0047378540039,-66.625732421875,-66.7650375366211,-66.8005905151367,-66.8611297607422,-66.9140625,-68.6885299682617,-68.72119140625,-68.5425796508789,-68.3176727294922,-65.7356948852539,-65.239990234375,-64.7800750732422,-64.832763671875,-65.4839782714844,-66.3128433227539,-66.7268600463867,-67.7743606567383,-68.6304702758789,-68.9593734741211,-69.4001007080078,-69.5506820678711,-69.7337875366211,-69.9493179321289,-70.1435089111328,-70.4026412963867,-70.638671875,-70.7126007080078,-70.6678237915039,-70.2127914428711,-70.264892578125,-71.1002960205078,-71.4700698852539,-71.6608428955078,-71.7958984375,-71.9276351928711,-72.0559539794922,-72.06298828125,-71.9486770629883,-71.6161117553711,-70.8770523071289,-70.7584991455078,-70.5684051513672,-70.55908203125,-70.7575225830078,-71.3558120727539,-71.2776336669922,-71.1063461303711,-71.1101608276367,-71.298583984375,-71.3878402709961,-71.9645538330078,-72.2155303955078,-72.4365234375,-73.4481430053711,-73.8050765991211,-74.1442337036133,-74.3944854736328,-74.6602020263672,-74.5407257080078,-74.051025390625,-73.64208984375,-73.4724578857422,-73.4059066772461,-73.2293930053711,-73.2011260986328,-73.2401351928711,-73.2935562133789,-73.3615264892578,-73.466064453125,-73.865966796875,-74.015380859375,-74.1886749267578,-74.406005859375,-74.7979507446289,-75.2594680786133,-75.50341796875,-75.7738265991211,-76.06689453125,-76.3760757446289,-76.8988265991211,-76.8550796508789,-76.6708984375,-76.2957077026367,-76.1163558959961,-75.947509765625,-75.6027374267578,-75.3536605834961,-75.0936050415039,-74.7272491455078,-74.481201171875,-74.5323257446289,-74.6409149169922,-75.233154296875,-75.5146484375,-75.6389617919922,-75.9118118286133,-76.1575622558594,-76.38037109375,-76.5314407348633,-76.7711410522461,-77.3980407714844,-77.729248046875,-77.9737777709961,-78.2579116821289,-78.5816421508789,-78.5589828491211,-78.4217834472656,-78.2219696044922,-78.0368194580078,-77.8827590942383,-77.6982421875,-77.5103988647461,-76.8248062133789,-76.5241241455078,-76.255859375,-76.0773468017578,-75.9526901245117,-75.7950668334961,-75.3998489379883,-75.0985412597656,-74.618408203125,-74.486328125,-74.43310546875,-74.5350570678711,-74.5465850830078,-74.8786163330078,-75.3965835571289,-75.9658203125,-76.3734893798828,-76.4161148071289,-76.1365203857422,-75.4883804321289,-75.2372055053711,-75.1934585571289,-75.5506820678711,-75.865966796875,-75.9696273803711,-76.0775375366211,-76.3555679321289,-76.7081069946289,-76.9740295410156,-77.4559555053711,-78.0125961303711,-78.056396484375,-78.0841293334961,-78.0810546875,-78.0471649169922,-78.076171875,-78.16796875,-78.2837448120117,-78.4932098388672,-78.70849609375,-78.8695297241211,-79.1375961303711,-79.9063949584961,-80.2816848754883,-80.5730438232422,-80.8746109008789,-81.3768539428711,-81.519287109375,-81.6590881347656,-81.65380859375,-81.5035629272461,-81.378173828125,-81.2777328491211,-81.3013687133789,-81.52294921875,-81.767333984375,-82.0567855834961,-82.0660171508789,-81.9678192138672,-81.8402328491211,-81.75634765625,-81.5344772338867,-81.2774429321289,-81.1171875,-80.7981948852539,-80.6725616455078,-80.2742156982422,-80.2186965942383,-79.9237289428711,-79.4972686767578,-79.3408737182617,-79.2811050415039,-79.2738265991211,-79.3189468383789,-79.2207565307617,-78.9792022705078,-78.7918014526367,-78.6585388183594,-78.4559555053711,-78.3700180053711,-78.2888641357422,-78.1650848388672,-77.9987335205078,-77.9832992553711,-78.1187057495117,-78.2843322753906,-78.9342727661133,-79.1307144165039,-79.285888671875,-79.5110397338867,-79.9535675048828,-80.1868133544922,-80.6902847290039,-80.7997055053711,-80.9629364013672,-80.9966812133789,-80.9551696777344,-80.9012222290039,-80.8348159790039,-80.8323745727539,-80.9745101928711,-81.0743637084961,-81.1707077026367,-81.3647918701172,-81.4744644165039,-81.5919952392578,-81.7173767089844,-81.8229446411133,-82.0341796875,-82.1137237548828,-82.2179183959961,-82.3111343383789,-82.3934631347656,-82.5298309326172,-82.493408203125,-82.3569793701172,-82.261962890625,-82.2083511352539,-82.233154296875,-83.3889694213867,-83.8856887817383,-83.986328125,-84.2237777709961,-84.2753372192383,-85.1412582397461,-85.3436050415039,-85.6805648803711,-86.1158142089844,-86.2961883544922,-86.3668441772461,-86.41943359375,-86.4537124633789,-86.5619201660156,-86.6802215576172,-86.9776840209961,-87.3542022705078,-87.4897918701172,-87.49755859375,-88.1043472290039,-88.39599609375,-88.4816360473633,-88.495849609375,-88.6141128540039,-88.5625457763672,-88.5457992553711,-88.8037109375,-89.36962890625,-89.570068359375,-89.5443344116211,-89.499755859375,-88.7708969116211,-88.5562057495117,-88.3981475830078,-88.14794921875,-87.8284225463867,-87.6104965209961,-87.3617172241211,-87.064453125,-86.8521957397461,-86.812255859375,-86.873779296875,-87.1008834838867,-87.1824188232422,-87.265380859375,-87.4296875,-87.5891571044922,-87.6814498901367,-87.7801742553711,-87.9379425048828,-88.0946807861328,-88.0169906616211,-87.7571258544922,-87.4967803955078,-87.2360382080078,-87.0179672241211,-86.7550735473633,-86.3851013183594,-86.1729965209961,-85.9066390991211,-85.731201171875,-85.5884780883789,-84.9508819580078,-84.7386703491211,-84.4870071411133,-83.9735794067383,-83.7212905883789,-83.6080551147461,-83.5498046875,-83.4773406982422,-83.2502899169922,-82.9027328491211,-82.7103500366211,-82.6647033691406,-82.6263198852539,-82.5953140258789,-82.7035675048828,-83.3037567138672,-83.42822265625,-83.7793960571289,-83.9281768798828,-84.1678237915039,-84.48583984375,-84.8605422973633,-85.087890625,-85.2893600463867,-85.2920455932617,-85.5475616455078,-85.2653274536133,-85.031494140625,-84.6154327392578,-84.524169921875,-84.2227096557617,-84.3881378173828,-84.5499954223633,-84.9103469848633,-84.783203125,-85.0243225097656,-85.2701644897461,-85.4189910888672,-85.5859375,-86.2177734375,-86.0625991821289,-85.9200744628906,-86.0709533691406,-86.4270477294922,-86.693603515625,-86.9132385253906,-87.33935546875,-87.5517578125,-87.4911117553711,-87.4913101196289,-87.3612823486328,-87.164306640625,-86.9529342651367,-86.8079147338867,-86.2418975830078,-85.6910171508789,-85.2296905517578,-85.0037612915039,-84.7872543334961,-83.9079132080078,-83.5470275878906,-83.3887176513672,-83.2714309692383,-83.1474075317383,-82.9897994995117,-82.4417953491211,-82.2906723022461,-82.1510772705078,-81.9811019897461,-81.7808151245117,-81.7500991821289,-81.8891143798828,-82.0283203125,-82.2374038696289,-82.4387741088867,-82.6440963745117,-83.0585403442383,-83.7786178588867,-84.1457977294922,-84.3161087036133,-84.4120101928711,-84.495849609375,-84.5677719116211,-84.5302734375,-84.3835906982422,-84.2566375732422,-84.0530242919922,-83.8246078491211,-83.5758819580078,-83.6620101928711,-83.9781265258789,-84.1973571777344,-84.3810501098633,-84.5224075317383,-84.83642578125,-85.0897903442383,-85.2685012817383,-85.4569396972656,-86.031494140625,-86.1466293334961,-86.4207534790039,-86.4943313598633,-86.614501953125,-86.49853515625,-86.3071823120117,-85.1596145629883,-84.6754379272461,-84.0565414428711,-83.7236328125,-83.34375,-83.0042953491211,-82.677490234375,-82.3769989013672,-82.0487823486328,-81.855712890625,-81.6883773803711,-81.4630355834961,-81.0380859375,-80.6678161621094,-80.4759216308594,-80.2705993652344,-80.1244659423828,-80.2874526977539,-80.7140121459961,-81.0101547241211,-81.1790542602539,-81.3586959838867,-81.6442337036133,-81.8602523803711,-82.3323669433594,-82.6812973022461,-82.9611282348633,-82.9870071411133,-82.7848129272461,-82.5361328125,-80.9796371459961,-80.0510711669922,-79.6743698120117,-79.6293411254883,-78.3861846923828,-77.5071258544922,-77.1691360473633,-76.8629837036133,-76.850341796875,-77.1185531616211,-77.3894500732422,-78.0038070678711,-78.5509796142578,-78.7162094116211,-78.6819381713867,-78.6292953491211,-78.4639663696289,-78.2868118286133,-77.5360336303711,-77.0307159423828,-76.8851089477539,-77.9723587036133,-78.3521499633789,-78.73388671875,-78.9315414428711,-79.0724563598633,-79.1983413696289,-79.3091812133789,-79.4021530151367,-79.4772491455078,-79.54541015625,-79.6066436767578,-79.7613296508789,-80.133544921875,-81.0070343017578,-81.3009796142578,-81.5526809692383,-82.3682174682617,-82.613037109375,-82.8843231201172,-82.768310546875,-82.3367691040039,-82.2223663330078,-82.4984436035156,-82.7799835205078,-83.4014205932617,-83.6471176147461,-83.8853530883789,-84.0762710571289,-84.2197799682617,-84.4178237915039,-85.1458511352539,-85.3074264526367,-85.7262268066406,-86.09716796875,-86.2503433227539,-86.5315933227539,-86.6154327392578,-86.6030731201172,-86.4404754638672,-86.2520980834961,-85.6393127441406,-85.2462844848633,-84.679931640625,-83.3492202758789,-83.288818359375,-84.6354522705078,-85.7808609008789,-85.966796875,-86.2334442138672,-87.0802764892578,-87.3298797607422,-87.711669921875,-88.003662109375,-88.2319793701172,-88.6250991821289,-88.9214324951172,-89.0616683959961,-89.1445846557617,-89.2117614746094,-89.2632827758789,-89.1669006347656,-88.4130859375,-87.388671875,-86.9290084838867,-86.4767608642578,-85.8095703125,-85.0832977294922,-84.9412078857422,-85.206298828125,-85.4024963378906,-85.8750457763672,-86.6227569580078,-87.2751007080078,-88.8868179321289,-89.3983840942383,-89.623046875,-89.7922821044922,-89.98095703125,-89.9473114013672,-89.5633773803711,-89.2625503540039,-89.2086944580078,-89.6356964111328,-89.6736831665039,-89.427001953125,-88.8922805786133,-88.6219253540039,-88.1265182495117,-87.61669921875,-87.5970153808594,-88.1013641357422,-88.4790573120117,-88.9783706665039,-90.3035125732422,-90.4163131713867,-90.6090316772461,-90.5537567138672,-89.84521484375,-89.8216781616211,-90.3308563232422,-90.4803695678711,-90.6263198852539,-90.833740234375,-91.1027374267578,-91.2923812866211,-91.4027786254883,-91.68408203125,-91.6475601196289,-91.423828125,-91.219482421875,-90.9419403076172,-90.4901809692383,-90.1630401611328,-89.6333541870117,-89.3810043334961,-89.1563491821289,-88.875244140625,-88.5668411254883,-88.0631866455078,-87.638916015625,-87.4043960571289,-87.2181625366211,-87.0182113647461,-86.9992218017578,-86.8340301513672,-86.6268005371094,-86.3775329589844,-86.1583480834961,-85.8748092651367,-85.6456527709961,-85.5379867553711,-85.4031753540039,-85.0448226928711,-85.05224609375,-85.1692352294922,-85.3105926513672,-86.5806121826172,-86.6156234741211,-86.1875991821289,-85.9200134277344,-85.79443359375,-85.4808654785156,-85.2759780883789,-84.8968276977539,-84.7447280883789,-84.5533676147461,-84.3681182861328,-83.8236312866211,-83.5906753540039,-83.1756820678711,-83.0101547241211,-82.7742156982422,-82.6336898803711,-82.3560028076172,-82.3274383544922,-82.6570816040039,-82.7474594116211,-82.7085876464844,-82.6383743286133,-82.5369110107422,-82.2765579223633,-81.58447265625,-80.5499038696289,-80.1533660888672,-79.9086456298828,-79.685546875,-79.4656219482422,-79.4248580932617,-79.6294937133789,-80.1298294067383,-81.46826171875,-81.9976119995117,-82.253662109375,-82.4475631713867,-82.4513626098633,-82.2688980102539,-82.0232467651367,-81.7177734375,-81.68115234375,-81.9585952758789,-82.1225128173828,-82.1168441772461,-81.7853546142578,-81.5796890258789,-81.1888656616211,-80.8625030517578,-80.8096694946289,-81.1466293334961,-81.1780776977539,-81.128173828125,-81.0101547241211,-80.6571273803711,-80.0757751464844,-79.0350570678711,-78.748779296875,-78.7918014526367,-79.2072219848633,-79.6419906616211,-79.8337936401367,-79.9743194580078,-80.1411666870117,-80.1549301147461,-79.8863296508789,-79.1805648803711,-78.5249481201172,-77.9686508178711,-77.6180648803711,-77.4795913696289,-77.2258758544922,-76.4209976196289,-76.3355484008789,-76.2440414428711,-76.146484375,-76.0093765258789,-75.7443389892578,-75.5656280517578,-75.6428680419922,-76.0869598388672,-76.1877975463867,-76.4099578857422,-76.908447265625,-77.0412063598633,-77.1249084472656,-75.7449188232422,-74.4141616821289,-74.19775390625,-74.0558624267578,-73.91650390625,-73.703125,-73.2720260620117,-72.65869140625,-72.7759246826172,-73.2346649169922,-73.44189453125,-73.4407196044922,-73.40380859375,-73.3311538696289,-72.8116683959961,-72.0692367553711,-71.9832000732422,-71.4059600830078,-71.1320266723633,-70.9403839111328,-70.9330139160156,-71.1983413696289,-71.4023971557617,-71.4235305786133,-71.0848083496094,-70.8705520629883,-69.9699172973633,-69.86767578125,-69.7821273803711,-69.5693817138672,-69.4888687133789],[9.52402305603027,9.55439472198486,9.62587833404541,9.60908222198486,9.52753925323486,9.48427772521973,9.4794921875,9.48769569396973,9.50234413146973,9.58027362823486,9.61992168426514,9.74501895904541,9.84531211853027,9.86464881896973,9.87773418426514,9.99687480926514,10.1334953308105,10.1797857284546,10.3494138717651,10.4149417877197,10.45458984375,10.4528331756592,10.4060554504395,10.39794921875,10.4382820129395,10.4424810409546,10.4306640625,10.3630857467651,10.2722654342651,10.1955080032349,10.1374998092651,10.0870122909546,10.0612306594849,10.0382814407349,10.0456056594849,10.0819339752197,10.1096677780151,10.1298828125,10.1452150344849,10.1283197402954,10.08056640625,10.041015625,9.97167873382568,9.93925762176514,9.88447189331055,9.78779220581055,9.63945388793945,9.57958984375,9.52871036529541,9.48105430603027,9.44062519073486,9.42763710021973,9.39931583404541,9.30439472198486,9.26015663146973,9.259765625,9.25107383728027,9.20341777801514,9.07099533081055,9.02236270904541,9.00302791595459,8.99892520904541,9.01914119720459,9.04667949676514,9.02373123168945,8.95371150970459,8.904296875,8.88515663146973,8.77802753448486,8.82675743103027,8.81855487823486,8.64169883728027,8.5654296875,8.45839881896973,8.4384765625,8.44296836853027,8.43681621551514,8.42255878448486,8.37070369720459,8.29853439331055,8.23193359375,8.095703125,8.08154296875,8.12724590301514,8.12519550323486,8.01425838470459,7.99316358566284,7.85234308242798,7.78788995742798,7.59257841110229,7.53857469558716,7.45156240463257,7.32792997360229,7.12900400161743,7.05576181411743,7.02109336853027,7.00390625,6.95371103286743,6.89726543426514,6.85800790786743,6.80566453933716,6.77207040786743,6.81679677963257,6.7841796875,6.76738262176514,6.77607440948486,6.75810527801514,6.57822227478027,6.42890596389771,6.32187509536743,6.23466730117798,6.22421884536743,6.22958993911743,6.27294969558716,6.19941425323486,6.08662128448486,6.00664091110229,5.97148418426514,5.97001934051514,6.0361328125,6.09589862823486,6.11591815948486,6.12324237823486,6.06025362014771,6.06796884536743,6.10703086853027,6.12968730926514,6.16074228286743,6.28515625,6.41015625,6.42900419235229,6.43857431411743,6.45625019073486,6.62480449676514,6.66689491271973,6.68808603286743,6.82070350646973,6.95205068588257,6.97851514816284,7.00058555603027,7.00058555603027,6.98408222198486,6.92148447036743,6.900390625,6.96835899353027,7.05341815948486,7.13603544235229,7.16923809051514,7.16747999191284,7.203125,7.26572227478027,7.34316349029541,7.42001962661743,7.46738243103027,7.49492216110229,7.61562490463257,7.69804668426514,7.92705106735229,8.09375,8.1982421875,8.32783222198486,8.41474628448486,8.43007850646973,8.45400333404541,8.47763633728027,8.55947303771973,8.57050800323486,8.56709003448486,8.55234336853027,8.45175838470459,8.41328144073486,8.40341854095459,8.43574237823486,8.50986289978027,8.57265663146973,8.61787128448486,8.72832012176514,8.75478553771973,8.77011775970459,8.79306602478027,8.83115291595459,8.8740234375,8.88115215301514,9.12753963470459,9.18281173706055,9.35000038146973,9.52402305603027],[-67.5751953125,-67.6114273071289,-67.6995086669922,-67.83154296875,-67.846435546875,-67.8497085571289,-67.8340835571289,-67.7620620727539,-67.5448303222656,-67.5172882080078,-67.5098114013672,-67.5452651977539,-67.5751953125],[-67.2888641357422,-67.3252944946289,-67.3522415161133,-67.3933639526367,-67.5599594116211,-67.5634765625,-67.546142578125,-67.5127944946289,-67.4488296508789,-67.3973617553711,-67.3740692138672,-67.3505859375,-67.3104476928711,-67.262451171875,-67.2672805786133,-67.2888641357422],[-66.4721145629883,-66.5517120361328,-66.6111297607422,-66.6301727294922,-66.6366195678711,-66.624755859375,-66.5997085571289,-66.5415573120117,-66.5231475830078,-66.435791015625,-66.4721145629883],[-67.0799331665039,-67.1094741821289,-67.1725616455078,-67.2574157714844,-67.3396987915039,-67.3992691040039,-67.4293975830078,-67.4432601928711,-67.4634780883789,-67.4947280883789,-67.5352478027344,-67.585205078125,-67.6914596557617,-67.7369613647461,-67.7677688598633,-68.0700225830078,-68.0995101928711,-68.1351089477539,-68.17431640625,-68.3013687133789,-68.10693359375,-67.8741226196289,-67.424560546875,-67.2452621459961,-67.1073226928711,-67.0854949951172,-67.0799331665039],[-69.7029724121094,-68.9007797241211,-68.6535186767578,-68.4580078125,-68.3998031616211,-68.5980987548828,-68.61328125,-68.5855484008789,-68.3817367553711,-68.330078125,-68.2826614379883,-68.32275390625,-68.3265609741211,-68.305419921875,-68.152587890625,-68.0898895263672,-68.0582962036133,-68.0455551147461,-68.04833984375,-68.0826644897461,-68.1570816040039,-68.2296371459961,-68.2933578491211,-68.3380355834961,-68.4666976928711,-68.5941925048828,-68.6935577392578,-68.7850112915039,-68.8670425415039,-68.8961486816406,-68.9312973022461,-68.9320755004883,-68.8889694213867,-68.8900833129883,-68.9126434326172,-69.0082015991211,-69.0468292236328,-69.1507797241211,-69.192626953125,-69.2970733642578,-69.3561553955078,-69.3592224121094,-69.2990264892578,-69.1808624267578,-69.2408142089844,-69.4118194580078,-69.4557189941406,-69.5086898803711,-69.6102523803711,-69.6458969116211,-69.6562957763672,-69.6573715209961,-69.6798324584961,-69.8240280151367,-69.8537139892578,-69.8657760620117,-69.8867721557617,-69.9797821044922,-69.9879837036133,-69.946533203125,-69.9208526611328,-69.8844223022461,-69.7029724121094],[-70.9916000366211,-70.9451217651367,-70.9279251098633,-70.8048324584961,-70.7493209838867,-70.6152801513672,-70.5347595214844,-70.4175262451172,-70.2830581665039,-70.2978515625,-70.4041519165039,-70.4755859375,-70.54345703125,-70.5387268066406,-70.5510711669922,-70.5974655151367,-70.6409149169922,-70.7109909057617,-70.7444381713867,-70.8154754638672,-70.9398422241211,-70.9645080566406,-70.96728515625,-70.99072265625,-71.120361328125,-71.2033233642578,-71.2736282348633,-71.29931640625,-71.3253402709961,-71.3885803222656,-71.4066467285156,-71.4269027709961,-71.4372100830078,-71.4105529785156,-71.374267578125,-71.1970748901367,-71.088623046875,-70.9916000366211],[-71.3904724121094,-71.1687469482422,-71.0219268798828,-71.0228576660156,-71.0048828125,-71.0280227661133,-71.0829620361328,-71.1175308227539,-71.1432647705078,-71.3046417236328,-71.4732894897461,-71.55810546875,-71.6706008911133,-71.76123046875,-71.8175811767578,-71.9485397338867,-71.9723587036133,-72.091552734375,-72.21044921875,-72.1460418701172,-72.0689468383789,-71.9964828491211,-71.7051315307617,-71.5541458129883,-71.3904724121094],[-72.9232482910156,-72.8962860107422,-72.8822250366211,-72.8093719482422,-72.6855010986328,-72.4822769165039,-72.459228515625,-72.3729019165039,-72.3066940307617,-72.2054214477539,-72.3062438964844,-72.365966796875,-72.369140625,-72.4085464477539,-72.4705047607422,-72.5628890991211,-72.6765594482422,-72.7886199951172,-72.8403854370117,-72.8709945678711,-72.9072799682617,-72.9460906982422,-72.9585952758789,-72.8817367553711,-72.7816925048828,-72.7637710571289,-72.8717269897461,-72.9361343383789,-72.9842224121094,-73.0394515991211,-73.0730438232422,-73.0855407714844,-73.0708541870117,-73.0807647705078,-73.1199722290039,-73.2106475830078,-73.3047409057617,-73.3121566772461,-73.2928695678711,-73.294921875,-73.3143539428711,-73.3248062133789,-73.360107421875,-73.470947265625,-73.5816421508789,-73.6415023803711,-73.845458984375,-73.6865234375,-73.4470748901367,-73.3658676147461,-73.099365234375,-73.1153335571289,-73.1108932495117,-73.0743179321289,-73.0536117553711,-73.0220718383789,-72.970947265625,-72.947265625,-72.9232482910156],[-74.3857421875,-74.3693313598633,-74.3299789428711,-74.2746124267578,-74.0659637451172,-73.8791961669922,-73.7817840576172,-73.6540069580078,-73.5492630004883,-73.5045394897461,-73.4505844116211,-73.3103561401367,-73.302490234375,-73.1433639526367,-73.1352081298828,-73.2257308959961,-73.4093780517578,-73.5010223388672,-73.5672836303711,-73.5828094482422,-73.595947265625,-73.6170883178711,-73.7935104370117,-73.866943359375,-73.9939956665039,-74.1385726928711,-74.2363739013672,-74.2702102661133,-74.4144058227539,-74.5583038330078,-74.6199188232422,-74.7115249633789,-74.7120132446289,-74.6699676513672,-74.571533203125,-74.4745559692383,-74.4222640991211,-74.3857421875],[-68.6299285888672,-68.6316909790039,-68.6334533691406,-68.6350631713867,-68.6366729736328,-68.6382293701172,-68.6397933959961,-68.6475143432617,-68.6532211303711,-68.8038558959961,-68.8435516357422,-69.0816421508789,-69.4862823486328,-69.5875549316406,-69.7234420776367,-69.7718200683594,-69.8994674682617,-70.030517578125,-70.1380844116211,-70.23779296875,-70.2590789794922,-70.28173828125,-70.4971694946289,-70.7351531982422,-70.9247055053711,-71.229248046875,-71.44091796875,-71.83154296875,-71.9015655517578,-71.927734375,-71.906982421875,-71.8234329223633,-71.8001403808594,-71.7158203125,-71.6062927246094,-71.57275390625,-71.5003890991211,-71.3934097290039,-71.355224609375,-71.1588363647461,-71.0799331665039,-70.9664611816406,-70.9461898803711,-70.9281768798828,-70.8982467651367,-70.7972640991211,-70.6988296508789,-70.6875457763672,-70.7011260986328,-70.572998046875,-70.4179153442383,-70.3109817504883,-70.2976608276367,-70.4683151245117,-70.5399856567383,-70.6361389160156,-70.7598648071289,-70.8630828857422,-70.8567352294922,-70.8677291870117,-70.6444854736328,-70.6956100463867,-70.6187515258789,-70.5313034057617,-70.4431610107422,-70.3797378540039,-70.4605484008789,-70.6298294067383,-70.5353012084961,-70.3799819946289,-70.24609375,-70.2433624267578,-70.1689910888672,-69.9901351928711,-69.8669967651367,-69.80908203125,-69.7418441772461,-69.6216812133789,-69.4192886352539,-69.3647918701172,-69.3250961303711,-69.3224563598633,-69.3120651245117,-69.253173828125,-69.169189453125,-69.1278839111328,-69.0772476196289,-69.0452117919922,-69.0443344116211,-69.1956558227539,-69.9881362915039,-70.0855941772461,-70.151123046875,-70.1488265991211,-70.0911178588867,-69.94970703125,-69.6897430419922,-69.3899383544922,-69.3524398803711,-69.35595703125,-69.3935546875,-69.5125503540039,-69.6370086669922,-69.755615234375,-69.8741226196289,-70.0903854370117,-70.2128448486328,-70.3292999267578,-70.4156723022461,-70.4602508544922,-70.4599609375,-70.443359375,-70.3906707763672,-70.3200225830078,-70.25634765625,-70.1964874267578,-70.160888671875,-70.130615234375,-70.1395568847656,-70.1627426147461,-70.2591781616211,-70.29736328125,-70.380126953125,-70.3349075317383,-70.1896438598633,-70.0882339477539,-69.9935531616211,-69.9354476928711,-69.8832015991211,-69.7635726928711,-69.6632766723633,-69.5718765258789,-69.4983901977539,-69.4140625,-69.1670455932617,-69.0799331665039,-68.789794921875,-68.7575225830078,-68.6592254638672,-68.6299285888672],[-74.1421890258789,-74.1720733642578,-74.2831115722656,-74.3386688232422,-74.4236297607422,-74.4371109008789,-74.4753952026367,-74.4507827758789,-74.3621063232422,-74.32568359375,-74.2770538330078,-74.1333999633789,-74.1154251098633,-74.118896484375,-74.1421890258789],[-74.8229446411133,-74.7801284790039,-74.74951171875,-74.6474609375,-74.5368194580078,-74.5318374633789,-74.6659698486328,-74.6944808959961,-74.851806640625,-74.917724609375,-75.0171356201172,-75.0506820678711,-75.1053695678711,-75.0081024169922,-74.9151916503906,-74.90966796875,-74.8229446411133],[-74.5586471557617,-74.5608825683594,-74.5925750732422,-74.620361328125,-74.6907196044922,-74.7309112548828,-74.79736328125,-74.8533172607422,-74.9366683959961,-75.04736328125,-75.1462860107422,-75.1924362182617,-75.2891159057617,-75.300048828125,-75.2384719848633,-75.2100067138672,-75.1536636352539,-75.0403289794922,-74.8814468383789,-74.7366638183594,-74.611572265625,-74.5705108642578,-74.5586471557617],[-75.302001953125,-75.3304672241211,-75.411376953125,-75.4385223388672,-75.45263671875,-75.4773941040039,-75.4426803588867,-75.4197769165039,-75.4276351928711,-75.3037109375,-75.1561508178711,-75.1153335571289,-75.1604461669922,-75.2034149169922,-75.2923278808594,-75.302001953125],[-75.0547866821289,-75.2503967285156,-75.307861328125,-75.4491195678711,-75.412109375,-75.3978500366211,-75.376708984375,-75.3688507080078,-75.32666015625,-75.2096633911133,-75.12255859375,-75.0042495727539,-74.8759765625,-74.8385772705078,-74.96337890625,-75.0547866821289],[-75.106689453125,-75.1150894165039,-75.2626953125,-75.3899383544922,-75.506103515625,-75.5803756713867,-75.6411590576172,-75.5728530883789,-75.4876480102539,-75.5145492553711,-75.5401382446289,-75.576171875,-75.6378402709961,-75.619140625,-75.5831069946289,-75.5352554321289,-75.490478515625,-75.2972640991211,-75.2361831665039,-75.1186065673828,-75.106689453125],[-74.4761734008789,-74.466796875,-74.4835968017578,-74.5220718383789,-74.5157699584961,-74.4708480834961,-74.4588394165039,-74.4719696044922,-74.49609375,-74.5425796508789,-74.56982421875,-74.5947265625,-74.703369140625,-74.7629928588867,-74.8108367919922,-74.82470703125,-74.8219223022461,-74.8804244995117,-74.8822250366211,-74.8593215942383,-74.81201171875,-74.8048324584961,-74.781005859375,-74.72705078125,-74.7188491821289,-74.7238235473633,-74.7438430786133,-74.9601135253906,-74.9813003540039,-74.9908218383789,-74.9934997558594,-75.0315399169922,-75.0660095214844,-75.1669387817383,-75.3000946044922,-75.451171875,-75.5498046875,-75.5701141357422,-75.520751953125,-75.3370590209961,-75.3058624267578,-75.3642120361328,-75.4288558959961,-75.4674758911133,-75.4331588745117,-75.32666015625,-75.2696304321289,-75.2168426513672,-75.0860366821289,-75.0937042236328,-75.2101593017578,-75.1842269897461,-75.037109375,-74.94921875,-74.9452133178711,-74.9807662963867,-74.9695281982422,-74.896240234375,-74.79345703125,-74.7466812133789,-74.6515655517578,-74.5666046142578,-74.5460891723633,-74.5306625366211,-74.4761734008789],[-75.51025390625,-75.6228485107422,-75.6509323120117,-75.5184555053711,-75.509033203125,-75.5535202026367,-75.5714874267578,-75.5606994628906,-75.3914108276367,-75.33837890625,-75.2754898071289,-75.155517578125,-75.1584930419922,-75.22509765625,-75.4339828491211,-75.51025390625],[-74.5672836303711,-74.5862808227539,-74.7095718383789,-74.9230499267578,-75.0128402709961,-75.0521469116211,-75.0789108276367,-75.1319274902344,-75.1584930419922,-75.212890625,-75.23388671875,-75.2472686767578,-75.1982955932617,-74.97509765625,-74.8956527709961,-74.8274383544922,-74.84619140625,-74.8052215576172,-74.7293014526367,-74.7152328491211,-74.702392578125,-74.6643600463867,-74.6151351928711,-74.6024398803711,-74.6001434326172,-74.6182098388672,-74.5672836303711],[-75.1122055053711,-75.1858444213867,-75.1943359375,-75.2610321044922,-75.203125,-75.08984375,-75.0039520263672,-74.9264678955078,-74.916015625,-75.05126953125,-75.08447265625,-75.1122055053711],[-74.3128890991211,-74.3684616088867,-74.4655303955078,-74.5616226196289,-74.677734375,-74.6898498535156,-74.6464309692383,-74.5584030151367,-74.4946823120117,-74.5023422241211,-74.4499969482422,-74.421875,-74.310546875,-74.285400390625,-74.3154296875,-74.2400360107422,-74.2291946411133,-74.243896484375,-74.3128890991211],[-75.04248046875,-75.0674819946289,-75.0987319946289,-75.1242218017578,-75.1421432495117,-75.107421875,-75.0794982910156,-75.0484390258789,-75.0322265625,-75.04248046875],[-73.6321792602539,-73.6648483276367,-73.694580078125,-73.7247543334961,-73.73486328125,-73.8001480102539,-73.8184585571289,-73.8169860839844,-73.7794952392578,-73.7239303588867,-73.6864776611328,-73.6412124633789,-73.6282272338867,-73.6166000366211,-73.6321792602539],[-72.9861297607422,-73.2284698486328,-73.3500061035156,-73.3970718383789,-73.4200668334961,-73.445068359375,-73.4036636352539,-73.31494140625,-73.281982421875,-73.2660217285156,-73.2715835571289,-73.260009765625,-73.2077178955078,-73.0284194946289,-72.8424301147461,-72.7763671875,-72.7640609741211,-72.8453140258789,-72.8971710205078,-72.9861297607422],[-73.7353515625,-73.7845764160156,-73.8623046875,-73.9832992553711,-73.9960403442383,-74.0020523071289,-73.9185562133789,-73.8773956298828,-73.827880859375,-73.7921447753906,-73.7953567504883,-73.7864685058594,-73.7271499633789,-73.7216796875,-73.7281799316406,-73.7521057128906,-73.77099609375,-73.8298873901367,-73.83447265625,-73.8489761352539,-74.0162582397461,-74.0990676879883,-74.0892639160156,-74.1952209472656,-74.2679672241211,-74.3498992919922,-74.4187469482422,-74.4988250732422,-74.6177749633789,-74.4805145263672,-74.5018081665039,-74.4216766357422,-74.3012237548828,-74.2125015258789,-74.1328125,-74.0972213745117,-74.1081085205078,-74.0828170776367,-73.9949264526367,-73.9001922607422,-73.8645477294922,-73.8177719116211,-73.7032241821289,-73.7037124633789,-73.7353515625],[-73.8106460571289,-73.7896499633789,-73.8336410522461,-73.9041519165039,-73.9382781982422,-73.9556655883789,-74.1177673339844,-74.1429672241211,-74.1399383544922,-73.9671859741211,-73.85693359375,-73.8414077758789,-73.8106460571289],[-74.6687469482422,-74.8104476928711,-74.8426818847656,-74.8419876098633,-74.8176727294922,-74.7450180053711,-74.6974639892578,-74.6726531982422,-74.664794921875,-74.6687469482422],[-73.7733840942383,-73.8485870361328,-73.918701171875,-73.9899444580078,-74.1144027709961,-74.2385787963867,-74.3549270629883,-74.3873519897461,-74.3731460571289,-74.2893600463867,-74.20947265625,-74.1562957763672,-74.1988296508789,-74.1935501098633,-74.174072265625,-74.1643524169922,-74.1703186035156,-74.1602096557617,-74.0723190307617,-74.0593719482422,-74.0568389892578,-74.018798828125,-74.030517578125,-74.0630416870117,-74.0366668701172,-73.73095703125,-73.52783203125,-73.5169448852539,-73.477783203125,-73.4544906616211,-73.4229049682617,-73.4392623901367,-73.5328140258789,-73.5245590209961,-73.4708023071289,-73.5492630004883,-73.6338882446289,-73.6534652709961,-73.7147445678711,-73.7892608642578,-73.766845703125,-73.6730499267578,-73.5682601928711,-73.5107421875,-73.4363327026367,-73.47265625,-73.5408172607422,-73.6496124267578,-73.7496566772461,-73.7378921508789,-73.7733840942383],[-78.8041458129883,-78.9833526611328,-78.9894485473633,-78.9792938232422,-78.9381332397461,-78.8882827758789,-78.87744140625,-78.8590316772461,-78.8381805419922,-78.78466796875,-78.7689437866211,-78.7747116088867,-78.8041458129883],[-109.279983520508,-109.434127807617,-109.429153442383,-109.390480041504,-109.276466369629,-109.222854614258,-109.279983520508],[-67.1948776245117,-67.0087890625,-67.0897445678711,-67.2191390991211,-67.3191452026367,-67.3355941772461,-67.356201171875,-67.57177734375,-67.88623046875,-68.04736328125,-68.2502975463867,-68.2995147705078,-68.3581008911133,-68.4225616455078,-68.4471206665039,-68.5072784423828,-68.56201171875,-68.56201171875,-68.5270462036133,-68.46630859375,-68.4471206665039,-68.4280242919922,-68.3842315673828,-68.3952102661133,-68.4307174682617,-68.496337890625,-68.5408172607422,-68.5920944213867,-68.6002960205078,-68.5419006347656,-68.5108413696289,-68.4267578125,-68.4144973754883,-68.5298385620117,-68.5757827758789,-68.5921859741211,-68.5915985107422,-68.5811538696289,-68.485107421875,-68.3733444213867,-68.3186492919922,-68.3186492919922,-68.3459930419922,-68.4053726196289,-68.537353515625,-68.5920944213867,-68.6521987915039,-68.7096176147461,-68.769775390625,-68.8463363647461,-68.8750915527344,-68.9419860839844,-68.9994125366211,-69.0421905517578,-69.1185073852539,-69.1552734375,-69.1744155883789,-69.251220703125,-69.3407211303711,-69.4095687866211,-69.4369125366211,-69.4888687133789,-69.5271453857422,-69.6569366455078,-69.6878890991211,-69.7349166870117,-69.7431640625,-69.8148498535156,-69.827880859375,-69.9003372192383,-69.99560546875,-70.0268096923828,-69.9826202392578,-69.9276351928711,-69.9241256713867,-69.9454574584961,-69.9599609375,-69.9235382080078,-69.8633804321289,-69.8442840576172,-69.8880386352539,-69.9071273803711,-69.9563446044922,-70.1020050048828,-70.1532211303711,-70.1696243286133,-70.1614227294922,-70.1939392089844,-70.2693786621094,-70.3192367553711,-70.34814453125,-70.33642578125,-70.3118209838867,-70.30908203125,-70.3505859375,-70.3883743286133,-70.4293899536133,-70.4730911254883,-70.5195770263672,-70.529052734375,-70.5546875,-70.56640625,-70.585205078125,-70.525634765625,-70.4501419067383,-70.3938522338867,-70.3309555053711,-70.28173828125,-70.25439453125,-70.2909164428711,-70.3555679321289,-70.36376953125,-70.3446273803711,-70.3200225830078,-70.2578125,-70.2297897338867,-70.1696243286133,-70.1769561767578,-70.1161575317383,-70.0520553588867,-70.02197265625,-70.0421371459961,-70.0930709838867,-70.10400390625,-70.0848617553711,-70.0198211669922,-69.9690475463867,-69.8961944580078,-69.8196258544922,-69.8086929321289,-69.7977523803711,-69.8387680053711,-69.882568359375,-69.8943328857422,-69.8814926147461,-69.8615264892578,-69.8573684692383,-69.8524398803711,-69.8797912597656,-69.9463348388672,-70.0028305053711,-70.0520553588867,-70.06298828125,-70.1014633178711,-70.1412582397461,-70.210693359375,-70.2546920776367,-70.2899398803711,-70.2867660522461,-70.3121109008789,-70.338134765625,-70.3931655883789,-70.4666061401367,-70.5251007080078,-70.55517578125,-70.5323257446289,-70.4704055786133,-70.4485397338867,-70.4567413330078,-70.4157257080078,-70.4197311401367,-70.3801727294922,-70.4157257080078,-70.4036636352539,-70.40478515625,-70.4567413330078,-70.5633773803711,-70.6218719482422,-70.721923828125,-70.7328567504883,-70.749267578125,-70.790283203125,-70.8531723022461,-70.9051284790039,-70.9779357910156,-71.0555191040039,-71.0732421875,-71.06640625,-71.107421875,-71.1593780517578,-71.1921920776367,-71.1593780517578,-71.1238250732422,-71.118408203125,-71.1634750366211,-71.2003936767578,-71.1648941040039,-71.1348114013672,-71.162841796875,-71.1867141723633,-71.1675796508789,-71.09619140625,-71.0281753540039,-71.0181655883789,-71.00048828125,-70.9679641723633,-70.899658203125,-70.84765625,-70.858642578125,-70.8969268798828,-70.9516067504883,-71.0871047973633,-71.197265625,-71.2857437133789,-71.3531799316406,-71.4015579223633,-71.4255828857422,-71.4093780517578,-71.4200134277344,-71.4653854370117,-71.5077667236328,-71.5257797241211,-71.53125,-71.5394592285156,-71.5870132446289,-71.654296875,-71.6925735473633,-71.7199249267578,-71.6968231201172,-71.6720733642578,-71.6378936767578,-71.6471176147461,-71.6597595214844,-71.7043914794922,-71.763671875,-71.8019561767578,-71.8183135986328,-71.8005905151367,-71.72265625,-71.6953125,-71.708984375,-71.7691421508789,-71.8046417236328,-71.8385238647461,-71.8837890625,-71.93212890625,-71.9413604736328,-71.873046875,-71.8807144165039,-71.8855972290039,-71.8921890258789,-71.8711471557617,-71.8976058959961,-71.9112777709961,-71.844482421875,-71.77001953125,-71.75,-71.7609405517578,-71.8607864379883,-71.944091796875,-71.9933090209961,-72.026123046875,-72.0644073486328,-72.1082077026367,-72.1246109008789,-72.078125,-72.053466796875,-72.1054153442383,-72.1436996459961,-72.1300277709961,-72.1136245727539,-72.1464309692383,-72.1023941040039,-72.0546875,-71.8985900878906,-71.781494140625,-71.7506332397461,-71.7638702392578,-71.8202209472656,-71.9049835205078,-71.9049835205078,-71.8324203491211,-71.7506332397461,-71.7327651977539,-71.7374038696289,-71.7947311401367,-71.7159652709961,-71.6800765991211,-71.7161636352539,-71.7671890258789,-71.8123550415039,-71.8121109008789,-71.8307647705078,-71.8350601196289,-71.8200225830078,-71.3257293701172,-71.2126007080078,-71.15087890625,-71.1597213745117,-71.2214813232422,-71.2611312866211,-71.358154296875,-71.4551773071289,-71.5603942871094,-71.6516647338867,-71.7828140258789,-71.95703125,-72.063720703125,-72.072509765625,-72.0417022705078,-71.8123550415039,-71.5962905883789,-71.5313034057617,-71.4434585571289,-71.353759765625,-71.3493194580078,-71.4904251098633,-71.5081024169922,-71.6933135986328,-71.7461929321289,-71.7726593017578,-71.7506332397461,-71.6800765991211,-71.6315383911133,-71.6844711303711,-71.8092727661133,-71.8756866455078,-71.8341293334961,-71.7776412963867,-71.7621078491211,-71.7312927246094,-71.6952133178711,-71.6996612548828,-71.7327194213867,-71.8564453125,-71.9402313232422,-71.9566421508789,-71.9629898071289,-71.9542465209961,-71.9005355834961,-71.9049835205078,-71.978515625,-72.0417022705078,-72.1034164428711,-72.2829055786133,-72.345947265625,-72.3415069580078,-72.41259765625,-72.4722137451172,-72.5179214477539,-72.5090866088867,-72.4079055786133,-72.3283233642578,-72.2930145263672,-72.354736328125,-72.4981384277344,-72.5828628540039,-72.6083984375,-72.5859375,-72.5917510986328,-72.6144027709961,-72.6512680053711,-72.7284698486328,-72.8654251098633,-72.9817428588867,-73.0336456298828,-73.0945816040039,-73.1488723754883,-73.13525390625,-73.4615707397461,-73.483642578125,-73.55419921875,-73.5762634277344,-73.5045394897461,-73.4704132080078,-73.5289077758789,-73.5077133178711,-73.5012741088867,-73.3866195678711,-73.3117218017578,-73.274169921875,-73.2516174316406,-73.2216262817383,-73.1745147705078,-73.1529312133789,-73.0823745727539,-72.95556640625,-72.8659210205078,-72.8036117553711,-72.6204071044922,-72.5098114013672,-72.4601593017578,-72.392578125,-72.3402328491211,-72.3006362915039,-72.2763137817383,-72.307373046875,-72.3591842651367,-72.3768081665039,-72.3591842651367,-72.3018569946289,-72.30322265625,-72.3664093017578,-72.4076690673828,-72.3345260620117,-72.2689971923828,-72.136962890625,-72.0284118652344,-71.9534683227539,-71.9710998535156,-71.9186553955078,-71.7166061401367,-71.4147415161133,-70.9431686401367,-70.4828567504883,-69.9602584838867,-69.7126007080078,-69.4884262084961,-69.2061996459961,-68.924560546875,-68.7151870727539,-68.5897979736328,-68.4609909057617,-68.443359375,-69.0072250366211,-69.1337890625,-69.2410125732422,-69.4468765258789,-69.5605926513672,-69.6203079223633,-69.7633361816406,-69.9072265625,-70.3909683227539,-70.5629348754883,-70.6803207397461,-70.7951126098633,-70.8390579223633,-70.8211898803711,-70.9520492553711,-70.9843292236328,-70.985107421875,-70.9478073120117,-70.995849609375,-71.0828094482422,-71.2977523803711,-71.4438934326172,-71.6937942504883,-71.8718719482422,-72.1009216308594,-72.1744079589844,-72.3768081665039,-72.3982467651367,-72.4128952026367,-72.3060531616211,-72.2486343383789,-72.0811538696289,-71.9416961669922,-71.8527374267578,-71.8282241821289,-71.8673324584961,-71.9027786254883,-71.8917007446289,-71.7914581298828,-71.7405319213867,-71.4003372192383,-71.2889633178711,-71.1802215576172,-71.1632843017578,-71.1550827026367,-71.2271423339844,-71.3877410888672,-71.8980026245117,-72.1290969848633,-72.2780227661133,-72.4582977294922,-72.4925765991211,-72.5308074951172,-72.5489273071289,-72.726806640625,-72.9983901977539,-73.052734375,-72.9982376098633,-72.91552734375,-72.909912109375,-72.88916015625,-72.8318862915039,-72.7276840209961,-72.6759796142578,-72.632080078125,-72.6266098022461,-72.4534683227539,-72.1175765991211,-71.9793014526367,-71.7970733642578,-71.5912094116211,-71.5541458129883,-71.5112838745117,-71.6647491455078,-71.8119125366211,-72.2256851196289,-72.3153762817383,-72.4376907348633,-72.478271484375,-72.50439453125,-72.6448287963867,-72.7121124267578,-72.7765579223633,-72.7660140991211,-72.8019104003906,-72.931884765625,-73.020263671875,-73.01611328125,-73.0230026245117,-73.0554733276367,-73.1224594116211,-73.3381881713867,-73.4598693847656,-73.5075225830078,-73.6452178955078,-73.345947265625,-73.2408218383789,-73.1448211669922,-73.0731964111328,-73.1239242553711,-73.1837921142578,-73.1781768798828,-73.244140625,-73.3821334838867,-73.585693359375,-73.7108383178711,-73.9146957397461,-74.0144500732422,-74.0358428955078,-73.9999008178711,-74.037353515625,-74.093505859375,-74.1508255004883,-74.1765670776367,-74.2384719848633,-74.2659683227539,-74.295654296875,-74.2649383544922,-74.1949234008789,-74.133544921875,-74.0402374267578,-73.83447265625,-73.7491149902344,-73.7027816772461,-73.6854019165039,-73.684326171875,-73.6490249633789,-73.5322265625,-73.4579620361328,-73.3267593383789,-73.2604522705078,-73.1373519897461,-72.9437026977539,-72.8432083129883,-72.7950210571289,-72.7354049682617,-72.6954574584961,-72.69482421875,-72.6495590209961,-72.5879821777344,-72.5708465576172,-72.5834503173828,-72.693603515625,-72.7140197753906,-72.6770477294922,-72.6314926147461,-72.5687026977539,-72.5329055786133,-72.5233383178711,-72.5193328857422,-72.5241241455078,-72.6135711669922,-72.624755859375,-72.6246109008789,-72.5228500366211,-72.494140625,-72.4896545410156,-72.5425338745117,-72.76123046875,-73.1267547607422,-73.1687469482422,-73.197021484375,-73.1633834838867,-73.114990234375,-72.7893524169922,-72.70458984375,-72.6490783691406,-72.5832977294922,-72.6000442504883,-72.9283676147461,-73.1886749267578,-73.3832550048828,-73.5181655883789,-73.5823211669922,-73.6502914428711,-73.7526321411133,-73.8106460571289,-73.8575668334961,-73.8944396972656,-73.9732437133789,-74.1464385986328,-74.1966781616211,-74.069580078125,-73.9297866821289,-73.8958969116211,-73.9394989013672,-74.1212387084961,-74.2104949951172,-74.3323287963867,-74.4143600463867,-74.5078659057617,-74.5868682861328,-74.6900863647461,-74.8147430419922,-74.9831085205078,-75.0553207397461,-75.0946807861328,-74.8366241455078,-74.6857452392578,-74.64892578125,-74.7020492553711,-74.77587890625,-74.7216796875,-74.6444778442383,-74.5641098022461,-74.3655776977539,-74.3314437866211,-74.190185546875,-74.1561050415039,-74.139404296875,-73.8474578857422,-73.8065490722656,-73.8246154785156,-73.7405700683594,-73.6590347290039,-73.6181640625,-73.6139602661133,-73.6932601928711,-73.6544418334961,-73.679931640625,-73.7501983642578,-73.8915100097656,-73.97802734375,-74.0967254638672,-74.1641159057617,-74.1972198486328,-74.1855926513672,-73.9503402709961,-74.0310516357422,-74.3055648803711,-74.3741226196289,-74.4250946044922,-74.5164108276367,-74.6295928955078,-74.434326171875,-74.333740234375,-74.0194396972656,-73.9585952758789,-74.01123046875,-74.0732955932617,-74.1713409423828,-74.3239288330078,-74.3187484741211,-74.2908172607422,-74.2303695678711,-74.1019973754883,-73.9555206298828,-73.8915557861328,-73.8363800048828,-73.8924865722656,-73.988037109375,-74.0944290161133,-74.08349609375,-74.0492172241211,-74.0234375,-74.005615234375,-74.015380859375,-73.9847640991211,-73.9378890991211,-73.9349594116211,-74.0277328491211,-74.0613250732422,-74.0738830566406,-74.1398010253906,-74.1678695678711,-74.1845703125,-74.2212905883789,-74.301513671875,-74.3485336303711,-74.3665542602539,-74.358154296875,-74.3799819946289,-74.3821258544922,-74.3410186767578,-74.2276916503906,-74.1762237548828,-74.1293029785156,-74.0569305419922,-74.0090789794922,-74.1715316772461,-74.2700729370117,-74.3429718017578,-74.4744186401367,-74.4994125366211,-74.5771942138672,-74.5907211303711,-74.5846710205078,-74.4004821777344,-74.2504425048828,-73.8538131713867,-73.5281753540039,-73.3844757080078,-73.3910675048828,-73.5009765625,-73.569580078125,-73.6099090576172,-73.62890625,-73.6351089477539,-73.7158660888672,-73.7482452392578,-73.7792434692383,-73.8466796875,-73.9408721923828,-74.0847702026367,-74.22705078125,-74.3505859375,-74.3793487548828,-74.376220703125,-74.4296875,-74.5692367553711,-74.60888671875,-74.6549301147461,-74.5884246826172,-74.5337905883789,-74.4668960571289,-74.403564453125,-74.3225631713867,-74.2429656982422,-74.1519546508789,-74.1340789794922,-74.1910171508789,-74.242919921875,-74.3236770629883,-74.482666015625,-74.4032745361328,-74.2156753540039,-74.1583938598633,-74.2080612182617,-74.1519088745117,-74.20947265625,-74.3135681152344,-74.4542007446289,-74.4844207763672,-74.4893569946289,-74.4669876098633,-74.4801712036133,-74.5122528076172,-74.6908264160156,-74.8106460571289,-75.0059509277344,-75.0314483642578,-75.0525360107422,-74.9841766357422,-75.0187454223633,-75.145751953125,-75.33740234375,-75.4784164428711,-75.5403366088867,-75.5650863647461,-75.527587890625,-75.4459991455078,-75.3864212036133,-75.4012145996094,-75.4303665161133,-75.4966278076172,-75.63525390625,-75.7081069946289,-75.7063980102539,-75.6567916870117,-75.4369125366211,-75.3760299682617,-75.2470169067383,-75.0748519897461,-74.9244613647461,-74.9976577758789,-75.0745086669922,-75.0667037963867,-74.7631378173828,-74.6306610107422,-74.4627914428711,-74.369140625,-74.3011779785156,-74.1578674316406,-74.09619140625,-74.0818405151367,-74.08251953125,-74.0992660522461,-74.1227035522461,-74.0989227294922,-74.0375518798828,-73.9571762084961,-73.9203109741211,-73.8250045776367,-73.8441390991211,-73.88232421875,-73.9602508544922,-73.99951171875,-74.06103515625,-74.0199203491211,-74.08154296875,-74.3567886352539,-74.3929748535156,-74.3724594116211,-74.213134765625,-74.0897445678711,-73.9675827026367,-73.92919921875,-73.8787155151367,-73.8125534057617,-73.7352523803711,-73.6948699951172,-73.7079086303711,-73.7081604003906,-73.8106460571289,-73.934814453125,-73.9486312866211,-73.9437484741211,-73.8453598022461,-73.770263671875,-73.7162094116211,-73.6620635986328,-73.668212890625,-73.6516571044922,-73.6294479370117,-73.5918426513672,-73.5943374633789,-73.6619644165039,-73.756591796875,-73.7803726196289,-73.7307586669922,-73.5499038696289,-73.3785629272461,-73.2662124633789,-73.2023391723633,-72.9781723022461,-72.933837890625,-72.9408187866211,-72.9755401611328,-73.0638656616211,-73.2263717651367,-73.4449691772461,-73.4048843383789,-73.3624572753906,-73.2564392089844,-73.0784225463867,-72.7389678955078,-72.6800765991211,-72.6638717651367,-72.8275375366211,-73.0010299682617,-73.1409683227539,-73.2650909423828,-73.24072265625,-73.2244644165039,-73.0687942504883,-72.99658203125,-73.1009750366211,-73.0759735107422,-72.9399948120117,-72.9154815673828,-72.8780288696289,-72.7580108642578,-72.75537109375,-72.7660140991211,-72.844970703125,-72.8480453491211,-72.77392578125,-72.6548309326172,-72.6318359375,-72.71630859375,-72.78515625,-72.7732391357422,-72.7073669433594,-72.6310577392578,-72.5484390258789,-72.4302749633789,-72.412353515625,-72.4601058959961,-72.4994125366211,-72.6239776611328,-72.7381820678711,-72.7811965942383,-72.8240737915039,-72.7838363647461,-72.7436981201172,-72.6598663330078,-72.4860382080078,-72.3603973388672,-72.3182601928711,-72.3594741821289,-72.427734375,-72.5423812866211,-72.600830078125,-72.6697769165039,-72.8051223754883,-72.8799819946289,-72.9528350830078,-73.0149917602539,-73.174072265625,-73.2417984008789,-73.5212936401367,-73.6240234375,-73.7351531982422,-73.7218704223633,-73.6880798339844,-73.6250457763672,-73.6239242553711,-73.7106399536133,-73.8107376098633,-73.8550262451172,-73.8761749267578,-73.9658660888672,-73.9835891723633,-73.9203109741211,-73.7840347290039,-73.7424774169922,-73.66943359375,-73.6709976196289,-73.4822311401367,-73.410400390625,-73.2499008178711,-73.2264633178711,-73.4807586669922,-73.5202178955078,-73.5325698852539,-73.4718704223633,-73.4647979736328,-73.5167465209961,-73.6618118286133,-73.6646041870117,-73.6034164428711,-73.6623992919922,-73.6336364746094,-73.6016616821289,-73.3745651245117,-73.2710952758789,-73.2159652709961,-73.1728515625,-73.1512680053711,-73.1377944946289,-73.1180648803711,-73.006591796875,-72.9678192138672,-72.8745574951172,-72.7784194946289,-72.6833953857422,-72.5873489379883,-72.6239242553711,-72.5620574951172,-72.5051727294922,-72.4549789428711,-72.38671875,-72.2237777709961,-72.1824264526367,-72.0559539794922,-72.03076171875,-71.9915084838867,-72.0028305053711,-71.9268569946289,-71.8539505004883,-71.8309555053711,-71.6643524169922,-71.6362838745117,-71.6955032348633,-71.6965866088867,-71.7429656982422,-71.6355514526367,-71.592041015625,-71.4522476196289,-71.46142578125,-71.4212875366211,-71.5130386352539,-71.52587890625,-71.5772933959961,-71.6619644165039,-71.6539077758789,-71.7056655883789,-71.7089309692383,-71.6694793701172,-71.400390625,-71.3480453491211,-71.3157196044922,-71.3267059326172,-71.353271484375,-71.48583984375,-71.5192413330078,-71.4936065673828,-71.3840866088867,-71.3067398071289,-71.266845703125,-71.1864242553711,-71.1544952392578,-71.0865249633789,-71.0526351928711,-70.94580078125,-70.92578125,-70.9092788696289,-70.9142608642578,-70.8979034423828,-70.812744140625,-70.8029327392578,-70.7083969116211,-70.6869583129883,-70.6465759277344,-70.6622543334961,-70.6354446411133,-70.6996078491211,-70.7137222290039,-70.6330108642578,-70.578125,-70.489501953125,-70.4522018432617,-70.4453659057617,-70.5586395263672,-70.5741195678711,-70.5464324951172,-70.5074234008789,-70.5200729370117,-70.5074234008789,-70.48779296875,-70.4099578857422,-70.392333984375,-70.4196243286133,-70.511962890625,-70.588134765625,-70.5933609008789,-70.56884765625,-70.5631866455078,-70.4496612548828,-70.38916015625,-70.3316879272461,-70.259521484375,-70.228515625,-70.1854476928711,-70.1550827026367,-70.1295852661133,-70.0875473022461,-70.0800323486328,-70.08837890625,-70.197021484375,-70.1936492919922,-70.1474609375,-70.1481475830078,-70.1574249267578,-70.1983413696289,-70.2104034423828,-70.2757797241211,-70.3348617553711,-70.3360824584961,-70.3616256713867,-70.4182586669922,-70.3774871826172,-70.2822799682617,-70.1837844848633,-70.05908203125,-69.9263687133789,-69.8396987915039,-69.8025894165039,-69.8024444580078,-69.8415069580078,-69.8520965576172,-69.8061065673828,-69.6847686767578,-69.58642578125,-69.5109405517578,-69.4950180053711,-69.3580093383789,-69.3133850097656,-69.2823257446289,-69.0939483642578,-69.0904312133789,-69.1180648803711,-69.1454620361328,-69.1263732910156,-69.09228515625,-69.0809555053711,-69.0601577758789,-69.0393981933594,-69.0268096923828,-68.9788589477539,-68.9690933227539,-68.9683151245117,-68.9310073852539,-68.8579559326172,-68.7590866088867,-68.6806640625,-68.6205520629883,-68.5478515625,-68.4919891357422,-68.4701690673828,-68.462890625,-68.4870071411133,-68.5752868652344,-68.6982879638672,-68.6961898803711,-68.5782699584961,-68.5593795776367,-68.5606918334961,-68.6001968383789,-68.7274856567383,-68.7558135986328,-68.7593231201172,-68.7300262451172,-68.7345733642578,-68.6885757446289,-68.7123031616211,-68.7592239379883,-68.7605438232422,-68.7451629638672,-68.69580078125,-68.4998474121094,-68.4843292236328,-68.4873962402344,-68.5631790161133,-68.571044921875,-68.5689468383789,-68.5582504272461,-68.5338363647461,-68.4354934692383,-68.3138656616211,-68.197021484375,-68.1985397338867,-68.1864242553711,-68.1121520996094,-68.101806640625,-68.0767517089844,-67.9883804321289,-67.9539108276367,-67.9449157714844,-67.9503860473633,-67.8817367553711,-67.8737258911133,-67.8899917602539,-67.88916015625,-67.8794479370117,-67.8205032348633,-67.79443359375,-67.7073211669922,-67.5799331665039,-67.3622589111328,-67.1948776245117],[110.888771057129,110.938285827637,110.970703125,110.997657775879,111.013671875,110.912696838379,110.822265625,110.640914916992,110.603126525879,110.572174072266,110.5625,110.566017150879,110.519340515137,110.477630615234,110.451271057129,110.399513244629,110.333694458008,110.290824890137,110.251762390137,110.15625,110.048530578613,110.06640625,110.0673828125,110.020217895508,109.967674255371,109.815628051758,109.759765625,109.702735900879,109.681053161621,109.589553833008,109.519340515137,109.400093078613,109.340919494629,109.183204650879,109.029884338379,108.922264099121,108.701568603516,108.676078796387,108.638084411621,108.635650634766,108.650001525879,108.66552734375,108.693557739258,108.791015625,108.90283203125,109.062889099121,109.179100036621,109.276664733887,109.219535827637,109.177436828613,109.218948364258,109.263481140137,109.314842224121,109.418159484863,109.513671875,109.584281921387,109.6513671875,109.90625,110.0830078125,110.171577453613,110.21337890625,110.343948364258,110.392280578613,110.387985229492,110.3935546875,110.417572021484,110.588188171387,110.588768005371,110.598335266113,110.651756286621,110.678512573242,110.744529724121,110.80908203125,110.888771057129],[110.385154724121,110.42236328125,110.521575927734,110.53955078125,110.538864135742,110.50390625,110.421875,110.339942932129,110.280960083008,110.2646484375,110.309860229492,110.385154724121],[112.643753051758,112.545600891113,112.525001525879,112.558982849121,112.647659301758,112.643753051758],[112.790229797363,112.771095275879,112.741989135742,112.733497619629,112.712699890137,112.760551452637,112.782028198242,112.839065551758,112.862602233887,112.812591552734,112.800682067871,112.790229797363],[113.555274963379,113.563667297363,113.485641479492,113.46337890625,113.426078796387,113.404396057129,113.464942932129,113.5205078125,113.555274963379],[118.183006286621,118.149513244629,118.090530395508,118.088768005371,118.076759338379,118.092971801758,118.103805541992,118.170700073242,118.183006286621],[119.820899963379,119.746688842773,119.700294494629,119.699417114258,119.723052978516,119.695991516113,119.722564697266,119.777923583984,119.797462463379,119.828712463379,119.838371276855,119.838676452637,119.809089660645,119.832420349121,119.820899963379],[121.251365661621,121.164260864258,121.131546020508,121.133979797363,121.205467224121,121.234367370605,121.2509765625,121.251365661621],[122.172561645508,122.169036865234,122.083786010742,122.042678833008,122.062309265137,122.11962890625,122.1650390625,122.172561645508],[122.403900146484,122.39404296875,122.367576599121,122.331832885742,122.350982666016,122.401565551758,122.403900146484],[122.295906066895,122.281547546387,122.157814025879,122.024017333984,121.977836608887,121.969436645508,122.110542297363,122.284469604492,122.322265625,122.295906066895],[121.862693786621,121.780471801758,121.519927978516,121.33642578125,121.226852416992,121.211135864258,121.338966369629,121.464164733887,121.491798400879,121.542289733887,121.576560974121,121.808296203613,121.843650817871,121.862693786621],[130.526947021484,130.498229980469,130.450286865234,130.360748291016,130.295608520508,130.246673583984,130.248825073242,130.240325927734,130.151275634766,130.124801635742,130.082626342773,130.022262573242,129.976959228516,129.941207885742,129.898239135742,129.861038208008,129.841506958008,129.779190063477,129.7734375,129.746475219727,129.7197265625,129.697845458984,129.6279296875,129.603912353516,129.567581176758,129.523727416992,129.48486328125,129.423629760742,129.365814208984,129.313674926758,129.252532958984,129.2177734375,129.205383300781,129.195510864258,129.133682250977,129.077239990234,128.960632324219,128.923446655273,128.83984375,128.7490234375,128.626770019531,128.42724609375,128.307815551758,128.16015625,128.045211791992,128.028717041016,128.032913208008,128.056060791016,128.084167480469,128.131927490234,128.181732177734,128.2578125,128.289260864258,128.290924072266,128.2548828125,128.200286865234,128.1494140625,128.111236572266,128.052734375,128.013092041016,127.918647766113,127.687698364258,127.572166442871,127.516998291016,127.420310974121,127.270797729492,127.179695129395,127.13671875,127.128410339355,127.085357666016,127.061325073242,127.006927490234,126.954788208008,126.903518676758,126.847267150879,126.787696838379,126.743064880371,126.721580505371,126.696975708008,126.601272583008,126.578323364258,126.540138244629,126.513572692871,126.490432739258,126.451461791992,126.411819458008,126.328704833984,126.253616333008,126.144538879395,126.093162536621,126.066795349121,125.989067077637,125.874908447266,125.783981323242,125.728317260742,125.688278198242,125.659187316895,125.645111083984,125.593849182129,125.542579650879,125.416893005371,125.314453125,125.185935974121,125.072952270508,125.025970458984,125.013374328613,124.996871948242,124.942283630371,124.889350891113,124.77197265625,124.712394714355,124.481056213379,124.386627197266,124.362106323242,124.349998474121,124.267478942871,124.105766296387,123.760162353516,123.65087890625,123.611236572266,123.580657958984,123.490036010742,123.348152160645,123.268943786621,123.226554870605,123.0322265625,122.960929870605,122.840034484863,122.334861755371,122.225006103516,122.120903015137,122.047653198242,121.982330322266,121.922660827637,121.864356994629,121.805183410645,121.744827270508,121.677253723145,121.6328125,121.67041015625,121.64990234375,121.517196655273,121.320114135742,121.236320495605,121.207420349121,121.163581848145,121.121681213379,121.106742858887,121.188278198242,121.263290405273,121.679885864258,121.627639770508,121.66455078125,121.757804870605,121.818458557129,121.785453796387,121.512496948242,121.355659484863,121.275489807129,121.2998046875,121.286331176758,121.267478942871,121.406448364258,121.469535827637,121.517578125,121.514259338379,121.474220275879,121.517387390137,121.800971984863,121.868949890137,121.982810974121,122.190925598145,122.203323364258,122.263870239258,122.275001525879,122.1787109375,122.140434265137,121.858787536621,121.834861755371,121.80859375,121.765617370605,121.729301452637,121.598922729492,121.537109375,121.174507141113,121.0859375,121.002937316895,120.922271728516,120.84130859375,120.770706176758,120.479103088379,120.368942260742,119.850395202637,119.591110229492,119.391120910645,119.322364807129,119.261329650879,119.224609375,119.040145874023,118.976951599121,118.912307739258,118.826469421387,118.75244140625,118.626373291016,118.471969604492,118.2978515625,118.147850036621,118.040916442871,117.86572265625,117.784675598145,117.616706848145,117.553810119629,117.557815551758,117.656059265137,117.766693115234,118.014945983887,118.543258666992,118.667091369629,118.799995422363,118.940040588379,119.027534484863,119.035652160645,119.038475036621,119.070320129395,119.089157104492,119.033493041992,118.990814208984,118.954887390137,118.95263671875,118.998153686523,119.11181640625,119.287399291992,119.449897766113,119.760551452637,119.887496948242,119.879981994629,119.882904052734,120.155853271484,120.311531066895,120.287101745605,120.257225036621,120.284675598145,120.3701171875,120.75,121.049026489258,121.219535827637,121.388092041016,121.505264282227,121.640235900879,121.816413879395,121.96484375,122.010154724121,122.056640625,122.109573364258,122.169136047363,122.337692260742,122.493270874023,122.602348327637,122.666984558105,122.573348999023,122.587303161621,122.515518188477,122.446685791016,122.487411499023,122.523445129395,122.519729614258,122.457023620605,122.340911865234,122.274223327637,122.242286682129,122.219734191895,122.203216552734,122.162399291992,122.049507141113,121.932716369629,121.669631958008,121.4130859375,121.14404296875,121.053802490234,120.989944458008,120.878517150879,120.810844421387,120.796676635742,120.882614135742,120.904975891113,120.895797729492,120.847068786621,120.776168823242,120.711517333984,120.682228088379,120.680961608887,120.637886047363,120.519332885742,120.39306640625,120.348243713379,120.330268859863,120.343452453613,120.327728271484,120.270126342773,120.183296203613,120.116989135742,120.094139099121,120.181442260742,120.264739990234,120.284767150879,120.219047546387,120.0546875,120.027450561523,119.978706359863,119.911712646484,119.8662109375,119.810546875,119.719734191895,119.6083984375,119.526466369629,119.429679870605,119.352828979492,119.215812683105,119.165328979492,119.200981140137,119.351364135742,119.4267578125,119.582908630371,119.769729614258,119.963668823242,120.201469421387,120.266693115234,120.322662353516,120.425689697266,120.499809265137,120.504783630371,120.615623474121,120.734466552734,120.871101379395,120.897369384766,120.85302734375,120.853218078613,120.989944458008,121.293357849121,121.341697692871,121.400970458984,121.403907775879,121.450775146484,121.490531921387,121.674217224121,121.751075744629,121.832420349121,121.856353759766,121.866310119629,121.763572692871,121.680854797363,121.351959228516,121.266403198242,121.145797729492,120.973541259766,120.791702270508,120.660552978516,120.520118713379,120.184074401855,120.098724365234,120.073921203613,120.035942077637,120.191604614258,120.347457885742,120.497169494629,120.715530395508,120.752243041992,120.787796020508,120.9375,121.055374145508,121.204879760742,121.350975036621,121.66064453125,121.785934448242,121.83447265625,121.877937316895,121.769432067871,121.675186157227,121.527534484863,121.4189453125,121.309967041016,120.997665405273,120.938278198242,120.8974609375,120.821487426758,120.629981994629,120.449798583984,120.245506286621,120.194625854492,120.228515625,120.260551452637,120.3525390625,120.494537353516,120.633392333984,120.904495239258,121.159370422363,121.258003234863,121.340621948242,121.432716369629,121.677932739258,121.812301635742,121.9443359375,122.017288208008,122.082908630371,121.90576171875,121.676567077637,121.574607849121,121.506248474121,121.690437316895,121.821876525879,121.887992858887,121.941207885742,121.968353271484,121.917770385742,121.853507995605,121.790824890137,121.717483520508,121.655952453613,121.53369140625,121.487113952637,121.447662353516,121.520896911621,121.664947509766,121.679679870605,121.641014099121,121.540031433105,121.662498474121,121.630081176758,121.59033203125,121.519149780273,121.475204467773,121.538078308105,121.60205078125,121.609962463379,121.509963989258,121.354591369629,121.272262573242,121.216804504395,121.145698547363,121.098434448242,121.035446166992,120.958595275879,120.892478942871,120.81298828125,120.747665405273,120.763481140137,120.833015441895,120.833015441895,120.685150146484,120.661331176758,120.664840698242,120.587493896484,120.629104614258,120.607521057129,120.539848327637,120.468650817871,120.384567260742,120.278717041016,120.138572692871,120.097457885742,120.086715698242,120.04296875,119.967781066895,119.882225036621,119.879493713379,119.842376708984,119.821281433105,119.815139770508,119.82421875,119.788681030273,119.766700744629,119.710441589355,119.651565551758,119.588180541992,119.589942932129,119.623634338379,119.63818359375,119.725975036621,119.784767150879,119.831146240234,119.840324401855,119.876457214355,119.881050109863,119.797271728516,119.692672729492,119.567092895508,119.463081359863,119.369728088379,119.313087463379,119.232124328613,119.139457702637,119.263763427734,119.332038879395,119.417770385742,119.500877380371,119.618759155273,119.648246765137,119.616897583008,119.552833557129,119.539459228516,119.619140625,119.622459411621,119.5927734375,119.499214172363,119.421775817871,119.343742370605,119.263076782227,119.180076599121,119.146286010742,119.169326782227,119.243560791016,119.285552978516,119.235542297363,119.024612426758,118.9775390625,118.914451599121,118.955665588379,118.909088134766,118.822067260742,118.707511901855,118.636917114258,118.640235900879,118.691802978516,118.719139099121,118.657035827637,118.560348510742,118.412010192871,118.295310974121,118.194526672363,118.087112426758,118.013862609863,118.005958557129,117.93505859375,117.896881103516,117.842674255371,117.848251342773,117.878997802734,118.024223327637,118.050582885742,118.056053161621,117.904106140137,117.839447021484,117.74169921875,117.667877197266,117.628219604492,117.579193115234,117.466415405273,117.43310546875,117.459571838379,117.462203979492,117.4169921875,117.36767578125,117.3466796875,117.330764770508,117.290824890137,117.225006103516,117.148139953613,117.082511901855,117.032821655273,116.910636901855,116.860946655273,116.759567260742,116.712112426758,116.629402160645,116.682319641113,116.698822021484,116.669136047363,116.58642578125,116.538284301758,116.519828796387,116.470710754395,116.345512390137,116.251853942871,116.222076416016,116.206352233887,116.157417297363,116.062606811523,115.852149963379,115.755859375,115.640426635742,115.561134338379,115.53466796875,115.498336791992,115.382522583008,115.289947509766,115.19580078125,115.091499328613,115.012107849121,114.914451599121,114.896392822266,114.853813171387,114.750389099121,114.711128234863,114.651664733887,114.5927734375,114.571968078613,114.54443359375,114.55419921875,114.496185302734,114.420112609863,114.340621948242,114.266021728516,114.228218078613,114.188179016113,114.122848510742,114.097854614258,114.050392150879,114.018257141113,114.015434265137,113.93115234375,113.828315734863,113.754493713379,113.6611328125,113.61962890625,113.603416442871,113.586326599121,113.592185974121,113.620513916016,113.519721984863,113.4453125,113.460350036621,113.44189453125,113.3310546875,113.337799072266,113.344825744629,113.432029724121,113.449806213379,113.484764099121,113.553031921387,113.551467895508,113.5888671875,113.576461791992,113.549125671387,113.546775817871,113.527053833008,113.494140625,113.481048583984,113.478904724121,113.473434448242,113.415718078613,113.367385864258,113.327735900879,113.266403198242,113.149024963379,113.088768005371,113.008201599121,112.983787536621,112.953903198242,112.90380859375,112.80859375,112.725387573242,112.660736083984,112.634078979492,112.586334228516,112.494728088379,112.421287536621,112.439453125,112.429298400879,112.396087646484,112.359664916992,112.37744140625,112.389747619629,112.3564453125,112.304985046387,112.193359375,112.1171875,112.025199890137,111.943946838379,111.926467895508,111.873435974121,111.824607849121,111.775978088379,111.7119140625,111.681640625,111.602737426758,111.392379760742,111.319137573242,111.220611572266,111.144241333008,111.1005859375,111.061126708984,111.016899108887,110.996772766113,110.878028869629,110.771095275879,110.652145385742,110.567192077637,110.504295349121,110.4580078125,110.4345703125,110.410934448242,110.3974609375,110.374603271484,110.331153869629,110.193550109863,110.154006958008,110.180366516113,110.365425109863,110.388481140137,110.370506286621,110.326171875,110.313087463379,110.511528015137,110.517578125,110.486915588379,110.449516296387,110.3447265625,110.123146057129,109.9384765625,109.882522583008,109.885841369629,109.931640625,109.98388671875,109.968360900879,109.946388244629,109.861038208008,109.7919921875,109.805274963379,109.767379760742,109.726272583008,109.684761047363,109.66259765625,109.704490661621,109.681243896484,109.760154724121,109.779586791992,109.921096801758,109.930763244629,109.82958984375,109.759376525879,109.743362426758,109.686912536621,109.594329833984,109.56640625,109.521484375,109.544044494629,109.435546875,109.3466796875,109.220405578613,109.148635864258,109.08154296875,109.09814453125,109.133491516113,109.101760864258,109.030570983887,108.921775817871,108.846389770508,108.771675109863,108.743949890137,108.674514770508,108.615821838379,108.58935546875,108.615821838379,108.59375,108.479888916016,108.480865478516,108.492576599121,108.525680541992,108.502151489258,108.4443359375,108.3828125,108.354591369629,108.324806213379,108.302146911621,108.246284484863,108.145606994629,108.0673828125,107.97265625,107.908393859863,107.802047729492,107.75927734375,107.641014099121,107.471389770508,107.433494567871,107.351165771484,107.272071838379,107.178512573242,107.061622619629,107.019821166992,107.006439208984,106.970993041992,106.925201416016,106.87451171875,106.794143676758,106.7294921875,106.697654724121,106.66357421875,106.65771484375,106.660057067871,106.654197692871,106.636528015137,106.593162536621,106.553611755371,106.536331176758,106.550392150879,106.582427978516,106.633102416992,106.701560974121,106.736328125,106.7802734375,106.6240234375,106.541801452637,106.450881958008,106.338088989258,106.279006958008,106.249412536621,106.183982849121,106.1484375,106.068458557129,106.0009765625,105.962303161621,105.902633666992,105.842964172363,105.782325744629,105.691207885742,105.548141479492,105.530860900879,105.494529724121,105.440139770508,105.350494384766,105.275390625,105.23876953125,105.189064025879,104.995704650879,104.91015625,104.86474609375,104.826560974121,104.814743041992,104.795700073242,104.740036010742,104.687309265137,104.631736755371,104.577537536621,104.52685546875,104.371780395508,104.29833984375,104.23828125,104.212501525879,104.14306640625,104.053909301758,104.0126953125,103.990821838379,103.971389770508,103.941505432129,103.9150390625,103.637298583984,103.620208740234,103.570701599121,103.525390625,103.492965698242,103.470993041992,103.356056213379,103.32666015625,103.300590515137,103.266304016113,103.193359375,103.137596130371,103.136329650879,103.075881958008,103.00537109375,102.98193359375,102.935157775879,102.874221801758,102.830078125,102.721000671387,102.598533630371,102.517181396484,102.470893859863,102.427925109863,102.406448364258,102.375778198242,102.30224609375,102.237007141113,102.175979614258,102.12744140625,102.091506958008,102.0244140625,101.945411682129,101.841796875,101.759963989258,101.73876953125,101.70751953125,101.671485900879,101.646194458008,101.619926452637,101.56787109375,101.524513244629,101.537307739258,101.561820983887,101.560256958008,101.575782775879,101.602928161621,101.699607849121,101.736518859863,101.743942260742,101.747268676758,101.743453979492,101.724220275879,101.722946166992,101.763084411621,101.802047729492,101.800590515137,101.783493041992,101.728126525879,101.704788208008,101.668556213379,101.621681213379,101.583885192871,101.542381286621,101.443557739258,101.281448364258,101.247848510742,101.224411010742,101.211822509766,101.219917297363,101.20556640625,101.175392150879,101.196678161621,101.138862609863,101.147270202637,101.128120422363,101.130859375,101.120704650879,101.079788208008,101.019340515137,100.835151672363,100.677146911621,100.604591369629,100.531349182129,100.445701599121,100.3505859375,100.214752197266,100.147659301758,100.116798400879,100.089256286621,100.105758666992,100.095504760742,100.041213989258,99.9782257080078,99.9407196044922,99.9255828857422,99.9404296875,99.9478530883789,99.9176788330078,99.8253936767578,99.5926742553711,99.388671875,99.3031234741211,99.2333984375,99.1929626464844,99.1734390258789,99.17236328125,99.2053680419922,99.2430648803711,99.3376998901367,99.3431625366211,99.3382797241211,99.3851547241211,99.466796875,99.5071258544922,99.4972610473633,99.4645462036133,99.4180679321289,99.3408203125,99.2203140258789,99.0550765991211,98.86376953125,98.8855438232422,98.8826141357422,98.85888671875,98.8197250366211,98.7978515625,98.8322296142578,98.7876968383789,98.7350540161133,98.6808624267578,98.6767578125,98.7015609741211,98.833984375,98.8350601196289,98.8023452758789,98.7643585205078,98.5833969116211,98.5641555786133,98.4994125366211,98.3672866821289,98.2125015258789,98.0168991088867,97.8376922607422,97.7556610107422,97.68603515625,97.6296844482422,97.5645523071289,97.5682601928711,97.6906280517578,97.7082061767578,97.6707000732422,97.6666030883789,97.6236343383789,97.5632858276367,97.5314483642578,97.5293884277344,97.5833053588867,97.6707000732422,97.7238235473633,97.7378921508789,97.7107391357422,97.7149429321289,97.7673797607422,97.8195343017578,97.91796875,97.9620056152344,98.0107421875,98.0640640258789,98.099609375,98.1428756713867,98.1725540161133,98.2965774536133,98.3337860107422,98.4016647338867,98.4655303955078,98.5584030151367,98.6253890991211,98.65625,98.6546859741211,98.5910110473633,98.5640640258789,98.5719680786133,98.6631851196289,98.685546875,98.671875,98.70947265625,98.7318344116211,98.7393493652344,98.7384796142578,98.7294921875,98.7165069580078,98.6748046875,98.6824264526367,98.6767578125,98.6511764526367,98.5998077392578,98.5044860839844,98.4525375366211,98.4088897705078,98.3923873901367,98.3504867553711,98.298828125,98.2742156982422,98.2410125732422,98.1304702758789,98.1183547973633,98.0989303588867,98.0616226196289,98.0222625732422,97.93408203125,97.8876037597656,97.8649368286133,97.8165054321289,97.76904296875,97.7300796508789,97.6946258544922,97.6588821411133,97.5992126464844,97.5378875732422,97.5021438598633,97.4777374267578,97.4314498901367,97.3564453125,97.3224639892578,97.2894515991211,97.1451187133789,97.0753860473633,96.9808578491211,96.8330078125,96.7757797241211,96.65283203125,96.6026382446289,96.427734375,96.3890609741211,96.3664093017578,96.31982421875,96.2814483642578,96.2789077758789,96.326171875,96.3298797607422,96.3273468017578,96.3956069946289,96.5808639526367,96.5499954223633,96.4771423339844,96.4670944213867,96.4357376098633,96.3468704223633,96.1622085571289,96.1371154785156,96.1414108276367,96.1223678588867,96.1808624267578,96.2705078125,96.3397445678711,96.3558654785156,96.3372116088867,96.2349624633789,96.1947250366211,96.1285095214844,96.07958984375,96.0353469848633,95.8850555419922,95.7103500366211,95.5158157348633,95.5169906616211,95.4937515258789,95.45654296875,95.4202117919922,95.3892593383789,95.3531188964844,95.2791061401367,95.1447296142578,94.9988250732422,94.9674835205078,94.7694320678711,94.7630844116211,94.7333984375,94.6770477294922,94.623046875,94.4680633544922,94.2932586669922,94.1934585571289,94.1115264892578,94.0176773071289,94.0132751464844,93.9736328125,93.9022445678711,93.7607421875,93.6649398803711,93.3605422973633,93.251953125,93.20654296875,93.1578140258789,93.1192398071289,93.0349655151367,92.8818359375,92.7018508911133,92.6525421142578,92.6434555053711,92.6656265258789,92.6875,92.6877899169922,92.6643600463867,92.5466766357422,92.4806671142578,92.4148406982422,92.3410186767578,92.2701187133789,92.25048828125,92.2222671508789,92.1576156616211,92.1012725830078,91.9776382446289,91.9093780517578,91.82470703125,91.7126007080078,91.6319351196289,91.62939453125,91.6418914794922,91.6055679321289,91.4933624267578,91.3675765991211,91.3068389892578,91.2730484008789,91.2258758544922,91.14990234375,91.0777282714844,91.0208053588867,90.9625015258789,90.9066390991211,90.7157211303711,90.6300811767578,90.4773406982422,90.3527374267578,90.3331069946289,90.3337860107422,90.3521499633789,90.3629837036133,90.3482360839844,90.2208023071289,90.1044921875,89.9810562133789,89.8978500366211,89.81689453125,89.7498092651367,89.6527328491211,89.5369186401367,89.4806594848633,89.3958969116211,89.2726593017578,89.1604461669922,89.1023483276367,89.0254898071289,88.9475555419922,88.8914108276367,88.83251953125,88.7648468017578,88.7490234375,88.8298797607422,88.8488311767578,88.82861328125,88.8037109375,88.7562561035156,88.62109375,88.5779266357422,88.5316390991211,88.4861297607422,88.4259796142578,88.2751922607422,88.14111328125,88.1089859008789,88.0989303588867,88.1097717285156,88.0233383178711,87.9333953857422,87.8607406616211,87.6827163696289,87.62255859375,87.5552749633789,87.4641571044922,87.2907257080078,87.1414031982422,87.0201110839844,86.9337844848633,86.8423843383789,86.7503890991211,86.7196273803711,86.6905288696289,86.6144561767578,86.5544967651367,86.5168914794922,86.4849624633789,86.40869140625,86.32861328125,86.2179718017578,86.1742248535156,86.1370086669922,86.0787124633789,86.0754928588867,86.0641632080078,85.9945373535156,85.9541015625,85.9216842651367,85.8402328491211,85.7594757080078,85.6783218383789,85.41064453125,85.2121047973633,85.1224594116211,85.0885772705078,85.1214828491211,85.16015625,85.1590881347656,85.1263656616211,85.0691375732422,84.8550796508789,84.796875,84.7593765258789,84.7142562866211,84.6767578125,84.6505813598633,84.4654312133789,84.4107437133789,84.3121109008789,84.2287139892578,84.1755828857422,84.1278305053711,84.1013717651367,84.02197265625,83.9359359741211,83.7904281616211,83.6710968017578,83.58349609375,83.4566421508789,83.3551788330078,83.2351608276367,83.1554718017578,83.0139617919922,82.8543014526367,82.6408233642578,82.4865188598633,82.220703125,82.1589889526367,82.1353530883789,82.0989303588867,82.0433578491211,81.8548889160156,81.6418914794922,81.4171905517578,81.2550811767578,81.1771469116211,81.1103515625,81.0555725097656,81.01025390625,80.9854507446289,80.87353515625,80.7467727661133,80.68212890625,80.60888671875,80.541015625,80.4095687866211,80.2609405517578,80.1912078857422,80.1862258911133,80.2071304321289,80.1943359375,80.1494140625,80.0814514160156,79.9245071411133,79.9166030883789,79.8718795776367,79.7946243286133,79.6642608642578,79.5654296875,79.4931640625,79.3884735107422,79.36962890625,79.3387680053711,79.2326202392578,79.1071243286133,79.0437469482422,79.0111312866211,78.9739227294922,78.9459991455078,78.8995132446289,78.8445358276367,78.7916030883789,78.7578125,78.7435607910156,78.7585983276367,78.7267532348633,78.7550811767578,78.8029327392578,78.75390625,78.6934585571289,78.68701171875,78.7197265625,78.7354507446289,78.7255859375,78.677734375,78.4959030151367,78.4861373901367,78.4552764892578,78.4413070678711,78.41748046875,78.3896484375,78.3917007446289,78.4124984741211,78.5263671875,78.6315383911133,78.7008819580078,78.7367172241211,78.7535171508789,78.7712860107422,78.837890625,78.9189453125,78.9976501464844,79.0669937133789,79.1273422241211,79.169921875,79.2193374633789,79.2190399169922,79.2165069580078,79.23388671875,79.2279281616211,79.20556640625,79.2095642089844,79.2022476196289,79.1455078125,79.1085968017578,79.1028289794922,79.1216735839844,79.1351623535156,79.1156234741211,79.1124954223633,79.0665054321289,79.0125961303711,78.9484329223633,78.9167022705078,78.8650360107422,78.8018493652344,78.7899398803711,78.7837905883789,78.76171875,78.7266616821289,78.7317428588867,78.7530288696289,78.9317321777344,78.9706039428711,78.9769515991211,78.9701156616211,78.9364242553711,78.8648376464844,78.7630844116211,78.6707992553711,78.5157241821289,78.3269577026367,78.2820358276367,78.2361373901367,78.1584930419922,78.0757827758789,78.01220703125,78.0091781616211,78.0474548339844,78.0426788330078,78.0094757080078,77.9458999633789,77.8949203491211,77.8515625,77.8109359741211,77.8025436401367,77.7994155883789,77.7240219116211,77.5725631713867,77.52001953125,77.4464874267578,77.2948226928711,77.0900344848633,76.87890625,76.7668914794922,76.7275390625,76.6318359375,76.5634765625,76.55126953125,76.5020523071289,76.3857421875,76.2516555786133,76.1778335571289,76.1478500366211,76.1033172607422,76.0708999633789,76.0104522705078,75.9451217651367,75.9123077392578,75.9048843383789,75.93408203125,75.9686508178711,75.9744110107422,75.9518508911133,75.9330062866211,75.8849563598633,75.8402328491211,75.7721633911133,75.6671829223633,75.57373046875,75.4602508544922,75.4242248535156,75.3768539428711,75.3466796875,75.1452178955078,75.0539093017578,74.9491195678711,74.8892593383789,74.8412094116211,74.7660140991211,74.6921920776367,74.6005859375,74.5414047241211,74.5264587402344,74.4979553222656,74.3761672973633,74.3721694946289,74.5589904785156,74.6689453125,74.7266616821289,74.7389602661133,74.7673797607422,74.8402328491211,74.8913116455078,74.9181671142578,75.0083999633789,75.0790023803711,75.1187515258789,75.0974578857422,74.9864273071289,74.9158248901367,74.8942413330078,74.9123077392578,74.9382781982422,74.9212875366211,74.9002914428711,74.8908157348633,74.8424835205078,74.7896499633789,74.7750930786133,74.7720718383789,74.8359375,74.8123016357422,74.7450180053711,74.5140686035156,74.2774429321289,74.1873016357422,74.13134765625,74.0653381347656,74.0255813598633,73.9700164794922,73.869140625,73.8016662597656,73.7541046142578,73.716796875,73.6960983276367,73.7068405151367,73.72998046875,73.7945327758789,73.8052749633789,73.7956008911133,73.7437515258789,73.6904296875,73.6073226928711,73.6231384277344,73.6363296508789,73.6316375732422,73.7157287597656,73.8229522705078,73.8727493286133,73.9071273803711,73.9146499633789,73.8825149536133,73.8397445678711,73.8353500366211,73.8562469482422,73.8845672607422,73.9387741088867,73.9916000366211,74.0205078125,74.0851593017578,74.24267578125,74.4119110107422,74.6130905151367,74.6798782348633,74.7677688598633,74.8304672241211,74.841796875,74.80126953125,74.8111267089844,74.8351593017578,74.8656234741211,75.0044937133789,75.111328125,75.2410125732422,75.5207977294922,75.5555648803711,75.58349609375,75.6173782348633,75.6559600830078,75.6771469116211,75.8719711303711,76.0042953491211,76.0623016357422,76.1566467285156,76.2060546875,76.25830078125,76.3185501098633,76.3963851928711,76.4801788330078,76.5208969116211,76.5779266357422,76.6221694946289,76.6398468017578,76.6611328125,76.7083969116211,76.8240203857422,76.90771484375,76.9866180419922,77.1820297241211,77.2839889526367,77.5817337036133,77.7193374633789,77.8152313232422,77.9564437866211,78.1234359741211,78.3462829589844,78.3488235473633,78.3624038696289,78.44287109375,78.5431671142578,78.7425765991211,79.1484375,79.2935485839844,79.3543930053711,79.50390625,79.76611328125,79.8404312133789,79.90966796875,80.2162094116211,80.2351531982422,80.2461929321289,80.2291946411133,80.2093734741211,80.2330093383789,80.2590789794922,80.2550811767578,80.2057647705078,80.1792984008789,80.1619110107422,80.1650390625,80.2022476196289,80.2502899169922,80.4240188598633,80.5389709472656,80.5437469482422,80.45068359375,80.3833999633789,80.3712844848633,80.37451171875,80.3902359008789,80.5070266723633,80.6169891357422,80.7511749267578,80.7777404785156,80.7857437133789,80.7570266723633,80.7297821044922,80.6677703857422,80.6654281616211,80.7038040161133,80.6507797241211,80.5934600830078,80.4960021972656,80.4315414428711,80.3957977294922,80.3552703857422,80.3589859008789,80.3653335571289,80.3548812866211,80.3363265991211,80.3550796508789,80.3910140991211,80.3814468383789,80.4005889892578,80.4554672241211,80.4815368652344,80.4554672241211,80.36083984375,80.2550811767578,80.1278305053711,79.9971694946289,79.93212890625,79.8752899169922,79.8718795776367,79.9501953125,80.0591812133789,80.2282180786133,80.4149398803711,80.5091781616211,80.634765625,80.7800827026367,80.8533172607422,81.0403289794922,81.3347702026367,81.60205078125,81.6919937133789,81.7588882446289,81.7896499633789,81.8674774169922,81.9449234008789,81.9892578125,82.1227493286133,82.2666015625,82.3234405517578,82.3966751098633,82.4787139892578,82.521484375,82.5589828491211,82.5969696044922,82.62109375,82.6257781982422,82.6116256713867,82.58251953125,82.45166015625,82.32666015625,82.3122100830078,82.3152313232422,82.34814453125,82.4296875,82.51171875,82.5550765991211,82.6921844482422,82.8000030517578,82.9748992919922,83.0040969848633,83.0201187133789,83.0294952392578,83.09033203125,83.1930618286133,83.4435501098633,83.6340789794922,83.7139663696289,83.8326110839844,84.0160217285156,84.1220703125,84.2151412963867,84.3388671875,84.5324249267578,84.59228515625,84.6666030883789,84.7195281982422,84.7459945678711,84.7861328125,84.8582000732422,85.01220703125,85.1105499267578,85.2334899902344,85.3553695678711,85.4847640991211,85.5296859741211,85.5772399902344,85.6566467285156,85.6698226928711,85.6417922973633,85.5866241455078,85.5882797241211,85.5616226196289,85.5259780883789,85.5623092651367,85.6263656616211,85.6515655517578,85.6921920776367,85.7494125366211,85.8298873901367,86.05615234375,86.265625,86.37255859375,86.4832992553711,86.5494079589844,86.6637725830078,86.7179641723633,86.7578125,86.7286148071289,86.7531204223633,86.8083038330078,86.8859329223633,86.93798828125,87.0485305786133,87.22998046875,87.3228454589844,87.4167022705078,87.4761734008789,87.5158233642578,87.5765609741211,87.6683578491211,87.7625045776367,87.8142623901367,87.8251953125,87.8163070678711,87.8346710205078,87.8721694946289,87.85986328125,87.8068389892578,87.7546844482422,87.7431640625,87.8091812133789,87.8318328857422,87.9421920776367,88.0279312133789,88.06005859375,88.0501937866211,88.0106430053711,87.9722595214844,87.9673767089844,87.9796905517578,88.0625991821289,88.158203125,88.3099594116211,88.4139633178711,88.51708984375,88.5667953491211,88.5759735107422,88.6818389892578,88.8382797241211,88.9177703857422,88.9710922241211,89.0476531982422,89.1156234741211,89.1962890625,89.3298873901367,89.4792022705078,89.5609359741211,89.6384735107422,89.6931610107422,89.7255859375,89.7781219482422,89.8313446044922,89.9104537963867,89.9586944580078,90.0279312133789,90.0539093017578,90.0666046142578,90.1032180786133,90.1910171508789,90.3132858276367,90.3306655883789,90.3474578857422,90.3806610107422,90.4251937866211,90.4674835205078,90.4764633178711,90.4961853027344,90.5529251098633,90.6433563232422,90.7155227661133,90.7990264892578,90.8699264526367,90.9105453491211,90.9857406616211,90.9978485107422,91.0043029785156,91.0289077758789,91.0338897705078,90.9714889526367,90.9182586669922,90.9115219116211,90.9475631713867,90.9967803955078,91.0017623901367,90.9597702026367,90.8871078491211,90.8524475097656,90.7958984375,90.7096633911133,90.6707000732422,90.6618118286133,90.6944351196289,90.7496109008789,90.76318359375,90.8532257080078,90.8772430419922,90.9139633178711,90.95361328125,91.0500030517578,91.1376953125,91.2217788696289,91.3121109008789,91.4410171508789,91.5100631713867,91.5843734741211,91.73779296875,91.8528366088867,92.02978515625,92.1726608276367,92.423828125,92.5789031982422,92.7878875732422,92.916015625,93.2943420410156,93.5162124633789,93.6564483642578,93.7552719116211,93.8681640625,93.9579162597656,94.1993179321289,94.36474609375,94.4943389892578,94.7120132446289,94.8660125732422,95.0498046875,95.3502960205078,95.3667984008789,95.3436508178711,95.3255844116211,95.3255844116211,95.3564453125,95.4712905883789,95.5255813598633,95.5671844482422,95.5912094116211,95.6873016357422,95.8419952392578,95.8595657348633,95.9124984741211,96.0802688598633,96.16845703125,96.2995147705078,96.3424835205078,96.3523483276367,96.3854522705078,96.6252899169922,96.8330078125,97.2056579589844,97.7189407348633,98.2482376098633,98.71630859375,98.9468765258789,99.4678726196289,99.7574157714844,99.9837951660156,100.086326599121,100.51904296875,100.772560119629,101.091987609863,101.313766479492,101.495315551758,101.5791015625,101.659965515137,101.7138671875,101.8798828125,101.972946166992,102.156639099121,102.5751953125,102.806831359863,103.072853088379,103.247848510742,103.44970703125,103.711135864258,103.997268676758,104.30517578125,104.498237609863,104.498237609863,104.773635864258,104.8603515625,104.982032775879,105.050582885742,105.115432739258,105.197067260742,105.314353942871,105.51708984375,105.56640625,105.867576599121,106.317184448242,106.518745422363,106.5791015625,106.693161010742,106.77001953125,106.906051635742,107.090721130371,107.292381286621,107.748725891113,107.805953979492,108.062301635742,108.171195983887,108.333984375,108.546485900879,108.687301635742,108.87451171875,109.131645202637,109.33984375,109.443168640137,109.595504760742,109.698051452637,109.858787536621,110.058006286621,110.196868896484,110.288864135742,110.400390625,110.429588317871,110.461715698242,110.520896911621,110.627532958984,110.708595275879,110.74853515625,110.839553833008,110.913284301758,111.007225036621,111.086524963379,111.186820983887,111.451072692871,111.503517150879,111.54736328125,111.640823364258,111.7197265625,111.771095275879,111.878120422363,111.933204650879,111.94287109375,111.931739807129,111.880271911621,111.8369140625,111.683792114258,111.602630615234,111.519729614258,111.486228942871,111.429588317871,111.402244567871,111.410934448242,111.489448547363,111.514739990234,111.547462463379,111.621284484863,111.681449890137,111.751075744629,111.898048400879,112.032615661621,112.112892150879,112.292083740234,112.411331176758,112.499313354492,112.596771240234,112.706741333008,113.049407958984,113.196098327637,113.300971984863,113.455665588379,113.507911682129,113.587005615234,113.652633666992,113.752151489258,113.877052307129,113.930862426758,114.0302734375,114.080268859863,114.167381286621,114.281051635742,114.419136047363,114.4873046875,114.502250671387,114.517181396484,114.560150146484,114.644325256348,114.738761901855,114.919242858887,115.162605285645,115.217475891113,115.439460754395,115.539451599121,115.681053161621,115.789161682129,115.934181213379,116.039543151855,116.109870910645,116.197647094727,116.240623474121,116.229095458984,116.212989807129,116.264549255371,116.357612609863,116.44482421875,116.516700744629,116.562599182129,116.619331359863,116.688865661621,116.787017822266,116.859085083008,116.978805541992,117.155952453613,117.269050598145,117.333404541016,117.356941223145,117.356353759766,117.392181396484,117.405570983887,117.438079833984,117.546875,117.620498657227,117.671096801758,117.741203308105,117.813484191895,117.910446166992,118.071296691895,118.156837463379,118.308700561523,118.404388427734,118.580474853516,118.648735046387,118.722946166992,118.790336608887,118.843940734863,118.957130432129,119.028518676758,119.162101745605,119.331832885742,119.474021911621,119.620216369629,119.706649780273,119.747467041016,119.867179870605,119.895896911621,119.884178161621,119.897850036621,119.862686157227,119.788475036621,119.759864807129,119.757232666016,119.711128234863,119.600189208984,119.526954650879,119.376663208008,119.325973510742,119.30859375,119.290824890137,119.235252380371,119.162406921387,119.122955322266,119.097267150879,119.081932067871,119.017578125,118.953125,118.880279541016,118.759956359863,118.690536499023,118.567764282227,118.498443603516,118.239646911621,118.147071838379,118.041900634766,117.979194641113,117.840431213379,117.768363952637,117.676658630371,117.555366516113,117.455078125,117.383987426758,117.350776672363,117.285934448242,117.197074890137,117.069732666016,116.95166015625,116.901176452637,116.760543823242,116.651954650879,116.513481140137,116.378227233887,116.317192077637,116.231147766113,116.074806213379,115.993850708008,115.898239135742,115.811714172363,115.711723327637,115.616401672363,115.5576171875,115.525093078613,115.639450073242,115.785545349121,115.796585083008,115.791702270508,115.820503234863,115.953811645508,116.025489807129,116.034378051758,116.098243713379,116.159660339355,116.243354797363,116.402145385742,116.589744567871,116.683303833008,116.888961791992,117.021682739258,117.24560546875,117.477149963379,117.698432922363,117.812591552734,117.873435974121,118.186614990234,118.451560974121,118.755950927734,118.9794921875,119.1474609375,119.259857177734,119.326065063477,119.346282958984,119.301559448242,119.19189453125,119.163673400879,119.216705322266,119.255851745605,119.280662536621,119.344047546387,119.445709228516,119.501754760742,119.512298583984,119.573432922363,119.684951782227,119.746002197266,119.756645202637,119.813186645508,119.966987609863,120.06689453125,120.237007141113,120.510543823242,120.681449890137,120.749809265137,120.744537353516,120.665428161621,120.650390625,120.699226379395,120.656150817871,120.521095275879,120.360061645508,120.172752380371,120.067581176758,120.044334411621,120.094535827637,120.218162536621,120.421287536621,120.7041015625,120.985450744629,121.405464172363,121.743942260742,122.0888671875,122.337799072266,122.380172729492,122.515823364258,122.744728088379,122.957611083984,123.154106140137,123.3095703125,123.424018859863,123.489448547363,123.534759521484,123.559761047363,123.607818603516,123.740913391113,123.994720458984,124.154304504395,124.219924926758,124.291412353516,124.369140625,124.465919494629,124.639846801758,124.812309265137,124.882125854492,124.906646728516,124.970893859863,125.074996948242,125.225593566895,125.422462463379,125.545997619629,125.595993041992,125.649017333984,125.691696166992,125.6953125,125.680770874023,125.728126525879,125.782814025879,125.871879577637,125.941596984863,126.004302978516,126.048149108887,126.056060791016,126.060157775879,126.047073364258,126.023246765137,126.016021728516,126.0458984375,126.156639099121,126.194435119629,126.202934265137,126.237594604492,126.312889099121,126.341697692871,126.324211120605,126.346290588379,126.383499145508,126.391502380371,126.394821166992,126.455574035645,126.468070983887,126.510551452637,126.653717041016,126.700790405273,126.688667297363,126.709175109863,126.774513244629,126.805465698242,126.801750183105,126.827339172363,126.847763061523,126.833786010742,126.854400634766,126.887687683105,126.911529541016,126.9248046875,127.020416259766,127.1982421875,127.307037353516,127.346870422363,127.34716796875,127.308197021484,127.306053161621,127.3408203125,127.351173400879,127.337211608887,127.395309448242,127.590225219727,127.512306213379,127.491790771484,127.502449035645,127.55078125,127.636711120605,127.690139770508,127.711135864258,127.814262390137,127.999603271484,128.237121582031,128.526763916016,128.703994750977,128.76904296875,128.791015625,128.770309448242,128.8193359375,128.938278198242,129.020309448242,129.065139770508,129.1201171875,129.185165405273,129.248428344727,129.309860229492,129.35009765625,129.384674072266,129.440719604492,129.498153686523,129.53369140625,129.591400146484,129.671096801758,129.792587280273,130.037109375,130.195999145508,130.355270385742,130.553131103516,130.6171875,130.565628051758,130.552154541016,130.597259521484,130.6591796875,130.746871948242,130.763473510742,130.804290771484,130.787200927734,130.712097167969,130.732620239258,130.8486328125,130.915420532227,130.932815551758,130.9619140625,131.002731323242,131.121871948242,131.3193359375,131.464263916016,131.556732177734,131.785247802734,132.149810791016,132.380172729492,132.476272583008,132.561904907227,132.636917114258,132.707229614258,132.772857666016,132.877151489258,133.020126342773,133.14404296875,133.301162719727,133.468353271484,133.5732421875,133.671768188477,133.842193603516,134.205856323242,134.293365478516,134.3349609375,134.456146240234,134.563583374023,134.665237426758,134.680847167969,134.669342041016,134.647262573242,134.605361938477,134.566009521484,134.59130859375,134.650299072266,134.698638916016,134.752334594727,134.728134155273,134.69580078125,134.59619140625,134.541900634766,134.483505249023,134.382522583008,134.339447021484,134.290817260742,134.260055541992,134.225204467773,134.167678833008,134.162994384766,134.189254760742,134.2021484375,134.136917114258,134.08642578125,134.071380615234,134.046005249023,134.03857421875,134.022659301758,133.95751953125,133.866592407227,133.88671875,133.902725219727,133.880279541016,133.874801635742,133.861328125,133.832809448242,133.750198364258,133.70068359375,133.711135864258,133.685729980469,133.647857666016,133.608001708984,133.551162719727,133.513076782227,133.484664916992,133.475769042969,133.449127197266,133.465621948242,133.436416625977,133.35546875,133.3095703125,133.266983032227,133.18603515625,133.113372802734,133.096878051758,133.113479614258,133.01171875,132.93603515625,132.888778686523,132.838668823242,132.72314453125,132.665618896484,132.549026489258,132.362991333008,132.181350708008,132.0673828125,131.9775390625,131.909271240234,131.851852416992,131.794921875,131.742095947266,131.654006958008,131.613967895508,131.578720092773,131.487487792969,131.446884155273,131.268264770508,131.227935791016,131.082321166992,131.033020019531,130.981643676758,130.9677734375,131.00390625,131.060653686523,131.0869140625,131.125778198242,131.255264282227,131.213272094727,131.174224853516,131.18359375,131.180084228516,131.182418823242,131.209182739258,131.243942260742,131.261810302734,131.25732421875,131.239349365234,131.2119140625,131.175582885742,131.135543823242,131.108978271484,131.086135864258,131.08349609375,131.068557739258,131.005569458008,130.94287109375,130.868560791016,130.803314208984,130.722457885742,130.577255249023,130.492965698242,130.452728271484,130.4248046875,130.419921875,130.439270019531,130.520614624023,130.576568603516,130.58447265625,130.526947021484],[-3.08671903610229,-3.114013671875,-3.24638700485229,-3.21489262580872,-3.08671903610229],[-5.52353477478027,-5.46127986907959,-5.38227558135986,-5.26230430603027,-5.17529296875,-5.09985399246216,-5.04931688308716,-4.99404287338257,-4.96992206573486,-4.88271474838257,-4.814453125,-4.72177743911743,-4.62583017349243,-4.526611328125,-4.48027324676514,-4.40620136260986,-4.33222675323486,-4.26718759536743,-4.18115186691284,-3.96347689628601,-3.87763667106628,-3.79062533378601,-3.58115220069885,-3.38627910614014,-3.28969740867615,-3.22353506088257,-3.16069340705872,-3.09580087661743,-3.04262709617615,-2.98828148841858,-2.94814467430115,-2.90087914466858,-2.87514638900757,-2.81674814224243,-2.76660180091858,-2.71718764305115,-2.69584941864014,-2.68613266944885,-2.70576167106628,-2.70180654525757,-2.67421865463257,-2.689208984375,-2.74667954444885,-2.74692416191101,-2.68989253044128,-2.64921879768372,-2.62490248680115,-2.60039067268372,-2.59799790382385,-2.55688500404358,-2.50585961341858,-2.53828144073486,-2.582763671875,-2.61171865463257,-2.61997103691101,-2.60097646713257,-2.61337924003601,-2.66884779930115,-2.78974580764771,-2.79814481735229,-2.83012676239014,-2.85688495635986,-2.89633774757385,-2.95908164978027,-2.98232436180115,-2.98579072952271,-3.01015639305115,-3.03769493103027,-3.16889667510986,-3.23579120635986,-3.22714877128601,-3.22412133216858,-3.24389624595642,-3.24028325080872,-3.22402334213257,-3.20058584213257,-3.10556650161743,-3.05615234375,-3.02529263496399,-2.998291015625,-2.97280287742615,-2.96225595474243,-2.82119154930115,-2.79365205764771,-2.75498056411743,-2.76191377639771,-2.78959965705872,-2.78867220878601,-2.79521489143372,-2.81567406654358,-2.89472651481628,-2.94833993911743,-3.01914048194885,-3.02587914466858,-3.06396460533142,-3.168701171875,-3.15141630172729,-3.19995093345642,-3.23759770393372,-3.31201171875,-3.34755849838257,-3.87060546875,-3.98417949676514,-4.12016582489014,-4.35727500915527,-4.55283212661743,-4.60888719558716,-4.11518573760986,-4.06206035614014,-4.03720760345459,-4.66152334213257,-4.89970684051514,-4.97011756896973,-5.023681640625,-5.28237295150757,-5.33544969558716,-5.36752939224243,-5.26577138900757,-5.10488271713257,-5.06181621551514,-5.56474590301514,-5.91376924514771,-6.06171846389771,-6.54843807220459,-6.84516572952271,-6.92290067672729,-7.05795907974243,-7.23139619827271,-7.42607450485229,-7.54497098922729,-7.57158231735229,-7.57465839385986,-7.59121131896973,-7.58505916595459,-7.5693359375,-7.56889629364014,-7.509765625,-7.494140625,-7.48520517349243,-7.42983436584473,-7.42890548706055,-7.41245079040527,-7.39990186691284,-7.42373085021973,-7.45439481735229,-7.46943330764771,-7.48281288146973,-7.513916015625,-7.63613271713257,-7.73037147521973,-7.79653310775757,-7.80092716217041,-7.833251953125,-7.85551738739014,-7.88862323760986,-7.98159170150757,-8.06894588470459,-8.13100528717041,-8.203857421875,-8.287109375,-8.34487342834473,-8.39931678771973,-8.44990253448486,-8.49033260345459,-8.53955078125,-8.587890625,-8.60356521606445,-8.40122032165527,-8.33256816864014,-8.32509708404541,-8.32451152801514,-8.30234336853027,-8.296630859375,-8.40874004364014,-8.43715858459473,-8.46728515625,-8.48642539978027,-8.42998123168945,-8.35175800323486,-8.23188495635986,-8.20595741271973,-8.11542987823486,-8.11782264709473,-8.12685489654541,-8.07382869720459,-8.03173828125,-8.00986385345459,-8.01674842834473,-8.048583984375,-8.09052753448486,-8.140625,-8.21713829040527,-8.256103515625,-8.244140625,-8.23696327209473,-8.2099609375,-8.16777324676514,-8.04912090301514,-7.953125,-7.86875009536743,-7.82358360290527,-7.78740262985229,-7.73896455764771,-7.69609403610229,-7.68120098114014,-7.69096612930298,-7.71958065032959,-7.78403377532959,-7.95097637176514,-7.95498037338257,-7.93818330764771,-7.902099609375,-7.77797889709473,-7.7998046875,-7.83940410614014,-7.91806650161743,-7.90000009536743,-7.89619112014771,-7.96269512176514,-8.031005859375,-8.08867168426514,-8.13696193695068,-8.14604473114014,-8.14585018157959,-8.15517616271973,-8.13662052154541,-8.07783222198486,-8.01352500915527,-7.99062538146973,-7.96093702316284,-7.88408231735229,-7.81420946121216,-7.74907255172729,-7.6611328125,-7.56210947036743,-7.53281259536743,-7.49794960021973,-7.45654296875,-7.41479444503784,-7.38505840301514,-7.36318397521973,-7.18232488632202,-7.10488271713257,-7.03974628448486,-7.01708984375,-6.98945283889771,-6.96816396713257,-6.96381855010986,-6.99174785614014,-6.9794921875,-6.95034217834473,-6.90380859375,-6.83364248275757,-6.75322294235229,-6.69326114654541,-6.66933631896973,-6.69199228286743,-6.68613290786743,-6.67636728286743,-6.65415000915527,-6.56459999084473,-6.48261690139771,-6.42392587661743,-6.40751934051514,-6.4326171875,-6.42587852478027,-6.40415048599243,-6.36562538146973,-6.26113271713257,-6.250244140625,-6.23066377639771,-6.23974561691284,-6.2177734375,-6.190673828125,-6.192626953125,-6.21499013900757,-6.24130821228027,-6.23837900161743,-6.19687461853027,-6.1171875,-6.03457021713257,-5.98867177963257,-5.940673828125,-5.90756845474243,-5.89619112014771,-5.84384775161743,-5.69428730010986,-5.55659151077271,-5.52353477478027],[15.4800777435303,15.3791017532349,15.2458992004395,15.2067384719849,15.1858396530151,15.1571292877197,15.0863285064697,15.0345697402954,14.9827146530151,14.8619146347046,14.7803707122803,14.7640619277954,14.7392578125,14.6995115280151,14.5593748092651,14.5121097564697,14.4749994277954,14.4407224655151,14.4311513900757,14.4638662338257,14.5031242370605,14.54248046875,14.5772457122803,14.5988273620605,14.6168947219849,14.6168947219849,14.5835943222046,14.5843744277954,14.5680656433105,14.56298828125,14.5735349655151,14.6017580032349,14.640625,14.6617183685303,14.708984375,14.7312498092651,14.7704105377197,14.8935556411743,15.0227537155151,15.0635738372803,15.0875005722046,15.1369152069092,15.1358394622803,15.1154308319092,15.0673837661743,15.0348634719849,15.0621089935303,15.1287117004395,15.2398443222046,15.3601560592651,15.4583978652954,15.5808601379395,15.6765632629395,15.7749996185303,15.8493175506592,15.9048824310303,15.9287118911743,15.9580078125,16.0082035064697,16.0634765625,16.0824222564697,16.0592765808105,16.0821304321289,16.08349609375,16.1018543243408,16.0955085754395,16.1067390441895,16.1361331939697,16.1833992004395,16.1826171875,16.1765613555908,16.11572265625,16.080078125,16.0696277618408,16.087890625,16.1349601745605,16.1361331939697,16.1195316314697,16.09033203125,16.0593738555908,15.9751958847046,15.8816413879395,15.7416019439697,15.6002941131592,15.41748046875,15.3387689590454,15.2824220657349,15.2035160064697,15.1600589752197,15.0996103286743,15.0578126907349,15.0064449310303,14.9024410247803,14.8927736282349,14.8749990463257,14.7628908157349,14.7283191680908,14.7132806777954,14.6691408157349,14.5789070129395,14.4840822219849,14.2870111465454,14.0343751907349,13.7727537155151,13.5334968566895,13.2935543060303,13.2699222564697,13.2203130722046,13.130859375,12.8674802780151,12.6657228469849,12.6013669967651,12.5297861099243,12.361328125,12.1534185409546,12.1061525344849,11.9397459030151,11.5589847564697,11.3484382629395,11.3533201217651,11.3399410247803,11.3287115097046,11.0965824127197,10.7909173965454,10.5022459030151,10.3070316314697,9.97988224029541,9.8701171875,9.8369140625,9.83037090301514,9.82617092132568,9.80078125,9.82177734375,9.86757850646973,9.88544940948486,9.94843769073486,9.9150390625,9.87617206573486,9.67207050323486,9.76572227478027,9.64238262176514,9.61591720581055,9.55615139007568,9.5927734375,9.62812519073486,9.73964786529541,9.73613262176514,9.63994121551514,9.64921951293945,9.68886661529541,9.66953086853027,9.60039138793945,9.55058574676514,9.51181602478027,9.48369121551514,9.50078105926514,9.46201229095459,9.42529296875,9.3623046875,9.31093788146973,9.29736328125,9.24912071228027,9.11386680603027,9.00009727478027,8.97705078125,8.93203163146973,8.91357421875,8.90283203125,8.91826152801514,8.88945388793945,8.8564453125,8.80712890625,8.76191425323486,8.70790958404541,8.66035175323486,8.68964862823486,8.65625,8.57441425323486,8.53955078125,8.53281307220459,8.57050800323486,8.55585956573486,8.58515548706055,8.64052772521973,8.71562480926514,8.80097675323486,8.85917949676514,8.89882850646973,8.93505859375,8.99716758728027,9.06015586853027,9.23876953125,9.37333965301514,9.44218730926514,9.490234375,9.57402324676514,9.65996074676514,9.7255859375,9.77988338470459,9.82070255279541,9.87421894073486,10.0388679504395,10.1435546875,10.1677732467651,10.185546875,10.2054691314697,10.2930660247803,10.4131832122803,10.4823236465454,10.51904296875,10.5563478469849,10.578125,10.6062498092651,10.7375974655151,10.8464841842651,10.9541997909546,11.0086917877197,11.0325193405151,11.0796880722046,11.1064453125,11.1533203125,11.2373046875,11.3246088027954,11.4017581939697,11.4775390625,11.5291013717651,11.5516595840454,11.56298828125,11.580078125,11.6575193405151,11.7870111465454,11.8614253997803,11.8547849655151,11.80859375,11.7673826217651,11.8091802597046,11.8524417877197,12.0160160064697,12.0166015625,12.0251951217651,12.1559562683105,12.2311525344849,12.2333984375,12.3113279342651,12.4035158157349,12.5827140808105,12.6515626907349,12.7311525344849,12.7822265625,12.8065433502197,12.8244142532349,12.85595703125,12.8756837844849,12.9294929504395,13.0194330215454,13.1754875183105,13.19873046875,13.22119140625,13.23876953125,13.2437496185303,13.2498044967651,13.2699222564697,13.41455078125,13.478515625,13.5353517532349,13.6999025344849,13.89208984375,13.9814453125,14.0567388534546,14.1432619094849,14.2023439407349,14.4094724655151,14.4960927963257,14.5597658157349,14.5753917694092,14.5816411972046,14.5618162155151,14.5973625183105,14.6181640625,14.6271486282349,14.6197261810303,14.5870113372803,14.5809564590454,14.5189447402954,14.4154300689697,14.2728519439697,14.1974611282349,14.1848630905151,14.1776361465454,14.1703128814697,14.1600580215454,14.06396484375,14.2448244094849,14.4617185592651,14.5162115097046,14.5447273254395,14.6232423782349,14.76123046875,14.8470706939697,14.8806629180908,14.9567384719849,14.9738292694092,15.0598640441895,15.0812501907349,15.0876951217651,15.0780267715454,15.1219720840454,15.0554685592651,15.0357427597046,15.0298824310303,15.0686521530151,15.1322259902954,15.2009773254395,15.2760753631592,15.3999032974243,15.5319337844849,15.6548824310303,15.540919303894,15.3200187683105,15.1931648254395,15.1327142715454,15.0715827941895,14.8358402252197,14.5979499816895,14.3772459030151,14.2432613372803,14.1397457122803,14.0559568405151,13.9772462844849,14.0049800872803,14.0641603469849,14.1779289245605,14.2800779342651,14.3323240280151,14.5361328125,14.7328119277954,14.7712898254395,14.8262691497803,14.8607416152954,14.9679689407349,15.1162109375,15.2523431777954,15.3490228652954,15.4429693222046,15.4844722747803,15.5498046875,15.5578117370605,15.5526361465454,15.5324220657349,15.4800777435303],[27.4033203125,27.4392566680908,27.4910163879395,27.6641597747803,27.71923828125,27.7614250183105,27.7880859375,27.8416004180908,27.9166011810303,27.9806632995605,28.0198249816895,28.07861328125,28.1920890808105,28.2472648620605,28.31103515625,28.3671875,28.4275398254395,28.5248050689697,28.6395492553711,28.72705078125,28.9393539428711,29.0574207305908,29.1514663696289,29.2249011993408,29.3848628997803,29.4696292877197,29.5520496368408,29.6768550872803,29.7798843383789,29.8702144622803,29.9339828491211,30.0213871002197,30.1949214935303,30.4207019805908,30.50830078125,30.5369148254395,30.5535163879395,30.5593757629395,30.5867176055908,30.6476573944092,30.6999034881592,30.7572269439697,30.7969703674316,30.81689453125,30.8385734558105,30.8953113555908,30.9064445495605,30.8675785064697,30.8278331756592,30.7792987823486,30.7540016174316,30.7865238189697,30.8213863372803,30.8399391174316,30.8507823944092,30.8466777801514,30.76953125,30.7298812866211,30.7286148071289,30.8300762176514,30.9619140625,31.0036125183105,31.0453128814697,31.0821285247803,31.1375961303711,31.1763668060303,31.1914081573486,31.2363262176514,31.2740230560303,31.2560558319092,31.2527351379395,31.1587905883789,30.9425773620605,30.4781246185303,30.4778308868408,30.3210945129395,30.2401371002197,30.1829090118408,30.0473651885986,29.9428691864014,29.931640625,29.9238262176514,29.9344711303711,29.8854484558105,29.8146495819092,29.77783203125,29.7497081756592,29.7176742553711,29.6978511810303,29.6843738555908,29.6332035064697,29.6478538513184,29.6082038879395,29.6064453125,29.5900382995605,29.5619125366211,29.5640640258789,29.5799808502197,29.5769519805908,29.5377922058105,29.4679698944092,29.4019527435303,29.3516578674316,29.2681636810303,29.1965827941895,29.1432628631592,29.12939453125,29.1406269073486,29.1480464935303,29.1315422058105,29.1064453125,28.9895496368408,28.9126968383789,28.8763656616211,28.8576183319092,28.8914070129395,28.8939437866211,28.9217777252197,29.0143566131592,29.0141582489014,29.0165996551514,29.0647468566895,29.1532211303711,29.2244148254395,29.2260761260986,29.2123050689697,29.2100582122803,29.2171859741211,29.216796875,29.2118167877197,29.2232418060303,29.3313484191895,29.3791980743408,29.4032230377197,29.4041996002197,29.3675765991211,29.32568359375,29.3234367370605,29.3427734375,29.4201183319092,29.4764633178711,29.5037117004395,29.5423812866211,29.5941410064697,29.6070327758789,29.5963878631592,29.4908199310303,29.4800758361816,29.5062503814697,29.5408210754395,29.5906238555908,29.7096672058105,29.7981452941895,29.9618167877197,30.1062507629395,30.1618156433105,30.2126960754395,30.3131828308105,30.3745098114014,30.40673828125,30.4856452941895,30.5588874816895,30.6538066864014,30.7208995819092,30.7511711120605,30.5779304504395,30.3275394439697,30.0513668060303,29.7662105560303,29.4837894439697,29.2156257629395,28.9722652435303,28.8981437683105,28.9344749450684,28.9177742004395,28.8695297241211,28.7935543060303,28.7587890625,28.6812515258789,28.6165027618408,28.4842777252197,28.4006824493408,28.4001960754395,28.54052734375,28.6041984558105,28.6300792694092,28.62890625,28.62353515625,28.6171875,28.607421875,28.6455078125,28.6388664245605,28.5442390441895,28.5179691314697,28.4703121185303,28.4041996002197,28.3572254180908,28.3833999633789,28.4070320129395,28.4318351745605,28.4825210571289,28.5416011810303,28.5746097564697,28.7694320678711,28.8499984741211,28.9734363555908,29.0643577575684,29.1912117004395,29.34375,29.4275379180908,29.4855480194092,29.5048847198486,29.5022468566895,29.4919910430908,29.5082035064697,29.5597648620605,29.6919937133789,29.7496089935303,29.7951183319092,29.7953128814697,29.7955093383789,29.7956047058105,29.7958011627197,29.7960929870605,29.7962913513184,29.7964859008789,29.7953128814697,29.7751960754395,29.7226581573486,29.6517562866211,29.6476554870605,29.6302719116211,29.5971660614014,29.5541973114014,29.4814453125,29.3818340301514,29.2537097930908,29.2018566131592,29.1116218566895,29.0142574310303,28.9422836303711,28.9216804504395,28.8587894439697,28.7731456756592,28.7300777435303,28.6729488372803,28.6154289245605,28.5508785247803,28.51123046875,28.4744148254395,28.4514656066895,28.4128894805908,28.3577136993408,28.2373046875,28.06884765625,27.857421875,27.7568378448486,27.6443347930908,27.5738296508789,27.5333995819092,27.4870128631592,27.4236335754395,27.2380867004395,27.1963863372803,27.1591796875,27.0954093933105,27.0460948944092,27.0266609191895,26.9768543243408,26.9496097564697,26.9308586120605,26.8904285430908,26.8240242004395,26.7296867370605,26.5963859558105,26.4296894073486,26.3396472930908,26.0963878631592,26.0259761810303,25.9265632629395,25.8548812866211,25.6188468933105,25.5119152069092,25.4599609375,25.4133777618408,25.3494129180908,25.3207035064697,25.2826156616211,25.2917976379395,25.3193359375,25.2887687683105,25.2459964752197,25.1848640441895,25.0759754180908,24.8768558502197,24.8063468933105,24.7281246185303,24.6682605743408,24.5185546875,24.4666023254395,24.3779296875,24.3351554870605,24.3779296875,24.3962879180908,24.36572265625,24.3199214935303,24.1872081756592,24.1365222930908,24.1151371002197,24.0784187316895,24.0027351379395,23.9665031433105,23.9287109375,23.9073238372803,23.9011707305908,23.8338871002197,23.6963863372803,23.5599613189697,23.4639644622803,23.4001960754395,23.15673828125,23.0762691497803,22.8147468566895,22.66650390625,22.56103515625,22.4861316680908,22.3929672241211,22.31494140625,22.27880859375,22.2566413879395,22.2261714935303,22.2166996002197,22.1779308319092,22.2035160064697,22.2804679870605,22.3070316314697,22.283203125,22.2816410064697,22.3024406433105,22.2745113372803,22.19775390625,22.0891590118408,21.9486331939697,21.8566417694092,21.8131847381592,21.8294906616211,21.8718738555908,21.9053707122803,21.8958969116211,21.8008804321289,21.7800788879395,21.8335952758789,21.8416004180908,21.8060550689697,21.7816410064697,21.7510738372803,21.5108413696289,21.1903324127197,20.9109382629395,20.6078128814697,20.5583972930908,20.5358390808105,20.5369129180908,20.5987300872803,20.5900382995605,20.4822254180908,20.1900386810303,19.9974613189697,19.8751945495605,19.6603507995605,19.5276374816895,19.4837894439697,19.4874019622803,19.4798812866211,19.4193363189697,19.3716793060303,19.3699207305908,19.3408203125,19.1426753997803,18.9444332122803,18.8983402252197,18.6534175872803,18.5626945495605,18.4846668243408,18.3348617553711,18.1915035247803,18.0471687316895,18.0087890625,17.9130859375,17.77880859375,17.6433582305908,17.57958984375,17.5360336303711,17.4113292694092,17.2450180053711,17.1550788879395,17.12158203125,17.0637702941895,16.9847660064697,16.9520511627197,16.9658203125,16.91943359375,16.8130855560303,16.7429695129395,16.7093753814697,16.7009773254395,16.7177734375,16.697265625,16.6395511627197,16.6080074310303,16.5851554870605,16.537109375,16.4314460754395,16.3152351379395,16.0601558685303,15.7269525527954,15.4250001907349,15.0893545150757,14.7494144439697,14.6579103469849,14.3986330032349,14.1908206939697,14.11376953125,13.978515625,13.7645502090454,13.6490230560303,13.3714847564697,13.3464841842651,13.3026371002197,13.1843748092651,13.0681638717651,13.0033197402954,12.86083984375,12.7916011810303,12.6806640625,12.5145511627197,12.4529294967651,12.4117193222046,12.3150396347046,12.2404298782349,12.2136716842651,12.2552738189697,12.3860349655151,12.4845705032349,12.5037107467651,12.5189456939697,12.5223627090454,12.4874029159546,12.4532222747803,12.4514646530151,12.5027351379395,12.5735359191895,12.59619140625,12.6748046875,12.8296871185303,12.9474611282349,13.0573234558105,13.07275390625,13.08740234375,13.1366205215454,13.15234375,13.1764650344849,13.2196283340454,13.2972660064697,13.3757810592651,13.4149417877197,13.4784173965454,13.5516605377197,13.6595706939697,13.6853513717651,13.6994132995605,13.7076168060303,13.7170896530151,13.7390632629395,13.7780275344849,13.8495111465454,13.88232421875,13.94091796875,13.9784183502197,14.046875,14.1338872909546,14.22705078125,14.3162117004395,14.3583011627197,14.4029293060303,14.4427728652954,14.4498043060303,14.4099607467651,14.3654298782349,14.4029293060303,14.4119138717651,14.4107427597046,14.4409189224243,14.4616212844849,14.4939451217651,14.5576171875,14.6339845657349,14.7079105377197,14.779296875,14.912109375,15.1062507629395,15.2671871185303,15.3946285247803,15.48095703125,15.5259761810303,15.60009765625,15.7545890808105,15.8724613189697,15.9900398254395,16.1467781066895,16.1906261444092,16.2173824310303,16.2018566131592,16.1916027069092,16.21533203125,16.27392578125,16.43359375,16.5407218933105,16.6224613189697,16.7800769805908,16.84912109375,16.8798828125,16.9747066497803,17.1076183319092,17.27880859375,17.5428714752197,17.7528305053711,17.72412109375,17.7731437683105,17.8876953125,17.9251956939697,17.8857421875,17.9024410247803,18.01171875,18.0578136444092,18.0728511810303,18.0721683502197,18.2116203308105,18.3434562683105,18.4909191131592,18.5470695495605,18.6221675872803,18.6103515625,18.5966796875,18.6336917877197,18.6199226379395,18.5674800872803,18.5941410064697,18.6999015808105,18.8317375183105,19.0685539245605,19.3234386444092,19.5009765625,19.68603515625,19.8065433502197,19.8624992370605,20.0023441314697,20.2263679504395,20.3935546875,20.4865245819092,20.55810546875,20.6474609375,20.79296875,20.9557609558105,21.1255855560303,21.2297859191895,21.2683582305908,21.3501949310303,21.53759765625,21.68701171875,21.908203125,22.4221687316895,22.44970703125,22.4618167877197,22.5056648254395,22.6171875,22.7117195129395,22.7557621002197,22.8645496368408,22.9928722381592,23.1159172058105,23.2188472747803,23.3128910064697,23.4171867370605,23.5236320495605,23.6818351745605,23.8484363555908,23.99169921875,24.2277336120605,24.31982421875,24.4371089935303,24.7655258178711,24.9784183502197,25.0652351379395,25.2493171691895,25.2831058502197,25.4001941680908,25.5250988006592,25.7138671875,25.8199214935303,26.1735363006592,26.6326160430908,26.767578125,26.8220710754395,26.8701171875,27.0206050872803,27.0718746185303,27.1149406433105,27.4033203125],[18.6103515625,18.6221675872803,18.5470695495605,18.4909191131592,18.3434562683105,18.2116203308105,18.0721683502197,18.0728511810303,18.0578136444092,18.01171875,17.9024410247803,17.8857421875,17.9251956939697,17.8876953125,17.7731437683105,17.72412109375,17.7528305053711,17.5428714752197,17.27880859375,17.1076183319092,16.9747066497803,16.8798828125,16.84912109375,16.7800769805908,16.6224613189697,16.5407218933105,16.43359375,16.27392578125,16.21533203125,16.1916027069092,16.2018566131592,16.2173824310303,16.1906261444092,16.1467781066895,15.9900398254395,15.8724613189697,15.7545890808105,15.60009765625,15.5259761810303,15.48095703125,15.3946285247803,15.2671871185303,15.1062507629395,14.912109375,14.779296875,14.7079105377197,14.6339845657349,14.5576171875,14.4939451217651,14.4616212844849,14.4409189224243,14.4107427597046,14.4119138717651,14.4029293060303,14.3654298782349,14.4099607467651,14.4498043060303,14.4427728652954,14.4029293060303,14.3583011627197,14.3162117004395,14.22705078125,14.1338872909546,14.046875,13.9784183502197,13.94091796875,13.88232421875,13.8495111465454,13.7780275344849,13.7390632629395,13.7170896530151,13.7076168060303,13.6994132995605,13.6853513717651,13.6595706939697,13.5516605377197,13.4784173965454,13.4149417877197,13.3757810592651,13.2972660064697,13.2196283340454,13.1764650344849,13.15234375,13.1366205215454,13.08740234375,13.07275390625,13.0480470657349,12.9713869094849,12.8810548782349,12.84814453125,12.7982416152954,12.7194337844849,12.6416988372803,12.5014638900757,12.3845710754395,12.3740234375,12.3466796875,12.3079099655151,12.2042961120605,12.1670894622803,12.0775394439697,12.0183591842651,12.0027341842651,11.966796875,11.8932619094849,11.8207025527954,11.80126953125,11.7808589935303,11.7775382995605,11.6680669784546,11.3938474655151,11.3644533157349,11.1301755905151,11.1900386810303,11.2344722747803,11.2882804870605,11.5042972564697,11.5368165969849,11.6857423782349,11.7333984375,11.7864265441895,11.84912109375,11.8798828125,11.884765625,11.8394536972046,11.8329095840454,11.8647451400757,11.8828125,11.9292964935303,11.9341793060303,11.8850584030151,11.7843742370605,11.7154302597046,11.6890630722046,11.7080078125,11.7634773254395,11.7601566314697,11.7113285064697,11.6756830215454,11.6390619277954,11.5377931594849,11.55712890625,11.5945310592651,11.6034183502197,11.5751953125,11.5777339935303,11.60546875,11.6659173965454,11.7267580032349,11.8923826217651,11.9502925872803,11.9982423782349,12.064453125,12.4463863372803,12.4538087844849,12.4756841659546,12.478515625,12.4625968933105,12.4437503814697,12.4324226379395,12.43212890625,12.4686527252197,12.5904293060303,12.62841796875,12.7136716842651,12.7935543060303,12.8644533157349,12.91357421875,12.9919919967651,13.1585941314697,13.3573246002197,13.4649410247803,13.6185541152954,13.70556640625,13.7337894439697,13.7843751907349,13.8416013717651,13.8785161972046,13.8876953125,13.86181640625,13.8869132995605,13.9938478469849,14.0874013900757,14.1297845840454,14.1998043060303,14.2003898620605,14.1628904342651,14.1628904342651,14.2017583847046,14.2396488189697,14.25146484375,14.2883796691895,14.3585948944092,14.3839836120605,14.4232425689697,14.4029293060303,14.4029293060303,14.447265625,14.45556640625,14.4369144439697,14.4240245819092,14.4106435775757,14.4449224472046,14.4805660247803,14.4741220474243,14.4247074127197,14.36376953125,14.2067375183105,14.1483402252197,14.1028318405151,14.0694332122803,13.8980464935303,13.8600587844849,13.87548828125,13.890625,13.8845710754395,13.9151363372803,13.9496097564697,14.0252933502197,14.0655269622803,14.0874996185303,14.23095703125,14.2831058502197,14.3242177963257,14.3415031433105,14.390625,14.4344730377197,14.4391603469849,14.4298830032349,14.3864259719849,14.3344717025757,14.3030271530151,14.23974609375,14.1808595657349,14.0662107467651,13.8513669967651,13.72119140625,13.5233402252197,13.3723640441895,13.2741212844849,13.21630859375,13.1901359558105,13.2283201217651,13.2473640441895,13.2227535247803,13.1845703125,13.1626949310303,13.1721677780151,13.20947265625,13.2886714935303,13.2935543060303,13.5334968566895,13.7727537155151,14.0343751907349,14.2870111465454,14.4840822219849,14.5789070129395,14.6691408157349,14.7132806777954,14.7283191680908,14.7628908157349,14.8749990463257,14.8927736282349,14.9024410247803,15.0064449310303,15.0578126907349,15.0996103286743,15.1600589752197,15.2035160064697,15.2824220657349,15.3387689590454,15.41748046875,15.6002941131592,15.7416019439697,15.8816413879395,15.9751958847046,16.0593738555908,16.09033203125,16.1195316314697,16.1361331939697,16.1349601745605,16.087890625,16.0696277618408,16.080078125,16.11572265625,16.1765613555908,16.1826171875,16.1833992004395,16.2517566680908,16.3196296691895,16.4012699127197,16.4685535430908,16.4595699310303,16.4662113189697,16.4800777435303,16.4767570495605,16.4962902069092,16.5430660247803,16.5704116821289,16.6107425689697,16.67333984375,16.7643566131592,17.0025386810303,17.2247066497803,17.2984371185303,17.43798828125,17.4916019439697,17.5376949310303,17.806640625,17.88037109375,17.9071292877197,17.9479484558105,18.0107421875,18.072265625,18.111328125,18.1609363555908,18.1939449310303,18.2371082305908,18.3181648254395,18.4744129180908,18.4998054504395,18.5538101196289,18.6103515625],[-159.740524291992,-159.772567749023,-159.813079833984,-159.839599609375,-159.842483520508,-159.83203125,-159.810592651367,-159.768356323242,-159.739501953125,-159.736862182617,-159.740524291992],[-78.1137161254883,-78.1408233642578,-78.1924743652344,-78.2101058959961,-78.1784210205078,-78.1376419067383,-78.119140625,-78.1137161254883],[-71.3197250366211,-71.3555679321289,-71.4001922607422,-71.5360794067383,-71.719482421875,-71.9581069946289,-72.0123062133789,-72.2484817504883,-72.4460983276367,-72.5180206298828,-72.572265625,-72.6900863647461,-72.7391586303711,-72.8693313598633,-72.9403839111328,-72.9673843383789,-73.0065383911133,-73.0640640258789,-73.1412582397461,-73.2242660522461,-73.295654296875,-73.3563461303711,-73.3662109375,-73.3367156982422,-73.1931610107422,-73.13671875,-73.0584030151367,-73.00927734375,-72.9601593017578,-72.9044418334961,-72.8521041870117,-72.79638671875,-72.7255401611328,-72.6654281616211,-72.5257339477539,-72.4165573120117,-72.3903350830078,-72.3641586303711,-72.3576202392578,-72.3917007446289,-72.446044921875,-72.4595718383789,-72.4688949584961,-72.4789581298828,-72.4719772338867,-72.4429702758789,-72.3946304321289,-72.2963333129883,-72.2077178955078,-72.1566848754883,-72.0842819213867,-72.0066375732422,-71.8926696777344,-71.811279296875,-71.6208953857422,-71.4571304321289,-71.2178192138672,-71.1286163330078,-71.0132751464844,-70.8106918334961,-70.7371520996094,-70.6550750732422,-70.5355453491211,-70.4706573486328,-70.3874969482422,-70.26611328125,-70.1881408691406,-70.1291961669922,-70.0950241088867,-69.9041976928711,-69.7389602661133,-69.5948257446289,-69.4392547607422,-69.4271545410156,-69.3570785522461,-69.3108367919922,-69.2681655883789,-69.1945266723633,-69.0899353027344,-68.9372100830078,-68.7364730834961,-68.4717788696289,-68.14306640625,-67.9388656616211,-67.8591766357422,-67.7271423339844,-67.5680694580078,-67.4819793701172,-67.4715805053711,-67.4393539428711,-67.473876953125,-67.5751953125,-67.63134765625,-67.6422882080078,-67.6946258544922,-67.7884216308594,-67.8249053955078,-67.80419921875,-67.8143081665039,-67.8552703857422,-67.8552703857422,-67.8143081665039,-67.79541015625,-67.7986297607422,-67.783203125,-67.7323226928711,-67.66162109375,-67.6025390625,-67.5511245727539,-67.4986801147461,-67.3477096557617,-67.3111343383789,-67.3221664428711,-67.3362808227539,-67.3536148071289,-67.5148391723633,-67.8347702026367,-67.8612289428711,-67.8590850830078,-67.7664489746094,-67.667236328125,-67.6187057495117,-67.5966796875,-67.5680160522461,-67.5349655151367,-67.4864273071289,-67.3916015625,-67.312255859375,-67.2527313232422,-67.2108383178711,-67.1976089477539,-67.2152328491211,-67.1654815673828,-67.1314468383789,-67.1138229370117,-67.1314468383789,-67.0895538330078,-67.0438919067383,-66.9881286621094,-66.9815444946289,-66.9583511352539,-66.9311065673828,-66.8844680786133,-66.8955078125,-66.8760299682617,-67.0652313232422,-67.082275390625,-67.0936508178711,-67.0882797241211,-67.0901336669922,-67.1192398071289,-67.205810546875,-67.3206024169922,-67.3519592285156,-67.4004364013672,-67.457763671875,-67.4996566772461,-67.5560531616211,-67.6092300415039,-67.7118682861328,-67.8150939941406,-67.8755340576172,-67.9362335205078,-67.98974609375,-68.0328674316406,-68.0770492553711,-68.1302795410156,-68.1937942504883,-68.2183609008789,-68.2394485473633,-68.2559585571289,-68.2132873535156,-68.1765670776367,-68.2395553588867,-68.4434509277344,-68.678466796875,-68.9131851196289,-69.124267578125,-69.3197250366211,-69.3943328857422,-69.4701690673828,-69.5429153442383,-69.5812454223633,-69.6500473022461,-69.7396011352539,-69.7999496459961,-69.8485794067383,-69.8494644165039,-69.8507843017578,-69.8521499633789,-69.7981491088867,-69.7513198852539,-69.7169876098633,-69.6208953857422,-69.5675811767578,-69.5171432495117,-69.4703140258789,-69.4415054321289,-69.4027786254883,-69.3613739013672,-69.3118133544922,-69.2586898803711,-69.2244644165039,-69.19384765625,-69.1632308959961,-69.1650390625,-69.1659622192383,-69.1767578125,-69.1632308959961,-69.1533203125,-69.1560516357422,-69.174072265625,-69.2127914428711,-69.2541961669922,-69.2830047607422,-69.3055191040039,-69.3271484375,-69.358642578125,-69.3919906616211,-69.4207992553711,-69.4721221923828,-69.5270538330078,-69.5648422241211,-69.6036148071289,-69.6387252807617,-69.673828125,-69.7188949584961,-69.7567367553711,-69.80712890625,-69.862060546875,-69.9251022338867,-69.9854507446289,-70.0539016723633,-70.0579147338867,-70.0657196044922,-70.0709457397461,-70.0705108642578,-70.0440444946289,-69.9227600097656,-69.8279266357422,-69.7474594116211,-69.66748046875,-69.6339797973633,-69.6119155883789,-69.6008758544922,-69.592041015625,-69.6008758544922,-69.6207046508789,-69.6119155883789,-69.583251953125,-69.5744171142578,-69.5545883178711,-69.5435485839844,-69.519287109375,-69.4884262084961,-69.44873046875,-69.4443359375,-69.44873046875,-69.4491271972656,-69.4114303588867,-69.4002380371094,-69.4178695678711,-69.4349136352539,-69.4786071777344,-69.5064468383789,-69.5518569946289,-69.6046905517578,-69.6690444946289,-69.7326202392578,-69.7941360473633,-69.8497543334961,-69.9110336303711,-69.9481964111328,-69.9659194946289,-70.0171813964844,-70.0947723388672,-70.1675338745117,-70.1983871459961,-70.2402877807617,-70.2984390258789,-70.3395004272461,-70.3791961669922,-70.4210968017578,-70.48583984375,-70.5296859741211,-70.7061996459961,-70.735107421875,-70.6216735839844,-70.4189910888672,-70.2901382446289,-70.1470642089844,-70.0740280151367,-70.064453125,-70.0647430419922,-70.0958480834961,-70.1647415161133,-70.2444305419922,-70.2946243286133,-70.3641586303711,-70.418212890625,-70.5167999267578,-70.5758819580078,-70.6480026245117,-70.7053756713867,-70.91455078125,-70.9685516357422,-71.0272979736328,-71.1133804321289,-71.1963882446289,-71.3001022338867,-71.39697265625,-71.4474639892578,-71.49609375,-71.5594787597656,-71.6714859008789,-71.7525405883789,-71.802734375,-71.8672866821289,-71.9324722290039,-71.9842758178711,-72.0538101196289,-72.1368179321289,-72.2184600830078,-72.3007354736328,-72.3956069946289,-72.5006790161133,-72.5867233276367,-72.6253433227539,-72.66015625,-72.7141571044922,-72.8112335205078,-72.8871536254883,-72.9411163330078,-72.9896469116211,-73.0681610107422,-73.1544952392578,-73.1726531982422,-73.1602020263672,-73.1265106201172,-73.1452102661133,-73.1814956665039,-73.1969757080078,-73.2239685058594,-73.2664566040039,-73.3495101928711,-73.4402847290039,-73.4962921142578,-73.5252456665039,-73.4943389892578,-73.5213851928711,-73.5754852294922,-73.6102523803711,-73.664306640625,-73.7357406616211,-73.8071746826172,-73.8631820678711,-73.9269561767578,-73.98681640625,-74.0543899536133,-74.1807556152344,-74.2463912963867,-74.2838897705078,-74.3344268798828,-74.32861328125,-74.3531265258789,-74.3749008178711,-74.4178695678711,-74.4651870727539,-74.5138702392578,-74.5550765991211,-74.6163635253906,-74.691650390625,-74.75537109375,-74.7804718017578,-74.8017578125,-74.8375015258789,-74.8888168334961,-74.9453125,-75.0049743652344,-75.0546875,-75.1383819580078,-75.18408203125,-75.224609375,-75.2844696044922,-75.4639663696289,-75.6173324584961,-75.7766571044922,-75.8797912597656,-75.974853515625,-76.0261764526367,-76.0679168701172,-76.2706069946289,-76.31103515625,-76.38818359375,-76.4133758544922,-76.41796875,-76.4272994995117,-76.49462890625,-76.60302734375,-76.6785202026367,-76.72900390625,-76.7393035888672,-76.7677230834961,-76.829345703125,-76.9201126098633,-77.00244140625,-77.1141128540039,-77.1657180786133,-77.2926712036133,-77.3963394165039,-77.4227523803711,-77.4676742553711,-77.4813995361328,-77.526123046875,-77.601318359375,-77.6486282348633,-77.6731948852539,-77.702880859375,-77.8295440673828,-78.0370178222656,-78.1806640625,-78.3121109008789,-78.5115203857422,-78.587646484375,-78.681640625,-78.7371063232422,-78.828857421875,-78.8596725463867,-78.8884811401367,-79.0254364013672,-78.9576644897461,-78.79296875,-78.576904296875,-78.5504379272461,-78.6286087036133,-78.6170425415039,-78.5916976928711,-78.5347137451172,-78.46044921875,-78.4168930053711,-78.3428726196289,-78.296142578125,-78.1200256347656,-78.066650390625,-78.0301742553711,-77.9872055053711,-77.9322738647461,-77.9007797241211,-77.87451171875,-77.8135757446289,-77.8079605102539,-77.7766647338867,-77.6700210571289,-77.6710968017578,-77.7009735107422,-77.6936569213867,-77.6320343017578,-77.5591354370117,-77.520263671875,-77.4722213745117,-77.4171447753906,-77.3565444946289,-77.3244171142578,-77.2427749633789,-77.0768051147461,-77.1268615722656,-77.1666030883789,-77.2120132446289,-77.2635269165039,-77.2483901977539,-77.2780303955078,-77.3582000732422,-77.4272994995117,-77.4335479736328,-77.4044952392578,-77.4087371826172,-77.5206985473633,-77.5155334472656,-77.4458465576172,-77.4142608642578,-77.353515625,-77.3283233642578,-77.3136672973633,-77.2863311767578,-77.3065414428711,-77.3394546508789,-77.3667449951172,-77.3591766357422,-77.373291015625,-77.4017562866211,-77.534423828125,-77.3246154785156,-77.249267578125,-77.3446807861328,-77.4694366455078,-77.4730453491211,-77.4400939941406,-77.3982391357422,-77.35986328125,-77.3687973022461,-77.4388656616211,-77.5259704589844,-77.6021499633789,-77.6458511352539,-77.6809539794922,-77.8037109375,-77.9011688232422,-77.8283157348633,-77.7646942138672,-77.743896484375,-77.7687530517578,-77.7619171142578,-77.7469253540039,-77.7320327758789,-77.7063446044922,-77.6585922241211,-77.6186065673828,-77.5865707397461,-77.5382843017578,-77.3507766723633,-77.3627471923828,-77.3456115722656,-77.282958984375,-77.2159652709961,-77.1959915161133,-77.2123031616211,-77.2826156616211,-77.3455047607422,-77.3858871459961,-77.4072799682617,-77.478515625,-77.4483413696289,-77.39306640625,-77.3742218017578,-77.3441390991211,-77.2615661621094,-77.130126953125,-76.9922866821289,-76.9358367919922,-76.8909606933594,-76.8518524169922,-76.8690948486328,-76.9122085571289,-76.9246597290039,-76.8966293334961,-76.8668975830078,-76.7865676879883,-76.7423324584961,-76.7720718383789,-76.818603515625,-76.8722152709961,-76.9204559326172,-76.8879852294922,-76.80224609375,-76.6893539428711,-76.27685546875,-76.135498046875,-76.0272445678711,-75.905029296875,-75.7555694580078,-75.6393585205078,-75.6036148071289,-75.6353530883789,-75.6800231933594,-75.6371154785156,-75.5926742553711,-75.5959014892578,-75.53857421875,-75.5584030151367,-75.6421890258789,-75.7083511352539,-75.6708984375,-75.5537109375,-75.4927749633789,-75.4459991455078,-75.2806167602539,-75.2479476928711,-75.123046875,-74.9215850830078,-74.8445739746094,-74.4542465209961,-74.3302230834961,-74.3523941040039,-74.4095687866211,-74.4922866821289,-74.5162582397461,-74.4602584838867,-74.40087890625,-74.3501968383789,-74.2999496459961,-74.2191390991211,-74.2001953125,-74.1429214477539,-74.0591354370117,-73.9095764160156,-73.7957000732422,-73.6769027709961,-73.3133773803711,-72.7218246459961,-72.4470748901367,-72.2750015258789,-72.1652374267578,-72.1357421875,-72.0550842285156,-71.9701232910156,-71.9312515258789,-71.9191360473633,-71.7145462036133,-71.5974578857422,-71.4939956665039,-71.2621078491211,-71.155029296875,-71.1373062133789,-71.2841796875,-71.3197250366211],[43.7886695861816,43.8589859008789,43.6636695861816,43.6329116821289,43.63134765625,43.7042961120605,43.7886695861816],[44.4763679504395,44.5267562866211,44.5262680053711,44.5049781799316,44.4601554870605,44.37744140625,44.2201156616211,44.2922859191895,44.33447265625,44.3791046142578,44.4070320129395,44.41259765625,44.4518547058105,44.4763679504395],[43.4658203125,43.44677734375,43.35546875,43.3033218383789,43.2266578674316,43.2560577392578,43.2806663513184,43.2990264892578,43.3415031433105,43.3929710388184,43.37939453125,43.4476547241211,43.4915008544922,43.4658203125],[-24.3082523345947,-24.3861331939697,-24.4405269622803,-24.4921875,-24.51708984375,-24.4968738555908,-24.3919925689697,-24.3294925689697,-24.2958011627197,-24.3082523345947],[-23.1821308135986,-23.2099609375,-23.2518062591553,-23.2424793243408,-23.2471675872803,-23.2102527618408,-23.1377429962158,-23.1193370819092,-23.1158676147461,-23.1821308135986],[-23.4442386627197,-23.5046863555908,-23.63720703125,-23.7053718566895,-23.7850093841553,-23.7825183868408,-23.7544937133789,-23.7593746185303,-23.7480964660645,-23.7072257995605,-23.7006359100342,-23.5799789428711,-23.5352535247803,-23.4442386627197],[-22.917724609375,-22.8343257904053,-22.8026351928711,-22.7494144439697,-22.692626953125,-22.681884765625,-22.7101078033447,-22.8205070495605,-22.8840827941895,-22.9592781066895,-22.9161128997803,-22.917724609375],[-24.0876960754395,-24.04638671875,-24.03271484375,-24.0941410064697,-24.2430667877197,-24.2828140258789,-24.3223628997803,-24.3980941772461,-24.3929195404053,-24.376708984375,-24.2710933685303,-24.0876960754395],[-22.8883285522461,-22.9202632904053,-22.9594230651855,-22.9806156158447,-22.9909191131592,-22.9329109191895,-22.9047355651855,-22.9039077758789,-22.8883285522461],[-24.8870620727539,-24.9691410064697,-25.0199718475342,-25.0930671691895,-25.0701179504395,-24.9910144805908,-24.9364738464355,-24.8918952941895,-24.8870620727539],[-25.1698246002197,-25.2672367095947,-25.3083019256592,-25.3219242095947,-25.341552734375,-25.3371086120605,-25.1134777069092,-25.03466796875,-24.9796867370605,-25.01708984375,-25.1698246002197],[-83.6419906616211,-83.6172866821289,-83.5881805419922,-83.5752944946289,-83.4482421875,-83.3468246459961,-83.1246109008789,-83.0285186767578,-82.8663101196289,-82.810302734375,-82.7784194946289,-82.6101531982422,-82.5635757446289,-82.5692443847656,-82.5865249633789,-82.6112823486328,-82.6440963745117,-82.723388671875,-82.801025390625,-82.843994140625,-82.8601608276367,-82.8889617919922,-82.925048828125,-82.9398422241211,-82.9428253173828,-82.9403305053711,-82.88134765625,-82.7830505371094,-82.7411651611328,-82.7278289794922,-82.739990234375,-82.8119125366211,-82.8819808959961,-82.9170379638672,-82.855712890625,-82.8426284790039,-82.8447799682617,-82.8616180419922,-82.99755859375,-83.02734375,-83.0233840942383,-82.9484405517578,-82.9128875732422,-82.88330078125,-82.8793487548828,-82.947265625,-83.0414505004883,-83.1233444213867,-83.1295928955078,-83.1623992919922,-83.2857894897461,-83.3914031982422,-83.4697265625,-83.4217758178711,-83.2977600097656,-83.28955078125,-83.29150390625,-83.3768081665039,-83.4520492553711,-83.5437545776367,-83.604736328125,-83.7340850830078,-83.6421890258789,-83.6137237548828,-83.6161575317383,-83.6372604370117,-83.7369155883789,-83.8955535888672,-84.1178741455078,-84.2223663330078,-84.482666015625,-84.5815963745117,-84.6588821411133,-84.6704635620117,-84.64306640625,-84.7149353027344,-85.0250473022461,-85.1984329223633,-85.2356414794922,-85.26318359375,-85.2365264892578,-85.1607437133789,-84.9627914428711,-84.9083480834961,-84.8864212036133,-85.0012664794922,-85.0597229003906,-85.0770492553711,-85.114501953125,-85.1540069580078,-85.3145446777344,-85.6248550415039,-85.6810073852539,-85.7964859008789,-85.8496551513672,-85.8306121826172,-85.703125,-85.663330078125,-85.6714401245117,-85.667236328125,-85.71484375,-85.8328628540039,-85.9080123901367,-85.8874053955078,-85.7522430419922,-85.7436981201172,-85.7443389892578,-85.7222671508789,-85.70263671875,-85.6905288696289,-85.6536560058594,-85.6213836669922,-85.5841827392578,-85.5387191772461,-85.3683624267578,-85.178955078125,-84.9091796875,-84.79736328125,-84.701171875,-84.6341781616211,-84.4891586303711,-84.40185546875,-84.3482971191406,-84.2555694580078,-84.2049865722656,-84.1965789794922,-84.1683578491211,-84.09619140625,-83.9192886352539,-83.8111801147461,-83.7129364013672,-83.658935546875,-83.6419906616211],[-82.561767578125,-82.6548385620117,-82.8531723022461,-82.9596176147461,-83.0672836303711,-83.1415023803711,-83.1837844848633,-83.1802215576172,-83.1129379272461,-83.0548782348633,-83.0072250366211,-82.9735794067383,-83.08251953125,-83.0777359008789,-82.9912109375,-82.7557601928711,-82.7145538330078,-82.6818389892578,-82.62939453125,-82.5678176879883,-82.561767578125],[-77.6689910888672,-77.7100601196289,-77.7550277709961,-77.7836456298828,-77.8231964111328,-77.9000015258789,-77.9185485839844,-77.854736328125,-77.7744140625,-77.6336898803711,-77.64599609375,-77.6689910888672],[-77.87939453125,-77.912353515625,-78.0119171142578,-78.0416488647461,-78.0066909790039,-77.9992141723633,-77.9856414794922,-77.9695816040039,-77.8936462402344,-77.8891143798828,-77.8424758911133,-77.87939453125],[-78.027099609375,-78.0475158691406,-78.1016616821289,-78.1800308227539,-78.2261199951172,-78.27001953125,-78.2735366821289,-78.2009735107422,-78.1505889892578,-78.0941390991211,-78.0616683959961,-78.027099609375],[-78.630126953125,-78.4928665161133,-78.4453125,-78.3995590209961,-78.3512191772461,-78.2838821411133,-78.343017578125,-78.3899459838867,-78.424560546875,-78.5476531982422,-78.6289978027344,-78.6736297607422,-78.6955032348633,-78.630126953125],[-79.3495559692383,-79.347900390625,-79.5227508544922,-79.5978469848633,-79.628173828125,-79.5791473388672,-79.3821792602539,-79.3495559692383],[-81.8374481201172,-81.575439453125,-81.3636245727539,-81.2623519897461,-81.2716293334961,-81.1786117553711,-81.1446304321289,-81.0076675415039,-80.650146484375,-80.6134338378906,-80.5504913330078,-80.459228515625,-80.3648910522461,-80.2661666870117,-80.1676254272461,-80.0752487182617,-79.9599075317383,-79.9235305786133,-79.8202590942383,-79.8507308959961,-79.6766662597656,-79.5492172241211,-79.4564895629883,-79.3582992553711,-79.2756805419922,-79.1830062866211,-78.9019012451172,-78.83544921875,-78.7759704589844,-78.71923828125,-78.6864776611328,-78.1431121826172,-77.9705123901367,-77.8650360107422,-77.6368103027344,-77.5451202392578,-77.4971160888672,-77.5065383911133,-77.5733413696289,-77.5831527709961,-77.4972686767578,-77.3421401977539,-77.2995147705078,-77.2220687866211,-77.1441421508789,-77.1812438964844,-77.2445297241211,-77.3661575317383,-77.2695846557617,-77.2528839111328,-77.2079086303711,-77.1409683227539,-77.0986328125,-76.9280776977539,-76.8363265991211,-76.8598098754883,-76.867431640625,-76.7649841308594,-76.72607421875,-76.6885223388672,-76.6474151611328,-76.5517120361328,-76.4551773071289,-76.2592239379883,-76.0736389160156,-75.8990249633789,-75.7229461669922,-75.6337356567383,-75.5958023071289,-75.6385269165039,-75.6629333496094,-75.5972671508789,-75.740234375,-75.7604064941406,-75.7529754638672,-75.7245559692383,-75.6427764892578,-75.5246124267578,-75.338134765625,-75.2132797241211,-74.959716796875,-74.882568359375,-74.7320785522461,-74.6624526977539,-74.5131378173828,-74.3843765258789,-74.2728042602539,-74.23388671875,-74.198486328125,-74.16748046875,-74.1368103027344,-74.1537094116211,-74.2174301147461,-74.2528305053711,-74.4121551513672,-74.634765625,-74.8500518798828,-74.9551239013672,-75.003173828125,-75.1164093017578,-75.1241226196289,-75.1219711303711,-75.151611328125,-75.1772994995117,-75.2194290161133,-75.2904815673828,-75.5519561767578,-75.6572265625,-75.76513671875,-76.158447265625,-76.2528305053711,-76.515625,-76.7797317504883,-76.8902359008789,-76.9994583129883,-77.2119598388672,-77.4631805419922,-77.715087890625,-77.5537567138672,-77.21337890625,-77.1494140625,-77.1038055419922,-77.093017578125,-77.10791015625,-77.18896484375,-77.2054672241211,-77.2295913696289,-77.3475646972656,-77.467041015625,-77.5927276611328,-77.8568801879883,-77.997314453125,-78.1163558959961,-78.3138732910156,-78.4063491821289,-78.453857421875,-78.4907684326172,-78.5372543334961,-78.5765686035156,-78.636474609375,-78.7276840209961,-78.8229522705078,-79.189208984375,-79.2744140625,-79.357421875,-79.9103012084961,-80.1383285522461,-80.2313461303711,-80.3106994628906,-80.3929214477539,-80.4854431152344,-80.4848098754883,-80.4990692138672,-80.9619140625,-81.03564453125,-81.0831069946289,-81.1166534423828,-81.1414031982422,-81.1855010986328,-81.1995620727539,-81.222412109375,-81.2843704223633,-81.3552703857422,-81.4411163330078,-81.8162155151367,-81.8494110107422,-81.9726104736328,-82.0777359008789,-81.9730453491211,-81.757080078125,-81.7103500366211,-81.6832504272461,-81.7027359008789,-81.7456588745117,-81.7898864746094,-81.8388214111328,-81.9034194946289,-82.738037109375,-82.786376953125,-82.8612289428711,-83.0094299316406,-83.1071243286133,-83.1437530517578,-83.1893997192383,-83.2921447753906,-83.379638671875,-83.4859313964844,-83.5440444946289,-83.6015167236328,-83.64306640625,-83.6866226196289,-83.9007339477539,-83.9327087402344,-83.9633331298828,-83.998046875,-84.0309600830078,-84.1383285522461,-84.2406692504883,-84.4488296508789,-84.5025863647461,-84.4909133911133,-84.5013656616211,-84.5600128173828,-84.6269073486328,-84.6826629638672,-84.7858428955078,-84.8382339477539,-84.88720703125,-84.8772430419922,-84.5327606201172,-84.4942398071289,-84.4330596923828,-84.3731460571289,-84.3263702392578,-84.3830032348633,-84.3612747192383,-84.2813415527344,-84.1217803955078,-84.044921875,-83.2578125,-83.17724609375,-82.6658172607422,-82.5877914428711,-82.3505401611328,-82.1013641357422,-81.8374481201172],[-68.7510757446289,-68.8033218383789,-68.9951171875,-69.15380859375,-69.1588897705078,-69.1184539794922,-69.0767593383789,-69.0131301879883,-68.827392578125,-68.7510757446289],[-81.3695297241211,-81.3372497558594,-81.2964859008789,-81.2848129272461,-81.1304702758789,-81.1071319580078,-81.224609375,-81.2772979736328,-81.3037109375,-81.40478515625,-81.4190902709961,-81.3910140991211,-81.3695297241211],[-79.97900390625,-79.98876953125,-80.020751953125,-80.0941925048828,-80.1258773803711,-80.1161117553711,-80.1009292602539,-80.0836410522461,-80.0675735473633,-80.0162124633789,-79.9918441772461,-79.97509765625,-79.97900390625],[-79.8233871459961,-79.8700637817383,-79.9061965942383,-79.82421875,-79.8031234741211,-79.78515625,-79.766357421875,-79.7422866821289,-79.7422866821289,-79.8233871459961],[34.0044937133789,33.9657211303711,33.9033203125,33.8664054870605,33.83203125,33.7922859191895,33.7569313049316,33.7257804870605,33.6753921508789,33.6144523620605,33.5256843566895,33.4757804870605,33.4638671875,33.4559555053711,33.4242210388184,33.3837890625,33.3255844116211,33.2483406066895,33.1910133361816,33.0775413513184,32.9859390258789,32.9195327758789,32.8694343566895,32.7840805053711,32.72021484375,32.7126960754395,32.7723655700684,32.8798828125,32.9263687133789,32.9416046142578,33.1234359741211,33.3078117370605,33.4587898254395,33.6076164245605,34.0634765625,34.1924819946289,34.2723617553711,34.4111328125,34.5560531616211,34.4631843566895,33.9419937133789,33.9079093933105,33.9312477111816,34.0044937133789],[32.7126960754395,32.72021484375,32.7840805053711,32.8694343566895,32.9195327758789,32.9859390258789,33.0775413513184,33.1910133361816,33.2483406066895,33.3255844116211,33.3837890625,33.4242210388184,33.4559555053711,33.4638671875,33.4757804870605,33.5256843566895,33.6144523620605,33.6753921508789,33.7257804870605,33.7569313049316,33.7922859191895,33.83203125,33.8664054870605,33.9033203125,33.9657211303711,34.0044937133789,34.0236358642578,34.0501937866211,33.9365234375,33.8224601745605,33.7589836120605,33.6994132995605,33.5144538879395,33.4149398803711,33.2965812683105,33.1760711669922,33.1155281066895,33.0623054504395,33.02490234375,33.02392578125,33.0079116821289,32.9417991638184,32.9142570495605,32.8671875,32.7500991821289,32.6929702758789,32.5055656433105,32.4490242004395,32.4137687683105,32.3171882629395,32.3009757995605,32.3909187316895,32.4749984741211,32.5559577941895,32.65234375,32.7126960754395],[14.8093748092651,14.8958005905151,14.9829092025757,14.9899415969849,14.9844722747803,14.993748664856,15.1259765625,15.2585935592651,15.2770519256592,15.3125972747803,15.3543949127197,15.3946285247803,15.4639644622803,15.6439456939697,15.7305660247803,15.8192386627197,15.8939456939697,15.9485340118408,15.9738283157349,16.0072269439697,16.06640625,16.2822265625,16.3599605560303,16.4125003814697,16.4197273254395,16.3922843933105,16.3791007995605,16.3566398620605,16.2825202941895,16.24072265625,16.2103500366211,16.2307605743408,16.2913093566895,16.3341808319092,16.3504867553711,16.4875965118408,16.5966796875,16.63916015625,16.6791019439697,16.7252922058105,16.7786140441895,16.841796875,16.8953132629395,16.9896488189697,16.9933586120605,16.9147472381592,16.869140625,16.8800773620605,16.9807624816895,17.1519546508789,17.4152336120605,17.4623031616211,17.5545902252197,17.6546878814697,17.7022476196289,17.7201156616211,17.7354488372803,17.7092761993408,17.58935546875,17.5962886810303,17.6270503997803,17.6810550689697,17.74658203125,17.7916984558105,17.8312511444092,17.8748054504395,17.9837894439697,18.0146484375,18.0283203125,18.0495128631592,18.0876941680908,18.0992183685303,18.2052726745605,18.2663097381592,18.3052749633789,18.3484382629395,18.5162105560303,18.5624008178711,18.5771484375,18.56884765625,18.5946273803711,18.8069324493408,18.8292961120605,18.8322257995605,18.8070316314697,18.7497081756592,18.6761722564697,18.5964851379395,18.5345706939697,18.47607421875,18.4158210754395,18.3831062316895,18.3648433685303,18.1609363555908,18.1326179504395,18.1099605560303,18.1003913879395,18.0859375,18.0508785247803,17.9407234191895,17.9132804870605,17.8926753997803,17.8308601379395,17.7584972381592,17.6253910064697,17.4826164245605,17.296875,17.1884765625,17.1356449127197,17.0632801055908,16.9852523803711,16.953125,16.9283199310303,16.8836917877197,16.8332023620605,16.7644538879395,16.7126960754395,16.6009769439697,16.5435543060303,16.4779300689697,16.4148426055908,16.3672847747803,16.2193355560303,16.0572261810303,15.8251953125,15.7650394439697,15.7007808685303,15.5994148254395,15.4029302597046,15.3109378814697,15.2527341842651,15.1996097564697,15.1617183685303,15.1397457122803,15.0667963027954,14.9934568405151,14.9721670150757,14.9473638534546,14.9225578308105,14.8218746185303,14.7859373092651,14.7066402435303,14.6913080215454,14.5539054870605,14.4886722564697,14.4310541152954,14.3675775527954,14.1898431777954,14.0491209030151,13.98876953125,13.92431640625,13.8431634902954,13.8147459030151,13.7699213027954,13.6849613189697,13.5476560592651,13.4407224655151,13.4011716842651,13.3836917877197,13.3390626907349,13.2887687683105,13.2278318405151,13.1405277252197,13.0237302780151,12.9166984558105,12.8133783340454,12.7478513717651,12.68115234375,12.6320314407349,12.5557613372803,12.5002927780151,12.45703125,12.408203125,12.3905277252197,12.4501953125,12.4718751907349,12.4975595474243,12.5124998092651,12.5120115280151,12.4576168060303,12.3841800689697,12.2764654159546,12.2078123092651,12.1825199127197,12.1750001907349,12.1278324127197,12.0897464752197,12.08984375,12.0992183685303,12.1348628997803,12.1748046875,12.2311525344849,12.27734375,12.3056640625,12.3585939407349,12.45263671875,12.5490236282349,12.6355466842651,12.7064447402954,12.7654294967651,12.8682613372803,12.9426755905151,12.966796875,12.9970703125,13.0164070129395,13.18115234375,13.2376956939697,13.26953125,13.3060550689697,13.3410158157349,13.3746089935303,13.4011716842651,13.4361333847046,13.4725589752197,13.5265617370605,13.5567388534546,13.7013673782349,13.8985347747803,13.9984369277954,14.0964851379395,14.2017583847046,14.3690433502197,14.3770513534546,14.2994146347046,14.2733402252197,14.255859375,14.283203125,14.3197259902954,14.3672857284546,14.5073232650757,14.5457019805908,14.5596685409546,14.59521484375,14.6238279342651,14.6135740280151,14.6582040786743,14.7233400344849,14.7665033340454,14.7974615097046,14.8093748092651],[14.2136716842651,14.1721677780151,14.04833984375,13.9257802963257,13.9021482467651,13.9216804504395,13.8724603652954,13.8271484375,13.8204097747803,13.8277339935303,14.0388669967651,14.21142578125,14.1982421875,14.2136716842651],[11.2828121185303,11.1292963027954,11.0707025527954,11.01171875,11.04345703125,11.0849618911743,11.2335939407349,11.2802734375,11.2828121185303],[13.7091789245605,13.7341804504395,13.7073240280151,13.5949220657349,13.4820318222046,13.41455078125,13.3643550872803,13.1900386810303,13.1621103286743,13.1563472747803,13.1812496185303,13.1766605377197,13.2314462661743,13.2399415969849,13.3368167877197,13.4227533340454,13.4500970840454,13.4912109375,13.6360359191895,13.6576175689697,13.6707029342651,13.6033210754395,13.5804681777954,13.6018552780151,13.7091789245605],[8.587890625,8.54892635345459,8.45380783081055,8.400390625,8.41767597198486,8.46816444396973,8.50996112823486,8.57343673706055,8.587890625],[9.73974609375,9.74589920043945,9.89228534698486,9.95380783081055,10.0221681594849,10.02880859375,9.94130897521973,9.86865234375,10.1434564590454,10.1708002090454,10.21240234375,10.3604488372803,10.7315425872803,10.9559574127197,11.0133790969849,11.0643548965454,11.0085945129395,10.8107423782349,10.8545894622803,10.9177732467651,11.1042966842651,11.3996095657349,11.4611330032349,11.7005863189697,11.7962894439697,12.111328125,12.1686525344849,12.2962884902954,12.3785161972046,12.5753908157349,12.7791023254395,12.8980474472046,13.0286130905151,13.1474609375,13.4480476379395,13.7242183685303,13.822265625,13.8655271530151,13.9503908157349,14.0250005722046,14.25,14.2588863372803,14.26611328125,14.2798824310303,14.2987308502197,14.41455078125,14.4123048782349,14.4109373092651,14.3685541152954,14.2931642532349,14.1936521530151,14.1388673782349,14.1286134719849,14.2537107467651,14.5140628814697,14.6194343566895,14.5697259902954,14.5545902252197,14.5739259719849,14.6156244277954,14.6798830032349,14.7053718566895,14.6923828125,14.70458984375,14.7525396347046,14.7481441497803,14.7248048782349,14.6929683685303,14.6749019622803,14.6016607284546,14.6239252090454,14.6813478469849,14.7249021530151,14.7386722564697,14.7109384536743,14.7247076034546,14.9059572219849,14.935546875,14.953125,15.0166015625,14.9638671875,14.9174795150757,14.8142585754395,14.8093748092651,14.7974615097046,14.7665033340454,14.7233400344849,14.6582040786743,14.6135740280151,14.6238279342651,14.59521484375,14.5596685409546,14.5457019805908,14.5073232650757,14.3672857284546,14.3197259902954,14.283203125,14.255859375,14.2733402252197,14.2994146347046,14.3770513534546,14.3690433502197,14.2017583847046,14.0964851379395,13.9984369277954,13.8985347747803,13.7013673782349,13.5567388534546,13.5265617370605,13.4725589752197,13.4361333847046,13.4011716842651,13.3746089935303,13.3410158157349,13.3060550689697,13.26953125,13.2376956939697,13.18115234375,13.0164070129395,12.9970703125,12.966796875,12.9426755905151,12.8682613372803,12.7654294967651,12.7064447402954,12.6355466842651,12.5490236282349,12.45263671875,12.3585939407349,12.3056640625,12.27734375,12.2311525344849,12.1748046875,12.1348628997803,12.0992183685303,12.08984375,12.0897464752197,12.1278324127197,12.1750001907349,12.1825199127197,12.2078123092651,12.2764654159546,12.3841800689697,12.4576168060303,12.5120115280151,12.5124998092651,12.4975595474243,12.4718751907349,12.4501953125,12.3905277252197,12.408203125,12.45703125,12.5002927780151,12.5557613372803,12.6320314407349,12.68115234375,12.7478513717651,12.8133783340454,12.9166984558105,13.0237302780151,13.1405277252197,13.2278318405151,13.2887687683105,13.3390626907349,13.3836917877197,13.4011716842651,13.4407224655151,13.5476560592651,13.6849613189697,13.7699213027954,13.8147459030151,13.8029298782349,13.7974615097046,13.798828125,13.7853507995605,13.7239255905151,13.6921873092651,13.6751947402954,13.4866209030151,13.4716796875,13.4598636627197,13.4093751907349,13.3746089935303,13.3228521347046,13.2152347564697,13.1404294967651,13.0821285247803,12.8974609375,12.8142576217651,12.7603511810303,12.7600584030151,12.8499021530151,12.9535160064697,12.9541988372803,12.9083013534546,12.8976564407349,12.9281244277954,12.9855470657349,13.0335931777954,13.0541019439697,13.0479488372803,13.0315437316895,13.0143547058105,12.9680671691895,12.87890625,12.8093748092651,12.7828121185303,12.7811517715454,12.7961912155151,12.7713861465454,12.6858396530151,12.59423828125,12.5265626907349,12.48291015625,12.4357423782349,12.3631839752197,12.2683591842651,12.2092771530151,12.1968755722046,12.2038087844849,12.1856451034546,11.716796875,11.5739250183105,11.4699220657349,11.3929691314697,11.3741207122803,11.2979488372803,11.2119140625,11.1912107467651,11.1360349655151,11.0419921875,10.9808588027954,10.9521484375,10.8939447402954,10.87060546875,10.873046875,10.7416009902954,10.65869140625,10.4828119277954,10.439453125,10.4303712844849,10.4039068222046,10.369140625,10.3127927780151,10.2406244277954,10.1830081939697,10.1857423782349,10.2002935409546,10.1587886810303,10.0964851379395,10.0663089752197,10.07421875,10.0598640441895,10.0340824127197,9.97158145904541,9.83915996551514,9.74892520904541,9.71513652801514,9.65058612823486,9.54892539978027,9.52402305603027,9.35000038146973,9.18281173706055,9.12753963470459,8.88115215301514,8.8740234375,8.83115291595459,8.79306602478027,8.77011775970459,8.75478553771973,8.72832012176514,8.61787128448486,8.57265663146973,8.50986289978027,8.43574237823486,8.40341854095459,8.41328144073486,8.45175838470459,8.55234336853027,8.56709003448486,8.57050800323486,8.55947303771973,8.47763633728027,8.45400333404541,8.43007850646973,8.41474628448486,8.32783222198486,8.1982421875,8.09375,7.92705106735229,7.69804668426514,7.61562490463257,7.56543016433716,7.52939462661743,7.53857469558716,7.59326219558716,7.60849618911743,7.58417987823486,7.61660194396973,7.70566415786743,7.76513624191284,7.79482460021973,7.83798789978027,7.92275381088257,8.1240234375,8.14033222198486,8.13486385345459,8.08066368103027,8.00126934051514,7.79921865463257,7.61093807220459,7.52548885345459,7.45058631896973,7.40419912338257,7.31337881088257,7.19990205764771,7.11738252639771,7.06572246551514,7.03671884536743,7.02216768264771,7.00146484375,6.95830106735229,6.89121103286743,6.84951162338257,6.82070350646973,6.77626895904541,6.73544931411743,6.60761785507202,6.57470703125,6.56630802154541,6.53427696228027,6.45810508728027,6.38222646713257,6.34433603286743,6.34843778610229,6.37832069396973,6.40673828125,6.44462871551514,6.48476552963257,6.49375009536743,6.4873046875,6.44091844558716,6.32460975646973,6.25605440139771,6.20488214492798,6.13818359375,6.10976600646973,6.10830116271973,6.11650419235229,6.12128877639771,6.17509794235229,6.36445331573486,6.34365224838257,6.34091854095459,6.294921875,6.20302724838257,6.1787109375,6.16845655441284,6.23593759536743,6.15449237823486,6.11943340301514,6.00595712661743,5.99394512176514,6.04843759536743,6.00683546066284,5.955078125,5.89472675323486,5.8671875,5.85751962661743,5.86835956573486,5.93925762176514,5.96103525161743,6.12998056411743,6.13691425323486,6.11337852478027,6.08242177963257,6.07480430603027,6.07587909698486,6.16621065139771,6.19287109375,6.19882822036743,6.19326162338257,6.1416015625,6.09111309051514,6.08935546875,6.052734375,5.94853496551514,5.94872999191284,6.00761699676514,6.08984375,6.1171875,6.16650390625,6.29707050323486,6.35566377639771,6.37216758728027,6.42499971389771,6.517578125,6.74179649353027,6.77519559860229,6.80039072036743,6.80244159698486,6.71562528610229,6.71298837661743,6.72451162338257,6.7490234375,6.80039072036743,6.85507774353027,6.97724628448486,7.01962900161743,7.03261661529541,7.03515625,7.00185537338257,6.96816396713257,6.92207050323486,6.83251953125,6.74882793426514,6.70292949676514,6.69160175323486,6.71240234375,6.71875,6.70537090301514,6.71074247360229,6.74843740463257,7.01318311691284,7.03300762176514,7.05087900161743,7.11708974838257,7.17949295043945,7.18994140625,7.18896532058716,7.197265625,7.15205097198486,7.05332040786743,7.07431650161743,7.10712909698486,7.20644569396973,7.28525400161743,7.62919902801514,8.00927734375,8.16709041595459,8.10849571228027,8.20078086853027,8.24521446228027,8.27900409698486,8.30156230926514,8.33388710021973,8.45136737823486,8.49267482757568,8.49521541595459,8.53847694396973,8.50625038146973,8.52841758728027,8.57558631896973,8.61894512176514,8.89775371551514,9.20556545257568,9.32197284698486,9.58535099029541,9.67314434051514,9.78398418426514,9.63124942779541,9.31201171875,9.21640586853027,9.06962871551514,8.97812461853027,8.92041015625,8.90351486206055,8.90664005279541,8.8515625,8.78037166595459,8.73603534698486,8.64492130279541,8.62578105926514,8.64804649353027,8.83115291595459,8.95185470581055,8.95722675323486,8.88095664978027,8.78964805603027,8.68232345581055,8.67031192779541,8.67070293426514,8.85722732543945,8.90292930603027,9.18583965301514,9.25498104095459,9.34199237823486,9.49873065948486,9.61581993103027,9.66123008728027,9.72500038146973,9.73974609375],[8.30771541595459,8.28466796875,8.29570293426514,8.40517520904541,8.45146560668945,8.40410137176514,8.39042949676514,8.37119197845459,8.3798828125,8.62959003448486,8.6005859375,8.34736251831055,8.30771541595459],[43.2459945678711,43.1593742370605,43.0486335754395,42.9227523803711,42.8441390991211,42.7830085754395,42.7412109375,42.6549797058105,42.5577125549316,42.4651374816895,42.30810546875,42.1662101745605,42.0521507263184,41.9574203491211,41.8721656799316,41.7982406616211,41.7820320129395,41.7646484375,41.7665061950684,41.7926788330078,41.8156242370605,41.9496078491211,41.9958992004395,42.1491203308105,42.2803688049316,42.3785171508789,42.4085922241211,42.4500007629395,42.4793968200684,42.6701164245605,42.7037124633789,42.7674827575684,42.8252944946289,42.8659172058105,42.88330078125,43.0056648254395,43.11669921875,43.130859375,43.2986335754395,43.353515625,43.4097671508789,43.3802719116211,43.3367195129395,43.2720680236816,43.0480461120605,42.7990226745605,42.6400375366211,42.5217781066895,42.5397453308105,42.5837860107422,42.6527366638184,42.7897453308105,42.9115257263184,43.0427742004395,43.1617164611816,43.2459945678711],[-61.2816848754883,-61.3753890991211,-61.4157218933105,-61.4811553955078,-61.4699211120605,-61.4581031799316,-61.3200187683105,-61.2772483825684,-61.2510719299316,-61.2816848754883],[11.3614263534546,11.5383787155151,11.6581048965454,11.7395505905151,11.7589845657349,11.7659177780151,11.6803703308105,11.5859384536743,11.4574222564697,11.0355472564697,11.0416994094849,11.05859375,11.2584962844849,11.3614263534546],[10.484375,10.4172859191895,10.3405275344849,10.2156248092651,10.1999025344849,10.2655277252197,10.3469724655151,10.4136714935303,10.5048828125,10.484375],[12.5492191314697,12.5110349655151,12.3575201034546,12.1844730377197,12.1188468933105,12.1436529159546,12.1617183685303,12.2199220657349,12.2587890625,12.2740230560303,12.31005859375,12.4171876907349,12.4695310592651,12.5132818222046,12.5492191314697],[10.0612306594849,9.95712852478027,9.90390586853027,9.80625057220459,9.77119064331055,9.78125,9.83037090301514,9.99882793426514,10.0577144622803,10.0612306594849],[10.7340822219849,10.6897459030151,10.6294918060303,10.6216793060303,10.6924800872803,10.73828125,10.8567380905151,10.9250001907349,10.9510746002197,10.9208002090454,10.7652339935303,10.7340822219849],[15.0876951217651,15.05078125,14.8855466842651,14.6841793060303,14.7136716842651,14.7653322219849,15.1326169967651,15.1371097564697,15.0876951217651],[10.6451168060303,10.6868162155151,10.7380857467651,10.8192386627197,10.7853517532349,10.8083982467651,10.7852535247803,10.6238279342651,10.4427738189697,10.2545900344849,9.98877048492432,9.96738338470459,9.93007850646973,9.85898494720459,9.86064434051514,9.99423885345459,10.2861328125,10.3536128997803,10.4240236282349,10.5050773620605,10.6227540969849,10.6451168060303],[12.6657228469849,12.5715818405151,12.5508785247803,12.5203132629395,12.5699214935303,12.5992183685303,12.6200189590454,12.6484375,12.6657228469849],[10.6073236465454,10.59033203125,10.5269527435303,10.5203123092651,10.5443363189697,10.51611328125,10.5471677780151,10.6363286972046,10.6617183685303,10.6273441314697,10.6073236465454],[12.5687503814697,12.5711908340454,12.5452146530151,12.5070304870605,12.4071292877197,12.3206052780151,12.2434568405151,12.2150392532349,12.2753896713257,12.3851556777954,12.4130859375,12.3224611282349,12.0899410247803,12.0655279159546,12.0730466842651,12.06884765625,12.0503902435303,11.8623046875,11.7409181594849,11.7398433685303,11.70361328125,11.69677734375,11.65380859375,11.4758787155151,11.4068355560303,11.3102540969849,11.2863283157349,11.1707038879395,11.1897459030151,11.1280279159546,11.1195306777954,11.1209955215454,11.0703125,11.0087900161743,10.9789056777954,11.0496091842651,11.2244138717651,11.2754888534546,11.322265625,11.4636716842651,11.4595699310303,11.4747066497803,11.6277341842651,11.6958990097046,11.6822261810303,11.69091796875,11.7835931777954,11.8197269439697,11.8583002090454,11.8853511810303,11.9220705032349,11.9345703125,11.9127931594849,11.8664064407349,12.0396490097046,12.2189455032349,12.3232421875,12.4282236099243,12.5257806777954,12.5787115097046,12.6083984375,12.54296875,12.5248050689697,12.5687503814697],[11.0521488189697,11.0114259719849,10.8738279342651,10.9345703125,11.0857419967651,11.1745109558105,11.0768556594849,11.0521488189697],[9.73974609375,9.72500038146973,9.66123008728027,9.61581993103027,9.49873065948486,9.34199237823486,9.25498104095459,9.18583965301514,8.90292930603027,8.85722732543945,8.67070293426514,8.67031192779541,8.66142559051514,8.63828086853027,8.57294940948486,8.66982460021973,8.65107440948486,8.61591815948486,8.34531211853027,8.13212871551514,8.18134784698486,8.20234298706055,8.12148380279541,8.1298828125,8.16396522521973,8.23173904418945,8.28144550323486,8.47314453125,8.55292892456055,8.60761737823486,8.67167949676514,8.71806621551514,8.73613262176514,8.88808631896973,8.99453067779541,9.06709003448486,9.14033222198486,9.19638729095459,9.20966815948486,9.2548828125,9.11044979095459,8.99277400970459,8.87607383728027,8.77197265625,8.60312557220459,8.46835899353027,8.3466796875,8.26826190948486,8.26630783081055,8.28408241271973,8.42705059051514,8.61855506896973,8.8115234375,8.95224571228027,9.03632831573486,9.298828125,9.43359375,9.55429649353027,9.81513595581055,9.96230506896973,10.2590827941895,10.5333013534546,10.6099615097046,10.48095703125,10.4602537155151,10.4446287155151,10.537109375,10.5175790786743,10.5241212844849,10.4369144439697,10.3384761810303,10.2960939407349,10.2870111465454,10.2966794967651,10.28271484375,10.3835935592651,10.490234375,10.8458976745605,10.8828125,10.9261713027954,10.8944330215454,10.8564453125,10.75341796875,10.6211919784546,10.5389652252197,10.4269533157349,10.3737306594849,10.3187503814697,10.2266607284546,10.1830081939697,10.1593742370605,10.1073246002197,10.0173826217651,9.90371131896973,9.96201133728027,10.0236330032349,9.9990234375,9.89902305603027,9.81035137176514,9.77324199676514,9.66142559051514,9.59111404418945,9.62558555603027,9.64023399353027,9.67099571228027,9.64326190948486,9.50478553771973,9.45371055603027,9.57236289978027,9.64540958404541,9.68818378448486,9.73232364654541,9.70527362823486,9.73974609375],[-71.768310546875,-71.7637634277344,-71.7372589111328,-71.7619171142578,-71.87255859375,-71.9403839111328,-72.0003890991211,-71.9868698120117,-71.8665084838867,-71.82421875,-71.7432174682617,-71.72705078125,-71.733642578125,-71.786376953125,-71.80712890625,-71.7420425415039,-71.6570281982422,-71.6453094482422,-71.647216796875,-71.7464828491211,-71.753173828125,-71.7069320678711,-71.7114715576172,-71.7574234008789,-71.7792510986328,-71.735107421875,-71.7060546875,-71.6673889160156,-71.615966796875,-71.5577621459961,-71.4417037963867,-71.2813491821289,-71.2359390258789,-71.0819320678711,-70.9541473388672,-70.8338851928711,-70.7852554321289,-70.6859359741211,-70.6361846923828,-70.4793472290039,-70.4364318847656,-70.3047332763672,-70.19384765625,-70.1294479370117,-70.0140609741211,-69.9568328857422,-69.8912048339844,-69.8783187866211,-69.8234405517578,-69.7394104003906,-69.324951171875,-69.2324676513672,-69.2642593383789,-69.32275390625,-69.5197296142578,-69.60595703125,-69.6232376098633,-69.6236343383789,-69.5083465576172,-69.395263671875,-69.2802276611328,-69.1630325317383,-69.0313034057617,-68.9013671875,-68.6847610473633,-68.4454040527344,-68.3813934326172,-68.3391571044922,-68.3592758178711,-68.44482421875,-68.4932098388672,-68.5637664794922,-68.6122055053711,-68.6588363647461,-68.6874008178711,-68.7210006713867,-68.7784652709961,-68.8195343017578,-68.9349594116211,-69.072265625,-69.2745132446289,-69.39697265625,-69.5194320678711,-69.6447296142578,-69.7706527709961,-69.8963851928711,-70.018310546875,-70.0633316040039,-70.1416015625,-70.18310546875,-70.4799270629883,-70.5654296875,-70.6446838378906,-70.7588424682617,-70.92431640625,-71.02783203125,-71.0699691772461,-71.0822296142578,-71.0826187133789,-71.1060028076172,-71.2672882080078,-71.3582992553711,-71.3956985473633,-71.43896484375,-71.5183563232422,-71.5690460205078,-71.6317367553711,-71.6583023071289,-71.6572265625,-71.6737289428711,-71.7124481201172,-71.768310546875],[8.57656288146973,8.59765625,8.60126876831055,8.50673866271973,8.44423770904541,8.36962890625,8.23076152801514,8.20761680603027,8.20878982543945,8.302734375,8.333984375,8.34873104095459,8.30673789978027,8.2802734375,8.24570274353027,8.2470703125,8.28290939331055,8.31806659698486,8.32900428771973,8.31640625,8.35986328125,8.39423847198486,8.31210994720459,8.27685546875,8.25468730926514,8.24560546875,8.19277381896973,8.12343788146973,8.04560565948486,7.94941377639771,7.83828163146973,7.74853563308716,7.55449247360229,7.51386737823486,7.49560499191284,7.50019502639771,7.53437471389771,7.62753868103027,7.70917987823486,7.73134708404541,7.7626953125,7.87724590301514,8.07558536529541,8.11250019073486,8.2109375,8.30419921875,8.33339881896973,8.51513576507568,8.68290996551514,8.84404277801514,9.01894569396973,9.04404258728027,9.10234355926514,9.16025352478027,9.22402381896973,9.28789043426514,9.36328220367432,9.40605449676514,9.4580078125,9.51874923706055,9.42099571228027,9.31025409698486,9.39101600646973,9.54619216918945,9.64013671875,9.67265605926514,9.74589920043945,9.80527400970459,9.82070255279541,9.84257793426514,9.81562519073486,9.85820293426514,9.916015625,9.82529258728027,9.74755859375,9.75253963470459,9.79541015625,9.83710956573486,9.89443302154541,9.88320350646973,9.85937595367432,9.68496131896973,9.49140548706055,9.43789005279541,9.42236328125,9.44824123382568,9.58125019073486,9.78105449676514,10.0006837844849,10.01904296875,10.0281248092651,10.1195316314697,10.2186527252197,10.255859375,10.3257808685303,10.3958988189697,10.43896484375,10.6861324310303,11.1082029342651,11.5076169967651,11.5369148254395,11.6242189407349,11.7669925689697,11.873046875,11.9678707122803,11.4499998092651,10.9322261810303,10.4143552780151,9.896484375,9.37871074676514,8.86093807220459,8.34306621551514,7.82519483566284,7.48173904418945,7.26337957382202,6.98935556411743,6.73066377639771,6.52705097198486,6.26337862014771,6.13066387176514,5.83662128448486,5.74834012985229,5.35869169235229,5.00136661529541,4.67128896713257,4.44570302963257,4.22763633728027,3.91015601158142,3.68349575996399,3.43876957893372,3.40087866783142,3.3564453125,3.32343769073486,3.25595688819885,3.17421889305115,3.11972641944885,3.10605454444885,3.13789057731628,3.17724633216858,3.1923828125,3.21962904930115,3.25439453125,3.25585961341858,3.22705078125,3.20166015625,3.20273447036743,3.20341777801514,3.20371079444885,3.13027358055115,2.99248075485229,2.86572289466858,2.80791044235229,2.66777348518372,2.47421860694885,2.40615224838257,2.28085970878601,2.21933603286743,1.92880845069885,1.83242213726044,1.75322282314301,1.68544924259186,1.64736306667328,1.63603508472443,1.61064445972443,1.29023444652557,1.20888650417328,1.16572284698486,1.16406226158142,1.17275404930115,1.15917956829071,1.14550769329071,0.999414324760437,0.671874940395355,0.344433575868607,0.0169922094792128,0,-0.310547143220901,-0.63798850774765,-0.965478599071503,-1.29296863079071,-1.62040996551514,-1.94790053367615,-2.275390625,-2.60292959213257,-2.93037080764771,-3.25786161422729,-3.58535146713257,-3.91279292106628,-4.24033212661743,-4.51699256896973,-4.82260751724243,-5.04951190948486,-5.27500009536743,-5.51694345474243,-5.67451143264771,-5.862548828125,-6.05058622360229,-6.23867225646973,-6.42670917510986,-6.61474609375,-6.80283212661743,-6.99086904525757,-7.17890596389771,-7.36699247360229,-7.55507802963257,-7.74311590194702,-7.93115234375,-8.11923789978027,-8.30727577209473,-8.49531269073486,-8.683349609375,-8.683349609375,-8.683349609375,-8.683349609375,-8.683349609375,-8.683349609375,-8.683349609375,-8.683349609375,-8.67841815948486,-8.659912109375,-8.558349609375,-8.39931678771973,-8.34047794342041,-8.26518535614014,-7.99892568588257,-7.94384813308716,-7.68515634536743,-7.62460947036743,-7.48574209213257,-7.42768573760986,-7.34975576400757,-7.23491191864014,-7.16020488739014,-7.14243173599243,-7.09492206573486,-6.85556650161743,-6.755126953125,-6.63535165786743,-6.59775400161743,-6.565673828125,-6.52055644989014,-6.51069355010986,-6.50791025161743,-6.50087881088257,-6.47973585128784,-6.42763662338257,-6.35761690139771,-6.21479463577271,-6.16650390625,-6.00429677963257,-5.77499961853027,-5.59331083297729,-5.44877958297729,-5.29365253448486,-5.18012666702271,-5.06191444396973,-4.96826219558716,-4.77851581573486,-4.61962890625,-4.52915048599243,-4.32285165786743,-4.14877939224243,-3.98535180091858,-3.86005830764771,-3.70200181007385,-3.66679692268372,-3.62690424919128,-3.62451171875,-3.67250990867615,-3.73017597198486,-3.77099609375,-3.81181669235229,-3.83339810371399,-3.82138657569885,-3.81513714790344,-3.78916025161743,-3.79643559455872,-3.83710932731628,-3.84956073760986,-3.8466796875,-3.82675766944885,-3.76816415786743,-3.70024394989014,-3.60458970069885,-3.43979501724243,-3.01738238334656,-2.98823213577271,-2.96113300323486,-2.93085956573486,-2.88720726966858,-2.86342763900757,-2.72260713577271,-2.52324199676514,-2.44838833808899,-2.23125004768372,-2.07280254364014,-1.81699228286743,-1.63515627384186,-1.47705078125,-1.27534151077271,-1.22592806816101,-1.22592806816101,-1.26210916042328,-1.24033212661743,-1.16259789466858,-1.06552731990814,-1.11103510856628,-1.18823254108429,-1.29638683795929,-1.35214829444885,-1.44999969005585,-1.51000964641571,-1.55073261260986,-1.62509739398956,-1.67919945716858,-1.63124978542328,-1.70297837257385,-1.71411156654358,-1.71469712257385,-1.69267606735229,-1.70693349838257,-1.79179656505585,-1.75185573101044,-1.73330104351044,-1.73945295810699,-1.81660175323486,-1.84965860843658,-1.83242213726044,-1.79218745231628,-1.79560554027557,-1.92089831829071,-2.13178682327271,-2.19077157974243,-2.21962881088257,-2.01777338981628,-1.91328132152557,-1.67363262176514,-1.48374009132385,-1.33583962917328,-1.20537090301514,-1.08769500255585,-0.917480766773224,-0.426123052835464,-0.350781381130219,-0.18916030228138,-0.0482424721121788,0,0.0479491204023361,0.151660218834877,0.312206983566284,0.514941275119781,0.790820419788361,0.9716796875,1.25722646713257,1.97451162338257,2.34287071228027,2.59335947036743,2.84648418426514,2.97285151481628,3.52050757408142,3.77900409698486,4.75810575485229,4.87783193588257,4.99540996551514,5.19560527801514,5.29540967941284,5.42460918426514,5.72548818588257,6.06474590301514,6.24912118911743,6.32783174514771,6.48652362823486,6.57587909698486,6.92753934860229,7.14345741271973,7.23847675323486,7.20429658889771,7.43242168426514,7.60771465301514,7.79160165786743,7.91044950485229,8.12714862823486,8.57656288146973],[-80.131591796875,-80.1506881713867,-80.2456970214844,-80.27294921875,-80.2721633911133,-80.2498016357422,-80.2236785888672,-80.1457061767578,-80.0807647705078,-79.9972686767578,-79.9090347290039,-80.0132369995117,-80.0711898803711,-80.0934066772461,-80.131591796875],[-90.4239273071289,-90.4644088745117,-90.51953125,-90.4772033691406,-90.4319839477539,-90.3987274169922,-90.379150390625,-90.4239273071289],[-89.4188995361328,-89.53662109375,-89.5772933959961,-89.6026382446289,-89.6085891723633,-89.54345703125,-89.4799346923828,-89.4231414794922,-89.3184127807617,-89.287841796875,-89.2674331665039,-89.2593765258789,-89.2948760986328,-89.3583450317383,-89.4188995361328],[-90.3348693847656,-90.3871078491211,-90.5421447753906,-90.5316925048828,-90.4697265625,-90.2693862915039,-90.185302734375,-90.1927261352539,-90.2610855102539,-90.3154296875,-90.3348693847656],[-91.4259719848633,-91.5263671875,-91.6107406616211,-91.6465835571289,-91.6541519165039,-91.6466827392578,-91.4601516723633,-91.3993682861328,-91.3999557495117,-91.4259719848633],[-90.5739212036133,-90.6204605102539,-90.8090286254883,-90.8677749633789,-90.8203659057617,-90.7803726196289,-90.6675338745117,-90.5533218383789,-90.5739212036133],[-91.2721633911133,-91.2100601196289,-91.1762237548828,-90.9755401611328,-90.9506378173828,-90.9684524536133,-90.9589309692383,-90.862548828125,-90.7996597290039,-90.905517578125,-91.1310577392578,-91.3715286254883,-91.4190444946289,-91.4835433959961,-91.4954147338867,-91.4583053588867,-91.3340835571289,-91.1446838378906,-91.1209487915039,-91.197021484375,-91.24951171875,-91.3691864013672,-91.4288558959961,-91.4686965942383,-91.5499954223633,-91.590087890625,-91.5968246459961,-91.5091781616211,-91.4910125732422,-91.3613739013672,-91.3057632446289,-91.2721633911133],[-78.9092330932617,-78.9656295776367,-78.99169921875,-78.9232406616211,-78.8998031616211,-78.9092330932617],[-75.2844696044922,-75.3404769897461,-75.3983917236328,-75.4759750366211,-75.583740234375,-75.6262741088867,-75.6320266723633,-75.5605926513672,-75.4910583496094,-75.4659652709961,-75.4247055053711,-75.3252487182617,-75.2632369995117,-75.2593765258789,-75.2787094116211,-75.2835922241211,-75.2496109008789,-75.2724151611328,-75.3091812133789,-75.3481979370117,-75.380126953125,-75.4080581665039,-75.42041015625,-75.4491729736328,-75.5138702392578,-75.570556640625,-75.6416473388672,-75.7445297241211,-75.8854522705078,-76.0897979736328,-76.2409210205078,-76.3601531982422,-76.4993667602539,-76.6791000366211,-76.8807601928711,-77.1614761352539,-77.3600616455078,-77.5064926147461,-77.6589813232422,-77.860595703125,-77.9384765625,-78.0679244995117,-78.1282272338867,-78.1833038330078,-78.1946258544922,-78.1874465942383,-78.1609878540039,-78.1584930419922,-78.1948776245117,-78.226318359375,-78.2403869628906,-78.250732421875,-78.2841796875,-78.3230514526367,-78.3453674316406,-78.3472595214844,-78.3980484008789,-78.3999481201172,-78.4214401245117,-78.4197769165039,-78.4710464477539,-78.4934539794922,-78.5090866088867,-78.5504379272461,-78.5651397705078,-78.6033706665039,-78.6479949951172,-78.6793975830078,-78.6851577758789,-78.6612319946289,-78.6529235839844,-78.6744613647461,-78.68603515625,-78.7430648803711,-78.8615188598633,-78.9076156616211,-78.92578125,-78.9142074584961,-78.919189453125,-78.9753875732422,-78.9952621459961,-79.0333023071289,-79.0762710571289,-79.1866760253906,-79.2681121826172,-79.3309631347656,-79.3994140625,-79.4557647705078,-79.5019073486328,-79.5161590576172,-79.5776901245117,-79.6385269165039,-79.7109832763672,-79.7972640991211,-79.8451156616211,-79.962890625,-80.0635223388672,-80.1395492553711,-80.1974563598633,-80.232177734375,-80.2933578491211,-80.3834991455078,-80.4241714477539,-80.4785614013672,-80.4884796142578,-80.44384765625,-80.3528823852539,-80.4537582397461,-80.4884796142578,-80.4934616088867,-80.510009765625,-80.4901351928711,-80.4372024536133,-80.3578567504883,-80.3032684326172,-80.2668914794922,-80.2305145263672,-80.1941375732422,-80.1792526245117,-80.2175827026367,-80.2288589477539,-80.21728515625,-80.2189483642578,-80.2206039428711,-80.2437438964844,-80.2454147338867,-80.2652359008789,-80.2718734741211,-80.2735366821289,-80.29833984375,-80.3246536254883,-80.3031311035156,-80.1598663330078,-80.100341796875,-80.0266647338867,-79.9633331298828,-79.9215774536133,-79.8227081298828,-79.7298812866211,-79.7455062866211,-79.8224105834961,-79.8397445678711,-79.8326187133789,-79.8421401977539,-79.8934020996094,-79.8803176879883,-79.9255828857422,-79.989013671875,-80.0301742553711,-80.0066452026367,-80.0530776977539,-80.1271514892578,-80.2486267089844,-80.2552795410156,-80.2847137451172,-80.3785705566406,-80.4501037597656,-80.6848602294922,-80.8390579223633,-80.9321746826172,-80.9516143798828,-80.9627914428711,-80.8676300048828,-80.7703628540039,-80.7605972290039,-80.7631378173828,-80.8350067138672,-80.8014144897461,-80.8200149536133,-80.9023971557617,-80.8414077758789,-80.6236801147461,-80.5539093017578,-80.5050735473633,-80.4554672241211,-80.3583984375,-80.2823715209961,-80.384765625,-80.4683074951172,-80.4822769165039,-80.3212890625,-80.2370147705078,-80.1333999633789,-80.046142578125,-80.0250015258789,-80.06103515625,-80.0882797241211,-80.0359344482422,-79.9035110473633,-79.7958526611328,-79.7412109375,-79.6131820678711,-79.4653778076172,-79.2290573120117,-78.899658203125,-78.8270492553711,-78.8596725463867,-78.828857421875,-78.7371063232422,-78.681640625,-78.587646484375,-78.5115203857422,-78.3121109008789,-78.1806640625,-78.0370178222656,-77.8295440673828,-77.702880859375,-77.6731948852539,-77.6486282348633,-77.601318359375,-77.526123046875,-77.4813995361328,-77.4676742553711,-77.4227523803711,-77.3963394165039,-77.2926712036133,-77.1657180786133,-77.1141128540039,-77.00244140625,-76.9201126098633,-76.829345703125,-76.7677230834961,-76.7393035888672,-76.72900390625,-76.6785202026367,-76.60302734375,-76.49462890625,-76.4272994995117,-76.41796875,-76.4133758544922,-76.38818359375,-76.31103515625,-76.2706069946289,-76.0679168701172,-76.0261764526367,-75.974853515625,-75.8797912597656,-75.7766571044922,-75.6173324584961,-75.4639663696289,-75.2844696044922],[34.1981468200684,34.2124977111816,34.2453117370605,34.3285140991211,34.4009742736816,34.4899406433105,34.5177764892578,34.5296859741211,34.6585922241211,34.7350616455078,34.7911109924316,34.8698234558105,34.904296875,34.8485374450684,34.7364273071289,34.6171875,34.4464836120605,34.4271469116211,34.3997077941895,34.3185539245605,34.2201194763184,34.0451202392578,33.76025390625,33.5941429138184,33.4161148071289,33.2477531433105,33.2019538879395,33.2037124633789,33.1301765441895,33.0757827758789,32.87060546875,32.8117218017578,32.7666969299316,32.7214813232422,32.6471672058105,32.5657234191895,32.4730491638184,32.4894561767578,32.4085960388184,32.3597679138184,32.3972625732422,32.5650367736816,32.5990257263184,32.6380844116211,32.6318359375,32.6588859558105,32.7844696044922,32.8294906616211,32.8565444946289,32.8982429504395,33.0228500366211,33.2021484375,33.3722648620605,33.4949226379395,33.5470733642578,33.5587882995605,33.5498046875,33.6574211120605,33.697265625,33.8016586303711,33.8493194580078,33.89306640625,33.9590835571289,34.0495109558105,34.3292961120605,34.5651359558105,34.6792945861816,34.8532218933105,35.1941413879395,35.3970680236816,35.4778327941895,35.6247062683105,35.7838859558105,35.6320304870605,35.5938453674316,35.5408210754395,35.5152359008789,35.50439453125,35.5227546691895,35.5643539428711,35.6978492736816,35.79736328125,35.8458023071289,35.9133796691895,36.2296867370605,36.41455078125,36.8296890258789,36.8704109191895,36.8713874816895,36.5432624816895,36.2152328491211,35.8870124816895,35.5589866638184,35.2308616638184,34.9027328491211,34.5746078491211,34.2464828491211,33.91845703125,33.59033203125,33.26220703125,32.93408203125,32.6060562133789,32.27783203125,31.9498043060303,31.6216793060303,31.4344730377197,31.4664039611816,31.4861335754395,31.4642562866211,31.4002914428711,31.3584938049316,31.2606449127197,31.2091808319092,31.0926780700684,30.7106437683105,30.3286151885986,29.9466800689697,29.5645484924316,29.1825199127197,28.8005847930908,28.4185543060303,28.0364265441895,27.6544914245605,27.2724609375,26.8904285430908,26.5083980560303,26.1263675689697,25.7443351745605,25.3623046875,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9716796875,24.9161128997803,24.8659191131592,24.8108406066895,24.8037109375,24.7116203308105,24.7032241821289,24.7264652252197,24.8775405883789,24.9230480194092,24.96142578125,24.9739265441895,24.9294929504395,24.8775405883789,24.8599605560303,24.8527336120605,24.9299812316895,25.0226554870605,25.0572261810303,25.1120109558105,25.1504878997803,25.2254886627197,25.3822269439697,25.8932609558105,26.4573230743408,26.7686519622803,27.2480449676514,27.5400390625,27.6201171875,27.8299789428711,27.9675769805908,28.5148448944092,28.8069343566895,28.9727535247803,29.0720710754395,29.1599636077881,29.2789077758789,29.4285163879395,29.5916042327881,29.9297866821289,30.0494136810303,30.1275367736816,30.2226543426514,30.2623043060303,30.3123035430908,30.34375,30.3951168060303,30.5709972381592,30.9235343933105,30.8841781616211,30.5629901885986,30.7004871368408,30.8414058685303,31.0017566680908,31.0308609008789,31.0519523620605,31.0829105377197,31.1939449310303,31.5244140625,31.6065425872803,31.8392581939697,31.8889656066895,31.9642581939697,32.1360359191895,32.0760726928711,31.8921890258789,31.8758773803711,31.7710933685303,31.9020519256592,32.0084953308105,32.0656242370605,32.1017608642578,32.20654296875,32.2818374633789,32.2427711486816,32.2162094116211,32.2505874633789,32.3235359191895,32.5328140258789,32.6033210754395,32.6845703125,32.8544921875,32.9015617370605,33.1298828125,33.15673828125,33.1943359375,33.3779296875,33.66650390625,33.9025382995605,34.17626953125,34.1981468200684],[40.1412086486816,40.1825180053711,40.21142578125,40.2340812683105,40.2500953674316,40.408203125,40.3990211486816,40.3046875,40.19580078125,40.0951156616211,39.9751968383789,39.9474601745605,40.02392578125,40.0634765625,40.0705070495605,40.0163078308105,39.93994140625,39.9452133178711,39.9793968200684,40.00048828125,39.9567413330078,40.0425796508789,40.0967750549316,40.1324234008789,40.1412086486816],[40.0764656066895,40.1100616455078,40.0124015808105,39.99609375,40.0390625,40.0481452941895,40.0764656066895],[38.6094703674316,38.9117164611816,39.0344734191895,39.142578125,39.2225608825684,39.2989234924316,39.4222640991211,39.5065422058105,39.5788078308105,39.6312522888184,39.7208023071289,39.7855491638184,39.8194313049316,39.8156242370605,39.7903327941895,39.8134765625,39.86376953125,39.9783210754395,40.041015625,40.0578117370605,40.0840835571289,40.2041015625,40.3052749633789,40.4365234375,40.5462875366211,40.6343727111816,40.79931640625,41.1764640808105,41.4796867370605,41.658203125,42.2451171875,42.3464813232422,42.3993186950684,42.5228538513184,42.7344703674316,42.7961921691895,42.9695320129395,42.9990234375,43.0829124450684,43.11669921875,43.0056648254395,42.88330078125,42.8659172058105,42.8252944946289,42.7674827575684,42.7037124633789,42.6701164245605,42.4793968200684,42.4500007629395,42.4085922241211,42.3785171508789,42.2899436950684,42.2250022888184,42.13427734375,42.0465812683105,41.9521484375,41.8595695495605,41.7650375366211,41.625,41.3628921508789,41.1223640441895,40.9385719299316,40.8201179504395,40.76953125,40.5244140625,40.3531227111816,40.2214851379395,40.140625,40.0621109008789,39.8951187133789,39.7561531066895,39.6979522705078,39.6048812866211,39.5318374633789,39.4460945129395,39.2701187133789,39.1980476379395,39.1585960388184,39.1354522705078,39.07421875,39.0238265991211,38.9957008361816,38.81201171875,38.50439453125,38.4314460754395,38.376953125,38.2214851379395,38.1770515441895,38.1419944763184,38.0699195861816,38.0025367736816,37.9434547424316,37.8841819763184,37.8203125,37.7083969116211,37.6484375,37.5711898803711,37.5467758178711,37.5072250366211,37.3537101745605,37.2572288513184,37.1851539611816,37.1326179504395,37.0994148254395,37.0634765625,37.0245132446289,36.9407234191895,36.8119125366211,36.6791038513184,36.5423812866211,36.5243186950684,36.4922866821289,36.4707984924316,36.4481430053711,36.4267578125,36.5217781066895,36.5660171508789,36.67919921875,36.7245140075684,36.8134765625,36.8258819580078,36.9137725830078,36.9054679870605,36.8877906799316,36.9357414245605,36.9787139892578,36.9757804870605,36.9952125549316,37.0089836120605,37.0615234375,37.1695327758789,37.2488288879395,37.3404273986816,37.4110374450684,37.4529304504395,37.5101547241211,37.5474624633789,37.5759735107422,37.65673828125,37.7259750366211,37.7824211120605,37.8033218383789,37.8629913330078,37.9225578308105,37.9500999450684,38.0252914428711,38.0989265441895,38.1485366821289,38.1815414428711,38.2190437316895,38.2535171508789,38.2672843933105,38.2898406982422,38.3473625183105,38.3737297058105,38.3855476379395,38.3971672058105,38.4224624633789,38.5228500366211,38.6094703674316],[-17.887939453125,-17.9847660064697,-18.1065921783447,-18.1359367370605,-18.1605472564697,-18.0433597564697,-17.9245109558105,-17.887939453125],[-15.4005861282349,-15.4066905975342,-15.3831539154053,-15.3891611099243,-15.436767578125,-15.5593738555908,-15.65576171875,-15.7103023529053,-15.8073234558105,-15.8094730377197,-15.720947265625,-15.6827630996704,-15.4527835845947,-15.4327154159546,-15.4154787063599,-15.4005861282349],[-17.1846694946289,-17.2253913879395,-17.27392578125,-17.3249034881592,-17.2903327941895,-17.2585945129395,-17.21435546875,-17.129638671875,-17.103759765625,-17.10107421875,-17.1846694946289],[-16.33447265625,-16.418212890625,-16.4962406158447,-16.5427722930908,-16.6580085754395,-16.7947273254395,-16.8660144805908,-16.9053230285645,-16.8430652618408,-16.7520503997803,-16.5568370819092,-16.5174312591553,-16.3189945220947,-16.1236324310303,-16.119140625,-16.33447265625],[-14.1967782974243,-14.3326177597046,-14.4686031341553,-14.4917964935303,-14.3555669784546,-14.2319822311401,-14.152587890625,-14.0283689498901,-14.0033693313599,-13.9541511535645,-13.8862791061401,-13.8572273254395,-13.8271484375,-13.8275880813599,-13.8629884719849,-13.9280281066895,-14.1967782974243],[-17.8342781066895,-17.859375,-17.8821277618408,-18.0007801055908,-17.9288082122803,-17.7975597381592,-17.7445316314697,-17.7265625,-17.7516117095947,-17.744384765625,-17.7580070495605,-17.8342781066895],[-13.7159662246704,-13.7839841842651,-13.8599119186401,-13.8236322402954,-13.7881832122803,-13.6500978469849,-13.5350589752197,-13.5014162063599,-13.4635744094849,-13.4229488372803,-13.4537591934204,-13.4779291152954,-13.5546884536743,-13.7159662246704],[1.59394514560699,1.57119119167328,1.50498020648956,1.40576171875,1.40195322036743,1.41718757152557,1.43632781505585,1.49687516689301,1.59267604351044,1.59394514560699],[1.44521498680115,1.40898454189301,1.25693356990814,1.22333991527557,1.25625014305115,1.2998046875,1.30253934860229,1.34863257408142,1.56445324420929,1.61318385601044,1.62363290786743,1.49453115463257,1.44521498680115],[3.14531254768372,3.24111342430115,3.34218764305115,3.39589858055115,3.44892573356628,3.46181607246399,3.41464853286743,3.34873032569885,3.29296898841858,3.24472641944885,3.15458989143372,3.07285165786743,2.90009737014771,2.7998046875,2.76982402801514,2.74599599838257,2.70058608055115,2.63408184051514,2.57587909698486,2.49951171875,2.45878863334656,2.39433622360229,2.37001919746399,2.37128901481628,2.78496098518372,2.90478563308716,3.15869164466858,3.19755887985229,3.16445326805115,3.1669921875,3.19863271713257,3.19091773033142,3.15869164466858,3.14531254768372],[4.29365253448486,4.27529287338257,3.96767616271973,3.86718726158142,3.84267544746399,3.84541010856628,3.85341763496399,4.05917978286743,4.22578144073486,4.31513643264771,4.32207012176514,4.29365253448486],[-1.79404306411743,-1.79272484779358,-1.75327146053314,-1.71284186840057,-1.62714862823486,-1.56147444248199,-1.47172856330872,-1.41069316864014,-1.40732419490814,-1.42260730266571,-1.45942378044128,-1.48046851158142,-1.46084010601044,-1.42875969409943,-1.39404296875,-1.37050759792328,-1.35273444652557,-1.31884741783142,-1.30004870891571,-1.30156242847443,-1.28544914722443,-1.17543947696686,-0.933838069438934,-0.839209020137787,-0.762646377086639,-0.740185260772705,-0.586425721645355,-0.54980456829071,-0.481152206659317,-0.398437738418579,-0.338573932647705,-0.299316585063934,-0.256054818630219,-0.205322295427322,-0.140038907527924,-0.0814940854907036,-0.0411618612706661,0,0.201367154717445,0.255468726158142,0.312890529632568,0.377246290445328,0.517675936222076,0.631640672683716,0.641992330551147,0.651757836341858,0.669824302196503,0.696874797344208,0.764452934265137,1.01005876064301,1.11113309860229,1.20830094814301,1.29326200485229,1.34941411018372,1.42832016944885,1.41484367847443,1.42197239398956,1.43027341365814,1.42812478542328,1.44882810115814,1.48623073101044,1.53408193588257,1.58642566204071,1.67851531505585,1.70605456829071,1.85976552963257,1.92792952060699,1.95146512985229,1.98652315139771,2.03271508216858,2.09834003448486,2.20039057731628,2.37441396713257,2.56796884536743,2.65166020393372,2.65478491783142,2.67001938819885,2.70185518264771,2.74941420555115,2.81562519073486,2.89140629768372,2.97001957893372,3.05263662338257,3.15214848518372,3.21142554283142,3.23984360694885,3.28789091110229,3.30673837661743,3.21865248680115,3.16640591621399,3.150390625,3.17519497871399,3.224609375,3.23808598518372,3.24804663658142,3.14687490463257,3.0048828125,2.31093740463257,2.14560556411743,2.08261728286743,1.56660187244415,1.20585954189301,1.03290998935699,0.816894590854645,0.714648544788361,0.796093940734863,0.891113221645355,0.859179437160492,0.720605552196503,0.660058856010437,0.627148687839508,0.596093833446503,0.363671809434891,0.158398196101189,0.0430663749575615,0,-0.0751462802290916,-0.327001929283142,-0.328955113887787,-0.204931527376175,-0.133789077401161,-0.0341310948133469,0,0.154883086681366,0.201562538743019,0.136327847838402,0,-0.0527344644069672,-0.381250143051147,-0.520800828933716,-0.550683438777924,-0.646777093410492,-0.683202922344208,-0.741552948951721,-0.752734184265137,-0.814648568630219,-0.823095679283142,-0.721581876277924,-0.771874964237213,-0.822168052196503,-0.938086032867432,-1.32753920555115,-1.64096665382385,-1.79760766029358,-1.93930697441101,-2.11152362823486,-2.18769550323486,-2.30556631088257,-2.45283174514771,-2.59570336341858,-2.67060542106628,-2.78754925727844,-2.90185546875,-3.14916968345642,-3.25913071632385,-3.43124985694885,-3.57880854606628,-3.82778310775757,-4.36684608459473,-4.43486356735229,-4.50224637985229,-4.67412090301514,-4.935302734375,-5.17148447036743,-5.23051738739014,-5.32968711853027,-5.36093759536743,-5.381591796875,-5.40722703933716,-5.44360303878784,-5.46250009536743,-5.55126953125,-5.62548828125,-5.80839824676514,-5.960693359375,-6.04067373275757,-6.17045927047729,-6.22626924514771,-6.26591825485229,-6.25771522521973,-6.26894569396973,-6.38413095474243,-6.41225576400757,-6.32832050323486,-6.25942420959473,-6.216796875,-6.32094717025757,-6.39619112014771,-6.492431640625,-6.88461875915527,-6.859375,-6.86377000808716,-6.92949199676514,-6.97465848922729,-7.17495059967041,-7.40615272521973,-7.46718788146973,-7.49604463577271,-7.50351524353027,-7.44394588470459,-7.37890625,-7.292236328125,-7.18544960021973,-7.072509765625,-7.02285146713257,-6.98110342025757,-6.95756864547729,-6.97480487823486,-7.10639667510986,-7.343017578125,-7.33579158782959,-7.30595731735229,-7.28637647628784,-7.28154277801514,-7.21992254257202,-7.12548828125,-7.04604482650757,-7.00624990463257,-6.99794864654541,-7.04296827316284,-7.17241239547729,-7.30576181411743,-7.33544921875,-7.36269569396973,-7.44511699676514,-7.52421855926514,-7.53569316864014,-7.45410203933716,-7.11767625808716,-7.04741239547729,-7.03671884536743,-6.97539043426514,-6.91118144989014,-6.89609384536743,-6.91640615463257,-7.02783203125,-7.03261661529541,-7.01469707489014,-6.94843721389771,-6.85888719558716,-6.81015634536743,-6.82177782058716,-6.84794950485229,-6.85205078125,-6.835693359375,-6.829833984375,-6.818359375,-6.83588838577271,-6.85771465301514,-6.92846727371216,-6.91552734375,-6.8828125,-6.77578115463257,-6.69013643264771,-6.56591796875,-6.40312480926514,-6.28935575485229,-6.24433565139771,-6.21250009536743,-6.22168016433716,-6.24311542510986,-6.30805683135986,-6.39169979095459,-6.48466825485229,-6.54218721389771,-6.55898427963257,-6.55258750915527,-6.55751991271973,-6.57534170150757,-6.61826229095459,-6.70361280441284,-6.77729511260986,-6.83320283889771,-6.86552762985229,-7.03046846389771,-7.09912157058716,-7.14711904525757,-7.17792940139771,-7.19536161422729,-7.19833993911743,-7.20961904525757,-7.26855516433716,-7.40361356735229,-7.51259756088257,-7.61259746551514,-7.64467811584473,-7.69306659698486,-7.89638662338257,-7.92084980010986,-7.990966796875,-8.09443378448486,-8.15249061584473,-8.17353534698486,-8.18125057220459,-8.22475624084473,-8.21333026885986,-8.12998104095459,-8.13930702209473,-8.173583984375,-8.20419979095459,-8.21308517456055,-8.26606369018555,-8.32255935668945,-8.5380859375,-8.58964824676514,-8.68295955657959,-8.77714824676514,-8.85234355926514,-8.87832069396973,-8.88720607757568,-8.7724609375,-8.69091796875,-8.72919940948486,-8.81581974029541,-8.80996131896973,-8.76938533782959,-8.73002910614014,-8.77617168426514,-8.81210899353027,-8.80991172790527,-8.79990196228027,-8.8115234375,-8.98779296875,-9.03310489654541,-9.03505897521973,-8.93720722198486,-8.92719745635986,-9.04160213470459,-9.127197265625,-9.179443359375,-9.23520565032959,-9.23564529418945,-9.17807579040527,-9.09555625915527,-9.02451133728027,-8.87368202209473,-8.66562461853027,-8.53706073760986,-8.42158126831055,-8.35546875,-8.24892520904541,-8.25229549407959,-8.28886699676514,-8.25673866271973,-8.13715744018555,-8.00468730926514,-7.85273408889771,-7.69814491271973,-7.59458017349243,-7.50361347198486,-7.39931631088257,-7.261962890625,-7.06098651885986,-6.90068340301514,-6.61728477478027,-6.47568368911743,-6.22412109375,-6.08012676239014,-5.8466796875,-5.66582012176514,-5.31572246551514,-5.10527372360229,-4.52304697036743,-4.31279325485229,-4.01533174514771,-3.88935565948486,-3.77402353286743,-3.60463833808899,-3.52363252639771,-3.41787075996399,-3.04560518264771,-2.94770526885986,-2.87504911422729,-2.60707998275757,-2.33710908889771,-2.19667959213257,-1.99130856990814,-1.82851564884186,-1.79404306411743],[22.6173820495605,22.6883792877197,22.7538089752197,22.8201179504395,22.9642562866211,23.2928714752197,23.3232421875,23.1271476745605,23.0826168060303,23.0354480743408,22.9798831939697,22.8851566314697,22.7570304870605,22.7302742004395,22.4984378814697,22.3716793060303,22.2693347930908,22.2273426055908,22.1524410247803,22.0762691497803,21.9968757629395,21.97802734375,21.9855480194092,22.1529293060303,22.1876945495605,22.1043949127197,22.0345706939697,21.8821296691895,21.8544921875,21.8910160064697,21.9244136810303,21.9650402069092,21.9840831756592,21.8623046875,21.9244136810303,22.0018558502197,22.0813484191895,22.1685562133789,22.20556640625,22.2666015625,22.328125,22.4744148254395,22.5469741821289,22.6173820495605],[23.3435554504395,23.2603511810303,23.0634765625,23.1090831756592,23.1654300689697,23.3328132629395,23.3564453125,23.3435554504395],[22.9237308502197,22.8416996002197,22.7928714752197,22.7672843933105,22.6614246368408,22.5421867370605,22.47265625,22.4789066314697,22.4110355377197,22.3074226379395,22.1619148254395,22.0562496185303,22.4625988006592,22.5045909881592,22.5872058868408,22.6494140625,22.7022457122803,22.7122058868408,22.7254867553711,22.9098625183105,22.9816417694092,23.0086917877197,22.9237308502197],[28.0124988555908,28.0658206939697,28.1330070495605,28.1510734558105,28.1283206939697,28.0613288879395,28.0460929870605,28.0164070129395,27.9381828308105,27.8976573944092,27.8495101928711,27.7576179504395,27.6217765808105,27.5130863189697,27.4644527435303,27.4341793060303,27.4270515441895,27.5313472747803,27.5300788879395,27.5055656433105,27.48779296875,27.50244140625,27.5710926055908,27.6441402435303,27.6734371185303,27.7219715118408,27.7687511444092,27.7785167694092,27.7769527435303,27.7528324127197,27.5420894622803,27.5147457122803,27.4919929504395,27.3999996185303,27.3717784881592,27.3542957305908,27.3519535064697,27.3265628814697,27.1871089935303,27.0333995819092,26.9660148620605,26.8998050689697,26.8197269439697,26.5326175689697,26.4621105194092,26.2980461120605,26.2150402069092,26.0303726196289,26.0152339935303,25.9911136627197,25.7937507629395,25.7208976745605,25.66015625,25.5712890625,25.3400402069092,25.2826156616211,25.2686519622803,25.2726554870605,25.25830078125,25.2287101745605,25.1751956939697,25.1110343933105,24.9113292694092,24.8390617370605,24.7757816314697,24.4588871002197,24.3624992370605,24.3225574493408,24.33203125,24.4638671875,24.4875011444092,24.5357418060303,24.5497074127197,24.5291004180908,24.3921871185303,24.3369140625,24.2872066497803,24.2356433868408,24.1148433685303,24.0109367370605,23.767578125,23.7060546875,23.6915035247803,23.5627937316895,23.50927734375,23.5306644439697,23.6474609375,23.6807613372803,23.5335941314697,23.5036125183105,23.4971694946289,23.4320316314697,23.4896488189697,23.5150394439697,23.4677734375,23.4801750183105,23.5169925689697,23.4944343566895,23.6405277252197,23.7825183868408,24.0833988189697,24.0536117553711,24.1753902435303,24.38037109375,24.5835933685303,24.8775405883789,25.4437503814697,25.5208988189697,25.5074214935303,25.50927734375,25.61572265625,25.7937507629395,26.4608402252197,26.625,26.85205078125,26.9747066497803,27.3358402252197,27.892578125,28.0018539428711,28.0124988555908],[42.3785171508789,42.2803688049316,42.1491203308105,41.9958992004395,41.9496078491211,41.8156242370605,41.7926788330078,41.7665061950684,41.7646484375,41.7820320129395,41.7982406616211,41.8721656799316,41.9574203491211,42.0521507263184,42.1662101745605,42.30810546875,42.4651374816895,42.5577125549316,42.6549797058105,42.7412109375,42.7830085754395,42.8441390991211,42.9227523803711,42.9061508178711,42.8628883361816,42.8097648620605,42.7630882263184,42.6595726013184,42.6564445495605,42.6692352294922,42.7251968383789,42.78369140625,42.81640625,42.8415985107422,42.9125022888184,43.0147438049316,43.0689468383789,43.181640625,43.2184562683105,43.3031272888184,43.3943367004395,43.4825210571289,43.5810546875,43.6205062866211,43.8267593383789,43.9837875366211,44.0228500366211,44.3062477111816,44.6320304870605,44.8935546875,45.2269515991211,45.5554695129395,45.86328125,46.2959938049316,46.6447296142578,46.9195289611816,46.9782218933105,47.3056640625,47.6376953125,47.9782218933105,47.7316398620605,47.4528312683105,47.1597633361816,46.97119140625,46.6717796325684,46.4229469299316,46.1667938232422,45.9349632263184,45.6335945129395,45.4384765625,45.1328125,44.9405288696289,44.91162109375,44.6366233825684,44.3695297241211,44.0281219482422,43.9888687133789,43.8894538879395,43.8291969299316,43.58349609375,43.5382843017578,43.333984375,43.1256828308105,43.0160140991211,42.9309577941895,42.8947257995605,42.8566398620605,42.7916030883789,42.3551750183105,42.2284164428711,42.0241203308105,41.9153289794922,41.8839836120605,41.7376937866211,41.48193359375,41.3724594116211,41.3189468383789,41.2208976745605,41.1404304504395,41.0872077941895,41.0208015441895,40.8726577758789,40.7652320861816,40.5287094116211,40.3160133361816,40.01416015625,39.8421859741211,39.7903327941895,39.6575202941895,39.5388641357422,39.4944343566895,39.2254905700684,39.1283187866211,38.9677734375,38.7527351379395,38.6080055236816,38.4515647888184,38.2252922058105,38.0861320495605,37.9449195861816,37.7628898620605,37.5754890441895,37.3825187683105,37.1545906066895,36.9055671691895,36.8482437133789,36.8236312866211,36.5530281066895,36.2718734741211,36.0819320678711,36.02197265625,35.9787101745605,35.9198265075684,35.8456039428711,35.7630844116211,35.7561531066895,35.779296875,35.80029296875,35.7884750366211,35.7914085388184,35.7450180053711,35.4686546325684,35.4240264892578,35.3779296875,35.3252944946289,35.28759765625,35.2646484375,35.2638664245605,35.2683563232422,35.25244140625,35.1644554138184,35.0819358825684,35.0319328308105,34.9835929870605,34.958984375,34.8978500366211,34.8380851745605,34.7492179870605,34.7106475830078,34.6387672424316,34.5627937316895,34.484375,34.279296875,34.2003898620605,34.0642585754395,34.0301742553711,34.0204124450684,33.9779319763184,33.9024429321289,33.6661109924316,33.6009750366211,33.5163078308105,33.3922843933105,33.2259788513184,33.0807609558105,33.0146484375,32.9989242553711,33.0125961303711,33.0652351379395,33.1652336120605,33.2342758178711,33.2810516357422,33.4093742370605,33.5453147888184,33.6448249816895,33.7850608825684,33.9533195495605,34.0197257995605,34.07275390625,34.0945320129395,34.1017570495605,34.1015625,34.0910148620605,34.0845718383789,34.0771484375,34.078125,34.0792961120605,34.1203117370605,34.1590805053711,34.1852531433105,34.29150390625,34.3112297058105,34.3148422241211,34.2756805419922,34.3439483642578,34.4314460754395,34.5080070495605,34.5718765258789,34.6017570495605,34.6749992370605,34.7712860107422,34.8162078857422,34.88232421875,34.9314460754395,34.9249000549316,34.9691429138184,34.9607429504395,35.0079078674316,35.0596694946289,35.0827140808105,35.1123046875,35.25244140625,35.3727531433105,35.4496116638184,35.5960960388184,35.6702156066895,35.7305679321289,35.8206062316895,35.9875984191895,36.1075172424316,36.1251945495605,36.1353530883789,36.1371116638184,36.16015625,36.2122077941895,36.2735366821289,36.3068351745605,36.3462867736816,36.390625,36.4470710754395,36.4439468383789,36.5243186950684,36.5423812866211,36.6791038513184,36.8119125366211,36.9407234191895,37.0245132446289,37.0634765625,37.0994148254395,37.1326179504395,37.1851539611816,37.2572288513184,37.3537101745605,37.5072250366211,37.5467758178711,37.5711898803711,37.6484375,37.7083969116211,37.8203125,37.8841819763184,37.9434547424316,38.0025367736816,38.0699195861816,38.1419944763184,38.1770515441895,38.2214851379395,38.376953125,38.4314460754395,38.50439453125,38.81201171875,38.9957008361816,39.0238265991211,39.07421875,39.1354522705078,39.1585960388184,39.1980476379395,39.2701187133789,39.4460945129395,39.5318374633789,39.6048812866211,39.6979522705078,39.7561531066895,39.8951187133789,40.0621109008789,40.140625,40.2214851379395,40.3531227111816,40.5244140625,40.76953125,40.8201179504395,40.9385719299316,41.1223640441895,41.3628921508789,41.625,41.7650375366211,41.8595695495605,41.9521484375,42.0465812683105,42.13427734375,42.2250022888184,42.2899436950684,42.3785171508789],[21.6283206939697,21.5406246185303,21.4860343933105,21.5067367553711,21.5679683685303,21.6340808868408,21.6481437683105,21.6283206939697],[21.8332042694092,21.7331047058105,21.6950206756592,21.7047863006592,21.7642574310303,21.8643550872803,21.8332042694092],[22.1750965118408,22.3017578125,22.35498046875,22.41552734375,22.3128910064697,22.3057613372803,22.3462886810303,22.3605461120605,22.25830078125,22.2093734741211,22.1880855560303,22.1405277252197,22.0771484375,22.1082038879395,22.1258792877197,22.1750965118408],[21.9942378997803,21.9214839935303,21.8186511993408,21.8056640625,21.8459968566895,21.8193359375,21.8272457122803,21.9068355560303,21.9502925872803,21.9078121185303,21.9797840118408,21.9942378997803],[21.4508781433105,21.4369144439697,21.3690433502197,21.2999992370605,21.2443351745605,21.2145500183105,21.2247066497803,21.26806640625,21.30126953125,21.4508781433105],[21.2177734375,21.228515625,21.287109375,21.3660144805908,21.4219722747803,21.4156246185303,21.3776359558105,21.3671855926514,21.3184566497803,21.3097648620605,21.25341796875,21.1493167877197,21.0838871002197,21.236328125,21.2217769622803,21.2177734375],[24.8482418060303,24.6989250183105,24.57861328125,24.5765628814697,24.6511726379395,24.7860336303711,24.9706039428711,24.99755859375,24.8917961120605,24.8482418060303],[28.9658222198486,28.89892578125,28.6921863555908,28.5660171508789,28.4140625,28.4535160064697,28.7059574127197,28.7448234558105,28.7728519439697,28.7776355743408,28.7520503997803,28.4792976379395,28.470703125,28.5601558685303,28.6851558685303,29.06298828125,29.3438491821289,29.5242195129395,29.8215827941895,29.9792003631592,29.9880847930908,29.9412097930908,29.7505855560303,29.5722675323486,29.3876953125,29.2433586120605,29.0870132446289,29.0690422058105,29.0662117004395,29.0930671691895,29.2932605743408,29.3711929321289,29.4643535614014,29.5443344116211,29.5907230377197,29.6708984375,29.7207050323486,29.8035163879395,29.9034194946289,29.9366207122803,30.0874996185303,30.1027336120605,30.0953121185303,30.0290050506592,29.8826179504395,29.7239246368408,29.7159194946289,29.8194351196289,29.7280254364014,29.71484375,29.6171894073486,29.6080055236816,29.6124019622803,29.6296882629395,29.8105449676514,29.826171875,29.8269538879395,29.8108386993408,29.7200183868408,29.6224613189697,29.6008796691895,29.6041984558105,29.6374988555908,29.7016620635986,29.783203125,30.0728511810303,30.1102542877197,30.1261711120605,30.1201171875,29.9855480194092,29.9866199493408,30.0418930053711,30.1081047058105,30.3906269073486,30.4878921508789,30.5137710571289,30.5279293060303,30.5260753631592,30.5039043426514,30.4153308868408,30.2102546691895,30.0041027069092,29.9915046691895,30.0553703308105,30.4185543060303,30.6552715301514,30.9748058319092,31.1808586120605,31.2474594116211,31.3367195129395,31.43701171875,31.50927734375,31.5365238189697,31.5339832305908,31.4373054504395,31.3824214935303,31.28564453125,31.1867179870605,30.9357433319092,30.5656261444092,30.4796886444092,30.3064441680908,30.0099620819092,29.9332008361816,29.6901359558105,29.5793952941895,29.4923820495605,29.2516613006592,28.9929676055908,28.7390632629395,28.6628894805908,28.5681648254395,28.455078125,28.4074211120605,28.1519527435303,27.7976551055908,27.7616214752197,27.6693363189697,27.5250968933105,27.46240234375,27.2418937683105,27.2052726745605,27.0755844116211,26.951171875,26.7214851379395,26.607421875,26.53466796875,26.5197277069092,26.5511722564697,26.6017589569092,26.6064453125,26.5693359375,26.4958019256592,26.4564456939697,26.3777351379395,26.2046890258789,26.0360355377197,25.9559574127197,26.0062503814697,26.0402336120605,26.0358390808105,25.9458999633789,25.8458003997803,25.7580070495605,25.7154293060303,25.6564464569092,25.5482425689697,25.4557609558105,25.2678718566895,25.1558589935303,24.9576168060303,24.8487300872803,24.6004886627197,24.5179672241211,24.4456062316895,24.3425788879395,24.0251960754395,23.7217788696289,23.5926761627197,23.4635753631592,23.3267574310303,23.1814460754395,23.0212898254395,22.9638671875,23.009765625,23.11572265625,23.1884765625,23.1984367370605,23.1484375,23.0801753997803,22.994140625,22.9117183685303,22.8670883178711,22.8444328308105,22.8191413879395,22.79345703125,22.7498054504395,22.6973648071289,22.6461906433105,22.4626960754395,22.4385757446289,22.4385757446289,22.4710941314697,22.4426765441895,22.4697265625,22.5129890441895,22.5642566680908,22.5899429321289,22.5879878997803,22.5166988372803,22.5123043060303,22.5758781433105,22.5849609375,22.5603523254395,22.5205078125,22.2579097747803,21.9339847564697,21.8542976379395,21.8052730560303,21.7271499633789,21.61328125,21.52783203125,21.43603515625,21.41064453125,21.4119148254395,21.4040050506592,21.37890625,21.3605480194092,21.3777332305908,21.4509773254395,21.4791011810303,21.5134773254395,21.5211925506592,21.5017585754395,21.5066394805908,21.5650386810303,21.5523433685303,21.5266609191895,21.4982433319092,21.5224609375,21.5923824310303,21.5980472564697,21.60595703125,21.5518550872803,21.5456047058105,21.4705085754395,21.3848628997803,21.2559566497803,21.3016605377197,21.3537101745605,21.3433589935303,21.3234367370605,21.1656246185303,21.1421871185303,21.1036128997803,21.1181640625,21.1438484191895,21.1957035064697,21.45751953125,21.4735355377197,21.6509780883789,21.5686531066895,21.5492191314697,21.5451164245605,21.8003902435303,21.8957023620605,22.1203117370605,22.3197269439697,22.3162097930908,22.2855472564697,22.2432632446289,22.2732410430908,22.3459968566895,22.3125972747803,22.3186531066895,22.3980464935303,22.5276374816895,22.5323238372803,22.7562503814697,23.0144519805908,23.1335945129395,23.2487316131592,23.4939460754395,23.5989246368408,23.6529293060303,23.8614273071289,23.9248027801514,24.0222644805908,24.2783203125,24.4406242370605,24.5301761627197,24.5579109191895,24.6576175689697,24.74755859375,24.9421882629395,25.13427734375,25.2142581939697,25.2881851196289,25.28076171875,25.22802734375,25.2710933685303,25.3726577758789,25.3623046875,25.3402347564697,25.255859375,25.2978515625,25.3079109191895,25.3478507995605,25.2417964935303,24.83935546875,24.7642574310303,24.6749019622803,24.58154296875,24.6232414245605,24.6280288696289,24.5916023254395,24.5326175689697,24.404296875,24.2374992370605,24.1554698944092,24.0490226745605,23.99462890625,23.9073238372803,23.75146484375,23.7209968566895,23.7002944946289,23.6935539245605,23.673828125,23.6820316314697,23.701171875,23.7683601379395,23.8655281066895,23.8858394622803,23.8941421508789,23.9388675689697,23.9885730743408,23.97607421875,23.9417972564697,23.8693351745605,23.7589836120605,23.6773433685303,23.6415042877197,23.623046875,23.6260738372803,23.6566410064697,23.7609367370605,23.77490234375,23.7335948944092,23.6608390808105,23.537109375,23.4680671691895,23.4548835754395,23.4514656066895,23.4654293060303,23.5044918060303,23.5370121002197,23.5413093566895,23.5001945495605,23.48779296875,23.5018558502197,23.6329097747803,23.6388664245605,23.4742202758789,23.35546875,23.3185539245605,23.1825199127197,23.0978527069092,22.9753894805908,22.8541011810303,22.7824230194092,22.3621101379395,22.1951160430908,21.9974613189697,21.8501949310303,21.7240219116211,21.6160144805908,21.4654293060303,21.42236328125,21.259765625,21.1833972930908,20.9185543060303,20.9089851379395,20.9070320129395,20.8951168060303,20.6221675872803,20.6758785247803,20.8892574310303,21.0657215118408,21.1044921875,21.1278324127197,21.0526371002197,21.0661125183105,21.1437511444092,21.2667961120605,21.4612293243408,21.59375,21.6217765808105,21.8197250366211,21.9894523620605,22.0796871185303,22.3003902435303,22.3829097747803,22.4109382629395,22.5006828308105,22.81103515625,23.0716781616211,23.1443367004395,23.3240242004395,23.4625015258789,23.70703125,23.7725582122803,23.85400390625,23.9973621368408,24.1541023254395,24.33203125,24.4905281066895,24.7032241821289,24.8024425506592,24.94140625,25.0869140625,25.1728515625,25.2491207122803,25.3571281433105,25.4808597564697,25.5752925872803,25.6466789245605,25.7483386993408,25.7681636810303,25.7486324310303,25.7671871185303,25.8501949310303,25.9615230560303,26.0115242004395,26.0724601745605,26.1561527252197,26.3082027435303,26.525390625,26.5842781066895,26.740234375,26.9342765808105,27.1086921691895,27.1275386810303,27.2056636810303,27.3480472564697,27.5916996002197,27.7478523254395,27.8899421691895,28.0472660064697,28.2691402435303,28.4117183685303,28.8003902435303,29.1416034698486,29.3333988189697,29.2388687133789,29.1917991638184,29.02490234375,28.8462905883789,28.8326148986816,28.8918952941895,28.9658222198486],[174.629684448242,174.621871948242,174.592971801758,174.587203979492,174.604202270508,174.627731323242,174.629684448242],[-178.711624145508,-178.709518432617,-178.714950561523,-178.723098754883,-178.729095458984,-178.730560302734,-178.7275390625,-178.724563598633,-178.719192504883,-178.714202880859,-178.711624145508],[-178.535110473633,-178.546340942383,-178.57373046875,-178.595947265625,-178.598678588867,-178.589309692383,-178.567672729492,-178.556686401367,-178.56298828125,-178.576171875,-178.574066162109,-178.55712890625,-178.540618896484,-178.535110473633],[178.487899780273,178.487686157227,178.358978271484,178.315628051758,178.287994384766,178.211334228516,178.189163208008,178.181838989258,178.162109375,178.020812988281,177.958694458008,178.000778198242,178.051956176758,178.104095458984,178.156646728516,178.208404541016,178.2822265625,178.334274291992,178.420318603516,178.487899780273],[-179.799850463867,-179.797607421875,-179.812454223633,-179.830215454102,-179.83935546875,-179.845504760742,-179.848587036133,-179.851211547852,-179.865036010742,-179.867340087891,-179.86279296875,-179.856201171875,-179.831100463867,-179.799850463867],[-178.761917114258,-178.773635864258,-178.827346801758,-178.847900390625,-178.790863037109,-178.763092041016,-178.761917114258],[179.349319458008,179.340423583984,179.253524780273,179.256439208984,179.271789550781,179.306442260742,179.337890625,179.362411499023,179.349319458008],[-178.988082885742,-179.018417358398,-179.039215087891,-179.063812255859,-179.079010009766,-179.047607421875,-178.999130249023,-178.988082885742],[-178.251113891602,-178.306838989258,-178.357223510742,-178.325393676758,-178.280334472656,-178.254592895508,-178.251113891602],[178.827529907227,178.776077270508,178.747665405273,178.787109375,178.8310546875,178.8525390625,178.827529907227],[178.280181884766,178.280181884766,178.309478759766,178.338577270508,178.410934448242,178.523239135742,178.591613769531,178.595703125,178.574905395508,178.603805541992,178.617874145508,178.667678833008,178.597351074219,178.486724853516,178.461135864258,178.423431396484,178.33154296875,178.243743896484,178.16015625,178.06396484375,177.955474853516,177.847076416016,177.770797729492,177.636428833008,177.457336425781,177.383209228516,177.321380615234,177.263473510742,177.2548828125,177.263961791992,177.316314697266,177.360153198242,177.366409301758,177.3857421875,177.410934448242,177.423248291016,177.405578613281,177.400680541992,177.504486083984,177.617965698242,177.817962646484,177.940231323242,178.127624511719,178.187606811523,178.247177124023,178.280181884766],[179.42236328125,179.388961791992,179.373138427734,179.407623291016,179.432815551758,179.447174072266,179.42236328125],[-178.956481933594,-178.981842041016,-179.00390625,-178.975524902344,-178.971481323242,-179.014938354492,-179.017669677734,-179.005020141602,-178.952835083008,-178.921142578125,-178.914840698242,-178.924560546875,-178.956481933594],[177.234176635742,177.182815551758,177.210159301758,177.2392578125,177.257522583008,177.287399291992,177.275787353516,177.234176635742],[180,179.925888061523,179.89697265625,179.930953979492,180,180.106399536133,180.139007568359,180.177688598633,180.132217407227,180.025100708008,180],[180,179.987197875977,179.984664916992,180,180.056350708008,180.099075317383,180.072647094727,180.070556640625,180],[180,179.848236083984,179.793838500977,179.748153686523,179.619140625,179.564163208008,179.568161010742,179.697067260742,179.841018676758,179.884963989258,179.930358886719,179.926559448242,179.905960083008,179.890045166016,179.927932739258,179.82080078125,179.714752197266,179.588958740234,179.465438842773,179.419326782227,179.375,179.346008300781,179.323333740234,179.300491333008,179.202346801758,179.055465698242,179.0068359375,178.950393676758,178.883697509766,178.802932739258,178.706634521484,178.6650390625,178.638092041016,178.603713989258,178.497451782227,178.513763427734,178.5419921875,178.567764282227,178.583587646484,178.63427734375,178.686325073242,178.744537353516,178.805084228516,178.86572265625,178.960540771484,179.091415405273,179.224609375,179.293563842773,179.359176635742,179.47509765625,179.5517578125,179.63525390625,179.715042114258,179.788864135742,179.84814453125,180,180.030624389648,180.055419921875,180.043838500977,180],[177.121490478516,177.082427978516,177.019332885742,177.006256103516,177.0263671875,177.067565917969,177.118057250977,177.126953125,177.121490478516],[-59.6826629638672,-59.74658203125,-59.7644538879395,-59.7848625183105,-59.7859382629395,-59.7933120727539,-59.7532196044922,-59.6810035705566,-59.6826629638672],[-58.4388198852539,-58.4327125549316,-58.5128440856934,-58.5414085388184,-58.4970741271973,-58.4605445861816,-58.4388198852539],[-61.0187530517578,-60.947265625,-60.8759727478027,-60.9161605834961,-60.9475593566895,-61.031982421875,-61.1157760620117,-61.14501953125,-61.0516586303711,-61.0187530517578],[-60.2862281799316,-60.1415519714355,-60.0086898803711,-59.9170913696289,-59.8416023254395,-59.7884292602539,-59.7113304138184,-59.4934539794922,-59.465087890625,-59.3875999450684,-59.3208503723145,-59.26806640625,-59.2939453125,-59.3542022705078,-59.3924293518066,-59.4370079040527,-59.5142097473145,-59.5731887817383,-59.7148971557617,-59.92138671875,-59.9897499084473,-60.1322746276855,-60.1937484741211,-60.246337890625,-60.2882843017578,-60.3534622192383,-60.3842277526855,-60.4520034790039,-60.4840850830078,-60.5083999633789,-60.6863746643066,-60.8122062683105,-60.96142578125,-60.7625045776367,-60.591064453125,-60.4497566223145,-60.3344764709473,-60.2886734008789,-60.2384796142578,-60.2381362915039,-60.2765159606934,-60.3283233642578,-60.3795890808105,-60.5000953674316,-60.58251953125,-60.528076171875,-60.4672355651855,-60.2809562683105,-60.2451629638672,-60.3026351928711,-60.4149436950684,-60.5058097839355,-60.5227584838867,-60.5182609558105,-60.5684585571289,-60.5157203674316,-60.4454612731934,-60.2862281799316],[-60.1117172241211,-60.2488250732422,-60.27587890625,-60.2753410339355,-60.1713829040527,-60.0698280334473,-60.0764656066895,-60.1117172241211],[-58.8501930236816,-58.6975135803223,-58.5062522888184,-58.4258270263672,-58.3787117004395,-58.40673828125,-58.4674301147461,-58.5192375183105,-58.5089378356934,-58.4737281799316,-58.2715797424316,-58.2345199584961,-58.2411117553711,-58.2762184143066,-58.289306640625,-58.2592239379883,-58.2064476013184,-57.9765090942383,-57.9225120544434,-57.8084945678711,-57.9154281616211,-57.9604454040527,-57.8663597106934,-57.7917938232422,-57.8311500549316,-57.8381843566895,-58.0039558410645,-58.1509246826172,-58.2176284790039,-58.3359832763672,-58.6834945678711,-58.6430702209473,-58.6376914978027,-58.6527824401855,-59.1312484741211,-59.1958503723145,-59.0680160522461,-59.1627922058105,-59.2563438415527,-59.3415031433105,-59.3956527709961,-59.5322303771973,-59.6487312316895,-59.649169921875,-59.5366706848145,-59.57080078125,-59.3087387084961,-59.2617683410645,-59.1800308227539,-59.0954132080078,-59.0595169067383,-59.0653800964355,-59.0994606018066,-59.0966339111328,-58.8866653442383,-58.9174766540527,-58.8501930236816],[55.79736328125,55.6561508178711,55.5576171875,55.3626976013184,55.3103523254395,55.2328147888184,55.25,55.3113288879395,55.4504890441895,55.5964813232422,55.6619110107422,55.7391586303711,55.8390655517578,55.8224601745605,55.79736328125],[45.1802749633789,45.1175804138184,45.0876922607422,45.0694351196289,45.0882797241211,45.0935554504395,45.0425758361816,45.0923843383789,45.134765625,45.1587905883789,45.22314453125,45.2042961120605,45.2085952758789,45.1793975830078,45.1802749633789],[-51.6525382995605,-51.6834487915039,-51.76708984375,-51.8052749633789,-51.8274879455566,-51.8794898986816,-51.9289054870605,-51.9443359375,-51.9906234741211,-51.99951171875,-52.1161117553711,-52.16259765625,-52.2294425964355,-52.271240234375,-52.327880859375,-52.3566398620605,-52.3566398620605,-52.3963851928711,-52.4184112548828,-52.4558601379395,-52.5546875,-52.5594749450684,-52.5830078125,-52.6531753540039,-52.7006340026855,-52.7833976745605,-52.8704109191895,-52.9034652709961,-52.96484375,-53.0097160339355,-53.082275390625,-53.1800765991211,-53.2297859191895,-53.252197265625,-53.2854995727539,-53.3344230651855,-53.3660163879395,-53.4318351745605,-53.5089874267578,-53.56396484375,-53.6836929321289,-53.7347183227539,-53.7501449584961,-53.7677764892578,-53.7942390441895,-53.8295402526855,-53.8766098022461,-53.9464340209961,-54.0897483825684,-54.1300773620605,-54.1673812866211,-54.2279777526855,-54.2930679321289,-54.43310546875,-54.5150871276855,-54.5504875183105,-54.5919418334961,-54.6162605285645,-54.604736328125,-54.5684089660645,-54.5359382629395,-54.4855461120605,-54.4020042419434,-54.2567367553711,-54.1955070495605,-54.1880836486816,-54.1707000732422,-54.203125,-54.1880378723145,-54.0631828308105,-54.0095710754395,-54.0059585571289,-53.990478515625,-54.0059089660645,-54.0342292785645,-54.0819816589355,-54.11279296875,-54.1974639892578,-54.2555198669434,-54.3507308959961,-54.3421401977539,-54.369140625,-54.3983879089355,-54.396240234375,-54.416015625,-54.440673828125,-54.4496078491211,-54.4260749816895,-54.4402351379395,-54.4711456298828,-54.4796867370605,-54.4733390808105,-54.4468765258789,-54.4521980285645,-54.3316421508789,-54.2401847839355,-54.1559600830078,-54.0853042602539,-53.9895973205566,-53.919921875,-53.84716796875,-53.4544448852539,-53.2703590393066,-52.8993148803711,-52.7649917602539,-52.4539527893066,-52.29052734375,-52.2889175415039,-52.3246116638184,-52.219970703125,-52.05810546875,-52.0123023986816,-51.9619140625,-51.9793434143066,-52.001708984375,-52.0029296875,-51.9547843933105,-51.9276847839355,-51.9195823669434,-51.8802719116211,-51.8275413513184,-51.78564453125,-51.6986351013184,-51.6658210754395,-51.6532707214355,-51.6581077575684,-51.6525382995605],[-60.8262710571289,-60.8366165161133,-60.8621063232422,-60.8994140625,-61.063720703125,-61.0888710021973,-61.09033203125,-61.0113258361816,-61.1043014526367,-61.14111328125,-61.2197303771973,-61.2133293151855,-61.1808128356934,-61.1273918151855,-61.027099609375,-60.9525413513184,-60.9271469116211,-60.9186553955078,-60.9336891174316,-60.8891639709473,-60.8699684143066,-60.8262710571289],[-61.23046875,-61.2862281799316,-61.3107414245605,-61.3184051513672,-61.2752952575684,-61.25,-61.2123527526855,-61.2034187316895,-61.23046875],[-61.5895500183105,-61.6704597473145,-61.7102546691895,-61.7594261169434,-61.7940940856934,-61.7671356201172,-61.748046875,-61.6415023803711,-61.5970230102539,-61.5523452758789,-61.5750503540039,-61.5638694763184,-61.5895500183105],[-61.3271484375,-61.44482421875,-61.5221710205078,-61.5399894714355,-61.5005874633789,-61.5289039611816,-61.5106430053711,-61.47119140625,-61.4064483642578,-61.3961410522461,-61.3554649353027,-61.172607421875,-61.3271484375],[9.48037052154541,9.45419883728027,9.47324275970459,9.50937461853027,9.52617168426514,9.55644512176514,9.55068302154541,9.42841815948486,9.40087890625,9.39482402801514,9.37421894073486,9.33086013793945,9.25341796875,9.18613243103027,9.00302791595459,8.89501953125,8.84208965301514,8.80751991271973,8.82978534698486,8.87900352478027,8.88681602478027,8.77099609375,8.71796894073486,8.71865177154541,8.75869178771973,8.74042987823486,8.67363262176514,8.62187480926514,8.61513710021973,8.65341854095459,8.70253849029541,8.70097637176514,8.6416015625,8.58779335021973,8.56621170043945,8.60791015625,8.67548751831055,8.62587928771973,8.59238338470459,8.56562519073486,8.58750057220459,8.64003944396973,8.71308612823486,8.81484413146973,8.99492168426514,9.04365158081055,9.08837985992432,9.13789081573486,9.19804668426514,9.25351524353027,9.28769588470459,9.31337833404541,9.33837890625,9.32304668426514,9.33095741271973,9.36318397521973,9.41523456573486,9.46328163146973,9.46084022521973,9.47861289978027,9.48037052154541],[-1.17832028865814,-1.21357440948486,-1.28027355670929,-1.36870110034943,-1.38886737823486,-1.38867211341858,-1.28505837917328,-1.17832028865814],[5.78974580764771,5.82343769073486,5.9013671875,5.92890644073486,5.95947265625,6.01142597198486,6.07412099838257,6.11992216110229,6.18105459213257,6.24218702316284,6.27734375,6.34433603286743,6.38222646713257,6.45810508728027,6.53427696228027,6.56630802154541,6.57470703125,6.60761785507202,6.73544931411743,6.77626895904541,6.82070350646973,6.84951162338257,6.89121103286743,6.95830106735229,7.00146484375,7.02216768264771,7.03671884536743,7.06572246551514,7.11738252639771,7.19990205764771,7.31337881088257,7.40419912338257,7.45058631896973,7.52548885345459,7.61093807220459,7.79921865463257,8.00126934051514,8.08066368103027,8.13486385345459,8.14033222198486,8.1240234375,7.92275381088257,7.83798789978027,7.79482460021973,7.76513624191284,7.70566415786743,7.61660194396973,7.58417987823486,7.60849618911743,7.59326219558716,7.53857469558716,7.52939462661743,7.56543016433716,7.61562490463257,7.49492216110229,7.46738243103027,7.42001962661743,7.34316349029541,7.26572227478027,7.203125,7.16747999191284,7.16923809051514,7.13603544235229,7.05341815948486,6.96835899353027,6.900390625,6.92148447036743,6.98408222198486,7.00058555603027,7.00058555603027,6.97851514816284,6.95205068588257,6.82070350646973,6.68808603286743,6.66689491271973,6.62480449676514,6.45625019073486,6.43857431411743,6.42900419235229,6.41015625,6.28515625,6.16074228286743,6.12968730926514,6.10703086853027,6.06796884536743,6.06025362014771,6.12324237823486,6.11591815948486,6.09589862823486,6.0361328125,5.97001934051514,5.97148418426514,6.00664091110229,6.08662128448486,6.19941425323486,6.27294969558716,6.22958993911743,6.22421884536743,6.23466730117798,6.32187509536743,6.42890596389771,6.57822227478027,6.75810527801514,6.77607440948486,6.76738262176514,6.7841796875,6.81679677963257,6.77207040786743,6.80566453933716,6.85800790786743,6.89726543426514,6.95371103286743,7.00390625,7.02109336853027,6.94082021713257,6.80449199676514,6.78916025161743,6.79091835021973,6.80624961853027,6.88144540786743,6.96240186691284,7.013671875,7.12607431411743,7.15341806411743,7.14638662338257,7.11679697036743,7.07832002639771,7.03242206573486,6.98124980926514,6.84228515625,6.78037118911743,6.69228506088257,6.62773466110229,6.63476610183716,6.69140625,6.72470712661743,6.73818349838257,6.80107402801514,6.88935565948486,6.93984365463257,6.97285175323486,6.99267578125,7.03066396713257,7.00791025161743,6.96035146713257,6.93193340301514,6.87519502639771,6.84296894073486,6.87861347198486,6.89384794235229,6.87480497360229,6.90019559860229,6.96728563308716,7.1494140625,7.31855440139771,7.37089872360229,7.59941434860229,7.63720655441284,7.6650390625,7.67714881896973,7.65146493911743,7.58964824676514,7.52265596389771,7.48203086853027,7.49052667617798,7.4931640625,7.43867206573486,7.43691396713257,7.41445302963257,7.39501953125,7.38007831573486,7.37773466110229,7.26152324676514,7.18144512176514,6.86474609375,6.71660137176514,6.68740224838257,6.6572265625,6.57021474838257,6.49404287338257,6.30537128448486,6.11591815948486,6.03056669235229,5.80947303771973,5.67158222198486,5.40654325485229,5.32021474838257,5.19951200485229,5.12041044235229,5.07314491271973,5.06083965301514,5.05976581573486,4.97597646713257,4.91191387176514,4.87373018264771,4.84355497360229,4.80790996551514,4.78720664978027,4.7890625,4.71210908889771,4.62871122360229,4.40976572036743,4.37617206573486,4.22421884536743,4.16279268264771,4.11308574676514,4.07509756088257,4.05263662338257,3.91083955764771,3.86162066459656,3.78476524353027,3.25888633728027,3.16289091110229,3.0517578125,3.04306650161743,3.09091830253601,3.19785165786743,3.21142554283142,3.15214848518372,3.05263662338257,2.97001957893372,2.89140629768372,2.81562519073486,2.74941420555115,2.70185518264771,2.67001938819885,2.65478491783142,2.65166020393372,2.56796884536743,2.37441396713257,2.20039057731628,2.09834003448486,2.03271508216858,1.98652315139771,1.95146512985229,1.92792952060699,1.85976552963257,1.70605456829071,1.71396493911743,1.74023449420929,1.73945295810699,1.70986306667328,1.56816422939301,1.50136709213257,1.45888686180115,1.42832016944885,1.34941411018372,1.29326200485229,1.20830094814301,1.11113309860229,1.01005876064301,0.764452934265137,0.696874797344208,0.669824302196503,0.651757836341858,0.641992330551147,0.631640672683716,0.517675936222076,0.377246290445328,0.312890529632568,0.255468726158142,0.201367154717445,0,-0.0411618612706661,-0.0814940854907036,-0.140038907527924,-0.205322295427322,-0.256054818630219,-0.299316585063934,-0.338573932647705,-0.398437738418579,-0.481152206659317,-0.54980456829071,-0.586425721645355,-0.740185260772705,-0.762646377086639,-0.839209020137787,-0.933838069438934,-1.17543947696686,-1.28544914722443,-1.30156242847443,-1.30004870891571,-1.31884741783142,-1.35273444652557,-1.37050759792328,-1.39404296875,-1.42875969409943,-1.46084010601044,-1.48046851158142,-1.45942378044128,-1.42260730266571,-1.40732419490814,-1.41069316864014,-1.47172856330872,-1.56147444248199,-1.62714862823486,-1.71284186840057,-1.75327146053314,-1.79272484779358,-1.79404306411743,-1.63144516944885,-1.48486304283142,-1.34599590301514,-1.24550771713257,-1.17080080509186,-1.07695329189301,-1.15288102626801,-1.20039069652557,-1.22031247615814,-1.24521493911743,-1.18906271457672,-1.14907193183899,-1.08100581169128,-0.941747784614563,-0.826318562030792,-0.766650199890137,-0.691113173961639,-0.633984088897705,-0.548486173152924,-0.582275211811066,-0.64111340045929,-0.73310524225235,-0.790771722793579,-0.880664229393005,-1.16997063159943,-1.19599616527557,-1.20996081829071,-1.11435544490814,-1.03173828125,-1.04150378704071,-1.06601548194885,-1.10439455509186,-1.13637697696686,-1.13203108310699,-1.14628922939301,-1.23881840705872,-1.31279301643372,-1.39248049259186,-1.78652369976044,-1.92143571376801,-2.05937480926514,-2.09248042106628,-2.09028291702271,-2.01889634132385,-2.08193397521973,-2.1435546875,-2.19707036018372,-2.14858365058899,-2.10830068588257,-2.02758765220642,-1.92172861099243,-1.82128894329071,-1.74252903461456,-1.97539055347443,-2.35302710533142,-2.43442368507385,-2.50312471389771,-2.530029296875,-2.476318359375,-2.42768573760986,-2.48271489143372,-2.55405282974243,-2.66591835021973,-2.77031230926514,-2.79677724838257,-2.73310542106628,-2.78720712661743,-2.859375,-2.96406245231628,-3.06420922279358,-3.15883779525757,-3.22158193588257,-3.26469755172729,-3.32861328125,-3.39589858055115,-3.44394516944885,-3.50781273841858,-3.90092754364014,-4.07070302963257,-4.22641658782959,-4.31210994720459,-4.37509775161743,-4.427978515625,-4.67880916595459,-4.62919902801514,-4.51240253448486,-4.37783193588257,-4.32944345474243,-4.43461942672729,-4.51220703125,-4.54433584213257,-4.5771484375,-4.53066444396973,-4.49790048599243,-4.4033203125,-4.24140596389771,-4.30175828933716,-4.36440420150757,-4.39316415786743,-4.52480459213257,-4.584716796875,-4.71938467025757,-4.74853467941284,-4.76250028610229,-4.72075176239014,-4.53120136260986,-4.05888700485229,-3.85566401481628,-3.71479463577271,-3.54599618911743,-3.47148418426514,-3.2314453125,-3.00322246551514,-2.79287147521973,-2.69233369827271,-2.44619131088257,-2.07944321632385,-2.00371074676514,-1.97314453125,-1.90571260452271,-1.85195302963257,-1.82470726966858,-1.43764615058899,-1.37646460533142,-1.48046851158142,-1.56547832489014,-1.58310532569885,-1.69033205509186,-1.81342744827271,-1.87006843090057,-1.87539064884186,-1.85644519329071,-1.70512700080872,-1.58823275566101,-1.36572277545929,-1.25864267349243,-1.26494133472443,-1.23227524757385,-1.19497048854828,-1.13852560520172,-0.959130823612213,-0.765527188777924,-0.520898282527924,-0.163476318120956,-0.0111818443983793,0,0.136133044958115,0.416894406080246,0.439258068799973,0.27763643860817,0.129394486546516,0.109374783933163,0.126562371850014,0.186718925833702,0.616210997104645,0.924121260643005,1.24550771713257,1.40722680091858,1.51406276226044,1.54843735694885,1.5927734375,1.55156230926514,1.57949221134186,1.60957026481628,1.67226552963257,1.76767563819885,1.91249990463257,2.44570302963257,2.52490234375,2.53603482246399,2.57480478286743,2.60146498680115,2.57929682731628,2.59677720069885,2.66914057731628,2.75937509536743,2.83974623680115,2.86240243911743,2.92197251319885,3.02285122871399,3.10683608055115,3.15488266944885,3.18203091621399,3.23496103286743,3.24980473518372,3.27333998680115,3.31621098518372,3.47695302963257,3.59541034698486,3.62675833702087,3.66728520393372,3.68935537338257,3.71884799003601,3.74804711341858,3.78857398033142,3.85810542106628,3.94970679283142,4.04414033889771,4.17460966110229,4.16962862014771,4.14414072036743,4.13525390625,4.15771436691284,4.19218730926514,4.18388652801514,4.15029335021973,4.13681697845459,4.13701200485229,4.14931631088257,4.17607402801514,4.36875009536743,4.54501962661743,4.65615224838257,4.67509794235229,4.70664072036743,4.77285146713257,4.81865215301514,4.86054706573486,4.7900390625,4.84150362014771,4.84912109375,4.86757802963257,4.93056678771973,5.00693368911743,5.06103515625,5.12412118911743,5.21503877639771,5.27880859375,5.30195331573486,5.353515625,5.43466806411743,5.50732421875,5.54238319396973,5.61005830764771,5.71044921875,5.78974580764771],[-6.69946336746216,-6.6796875,-6.70302772521973,-6.7705078125,-6.88813495635986,-6.92924785614014,-6.93486356735229,-6.90590810775757,-6.88164091110229,-6.77001953125,-6.74062490463257,-6.74106454849243,-6.70351552963257,-6.69946336746216],[-6.62319326400757,-6.64277362823486,-6.670166015625,-6.76425790786743,-6.83915996551514,-6.86396503448486,-6.884765625,-6.841796875,-6.790771484375,-6.662109375,-6.62582969665527,-6.62319326400757],[-7.18686485290527,-7.09711933135986,-7.065185546875,-7.11679697036743,-7.17939472198486,-7.25493144989014,-7.37910175323486,-7.422607421875,-7.33676719665527,-7.23530292510986,-7.18686485290527],[-6.63105487823486,-6.65581083297729,-6.69643592834473,-6.76889610290527,-6.82343769073486,-6.84052705764771,-6.83769559860229,-6.80947256088257,-6.72255849838257,-6.71440410614014,-6.72519493103027,-6.80971717834473,-7.01357412338257,-7.17216825485229,-6.95864248275757,-6.80366230010986,-6.63105487823486],[-6.40605497360229,-6.453857421875,-6.52470684051514,-6.54414033889771,-6.55947303771973,-6.55205059051514,-6.55458974838257,-6.47304677963257,-6.40605497360229],[162.983200073242,162.993469238281,162.929885864258,162.921096801758,162.958206176758,162.983200073242],[158.314834594727,158.256546020508,158.183395385742,158.160842895508,158.127639770508,158.134765625,158.186126708984,158.294631958008,158.3349609375,158.309371948242,158.314834594727],[151.647750854492,151.639450073242,151.578323364258,151.569732666016,151.575088500977,151.604293823242,151.607803344727,151.592864990234,151.605667114258,151.629486083984,151.643264770508,151.650482177734,151.647750854492],[151.881439208984,151.8642578125,151.85595703125,151.859970092773,151.865341186523,151.8818359375,151.910552978516,151.91259765625,151.881439208984],[138.142684936523,138.067092895508,138.061920166016,138.085067749023,138.116882324219,138.14697265625,138.185836791992,138.213577270508,138.182510375977,138.142684936523],[13.2935543060303,13.2886714935303,13.20947265625,13.1721677780151,13.1626949310303,13.1845703125,13.2227535247803,13.2473640441895,13.2283201217651,13.1901359558105,13.21630859375,13.2741212844849,13.3723640441895,13.5233402252197,13.72119140625,13.8513669967651,14.0662107467651,14.1808595657349,14.23974609375,14.3030271530151,14.3344717025757,14.3864259719849,14.4298830032349,14.4391603469849,14.4344730377197,14.390625,14.3415031433105,14.3242177963257,14.2831058502197,14.23095703125,14.0874996185303,14.0655269622803,14.0252933502197,13.9496097564697,13.9151363372803,13.8845710754395,13.890625,13.87548828125,13.8600587844849,13.8980464935303,14.0694332122803,14.1028318405151,14.1483402252197,14.2067375183105,14.36376953125,14.4247074127197,14.4741220474243,14.4805660247803,14.4449224472046,14.4106435775757,14.4240245819092,14.4369144439697,14.45556640625,14.447265625,14.4029293060303,14.4029293060303,14.4232425689697,14.3839836120605,14.3585948944092,14.2883796691895,14.25146484375,14.2396488189697,14.2017583847046,14.1628904342651,14.1628904342651,14.2003898620605,14.1998043060303,14.1297845840454,14.0874013900757,13.9938478469849,13.8869132995605,13.86181640625,13.8876953125,13.8785161972046,13.8416013717651,13.7843751907349,13.7337894439697,13.70556640625,13.6185541152954,13.4649410247803,13.3573246002197,13.1585941314697,12.9919919967651,12.91357421875,12.8644533157349,12.7935543060303,12.7136716842651,12.62841796875,12.5904293060303,12.4686527252197,12.43212890625,12.4324226379395,12.4437503814697,12.4625968933105,12.478515625,12.4756841659546,12.4538087844849,12.4463863372803,12.064453125,11.9982423782349,11.9502925872803,11.8923826217651,11.7267580032349,11.6659173965454,11.60546875,11.5777339935303,11.5751953125,11.6034183502197,11.5945310592651,11.55712890625,11.5377931594849,11.6390619277954,11.6756830215454,11.7113285064697,11.7601566314697,11.7634773254395,11.7080078125,11.6890630722046,11.7154302597046,11.7843742370605,11.8850584030151,11.9341793060303,11.9292964935303,11.8828125,11.8647451400757,11.8329095840454,11.8394536972046,11.884765625,11.8798828125,11.84912109375,11.7864265441895,11.7333984375,11.6857423782349,11.5368165969849,11.5042972564697,11.2882804870605,11.2344722747803,11.1900386810303,11.1301755905151,11.0320310592651,10.947265625,10.8485355377197,10.6407222747803,10.58544921875,10.34765625,10.0061521530151,9.75947284698486,9.72207069396973,9.763671875,10.001953125,10.0344724655151,10.06201171875,9.95908164978027,9.86083984375,9.76865291595459,9.67636775970459,9.62460899353027,9.59101581573486,9.57402324676514,9.533203125,9.40224647521973,9.37050724029541,9.29892635345459,9.34248065948486,9.48281288146973,9.49531269073486,9.48320293426514,9.34218788146973,9.265625,9.24794864654541,9.25839805603027,9.15751934051514,9.05283260345459,9.03632831573486,9.31884670257568,9.35664081573486,9.40634822845459,9.52334022521973,9.50107383728027,9.44833946228027,9.39716815948486,9.33066463470459,9.29580020904541,9.28017520904541,9.3466796875,9.31787109375,9.29667949676514,9.26015663146973,9.20380783081055,9.06464767456055,8.94189453125,8.90937519073486,8.87656211853027,8.84423828125,8.703125,8.75722694396973,8.82138633728027,8.94638633728027,8.99521446228027,9.03789043426514,9.08154296875,9.13652420043945,9.29667949676514,9.33906269073486,9.32529354095459,9.30185508728027,9.35488319396973,9.37578105926514,9.38613319396973,9.4111328125,9.46816444396973,9.57431602478027,9.73837947845459,9.79677772521973,9.81269550323486,10.00146484375,9.94443321228027,9.77665996551514,9.54648494720459,9.47011661529541,9.39882850646973,9.32480430603027,9.32997989654541,9.49531269073486,9.53896427154541,9.556640625,9.60107421875,9.61796855926514,9.62529277801514,9.62587833404541,9.57539081573486,9.5908203125,9.63613224029541,9.67646503448486,9.70458984375,9.76054668426514,9.78867149353027,9.80390548706055,9.8603515625,9.90673828125,9.94668006896973,9.97978496551514,10.0285158157349,10.1789064407349,10.3154296875,10.5872068405151,10.85888671875,11.1306638717651,11.3353519439697,11.3346681594849,11.3335943222046,11.3323249816895,11.3311519622803,11.330078125,11.3287115097046,11.3399410247803,11.3533201217651,11.3484382629395,11.5589847564697,11.9397459030151,12.1061525344849,12.1534185409546,12.361328125,12.5297861099243,12.6013669967651,12.6657228469849,12.8674802780151,13.130859375,13.2203130722046,13.2699222564697,13.2935543060303],[-1.06557595729828,-1.14936530590057,-1.17583024501801,-1.19609355926514,-1.25146472454071,-1.30629909038544,-1.51533186435699,-1.56342768669128,-1.51567399501801,-1.38583993911743,-1.31279301643372,-1.14423859119415,-1.06557595729828],[-4.19677734375,-4.15488290786743,-4.04936552047729,-4.08427762985229,-4.20039081573486,-4.27861309051514,-4.373046875,-4.41884756088257,-4.47197294235229,-4.55322265625,-4.56787109375,-4.46171855926514,-4.31508779525757,-4.19677734375],[-6.218017578125,-6.30366230010986,-6.36367177963257,-6.402587890625,-6.44028377532959,-6.54814481735229,-6.64980459213257,-6.66420936584473,-6.64687442779541,-6.66953134536743,-6.76660203933716,-6.80258798599243,-6.85834980010986,-6.86923885345459,-6.87724637985229,-6.93613243103027,-7.00771522521973,-7.04970741271973,-7.13349580764771,-7.20258712768555,-7.17807626724243,-7.15546894073486,-7.19306612014771,-7.30673837661743,-7.32451200485229,-7.35517549514771,-7.40942335128784,-7.54443407058716,-7.60654258728027,-7.67876005172729,-7.85493183135986,-7.88447237014771,-7.91845703125,-8.11826229095459,-8.14482402801514,-8.11894512176514,-8.04433631896973,-7.79379892349243,-7.75439500808716,-7.74628925323486,-7.81982469558716,-7.88613319396973,-7.90874004364014,-7.91059541702271,-7.87295007705688,-7.79726552963257,-7.73749971389771,-7.68999052047729,-7.6064453125,-7.55039072036743,-7.50219678878784,-7.45126914978027,-7.44599628448486,-7.40141582489014,-7.37690401077271,-7.21865177154541,-7.17861318588257,-7.10063505172729,-7.03076171875,-6.94716787338257,-6.88896512985229,-6.82485294342041,-6.69882822036743,-6.47504901885986,-6.37529277801514,-6.23422813415527,-6.12914991378784,-6.03579139709473,-5.98574209213257,-5.86918926239014,-5.71684551239014,-5.71074199676514,-5.76518535614014,-5.87910175323486,-5.87861347198486,-5.803466796875,-5.73862314224243,-5.58252000808716,-5.52792930603027,-5.49018526077271,-5.47041034698486,-5.48388671875,-5.52587890625,-5.56855440139771,-5.615966796875,-5.67109346389771,-5.64609384536743,-5.65595674514771,-5.63188457489014,-5.55781221389771,-5.60678768157959,-5.70805644989014,-5.826171875,-5.85463857650757,-5.87607383728027,-5.937744140625,-6.01904296875,-6.11953163146973,-6.218017578125],[-5.10542011260986,-5.23149394989014,-5.27705049514771,-5.33149433135986,-5.39267587661743,-5.37080097198486,-5.345703125,-5.318115234375,-5.25161123275757,-5.18544912338257,-5.160400390625,-5.10498094558716,-5.0947265625,-5.10542011260986],[-6.12890577316284,-6.09282255172729,-6.0576171875,-6.05532217025757,-6.08837890625,-6.25317335128784,-6.30507802963257,-6.30722618103027,-6.27001953125,-6.30205106735229,-6.28642606735229,-6.30175733566284,-6.33388662338257,-6.45195341110229,-6.49135780334473,-6.49565410614014,-6.46645498275757,-6.46284198760986,-6.44526338577271,-6.41318368911743,-6.37495136260986,-6.34414052963257,-6.311279296875,-6.21567344665527,-6.12890577316284],[-5.97006845474243,-5.99091768264771,-6.04155302047729,-6.06035184860229,-6.07070350646973,-6.07197237014771,-6.04130887985229,-5.91176795959473,-5.97031211853027,-5.97265625,-5.93906307220459,-5.79960918426514,-5.76225614547729,-5.72514629364014,-5.79721689224243,-5.97006845474243],[-5.77788066864014,-6.17617177963257,-6.31342792510986,-6.32582998275757,-6.29848623275757,-6.18486356735229,-6.13886690139771,-6.31064462661743,-6.31967735290527,-6.30625009536743,-6.28632831573486,-6.18207979202271,-6.13828134536743,-6.10273408889771,-6.02959012985229,-5.94668006896973,-5.83603525161743,-5.76083993911743,-5.77788066864014],[-6.60761785507202,-6.66445302963257,-6.66855478286743,-6.56992197036743,-6.50605487823486,-6.48369121551514,-6.53007793426514,-6.60761785507202],[-7.41689491271973,-7.50478458404541,-7.53740215301514,-7.54296875,-7.52294874191284,-7.45546913146973,-7.40668964385986,-7.39892625808716,-7.41689491271973],[-6.279052734375,-6.30874061584473,-6.34624052047729,-6.38339853286743,-6.4326171875,-6.32236337661743,-6.27822303771973,-6.26127910614014,-6.26054716110229,-6.279052734375],[-7.24985313415527,-7.29204034805298,-7.347412109375,-7.38149452209473,-7.41591787338257,-7.42236328125,-7.40703105926514,-7.41054677963257,-7.29638624191284,-7.26713848114014,-7.24755907058716,-7.24985313415527],[-6.14472675323486,-6.14614248275757,-6.16376924514771,-6.14082050323486,-6.13554668426514,-6.09340858459473,-6.067626953125,-5.88027334213257,-5.70600557327271,-5.67246103286743,-5.66865253448486,-5.69619131088257,-5.79541015625,-5.91376924514771,-5.94907236099243,-5.9873046875,-6.01474618911743,-6.03437471389771,-6.16274452209473,-6.26611328125,-6.32270479202271,-6.36240243911743,-6.44243144989014,-6.67543983459473,-6.74130821228027,-6.76113319396973,-6.75273418426514,-6.70419883728027,-6.64345693588257,-6.60585975646973,-6.5830078125,-6.58349609375,-6.61528301239014,-6.61679697036743,-6.37851572036743,-6.35766553878784,-6.30595684051514,-6.24692392349243,-6.16606473922729,-6.14472675323486],[-7.20556592941284,-7.0927734375,-7.18261766433716,-7.320556640625,-7.51474571228027,-7.515625,-7.49941444396973,-7.47031259536743,-7.44003868103027,-7.39189434051514,-7.32485389709473,-7.27119159698486,-7.20556592941284],[-6.19868135452271,-6.32582998275757,-6.37558650970459,-6.41928768157959,-6.55458974838257,-6.4365234375,-6.40336894989014,-6.40244150161743,-6.42519521713257,-6.578125,-6.68330049514771,-6.79658174514771,-6.853759765625,-6.91035175323486,-6.95693349838257,-6.98310518264771,-7.01318311691284,-7.08344745635986,-6.95595693588257,-6.94414043426514,-6.85683584213257,-6.86416053771973,-7.00253868103027,-7.05707979202271,-7.05190420150757,-6.98530292510986,-7.01689434051514,-7.03823232650757,-7.076904296875,-7.08847618103027,-7.09560585021973,-7.08525419235229,-7.044921875,-7.02841806411743,-7.01206064224243,-6.94956016540527,-6.88622999191284,-6.81230497360229,-6.72646522521973,-6.72470712661743,-6.78774404525757,-6.77646446228027,-6.74228525161743,-6.54418897628784,-6.29716777801514,-6.23745107650757,-6.21943330764771,-6.19423770904541,-6.19868135452271],[-3.10966777801514,-3.10112309455872,-3.11289048194885,-3.13676738739014,-3.21235394477844,-3.41098618507385,-3.77499985694885,-3.99003911018372,-4.01962900161743,-4.03559541702271,-3.90683579444885,-3.85712909698486,-3.88793921470642,-4.07841777801514,-4.134521484375,-3.98847651481628,-3.8681640625,-3.62822246551514,-3.40278315544128,-3.29453134536743,-3.08393573760986,-3.03603506088257,-2.94667959213257,-2.85629892349243,-2.24414038658142,-2.07407236099243,-1.96152341365814,-1.86738288402557,-1.77792978286743,-1.78066408634186,-1.83471691608429,-1.93447279930115,-2.02031278610229,-2.04550814628601,-2.06235361099243,-2.08955073356628,-2.26025366783142,-2.42666006088257,-2.50097680091858,-2.59267568588257,-2.68095731735229,-2.77519559860229,-3.04741191864014,-3.12358355522156,-3.21445298194885,-3.30996108055115,-3.19799828529358,-3.08701181411743,-2.88515639305115,-2.65273451805115,-2.67426729202271,-2.767578125,-2.97978496551514,-3.17822265625,-3.26777362823486,-3.36225628852844,-3.48042011260986,-3.69511699676514,-3.78906273841858,-3.70415043830872,-3.60781264305115,-3.04873037338257,-3.01508784294128,-2.83686518669128,-2.59931635856628,-2.14707064628601,-2.016845703125,-1.83027362823486,-1.728759765625,-1.65537142753601,-1.61015641689301,-1.52255856990814,-1.42265641689301,-1.29174816608429,-1.23242199420929,-1.15439486503601,-0.759326040744781,-0.671386778354645,-0.518115401268005,-0.370361059904099,-0.232861503958702,-0.0843749046325684,-0.156298875808716,-0.205566376447678,-0.168749809265137,-0.10825178027153,0,0.0105470065027475,0.115331828594208,0.0767087340354919,0.0360837355256081,0,-0.0194335822016001,-0.0737304985523224,-0.173827946186066,-0.270019561052322,-0.461377143859863,-0.567675709724426,-0.659912168979645,-0.485058635473251,-0.29370105266571,0,0.128320202231407,0.270995855331421,0.355761528015137,0.298046916723251,0.208203122019768,0.124414347112179,0.0458985045552254,0.27978503704071,0.330175518989563,0.381933659315109,0.431640595197678,0.515527367591858,0.558789134025574,0.704492270946503,0.826758086681366,0.948535025119781,1.05556643009186,1.27128899097443,1.38212895393372,1.65673840045929,1.71611344814301,1.74335932731628,1.74658226966858,1.70039033889771,1.64736306667328,1.61464834213257,1.59140634536743,1.55898439884186,1.41347658634186,1.31679689884186,1.27597641944885,1.23242199420929,1.22783195972443,1.27382779121399,1.27441394329071,1.1884765625,1.10117161273956,0.95507824420929,0.752246022224426,0.898046612739563,0.927441596984863,0.890917897224426,0.799218833446503,0.697558343410492,0.593457102775574,0.507226347923279,0.424511879682541,0.528320372104645,0.600293040275574,0.645507991313934,0.686523199081421,0.889355421066284,1.01494145393372,1.25712895393372,1.37343776226044,1.41494166851044,1.41562509536743,1.39755856990814,1.36552751064301,1.04443395137787,0.978613078594208,0.960156381130219,0.772363126277924,0.684375166893005,0.532324194908142,0.414745807647705,0.299706786870956,0.205078199505806,0,-0.203906506299973,-0.450781434774399,-0.785254120826721,-0.871386885643005,-1.00058603286743,-1.13286137580872,-1.28505837917328,-1.41645479202271,-1.33447253704071,-1.51674818992615,-1.60083031654358,-1.68789064884186,-1.86601579189301,-2.03105449676514,-2.00625014305115,-1.96206068992615,-1.99790036678314,-2.03583979606628,-2.35014629364014,-2.39467787742615,-2.43344736099243,-2.54775357246399,-2.65883803367615,-2.77695345878601,-2.90087914466858,-2.99941396713257,-3.40458965301514,-3.48544883728027,-3.52587914466858,-3.58437538146973,-3.67978525161743,-3.79335951805115,-3.90019559860229,-4.10341739654541,-4.17255878448486,-4.194580078125,-4.21728515625,-4.29697275161743,-4.37949228286743,-4.50668954849243,-4.72797870635986,-4.8173828125,-5.00952100753784,-5.04863309860229,-5.11850595474243,-5.22524356842041,-5.32285165786743,-5.43398475646973,-5.55122089385986,-5.62211894989014,-5.65517616271973,-5.65625047683716,-5.57065439224243,-5.34228515625,-5.14179658889771,-5.04345703125,-5.00444316864014,-4.95639657974243,-4.8935546875,-4.86127901077271,-4.58291006088257,-4.55996084213257,-4.54609346389771,-4.52309560775757,-4.29648447036743,-4.18818330764771,-4.15839815139771,-3.84233403205872,-3.60790991783142,-3.37509775161743,-3.25576162338257,-3.13598656654358,-3.04204106330872,-2.88124990463257,-2.79082012176514,-2.68720698356628,-2.59028315544128,-2.43305659294128,-2.53935527801514,-2.66767597198486,-2.74213862419128,-2.97851538658142,-3.08037114143372,-3.2587890625,-3.29311513900757,-3.56235361099243,-3.7626953125,-3.89077115058899,-3.94365262985229,-3.99833941459656,-4.11528348922729,-4.23457002639771,-4.17368173599243,-4.09101581573486,-4.27617168426514,-4.32763671875,-4.38627958297729,-4.531494140625,-4.60078144073486,-4.71762657165527,-4.90229511260986,-5.124755859375,-5.16835927963257,-5.16723680496216,-5.20058584213257,-5.26230430603027,-5.18335008621216,-5.08808565139771,-4.87851572036743,-4.56113290786743,-4.38315391540527,-4.21772480010986,-4.14936542510986,-4.09975624084473,-4.050537109375,-3.98032212257385,-4.04843711853027,-4.07890605926514,-4.07070302963257,-4.03925752639771,-4.06743144989014,-4.11752939224243,-4.11474609375,-4.10146522521973,-4.22915029525757,-4.35644483566284,-4.47182607650757,-4.58369159698486,-4.68305683135986,-4.68144512176514,-4.63832998275757,-4.52568387985229,-4.40507841110229,-4.36220741271973,-4.32841777801514,-4.2685546875,-4.11103534698486,-3.80927729606628,-3.76420903205872,-3.64589810371399,-3.52958965301514,-3.427734375,-3.326171875,-3.09755897521973,-3.16557621955872,-3.06474590301514,-2.91855478286743,-2.86416053771973,-2.74951171875,-2.79375004768372,-2.84541058540344,-2.9130859375,-2.96997094154358,-3.06459927558899,-3.05947256088257,-2.99570298194885,-2.92509794235229,-2.98432636260986,-3.03178739547729,-3.04536128044128,-3.02675771713257,-2.89985346794128,-2.86240243911743,-2.84648418426514,-2.86757802963257,-2.99350571632385,-3.05473661422729,-3.10966777801514,-3.16596674919128,-3.321533203125,-3.41025400161743,-3.56938481330872,-3.59204077720642,-3.464599609375,-3.26791977882385,-3.03623056411743,-3.0810546875,-3.43408203125,-3.55043959617615,-3.65830063819885,-3.71923828125,-3.78325152397156,-3.84160184860229,-3.89858412742615,-3.95790982246399,-4.07578086853027,-4.13295888900757,-4.17402315139771,-4.20839834213257,-4.25341796875,-4.30366230010986,-4.40991258621216,-4.51748085021973,-4.64755868911743,-4.81806612014771,-4.85170936584473,-4.88950204849243,-4.91123056411743,-5.03232479095459,-5.135498046875,-5.17011737823486,-5.17270469665527,-5.11669921875,-5.05585908889771,-4.96518611907959,-4.78481483459473,-4.72114229202271,-4.6767578125,-4.68437528610229,-4.72416973114014,-4.891845703125,-4.8896484375,-4.87167978286743,-4.82607460021973,-4.80683565139771,-4.58408212661743,-4.67094755172729,-4.84409189224243,-4.84101581573486,-4.80029344558716,-4.81914091110229,-4.85624980926514,-4.92709970474243,-4.97036170959473,-5.09282255172729,-5.114990234375,-5.13466787338257,-5.19584941864014,-5.21459913253784,-5.22822237014771,-5.24560594558716,-5.24731397628784,-5.22294902801514,-5.17641592025757,-4.99697303771973,-5.08432626724243,-5.28232431411743,-5.38344717025757,-5.41044902801514,-5.41889667510986,-5.41831064224243,-5.37290048599243,-5.38583993911743,-5.55644559860229,-5.58876943588257,-5.61845684051514,-5.64653348922729,-5.73066425323486,-5.76821279525757,-5.76787090301514,-5.75209951400757,-5.68134737014771,-5.650634765625,-5.60502958297729,-5.50449228286743,-5.50693368911743,-5.57387638092041,-5.60239267349243,-5.62285137176514,-5.60957050323486,-5.55527353286743,-5.53496074676514,-5.48789072036743,-5.43339824676514,-5.39194297790527,-5.32944297790527,-5.31269550323486,-5.24257850646973,-5.18837881088257,-5.21757793426514,-5.56420946121216,-5.65244150161743,-5.77280235290527,-5.86484336853027,-5.936767578125,-5.96889686584473,-6.05771493911743,-6.13369131088257,-6.13276386260986,-6.03471660614014,-5.87763643264771,-5.73061513900757,-5.86142587661743,-5.85039091110229,-5.73627948760986,-5.59130859375,-5.56191396713257,-5.63124990463257,-5.65634775161743,-5.794921875,-5.81806612014771,-5.80195331573486,-5.75673818588257,-5.68862295150757,-5.58178663253784,-5.67876005172729,-5.71494150161743,-5.74238300323486,-5.74492168426514,-5.69472646713257,-5.66547822952271,-5.60834980010986,-5.34902334213257,-5.31918954849243,-5.289794921875,-5.1572265625,-5.17690467834473,-5.39375019073486,-5.41318368911743,-5.35136747360229,-5.34687471389771,-5.35595703125,-5.33828115463257,-5.26953125,-5.05996131896973,-5.00830078125,-5.03183555603027,-5.08061504364014,-5.09013700485229,-5.07871103286743,-5.07602548599243,-5.06650352478027,-5.01674795150757,-4.97563505172729,-4.92465829849243,-4.80961894989014,-4.76577138900757,-4.71542930603027,-4.67822265625,-4.53496074676514,-4.49189424514771,-4.43325185775757,-4.18862295150757,-3.85952186584473,-3.66181635856628,-3.45356440544128,-3.25913071632385,-3.05307602882385,-3.04619145393372,-3.05698251724243,-3.10966777801514],[-2.92939448356628,-2.93896460533142,-2.97539043426514,-3.03544902801514,-2.94121074676514,-2.89643573760986,-2.9130859375,-2.92939448356628],[-3.16494154930115,-3.22211933135986,-3.27880883216858,-3.36718773841858,-3.40083003044128,-3.39472675323486,-3.35742211341858,-3.27192354202271,-3.22763681411743,-3.22211933135986,-3.21162080764771,-3.15854501724243,-3.16494154930115],[-3.05742192268372,-3.07070326805115,-2.99467778205872,-2.88457036018372,-2.81791973114014,-2.762451171875,-2.79301738739014,-2.82622075080872,-2.86376953125,-2.99482417106628,-3.16660141944885,-3.20078110694885,-3.22333955764771,-3.23261690139771,-3.23281240463257,-3.24213838577271,-3.30434560775757,-3.33164072036743,-3.34707045555115,-3.35371112823486,-3.34682631492615,-3.31035137176514,-3.24858427047729,-3.15649437904358,-3.05112266540527,-3.01923847198486,-3.02001929283142,-3.05742192268372],[-2.54887723922729,-2.66206049919128,-2.60361313819885,-2.53564476966858,-2.40698266029358,-2.42983388900757,-2.54887723922729],[-2.72939443588257,-2.81523442268372,-2.85185551643372,-2.86142587661743,-2.96376967430115,-3.01347661018372,-3.05205106735229,-3.042236328125,-2.97553706169128,-2.86162114143372,-2.81503891944885,-2.73066425323486,-2.71992182731628,-2.72939443588257],[-1.30810570716858,-1.28740239143372,-1.23574221134186,-1.15776383876801,-1.11796903610229,-1.05244159698486,-1.06567394733429,-1.13369154930115,-1.15278315544128,-1.16572284698486,-1.17924797534943,-1.19931638240814,-1.24531233310699,-1.28378927707672,-1.29946315288544,-1.35585939884186,-1.29951179027557,-1.27617180347443,-1.29091799259186,-1.32280254364014,-1.40903317928314,-1.48149406909943,-1.49687516689301,-1.49912118911743,-1.5166015625,-1.61303722858429,-1.64135730266571,-1.66005873680115,-1.66376972198486,-1.57666003704071,-1.49443376064301,-1.37460935115814,-1.44956028461456,-1.548828125,-1.57177734375,-1.55263662338257,-1.49814426898956,-1.41420876979828,-1.36396503448486,-1.30170905590057,-1.30810570716858],[-1.04252934455872,-1.06787121295929,-1.16552734375,-1.09331059455872,-1.00561535358429,-0.99165016412735,-1.00034189224243,-1.04501950740814,-1.04902327060699,-1.03510725498199,-1.03422832489014,-1.04252934455872],[-0.774267673492432,-0.774316370487213,-0.826171875,-0.825488388538361,-0.909131050109863,-0.922265529632568,-0.938086032867432,-0.927539050579071,-0.915820360183716,-0.891405999660492,-0.864941656589508,-0.823437750339508,-0.801806926727295,-0.774267673492432],[46.4298858642578,46.4054679870605,46.3482437133789,46.3025398254395,46.2518539428711,46.2018547058105,46.1842765808105,46.18212890625,46.1905288696289,46.2035140991211,46.2546882629395,46.3054695129395,46.3849639892578,46.5087890625,46.6189422607422,46.6725578308105,46.6623992919922,46.6263694763184,46.5343742370605,46.4579124450684,46.4309577941895,46.3807601928711,46.2799835205078,46.1707038879395,46.0865249633789,46.03125,45.9219703674316,45.7927742004395,45.7254867553711,45.6957054138184,45.7156257629395,45.4222640991211,45.2809562683105,45.2171859741211,45.0013656616211,44.9758796691895,44.8113288879395,44.8109397888184,44.8485336303711,44.8414077758789,44.5648422241211,44.4730491638184,44.2273445129395,44.146484375,44.0772476196289,43.9091796875,43.7931632995605,43.64501953125,43.4919929504395,43.439453125,43.4416007995605,43.4333992004395,43.40234375,43.3589820861816,43.279296875,43.2054672241211,43.15283203125,43.1410140991211,43.1712913513184,43.1490249633789,43.05712890625,42.90673828125,42.8216781616211,42.7878913879395,42.7541007995605,42.6824226379395,42.6068344116211,42.5904312133789,42.5673828125,42.5079116821289,42.4664077758789,42.3643531799316,42.2799797058105,42.2111320495605,42.0777359008789,41.92578125,41.8235359191895,41.7793922424316,41.7017593383789,41.5765647888184,41.5100593566895,41.7017593383789,41.7587890625,41.7607421875,41.7629852294922,41.6632804870605,41.5777359008789,41.48876953125,41.41943359375,41.1287117004395,41.0616226196289,40.8366203308105,40.5240249633789,40.4621086120605,40.1906242370605,39.9783210754395,40.0237312316895,40.0845718383789,40.1501960754395,40.34228515625,40.5189476013184,40.6480484008789,40.8016624450684,40.9419898986816,41.0831069946289,41.3582038879395,41.4607391357422,41.58056640625,42.0500030517578,42.0877914428711,42.1222648620605,42.2796859741211,42.4190444946289,42.5660171508789,42.6602554321289,42.7606468200684,42.8900413513184,42.9916000366211,43.0001983642578,43.0891609191895,43.3479499816895,43.5578117370605,43.623046875,43.7826194763184,43.7987327575684,43.79541015625,43.7499008178711,43.7383804321289,43.7598648071289,43.8259773254395,43.9574203491211,44.0046882629395,44.1027336120605,44.19970703125,44.3294906616211,44.505859375,44.5764656066895,44.6443367004395,44.6917953491211,44.7710952758789,44.8504867553711,44.8709945678711,44.943359375,45.0715827941895,45.1602554321289,45.2082023620605,45.34375,45.5628890991211,45.6555671691895,45.7052726745605,45.7275390625,45.6883773803711,45.63427734375,45.6385765075684,45.7265625,45.8459968566895,45.9103546142578,45.9540061950684,46.0484390258789,46.1597633361816,46.2126960754395,46.2677726745605,46.4115257263184,46.4298858642578],[-2.51230478286743,-2.54736351966858,-2.63901400566101,-2.64614248275757,-2.54218745231628,-2.52089858055115,-2.51230478286743],[-0.0686036720871925,-0.004736069124192,0,0.00942401029169559,0,-0.0138672972097993,-0.0605954453349113,-0.0901852771639824,-0.0863281190395355,-0.0577146112918854,0,0.0394533015787601,0.0892576575279236,0.148241952061653,0.21601539850235,0.331835955381393,0.380859375,0.378613352775574,0.362695455551147,0.351855665445328,0.343066513538361,0.334570109844208,0.323925703763962,0.311718851327896,0.289648503065109,0.26953136920929,0.264550656080246,0.272753715515137,0.342578321695328,0.327343970537186,0.275488436222076,0.251562267541885,0.261913895606995,0.233398362994194,0.241503983736038,0.259960681200027,0.28935569524765,0.370995879173279,0.405273675918579,0.447558522224426,0.525683641433716,0.529003918170929,0.497168093919754,0.466113179922104,0.460351556539536,0.493261635303497,0.488769680261612,0.453124821186066,0.372558355331421,0.378613352775574,0.415331959724426,0.483300775289536,0.616210997104645,0.68632835149765,0.688086211681366,0.647070467472076,0.599218726158142,0.583593606948853,0.605175793170929,0.500000238418579,0.498925894498825,0.509570300579071,0.537304937839508,0.591015696525574,0.634765625,0.619531214237213,0.59619128704071,0.592480182647705,0.579492330551147,0.538085877895355,0.523046851158142,0.533398509025574,0.525586187839508,0.548046767711639,0.595703065395355,0.672753870487213,0.702246248722076,0.715429484844208,0.707226455211639,0.736914217472076,0.822460830211639,0.912207305431366,0.984960854053497,1.00214838981628,1.04990220069885,1.08447277545929,1.13964855670929,1.18505835533142,1.18720698356628,1.10556626319885,1.05029296875,1.00800812244415,0.949706733226776,0.74882835149765,0.671874940395355,0.259667903184891,0,-0.126513659954071,-0.348730206489563,-0.485449403524399,-0.66943359375,-0.797705054283142,-1.06430697441101,-1.50165987014771,-1.63847649097443,-1.77685558795929,-2.00185537338257,-2.09018564224243,-2.26640605926514,-2.39892554283142,-2.72304654121399,-2.96499013900757,-3.08188486099243,-3.114013671875,-3.08671903610229,-3.01914048194885,-2.94833993911743,-2.89472651481628,-2.81567406654358,-2.79521489143372,-2.78867220878601,-2.78959965705872,-2.76191377639771,-2.75498056411743,-2.79365205764771,-2.82119154930115,-2.96225595474243,-2.97280287742615,-2.998291015625,-3.02529263496399,-3.05615234375,-3.10556650161743,-3.20058584213257,-3.22402334213257,-3.24028325080872,-3.24389624595642,-3.22412133216858,-3.22714877128601,-3.23579120635986,-3.16889667510986,-3.03769493103027,-3.01015639305115,-2.98579072952271,-2.98232436180115,-2.95908164978027,-2.89633774757385,-2.85688495635986,-2.83012676239014,-2.79814481735229,-2.78974580764771,-2.66884779930115,-2.61337924003601,-2.60097646713257,-2.61997103691101,-2.61171865463257,-2.582763671875,-2.53828144073486,-2.50585961341858,-2.55688500404358,-2.59799790382385,-2.60039067268372,-2.62490248680115,-2.64921879768372,-2.68989253044128,-2.74692416191101,-2.74667954444885,-2.689208984375,-2.67421865463257,-2.70180654525757,-2.70576167106628,-2.68613266944885,-2.69584941864014,-2.70620131492615,-2.76596689224243,-2.78051781654358,-2.74980497360229,-2.75073266029358,-2.78320336341858,-2.78847670555115,-2.76650381088257,-2.77709937095642,-2.8203125,-2.82343745231628,-2.78662085533142,-2.79116225242615,-2.83720684051514,-2.87841796875,-2.91489243507385,-2.90732431411743,-2.83857464790344,-2.82993125915527,-2.75209975242615,-2.75166010856628,-2.50917983055115,-2.23193359375,-1.90063464641571,-1.59965801239014,-1.58647477626801,-1.53676736354828,-1.23261725902557,-1.04248070716858,-0.961816251277924,-0.902929902076721,-0.771582186222076,-0.701416075229645,-0.648534953594208,-0.627148687839508,-0.597656309604645,-0.545214593410492,-0.491699188947678,-0.453515589237213,-0.430322259664536,-0.395605593919754,-0.345751971006393,-0.31254905462265,-0.299462705850601,-0.0686036720871925],[-11.3894033432007,-11.4180669784546,-11.4475584030151,-11.4745607376099,-11.502197265625,-11.492431640625,-11.4146490097046,-11.30517578125,-11.2606935501099,-11.2096681594849,-11.1292476654053,-11.0658197402954,-11.0045413970947,-10.9332036972046,-10.8761720657349,-10.8064937591553,-10.7430171966553,-10.7349119186401,-10.709228515625,-10.6773433685303,-10.6437015533447,-10.6189937591553,-10.5895023345947,-10.4658203125,-10.3727540969849,-10.3398933410645,-10.2748537063599,-10.1670904159546,-10.0106449127197,-9.82070255279541,-9.75400352478027,-9.71474647521973,-9.65830039978027,-9.58774375915527,-9.48681640625,-9.40498065948486,-9.35810565948486,-9.34018516540527,-9.33154296875,-9.3408203125,-9.39365196228027,-9.39536190032959,-9.36518573760986,-9.30000019073486,-9.21552658081055,-9.12045955657959,-9.04306602478027,-8.99892520904541,-8.95083045959473,-8.91386699676514,-8.81831073760986,-8.820068359375,-8.822021484375,-8.77973651885986,-8.73310565948486,-8.71142578125,-8.66494178771973,-8.62114238739014,-8.56874942779541,-8.470703125,-8.407470703125,-8.39853572845459,-8.40068340301514,-8.42529296875,-8.46352481842041,-8.52031230926514,-8.56728458404541,-8.66391563415527,-8.66669940948486,-8.64619159698486,-8.606201171875,-8.56352519989014,-8.47470664978027,-8.40449237823486,-8.33740234375,-8.312744140625,-8.30634784698486,-8.32168006896973,-8.32412147521973,-8.30156230926514,-8.26665019989014,-8.23149490356445,-8.00727558135986,-7.98569345474243,-7.97446250915527,-7.99062538146973,-8.01352500915527,-8.07783222198486,-8.13662052154541,-8.15517616271973,-8.14585018157959,-8.14604473114014,-8.13696193695068,-8.08867168426514,-8.031005859375,-7.96269512176514,-7.89619112014771,-7.90000009536743,-7.91806650161743,-7.83940410614014,-7.7998046875,-7.77797889709473,-7.902099609375,-7.93818330764771,-7.95498037338257,-7.95097637176514,-7.78403377532959,-7.71958065032959,-7.69096612930298,-7.68120098114014,-7.69609403610229,-7.73896455764771,-7.78740262985229,-7.82358360290527,-7.86875009536743,-7.953125,-8.04912090301514,-8.16777324676514,-8.2099609375,-8.23696327209473,-8.244140625,-8.256103515625,-8.21713829040527,-8.140625,-8.09052753448486,-8.048583984375,-8.01674842834473,-8.00986385345459,-8.03173828125,-8.07382869720459,-8.12685489654541,-8.11782264709473,-8.11542987823486,-8.20595741271973,-8.23188495635986,-8.35175800323486,-8.42998123168945,-8.48642539978027,-8.52226543426514,-8.56440448760986,-8.578857421875,-8.60732460021973,-8.65976524353027,-8.70830059051514,-8.72944355010986,-8.73261737823486,-8.740234375,-8.76914024353027,-8.82792949676514,-8.85551738739014,-8.8896484375,-8.93842792510986,-8.96098613739014,-8.9765625,-9.05234336853027,-9.11757850646973,-9.13481426239014,-9.1728515625,-9.21518516540527,-9.26328086853027,-9.35532283782959,-9.39165019989014,-9.43510723114014,-9.46381855010986,-9.45976543426514,-9.41147518157959,-9.38398456573486,-9.36894512176514,-9.369140625,-9.39492225646973,-9.43632793426514,-9.44638633728027,-9.44155216217041,-9.45112323760986,-9.46455001831055,-9.47114276885986,-9.484130859375,-9.50849628448486,-9.522216796875,-9.51826190948486,-9.55390644073486,-9.61015605926514,-9.64321327209473,-9.66357421875,-9.68388652801514,-9.701171875,-9.71689414978027,-9.735595703125,-9.76826190948486,-9.781982421875,-9.80473613739014,-10.0643558502197,-10.07568359375,-10.09765625,-10.1474123001099,-10.2330570220947,-10.283203125,-10.3600578308105,-10.3944339752197,-10.4964361190796,-10.5577154159546,-10.60400390625,-10.6526365280151,-10.6869630813599,-10.7121095657349,-10.7021484375,-10.6773433685303,-10.6284666061401,-10.5031251907349,-10.5005369186401,-10.5517578125,-10.6056156158447,-10.6057615280151,-10.615966796875,-10.7268552780151,-10.7470216751099,-10.7499513626099,-10.7212400436401,-10.6876459121704,-10.6827144622803,-10.6905279159546,-10.7585935592651,-10.8647947311401,-10.9630861282349,-11.0474615097046,-11.1156740188599,-11.1808595657349,-11.2056636810303,-11.2736330032349,-11.471923828125,-11.7100591659546,-11.9110841751099,-11.9227533340454,-12.142333984375,-12.2777347564697,-12.427978515625,-12.5014638900757,-12.5243654251099,-12.557861328125,-12.5898447036743,-12.6036128997803,-12.6221685409546,-12.6516599655151,-12.6844234466553,-12.755859375,-12.8311033248901,-12.9587888717651,-12.9986324310303,-13.0280275344849,-13.0772943496704,-13.1298828125,-13.1783695220947,-13.2342281341553,-13.2926759719849,-13.3026371002197,-13.2694826126099,-13.2958984375,-13.3960933685303,-13.4055662155151,-13.436279296875,-13.5682611465454,-13.6913576126099,-13.6571292877197,-13.65869140625,-13.7004890441895,-13.6897954940796,-13.712646484375,-13.7537107467651,-13.8201179504395,-13.9546384811401,-14.0218753814697,-14.0299320220947,-14.0450191497803,-14.0862789154053,-14.17041015625,-14.4269046783447,-14.6095695495605,-14.6136236190796,-14.58740234375,-14.5935068130493,-14.6773443222046,-14.693359375,-14.7573728561401,-14.7759275436401,-14.8374519348145,-14.88671875,-14.9248046875,-14.9750003814697,-15.0124025344849,-15.0512199401855,-15.0430173873901,-14.9990234375,-14.9444332122803,-14.779296875,-14.7202634811401,-14.6829586029053,-14.6047849655151,-14.4524412155151,-14.3278312683105,-14.2655754089355,-14.1223154067993,-13.9532222747803,-13.7327642440796,-13.7285642623901,-13.7306642532349,-13.7379884719849,-13.8163089752197,-13.8619146347046,-13.9011716842651,-13.9488773345947,-13.9473142623901,-13.8874998092651,-13.8494625091553,-13.759765625,-13.7300777435303,-13.7079105377197,-13.682373046875,-13.6735353469849,-13.7326173782349,-13.729248046875,-13.40576171875,-13.37255859375,-13.2280759811401,-13.1384763717651,-13.0829105377197,-13.0597658157349,-13.0644044876099,-13.079833984375,-13.061279296875,-13.0119142532349,-12.9856443405151,-12.9605464935303,-12.9307126998901,-12.88818359375,-12.7973136901855,-12.7130374908447,-12.6208009719849,-12.5342283248901,-12.4573736190796,-12.3990726470947,-12.2912101745605,-12.1519536972046,-12.0423822402954,-11.8885746002197,-11.80810546875,-11.5736808776855,-11.4567384719849,-11.3894033432007],[-16.7633304595947,-16.7693367004395,-16.8248043060303,-16.7503910064697,-16.6693363189697,-16.6147956848145,-16.5983390808105,-16.5564460754395,-16.4133777618408,-16.2716789245605,-16.18505859375,-16.1878890991211,-16.1583995819092,-15.9864263534546,-15.8044929504395,-15.6176748275757,-15.4712886810303,-15.427490234375,-15.4381351470947,-15.5695304870605,-15.8499031066895,-16.1354484558105,-16.351806640625,-16.4405269622803,-16.5300788879395,-16.5623054504395,-16.3087406158447,-16.0016117095947,-15.6671867370605,-15.5096673965454,-15.4268550872803,-15.26953125,-15.1083498001099,-15.0244626998901,-14.9357919692993,-14.7660150527954,-14.66015625,-14.5708494186401,-14.5069818496704,-14.4054679870605,-14.3255367279053,-14.2780275344849,-14.1990232467651,-14.1469717025757,-13.9773921966553,-13.8528318405151,-13.8267087936401,-13.8475103378296,-14.014892578125,-14.2467775344849,-14.4385738372803,-14.6719245910645,-14.8082513809204,-14.8650388717651,-14.9502935409546,-15.0246095657349,-15.0963859558105,-15.151123046875,-15.1916027069092,-15.2121086120605,-15.2445316314697,-15.2862300872803,-15.4818363189697,-15.6573238372803,-15.7515621185303,-15.8144044876099,-15.8342771530151,-16.0330562591553,-16.2283191680908,-16.4308605194092,-16.6487789154053,-16.7045402526855,-16.7633304595947],[-15.8958988189697,-15.9051761627197,-15.9639654159546,-15.9506349563599,-15.9634771347046,-15.9464855194092,-15.9376945495605,-15.9091310501099,-15.9052734375,-15.8958988189697],[-16.114501953125,-16.1945304870605,-16.2310047149658,-16.2364253997803,-16.1946296691895,-16.1758785247803,-16.14404296875,-16.1047859191895,-16.0874519348145,-16.0673332214355,-16.0527839660645,-16.0722179412842,-16.114501953125],[-15.7251462936401,-15.7251462936401,-15.7674798965454,-15.7797861099243,-15.7546863555908,-15.7174806594849,-15.6719236373901,-15.6583499908447,-15.6671867370605,-15.6871089935303,-15.7251462936401],[-15.9018058776855,-15.94873046875,-15.9972162246704,-16.023193359375,-16.0193367004395,-15.9645509719849,-15.9153327941895,-15.9018058776855],[-15.5534181594849,-15.5627927780151,-15.6196279525757,-15.5365715026855,-15.4824714660645,-15.4844245910645,-15.5262203216553,-15.5534181594849],[-15.9864263534546,-16.038330078125,-16.1024398803711,-16.1473636627197,-16.1524410247803,-16.0218753814697,-15.9864263534546],[-15.0430173873901,-15.09375,-15.0545892715454,-15.0967769622803,-15.1810550689697,-15.2221202850342,-15.2166986465454,-15.2633790969849,-15.3174810409546,-15.3931150436401,-15.4005861282349,-15.39453125,-15.3484373092651,-15.3546867370605,-15.399169921875,-15.448974609375,-15.4794912338257,-15.4291019439697,-15.2525873184204,-15.1637697219849,-15.0726566314697,-15.1224126815796,-15.2303705215454,-15.3166990280151,-15.3596677780151,-15.4129877090454,-15.5019044876099,-15.5002450942993,-15.4671869277954,-15.4157228469849,-15.2108402252197,-15.1331052780151,-15.1017084121704,-15.0719738006592,-15.0782709121704,-15.1115236282349,-15.1880865097046,-15.4347658157349,-15.5134773254395,-15.6506834030151,-15.8193836212158,-15.9417486190796,-15.9027347564697,-15.9202146530151,-15.9587879180908,-16.138427734375,-16.2743148803711,-16.3280754089355,-16.31884765625,-16.2547378540039,-16.2445812225342,-16.3123054504395,-16.4368152618408,-16.7118167877197,-16.6569328308105,-16.5213375091553,-16.4163093566895,-16.34228515625,-16.2415027618408,-16.1441879272461,-15.8395500183105,-15.5748052597046,-15.3779296875,-15.1960935592651,-14.9605951309204,-14.7081546783447,-14.3492183685303,-14.0648431777954,-13.729248046875,-13.7326173782349,-13.6735353469849,-13.682373046875,-13.7079105377197,-13.7300777435303,-13.759765625,-13.8494625091553,-13.8874998092651,-13.9473142623901,-13.9488773345947,-13.9011716842651,-13.8619146347046,-13.8163089752197,-13.7379884719849,-13.7306642532349,-13.7285642623901,-13.7327642440796,-13.9532222747803,-14.1223154067993,-14.2655754089355,-14.3278312683105,-14.4524412155151,-14.6047849655151,-14.6829586029053,-14.7202634811401,-14.779296875,-14.9444332122803,-14.9990234375,-15.0430173873901],[9.5908203125,9.59941387176514,9.50986385345459,9.4453125,9.38593769073486,9.43408203125,9.49423885345459,9.58427810668945,9.63212871551514,9.64765644073486,9.71884822845459,9.80703163146973,9.77968788146973,9.80078125,9.82617092132568,9.83037090301514,9.8369140625,9.8701171875,9.97988224029541,10.3070316314697,10.5022459030151,10.7909173965454,11.0965824127197,11.3287115097046,11.330078125,11.3311519622803,11.3323249816895,11.3335943222046,11.3346681594849,11.3353519439697,11.1306638717651,10.85888671875,10.5872068405151,10.3154296875,10.1789064407349,10.0285158157349,9.97978496551514,9.94668006896973,9.90673828125,9.8603515625,9.80390548706055,9.78867149353027,9.76054668426514,9.70458984375,9.67646503448486,9.63613224029541,9.5908203125],[8.73574161529541,8.76044940948486,8.91005897521973,8.95068359375,8.94609355926514,8.79218769073486,8.76347637176514,8.70400333404541,8.65234375,8.47490215301514,8.44492149353027,8.43427658081055,8.45175838470459,8.46464824676514,8.5498046875,8.57724666595459,8.62275409698486,8.6376953125,8.67587852478027,8.73574161529541],[23.8522472381592,23.9206047058105,24.0132827758789,24.0343761444092,24.0933589935303,24.166015625,24.19775390625,24.1240234375,24.1089839935303,24.1231441497803,24.1785163879395,24.25537109375,24.2577133178711,24.27490234375,24.3128890991211,24.35400390625,24.4449234008789,24.5345706939697,24.626953125,24.7212886810303,25.0031242370605,25.1042976379395,25.2967777252197,25.4756832122803,25.5696296691895,25.7301769256592,25.755859375,25.7351551055908,25.7450199127197,25.7913074493408,25.8371105194092,25.8933601379395,26.0280265808105,26.1678714752197,26.2855472564697,26.3202133178711,26.2986316680908,26.2808609008789,26.2555656433105,26.2443351745605,26.1656246185303,26.0466804504395,25.8296871185303,25.6109371185303,25.2057609558105,24.7998046875,24.7452144622803,24.7439441680908,24.7351570129395,24.7088871002197,24.5833969116211,24.4636707305908,23.9943351745605,23.8835945129395,23.7039070129395,23.6380863189697,23.5927734375,23.5616226196289,23.5475578308105,23.56982421875,23.6086921691895,23.6265640258789,23.6726551055908,23.7154293060303,23.7150382995605,23.7369136810303,23.7707996368408,23.7933597564697,23.8522472381592],[27.1760749816895,27.1378898620605,27.09912109375,27.1158199310303,27.0707035064697,27.1560554504395,27.1580066680908,27.22314453125,27.2070331573486,27.1572265625,27.2088871002197,27.2335929870605,27.1760749816895],[23.0538101196289,23.0421886444092,22.939453125,22.9108390808105,22.9056644439697,22.9326171875,22.9504890441895,22.9978523254395,23.0970706939697,23.0538101196289],[27.8427734375,27.7706050872803,27.7457046508789,27.7155284881592,27.75732421875,27.7186527252197,27.71630859375,27.7744140625,27.8152351379395,27.9144535064697,28.1714839935303,28.2318363189697,28.2300777435303,28.14404296875,28.0676746368408,28.0877933502197,27.9655265808105,27.8427734375],[25.482421875,25.4359378814697,25.3705062866211,25.3971672058105,25.4128894805908,25.4146480560303,25.3968753814697,25.4089851379395,25.4673824310303,25.482421875],[26.4606437683105,26.3816394805908,26.3314456939697,26.27001953125,26.2698249816895,26.3370113372803,26.3841800689697,26.3700199127197,26.4212894439697,26.4606437683105],[27.8601570129395,27.8382816314697,27.7880859375,27.7857418060303,27.8368167877197,27.8624992370605,27.8698234558105,27.8690433502197,27.8601570129395],[24.5357418060303,24.5375003814697,24.5306644439697,24.3259773254395,24.3449211120605,24.357421875,24.4251956939697,24.4501953125,24.4603519439697,24.5357418060303],[25.3817367553711,25.3643569946289,25.2886714935303,25.2599601745605,25.2958984375,25.4069328308105,25.3817367553711],[26.9496097564697,26.9183597564697,26.95556640625,27.0611324310303,27.2149410247803,27.265625,27.3521480560303,27.1931629180908,27.1509761810303,27.0335941314697,26.9496097564697],[25.859375,25.7710952758789,25.7431640625,25.7967777252197,25.8343734741211,25.8524417877197,25.9419918060303,26.0006847381592,26.064453125,25.9846668243408,25.859375],[24.7208976745605,24.70263671875,24.6764640808105,24.6709976196289,24.6814460754395,24.7161140441895,24.76318359375,24.7208976745605],[27.0197257995605,26.919921875,26.9376964569092,26.888671875,26.9666004180908,27.0160160064697,27.0401363372803,27.0345706939697,27.0197257995605],[25.2789058685303,25.1994152069092,25.13330078125,25.10546875,25.146484375,25.2350597381592,25.2752933502197,25.271484375,25.2789058685303],[25.5458984375,25.4567375183105,25.3958988189697,25.3619136810303,25.5252933502197,25.5643539428711,25.587890625,25.5842761993408,25.5458984375],[24.5235347747803,24.4865226745605,24.4248046875,24.4412097930908,24.4837875366211,24.5291004180908,24.5359363555908,24.5235347747803],[24.4357433319092,24.37890625,24.3977527618408,24.3697280883789,24.3948249816895,24.4312496185303,24.4485359191895,24.4814453125,24.4357433319092],[24.94287109375,24.9378890991211,24.9115238189697,24.8961906433105,24.8953113555908,24.9065418243408,24.94287109375],[25.4027328491211,25.30712890625,25.3126945495605,25.34814453125,25.4629878997803,25.4574222564697,25.4027328491211],[26.029296875,25.982421875,25.9967765808105,26.0863285064697,26.2115230560303,26.3255863189697,26.3513660430908,26.296875,26.2048835754395,26.029296875],[25.255859375,25.21875,25.1563472747803,25.0519542694092,25.0163097381592,24.9964847564697,25.0393562316895,25.091796875,25.2253913879395,25.255859375],[24.35595703125,24.2889652252197,24.2774410247803,24.3204097747803,24.3791027069092,24.4007816314697,24.35595703125],[26.8244132995605,26.9473648071289,26.9815425872803,27.0396480560303,27.0550785064697,26.9781246185303,26.8449211120605,26.7882823944092,26.7205085754395,26.6128902435303,26.5810546875,26.638671875,26.7433605194092,26.8244132995605],[20.8884773254395,20.9939460754395,20.9090824127197,20.8185558319092,20.7038097381592,20.6350593566895,20.6195316314697,20.6915035247803,20.7586917877197,20.83984375,20.8884773254395],[24.99169921875,24.9622077941895,24.8840808868408,24.7985343933105,24.7665023803711,24.7143573760986,24.7001953125,24.7633781433105,24.7904300689697,24.8550796508789,24.9563484191895,24.9484367370605,24.98046875,24.99169921875],[23.5509777069092,23.5114250183105,23.466796875,23.4352531433105,23.4193363189697,23.4390621185303,23.4622077941895,23.4836921691895,23.5155277252197,23.5348625183105,23.5509777069092],[20.6123027801514,20.6247081756592,20.6952152252197,20.7888679504395,20.78076171875,20.7613277435303,20.6061515808105,20.5689449310303,20.5235347747803,20.4955081939697,20.4987316131592,20.4733409881592,20.4521484375,20.3910160064697,20.3525390625,20.3522453308105,20.40869140625,20.43505859375,20.4814453125,20.5196304321289,20.54833984375,20.55029296875,20.5631847381592,20.6123027801514],[20.7586917877197,20.7092761993408,20.6463871002197,20.6236324310303,20.6497077941895,20.6748046875,20.7012691497803,20.7010746002197,20.7116222381592,20.7391605377197,20.7586917877197],[26.0940418243408,25.99853515625,25.8918952941895,25.8743152618408,25.95263671875,25.9914073944092,25.9599609375,25.8512687683105,25.8460941314697,26.0125007629395,26.1104488372803,26.1603527069092,26.1412105560303,26.1496105194092,26.1570301055908,26.1107425689697,26.1031246185303,26.0940418243408],[20.6867179870605,20.6478519439697,20.6143569946289,20.583984375,20.5546875,20.5579090118408,20.5924816131592,20.6346683502197,20.6941413879395,20.7196273803711,20.71484375,20.6867179870605],[24.6747074127197,24.5690422058105,24.541015625,24.5645503997803,24.56640625,24.4610347747803,24.4734363555908,24.4856433868408,24.5640640258789,24.5812492370605,24.6747074127197],[23.4154281616211,23.4719715118408,23.5249996185303,23.63623046875,23.6884765625,23.8782215118408,24.0990238189697,24.1275386810303,24.1546878814697,24.19970703125,24.2110347747803,24.1875,24.2201175689697,24.2757816314697,24.3596687316895,24.4639644622803,24.5632820129395,24.58837890625,24.5785160064697,24.5365238189697,24.5023441314697,24.47265625,24.44580078125,24.41650390625,24.3594722747803,24.3177738189697,24.2120113372803,24.1925773620605,24.1890621185303,24.1441402435303,24.1028327941895,24.0635738372803,24.0418949127197,24.0401363372803,23.88623046875,23.7587890625,23.6507816314697,23.6173820495605,23.5533199310303,23.5052738189697,23.4652347564697,23.3640613555908,23.2521476745605,23.1439456939697,23.0291004180908,22.9357433319092,22.88134765625,22.8703136444092,22.986328125,23.1457996368408,23.2582035064697,23.3126964569092,23.4154281616211],[23.77978515625,23.7351570129395,23.6661128997803,23.5939464569092,23.77978515625],[23.8879890441895,23.8412113189697,23.8880863189697,23.9708995819092,23.9397468566895,23.8879890441895],[26.41015625,26.3927726745605,26.5310554504395,26.5782222747803,26.5956058502197,26.583984375,26.5315437316895,26.4886722564697,26.5031242370605,26.5471668243408,26.46875,26.39013671875,26.1608409881592,26.10791015625,26.2451171875,26.2731437683105,26.1759757995605,26.0723628997803,25.90625,25.85546875,25.8441410064697,25.9095687866211,26.0264644622803,26.08837890625,26.1648445129395,26.1654300689697,26.3477535247803,26.41015625],[20.0779304504395,20.099609375,19.9750003814697,19.8839855194092,19.8088874816895,19.64892578125,19.646484375,19.7073249816895,19.8385753631592,19.8917007446289,19.9260749816895,19.9368171691895,19.8622074127197,19.8466796875,19.9041023254395,19.9031257629395,19.9273433685303,19.9552745819092,20.0277347564697,20.0779304504395],[25.4376964569092,25.39990234375,25.3720722198486,25.3570308685303,25.2987308502197,25.2633781433105,25.2517566680908,25.2494144439697,25.2238292694092,25.2032222747803,25.1851558685303,25.12646484375,25.0622081756592,25.0652351379395,25.0523433685303,25.0580062866211,25.2341804504395,25.2857418060303,25.3480472564697,25.3736343383789,25.4491214752197,25.4376964569092],[25.6857433319092,25.5726566314697,25.4480476379395,25.5685539245605,25.6243171691895,25.6642589569092,25.6857433319092],[24.7742195129395,24.6458969116211,24.5155277252197,24.5166988372803,24.5855464935303,24.6233386993408,24.7191410064697,24.7736320495605,24.7863273620605,24.7686519622803,24.7742195129395],[26.0389633178711,26.0107421875,25.8556632995605,25.4967765808105,25.3252925872803,25.2500972747803,25.1044921875,25.0046882629395,24.79296875,24.6787109375,24.5565433502197,24.47705078125,24.3837890625,24.234375,24.0823249816895,23.9460926055908,23.7627925872803,23.7432632446289,23.7787113189697,23.87890625,23.8319339752197,23.8667964935303,23.9320316314697,24.0305652618408,24.2127933502197,24.29248046875,24.3433589935303,24.232421875,24.1587886810303,24.0560550689697,23.9131832122803,23.8234367370605,23.7279300689697,23.7205066680908,23.8234367370605,23.9175796508789,23.9674797058105,24.0007801055908,23.9818363189697,23.9470691680908,23.8353519439697,23.66455078125,23.42626953125,23.3864269256592,23.4332027435303,23.4670886993408,23.6741199493408,23.6575202941895,23.6273441314697,23.3956050872803,23.3282222747803,23.31201171875,23.09814453125,22.896484375,22.8513679504395,22.8928718566895,22.9222660064697,22.8114261627197,22.7418956756592,22.6294918060303,22.6249027252197,22.6426753997803,22.60546875,22.5693359375,22.5921878814697,22.8357410430908,22.9190425872803,22.9788093566895,23.1034164428711,23.2333984375,23.2884750366211,23.3277339935303,23.2183589935303,23.1546878814697,23.1194343566895,23.1687488555908,23.1617183685303,22.9928722381592,22.92138671875,22.8389644622803,22.8860359191895,22.93896484375,22.9655284881592,23.0666999816895,22.9304676055908,22.8026351928711,22.6768550872803,22.5967769622803,22.5691413879395,22.63427734375,22.6875,22.7740230560303,23.0203132629395,23.1376953125,23.2529296875,23.3689460754395,23.5696296691895,23.6839847564697,23.8360347747803,23.9669914245605,24.00537109375,24.0245132446289,24.0330066680908,24.0613288879395,24.0623035430908,24.0553722381592,24.0197257995605,23.9715824127197,23.8773441314697,23.7328128814697,23.5804691314697,23.5372066497803,23.5017566680908,23.4202156066895,23.1936531066895,23.08740234375,23.0474605560303,23.0363273620605,23.0861339569092,23.1471672058105,23.1471672058105,23.1975593566895,23.2626953125,23.3475589752197,23.3961906433105,23.4087905883789,23.4581050872803,23.4906253814697,23.4892578125,23.2525386810303,23.2030277252197,23.1615238189697,23.1000003814697,23.0964851379395,23.01513671875,22.9405269622803,22.85107421875,22.7749996185303,22.7253894805908,22.7650394439697,22.85107421875,22.9950180053711,23.0603504180908,23.0735359191895,23.041015625,23.1117191314697,23.16015625,23.1068363189697,23.060546875,22.98291015625,22.8323230743408,22.7798824310303,22.7171878814697,22.6083984375,22.4890613555908,22.4894523620605,22.427734375,22.3748054504395,22.3812484741211,22.3759765625,22.2311515808105,22.1647453308105,22.1337890625,22.0804691314697,22.01171875,21.95556640625,21.9400386810303,21.9342784881592,21.8923835754395,21.7380867004395,21.5829086303711,21.5788078308105,21.6924800872803,21.6789054870605,21.5712890625,21.4162101745605,21.3292980194092,21.2884769439697,21.2052726745605,21.1379871368408,21.1247062683105,21.14501953125,21.30810546875,21.4037113189697,21.451171875,21.5487308502197,21.6583976745605,21.7484378814697,21.82470703125,21.9533214569092,22.2437515258789,22.5558605194092,22.7115230560303,22.7996082305908,22.8463859558105,22.9203128814697,22.9169921875,22.8931636810303,22.9547863006592,23.1220703125,23.1525402069092,23.1834964752197,23.14892578125,23.0935535430908,23.0343742370605,22.9954090118408,22.9325180053711,22.8343753814697,22.78369140625,22.75390625,22.5833988189697,22.4216785430908,22.38525390625,22.3199234008789,22.2268562316895,21.96533203125,21.8046875,21.7170906066895,21.6500988006592,21.5676746368408,21.4725589752197,21.39013671875,21.35546875,21.3310546875,21.3297863006592,21.3033199310303,21.1826171875,21.1131839752197,21.0597667694092,20.9921875,20.8732433319092,20.77685546875,20.7685546875,20.77734375,20.8931636810303,21.07421875,21.1116218566895,21.15234375,21.14453125,21.1183586120605,21.0685558319092,21.0340824127197,20.9227523803711,20.7796878814697,20.71337890625,20.6913089752197,20.5716800689697,20.46826171875,20.30078125,20.19140625,20.0994148254395,20.0012702941895,20.0225582122803,20.0597648620605,20.1310558319092,20.2068367004395,20.2482433319092,20.2720699310303,20.28759765625,20.2938480377197,20.30615234375,20.3640613555908,20.3824214935303,20.3816413879395,20.34423828125,20.3113288879395,20.3111324310303,20.3384761810303,20.3836917877197,20.4080066680908,20.4560546875,20.5270500183105,20.6062507629395,20.6574230194092,20.6649417877197,20.6969718933105,20.7178707122803,20.7516593933105,20.77001953125,20.8060531616211,20.8816394805908,20.9501953125,21.001953125,21.0308589935303,21.0310535430908,20.9878902435303,20.9557609558105,20.9642581939697,21.1000003814697,21.1475582122803,21.32373046875,21.4041023254395,21.4596672058105,21.5757808685303,21.6275386810303,21.7794933319092,21.9294929504395,21.9933586120605,22.1388683319092,22.1844730377197,22.2376956939697,22.4007816314697,22.4935550689697,22.6036128997803,22.7248039245605,22.7550773620605,22.7838878631592,22.8592777252197,22.916015625,23.0255851745605,23.1559562683105,23.2398433685303,23.3720703125,23.4333992004395,23.5358390808105,23.6351566314697,23.7623043060303,23.880859375,23.9735355377197,24.0113277435303,24.0329093933105,24.0560550689697,24.2303714752197,24.2894535064697,24.38671875,24.4878921508789,24.5182609558105,24.5693359375,24.5959968566895,24.6510753631592,24.7737293243408,24.7958011627197,24.8468761444092,24.9935550689697,25.1333980560303,25.2511711120605,25.3819332122803,25.5270500183105,25.6214847564697,25.7239246368408,25.7849617004395,25.92333984375,26.06640625,26.1353511810303,26.1551761627197,26.1435546875,26.1112308502197,26.0769538879395,26.0660152435303,26.0855484008789,26.107421875,26.2005844116211,26.3208980560303,26.4105472564697,26.4624996185303,26.4950199127197,26.5445308685303,26.5813484191895,26.6097640991211,26.6249027252197,26.6023445129395,26.5364246368408,26.3306636810303,26.32568359375,26.3284168243408,26.3326187133789,26.3541030883789,26.3541030883789,26.3310546875,26.2412109375,26.1789054870605,26.1091804504395,26.0697250366211,26.0389633178711],[-61.7155265808105,-61.7821769714355,-61.7557640075684,-61.7499046325684,-61.7149887084961,-61.6604461669922,-61.6070289611816,-61.6271514892578,-61.7155265808105],[-46.2667007446289,-46.3815422058105,-46.496337890625,-46.5531234741211,-46.6662139892578,-46.7880859375,-46.7899894714355,-46.3938941955566,-46.2052268981934,-46.2186012268066,-46.2544937133789,-46.2667007446289],[-37.03125,-37.1868133544922,-37.2384262084961,-37.222900390625,-37.0475082397461,-36.9530754089355,-36.9869155883789,-37.03125],[-51.013671875,-51.17041015625,-51.2020530700684,-51.2339820861816,-51.3148918151855,-51.3388671875,-51.3189430236816,-51.3502922058105,-51.2088851928711,-51.0945777893066,-50.9402351379395,-50.6790046691895,-50.6979026794434,-50.75439453125,-50.9117164611816,-50.9672355651855,-50.9778823852539,-50.9704093933105,-51.013671875],[-52.7311553955078,-52.3982429504395,-52.0453109741211,-52.0107917785645,-51.9833984375,-51.97705078125,-51.985107421875,-52.0074729919434,-51.981689453125,-51.9077644348145,-51.9001960754395,-51.9884300231934,-52.1125984191895,-52.7704582214355,-53.0031242370605,-53.5784187316895,-53.7543487548828,-53.7931632995605,-53.9020500183105,-54.0511741638184,-54.1210479736328,-54.1827163696289,-54.1581535339355,-54.04736328125,-53.8895988464355,-53.6583023071289,-53.7222671508789,-53.7830581665039,-53.8249969482422,-53.9214859008789,-53.9937477111816,-54.1332054138184,-54.4969711303711,-54.734130859375,-54.8041038513184,-54.8657722473145,-54.9191398620605,-54.8412590026855,-54.7878913879395,-54.6646003723145,-54.3630867004395,-54.3226089477539,-54.6524429321289,-54.7736320495605,-54.809326171875,-54.8307609558105,-54.8304672241211,-54.8155784606934,-54.7862319946289,-54.7059555053711,-54.3716316223145,-54.0072288513184,-53.3751449584961,-53.2967262268066,-53.1029319763184,-52.7311553955078],[-51.6751441955566,-51.8086929321289,-52.119384765625,-52.1441879272461,-52.1480484008789,-52.1067390441895,-51.9698219299316,-51.8069343566895,-51.63134765625,-51.60693359375,-51.6751441955566],[-25.4323234558105,-25.397216796875,-25.3936538696289,-25.4013671875,-25.4208011627197,-25.4677734375,-25.380126953125,-25.3516616821289,-25.3463382720947,-25.4022445678711,-25.8005847930908,-25.9113292694092,-26.0497074127197,-26.2178707122803,-26.2738761901855,-26.3391609191895,-26.6046867370605,-27.1047840118408,-27.6900386810303,-27.8979988098145,-28.0030269622803,-28.0352535247803,-28.0368175506592,-27.967529296875,-27.9395503997803,-27.8052730560303,-27.7142105102539,-27.7439937591553,-27.7089347839355,-27.6172370910645,-27.3874988555908,-27.2388687133789,-26.9755859375,-26.6217765808105,-26.3374519348145,-25.8188972473145,-25.7268047332764,-25.6608390808105,-25.6123046875,-25.458251953125,-25.4323234558105],[-53.5352020263672,-53.6288070678711,-53.8975601196289,-53.941162109375,-53.9578132629395,-53.9474601745605,-53.8618659973145,-53.7009773254395,-53.58447265625,-53.5123519897461,-53.44140625,-53.43212890625,-53.4369125366211,-53.4554672241211,-53.48779296875,-53.5352020263672],[-55.0168952941895,-55.158203125,-55.2735824584961,-55.5236320495605,-55.5666007995605,-55.6341323852539,-55.6865234375,-55.781005859375,-55.8139152526855,-55.8271484375,-55.8690414428711,-55.9356956481934,-56.0426750183105,-56.140869140625,-56.2147979736328,-56.0780754089355,-55.9935569763184,-55.6664543151855,-55.57421875,-55.5162582397461,-55.4279327392578,-55.2346687316895,-55.205810546875,-55.0330047607422,-55.0168952941895],[-18.0005359649658,-17.9211921691895,-17.8858890533447,-17.7624015808105,-17.4978523254395,-17.3919925689697,-17.5860347747803,-18.3533210754395,-18.6708011627197,-18.8913078308105,-18.8824710845947,-18.8560562133789,-18.6355457305908,-18.4503421783447,-18.2298831939697,-18.0005359649658],[-18.5826168060303,-18.697265625,-19.0853519439697,-19.0853519439697,-19.0588855743408,-18.8824710845947,-18.7326164245605,-18.6620121002197,-18.5826168060303],[-71.6673278808594,-72.0235290527344,-72.3744125366211,-72.4949264526367,-72.4895477294922,-72.4364242553711,-72.2467803955078,-72.0890579223633,-71.9827651977539,-71.73291015625,-71.5521469116211,-71.4334411621094,-71.4670867919922,-71.6673278808594],[-17.9537105560303,-18.1479988098145,-18.2200202941895,-18.1740226745605,-17.9037094116211,-17.8135738372803,-17.6814460754395,-17.641357421875,-17.7294921875,-17.9537105560303],[-18.9971675872803,-19.1294918060303,-19.2176265716553,-19.2970218658447,-19.314697265625,-19.11181640625,-19.0059566497803,-18.9354000091553,-18.9530277252197,-18.9530277252197,-18.8824710845947,-18.9971675872803],[-17.6124992370605,-18.0358409881592,-18.6620121002197,-19.0324230194092,-19.1382808685303,-18.9971675872803,-18.54736328125,-17.98291015625,-17.4713859558105,-17.4008312225342,-17.6124992370605],[-18.6647453308105,-18.7676753997803,-19.0314445495605,-19.3692855834961,-19.594482421875,-19.6105480194092,-19.4947280883789,-19.3145503997803,-19.06689453125,-18.8126964569092,-18.6647453308105],[-44.8645515441895,-45.0674324035645,-45.4907722473145,-46.1610336303711,-46.7519035339355,-47.3075180053711,-47.3512229919434,-47.2722663879395,-46.7872047424316,-46.399169921875,-45.411376953125,-44.91748046875,-44.7499046325684,-44.7763671875,-44.8645515441895],[-29.9528789520264,-28.9919910430908,-28.4837894439697,-28.3770503997803,-27.6883792877197,-27.034423828125,-25.9474124908447,-25.7950687408447,-25.9124526977539,-26.1827144622803,-27.5718746185303,-30.0919933319092,-31.5339832305908,-31.9926738739014,-32.03271484375,-31.8367691040039,-31.5155754089355,-30.3860359191895,-29.9635753631592,-29.1749992370605,-28.1514644622803,-27.738525390625,-27.0020523071289,-26.1408195495605,-25.1233882904053,-24.8451671600342,-24.4703121185303,-24.1736316680908,-23.9195308685303,-23.8335456848145,-23.6946296691895,-23.4069328308105,-22.52490234375,-21.9196796417236,-21.6917953491211,-21.58251953125,-21.5206546783447,-21.6157703399658,-21.9939441680908,-22.4725589752197,-23.1180667877197,-23.8622074127197,-29.5793952941895,-29.7727527618408,-29.8874015808105,-29.8109855651855,-29.5438461303711,-28.91943359375,-27.8395023345947,-27.0459480285645,-25.1488285064697,-24.5891609191895,-24.2930679321289,-23.6365242004395,-23.4961433410645,-23.3929691314697,-23.310546875,-23.248779296875,-23.1798324584961,-23.1037101745605,-22.9400863647461,-22.5633773803711,-21.5755386352539,-21.3379878997803,-21.1673831939697,-21.1303215026855,-21.1179676055908,-21.1234378814697,-21.1465816497803,-21.2305183410645,-21.50390625,-21.7236328125,-21.9607410430908,-22.415283203125,-22.57275390625,-23.0724601745605,-23.1963863372803,-23.2036628723145,-23.1177253723145,-22.9728507995605,-22.9189453125,-22.82568359375,-22.0894031524658,-21.9313488006592,-21.4497547149658,-21.1424331665039,-20.8897457122803,-20.755859375,-20.0157222747803,-19.6299304962158,-19.2247562408447,-19.1529788970947,-18.6673831939697,-18.45654296875,-18.1178703308105,-17.9693832397461,-17.7166500091553,-17.4560546875,-17.2262210845947,-17.1590328216553,-16.9370613098145,-16.6371097564697,-16.3589839935303,-16.2667980194092,-16.1207027435303,-15.9688968658447,-15.5555181503296,-15.4506340026855,-15.2274904251099,-14.2419929504395,-13.7044916152954,-12.9560060501099,-12.4344244003296,-12.19287109375,-11.8411130905151,-11.5574703216553,-11.425537109375,-11.4306640625,-11.52880859375,-12.2313470840454,-12.4612302780151,-13.126220703125,-13.451171875,-13.8042964935303,-14.1973628997803,-14.4523439407349,-14.4901371002197,-14.3084955215454,-14.2285642623901,-14.2401857376099,-14.4312505722046,-14.5035638809204,-15.1942386627197,-15.5426759719849,-15.9975099563599,-16.3189449310303,-16.7605953216553,-16.5877914428711,-16.429443359375,-15.937255859375,-15.9326162338257,-16.1677722930908,-16.48876953125,-16.8684062957764,-17.0111331939697,-17.191162109375,-17.3572273254395,-17.7228527069092,-18.0709476470947,-18.6925773620605,-19.0290050506592,-19.2060050964355,-19.42919921875,-19.5150394439697,-19.8667964935303,-20.0395011901855,-20.150146484375,-20.1974124908447,-20.1813468933105,-20.1384773254395,-20.06884765625,-19.9853992462158,-19.8393077850342,-19.5178718566895,-19.3915042877197,-19.35302734375,-19.2835922241211,-19.2959957122803,-19.3542003631592,-19.3993167877197,-19.4312019348145,-19.4140129089355,-19.2839832305908,-19.2229499816895,-19.1521968841553,-19.07177734375,-19.0113277435303,-18.9708003997803,-18.9919929504395,-19.074951171875,-19.26220703125,-19.7230472564697,-19.7698230743408,-19.8060550689697,-19.8315906524658,-19.88720703125,-19.972900390625,-20.0504875183105,-20.1999015808105,-20.3957023620605,-20.6155757904053,-21.1337413787842,-21.1414546966553,-20.9474601745605,-20.9556636810303,-21.1947746276855,-21.2602043151855,-21.31201171875,-21.3972663879395,-21.5159664154053,-21.6326656341553,-21.74755859375,-21.7295894622803,-21.5789070129395,-21.3796882629395,-21.1318855285645,-20.8625984191895,-20.5718269348145,-20.318603515625,-19.9951171875,-19.7243156433105,-19.4904289245605,-19.3939933776855,-19.296875,-19.2960929870605,-19.467529296875,-19.5241222381592,-19.9532222747803,-20.1620616912842,-20.439208984375,-20.6808109283447,-20.4637699127197,-20.23193359375,-19.8086433410645,-19.5875968933105,-19.4264163970947,-19.30029296875,-19.1310062408447,-18.9034175872803,-18.5858898162842,-18.442626953125,-18.3390140533447,-18.2923812866211,-18.302734375,-18.3372573852539,-18.3960437774658,-18.5103034973145,-18.6057605743408,-18.7402839660645,-18.8653316497803,-18.9810047149658,-19.1563472747803,-19.5087890625,-19.8649406433105,-20.0643558502197,-20.4867191314697,-20.9420890808105,-20.9599132537842,-21.6146965026855,-21.7490234375,-21.9308109283447,-22.1852531433105,-22.3343257904053,-22.5545406341553,-22.6093254089355,-22.6066398620605,-22.4444332122803,-22.3786125183105,-22.2948722839355,-22.0037097930908,-21.8773441314697,-21.7581062316895,-21.569091796875,-21.4882335662842,-21.4168453216553,-21.1854496002197,-20.8874015808105,-20.7833003997803,-20.5638179779053,-20.4354000091553,-20.279296875,-20.1036128997803,-19.8628902435303,-19.9577159881592,-19.806884765625,-19.5660152435303,-19.5089836120605,-19.4856929779053,-19.4802742004395,-19.462158203125,-19.4314460754395,-19.3995113372803,-19.366455078125,-19.3752937316895,-19.4259757995605,-19.5263671875,-19.67626953125,-19.7984867095947,-19.8931636810303,-20.0265617370605,-20.1986808776855,-20.4849624633789,-20.9058609008789,-21.0938472747803,-21.2465324401855,-21.409423828125,-21.6493167877197,-21.8610343933105,-22.2328624725342,-22.0977535247803,-21.9043445587158,-21.783935546875,-21.6951179504395,-21.59765625,-21.4573249816895,-21.1405754089355,-21.0566902160645,-20.9857902526855,-20.9277839660645,-20.861083984375,-20.7856922149658,-20.7953109741211,-20.8899898529053,-20.9709968566895,-21.0382823944092,-21.0384769439697,-20.861572265625,-20.6111316680908,-20.53173828125,-20.4170894622803,-20.2142581939697,-19.9849109649658,-19.7997074127197,-19.5377941131592,-19.4273452758789,-19.2870121002197,-19.22509765625,-19.2416496276855,-19.2715816497803,-19.31494140625,-19.369140625,-19.4667472839355,-19.646240234375,-20.0475578308105,-20.2564449310303,-20.2305660247803,-20.6531238555908,-21.1294422149658,-21.58056640625,-21.9549312591553,-21.83203125,-21.761962890625,-21.9429206848145,-21.9826164245605,-21.920166015625,-21.9727058410645,-22.1771965026855,-22.3215827941895,-22.3343257904053,-22.2635250091553,-22.2173328399658,-22.1956539154053,-22.2200183868408,-22.2905769348145,-22.3289546966553,-22.3352546691895,-22.2705574035645,-22.1348152160645,-21.9876956939697,-21.2982902526855,-21.022216796875,-20.3672866821289,-20.3379878997803,-20.4489250183105,-20.5096683502197,-20.63671875,-21.3258800506592,-21.5479984283447,-21.8728504180908,-22.18505859375,-22.3468742370605,-22.9875011444092,-23.2332019805908,-23.7605953216553,-24.15771484375,-24.3398933410645,-24.4512691497803,-24.5663089752197,-24.67724609375,-24.7841796875,-24.9054679870605,-25.1088371276855,-25.3514652252197,-25.5212879180908,-25.5277347564697,-25.4274425506592,-25.280517578125,-24.9088878631592,-24.7783203125,-24.791259765625,-25.02587890625,-25.3107433319092,-25.4500980377197,-25.6654300689697,-25.7401866912842,-26.0623054504395,-26.1685543060303,-26.40673828125,-26.7654800415039,-26.9767074584961,-27.2704105377197,-27.1693840026855,-26.6036128997803,-26.5418453216553,-26.6576175689697,-26.7286128997803,-26.8633308410645,-27.0618648529053,-27.264892578125,-27.4723625183105,-27.5616207122803,-27.5325679779053,-27.483154296875,-27.413330078125,-27.3480472564697,-27.1898918151855,-27.0700187683105,-26.7532234191895,-26.432861328125,-26.2020015716553,-26.0287590026855,-25.3990230560303,-25.268310546875,-25.0570316314697,-24.5872077941895,-24.1326675415039,-23.8989734649658,-23.7096195220947,-23.4557609558105,-23.2440910339355,-22.9960441589355,-22.852294921875,-22.4501953125,-22.1942386627197,-22.0363273620605,-22.0234851837158,-22.0067386627197,-22.0748043060303,-22.2802238464355,-22.2392578125,-22.293212890625,-22.4975090026855,-22.7068367004395,-23.2080078125,-23.6743640899658,-23.8555660247803,-24.0690441131592,-24.3585948944092,-24.5472164154053,-24.6299800872803,-24.78857421875,-24.9924793243408,-25.1705570220947,-25.255859375,-25.86083984375,-26.0804691314697,-26.2057609558105,-26.6576175689697,-26.4765625,-26.39208984375,-26.20947265625,-26.0998058319092,-25.68798828125,-25.357421875,-25.2374992370605,-24.9848155975342,-24.8133296966553,-24.7894535064697,-24.7710456848145,-24.6500015258789,-24.70068359375,-24.8369140625,-25.1280269622803,-25.2037105560303,-25.1178703308105,-24.8441905975342,-24.6668453216553,-24.5723628997803,-24.4171867370605,-24.2422847747803,-23.7977066040039,-23.5871086120605,-23.2909183502197,-22.9557628631592,-22.8685054779053,-22.5621585845947,-22.4968757629395,-22.3702144622803,-22.2645034790039,-21.9596672058105,-22.0133304595947,-22.31103515625,-22.4649906158447,-22.5032234191895,-22.4885730743408,-22.4796390533447,-22.4175796508789,-22.3477535247803,-22.2990245819092,-22.2337894439697,-22.1695804595947,-21.96142578125,-21.7522449493408,-21.6979484558105,-21.6711921691895,-21.6896476745605,-21.6666011810303,-21.6745109558105,-21.6251468658447,-21.5739269256592,-21.5226573944092,-21.6255359649658,-21.9435062408447,-22.0692863464355,-22.3841304779053,-22.384521484375,-22.3998527526855,-22.401123046875,-22.422119140625,-22.43701171875,-22.5260753631592,-22.5313472747803,-22.5550289154053,-22.6096687316895,-22.690673828125,-22.9425773620605,-23.1906242370605,-23.3278312683105,-23.7917957305908,-23.9713878631592,-24.13037109375,-24.228515625,-24.2657222747803,-24.3770008087158,-24.5622062683105,-24.781005859375,-25.0333976745605,-25.2549800872803,-25.4457988739014,-25.6558589935303,-25.8851566314697,-26.21142578125,-26.6885261535645,-27.0106430053711,-27.0872077941895,-27.1623039245605,-27.1070308685303,-26.7372074127197,-26.4520034790039,-26.0740737915039,-25.8427238464355,-25.7578125,-25.6994132995605,-25.6675777435303,-25.7422370910645,-26.0141105651855,-26.1575183868408,-26.5759754180908,-26.7179183959961,-27.0673351287842,-27.335693359375,-27.6887683868408,-27.888916015625,-28.3031253814697,-28.3984375,-28.2915534973145,-28.1158695220947,-27.9921875,-27.9792957305908,-28.0238761901855,-28.0698738098145,-28.1456546783447,-28.41748046875,-28.5300769805908,-29.0368175506592,-29.0720710754395,-28.9534664154053,-28.6331062316895,-28.5409183502197,-28.0150394439697,-27.5960941314697,-26.7472648620605,-26.677490234375,-26.6217765808105,-26.5654296875,-26.5083980560303,-26.5768070220947,-26.7706546783447,-27.072509765625,-27.2032222747803,-27.328125,-27.5608386993408,-27.6288585662842,-27.3841800689697,-27.2742195129395,-27.1444835662842,-27.0277347564697,-26.7521476745605,-26.4156723022461,-26.1557121276855,-25.6248531341553,-25.5298824310303,-24.7488288879395,-24.041015625,-23.6673355102539,-23.1732425689697,-22.2844715118408,-22.2065906524658,-22.2354488372803,-22.2870616912842,-22.4351062774658,-22.6149406433105,-22.7262210845947,-22.8208980560303,-23.0336437225342,-23.0882320404053,-23.0145511627197,-23.049560546875,-23.2369632720947,-23.5525398254395,-23.8116207122803,-23.86572265625,-23.8165512084961,-23.7642593383789,-23.708984375,-23.7394046783447,-23.8555660247803,-23.9436511993408,-24.2475090026855,-24.2966804504395,-24.2522964477539,-24.2270488739014,-24.2208995819092,-24.2955570220947,-24.4510746002197,-24.7405757904053,-24.8666019439697,-25.1325187683105,-25.1885757446289,-25.0804691314697,-25.0924320220947,-25.272216796875,-25.5440425872803,-25.5811519622803,-25.6063480377197,-25.6261234283447,-25.697998046875,-25.9558601379395,-26.1386241912842,-26.229248046875,-26.3414058685303,-26.48291015625,-26.6537094116211,-26.8153305053711,-27.0811519622803,-27.2662601470947,-27.8512191772461,-28.12646484375,-28.3645515441895,-28.8543453216553,-29.0876960754395,-29.2495098114014,-29.4262199401855,-29.7135734558105,-29.8685054779053,-29.9637699127197,-30.0511207580566,-30.1955089569092,-30.3181133270264,-30.7200183868408,-30.7118663787842,-30.6056652069092,-30.6107425689697,-30.8497543334961,-30.9785652160645,-31.1684589385986,-31.4194812774658,-31.7419929504395,-32.1372566223145,-32.3274421691895,-32.3136711120605,-32.2696266174316,-32.1952133178711,-32.1801261901855,-32.2242164611816,-32.2823753356934,-32.3545875549316,-32.3668441772461,-32.2489242553711,-32.1559562683105,-32.16455078125,-32.2748069763184,-32.3695297241211,-32.44873046875,-32.9180183410645,-33.0487327575684,-33.108154296875,-33.156982421875,-33.2936019897461,-33.348876953125,-33.45849609375,-33.5044441223145,-33.517578125,-33.4977531433105,-33.5279769897461,-33.6082000732422,-33.88134765625,-34.1016578674316,-34.1982421875,-34.2688980102539,-34.3136253356934,-34.4228515625,-34.4758796691895,-34.52392578125,-34.5762672424316,-34.6328125,-35.0746574401855,-35.1885757446289,-35.2908706665039,-35.4117202758789,-35.6620597839355,-35.7054710388184,-35.8672370910645,-35.8618621826172,-35.834716796875,-35.8121109008789,-35.7555160522461,-35.6300773620605,-35.7292976379395,-35.8179168701172,-36.044189453125,-36.2887191772461,-36.3791999816895,-36.3992195129395,-36.3889656066895,-36.5272483825684,-36.5227546691895,-36.5370101928711,-36.6372528076172,-36.6651878356934,-36.7145004272461,-36.8221664428711,-36.9324226379395,-37.02587890625,-37.0627937316895,-37.2332000732422,-37.3160133361816,-37.329833984375,-37.4100608825684,-37.5160636901855,-37.6637687683105,-37.7541999816895,-37.9547843933105,-38.0012702941895,-37.84228515625,-37.79736328125,-37.8265113830566,-37.787841796875,-37.4844741821289,-37.2787132263184,-37.2906761169434,-37.5699234008789,-37.8139152526855,-38.0516586303711,-38.1566390991211,-37.9891586303711,-37.7523422241211,-37.868896484375,-37.9694366455078,-38.0734367370605,-38.1399383544922,-38.3981437683105,-38.5203628540039,-38.4426765441895,-38.2163581848145,-38.2018547058105,-38.203369140625,-38.63671875,-39.0889625549316,-39.4133796691895,-39.9609375,-40.1735343933105,-40.1915512084961,-39.6559600830078,-39.5779304504395,-39.6525421142578,-39.76318359375,-39.937255859375,-40.0280265808105,-40.2531242370605,-40.6675796508789,-40.8805656433105,-41.0844230651855,-41.0886726379395,-41.0277328491211,-40.9660148620605,-40.8292961120605,-40.6554679870605,-40.5210952758789,-40.4327163696289,-40.2784194946289,-40.2098617553711,-40.1822242736816,-40.2784652709961,-40.4776382446289,-40.6985359191895,-40.6864242553711,-40.78173828125,-40.9845695495605,-41.0793952941895,-41.177734375,-41.5810050964355,-41.1749992370605,-41.0305633544922,-40.96630859375,-40.82568359375,-40.6177711486816,-40.65234375,-40.561279296875,-40.5503883361816,-40.7715339660645,-40.7751960754395,-40.9068374633789,-41.0487289428711,-41.05615234375,-41.1522445678711,-41.1352043151855,-41.1077156066895,-41.1954612731934,-41.2747077941895,-41.3878898620605,-41.4478530883789,-41.6279296875,-41.844482421875,-42.0197257995605,-42.0923843383789,-42.1745109558105,-42.1429672241211,-42.093994140625,-41.9322776794434,-41.6344718933105,-41.6436004638672,-41.7232437133789,-41.9089851379395,-41.9749031066895,-42.0582542419434,-42.3156242370605,-42.4237289428711,-42.7411155700684,-42.8490715026855,-42.941650390625,-42.8552742004395,-42.6736831665039,-42.4671401977539,-42.1529769897461,-42.1643562316895,-42.2431640625,-42.1979484558105,-42.2331047058105,-42.2481460571289,-42.3214836120605,-42.2361335754395,-42.14306640625,-42.1538581848145,-42.1102066040039,-42.2497062683105,-42.3654289245605,-42.5304183959961,-42.5853042602539,-42.3236312866211,-42.3473625183105,-42.4187507629395,-42.4937477111816,-42.64599609375,-42.717041015625,-43.0440902709961,-43.1599617004395,-43.1890602111816,-43.3483390808105,-43.5983390808105,-43.7919921875,-43.9227066040039,-43.9395484924316,-43.8311500549316,-43.6654777526855,-43.5330581665039,-43.295654296875,-43.2129898071289,-43.156494140625,-43.1648406982422,-43.1653327941895,-43.1229019165039,-43.2348136901855,-43.3201179504395,-43.6168937683105,-43.6685066223145,-43.9550285339355,-43.9374008178711,-43.7301254272461,-43.6579093933105,-43.7061996459961,-43.7898406982422,-43.9065437316895,-44.1169891357422,-44.1054191589355,-44.0654792785645,-44.1617202758789,-44.2689437866211,-44.3312492370605,-44.3835945129395,-44.4129409790039,-44.4534683227539,-44.4049301147461,-44.2313499450684,-44.1761245727539,-44.224365234375,-44.3483390808105,-44.4763679504395,-44.5331535339355,-44.61328125,-44.8122100830078,-45.3792457580566,-45.3623542785645,-45.3677711486816,-45.2024879455566,-45.082275390625,-44.9747047424316,-44.853515625,-44.742431640625,-44.7567367553711,-45.0827140808105,-45.2833023071289,-45.3805160522461,-45.4288597106934,-45.5902824401855,-45.6952171325684,-45.934326171875,-45.9765129089355,-46.046630859375,-46.1419448852539,-46.0186538696289,-45.9337387084961,-45.8798828125,-45.8494148254395,-45.8702163696289,-45.9422836303711,-45.9756851196289,-45.9704093933105,-46.01171875,-46.2966766357422,-46.5824241638184,-46.7177734375,-46.8056640625,-46.8744621276855,-46.9796867370605,-47.1248512268066,-47.2244110107422,-47.3697242736816,-47.4646492004395,-47.5790519714355,-47.7074699401855,-47.7962379455566,-47.7888679504395,-47.7299308776855,-47.8279266357422,-48.0139656066895,-48.1075172424316,-48.1808090209961,-48.241943359375,-48.2051773071289,-47.9059562683105,-47.7703132629395,-47.8587913513184,-48.1461448669434,-48.1939430236816,-48.3864250183105,-48.3781242370605,-48.4249534606934,-48.4281730651855,-48.4948234558105,-48.5579109191895,-48.59716796875,-48.9220695495605,-48.9645042419434,-48.9872055053711,-49.0492172241211,-49.2047386169434,-49.2890625,-49.2222671508789,-49.193115234375,-49.2652359008789,-49.3112297058105,-49.3044929504395,-49.3628921508789,-49.3802719116211,-49.3134765625,-49.1297874450684,-49.070556640625,-49.0396499633789,-48.8287124633789,-49.0081558227539,-49.1204605102539,-49.2022972106934,-49.2779312133789,-49.3485374450684,-49.623779296875,-49.6642570495605,-49.6833953857422,-49.667724609375,-49.553466796875,-49.6852531433105,-49.8060531616211,-49.943359375,-50.0702133178711,-50.1791496276855,-50.2852058410645,-50.3192405700684,-50.2809104919434,-50.2593269348145,-50.2560043334961,-50.2987327575684,-50.2037582397461,-50.0760269165039,-49.7931137084961,-50.0922355651855,-50.3383293151855,-50.3900871276855,-50.408203125,-50.5015602111816,-50.572021484375,-50.603515625,-50.7435073852539,-50.8042945861816,-50.8904800415039,-51.0130844116211,-51.1875991821289,-51.4688453674316,-51.5381813049316,-51.4510726928711,-51.5475082397461,-51.2800750732422,-50.8975601196289,-50.6993637084961,-50.5850105285645,-50.3418960571289,-50.2606925964355,-50.3959465026855,-50.4866218566895,-50.4922866821289,-50.458740234375,-50.4370613098145,-50.4833984375,-50.7210464477539,-51.0723114013672,-51.3466796875,-51.39111328125,-51.4871101379395,-51.5422859191895,-51.5849113464355,-51.6820297241211,-51.7078628540039,-51.5337905883789,-51.4037628173828,-51.2315444946289,-51.1099128723145,-50.9065437316895,-50.8349113464355,-50.8577156066895,-50.8492164611816,-50.684326171875,-50.4920883178711,-50.3551254272461,-50.2689476013184,-50.158203125,-50.0089836120605,-50.0155296325684,-50.0929679870605,-50.1216316223145,-50.2199211120605,-50.2988739013672,-50.5169906616211,-50.6481437683105,-50.6778793334961,-50.6812973022461,-50.8121566772461,-50.854248046875,-50.9237289428711,-50.9606475830078,-50.9137191772461,-50.8523445129395,-50.7648429870605,-50.7215843200684,-50.7801742553711,-50.8910636901855,-50.9890632629395,-51.2206077575684,-51.1708488464355,-51.1389656066895,-51.25537109375,-51.3636245727539,-51.4009284973145,-51.470458984375,-51.6767578125,-51.7581062316895,-51.8349609375,-51.922607421875,-51.9986801147461,-52.0631828308105,-52.0934104919434,-52.0970230102539,-52.0888671875,-52.1240234375,-52.2354507446289,-52.259033203125,-52.4476089477539,-52.4503402709961,-52.4997062683105,-52.5376968383789,-52.5062484741211,-52.46142578125,-52.1795883178711,-51.970703125,-51.7210960388184,-51.619140625,-51.2529296875,-51.0903816223145,-51.0918960571289,-51.1463851928711,-51.393798828125,-51.7234382629395,-51.7798843383789,-51.9241180419922,-52.0353507995605,-52.3482437133789,-52.55126953125,-52.7609367370605,-52.9949226379395,-53.1529273986816,-53.198974609375,-53.2337379455566,-53.1063461303711,-53.2437477111816,-53.272216796875,-53.3447265625,-53.3920402526855,-53.35693359375,-53.0178718566895,-52.5108871459961,-52.2926292419434,-52.1579093933105,-52.0561027526855,-51.9321784973145,-51.8912124633789,-51.8221206665039,-51.6763687133789,-51.51708984375,-51.2585906982422,-51.2250022888184,-51.2810516357422,-51.4019508361816,-51.647705078125,-51.8230476379395,-52.4212417602539,-52.6758766174316,-52.814453125,-52.9219245910645,-53.0357894897461,-53.1560516357422,-53.4127464294434,-53.5387687683105,-53.6146965026855,-53.6480979919434,-53.6226043701172,-53.6347160339355,-53.5707054138184,-53.4756813049316,-53.435791015625,-53.4187507629395,-53.2227058410645,-53.1146469116211,-53.0382804870605,-52.6031265258789,-52.4910659790039,-52.4314460754395,-52.3868637084961,-52.4297370910645,-52.5601081848145,-52.9066886901855,-53.2269554138184,-53.3730964660645,-53.443603515625,-53.5600128173828,-53.6871566772461,-53.8844261169434,-53.8054656982422,-53.798583984375,-53.5479011535645,-53.4138679504395,-53.2235832214355,-52.9695320129395,-52.6664543151855,-52.5120124816895,-52.3835945129395,-51.9090843200684,-51.6650390625,-51.4505844116211,-51.1814460754395,-50.7053718566895,-50.6134757995605,-50.64013671875,-51.1710472106934,-51.16796875,-51.0320777893066,-50.8870124816895,-50.9688491821289,-51.3214836120605,-51.4232406616211,-51.7652359008789,-51.94384765625,-52.1042022705078,-52.3448257446289,-52.5461921691895,-52.6732406616211,-52.8983383178711,-52.9795875549316,-53.4188003540039,-53.6036109924316,-53.7352066040039,-53.642822265625,-53.6162109375,-53.6163558959961,-53.5779762268066,-53.3529281616211,-53.2113761901855,-53.1515617370605,-53.0409660339355,-52.8898429870605,-52.4360847473145,-52.0584945678711,-51.7799835205078,-51.5968742370605,-51.5183563232422,-51.4564933776855,-51.4326667785645,-51.4146957397461,-51.3936996459961,-51.33251953125,-51.207275390625,-51.1691398620605,-51.2101554870605,-51.29345703125,-51.4561042785645,-51.47802734375,-51.4750480651855,-51.6324234008789,-51.8040046691895,-52.1985359191895,-52.3785133361816,-52.6983909606934,-52.7467765808105,-52.780029296875,-53.1725120544434,-53.2898445129395,-53.3831520080566,-53.33740234375,-53.2132797241211,-53.0394515991211,-52.8938484191895,-52.6045913696289,-52.3027801513672,-51.7806663513184,-51.6231422424316,-51.4787139892578,-51.13330078125,-51.0699234008789,-50.9457054138184,-50.8006362915039,-50.8077163696289,-51.0302238464355,-51.1488761901855,-51.2494163513184,-51.1560554504395,-51.1197280883789,-51.0848617553711,-50.7922859191895,-50.3926734924316,-50.29736328125,-50.2988739013672,-50.4593734741211,-50.53662109375,-50.6710472106934,-50.85107421875,-51.0769538879395,-51.0578117370605,-50.8922386169434,-50.8751983642578,-50.8105964660645,-50.8041000366211,-50.7202644348145,-50.4590797424316,-50.3494644165039,-50.3434562683105,-50.5,-50.4602508544922,-50.3375015258789,-50.2917022705078,-50.3229484558105,-50.4360847473145,-50.60986328125,-50.8023414611816,-50.9727058410645,-51.105712890625,-51.18994140625,-51.4188499450684,-51.4990730285645,-51.5980987548828,-52.254638671875,-52.3360328674316,-52.5712394714355,-52.7650375366211,-53.0230445861816,-53.3575210571289,-53.7685050964355,-54.0144538879395,-54.1356468200684,-54.3433113098145,-54.5011749267578,-54.53076171875,-54.43798828125,-54.3435516357422,-54.1658210754395,-53.8591804504395,-53.6944313049316,-53.5130844116211,-53.3760261535645,-53.09130859375,-52.8019523620605,-52.6304206848145,-52.4052238464355,-51.7837905883789,-51.5244636535645,-51.4117202758789,-50.9468765258789,-50.8723640441895,-50.68212890625,-50.6632804870605,-50.7275390625,-50.9326667785645,-51.17333984375,-51.3228492736816,-51.3398933410645,-51.3204078674316,-51.2828140258789,-51.256591796875,-51.3960952758789,-51.4935531616211,-51.752685546875,-51.7743148803711,-51.6501007080078,-51.5284194946289,-51.26708984375,-51.1300773620605,-51.0304183959961,-51.0189437866211,-51.1777839660645,-51.3766593933105,-51.7918930053711,-52.0613746643066,-52.2335968017578,-52.4168930053711,-52.5345726013184,-52.7750015258789,-52.8966789245605,-53.007568359375,-53.1170387268066,-53.087890625,-53.0021018981934,-52.9373054504395,-52.891845703125,-52.7494125366211,-51.96728515625,-51.7699203491211,-51.7786140441895,-51.9117202758789,-52.0819358825684,-52.19580078125,-52.6562995910645,-52.7280769348145,-52.91455078125,-53.1675262451172,-53.2840805053711,-53.4400863647461,-53.46484375,-53.4760246276855,-53.3047370910645,-53.2497062683105,-53.1388664245605,-53.14453125,-53.3336944580078,-53.3583488464355,-53.3552742004395,-53.3736305236816,-53.4201202392578,-53.5753898620605,-53.6397933959961,-53.69287109375,-53.8098640441895,-53.7759780883789,-53.6720237731934,-53.6521453857422,-53.9005355834961,-53.927734375,-53.8809051513672,-53.8474578857422,-53.8275375366211,-53.7928695678711,-53.7029304504395,-53.6309547424316,-53.513671875,-53.4625015258789,-53.4775886535645,-53.5686492919922,-53.7154273986816,-53.7598609924316,-53.7796859741211,-53.8940925598145,-53.96435546875,-54.0199203491211,-53.9542961120605,-53.9120597839355,-53.9629898071289,-54.0989265441895,-54.1727066040039,-54.3177261352539,-54.6890602111816,-54.8183097839355,-55.0553703308105,-55.33642578125,-55.4478988647461,-55.5940399169922,-55.6678199768066,-55.6693382263184,-55.6297836303711,-55.5492172241211,-55.4524421691895,-55.3155746459961,-54.9708976745605,-54.8726043701172,-54.8401336669922,-54.8406219482422,-54.8963356018066,-55.3201179504395,-55.5814476013184,-55.6594696044922,-55.6358375549316,-55.5893096923828,-55.3779296875,-55.2957038879395,-55.427978515625,-55.5687484741211,-55.6017074584961,-55.456787109375,-55.1218719482422,-55.0462379455566,-54.9249534606934,-54.7903823852539,-54.7400398254395,-54.7287101745605,-54.7577171325684,-54.7608413696289,-54.7379379272461,-54.7730979919434,-54.8662109375,-55.0730972290039,-55.1339836120605,-55.1984367370605,-55.2889175415039,-55.3724136352539,-55.4595222473145,-55.5451164245605,-55.6339836120605,-55.6685562133789,-55.6908226013184,-55.6927757263184,-55.59228515625,-55.4523468017578,-55.3586883544922,-55.2971687316895,-55.2882804870605,-55.33203125,-55.4457015991211,-55.6562042236328,-55.7388687133789,-55.7579116821289,-55.7870635986328,-55.8755378723145,-55.9919929504395,-56.1040496826172,-56.1091804504395,-56.0826187133789,-56.0330085754395,-55.9684066772461,-55.8973159790039,-55.8382835388184,-55.8724136352539,-55.9294929504395,-55.9965324401855,-55.9989242553711,-56.0144538879395,-56.0662117004395,-56.1242179870605,-56.2253913879395,-56.2984848022461,-56.3921852111816,-56.4931640625,-56.6551742553711,-56.9542961120605,-57.1911125183105,-57.2305679321289,-57.1121101379395,-56.9375,-56.7063484191895,-56.6389656066895,-56.6639137268066,-56.654296875,-56.7176742553711,-56.656005859375,-56.445556640625,-56.3502922058105,-56.2554702758789,-56.5220680236816,-56.8013191223145,-56.8711433410645,-56.9325675964355,-56.9855461120605,-57.0716819763184,-57.1908683776855,-57.3647918701172,-57.8131790161133,-57.9670906066895,-58.1088371276855,-58.1796875,-58.2533187866211,-58.5655250549316,-58.6034660339355,-58.2811965942383,-58.2496566772461,-58.3813018798828,-58.5162124633789,-58.6630859375,-58.8814430236816,-59.0815925598145,-59.2636260986328,-59.4453163146973,-59.7174301147461,-60.1727523803711,-60.8746070861816,-61.1882286071777,-61.3748016357422,-61.6208457946777,-62.0966796875,-62.4961891174316,-62.7428703308105,-62.8234367370605,-63.0058097839355,-63.2913055419922,-63.4388656616211,-63.6219749450684,-63.8430709838867,-63.9603500366211,-64.1352005004883,-64.2231979370117,-64.3072738647461,-64.3873062133789,-64.5434112548828,-64.6920852661133,-64.9119644165039,-65.087646484375,-65.313232421875,-65.3699264526367,-65.456787109375,-65.57373046875,-65.6832962036133,-65.7854537963867,-65.875732421875,-65.9540557861328,-66.134033203125,-66.3617706298828,-66.4657745361328,-66.55322265625,-66.6599655151367,-66.8740234375,-66.9925765991211,-67.0787124633789,-67.0547866821289,-66.8539047241211,-66.6748046875,-66.826171875,-68.1487350463867,-68.3172836303711,-68.5606460571289,-68.7630844116211,-69.1075668334961,-69.3729019165039,-69.4608840942383,-69.4840850830078,-69.399658203125,-68.864990234375,-68.6607437133789,-68.2454147338867,-68.147216796875,-68.1142578125,-68.223388671875,-68.7673797607422,-69.2520523071289,-69.673828125,-69.7472152709961,-69.8186492919922,-69.8880844116211,-69.8722152709961,-69.7710418701172,-69.7117156982422,-69.6942367553711,-70.2244567871094,-70.4413070678711,-70.6131286621094,-70.7336883544922,-70.7928237915039,-70.7906265258789,-70.771240234375,-70.7346649169922,-71.0150375366211,-71.1414566040039,-71.1548843383789,-71.0554656982422,-70.9581069946289,-70.8628463745117,-70.6037139892578,-69.6565399169922,-68.9783172607422,-68.7474594116211,-68.5915985107422,-68.1355514526367,-67.4337921142578,-66.93798828125,-66.7057571411133,-66.3894500732422,-66.3712921142578,-66.4476623535156,-66.4530792236328,-66.3252868652344,-66.2664566040039,-66.3064422607422,-66.4453659057617,-66.6912078857422,-66.8235397338867,-66.9706573486328,-67.1473617553711,-67.5146484375,-67.6880798339844,-67.9773406982422,-68.1373062133789,-68.2918930053711,-68.5334930419922,-68.6215286254883,-68.7282257080078,-68.8536605834961,-68.9745635986328,-69.0909118652344,-69.1996612548828,-69.3513641357422,-69.9767532348633,-70.1181640625,-70.1263656616211,-70.3183059692383,-70.535400390625,-70.5619125366211,-70.28662109375,-70.0814971923828,-70.1144561767578,-70.4123992919922,-70.613525390625,-70.7287139892578,-70.9935989379883,-71.2716293334961,-71.3898391723633,-71.5124053955078,-71.64990234375,-72.06494140625,-72.1585464477539,-72.2472686767578,-72.5863265991211,-72.79150390625,-72.8180694580078,-72.581298828125,-72.5709457397461,-72.6724624633789,-72.7147979736328,-72.6797332763672,-72.4725112915039,-72.3956069946289,-72.023681640625,-71.6513214111328,-71.515625,-71.394775390625,-70.90576171875,-70.7540969848633,-70.6253890991211,-70.4142074584961,-69.9735336303711,-68.9934539794922,-68.9296417236328,-68.9239273071289,-69.011962890625,-69.030517578125,-68.829833984375,-68.3770523071289,-68.0675277709961,-67.8683624267578,-67.707763671875,-67.4822311401367,-67.3545455932617,-66.583740234375,-66.2427749633789,-66.0753402709961,-65.9678726196289,-65.8255386352539,-65.5597686767578,-65.4198760986328,-65.2877883911133,-65.116943359375,-64.9892578125,-64.9046401977539,-64.8389663696289,-64.7922897338867,-64.6324234008789,-64.4657211303711,-64.1791458129883,-64.2052764892578,-64.3268051147461,-64.43994140625,-64.5446319580078,-64.7352523803711,-64.9822311401367,-65.2221221923828,-65.3949203491211,-65.5534133911133,-65.8104476928711,-65.98193359375,-66.29150390625,-66.4477081298828,-66.8436508178711,-66.95947265625,-67.0606384277344,-67.1413040161133,-67.2014694213867,-67.1931686401367,-67.0506362915039,-66.9958953857422,-66.6100616455078,-66.372314453125,-66.1356887817383,-65.9632797241211,-65.8009796142578,-65.6452178955078,-65.3582000732422,-65.0621566772461,-64.6937942504883,-64.5155258178711,-63.8915481567383,-63.7219696044922,-63.5780296325684,-63.4416999816895,-63.05859375,-63.0286636352539,-63.0954551696777,-63.2352027893066,-63.2124977111816,-62.9932632446289,-62.9033699035645,-62.6719245910645,-62.2988777160645,-62.0494117736816,-61.8603553771973,-61.6355934143066,-61.5190887451172,-61.4359855651855,-61.3169937133789,-61.1620635986328,-61.0999984741211,-61.1307601928711,-61.1759757995605,-61.2356910705566,-61.2029266357422,-61.0150375366211,-60.8428726196289,-60.432373046875,-60.0994644165039,-59.9019050598145,-59.6422843933105,-59.2819366455078,-58.956787109375,-58.4297828674316,-58.0799331665039,-57.7903327941895,-57.5048866271973,-57.0828628540039,-56.862060546875,-56.7306632995605,-56.6151390075684,-56.6581535339355,-56.8596687316895,-57.1684074401855,-57.85302734375,-58.2300796508789,-58.5682106018066,-58.8167457580566,-59.2680168151855,-59.2618141174316,-58.7173805236816,-57.7168960571289,-56.5893516540527,-56.2119598388672,-55.5486831665039,-55.4862289428711,-55.3436012268066,-54.7258796691895,-54.5488739013672,-54.2770500183105,-53.9873046875,-53.8532257080078,-53.6713371276855,-53.58203125,-53.5955085754395,-53.5907707214355,-53.5556640625,-53.4301261901855,-53.2799797058105,-53.14501953125,-53.0412101745605,-52.968505859375,-52.925537109375,-53.1019554138184,-53.1107406616211,-53.0225601196289,-52.7755889892578,-51.7540016174316,-51.3518562316895,-50.8944358825684,-50.3600578308105,-49.8670387268066,-49.6488304138184,-49.5410614013672,-49.6942863464355,-50.3948211669922,-50.713134765625,-50.935546875,-50.9899406433105,-50.8195304870605,-50.037109375,-48.8611831665039,-47.357421875,-46.6173362731934,-45.2910652160645,-44.8909683227539,-44.7294921875,-44.6076164245605,-44.5324249267578,-44.5269050598145,-44.5910148620605,-44.6277351379395,-44.6371116638184,-44.5470695495605,-44.3332023620605,-44.2388687133789,-44.3265609741211,-44.5772476196289,-45.5524406433105,-45.5565414428711,-45.359619140625,-45.0674324035645,-42.6507339477539,-42.2329597473145,-42.0546379089355,-41.9765625,-41.87646484375,-41.3572769165039,-41.36962890625,-41.4344253540039,-44.2392120361328,-44.761962890625,-45.0279312133789,-45.302978515625,-45.8733406066895,-46.1368179321289,-46.4781723022461,-46.1690444946289,-45.9088859558105,-45.4146003723145,-45.1217765808105,-44.6569328308105,-44.1973152160645,-43.194580078125,-43.00927734375,-42.7755393981934,-42.259521484375,-42.0545921325684,-41.8197746276855,-41.6834983825684,-41.52197265625,-41.3001441955566,-40.9793930053711,-40.689453125,-40.3568344116211,-39.8863258361816,-39.5884284973145,-39.3160171508789,-38.9311065673828,-38.2783660888672,-38.15625,-38.0985832214355,-38.0370140075684,-38.014892578125,-37.9347648620605,-37.9927749633789,-38.53955078125,-38.6429176330566,-38.747802734375,-38.7495613098145,-38.6482429504395,-38.5414581298828,-38.1879386901855,-38.0710945129395,-37.9608383178711,-37.8280258178711,-37.7233390808105,-37.4869117736816,-37.1229972839355,-36.8044929504395,-36.6896018981934,-36.672119140625,-36.6444320678711,-36.6064949035645,-35.4518547058105,-35.16552734375,-34.941650390625,-34.6677742004395,-34.4283180236816,-34.1319351196289,-33.8373565673828,-33.3983421325684,-32.9844245910645,-30.7029304504395,-29.9528789520264],[-89.1614761352539,-89.1710891723633,-89.1821746826172,-89.1903762817383,-89.2012634277344,-89.2124557495117,-89.2276382446289,-89.2375030517578,-89.2328186035156,-89.1135711669922,-88.9371566772461,-88.89404296875,-88.8399887084961,-88.79833984375,-88.7086486816406,-88.6033706665039,-88.5362319946289,-88.5715866088867,-88.5979995727539,-88.5939483642578,-88.2283172607422,-88.2714385986328,-88.3645477294922,-88.5333023071289,-88.6844711303711,-88.8299331665039,-88.9609909057617,-88.9764175415039,-89.142578125,-89.1703109741211,-89.2061004638672,-89.2223663330078,-89.1922378540039,-89.1622009277344,-89.1717758178711,-89.2867202758789,-89.3394088745117,-89.3625946044922,-89.3832473754883,-89.4188461303711,-89.5008773803711,-89.54052734375,-89.5736312866211,-89.5769577026367,-89.5550308227539,-89.5471649169922,-89.5702590942383,-89.6712875366211,-89.7111358642578,-89.7493591308594,-89.793701171875,-89.8399429321289,-89.8727111816406,-89.9426803588867,-90.0481414794922,-90.104736328125,-90.1059036254883,-90.09521484375,-90.4791030883789,-90.60693359375,-91.1460494995117,-91.3773422241211,-91.6409225463867,-91.819091796875,-92.2351531982422,-92.2090377807617,-92.1870651245117,-92.159912109375,-92.1764602661133,-92.1863708496094,-92.1556625366211,-92.1585464477539,-92.1442413330078,-92.0987319946289,-92.0747985839844,-92.204345703125,-92.2042465209961,-92.1871566772461,-92.0821304321289,-91.9572296142578,-91.8194351196289,-91.736572265625,-91.4339904785156,-91.2337875366211,-90.9795913696289,-90.7032241821289,-90.52197265625,-90.4471664428711,-90.4598617553711,-90.4501495361328,-90.4169921875,-90.4169921875,-90.4710922241211,-90.5757827758789,-90.6340866088867,-90.6343765258789,-90.6599655151367,-90.710693359375,-90.8160171508789,-90.975830078125,-91.1118698120117,-91.2241668701172,-91.3191909790039,-91.392333984375,-91.4096145629883,-91.1955032348633,-90.9929656982422,-90.9916000366211,-90.9904251098633,-90.9891586303711,-90.6220245361328,-90.18359375,-89.7288131713867,-89.3715362548828,-89.1614761352539],[144.741806030273,144.699508666992,144.662796020508,144.650009155273,144.649322509766,144.790344238281,144.836715698242,144.875381469727,144.90966796875,144.940826416016,144.779891967773,144.741806030273],[-57.1947746276855,-57.2478981018066,-57.2575187683105,-57.2918930053711,-57.3185501098633,-57.2796401977539,-57.2353019714355,-57.2184600830078,-57.2073249816895,-57.2098159790039,-57.2268562316895,-57.269287109375,-57.3095703125,-57.3057594299316,-57.3310012817383,-57.4121589660645,-57.5708999633789,-57.6488265991211,-57.7110824584961,-57.7520027160645,-57.8041038513184,-57.8449211120605,-57.8811073303223,-57.9170379638672,-57.9048805236816,-57.8678703308105,-57.8460006713867,-57.8747062683105,-57.90625,-57.9247093200684,-57.9497566223145,-58.0107421875,-58.0544891357422,-58.0542984008789,-58.0322227478027,-57.90771484375,-57.8665504455566,-57.8326683044434,-57.7203636169434,-57.6494598388672,-57.6561050415039,-57.646728515625,-57.6027336120605,-57.5496101379395,-57.4905776977539,-57.4378890991211,-57.4255867004395,-57.3036613464355,-57.2899436950684,-57.2829132080078,-57.2779273986816,-57.2489738464355,-57.2316398620605,-57.2305679321289,-57.2249984741211,-57.2069320678711,-57.2098159790039,-57.1973648071289,-57.1636238098145,-57.1211433410645,-57.1051292419434,-57.0968742370605,-57.0604476928711,-57.0419425964355,-57.0289535522461,-57.0234870910645,-56.9971199035645,-56.9792976379395,-56.9452133178711,-56.9314956665039,-56.8864250183105,-56.8405265808105,-56.81982421875,-56.7611312866211,-56.704345703125,-56.627197265625,-56.5626945495605,-56.5223617553711,-56.4828109741211,-56.5254859924316,-56.5635719299316,-56.616455078125,-56.6898422241211,-56.7662582397461,-56.8367195129395,-56.9695320129395,-57.0100593566895,-57.03759765625,-57.0926780700684,-57.118896484375,-57.1896018981934,-57.2755851745605,-57.3174781799316,-57.3667984008789,-57.4126968383789,-57.5004386901855,-57.5457496643066,-57.5944366455078,-57.6917457580566,-57.795654296875,-57.8734359741211,-57.9463424682617,-57.9828147888184,-57.9951171875,-58.0117683410645,-58.03466796875,-58.0913047790527,-58.1422386169434,-58.173095703125,-58.2304229736328,-58.2811508178711,-58.314208984375,-58.3406753540039,-58.3626976013184,-58.38037109375,-58.3958015441895,-58.4729499816895,-58.5060539245605,-58.4868621826172,-58.4957008361816,-58.5118675231934,-58.6050796508789,-58.6846160888672,-58.7303237915039,-58.7872047424316,-58.8217811584473,-58.8625030517578,-58.9166030883789,-58.968505859375,-59.1003913879395,-59.231201171875,-59.3169898986816,-59.3372535705566,-59.3776817321777,-59.4794464111328,-59.5356941223145,-59.5966300964355,-59.6665992736816,-59.6637725830078,-59.6685066223145,-59.6985359191895,-59.74072265625,-59.7561988830566,-59.7517585754395,-59.7435073852539,-59.7552223205566,-59.84912109375,-59.8896446228027,-59.9607887268066,-59.9943313598633,-59.9958953857422,-59.9723129272461,-59.9456520080566,-59.8730506896973,-59.8311538696289,-59.8288116455078,-59.8330574035645,-59.8543968200684,-59.7316436767578,-59.6790046691895,-59.6702117919922,-59.6044464111328,-59.5753898620605,-59.5511207580566,-59.5577621459961,-59.58642578125,-59.6202163696289,-59.6912078857422,-59.7168960571289,-59.7385749816895,-59.7274894714355,-59.69970703125,-59.7032699584961,-59.7458038330078,-59.8333511352539,-59.9061012268066,-59.9623527526855,-60.0450210571289,-60.1111373901367,-60.1486320495605,-60.1409149169922,-60.1245574951172,-60.0689468383789,-60.0317878723145,-60.0267562866211,-60.0154800415039,-59.9993667602539,-59.9906730651855,-60.0780792236328,-60.10595703125,-60.1420440673828,-60.1817398071289,-60.2416534423828,-60.335205078125,-60.4087867736816,-60.4595222473145,-60.576416015625,-60.6513710021973,-60.7421340942383,-60.9540023803711,-61.1671867370605,-61.3768043518066,-61.3908195495605,-61.3031272888184,-61.2249488830566,-61.1594734191895,-61.1287117004395,-61.1522941589355,-61.1510238647461,-61.1815910339355,-61.2036094665527,-61.17724609375,-61.1456069946289,-61.1047859191895,-61.007080078125,-60.9379920959473,-60.91357421875,-60.8733367919922,-60.8208465576172,-60.7179222106934,-60.6710472106934,-60.5860862731934,-60.39501953125,-60.3520965576172,-60.3220710754395,-60.3254890441895,-60.3450698852539,-60.3923835754395,-60.4649391174316,-60.523193359375,-60.5832023620605,-60.6333045959473,-60.6361808776855,-60.6065444946289,-60.6237258911133,-60.7192420959473,-60.7186546325684,-60.6494636535645,-60.610107421875,-60.5563468933105,-60.5136260986328,-60.380615234375,-60.3467788696289,-60.2789039611816,-60.1781730651855,-60.0324249267578,-59.9907188415527,-59.9648475646973,-59.8490753173828,-59.8289070129395,-59.8316383361816,-60.0175285339355,-59.9806175231934,-59.8365211486816,-59.7567367553711,-59.7399406433105,-59.7393074035645,-59.6661148071289,-59.4769058227539,-59.2002410888672,-58.8115692138672,-58.7010765075684,-58.6266136169434,-58.5110816955566,-58.4772911071777,-58.4805641174316,-58.5829086303711,-58.6079139709473,-58.6134719848633,-58.6729507446289,-58.593994140625,-58.5694808959961,-58.5022964477539,-58.4149894714355,-58.2984390258789,-58.1728515625,-58.07177734375,-57.9825668334961,-57.7928733825684,-57.6075706481934,-57.5401382446289,-57.3436508178711,-57.2275390625,-57.1902351379395,-57.167236328125,-57.2052726745605,-57.1947746276855],[114.232032775879,114.207221984863,114.138771057129,114.134475708008,114.187400817871,114.246871948242,114.243553161621,114.232032775879],[113.997756958008,113.877349853516,113.8515625,113.8388671875,113.881538391113,114.0439453125,114.003326416016,113.997756958008],[114.015434265137,114.018257141113,114.050392150879,114.097854614258,114.122848510742,114.188179016113,114.228218078613,114.266021728516,114.269630432129,114.291114807129,114.284568786621,114.3251953125,114.335258483887,114.29052734375,114.287887573242,114.267967224121,114.139068603516,114.032814025879,113.937301635742,113.902534484863,113.896484375,114.006736755371,114.015434265137],[73.7074203491211,73.5879898071289,73.4651412963867,73.4132843017578,73.3363265991211,73.2854537963867,73.25390625,73.2511672973633,73.3052749633789,73.3880844116211,73.5857391357422,73.7312469482422,73.8377914428711,73.7951202392578,73.7074203491211],[-83.1575164794922,-83.4150390625,-83.5365295410156,-83.5897445678711,-83.635498046875,-83.6736373901367,-83.7509231567383,-83.8672866821289,-83.9722671508789,-84.0658187866211,-84.0929718017578,-84.1002883911133,-84.1144027709961,-84.1507797241211,-84.1923828125,-84.2392120361328,-84.2639617919922,-84.2666549682617,-84.2919464111328,-84.3397979736328,-84.3936538696289,-84.4535598754883,-84.5376510620117,-84.6459503173828,-84.7297897338867,-84.7891616821289,-84.8604507446289,-84.9851531982422,-85.037353515625,-85.0486373901367,-85.0365219116211,-85.0593719482422,-85.1172866821289,-85.1613235473633,-85.1914978027344,-85.1975631713867,-85.1794967651367,-85.2083511352539,-85.2841796875,-85.373779296875,-85.47705078125,-85.5797805786133,-85.6819381713867,-85.731201171875,-85.7277297973633,-85.7339324951172,-85.75341796875,-85.7867202758789,-85.9837875366211,-86.0403747558594,-86.0892562866211,-86.1512145996094,-86.2382354736328,-86.3317413330078,-86.376953125,-86.6102523803711,-86.733642578125,-86.7589797973633,-86.7706069946289,-86.7635269165039,-86.7295913696289,-86.710693359375,-86.7293014526367,-86.7921371459961,-86.87353515625,-86.918212890625,-86.9288101196289,-86.9331512451172,-86.9588851928711,-87.0093307495117,-87.0591812133789,-87.3372497558594,-87.33251953125,-87.4127960205078,-87.4584503173828,-87.4983901977539,-87.4851531982422,-87.4891052246094,-87.6022415161133,-87.7083511352539,-87.7693862915039,-87.814208984375,-87.7370071411133,-87.7316436767578,-87.7564392089844,-87.7818832397461,-87.7742233276367,-87.758544921875,-87.71533203125,-87.7314453125,-87.80224609375,-87.8919906616211,-87.9910202026367,-88.0387191772461,-88.0804672241211,-88.1510238647461,-88.2762222290039,-88.4084930419922,-88.4491195678711,-88.482666015625,-88.4976577758789,-88.5043487548828,-88.5125503540039,-88.5831527709961,-88.6656188964844,-88.7076110839844,-88.7473678588867,-88.845947265625,-88.8683090209961,-89.0001907348633,-89.02685546875,-89.05712890625,-89.1205062866211,-89.1701126098633,-89.3372573852539,-89.3625946044922,-89.3394088745117,-89.2867202758789,-89.1717758178711,-89.1622009277344,-89.1922378540039,-89.2223663330078,-89.2061004638672,-89.1703109741211,-89.142578125,-88.9764175415039,-88.9609909057617,-88.8299331665039,-88.6844711303711,-88.5333023071289,-88.3645477294922,-88.2714385986328,-88.2283172607422,-88.131103515625,-88.0545883178711,-88.0103988647461,-87.9070358276367,-87.8749542236328,-87.7018585205078,-87.6181640625,-87.5449752807617,-87.4869155883789,-87.3774871826172,-87.285888671875,-86.9072265625,-86.7570266723633,-86.4808120727539,-86.3566436767578,-86.1812057495117,-86.0685501098633,-85.936279296875,-85.9539108276367,-85.9856491088867,-85.7839889526367,-85.4836883544922,-85.1636734008789,-85.0482482910156,-84.9737243652344,-84.6460876464844,-84.5596694946289,-84.4922866821289,-84.4400405883789,-84.4259719848633,-84.4903793334961,-84.5196304321289,-84.2614288330078,-83.7754898071289,-83.7652816772461,-83.9728012084961,-84.082763671875,-84.111328125,-84.1051712036133,-84.0950698852539,-84.0479507446289,-84.01318359375,-83.9274368286133,-83.8706512451172,-83.8016586303711,-83.7604446411133,-83.7159118652344,-83.6721649169922,-83.5896530151367,-83.5359420776367,-83.4979476928711,-83.5510711669922,-83.6761245727539,-83.6463928222656,-83.3691864013672,-83.2908706665039,-83.2255859375,-83.1575164794922],[-86.419921875,-86.5802688598633,-86.6304168701172,-86.5569305419922,-86.4382858276367,-86.3378372192383,-86.2555160522461,-86.419921875],[-85.8709487915039,-85.9472122192383,-85.9609832763672,-85.9242172241211,-85.8782196044922,-85.8337936401367,-85.8444366455078,-85.8709487915039],[17.6078128814697,17.7442398071289,17.3441410064697,17.3895511627197,17.4319343566895,17.6078128814697],[16.6506824493408,16.8355464935303,16.9710941314697,17.0936508178711,17.1698226928711,17.1882820129395,17.08935546875,16.9775390625,16.8506832122803,16.7388668060303,16.6963863372803,16.6663093566895,16.6506824493408],[17.6675777435303,17.740234375,17.8019542694092,17.84130859375,17.9188480377197,18.0445308685303,18.1239261627197,18.3040027618408,18.3465805053711,18.4363288879395,18.4380855560303,18.4766597747803,18.5174789428711,18.3330078125,18.16064453125,17.8238277435303,17.5849609375,17.2582035064697,17.04541015625,17.12646484375,17.2198238372803,17.7236328125,17.6675777435303],[17.1940441131592,17.1241207122803,16.67919921875,16.5498046875,16.4058609008789,16.37646484375,16.5213871002197,16.6559581756592,16.697265625,17.0611324310303,17.1940441131592],[16.7852554321289,16.62744140625,16.4903316497803,16.4231452941895,16.4281253814697,16.4489269256592,16.6015625,16.8343734741211,16.8913097381592,16.8736324310303,16.7852554321289],[15.3713865280151,15.4372072219849,15.3742198944092,15.30859375,15.27001953125,15.3713865280151],[15.231053352356,15.2466793060303,15.1218748092651,15.0746097564697,15.0658197402954,15.231053352356],[15.1887693405151,15.2030277252197,15.20166015625,15.1498050689697,15.1358394622803,14.8913078308105,14.8650388717651,14.9525394439697,15.1887693405151],[15.1884756088257,15.1625986099243,15.0979490280151,15.03857421875,14.99609375,14.9127922058105,14.8846673965454,14.7604484558105,14.7418947219849,14.8038082122803,14.8553714752197,14.8980464935303,15.0064449310303,15.1129884719849,15.2399415969849,15.2135734558105,15.1884756088257],[14.8314447402954,14.8566398620605,14.7624998092651,14.6782217025757,14.6603507995605,14.6724605560303,14.6905279159546,14.7541990280151,14.7637701034546,14.8314447402954],[14.4880867004395,14.4803714752197,14.4195318222046,14.3888673782349,14.3124017715454,14.3025388717651,14.3421869277954,14.3400382995605,14.2858400344849,14.3312501907349,14.3582029342651,14.369140625,14.3937492370605,14.4673824310303,14.4525384902954,14.4675788879395,14.4825201034546,14.4880867004395],[14.8102540969849,14.68701171875,14.6283206939697,14.6129884719849,14.51171875,14.4503908157349,14.4378910064697,14.5246086120605,14.5710935592651,14.6299800872803,14.7011709213257,14.7391605377197,14.8102540969849],[18.9053707122803,18.9010753631592,18.8935546875,18.8390617370605,18.89453125,18.947265625,18.9178714752197,18.9537105560303,19.0550785064697,19.0642585754395,19.0333003997803,19.0076179504395,19.0046863555908,19.0930671691895,19.2728519439697,19.3302745819092,19.3522472381592,19.3823223114014,19.3999996185303,19.4009761810303,19.3880863189697,19.3030261993408,19.2059574127197,19.1369152069092,19.1307621002197,19.1296882629395,19.0628910064697,19.1000003814697,19.0852546691895,19.060546875,19.03759765625,19.0095710754395,18.9955081939697,19.0071296691895,18.9413089752197,18.83642578125,18.7883777618408,18.7801761627197,18.7793941497803,18.74609375,18.66259765625,18.48828125,18.4239273071289,18.3576164245605,18.2849617004395,18.2179679870605,18.13720703125,17.9962902069092,17.9486331939697,17.8744144439697,17.8127937316895,17.6901378631592,17.6535148620605,17.5462894439697,17.5026359558105,17.4691410064697,17.3241214752197,17.2586917877197,17.2106437683105,17.1253890991211,16.9186515808105,16.7908191680908,16.5306644439697,16.4535160064697,16.3650398254395,16.2933597564697,16.2310543060303,16.1573238372803,16.0283203125,15.9631834030151,15.8882818222046,15.8228521347046,15.7880868911743,15.7615242004395,15.7379884719849,15.7366209030151,15.8800792694092,16.0490226745605,16.1034183502197,16.1302738189697,16.1698246002197,16.2142581939697,16.3000984191895,16.3775386810303,16.4720706939697,16.5905284881592,16.6876964569092,16.7134761810303,16.90185546875,17.0845699310303,17.248046875,17.2738285064697,17.2752933502197,17.2930679321289,17.4022464752197,17.6248054504395,17.6504878997803,17.6578140258789,17.6434574127197,17.5851573944092,17.5373039245605,17.3298816680908,17.12939453125,16.9031257629395,16.6002941131592,16.3939437866211,16.2689456939697,16.1310539245605,16.0459957122803,15.9855470657349,15.9425783157349,15.9491214752197,15.9415044784546,15.8206052780151,15.6556644439697,15.4994134902954,15.1858396530151,15.1229486465454,15.1846685409546,15.2313470840454,15.284276008606,15.3697271347046,15.4709968566895,15.3813467025757,15.2698240280151,14.9813470840454,14.8952150344849,14.8852529525757,14.9065427780151,14.8545894622803,14.6320304870605,14.5504884719849,14.3861322402954,14.3126955032349,14.2685546875,14.236328125,14.0906248092651,14.0419921875,13.9658203125,13.8998050689697,13.8607416152954,13.7424802780151,13.6292972564697,13.6134767532349,13.6033210754395,13.5171880722046,13.5779294967651,13.615234375,13.8787107467651,13.9356441497803,13.9701175689697,13.9703130722046,13.9927740097046,14.0855464935303,14.1612310409546,14.2830076217651,14.3699226379395,14.4273433685303,14.5051755905151,14.5339851379395,14.5688486099243,14.5917978286743,14.6085929870605,14.6495113372803,14.7335939407349,14.7930660247803,14.8470706939697,14.8999996185303,14.95458984375,15.1104497909546,15.2420892715454,15.3394527435303,15.3266592025757,15.2912111282349,15.2835931777954,15.2901363372803,15.3569345474243,15.3537101745605,15.2729482650757,15.2770519256592,15.4541025161743,15.6248054504395,15.6521482467651,15.6680660247803,15.6755867004395,15.6662101745605,15.5968751907349,15.5925788879395,15.6089849472046,15.6359367370605,15.7041997909546,15.7842769622803,15.8475589752197,15.9333009719849,16.0006828308105,16.0665035247803,16.1064453125,16.2274417877197,16.2533187866211,16.2367191314697,16.2583980560303,16.3011722564697,16.3211917877197,16.4276371002197,16.5162105560303,16.5699214935303,16.748046875,16.8714847564697,16.93994140625,17.03271484375,17.1496086120605,17.2421875,17.3106441497803,17.4063472747803,17.5291996002197,17.6070308685303,17.6396484375,17.7064456939697,17.80712890625,17.9638671875,18.2639656066895,18.2906246185303,18.3583011627197,18.4373054504395,18.5335941314697,18.5646495819092,18.666015625,18.7217769622803,18.8330078125,18.9002933502197,18.9053707122803],[-72.8045883178711,-72.8222122192383,-73.0779800415039,-73.2852554321289,-73.2764129638672,-73.1706085205078,-73.0691375732422,-72.9192352294922,-72.8045883178711],[-71.7792510986328,-71.7574234008789,-71.7114715576172,-71.7069320678711,-71.753173828125,-71.7464828491211,-71.647216796875,-71.6453094482422,-71.6570281982422,-71.7420425415039,-71.80712890625,-71.786376953125,-71.733642578125,-71.72705078125,-71.7432174682617,-71.82421875,-71.8665084838867,-71.9868698120117,-72.0003890991211,-71.9403839111328,-71.87255859375,-71.7619171142578,-71.7372589111328,-71.7637634277344,-71.768310546875,-71.8529281616211,-71.9460906982422,-72.0020523071289,-72.0598602294922,-72.5035629272461,-72.55322265625,-72.5918960571289,-72.63330078125,-72.7552719116211,-72.8766632080078,-73.1600570678711,-73.2722702026367,-73.3851547241211,-73.5148391723633,-73.64404296875,-73.747314453125,-73.82470703125,-73.8391571044922,-73.8849639892578,-73.9894485473633,-74.0854034423828,-74.1946334838867,-74.4190444946289,-74.4599609375,-74.4781265258789,-74.3875045776367,-74.2844772338867,-74.2277374267578,-74.100341796875,-73.9759750366211,-73.8624954223633,-73.68701171875,-73.5915985107422,-72.9172821044922,-72.7893524169922,-72.7394485473633,-72.6959991455078,-72.6597671508789,-72.6180648803711,-72.4181137084961,-72.3760757446289,-72.3467330932617,-72.34765625,-72.4652328491211,-72.6491241455078,-72.8110885620117,-72.7412109375,-72.7679672241211,-72.7417984008789,-72.7032241821289,-72.8634262084961,-73.052734375,-73.3155212402344,-73.3963394165039,-73.4383773803711,-73.4005355834961,-73.3153305053711,-73.2177734375,-73.1177749633789,-72.8765106201172,-72.6370162963867,-72.429931640625,-72.2198257446289,-71.9542922973633,-71.834716796875,-71.7792510986328],[-72.6640625,-72.6234893798828,-72.6388702392578,-72.7397918701172,-72.84423828125,-72.87841796875,-72.8993225097656,-72.9603576660156,-72.90673828125,-72.8514633178711,-72.791015625,-72.6640625],[22.1318359375,22.2271480560303,22.2311515808105,22.2537117004395,22.2694339752197,22.2721672058105,22.2951164245605,22.3166980743408,22.3501949310303,22.423828125,22.5201168060303,22.5824222564697,22.6763668060303,22.68310546875,22.7015628814697,22.7691402435303,22.7822265625,22.8362312316895,22.8572254180908,22.8464832305908,22.8560543060303,22.8766593933105,22.8517570495605,22.6767578125,22.6083984375,22.5628910064697,22.4914073944092,22.41748046875,22.3514652252197,22.2906246185303,22.24462890625,22.18505859375,22.1119136810303,22.0379886627197,21.9997062683105,21.9953136444092,21.9542980194092,21.8992195129395,21.8693370819092,21.7854499816895,21.7217769622803,21.6614265441895,21.6514644622803,21.6526374816895,21.5841789245605,21.4944343566895,21.47705078125,21.4970703125,21.4110336303711,21.361328125,21.3202152252197,21.2945308685303,21.2522468566895,21.2632827758789,21.2645511627197,21.1917972564697,21.17041015625,21.1519527435303,21.1216793060303,21.0398445129395,20.8370113372803,20.76025390625,20.7374019622803,20.7327156066895,20.7074222564697,20.6610355377197,20.6136722564697,20.5081062316895,20.2809581756592,20.2417964935303,20.2101554870605,20.1614265441895,19.93408203125,19.8444328308105,19.7245121002197,19.6134757995605,19.53076171875,19.45751953125,19.4212894439697,19.3928699493408,19.3302745819092,19.2781257629395,19.2083988189697,19.1462898254395,19.0873050689697,19.0662117004395,19.0476551055908,19.0157222747803,18.9278316497803,18.9053707122803,18.9002933502197,18.8330078125,18.7217769622803,18.666015625,18.5646495819092,18.5335941314697,18.4373054504395,18.3583011627197,18.2906246185303,18.2639656066895,17.9638671875,17.80712890625,17.7064456939697,17.6396484375,17.6070308685303,17.5291996002197,17.4063472747803,17.3106441497803,17.2421875,17.1496086120605,17.03271484375,16.93994140625,16.8714847564697,16.748046875,16.5699214935303,16.5162105560303,16.5056648254395,16.41845703125,16.3812503814697,16.3845710754395,16.3671875,16.33544921875,16.3184566497803,16.3084964752197,16.2835922241211,16.0930671691895,16.2525386810303,16.3318367004395,16.4239253997803,16.4534187316895,16.4612312316895,16.4769535064697,16.4847660064697,16.49267578125,16.4828128814697,16.4383792877197,16.4168949127197,16.4397468566895,16.4625988006592,16.4343738555908,16.44287109375,16.5147457122803,16.5744132995605,16.623046875,16.6366214752197,16.6765632629395,16.6397457122803,16.43212890625,16.4212894439697,16.4696273803711,16.5210933685303,16.5509777069092,16.5909175872803,16.6474609375,16.74755859375,16.7859382629395,16.8230457305908,16.8626956939697,16.9734363555908,17.0666027069092,17.0456066131592,17.0458984375,17.0300769805908,17.0399398803711,17.0777339935303,17.0890636444092,17.1473636627197,17.1746101379395,17.2772445678711,17.3015632629395,17.3172855377197,17.4806632995605,17.63525390625,17.7619132995605,17.9479484558105,18.1456050872803,18.4762687683105,18.7242183685303,18.7406253814697,18.7780265808105,18.7483406066895,18.7500972747803,18.7918949127197,18.9141597747803,19.26513671875,19.4669914245605,19.4974613189697,19.5642566680908,19.6253910064697,19.7091808319092,19.81005859375,19.8986320495605,19.9503898620605,20.1286144256592,20.3337898254395,20.4750003814697,20.4900398254395,20.6431636810303,20.8666019439697,20.9811534881592,21.0672855377197,21.1963882446289,21.3824214935303,21.4513664245605,21.5046863555908,21.5631847381592,21.6026363372803,21.6325187683105,21.6486320495605,21.6746101379395,21.7214832305908,21.7669906616211,22.111328125,22.1318359375],[122.948921203613,122.855857849121,122.826171875,122.818450927734,122.845703125,123.061431884766,123.145797729492,123.265434265137,123.339653015137,123.358505249023,123.371101379395,123.383102416992,123.412887573242,123.418159484863,123.310745239258,123.214851379395,123.005271911621,122.948921203613],[121.883003234863,121.833099365234,121.726173400879,121.704681396484,121.796287536621,121.866996765137,121.949516296387,121.998336791992,121.981346130371,121.883003234863],[123.416206359863,123.325981140137,123.325584411621,123.395317077637,123.458786010742,123.493942260742,123.496772766113,123.405082702637,123.416206359863],[120.012504577637,120.057624816895,120.221099853516,120.248046875,120.258308410645,120.291122436523,120.36474609375,120.443649291992,120.503715515137,120.555572509766,120.632614135742,120.700386047363,120.784477233887,120.832618713379,120.804206848145,120.698051452637,120.640426635742,120.561721801758,120.439155578613,120.39453125,120.255462646484,120.144828796387,120.051956176758,119.998435974121,119.9306640625,119.812789916992,119.601081848145,119.470306396484,119.41650390625,119.362594604492,119.085456848145,119.042381286621,119.008399963379,118.977348327637,118.958793640137,118.994132995605,119.031440734863,119.185638427734,119.295890808105,119.423934936523,119.614738464355,119.795112609863,119.850776672363,119.942085266113,119.973831176758,120.012504577637],[125.068168640137,124.997955322266,124.963088989258,124.841804504395,124.708396911621,124.601860046387,124.508201599121,124.427536010742,124.326766967773,124.175979614258,123.971092224121,123.857612609863,123.747261047363,123.644142150879,123.604789733887,123.614059448242,123.648239135742,123.690139770508,123.716407775879,123.599418640137,123.589256286621,123.6357421875,123.665817260742,123.709373474121,123.876762390137,123.977157592773,124.036331176758,124.052436828613,124.090141296387,124.115531921387,124.134574890137,124.282325744629,124.3193359375,124.375679016113,124.412986755371,124.438285827637,124.444435119629,124.575492858887,124.645896911621,124.708206176758,124.889747619629,124.922264099121,124.915031433105,124.936805725098,124.973243713379,125.100395202637,125.124404907227,125.149024963379,125.149421691895,125.100486755371,124.977546691895,124.960151672363,124.958595275879,124.968254089355,124.996971130371,125.033592224121,125.068168640137],[115.609970092773,115.581939697266,115.500877380371,115.48046875,115.540626525879,115.561431884766,115.613288879395,115.609970092773],[122.977340698242,122.945510864258,122.887794494629,122.903518676758,122.932815551758,123.010543823242,123.089447021484,123.137886047363,123.153121948242,123.030082702637,122.977340698242],[119.464065551758,119.424903869629,119.385543823242,119.401664733887,119.37890625,119.419914245605,119.43017578125,119.446487426758,119.470512390137,119.481742858887,119.502151489258,119.546974182129,119.557228088379,119.555465698242,119.536331176758,119.482810974121,119.444038391113,119.464065551758],[123.317481994629,123.297264099121,123.025001525879,123.032623291016,123.108299255371,123.133499145508,123.217094421387,123.336029052734,123.317481994629],[116.640823364258,116.514259338379,116.559379577637,116.586517333984,116.377243041992,116.289848327637,116.239356994629,116.026756286621,115.874610900879,115.857330322266,115.869338989258,115.914459228516,116.031639099121,116.076469421387,116.077743530273,116.061126708984,116.219818115234,116.304298400879,116.401557922363,116.646965026855,116.6875,116.718940734863,116.734077453613,116.640823364258],[124.286613464355,124.225784301758,124.184371948242,124.146682739258,124.065727233887,124.017288208008,123.927734375,123.971481323242,124.013763427734,124.068748474121,124.095794677734,124.110549926758,124.239555358887,124.265617370605,124.287109375,124.304496765137,124.286613464355],[123.9248046875,123.783889770508,123.697853088379,123.629196166992,123.591606140137,123.582618713379,123.587890625,123.580169677734,123.553024291992,123.488670349121,123.433784484863,123.410743713379,123.329978942871,123.253318786621,123.230079650879,123.324996948242,123.45458984375,123.475875854492,123.425193786621,123.394920349121,123.391204833984,123.473236083984,123.529983520508,123.573150634766,123.600593566895,123.775978088379,123.845512390137,123.896095275879,123.9248046875],[138.895126342773,138.845504760742,138.59423828125,138.567184448242,138.563385009766,138.621002197266,138.676666259766,138.7626953125,138.796188354492,138.897659301758,138.895126342773],[117.556350708008,117.533592224121,117.490425109863,117.505958557129,117.482124328613,117.490524291992,117.546096801758,117.665031433105,117.669242858887,117.556350708008],[119.073829650879,119.029983520508,119.020896911621,119.03662109375,119.078704833984,119.097747802734,119.128318786621,119.134864807129,119.106735229492,119.073829650879],[124.575584411621,124.599601745605,124.676849365234,124.752243041992,124.924118041992,125.050285339355,125.124610900879,125.131744384766,125.096771240234,124.444244384766,124.380668640137,124.355560302734,124.425979614258,124.393547058105,124.4306640625,124.508598327637,124.575584411621],[127.823440551758,127.998443603516,128.098831176758,128.119247436523,128.023529052734,127.820899963379,127.786224365234,127.823440551758],[122.782905578613,122.641494750977,122.553810119629,122.470207214355,122.417289733887,122.321487426758,122.185745239258,122.094139099121,121.838676452637,121.738288879395,121.6513671875,121.621284484863,121.584564208984,121.499610900879,121.414642333984,121.328323364258,121.190818786621,121.137504577637,121.086128234863,121.035255432129,120.981826782227,120.780952453613,120.550491333008,120.319534301758,120.120895385742,120.012107849121,119.909378051758,119.879096984863,119.841407775879,119.807914733887,119.807037353516,119.818168640137,119.847663879395,119.866119384766,119.874809265137,119.918258666992,119.963775634766,120.099220275879,120.231147766113,120.354110717773,120.424896240234,120.485549926758,120.547172546387,120.610252380371,120.709564208984,120.751365661621,120.886138916016,121.008682250977,121.118156433105,121.276664733887,121.371971130371,121.444534301758,121.498435974121,121.547943115234,121.610359191895,121.683395385742,121.7470703125,121.862899780273,121.911720275879,121.966506958008,122.020111083984,122.067092895508,122.263092041016,122.323234558105,122.433486938477,122.466606140137,122.483596801758,122.513771057129,122.555862426758,122.603523254395,122.75,122.850486755371,122.919143676758,122.758590698242,122.792388916016,122.845703125,122.916984558105,122.978317260742,123.005958557129,122.955467224121,122.923637390137,122.902153015137,122.811134338379,122.846771240234,122.820022583008,122.782905578613],[130.862197875977,130.775207519531,130.833389282227,131.020111083984,131.08740234375,131.176361083984,131.043746948242,130.908096313477,130.862197875977],[118.242378234863,118.292388916016,118.337890625,118.433197021484,118.490623474121,118.552154541016,118.611915588379,118.670600891113,118.691802978516,118.7138671875,118.748344421387,118.794242858887,118.845703125,118.926177978516,118.98779296875,119.043846130371,119.042098999023,119.062507629395,119.101066589355,119.129684448242,119.104194641113,119.078910827637,119.006248474121,118.971488952637,118.939353942871,118.9033203125,118.821197509766,118.745895385742,118.756256103516,118.818069458008,118.836715698242,118.832611083984,118.808296203613,118.727935791016,118.673637390137,118.478622436523,118.426956176758,118.397842407227,118.399909973145,118.378913879395,118.233985900879,118.189933776855,118.131538391113,118.070709228516,117.861228942871,117.79541015625,117.731643676758,117.507911682129,117.387893676758,117.326370239258,117.265045166016,117.210250854492,117.161224365234,117.061325073242,116.958206176758,116.87109375,116.788475036621,116.767974853516,116.772071838379,116.806930541992,116.783103942871,116.801277160645,116.835060119629,116.88623046875,116.953125,117.063674926758,117.164848327637,117.223625183105,117.356636047363,117.4345703125,117.567092895508,117.621772766113,117.643356323242,117.6728515625,117.712104797363,117.806053161621,117.893165588379,117.969528198242,118.104103088379,118.205955505371,118.23486328125,118.174018859863,118.100486755371,118.06103515625,118.017868041992,117.979103088379,117.814849853516,117.766410827637,117.738380432129,117.755271911621,117.868263244629,117.920989990234,118.117477416992,118.150688171387,118.202827453613,118.242378234863],[115.447845458984,115.549407958984,115.69091796875,115.704292297363,115.661422729492,115.559959411621,115.333793640137,115.295013427734,115.247161865234,115.236137390137,115.220207214355,115.194244384766,115.144927978516,115.091499328613,115.139747619629,115.141609191895,115.105667114258,115.055084228516,114.952056884766,114.842086791992,114.731346130371,114.613182067871,114.570899963379,114.501754760742,114.478912353516,114.467575073242,114.475296020508,114.504295349121,114.620010375977,114.8330078125,114.9384765625,114.998146057129,115.153999328613,115.191017150879,115.340232849121,115.447845458984],[129.8388671875,129.77978515625,129.713470458984,129.591903686523,129.598724365234,129.608993530273,129.655471801758,129.81298828125,129.843551635742,129.8388671875],[126.800971984863,126.814453125,126.812690734863,126.692863464355,126.577339172363,126.518165588379,126.472068786621,126.312889099121,126.171089172363,126.108390808105,126.040031433105,125.951553344727,125.826171875,125.798248291016,125.808395385742,125.843162536621,125.975296020508,126.085350036621,126.213668823242,126.359375,126.462890625,126.609565734863,126.726371765137,126.800971984863],[127.41943359375,127.355270385742,127.374992370605,127.370704650879,127.475196838379,127.474021911621,127.463966369629,127.41943359375],[138.535354614258,138.296295166016,137.982818603516,137.871871948242,137.687698364258,137.650390625,137.685150146484,137.83251953125,138.007522583008,138.081832885742,138.185348510742,138.295516967773,138.543838500977,138.769821166992,138.801956176758,138.8994140625,138.962600708008,138.989059448242,138.892959594727,138.785934448242,138.611724853516,138.535354614258],[131.325592041016,131.309188842773,131.184951782227,131.11376953125,131.123428344727,131.086822509766,131.136825561523,131.137786865234,131.190032958984,131.197357177734,131.260055541992,131.296875,131.349227905273,131.411041259766,131.446197509766,131.482620239258,131.535247802734,131.530868530273,131.560745239258,131.643447875977,131.700790405273,131.736129760742,131.643844604492,131.691116333008,131.624404907227,131.580276489258,131.498428344727,131.473526000977,131.377044677734,131.347747802734,131.343460083008,131.325592041016],[131.982025146484,131.969528198242,131.926849365234,131.884475708008,131.822860717773,131.777542114258,131.750778198242,131.922256469727,131.982025146484],[114.41259765625,114.397659301758,114.346878051758,114.298828125,114.322166442871,114.348922729492,114.383590698242,114.41259765625],[128.670120239258,128.625,128.550201416016,128.52978515625,128.577346801758,128.627731323242,128.658309936523,128.673233032227,128.666900634766,128.670120239258],[120.774406433105,120.672370910645,120.640823364258,120.633392333984,120.745506286621,120.781730651855,120.774406433105],[113.844535827637,113.825584411621,113.655860900879,113.54638671875,113.470703125,113.198440551758,113.166015625,113.141891479492,113.126953125,113.040428161621,112.763771057129,112.725875854492,112.768753051758,112.868064880371,113.0673828125,113.974700927734,114.073631286621,114.0830078125,113.885353088379,113.844535827637],[115.377044677734,115.295806884766,115.220321655273,115.222160339355,115.240524291992,115.353713989258,115.414443969727,115.479194641113,115.524223327637,115.546089172363,115.424118041992,115.377044677734],[134.674423217773,134.657424926758,134.631439208984,134.629104614258,134.663482666016,134.697662353516,134.735733032227,134.72607421875,134.674423217773],[105.252830505371,105.1904296875,105.142768859863,105.121383666992,105.192283630371,105.225685119629,105.260543823242,105.277442932129,105.252830505371],[134.716110229492,134.660842895508,134.633697509766,134.679107666016,134.728515625,134.716110229492],[134.819534301758,134.795120239258,134.795318603516,134.822952270508,134.851852416992,134.885833740234,134.819534301758],[134.536819458008,134.520416259766,134.504287719727,134.412490844727,134.35595703125,134.32275390625,134.199996948242,134.0908203125,134.059188842773,134.107040405273,134.154205322266,134.184768676758,134.194625854492,134.124618530273,134.111221313477,134.114639282227,134.168075561523,134.234191894531,134.317779541016,134.4150390625,134.536819458008],[107.373924255371,107.474708557129,107.56298828125,107.666801452637,107.776077270508,107.8837890625,108.0087890625,108.137596130371,108.197463989258,108.254486083984,108.295021057129,108.330177307129,108.419136047363,108.515922546387,108.537986755371,108.603614807129,108.677833557129,108.779685974121,108.8994140625,109.018356323242,109.294235229492,109.403709411621,109.500587463379,109.5869140625,109.820991516113,109.936233520508,110.067092895508,110.198432922363,110.260940551758,110.321090698242,110.372756958008,110.42626953125,110.520896911621,110.583587646484,110.63427734375,110.674018859863,110.700782775879,110.736915588379,110.7841796875,110.834762573242,110.972259521484,111.000679016113,111.154396057129,111.181549072266,111.342086791992,111.386528015137,111.484474182129,111.540328979492,111.6435546875,111.688087463379,111.737602233887,111.989845275879,112.087303161621,112.13671875,112.312301635742,112.43359375,112.539260864258,112.5869140625,112.6259765625,112.648735046387,112.751953125,112.794334411621,112.782913208008,112.794532775879,113.013572692871,113.248435974121,113.497657775879,113.747467041016,113.876266479492,114.037300109863,114.070701599121,114.382713317871,114.409271240234,114.444236755371,114.443260192871,114.384956359863,114.38134765625,114.386909484863,114.448829650879,114.481735229492,114.595016479492,114.599220275879,114.583786010742,114.459175109863,114.383201599121,114.339256286621,114.276954650879,114.15966796875,113.940338134766,113.692573547363,113.253318786621,113.133689880371,113.018943786621,112.897750854492,112.771682739258,112.678802490234,112.586036682129,112.3515625,112.115135192871,111.509963989258,111.338569641113,111.055374145508,110.830177307129,110.607231140137,110.038673400879,109.852638244629,109.281639099121,109.193550109863,108.986717224121,108.856246948242,108.7412109375,108.570510864258,108.517967224121,108.451759338379,108.335548400879,108.220512390137,107.91748046875,107.804389953613,107.69580078125,107.597846984863,107.546875,107.284965515137,107.071189880371,106.631446838379,106.535354614258,106.455276489258,106.411331176758,106.416893005371,106.448440551758,106.491500854492,106.519721984863,106.1982421875,105.9443359375,105.834762573242,105.724807739258,105.600975036621,105.478416442871,105.420799255371,105.361915588379,105.302932739258,105.255470275879,105.2431640625,105.2734375,105.335639953613,105.370903015137,105.387008666992,105.404685974121,105.459770202637,105.483695983887,105.580856323242,105.608009338379,105.655075073242,105.7060546875,105.757423400879,105.786911010742,105.868263244629,105.936134338379,106.02880859375,106.075004577637,106.165817260742,106.349708557129,106.459083557129,106.568748474121,106.675880432129,106.8251953125,106.8779296875,106.931640625,107.011619567871,107.046287536621,107.162109375,107.331832885742,107.373924255371],[124.051277160645,124.042098999023,124.005661010742,123.972267150879,123.975791931152,124.022941589355,124.051277160645],[120.5283203125,120.487312316895,120.467964172363,120.460739135742,120.435554504395,120.451560974121,120.446479797363,120.477340698242,120.5341796875,120.549217224121,120.5283203125],[112.719429016113,112.697952270508,112.602149963379,112.586036682129,112.648536682129,112.690040588379,112.727340698242,112.719429016113],[132.80712890625,132.746276855469,132.704879760742,132.681457519531,132.667282104492,132.681335449219,132.630172729492,132.697860717773,132.716506958008,132.73779296875,132.804306030273,132.80712890625],[134.746963500977,134.739059448242,134.738372802734,134.754974365234,134.712203979492,134.752151489258,134.758102416992,134.755859375,134.744430541992,134.714157104492,134.683883666992,134.6611328125,134.637603759766,134.441116333008,134.356140136719,134.280471801758,134.264450073242,134.175384521484,134.154891967773,134.153121948242,134.22509765625,134.301956176758,134.298629760742,134.343078613281,134.226165771484,134.205383300781,134.247268676758,134.34130859375,134.456344604492,134.490325927734,134.506454467773,134.57080078125,134.616500854492,134.646087646484,134.657806396484,134.6455078125,134.700790405273,134.746963500977],[132.92626953125,132.845016479492,132.921676635742,132.937698364258,133.0087890625,133.114654541016,133.138473510742,133.1728515625,133.11962890625,132.971099853516,132.92626953125],[102.3671875,102.285934448242,102.135551452637,102.110740661621,102.153518676758,102.198440551758,102.371780395508,102.405471801758,102.3671875],[123.626754760742,123.622749328613,123.582618713379,123.550102233887,123.540924072266,123.542770385742,123.560646057129,123.626754760742],[122.04296875,121.979583740234,121.859375,121.808502197266,121.820701599121,121.856643676758,121.873725891113,121.866310119629,121.913673400879,121.965721130371,121.999893188477,122.041015625,122.061820983887,122.04296875],[122.645118713379,122.619338989258,122.563865661621,122.519729614258,122.473640441895,122.391990661621,122.371292114258,122.307037353516,122.283103942871,122.329002380371,122.396293640137,122.390037536621,122.334465026855,122.368942260742,122.5244140625,122.659965515137,122.701957702637,122.73974609375,122.759864807129,122.614067077637,122.645118713379],[123.179786682129,123.203033447266,123.195701599121,123.139457702637,123.119239807129,123.103805541992,123.083885192871,123.05517578125,123.017959594727,123.014640808105,122.986518859863,122.9716796875,122.981056213379,123.024604797363,123.051460266113,123.149909973145,123.201957702637,123.187309265137,123.120704650879,123.043357849121,122.985740661621,122.968742370605,122.934661865234,122.908790588379,122.916206359863,122.850196838379,122.812118530273,122.733100891113,122.684379577637,122.64501953125,122.584953308105,122.586433410645,122.642196655273,122.642578125,122.670112609863,122.731452941895,122.766502380371,122.767585754395,122.793647766113,122.803810119629,122.821487426758,122.849411010742,122.853317260742,122.946876525879,123.038276672363,123.074615478516,123.068946838379,123.179786682129],[133.57080078125,133.621871948242,133.5029296875,133.3330078125,133.320892333984,133.46435546875,133.57080078125],[123.242378234863,123.144523620605,123.076171875,122.994728088379,122.970901489258,122.969047546387,123.02490234375,123.2119140625,123.246978759766,123.242378234863],[128.562606811523,128.3916015625,128.428314208984,128.451553344727,128.536331176758,128.562606811523],[128.275588989258,128.249908447266,128.191787719727,128.143157958984,128.158981323242,128.146881103516,128.11083984375,128.05224609375,127.978034973145,127.934371948242,127.925003051758,127.927543640137,128.016220092773,128.119140625,128.264358520508,128.3291015625,128.313674926758,128.291015625,128.277435302734,128.275588989258],[128.722457885742,128.720123291016,128.713272094727,128.658782958984,128.619522094727,128.585159301758,128.594924926758,128.66650390625,128.693542480469,128.722457885742],[116.424125671387,116.387786865234,116.326560974121,116.395317077637,116.426956176758,116.424125671387],[127.606254577637,127.629295349121,127.531051635742,127.487693786621,127.530464172363,127.554496765137,127.606254577637],[116.303314208984,116.093353271484,116.058792114258,116.076950073242,116.018356323242,116.022453308105,116.063575744629,116.117385864258,116.239356994629,116.269721984863,116.262115478516,116.286521911621,116.295120239258,116.282028198242,116.305168151855,116.318656921387,116.289260864258,116.303314208984],[126.861129760742,127.025482177734,127.062896728516,127.092376708984,127.124710083008,127.163475036621,127.227340698242,127.244239807129,127.229591369629,127.155174255371,127.085052490234,126.940925598145,126.869918823242,126.794143676758,126.740333557129,126.686325073242,126.546684265137,126.4111328125,126.214546203613,126.178321838379,126.146682739258,126.056533813477,126.033981323242,126.026473999023,126.050102233887,126.088287353516,126.219627380371,126.306251525879,126.555068969727,126.808296203613,126.861129760742],[127.987892150879,127.937690734863,127.849609375,127.834274291992,127.938377380371,127.987892150879],[106.886421203613,106.869728088379,106.814254760742,106.774322509766,106.749214172363,106.742874145508,106.796875,106.91064453125,106.886421203613],[107.473342895508,107.432807922363,107.409278869629,107.402442932129,107.419334411621,107.474411010742,107.499702453613,107.473342895508],[129.754684448242,129.984375,130.103424072266,130.303619384766,130.379104614258,130.569931030273,130.625595092773,130.641693115234,130.671096801758,130.718078613281,130.7734375,130.845611572266,130.859970092773,130.805068969727,130.580383300781,130.363082885742,130.269729614258,130.01953125,129.981155395508,129.953125,129.844131469727,129.626663208008,129.545013427734,129.51171875,129.520416259766,129.521682739258,129.467666625977,129.332809448242,129.212112426758,129.107620239258,128.967483520508,128.952041625977,128.964065551758,128.957809448242,128.925399780273,128.862503051758,128.8017578125,128.751266479492,128.676956176758,128.638961791992,128.5166015625,128.465911865234,128.419235229492,128.279983520508,128.233001708984,128.1806640625,128.132019042969,128.082122802734,128.055755615234,128.0439453125,128.030075073242,127.970016479492,127.92041015625,127.90234375,127.927833557129,127.927925109863,127.897163391113,127.8779296875,128.113372802734,128.198532104492,128.56982421875,128.79052734375,128.910736083984,128.991119384766,129.057708740234,129.074325561523,129.116317749023,129.174407958984,129.279586791992,129.37109375,129.427337646484,129.484191894531,129.54296875,129.600494384766,129.754684448242],[100.425094604492,100.465133666992,100.346092224121,100.348434448242,100.33203125,100.259963989258,100.204299926758,100.179298400879,100.198532104492,100.24560546875,100.45458984375,100.468841552734,100.464256286621,100.433891296387,100.425094604492],[108.207221984863,108.191795349121,108.167289733887,108.083587646484,108.055274963379,107.977149963379,107.96728515625,107.941116333008,107.858200073242,107.836616516113,107.82177734375,107.659576416016,107.614456176758,107.63671875,107.594924926758,107.591598510742,107.583885192871,107.5634765625,107.604881286621,107.59814453125,107.6416015625,107.666305541992,107.837791442871,107.874710083008,108.074409484863,108.215141296387,108.290626525879,108.207221984863],[100.2041015625,100.132720947266,100.014938354492,99.9918899536133,99.9968719482422,99.9681625366211,99.9693298339844,99.9878921508789,100.011909484863,100.201957702637,100.2041015625],[134.374221801758,134.34521484375,134.335052490234,134.350784301758,134.369537353516,134.391021728516,134.419036865234,134.374221801758],[99.8430633544922,99.8478546142578,99.6851577758789,99.6070327758789,99.5374069213867,99.5588836669922,99.5618133544922,99.5721740722656,99.6220703125,99.6864242553711,99.7347640991211,99.8157196044922,99.8430633544922],[126.055076599121,126.037887573242,125.977935791016,125.937599182129,125.903228759766,125.862884521484,125.873245239258,125.922744750977,125.962799072266,125.99267578125,125.975982666016,126.065719604492,126.055076599121],[123.848243713379,123.866012573242,123.803520202637,123.777244567871,123.783493041992,123.848243713379],[126.024215698242,126.331741333008,126.288078308105,125.956443786621,125.838874816895,125.479194641113,125.432609558105,125.425971984863,125.38720703125,125.444732666016,125.520896911621,125.720314025879,126.024215698242],[123.152542114258,123.078804016113,123.070899963379,123.085838317871,123.1064453125,123.137504577637,123.152542114258],[130.353332519531,130.365432739258,130.425003051758,130.404296875,130.380569458008,130.393371582031,130.418838500977,130.372650146484,130.338973999023,130.2841796875,130.248046875,130.133483886719,130.093353271484,129.886520385742,129.75439453125,129.737701416016,129.99365234375,130.105758666992,130.199600219727,130.317977905273,130.353332519531],[124.969528198242,125.06298828125,125.095901489258,125.126754760742,125.145797729492,125.187889099121,125.197654724121,125.258193969727,125.305366516113,125.320213317871,125.314071655273,125.134765625,125.006736755371,124.834480285645,124.63916015625,124.520606994629,124.417770385742,124.329689025879,124.380859375,124.417579650879,124.483009338379,124.663970947266,124.969528198242],[135.474227905273,135.869140625,135.976165771484,136.201568603516,136.3896484375,136.718551635742,136.816696166992,136.892578125,136.708602905273,136.621871948242,136.460845947266,136.326065063477,136.228118896484,136.192581176758,136.049209594727,135.86572265625,135.487594604492,135.4697265625,135.474227905273],[108.953125,108.837890625,108.8037109375,108.877243041992,108.956840515137,108.953125],[106.045700073242,106.080078125,106.127151489258,106.161720275879,106.208786010742,106.365921020508,106.818458557129,106.744331359863,106.706642150879,106.678802490234,106.612014770508,106.618553161621,106.657615661621,106.667190551758,106.610542297363,106.546775817871,106.49609375,106.44873046875,106.397361755371,106.341598510742,106.250091552734,106.125877380371,105.998733520508,105.937202453613,105.908004760742,105.939064025879,105.907615661621,105.862396240234,105.806838989258,105.785842895508,105.705268859863,105.599021911621,105.552734375,105.342864990234,105.292579650879,105.247657775879,105.133399963379,105.1376953125,105.191009521484,105.316207885742,105.374809265137,105.386528015137,105.3642578125,105.373146057129,105.412696838379,105.459571838379,105.58544921875,105.640434265137,105.667572021484,105.700881958008,105.754486083984,105.720405578613,105.816108703613,105.910057067871,105.98095703125,106.02734375,106.045700073242],[123.597557067871,123.528617858887,123.482528686523,123.486625671387,123.528518676758,123.548530578613,123.561332702637,123.58203125,123.616409301758,123.597557067871],[128.153015136719,128.091796875,128.061218261719,127.913772583008,127.741012573242,127.561622619629,127.457611083984,127.392196655273,127.39501953125,127.456741333008,127.591796875,127.646675109863,127.742965698242,127.905075073242,128.032806396484,128.148727416992,128.153015136719],[123.212310791016,123.234275817871,123.198051452637,123.237785339355,123.338569641113,123.434768676758,123.489356994629,123.52685546875,123.547264099121,123.511909484863,123.44873046875,123.366989135742,123.32861328125,123.27490234375,123.237403869629,123.220504760742,123.172950744629,123.13037109375,123.122940063477,123.182914733887,123.150390625,123.105171203613,122.984375,122.890426635742,122.858489990234,122.810844421387,122.832221984863,122.908012390137,122.972457885742,123.158294677734,123.212310791016],[109.710258483887,109.510841369629,109.463668823242,109.428123474121,109.450294494629,109.475982666016,109.614646911621,109.699508666992,109.743362426758,109.760543823242,109.750778198242,109.710258483887],[134.96533203125,134.917388916016,134.861709594727,134.808898925781,134.827926635742,134.889251708984,134.940826416016,134.956741333008,134.996292114258,134.96533203125],[99.1638717651367,99.07177734375,98.8743133544922,98.8277359008789,98.8163070678711,98.626953125,98.6017608642578,98.6760787963867,98.8690414428711,98.9326171875,98.9547805786133,99.0650329589844,99.1014633178711,99.12890625,99.1404266357422,99.1306610107422,99.2103500366211,99.2672882080078,99.271484375,99.1638717651367],[131.001846313477,130.966598510742,130.845108032227,130.782318115234,130.739349365234,130.712097167969,130.704391479492,130.66796875,130.672958374023,130.897171020508,130.939453125,131.033020019531,131.073928833008,131.046188354492,131.001846313477],[130.9052734375,130.879791259766,130.832427978516,130.402435302734,130.439056396484,130.457321166992,130.484268188477,130.526947021484,130.548141479492,130.569534301758,130.59375,130.635452270508,130.723236083984,130.8134765625,130.807022094727,130.9052734375],[135.383010864258,135.595703125,135.673233032227,135.7490234375,135.841201782227,135.8935546875,136.068740844727,136.154678344727,136.282608032227,136.375289916992,136.305374145508,136.164749145508,136.1103515625,136.002532958984,135.9150390625,135.838775634766,135.825576782227,135.7470703125,135.645706176758,135.523834228516,135.491119384766,135.4833984375,135.431640625,135.3876953125,135.383010864258],[127.300399780273,127.289054870605,127.1845703125,127.156440734863,127.209075927734,127.258201599121,127.30126953125,127.300399780273],[130.626663208008,130.569152832031,130.465423583984,130.52587890625,130.564163208008,130.597457885742,130.615921020508,130.656921386719,130.684280395508,130.626663208008],[121.864356994629,121.906845092773,121.881256103516,121.846870422363,121.756057739258,121.721771240234,121.680953979492,121.6552734375,121.67236328125,121.749313354492,121.797370910645,121.864356994629],[140.973434448242,140.973526000977,140.9736328125,140.973724365234,140.973831176758,140.973922729492,140.974029541016,140.974029541016,140.974212646484,140.974212646484,140.974319458008,140.974411010742,140.974517822266,140.974609375,140.974609375,140.974700927734,140.974807739258,140.974899291992,140.974990844727,140.974990844727,140.944046020508,140.874603271484,140.8623046875,140.919540405273,140.975204467773,140.975204467773,140.975296020508,140.975387573242,140.975494384766,140.9755859375,140.9755859375,140.975677490234,140.975784301758,140.975875854492,140.975982666016,140.975982666016,140.976165771484,140.924621582031,140.786529541016,140.661514282227,140.5810546875,140.48974609375,140.101654052734,140.0029296875,139.983306884766,139.992584228516,140.037399291992,140.116989135742,140.033798217773,139.934768676758,139.790817260742,139.6494140625,139.5185546875,139.385650634766,139.319137573242,139.279098510742,139.25830078125,139.248825073242,139.192977905273,139.083190917969,138.933486938477,138.890625,138.86474609375,138.856140136719,138.885055541992,138.905471801758,138.935943603516,139.003021240234,139.045700073242,139.073638916016,139.087997436523,139.048919677734,138.983016967773,138.937896728516,138.885543823242,138.853134155273,138.793655395508,138.747940063477,138.798431396484,138.864852905273,138.919342041016,139.017974853516,139.0625,139.176849365234,139.112594604492,139.049011230469,138.845703125,138.720016479492,138.601364135742,138.600189208984,138.683792114258,138.864547729492,138.808502197266,138.726654052734,138.698135375977,138.642181396484,138.521575927734,138.438674926758,138.368362426758,138.296295166016,138.313873291016,138.374603271484,138.282806396484,138.199615478516,138.243560791016,138.339645385742,138.252151489258,138.16650390625,138.12744140625,138.087112426758,138.06591796875,138.063079833984,138.075576782227,138.060836791992,137.984954833984,137.922271728516,137.886825561523,137.84033203125,137.795211791992,137.759078979492,137.306640625,137.27978515625,137.237884521484,137.195892333984,137.143753051758,137.089263916016,137.029693603516,136.974609375,136.9169921875,136.856842041016,136.618850708008,136.393753051758,136.210632324219,136.097457885742,135.979675292969,135.716598510742,135.4501953125,135.353897094727,135.273132324219,135.195602416992,134.754196166992,134.6796875,134.686920166016,134.70654296875,134.886520385742,134.759765625,134.707611083984,134.603408813477,134.546875,134.467193603516,134.391021728516,134.266204833984,134.202346801758,134.180465698242,134.147079467773,134.100006103516,134.036911010742,133.973831176758,133.933212280273,133.904006958008,133.860733032227,133.808502197266,133.723037719727,133.678314208984,133.683395385742,133.697174072266,133.781646728516,133.841506958008,133.767379760742,133.700393676758,133.671981811523,133.660736083984,133.653121948242,133.599411010742,133.518157958984,133.542282104492,133.509170532227,133.415130615234,133.4072265625,133.422271728516,133.40087890625,133.248733520508,133.198043823242,133.085159301758,132.968551635742,132.914459228516,132.8701171875,132.837112426758,132.790908813477,132.75390625,132.869735717773,132.829788208008,132.751358032227,132.553527832031,132.348251342773,132.254974365234,132.10205078125,132.053909301758,132.00634765625,131.97119140625,132.06689453125,132.230651855469,132.323348999023,132.575485229492,132.652923583984,132.725006103516,132.897277832031,133.033798217773,133.118850708008,133.191009521484,133.264938354492,133.411422729492,133.526565551758,133.608688354492,133.651550292969,133.700088500977,133.7109375,133.753311157227,133.834671020508,133.877639770508,133.904891967773,133.89892578125,133.791015625,133.849700927734,133.902450561523,133.920501708984,133.921585083008,133.710357666016,133.48779296875,133.356246948242,133.224899291992,132.962783813477,132.86328125,132.631057739258,132.502639770508,132.4033203125,132.3076171875,132.207427978516,132.122161865234,132.079895019531,132.0234375,131.998428344727,131.936126708984,131.930374145508,131.829788208008,131.7314453125,131.293746948242,131.240814208984,131.17919921875,131.117782592773,131.056747436523,130.995895385742,131.0009765625,131.046188354492,131.090530395508,131.15185546875,131.190811157227,131.254104614258,131.258987426758,131.252059936523,131.257217407227,131.29638671875,131.461532592773,131.804290771484,131.890914916992,131.96240234375,132.046005249023,132.08447265625,132.12841796875,132.393753051758,132.508010864258,132.625106811523,132.8564453125,133.0771484375,133.268447875977,133.47265625,133.7236328125,133.850280761719,133.974517822266,134.02490234375,134.111526489258,134.086715698242,134.071975708008,134.1162109375,134.188293457031,134.247161865234,134.259567260742,134.237197875977,134.216979980469,134.145401000977,134.105865478516,134.131256103516,134.145401000977,134.142776489258,134.155670166016,134.19482421875,134.362121582031,134.4599609375,134.4912109375,134.483291625977,134.517959594727,134.56689453125,134.62744140625,134.644714355469,134.649017333984,134.7021484375,134.769821166992,134.843353271484,134.855377197266,134.852722167969,134.886810302734,134.917190551758,135.037399291992,135.092193603516,135.251556396484,135.37158203125,135.486618041992,135.560745239258,135.627731323242,135.859176635742,135.926177978516,135.99072265625,136.012985229492,136.243255615234,136.26953125,136.302551269531,136.352447509766,136.389938354492,136.6123046875,136.84326171875,137.072067260742,137.171096801758,137.17578125,137.12548828125,137.123443603516,137.176467895508,137.380569458008,137.616607666016,137.806259155273,137.9111328125,138.0078125,138.110931396484,138.649810791016,138.736129760742,138.811416625977,138.919128417969,139.039459228516,139.148834228516,139.252639770508,139.481826782227,139.78955078125,139.868347167969,140.154586791992,140.204010009766,140.2509765625,140.294631958008,140.62255859375,140.673049926758,140.720504760742,140.747467041016,140.973434448242],[104.474220275879,104.567771911621,104.590133666992,104.5439453125,104.506546020508,104.4853515625,104.413864135742,104.363189697266,104.329788208008,104.257125854492,104.302345275879,104.318748474121,104.340721130371,104.363578796387,104.474220275879],[127.566993713379,127.682418823242,127.60498046875,127.658599853516,127.804298400879,127.837890625,127.863288879395,127.880172729492,127.842292785645,127.761131286621,127.667572021484,127.642875671387,127.623832702637,127.497856140137,127.462692260742,127.438285827637,127.468650817871,127.380569458008,127.300003051758,127.297073364258,127.329490661621,127.325096130371,127.371185302734,127.455184936523,127.49169921875,127.527351379395,127.566993713379],[127.249908447266,127.187301635742,127.119140625,127.104400634766,127.126457214355,127.189651489258,127.2900390625,127.253021240234,127.280563354492,127.249908447266],[103.736526489258,103.606346130371,103.461326599121,103.47900390625,103.548927307129,103.610939025879,103.723930358887,103.764251708984,103.736526489258],[130.813278198242,130.986526489258,131.025787353516,131.27685546875,131.31689453125,131.302734375,131.339736938477,131.257522583008,131.217864990234,131.177734375,131.097747802734,131.00537109375,130.946487426758,130.896682739258,130.808395385742,130.683502197266,130.622161865234,130.638275146484,130.691314697266,130.761322021484,130.801559448242,130.843154907227,130.899215698242,130.896286010742,130.750198364258,130.699813842773,130.688674926758,130.606552124023,130.574905395508,130.55078125,130.496276855469,130.340530395508,130.236633300781,130.287689208984,130.294921875,130.362503051758,130.430953979492,130.499618530273,130.54833984375,130.584289550781,130.722366333008,130.813278198242],[98.4592819213867,98.3997039794922,98.3096618652344,98.3399429321289,98.3547821044922,98.4087905883789,98.4271545410156,98.3229522705078,98.37451171875,98.4154281616211,98.484375,98.5441360473633,98.5201187133789,98.4592819213867],[104.778610229492,104.807518005371,104.843162536621,104.908988952637,104.94970703125,105.00537109375,104.950584411621,104.928512573242,104.914260864258,104.702247619629,104.566604614258,104.473533630371,104.447067260742,104.4970703125,104.542678833008,104.635643005371,104.658401489258,104.652732849121,104.713478088379,104.778610229492],[129.548919677734,129.505661010742,129.46923828125,129.3701171875,129.308792114258,129.5419921875,129.548919677734],[127.453422546387,127.448638916016,127.417877197266,127.396781921387,127.419525146484,127.431343078613,127.449417114258,127.453422546387],[104.689262390137,104.698143005371,104.65087890625,104.622367858887,104.603515625,104.499214172363,104.543846130371,104.659866333008,104.689262390137],[103.284477233887,103.172164916992,103.139549255371,103.1533203125,103.187400817871,103.238189697266,103.295112609863,103.284477233887],[127.419723510742,127.383987426758,127.373626708984,127.362892150879,127.382621765137,127.4248046875,127.442573547363,127.445892333984,127.419723510742],[127.372657775879,127.338386535645,127.306053161621,127.286422729492,127.292770385742,127.31982421875,127.353805541992,127.372657775879],[103.4501953125,103.4296875,103.344436645508,103.36572265625,103.386131286621,103.43310546875,103.470314025879,103.497459411621,103.4501953125],[103.82861328125,103.833984375,103.742385864258,103.740043640137,103.751953125,103.806640625,103.82861328125],[104.239356994629,104.1767578125,104.09814453125,104.10107421875,104.108299255371,104.122749328613,104.170509338379,104.22705078125,104.239356994629],[103.027534484863,103.0087890625,102.971481323242,102.776268005371,102.710548400879,102.541603088379,102.490432739258,102.453910827637,102.466407775879,102.491409301758,102.506637573242,102.549217224121,102.633201599121,102.726173400879,102.780075073242,102.944137573242,103.00244140625,103.027534484863],[103.423927307129,103.4296875,103.36328125,103.3154296875,103.35498046875,103.379981994629,103.404884338379,103.423927307129],[103.166404724121,103.13720703125,103.086723327637,103.033393859863,102.963966369629,102.886329650879,102.787986755371,102.726463317871,102.701850891113,102.7255859375,102.790138244629,102.999420166016,103.067573547363,103.166404724121],[104.024803161621,104.088088989258,104.139846801758,104.137786865234,104.127342224121,104.066116333008,103.963569641113,103.939842224121,103.932228088379,103.946975708008,103.955368041992,103.999809265137,104.024803161621],[104.585350036621,104.591011047363,104.648139953613,104.662887573242,104.65283203125,104.59912109375,104.5751953125,104.504295349121,104.480667114258,104.47119140625,104.481056213379,104.428611755371,104.46240234375,104.439254760742,104.2939453125,104.251953125,104.244239807129,104.250198364258,104.36181640625,104.428413391113,104.500099182129,104.585350036621],[102.427146911621,102.380859375,102.325286865234,102.279586791992,102.255462646484,102.234184265137,102.228614807129,102.25634765625,102.276458740234,102.358596801758,102.412887573242,102.44287109375,102.448829650879,102.428901672363,102.427146911621],[97.4815444946289,97.6983413696289,97.7864227294922,97.9032211303711,97.9319381713867,97.9020538330078,97.87646484375,97.8204116821289,97.6839828491211,97.6825180053711,97.6039123535156,97.4612350463867,97.4053649902344,97.3688507080078,97.296875,97.0792007446289,97.2442398071289,97.3244171142578,97.3427734375,97.35595703125,97.4815444946289],[102.491889953613,102.499412536621,102.425193786621,102.366889953613,102.274223327637,102.161323547363,102.078712463379,102.020896911621,102.018363952637,102.024024963379,102.042182922363,102.469528198242,102.491889953613],[124.888862609863,124.698150634766,124.639846801758,124.589057922363,124.514060974121,124.427536010742,124.384368896484,124.278022766113,124.216796875,124.101371765137,123.753807067871,123.6396484375,123.525978088379,123.310447692871,123.265434265137,123.179496765137,123.08251953125,122.996871948242,122.909568786621,122.280769348145,122.060928344727,121.841995239258,121.722755432129,121.604591369629,121.515716552734,121.42578125,121.012985229492,120.9091796875,120.700386047363,120.578994750977,120.4599609375,120.349029541016,120.307037353516,120.192283630371,120.127342224121,120.078315734863,120.036041259766,120.013282775879,120.012107849121,120.03173828125,120.062896728516,120.097457885742,120.240631103516,120.269821166992,120.425384521484,120.517578125,120.605079650879,120.667381286621,120.728607177734,120.796974182129,120.915824890137,121.033699035645,121.148544311523,121.212593078613,121.27685546875,121.431350708008,121.519340515137,121.575592041016,121.632720947266,121.681159973145,121.737701416016,121.853126525879,121.969627380371,122.093650817871,122.138084411621,122.174896240234,122.279975891113,122.529685974121,122.658790588379,122.888763427734,122.885551452637,122.841117858887,122.829490661621,122.872261047363,123.020408630371,123.171478271484,123.281448364258,123.379692077637,123.417381286621,123.434181213379,123.396286010742,123.3779296875,123.299613952637,123.225776672363,123.152740478516,123.049407958984,122.90283203125,122.8525390625,122.807418823242,122.724609375,122.655662536621,122.506645202637,122.334182739258,122.250686645508,122.157615661621,121.858596801758,121.779884338379,121.71875,121.650985717773,121.572654724121,121.513870239258,121.394729614258,121.35546875,121.348831176758,121.407516479492,121.501945495605,121.575004577637,121.621871948242,121.725975036621,121.769729614258,121.848236083984,121.971878051758,122.013969421387,122.082626342773,122.291694641113,122.303321838379,122.290428161621,122.306541442871,122.381256103516,122.399024963379,122.317283630371,122.312789916992,122.2626953125,122.251373291016,122.2529296875,122.2880859375,122.3291015625,122.385353088379,122.4345703125,122.529197692871,122.578620910645,122.609954833984,122.606742858887,122.649894714355,122.689651489258,122.750381469727,122.77880859375,122.798233032227,122.847946166992,122.877342224121,122.894340515137,122.899803161621,122.897369384766,122.872261047363,122.817581176758,122.7197265625,122.715042114258,122.7216796875,122.671875,122.614753723145,122.471382141113,122.207122802734,122.1142578125,122.05419921875,122.050003051758,122.0732421875,122.038093566895,121.9169921875,121.748054504395,121.645698547363,121.588676452637,121.514350891113,121.486526489258,121.541206359863,121.556739807129,121.583404541016,121.611518859863,121.618064880371,121.537399291992,121.415817260742,121.312698364258,120.914253234863,120.891792297363,120.890922546387,120.906929016113,121.037895202637,121.054298400879,121.070304870605,121.066795349121,121.052154541016,120.990135192871,120.879386901855,120.765037536621,120.653610229492,120.5439453125,120.341400146484,120.261039733887,120.254096984863,120.300491333008,120.360443115234,120.392387390137,120.436614990234,120.435157775879,120.383010864258,120.362495422363,120.384567260742,120.420112609863,120.404975891113,120.310165405273,120.281448364258,120.279304504395,120.390922546387,120.416603088379,120.430374145508,120.311622619629,120.256446838379,120.200782775879,120.077049255371,119.951560974121,119.907615661621,119.818458557129,119.764457702637,119.71728515625,119.557418823242,119.463081359863,119.376174926758,119.360359191895,119.390625,119.433586120605,119.519523620605,119.515525817871,119.544921875,119.594047546387,119.611717224121,119.623634338379,119.611419677734,119.49365234375,119.480072021484,119.479293823242,119.491989135742,119.494529724121,119.467483520508,119.419822692871,119.362106323242,119.240036010742,118.99462890625,118.922172546387,118.867683410645,118.832809448242,118.812507629395,118.821876525879,118.858108520508,118.828910827637,118.783699035645,118.783302307129,118.808982849121,118.853324890137,118.907516479492,118.958206176758,119.092185974121,119.135353088379,119.13818359375,119.172264099121,119.240814208984,119.321876525879,119.348236083984,119.308296203613,119.324111938477,119.310348510742,119.308982849121,119.359184265137,119.508201599121,119.653518676758,119.711334228516,119.786720275879,119.844329833984,119.84521484375,119.829879760742,119.771682739258,119.721878051758,119.73583984375,119.786521911621,119.838279724121,119.865623474121,119.811721801758,119.809280395508,119.913284301758,119.998046875,120.035148620605,120.056449890137,120.1005859375,120.156539916992,120.229782104492,120.269538879395,120.293853759766,120.322456359863,120.366500854492,120.416015625,120.5166015625,120.6025390625,120.62646484375,120.658882141113,120.711036682129,120.754875183105,120.803619384766,120.867973327637,120.912109375,120.965423583984,121.024612426758,121.081741333008,121.208396911621,121.281745910645,121.356735229492,121.404098510742,121.440040588379,121.472755432129,121.513282775879,121.550689697266,121.591789245605,121.867378234863,122.108207702637,122.436614990234,122.54931640625,122.657424926758,122.789848327637,122.838272094727,122.892478942871,122.960060119629,123.012802124023,123.066497802734,123.278121948242,123.846687316895,123.930763244629,124.273635864258,124.410842895508,124.53369140625,124.575393676758,124.600189208984,124.643753051758,124.746681213379,124.787689208984,124.860641479492,124.947067260742,124.989250183105,125.110939025879,125.164840698242,125.233787536621,125.2216796875,125.140922546387,125.117485046387,125.028030395508,124.966796875,124.888862609863],[101.708106994629,101.762306213379,101.773536682129,101.734085083008,101.719429016113,101.602729797363,101.500785827637,101.4677734375,101.403419494629,101.40966796875,101.450294494629,101.544731140137,101.640724182129,101.708106994629],[127.732719421387,127.805366516113,127.881057739258,127.918647766113,127.929100036621,127.96728515625,128.055267333984,128.116989135742,128.160736083984,128.153137207031,128.157424926758,128.222457885742,128.424133300781,128.539260864258,128.688369750977,128.705169677734,128.688079833984,128.716888427734,128.70263671875,128.668762207031,128.514541625977,128.345993041992,128.298828125,128.257217407227,128.260635375977,128.39794921875,128.611236572266,128.6552734375,128.683776855469,128.691589355469,128.743255615234,128.8154296875,128.86328125,128.899597167969,128.540435791016,128.446487426758,128.332809448242,128.220596313477,128.106063842773,127.983100891113,127.924415588379,127.901374816895,127.887397766113,127.914649963379,127.912200927734,127.888969421387,127.977828979492,128.089447021484,128.253524780273,128.334564208984,128.425491333008,128.278121948242,128.2333984375,128.04638671875,128.010848999023,127.888969421387,127.853317260742,127.740821838379,127.691604614258,127.674797058105,127.687408447266,127.681343078613,127.685455322266,127.708694458008,127.668655395508,127.6162109375,127.555366516113,127.537101745605,127.541801452637,127.566993713379,127.600677490234,127.608016967773,127.520408630371,127.428520202637,127.420310974121,127.537101745605,127.534675598145,127.557907104492,127.570709228516,127.631744384766,127.731452941895,127.899894714355,127.964263916016,128.036437988281,128.042770385742,128.03125,127.90673828125,127.890144348145,127.886825561523,127.946479797363,128.010940551758,128.023727416992,128.02587890625,128.01171875,127.987693786621,127.885345458984,127.65283203125,127.633010864258,127.634376525879,127.677436828613,127.732719421387],[97.3341751098633,97.3283157348633,97.22509765625,97.1083068847656,97.1566390991211,97.2528305053711,97.2914047241211,97.3287124633789,97.3341751098633],[128.453903198242,128.2958984375,128.259948730469,128.217956542969,128.330368041992,128.472076416016,128.568649291992,128.602142333984,128.6884765625,128.623245239258,128.547561645508,128.453903198242],[125.407424926758,125.397262573242,125.360061645508,125.390823364258,125.435249328613,125.446479797363,125.403915405273,125.407424926758],[96.4636688232422,96.4009780883789,96.3406219482422,96.2904281616211,96.02197265625,95.9384765625,95.8797836303711,95.80859375,95.7330093383789,95.7171859741211,95.7721710205078,95.8062515258789,95.8957977294922,95.9978485107422,96.1015625,96.1297836303711,96.1799774169922,96.4172897338867,96.4430618286133,96.4593734741211,96.4636688232422],[108.887496948242,108.8388671875,108.786529541016,108.867088317871,108.8857421875,108.887496948242],[105.760353088379,105.718559265137,105.706153869629,105.707908630371,105.704200744629,105.692184448242,105.692184448242,105.730667114258,105.760353088379,105.794525146484,105.822166442871,105.836715698242,105.809379577637,105.760353088379],[106.285255432129,106.28369140625,106.214553833008,106.200981140137,106.223731994629,106.271194458008,106.285255432129],[117.658393859863,117.645797729492,117.560356140137,117.537498474121,117.547859191895,117.63671875,117.680854797363,117.658393859863],[125.658103942871,125.633201599121,125.511528015137,125.517578125,125.501174926758,125.468551635742,125.455276489258,125.468856811523,125.543449401855,125.585647583008,125.6435546875,125.658103942871],[126.851860046387,126.835548400879,126.799606323242,126.777542114258,126.778907775879,126.804496765137,126.857032775879,126.857810974121,126.851860046387],[126.719337463379,126.721786499023,126.661224365234,126.637496948242,126.685539245605,126.739654541016,126.719337463379],[117.884765625,117.917877197266,117.922843933105,117.73681640625,117.625091552734,117.649024963379,117.745414733887,117.884765625],[108.316017150879,108.179588317871,108.100395202637,108.186134338379,108.216400146484,108.236137390137,108.243263244629,108.088478088379,108.044532775879,108.002342224121,108.003517150879,108.201950073242,108.248344421387,108.255569458008,108.392868041992,108.398834228516,108.3935546875,108.316017150879],[117.574409484863,117.566215515137,117.497459411621,117.46533203125,117.559371948242,117.566009521484,117.639068603516,117.728225708008,117.731735229492,117.762008666992,117.777244567871,117.714454650879,117.629890441895,117.567375183105,117.509658813477,117.494918823242,117.450386047363,117.287887573242,117.171577453613,117.055961608887,117.113868713379,117.166404724121,117.346282958984,117.384666442871,117.321868896484,117.352447509766,117.422073364258,117.5068359375,117.567184448242,117.610641479492,117.612403869629,117.637893676758,117.569137573242,117.63720703125,117.697654724121,117.664558410645,117.638862609863,117.666793823242,117.749702453613,117.785942077637,117.804878234863,117.8857421875,118.0341796875,118.066596984863,118.066307067871,118.041595458984,117.957023620605,117.889259338379,117.881057739258,117.789260864258,117.831253051758,117.864654541016,117.928421020508,118.080368041992,118.156837463379,118.4716796875,118.638961791992,118.852546691895,118.963478088379,118.984954833984,118.892379760742,118.757431030273,118.534767150879,118.3115234375,118.196098327637,118.095512390137,118.016311645508,117.91162109375,117.951957702637,117.977340698242,117.964248657227,117.923049926758,117.8525390625,117.776954650879,117.745109558105,117.553321838379,117.522171020508,117.463760375977,117.462890625,117.548919677734,117.556831359863,117.573822021484,117.5625,117.521781921387,117.357131958008,117.240715026855,117.146484375,117.070220947266,117.003219604492,116.913963317871,116.849411010742,116.797065734863,116.760543823242,116.739845275879,116.726173400879,116.728713989258,116.759284973145,116.770988464355,116.753425598145,116.715225219727,116.611618041992,116.554489135742,116.545112609863,116.517578125,116.477935791016,116.332122802734,116.299613952637,116.275489807129,116.353225708008,116.42431640625,116.429588317871,116.451950073242,116.423538208008,116.313972473145,116.36865234375,116.418167114258,116.528121948242,116.5654296875,116.549217224121,116.529296875,116.450393676758,116.401275634766,116.352546691895,116.316795349121,116.307220458984,116.375495910645,116.371681213379,116.353225708008,116.330665588379,116.288864135742,116.225776672363,116.166313171387,116.154098510742,116.172271728516,116.257225036621,116.205078125,116.167091369629,116.150001525879,116.057525634766,116.016708374023,115.999412536621,115.956146240234,115.258201599121,114.693557739258,114.652542114258,114.625289916992,114.60595703125,114.5361328125,114.525588989258,114.445991516113,114.397163391113,114.344329833984,114.304588317871,114.301658630371,114.344329833984,114.292678833008,114.236328125,114.177932739258,114.127639770508,114.108985900879,114.082221984863,113.958786010742,113.795799255371,113.705078125,113.633590698242,113.637306213379,113.630081176758,113.610061645508,113.566307067871,113.525978088379,113.408988952637,113.3671875,113.343162536621,113.033981323242,112.971488952637,112.758010864258,112.600296020508,112.443939208984,112.284965515137,112.126663208008,111.954879760742,111.907424926758,111.858100891113,111.822067260742,111.834373474121,111.8359375,111.823043823242,111.809379577637,111.760162353516,111.694725036621,111.658302307129,111.62548828125,111.494918823242,111.367576599121,111.259178161621,111.044334411621,110.930076599121,110.868751525879,110.829689025879,110.85205078125,110.899314880371,110.811134338379,110.73583984375,110.703125,110.668159484863,110.574020385742,110.377540588379,110.350975036621,110.302543640137,110.256057739258,110.232612609863,110.224311828613,110.124420166016,110.096580505371,110.075004577637,109.959861755371,109.963768005371,110.0234375,110.0361328125,110.019241333008,109.983299255371,109.938079833984,109.873435974121,109.787399291992,109.681739807129,109.453811645508,109.33349609375,109.288864135742,109.2587890625,109.27099609375,109.311721801758,109.366310119629,109.372756958008,109.257034301758,109.160545349121,109.130279541016,109.12109375,109.121780395508,109.149612426758,109.164749145508,109.194633483887,109.257522583008,109.247261047363,109.22021484375,109.180755615234,109.148536682129,109.074798583984,108.944526672363,108.922752380371,108.905853271484,108.916801452637,108.958595275879,109.030853271484,109.088478088379,109.131538391113,109.096092224121,109.0654296875,109.01025390625,109.055473327637,109.075881958008,109.166702270508,109.273139953613,109.318161010742,109.378509521484,109.62890625,109.538970947266,109.548927307129,109.57080078125,109.635841369629,109.653999328613,109.735748291016,109.818069458008,109.878517150879,109.944923400879,109.99169921875,110.040824890137,110.11474609375,110.315231323242,110.399024963379,110.46142578125,110.505760192871,110.61474609375,110.938087463379,110.99609375,111.101364135742,111.286720275879,111.483200073242,111.546684265137,111.607421875,111.691307067871,111.769729614258,111.808982849121,111.923149108887,112.078514099121,112.128616333008,112.167381286621,112.185737609863,112.250679016113,112.341598510742,112.476173400879,112.942970275879,112.98828125,112.998046875,112.98828125,113.006546020508,113.068649291992,113.126274108887,113.358985900879,113.458206176758,113.51318359375,113.622268676758,113.681640625,113.760353088379,113.835250854492,113.90234375,114,114.1259765625,114.274703979492,114.387107849121,114.512496948242,114.5458984375,114.567481994629,114.632225036621,114.660942077637,114.686126708984,114.703521728516,114.751068115234,114.800003051758,114.812690734863,114.830558776855,114.815818786621,114.787994384766,114.758697509766,114.768363952637,114.786422729492,114.836326599121,114.969146728516,115.086532592773,115.150787353516,115.179107666016,115.180854797363,115.129890441895,115.080764770508,115.077049255371,115.078903198242,115.093643188477,115.086532592773,115.086326599121,115.117576599121,115.18994140625,115.246971130371,115.310150146484,115.384178161621,115.454391479492,115.493156433105,115.499130249023,115.48974609375,115.514259338379,115.519920349121,115.566116333008,115.570709228516,115.544532775879,115.560943603516,115.568450927734,115.596092224121,115.627532958984,115.678802490234,115.782417297363,115.836814880371,115.860748291016,115.896194458008,116.021583557129,116.134468078613,116.236228942871,116.320304870605,116.367683410645,116.41455078125,116.514747619629,116.553123474121,116.589057922363,116.638679504395,116.697853088379,116.843551635742,117.1005859375,117.277542114258,117.450881958008,117.537307739258,117.574409484863],[126.816596984863,126.776275634766,126.711227416992,126.704490661621,126.770111083984,126.813568115234,126.767288208008,126.722068786621,126.720512390137,126.757331848145,126.8125,126.865135192871,126.88671875,126.921089172363,126.847648620605,126.816596984863],[96.4925765991211,96.615234375,96.8426742553711,96.9677734375,97.0857467651367,97.1904296875,97.451171875,97.5001907348633,97.5471725463867,97.5875015258789,97.7067337036133,97.9084014892578,97.9665985107422,97.9998092651367,98.0206985473633,98.2484359741211,98.2733459472656,98.2412109375,98.3073196411133,98.5283203125,98.65869140625,98.6865234375,98.7057647705078,98.7779312133789,98.86865234375,99.1511688232422,99.521484375,99.7323226928711,99.9066390991211,99.9694366455078,100.021293640137,100.127250671387,100.307228088379,100.352729797363,100.401168823242,100.45703125,100.523826599121,100.603614807129,100.685249328613,100.816795349121,100.887893676758,100.876663208008,100.81689453125,100.817771911621,100.828224182129,100.877052307129,100.935935974121,101.046195983887,101.225189208984,101.30078125,101.357620239258,101.405082702637,101.476661682129,101.575004577637,101.684280395508,101.784759521484,102.019920349121,102.098052978516,102.1572265625,102.197952270508,102.223335266113,102.239059448242,102.389938354492,102.46923828125,102.56640625,102.849411010742,102.949317932129,103.031829833984,103.066505432129,103.007514953613,102.786331176758,102.549995422363,102.779586791992,102.895896911621,103.002830505371,103.108695983887,103.276565551758,103.338966369629,103.412300109863,103.478912353516,103.578712463379,103.672653198242,103.742774963379,103.786720275879,103.706443786621,103.589454650879,103.428520202637,103.41162109375,103.444427490234,103.405174255371,103.495414733887,103.509178161621,103.43115234375,103.438575744629,103.53271484375,103.577537536621,103.721092224121,103.940032958984,104.061134338379,104.198539733887,104.257514953613,104.360542297363,104.381256103516,104.425682067871,104.446876525879,104.478324890137,104.5185546875,104.515914916992,104.568748474121,104.676361083984,104.791015625,104.84521484375,104.844528198242,104.826072692871,104.787307739258,104.66845703125,104.647270202637,104.630569458008,104.650779724121,104.698333740234,104.735740661621,104.87841796875,104.9169921875,104.970802307129,105.02587890625,105.286529541016,105.39697265625,105.495307922363,105.58203125,105.899116516113,106.044334411621,106.055763244629,106.058395385742,106.03369140625,105.901458740234,105.885055541992,105.84375,105.8515625,105.8955078125,105.930465698242,105.927734375,105.840621948242,105.831443786621,105.886520385742,105.890525817871,105.879295349121,105.88720703125,105.816108703613,105.802734375,105.748344421387,105.676567077637,105.618553161621,105.577926635742,105.555564880371,105.522659301758,105.349411010742,105.304000854492,105.128128051758,105.081344604492,105.022651672363,104.930274963379,104.639549255371,104.621681213379,104.6181640625,104.675971984863,104.683990478516,104.631050109863,104.6015625,104.480857849121,104.369529724121,104.242965698242,104.150489807129,104.066795349121,103.831443786621,103.770309448242,103.405662536621,103.332130432129,103.238868713379,103.138671875,102.9189453125,102.537696838379,102.371971130371,102.187698364258,102.127532958984,101.81787109375,101.649024963379,101.57861328125,101.414260864258,101.3662109375,101.3056640625,101.206253051758,101.118553161621,100.944427490234,100.889549255371,100.848045349121,100.855278015137,100.486518859863,100.393943786621,100.308204650879,100.2890625,100.087890625,100.016700744629,99.9306640625,99.8600540161133,99.7212905883789,99.6698226928711,99.59765625,99.3345718383789,99.2364273071289,99.1591796875,99.1117172241211,99.0595703125,98.935546875,98.79638671875,98.7025375366211,98.5953140258789,98.5642623901367,98.0865249633789,98.0050811767578,97.9185562133789,97.7950210571289,97.7007827758789,97.6620101928711,97.640625,97.6167984008789,97.5908203125,97.3913040161133,97.3131790161133,97.2479476928711,97.1883773803711,96.9689483642578,96.8939437866211,96.8009796142578,96.525390625,96.4447250366211,96.3108367919922,96.2300796508789,95.9879913330078,95.57861328125,95.4947280883789,95.4319381713867,95.3812484741211,95.2066421508789,95.220703125,95.2470703125,95.2429733276367,95.2238235473633,95.2278289794922,95.2795867919922,95.3960876464844,95.5169906616211,95.62890625,95.7373046875,95.84130859375,96.02734375,96.13330078125,96.2508773803711,96.4925765991211],[95.3621139526367,95.3425750732422,95.283203125,95.2176742553711,95.2419967651367,95.2825164794922,95.3591842651367,95.3660202026367,95.3621139526367],[-4.41206026077271,-4.6142578125,-4.69609403610229,-4.76577138900757,-4.78535127639771,-4.74555683135986,-4.69873046875,-4.61484384536743,-4.50864267349243,-4.42470693588257,-4.39555692672729,-4.377197265625,-4.33798837661743,-4.39228534698486,-4.41206026077271],[93.8900375366211,93.8288116455078,93.7092819213867,93.6580047607422,93.6563491821289,93.6841812133789,93.8224639892578,93.8589859008789,93.9295883178711,93.8900375366211],[93.7335968017578,93.6384811401367,93.5972671508789,93.6142578125,93.6546859741211,93.6924743652344,93.7335968017578],[93.4425735473633,93.3650360107422,93.3419952392578,93.3093719482422,93.33447265625,93.37548828125,93.4336853027344,93.4473648071289,93.4425735473633],[93.5369110107422,93.4900360107422,93.4782180786133,93.4717712402344,93.4697265625,93.4612274169922,93.4564437866211,93.4940414428711,93.5316390991211,93.5116195678711,93.5369110107422],[73.0673828125,73.0533218383789,73.0388641357422,73.0285186767578,73.0234375,73.0260772705078,73.0389633178711,73.0558624267578,73.0751953125,73.0794906616211,73.0835876464844,73.0797882080078,73.0673828125],[93.1407241821289,93.1706008911133,93.115234375,93.0642547607422,93.0775451660156,93.0969772338867,93.1407241821289],[92.7874984741211,92.7435531616211,92.7165985107422,92.7132797241211,92.7385711669922,92.7621078491211,92.7857437133789,92.8092803955078,92.7874984741211],[92.5028305053711,92.47265625,92.3695297241211,92.3771438598633,92.3528289794922,92.3706970214844,92.4478530883789,92.5103530883789,92.5540008544922,92.5743179321289,92.5028305053711],[72.7803726196289,72.7730484008789,72.7724609375,72.7818298339844,72.7926712036133,72.7958984375,72.7928695678711,72.7878952026367,72.7803726196289],[92.6931686401367,92.64453125,92.595703125,92.6338882446289,92.6402359008789,92.6900405883789,92.6872024536133,92.6931686401367],[93.0173797607422,93.0621109008789,92.9817352294922,92.9553680419922,92.9957962036133,93.0173797607422],[92.7175750732422,92.6857452392578,92.6796875,92.6944274902344,92.7106475830078,92.7308654785156,92.7175750732422],[92.7227554321289,92.7007827758789,92.6683578491211,92.5755844116211,92.5596694946289,92.5338897705078,92.5664978027344,92.6075210571289,92.6318359375,92.640625,92.6764678955078,92.6947250366211,92.7692413330078,92.7882843017578,92.7776336669922,92.7340774536133,92.7189407348633,92.720703125,92.7320327758789,92.7591781616211,92.7400436401367,92.7531204223633,92.8070297241211,92.8308563232422,92.8089828491211,92.8601608276367,92.8573226928711,92.9246063232422,93.0293960571289,93.0623016357422,93.0667037963867,93.07666015625,93.0160140991211,93.0738296508789,93.0661163330078,93.04296875,93.0046844482422,92.9513702392578,92.9099655151367,92.88623046875,92.9650421142578,92.990234375,92.9326171875,92.8636703491211,92.8794937133789,92.8671875,92.798828125,92.7862243652344,92.7476577758789,92.7639617919922,92.7967834472656,92.7975616455078,92.7669906616211,92.7646484375,92.7227554321289],[77.7994155883789,77.8025436401367,77.8109359741211,77.8515625,77.8949203491211,77.9458999633789,78.0094757080078,78.0426788330078,78.0474548339844,78.0091781616211,78.01220703125,78.0757827758789,78.1584930419922,78.2361373901367,78.2820358276367,78.3269577026367,78.5157241821289,78.6707992553711,78.7630844116211,78.8648376464844,78.9364242553711,78.9701156616211,78.9769515991211,78.9706039428711,78.9317321777344,78.7530288696289,78.7317428588867,78.7266616821289,78.76171875,78.7837905883789,78.7899398803711,78.8018493652344,78.8650360107422,78.9167022705078,78.9484329223633,79.0125961303711,79.0665054321289,79.1124954223633,79.1156234741211,79.1351623535156,79.1216735839844,79.1028289794922,79.1085968017578,79.1455078125,79.2022476196289,79.2095642089844,79.20556640625,79.2279281616211,79.23388671875,79.2165069580078,79.2190399169922,79.2193374633789,79.169921875,79.1273422241211,79.0669937133789,78.9976501464844,78.9189453125,78.837890625,78.7712860107422,78.7535171508789,78.7367172241211,78.7008819580078,78.6315383911133,78.5263671875,78.4124984741211,78.3917007446289,78.3896484375,78.41748046875,78.4413070678711,78.4552764892578,78.4861373901367,78.4959030151367,78.677734375,78.7255859375,78.7354507446289,78.7197265625,78.68701171875,78.6934585571289,78.75390625,78.8029327392578,78.7550811767578,78.7267532348633,78.7585983276367,78.7435607910156,78.7578125,78.7916030883789,78.8445358276367,78.8995132446289,78.9459991455078,78.9739227294922,79.0111312866211,79.0437469482422,79.1071243286133,79.2326202392578,79.3387680053711,79.36962890625,79.3884735107422,79.4931640625,79.5654296875,79.6642608642578,79.7946243286133,79.8718795776367,79.9166030883789,79.9245071411133,80.0814514160156,80.1494140625,80.1943359375,80.2071304321289,80.1862258911133,80.1912078857422,80.2609405517578,80.4095687866211,80.541015625,80.60888671875,80.68212890625,80.7467727661133,80.87353515625,80.9854507446289,81.01025390625,80.9661178588867,80.9076232910156,80.84814453125,80.8199234008789,80.68408203125,80.6128921508789,80.5490188598633,80.40185546875,80.31689453125,80.2548828125,80.2559585571289,80.2330093383789,80.1695327758789,80.1304702758789,80.0845718383789,80.0516586303711,80.0707015991211,80.1496047973633,80.2265625,80.3248062133789,80.4185485839844,80.4791030883789,80.4958038330078,80.5178680419922,80.5870132446289,80.6712875366211,80.7261734008789,80.7507781982422,80.8960876464844,81.0166015625,81.1689453125,81.2062530517578,81.2389678955078,81.3108444213867,81.4860382080078,81.6355514526367,81.7572250366211,81.8526382446289,81.8968811035156,81.9452133178711,81.9876937866211,82.0370101928711,82.1119155883789,82.2876968383789,82.4513626098633,82.6298828125,82.6773452758789,82.7108383178711,82.7333984375,82.9328155517578,83.0640640258789,83.2138671875,83.2897415161133,83.3694305419922,83.3839874267578,83.4471664428711,83.5516586303711,83.7469711303711,83.8288116455078,83.8971633911133,84.0248031616211,84.0910110473633,84.2297821044922,84.4808578491211,84.6101531982422,84.6407241821289,84.65478515625,84.65380859375,84.6853485107422,84.9372100830078,85.0201187133789,85.0873107910156,85.1253890991211,85.1515655517578,85.1741180419922,85.1917953491211,85.240234375,85.29296875,85.4564437866211,85.5684585571289,85.6484375,85.6998977661133,85.7074203491211,85.7373046875,85.7945327758789,85.8556594848633,86.00732421875,86.12939453125,86.2416000366211,86.3661193847656,86.4144515991211,86.5436553955078,86.7013702392578,86.7624969482422,87.0164031982422,87.0378952026367,87.0895462036133,87.1668014526367,87.2873992919922,87.41357421875,87.5130844116211,87.6333999633789,87.7488250732422,87.8492202758789,87.9951171875,88.0269546508789,88.0548782348633,88.1115264892578,88.1615219116211,88.1572265625,88.1110305786133,87.9931640625,87.984375,88.0241165161133,88.06787109375,88.1055679321289,88.14697265625,88.154296875,88.1502914428711,88.1097717285156,88.0989303588867,88.1089859008789,88.14111328125,88.2751922607422,88.4259796142578,88.4861297607422,88.5316390991211,88.5779266357422,88.62109375,88.7562561035156,88.8037109375,88.82861328125,88.8488311767578,88.8298797607422,88.7490234375,88.7648468017578,88.83251953125,88.8914108276367,88.8816375732422,88.7603530883789,88.73876953125,88.765625,88.8135681152344,88.8351516723633,88.8576202392578,88.9191360473633,89.0409240722656,89.1482467651367,89.3321228027344,89.3841781616211,89.474609375,89.5451202392578,89.5861358642578,89.6091766357422,89.6061553955078,89.6099548339844,89.7109375,89.7638702392578,89.9431610107422,90.1229553222656,90.2060546875,90.2423858642578,90.3458938598633,90.4476547241211,90.5598678588867,90.6203155517578,90.7396469116211,90.8557662963867,91.1338882446289,91.2865219116211,91.4267578125,91.4558563232422,91.517578125,91.6715850830078,91.7537078857422,91.8420944213867,91.8986282348633,91.9437484741211,91.9983367919922,92.0497055053711,92.0734329223633,92.0681610107422,92.0308532714844,91.9986343383789,91.9922866821289,92.0025405883789,92.0311508178711,92.0834045410156,92.044921875,91.9908218383789,91.9509811401367,91.8512725830078,91.7430648803711,91.6581039428711,91.5947265625,91.5792999267578,91.59765625,91.6258850097656,91.6319351196289,91.7126007080078,91.82470703125,91.9093780517578,91.9776382446289,92.1012725830078,92.1576156616211,92.2222671508789,92.25048828125,92.2701187133789,92.3410186767578,92.4148406982422,92.4806671142578,92.5466766357422,92.6643600463867,92.6877899169922,92.6875,92.6656265258789,92.6434555053711,92.6525421142578,92.7018508911133,92.8818359375,93.0349655151367,93.1192398071289,93.1578140258789,93.20654296875,93.251953125,93.3605422973633,93.6649398803711,93.7607421875,93.9022445678711,93.9736328125,94.0132751464844,94.0176773071289,94.1115264892578,94.1934585571289,94.2932586669922,94.4680633544922,94.623046875,94.6770477294922,94.7333984375,94.7630844116211,94.7694320678711,94.9674835205078,94.9988250732422,95.1447296142578,95.2791061401367,95.3531188964844,95.3892593383789,95.4202117919922,95.45654296875,95.4937515258789,95.5169906616211,95.5158157348633,95.7103500366211,95.8850555419922,96.0353469848633,96.07958984375,96.1285095214844,96.1947250366211,96.2349624633789,96.3372116088867,96.3558654785156,96.3397445678711,96.2705078125,96.1808624267578,96.1223678588867,96.1414108276367,96.1371154785156,96.1622085571289,96.3468704223633,96.4357376098633,96.4670944213867,96.4771423339844,96.5499954223633,96.5808639526367,96.3956069946289,96.3273468017578,96.3298797607422,96.326171875,96.2789077758789,96.2814483642578,96.31982421875,96.3664093017578,96.3890609741211,96.427734375,96.6026382446289,96.65283203125,96.7757797241211,96.8330078125,96.9808578491211,97.0753860473633,97.1451187133789,97.2894515991211,97.3224639892578,97.3102493286133,97.302734375,97.3391647338867,97.3435592651367,97.3351516723633,97.30615234375,97.22607421875,97.1578140258789,97.0497055053711,96.9627914428711,96.8997116088867,96.8768539428711,96.8835983276367,96.9019470214844,97.1037139892578,97.10205078125,97.0380859375,96.9534149169922,96.8802719116211,96.7978515625,96.7316360473633,96.6657180786133,96.2742233276367,96.1908187866211,96.0614242553711,95.9709014892578,95.9052734375,95.8373107910156,95.7383804321289,95.4638671875,95.3050765991211,95.2014617919922,95.1287078857422,95.0894470214844,95.0597686767578,95.0508804321289,95.0689468383789,95.1083984375,95.1292953491211,95.1324234008789,95.0929641723633,95.0407257080078,95.0152359008789,94.9919891357422,94.9457015991211,94.8611297607422,94.7858428955078,94.6677780151367,94.6228485107422,94.5798797607422,94.5543899536133,94.5530242919922,94.5665054321289,94.6156234741211,94.67529296875,94.7037048339844,94.7076110839844,94.6632766723633,94.5840835571289,94.4931640625,94.3994140625,94.3772430419922,94.2930603027344,94.2197265625,94.1703109741211,94.1276397705078,94.0748062133789,94.0108413696289,93.85546875,93.755859375,93.6834030151367,93.63330078125,93.5640640258789,93.4937515258789,93.4521484375,93.3555679321289,93.3262710571289,93.3073272705078,93.37255859375,93.4149398803711,93.4081039428711,93.3913040161133,93.3660125732422,93.3494110107422,93.3080062866211,93.2535171508789,93.2039108276367,93.1641616821289,93.1509704589844,93.1625061035156,93.1142578125,93.0787124633789,93.0881805419922,93.1050720214844,93.1620101928711,93.1623992919922,93.1511688232422,93.1214904785156,93.0706024169922,93.04296875,93.02197265625,92.9645538330078,92.9094696044922,92.8542938232422,92.7713851928711,92.7210006713867,92.68896484375,92.6747055053711,92.6526336669922,92.63037109375,92.5749053955078,92.5612258911133,92.5318374633789,92.5095748901367,92.4914093017578,92.4644546508789,92.4304733276367,92.3931655883789,92.3616256713867,92.3412170410156,92.3337860107422,92.3341751098633,92.2893600463867,92.24609375,92.1871109008789,92.15234375,92.1270523071289,92.0440444946289,91.978515625,91.9295959472656,91.9294891357422,91.9378890991211,91.9191436767578,91.7900390625,91.7542037963867,91.7579116821289,91.7738265991211,91.7509765625,91.6949234008789,91.6195297241211,91.5535125732422,91.51123046875,91.4713897705078,91.4362258911133,91.3994140625,91.37060546875,91.3667984008789,91.36865234375,91.359375,91.3388671875,91.3152389526367,91.2538070678711,91.16552734375,91.1604537963867,91.1924819946289,91.2320327758789,91.33642578125,91.3501968383789,91.3670959472656,91.3926773071289,91.5263671875,91.5713882446289,91.6111373901367,91.6687469482422,91.7265625,91.7724609375,91.84619140625,91.876953125,91.8990173339844,91.9310531616211,91.95166015625,92.0010757446289,92.0641632080078,92.0850601196289,92.1019515991211,92.1174774169922,92.1980438232422,92.2266540527344,92.2305679321289,92.2283172607422,92.2512664794922,92.3849639892578,92.4431610107422,92.4750061035156,92.4854431152344,92.4683609008789,92.3734359741211,92.2046813964844,92.0497055053711,91.7634735107422,91.4796905517578,91.3966827392578,91.2931594848633,91.0382766723633,90.7301788330078,90.6130905151367,90.5552749633789,90.4393539428711,90.2503890991211,90.11962890625,90.0038070678711,89.8663101196289,89.8332977294922,89.8140640258789,89.8008804321289,89.7962951660156,89.8249053955078,89.7996139526367,89.8229522705078,89.7098617553711,89.6708984375,89.6190414428711,89.5857467651367,89.57275390625,89.5914077758789,89.5498962402344,89.4668960571289,89.3697280883789,89.2892608642578,89.1864242553711,89.1082992553711,89.1019592285156,89.0667953491211,89.0186538696289,88.9833984375,88.9519500732422,88.9241180419922,88.9482421875,88.9815444946289,88.9704132080078,88.9407196044922,88.896484375,88.8280334472656,88.7619171142578,88.72216796875,88.6828155517578,88.6806640625,88.6201171875,88.5182647705078,88.4181671142578,88.3699264526367,88.3459014892578,88.3514709472656,88.38623046875,88.4367141723633,88.4478454589844,88.4404296875,88.3780212402344,88.333984375,88.2351531982422,88.1507797241211,88.1290054321289,88.0973663330078,88.0845718383789,88.1066436767578,88.1474609375,88.2529296875,88.3630828857422,88.4523468017578,88.50244140625,88.5934524536133,88.7691421508789,88.79541015625,88.8203125,88.8547821044922,88.9441375732422,88.95166015625,88.9297866821289,88.89013671875,88.8172836303711,88.74755859375,88.6775436401367,88.5738220214844,88.4562530517578,88.3729476928711,88.3133773803711,88.2794952392578,88.1888656616211,88.1498031616211,88.0451202392578,88.0302734375,88.0234375,88.0791015625,88.1455078125,88.2249984741211,88.287109375,88.3375015258789,88.39697265625,88.49853515625,88.6422805786133,88.7235336303711,88.7335968017578,88.7265625,88.7137680053711,88.6998062133789,88.62255859375,88.5673828125,88.5959930419922,88.6164016723633,88.6357421875,88.6976547241211,88.7408218383789,88.7040023803711,88.7244186401367,88.8076171875,88.8970718383789,88.9281234741211,88.8505859375,88.8669891357422,88.8997039794922,88.9234313964844,88.9269561767578,88.9207000732422,88.9714889526367,89.0499954223633,89.0558624267578,89.0514678955078,89.0279312133789,88.9493179321289,89.0196304321289,89.0419921875,89.0516586303711,88.9670867919922,88.9074249267578,88.8575134277344,88.8343734741211,88.7450180053711,88.7129898071289,88.6947250366211,88.6912078857422,88.740234375,88.7302703857422,88.7082977294922,88.6595687866211,88.6416015625,88.5667953491211,88.5998077392578,88.5846710205078,88.4459991455078,88.3054656982422,88.2874984741211,88.2791976928711,88.2537078857422,88.1220703125,88.0568313598633,88.0994110107422,88.1810531616211,88.1962890625,88.0871124267578,87.9944305419922,87.94140625,87.9616241455078,88.0107421875,88.0830078125,88.1592788696289,88.1040954589844,88.05078125,87.9484405517578,87.82373046875,87.67822265625,87.20068359375,87.1006851196289,86.9541015625,86.8595733642578,86.84228515625,86.8957977294922,86.9393539428711,86.9754867553711,86.9245147705078,86.8359375,86.7624969482422,86.7692413330078,86.7503890991211,86.4987335205078,86.44580078125,86.3765563964844,86.2936553955078,86.2452163696289,86.3119125366211,86.3029327392578,86.2794876098633,86.2162170410156,85.8529281616211,85.5750045776367,85.4968719482422,85.5111312866211,85.5597610473633,85.5550765991211,85.5040969848633,85.4599609375,85.2486343383789,85.1627960205078,85.1807632446289,85.228515625,85.3708953857422,85.4369201660156,85.4416046142578,85.2255859375,84.77099609375,84.7498092651367,84.6908187866211,84.609375,84.4627990722656,84.1817398071289,84.1041030883789,83.654296875,83.572265625,83.3879928588867,83.1983413696289,82.9768524169922,82.5931625366211,82.3595657348633,82.2865295410156,82.2819366455078,82.3072280883789,82.3499984741211,82.3597640991211,82.3386764526367,82.3271484375,82.2587890625,82.1415023803711,81.7619094848633,81.7117156982422,81.40185546875,81.2861328125,81.2385787963867,81.1321258544922,81.0300827026367,80.9934539794922,80.9787063598633,80.9177703857422,80.86474609375,80.8259811401367,80.7818374633789,80.7078094482422,80.6465835571289,80.3848648071289,80.29345703125,80.10107421875,80.0534133911133,80.0986328125,80.1654281616211,80.1787109375,80.1701202392578,80.13623046875,80.1117172241211,80.1436538696289,80.2244186401367,80.244140625,80.2458038330078,80.3065414428711,80.265625,80.2333984375,80.15625,80.0621109008789,80.1142578125,80.2903366088867,80.3423843383789,80.2291030883789,80.14306640625,80.0374984741211,79.9817428588867,79.8584976196289,79.7713851928711,79.7541046142578,79.7933578491211,79.7489242553711,79.6931686401367,79.7990264892578,79.8352508544922,79.8486328125,79.8501968383789,79.8381805419922,79.7569351196289,79.6673812866211,79.5885772705078,79.5316390991211,79.3905258178711,79.3145523071289,79.2536163330078,79.2578125,78.9962844848633,78.93994140625,78.9191436767578,78.953125,79.0199203491211,79.1070327758789,79.2754821777344,79.3563461303711,79.4114303588867,79.212890625,78.9795913696289,78.4214859008789,78.2745132446289,78.1924743652344,78.1360321044922,78.1263656616211,78.0601577758789,77.7703094482422,77.5872039794922,77.517578125,77.3014678955078,77.06591796875,76.9668960571289,76.6172866821289,76.5534133911133,76.48291015625,76.4717712402344,76.4523391723633,76.4190444946289,76.4031219482422,76.3246078491211,76.2923812866211,76.2423858642578,76.28466796875,76.3430633544922,76.3722686767578,76.3755874633789,76.4195327758789,76.4587936401367,76.3464813232422,76.2487258911133,76.2227554321289,76.1956024169922,76.1926727294922,76.2014617919922,76.1233367919922,76.0960922241211,75.9225616455078,75.8446273803711,75.7238311767578,75.6460952758789,75.5245056152344,75.4226608276367,75.3146514892578,75.2297897338867,75.1966781616211,74.9455108642578,74.8682632446289,74.8029251098633,74.7705078125,74.6823272705078,74.681640625,74.6708984375,74.6084976196289,74.49853515625,74.4666976928711,74.4669876098633,74.3971633911133,74.3822250366211,74.3350601196289,74.2803726196289,74.2230453491211,74.0887680053711,74.0406265258789,73.94921875,73.88427734375,73.80078125,73.9319381713867,73.8519515991211,73.8138656616211,73.7717819213867,73.8328170776367,73.7328109741211,73.6798782348633,73.6077117919922,73.47607421875,73.4537124633789,73.3376007080078,73.2391586303711,73.1490249633789,73.1560516357422,73.0471649169922,72.9939422607422,72.9720687866211,72.9431610107422,72.9171905517578,72.87548828125,72.8708953857422,72.8987350463867,72.9768524169922,73.0055694580078,72.9720687866211,72.9006881713867,72.8346710205078,72.8030242919922,72.802734375,72.7945327758789,72.8116226196289,72.9872055053711,72.7878952026367,72.7639617919922,72.7564468383789,72.7994155883789,72.7265625,72.6974563598633,72.6759796142578,72.6677780151367,72.708984375,72.8811492919922,72.8937530517578,72.87890625,72.8405227661133,72.8243179321289,72.8138656616211,72.7515640258789,72.6923828125,72.6238327026367,72.6865234375,72.7347640991211,72.6683578491211,72.61328125,72.7175750732422,72.810546875,73.0224609375,73.1125030517578,72.9791030883789,72.8397445678711,72.5430603027344,72.5924835205078,72.64404296875,72.7001953125,72.6174774169922,72.5222625732422,72.5530319213867,72.6279296875,72.7087936401367,72.8091812133789,72.7019577026367,72.5901336669922,72.4559555053711,72.3326187133789,72.1828155517578,72.2425765991211,72.3064422607422,72.2744140625,72.2443389892578,72.1617202758789,72.0944366455078,72.0755844116211,72.0372085571289,72.1029281616211,72.1708984375,72.2103500366211,72.2566375732422,72.2540054321289,72.0765609741211,72.0152359008789,71.5710906982422,71.396484375,71.0246124267578,70.8796844482422,70.7193374633789,70.4850616455078,70.1273422241211,70.0343780517578,69.7484359741211,69.5419921875,69.3854446411133,69.1917037963867,69.0087890625,68.9699249267578,68.9834976196289,69.0516586303711,69.13134765625,69.1942367553711,69.2388610839844,69.2765579223633,69.5492172241211,69.6551742553711,69.7275390625,69.8190383911133,70.005859375,70.0841827392578,70.17724609375,70.3277359008789,70.4404296875,70.5134811401367,70.5093765258789,70.4892578125,70.4345703125,70.3962860107422,70.3679656982422,70.3394546508789,70.2511749267578,70.1917037963867,70.1182632446289,69.8498001098633,69.7396469116211,69.6646499633789,69.2359390258789,68.8170928955078,68.6407241821289,68.5291976928711,68.41748046875,68.4538116455078,68.6271438598633,68.7767639160156,68.6423797607422,68.4968719482422,68.4249038696289,68.3433609008789,68.2349624633789,68.1919937133789,68.1650390625,68.2341766357422,68.2825241088867,68.3812484741211,68.4886703491211,68.5866241455078,68.72412109375,68.7281265258789,68.7396469116211,68.7589797973633,68.7811508178711,68.7999954223633,68.8283233642578,68.8634796142578,68.9007797241211,68.9845657348633,69.0515594482422,69.1195373535156,69.2350616455078,69.4434585571289,69.5591812133789,69.6341781616211,69.7162094116211,69.80517578125,69.9337844848633,70.0210876464844,70.0651397705078,70.0982437133789,70.2890625,70.4892578125,70.5467758178711,70.5650405883789,70.5558624267578,70.5792922973633,70.6594772338867,70.71630859375,70.7672805786133,70.8050842285156,70.88623046875,70.9281234741211,70.9828109741211,71.0440444946289,71.0453109741211,71.0062484741211,70.9732437133789,70.9793014526367,70.9698257446289,70.9763717651367,71.0023422241211,71.0478515625,71.0207061767578,70.9508743286133,70.8777313232422,70.8004913330078,70.7025375366211,70.6520538330078,70.6572265625,70.6484375,70.6148452758789,70.5695266723633,70.505859375,70.4485397338867,70.3251953125,70.2646484375,70.1001892089844,70.07861328125,70.0777359008789,70.1326141357422,70.1492156982422,70.1568374633789,70.1476516723633,70.1146545410156,70.0593719482422,69.9114227294922,69.7359390258789,69.6005859375,69.5069351196289,69.4812469482422,69.4700164794922,69.4945297241211,69.5370178222656,69.5679702758789,69.62158203125,69.6613311767578,69.7248077392578,69.8962936401367,70.0498046875,70.14453125,70.1939392089844,70.2443389892578,70.3184585571289,70.4037094116211,70.4885787963867,70.5692443847656,70.6291046142578,70.6491241455078,70.6916046142578,70.7374038696289,70.7979507446289,70.8749008178711,71.1847610473633,71.2901306152344,71.54296875,71.7166976928711,71.8703079223633,71.8888702392578,71.9480438232422,72.1285171508789,72.17919921875,72.23388671875,72.2919921875,72.3418960571289,72.6255874633789,72.9033203125,72.94873046875,73.1283187866211,73.2311477661133,73.2578125,73.3172836303711,73.3816375732422,73.4674835205078,73.6580123901367,73.8091812133789,73.8865203857422,73.9333953857422,73.9246063232422,73.8827133178711,73.8916015625,73.8993148803711,74.0089874267578,74.2156295776367,74.33935546875,74.38037109375,74.509765625,74.6328125,74.6257781982422,74.6103515625,74.5397491455078,74.5176773071289,74.5349578857422,74.5939407348633,74.5818328857422,74.5099563598633,74.5259780883789,74.5555648803711,74.6357421875,74.7394561767578,75.0714797973633,75.1387710571289,75.2541046142578,75.32470703125,75.33349609375,75.3026351928711,75.2336959838867,75.1041030883789,74.9873046875,74.7888717651367,74.6857452392578,74.6578140258789,74.6433639526367,74.6632843017578,74.6324234008789,74.5882797241211,74.4833984375,74.3545837402344,74.3054733276367,74.3299789428711,74.32275390625,74.3036117553711,74.2835922241211,74.2220687866211,74.1262664794922,74.0491180419922,74.0038070678711,73.9898376464844,73.9942398071289,74.0503921508789,74.1177673339844,74.142578125,74.1500015258789,74.1312561035156,74.0697250366211,74.0040054321289,73.9775390625,73.9764709472656,74.0009765625,74.0784149169922,74.2156295776367,74.2508773803711,74.2464828491211,74.208984375,74.1126022338867,73.9499053955078,73.92236328125,73.9040985107422,73.9039001464844,73.9382781982422,73.9794921875,73.9723663330078,73.9246063232422,73.8099594116211,73.7945327758789,73.8121109008789,73.85009765625,73.8831024169922,73.9612274169922,74.0558624267578,74.1719741821289,74.3003921508789,74.4979553222656,74.5941390991211,74.7887649536133,74.9518508911133,75.1184539794922,75.1875,75.2640609741211,75.4525375366211,75.6055603027344,75.7091751098633,75.8621139526367,75.9382858276367,76.041015625,76.1724624633789,76.4567413330078,76.5099639892578,76.5944290161133,76.6962890625,76.7490234375,76.7575225830078,76.7829132080078,76.8917007446289,77.0008850097656,77.0306625366211,77.0486297607422,77.1685562133789,77.29296875,77.4234390258789,77.5715789794922,77.6969757080078,77.7994155883789],[96.9182586669922,96.9065399169922,96.8967819213867,96.8939437866211,96.8929672241211,96.9043960571289,96.9133758544922,96.9189453125,96.9205093383789,96.92529296875,96.92431640625,96.9182586669922],[96.8404312133789,96.8519515991211,96.8673858642578,96.8736343383789,96.8495178222656,96.8348617553711,96.8277359008789,96.8258819580078,96.8326187133789,96.8326187133789,96.8341827392578,96.8394546508789,96.8356399536133,96.8404312133789],[105.725395202637,105.696876525879,105.644340515137,105.584083557129,105.595703125,105.6455078125,105.669822692871,105.705467224121,105.725395202637],[72.4919967651367,72.46875,72.4291000366211,72.4076156616211,72.3497085571289,72.3728485107422,72.4274444580078,72.447265625,72.4671859741211,72.4622039794922,72.4737319946289,72.4654312133789,72.4357376098633,72.4456024169922,72.4935607910156,72.49853515625,72.4919967651367],[-9.94819355010986,-9.95244121551514,-10.0265140533447,-10.0625,-10.2657222747803,-10.1810550689697,-10.1397466659546,-9.99638652801514,-9.95615291595459,-9.94819355010986],[-6.218017578125,-6.17573213577271,-6.15693378448486,-6.23066377639771,-6.3076171875,-6.34516572952271,-6.34760713577271,-6.32158231735229,-6.27011728286743,-6.22900390625,-6.19487333297729,-6.141845703125,-6.13095712661743,-6.13876962661743,-6.12910127639771,-6.15165996551514,-6.13471698760986,-6.072265625,-6.04501914978027,-6.02739286422729,-6.07148408889771,-6.13066387176514,-6.16933536529541,-6.19921875,-6.21723604202271,-6.34540987014771,-6.39995145797729,-6.46318340301514,-6.32500028610229,-6.43793964385986,-6.56108379364014,-6.69731473922729,-6.78222608566284,-6.85971641540527,-6.89023447036743,-6.91464853286743,-6.96577167510986,-7.00327157974243,-7.081787109375,-7.21621084213257,-7.44086933135986,-7.52729463577271,-7.56318378448486,-7.58984375,-7.62490224838257,-7.66455078125,-7.83798789978027,-7.87216758728027,-7.95248985290527,-8.05781269073486,-8.14501953125,-8.22246074676514,-8.25429725646973,-8.29023456573486,-8.4091796875,-8.37163066864014,-8.34736251831055,-8.33559513092041,-8.34912109375,-8.40781307220459,-8.47783184051514,-8.58828067779541,-8.73447322845459,-8.81342792510986,-9.29648399353027,-9.32387638092041,-9.39057636260986,-9.462890625,-9.53486251831055,-9.7373046875,-9.83535194396973,-9.71035194396973,-9.54238319396973,-9.52490329742432,-9.579833984375,-9.89902305603027,-10.0099124908447,-10.1207513809204,-10.0694341659546,-9.92641544342041,-9.84970760345459,-9.80288124084473,-9.74951171875,-9.59882831573486,-10.084228515625,-10.2117185592651,-10.2417478561401,-10.341064453125,-10.3787107467651,-10.2315921783447,-10.1458501815796,-10.0440435409546,-9.946044921875,-9.90966796875,-9.955810546875,-10.24951171875,-10.3902339935303,-10.3826169967651,-10.356689453125,-10.2109375,-10.132080078125,-10.061767578125,-9.99311447143555,-9.93730449676514,-9.77211856842041,-9.841064453125,-9.85322284698486,-9.90605449676514,-9.83847713470459,-9.76113224029541,-9.63222599029541,-9.58632850646973,-9.33125019073486,-9.05615329742432,-8.783447265625,-8.92329120635986,-8.99028301239014,-9.097900390625,-9.17539119720459,-9.39423847198486,-9.46347618103027,-9.56103420257568,-9.59135818481445,-9.61953067779541,-9.76435565948486,-9.91660118103027,-9.73959922790527,-9.51499080657959,-9.46489238739014,-9.39365196228027,-9.41572189331055,-9.46196365356445,-9.29921913146973,-9.24189472198486,-9.13759708404541,-9.06113338470459,-9.02744102478027,-8.99716758728027,-8.93012714385986,-9.03354454040527,-9.14033222198486,-9.47075176239014,-9.51420879364014,-9.55517578125,-9.58173847198486,-9.60175800323486,-9.62597560882568,-9.70058536529541,-9.77407264709473,-9.82539081573486,-9.87578201293945,-9.79541015625,-9.89902305603027,-10.0039052963257,-10.0912599563599,-10.093994140625,-10.0543947219849,-10.1062498092651,-10.1169919967651,-10.0617189407349,-10.0013675689697,-9.87827110290527,-9.72065448760986,-9.85585975646973,-9.90971755981445,-9.91230392456055,-9.901611328125,-9.74506759643555,-9.57822227478027,-9.59052753448486,-9.578857421875,-9.74750995635986,-9.9140625,-9.896240234375,-9.85634803771973,-9.84848690032959,-9.8564453125,-9.93447303771973,-9.943603515625,-9.97709941864014,-10.0926761627197,-10.0896968841553,-10.056396484375,-9.99594688415527,-9.93593788146973,-9.82456016540527,-9.71713829040527,-9.56230449676514,-9.31552696228027,-9.14589786529541,-9.10209941864014,-9.03427791595459,-9.00244140625,-8.74677658081055,-8.58803653717041,-8.54555702209473,-8.56845664978027,-8.62314414978027,-8.554443359375,-8.47099590301514,-8.41523456573486,-8.28652381896973,-8.23037147521973,-8.19296932220459,-8.13344764709473,-8.45654296875,-8.763916015625,-8.71518516540527,-8.65029239654541,-8.53828144073486,-8.52768611907959,-8.47099590301514,-8.37729549407959,-8.41171836853027,-8.39326190948486,-8.32578086853027,-8.3046875,-8.27460956573486,-8.1376953125,-8.006103515625,-7.95859384536743,-7.80317449569702,-7.75053644180298,-7.76254940032959,-7.66708946228027,-7.62978506088257,-7.61337900161743,-7.57001972198486,-7.556640625,-7.585693359375,-7.63427734375,-7.58984375,-7.65873956680298,-7.58437538146973,-7.47841835021973,-7.48393583297729,-7.50195264816284,-7.53144550323486,-7.51787042617798,-7.45830106735229,-7.30175828933716,-7.36596632003784,-7.30878925323486,-7.24667978286743,-7.15532207489014,-7.06025409698486,-6.961669921875,-7.056396484375,-7.17285203933716,-7.21865177154541,-7.37690401077271,-7.40141582489014,-7.44599628448486,-7.45126914978027,-7.50219678878784,-7.55039072036743,-7.6064453125,-7.68999052047729,-7.73749971389771,-7.79726552963257,-7.87295007705688,-7.91059541702271,-7.90874004364014,-7.88613319396973,-7.81982469558716,-7.74628925323486,-7.75439500808716,-7.79379892349243,-8.04433631896973,-8.11894512176514,-8.14482402801514,-8.11826229095459,-7.91845703125,-7.88447237014771,-7.85493183135986,-7.67876005172729,-7.60654258728027,-7.54443407058716,-7.40942335128784,-7.35517549514771,-7.32451200485229,-7.30673837661743,-7.19306612014771,-7.15546894073486,-7.17807626724243,-7.20258712768555,-7.13349580764771,-7.04970741271973,-7.00771522521973,-6.93613243103027,-6.87724637985229,-6.86923885345459,-6.85834980010986,-6.80258798599243,-6.76660203933716,-6.66953134536743,-6.64687442779541,-6.66420936584473,-6.64980459213257,-6.54814481735229,-6.44028377532959,-6.402587890625,-6.36367177963257,-6.30366230010986,-6.218017578125],[56.18798828125,56.0949211120605,55.9542961120605,55.8941383361816,55.84765625,55.7472648620605,55.6746101379395,55.5431632995605,55.4237289428711,55.3404312133789,55.3115234375,55.2950172424316,55.2964859008789,55.3469734191895,55.53173828125,55.7625961303711,55.7845687866211,55.7474594116211,55.7576179504395,55.9071311950684,56.0741233825684,56.2139663696289,56.2793960571289,56.18798828125],[46.1144523620605,46.1701202392578,46.3177719116211,46.4906234741211,46.5547828674316,46.783203125,46.8525390625,46.9888687133789,47.0654296875,47.1883811950684,47.3384780883789,47.4761734008789,47.5818367004395,47.7728500366211,47.8922843933105,47.9958992004395,48.1510734558105,48.28173828125,48.3221664428711,48.2572250366211,48.1360359191895,48.1128921508789,48.1043930053711,48.1091766357422,48.12548828125,48.2741203308105,48.291015625,48.2920913696289,48.2750968933105,48.2419929504395,48.1385765075684,48.0500984191895,48.0193367004395,47.99267578125,47.9964828491211,48.0232429504395,48.2046890258789,48.2251968383789,48.2613296508789,48.3055686950684,48.3812484741211,48.4173851013184,48.5926742553711,48.6355476379395,48.84033203125,48.8687515258789,48.8707046508789,48.9013671875,48.9250984191895,48.9599609375,49.0153312683105,49.0809555053711,49.1711921691895,49.3724594116211,49.4701194763184,49.7269515991211,49.9806632995605,50.1304702758789,50.17626953125,50.2140617370605,50.337890625,50.533203125,50.9274406433105,51.1185569763184,51.7620124816895,52.1901397705078,53.3741188049316,53.7676773071289,53.9154319763184,53.8274383544922,53.6794929504395,53.7687492370605,53.90625,53.9701156616211,54.0162086486816,54.0238304138184,54.0171852111816,53.9519538879395,53.9141578674316,54.1916007995605,54.2998046875,54.4586906433105,54.5789070129395,54.6396484375,54.6994132995605,54.7452163696289,54.8486328125,54.9000968933105,55.0755844116211,55.2247047424316,55.380859375,55.5784187316895,55.84130859375,56.05029296875,56.1711921691895,56.2288093566895,56.2720718383789,56.2969741821289,56.3241195678711,56.3668937683105,56.4406280517578,56.5440406799316,56.669921875,56.7746086120605,56.9066390991211,57.0790023803711,57.1935539245605,57.2601585388184,57.3081092834473,57.3314476013184,57.3367195129395,57.3357429504395,57.3537139892578,57.4238243103027,57.5209999084473,57.7105484008789,57.88818359375,57.9805679321289,58.1087913513184,58.2616233825684,58.3181648254395,58.38671875,58.4357452392578,58.5504837036133,58.6501922607422,58.7007827758789,58.8154296875,58.9372024536133,59.2408180236816,59.2741241455078,59.3017578125,59.3269538879395,59.3447265625,59.3673820495605,59.4549789428711,59.5622062683105,59.6872062683105,59.9486312866211,60.0627899169922,60.1783180236816,60.3207054138184,60.3413047790527,60.7079086303711,61.1196250915527,61.1699256896973,61.1603469848633,61.1750946044922,61.2120132446289,61.21240234375,61.1826133728027,61.1594734191895,61.1529312133789,61.2058563232422,61.2521476745605,61.2586898803711,61.2355499267578,61.2388687133789,61.2620086669922,61.2818374633789,61.2785186767578,61.2455062866211,61.2256851196289,61.19921875,61.1892623901367,61.1396446228027,61.0999984741211,61.1066436767578,61.12646484375,61.1496124267578,61.1231460571289,61.1066436767578,61.0702171325684,61.080078125,61.0404319763184,60.9908218383789,60.9578132629395,60.9511756896973,60.9147491455078,60.8453140258789,60.8023452758789,60.7394523620605,60.7262687683105,60.7361335754395,60.7625999450684,60.8039093017578,60.8894577026367,60.6426734924316,60.5702133178711,60.4857444763184,60.5270538330078,60.4859352111816,60.5108375549316,60.5738258361816,60.6545867919922,60.8064422607422,60.9069328308105,60.9169883728027,60.8592796325684,60.7668952941895,60.7180671691895,60.560546875,60.5619163513184,60.5765647888184,60.64453125,60.71044921875,60.8292961120605,60.8272476196289,60.7899398803711,60.7875022888184,60.8042984008789,60.7915992736816,60.8207054138184,60.8541030883789,61.1107406616211,61.3464851379395,61.6601524353027,61.7550811767578,61.8142585754395,61.8108406066895,61.7841758728027,61.5594711303711,61.3316421508789,61.1041030883789,60.8433609008789,61.0341796875,61.1521453857422,61.318359375,61.3394546508789,61.337890625,61.5085906982422,61.5687522888184,61.623046875,61.7580108642578,61.8898468017578,62.0330085754395,62.1305618286133,62.35302734375,62.4338836669922,62.5645523071289,62.7175750732422,62.7494125366211,62.7580032348633,62.7625007629395,62.73974609375,62.7823257446289,62.81201171875,62.8008766174316,62.7642555236816,62.7625007629395,62.7527313232422,62.7629890441895,62.8116226196289,62.9154281616211,63.1667976379395,63.1960945129395,63.2562446594238,63.3015632629395,63.3051795959473,63.2420883178711,63.2314453125,63.2503929138184,63.2416038513184,63.1861343383789,63.1680679321289,63.1578102111816,63.0929718017578,62.7866249084473,62.7515640258789,62.6364288330078,62.4392585754395,62.3850555419922,62.3123092651367,62.2596664428711,62.2496070861816,62.2393531799316,62.1259727478027,62.0890617370605,61.8698272705078,61.8423843383789,61.8099594116211,61.78076171875,61.7543983459473,61.7376976013184,61.6686515808105,61.6618156433105,61.6713905334473,61.64013671875,61.6154289245605,61.587890625,61.5331077575684,61.4903335571289,61.4122047424316,61.2429656982422,61.1085968017578,60.6638679504395,60.6151351928711,60.5875015258789,60.5105438232422,60.4001960754395,60.0247077941895,59.8970718383789,59.818359375,59.6160163879395,59.4560508728027,59.2272491455078,59.0460891723633,58.7978553771973,58.5308609008789,58.3142547607422,58.2029304504395,58.0223617553711,57.9366226196289,57.7960891723633,57.7325210571289,57.3345680236816,57.2609405517578,57.20556640625,57.2013664245605,57.1042976379395,57.0719718933105,57.0360336303711,56.9822273254395,56.9104499816895,56.8128929138184,56.7281265258789,56.3561553955078,56.2843742370605,56.1180686950684,55.9411125183105,55.6502914428711,55.5916023254395,55.5185546875,55.4240226745605,55.2941398620605,55.1545906066895,54.8958015441895,54.75927734375,54.6449203491211,54.5220718383789,54.2470703125,54.0693359375,53.8225593566895,53.7057609558105,53.5071296691895,53.4549789428711,53.3416976928711,52.9825172424316,52.6916007995605,52.63818359375,52.6026382446289,52.4758796691895,52.19189453125,52.03076171875,51.8419914245605,51.6663093566895,51.5890655517578,51.5185546875,51.2789039611816,51.2760734558105,51.12841796875,51.0938491821289,51.06201171875,51.0211906433105,50.8669891357422,50.8429679870605,50.8757820129395,50.8757820129395,50.7955093383789,50.6751937866211,50.6460952758789,50.66796875,50.6496086120605,50.5435523986816,50.3869132995605,50.2301750183105,50.1689453125,50.12890625,50.0715789794922,49.9831047058105,49.5548820495605,49.4299812316895,49.0542945861816,49.0281257629395,49.001953125,49.0490226745605,49.09619140625,49.1903343200684,49.2472648620605,49.2245101928711,49.13037109375,49.001953125,49.037109375,48.9167976379395,48.8912124633789,48.90869140625,48.9191398620605,48.8701171875,48.8324203491211,48.6708984375,48.5955085754395,48.5464859008789,48.478515625,48.4345703125,48.3986320495605,48.3875961303711,48.4013671875,48.3826179504395,48.3310546875,48.2789039611816,48.2261734008789,48.1824226379395,48.1475563049316,48.0661125183105,48.0149421691895,48.0134773254395,48.0120124816895,48.0106430053711,47.8363304138184,47.6794929504395,47.6794929504395,47.6794929504395,47.75390625,47.8299827575684,47.7145500183105,47.5915031433105,47.5119132995605,47.4181632995605,47.3712882995605,47.3297843933105,47.28515625,47.1213874816895,46.9685554504395,46.7890625,46.5699195861816,46.3770484924316,46.2982444763184,46.11279296875,46.0930671691895,46.0804672241211,46.0807609558105,46.14111328125,46.1458969116211,46.0199241638184,45.9810562133789,45.8732414245605,45.8947257995605,45.8947257995605,45.87939453125,45.8544921875,45.8228530883789,45.73828125,45.6737327575684,45.4732398986816,45.4089851379395,45.3970718383789,45.4460945129395,45.5286140441895,45.5427742004395,45.52685546875,45.4375991821289,45.4593772888184,45.4977531433105,45.5007820129395,45.5608406066895,45.6375007629395,45.6615219116211,45.6600570678711,45.6781272888184,45.9208984375,46.0417976379395,46.1337890625,46.1546859741211,46.1357421875,46.1177711486816,46.1121101379395,46.0106430053711,45.9753913879395,45.9710922241211,45.9950180053711,46.0373992919922,46.1809577941895,46.2625007629395,46.2734375,46.16748046875,45.94140625,45.7763671875,45.7234382629395,45.64501953125,45.5616188049316,45.4837913513184,45.40771484375,45.3616218566895,45.3508796691895,45.2411155700684,45.20654296875,45.1552772521973,45.1124038696289,45.0837898254395,45.0531272888184,45.0310554504395,45.0293960571289,45.0339851379395,45.0192375183105,44.9814453125,44.9278297424316,44.880859375,44.7984390258789,44.7654304504395,44.76513671875,44.7667007446289,44.75830078125,44.7967758178711,44.7941398620605,44.7151374816895,44.6041030883789,44.5740242004395,44.5731468200684,44.5771484375,44.5671844482422,44.5460929870605,44.5453109741211,44.5899391174316,44.5612297058105,44.3977508544922,44.3362312316895,44.2229461669922,44.2113304138184,44.2289085388184,44.2679672241211,44.3293952941895,44.3489265441895,44.3727531433105,44.380859375,44.4496078491211,44.4499015808105,44.4308586120605,44.3757781982422,44.3196296691895,44.2985343933105,44.2908210754395,44.2978515625,44.2801742553711,44.2570304870605,44.2716827392578,44.232421875,44.1708030700684,44.14453125,44.1587867736816,44.171875,44.1805648803711,44.1780242919922,44.1212882995605,44.0791015625,44.0743141174316,44.0575218200684,44.0337905883789,44.0232429504395,44.0439453125,44.1240234375,44.2404289245605,44.33544921875,44.3893547058105,44.4559555053711,44.5167007446289,44.5871086120605,44.7250022888184,44.7821273803711,44.8171882629395,44.8381843566895,45.0001945495605,45.0716819763184,45.1130867004395,45.1412124633789,45.1906242370605,45.2559547424316,45.3355484008789,45.3892555236816,45.4796905517578,45.5750007629395,45.921875,46.1144523620605],[44.76513671875,44.7654304504395,44.7984390258789,44.880859375,44.9278297424316,44.9814453125,45.0192375183105,45.0339851379395,45.0293960571289,45.0310554504395,45.0531272888184,45.0837898254395,45.1124038696289,45.1552772521973,45.20654296875,45.2411155700684,45.3508796691895,45.3616218566895,45.40771484375,45.4837913513184,45.5616188049316,45.64501953125,45.7234382629395,45.7763671875,45.94140625,46.16748046875,46.2734375,46.2625007629395,46.1809577941895,46.0373992919922,45.9950180053711,45.9710922241211,45.9753913879395,46.0106430053711,46.1121101379395,46.1177711486816,46.1357421875,46.1546859741211,46.1337890625,46.0417976379395,45.9208984375,45.6781272888184,45.6600570678711,45.6615219116211,45.6375007629395,45.5608406066895,45.5007820129395,45.4977531433105,45.4593772888184,45.4375991821289,45.52685546875,45.5427742004395,45.5286140441895,45.4460945129395,45.3970718383789,45.4089851379395,45.4732398986816,45.6737327575684,45.73828125,45.8228530883789,45.8544921875,45.87939453125,45.8947257995605,45.8947257995605,45.8732414245605,45.9810562133789,46.0199241638184,46.1458969116211,46.14111328125,46.0807609558105,46.0804672241211,46.0930671691895,46.11279296875,46.2982444763184,46.3770484924316,46.5699195861816,46.7890625,46.9685554504395,47.1213874816895,47.28515625,47.3297843933105,47.3712882995605,47.4181632995605,47.5119132995605,47.5915031433105,47.7145500183105,47.8299827575684,47.75390625,47.6794929504395,47.6794929504395,47.6794929504395,47.8363304138184,48.0106430053711,48.0120124816895,48.0134773254395,48.0149421691895,48.0661125183105,48.1475563049316,48.1824226379395,48.2261734008789,48.2789039611816,48.3310546875,48.3826179504395,48.4013671875,48.3875961303711,48.3986320495605,48.4345703125,48.478515625,48.5464859008789,48.4541969299316,48.3545913696289,48.1417007446289,48.07275390625,47.9825210571289,47.9787101745605,47.75390625,47.6727523803711,47.6437492370605,47.5148429870605,47.3313446044922,47.2232398986816,47.1482429504395,47.1143531799316,47.10205078125,47.0436515808105,46.9759750366211,46.9058609008789,46.7693328857422,46.6937522888184,46.5314445495605,46.3564453125,45.94970703125,45.4989242553711,45.05029296875,44.7165031433105,44.6908187866211,44.3607444763184,44.099609375,43.7737312316895,43.4408187866211,43.1031227111816,42.8577156066895,42.5597648620605,42.28857421875,42.0744132995605,41.7997055053711,41.5850563049316,41.2724609375,41.0224609375,40.8083992004395,40.4789085388184,40.3693351745605,40.02783203125,39.7041015625,39.36865234375,39.1454086303711,39.2927742004395,39.2474632263184,39.1400413513184,39.0414047241211,38.9816398620605,39.0578117370605,38.9874038696289,38.9148406982422,38.8450202941895,38.7735366821289,39.0567398071289,39.2683601379395,39.564453125,39.8500022888184,40.1219711303711,40.4214820861816,40.689453125,40.93505859375,40.9870109558105,41.0990257263184,41.1947250366211,41.19921875,41.1996078491211,41.2164039611816,41.2483406066895,41.3033218383789,41.3540992736816,41.359375,41.3526344299316,41.3001937866211,41.24560546875,41.2517585754395,41.2618141174316,41.2959976196289,41.3541984558105,41.4167938232422,41.6501960754395,41.78857421875,41.9740219116211,42.083984375,42.2373046875,42.35009765625,42.3590812683105,42.3589859008789,42.4558601379395,42.6354484558105,42.7411155700684,42.7746086120605,42.869140625,42.9366226196289,43.0924797058105,43.1851539611816,43.2630844116211,43.3067398071289,43.5158233642578,43.5679702758789,43.67578125,43.83642578125,43.9400367736816,44.01318359375,44.0646476745605,44.1144523620605,44.15625,44.1917991638184,44.2084007263184,44.20166015625,44.2174797058105,44.2457008361816,44.2818374633789,44.3255844116211,44.4019546508789,44.4959945678711,44.5660133361816,44.60595703125,44.6693344116211,44.73095703125,44.76513671875],[-15.5431146621704,-15.428466796875,-15.2409181594849,-15.1624021530151,-14.9699716567993,-14.8561038970947,-14.6806631088257,-14.5958499908447,-14.5938959121704,-14.7016611099243,-14.7882318496704,-14.9122066497803,-15.0299816131592,-15.1173820495605,-15.1164054870605,-15.0103015899658,-14.8940439224243,-14.7871580123901,-14.7404289245605,-14.6981935501099,-14.6743659973145,-14.6689939498901,-14.6882333755493,-14.7525396347046,-14.8393068313599,-14.8270988464355,-14.7575197219849,-14.4262199401855,-14.391845703125,-14.372802734375,-14.3690919876099,-14.3508796691895,-14.3181638717651,-14.328369140625,-14.4733896255493,-14.3022947311401,-14.1669435501099,-13.9354486465454,-13.8407230377197,-13.7852535247803,-13.7051267623901,-13.6703119277954,-13.6178712844849,-13.6160154342651,-13.6544427871704,-13.6677742004395,-13.7080078125,-13.7832517623901,-13.8047847747803,-13.7716312408447,-13.7228517532349,-13.6534671783447,-13.64111328125,-13.6395502090454,-13.64892578125,-13.6715822219849,-13.7074213027954,-13.7548828125,-13.580810546875,-13.55859375,-13.5561037063599,-13.5696773529053,-13.5993165969849,-13.6518564224243,-13.7772464752197,-13.85400390625,-13.8278322219849,-13.8298349380493,-13.8529291152954,-13.95166015625,-14.04443359375,-14.1351566314697,-14.2969732284546,-14.3851070404053,-14.3752927780151,-14.4653806686401,-14.4483394622803,-14.4169921875,-14.4325685501099,-14.4753904342651,-14.5470705032349,-14.6282234191895,-14.78955078125,-14.9273920059204,-15.0215816497803,-15.255859375,-15.4949703216553,-15.8329095840454,-16.0604496002197,-16.2360363006592,-16.4680652618408,-16.6403312683105,-16.7396965026855,-16.9330558776855,-17.0951156616211,-17.6334457397461,-17.8157215118408,-17.8392562866211,-17.9148426055908,-17.9195804595947,-17.8863773345947,-17.8802738189697,-17.9469223022461,-18.0800304412842,-18.1429214477539,-18.2190437316895,-18.252197265625,-18.2652339935303,-18.2660160064697,-18.2222652435303,-18.3028316497803,-18.6536140441895,-19.2501945495605,-19.486572265625,-19.7782707214355,-19.9519519805908,-20.1981449127197,-20.4004402160645,-20.4940414428711,-20.5015621185303,-20.4910144805908,-20.469970703125,-20.4384765625,-20.3717288970947,-20.363037109375,-20.4139652252197,-20.4626960754395,-20.5929698944092,-20.6509284973145,-20.72705078125,-20.7299308776855,-20.8787593841553,-21.0081062316895,-21.1365718841553,-21.15576171875,-21.0940418243408,-21.10595703125,-21.1523914337158,-21.2462406158447,-21.3875980377197,-21.4486331939697,-22.37255859375,-22.6068840026855,-22.6521968841553,-22.6930160522461,-22.7293949127197,-22.7429695129395,-22.733642578125,-22.701171875,-22.6509265899658,-22.6030769348145,-22.559814453125,-22.5100574493408,-22.1875972747803,-22.056640625,-22.0009765625,-21.9354496002197,-21.8659191131592,-21.832763671875,-21.767578125,-21.7225589752197,-21.6686515808105,-21.6060066223145,-21.4633293151855,-21.5571784973145,-21.6466789245605,-21.9512214660645,-22.0533695220947,-22.049072265625,-22.0060062408447,-21.9012699127197,-21.9731941223145,-22.0006828308105,-22.0038070678711,-21.9503421783447,-21.702392578125,-21.6166496276855,-21.5906238555908,-21.6231441497803,-21.6749515533447,-21.9244136810303,-22.1060066223145,-22.1599617004395,-22.25390625,-22.2841796875,-22.32470703125,-22.3201179504395,-22.2335948944092,-22.24755859375,-22.3070316314697,-22.467041015625,-22.7203121185303,-23.3469734191895,-23.4764652252197,-23.68994140625,-23.8189945220947,-23.8785648345947,-23.9327640533447,-23.9819831848145,-24.0261726379395,-24.0070304870605,-23.9244136810303,-23.8638191223145,-23.6932125091553,-23.4853038787842,-23.3526840209961,-23.3145999908447,-23.2365245819092,-23.197998046875,-23.1378898620605,-23.1088390350342,-22.8995113372803,-22.8276863098145,-22.819580078125,-22.7880859375,-22.6839828491211,-22.5997085571289,-22.4944820404053,-22.3084487915039,-21.8921394348145,-21.8297843933105,-21.8004398345947,-21.7637195587158,-21.7799816131592,-22.0399913787842,-22.0993156433105,-22.4002933502197,-22.5090808868408,-22.4734382629395,-22.31396484375,-22.1493167877197,-21.906982421875,-21.8502445220947,-21.8443851470947,-22.0057621002197,-22.3114738464355,-22.3896980285645,-22.6436042785645,-22.8126449584961,-22.9024906158447,-23.1220703125,-23.6045417785645,-23.7964839935303,-23.89990234375,-24.0189933776855,-24.2239742279053,-24.4547863006592,-24.4756832122803,-24.341064453125,-24.2489242553711,-24.1561031341553,-23.97900390625,-23.8567371368408,-24.010009765625,-24.0060062408447,-24.017578125,-24.0650386810303,-24.1119136810303,-24.0926265716553,-24.0324211120605,-23.9090824127197,-23.6159172058105,-23.4719715118408,-23.3929691314697,-23.2853507995605,-23.31591796875,-23.5692863464355,-23.7047367095947,-23.7732410430908,-23.8326168060303,-23.8117179870605,-23.7413082122803,-23.5249500274658,-23.66748046875,-23.7665538787842,-23.77734375,-23.7705574035645,-23.7571296691895,-23.7371578216553,-23.4888668060303,-23.4344730377197,-23.4846687316895,-23.5935554504395,-23.5985355377197,-23.5526371002197,-23.5299816131592,-23.5279312133789,-23.4525394439697,-23.3765621185303,-23.3000011444092,-23.0625495910645,-23.0285148620605,-23.0172843933105,-23.0289058685303,-23.0189933776855,-22.9262218475342,-22.8522472381592,-22.8153324127197,-22.7233390808105,-22.6598625183105,-22.62158203125,-22.6097183227539,-22.6040515899658,-22.6202144622803,-22.6160163879395,-22.5515632629395,-22.4416999816895,-22.4275398254395,-22.4242172241211,-22.4331531524658,-22.4453125,-22.8064460754395,-22.8692398071289,-22.9479007720947,-22.9319820404053,-22.8616218566895,-22.7555179595947,-22.5093746185303,-22.4844245910645,-22.5321292877197,-22.6460933685303,-22.6727542877197,-22.6862316131592,-22.8213386535645,-22.9720211029053,-23.116943359375,-23.1199226379395,-23.0626964569092,-22.9443359375,-22.8892078399658,-22.7237300872803,-22.559326171875,-22.4261226654053,-22.3204593658447,-22.1702156066895,-21.9669914245605,-21.9483890533447,-21.8402347564697,-21.6252918243408,-21.4068851470947,-21.3967761993408,-21.4327144622803,-21.5166492462158,-21.4974594116211,-21.3877925872803,-21.3087882995605,-21.303466796875,-21.3749027252197,-21.412841796875,-21.4566421508789,-21.658447265625,-21.6103515625,-21.4662609100342,-21.4336433410645,-21.4551277160645,-21.4394035339355,-21.3866195678711,-21.36474609375,-21.3738765716553,-21.3963375091553,-21.4321765899658,-21.421875,-21.365478515625,-21.3125476837158,-21.22998046875,-21.1629886627197,-21.1296863555908,-21.105712890625,-21.0755863189697,-21.0473155975342,-21.0208492279053,-20.9979972839355,-20.9788570404053,-20.9397468566895,-20.8043460845947,-20.7396965026855,-20.678955078125,-20.6494140625,-20.5481433868408,-20.4865245819092,-20.454833984375,-20.4115238189697,-20.3566417694092,-20.3440933227539,-20.3739261627197,-20.3565921783447,-20.2921390533447,-20.20751953125,-20.1026859283447,-20.0260734558105,-19.874755859375,-19.7526378631592,-19.6478519439697,-19.5935554504395,-19.4896965026855,-19.4618167877197,-19.4432621002197,-19.4338855743408,-19.4562492370605,-19.4270515441895,-19.3829593658447,-19.1953125,-19.0932140350342,-18.9937496185303,-18.9113292694092,-18.8458995819092,-18.7775382995605,-18.7062015533447,-18.5949211120605,-18.4549312591553,-18.2769527435303,-18.1836414337158,-18.1637210845947,-18.1419429779053,-18.118408203125,-18.1033210754395,-18.0990238189697,-18.1488761901855,-18.3153324127197,-18.3182125091553,-18.2971687316895,-18.1798820495605,-17.906982421875,-17.81982421875,-17.6343250274658,-17.5822277069092,-17.5504398345947,-17.5390148162842,-17.467041015625,-17.417236328125,-17.3342781066895,-17.1530265808105,-17.1153812408447,-17.0624504089355,-16.9695320129395,-16.9254398345947,-16.8380374908447,-16.7484378814697,-16.624755859375,-16.4850101470947,-16.4371089935303,-16.4280776977539,-16.5406742095947,-16.4933605194092,-16.2493152618408,-16.035888671875,-15.9854001998901,-15.8509273529053,-15.7597646713257,-15.713770866394,-15.7027826309204,-15.6473636627197,-15.5431146621704],[35.7873039245605,35.7344741821289,35.6112289428711,35.5945320129395,35.5728530883789,35.5690422058105,35.5514640808105,35.484375,35.4026374816895,35.38671875,35.3621101379395,35.3038101196289,35.1932601928711,35.0650405883789,35.0105438232422,34.99951171875,34.9559555053711,34.9713897705078,34.9788093566895,34.98974609375,34.9783210754395,34.9538078308105,34.9611320495605,34.9830055236816,35.05322265625,35.1271476745605,35.1980476379395,35.2037086486816,35.1534194946289,35.03466796875,34.9509773254395,34.92919921875,34.8727531433105,34.8804664611816,34.9078140258789,35.1011734008789,35.2766609191895,35.40869140625,35.4505882263184,35.4228515625,35.4235382080078,35.4006843566895,35.40966796875,35.4392547607422,35.3830108642578,35.3201179504395,35.2978515625,35.2366218566895,35.1740226745605,35.140625,35.1481437683105,35.1326179504395,35.1416015625,35.0681648254395,35.0534172058105,35.02392578125,34.9734382629395,34.904296875,34.8698234558105,34.7911109924316,34.7350616455078,34.6585922241211,34.5296859741211,34.5177764892578,34.4899406433105,34.4009742736816,34.3285140991211,34.2453117370605,34.3483390808105,34.3501968383789,34.5255851745605,34.5241203308105,34.4773445129395,34.4839859008789,34.6784172058105,34.8038101196289,34.921875,35.005859375,35.0770530700684,35.1085929870605,35.2233390808105,35.3088874816895,35.4112319946289,35.4931640625,35.5325202941895,35.5792961120605,35.6029319763184,35.6272468566895,35.7344741821289,35.7874984741211,35.8407249450684,35.869140625,35.8515625,35.8371086120605,35.8370132446289,35.8587913513184,35.8884773254395,35.9066390991211,35.8680648803711,35.8717765808105,35.8820304870605,35.9134750366211,35.8568382263184,35.8014640808105,35.7873039245605],[12.05126953125,12.0033206939697,11.9406242370605,11.9364261627197,11.9480476379395,12.0242195129395,12.0480470657349,12.05126953125],[15.5765619277954,15.5088872909546,15.4756841659546,15.2344722747803,15.2068357467651,15.1898441314697,15.1648435592651,15.1310548782349,15.0995111465454,15.1056632995605,15.1169929504395,15.14599609375,15.1936511993408,15.2302732467651,15.1741209030151,15.2360343933105,15.2886714935303,15.2957029342651,15.2945318222046,15.1851568222046,15.1423826217651,15.1158199310303,15.1042976379395,15.1163082122803,15.1125974655151,15.0024423599243,14.8896484375,14.7759771347046,14.6143560409546,14.5554676055908,14.5018558502197,14.3672857284546,14.2590818405151,14.1429691314697,14.0243167877197,13.9054689407349,13.8005857467651,13.5871095657349,13.3609380722046,13.2649412155151,13.2210941314697,13.169921875,13.0403318405151,12.9241218566895,12.8711919784546,12.75732421875,12.6990232467651,12.6402339935303,12.5267581939697,12.4543952941895,12.435546875,12.48681640625,12.5476560592651,12.6016597747803,12.6643562316895,12.7023439407349,12.734375,12.8506832122803,12.9027347564697,12.9554691314697,13.0490236282349,13.0568361282349,13.1599607467651,13.2911138534546,13.3516597747803,13.3834962844849,13.4334964752197,13.4913091659546,13.6815423965454,13.73486328125,13.7888669967651,13.9366207122803,14.0499992370605,14.2876949310303,14.4162101745605,14.5059576034546,14.6367177963257,14.7372074127197,14.7896480560303,14.8458995819092,14.9819326400757,15.1187496185303,15.1760740280151,15.2240238189697,15.2795886993408,15.3407230377197,15.4987306594849,15.568359375,15.6346683502197,15.5765619277954],[8.47890663146973,8.42148399353027,8.36093807220459,8.35859298706055,8.36679649353027,8.44062519073486,8.47890663146973],[13.9382820129395,13.8936519622803,13.86767578125,13.853515625,13.8711910247803,13.9621095657349,13.9608402252197,13.9382820129395],[8.28603553771973,8.25273513793945,8.20566463470459,8.22402286529541,8.26738262176514,8.32021427154541,8.34375,8.31894588470459,8.28603553771973],[9.63203048706055,9.68203163146973,9.79433631896973,9.80527400970459,9.78281211853027,9.75419902801514,9.64296817779541,9.65947341918945,9.70078086853027,9.70673751831055,9.68603515625,9.61699295043945,9.58359336853027,9.5625,9.486328125,9.38808536529541,9.26416015625,9.20693397521973,9.14931678771973,9.10175800323486,9.05634784698486,9.02265644073486,8.96660137176514,8.88134765625,8.80117130279541,8.71855449676514,8.64853477478027,8.59541034698486,8.55331993103027,8.48622989654541,8.41816329956055,8.41074180603027,8.39912128448486,8.41865253448486,8.44707012176514,8.46103477478027,8.451171875,8.47109413146973,8.5107421875,8.54052734375,8.53867244720459,8.54775333404541,8.49589824676514,8.40781307220459,8.39931678771973,8.40859413146973,8.455078125,8.47080039978027,8.47128868103027,8.4091796875,8.38535118103027,8.35322189331055,8.29550743103027,8.23027420043945,8.18994140625,8.18085861206055,8.20380878448486,8.22421836853027,8.24521446228027,8.31015586853027,8.36328029632568,8.46845722198486,8.57187461853027,8.69892597198486,8.82119083404541,8.99814510345459,9.10722637176514,9.1630859375,9.18212890625,9.22841835021973,9.28300762176514,9.35078048706055,9.45517635345459,9.50019550323486,9.53876876831055,9.57568359375,9.61533164978027,9.62119102478027,9.58974647521973,9.5537109375,9.57402324676514,9.63203048706055],[10.3951168060303,10.4283199310303,10.4322261810303,10.4099607467651,10.4193353652954,10.3356447219849,10.208984375,10.1312503814697,10.1097660064697,10.1275386810303,10.2482423782349,10.2857427597046,10.3589839935303,10.3951168060303],[13.6999998092651,13.6796875,13.6371097564697,13.5632810592651,13.478515625,13.3995122909546,13.3782224655151,13.3996095657349,13.4209966659546,13.4498052597046,13.4917974472046,13.5447263717651,13.6325197219849,13.6349601745605,13.6166019439697,13.5480470657349,13.4864253997803,13.4802742004395,13.4876947402954,13.5091791152954,13.6005859375,13.6139650344849,13.5696287155151,13.5833988189697,13.6634769439697,13.7216796875,13.8311529159546,13.8747072219849,13.8447256088257,13.7759771347046,13.7198247909546,13.7833003997803,13.6283197402954,13.5582027435303,13.4651365280151,13.2063474655151,13.15673828125,13.1201171875,13.0302724838257,12.9030275344849,12.76123046875,12.6117191314697,12.4975595474243,12.43212890625,12.5361328125,12.4917974472046,12.3538093566895,12.2743158340454,12.2488279342651,12.2256832122803,12.2863283157349,12.3924808502197,12.5234375,12.4979496002197,12.4635744094849,12.3844728469849,12.3190431594849,12.2789058685303,12.2483396530151,12.3049802780151,12.3962888717651,12.48681640625,12.6911134719849,12.9070310592651,13.2953128814697,13.5082035064697,13.5641593933105,13.6932611465454,13.8046875,13.9249019622803,14.0104494094849,14.1827144622803,14.5407228469849,14.8661136627197,15.1687507629395,15.4049806594849,15.9640626907349,16.0615234375,16.1646480560303,16.1891593933105,16.1512699127197,16.03369140625,15.9137687683105,15.9004878997803,16.0125980377197,16.5518550872803,17.1034183502197,17.2751960754395,17.4742202758789,17.9549808502197,18.0361328125,18.3282222747803,18.4606437683105,18.48583984375,18.4225578308105,18.3934574127197,18.34375,18.2193374633789,18.0779304504395,17.8650398254395,17.4761714935303,17.3957996368408,17.2577152252197,17.2494144439697,17.2153339385986,17.1799793243408,17.03125,16.92822265625,16.8070316314697,16.6696281433105,16.5299797058105,16.5218753814697,16.5977535247803,16.8243160247803,16.9992198944092,17.1145496368408,17.1229496002197,17.1746101379395,17.0985355377197,16.9514636993408,16.7554683685303,16.61669921875,16.5589847564697,16.57421875,16.5456047058105,16.2824211120605,16.1441402435303,16.1097660064697,16.0568351745605,15.7245111465454,15.6458015441895,15.64306640625,15.7001953125,15.8223638534546,15.90478515625,15.8789052963257,15.9269523620605,15.9723634719849,16.0655269622803,16.19677734375,16.2099609375,16.107421875,16.0714855194092,16.0236320495605,15.854395866394,15.763671875,15.692774772644,15.5851564407349,15.3909177780151,15.2945318222046,14.9508790969849,14.9269533157349,14.9291009902954,14.9861335754395,14.947657585144,14.9069337844849,14.8395509719849,14.7657232284546,14.6112308502197,14.5569324493408,14.4593753814697,14.3827152252197,14.3399410247803,14.4605464935303,14.4281253814697,14.3088865280151,14.1471672058105,14.1023435592651,14.0758781433105,14.0443353652954,14.0476560592651,13.8597650527954,13.7333984375,13.6697273254395,13.5547857284546,13.3619136810303,13.2468748092651,13.1833982467651,13.0886716842651,13.041015625,13.0242185592651,12.8492183685303,12.6308584213257,12.2056646347046,12.0752925872803,11.8070316314697,11.6373043060303,11.4984369277954,11.2962894439697,11.2497072219849,11.1888666152954,11.1412105560303,11.1032218933105,11.1417961120605,11.1847658157349,11.1677732467651,10.9377927780151,10.8031253814697,10.76513671875,10.7371091842651,10.7083988189697,10.6446285247803,10.5902347564697,10.5148429870605,10.5172853469849,10.5323238372803,10.5208005905151,10.4475593566895,10.3205080032349,10.2458009719849,10.1880865097046,10.0476560592651,9.73085975646973,9.28935527801514,9.19599628448486,8.93037033081055,8.76582050323486,8.55195331573486,8.29238224029541,8.08164024353027,8.00498104095459,7.73330068588257,7.4931640625,7.49052667617798,7.48203086853027,7.52265596389771,7.58964824676514,7.65146493911743,7.67714881896973,7.6650390625,7.63720655441284,7.59941434860229,7.37089872360229,7.31855440139771,7.1494140625,6.96728563308716,6.90019559860229,6.87480497360229,6.89384794235229,6.87861347198486,6.84296894073486,6.87519502639771,6.93193340301514,6.96035146713257,7.00791025161743,7.03066396713257,6.99267578125,6.97285175323486,6.93984365463257,6.88935565948486,6.80107402801514,6.73818349838257,6.72470712661743,6.69140625,6.63476610183716,6.62773466110229,6.69228506088257,6.78037118911743,6.84228515625,6.98124980926514,7.03242206573486,7.07832002639771,7.11679697036743,7.14638662338257,7.15341806411743,7.12607431411743,7.013671875,6.96240186691284,6.88144540786743,6.80624961853027,6.79091835021973,6.78916025161743,6.80449199676514,6.94082021713257,7.02109336853027,7.05576181411743,7.12900400161743,7.32792997360229,7.45156240463257,7.53857469558716,7.59257841110229,7.78788995742798,7.85234308242798,7.99316358566284,8.01425838470459,8.12519550323486,8.12724590301514,8.08154296875,8.095703125,8.23193359375,8.29853439331055,8.37070369720459,8.42255878448486,8.43681621551514,8.44296836853027,8.4384765625,8.45839881896973,8.5654296875,8.64169883728027,8.81855487823486,8.82675743103027,8.77802753448486,8.88515663146973,8.904296875,8.95371150970459,9.02373123168945,9.04667949676514,9.01914119720459,8.99892520904541,9.00302791595459,9.02236270904541,9.07099533081055,9.20341777801514,9.25107383728027,9.259765625,9.26015663146973,9.30439472198486,9.39931583404541,9.42763710021973,9.44062519073486,9.48105430603027,9.52871036529541,9.57958984375,9.63945388793945,9.78779220581055,9.88447189331055,9.93925762176514,9.97167873382568,10.041015625,10.08056640625,10.1283197402954,10.1452150344849,10.1298828125,10.1096677780151,10.0819339752197,10.0456056594849,10.0382814407349,10.0612306594849,10.0870122909546,10.1374998092651,10.1955080032349,10.2722654342651,10.3630857467651,10.4306640625,10.4424810409546,10.4382820129395,10.39794921875,10.4060554504395,10.4528331756592,10.4793949127197,10.5797853469849,10.6892576217651,10.759765625,10.8289060592651,10.9273443222046,10.9932613372803,11.0250978469849,11.0634765625,11.1338863372803,11.2444334030151,11.4332027435303,11.5275392532349,11.62548828125,11.6994142532349,11.7756834030151,11.9695310592651,12.16943359375,12.1971683502197,12.2012701034546,12.16552734375,12.1307621002197,12.1541013717651,12.2679691314697,12.330078125,12.3882808685303,12.4791984558105,12.5986318588257,12.6998052597046,12.8055667877197,13.1687498092651,13.3515625,13.4900398254395,13.6999998092651],[12.3968753814697,12.4263668060303,12.4852533340454,12.5146484375,12.5037107467651,12.4411134719849,12.3968753814697],[-77.261474609375,-77.1395492553711,-77.0137710571289,-76.9593734741211,-76.908203125,-76.7932662963867,-76.7007293701172,-76.349853515625,-76.2327651977539,-76.2107925415039,-76.3014602661133,-76.41552734375,-76.5246047973633,-76.6253890991211,-76.6693878173828,-76.7743148803711,-76.748291015625,-76.7948226928711,-76.8532180786133,-76.896240234375,-76.9441452026367,-77.0359344482422,-77.0712890625,-77.1194839477539,-77.1584014892578,-77.2049789428711,-77.2798843383789,-77.3614273071289,-77.4638671875,-77.6707534790039,-77.7681655883789,-77.8494186401367,-77.8813018798828,-77.9629898071289,-78.0444793701172,-78.0736312866211,-78.2940902709961,-78.3395004272461,-78.3259811401367,-78.25244140625,-78.2166976928711,-78.0945281982422,-77.9781723022461,-77.9268569946289,-77.8734359741211,-77.4516143798828,-77.354248046875,-77.261474609375],[-2.01865243911743,-2.00991177558899,-2.05375981330872,-2.09101557731628,-2.16567397117615,-2.23584008216858,-2.22050762176514,-2.08222651481628,-2.01865243911743],[39.1454086303711,38.9970703125,38.9623069763184,38.7696304321289,38.37548828125,38.1114234924316,37.7738265991211,37.4933586120605,37.2156257629395,36.9585952758789,37.1052742004395,37.3294944763184,37.47900390625,37.6554679870605,37.81298828125,37.9800758361816,37.8628921508789,37.6697273254395,37.64990234375,37.6335945129395,37.5536117553711,37.49072265625,37.46923828125,37.1994132995605,36.9270515441895,36.7552719116211,36.7039070129395,36.591796875,36.47607421875,36.2828102111816,36.0684547424316,36.0154304504395,35.8603515625,35.5953140258789,35.3391609191895,35.1637725830078,34.9507827758789,34.9822273254395,34.9734382629395,35.02392578125,35.0534172058105,35.0681648254395,35.1416015625,35.1326179504395,35.1481437683105,35.140625,35.1740226745605,35.2366218566895,35.2978515625,35.3201179504395,35.3830108642578,35.4392547607422,35.40966796875,35.4006843566895,35.4235382080078,35.4228515625,35.4505882263184,35.4654312133789,35.4994125366211,35.5589866638184,35.5314483642578,35.5347671508789,35.5720672607422,35.5514640808105,35.5690422058105,35.5728530883789,35.5945320129395,35.6112289428711,35.7344741821289,35.7873039245605,35.8947257995605,35.9564437866211,36.0594711303711,36.2197265625,36.2842750549316,36.3720703125,36.4791984558105,36.818359375,37.0889663696289,37.3175773620605,37.5774383544922,37.7541046142578,38.0557632446289,38.2542991638184,38.515625,38.7735366821289,38.8450202941895,38.9148406982422,38.9874038696289,39.0578117370605,38.9816398620605,39.0414047241211,39.1400413513184,39.2474632263184,39.2927742004395,39.1454086303711],[123.888679504395,123.825592041016,123.749801635742,123.680671691895,123.679779052734,123.752342224121,123.753707885742,123.771484375,123.934860229492,123.928123474121,123.888679504395],[124.293159484863,124.234283447266,124.185638427734,124.135734558105,124.084770202637,124.120407104492,124.170219421387,124.210540771484,124.301956176758,124.324020385742,124.293159484863],[125.444145202637,125.359382629395,125.268951416016,125.283592224121,125.31494140625,125.334571838379,125.401847839355,125.444145202637],[142.188186645508,142.169921875,142.107131958008,142.125289916992,142.161712646484,142.2021484375,142.188186645508],[128.2587890625,128.162506103516,128.126953125,128.037887573242,127.951271057129,127.867080688477,127.869239807129,127.90478515625,127.848724365234,127.790138244629,127.785545349121,127.806442260742,127.803619384766,127.729400634766,127.653121948242,127.649703979492,127.654884338379,127.72705078125,127.728912353516,127.7958984375,127.820404052734,127.925971984863,127.945510864258,127.890823364258,127.894828796387,127.907234191895,127.994331359863,128.029678344727,128.046768188477,128.09765625,128.12158203125,128.216506958008,128.2548828125,128.331649780273,128.310943603516,128.2587890625],[128.998153686523,128.956253051758,128.899993896484,128.8828125,128.907623291016,128.95166015625,128.98974609375,129.016418457031,128.998153686523],[129.324020385742,129.33056640625,129.232421875,129.192474365234,129.257415771484,129.27734375,129.324020385742],[129.452529907227,129.366409301758,129.27490234375,129.164642333984,129.217086791992,129.247848510742,129.250885009766,129.322463989258,129.464553833008,129.509658813477,129.560546875,129.5771484375,129.598037719727,129.689559936523,129.714645385742,129.71044921875,129.641708374023,129.574600219727,129.5126953125,129.456741333008,129.439071655273,129.452529907227],[130.622756958008,130.508209228516,130.445602416992,130.388076782227,130.497177124023,130.6435546875,130.673233032227,130.622756958008],[130.959762573242,130.872161865234,130.870315551758,130.93994140625,130.947357177734,131.01220703125,131.039840698242,131.060348510742,131.082611083984,131.057418823242,130.992584228516,130.959762573242],[129.717971801758,129.686813354492,129.706832885742,129.787307739258,129.793640136719,129.717971801758],[130.381057739258,130.292572021484,130.256057739258,130.24169921875,130.365432739258,130.46142578125,130.418548583984,130.381057739258],[130.08251953125,130.003509521484,129.993362426758,129.960159301758,130.017288208008,130.015335083008,129.979293823242,130.021286010742,130.009765625,130.167785644531,130.196685791016,130.199508666992,130.08251953125],[128.665328979492,128.7041015625,128.761032104492,128.806060791016,128.838562011719,128.87939453125,128.89453125,128.8212890625,128.790435791016,128.75048828125,128.692962646484,128.657318115234,128.649124145508,128.665328979492],[139.841110229492,139.823822021484,139.775680541992,139.768951416016,139.777450561523,139.808883666992,139.873626708984,139.841110229492],[129.076950073242,129.051956176758,129.019622802734,128.997268676758,129.034957885742,129.109771728516,129.123626708984,129.152725219727,129.181930541992,129.153518676758,129.111633300781,129.076950073242],[129.491790771484,129.42138671875,129.370407104492,129.4169921875,129.423156738281,129.4619140625,129.537994384766,129.569915771484,129.508102416992,129.491790771484],[129.795700073242,129.7265625,129.6748046875,129.700012207031,129.71728515625,129.7763671875,129.795700073242],[131.174606323242,131.309188842773,131.366012573242,131.418746948242,131.498825073242,131.5830078125,131.64306640625,131.6962890625,131.724212646484,131.710647583008,131.615539550781,131.537414550781,131.717086791992,131.896575927734,131.854690551758,131.847854614258,131.902725219727,131.949310302734,131.937210083008,131.910446166992,132.008590698242,132.002151489258,131.976654052734,131.732116699219,131.660354614258,131.6103515625,131.564651489258,131.531158447266,131.505661010742,131.460250854492,131.475296020508,131.4599609375,131.337203979492,131.249710083008,131.139739990234,131.07080078125,131.03515625,131.098434448242,130.902236938477,130.685745239258,130.684188842773,130.704498291016,130.735733032227,130.758392333984,130.789749145508,130.774215698242,130.708984375,130.704193115234,130.749404907227,130.77978515625,130.796295166016,130.796875,130.776260375977,130.714553833008,130.655090332031,130.613479614258,130.556060791016,130.528121948242,130.540435791016,130.56591796875,130.64453125,130.621475219727,130.588760375977,130.310638427734,130.250579833984,130.20068359375,130.147262573242,130.260543823242,130.294143676758,130.306640625,130.321975708008,130.268951416016,130.224227905273,130.187881469727,130.2109375,130.19580078125,130.194427490234,130.214065551758,130.319137573242,130.394927978516,130.462005615234,130.560348510742,130.640518188477,130.563293457031,130.497848510742,130.569427490234,130.547256469727,130.4404296875,130.381744384766,130.287307739258,130.237503051758,130.176864624023,130.126846313477,130.173141479492,130.167785644531,130.175003051758,130.22216796875,130.280075073242,130.326477050781,130.353515625,130.36083984375,130.340438842773,130.297668457031,130.245498657227,130.192962646484,130.152053833008,130.054092407227,129.950790405273,129.8525390625,129.7685546875,129.80810546875,129.826766967773,129.785949707031,129.690032958984,129.667770385742,129.662307739258,129.679107666016,129.777740478516,129.828323364258,129.900787353516,129.99169921875,129.921875,129.896789550781,129.798736572266,129.6650390625,129.580078125,129.610168457031,129.659973144531,129.7021484375,129.844131469727,129.857528686523,129.836624145508,129.82568359375,129.919143676758,130.072067260742,130.103424072266,130.130569458008,130.16796875,130.275100708008,130.365036010742,130.439453125,130.457229614258,130.483779907227,130.669540405273,130.715621948242,130.839645385742,130.953125,131.009078979492,131.05810546875,131.174606323242],[132.266006469727,132.314544677734,132.430465698242,132.444915771484,132.411026000977,132.359954833984,132.267288208008,132.208786010742,132.200592041016,132.2080078125,132.266006469727],[132.578414916992,132.549407958984,132.4609375,132.496292114258,132.523544311523,132.54345703125,132.560165405273,132.578414916992],[129.279495239258,129.214447021484,129.186431884766,129.21484375,129.337097167969,129.335052490234,129.279495239258],[134.357421875,134.495697021484,134.637496948242,134.63525390625,134.655380249023,134.6953125,134.6748046875,134.738861083984,134.548736572266,134.377044677734,134.306549072266,134.24267578125,134.205657958984,134.181640625,134.124130249023,133.958694458008,133.85400390625,133.685638427734,133.632034301758,133.285934448242,133.239944458008,133.145599365234,133.100875854492,133.051162719727,133.016021728516,132.977233886719,132.869918823242,132.804306030273,132.692184448242,132.641799926758,132.708984375,132.601943969727,132.4951171875,132.492584228516,132.427825927734,132.475784301758,132.477142333984,132.505279541016,132.515045166016,132.511413574219,132.445419311523,132.405166625977,132.412796020508,132.374893188477,132.281051635742,132.085845947266,132.032608032227,132.114456176758,132.287902832031,132.365921020508,132.536041259766,132.64306640625,132.698928833008,132.716217041016,132.752349853516,132.784271240234,132.839447021484,132.935150146484,132.990142822266,133.05126953125,133.133697509766,133.193054199219,133.298828125,133.349792480469,133.472076416016,133.58203125,133.626770019531,133.643463134766,133.602645874023,133.655578613281,133.706253051758,133.825592041016,133.948348999023,134.075881958008,134.21923828125,134.357421875],[134.351852416992,134.333206176758,134.315322875977,134.251846313477,134.238082885742,134.188293457031,134.18212890625,134.325973510742,134.372268676758,134.351852416992],[134.932815551758,134.824417114258,134.730651855469,134.683486938477,134.667877197266,134.757232666016,134.834274291992,134.904098510742,134.960739135742,135.004684448242,134.905471801758,134.932815551758],[129.385635375977,129.365325927734,129.297454833984,129.266693115234,129.329406738281,129.322067260742,129.325881958008,129.451080322266,129.472457885742,129.480163574219,129.469146728516,129.475387573242,129.381439208984,129.385635375977],[139.456436157227,139.445709228516,139.392379760742,139.366897583008,139.370025634766,139.426162719727,139.456436157227],[133.370513916016,133.32470703125,133.2392578125,133.18994140625,133.206146240234,133.295700073242,133.381240844727,133.370513916016],[138.344039916992,138.2490234375,138.225204467773,138.282806396484,138.287887573242,138.322265625,138.321685791016,138.246200561523,138.25,138.306350708008,138.461318969727,138.503616333008,138.510055541992,138.462799072266,138.45361328125,138.5751953125,138.496978759766,138.344039916992],[141.229293823242,141.268753051758,141.455474853516,141.419921875,141.399993896484,141.413665771484,141.430465698242,141.462799072266,141.542282104492,141.646286010742,141.797073364258,141.8779296875,141.93505859375,141.977828979492,141.990829467773,141.991897583008,141.979095458984,141.9931640625,141.976959228516,141.909469604492,141.900787353516,141.842086791992,141.806549072266,141.776168823242,141.742477416992,141.693557739258,141.658584594727,141.644729614258,141.622268676758,141.579681396484,141.546279907227,141.518753051758,141.5087890625,141.467483520508,141.3681640625,141.254287719727,141.1083984375,141.077331542969,140.962112426758,140.929000854492,140.927932739258,140.9599609375,141.00341796875,141.036331176758,141.001663208008,140.968353271484,140.895126342773,140.839645385742,140.791793823242,140.729888916016,140.627334594727,140.618835449219,140.619232177734,140.591598510742,140.573532104492,140.590423583984,140.621963500977,140.759567260742,140.8134765625,140.8740234375,140.639266967773,140.596878051758,140.457412719727,140.412887573242,140.41650390625,140.392974853516,140.354690551758,140.314743041992,140.158889770508,140.059173583984,139.959762573242,139.92041015625,139.843948364258,139.799224853516,139.84326171875,139.829681396484,139.851470947266,139.826461791992,139.906158447266,139.944137573242,140.027145385742,140.086334228516,140.096878051758,140.043655395508,139.987503051758,139.909774780273,139.834762573242,139.786331176758,139.770126342773,139.77392578125,139.767776489258,139.650009155273,139.66552734375,139.699996948242,139.744049072266,139.730865478516,139.675003051758,139.635940551758,139.564071655273,139.474411010742,139.363479614258,139.249420166016,139.162704467773,139.134078979492,139.115814208984,139.121978759766,139.086044311523,139.015625,138.982620239258,138.896682739258,138.837493896484,138.795120239258,138.761032104492,138.804504394531,138.802734375,138.903610229492,138.820907592773,138.719619750977,138.5771484375,138.537017822266,138.509567260742,138.43310546875,138.3486328125,138.253219604492,138.189056396484,137.97900390625,137.8642578125,137.74853515625,137.543365478516,137.318069458008,137.061721801758,137.077041625977,137.287796020508,137.29541015625,137.275192260742,137.22265625,137.096588134766,137.0322265625,137.005966186523,136.963287353516,136.934753417969,136.944152832031,136.912887573242,136.871292114258,136.884567260742,136.856155395508,136.852935791016,136.897064208984,136.851852416992,136.80419921875,136.743560791016,136.690032958984,136.576950073242,136.533004760742,136.615814208984,136.841598510742,136.880264282227,136.881164550781,136.853713989258,136.792190551758,136.54443359375,136.329879760742,136.267868041992,136.072555541992,135.916213989258,135.6953125,135.452835083008,135.394241333008,135.346786499023,135.256637573242,135.175384521484,135.1279296875,135.135345458984,135.10009765625,135.131927490234,135.265625,135.309265136719,135.384765625,135.411819458008,135.415924072266,135.355163574219,135.1982421875,135.041702270508,134.929885864258,134.784957885742,134.740036010742,134.583801269531,134.472259521484,134.362686157227,134.246871948242,134.208297729492,134.074508666992,133.96826171875,133.876373291016,133.677932739258,133.57861328125,133.474411010742,133.4453125,133.335647583008,133.209762573242,133.142379760742,133.018951416016,132.774612426758,132.656555175781,132.534469604492,132.421295166016,132.312591552734,132.238082885742,132.201965332031,132.159378051758,132.146484375,132.090225219727,131.76318359375,131.740524291992,131.476165771484,131.407913208008,131.32275390625,131.232620239258,131.150390625,131.071884155273,130.996383666992,130.918853759766,130.889251708984,130.904296875,130.951858520508,131.004196166992,131.132232666016,131.261810302734,131.354400634766,131.432525634766,131.515029907227,131.608001708984,131.734069824219,131.856048583984,131.963073730469,132.064743041992,132.158203125,132.259567260742,132.4140625,132.619033813477,132.697647094727,132.74658203125,132.922958374023,133.156921386719,133.267181396484,133.37646484375,133.435348510742,133.494934082031,133.615432739258,133.739349365234,133.860260009766,133.981246948242,134.214065551758,134.336532592773,134.4560546875,134.882217407227,135.17431640625,135.220504760742,135.265441894531,135.268753051758,135.232040405273,135.267776489258,135.326950073242,135.601852416992,135.680267333984,135.795013427734,135.903121948242,136.016204833984,136.095321655273,136.022262573242,136.006256103516,136.067474365234,136.15625,136.261825561523,136.358993530273,136.555862426758,136.698150634766,136.749328613281,136.71923828125,136.843460083008,136.962295532227,137.198638916016,137.322662353516,137.341217041016,137.337493896484,137.152053833008,137.045806884766,136.982223510742,136.924026489258,136.89990234375,136.994430541992,137.0185546875,137.0126953125,137.016693115234,137.123718261719,137.246292114258,137.297668457031,137.342575073242,137.482604980469,137.514053344727,137.913192749023,138.109573364258,138.218078613281,138.319915771484,138.54833984375,138.6328125,138.709381103516,138.770599365234,138.81884765625,138.885055541992,139.247161865234,139.363861083984,139.400985717773,139.44580078125,139.476745605469,139.520812988281,139.580169677734,139.659759521484,139.749130249023,139.801956176758,139.878616333008,139.912292480469,139.938568115234,139.977142333984,140.010848999023,140.036529541016,140.048141479492,140.064743041992,140.0546875,139.994720458984,139.945220947266,139.891204833984,139.810348510742,139.741500854492,139.755462646484,139.82568359375,139.873626708984,139.908004760742,139.972457885742,140.011138916016,140.014450073242,139.964065551758,139.923919677734,139.9228515625,139.966690063477,140.029296875,140.085342407227,140.146102905273,140.201278686523,140.252349853516,140.28125,140.326263427734,140.343551635742,140.315246582031,140.344436645508,140.385925292969,140.441314697266,140.498046875,140.564163208008,140.627639770508,140.6396484375,140.679397583008,140.702438354492,140.748626708984,140.80078125,140.845794677734,140.876174926758,140.93603515625,141.118545532227,141.183212280273,141.225387573242,141.262115478516,141.244247436523,141.200393676758,141.155075073242,141.115036010742,141.070419311523,140.800582885742,140.801849365234,140.859558105469,140.891510009766,140.936904907227,141.050201416016,141.105865478516,141.229293823242],[139.481246948242,139.458389282227,139.4345703125,139.411529541016,139.431350708008,139.495788574219,139.558395385742,139.505081176758,139.481246948242],[141.29541015625,141.225982666016,141.145309448242,141.135360717773,141.193756103516,141.251846313477,141.31005859375,141.329193115234,141.29541015625],[141.07275390625,141.033981323242,140.982131958008,140.9716796875,141.001663208008,141.056732177734,141.069915771484,141.07275390625],[143.824310302734,143.949508666992,144.005462646484,144.101364135742,144.481826782227,144.5966796875,144.715240478516,144.798538208008,144.871871948242,145.1015625,145.3427734375,145.369522094727,145.36962890625,145.351959228516,145.245208740234,145.126373291016,145.10107421875,145.1396484375,145.214050292969,145.27294921875,145.3408203125,145.436126708984,145.487899780273,145.583297729492,145.673629760742,145.751266479492,145.8330078125,145.7255859375,145.624221801758,145.537399291992,145.505081176758,145.40478515625,145.347457885742,145.230087280273,145.127151489258,145.02880859375,144.92138671875,144.80712890625,144.630752563477,144.516220092773,144.301559448242,144.197265625,143.969329833984,143.762115478516,143.580963134766,143.429489135742,143.36865234375,143.332138061523,143.3271484375,143.313659667969,143.278701782227,143.236526489258,143.111724853516,142.906356811523,142.508209228516,142.087890625,141.851364135742,141.406631469727,140.986129760742,140.948440551758,140.78759765625,140.709762573242,140.616790771484,140.547653198242,140.48046875,140.385452270508,140.3505859375,140.323532104492,140.315246582031,140.32666015625,140.416595458984,140.527542114258,140.577743530273,140.684280395508,140.733795166016,140.912307739258,141.107727050781,141.150985717773,141.078704833984,140.99951171875,140.907516479492,140.81640625,140.659866333008,140.592971801758,140.489166259766,140.431640625,140.384963989258,140.270126342773,140.148635864258,140.085159301758,140.03662109375,140.009185791016,139.995315551758,140.021301269531,140.084167480469,140.1083984375,140.056838989258,140.024124145508,139.895111083984,139.83544921875,139.820892333984,139.828521728516,139.860153198242,139.89111328125,139.950592041016,140.015045166016,140.114654541016,140.22412109375,140.32861328125,140.432220458984,140.486434936523,140.3974609375,140.379287719727,140.392379760742,140.486907958984,140.584564208984,140.780670166016,140.819137573242,140.953811645508,141.13818359375,141.245025634766,141.296279907227,141.374130249023,141.412307739258,141.398345947266,141.397659301758,141.44677734375,141.600677490234,141.644729614258,141.660842895508,141.71630859375,141.760925292969,141.7822265625,141.719055175781,141.65576171875,141.5830078125,141.59375,141.652542114258,141.653991699219,141.66796875,141.778137207031,141.829498291016,141.878707885742,141.937698364258,141.980865478516,142.016403198242,142.171569824219,142.416015625,142.7041015625,142.884765625,143.075103759766,143.28857421875,143.511917114258,143.654586791992,143.759078979492,143.824310302734],[77.7994155883789,77.6969757080078,77.5715789794922,77.4234390258789,77.29296875,77.1685562133789,77.0486297607422,77.0044937133789,76.9789123535156,76.927734375,76.8822250366211,76.8127899169922,76.7668914794922,76.87890625,77.0900344848633,77.2948226928711,77.4464874267578,77.52001953125,77.5725631713867,77.7240219116211,77.7994155883789],[50.1844749450684,50.1487312316895,50.0953140258789,49.9951171875,50.0230445861816,50.0593757629395,50.10986328125,50.1166000366211,50.0453109741211,50.0388679504395,50.09814453125,50.1844749450684],[50.3117179870605,50.2772445678711,50.2561531066895,50.294921875,50.3497047424316,50.3308563232422,50.3117179870605],[52.6824188232422,52.6648445129395,52.5983390808105,52.5542945861816,52.60888671875,52.6595687866211,52.6929702758789,52.6824188232422],[87.3228454589844,87.22998046875,87.0485305786133,86.93798828125,86.8859329223633,86.8083038330078,86.7531204223633,86.7286148071289,86.7578125,86.7179641723633,86.6637725830078,86.5494079589844,86.4832992553711,86.37255859375,86.265625,86.05615234375,85.8298873901367,85.7494125366211,85.6921920776367,85.6515655517578,85.6263656616211,85.5623092651367,85.5259780883789,85.5616226196289,85.5882797241211,85.5866241455078,85.6417922973633,85.6698226928711,85.6566467285156,85.5772399902344,85.5296859741211,85.4847640991211,85.3553695678711,85.2334899902344,85.1105499267578,85.01220703125,84.8582000732422,84.7861328125,84.7459945678711,84.7195281982422,84.6666030883789,84.59228515625,84.5324249267578,84.3388671875,84.2151412963867,84.1220703125,84.0160217285156,83.8326110839844,83.7139663696289,83.6340789794922,83.4435501098633,83.1930618286133,83.09033203125,83.0294952392578,83.0201187133789,83.0040969848633,82.9748992919922,82.8000030517578,82.6921844482422,82.5550765991211,82.51171875,82.4296875,82.34814453125,82.3152313232422,82.3122100830078,82.32666015625,82.45166015625,82.58251953125,82.6116256713867,82.6257781982422,82.62109375,82.5969696044922,82.5589828491211,82.521484375,82.4787139892578,82.3966751098633,82.3234405517578,82.2666015625,82.1227493286133,81.9892578125,81.9449234008789,81.8674774169922,81.7896499633789,81.7588882446289,81.6919937133789,81.60205078125,81.3347702026367,81.0403289794922,80.8533172607422,80.7800827026367,80.634765625,80.5091781616211,80.4149398803711,80.2282180786133,80.0591812133789,79.9501953125,79.8718795776367,79.8752899169922,79.93212890625,79.9971694946289,80.1278305053711,80.2550811767578,80.36083984375,80.4554672241211,80.4815368652344,80.4554672241211,80.4005889892578,80.3814468383789,80.3910140991211,80.3550796508789,80.3363265991211,80.3548812866211,80.3653335571289,80.3589859008789,80.3552703857422,80.3957977294922,80.4315414428711,80.4960021972656,80.5934600830078,80.6507797241211,80.7038040161133,80.6654281616211,80.6677703857422,80.7297821044922,80.7570266723633,80.7857437133789,80.7777404785156,80.7511749267578,80.6169891357422,80.5070266723633,80.3902359008789,80.37451171875,80.3712844848633,80.3833999633789,80.45068359375,80.5437469482422,80.5389709472656,80.4240188598633,80.2502899169922,80.2022476196289,80.1650390625,80.1619110107422,80.1792984008789,80.2057647705078,80.2550811767578,80.2590789794922,80.2330093383789,80.2093734741211,80.0712890625,79.9210968017578,79.8034210205078,79.5984344482422,79.4901351928711,79.42822265625,79.3677749633789,79.2955017089844,79.2030258178711,79.1648406982422,79.1266632080078,79.0598602294922,78.9479522705078,78.8851547241211,78.79150390625,78.6422882080078,78.5242156982422,78.3759765625,78.2900390625,78.0231399536133,77.8016586303711,77.6224670410156,77.51220703125,77.4592742919922,77.3685607910156,77.2355499267578,77.0573272705078,76.9880828857422,76.9440383911133,76.646484375,76.5091781616211,76.2181625366211,75.9322204589844,75.84033203125,75.78955078125,75.6817398071289,75.6356430053711,75.3662109375,75.0476608276367,74.8175811767578,74.6222686767578,74.3638687133789,74.2090835571289,74.1868133544922,74.1458969116211,74.0862274169922,73.94921875,73.8860321044922,73.7185516357422,73.6120071411133,73.5562438964844,73.4501953125,73.421875,73.4929656982422,73.41162109375,73.3160171508789,73.2829132080078,73.1908187866211,72.8550796508789,72.7923889160156,72.7529296875,72.6661148071289,72.5431671142578,72.2757797241211,72.1618194580078,71.8167953491211,71.7605514526367,71.7347640991211,71.6007843017578,71.5142593383789,71.4220733642578,71.2566375732422,71.1673812866211,71.0935516357422,71.0227508544922,71.001953125,70.9523468017578,70.8928680419922,70.8928680419922,70.94677734375,70.8603515625,70.7645492553711,70.7152328491211,70.6624984741211,70.61328125,70.5842742919922,70.5401382446289,70.4890594482422,70.416015625,70.3289108276367,70.2258834838867,70.0956115722656,69.9599609375,69.7880859375,69.6638641357422,69.5651321411133,69.4009780883789,69.3683547973633,69.2493133544922,69.1536178588867,69.06494140625,69.04345703125,68.9869155883789,68.8511657714844,68.7371139526367,68.6627883911133,68.5840835571289,68.5592803955078,68.5565414428711,68.5936508178711,68.6006851196289,68.5726547241211,68.4957046508789,68.4150390625,68.2918930053711,68.1602554321289,68.1123046875,68.0476608276367,68.0570297241211,68.09033203125,68.1130828857422,68.0593719482422,68.0197219848633,67.9914016723633,67.9357376098633,67.86572265625,67.8050765991211,67.7350616455078,67.5280227661133,67.37158203125,67.2250061035156,67.0386657714844,66.8142547607422,66.7498092651367,66.7096633911133,66.6686477661133,66.6453094482422,66.6016616821289,66.5725555419922,66.5378952026367,66.5150375366211,66.4986343383789,66.3288040161133,66.1931686401367,66.0095672607422,66.01123046875,66.01318359375,66.0155258178711,66.0498046875,66.0626983642578,66.0785140991211,66.0888671875,66.1002960205078,66.0056610107422,65.9010696411133,65.8030242919922,65.7356491088867,65.6702117919922,65.5707015991211,65.4961929321289,65.3665008544922,65.2705078125,65.1708984375,65.0848693847656,65.0031204223633,64.9054718017578,64.8118133544922,64.7060546875,64.6041030883789,64.49609375,64.4431610107422,64.3181610107422,64.2087936401367,64.0132827758789,63.84814453125,63.6796836853027,63.4448204040527,63.2070274353027,63.0476608276367,62.8461875915527,62.6344757080078,62.4593734741211,62.2378921508789,62.0719718933105,61.9902381896973,61.8875999450684,61.7232437133789,61.6236343383789,61.52587890625,61.3850631713867,61.2714805603027,61.1607437133789,61.0970726013184,61.0653343200684,61.0079116821289,60.8791961669922,60.7411117553711,60.6029281616211,60.4647483825684,60.3266639709473,60.1884727478027,60.05029296875,59.9122047424316,59.7740249633789,59.6359405517578,59.4978523254395,59.3595695495605,59.2214813232422,59.0833969116211,58.9451217651367,58.8070335388184,58.6689491271973,58.5552749633789,58.4494132995605,58.2911148071289,58.1251983642578,57.9610328674316,57.6666984558105,57.4773483276367,57.3292961120605,57.1716766357422,56.9650382995605,56.7918968200684,56.5887680053711,56.4091796875,56.2579078674316,56.1004905700684,55.9756851196289,55.9757804870605,55.9759750366211,55.97607421875,55.9761734008789,55.9762687683105,55.9763679504395,55.9764633178711,55.9765625,55.9766578674316,55.9767570495605,55.9768562316895,55.9769515991211,55.97705078125,55.9771499633789,55.9773445129395,55.9774398803711,55.9349594116211,55.8390655517578,55.6786117553711,55.5452156066895,55.4870109558105,55.4343757629395,55.3883781433105,55.3197288513184,55.2496070861816,55.1623039245605,55.1018562316895,54.9523468017578,54.931640625,54.9037094116211,54.8538093566895,54.6779289245605,54.4728507995605,54.2718734741211,54.2149391174316,54.1209945678711,54.0051765441895,53.9263687133789,53.6853523254395,53.5007820129395,53.2500991821289,53.0558586120605,53.0125007629395,52.8705062866211,52.6968727111816,52.4938507080078,52.4675750732422,52.4585952758789,52.4621086120605,52.5171890258789,52.5732421875,52.6183586120605,52.6384735107422,52.5965805053711,52.5499992370605,52.4939460754395,52.4342765808105,52.3244132995605,52.2730445861816,52.1836891174316,52.0755844116211,52.0185546875,51.9607429504395,51.8982429504395,51.8441429138184,51.81103515625,51.78515625,51.7003898620605,51.6160163879395,51.5140609741211,51.3478546142578,51.29541015625,51.2923812866211,51.3133773803711,51.3138656616211,51.3017578125,51.2741203308105,51.2389640808105,51.1396484375,51.0648422241211,50.9398460388184,50.8307647705078,50.7826156616211,50.6849632263184,50.4717788696289,50.3311500549316,50.2755851745605,50.2525405883789,50.2529296875,50.2645492553711,50.2974624633789,50.4094734191895,50.6524391174316,50.8603515625,51.048828125,51.1107444763184,51.1771507263184,51.3107414245605,51.3766593933105,51.5435562133789,51.4937477111816,51.4310531616211,51.3663063049316,51.3102531433105,51.2181663513184,51.0579109191895,51.0207023620605,51.0093765258789,51.0403327941895,51.1537132263184,51.2499008178711,51.2940444946289,51.3334007263184,51.4157257080078,51.5396461486816,51.7326164245605,52.0487289428711,52.4267578125,52.5310516357422,52.77197265625,52.9107437133789,53.0789070129395,53.2003898620605,53.0857429504395,52.8375015258789,52.7738304138184,52.8875007629395,53.0416030883789,53.13525390625,53.1089820861816,53.06396484375,53.0785179138184,53.1324195861816,53.1702156066895,53.1375007629395,53.0694313049316,53.0345726013184,52.916015625,52.6776351928711,52.4832038879395,52.4203109741211,52.3848648071289,52.34033203125,52.1887702941895,52.1382789611816,52.0855484008789,52.0111312866211,51.9451179504395,51.7445297241211,51.6501007080078,51.615234375,51.2908210754395,51.1780281066895,50.9200210571289,50.7327156066895,50.6798858642578,50.5829086303711,50.5284194946289,50.4722671508789,50.4193344116211,50.3062477111816,50.1015625,49.9998054504395,49.8863296508789,49.7605476379395,49.6315422058105,49.5843772888184,49.4372062683105,49.3474617004395,49.34423828125,49.3621101379395,49.2858390808105,49.2056617736816,49.2322273254395,49.1842765808105,48.958984375,48.7743148803711,48.6101570129395,48.5860366821289,48.5412101745605,48.5091781616211,48.5023460388184,48.5185546875,48.5583953857422,48.6052703857422,48.6470680236816,48.6935539245605,48.7763671875,48.8835906982422,48.9502906799316,48.9593734741211,48.8318367004395,48.71435546875,48.6006813049316,48.5525398254395,48.4130859375,48.2756843566895,48.1669921875,48.1099586486816,47.9346656799316,47.6001968383789,47.48193359375,47.3873062133789,47.2923851013184,47.2020492553711,47.1307601928711,47.09326171875,47.1115226745605,47.1190452575684,47.0646476745605,47.0042953491211,46.8531227111816,46.6609344482422,46.6091804504395,46.70263671875,46.8529281616211,46.9622077941895,47.0142593383789,47.0313491821289,47.0181655883789,46.9534187316895,46.8529281616211,46.8020515441895,46.8231468200684,46.8895530700684,46.9919929504395,47.1295890808105,47.2483406066895,47.2952156066895,47.2976570129395,47.2947273254395,47.3264656066895,47.3763656616211,47.42919921875,47.5036125183105,47.599609375,47.7057609558105,47.849609375,48.0607414245605,48.1813468933105,48.2248039245605,48.3349609375,48.4342765808105,48.5999984741211,48.7589874267578,48.8102531433105,48.84326171875,48.8179664611816,48.7847671508789,48.7494125366211,48.7004890441895,48.666015625,48.6250953674316,48.6551780700684,48.7347640991211,48.8083992004395,48.9137687683105,49.0586891174316,49.3234367370605,49.3794898986816,49.4246063232422,49.498046875,49.6663093566895,49.822265625,49.9323234558105,50.1048851013184,50.2468757629395,50.3092765808105,50.3537101745605,50.5163078308105,50.6439437866211,50.7561531066895,50.7939453125,50.8824234008789,51.0178718566895,51.1634750366211,51.2699203491211,51.2907218933105,51.3010711669922,51.3445281982422,51.39599609375,51.4734344482422,51.6090850830078,51.775390625,52.0071296691895,52.2191429138184,52.3310546875,52.4230461120605,52.4961929321289,52.5711898803711,52.6177711486816,52.6351547241211,52.7281265258789,52.7350578308105,52.8205070495605,52.9026374816895,53.0383796691895,53.2273445129395,53.2472686767578,53.3380851745605,53.4486312866211,53.53466796875,53.6880874633789,53.7764625549316,53.9568328857422,54.04150390625,54.1397476196289,54.1911125183105,54.2978515625,54.4214859008789,54.443359375,54.4714813232422,54.5173797607422,54.5552711486816,54.59619140625,54.6361351013184,54.6500015258789,54.6378898620605,54.6062507629395,54.5656242370605,54.5460929870605,54.5729484558105,54.6416015625,54.7271461486816,54.8679695129395,55.0148429870605,55.1952171325684,55.3611335754395,55.5422859191895,55.6862335205078,55.7976570129395,55.92919921875,56.0497055053711,56.1044921875,56.1439476013184,56.3255844116211,56.4914054870605,56.56689453125,56.6202163696289,56.7903327941895,56.849609375,57.01171875,57.1790046691895,57.3125,57.4421844482422,57.5578117370605,57.6538124084473,57.7169914245605,57.7648429870605,57.8289031982422,57.8388671875,58.0451202392578,58.1747093200684,58.1884765625,58.3591766357422,58.5474586486816,58.66455078125,58.8140640258789,58.8836898803711,58.98486328125,59.0643539428711,59.1708984375,59.4523429870605,59.4951133728027,59.5239219665527,59.4978523254395,59.5230484008789,59.7511711120605,59.8124046325684,59.8877944946289,59.9551734924316,60.0052757263184,60.0585899353027,60.1121139526367,60.1867179870605,60.2880859375,60.4248046875,60.5084953308105,60.6379890441895,60.9422874450684,61.2268524169922,61.3894538879395,61.4650344848633,61.51220703125,61.5850563049316,61.5546875,61.4113311767578,61.3630905151367,61.0148429870605,60.9933586120605,60.9735374450684,60.6303749084473,60.4647483825684,60.4183578491211,60.3874969482422,60.2803726196289,60.0674858093262,60.0302734375,60.0655288696289,60.2336883544922,60.4254875183105,60.4993171691895,60.6703109741211,60.8284149169922,60.9375953674316,60.9945335388184,60.9794921875,60.8212890625,60.7744140625,60.8023452758789,60.8932609558105,60.9447250366211,61.0065460205078,61.0474586486816,61.2069358825684,61.4007835388184,61.5335922241211,61.7193374633789,61.8885765075684,61.9742202758789,62.0371131896973,62.0827178955078,62.0810585021973,62.0146484375,61.7662124633789,61.6598587036133,61.576171875,61.4368133544922,61.3109397888184,61.19921875,61.1627922058105,61.1859397888184,61.2289047241211,61.3116226196289,61.4009742736816,61.4985389709473,61.5265617370605,61.5349655151367,61.5191383361816,61.47412109375,61.4099578857422,61.3361282348633,61.2479515075684,61.0985336303711,60.9794921875,60.9855499267578,61.0735359191895,61.1131858825684,61.1131858825684,61.1131858825684,61.1437492370605,61.2310562133789,61.3336906433105,61.59814453125,61.9287109375,61.9856452941895,62.0023460388184,62.0402336120605,62.4990234375,62.5882797241211,62.6327133178711,63.0739288330078,63.1265602111816,63.1913032531738,63.2926788330078,63.4136695861816,63.58203125,63.7013702392578,63.72119140625,63.8470726013184,64.00390625,64.0373992919922,64.0628890991211,64.1994171142578,64.4612274169922,64.5251007080078,64.64990234375,64.8092727661133,64.9267578125,64.9954071044922,65.08837890625,65.1578140258789,65.1921920776367,65.2374038696289,65.31591796875,65.3781280517578,65.4343719482422,65.4769515991211,65.7078094482422,65.9142532348633,65.9546890258789,66.22265625,66.5553741455078,66.7544937133789,67.0983352661133,67.25732421875,67.4846649169922,67.693359375,67.8298873901367,67.93994140625,68.0738296508789,68.1558609008789,68.2093734741211,68.2440414428711,68.2252883911133,68.2062454223633,68.3019561767578,68.4384765625,68.5248031616211,68.712890625,68.8429641723633,68.9772491455078,69.2469711303711,69.4932632446289,69.740234375,69.8702087402344,69.9817352294922,70.08740234375,70.1824188232422,70.2933578491211,70.3714904785156,70.4171905517578,70.486328125,70.7380828857422,70.7903366088867,70.91015625,70.9917984008789,71.1262741088867,71.185546875,71.1591796875,71.1597671508789,71.1521530151367,71.052734375,71.0931625366211,71.33642578125,71.6771469116211,71.8873977661133,72.0044860839844,72.0656280517578,72.1053695678711,72.18603515625,72.2691421508789,72.3294906616211,72.3873062133789,72.3830032348633,72.404296875,72.44677734375,72.5302734375,72.5859375,72.5992202758789,72.5755920410156,72.5642547607422,72.5827102661133,72.6222686767578,72.7410125732422,72.9140625,73.1193313598633,73.2298812866211,73.2765655517578,73.3806610107422,73.5056610107422,73.5899429321289,73.6179733276367,73.6664047241211,73.71240234375,73.7155303955078,73.6789016723633,73.55419921875,73.3994140625,73.3056640625,73.2857437133789,73.3268585205078,73.3619155883789,73.3718719482422,73.4069290161133,73.4699172973633,73.6429672241211,73.7311553955078,73.8589782714844,74.0686569213867,74.2099609375,74.27734375,74.3515625,74.4027328491211,74.4292984008789,74.4304733276367,74.4519500732422,74.6814422607422,74.8341827392578,74.8868179321289,74.9889678955078,75.0521469116211,75.22021484375,75.3770523071289,75.3923797607422,75.3981475830078,75.4372024536133,75.6568374633789,75.69287109375,75.8806610107422,76.1405258178711,76.2666015625,76.4964828491211,76.5391616821289,76.6155319213867,76.7593765258789,76.8373031616211,76.7889633178711,76.7030258178711,76.6545944213867,76.4216842651367,76.4220733642578,76.4585952758789,76.4847640991211,76.5130844116211,76.57568359375,76.8207015991211,77.1324234008789,77.46923828125,77.7043914794922,77.7994155883789,77.8599624633789,78.0335006713867,78.1980514526367,78.4754867553711,78.7214813232422,78.9920883178711,79.1487274169922,79.4688491821289,79.5542907714844,79.7164077758789,79.8596725463867,79.9862289428711,80.06591796875,80.0720672607422,80.0863342285156,80.1272430419922,80.22021484375,80.2704086303711,80.34521484375,80.4236297607422,80.4522476196289,80.43359375,80.4214782714844,80.4480438232422,80.4910125732422,80.5506820678711,80.60546875,80.6504898071289,80.7352523803711,80.8130874633789,80.8773422241211,80.93408203125,80.9656219482422,81.0267562866211,81.1272430419922,81.1410140991211,81.1124038696289,81.0775451660156,81.0714797973633,81.1246109008789,81.3191375732422,81.3882827758789,81.41015625,81.4376983642578,81.4515609741211,81.4314498901367,81.4659194946289,81.6338882446289,81.7520523071289,81.9336929321289,82.0980453491211,82.2119140625,82.3263702392578,82.4939422607422,82.6117172241211,82.6929702758789,82.7185592651367,82.7608413696289,82.9190444946289,83.0192337036133,83.0927734375,83.1602478027344,83.2737274169922,83.3573226928711,83.5814514160156,83.7177734375,83.8597640991211,83.9451141357422,84.0023422241211,84.0993118286133,84.1759719848633,84.1945343017578,84.2578125,84.3232421875,84.4009780883789,84.4990234375,84.6073226928711,84.8389663696289,84.9240264892578,84.9894485473633,84.9997100830078,84.9751968383789,85.0007858276367,85.0764694213867,85.1365280151367,85.2101516723633,85.2326202392578,85.2918930053711,85.37158203125,85.4984359741211,85.8804702758789,85.93359375,85.9744110107422,86.0295944213867,86.0929718017578,86.1808624267578,86.2421875,86.2923812866211,86.41796875,86.5222702026367,86.6101608276367,86.6754913330078,86.7287139892578,86.7306671142578,86.6653366088867,86.6142578125,86.62646484375,86.71435546875,86.8121109008789,86.9529342651367,87.0009765625,87.0706024169922,87.1480407714844,87.2336959838867,87.296875,87.3228454589844],[40.9944343566895,40.9573249816895,40.9764671325684,41.0860366821289,41.1306648254395,41.1392555236816,41.1368141174316,41.1181640625,40.9944343566895],[41.8839836120605,41.7609367370605,41.6134757995605,41.341796875,41.1349601745605,40.9787101745605,40.9644508361816,40.9650382995605,40.9666976928711,40.9700202941895,40.9732437133789,40.9765625,40.9782218933105,40.9787101745605,41.1158180236816,41.2498054504395,41.4269523620605,41.5218734741211,41.53759765625,41.53271484375,41.3869132995605,41.2674789428711,41.1068344116211,41.0586929321289,40.9955101013184,40.970703125,40.9521484375,40.9166030883789,40.8897476196289,40.9058609008789,40.92236328125,40.8982391357422,40.8201179504395,40.8131828308105,40.6441421508789,40.4044914245605,40.2785186767578,40.2224617004395,40.1797866821289,40.1947288513184,40.1281280517578,40.1154289245605,39.99169921875,39.9368171691895,39.8962898254395,39.8609352111816,39.8191413879395,39.7614288330078,39.7458000183105,39.7316398620605,39.6869125366211,39.6580085754395,39.6371078491211,39.4909172058105,39.376953125,39.2875022888184,39.2281265258789,39.2217788696289,39.1901359558105,39.1154327392578,38.9619140625,38.8083953857422,38.6548805236816,38.5013656616211,38.3478507995605,38.1943359375,38.0408210754395,37.8873062133789,37.7972679138184,37.7574234008789,37.7261695861816,37.7110366821289,37.6701164245605,37.6220703125,37.6082038879395,37.6086921691895,37.6253929138184,37.6818351745605,37.68798828125,37.6768569946289,37.6591796875,37.6438484191895,37.5421867370605,37.3290023803711,37.1158218383789,36.9026374816895,36.689453125,36.4763641357422,36.2630844116211,36.0500030517578,35.8369140625,35.6237297058105,35.4105453491211,35.1974601745605,34.9842758178711,34.7710914611816,34.5579071044922,34.3447265625,34.1316413879395,34.0515632629395,33.9793968200684,33.9032211303711,33.9000015258789,33.9244117736816,33.9214859008789,33.9431648254395,34.0372047424316,34.08056640625,34.1117210388184,34.1609382629395,34.2725563049316,34.2925758361816,34.4108390808105,34.4817390441895,34.5352516174316,34.6019515991211,34.6491241455078,34.7267570495605,34.78759765625,34.7986335754395,34.8038101196289,34.7835960388184,34.8095703125,34.8509750366211,34.8983383178711,34.9412117004395,34.9652328491211,34.9764671325684,34.9782218933105,34.9775390625,34.9640617370605,34.9139633178711,34.8830070495605,34.90576171875,34.8662109375,34.8466796875,34.814453125,34.7734375,34.7424812316895,34.7232398986816,34.5891609191895,34.5225601196289,34.4478492736816,34.4072265625,34.3994140625,34.4417953491211,34.4376945495605,34.3928718566895,34.26708984375,34.1650390625,34.17822265625,34.1857414245605,34.1320343017578,33.97607421875,34.1768569946289,34.3801727294922,34.6398429870605,34.8783187866211,35.08447265625,35.2683563232422,35.2638664245605,35.2646484375,35.28759765625,35.3252944946289,35.3779296875,35.4240264892578,35.4686546325684,35.7450180053711,35.7914085388184,35.7884750366211,35.80029296875,35.779296875,35.7561531066895,35.7630844116211,35.8456039428711,35.9198265075684,35.9787101745605,36.02197265625,36.0819320678711,36.2718734741211,36.5530281066895,36.8236312866211,36.8482437133789,36.9055671691895,37.1545906066895,37.3825187683105,37.5754890441895,37.7628898620605,37.9449195861816,38.0861320495605,38.2252922058105,38.4515647888184,38.6080055236816,38.7527351379395,38.9677734375,39.1283187866211,39.2254905700684,39.4944343566895,39.5388641357422,39.6575202941895,39.7903327941895,39.8421859741211,40.01416015625,40.3160133361816,40.5287094116211,40.7652320861816,40.8726577758789,41.0208015441895,41.0872077941895,41.1404304504395,41.2208976745605,41.3189468383789,41.3724594116211,41.48193359375,41.7376937866211,41.8839836120605],[80.2093734741211,80.2291946411133,80.2461929321289,80.2351531982422,80.2162094116211,79.90966796875,79.8404312133789,79.76611328125,79.50390625,79.3543930053711,79.2935485839844,79.1484375,78.7425765991211,78.5431671142578,78.44287109375,78.3624038696289,78.3488235473633,78.3462829589844,78.1234359741211,77.9564437866211,77.8152313232422,77.7193374633789,77.5817337036133,77.2839889526367,77.1820297241211,76.9866180419922,76.90771484375,76.8240203857422,76.7083969116211,76.6611328125,76.6398468017578,76.6221694946289,76.5779266357422,76.5208969116211,76.4801788330078,76.3963851928711,76.3185501098633,76.25830078125,76.2060546875,76.1566467285156,76.0623016357422,76.0042953491211,75.8719711303711,75.6771469116211,75.6559600830078,75.6173782348633,75.58349609375,75.5555648803711,75.5207977294922,75.2410125732422,75.111328125,75.0044937133789,74.8656234741211,74.8351593017578,74.8111267089844,74.80126953125,74.841796875,74.8304672241211,74.7677688598633,74.6798782348633,74.6130905151367,74.4119110107422,74.24267578125,74.0851593017578,74.0205078125,73.9916000366211,73.9387741088867,73.8845672607422,73.8562469482422,73.8353500366211,73.8397445678711,73.8825149536133,73.9146499633789,73.9071273803711,73.8727493286133,73.8229522705078,73.7157287597656,73.6316375732422,73.5755844116211,73.4704132080078,73.3874053955078,73.3361282348633,73.2349624633789,73.1092758178711,72.9494094848633,72.8724594116211,72.6399459838867,72.5633773803711,72.490234375,72.3577117919922,72.2872009277344,72.2498016357422,72.22998046875,72.1473617553711,72.0841751098633,72.0427703857422,71.9910125732422,71.8059539794922,71.7786178588867,71.7256774902344,71.7353515625,71.7322235107422,71.6726608276367,71.5462875366211,71.5033187866211,71.505859375,71.5173797607422,71.5030288696289,71.4703140258789,71.404296875,71.3285140991211,71.2728500366211,71.2027359008789,71.1180648803711,71.0650405883789,71.0048828125,70.79931640625,70.7331008911133,70.6786117553711,70.6078109741211,70.5679702758789,70.5011749267578,70.39208984375,70.2448272705078,70.2092742919922,70.1710968017578,70.1368179321289,70.1016616821289,69.9559555053711,69.7720718383789,69.6669921875,69.5988311767578,69.4632797241211,69.3915023803711,69.2976608276367,69.2802734375,69.2291030883789,69.2447204589844,69.27880859375,69.3072280883789,69.3654251098633,69.4319305419922,69.4762649536133,69.4878921508789,69.4709930419922,69.46875,69.49365234375,69.5302734375,69.7652359008789,69.966796875,70.0714874267578,70.2744140625,70.37890625,70.4513626098633,70.51513671875,70.5568389892578,70.5992202758789,70.6241226196289,70.6443405151367,70.7385787963867,70.9463806152344,70.9609375,70.9580078125,70.9906234741211,71.0945281982422,71.3046875,71.3761749267578,71.4574279785156,71.5204162597656,71.5804748535156,71.6298828125,71.65087890625,71.6668014526367,71.6924819946289,71.7726593017578,71.8454055786133,71.9027328491211,71.9556655883789,71.9710998535156,72.0125961303711,72.1312484741211,72.19287109375,72.2328186035156,72.2346649169922,72.2540054321289,72.3577117919922,72.3892517089844,72.4059524536133,72.3697280883789,72.3690414428711,72.3826141357422,72.4020462036133,72.5674819946289,72.6040954589844,72.6795959472656,72.7488327026367,72.7738265991211,73.1128921508789,73.1369171142578,73.1321258544922,72.9900360107422,72.9259796142578,72.8666000366211,72.8309555053711,72.6583023071289,72.6204071044922,72.5059585571289,72.4273452758789,72.3640670776367,72.294921875,72.2130813598633,72.1873092651367,72.1806640625,72.1809539794922,72.1642532348633,72.1154327392578,72.0524444580078,71.95849609375,71.8786163330078,71.8580093383789,71.8257827758789,71.79248046875,71.7577133178711,71.70068359375,71.697265625,71.6851577758789,71.6649398803711,71.6374969482422,71.6022491455078,71.61962890625,71.6062469482422,71.5855484008789,71.5456008911133,71.5,71.4208984375,71.4084014892578,71.39306640625,71.298828125,71.2234344482422,71.1107406616211,71.0259780883789,70.9626007080078,70.8604431152344,70.7824172973633,70.734375,70.6888656616211,70.6458969116211,70.4713821411133,70.4078063964844,70.2900390625,70.2008743286133,70.1769561767578,70.1809616088867,70.4549789428711,70.5628890991211,70.630859375,70.7277374267578,70.8033218383789,70.8418960571289,70.8566436767578,70.9103546142578,71.0322265625,71.228515625,71.2323226928711,71.2126922607422,71.1299819946289,71.0360336303711,70.97900390625,70.94677734375,70.8928680419922,70.8928680419922,70.9523468017578,71.001953125,71.0227508544922,71.0935516357422,71.1673812866211,71.2566375732422,71.4220733642578,71.5142593383789,71.6007843017578,71.7347640991211,71.7605514526367,71.8167953491211,72.1618194580078,72.2757797241211,72.5431671142578,72.6661148071289,72.7529296875,72.7923889160156,72.8550796508789,73.1908187866211,73.2829132080078,73.3160171508789,73.41162109375,73.4929656982422,73.421875,73.4501953125,73.5562438964844,73.6120071411133,73.7185516357422,73.8860321044922,73.94921875,74.0862274169922,74.1458969116211,74.1868133544922,74.2090835571289,74.3638687133789,74.6222686767578,74.8175811767578,75.0476608276367,75.3662109375,75.6356430053711,75.6817398071289,75.78955078125,75.84033203125,75.9322204589844,76.2181625366211,76.5091781616211,76.646484375,76.9440383911133,76.9880828857422,77.0573272705078,77.2355499267578,77.3685607910156,77.4592742919922,77.51220703125,77.6224670410156,77.8016586303711,78.0231399536133,78.2900390625,78.3759765625,78.5242156982422,78.6422882080078,78.79150390625,78.8851547241211,78.9479522705078,79.0598602294922,79.1266632080078,79.1648406982422,79.2030258178711,79.2955017089844,79.3677749633789,79.42822265625,79.4901351928711,79.5984344482422,79.8034210205078,79.9210968017578,80.0712890625,80.2093734741211],[70.70166015625,70.6982421875,70.6641616821289,70.5670928955078,70.4977569580078,70.4828109741211,70.4892578125,70.5186538696289,70.5595703125,70.6121063232422,70.70166015625],[71.7796859741211,71.7899398803711,71.7653274536133,71.7365188598633,71.6812515258789,71.6689453125,71.7058563232422,71.7529296875,71.7796859741211],[71.2061462402344,71.2156219482422,71.1792984008789,71.2287063598633,71.1302719116211,71.0803680419922,71.0241165161133,71.0054702758789,70.9762649536133,70.9606475830078,70.9744186401367,71.0144500732422,71.041015625,71.0448303222656,71.0436553955078,71.01171875,71.0642623901367,71.15625,71.2061462402344],[103.317771911621,103.28125,103.222946166992,103.223442077637,103.317771911621],[103.045120239258,103.02734375,103.010551452637,102.993354797363,102.995018005371,103.007514953613,103.036819458008,103.045120239258],[104.426368713379,104.262397766113,103.937103271484,103.901756286621,103.870506286621,103.840530395508,103.661911010742,103.587104797363,103.5322265625,103.540428161621,103.592094421387,103.680854797363,103.721870422363,103.654296875,103.595024108887,103.532424926758,103.466697692871,103.411323547363,103.353614807129,103.272163391113,103.15283203125,103.1064453125,103.091110229492,103.107421875,103.12548828125,103.010551452637,103.004196166992,102.948631286621,102.932327270508,102.933891296387,102.918067932129,102.736618041992,102.706253051758,102.737403869629,102.755661010742,102.703323364258,102.629684448242,102.499610900879,102.49072265625,102.461715698242,102.422653198242,102.362983703613,102.330764770508,102.319725036621,102.336326599121,102.428512573242,102.546875,102.565528869629,102.544723510742,102.620407104492,102.728904724121,102.812797546387,102.873245239258,102.909278869629,103.031051635742,103.199409484863,103.3134765625,103.432426452637,103.54638671875,103.600387573242,103.741889953613,103.818359375,103.898628234863,103.981834411621,104.054298400879,104.227737426758,104.41162109375,104.575782775879,104.778999328613,104.878814697266,104.9697265625,104.982421875,105.00341796875,105.03369140625,105.074119567871,105.1259765625,105.169143676758,105.183303833008,105.185546875,105.20703125,105.245704650879,105.284866333008,105.350196838379,105.392677307129,105.531547546387,105.73974609375,105.764068603516,105.831443786621,105.904495239258,106.066802978516,106.124702453613,106.0966796875,106.004096984863,105.978912353516,106.008399963379,106.165237426758,106.190719604492,106.225387573242,106.267967224121,106.35498046875,106.446975708008,106.50146484375,106.531150817871,106.563667297363,106.599220275879,106.665428161621,106.738182067871,106.783500671387,106.819923400879,106.913185119629,106.938087463379,106.9921875,107.030174255371,107.062400817871,107.109375,107.206642150879,107.262298583984,107.292678833008,107.3798828125,107.414741516113,107.465141296387,107.519432067871,107.535255432129,107.4931640625,107.448440551758,107.364456176758,107.3603515625,107.331443786621,107.342575073242,107.362113952637,107.389450073242,107.462303161621,107.528610229492,107.593948364258,107.60546875,107.545509338379,107.475387573242,107.481544494629,107.511520385742,107.543556213379,107.555465698242,107.5380859375,107.506446838379,107.445991516113,107.393356323242,107.330078125,107.279685974121,107.212104797363,107.158981323242,107.050682067871,106.9306640625,106.7646484375,106.700096130371,106.630958557129,106.499610900879,106.413871765137,106.417778015137,106.410743713379,106.412506103516,106.399215698242,106.33984375,106.239158630371,106.102928161621,106.006050109863,105.956253051758,105.926559448242,105.889846801758,105.851463317871,105.838470458984,105.835350036621,105.85400390625,105.860939025879,105.856056213379,105.8916015625,106.099510192871,106.160942077637,106.16796875,106.131546020508,106.163963317871,106.098823547363,105.990135192871,105.938179016113,105.875190734863,105.853324890137,105.810745239258,105.755073547363,105.69775390625,105.576560974121,105.452728271484,105.40576171875,105.386528015137,105.314643859863,105.284278869629,105.159469604492,105.045700073242,105.022262573242,105.0361328125,105.061126708984,105.04638671875,104.98388671875,104.901268005371,104.8505859375,104.8154296875,104.689651489258,104.564262390137,104.514068603516,104.466995239258,104.426368713379],[-151.782623291016,-151.790878295898,-151.806686401367,-151.815963745117,-151.819137573242,-151.813278198242,-151.802780151367,-151.791122436523,-151.782623291016],[-155.863830566406,-155.887115478516,-155.914352416992,-155.927932739258,-155.928619384766,-155.919387817383,-155.910781860352,-155.872268676758,-155.862350463867,-155.863830566406],[-174.512939453125,-174.501129150391,-174.501022338867,-174.506744384766,-174.523880004883,-174.529235839844,-174.516738891602,-174.511474609375,-174.523040771484,-174.533111572266,-174.540679931641,-174.540832519531,-174.531402587891,-174.512939453125],[-172.214538574219,-172.208297729492,-172.193893432617,-172.180953979492,-172.188827514648,-172.215225219727,-172.228317260742,-172.212203979492,-172.197845458984,-172.196395874023,-172.197204589844,-172.203857421875,-172.214752197266,-172.214538574219],[-171.233200073242,-171.243026733398,-171.254531860352,-171.261764526367,-171.261962890625,-171.252395629883,-171.239395141602,-171.231887817383,-171.233200073242],[-154.956253051758,-154.959030151367,-154.971084594727,-154.99462890625,-155.014602661133,-155.015029907227,-154.986968994141,-154.951217651367,-154.943359375,-154.950057983398,-154.956253051758],[-171.085159301758,-171.089797973633,-171.096725463867,-171.091751098633,-171.087707519531,-171.081008911133,-171.085159301758],[-171.697616577148,-171.664993286133,-171.6396484375,-171.627639770508,-171.62841796875,-171.647369384766,-171.670608520508,-171.687316894531,-171.696090698242,-171.698287963867,-171.678375244141,-171.655364990234,-171.638534545898,-171.639739990234,-171.660202026367,-171.672653198242,-171.688034057617,-171.705963134766,-171.718170166016,-171.724807739258,-171.727630615234,-171.725143432617,-171.718887329102,-171.697616577148],[174.773239135742,174.778915405273,174.755966186523,174.748443603516,174.741012573242,174.716796875,174.744140625,174.7666015625,174.773239135742],[169.551071166992,169.541702270508,169.52294921875,169.525588989258,169.538665771484,169.555282592773,169.551071166992],[174.508697509766,174.476364135742,174.464065551758,174.479675292969,174.452728271484,174.407821655273,174.381057739258,174.39404296875,174.438766479492,174.474807739258,174.495407104492,174.508697509766],[173.032821655273,173.086532592773,173.079498291016,173.061416625977,172.991104125977,172.969924926758,173.03857421875,173.065048217773,173.025588989258,173.009963989258,173.003707885742,172.990051269531,173.003707885742,173.032821655273],[173.038375854492,173.011322021484,173.028610229492,173.143356323242,173.1533203125,173.171875,173.171478271484,173.1630859375,173.106353759766,173.061721801758,173.038375854492],[173.029403686523,172.993255615234,173.020416259766,173.02783203125,173.023635864258,173.037689208984,173.042678833008,173.045227050781,173.029403686523],[173.018753051758,173.023635864258,172.966598510742,172.932708740234,172.934768676758,172.950103759766,172.969146728516,172.981552124023,173.018753051758],[-157.342132568359,-157.17578125,-157.246139526367,-157.420120239258,-157.578948974609,-157.531494140625,-157.508209228516,-157.435852050781,-157.393218994141,-157.365188598633,-157.4921875,-157.44189453125,-157.321884155273,-157.342132568359],[172.84423828125,172.770324707031,172.75048828125,172.77734375,172.826950073242,172.887115478516,172.880264282227,172.84423828125],[172.969619750977,172.90625,172.887115478516,172.962509155273,172.962203979492,172.969619750977],[-159.339065551758,-159.259323120117,-159.274749755859,-159.332275390625,-159.358734130859,-159.313674926758,-159.306259155273,-159.326812744141,-159.354202270508,-159.373184204102,-159.377777099609,-159.409027099609,-159.390960693359,-159.369049072266,-159.339065551758],[-62.5322265625,-62.5824203491211,-62.6249046325684,-62.6152801513672,-62.5747108459473,-62.5341796875,-62.5322265625],[-62.6306610107422,-62.656494140625,-62.7020034790039,-62.7755393981934,-62.8389167785645,-62.8404808044434,-62.8394050598145,-62.8270492553711,-62.7946281433105,-62.7137184143066,-62.67578125,-62.6405258178711,-62.6306610107422],[126.326957702637,126.282028198242,126.240226745605,126.228996276855,126.178718566895,126.165626525879,126.199409484863,126.337692260742,126.695503234863,126.759872436523,126.901168823242,126.931251525879,126.905364990234,126.872856140137,126.709175109863,126.581741333008,126.326957702637],[126.753021240234,126.769920349121,126.689064025879,126.646095275879,126.65185546875,126.699996948242,126.753021240234],[126.233688354492,126.169723510742,126.1337890625,126.108596801758,126.122856140137,126.227043151855,126.247451782227,126.343841552734,126.379875183105,126.335456848145,126.233688354492],[127.799026489258,127.787796020508,127.737312316895,127.787117004395,127.799026489258],[126.171966552734,126.158782958984,126.115234375,126.070610046387,126.052055358887,126.007522583008,126.078422546387,126.168548583984,126.171966552734],[128.065811157227,128.0546875,127.983985900879,127.941795349121,127.896873474121,127.873435974121,127.838287353516,127.832221984863,127.915428161621,127.965629577637,128.037994384766,128.065811157227],[128.741012573242,128.646682739258,128.51953125,128.4892578125,128.5859375,128.66796875,128.721878051758,128.741012573242],[126.417579650879,126.40380859375,126.337501525879,126.318557739258,126.386627197266,126.417579650879],[130.916015625,130.87060546875,130.816802978516,130.810256958008,130.83837890625,130.903717041016,130.934280395508,130.916015625],[126.520706176758,126.516014099121,126.460845947266,126.423347473145,126.4072265625,126.369331359863,126.41162109375,126.493560791016,126.520706176758],[128.374603271484,128.618835449219,128.852447509766,129.051559448242,129.335144042969,129.418258666992,129.426162719727,129.473434448242,129.433013916016,129.445022583008,129.427154541016,129.392578125,129.391311645508,129.402435302734,129.403518676758,129.42578125,129.458297729492,129.509765625,129.572860717773,129.561721801758,129.485458374023,129.419128417969,129.329010009766,129.214157104492,129.076751708984,128.980072021484,128.795700073242,128.642578125,128.510940551758,128.458099365234,128.418853759766,128.447647094727,128.443954467773,128.3876953125,128.275985717773,128.15234375,128.094528198242,128.036224365234,127.976760864258,127.873245239258,127.714836120605,127.659080505371,127.639350891113,127.662506103516,127.7421875,127.715423583984,127.632423400879,127.565437316895,127.523635864258,127.476959228516,127.404296875,127.389656066895,127.423439025879,127.479103088379,127.401168823242,127.380569458008,127.324600219727,127.173439025879,127.194923400879,127.260551452637,127.268646240234,127.247062683105,127.030769348145,126.897453308105,126.826271057129,126.796478271484,126.754783630371,126.610832214355,126.584373474121,126.531440734863,126.50830078125,126.506439208984,126.481742858887,126.332611083984,126.264450073242,126.301071166992,126.425582885742,126.524513244629,126.504981994629,126.472846984863,126.53857421875,126.593360900879,126.547958374023,126.478523254395,126.420707702637,126.397857666016,126.327438354492,126.291114807129,126.360549926758,126.395904541016,126.46044921875,126.492767333984,126.582229614258,126.614059448242,126.564933776855,126.486526489258,126.488479614258,126.541893005371,126.6015625,126.717384338379,126.753021240234,126.719627380371,126.6474609375,126.663673400879,126.693450927734,126.682319641113,126.597061157227,126.540428161621,126.557228088379,126.544242858887,126.551956176758,126.548240661621,126.506645202637,126.487701416016,126.433006286621,126.388771057129,126.230667114258,126.180862426758,126.160545349121,126.217185974121,126.3515625,126.4287109375,126.487014770508,126.577735900879,126.686714172363,126.784469604492,126.838768005371,126.872077941895,126.879096984863,126.958015441895,126.976852416992,126.959762573242,126.868949890137,126.787406921387,126.776069641113,126.746383666992,126.790519714355,126.696197509766,126.650299072266,126.656845092773,126.607620239258,126.580177307129,126.563377380371,126.577735900879,126.620697021484,126.633895874023,126.664543151855,126.66650390625,126.666801452637,126.754295349121,126.87890625,126.940032958984,127.009658813477,127.090324401855,127.169532775879,127.294044494629,127.53271484375,127.579490661621,127.745506286621,127.78466796875,127.905281066895,128.038955688477,128.106246948242,128.168655395508,128.22314453125,128.279296875,128.339462280273,128.374603271484],[21.5625,21.5608386993408,21.3895492553711,21.3317375183105,21.2975578308105,21.28662109375,21.25634765625,21.2060546875,21.1424808502197,21.0597667694092,20.7781257629395,20.7503910064697,20.744140625,20.7250003814697,20.6949214935303,20.5785140991211,20.5662117004395,20.5814456939697,20.5753898620605,20.5228519439697,20.4854507446289,20.4083003997803,20.3482418060303,20.2405261993408,20.1857414245605,20.103515625,20.06396484375,20.0703125,20.0892581939697,20.0657234191895,20.0294914245605,20.0542984008789,20.1299819946289,20.1925773620605,20.2151374816895,20.3443374633789,20.4688472747803,20.48681640625,20.4583988189697,20.47509765625,20.6240234375,20.6485347747803,20.6576175689697,20.6375980377197,20.6096668243408,20.6231441497803,20.7005863189697,20.7633781433105,20.8005867004395,20.8238277435303,20.8238277435303,20.8444347381592,20.8907222747803,20.9676742553711,21.0570316314697,21.1270503997803,21.22265625,21.2371082305908,21.3231449127197,21.4030284881592,21.390625,21.6625003814697,21.7238273620605,21.7529296875,21.7521476745605,21.7306632995605,21.6190433502197,21.60986328125,21.5299797058105,21.5189437866211,21.5416030883789,21.5625],[48.275390625,48.21826171875,48.1796875,48.142578125,48.0814437866211,48.11474609375,48.1134796142578,48.1201171875,48.1388664245605,48.1585960388184,48.1847648620605,48.2277336120605,48.3482437133789,48.3473625183105,48.3402328491211,48.275390625],[48.4424819946289,48.2687492370605,48.0496101379395,47.8719711303711,47.6712875366211,47.5831069946289,47.55322265625,47.5212898254395,47.4332008361816,47.1387672424316,46.9822273254395,46.7248039245605,46.5314445495605,46.6937522888184,46.7693328857422,46.9058609008789,46.9759750366211,47.0436515808105,47.10205078125,47.1143531799316,47.1482429504395,47.2232398986816,47.3313446044922,47.5148429870605,47.6437492370605,47.6727523803711,47.75390625,47.9787101745605,47.9736328125,48.0056648254395,48.0773429870605,48.1361351013184,48.1434555053711,48.0894546508789,48.04833984375,47.9696273803711,47.8174819946289,47.7252960205078,47.72265625,47.8453102111816,47.9353485107422,47.9981422424316,48.0514640808105,48.0863265991211,48.1003913879395,48.1837882995605,48.2529296875,48.3392562866211,48.3712921142578,48.3896484375,48.4424819946289],[102.12744140625,102.183013916016,102.301361083984,102.442672729492,102.487503051758,102.58251953125,102.609664916992,102.631248474121,102.640815734863,102.662010192871,102.6953125,102.738571166992,102.771095275879,102.798240661621,102.81591796875,102.84521484375,102.876174926758,102.917671203613,102.949607849121,102.959182739258,102.948631286621,102.909568786621,102.887496948242,102.872268676758,102.851173400879,102.8837890625,103.1044921875,103.210739135742,103.463577270508,103.5546875,103.635063171387,103.714454650879,103.79052734375,103.882034301758,104.052047729492,104.101364135742,104.1953125,104.349609375,104.46142578125,104.530372619629,104.583198547363,104.5751953125,104.53271484375,104.478607177734,104.407814025879,104.367774963379,104.392189025879,104.496192932129,104.618850708008,104.656448364258,104.661911010742,104.676948547363,104.69873046875,104.812698364258,104.847846984863,104.888671875,104.92919921875,104.927925109863,104.845802307129,104.815132141113,104.8017578125,104.7431640625,104.587890625,104.546287536621,104.259864807129,104.127151489258,104.062797546387,104.032028198242,104.013473510742,104.051559448242,104.062889099121,104.027534484863,103.932029724121,103.896385192871,103.8916015625,103.918357849121,104.00634765625,104.108589172363,104.44580078125,104.517967224121,104.61328125,104.716506958008,104.9931640625,105.115135192871,105.146484375,105.145408630371,105.113479614258,105.087005615234,105.085838317871,105.114547729492,105.163284301758,105.273239135742,105.33349609375,105.400001525879,105.458206176758,105.5185546875,105.588478088379,105.59765625,105.627243041992,105.69140625,105.779495239258,105.902732849121,105.973533630371,106.006248474121,106.26953125,106.333396911621,106.425979614258,106.46533203125,106.502243041992,106.525978088379,106.53369140625,106.546188354492,106.593650817871,106.637504577637,106.656440734863,106.696090698242,106.739555358887,106.791595458984,106.832420349121,106.85107421875,106.892776489258,106.9306640625,107.001953125,107.069725036621,107.217384338379,107.296485900879,107.35009765625,107.396385192871,107.41015625,107.391990661621,107.360641479492,107.188873291016,107.165924072266,107.189552307129,107.232620239258,107.279396057129,107.338775634766,107.459571838379,107.564254760742,107.621681213379,107.653121948242,107.633689880371,107.589645385742,107.555274963379,107.496292114258,107.480369567871,107.504684448242,107.524513244629,107.513771057129,107.519432067871,107.465141296387,107.414741516113,107.3798828125,107.292678833008,107.262298583984,107.206642150879,107.109375,107.062400817871,107.030174255371,106.9921875,106.938087463379,106.913185119629,106.819923400879,106.783500671387,106.738182067871,106.665428161621,106.599220275879,106.563667297363,106.531150817871,106.50146484375,106.446975708008,106.35498046875,106.267967224121,106.225387573242,106.190719604492,106.165237426758,106.008399963379,105.978912353516,106.004096984863,106.0966796875,106.124702453613,106.066802978516,105.904495239258,105.831443786621,105.764068603516,105.73974609375,105.531547546387,105.392677307129,105.350196838379,105.284866333008,105.245704650879,105.20703125,105.185546875,105.183303833008,105.24365234375,105.342185974121,105.422660827637,105.4755859375,105.497360229492,105.500198364258,105.523048400879,105.546684265137,105.533393859863,105.490425109863,105.490425109863,105.505859375,105.51318359375,105.57373046875,105.615623474121,105.638870239258,105.641014099121,105.6220703125,105.562400817871,105.462013244629,105.39892578125,105.373245239258,105.375587463379,105.40625,105.330665588379,105.148735046387,105.047172546387,105.025779724121,104.949897766113,104.8193359375,104.750587463379,104.743553161621,104.758979797363,104.816017150879,104.739646911621,104.655860900879,104.539253234863,104.428123474121,104.322654724121,104.196189880371,104.048728942871,103.949615478516,103.898826599121,103.792282104492,103.629684448242,103.487983703613,103.366989135742,103.288276672363,103.251754760742,103.248924255371,103.279594421387,103.26318359375,103.19970703125,103.148536682129,103.091209411621,103.051368713379,102.991401672363,102.898628234863,102.807418823242,102.717575073242,102.675193786621,102.680076599121,102.66064453125,102.616798400879,102.596092224121,102.598236083984,102.552536010742,102.458786010742,102.351852416992,102.231643676758,102.148246765137,102.101463317871,102.034568786621,101.947456359863,101.87548828125,101.818656921387,101.774803161621,101.744140625,101.6875,101.563667297363,101.555076599121,101.413673400879,101.299713134766,101.16748046875,101.105171203613,101.045700073242,100.955856323242,100.908500671387,100.9990234375,101.11328125,101.143951416016,101.148727416992,101.137496948242,101.0927734375,101.050582885742,101.046974182129,101.060447692871,101.106346130371,101.16552734375,101.220504760742,101.286331176758,101.279884338379,101.2265625,101.197555541992,101.220802307129,101.2119140625,101.154685974121,100.966506958008,100.906051635742,100.858200073242,100.806838989258,100.743942260742,100.62548828125,100.513572692871,100.420120239258,100.397651672363,100.466209411621,100.514549255371,100.543067932129,100.539939880371,100.51953125,100.491600036621,100.431549072266,100.373146057129,100.317962646484,100.266014099121,100.218070983887,100.174125671387,100.139747619629,100.114936828613,100.122459411621,100.129684448242,100.183891296387,100.249313354492,100.326072692871,100.407424926758,100.493362426758,100.565139770508,100.622947692871,100.61767578125,100.54931640625,100.522262573242,100.5361328125,100.566596984863,100.613670349121,100.6591796875,100.703125,100.756645202637,100.819534301758,100.927543640137,101.080375671387,101.138862609863,101.196678161621,101.175392150879,101.20556640625,101.219917297363,101.211822509766,101.224411010742,101.247848510742,101.281448364258,101.443557739258,101.542381286621,101.583885192871,101.621681213379,101.668556213379,101.704788208008,101.728126525879,101.783493041992,101.800590515137,101.802047729492,101.763084411621,101.722946166992,101.724220275879,101.743453979492,101.747268676758,101.743942260742,101.736518859863,101.699607849121,101.602928161621,101.575782775879,101.560256958008,101.561820983887,101.537307739258,101.524513244629,101.56787109375,101.619926452637,101.646194458008,101.671485900879,101.70751953125,101.73876953125,101.759963989258,101.841796875,101.945411682129,102.0244140625,102.091506958008,102.12744140625],[35.869140625,35.8407249450684,35.7874984741211,35.7344741821289,35.6272468566895,35.6029319763184,35.5792961120605,35.5325202941895,35.4931640625,35.4112319946289,35.3088874816895,35.2233390808105,35.1085929870605,35.1550788879395,35.2035140991211,35.2513694763184,35.3357429504395,35.5108413696289,35.61181640625,35.6478500366211,35.8042945861816,35.92138671875,35.9779281616211,35.9762687683105,36.1510734558105,36.2635726928711,36.2962875366211,36.3838882446289,36.4330062866211,36.388671875,36.3262672424316,36.3298835754395,36.37646484375,36.45556640625,36.50439453125,36.5849609375,36.53515625,36.45751953125,36.4228515625,36.3548851013184,36.2978515625,36.27783203125,36.2822265625,36.36279296875,36.3650398254395,36.3485336303711,36.2833976745605,36.1994132995605,36.1498031616211,36.0921897888184,36.0188484191895,35.9861335754395,35.9684562683105,35.9423828125,35.9716796875,36.0266609191895,36.0344734191895,36.0222625732422,35.9675788879395,35.9265632629395,35.9147453308105,35.869140625],[-8.48642539978027,-8.46728515625,-8.43715858459473,-8.40874004364014,-8.296630859375,-8.30234336853027,-8.32451152801514,-8.32509708404541,-8.33256816864014,-8.40122032165527,-8.60356521606445,-8.587890625,-8.53955078125,-8.49033260345459,-8.44990253448486,-8.39931678771973,-8.34487342834473,-8.287109375,-8.203857421875,-8.13100528717041,-8.06894588470459,-7.98159170150757,-7.88862323760986,-7.85551738739014,-7.833251953125,-7.80092716217041,-7.79653310775757,-7.73037147521973,-7.63613271713257,-7.513916015625,-7.48281288146973,-7.46943330764771,-7.45439481735229,-7.42373085021973,-7.39990186691284,-7.41245079040527,-7.42890548706055,-7.42983436584473,-7.48520517349243,-7.494140625,-7.509765625,-7.56889629364014,-7.5693359375,-7.58505916595459,-7.59121131896973,-7.57465839385986,-7.57158231735229,-7.54497098922729,-7.66000938415527,-7.99824190139771,-8.259033203125,-9.13217830657959,-9.374755859375,-9.65439510345459,-10.2763671875,-10.4181632995605,-10.5970697402954,-10.7076168060303,-10.7855958938599,-10.8490238189697,-11.0045413970947,-11.2916021347046,-11.5075187683105,-11.4545412063599,-11.3766603469849,-11.2676763534546,-11.1661128997803,-11.0854005813599,-11.000244140625,-10.8780755996704,-10.6913089752197,-10.6474609375,-10.6175785064697,-10.5708494186401,-10.5167474746704,-10.3895511627197,-10.3598136901855,-10.3146486282349,-10.2857427597046,-10.283203125,-10.2330570220947,-10.1474123001099,-10.09765625,-10.07568359375,-10.0643558502197,-9.80473613739014,-9.781982421875,-9.76826190948486,-9.735595703125,-9.71689414978027,-9.701171875,-9.68388652801514,-9.66357421875,-9.64321327209473,-9.61015605926514,-9.55390644073486,-9.51826190948486,-9.522216796875,-9.50849628448486,-9.484130859375,-9.47114276885986,-9.46455001831055,-9.45112323760986,-9.44155216217041,-9.44638633728027,-9.43632793426514,-9.39492225646973,-9.369140625,-9.36894512176514,-9.38398456573486,-9.41147518157959,-9.45976543426514,-9.46381855010986,-9.43510723114014,-9.39165019989014,-9.35532283782959,-9.26328086853027,-9.21518516540527,-9.1728515625,-9.13481426239014,-9.11757850646973,-9.05234336853027,-8.9765625,-8.96098613739014,-8.93842792510986,-8.8896484375,-8.85551738739014,-8.82792949676514,-8.76914024353027,-8.740234375,-8.73261737823486,-8.72944355010986,-8.70830059051514,-8.65976524353027,-8.60732460021973,-8.578857421875,-8.56440448760986,-8.52226543426514,-8.48642539978027],[25.1504878997803,25.1120109558105,25.0572261810303,25.0226554870605,24.9299812316895,24.8527336120605,24.8599605560303,24.8775405883789,24.9294929504395,24.9739265441895,24.96142578125,24.9230480194092,24.8775405883789,24.7264652252197,24.7032241821289,24.7116203308105,24.8037109375,24.8108406066895,24.8659191131592,24.9161128997803,24.9716796875,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9802722930908,24.9800777435303,24.9798831939697,24.9796867370605,24.9794921875,24.9763679504395,24.9732437133789,24.97021484375,24.9669914245605,24.7204093933105,24.4736328125,24.2269515991211,23.9802742004395,23.9802742004395,23.9802742004395,23.9802742004395,23.9802742004395,23.5012683868408,23.0221691131592,22.5430660247803,22.0640621185303,21.5849609375,21.1058597564697,20.6267585754395,20.1476554870605,19.6685543060303,19.189453125,18.71044921875,18.2313480377197,17.7522449493408,17.2732429504395,16.7941417694092,16.3150386810303,15.9840822219849,15.6271486282349,15.3474607467651,14.9790048599243,14.9789056777954,14.5556640625,14.2307615280151,14.2155275344849,14.2006826400757,13.8626956939697,13.5986337661743,13.4812498092651,12.9835939407349,12.48876953125,11.9678707122803,11.873046875,11.7669925689697,11.6242189407349,11.5369148254395,11.5076169967651,11.1082029342651,10.6861324310303,10.43896484375,10.3958988189697,10.3257808685303,10.255859375,10.2186527252197,10.1195316314697,10.0281248092651,10.01904296875,10.0006837844849,9.78105449676514,9.58125019073486,9.44824123382568,9.42236328125,9.43789005279541,9.49140548706055,9.68496131896973,9.85937595367432,9.88320350646973,9.89443302154541,9.83710956573486,9.79541015625,9.75253963470459,9.74755859375,9.82529258728027,9.916015625,9.85820293426514,9.81562519073486,9.84257793426514,9.82070255279541,9.80527400970459,9.74589920043945,9.67265605926514,9.64013671875,9.54619216918945,9.39101600646973,9.31025409698486,9.42099571228027,9.51874923706055,9.63798904418945,9.80742168426514,9.89501953125,9.93251991271973,10.0597658157349,10.1259765625,10.2164068222046,10.2560548782349,10.2570314407349,10.2433595657349,10.1726570129395,10.1149415969849,10.1598634719849,10.1959962844849,10.2746095657349,10.3060541152954,10.4757814407349,10.5436515808105,10.5955076217651,10.60888671875,10.6830072402954,10.7715826034546,10.8263673782349,11.0051755905151,11.1682615280151,11.3580074310303,11.5049810409546,11.5359373092651,11.5337886810303,11.4539060592651,11.4539060592651,11.4591798782349,11.4671878814697,11.50244140625,11.5045900344849,11.6571292877197,11.8134765625,12.2798824310303,12.4270505905151,12.7535152435303,13.1380853652954,13.2834959030151,13.5363273620605,13.6477537155151,13.8353519439697,14.1556644439697,14.2371091842651,14.423828125,14.5133781433105,15.1765623092651,15.2668952941895,15.3590822219849,15.3630867004395,15.4140615463257,15.4963865280151,15.5958013534546,15.7059564590454,15.8322267532349,16.123046875,16.4509773254395,16.7815418243408,17.3492183685303,17.8304691314697,17.9493179321289,18.1904296875,18.6698226928711,18.9364261627197,19.1237297058105,19.2916984558105,19.58984375,19.7132816314697,20.01318359375,20.1115226745605,20.1509780883789,20.14111328125,20.1038074493408,20.02001953125,19.9612293243408,19.9263687133789,19.9734363555908,20.0309581756592,20.1214847564697,20.37060546875,20.62109375,21.0623035430908,21.3187503814697,21.4247074127197,21.6359367370605,21.7213859558105,21.8394527435303,22.1874008178711,22.3406257629395,22.5234375,22.7541007995605,22.9168949127197,23.0906238555908,23.1296882629395,23.1104488372803,23.1062507629395,23.2863273620605,23.7976570129395,23.8984375,24.0389652252197,24.1296863555908,24.4797859191895,24.6838874816895,24.8785152435303,24.95068359375,25.0249996185303,25.1150398254395,25.1504878997803],[-60.8952140808105,-60.9514122009277,-61.0606422424316,-61.0731468200684,-61.0635719299316,-60.9966773986816,-60.9445838928223,-60.9081039428711,-60.8867645263672,-60.8952140808105],[9.58027362823486,9.50234413146973,9.48769569396973,9.4794921875,9.48427772521973,9.52753925323486,9.53681659698486,9.54218769073486,9.55107402801514,9.55576133728027,9.57187461853027,9.60117244720459,9.61054706573486,9.595703125,9.58027362823486],[79.8748092651367,79.9037094116211,79.8210983276367,79.7667922973633,79.7476577758789,79.8599548339844,79.8748092651367],[79.9695281982422,79.9068374633789,79.857421875,79.845703125,79.8585891723633,79.8722686767578,79.8884735107422,79.9119186401367,79.9695281982422],[79.9823226928711,80.0784149169922,80.1809539794922,80.2528305053711,80.3759765625,80.7111282348633,80.8934555053711,80.9100570678711,80.9354476928711,80.9792022705078,81.0160140991211,81.1982421875,81.21923828125,81.2162094116211,81.2269515991211,81.2746047973633,81.333984375,81.3728485107422,81.4221649169922,81.4221649169922,81.4359359741211,81.6654281616211,81.6787109375,81.67626953125,81.6829071044922,81.7273483276367,81.7966766357422,81.83203125,81.8741226196289,81.876953125,81.8614273071289,81.8185577392578,81.7677764892578,81.7126922607422,81.6373977661133,81.3799819946289,81.3062515258789,80.9710922241211,80.72412109375,80.4958038330078,80.2673873901367,80.0953140258789,80.0072250366211,79.9469757080078,79.859375,79.7920837402344,79.7599639892578,79.7078170776367,79.7129898071289,79.7498092651367,79.7497024536133,79.7834930419922,79.8088912963867,79.8319320678711,79.8508834838867,79.9417953491211,79.9436569213867,79.9279327392578,79.9289093017578,80.0648422241211,80.099609375,80.1183547973633,80.1109390258789,80.0863342285156,80.1960906982422,80.2564468383789,80.3179702758789,80.3679656982422,80.4283218383789,80.3853530883789,80.2576217651367,80.0457992553711,79.9794921875,79.9540023803711,79.9669952392578,79.9823226928711],[28.7369136810303,28.6468753814697,28.6343746185303,28.57666015625,28.4996089935303,28.4390621185303,28.39208984375,28.3154296875,28.1761703491211,28.1390628814697,28.1287117004395,28.0963859558105,28.0568351745605,28.0181636810303,27.90185546875,27.7531242370605,27.6666011810303,27.5896472930908,27.5490226745605,27.5065422058105,27.4919929504395,27.4314441680908,27.4085922241211,27.3884754180908,27.3640632629395,27.3497066497803,27.3553695678711,27.3126945495605,27.23974609375,27.1935539245605,27.1304683685303,27.091796875,27.0517578125,27.0569343566895,27.09521484375,27.2074222564697,27.2945327758789,27.3568363189697,27.4249019622803,27.4580078125,27.4910163879395,27.5271472930908,27.5902347564697,27.6604499816895,27.7355480194092,27.8303699493408,27.9598617553711,28.0843753814697,28.2326183319092,28.4718742370605,28.5833969116211,28.6257801055908,28.6526374816895,28.68115234375,28.7217788696289,28.8162097930908,28.8562507629395,28.9537105560303,29.0580081939697,29.1780281066895,29.259765625,29.3013687133789,29.3359394073486,29.3708992004395,29.3907241821289,29.38671875,29.3488273620605,29.2935543060303,29.2492198944092,29.1951179504395,29.1421871185303,29.1219730377197,29.0980472564697,29.0290031433105,28.9752922058105,28.9010753631592,28.7369136810303],[20.9578113555908,20.8998050689697,21.0140628814697,21.0576171875,21.087890625,21.1148433685303,21.11572265625,21.1039066314697,21.03173828125,20.9578113555908],[26.5935554504395,26.5908203125,26.5666007995605,26.5192375183105,26.4695301055908,26.4576168060303,26.4953136444092,26.6812496185303,26.7601547241211,26.7756843566895,26.734375,26.6749992370605,26.6484375,26.6011714935303,26.2917976379395,26.2507801055908,26.2312488555908,26.2158203125,26.1751956939697,26.0929679870605,25.9644527435303,25.8592777252197,25.7808589935303,25.7224617004395,25.7239246368408,25.7316398620605,25.7248039245605,25.6851577758789,25.6203136444092,25.5675792694092,25.54736328125,25.5575199127197,25.6168937683105,25.7025394439697,25.7481441497803,25.7650394439697,25.7652339935303,25.7492198944092,25.6805667877197,25.5730457305908,25.5103511810303,25.4973621368408,25.52734375,25.5056648254395,25.4611339569092,25.37060546875,25.28369140625,25.1794929504395,25.1114253997803,25.0460929870605,24.8695316314697,24.82568359375,24.7892589569092,24.7681655883789,24.6207027435303,24.478515625,24.3179683685303,24.2366199493408,24.1913089752197,24.1039066314697,24.0084953308105,23.9444332122803,23.87255859375,23.7336921691895,23.55908203125,23.4846687316895,23.4776363372803,23.4830074310303,23.4813461303711,23.45361328125,23.3701171875,23.2823238372803,23.1703128814697,23.0874996185303,23.0421886444092,23.0319347381592,23.0155277252197,22.9767589569092,22.8939456939697,22.8237285614014,22.7662105560303,22.7243156433105,22.6798820495605,22.6844711303711,22.7096691131592,22.8312492370605,22.82470703125,22.7365226745605,22.62744140625,22.5672855377197,22.3463859558105,22.1378917694092,22.0723628997803,21.8739261627197,21.6827144622803,21.5546875,21.4470710754395,21.3892574310303,21.2975578308105,21.2357425689697,21.236328125,21.2010746002197,21.2378902435303,21.1710929870605,21.0619144439697,21.0538082122803,21.0460948944092,21.3146476745605,21.6535148620605,21.7305660247803,22.0428714752197,22.0845699310303,22.3659191131592,22.5869140625,22.7732429504395,22.8755855560303,22.96826171875,23.04296875,23.1198253631592,23.1958980560303,23.6126956939697,23.7067375183105,23.8126945495605,24.0082035064697,24.1207046508789,24.3678703308105,24.4736328125,24.5290031433105,24.6995105743408,24.7732410430908,24.8410148620605,24.9030265808105,24.94384765625,25.0699234008789,25.2069339752197,25.5857410430908,25.6631851196289,25.8763675689697,26.0041999816895,26.0855484008789,26.2095699310303,26.28125,26.4010734558105,26.5428714752197,26.5935554504395],[6.11650419235229,6.10830116271973,6.10976600646973,6.13818359375,6.20488214492798,6.25605440139771,6.32460975646973,6.44091844558716,6.4873046875,6.49375009536743,6.48476552963257,6.44462871551514,6.40673828125,6.37832069396973,6.34843778610229,6.34433603286743,6.27734375,6.24218702316284,6.18105459213257,6.11992216110229,6.07412099838257,6.01142597198486,5.95947265625,5.92890644073486,5.9013671875,5.82343769073486,5.78974580764771,5.81543016433716,5.83759737014771,5.85654306411743,5.88037109375,5.8037109375,5.78798818588257,5.72499990463257,5.72578191757202,5.74082040786743,5.73525333404541,5.74404287338257,5.78808641433716,5.8173828125,5.86689472198486,5.97626972198486,6.05478572845459,6.08906221389771,6.11005830764771,6.11650419235229],[27.3519535064697,27.4697265625,27.5111331939697,27.5386714935303,27.6727542877197,27.796875,27.82861328125,27.8382816314697,27.8302726745605,27.8145503997803,27.7627925872803,27.7173824310303,27.7111320495605,27.6394538879395,27.6556625366211,27.8060550689697,27.8486328125,27.8815422058105,27.89208984375,27.94140625,27.9916000366211,28.0075206756592,28.1031246185303,28.11083984375,28.1692390441895,28.1916999816895,28.2020511627197,28.17333984375,28.14794921875,28.1178703308105,28.0320301055908,27.8962898254395,27.6942386627197,27.6422843933105,27.5894527435303,27.5767574310303,27.4591808319092,27.4271488189697,27.3091793060303,27.0525379180908,26.9530277252197,26.8224620819092,26.7718753814697,26.6202144622803,26.5935554504395,26.5428714752197,26.4010734558105,26.28125,26.2095699310303,26.0855484008789,26.0041999816895,25.8763675689697,25.6631851196289,25.5857410430908,25.2069339752197,25.0699234008789,24.94384765625,24.9030265808105,24.8410148620605,24.7732410430908,24.6995105743408,24.5290031433105,24.4736328125,24.3678703308105,24.1207046508789,24.0082035064697,23.8126945495605,23.7067375183105,23.6126956939697,23.1958980560303,23.1198253631592,23.04296875,22.96826171875,22.8755855560303,22.7732429504395,22.5869140625,22.3659191131592,22.0845699310303,22.0428714752197,21.7305660247803,21.6535148620605,21.3146476745605,21.0460948944092,21.0149421691895,21.0314445495605,21.0712890625,21.2574214935303,21.3507823944092,21.4050788879395,21.4214839935303,21.4591789245605,21.7287120819092,21.9423828125,22.2314453125,22.5545883178711,22.6169910430908,22.6486320495605,23.0377922058105,23.1368160247803,23.2873039245605,23.6477546691895,23.93115234375,24.0542964935303,24.28125,24.3826179504395,24.4032230377197,24.3629875183105,24.3015613555908,24.3225574493408,24.3624992370605,24.4588871002197,24.7757816314697,24.8390617370605,24.9113292694092,25.1110343933105,25.1751956939697,25.2287101745605,25.25830078125,25.2726554870605,25.2686519622803,25.2826156616211,25.3400402069092,25.5712890625,25.66015625,25.7208976745605,25.7937507629395,25.9911136627197,26.0152339935303,26.0303726196289,26.2150402069092,26.2980461120605,26.4621105194092,26.5326175689697,26.8197269439697,26.8998050689697,26.9660148620605,27.0333995819092,27.1871089935303,27.3265628814697,27.3519535064697],[113.478904724121,113.481048583984,113.494140625,113.527053833008,113.548141479492,113.545509338379,113.498832702637,113.484184265137,113.478904724121],[-63.0111808776855,-63.1230430603027,-63.114990234375,-63.0630874633789,-63.0248069763184,-63.0094223022461,-63.0111808776855],[-2.21962881088257,-2.19077157974243,-2.13178682327271,-1.92089831829071,-1.79560554027557,-1.79218745231628,-1.83242213726044,-1.84965860843658,-1.81660175323486,-1.73945295810699,-1.73330104351044,-1.75185573101044,-1.79179656505585,-1.70693349838257,-1.69267606735229,-1.71469712257385,-1.71411156654358,-1.70297837257385,-1.63124978542328,-1.67919945716858,-1.62509739398956,-1.55073261260986,-1.51000964641571,-1.44999969005585,-1.35214829444885,-1.29638683795929,-1.18823254108429,-1.11103510856628,-1.06552731990814,-1.16259789466858,-1.24033212661743,-1.26210916042328,-1.22592806816101,-1.22592806816101,-1.27534151077271,-1.47705078125,-1.63515627384186,-1.81699228286743,-2.07280254364014,-2.23125004768372,-2.44838833808899,-2.52324199676514,-2.72260713577271,-2.86342763900757,-2.88720726966858,-2.93085956573486,-2.96113300323486,-2.98823213577271,-3.01738238334656,-3.43979501724243,-3.60458970069885,-3.70024394989014,-3.76816415786743,-3.82675766944885,-3.8466796875,-3.84956073760986,-3.83710932731628,-3.79643559455872,-3.78916025161743,-3.81513714790344,-3.82138657569885,-3.83339810371399,-3.81181669235229,-3.77099609375,-3.73017597198486,-3.67250990867615,-3.62451171875,-3.62690424919128,-3.66679692268372,-3.70200181007385,-3.86005830764771,-3.98535180091858,-4.14877939224243,-4.32285165786743,-4.52915048599243,-4.61962890625,-4.77851581573486,-4.96826219558716,-5.06191444396973,-5.18012666702271,-5.29365253448486,-5.44877958297729,-5.59331083297729,-5.77499961853027,-6.00429677963257,-6.16650390625,-6.21479463577271,-6.35761690139771,-6.42763662338257,-6.47973585128784,-6.50087881088257,-6.50791025161743,-6.51069355010986,-6.52055644989014,-6.565673828125,-6.59775400161743,-6.63535165786743,-6.755126953125,-6.85556650161743,-7.09492206573486,-7.14243173599243,-7.16020488739014,-7.23491191864014,-7.34975576400757,-7.42768573760986,-7.48574209213257,-7.62460947036743,-7.68515634536743,-7.94384813308716,-7.99892568588257,-8.26518535614014,-8.34047794342041,-8.39931678771973,-8.558349609375,-8.659912109375,-8.67841815948486,-8.683349609375,-8.683349609375,-8.683349609375,-8.683349609375,-8.683349609375,-8.683349609375,-8.81782245635986,-8.81777381896973,-8.81391620635986,-8.78457069396973,-8.77436542510986,-8.78896427154541,-8.80268573760986,-8.79682636260986,-8.77436542510986,-8.75385761260986,-8.75385761260986,-8.79487228393555,-8.88906288146973,-9.00190448760986,-9.08442401885986,-9.20844745635986,-9.28559589385986,-9.35297870635986,-9.41303730010986,-9.4873046875,-9.56982421875,-9.67333984375,-9.7353515625,-9.81787109375,-9.90034198760986,-9.98090839385986,-10.03271484375,-10.0668458938599,-10.123046875,-10.189453125,-10.2514638900757,-10.3549308776855,-10.4789543151855,-10.55126953125,-10.6542482376099,-10.7577638626099,-10.830078125,-10.9228029251099,-11.0468263626099,-11.1503419876099,-11.2636232376099,-11.392578125,-11.3612794876099,-11.3168458938599,-11.3168458938599,-11.337890625,-11.39990234375,-11.470703125,-11.5116701126099,-11.5531740188599,-11.583984375,-11.63720703125,-11.6845216751099,-11.6992197036743,-11.7182130813599,-11.7548828125,-11.8808584213257,-11.9608888626099,-12.03076171875,-12.0567874908447,-12.0609865188599,-12.0810537338257,-12.0810537338257,-12.1010255813599,-12.1308603286743,-12.1708498001099,-12.2011232376099,-12.23095703125,-12.2709474563599,-12.3109865188599,-12.3608388900757,-12.40087890625,-12.43115234375,-12.5009765625,-12.56103515625,-12.6308097839355,-12.7109375,-12.8207521438599,-12.9111328125,-12.9478511810303,-12.9911623001099,-13.06103515625,-13.12109375,-13.1611328125,-13.23095703125,-13.28076171875,-13.3109865188599,-13.39111328125,-13.48095703125,-13.5810546875,-13.6610832214355,-13.7709474563599,-13.8407716751099,-13.89111328125,-13.9311027526855,-13.9809083938599,-14.02099609375,-14.0409669876099,-14.10107421875,-14.12109375,-14.1410636901855,-14.1708984375,-14.1908693313599,-14.1908693313599,-14.2109375,-14.22119140625,-14.27099609375,-14.31103515625,-14.3808097839355,-14.4409189224243,-14.4608888626099,-14.52099609375,-14.5810060501099,-14.630859375,-14.62109375,-14.6107902526855,-14.64111328125,-14.6708498001099,-14.7509765625,-14.8408212661743,-14.9711427688599,-15.15087890625,-15.2909660339355,-15.4609375,-15.6107902526855,-15.7509288787842,-15.9208498001099,-16.041015625,-16.1908702850342,-16.5810070037842,-16.73095703125,-16.9511222839355,-17.0029792785645,-17.0030765533447,-16.9308586120605,-16.7932605743408,-16.6839828491211,-16.514404296875,-16.3587398529053,-16.3042964935303,-16.2018566131592,-16.1697273254395,-16.2102527618408,-16.1136703491211,-15.9967279434204,-15.9426259994507,-15.8059577941895,-15.7892580032349,-15.8016595840454,-15.8551759719849,-15.9125499725342,-15.980712890625,-15.9528322219849,-15.8993158340454,-15.7777833938599,-15.5863275527954,-15.1886224746704,-15.0388679504395,-14.9042978286743,-14.8560543060303,-14.8429193496704,-14.794921875,-14.70703125,-14.6022939682007,-14.5227546691895,-14.4705562591553,-14.4138679504395,-14.3124513626099,-14.1683588027954,-13.9520998001099,-13.6958990097046,-13.5757808685303,-13.4957523345947,-13.4098148345947,-13.2561521530151,-13.1773929595947,-13.1759757995605,-13.0407228469849,-12.9489259719849,-12.7936525344849,-12.4688959121704,-11.986083984375,-11.5526857376099,-11.43017578125,-11.299072265625,-11.0809564590454,-10.673828125,-10.4864749908447,-10.2005863189697,-10.010498046875,-9.85263633728027,-9.74345684051514,-9.66709041595459,-9.62382793426514,-9.65292930603027,-9.77314472198486,-9.85390663146973,-9.87548923492432,-9.83242225646973,-9.83335018157959,-9.80869102478027,-9.67495059967041,-9.34746074676514,-9.28657150268555,-9.24912071228027,-9.245849609375,-8.83623123168945,-8.59628868103027,-8.51284217834473,-8.30117225646973,-7.56235408782959,-7.14467763900757,-6.90097618103027,-6.75576210021973,-6.35312509536743,-5.95756816864014,-5.9248046875,-5.74794912338257,-5.62285137176514,-5.52226543426514,-5.39736318588257,-5.27783250808716,-5.33764696121216,-5.33764696121216,-5.252685546875,-5.10537147521973,-4.83720684051514,-4.62832069396973,-4.32998085021973,-3.98242163658142,-3.78798842430115,-3.69326162338257,-3.59062480926514,-3.39472675323486,-3.20600605010986,-3.06308555603027,-2.97221684455872,-2.95795869827271,-2.95361351966858,-2.92597627639771,-2.86953139305115,-2.83994150161743,-2.73139619827271,-2.63681650161743,-2.42373061180115,-2.21962881088257],[7.43867206573486,7.37773466110229,7.38007831573486,7.39501953125,7.41445302963257,7.43691396713257,7.43867206573486],[28.2125015258789,28.1625003814697,28.1119136810303,28.07470703125,28.09033203125,28.130859375,28.1597652435303,28.15625,28.1349620819092,28.1155281066895,28.1135749816895,28.0997066497803,28.119140625,28.1996097564697,28.2443370819092,28.22265625,28.2394523620605,28.2046871185303,28.1499996185303,28.07177734375,27.9742183685303,27.8538093566895,27.8023433685303,27.7679691314697,27.6961917877197,27.6140613555908,27.5158195495605,27.46484375,27.44921875,27.3369140625,27.2779293060303,27.2481441497803,27.2308597564697,27.1520519256592,27.0803718566895,27.01220703125,26.9807624816895,26.9009761810303,26.7873058319092,26.7137699127197,26.6189460754395,26.6404285430908,26.8470687866211,26.9005851745605,27.0084953308105,27.228515625,27.3369140625,27.40380859375,27.4583988189697,27.5492172241211,27.5622081756592,27.57373046875,27.7144546508789,27.8200187683105,27.890625,27.96337890625,28.0384769439697,28.080078125,28.0884761810303,28.1587886810303,28.291015625,28.34716796875,28.3269519805908,28.3405284881592,28.3875007629395,28.4230461120605,28.4419918060303,28.4630851745605,28.5304679870605,28.6016597747803,28.7738285064697,28.8658199310303,28.9231433868408,28.9733409881592,29.0369129180908,29.0929679870605,29.1253910064697,29.19482421875,29.2111339569092,29.2107410430908,29.1860332489014,29.15087890625,29.1229496002197,29.1348609924316,29.1597652435303,29.2005844116211,29.3337879180908,29.3833980560303,29.4556655883789,29.5106449127197,29.5391616821289,29.5493183135986,29.5417976379395,29.5109386444092,29.5150394439697,29.5634765625,29.5686531066895,29.5719738006592,29.5977535247803,29.7197265625,29.8778324127197,29.9180660247803,29.9424800872803,29.9347648620605,29.92431640625,30.1310558319092,30.1075191497803,30.07568359375,29.8780269622803,29.837890625,29.751953125,29.7068347930908,29.6645488739014,29.6149425506592,29.5550785064697,29.4910144805908,29.4587898254395,29.4328117370605,29.3928718566895,29.3395519256592,29.3048820495605,29.2545890808105,29.2238292694092,29.2045917510986,29.2007789611816,29.1862316131592,29.1462879180908,29.0499019622803,28.9583969116211,28.9274425506592,28.9305667877197,28.9437503814697,29.0062484741211,28.9718761444092,28.9477558135986,28.8495140075684,28.73876953125,28.7292957305908,28.6675777435303,28.5623035430908,28.4916019439697,28.5094718933105,28.5137691497803,28.5017585754395,28.4990234375,28.4713878631592,28.3103523254395,28.2648448944092,28.2125015258789],[49.9364242553711,49.8240242004395,49.8556632995605,49.9859352111816,50.0230445861816,49.9364242553711],[48.3421859741211,48.3435516357422,48.2119140625,48.1912117004395,48.2556648254395,48.2697257995605,48.3088874816895,48.35107421875,48.3421859741211],[49.5382804870605,49.5841789245605,49.6377906799316,49.8049812316895,49.87646484375,49.9375,49.9671897888184,50.0734367370605,50.173828125,50.20458984375,50.2353515625,50.3134765625,50.4413108825684,50.4827117919922,50.4045906066895,50.29150390625,50.2623023986816,50.208984375,50.1849632263184,50.0944328308105,50.0204086303711,49.9265632629395,49.892578125,49.8533172607422,49.7437477111816,49.6643562316895,49.64990234375,49.6669921875,49.6970710754395,49.71044921875,49.7127952575684,49.7422828674316,49.7859382629395,49.8310546875,49.8390655517578,49.8113250732422,49.7339859008789,49.7385749816895,49.7671852111816,49.73974609375,49.6369132995605,49.59521484375,49.53955078125,49.4493179321289,49.4371109008789,49.49365234375,49.4778327941895,49.3628921508789,49.296875,49.2033195495605,49.06005859375,48.9180679321289,48.7974586486816,48.7083015441895,48.6070327758789,48.4685554504395,48.3507843017578,48.1758804321289,47.9344711303711,47.9083976745605,47.8583030700684,47.8041000366211,47.7394523620605,47.6041030883789,47.5886688232422,47.5580062866211,47.4276390075684,47.37255859375,47.3335952758789,47.3117179870605,47.2728538513184,47.1773414611816,47.0349617004395,46.9381828308105,46.728515625,46.6222686767578,46.38671875,46.15869140625,45.9208984375,45.6921844482422,45.6045875549316,45.5080070495605,45.2057609558105,45.115234375,44.8128929138184,44.69580078125,44.4738273620605,44.40673828125,44.3458976745605,44.2561531066895,44.078125,44.0353546142578,44.00830078125,43.9898452758789,43.9437484741211,43.9095726013184,43.8515625,43.6875,43.6700210571289,43.6568374633789,43.662109375,43.6460952758789,43.6647453308105,43.7222633361816,43.69873046875,43.6375961303711,43.6146469116211,43.5695304870605,43.3978500366211,43.3575172424316,43.32958984375,43.2648429870605,43.2571296691895,43.2666015625,43.29052734375,43.3322257995605,43.3426780700684,43.3697280883789,43.4105491638184,43.4377937316895,43.5018539428711,43.5831069946289,43.70361328125,43.8001976013184,43.8556632995605,43.9111328125,44.0630874633789,44.1171875,44.2396469116211,44.34814453125,44.3810577392578,44.4046897888184,44.4322280883789,44.4529304504395,44.4488296508789,44.3865242004395,44.23876953125,44.2339859008789,44.2457008361816,44.2331047058105,44.1787109375,44.1087875366211,44.0400390625,44.0066375732422,44.013671875,43.9935569763184,43.9435539245605,43.9793930053711,44.42138671875,44.4357414245605,44.41796875,44.4270515441895,44.4424819946289,44.4761734008789,44.5518569946289,44.9091796875,44.955078125,45.0442352294922,45.1667976379395,45.2228546142578,45.2712898254395,45.3023414611816,45.3421859741211,45.486328125,45.5417976379395,45.5982437133789,45.6247062683105,45.6405258178711,45.6615219116211,45.7001953125,45.8859367370605,46.0042953491211,46.1575202941895,46.1905288696289,46.3140602111816,46.3515625,46.3996086120605,46.4416007995605,46.34130859375,46.326171875,46.3314437866211,46.3851585388184,46.47509765625,46.6747093200684,46.8820343017578,46.9422836303711,46.9932632446289,47.0323257446289,47.0274391174316,47.060546875,47.0992202758789,47.1333961486816,47.1351547241211,47.1073226928711,47.09375,47.0925750732422,47.1976547241211,47.2804679870605,47.3187522888184,47.3519515991211,47.4390602111816,47.4647483825684,47.4850578308105,47.4963874816895,47.4740219116211,47.4420928955078,47.42919921875,47.4783210754395,47.5247039794922,47.5925788879395,47.6700210571289,47.7160148620605,47.7740249633789,47.8704071044922,47.9641609191895,47.8115234375,47.7733383178711,47.9551734924316,47.9569320678711,47.9832038879395,47.9955062866211,47.9013671875,47.8835945129395,47.89599609375,47.9410171508789,47.9818382263184,48.0398445129395,48.0859375,48.1871109008789,48.2552719116211,48.3376960754395,48.4050788879395,48.5064468383789,48.6213874816895,48.7964820861816,48.9103546142578,48.91943359375,48.8942375183105,48.8538093566895,48.7863273620605,48.8039093017578,48.8996086120605,48.9317398071289,49.0357398986816,49.20703125,49.2634773254395,49.3121070861816,49.3301773071289,49.3639678955078,49.4797859191895,49.5382804870605],[73.4166030883789,73.3953170776367,73.3820266723633,73.3849563598633,73.4015655517578,73.427734375,73.4427719116211,73.4349670410156,73.4166030883789],[73.51220703125,73.4948196411133,73.4786148071289,73.4730453491211,73.4811553955078,73.4947280883789,73.5040969848633,73.5177688598633,73.5283203125,73.5271530151367,73.5221710205078,73.51904296875,73.51220703125],[-91.6836929321289,-91.796142578125,-91.8161163330078,-91.589111328125,-91.55029296875,-91.5367202758789,-91.6542434692383,-91.6836929321289],[-110.914451599121,-110.974800109863,-111.063667297363,-111.039939880371,-110.989402770996,-110.942092895508,-110.914451599121],[-86.9396438598633,-86.9914093017578,-87.0194320678711,-86.9779815673828,-86.9278335571289,-86.8285598754883,-86.7632751464844,-86.7550277709961,-86.8087844848633,-86.9396438598633],[-86.7140121459961,-86.6962890625,-86.713623046875,-86.7363739013672,-86.7528839111328,-86.7390670776367,-86.7269058227539,-86.7140121459961],[-106.502243041992,-106.531349182129,-106.607032775879,-106.634178161621,-106.639350891113,-106.597358703613,-106.536422729492,-106.523826599121,-106.502243041992],[-109.805076599121,-109.826759338379,-109.8779296875,-109.900489807129,-109.890335083008,-109.793800354004,-109.795608520508,-109.805076599121],[-111.698875427246,-111.712303161621,-112.013275146484,-111.940872192383,-111.856834411621,-111.698875427246],[-110.5673828125,-110.538864135742,-110.590187072754,-110.657424926758,-110.703422546387,-110.699264526367,-110.690238952637,-110.59521484375,-110.5673828125],[-112.057281494141,-112.077346801758,-112.162895202637,-112.175491333008,-112.210502624512,-112.296775817871,-112.222312927246,-112.159423828125,-112.131683349609,-112.198387145996,-112.195014953613,-112.163764953613,-112.130226135254,-112.126274108887,-112.067481994629,-112.057281494141],[-111.100296020508,-111.087745666504,-111.094436645508,-111.13525390625,-111.204490661621,-111.224655151367,-111.182914733887,-111.139259338379,-111.090866088867,-111.100296020508],[-115.170608520508,-115.184280395508,-115.352935791016,-115.260398864746,-115.273979187012,-115.233543395996,-115.196975708008,-115.148536682129,-115.170608520508],[-118.242774963379,-118.285499572754,-118.400100708008,-118.4013671875,-118.367820739746,-118.312301635742,-118.312057495117,-118.265533447266,-118.247360229492,-118.242774963379],[-112.203079223633,-112.278419494629,-112.355270385742,-112.514060974121,-112.531005859375,-112.469825744629,-112.423530578613,-112.285057067871,-112.263427734375,-112.248725891113,-112.203079223633],[-113.155616760254,-113.162796020508,-113.264747619629,-113.496337890625,-113.580612182617,-113.594383239746,-113.587203979492,-113.507957458496,-113.415916442871,-113.375831604004,-113.373825073242,-113.2021484375,-113.177925109863,-113.155616760254],[-114.694137573242,-114.727249145508,-114.789207458496,-114.784568786621,-114.771095275879,-114.709075927734,-114.687934875488,-114.694137573242],[-97.146240234375,-97.1644515991211,-97.2248992919922,-97.424072265625,-97.507080078125,-97.6676788330078,-97.717041015625,-97.7286148071289,-97.74267578125,-97.7273941040039,-97.765869140625,-97.7452087402344,-97.7583465576172,-97.8166961669922,-97.8578109741211,-97.8415985107422,-97.8424835205078,-97.7823715209961,-97.7632827758789,-97.5847625732422,-97.4845275878906,-97.3601608276367,-97.3144989013672,-97.3368606567383,-97.3875503540039,-97.4091796875,-97.4341278076172,-97.4244079589844,-97.3848190307617,-97.3834457397461,-97.4565887451172,-97.5903778076172,-97.7538070678711,-97.6375503540039,-97.5976028442383,-97.5665512084961,-97.5145492553711,-97.5010757446289,-97.5005874633789,-97.3571243286133,-97.1949691772461,-97.1863250732422,-97.1214370727539,-96.7086868286133,-96.4560546875,-96.3683624267578,-96.3153305053711,-96.28955078125,-96.1239776611328,-96.0733871459961,-95.9846725463867,-95.9130325317383,-95.7781219482422,-95.8103485107422,-95.92822265625,-95.9203643798828,-95.8210906982422,-95.6268081665039,-95.5783233642578,-95.6549377441406,-95.7198181152344,-95.6971206665039,-95.5614242553711,-95.1818313598633,-95.0147018432617,-94.7981414794922,-94.681640625,-94.5461883544922,-94.4597702026367,-94.3922882080078,-94.1890106201172,-93.8731460571289,-93.764404296875,-93.5523452758789,-93.2279281616211,-93.1273422241211,-92.884765625,-92.7690887451172,-92.7289581298828,-92.7101058959961,-92.4853057861328,-92.4410171508789,-92.2131805419922,-92.1032257080078,-91.9737777709961,-91.88037109375,-91.8804702758789,-91.9426803588867,-91.91357421875,-91.802978515625,-91.5997085571289,-91.5339813232422,-91.4404754638672,-91.2752456665039,-91.2787551879883,-91.3082962036133,-91.3563003540039,-91.3677749633789,-91.334228515625,-91.3430633544922,-91.445556640625,-91.4691925048828,-91.4578552246094,-91.4366683959961,-91.1359405517578,-91.0589370727539,-90.9550247192383,-90.7392578125,-90.6931610107422,-90.6501007080078,-90.507080078125,-90.49169921875,-90.482421875,-90.4863815307617,-90.4783172607422,-90.484130859375,-90.4584426879883,-90.4351501464844,-90.3531265258789,-90.1829071044922,-89.8876495361328,-89.8197708129883,-88.8787078857422,-88.7466735839844,-88.5849075317383,-88.4666976928711,-88.2510223388672,-88.1847610473633,-88.1717300415039,-88.17138671875,-88.1316375732422,-88.0068359375,-87.7737274169922,-87.6888198852539,-87.48046875,-87.2508773803711,-87.2179260253906,-87.1875991821289,-87.164306640625,-87.1882781982422,-87.2105941772461,-87.2494659423828,-87.2957534790039,-87.3866729736328,-87.3685073852539,-87.2757339477539,-87.2164535522461,-87.1284713745117,-87.0347671508789,-86.9117202758789,-86.8240737915039,-86.8170852661133,-86.8038558959961,-86.7717742919922,-86.8155288696289,-86.8647003173828,-86.9262161254883,-87.0595703125,-87.2212448120117,-87.42138671875,-87.4671936035156,-87.4658203125,-87.4319305419922,-87.4417495727539,-87.4662170410156,-87.5068893432617,-87.5857925415039,-87.6876983642578,-87.6900863647461,-87.6453170776367,-87.5873031616211,-87.5116653442383,-87.4693832397461,-87.4247512817383,-87.4347229003906,-87.482666015625,-87.5128936767578,-87.5669937133789,-87.6275329589844,-87.65869140625,-87.65576171875,-87.6220703125,-87.55078125,-87.5094680786133,-87.5010757446289,-87.5935592651367,-87.6530227661133,-87.7335510253906,-87.7618179321289,-87.8041000366211,-87.8532180786133,-87.8819808959961,-87.9596710205078,-88.0390625,-88.0564422607422,-88.0111312866211,-88.03173828125,-88.0737762451172,-88.1267547607422,-88.19677734375,-88.1953125,-88.2757339477539,-88.295654296875,-88.3724060058594,-88.4612731933594,-88.5230026245117,-88.586181640625,-88.7436065673828,-88.8063430786133,-88.8573760986328,-88.8978042602539,-88.942626953125,-89.0504455566406,-89.133544921875,-89.162353515625,-89.1614761352539,-89.3715362548828,-89.7288131713867,-90.18359375,-90.6220245361328,-90.9891586303711,-90.9904251098633,-90.9916000366211,-90.9929656982422,-91.1955032348633,-91.4096145629883,-91.392333984375,-91.3191909790039,-91.2241668701172,-91.1118698120117,-90.975830078125,-90.8160171508789,-90.710693359375,-90.6599655151367,-90.6343765258789,-90.6340866088867,-90.5757827758789,-90.4710922241211,-90.4169921875,-90.4169921875,-90.4501495361328,-90.4598617553711,-90.4471664428711,-90.52197265625,-90.7032241821289,-90.9795913696289,-91.2337875366211,-91.4339904785156,-91.736572265625,-91.8194351196289,-91.9572296142578,-92.0821304321289,-92.1871566772461,-92.2042465209961,-92.204345703125,-92.0747985839844,-92.0987319946289,-92.1442413330078,-92.1585464477539,-92.1556625366211,-92.1863708496094,-92.1764602661133,-92.159912109375,-92.1870651245117,-92.2090377807617,-92.2351531982422,-92.2645492553711,-92.5309600830078,-92.8089370727539,-92.9184036254883,-93.0244140625,-93.1668930053711,-93.5411605834961,-93.734375,-93.9160614013672,-94.0789566040039,-94.2398910522461,-94.311279296875,-94.3741683959961,-94.4090347290039,-94.4264144897461,-94.3701705932617,-94.3028335571289,-94.24951171875,-94.1934127807617,-94.0283203125,-94.0012741088867,-94.4707489013672,-94.6615219116211,-94.6822738647461,-94.5871124267578,-94.6168518066406,-94.6507797241211,-94.7528305053711,-94.7908172607422,-94.7974624633789,-94.7929153442383,-94.8586883544922,-94.9004364013672,-94.9347152709961,-95.0235366821289,-95.0208435058594,-94.8460388183594,-94.7857894897461,-94.7994155883789,-94.9493179321289,-95.1343765258789,-95.4644088745117,-95.7717819213867,-96.2135772705078,-96.4086380004883,-96.5108413696289,-96.8079605102539,-97.1846618652344,-97.7547912597656,-98.1389617919922,-98.5203170776367,-98.76220703125,-98.907958984375,-99.0016555786133,-99.3480453491211,-99.690673828125,-100.024513244629,-100.243019104004,-100.431884765625,-100.847801208496,-101.001953125,-101.147857666016,-101.385498046875,-101.487060546875,-101.600296020508,-101.762397766113,-101.847068786621,-101.918701171875,-101.995506286621,-102.216598510742,-102.546974182129,-102.699554443359,-103.018508911133,-103.441604614258,-103.580276489258,-103.698921203613,-103.912452697754,-104.045654296875,-104.277000427246,-104.405181884766,-104.602981567383,-104.9384765625,-105.045211791992,-105.107666015625,-105.286376953125,-105.482078552246,-105.532417297363,-105.570411682129,-105.615921020508,-105.66943359375,-105.642143249512,-105.542579650879,-105.377052307129,-105.260154724121,-105.244674682617,-105.252296447754,-105.327049255371,-105.420120239258,-105.492378234863,-105.510841369629,-105.456344604492,-105.393989562988,-105.301956176758,-105.237060546875,-105.224998474121,-105.233253479004,-105.208686828613,-105.431449890137,-105.457420349121,-105.527442932129,-105.649116516113,-105.6455078125,-105.791793823242,-105.943359375,-106.021728515625,-106.234573364258,-106.402244567871,-106.566505432129,-106.728759765625,-106.935501098633,-107.084861755371,-107.764938354492,-107.726608276367,-107.527244567871,-107.493698120117,-107.488914489746,-107.511917114258,-107.548873901367,-107.601997375488,-107.673683166504,-107.709518432617,-107.816703796387,-107.951171875,-108.0087890625,-108.015090942383,-108.207664489746,-108.28076171875,-108.243309020996,-108.192039489746,-108.140090942383,-108.079635620117,-108.035690307617,-108.051460266113,-108.092826843262,-108.373680114746,-108.466262817383,-108.696388244629,-108.7509765625,-108.787254333496,-108.843605041504,-108.893165588379,-109.02880859375,-109.0634765625,-109.068458557129,-108.972755432129,-108.884857177734,-108.886573791504,-108.935157775879,-109.008354187012,-109.084083557129,-109.196479797363,-109.253959655762,-109.304298400879,-109.384963989258,-109.425636291504,-109.354148864746,-109.270660400391,-109.19970703125,-109.158790588379,-109.11669921875,-109.146339416504,-109.216018676758,-109.240623474121,-109.243263244629,-109.276268005371,-109.482864379883,-109.676078796387,-109.754791259766,-109.828369140625,-109.890922546387,-109.921730041504,-109.925628662109,-109.943992614746,-110.277145385742,-110.377296447754,-110.477783203125,-110.519386291504,-110.560646057129,-110.592674255371,-110.615478515625,-110.578269958496,-110.529884338379,-110.759033203125,-110.8486328125,-110.920806884766,-110.986083984375,-111.121383666992,-111.282424926758,-111.4716796875,-111.680076599121,-111.747215270996,-111.832420349121,-111.907028198242,-111.918601989746,-111.940818786621,-112.044876098633,-112.161766052246,-112.192039489746,-112.223487854004,-112.301414489746,-112.378219604492,-112.393211364746,-112.388671875,-112.41455078125,-112.572906494141,-112.653121948242,-112.697166442871,-112.738380432129,-112.759231567383,-112.824806213379,-112.951759338379,-113.057670593262,-113.110443115234,-113.087013244629,-113.10498046875,-113.118598937988,-113.107963562012,-113.072807312012,-113.042922973633,-113.046730041504,-113.083641052246,-113.186180114746,-113.2314453125,-113.480812072754,-113.623489379883,-113.633010864258,-113.699951171875,-113.759422302246,-113.94775390625,-113.977493286133,-114.002685546875,-114.080909729004,-114.149314880371,-114.264068603516,-114.548675537109,-114.608787536621,-114.697601318359,-114.741302490234,-114.933586120605,-114.895065307617,-114.839508056641,-114.789894104004,-114.84814453125,-114.881881713867,-114.844673156738,-114.761032104492,-114.703369140625,-114.685447692871,-114.633308410645,-114.649749755859,-114.629928588867,-114.550491333008,-114.403411865234,-114.372611999512,-114.17919921875,-114.061912536621,-113.828956604004,-113.755470275879,-113.545310974121,-113.538475036621,-113.499702453613,-113.3818359375,-113.328903198242,-113.335006713867,-113.320701599121,-113.258888244629,-113.20556640625,-113.093650817871,-113.033592224121,-112.956642150879,-112.870849609375,-112.865234375,-112.868461608887,-112.795707702637,-112.808052062988,-112.749313354492,-112.758201599121,-112.734031677246,-112.552635192871,-112.329200744629,-112.282623291016,-112.191459655762,-112.09814453125,-112.003952026367,-112.015571594238,-112.009086608887,-111.883155822754,-111.862648010254,-111.754005432129,-111.723388671875,-111.699409484863,-111.778518676758,-111.816841125488,-111.82177734375,-111.795265197754,-111.569679260254,-111.5458984375,-111.470161437988,-111.464500427246,-111.418502807617,-111.404594421387,-111.332130432129,-111.330368041992,-111.291603088379,-111.149559020996,-111.034423828125,-111.013626098633,-110.893943786621,-110.755661010742,-110.686767578125,-110.67724609375,-110.72900390625,-110.734527587891,-110.659324645996,-110.546974182129,-110.421485900879,-110.399658203125,-110.409622192383,-110.367431640625,-110.319969177246,-110.296821594238,-110.320899963379,-110.325096130371,-110.303756713867,-110.262893676758,-110.022804260254,-109.982521057129,-109.893165588379,-109.811325073242,-109.775978088379,-109.710548400879,-109.676559448242,-109.509620666504,-109.420852661133,-109.414993286133,-109.458053588867,-109.495704650879,-109.630470275879,-109.728416442871,-109.823051452637,-109.923439025879,-110.006248474121,-110.086029052734,-110.180618286133,-110.244087219238,-110.288772583008,-110.362693786621,-110.629981994629,-110.764892578125,-110.895561218262,-111.036178588867,-111.419342041016,-111.578224182129,-111.682907104492,-111.750389099121,-111.802490234375,-111.822265625,-111.8251953125,-111.848243713379,-112.072555541992,-112.119041442871,-112.128517150879,-112.077980041504,-112.055763244629,-112.069869995117,-112.093360900879,-112.114601135254,-112.119773864746,-112.173828125,-112.377243041992,-112.526077270508,-112.658393859863,-113.020751953125,-113.119239807129,-113.143211364746,-113.155815124512,-113.205764770508,-113.272270202637,-113.425880432129,-113.598541259766,-113.701271057129,-113.756645202637,-113.840965270996,-113.935935974121,-113.996482849121,-114.110061645508,-114.201858520508,-114.333396911621,-114.445266723633,-114.479690551758,-114.498291015625,-114.539894104004,-114.715621948242,-114.858741760254,-114.993499755859,-115.033195495605,-115.036476135254,-114.823539733887,-114.570014953613,-114.448486328125,-114.372703552246,-114.300582885742,-114.2890625,-114.30224609375,-114.232666015625,-114.13720703125,-114.0693359375,-114.135055541992,-114.175392150879,-114.157325744629,-114.158393859863,-114.252639770508,-114.265869140625,-114.185249328613,-114.092720031738,-114.048484802246,-114.1455078125,-114.309226989746,-114.664016723633,-114.875930786133,-114.937301635742,-114.993499755859,-115.166358947754,-115.311187744141,-115.565284729004,-115.673828125,-115.748672485352,-115.808303833008,-115.789543151855,-115.815628051758,-115.858207702637,-115.995803833008,-116.028564453125,-116.035354614258,-116.062156677246,-116.296295166016,-116.309623718262,-116.309669494629,-116.333450317383,-116.45849609375,-116.609565734863,-116.662155151367,-116.66845703125,-116.722076416016,-116.701705932617,-116.65210723877,-116.623878479004,-116.620796203613,-116.847991943359,-116.913673400879,-117.034774780273,-117.063133239746,-117.128280639648,-116.842094421387,-116.555961608887,-116.269821166992,-115.983688354492,-115.69750213623,-115.41136932373,-115.125198364258,-114.839057922363,-114.724761962891,-114.787994384766,-114.835945129395,-114.361717224121,-113.887451171875,-113.413177490234,-112.93896484375,-112.464744567871,-111.990478515625,-111.516212463379,-111.0419921875,-110.688522338867,-110.335105895996,-109.981636047363,-109.628227233887,-109.274757385254,-108.921340942383,-108.56787109375,-108.214447021484,-108.213813781738,-108.213188171387,-108.212493896484,-108.211814880371,-107.992042541504,-107.772216796875,-107.552345275879,-107.33251953125,-107.112693786621,-106.892868041992,-106.673049926758,-106.453224182129,-106.445411682129,-106.43603515625,-106.346977233887,-106.255714416504,-106.148048400879,-106.024070739746,-105.812698364258,-105.514015197754,-105.275833129883,-105.09814453125,-104.978805541992,-104.917869567871,-104.835891723633,-104.681350708008,-104.681350708008,-104.622215270996,-104.504005432129,-104.400634765625,-104.312210083008,-104.215522766113,-104.110595703125,-103.98974609375,-103.852928161621,-103.663963317871,-103.422950744629,-103.257713317871,-103.168312072754,-103.089988708496,-103.022850036621,-102.956840515137,-102.891990661621,-102.865676879883,-102.877830505371,-102.833984375,-102.734176635742,-102.614936828613,-102.476272583008,-102.385643005371,-102.343070983887,-102.268951416016,-102.1630859375,-101.990921020508,-101.752349853516,-101.611625671387,-101.568702697754,-101.54638671875,-101.544624328613,-101.50927734375,-101.440376281738,-101.38037109375,-101.303512573242,-101.038970947266,-101.038619995117,-101.016304016113,-100.924118041992,-100.754592895508,-100.658645629883,-100.636329650879,-100.549705505371,-100.39892578125,-100.331733703613,-100.34814453125,-100.336280822754,-100.296051025391,-100.221290588379,-100.111961364746,-100.001411437988,-99.8896484375,-99.7542495727539,-99.5953140258789,-99.5053253173828,-99.4842834472656,-99.48583984375,-99.5100555419922,-99.4998016357422,-99.4551315307617,-99.4402313232422,-99.4577178955078,-99.45654296875,-99.4564971923828,-99.4435501098633,-99.3024368286133,-99.2299270629883,-99.17236328125,-99.1720733642578,-99.1077651977539,-99.0152816772461,-98.8731994628906,-98.7652282714844,-98.69140625,-98.5982894897461,-98.4858932495117,-98.3781280517578,-98.2750473022461,-98.0828170776367,-97.8014144897461,-97.5872573852539,-97.4402847290039,-97.3756332397461,-97.358154296875,-97.3497543334961,-97.3386688232422,-97.2817916870117,-97.146240234375],[169.635055541992,169.615432739258,169.590530395508,169.612197875977,169.627151489258,169.651077270508,169.700393676758,169.734573364258,169.726364135742,169.672561645508,169.635055541992],[171.577346801758,171.61474609375,171.688369750977,171.7568359375,171.73046875,171.693359375,171.659378051758,171.614151000977,171.5927734375,171.577346801758],[171.101943969727,171.226959228516,171.393753051758,171.366989135742,171.3046875,171.263275146484,171.2353515625,171.202331542969,171.095520019531,171.035736083984,171.050399780273,171.101943969727],[168.830261230469,168.8154296875,168.71923828125,168.675094604492,168.679290771484,168.755462646484,168.830261230469],[166.890335083008,166.864456176758,166.8447265625,166.85888671875,166.888092041016,166.8994140625,166.890335083008],[22.3440437316895,22.4982414245605,22.5827140808105,22.6823234558105,22.7960948944092,22.8368167877197,22.9091796875,22.9439449310303,22.9919929504395,23.0036125183105,23.0056629180908,22.9514636993408,22.9296875,22.916015625,22.8592777252197,22.7838878631592,22.7550773620605,22.7248039245605,22.6036128997803,22.4935550689697,22.4007816314697,22.2376956939697,22.1844730377197,22.1388683319092,21.9933586120605,21.9294929504395,21.7794933319092,21.6275386810303,21.5757808685303,21.4596672058105,21.4041023254395,21.32373046875,21.1475582122803,21.1000003814697,20.9642581939697,20.9585933685303,20.9334964752197,20.8702144622803,20.7408199310303,20.7092761993408,20.6560535430908,20.6144542694092,20.56787109375,20.4889659881592,20.4870128631592,20.4923820495605,20.4486331939697,20.4755859375,20.5162105560303,20.5166015625,20.5051746368408,20.5531253814697,20.5662117004395,20.5785140991211,20.6949214935303,20.7250003814697,20.744140625,20.7503910064697,20.7781257629395,21.0597667694092,21.1424808502197,21.2060546875,21.25634765625,21.28662109375,21.2975578308105,21.3317375183105,21.3895492553711,21.5608386993408,21.5625,21.6182632446289,21.7392578125,21.8146495819092,21.85302734375,21.9041004180908,21.9775390625,22.0520515441895,22.1466808319092,22.23974609375,22.2770519256592,22.3173828125,22.3440437316895],[4.22763633728027,4.22822284698486,4.22900390625,4.22998046875,4.23085927963257,4.23193311691284,4.23271465301514,4.23369121551514,4.23466825485229,4.20292949676514,4.19121122360229,4.18212890625,4.12128877639771,4.01484346389771,3.97617197036743,3.94707036018372,3.90722680091858,3.89794921875,3.87695336341858,3.84296870231628,3.81650400161743,3.70957040786743,3.52050757408142,3.50429701805115,3.2890625,3.06015610694885,3.02939462661743,3.01054716110229,3.00107431411743,2.68964838981628,2.42080044746399,2.08818364143372,1.85937511920929,1.56914055347443,1.30019533634186,1.12128937244415,0.960058450698853,0.947460770606995,0.718652367591858,0.433007687330246,0.286230236291885,0.228710994124413,0.217480450868607,0.0073241195641458,0,-0.235888451337814,-0.405419766902924,-0.432275474071503,-0.454492479562759,-0.536523401737213,-0.666455328464508,-0.760449111461639,-0.907958805561066,-1.01918935775757,-1.04956066608429,-1.20498073101044,-1.49365222454071,-1.65732395648956,-1.69506812095642,-1.76777362823486,-1.87978529930115,-1.97304654121399,-2.05712866783142,-2.11323213577271,-2.45722627639771,-2.52690434455872,-2.58671879768372,-2.77885723114014,-2.87392592430115,-2.92587876319885,-2.91850614547729,-2.91708970069885,-2.95082998275757,-2.99721670150757,-3.03867197036743,-3.19843769073486,-3.24863290786743,-3.27016592025757,-3.26674795150757,-3.30175757408142,-3.396728515625,-3.46992182731628,-3.52763700485229,-3.57578134536743,-3.85346674919128,-3.94731402397156,-4.05117225646973,-4.15102529525757,-4.19619178771973,-4.25869178771973,-4.32871103286743,-4.31025362014771,-4.26064491271973,-4.22524404525757,-4.22709989547729,-4.48061513900757,-4.45986318588257,-4.42192363739014,-4.42158174514771,-4.4287109375,-4.47988319396973,-4.54604482650757,-4.5869140625,-4.62724590301514,-4.69931602478027,-4.79794931411743,-4.968994140625,-5.10590839385986,-5.15751981735229,-5.23017597198486,-5.28813457489014,-5.302001953125,-5.29052686691284,-5.27031278610229,-5.24477529525757,-5.22939443588257,-5.250244140625,-5.29985332489014,-5.347412109375,-5.42421913146973,-5.490478515625,-5.46855401992798,-5.45707988739014,-5.47568368911743,-5.47900390625,-5.50703144073486,-5.52353477478027,-5.55659151077271,-5.69428730010986,-5.84384775161743,-5.89619112014771,-5.90756845474243,-5.940673828125,-5.98867177963257,-6.03457021713257,-6.1171875,-6.19687461853027,-6.23837900161743,-6.24130821228027,-6.21499013900757,-6.192626953125,-6.190673828125,-6.2177734375,-6.23974561691284,-6.23066377639771,-6.250244140625,-6.26113271713257,-6.36562538146973,-6.40415048599243,-6.42587852478027,-6.4326171875,-6.40751934051514,-6.42392587661743,-6.48261690139771,-6.56459999084473,-6.65415000915527,-6.67636728286743,-6.68613290786743,-6.69199228286743,-6.66933631896973,-6.69326114654541,-6.75322294235229,-6.83364248275757,-6.90380859375,-6.95034217834473,-6.9794921875,-6.99174785614014,-6.96381855010986,-6.96816396713257,-6.98945283889771,-7.01708984375,-7.03974628448486,-7.10488271713257,-7.18232488632202,-7.36318397521973,-7.38505840301514,-7.41479444503784,-7.45654296875,-7.49794960021973,-7.53281259536743,-7.56210947036743,-7.6611328125,-7.74907255172729,-7.81420946121216,-7.88408231735229,-7.96093702316284,-7.99062538146973,-7.97446250915527,-7.98569345474243,-8.00727558135986,-8.23149490356445,-8.26665019989014,-8.30156230926514,-8.32412147521973,-8.32168006896973,-8.30634784698486,-8.312744140625,-8.33740234375,-8.40449237823486,-8.47470664978027,-8.56352519989014,-8.606201171875,-8.64619159698486,-8.66669940948486,-8.66391563415527,-8.56728458404541,-8.52031230926514,-8.46352481842041,-8.42529296875,-8.40068340301514,-8.39853572845459,-8.407470703125,-8.470703125,-8.56874942779541,-8.62114238739014,-8.66494178771973,-8.71142578125,-8.73310565948486,-8.77973651885986,-8.822021484375,-8.820068359375,-8.81831073760986,-8.91386699676514,-8.95083045959473,-8.99892520904541,-9.04306602478027,-9.12045955657959,-9.21552658081055,-9.30000019073486,-9.36518573760986,-9.39536190032959,-9.39365196228027,-9.3408203125,-9.33154296875,-9.34018516540527,-9.35810565948486,-9.40498065948486,-9.48681640625,-9.58774375915527,-9.65830039978027,-9.71474647521973,-9.75400352478027,-9.82070255279541,-10.0106449127197,-10.1670904159546,-10.2748537063599,-10.3398933410645,-10.3727540969849,-10.4658203125,-10.5895023345947,-10.6189937591553,-10.6437015533447,-10.6773433685303,-10.709228515625,-10.7349119186401,-10.7430171966553,-10.8064937591553,-10.8761720657349,-10.9332036972046,-11.0045413970947,-11.0658197402954,-11.1292476654053,-11.2096681594849,-11.2606935501099,-11.30517578125,-11.4146490097046,-11.492431640625,-11.502197265625,-11.4745607376099,-11.4475584030151,-11.4180669784546,-11.3894033432007,-11.3824224472046,-11.4487791061401,-11.4505863189697,-11.444091796875,-11.4143552780151,-11.4174308776855,-11.390380859375,-11.4339351654053,-11.4441404342651,-11.4928216934204,-11.5487794876099,-11.5616703033447,-11.5813484191895,-11.6349611282349,-11.6744632720947,-11.7582521438599,-11.772216796875,-11.8033685684204,-11.8316898345947,-11.8777828216553,-11.8952150344849,-11.8945808410645,-11.9570798873901,-12.05419921875,-12.0441408157349,-11.9841794967651,-11.9663572311401,-11.9608888626099,-11.9880857467651,-12.0201168060303,-12.0111818313599,-12.0191898345947,-12.068359375,-12.1128902435303,-12.1752443313599,-12.2284183502197,-12.2068367004395,-12.1865234375,-12.2806148529053,-12.1046876907349,-12.08154296875,-12.0215816497803,-11.9409170150757,-11.8728513717651,-11.8422355651855,-11.8287601470947,-11.7984371185303,-11.7601566314697,-11.6758785247803,-11.5967292785645,-11.502685546875,-11.4552240371704,-11.3656253814697,-11.1693353652954,-11.0074214935303,-10.9482412338257,-10.8956060409546,-10.8150882720947,-10.7319822311401,-10.6965827941895,-10.5865716934204,-10.4931640625,-10.4118165969849,-10.2621097564697,-10.1937503814697,-10.1295413970947,-9.94140625,-9.75507736206055,-9.57783126831055,-9.44692325592041,-9.44033241271973,-9.44770431518555,-9.42656230926514,-9.38535213470459,-9.3505859375,-9.33544921875,-9.29370212554932,-9.17680644989014,-8.987060546875,-8.78310585021973,-8.57915019989014,-8.37519550323486,-8.17124080657959,-7.96728515625,-7.76337909698486,-7.55937480926514,-7.35546922683716,-7.15151357650757,-6.94755840301514,-6.74360322952271,-6.53964853286743,-6.33574199676514,-6.13178682327271,-5.92783260345459,-5.723876953125,-5.51250028610229,-5.45561552047729,-5.403564453125,-5.35991191864014,-5.50961923599243,-5.62866258621216,-5.65625047683716,-5.68476581573486,-5.71318340301514,-5.74169969558716,-5.77016639709473,-5.79863262176514,-5.82709932327271,-5.85556650161743,-5.88408184051514,-5.91249990463257,-5.94101572036743,-5.969482421875,-5.99794912338257,-6.02641582489014,-6.05488300323486,-6.08339834213257,-6.11181640625,-6.14033174514771,-6.16879892349243,-6.197265625,-6.22573232650757,-6.25419950485229,-6.28271484375,-6.31113290786743,-6.33964824676514,-6.36811542510986,-6.39658212661743,-6.425048828125,-6.45351552963257,-6.48203134536743,-6.51044893264771,-6.53896522521973,-6.5673828125,-6.59409189224243,-6.28720664978027,-5.95981454849243,-5.64077091217041,-5.17290019989014,-4.82260751724243,-4.51699256896973,-4.24033212661743,-3.91279292106628,-3.58535146713257,-3.25786161422729,-2.93037080764771,-2.60292959213257,-2.275390625,-1.94790053367615,-1.62040996551514,-1.29296863079071,-0.965478599071503,-0.63798850774765,-0.310547143220901,0,0.0169922094792128,0.344433575868607,0.671874940395355,0.999414324760437,1.14550769329071,1.15917956829071,1.17275404930115,1.16406226158142,1.16572284698486,1.20888650417328,1.29023444652557,1.61064445972443,1.63603508472443,1.64736306667328,1.68544924259186,1.75322282314301,1.83242213726044,1.92880845069885,2.21933603286743,2.28085970878601,2.40615224838257,2.47421860694885,2.66777348518372,2.80791044235229,2.86572289466858,2.99248075485229,3.13027358055115,3.20371079444885,3.20341777801514,3.20273447036743,3.20166015625,3.22705078125,3.25585961341858,3.25439453125,3.21962904930115,3.1923828125,3.17724633216858,3.13789057731628,3.10605454444885,3.11972641944885,3.17421889305115,3.25595688819885,3.32343769073486,3.3564453125,3.40087866783142,3.43876957893372,3.68349575996399,3.91015601158142,4.22763633728027],[14.5662117004395,14.53271484375,14.4364261627197,14.3523435592651,14.3512687683105,14.4483394622803,14.537013053894,14.5662117004395],[14.3134765625,14.2536134719849,14.1942377090454,14.1803712844849,14.2632808685303,14.3037109375,14.3208990097046,14.3134765625],[98.1826171875,98.1343765258789,98.1180648803711,98.1402359008789,98.220703125,98.2917022705078,98.2833938598633,98.2312469482422,98.1826171875],[98.2097702026367,98.29345703125,98.2843704223633,98.271484375,98.2517623901367,98.2181625366211,98.1553726196289,98.0804748535156,98.142578125,98.1672821044922,98.2097702026367],[98.5417022705078,98.5189437866211,98.498046875,98.4774398803711,98.5265655517578,98.5417022705078],[98.2216796875,98.2162094116211,98.2093811035156,98.1873016357422,98.2010726928711,98.2390594482422,98.2781219482422,98.2996139526367,98.3075180053711,98.2837829589844,98.2632827758789,98.2216796875],[98.5538101196289,98.5284194946289,98.46484375,98.4347686767578,98.3968734741211,98.3995132446289,98.37646484375,98.5235290527344,98.5538101196289],[98.0754852294922,98.0835952758789,98.0210952758789,98.0103530883789,98.0595703125,98.0807571411133,98.0754852294922],[98.5160217285156,98.4743118286133,98.4544906616211,98.4662094116211,98.5252914428711,98.6095733642578,98.5764694213867,98.5160217285156],[98.13671875,98.1250991821289,98.1084976196289,98.0753860473633,98.0373077392578,98.0573272705078,98.0713882446289,98.1048812866211,98.1224670410156,98.12841796875,98.1184539794922,98.1201171875,98.13671875],[98.0661163330078,98.0603485107422,98.0023422241211,97.9517593383789,97.9386672973633,97.990234375,98.0451202392578,98.0598678588867,98.0661163330078],[98.4139633178711,98.4364242553711,98.46826171875,98.45947265625,98.380859375,98.33447265625,98.3138656616211,98.3314437866211,98.3025436401367,98.3121109008789,98.396484375,98.4139633178711],[98.3154296875,98.3091812133789,98.25927734375,98.2507858276367,98.2545852661133,98.2653274536133,98.2685546875,98.2986373901367,98.3154296875],[94.8048858642578,94.7843704223633,94.7433547973633,94.7334976196289,94.8280258178711,94.8381805419922,94.8048858642578],[94.4767532348633,94.4119186401367,94.3878936767578,94.4937515258789,94.5459976196289,94.6012649536133,94.61865234375,94.5661163330078,94.4767532348633],[97.5750045776367,97.5372085571289,97.4803695678711,97.4691390991211,97.5164108276367,97.5419921875,97.5790023803711,97.59326171875,97.599609375,97.58935546875,97.5750045776367],[93.6908187866211,93.6740264892578,93.5699234008789,93.4874954223633,93.6182632446289,93.7447280883789,93.7455062866211,93.7183609008789,93.6908187866211],[93.71484375,93.8294906616211,93.8747100830078,93.9457015991211,93.9474639892578,93.9339828491211,93.9019546508789,93.8152389526367,93.755859375,93.7323226928711,93.6622085571289,93.64404296875,93.6883773803711,93.71484375],[93.4917984008789,93.5132827758789,93.4446334838867,93.4195327758789,93.4128875732422,93.4917984008789],[93.0101547241211,93.0232391357422,92.9751892089844,92.9126968383789,92.9146499633789,92.9595718383789,93.0101547241211],[101.138862609863,101.080375671387,100.927543640137,100.819534301758,100.756645202637,100.703125,100.6591796875,100.613670349121,100.566596984863,100.5361328125,100.522262573242,100.54931640625,100.61767578125,100.622947692871,100.565139770508,100.493362426758,100.407424926758,100.326072692871,100.249313354492,100.183891296387,100.129684448242,100.122459411621,100.003616333008,99.9542999267578,99.8903350830078,99.8251953125,99.7733383178711,99.7201232910156,99.638671875,99.5316390991211,99.4588851928711,99.4479522705078,99.4874954223633,99.5016632080078,99.4859390258789,99.4515609741211,99.3992156982422,99.337890625,99.28369140625,99.1968765258789,99.1307601928711,99.07421875,99.0397415161133,99.0206985473633,98.9874038696289,98.9580078125,98.9166946411133,98.8757781982422,98.8195343017578,98.7606430053711,98.4938507080078,98.4549789428711,98.3712844848633,98.2936553955078,98.2390594482422,98.1110382080078,98.0490264892578,98.0150375366211,97.9912109375,97.9164047241211,97.8167953491211,97.7935562133789,97.8039016723633,97.7141647338867,97.7060546875,97.7540054321289,97.7459030151367,97.7277297973633,97.6715850830078,97.5773468017578,97.51513671875,97.4849624633789,97.3970718383789,97.3739242553711,97.3806610107422,97.4507827758789,97.5238342285156,97.5993194580078,97.6322250366211,97.6224594116211,97.6515655517578,97.7197265625,97.7399368286133,97.6985321044922,97.7064437866211,97.7290954589844,97.79296875,97.9292984008789,98.0630874633789,98.1746139526367,98.2565383911133,98.4388656616211,98.47119140625,98.4781265258789,98.5231399536133,98.5647430419922,98.5936508178711,98.6607437133789,98.6892623901367,98.83544921875,98.8693389892578,98.8882751464844,98.8884735107422,98.8655242919922,98.8179702758789,98.5923767089844,98.5740203857422,98.5582046508789,98.5544967651367,98.5652389526367,98.5569305419922,98.5373001098633,98.4521484375,98.3293991088867,98.2861328125,98.2322235107422,98.1910171508789,98.1779327392578,98.2021484375,98.2459945678711,98.3321304321289,98.4001922607422,98.4950256347656,98.5700149536133,98.72119140625,98.93359375,99.0146484375,99.0862274169922,99.1368179321289,99.1560592651367,99.1716842651367,99.1761703491211,99.1371078491211,99.107421875,99.1239242553711,99.1735305786133,99.1735305786133,99.2198257446289,99.29736328125,99.3719711303711,99.4050750732422,99.3942413330078,99.4163055419922,99.4324188232422,99.462890625,99.52294921875,99.61474609375,99.6124954223633,99.5728530883789,99.5152359008789,99.4779281616211,99.4426803588867,99.3587875366211,99.1901397705078,99.025390625,98.8871078491211,98.7869110107422,98.7572250366211,98.7572250366211,98.775390625,98.7683639526367,98.7468719482422,98.7184600830078,98.7025375366211,98.6580047607422,98.5625991821289,98.5212936401367,98.4968719482422,98.5230484008789,98.4649429321289,98.5009765625,98.53564453125,98.5988311767578,98.6755828857422,98.6826171875,98.7447280883789,98.7300796508789,98.7332992553711,98.7463912963867,98.7414093017578,98.7907257080078,98.8759765625,98.8402328491211,98.8047866821289,98.6936492919922,98.6363296508789,98.6249008178711,98.6390686035156,98.6449279785156,98.689453125,98.6863327026367,98.6638641357422,98.6962890625,98.6305694580078,98.6002883911133,98.619140625,98.6787109375,98.6244125366211,98.6646499633789,98.6631851196289,98.6356430053711,98.6371078491211,98.5951156616211,98.5759735107422,98.4871063232422,98.4212875366211,98.2454071044922,98.2484359741211,98.2389678955078,98.2003860473633,98.1495132446289,98.1106491088867,98.0982437133789,98.0726547241211,98.1001968383789,97.9984436035156,97.9765625,97.9097671508789,97.9292984008789,98.0187454223633,97.9365234375,97.869140625,97.8123016357422,97.7998046875,97.7437515258789,97.7742156982422,97.7103500366211,97.5842819213867,97.6092758178711,97.640625,97.6336898803711,97.6646499633789,97.7259750366211,97.66845703125,97.61962890625,97.5050735473633,97.3758773803711,97.3310546875,97.2674789428711,97.2117156982422,97.1783218383789,97.2001953125,97.1001968383789,97.0745162963867,96.9701156616211,96.8514709472656,96.8777389526367,96.9097671508789,96.8508834838867,96.9085922241211,96.8580093383789,96.8106460571289,96.7654266357422,96.6224594116211,96.5066375732422,96.43115234375,96.3643493652344,96.2822265625,96.2621078491211,96.2489318847656,96.2203140258789,96.1890640258789,96.2376937866211,96.2367172241211,96.3243103027344,96.2930679321289,96.1350555419922,96.0809555053711,96.0428695678711,96.0321350097656,96.0123062133789,95.7632827758789,95.71142578125,95.6794891357422,95.5556640625,95.3895492553711,95.3484344482422,95.3014678955078,95.3078079223633,95.36474609375,95.3467788696289,95.3330078125,95.2258758544922,95.1769485473633,95.0783233642578,94.9425811767578,94.8912124633789,94.8921890258789,94.8822250366211,94.8978500366211,94.8931655883789,94.8601531982422,94.8477554321289,94.7981414794922,94.6615219116211,94.65625,94.6513671875,94.6807556152344,94.6765670776367,94.7199249267578,94.7166061401367,94.7033157348633,94.6790008544922,94.6652297973633,94.6376953125,94.5875015258789,94.4957046508789,94.4415969848633,94.2990264892578,94.2238235473633,94.2142562866211,94.2712860107422,94.3273468017578,94.3534164428711,94.3999938964844,94.4524383544922,94.47314453125,94.4943389892578,94.564453125,94.5889663696289,94.56005859375,94.4943389892578,94.4307632446289,94.2658157348633,94.2521514892578,94.1707000732422,94.2457046508789,94.09130859375,94.0700149536133,94.0389633178711,94.044921875,94.0224609375,94.0015640258789,93.9410171508789,93.9680709838867,93.9613265991211,93.92919921875,93.8000946044922,93.7054672241211,93.59814453125,93.4930648803711,93.5305709838867,93.57861328125,93.72802734375,93.8249053955078,93.8861312866211,93.9620132446289,93.9981460571289,93.9607391357422,93.8878860473633,93.8395538330078,93.7699279785156,93.7610321044922,93.7395553588867,93.6687469482422,93.6117172241211,93.6598663330078,93.70703125,93.5818328857422,93.4390640258789,93.4095687866211,93.3623046875,93.25,93.1566390991211,93.1990203857422,93.1906204223633,93.1294937133789,93.001953125,93.0403289794922,93.0955047607422,93.068359375,93.01513671875,93.0667953491211,93.0353546142578,93.0187454223633,92.99072265625,92.8821334838867,92.8283233642578,92.7912139892578,92.8435516357422,92.8716812133789,92.89111328125,92.8506851196289,92.7869110107422,92.7356414794922,92.708984375,92.7326202392578,92.7228469848633,92.6080093383789,92.3783187866211,92.3241195678711,92.3119125366211,92.2862319946289,92.2684555053711,92.2644577026367,92.2147445678711,92.1919937133789,92.1795883178711,92.2082061767578,92.2796859741211,92.33056640625,92.3726501464844,92.4718780517578,92.5391616821289,92.5685577392578,92.5998077392578,92.6316375732422,92.6252975463867,92.5934600830078,92.5842819213867,92.5828094482422,92.5749053955078,92.63037109375,92.6526336669922,92.6747055053711,92.68896484375,92.7210006713867,92.7713851928711,92.8542938232422,92.9094696044922,92.9645538330078,93.02197265625,93.04296875,93.0706024169922,93.1214904785156,93.1511688232422,93.1623992919922,93.1620101928711,93.1050720214844,93.0881805419922,93.0787124633789,93.1142578125,93.1625061035156,93.1509704589844,93.1641616821289,93.2039108276367,93.2535171508789,93.3080062866211,93.3494110107422,93.3660125732422,93.3913040161133,93.4081039428711,93.4149398803711,93.37255859375,93.3073272705078,93.3262710571289,93.3555679321289,93.4521484375,93.4937515258789,93.5640640258789,93.63330078125,93.6834030151367,93.755859375,93.85546875,94.0108413696289,94.0748062133789,94.1276397705078,94.1703109741211,94.2197265625,94.2930603027344,94.3772430419922,94.3994140625,94.4931640625,94.5840835571289,94.6632766723633,94.7076110839844,94.7037048339844,94.67529296875,94.6156234741211,94.5665054321289,94.5530242919922,94.5543899536133,94.5798797607422,94.6228485107422,94.6677780151367,94.7858428955078,94.8611297607422,94.9457015991211,94.9919891357422,95.0152359008789,95.0407257080078,95.0929641723633,95.1324234008789,95.1292953491211,95.1083984375,95.0689468383789,95.0508804321289,95.0597686767578,95.0894470214844,95.1287078857422,95.2014617919922,95.3050765991211,95.4638671875,95.7383804321289,95.8373107910156,95.9052734375,95.9709014892578,96.0614242553711,96.1908187866211,96.2742233276367,96.6657180786133,96.7316360473633,96.7978515625,96.8802719116211,96.9534149169922,97.0380859375,97.10205078125,97.1037139892578,96.9019470214844,96.8835983276367,96.8768539428711,96.8997116088867,96.9627914428711,97.0497055053711,97.1578140258789,97.22607421875,97.30615234375,97.3351516723633,97.3435592651367,97.3391647338867,97.302734375,97.3102493286133,97.3224639892578,97.3564453125,97.4314498901367,97.4777374267578,97.5021438598633,97.5378875732422,97.5992126464844,97.6588821411133,97.6946258544922,97.7300796508789,97.76904296875,97.8165054321289,97.8649368286133,97.8876037597656,97.93408203125,98.0222625732422,98.0616226196289,98.0989303588867,98.1183547973633,98.1304702758789,98.2410125732422,98.2742156982422,98.298828125,98.3504867553711,98.3923873901367,98.4088897705078,98.4525375366211,98.5044860839844,98.5998077392578,98.6511764526367,98.6767578125,98.6824264526367,98.6748046875,98.7165069580078,98.7294921875,98.7384796142578,98.7393493652344,98.7318344116211,98.70947265625,98.671875,98.685546875,98.6631851196289,98.5719680786133,98.5640640258789,98.5910110473633,98.6546859741211,98.65625,98.6253890991211,98.5584030151367,98.4655303955078,98.4016647338867,98.3337860107422,98.2965774536133,98.1725540161133,98.1428756713867,98.099609375,98.0640640258789,98.0107421875,97.9620056152344,97.91796875,97.8195343017578,97.7673797607422,97.7149429321289,97.7107391357422,97.7378921508789,97.7238235473633,97.6707000732422,97.5833053588867,97.5293884277344,97.5314483642578,97.5632858276367,97.6236343383789,97.6666030883789,97.6707000732422,97.7082061767578,97.6906280517578,97.5682601928711,97.5645523071289,97.6296844482422,97.68603515625,97.7556610107422,97.8376922607422,98.0168991088867,98.2125015258789,98.3672866821289,98.4994125366211,98.5641555786133,98.5833969116211,98.7643585205078,98.8023452758789,98.8350601196289,98.833984375,98.7015609741211,98.6767578125,98.6808624267578,98.7350540161133,98.7876968383789,98.8322296142578,98.7978515625,98.8197250366211,98.85888671875,98.8826141357422,98.8855438232422,98.86376953125,99.0550765991211,99.2203140258789,99.3408203125,99.4180679321289,99.4645462036133,99.4972610473633,99.5071258544922,99.466796875,99.3851547241211,99.3382797241211,99.3431625366211,99.3376998901367,99.2430648803711,99.2053680419922,99.17236328125,99.1734390258789,99.1929626464844,99.2333984375,99.3031234741211,99.388671875,99.5926742553711,99.8253936767578,99.9176788330078,99.9478530883789,99.9404296875,99.9255828857422,99.9407196044922,99.9782257080078,100.041213989258,100.095504760742,100.105758666992,100.089256286621,100.116798400879,100.147659301758,100.214752197266,100.3505859375,100.445701599121,100.531349182129,100.604591369629,100.677146911621,100.835151672363,101.019340515137,101.079788208008,101.120704650879,101.130859375,101.128120422363,101.147270202637,101.138862609863],[19.1943359375,19.1916027069092,19.1964836120605,19.21875,19.2982425689697,19.4146480560303,19.5515632629395,19.6144542694092,19.6709957122803,19.7811508178711,19.8580074310303,19.9440422058105,20.1678695678711,20.2684574127197,20.3399410247803,20.34765625,20.3443374633789,20.2151374816895,20.1925773620605,20.1299819946289,20.0542984008789,20.0294914245605,20.0657234191895,20.0892581939697,20.0703125,20.06396484375,20.0457038879395,19.9390621185303,19.8597660064697,19.7882804870605,19.7544918060303,19.7377910614014,19.74072265625,19.7278308868408,19.7034187316895,19.6544933319092,19.5974597930908,19.5445308685303,19.4651374816895,19.3996086120605,19.3290042877197,19.2806644439697,19.3308601379395,19.3614253997803,19.3521499633789,19.3611316680908,19.3455085754395,19.3423843383789,19.1864242553711,19.1222648620605,18.8942375183105,18.6329097747803,18.6190433502197,18.6333980560303,18.6458988189697,18.5916004180908,18.5535163879395,18.5174789428711,18.4766597747803,18.4380855560303,18.4363288879395,18.4539051055908,18.4800777435303,18.5349617004395,18.5458984375,18.5432624816895,18.4660148620605,18.455078125,18.44384765625,18.4601573944092,18.4884777069092,18.6236324310303,18.6218757629395,18.6299819946289,18.6568355560303,18.6742191314697,18.7492198944092,18.85107421875,18.8956050872803,18.9346675872803,18.9787101745605,19.0266590118408,19.0367183685303,18.9738292694092,18.9402332305908,18.9506816864014,18.9742202758789,19.0283203125,19.0800762176514,19.11279296875,19.1643543243408,19.1943359375],[116.683303833008,116.589744567871,116.402145385742,116.243354797363,116.159660339355,116.098243713379,116.034378051758,116.025489807129,115.953811645508,115.820503234863,115.791702270508,115.796585083008,115.785545349121,115.639450073242,115.525093078613,115.5576171875,115.616401672363,115.711723327637,115.811714172363,115.898239135742,115.993850708008,116.074806213379,116.231147766113,116.317192077637,116.378227233887,116.513481140137,116.651954650879,116.760543823242,116.901176452637,116.95166015625,117.069732666016,117.197074890137,117.285934448242,117.350776672363,117.383987426758,117.455078125,117.555366516113,117.676658630371,117.768363952637,117.840431213379,117.979194641113,118.041900634766,118.147071838379,118.239646911621,118.498443603516,118.567764282227,118.690536499023,118.759956359863,118.880279541016,118.953125,119.017578125,119.081932067871,119.097267150879,119.122955322266,119.162406921387,119.235252380371,119.290824890137,119.30859375,119.325973510742,119.376663208008,119.526954650879,119.600189208984,119.711128234863,119.757232666016,119.759864807129,119.788475036621,119.862686157227,119.897850036621,119.884178161621,119.895896911621,119.867179870605,119.747467041016,119.706649780273,119.620216369629,119.474021911621,119.331832885742,119.162101745605,119.028518676758,118.957130432129,118.843940734863,118.790336608887,118.722946166992,118.648735046387,118.580474853516,118.404388427734,118.308700561523,118.156837463379,118.071296691895,117.910446166992,117.813484191895,117.741203308105,117.671096801758,117.620498657227,117.546875,117.438079833984,117.405570983887,117.392181396484,117.356353759766,117.356941223145,117.333404541016,117.269050598145,117.155952453613,116.978805541992,116.859085083008,116.787017822266,116.688865661621,116.619331359863,116.562599182129,116.516700744629,116.44482421875,116.357612609863,116.264549255371,116.212989807129,116.229095458984,116.240623474121,116.197647094727,116.109870910645,116.039543151855,115.934181213379,115.789161682129,115.681053161621,115.539451599121,115.439460754395,115.217475891113,115.162605285645,114.919242858887,114.738761901855,114.644325256348,114.560150146484,114.517181396484,114.502250671387,114.4873046875,114.419136047363,114.281051635742,114.167381286621,114.080268859863,114.0302734375,113.930862426758,113.877052307129,113.752151489258,113.652633666992,113.587005615234,113.507911682129,113.455665588379,113.300971984863,113.196098327637,113.049407958984,112.706741333008,112.596771240234,112.499313354492,112.411331176758,112.292083740234,112.112892150879,112.032615661621,111.898048400879,111.751075744629,111.681449890137,111.621284484863,111.547462463379,111.514739990234,111.489448547363,111.410934448242,111.402244567871,111.429588317871,111.486228942871,111.519729614258,111.602630615234,111.683792114258,111.8369140625,111.880271911621,111.931739807129,111.94287109375,111.933204650879,111.878120422363,111.771095275879,111.7197265625,111.640823364258,111.54736328125,111.503517150879,111.451072692871,111.186820983887,111.086524963379,111.007225036621,110.913284301758,110.839553833008,110.74853515625,110.708595275879,110.627532958984,110.520896911621,110.461715698242,110.429588317871,110.400390625,110.288864135742,110.196868896484,110.058006286621,109.858787536621,109.698051452637,109.595504760742,109.443168640137,109.33984375,109.131645202637,108.87451171875,108.687301635742,108.546485900879,108.333984375,108.171195983887,108.062301635742,107.805953979492,107.748725891113,107.292381286621,107.090721130371,106.906051635742,106.77001953125,106.693161010742,106.5791015625,106.518745422363,106.317184448242,105.867576599121,105.56640625,105.51708984375,105.314353942871,105.197067260742,105.115432739258,105.050582885742,104.982032775879,104.8603515625,104.773635864258,104.498237609863,104.498237609863,104.30517578125,103.997268676758,103.711135864258,103.44970703125,103.247848510742,103.072853088379,102.806831359863,102.5751953125,102.156639099121,101.972946166992,101.8798828125,101.7138671875,101.659965515137,101.5791015625,101.495315551758,101.313766479492,101.091987609863,100.772560119629,100.51904296875,100.086326599121,99.9837951660156,99.7574157714844,99.4678726196289,98.9468765258789,98.71630859375,98.2482376098633,97.7189407348633,97.2056579589844,96.8330078125,96.6252899169922,96.3854522705078,96.3523483276367,96.3424835205078,96.2995147705078,96.16845703125,96.0802688598633,95.9124984741211,95.8595657348633,95.8419952392578,95.6873016357422,95.5912094116211,95.5671844482422,95.5255813598633,95.4712905883789,95.3564453125,95.3255844116211,95.3255844116211,95.3436508178711,95.3667984008789,95.3502960205078,95.0498046875,94.8660125732422,94.7120132446289,94.4943389892578,94.36474609375,94.1993179321289,93.9579162597656,93.8681640625,93.7552719116211,93.6564483642578,93.5162124633789,93.2943420410156,92.916015625,92.7878875732422,92.5789031982422,92.423828125,92.1726608276367,92.02978515625,91.8528366088867,91.73779296875,91.5843734741211,91.5100631713867,91.4410171508789,91.3121109008789,91.2217788696289,91.1376953125,91.0500030517578,90.95361328125,90.9139633178711,90.8772430419922,90.8532257080078,90.76318359375,90.7496109008789,90.6944351196289,90.6618118286133,90.6707000732422,90.7096633911133,90.7958984375,90.8524475097656,90.8871078491211,90.9597702026367,91.0017623901367,90.9967803955078,90.9475631713867,90.9115219116211,90.9182586669922,90.9714889526367,91.0338897705078,91.0289077758789,91.0043029785156,90.9978485107422,90.9857406616211,90.9105453491211,90.8699264526367,90.7990264892578,90.7155227661133,90.6433563232422,90.5529251098633,90.4961853027344,90.4764633178711,90.4674835205078,90.4251937866211,90.3806610107422,90.3474578857422,90.3306655883789,90.3132858276367,90.1910171508789,90.1032180786133,90.0666046142578,90.0539093017578,90.0279312133789,89.9586944580078,89.9104537963867,89.8313446044922,89.7781219482422,89.7255859375,89.6931610107422,89.6384735107422,89.5609359741211,89.4792022705078,89.3298873901367,89.1962890625,89.1156234741211,89.0476531982422,88.9710922241211,88.9177703857422,88.8382797241211,88.6818389892578,88.5759735107422,88.5667953491211,88.51708984375,88.4139633178711,88.3099594116211,88.158203125,88.0625991821289,87.9796905517578,87.9673767089844,87.9722595214844,88.0106430053711,88.0501937866211,88.06005859375,88.0279312133789,87.9421920776367,87.8318328857422,87.8091812133789,87.7431640625,87.7546844482422,87.8068389892578,87.85986328125,87.8721694946289,87.8346710205078,87.8163070678711,87.8251953125,87.8142623901367,87.8182601928711,87.9347686767578,87.9880905151367,88.0285186767578,88.11572265625,88.13427734375,88.1355438232422,88.1925735473633,88.3377914428711,88.3933563232422,88.4524383544922,88.5443420410156,88.6332015991211,88.6827087402344,88.7478485107422,88.8316421508789,88.8603515625,88.8638687133789,88.9001922607422,88.9454040527344,88.9706039428711,89.0083923339844,89.1094741821289,89.1799850463867,89.2029342651367,89.2439498901367,89.2992172241211,89.3956069946289,89.4749984741211,89.5792007446289,89.6540985107422,89.6695327758789,89.63427734375,89.6438522338867,89.7442398071289,89.8780288696289,89.9773483276367,90.0049819946289,90.0537109375,90.1037139892578,90.2245101928711,90.3113327026367,90.3648452758789,90.5168991088867,90.6550827026367,90.71435546875,90.7607421875,90.8380889892578,90.9171905517578,91.0215835571289,91.0627899169922,91.2337875366211,91.3005828857422,91.3408203125,91.4150390625,91.4464874267578,91.5216827392578,91.5968704223633,91.6341781616211,91.7063446044922,91.8042984008789,91.95654296875,92.10400390625,92.1923828125,92.2653274536133,92.2790069580078,92.2957992553711,92.3547821044922,92.4263687133789,92.4864273071289,92.5789031982422,92.6266632080078,92.6813507080078,92.7386703491211,92.779296875,92.8564453125,92.9413070678711,92.9635772705078,92.970703125,93.0098648071289,93.1031265258789,93.2225646972656,93.2705078125,93.3868103027344,93.5010757446289,93.6255798339844,93.6620101928711,93.79541015625,93.9898452758789,94.0757827758789,94.2510681152344,94.2870101928711,94.3193359375,94.3468780517578,94.3546829223633,94.4001922607422,94.45849609375,94.4968795776367,94.5646438598633,94.61474609375,94.6754837036133,94.7180633544922,94.8112258911133,94.9302749633789,95.0128936767578,95.0443344116211,95.1114273071289,95.1662139892578,95.3294906616211,95.3856506347656,95.4417953491211,95.5226516723633,95.5671844482422,95.7078170776367,95.7893524169922,95.8519515991211,95.8994140625,95.9357452392578,95.9895553588867,96.0185546875,96.0655212402344,96.1117172241211,96.2296905517578,96.3150405883789,96.3811569213867,96.4664077758789,96.5057678222656,96.5432586669922,96.5984344482422,96.6402359008789,96.7117233276367,96.9857406616211,97.0491180419922,97.09765625,97.1369171142578,97.2085876464844,97.3597640991211,97.4183578491211,97.5408172607422,97.58935546875,97.6509780883789,97.720703125,97.7855453491211,97.8539047241211,97.9366226196289,98.00390625,98.1034240722656,98.1219711303711,98.1701202392578,98.1999969482422,98.2502899169922,98.27734375,98.2926788330078,98.2794876098633,98.2205123901367,98.14501953125,98.0789108276367,98.02978515625,98.0011749267578,97.9619140625,97.9641571044922,97.953125,97.9198226928711,97.8561553955078,97.8252944946289,97.8357391357422,97.9108352661133,97.9178695678711,97.9273452758789,97.9232406616211,97.9468765258789,97.9891586303711,98.03759765625,98.1031265258789,98.1846694946289,98.2199249267578,98.2374954223633,98.27685546875,98.3031234741211,98.3527374267578,98.6405258178711,98.7601547241211,98.8025360107422,98.8486328125,98.8931579589844,98.9580993652344,99.0342788696289,99.0914077758789,99.1761703491211,99.4070281982422,99.5323257446289,99.6128921508789,99.71923828125,99.7878875732422,99.9216766357422,100.034568786621,100.230369567871,100.468948364258,100.536231994629,100.710746765137,100.903610229492,101.085350036621,101.223236083984,101.304489135742,101.381248474121,101.46435546875,101.570899963379,101.821189880371,101.979194641113,102.111526489258,102.155662536621,102.160064697266,102.142387390137,102.151954650879,102.194534301758,102.210258483887,102.226173400879,102.215034484863,102.235054016113,102.276565551758,102.316604614258,102.303314208984,102.285743713379,102.288383483887,102.33642578125,102.406837463379,102.469436645508,102.546287536621,102.683303833008,102.765426635742,102.859672546387,103.039451599121,103.161712646484,103.233787536621,103.304389953613,103.421195983887,103.496292114258,103.632911682129,103.723236083984,103.802635192871,103.856155395508,103.95849609375,104.078712463379,104.1796875,104.259963989258,104.353904724121,104.46630859375,104.596382141113,104.685356140137,104.976951599121,105.0947265625,105.185935974121,105.266700744629,105.383598327637,105.541603088379,105.692581176758,105.875190734863,105.996482849121,106.08251953125,106.217872619629,106.368453979492,106.574409484863,106.711135864258,106.853713989258,106.941307067871,107.040229797363,107.14306640625,107.233299255371,107.347068786621,107.630950927734,107.786819458008,107.916603088379,107.947853088379,107.934860229492,107.938774108887,107.936721801758,107.965431213379,108.009574890137,108.033790588379,108.098045349121,108.213081359863,108.406929016113,108.5224609375,108.613677978516,108.733001708984,108.919921875,109.236717224121,109.453712463379,109.528709411621,109.750389099121,109.994537353516,110.199905395508,110.321388244629,110.427833557129,110.529594421387,110.631050109863,110.709762573242,110.827926635742,111.204200744629,111.336616516113,111.429298400879,111.511917114258,111.574798583984,111.735549926758,111.833396911621,111.934471130371,112.079689025879,112.375198364258,112.494926452637,112.697364807129,112.806449890137,112.914840698242,113.055564880371,113.092094421387,113.164161682129,113.319038391113,113.445510864258,113.57421875,113.732421875,113.881149291992,114.070701599121,114.221778869629,114.297065734863,114.386322021484,114.554008483887,114.674903869629,114.7431640625,114.879585266113,115.003326416016,115.098045349121,115.274513244629,115.365036010742,115.42919921875,115.587989807129,115.7177734375,115.795219421387,115.925971984863,116.134567260742,116.216804504395,116.351165771484,116.551170349121,116.631546020508,116.683303833008],[145.264846801758,145.21533203125,145.179595947266,145.157424926758,145.152145385742,145.232421875,145.265426635742,145.264846801758],[145.662307739258,145.621002197266,145.591598510742,145.586730957031,145.624816894531,145.647369384766,145.662307739258],[145.751953125,145.749221801758,145.6982421875,145.684280395508,145.713180541992,145.786331176758,145.821868896484,145.78857421875,145.782333374023,145.751953125],[145.712112426758,145.690231323242,145.658309936523,145.636032104492,145.631057739258,145.695495605469,145.719528198242,145.712112426758],[145.777526855469,145.729110717773,145.789260864258,145.807418823242,145.83544921875,145.777526855469],[145.708404541016,145.678131103516,145.652542114258,145.6455078125,145.690139770508,145.706634521484,145.708404541016],[32.8861312866211,32.7765617370605,32.5887680053711,32.4777336120605,32.353515625,32.1996078491211,32.1128883361816,32.10595703125,32.0779304504395,32.04833984375,32.0414085388184,32.0599632263184,32.06884765625,32.060546875,31.9684581756592,31.9482440948486,31.9283199310303,31.9203128814697,31.9845695495605,31.9793930053711,31.9870128631592,31.9857425689697,31.984375,31.9832038879395,31.98583984375,31.9666004180908,31.9505863189697,31.9080066680908,31.8583011627197,31.7996082305908,31.7240219116211,31.7000007629395,31.6755867004395,31.6041011810303,31.5456047058105,31.5296859741211,31.53173828125,31.4666976928711,31.4193344116211,31.3480472564697,31.3001956939697,31.2931632995605,31.2878913879395,31.4294910430908,31.5714836120605,31.7376956939697,31.8859348297119,32.0163116455078,32.1947250366211,32.37109375,32.4124031066895,32.4297866821289,32.3536109924316,32.4761734008789,32.4828109741211,32.4776344299316,32.4923820495605,32.529296875,32.6725578308105,32.7808609008789,32.86962890625,32.9927749633789,33.0048828125,33.0067405700684,32.97265625,32.8904304504395,32.8307609558105,32.7776374816895,32.8309555053711,32.8499984741211,32.8498039245605,32.826171875,32.7662086486816,32.7165031433105,32.69970703125,32.69921875,32.7219696044922,32.8544921875,32.8845710754395,32.9002952575684,32.9016609191895,32.9424819946289,32.9930648803711,32.9963874816895,32.978515625,32.9646492004395,32.95556640625,32.9546852111816,32.9807586669922,32.9693374633789,32.8843765258789,32.8762702941895,32.9378890991211,32.9480476379395,32.9029312133789,32.8102531433105,32.7417945861816,32.6358375549316,32.4519538879395,32.2432594299316,31.9398422241211,31.6875972747803,31.4898433685303,31.4261722564697,31.2362308502197,30.9387702941895,30.6301746368408,30.4377918243408,30.4093742370605,30.3981456756592,30.3960933685303,30.3798828125,30.3505840301514,30.3056659698486,30.2521476745605,30.2250003814697,30.2217769622803,30.2318344116211,30.4460945129395,30.5376930236816,30.6733417510986,30.9151363372803,31.1308574676514,31.3285179138184,31.5378894805908,31.6230487823486,31.7289066314697,31.9821281433105,32.0544891357422,32.1999015808105,32.2728538513184,32.55322265625,32.87451171875,32.9871101379395,33.2017593383789,33.2435569763184,33.3899383544922,33.5052757263184,33.6364250183105,33.6583023071289,33.6960945129395,33.7614288330078,33.9698219299316,34.0494155883789,34.1018524169922,34.2087860107422,34.33251953125,34.375,34.5052719116211,34.5241203308105,34.5511703491211,34.5576171875,34.5554656982422,34.5408210754395,34.4349632263184,34.4147453308105,34.3580055236816,34.2830085754395,34.24609375,34.2482414245605,34.2882804870605,34.3759765625,34.4030265808105,34.3951187133789,34.3955078125,34.4164085388184,34.4413070678711,34.5281257629395,34.6126937866211,34.7587890625,34.9333992004395,35.0153312683105,35.0798835754395,35.1121101379395,35.09423828125,35.0439453125,35.0646476745605,35.0930633544922,35.1246109008789,35.2013664245605,35.2725601196289,35.2904281616211,35.2811508178711,35.2297859191895,35.1783180236816,35.1671867370605,35.1852569580078,35.2427711486816,35.29150390625,35.3224601745605,35.3584976196289,35.5993156433105,35.7088890075684,35.7552757263184,35.7912139892578,35.8199234008789,35.8302726745605,35.8053703308105,35.8399429321289,35.8927726745605,35.86669921875,35.84716796875,35.6904296875,35.4884796142578,35.3757781982422,35.2474594116211,35.0138702392578,34.9068374633789,34.8504867553711,34.66162109375,34.6115226745605,34.5636749267578,34.5457038879395,34.5425796508789,34.5212898254395,34.48291015625,34.4658203125,34.412109375,34.36083984375,34.3578109741211,34.3759765625,34.462890625,34.5248069763184,34.5539054870605,34.6062507629395,34.6185531616211,34.6595687866211,34.8265609741211,34.95947265625,35.1826171875,35.4182586669922,35.4513664245605,35.50439453125,35.5643539428711,35.6309547424316,35.7046890258789,35.7854461669922,35.9113273620605,36.0822257995605,36.1754875183105,36.1913070678711,36.3056640625,36.5186500549316,36.673828125,36.7710952758789,36.8726577758789,36.9789085388184,37.0591773986816,37.1138687133789,37.2183609008789,37.3728523254395,37.5416984558105,37.7248039245605,37.8292961120605,37.8550758361816,37.8853530883789,37.9202117919922,38.0172843933105,38.1765632629395,38.3151397705078,38.4917984008789,38.6033210754395,38.7947273254395,38.9874992370605,39.1709938049316,39.3215827941895,39.4391632080078,39.5634765625,39.6944351196289,39.8170890808105,39.9886703491211,40.1662101745605,40.3474617004395,40.4635734558105,40.5167007446289,40.6117172241211,40.5550765991211,40.4866218566895,40.59716796875,40.51611328125,40.5062484741211,40.52685546875,40.5445327758789,40.4914054870605,40.4209938049316,40.40283203125,40.4651374816895,40.43310546875,40.4935569763184,40.5104484558105,40.5315437316895,40.50146484375,40.5091781616211,40.5231437683105,40.4871101379395,40.54833984375,40.5808601379395,40.5720710754395,40.5533180236816,40.4476547241211,40.4351577758789,40.4368171691895,40.5687484741211,40.5732421875,40.564453125,40.5695304870605,40.5519523620605,40.5829124450684,40.5451164245605,40.5582046508789,40.5598640441895,40.5905265808105,40.595703125,40.6025390625,40.6495132446289,40.7156257629395,40.7130889892578,40.6399421691895,40.6355476379395,40.6460952758789,40.7266578674316,40.7750015258789,40.8181648254395,40.8121070861816,40.8269538879395,40.8206062316895,40.8445320129395,40.8351554870605,40.7759780883789,40.70068359375,40.6874046325684,40.6943359375,40.6421852111816,40.6177711486816,40.6531257629395,40.6509780883789,40.5589866638184,40.3138656616211,40.2080078125,40.1087913513184,40.10888671875,40.0992164611816,39.9835929870605,39.8597640991211,39.7909164428711,39.8446273803711,39.7645530700684,39.6253890991211,39.2422866821289,39.1817398071289,39.0843734741211,38.9560546875,38.884765625,38.7576179504395,38.7132797241211,38.669921875,38.63330078125,38.3807640075684,38.1449241638184,38.0869140625,38.0482406616211,37.8394546508789,37.5123023986816,37.2445297241211,37.0505867004395,36.99951171875,36.9393577575684,36.9192390441895,36.8996086120605,36.7561492919922,36.5401344299316,36.498046875,36.4122047424316,36.4037094116211,36.3272438049316,36.2628898620605,36.2356452941895,36.1832046508789,36.125,35.9800758361816,35.8537101745605,35.6512680053711,35.3653335571289,34.9478530883789,34.8908195495605,34.8523445129395,34.7209968566895,34.6494140625,34.7134780883789,34.7557640075684,34.7450180053711,34.75,34.6981430053711,34.705078125,34.7647476196289,34.8770484924316,34.9823226928711,35.1175765991211,35.1280288696289,35.2676734924316,35.27294921875,35.3292961120605,35.3255844116211,35.3157234191895,35.3830108642578,35.4078140258789,35.40087890625,35.4188461303711,35.4563484191895,35.4937515258789,35.5048828125,35.5300788879395,35.5402336120605,35.5419921875,35.490234375,35.5057601928711,35.5753936767578,35.4944343566895,35.376953125,35.3704109191895,35.3988265991211,35.4621086120605,35.4853515625,35.5224609375,35.5419921875,35.4896507263184,35.4380836486816,35.2548828125,35.1559562683105,34.9920921325684,34.6073265075684,33.8360366821289,33.5300788879395,33.3474578857422,32.9611320495605,32.7921905517578,32.7225570678711,32.6558570861816,32.5904273986816,32.6474609375,32.7035179138184,32.7696304321289,32.8039054870605,32.8488273620605,32.89404296875,32.9164047241211,32.9548835754395,32.93359375,32.88916015625,32.8861312866211],[34.6416015625,34.6242218017578,34.5914077758789,34.5540046691895,34.5416030883789,34.5804672241211,34.6217765808105,34.6416015625],[34.7193374633789,34.7459945678711,34.7562522888184,34.7559585571289,34.7389640808105,34.7149429321289,34.6798820495605,34.66748046875,34.662109375,34.6841773986816,34.7193374633789],[-16.3733406066895,-16.4375476837158,-16.4659671783447,-16.4770011901855,-16.420166015625,-16.3932609558105,-16.3436527252197,-16.3733406066895],[-12.2806148529053,-12.3025388717651,-12.40869140625,-12.4598636627197,-12.5435543060303,-12.6596193313599,-12.7352542877197,-12.7703123092651,-12.8131837844849,-12.8584966659546,-12.8626461029053,-12.8519039154053,-12.8626947402954,-12.9308595657349,-12.9943361282349,-13.0485353469849,-13.0792961120605,-13.097900390625,-13.1052732467651,-13.1423826217651,-13.2064447402954,-13.2580080032349,-13.2970218658447,-13.3475580215454,-13.40966796875,-13.4541015625,-13.4869632720947,-13.4981441497803,-13.5069828033447,-13.5555171966553,-13.6235342025757,-13.6846685409546,-13.7149410247803,-13.7566404342651,-13.809814453125,-13.8684577941895,-13.9326171875,-13.9681634902954,-13.9750490188599,-14.0856447219849,-14.3000984191895,-14.5337409973145,-14.7867193222046,-14.9286127090454,-14.9595212936401,-14.9906244277954,-15.0219240188599,-15.0552244186401,-15.0905752182007,-15.1126470565796,-15.1214361190796,-15.2105464935303,-15.3799810409546,-15.5166988372803,-15.6208009719849,-15.7682123184204,-15.958984375,-16.0740242004395,-16.11328125,-16.1683597564697,-16.239013671875,-16.3022956848145,-16.3581047058105,-16.4043445587158,-16.4410171508789,-16.4800777435303,-16.5020503997803,-16.5352554321289,-16.5357437133789,-16.4813003540039,-16.463623046875,-16.3466796875,-16.2074699401855,-16.0789051055908,-16.0303230285645,-16.0467281341553,-16.0849609375,-16.1500988006592,-16.2130870819092,-16.305908203125,-16.4761714935303,-16.5144538879395,-16.4748058319092,-16.3712882995605,-16.3052730560303,-16.4448719024658,-16.2833976745605,-16.2332038879395,-16.2411613464355,-16.2104473114014,-16.333740234375,-16.4297866821289,-16.4791984558105,-16.5304203033447,-16.534912109375,-16.5626945495605,-16.6225109100342,-16.7283687591553,-16.8760738372803,-16.9279308319092,-16.9711437225342,-16.9982414245605,-17.0480461120605,-17.06396484375,-17.0423831939697,-17.0059070587158,-16.9645500183105,-16.8363265991211,-16.6070308685303,-16.3777351379395,-16.1484375,-15.9191408157349,-15.6897945404053,-15.4605464935303,-15.2312002182007,-15.0019044876099,-14.7726078033447,-14.5432624816895,-14.31396484375,-14.0846681594849,-13.8553705215454,-13.6260261535645,-13.396728515625,-13.1674318313599,-13.0162115097046,-13.0250968933105,-13.0322265625,-13.041748046875,-13.0512199401855,-13.0606441497803,-13.069580078125,-13.0784664154053,-13.0867671966553,-13.0943355560303,-13.1073246002197,-13.1559572219849,-13.16650390625,-13.1532707214355,-13.1208982467651,-13.031494140625,-12.89599609375,-12.7395992279053,-12.6204099655151,-12.5593748092651,-12.3729009628296,-12.2261714935303,-12.0833501815796,-12.0234375,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-11.8666505813599,-11.6803703308105,-11.4940433502197,-11.3077144622803,-11.1213865280151,-10.9351072311401,-10.748779296875,-10.5624513626099,-10.3761224746704,-10.1897945404053,-10.0035152435303,-9.81718826293945,-9.630859375,-9.44453144073486,-9.25820350646973,-9.07192420959473,-8.88564395904541,-8.68222618103027,-8.68212890625,-8.68212890625,-8.68232345581055,-8.6826171875,-8.682861328125,-8.68310546875,-8.683349609375,-8.49531269073486,-8.30727577209473,-8.11923789978027,-7.93115234375,-7.74311590194702,-7.55507802963257,-7.36699247360229,-7.17890596389771,-6.99086904525757,-6.80283212661743,-6.61474609375,-6.42670917510986,-6.23867225646973,-6.05058622360229,-5.862548828125,-5.67451143264771,-5.51694345474243,-5.27500009536743,-5.04951190948486,-4.82260751724243,-5.17290019989014,-5.64077091217041,-5.95981454849243,-6.28720664978027,-6.59409189224243,-6.5673828125,-6.53896522521973,-6.51044893264771,-6.48203134536743,-6.45351552963257,-6.425048828125,-6.39658212661743,-6.36811542510986,-6.33964824676514,-6.31113290786743,-6.28271484375,-6.25419950485229,-6.22573232650757,-6.197265625,-6.16879892349243,-6.14033174514771,-6.11181640625,-6.08339834213257,-6.05488300323486,-6.02641582489014,-5.99794912338257,-5.969482421875,-5.94101572036743,-5.91249990463257,-5.88408184051514,-5.85556650161743,-5.82709932327271,-5.79863262176514,-5.77016639709473,-5.74169969558716,-5.71318340301514,-5.68476581573486,-5.65625047683716,-5.62866258621216,-5.50961923599243,-5.35991191864014,-5.403564453125,-5.45561552047729,-5.51250028610229,-5.723876953125,-5.92783260345459,-6.13178682327271,-6.33574199676514,-6.53964853286743,-6.74360322952271,-6.94755840301514,-7.15151357650757,-7.35546922683716,-7.55937480926514,-7.76337909698486,-7.96728515625,-8.17124080657959,-8.37519550323486,-8.57915019989014,-8.78310585021973,-8.987060546875,-9.17680644989014,-9.29370212554932,-9.33544921875,-9.3505859375,-9.38535213470459,-9.42656230926514,-9.44770431518555,-9.44033241271973,-9.44692325592041,-9.57783126831055,-9.75507736206055,-9.94140625,-10.1295413970947,-10.1937503814697,-10.2621097564697,-10.4118165969849,-10.4931640625,-10.5865716934204,-10.6965827941895,-10.7319822311401,-10.8150882720947,-10.8956060409546,-10.9482412338257,-11.0074214935303,-11.1693353652954,-11.3656253814697,-11.4552240371704,-11.502685546875,-11.5967292785645,-11.6758785247803,-11.7601566314697,-11.7984371185303,-11.8287601470947,-11.8422355651855,-11.8728513717651,-11.9409170150757,-12.0215816497803,-12.08154296875,-12.1046876907349,-12.2806148529053],[-62.1484375,-62.1542510986328,-62.2216300964355,-62.2230453491211,-62.1913604736328,-62.1757774353027,-62.1484375],[57.6512718200684,57.5248031616211,57.38330078125,57.3283195495605,57.3176765441895,57.3651313781738,57.3621063232422,57.3857383728027,57.4160194396973,57.4864234924316,57.5150413513184,57.5757827758789,57.6565437316895,57.7372093200684,57.7919921875,57.7806625366211,57.7250022888184,57.7066421508789,57.6512718200684],[34.7193374633789,34.6841773986816,34.662109375,34.66748046875,34.6798820495605,34.7149429321289,34.7389640808105,34.7559585571289,34.7562522888184,34.7459945678711,34.7193374633789],[34.6416015625,34.6217765808105,34.5804672241211,34.5416030883789,34.5540046691895,34.5914077758789,34.6242218017578,34.6416015625],[34.95947265625,34.8265609741211,34.6595687866211,34.6185531616211,34.6062507629395,34.5539054870605,34.5248069763184,34.462890625,34.3759765625,34.3578109741211,34.36083984375,34.412109375,34.4658203125,34.48291015625,34.5212898254395,34.5425796508789,34.5457038879395,34.5636749267578,34.6115226745605,34.66162109375,34.8504867553711,34.9068374633789,35.0138702392578,35.2474594116211,35.3757781982422,35.4884796142578,35.6904296875,35.84716796875,35.86669921875,35.8927726745605,35.8399429321289,35.8053703308105,35.8302726745605,35.8199234008789,35.7912139892578,35.7552757263184,35.7088890075684,35.5993156433105,35.3584976196289,35.3224601745605,35.29150390625,35.2427711486816,35.1852569580078,35.1671867370605,35.1783180236816,35.2297859191895,35.2811508178711,35.2904281616211,35.2725601196289,35.2013664245605,35.1246109008789,35.0930633544922,35.0646476745605,35.0439453125,35.09423828125,35.1121101379395,35.0798835754395,35.0153312683105,34.9333992004395,34.7587890625,34.6126937866211,34.5281257629395,34.4413070678711,34.4164085388184,34.3955078125,34.3951187133789,34.4030265808105,34.3759765625,34.2882804870605,34.2482414245605,34.24609375,34.2830085754395,34.3580055236816,34.4147453308105,34.4349632263184,34.5408210754395,34.5554656982422,34.5576171875,34.5511703491211,34.5241203308105,34.5052719116211,34.375,34.33251953125,34.2087860107422,34.1018524169922,34.0494155883789,33.9698219299316,33.7614288330078,33.6960945129395,33.6583023071289,33.6364250183105,33.5052757263184,33.3899383544922,33.2435569763184,33.2017593383789,33.1480445861816,33.1036148071289,33.0423812866211,33.00927734375,32.9920883178711,32.9812469482422,32.9675788879395,32.9203147888184,32.8671875,32.81103515625,32.76513671875,32.7853507995605,32.8067359924316,32.7974624633789,32.7717781066895,32.6720695495605,32.67041015625,32.7583999633789,32.8140640258789,32.8518524169922,32.8997077941895,32.9385719299316,32.9675788879395,32.9776344299316,32.9710922241211,32.9904289245605,33,32.9705085754395,32.9456062316895,32.9751930236816,33.0215797424316,33.2434577941895,33.39794921875,33.4306640625,33.4832038879395,33.5123062133789,33.4914054870605,33.3700180053711,33.3401374816895,33.2523422241211,33.3009757995605,33.3050765991211,33.3039054870605,33.2882843017578,33.25,33.2263679504395,33.2327156066895,33.2683601379395,33.3455047607422,33.3797836303711,33.3386726379395,33.2932624816895,33.2727546691895,33.2613258361816,33.2927742004395,33.3449211120605,33.4031257629395,33.4647483825684,33.6590843200684,33.6615257263184,33.6261711120605,33.5537109375,33.53759765625,33.5289077758789,33.5000953674316,33.3935546875,33.3115234375,33.3371086120605,33.3509788513184,33.3104515075684,33.25,33.2126960754395,33.1957015991211,33.1480445861816,33.1044921875,33.0724639892578,33.0377922058105,32.9959945678711,32.9821281433105,32.9798812866211,32.9510726928711,32.92333984375,32.919921875,32.9373016357422,32.9740219116211,33.1304664611816,33.2252960205078,33.3308601379395,33.4208984375,33.4677734375,33.5275421142578,33.6976585388184,33.7662124633789,33.8541984558105,33.8888664245605,33.9439468383789,33.9537086486816,33.9593772888184,33.9496078491211,33.9621086120605,33.99560546875,34.0885772705078,34.3208961486816,34.3278312683105,34.4759788513184,34.5242195129395,34.5242195129395,34.5699234008789,34.5799789428711,34.5697250366211,34.5715827941895,34.5895500183105,34.5835952758789,34.6365242004395,34.6618156433105,34.6670913696289,34.65234375,34.6056671142578,34.59765625,34.60791015625,34.6380844116211,34.6884765625,34.7264633178711,34.7521476745605,34.7738265991211,34.8008804321289,34.8505859375,34.890625,34.93701171875,34.95263671875,34.95947265625],[111.389259338379,111.358695983887,111.3115234375,111.300392150879,111.33349609375,111.355079650879,111.378318786621,111.376266479492,111.380470275879,111.389259338379],[104.221580505371,104.17333984375,104.146873474121,104.129104614258,104.169822692871,104.184768676758,104.223243713379,104.221580505371],[101.318557739258,101.26806640625,101.265426635742,101.274223327637,101.311233520508,101.328414916992,101.318557739258],[117.884765625,117.745414733887,117.649024963379,117.666793823242,117.662109375,117.7080078125,117.761421203613,117.884765625],[100.288963317871,100.263771057129,100.191017150879,100.203910827637,100.245506286621,100.310157775879,100.3388671875,100.288963317871],[99.8480529785156,99.9186553955078,99.8833923339844,99.8658218383789,99.8232421875,99.7826156616211,99.7437515258789,99.7046890258789,99.6566390991211,99.6462936401367,99.7105407714844,99.7492141723633,99.8216781616211,99.8480529785156],[102.10107421875,102.274024963379,102.340141296387,102.534378051758,102.790237426758,102.898536682129,102.982421875,103.097068786621,103.196975708008,103.415817260742,103.453903198242,103.46875,103.420509338379,103.362007141113,103.373344421387,103.453514099121,103.429496765137,103.445014953613,103.439453125,103.485160827637,103.537307739258,103.812210083008,103.832321166992,103.9677734375,104.218559265137,104.288475036621,104.280372619629,104.250099182129,104.176368713379,104.114944458008,104.09423828125,104.1005859375,104.076171875,104.016014099121,103.9814453125,103.9912109375,103.991500854492,103.915138244629,103.816795349121,103.694534301758,103.5498046875,103.480278015137,103.427345275879,103.400001525879,103.356834411621,102.896873474121,102.727149963379,102.548240661621,102.145599365234,101.889938354492,101.78125,101.519721984863,101.406837463379,101.351364135742,101.295509338379,101.354293823242,101.330169677734,101.299903869629,101.115432739258,101.024803161621,100.851264953613,100.781837463379,100.715431213379,100.757034301758,100.795509338379,100.76025390625,100.661033630371,100.614555358887,100.614555358887,100.473434448242,100.352638244629,100.3740234375,100.34326171875,100.263282775879,100.158401489258,100.119140625,100.137992858887,100.161224365234,100.1767578125,100.216598510742,100.261428833008,100.345413208008,100.563865661621,100.629493713379,100.715629577637,100.754493713379,100.793746948242,100.816505432129,100.873924255371,100.98876953125,101.029388427734,101.053512573242,101.075973510742,101.086524963379,101.075584411621,100.992774963379,100.981643676758,101.025199890137,101.081741333008,101.113960266113,101.147659301758,101.190628051758,101.229782104492,101.257026672363,101.404197692871,101.556053161621,101.576759338379,101.601364135742,101.649993896484,101.678413391113,101.719528198242,101.790725708008,101.873634338379,101.917190551758,101.936134338379,102.05517578125,102.068359375,102.10107421875],[117.574409484863,117.537307739258,117.450881958008,117.277542114258,117.1005859375,116.843551635742,116.697853088379,116.638679504395,116.589057922363,116.553123474121,116.514747619629,116.41455078125,116.367683410645,116.320304870605,116.236228942871,116.134468078613,116.021583557129,115.896194458008,115.860748291016,115.836814880371,115.782417297363,115.678802490234,115.627532958984,115.596092224121,115.568450927734,115.560943603516,115.544532775879,115.570709228516,115.566116333008,115.519920349121,115.514259338379,115.48974609375,115.499130249023,115.493156433105,115.454391479492,115.384178161621,115.310150146484,115.246971130371,115.18994140625,115.117576599121,115.086326599121,115.086532592773,115.093643188477,115.078903198242,115.077049255371,115.080764770508,115.129890441895,115.180854797363,115.179107666016,115.150787353516,115.086532592773,114.969146728516,114.836326599121,114.786422729492,114.768363952637,114.758697509766,114.787994384766,114.815818786621,114.830558776855,114.812690734863,114.800003051758,114.751068115234,114.703521728516,114.686126708984,114.660942077637,114.632225036621,114.567481994629,114.5458984375,114.512496948242,114.387107849121,114.274703979492,114.1259765625,114,113.90234375,113.835250854492,113.760353088379,113.681640625,113.622268676758,113.51318359375,113.458206176758,113.358985900879,113.126274108887,113.068649291992,113.006546020508,112.98828125,112.998046875,112.98828125,112.942970275879,112.476173400879,112.341598510742,112.250679016113,112.185737609863,112.167381286621,112.128616333008,112.078514099121,111.923149108887,111.808982849121,111.769729614258,111.691307067871,111.607421875,111.546684265137,111.483200073242,111.286720275879,111.101364135742,110.99609375,110.938087463379,110.61474609375,110.505760192871,110.46142578125,110.399024963379,110.315231323242,110.11474609375,110.040824890137,109.99169921875,109.944923400879,109.878517150879,109.818069458008,109.735748291016,109.653999328613,109.635841369629,109.57080078125,109.548927307129,109.538970947266,109.62890625,109.6943359375,109.719627380371,109.864845275879,109.984565734863,110.114059448242,110.245895385742,110.29833984375,110.349220275879,110.399513244629,110.675193786621,110.782035827637,110.894920349121,110.93994140625,111.098434448242,111.145217895508,111.223243713379,111.123435974121,111.058006286621,111.028717041016,111.046585083008,111.110153198242,111.154197692871,111.170013427734,111.198051452637,111.250877380371,111.268165588379,111.208885192871,111.195503234863,111.208595275879,111.2421875,111.2958984375,111.351364135742,111.406150817871,111.44384765625,111.450775146484,111.4404296875,111.443267822266,111.512504577637,111.623237609863,111.727737426758,112.118843078613,112.7373046875,112.920501708984,112.987892150879,113.044723510742,113.140235900879,113.320121765137,113.446098327637,113.712112426758,113.923927307129,113.952537536621,113.98779296875,113.990425109863,113.984283447266,114.012504577637,114.053611755371,114.063865661621,114.095123291016,114.168846130371,114.22412109375,114.261032104492,114.28759765625,114.289649963379,114.322944641113,114.416603088379,114.447074890137,114.51220703125,114.57177734375,114.608306884766,114.654098510742,114.724990844727,114.776168823242,114.810455322266,114.783500671387,114.8310546875,114.840232849121,114.818267822266,114.790138244629,114.779296875,114.759963989258,114.746688842773,114.784187316895,114.864547729492,114.944717407227,115.026748657227,115.028816223145,115.026748657227,115.051559448242,115.107032775879,115.170608520508,115.246681213379,115.290626525879,115.319236755371,115.326759338379,115.279289245605,115.266693115234,115.227928161621,115.168449401855,115.140037536621,115.374900817871,115.427642822266,115.519821166992,115.554489135742,115.58203125,115.466888427734,115.421684265137,115.419036865234,115.556449890137,115.603904724121,115.624519348145,115.68505859375,115.740821838379,115.796867370605,115.877151489258,115.91845703125,116.059761047363,116.110061645508,116.138381958008,116.494720458984,116.538284301758,116.749801635742,116.776168823242,116.8330078125,116.849807739258,116.841995239258,116.807708740234,116.788078308105,116.812408447266,116.913276672363,117.0185546875,117.077926635742,117.128509521484,117.229881286621,117.252433776855,117.245307922363,117.254974365234,117.294044494629,117.380363464355,117.499221801758,117.609672546387,117.645706176758,117.669624328613,117.693748474121,117.695610046387,117.615913391113,117.649803161621,117.64453125,117.617179870605,117.501167297363,117.817672729492,117.895797729492,118.003814697266,118.061721801758,118.115814208984,118.072265625,117.934761047363,117.928024291992,117.9736328125,118.031143188477,118.144622802734,118.249122619629,118.2998046875,118.353118896484,118.456352233887,118.514167785645,118.563079833984,118.594825744629,118.713676452637,118.957321166992,119.002532958984,119.050003051758,119.178413391113,119.223434448242,119.255569458008,119.266304016113,119.262794494629,119.249710083008,119.219627380371,119.132225036621,118.912498474121,118.672065734863,118.551361083984,118.3818359375,118.320022583008,118.260543823242,118.185348510742,118.32421875,118.562492370605,118.595115661621,118.586334228516,118.54833984375,118.498046875,118.364067077637,118.228713989258,118.117286682129,118.008193969727,117.895606994629,117.741012573242,117.696479797363,117.649803161621,117.603805541992,117.574409484863],[117.141593933105,117.080657958984,117.060150146484,117.064254760742,117.146865844727,117.264060974121,117.280769348145,117.266792297363,117.239349365234,117.141593933105],[23.3806648254395,23.5949211120605,23.7992191314697,24.0369148254395,24.2271480560303,24.27490234375,24.73291015625,24.9324226379395,25.0017566680908,25.0921878814697,25.2587890625,25.2160148620605,24.9090824127197,24.7921886444092,24.5305671691895,24.4749011993408,24.4122085571289,24.3589859008789,24.2439460754395,24.1292972564697,24.0026378631592,23.8983402252197,23.8642578125,23.7004890441895,23.6471691131592,23.5997066497803,23.58056640625,23.5601577758789,23.4597644805908,23.2986316680908,23.2515621185303,23.2193355560303,23.0999031066895,22.7527351379395,22.4600582122803,22.0114250183105,21.5296859741211,21.2325191497803,20.97412109375,20.9743156433105,20.9750003814697,20.9755859375,20.9761714935303,20.9768543243408,20.9774417877197,20.9781246185303,20.9787101745605,20.9792957305908,20.9794921875,20.9709968566895,20.82275390625,20.4875011444092,20.2053718566895,19.9773445129395,19.9776382446289,19.9779300689697,19.9782238006592,19.978515625,19.9789066314697,19.9792976379395,19.9795913696289,19.9798831939697,19.9801750183105,19.98046875,19.98046875,19.98046875,19.98046875,19.98046875,19.98046875,19.98046875,19.98046875,19.98046875,19.98046875,19.98046875,19.8778324127197,19.6714839935303,19.5398445129395,19.48291015625,19.4072265625,19.3126945495605,19.27099609375,19.2822265625,19.2458019256592,19.1617183685303,19.0260734558105,18.8387699127197,18.6003913879395,18.3108386993408,18.1027355194092,17.97607421875,17.8416023254395,17.6993160247803,17.6167964935303,17.4479484558105,17.4157218933105,17.3958969116211,17.3478527069092,17.3425769805908,17.3802738189697,17.3857421875,17.3586921691895,17.31201171875,17.2458000183105,17.20458984375,17.1884765625,17.1494140625,17.0562496185303,16.9333000183105,16.8752918243408,16.8412113189697,16.8101558685303,16.7945327758789,16.7875003814697,16.7557621002197,16.7230472564697,16.689453125,16.6261711120605,16.4871082305908,16.4475574493408,16.3350582122803,16.0071296691895,15.8909187316895,15.7190437316895,15.3415050506592,15.28759765625,15.2157230377197,15.1328115463257,15.1237306594849,15.1632814407349,15.1390628814697,15.0965824127197,14.9677734375,14.9312496185303,14.84521484375,14.8636713027954,14.8225593566895,14.8185548782349,14.8371086120605,14.7679681777954,14.6279296875,14.5015621185303,14.4833993911743,14.4968748092651,14.4724607467651,14.4738283157349,14.423828125,14.4033203125,14.4384765625,14.4592781066895,14.4957036972046,14.5199222564697,14.5259771347046,14.4627923965454,14.3218746185303,13.9732418060303,13.8880853652954,13.83935546875,13.4505853652954,13.2843751907349,13.1683597564697,13.0420904159546,12.4582023620605,12.3287105560303,12.095703125,12.0412111282349,11.9513673782349,11.77587890625,11.7334957122803,11.7216796875,11.7430667877197,11.9025392532349,12.0139646530151,12.1143560409546,12.21337890625,12.3184566497803,12.3592767715454,12.5481443405151,12.6565427780151,12.78515625,12.8592777252197,12.9631843566895,13.1011714935303,13.1794919967651,13.2756834030151,13.4037113189697,13.4759769439697,13.5617189407349,13.6943359375,13.7919921875,13.9041986465454,13.9379892349243,13.9874019622803,14.0174798965454,14.2258787155151,14.4147453308105,14.6179695129395,15.0005865097046,15.3832035064697,15.7658195495605,16.1484375,16.5310535430908,16.9136714935303,17.2962894439697,17.6788082122803,17.8353519439697,18.1087875366211,18.3963871002197,18.42822265625,18.4603519439697,18.4866199493408,18.5881843566895,18.7180671691895,18.8259773254395,18.9552726745605,19.0764636993408,19.189453125,19.3771476745605,19.6393547058105,19.9118156433105,20.1943359375,20.3929691314697,20.5076179504395,20.6250991821289,20.7455081939697,20.9083003997803,21.1134757995605,21.2878913879395,21.3687496185303,21.4168949127197,21.7184562683105,21.9608383178711,22.32421875,22.6240234375,23.0682621002197,23.3806648254395],[167.54443359375,167.5126953125,167.473434448242,167.443756103516,167.422073364258,167.443450927734,167.529479980469,167.54443359375],[168.010940551758,168.057907104492,168.139068603516,168.120712280273,168.006454467773,167.966796875,167.941299438477,167.875869750977,167.879089355469,167.8154296875,167.925979614258,167.988479614258,167.984970092773,167.99462890625,168.010940551758],[167.40087890625,167.34619140625,167.273239135742,167.133880615234,167.072662353516,167.03271484375,167.111724853516,167.189453125,167.136428833008,167.045028686523,167.055755615234,167.203994750977,167.268936157227,167.297943115234,167.29345703125,167.36083984375,167.430572509766,167.430267333984,167.40087890625],[166.546768188477,166.493545532227,166.557815551758,166.559661865234,166.58544921875,166.58251953125,166.624710083008,166.670791625977,166.617874145508,166.600296020508,166.602157592773,166.62255859375,166.5888671875,166.546768188477],[164.202331542969,164.315124511719,164.435943603516,164.588073730469,164.975677490234,165.111907958984,165.191802978516,165.252349853516,165.306640625,165.380569458008,165.412490844727,165.420501708984,165.447174072266,165.582412719727,165.662796020508,165.774612426758,165.822860717773,165.885345458984,165.949508666992,166.057815551758,166.303314208984,166.492965698242,166.587493896484,166.689651489258,166.820114135742,166.9423828125,167.004302978516,166.970306396484,166.900009155273,166.8349609375,166.774124145508,166.570602416992,166.522171020508,166.467971801758,166.437698364258,166.416412353516,166.292282104492,166.176651000977,166.143157958984,166.123718261719,166.096084594727,165.933013916016,165.823440551758,165.743835449219,165.620223999023,165.427627563477,165.32861328125,165.241989135742,165.010162353516,164.927444458008,164.855270385742,164.655670166016,164.559478759766,164.454681396484,164.37451171875,164.312896728516,164.169723510742,164.152145385742,164.158111572266,164.123626708984,164.065032958984,164.037307739258,164.04052734375,164.059661865234,164.202331542969],[159.951766967773,159.936416625977,159.92822265625,159.959854125977,159.97509765625,159.951766967773],[14.9790048599243,15.0889654159546,15.1722660064697,15.1778316497803,15.1818351745605,15.2158203125,15.2936525344849,15.6073236465454,15.5403318405151,15.5871095657349,15.66845703125,15.9292974472046,15.9631834030151,15.9488277435303,15.7662115097046,15.7350587844849,15.6986331939697,15.6729497909546,15.637598991394,15.5955076217651,15.5615243911743,15.5166988372803,15.4743156433105,15.2121086120605,14.7466793060303,14.3679685592651,14.17822265625,13.80712890625,13.6423835754395,13.513671875,13.4482421875,13.5057621002197,13.6063480377197,13.4269533157349,13.3238286972046,13.19384765625,13.0484380722046,12.8716793060303,12.7599601745605,12.65478515625,12.5101566314697,12.4631834030151,12.3190431594849,12.1179685592651,11.9900398254395,11.693359375,11.5010738372803,11.4119138717651,10.9588861465454,10.4758787155151,10.2295894622803,10.1846685409546,10.0451173782349,9.92929744720459,9.61591720581055,9.20156288146973,8.95761680603027,8.75058650970459,8.4560546875,8.09501934051514,7.95576143264771,7.83046865463257,7.78867197036743,7.35781192779541,7.27470731735229,7.17304658889771,7.10605430603027,7.05673837661743,7.00507831573486,6.93720674514771,6.87060546875,6.80429697036743,6.62656259536743,6.58994150161743,6.51406240463257,6.38632822036743,6.2998046875,6.24716806411743,6.18427753448486,5.83818387985229,5.49199247360229,5.41582059860229,5.36162090301514,5.24189424514771,5.10087871551514,4.92167997360229,4.82333946228027,4.66484355926514,4.55947256088257,4.42138671875,4.2421875,4.19082021713257,4.14755868911743,4.08740234375,4.03876972198486,3.94785141944885,3.76923823356628,3.64667987823486,3.64384794235229,3.63417983055115,3.63251948356628,3.64062476158142,3.62011671066284,3.61181640625,3.61845707893372,3.64736318588257,3.66474652290344,3.65312504768372,3.59541034698486,3.53173828125,3.44980478286743,3.35996079444885,3.29912137985229,3.26738309860229,3.14960908889771,2.87812519073486,2.85019516944885,2.80527353286743,2.728515625,2.68134737014771,2.64843726158142,2.59843754768372,2.46933579444885,2.36601543426514,2.36328125,2.41269540786743,2.38916015625,2.34335947036743,2.19443345069885,2.09140658378601,2.07294917106628,2.05839848518372,2.06855463981628,2.109375,2.20380878448486,2.22138667106628,2.22626948356628,2.21152377128601,2.15976548194885,2.10458970069885,2.07382822036743,2.01738262176514,1.95615231990814,1.84091806411743,1.78984391689301,1.67109382152557,1.56494140625,1.50048816204071,1.30869114398956,1.09677755832672,1.00791013240814,0.98730480670929,0.973046779632568,0.976757824420929,0.988476514816284,1.07685530185699,1.17089831829071,1.20117163658142,1.12597668170929,1.01787102222443,0.977734088897705,0.946581840515137,0.897949159145355,0.842285215854645,0.786035060882568,0.747754037380219,0.684570550918579,0.61816418170929,0.522363305091858,0.429199248552322,0.374023407697678,0.354882627725601,0.38251981139183,0.354589819908142,0.250585943460464,0.163867071270943,0.185058489441872,0.202734231948853,0.203808531165123,0.217480450868607,0.228710994124413,0.286230236291885,0.433007687330246,0.718652367591858,0.947460770606995,0.960058450698853,1.12128937244415,1.30019533634186,1.56914055347443,1.85937511920929,2.08818364143372,2.42080044746399,2.68964838981628,3.00107431411743,3.01054716110229,3.02939462661743,3.06015610694885,3.2890625,3.50429701805115,3.52050757408142,3.70957040786743,3.81650400161743,3.84296870231628,3.87695336341858,3.89794921875,3.90722680091858,3.94707036018372,3.97617197036743,4.01484346389771,4.12128877639771,4.18212890625,4.19121122360229,4.20292949676514,4.23466825485229,4.23369121551514,4.23271465301514,4.23193311691284,4.23085927963257,4.22998046875,4.22900390625,4.22822284698486,4.22763633728027,4.44570302963257,4.67128896713257,5.00136661529541,5.35869169235229,5.74834012985229,5.83662128448486,6.13066387176514,6.26337862014771,6.52705097198486,6.73066377639771,6.98935556411743,7.26337957382202,7.48173904418945,7.82519483566284,8.34306621551514,8.86093807220459,9.37871074676514,9.896484375,10.4143552780151,10.9322261810303,11.4499998092651,11.9678707122803,12.48876953125,12.9835939407349,13.4812498092651,13.5986337661743,13.8626956939697,14.2006826400757,14.2155275344849,14.2307615280151,14.5556640625,14.9789056777954,14.9790048599243],[167.939453125,167.959762573242,167.978118896484,167.990432739258,167.988662719727,167.97900390625,167.967666625977,167.964157104492,167.9619140625,167.960647583008,167.960739135742,167.954696655273,167.944427490234,167.933792114258,167.926559448242,167.92041015625,167.918258666992,167.914260864258,167.912399291992,167.916397094727,167.924026489258,167.924606323242,167.918548583984,167.906158447266,167.920608520508,167.939453125],[7.30078172683716,7.20390653610229,7.14042949676514,7.22734355926514,7.27138710021973,7.32792997360229,7.30078172683716],[13.6063480377197,13.7634763717651,13.9323244094849,14.06396484375,14.1600580215454,14.1703128814697,14.1776361465454,14.1848630905151,14.1974611282349,14.2728519439697,14.4154300689697,14.5189447402954,14.5809564590454,14.5870113372803,14.6197261810303,14.6271486282349,14.6181640625,14.5973625183105,14.5618162155151,14.5816411972046,14.5753917694092,14.5597658157349,14.4960927963257,14.4094724655151,14.2023439407349,14.1432619094849,14.0567388534546,13.9814453125,13.89208984375,13.6999025344849,13.5353517532349,13.478515625,13.41455078125,13.2699222564697,13.2498044967651,13.2437496185303,13.23876953125,13.22119140625,13.19873046875,13.1754875183105,13.0194330215454,12.9294929504395,12.8756837844849,12.85595703125,12.8244142532349,12.8065433502197,12.7822265625,12.7311525344849,12.6515626907349,12.5827140808105,12.4035158157349,12.3113279342651,12.2333984375,12.2311525344849,12.1559562683105,12.0251951217651,12.0166015625,12.0160160064697,11.8524417877197,11.8091802597046,11.7673826217651,11.80859375,11.8547849655151,11.8614253997803,11.7870111465454,11.6575193405151,11.580078125,11.56298828125,11.5516595840454,11.5291013717651,11.4775390625,11.4017581939697,11.3246088027954,11.2373046875,11.1533203125,11.1064453125,11.0796880722046,11.0325193405151,11.0086917877197,10.9541997909546,10.8464841842651,10.7375974655151,10.6062498092651,10.578125,10.5563478469849,10.51904296875,10.4823236465454,10.4131832122803,10.2930660247803,10.2054691314697,10.185546875,10.1677732467651,10.1435546875,10.0388679504395,9.87421894073486,9.82070255279541,9.77988338470459,9.7255859375,9.65996074676514,9.57402324676514,9.490234375,9.44218730926514,9.37333965301514,9.23876953125,9.06015586853027,8.99716758728027,8.93505859375,8.89882850646973,8.85917949676514,8.80097675323486,8.71562480926514,8.64052772521973,8.58515548706055,8.55585956573486,8.54375076293945,8.51484394073486,8.43134784698486,8.39365291595459,8.34208965301514,8.25273513793945,8.23378849029541,8.32802772521973,8.29306602478027,8.02851486206055,7.80078125,7.64423847198486,7.56562566757202,7.53076171875,7.51738309860229,7.50947284698486,7.45986366271973,7.28437519073486,7.20673847198486,7.14384746551514,7.07656240463257,7.08691358566284,7.16415977478027,7.15468740463257,7.01337862014771,6.92324209213257,6.86787080764771,6.83915996551514,6.82470703125,6.78759765625,6.76767587661743,6.78603553771973,6.79218769073486,6.79306650161743,6.86035108566284,6.75703096389771,6.71513700485229,6.63300800323486,6.61728477478027,6.60156202316284,6.57997989654541,6.55458974838257,6.5,6.46211004257202,6.2998046875,6.263671875,6.25595712661743,6.27529287338257,6.27099609375,6.21464872360229,6.20556688308716,6.17334032058716,6.07656240463257,5.970703125,5.90644550323486,5.79863262176514,5.58779335021973,5.55361366271973,5.49326181411743,5.44814443588257,5.38330078125,5.40322303771973,5.4521484375,5.47597646713257,5.38828134536743,5.37001943588257,5.36416053771973,5.36796855926514,5.43925762176514,5.50087881088257,5.53183603286743,5.54970693588257,5.38583993911743,5.232421875,5.19921875,5.21582078933716,5.2890625,5.39384794235229,5.45664119720459,5.41806650161743,5.35029315948486,5.32529306411743,5.32734394073486,5.30537080764771,5.27626991271973,5.1728515625,5.11240196228027,5.10624980926514,5.09306669235229,5.04208993911743,4.86103534698486,4.63359403610229,4.43134784698486,4.12587881088257,3.48662114143372,3.45078110694885,3.48994159698486,3.54609346389771,3.75165987014771,3.71699190139771,3.50332021713257,3.43017554283142,3.33554720878601,2.7724609375,2.70644521713257,2.70800757408142,2.73564434051514,2.75371074676514,2.77460932731628,2.75292992591858,2.73173832893372,2.72138690948486,2.74775362014771,2.75673818588257,2.75058579444885,2.75048851966858,2.76582026481628,2.78398418426514,2.78515648841858,2.75097680091858,2.71933603286743,2.72040987014771,2.70771503448486,2.68603539466858,2.70234417915344,2.71152353286743,2.70312476158142,2.7236328125,2.73466801643372,2.73291015625,2.77480483055115,2.89804697036743,3.044921875,3.11044907569885,3.14804673194885,3.13613247871399,3.16464829444885,3.22343754768372,3.32949209213257,3.3251953125,3.35449194908142,3.40478515625,3.47675776481628,3.55722665786743,3.60205101966858,3.64589810371399,3.57656240463257,3.57792949676514,3.60410165786743,3.64658212661743,3.68027377128601,3.75849604606628,3.77177739143372,3.78378915786743,3.83447241783142,3.82968783378601,3.75683617591858,3.74492192268372,3.73417973518372,3.71640658378601,3.6953125,3.65624976158142,3.63886690139771,3.48779296875,3.49052691459656,3.55390644073486,3.59541034698486,3.65312504768372,3.66474652290344,3.64736318588257,3.61845707893372,3.61181640625,3.62011671066284,3.64062476158142,3.63251948356628,3.63417983055115,3.64384794235229,3.64667987823486,3.76923823356628,3.94785141944885,4.03876972198486,4.08740234375,4.14755868911743,4.19082021713257,4.2421875,4.42138671875,4.55947256088257,4.66484355926514,4.82333946228027,4.92167997360229,5.10087871551514,5.24189424514771,5.36162090301514,5.41582059860229,5.49199247360229,5.83818387985229,6.18427753448486,6.24716806411743,6.2998046875,6.38632822036743,6.51406240463257,6.58994150161743,6.62656259536743,6.80429697036743,6.87060546875,6.93720674514771,7.00507831573486,7.05673837661743,7.10605430603027,7.17304658889771,7.27470731735229,7.35781192779541,7.78867197036743,7.83046865463257,7.95576143264771,8.09501934051514,8.4560546875,8.75058650970459,8.95761680603027,9.20156288146973,9.61591720581055,9.92929744720459,10.0451173782349,10.1846685409546,10.2295894622803,10.4758787155151,10.9588861465454,11.4119138717651,11.5010738372803,11.693359375,11.9900398254395,12.1179685592651,12.3190431594849,12.4631834030151,12.5101566314697,12.65478515625,12.7599601745605,12.8716793060303,13.0484380722046,13.19384765625,13.3238286972046,13.4269533157349,13.6063480377197],[-83.1575164794922,-83.1853485107422,-83.2159194946289,-83.2798843383789,-83.302001953125,-83.3063507080078,-83.3443832397461,-83.3890151977539,-83.4137191772461,-83.3748550415039,-83.3407287597656,-83.2992172241211,-83.187744140625,-83.2117156982422,-83.2808074951172,-83.3465881347656,-83.4123077392578,-83.4937515258789,-83.5673370361328,-83.5144500732422,-83.5412139892578,-83.5179672241211,-83.5109405517578,-83.5652313232422,-83.5959014892578,-83.627197265625,-83.6236801147461,-83.5912094116211,-83.5780715942383,-83.5933609008789,-83.6253356933594,-83.681640625,-83.7183609008789,-83.7542419433594,-83.7162094116211,-83.6673278808594,-83.6512680053711,-83.6692352294922,-83.680419921875,-83.6977081298828,-83.715576171875,-83.7671890258789,-83.7733459472656,-83.7693328857422,-83.8131866455078,-83.8289031982422,-83.79296875,-83.7533721923828,-83.70458984375,-83.664306640625,-83.6517562866211,-83.7449722290039,-83.776611328125,-83.8293914794922,-83.8590850830078,-83.8678741455078,-83.8318328857422,-83.7679214477539,-83.7140655517578,-83.6419906616211,-83.658935546875,-83.7129364013672,-83.8111801147461,-83.9192886352539,-84.09619140625,-84.1683578491211,-84.1965789794922,-84.2049865722656,-84.2555694580078,-84.3482971191406,-84.40185546875,-84.4891586303711,-84.6341781616211,-84.701171875,-84.79736328125,-84.9091796875,-85.178955078125,-85.3683624267578,-85.5387191772461,-85.5841827392578,-85.6213836669922,-85.6536560058594,-85.6905288696289,-85.70263671875,-85.7222671508789,-85.7443389892578,-85.7452163696289,-85.8285140991211,-85.9611358642578,-86.4688949584961,-86.655517578125,-86.755615234375,-86.8509750366211,-87.1251983642578,-87.1884307861328,-87.4601593017578,-87.6675262451172,-87.670166015625,-87.5850601196289,-87.5433120727539,-87.4979476928711,-87.4243621826172,-87.3896484375,-87.3385696411133,-87.3372497558594,-87.0591812133789,-87.0093307495117,-86.9588851928711,-86.9331512451172,-86.9288101196289,-86.918212890625,-86.87353515625,-86.7921371459961,-86.7293014526367,-86.710693359375,-86.7295913696289,-86.7635269165039,-86.7706069946289,-86.7589797973633,-86.733642578125,-86.6102523803711,-86.376953125,-86.3317413330078,-86.2382354736328,-86.1512145996094,-86.0892562866211,-86.0403747558594,-85.9837875366211,-85.7867202758789,-85.75341796875,-85.7339324951172,-85.7277297973633,-85.731201171875,-85.6819381713867,-85.5797805786133,-85.47705078125,-85.373779296875,-85.2841796875,-85.2083511352539,-85.1794967651367,-85.1975631713867,-85.1914978027344,-85.1613235473633,-85.1172866821289,-85.0593719482422,-85.0365219116211,-85.0486373901367,-85.037353515625,-84.9851531982422,-84.8604507446289,-84.7891616821289,-84.7297897338867,-84.6459503173828,-84.5376510620117,-84.4535598754883,-84.3936538696289,-84.3397979736328,-84.2919464111328,-84.2666549682617,-84.2639617919922,-84.2392120361328,-84.1923828125,-84.1507797241211,-84.1144027709961,-84.1002883911133,-84.0929718017578,-84.0658187866211,-83.9722671508789,-83.8672866821289,-83.7509231567383,-83.6736373901367,-83.635498046875,-83.5897445678711,-83.5365295410156,-83.4150390625,-83.1575164794922],[-169.803421020508,-169.90380859375,-169.948348999023,-169.908737182617,-169.861572265625,-169.834030151367,-169.793411254883,-169.803421020508],[-68.205810546875,-68.2543487548828,-68.2822265625,-68.2872543334961,-68.30712890625,-68.3484420776367,-68.37109375,-68.3692398071289,-68.219482421875,-68.205810546875],[-62.9375038146973,-62.9617156982422,-62.9831085205078,-62.9971694946289,-62.9996109008789,-62.9793434143066,-62.9717750549316,-62.9654312133789,-62.9375038146973],[-63.2326698303223,-63.2416038513184,-63.2544898986816,-63.2521476745605,-63.2416496276855,-63.2334938049316,-63.2269058227539,-63.2326698303223],[4.22617149353027,4.21142578125,4.17255878448486,4.0400390625,3.90205025672913,3.83076143264771,3.78193330764771,3.75566411018372,3.68183612823486,3.58027362823486,3.51709008216858,3.47197222709656,3.43251967430115,3.40283179283142,3.38007783889771,3.35009741783142,3.42578101158142,3.58945345878601,3.71650409698486,3.88339829444885,4.01103496551514,4.11152315139771,4.22617149353027],[3.94912147521973,4.04677724838257,4.06757831573486,4.07509756088257,3.95097661018372,3.81904268264771,3.73183560371399,3.69902348518372,3.69853544235229,3.78906273841858,3.94912147521973],[4.88613271713257,4.787109375,4.72675800323486,4.70917987823486,4.73984384536743,4.88642597198486,4.88613271713257],[5.10859394073486,4.92373037338257,4.90791034698486,5.02705097198486,5.10859394073486],[4.22617149353027,4.13886690139771,4.00654315948486,3.82187509536743,3.69355463981628,3.5869140625,3.52050757408142,3.44892573356628,3.49960970878601,3.54863309860229,3.74394512176514,3.88603520393372,4.14130830764771,4.20576143264771,4.27412128448486,4.23935556411743,4.17548847198486,4.08046865463257,4.00478553771973,4.1826171875,4.15800762176514,4.13457059860229,3.94687485694885,3.97890567779541,4.02607440948486,4.08486270904541,4.13173866271973,4.20878934860229,4.37626934051514,4.48281288146973,4.56210947036743,4.67831993103027,4.71269512176514,4.76875019073486,4.83906221389771,4.88798809051514,5.06123065948486,5.3583984375,5.44599580764771,5.53203153610229,5.87353515625,6.06220674514771,6.35322237014771,6.56357383728027,6.81621074676514,6.91240262985229,6.96816396713257,7.05800771713257,7.197265625,7.18896532058716,7.18994140625,7.17949295043945,7.11708974838257,7.05087900161743,7.03300762176514,7.01318311691284,6.74843740463257,6.71074247360229,6.70537090301514,6.71875,6.71240234375,6.69160175323486,6.70292949676514,6.74882793426514,6.83251953125,6.92207050323486,6.96816396713257,7.00185537338257,7.03515625,7.03261661529541,7.01962900161743,6.97724628448486,6.85507774353027,6.80039072036743,6.7490234375,6.72451162338257,6.71298837661743,6.71562528610229,6.80244159698486,6.80039072036743,6.77519559860229,6.74179649353027,6.517578125,6.42499971389771,6.37216758728027,6.35566377639771,6.29707050323486,6.16650390625,6.1171875,6.08984375,6.00761699676514,5.94872999191284,5.94853496551514,6.052734375,6.08935546875,6.09111309051514,6.1416015625,6.19326162338257,6.19882822036743,6.19287109375,6.16621065139771,6.07587909698486,6.07480430603027,6.08242177963257,6.11337852478027,6.13691425323486,6.12998056411743,5.96103525161743,5.93925762176514,5.86835956573486,5.85751962661743,5.8671875,5.89472675323486,5.955078125,6.00683546066284,6.04843759536743,5.99394512176514,5.89246129989624,5.79736328125,5.7470703125,5.69355487823486,5.66914081573486,5.63945341110229,5.64755868911743,5.73662137985229,5.75,5.74082040786743,5.74980449676514,5.81826162338257,5.8271484375,5.79648447036743,5.75234413146973,5.60878896713257,5.54042959213257,5.5087890625,5.47685527801514,5.42978525161743,5.31083965301514,5.21415996551514,5.09990262985229,5.07343769073486,5.05947303771973,5.03095674514771,4.99257850646973,4.94394493103027,4.84804677963257,4.82070302963257,4.81601524353027,4.81054639816284,4.78418016433716,4.75566434860229,4.63398408889771,4.58876943588257,4.53164052963257,4.50341796875,4.44091796875,4.38476610183716,4.40400409698486,4.37373018264771,4.30449199676514,4.22617149353027],[5.32578134536743,5.23261737823486,5.19023418426514,5.41513681411743,5.55742168426514,5.58261728286743,5.32578134536743],[5.92929649353027,5.73203134536743,5.66533184051514,5.654296875,5.70810508728027,5.87626981735229,5.92822265625,5.92929649353027],[6.33339881896973,6.19326162338257,6.15927696228027,6.16767597198486,6.29091787338257,6.33339881896973],[6.73476552963257,6.64208984375,6.66855478286743,6.75458955764771,6.80087900161743,6.73476552963257],[5.08583974838257,5.08906269073486,4.99697303771973,4.95556640625,4.94355487823486,4.95078086853027,4.93007850646973,4.95722675323486,4.99062490463257,5.05019521713257,5.08583974838257],[4.95869112014771,4.8701171875,4.79902315139771,4.82441425323486,4.86162137985229,4.91542959213257,4.97324228286743,4.95869112014771],[8.10273456573486,8.00468730926514,7.88828086853027,7.81533193588257,7.80400323867798,7.93837881088257,8.07353591918945,8.13613319396973,8.14091777801514,8.10273456573486],[8.47080039978027,8.35615253448486,8.287109375,8.45127010345459,8.70888614654541,8.7333984375,8.7646484375,8.80917930603027,8.81484413146973,8.78652381896973,8.47080039978027],[11.2314453125,11.1790037155151,11.0625,10.83251953125,10.7398433685303,10.8134765625,11.02099609375,11.1326169967651,11.2461919784546,11.2314453125],[11.9679679870605,11.90185546875,11.7783203125,11.76513671875,11.8003902435303,11.8753910064697,11.9723634719849,12.0032224655151,11.9679679870605],[12.5095701217651,12.4294919967651,12.43017578125,12.47607421875,12.548828125,12.6423826217651,12.7470703125,12.77880859375,12.7186527252197,12.5095701217651],[12.419921875,12.3273439407349,12.3427734375,12.4176759719849,12.4463863372803,12.4613275527954,12.5274410247803,12.6208009719849,12.6226558685303,12.5763673782349,12.419921875],[12.9717779159546,12.8240232467651,12.8779296875,12.9577150344849,13.0680665969849,13.1228513717651,13.0977544784546,13.0982418060303,13.0746097564697,12.9717779159546],[13.8728523254395,13.9323244094849,14.0876951217651,14.1188478469849,14.0967769622803,14.029296875,13.8876953125,13.8240232467651,13.7784175872803,13.6561517715454,13.583984375,13.4952154159546,13.4242181777954,13.4043951034546,13.3915033340454,13.35205078125,13.2293939590454,13.1995124816895,13.2559566497803,13.3001947402954,13.3679685592651,13.4287109375,13.5379877090454,13.6876955032349,13.7840824127197,13.8728523254395],[15.2071294784546,15.3372068405151,15.3965816497803,15.3484373092651,15.2220706939697,15.0270509719849,14.8902339935303,14.8040027618408,14.7932605743408,14.7434568405151,14.6121091842651,14.5208015441895,14.4046878814697,14.3734378814697,14.4966802597046,14.5537109375,14.6904306411743,14.7246084213257,14.8018550872803,14.8488283157349,14.8379878997803,14.8723630905151,15.0374994277954,15.0377931594849,15.1018552780151,15.1281242370605,15.1755857467651,15.2071294784546],[15.7603521347046,15.7723636627197,15.9085931777954,16.0595703125,16.0689449310303,16.12744140625,16.1208000183105,16.1505851745605,16.2275390625,16.2755851745605,16.3289051055908,16.4251956939697,16.4796867370605,16.54736328125,16.5192375183105,16.3379878997803,16.1939449310303,16.0484371185303,15.9752931594849,15.9125003814697,15.8727531433105,15.83740234375,15.763671875,15.6825189590454,15.4375,15.3414058685303,15.3370122909546,15.2796878814697,15.1878900527954,15.098048210144,15.0376958847046,14.9268560409546,14.62890625,14.3495121002197,14.2575197219849,14.2572269439697,14.4377918243408,14.5858402252197,15.0953121185303,15.4125986099243,15.4892568588257,15.5642576217651,15.5290040969849,15.4436521530151,15.4384765625,15.4830074310303,15.6495122909546,15.7419910430908,15.8926753997803,15.9653310775757,16.0480461120605,16.1294918060303,16.1148452758789,15.99267578125,15.8117189407349,15.8337888717651,15.9058589935303,15.9235363006592,15.9279298782349,15.7907228469849,15.7603521347046],[17.5030269622803,17.6232414245605,17.6773433685303,17.78369140625,17.86279296875,17.9274425506592,18.0041027069092,18.05224609375,18.0767574310303,18.0210933685303,17.9420890808105,17.9207038879395,17.95068359375,17.7735347747803,17.5681629180908,17.4878902435303,17.3236331939697,17.1609382629395,17.08251953125,17.0770511627197,16.9601573944092,16.8104496002197,16.8154296875,16.8425769805908,16.9717769622803,16.99755859375,16.97412109375,16.9968738555908,17.0017585754395,17.0830078125,17.36083984375,17.39453125,17.3734378814697,17.2298831939697,17.251953125,17.3555679321289,17.45361328125,17.4831047058105,17.4881839752197,17.5030269622803],[29.9561519622803,29.7662105560303,29.7442378997803,29.7859363555908,29.8358402252197,29.9139652252197,29.9929695129395,30.05517578125,29.9561519622803],[20.7791996002197,20.7252922058105,20.642578125,20.5980472564697,20.53466796875,20.4642581939697,20.4050769805908,20.4117183685303,20.4927730560303,20.6548824310303,20.7860355377197,20.8194332122803,20.7791996002197],[19.2550792694092,19.34375,19.4222660064697,19.4458980560303,19.49951171875,19.6078128814697,19.59228515625,19.4423828125,19.3347663879395,19.1970691680908,19.130859375,19.0078125,18.9091796875,18.8069324493408,18.8006839752197,18.7847652435303,18.4102535247803,18.2741222381592,18.1298828125,18.0615234375,18.08349609375,18.2274417877197,18.2320308685303,18.2684555053711,18.3150386810303,18.3493175506592,18.40625,18.5124034881592,18.583984375,18.6243152618408,18.6740245819092,18.6979503631592,18.6740245819092,18.6865234375,18.8238296508789,18.8832035064697,18.9686527252197,19.0509777069092,19.0749015808105,19.0509777069092,19.06005859375,19.1327152252197,19.2126960754395,19.2494144439697,19.2550792694092],[19.7674808502197,19.818359375,19.86865234375,19.9104480743408,19.994140625,20.0842761993408,20.0884761810303,20.0059566497803,19.8972663879395,19.7808589935303,19.7466793060303,19.7108402252197,19.6134757995605,19.5990238189697,19.6837882995605,19.7674808502197],[23.6153335571289,23.6339836120605,23.6410160064697,23.5477542877197,23.3328132629395,23.3451175689697,23.2707042694092,23.1591796875,23.1002922058105,23.1083984375,23.0906238555908,23.0059566497803,22.9178714752197,22.9177722930908,22.9410152435303,23.0224609375,23.1583976745605,23.248046875,23.5466785430908,23.5789051055908,23.6153335571289],[24.017578125,23.8271484375,23.7166004180908,23.6701183319092,23.6632804870605,23.6891613006592,23.7784175872803,23.8365230560303,23.85205078125,23.9564456939697,24.0783214569092,24.017578125],[23.4405288696289,23.4208984375,23.3871097564697,23.30517578125,23.0681648254395,22.9289054870605,22.884765625,22.8291015625,22.6560554504395,22.6053714752197,22.5575199127197,22.4322261810303,22.3586902618408,22.1687507629395,22.0557613372803,21.9945316314697,22.1700210571289,22.2326183319092,22.3502922058105,22.4209957122803,22.5707035064697,22.8581066131592,22.9635734558105,23.2046890258789,23.2801761627197,23.3956050872803,23.4405288696289],[30.8697261810303,30.9241218566895,30.9224624633789,30.8966789245605,30.8607406616211,30.7888679504395,30.6154289245605,30.3796882629395,30.2275390625,30.1801738739014,30.1597652435303,30.1964855194092,30.1867179870605,30.1637687683105,30.1318378448486,30.0873050689697,29.9940452575684,29.8327159881592,29.3882827758789,29.35302734375,29.2099590301514,29.1708984375,29.1185550689697,28.9658222198486,28.8918952941895,28.8326148986816,28.8462905883789,29.02490234375,29.1917991638184,29.2388687133789,29.3333988189697,29.1416034698486,28.8003902435303,28.4117183685303,28.2691402435303,28.0472660064697,27.8899421691895,27.7478523254395,27.5916996002197,27.3480472564697,27.2056636810303,27.1275386810303,27.1086921691895,26.9342765808105,26.740234375,26.5842781066895,26.525390625,26.3082027435303,26.1561527252197,26.0724601745605,26.0115242004395,25.9615230560303,25.8501949310303,25.7671871185303,25.7486324310303,25.7681636810303,25.7483386993408,25.6466789245605,25.5752925872803,25.4808597564697,25.3571281433105,25.2491207122803,25.1728515625,25.0869140625,24.94140625,24.8024425506592,24.7032241821289,24.4905281066895,24.33203125,24.1541023254395,23.9973621368408,23.85400390625,23.7725582122803,23.70703125,23.4625015258789,23.3240242004395,23.1443367004395,23.0716781616211,22.81103515625,22.5006828308105,22.4109382629395,22.3829097747803,22.3003902435303,22.0796871185303,21.9894523620605,21.8197250366211,21.6217765808105,21.59375,21.4612293243408,21.2667961120605,21.1437511444092,21.0661125183105,21.0526371002197,21.1278324127197,21.1044921875,21.0657215118408,20.8892574310303,20.6758785247803,20.6221675872803,20.4919910430908,20.11669921875,20.2823238372803,20.3371105194092,20.3480472564697,20.3194332122803,20.2400398254395,20.1474609375,19.9688472747803,20.2400398254395,20.0559577941895,19.9698238372803,19.8700199127197,19.6912097930908,19.2589836120605,19.0526371002197,18.8682613372803,18.7698249816895,18.3786125183105,18.3030261993408,18.16259765625,18.1470699310303,18.1559581756592,18.1766605377197,18.125,18.0732421875,17.9167003631592,17.5647468566895,17.3246097564697,17.1705074310303,16.7835941314697,16.5855464935303,16.5741214752197,16.4571285247803,16.30712890625,16.1935558319092,16.12744140625,16.2815437316895,16.3606433868408,16.4342765808105,16.4207038879395,16.4035148620605,16.2376956939697,15.8841791152954,15.5570316314697,15.4229497909546,15.4837894439697,15.3749027252197,15.1533193588257,15.0400390625,14.9179677963257,14.5432624816895,14.6099605560303,14.6351556777954,14.6345701217651,14.5958003997803,14.5495119094849,14.4796867370605,14.42626953125,14.3524417877197,14.1151361465454,13.9248046875,13.6502933502197,13.87353515625,14.07763671875,14.1199216842651,14.1480474472046,14.1412105560303,14.0632810592651,14.0027351379395,13.9605474472046,13.6707029342651,13.2996091842651,13.2035150527954,12.9875974655151,12.7927732467651,12.6900386810303,12.6625003814697,12.53271484375,12.3019533157349,12.1751956939697,12.2121095657349,11.9999027252197,12.138671875,12.1446285247803,12.2181644439697,12.1410160064697,12.1085939407349,12.1196298599243,12.1398429870605,12.1218748092651,12.1145515441895,12.3035163879395,12.3013668060303,12.2919921875,12.2336912155151,12.1553707122803,12.2920904159546,12.48681640625,12.5960941314697,12.7575197219849,12.8807621002197,12.8636713027954,12.8282222747803,12.7763662338257,12.7278318405151,12.7060546875,12.6830081939697,12.4675779342651,12.3537111282349,12.2941408157349,12.3146486282349,12.4453125,12.5538082122803,12.5886716842651,12.5528326034546,12.5158205032349,12.5146484375,12.4861326217651,12.4020509719849,12.2919921875,12.1692390441895,12.0718755722046,11.98828125,11.93212890625,11.8812494277954,11.8342771530151,11.6807613372803,11.6848630905151,11.7433595657349,11.7981443405151,11.7518548965454,11.7122068405151,11.6427736282349,11.5435552597046,11.470703125,11.3882808685303,11.3864259719849,11.3659181594849,11.1321287155151,11.0908203125,10.9989261627197,10.9450197219849,10.83447265625,10.7425775527954,10.6449222564697,10.6310548782349,10.6343746185303,10.6044921875,10.5953121185303,10.5338869094849,10.5695314407349,10.4937496185303,10.3981437683105,10.4071292877197,10.4463863372803,10.45458984375,10.4313478469849,10.2431640625,10.2051763534546,10.1793947219849,10.0831050872803,9.95956993103027,9.84257793426514,9.80019569396973,9.63515663146973,9.55722713470459,9.62714862823486,9.69609355926514,9.65693378448486,9.61845684051514,9.55107402801514,9.30996131896973,9.39580154418945,9.32294940948486,9.23867225646973,9.19375038146973,9.17812442779541,8.92841815948486,8.52138710021973,8.31220722198486,8.16611289978027,8.03740215301514,7.87558650970459,7.46591758728027,7.19414091110229,7.0048828125,6.90341758728027,6.89023447036743,6.89531278610229,6.91230535507202,6.87705087661743,6.80283212661743,6.77109384536743,6.76679754257202,6.7314453125,6.59052705764771,6.55507850646973,6.60576152801514,6.69248056411743,6.6767578125,6.65986299514771,6.61757802963257,6.49150419235229,6.38906288146973,6.0546875,5.9765625,5.70683622360229,5.5859375,5.51728487014771,5.5224609375,5.55556678771973,5.61220693588257,5.85429716110229,6.09902334213257,6.13730430603027,6.21415996551514,6.36328125,6.32109355926514,6.09941387176514,6.01699209213257,5.88915967941284,5.88915967941284,5.94872999191284,5.96855497360229,5.93730497360229,5.95185518264771,6.05068349838257,6.19892549514771,6.3056640625,6.41533184051514,6.40390634536743,6.27851581573486,6.15859365463257,6.01738262176514,5.84521436691284,5.71796894073486,5.65732383728027,5.56406259536743,5.46757793426514,5.3623046875,5.17324209213257,5.13164043426514,5.18505811691284,5.2421875,5.30488300323486,5.40351581573486,5.47246074676514,5.52968740463257,5.57949209213257,5.77216815948486,5.86728477478027,5.99101591110229,6.21660184860229,6.2119140625,6.05927705764771,5.96669960021973,5.83398485183716,5.76347637176514,5.73046875,5.78359365463257,5.99648475646973,6.06992149353027,6.11181640625,6.10517549514771,6.14052724838257,6.34873056411743,6.51806640625,6.57363319396973,6.5263671875,6.52685594558716,6.66093730926514,6.71992158889771,6.787109375,6.94970703125,6.99570322036743,6.80634784698486,6.34697246551514,6.1533203125,6.10175752639771,5.96738290786743,5.90439414978027,5.87656259536743,5.80087900161743,5.69882774353027,5.55703115463257,5.49453115463257,5.35341835021973,5.26386690139771,5.23447275161743,5.18603563308716,5.14580106735229,5.11074256896973,5.10488271713257,5.11923837661743,5.18710947036743,5.21953153610229,5.17441368103027,5.20566415786743,5.26542949676514,5.37646436691284,5.49453115463257,5.68857431411743,5.65761709213257,5.57382774353027,5.41738271713257,5.28584003448486,5.18359422683716,5.13710927963257,5.16816425323486,5.54648399353027,5.64834022521973,5.58935546875,5.44736337661743,5.24404335021973,5.11582040786743,5.04912137985229,5.0107421875,5.02460956573486,5.00859355926514,5.09540987014771,5.19248056411743,5.28818368911743,5.50527381896973,5.98398399353027,6.29257822036743,6.41796875,6.60986328125,6.77783250808716,6.90341758728027,6.97207021713257,6.98056650161743,7.03867149353027,7.07792997360229,7.04667949676514,7.04013681411743,7.54501962661743,7.60449266433716,7.40390634536743,7.34658241271973,7.45253896713257,7.44257783889771,7.33115291595459,7.27626991271973,7.29804706573486,7.27597665786743,7.17353487014771,6.94257831573486,6.79433584213257,6.65703105926514,6.61025428771973,6.62587881088257,6.59990262985229,6.54306650161743,6.49257850646973,6.38349580764771,6.08251953125,5.64677715301514,5.45126962661743,5.32460927963257,5.10673856735229,5.02167940139771,4.98994159698486,5.00273466110229,5.17246103286743,5.25830030441284,5.33867168426514,5.267578125,5.16757822036743,5.09941434860229,4.99667930603027,4.92783164978027,4.91035127639771,4.93007850646973,4.98505878448486,5.11699199676514,5.46533203125,5.79326152801514,6.01582002639771,6.46669864654541,6.73076152801514,6.68232393264771,6.39589881896973,6.13115262985229,5.66445350646973,5.47304677963257,5.26689481735229,5.15957021713257,5.09648418426514,5.14316415786743,5.24091768264771,5.29384756088257,5.35771465301514,5.42236280441284,5.48427724838257,5.53330087661743,5.71816396713257,5.79628944396973,5.90830039978027,5.97978544235229,6.02558565139771,6.08349609375,6.208984375,6.580078125,6.62001943588257,6.69238233566284,6.45712900161743,6.26171875,6.13613271713257,6.11845684051514,6.16474628448486,6.23749971389771,6.27285146713257,6.35292959213257,6.439453125,6.61835956573486,6.74462890625,6.96113300323486,7.28378868103027,7.49179697036743,7.57011699676514,7.65312528610229,7.69072198867798,7.52744150161743,7.51816415786743,7.53837919235229,7.8046875,8.09550857543945,8.04550743103027,7.40839862823486,7.24208974838257,7.11083984375,7.02490234375,6.77998065948486,6.73496055603027,6.78154277801514,6.92822265625,6.9404296875,7.00849580764771,7.38906240463257,7.57187509536743,7.654296875,7.73603439331055,7.8603515625,8.10058689117432,8.21113300323486,8.310546875,8.62314414978027,8.60917949676514,8.33857440948486,8.23515605926514,8.15800762176514,8.18447303771973,8.271484375,8.58017539978027,8.63554763793945,8.64101600646973,8.59375,8.48017597198486,8.38652420043945,8.36074256896973,8.39814472198486,8.57617282867432,8.67363262176514,8.84238243103027,9.13583946228027,9.15810585021973,9.07587814331055,9.08417987823486,9.15605449676514,9.32363224029541,9.52070331573486,9.60224628448486,9.69687461853027,9.83222675323486,9.89150428771973,9.93603515625,9.97919940948486,10.02099609375,10.08056640625,10.1885747909546,10.3400392532349,10.5909175872803,10.7044916152954,10.7601566314697,10.7067384719849,10.6736326217651,10.7252931594849,10.7791996002197,10.9525394439697,11.1178712844849,11.2257814407349,11.3707036972046,11.3479490280151,11.3076171875,11.2138671875,11.1755867004395,11.2946290969849,11.4576168060303,11.4292001724243,11.306640625,11.2135744094849,11.0751953125,10.9142570495605,10.9666996002197,11.0472660064697,10.9348630905151,10.3391599655151,10.0550785064697,9.92402362823486,9.89277362823486,9.83232498168945,9.76748085021973,9.6572265625,9.59462928771973,9.56728458404541,9.61474609375,9.7080078125,9.86445331573486,9.93945217132568,10.0099611282349,10.2362298965454,10.5656251907349,10.833984375,10.9323244094849,11.0904293060303,11.2253913879395,11.3313474655151,11.5238275527954,11.6329097747803,11.5617189407349,11.3924808502197,11.2967777252197,11.3035154342651,11.3499031066895,11.4893560409546,12.15966796875,12.2265615463257,12.3065423965454,12.5083990097046,12.7383785247803,12.91552734375,12.8198232650757,12.71533203125,12.51171875,12.4175777435303,12.3638677597046,12.333984375,12.2633790969849,12.1996088027954,12.1338863372803,12.1221675872803,12.2062501907349,12.2728519439697,12.3448238372803,12.6277341842651,12.6888666152954,12.8167972564697,12.9830083847046,13.0331058502197,12.97607421875,12.794921875,12.7837896347046,13.3871097564697,13.6744146347046,13.7596683502197,13.9158201217651,14.0341796875,13.97314453125,13.6813478469849,13.4989252090454,13.4164056777954,13.35205078125,13.1188478469849,13.0681638717651,13.1046867370605,13.1916007995605,13.21142578125,13.3118171691895,13.4503898620605,13.5202140808105,13.62109375,13.7879886627197,13.95947265625,13.9169931411743,13.7041015625,13.6515626907349,13.7266597747803,13.8083992004395,13.8801755905151,14.0223627090454,14.1087884902954,14.20556640625,14.34033203125,14.4726572036743,14.6006832122803,14.7755861282349,15.4157228469849,15.4347658157349,15.3000984191895,14.8244152069092,14.5815420150757,14.4792957305908,14.4416990280151,14.4483394622803,14.5366220474243,14.5785160064697,14.7549810409546,14.9619131088257,15.120509147644,15.2891597747803,15.4093742370605,15.4653310775757,15.5529298782349,15.5944337844849,15.5756826400757,15.6915044784546,15.6613283157349,15.4873056411743,15.35400390625,15.2487306594849,15.2186517715454,15.2840814590454,15.3458013534546,15.3039064407349,15.0408210754395,14.8546876907349,14.7813472747803,14.8210935592651,14.7989263534546,15.0484371185303,15.1342763900757,15.2744131088257,15.4008779525757,15.5066394805908,15.6213874816895,15.6057615280151,15.3569345474243,15.2928705215454,15.3160161972046,15.4868173599243,15.6566410064697,15.8512687683105,16.0079116821289,16.0380859375,16.0645503997803,16.1208000183105,16.2607421875,16.3123054504395,16.2585945129395,16.3086929321289,16.3721675872803,16.3919906616211,16.3192386627197,16.259765625,16.1748046875,16.2038097381592,16.3878917694092,16.6188468933105,16.8649406433105,16.9513664245605,17.0940418243408,17.3361339569092,17.478515625,17.5528316497803,17.5711917877197,17.5023441314697,17.4801750183105,17.4261722564697,17.2023429870605,16.5848636627197,16.5252933502197,16.5143566131592,16.5798835754395,16.65185546875,16.8846683502197,17.1311531066895,17.3908195495605,17.4900398254395,17.5462894439697,17.70458984375,18.1014652252197,18.1174812316895,18.0753898620605,18.0787124633789,18.1875,18.259765625,18.2931632995605,18.3787097930908,18.4826164245605,18.6455078125,18.8589839935303,18.9159183502197,18.75,18.6244144439697,18.6144523620605,18.6740245819092,18.7666015625,18.8828125,18.9911136627197,19.0068359375,19.0113277435303,19.0383796691895,19.197265625,19.68701171875,19.7224617004395,19.6959953308105,19.6397457122803,19.6415023803711,19.73681640625,19.8646488189697,19.9605464935303,20.0689449310303,20.1463871002197,20.2230453491211,20.32421875,20.3551769256592,20.38720703125,20.3327140808105,20.3381843566895,20.2771472930908,20.0437507629395,20.0544929504395,20.1072273254395,20.1976566314697,20.4867191314697,20.7394523620605,20.7425785064697,20.6615238189697,20.5625,20.53271484375,20.5459957122803,20.6220703125,20.84033203125,20.9710941314697,21.0321292877197,21.1630859375,21.2537117004395,21.4329090118408,21.5902347564697,21.7795906066895,21.9317378997803,21.9747085571289,21.892578125,21.802734375,21.6078128814697,21.400390625,21.3462886810303,21.3557605743408,21.5387687683105,21.7802734375,21.9955081939697,22.0543937683105,22.2194328308105,22.3219738006592,22.384765625,22.4211902618408,22.6845703125,22.8516597747803,22.9412097930908,22.9828128814697,23.0464839935303,23.1769523620605,23.2579116821289,23.3539066314697,23.4001960754395,23.3102531433105,23.2860336303711,23.3291015625,23.37939453125,23.6612300872803,23.8971672058105,24.0384769439697,24.2855472564697,24.3555660247803,24.4200191497803,24.4035167694092,24.2681636810303,24.2634773254395,24.4417953491211,24.6580066680908,24.7647457122803,24.8316402435303,25.0421867370605,25.1711921691895,25.2646484375,25.3253898620605,25.3755855560303,25.4359378814697,25.56982421875,25.6497058868408,25.7119140625,25.7681636810303,25.7814445495605,25.6656246185303,25.46826171875,25.2735347747803,25.2092761993408,25.1463871002197,24.9942398071289,24.9827136993408,25.0438480377197,25.2118167877197,25.4188480377197,25.4705085754395,25.9880867004395,26.2308597564697,26.5069351196289,26.6613273620605,26.7339839935303,26.6754875183105,26.5582027435303,26.6446285247803,26.6281242370605,26.6011714935303,26.583984375,26.5850582122803,26.6661128997803,26.9893550872803,27.0712890625,27.1472663879395,27.1836910247803,27.3093738555908,27.5464839935303,27.5556640625,27.26904296875,27.2352523803711,27.3316402435303,27.5970706939697,27.7334957122803,27.8150386810303,28.1416988372803,28.3922843933105,28.3827152252197,28.3268547058105,28.2718734741211,27.9509773254395,27.8980464935303,27.9988288879395,28.2156257629395,28.2717761993408,28.2027339935303,28.1910152435303,28.166015625,28.166015625,28.1929683685303,28.2800788879395,28.3098640441895,28.4373054504395,28.4847660064697,28.609375,28.7498035430908,28.83154296875,29.1023426055908,29.2185554504395,29.3210945129395,29.3976573944092,29.6390628814697,29.7219715118408,29.7374992370605,29.7964859008789,29.9593753814697,30.0651378631592,30.2376956939697,30.2030277252197,30.2131843566895,30.4220695495605,30.5958976745605,30.9263648986816,30.9606456756592,30.9441413879395,30.4689445495605,30.2629871368408,29.9258804321289,28.7811527252197,28.8042984008789,29.6013679504395,29.6468734741211,29.6213855743408,29.6209964752197,29.6359386444092,29.6946296691895,29.7920894622803,29.9903335571289,30.0882816314697,30.1551761627197,30.1800785064697,30.2375984191895,30.3488292694092,30.3972644805908,30.4283218383789,30.4843730926514,30.5945320129395,30.7144527435303,30.8697261810303],[25.5863265991211,25.853515625,25.9450206756592,26.07763671875,26.1468753814697,26.1337890625,25.9997081756592,25.7913074493408,25.7601566314697,25.58203125,25.4820327758789,25.31494140625,25.3152332305908,25.4234371185303,25.5863265991211],[-8.95356464385986,-9.04580116271973,-9.098876953125,-8.96464824676514,-8.52080059051514,-8.34370136260986,-8.00136661529541,-7.97880840301514,-8.00209999084473,-8.30234336853027,-8.63535213470459,-8.95356464385986],[19.2193355560303,19.0985355377197,18.9175777435303,18.7974605560303,18.8612308502197,19.1829109191895,19.2615222930908,19.2747058868408,19.2193355560303],[21.6081066131592,21.74560546875,22.0431632995605,22.2073249816895,22.2995128631592,22.4493179321289,22.7345695495605,22.9888668060303,23.1192378997803,23.3516597747803,23.4519538879395,23.3646488189697,23.1519527435303,23.11669921875,23.33056640625,23.6839847564697,23.8830089569092,24.23828125,24.5714836120605,24.90185546875,24.1297836303711,24.0619144439697,23.9549808502197,23.8412113189697,23.7361335754395,23.5051765441895,23.380859375,23.1013679504395,22.9966793060303,22.8995113372803,22.8017578125,22.5537109375,22.4269542694092,22.4688472747803,22.4866218566895,22.4424800872803,22.6789054870605,22.7326183319092,22.6853523254395,22.6203136444092,22.4482440948486,22.3972663879395,22.2545909881592,22.0568351745605,21.8561534881592,21.0499019622803,20.9281253814697,20.8731460571289,21.2010746002197,21.25146484375,21.3341789245605,21.4308586120605,21.6083984375,21.6531257629395,21.21044921875,21.0354499816895,20.8449211120605,20.7864265441895,20.5283203125,20.5602550506592,20.3727531433105,20.2279300689697,20.3626956939697,21.046875,21.4547843933105,21.6081066131592],[26.8759765625,26.7294921875,26.4595699310303,26.40771484375,26.4557609558105,26.5859375,26.7887706756592,27.0076160430908,26.8759765625],[11.2502927780151,11.26171875,11.4242191314697,11.6163082122803,11.82568359375,11.8848638534546,11.9293937683105,12.05615234375,12.1164064407349,11.9650392532349,11.7565422058105,11.5865230560303,11.3724613189697,11.19921875,11.1212892532349,10.8406248092651,10.7888669967651,10.62841796875,10.5576171875,10.5582027435303,10.7728519439697,10.9608402252197,11.1239261627197,11.1529293060303,11.0782232284546,11.1549806594849,11.2502927780151],[29.0470695495605,29.3454093933105,29.6451187133789,29.6966781616211,29.3105487823486,28.8811511993408,28.4945316314697,28.0378913879395,27.8890628814697,28.1209945678711,28.3740234375,28.4147472381592,28.5111331939697,28.84521484375,29.0470695495605],[16.7867183685303,16.8384780883789,16.8889636993408,16.9255867004395,16.9664058685303,17.2194328308105,17.5782222747803,17.6847648620605,17.8345699310303,17.9561519622803,17.8597640991211,17.7326164245605,17.6875,17.7339859008789,17.7150402069092,17.6687507629395,17.8610343933105,18.2720699310303,18.3332996368408,18.3973636627197,18.5814456939697,18.7484378814697,18.7852535247803,18.8152351379395,18.8324203491211,18.8229503631592,18.8074226379395,18.7200202941895,18.6778316497803,18.7722663879395,18.8800792694092,18.97900390625,19.0894527435303,19.490234375,19.7508792877197,19.8935546875,20.11376953125,20.1144523620605,20.1626949310303,20.4582023620605,20.6110343933105,20.7671871185303,20.5006828308105,20.7203121185303,21.0896492004395,21.3122062683105,21.3525390625,21.3887691497803,21.2439460754395,21.0962886810303,20.7248058319092,20.3870124816895,19.7687492370605,19.6767578125,19.6549816131592,19.6185531616211,19.3806629180908,19.1504878997803,19.0556640625,18.9837894439697,18.9576168060303,19.0086917877197,18.9951171875,18.8220691680908,18.7123050689697,18.5746097564697,18.4392585754395,18.4306640625,18.4386711120605,18.4040050506592,18.3619136810303,18.2987308502197,18.2279281616211,18.1374015808105,17.8470706939697,17.6233386993408,17.4424800872803,17.3486328125,17.1525402069092,17.1878910064697,17.2490234375,17.1419925689697,16.9766597747803,16.9798831939697,17.0355472564697,17.0626945495605,16.9351577758789,16.7004890441895,16.4619140625,16.3458003997803,16.2380847930908,16.1238269805908,16.0044918060303,15.5467777252197,15.1242189407349,14.7384757995605,14.4869146347046,14.3658208847046,14.24755859375,14.1453123092651,14.0503902435303,14.0041999816895,13.9957027435303,14.0260744094849,14.0712881088257,14.3776369094849,14.48779296875,14.5962886810303,14.6950187683105,14.9208011627197,16.2059574127197,16.619140625,17.0333003997803,16.96875,16.9140625,16.8529300689697,16.5396480560303,16.06005859375,15.8263683319092,15.5853509902954,15.3448238372803,15.0968742370605,14.8468742370605,14.6039056777954,14.0899419784546,13.9624996185303,13.7911128997803,13.7496089935303,13.7141599655151,13.6805667877197,13.7176761627197,13.7701168060303,13.8240232467651,13.9369134902954,14.0477542877197,14.3072261810303,14.2481441497803,14.9947261810303,15.3414058685303,15.5194330215454,15.6980476379395,15.6586923599243,15.6571292877197,15.7838859558105,15.875391960144,16.1502933502197,16.7769527435303,17.0029296875,17.1719741821289,16.9917964935303,16.8112297058105,16.7265625,16.5353527069092,16.4486331939697,16.6965827941895,16.7826156616211,16.5304679870605,16.4462890625,16.1575202941895,15.9440431594849,15.6806650161743,15.4173831939697,15.3599615097046,15.2793941497803,15.2541999816895,15.2649421691895,15.3483400344849,15.3916006088257,15.3841800689697,15.32275390625,15.2252922058105,15.1373043060303,15.0163087844849,14.8917961120605,14.8386716842651,14.7923831939697,14.7435541152954,14.6892576217651,14.57763671875,14.4671878814697,14.5052728652954,14.5154294967651,14.4677734375,14.4318361282349,14.5456047058105,14.6382808685303,14.49951171875,14.36328125,14.23828125,14.1104497909546,13.9076175689697,13.6549806594849,13.1501960754395,12.9127931594849,12.8695316314697,12.8221683502197,12.6646490097046,12.4347658157349,12.2579097747803,12.1382808685303,11.9617185592651,11.8655271530151,11.7738275527954,11.7462892532349,11.7551755905151,11.8610353469849,11.6110343933105,11.3654298782349,11.4561529159546,11.5475587844849,12.27490234375,12.3234367370605,12.4033203125,12.375,12.2531242370605,12.0873050689697,12.0458002090454,11.9818363189697,11.9256839752197,11.9019527435303,11.8927736282349,12.01611328125,12.083984375,11.9783201217651,11.6792964935303,11.5797853469849,11.6164064407349,11.5211915969849,11.3388671875,11.2081060409546,11.1072263717651,10.9753904342651,10.92578125,10.8880853652954,10.8343753814697,10.7375974655151,10.7249994277954,10.7370119094849,10.8107423782349,10.7545900344849,10.6862306594849,10.68212890625,10.7463865280151,10.8040037155151,10.8659172058105,11.0496091842651,11.150390625,11.1852540969849,11.2505855560303,11.3436527252197,11.7023439407349,12.1017580032349,12.2051753997803,12.2877931594849,12.2452154159546,12.2191410064697,12.2799797058105,12.6024417877197,12.7535152435303,13.1075201034546,13.6928701400757,13.9141597747803,13.9256830215454,13.9210939407349,13.9070310592651,13.7775392532349,13.0392580032349,12.5553703308105,13.2151374816895,13.3337898254395,13.3835945129395,13.431640625,13.6012697219849,13.7162113189697,13.8336906433105,13.9572267532349,14.0295906066895,14.0558595657349,14.0263671875,14.0111322402954,14.0198240280151,14.0398435592651,14.1784181594849,14.3797855377197,14.5936517715454,14.8318357467651,15.0523433685303,15.2512693405151,15.4439458847046,15.6601572036743,15.7640628814697,15.8584957122803,16.2945308685303,16.34375,16.2535152435303,16.0275382995605,15.8750982284546,15.8407230377197,15.8161125183105,15.8257808685303,15.8451166152954,15.9557628631592,16.1001949310303,16.056640625,16.0938491821289,16.2457046508789,16.3866214752197,16.5240230560303,16.7867183685303],[32.5259742736816,31.5776386260986,31.48193359375,33.0191383361816,33.0986328125,33.3839836120605,33.6292953491211,33.556640625,32.5259742736816],[18.7416019439697,18.5249996185303,18.1622066497803,18.20556640625,18.2916984558105,18.5193347930908,18.7416019439697],[20.8978519439697,20.9984378814697,21.5492191314697,21.6548824310303,21.6966800689697,21.7806644439697,21.8977546691895,22.1902351379395,22.2897472381592,22.3763675689697,22.4426765441895,22.4461898803711,22.4178714752197,22.4507808685303,22.548828125,22.6720695495605,22.7925777435303,22.8968753814697,23.0080070495605,23.2513675689697,23.3154296875,23.2500972747803,23.224609375,23.1145496368408,23.3533191680908,23.68798828125,23.77294921875,23.9529304504395,24.1429691314697,24.2340812683105,24.2801742553711,24.2975597381592,24.4026374816895,24.5466804504395,24.6136722564697,24.736328125,24.7859363555908,24.9070301055908,25.4712886810303,25.6668949127197,25.7511730194092,25.8363285064697,26.4367179870605,26.86083984375,27.0171871185303,27.1483402252197,27.1986331939697,27.0798835754395,26.2210941314697,26.005859375,25.9020519256592,25.7263660430908,25.6412105560303,25.2390613555908,25.1451168060303,24.8428707122803,24.7505855560303,24.3833980560303,24.2568359375,24.1329097747803,23.94775390625,23.7587890625,22.9037113189697,22.7891597747803,22.6957035064697,22.8655261993408,21.9114265441895,20.8611316680908,20.8055667877197,20.7608394622803,20.3995132446289,20.1282234191895,19.9001941680908,19.6746101379395,19.7466793060303,19.8210945129395,20.0148429870605,20.1871089935303,20.4934558868408,20.5648441314697,20.6868152618408,20.7840805053711,20.4607410430908,20.1234378814697,19.8986320495605,19.6380863189697,19.3999996185303,18.9420909881592,18.7250003814697,18.4280281066895,18.32470703125,18.2847652435303,18.25537109375,18.5946273803711,18.7264652252197,18.85595703125,18.3438472747803,18.1294918060303,17.9168949127197,18.0894546508789,18.779296875,18.9619140625,19.1429672241211,19.3433589935303,19.537109375,19.3546867370605,19.19140625,19.1569347381592,19.1783199310303,19.2637691497803,19.3274402618408,19.5684566497803,19.7510757446289,19.80224609375,19.8103504180908,19.7771492004395,19.6913089752197,19.6143550872803,19.7333011627197,19.8511714935303,20.1042957305908,20.359375,20.4758796691895,20.6934566497803,20.8978519439697],[88.1097717285156,88.1502914428711,88.154296875,88.14697265625,88.1055679321289,88.06787109375,88.0241165161133,87.984375,87.9931640625,88.1110305786133,88.1572265625,88.1615219116211,88.1115264892578,88.0548782348633,88.0269546508789,87.9951171875,87.8492202758789,87.7488250732422,87.6333999633789,87.5130844116211,87.41357421875,87.2873992919922,87.1668014526367,87.0895462036133,87.0378952026367,87.0164031982422,86.7624969482422,86.7013702392578,86.5436553955078,86.4144515991211,86.3661193847656,86.2416000366211,86.12939453125,86.00732421875,85.8556594848633,85.7945327758789,85.7373046875,85.7074203491211,85.6998977661133,85.6484375,85.5684585571289,85.4564437866211,85.29296875,85.240234375,85.1917953491211,85.1741180419922,85.1515655517578,85.1253890991211,85.0873107910156,85.0201187133789,84.9372100830078,84.6853485107422,84.65380859375,84.65478515625,84.6407241821289,84.6101531982422,84.4808578491211,84.2297821044922,84.0910110473633,84.0248031616211,83.8971633911133,83.8288116455078,83.7469711303711,83.5516586303711,83.4471664428711,83.3839874267578,83.3694305419922,83.2897415161133,83.2138671875,83.0640640258789,82.9328155517578,82.7333984375,82.7108383178711,82.6773452758789,82.6298828125,82.4513626098633,82.2876968383789,82.1119155883789,82.0370101928711,81.9876937866211,81.9452133178711,81.8968811035156,81.8526382446289,81.7572250366211,81.6355514526367,81.4860382080078,81.3108444213867,81.2389678955078,81.2062530517578,81.1689453125,81.0166015625,80.8960876464844,80.7507781982422,80.7261734008789,80.6712875366211,80.5870132446289,80.5178680419922,80.4958038330078,80.4791030883789,80.4185485839844,80.3248062133789,80.2265625,80.1496047973633,80.0707015991211,80.0516586303711,80.0845718383789,80.1304702758789,80.1695327758789,80.2330093383789,80.2559585571289,80.2548828125,80.31689453125,80.40185546875,80.5490188598633,80.6128921508789,80.68408203125,80.8199234008789,80.84814453125,80.9076232910156,80.9661178588867,81.01025390625,81.0555725097656,81.1103515625,81.1771469116211,81.2550811767578,81.4171905517578,81.6418914794922,81.8548889160156,82.0433578491211,82.0989303588867,82.1353530883789,82.1589889526367,82.220703125,82.4865188598633,82.6408233642578,82.8543014526367,83.0139617919922,83.1554718017578,83.2351608276367,83.3551788330078,83.4566421508789,83.58349609375,83.6710968017578,83.7904281616211,83.9359359741211,84.02197265625,84.1013717651367,84.1278305053711,84.1755828857422,84.2287139892578,84.3121109008789,84.4107437133789,84.4654312133789,84.6505813598633,84.6767578125,84.7142562866211,84.7593765258789,84.796875,84.8550796508789,85.0691375732422,85.1263656616211,85.1590881347656,85.16015625,85.1214828491211,85.0885772705078,85.1224594116211,85.2121047973633,85.41064453125,85.6783218383789,85.7594757080078,85.8402328491211,85.9216842651367,85.9541015625,85.9945373535156,86.0641632080078,86.0754928588867,86.0787124633789,86.1370086669922,86.1742248535156,86.2179718017578,86.32861328125,86.40869140625,86.4849624633789,86.5168914794922,86.5544967651367,86.6144561767578,86.6905288696289,86.7196273803711,86.7503890991211,86.8423843383789,86.9337844848633,87.0201110839844,87.1414031982422,87.2907257080078,87.4641571044922,87.5552749633789,87.62255859375,87.6827163696289,87.8607406616211,87.9333953857422,88.0233383178711,88.1097717285156],[166.958404541016,166.93896484375,166.916412353516,166.907043457031,166.91357421875,166.93896484375,166.955673217773,166.958404541016],[169.17822265625,169.233489990234,169.127532958984,169.075973510742,169.039840698242,169.021774291992,169.0791015625,169.128616333008,169.17822265625],[166.221099853516,166.242874145508,166.187881469727,166.0732421875,166.037689208984,166.013275146484,165.971389770508,165.904113769531,165.88916015625,165.915618896484,166.073822021484,166.103118896484,166.101364135742,166.22509765625,166.254287719727,166.267486572266,166.259368896484,166.209564208984,166.207611083984,166.220413208008,166.179489135742,166.200775146484,166.221099853516],[168.144927978516,168.145309448242,168.041015625,168.043151855469,168.12548828125,168.155944824219,168.241409301758,168.260650634766,168.240921020508,168.183883666992,168.015045166016,167.905563354492,167.810745239258,167.784957885742,167.676361083984,167.554885864258,167.52197265625,167.538772583008,167.628997802734,167.630950927734,167.654098510742,167.740921020508,167.741989135742,167.80078125,167.765243530273,167.783981323242,167.955764770508,168.144927978516],[166.746276855469,166.741012573242,166.729187011719,166.694534301758,166.642471313477,166.591705322266,166.559173583984,166.532028198242,166.567092895508,166.685638427734,166.7314453125,166.746276855469],[166.9794921875,167.022659301758,166.93115234375,166.892684936523,166.962692260742,166.9794921875],[-176.176467895508,-176.220809936523,-176.214599609375,-176.229309082031,-176.154693603516,-176.12255859375,-176.176467895508],[-176.177642822266,-176.213531494141,-176.274841308594,-176.381744384766,-176.375244140625,-176.407363891602,-176.499130249023,-176.516555786133,-176.454925537109,-176.441253662109,-176.500152587891,-176.439117431641,-176.385452270508,-176.333587646484,-176.333831787109,-176.452789306641,-176.515533447266,-176.571533203125,-176.597991943359,-176.629348754883,-176.631546020508,-176.562744140625,-176.523773193359,-176.555130004883,-176.634567260742,-176.807952880859,-176.84765625,-176.761093139648,-176.667236328125,-176.566116333008,-176.177642822266],[173.914642333984,173.780853271484,173.786224365234,173.812393188477,173.873336791992,173.9033203125,173.964447021484,173.9580078125,173.914642333984],[173.115341186523,173.230865478516,173.337799072266,173.447265625,173.5625,173.737899780273,173.783798217773,173.897552490234,173.947174072266,174.00244140625,173.952835083008,173.889846801758,173.8798828125,173.915145874023,173.8603515625,173.862396240234,173.7978515625,173.897064208984,173.933410644531,173.914642333984,173.957626342773,174.024032592773,173.99755859375,173.999404907227,174.08056640625,174.121185302734,174.153228759766,174.211822509766,174.223815917969,174.302536010742,174.27392578125,174.213668823242,174.199508666992,174.103118896484,174.03857421875,174.138092041016,174.283584594727,174.3701171875,174.367568969727,174.297256469727,174.237121582031,174.169540405273,174.10205078125,174.0693359375,174.072937011719,174.092376708984,174.1611328125,174.083694458008,174.169921875,174.217086791992,174.283111572266,174.243362426758,174.215438842773,174.047256469727,173.973922729492,173.887985229492,173.88916015625,173.83984375,173.589263916016,173.545120239258,173.347564697266,173.22119140625,173.148834228516,173.072357177734,172.888870239258,172.808013916016,172.718551635742,172.6240234375,172.626953125,172.6875,172.734756469727,172.69970703125,172.632232666016,172.562194824219,172.526657104492,172.693450927734,172.740432739258,172.749206542969,172.7666015625,172.807037353516,172.947265625,173.0732421875,173.098037719727,173.116897583008,173.093948364258,173.065628051758,173.023330688477,172.920608520508,172.817672729492,172.749328613281,172.5546875,172.502731323242,172.475982666016,172.583786010742,172.527252197266,172.480377197266,172.4296875,172.395599365234,172.38525390625,172.350387573242,172.296569824219,172.220703125,172.145812988281,172.035537719727,172.05224609375,172.13720703125,172.179779052734,172.080749511719,171.977630615234,171.890625,171.808395385742,171.712005615234,171.658981323242,171.517776489258,171.442581176758,171.41748046875,171.364562988281,171.24072265625,171.285354614258,171.31298828125,171.231048583984,171.213088989258,171.197860717773,171.146286010742,170.9990234375,171.022857666016,171.134170532227,171.11328125,170.99072265625,170.939651489258,170.889938354492,170.815231323242,170.700576782227,170.69970703125,170.739837646484,170.788467407227,170.791213989258,170.776275634766,170.721786499023,170.674209594727,170.419128417969,170.33544921875,170.266799926758,170.186126708984,169.918258666992,169.7607421875,169.729110717773,169.686630249023,169.34228515625,169.0986328125,168.9658203125,168.837799072266,168.766799926758,168.631439208984,168.572265625,168.466415405273,168.382125854492,168.357223510742,168.32568359375,168.343078613281,168.319732666016,168.266204833984,168.230270385742,168.189163208008,168.077346801758,167.900390625,167.841995239258,167.722076416016,167.682220458984,167.539459228516,167.490615844727,167.414260864258,167.368942260742,167.100296020508,166.830764770508,166.731536865234,166.712097167969,166.916702270508,166.8564453125,166.730270385742,166.64990234375,166.726959228516,166.733779907227,166.717971801758,166.612701416016,166.4931640625,166.477630615234,166.48828125,166.512893676758,166.836029052734,166.952545166016,167.003311157227,166.809967041016,166.797653198242,166.825576782227,166.990814208984,166.869049072266,166.733993530273,166.743072509766,166.7783203125,166.919921875,166.875595092773,166.869232177734,166.908599853516,167.052154541016,167.155670166016,167.112106323242,167.117782592773,167.145309448242,167.230072021484,167.206848144531,167.127334594727,167.032806396484,167.022659301758,167.02587890625,167.1279296875,167.188186645508,167.259475708008,167.205078125,167.171875,167.194534301758,167.410736083984,167.466217041016,167.479202270508,167.482131958008,167.456253051758,167.4599609375,167.484970092773,167.57763671875,167.698135375977,167.787017822266,167.859375,167.908981323242,167.901565551758,167.866409301758,167.856536865234,168.018371582031,168.196197509766,168.366607666016,168.457427978516,168.650970458984,168.774810791016,168.806442260742,168.990417480469,169.066497802734,169.135940551758,169.178909301758,169.1357421875,169.169525146484,169.323150634766,169.515228271484,169.661529541016,169.769241333008,169.833877563477,169.824020385742,169.835052490234,169.890823364258,169.908004760742,169.858993530273,170.017578125,170.103713989258,170.148818969727,170.189651489258,170.240234375,170.300003051758,170.355758666992,170.396087646484,170.374313354492,170.302825927734,170.379486083984,170.458694458008,170.535842895508,170.61181640625,170.535842895508,170.523635864258,170.615524291992,170.665435791016,170.735244750977,170.725296020508,170.741607666016,170.84033203125,170.969924926758,171.011428833008,171.017776489258,171.01171875,171.038375854492,171.047546386719,171.027725219727,171.189559936523,171.221282958984,171.257034301758,171.313385009766,171.296096801758,171.252243041992,171.296493530273,171.322662353516,171.360260009766,171.420608520508,171.486221313477,171.536315917969,171.672180175781,171.731643676758,171.830657958984,171.948043823242,172.0107421875,172.093368530273,172.139450073242,172.272750854492,172.468170166016,172.640625,172.711120605469,172.830184936523,172.943649291992,172.732620239258,172.711120605469,172.704391479492,172.728897094727,172.766799926758,172.869140625,172.988662719727,173.042282104492,173.052139282227,173.068649291992,173.115341186523],[175.543167114258,175.551177978516,175.474609375,175.444625854492,175.358779907227,175.34619140625,175.336624145508,175.381637573242,175.389541625977,175.409378051758,175.4443359375,175.512603759766,175.543167114258],[173.269424438477,173.284576416016,173.339935302734,173.381256103516,173.447845458984,173.438659667969,173.47265625,173.693740844727,173.7392578125,173.786224365234,173.812789916992,173.843948364258,173.923828125,174.10400390625,174.118942260742,174.109756469727,174.118743896484,174.143157958984,174.203216552734,174.282897949219,174.3203125,174.373336791992,174.393173217773,174.384963989258,174.419143676758,174.464752197266,174.54345703125,174.53173828125,174.508590698242,174.580657958984,174.533493041992,174.391006469727,174.395812988281,174.478713989258,174.548736572266,174.604873657227,174.802154541016,174.7724609375,174.777053833008,174.751754760742,174.819244384766,174.777145385742,174.749221801758,174.718658447266,174.722457885742,174.801956176758,174.849899291992,174.891403198242,174.917190551758,174.952041625977,175.047073364258,175.2451171875,175.299514770508,175.326461791992,175.3466796875,175.385345458984,175.4609375,175.54248046875,175.568161010742,175.551956176758,175.4931640625,175.492874145508,175.501266479492,175.487411499023,175.4580078125,175.426376342773,175.385543823242,175.399810791016,175.460845947266,175.497650146484,175.528030395508,175.681442260742,175.772171020508,175.780670166016,175.842178344727,175.876174926758,175.921096801758,175.990142822266,176.114562988281,176.129013061523,176.053314208984,176.029891967773,176.037887573242,176.1083984375,176.191116333008,176.2431640625,176.291702270508,176.61474609375,176.77001953125,177.161819458008,177.274017333984,177.3359375,177.453308105469,177.558288574219,177.64892578125,177.727340698242,177.812698364258,177.909469604492,177.9580078125,178.009185791016,178.272171020508,178.360748291016,178.475967407227,178.536224365234,178.516006469727,178.447067260742,178.393936157227,178.347259521484,178.3154296875,178.267684936523,178.1806640625,178.084854125977,177.976165771484,177.93212890625,177.910354614258,177.916610717773,177.951370239258,177.965621948242,177.908798217773,177.87548828125,177.828720092773,177.7861328125,177.655853271484,177.52294921875,177.407531738281,177.296585083008,177.128723144531,177.076751708984,177.03125,176.9541015625,176.935729980469,176.939254760742,176.966598510742,177.10986328125,176.967971801758,176.842193603516,176.770706176758,176.688766479492,176.611526489258,176.476455688477,176.385147094727,176.313858032227,176.251754760742,176.11865234375,176.059951782227,175.98291015625,175.839660644531,175.687301635742,175.447067260742,175.380264282227,175.309768676758,175.22216796875,175.204483032227,175.184661865234,175.165618896484,175.053909301758,174.906051635742,174.88134765625,174.875198364258,174.875,174.900192260742,174.865631103516,174.83154296875,174.819732666016,174.841217041016,174.757034301758,174.669540405273,174.642959594727,174.635360717773,174.656539916992,174.684860229492,174.847747802734,175.016799926758,175.162490844727,175.200485229492,175.254104614258,175.210159301758,175.155960083008,175.00927734375,174.813766479492,174.687301635742,174.567474365234,174.454681396484,174.35205078125,174.148635864258,173.934371948242,173.812118530273,173.783004760742,173.763671875,173.766403198242,173.781631469727,173.806045532227,173.844345092773,174.071380615234,174.311721801758,174.356048583984,174.3984375,174.45849609375,174.566207885742,174.597366333008,174.618560791016,174.653030395508,174.71533203125,174.809280395508,174.840042114258,174.801651000977,174.836822509766,174.879592895508,174.928024291992,174.845993041992,174.749420166016,174.729202270508,174.743942260742,174.767669677734,174.707427978516,174.672546386719,174.585830688477,174.609680175781,174.65966796875,174.734283447266,174.746383666992,174.803604125977,174.863861083984,174.928909301758,174.782028198242,174.73291015625,174.66796875,174.601470947266,174.536529541016,174.4755859375,174.444534301758,174.406051635742,174.381927490234,174.188873291016,174.245712280273,174.401565551758,174.431732177734,174.454299926758,174.446868896484,174.409576416016,174.354095458984,174.353118896484,174.395416259766,174.392776489258,174.303512573242,174.267868041992,174.252044677734,174.277542114258,174.253707885742,174.036422729492,173.969329833984,173.914459228516,173.908889770508,173.917282104492,174.003128051758,174.142379760742,174.166412353516,174.145797729492,174.097457885742,174.0546875,173.991012573242,173.945114135742,173.412200927734,173.480270385742,173.585845947266,173.6103515625,173.626174926758,173.581634521484,173.541702270508,173.49609375,173.454299926758,173.401657104492,173.376358032227,173.31396484375,173.290237426758,173.291213989258,173.274505615234,173.228118896484,173.16015625,173.11669921875,173.188781738281,173.190628051758,173.117294311523,173.029586791992,172.860733032227,172.705947875977,172.873733520508,173.0439453125,172.963760375977,172.999801635742,173.054397583008,173.171096801758,173.181243896484,173.240524291992,173.269424438477],[-171.186431884766,-171.188629150391,-171.193023681641,-171.200042724609,-171.204437255859,-171.20166015625,-171.194427490234,-171.189300537109,-171.186431884766],[-172.479141235352,-172.483703613281,-172.488235473633,-172.494033813477,-172.498687744141,-172.497009277344,-172.487258911133,-172.481094360352,-172.479141235352],[58.7220726013184,58.6590843200684,58.6409187316895,58.6412124633789,58.7879905700684,58.8843727111816,58.9507827758789,58.8351554870605,58.7722702026367,58.7220726013184],[56.3879890441895,56.4898452758789,56.640625,56.7741203308105,56.9124984741211,57.123046875,57.2198257446289,57.611328125,57.8250999450684,58.1204071044922,58.3245086669922,58.3931655883789,58.4999961853027,58.5780258178711,58.7730484008789,58.8303718566895,58.9115219116211,58.9833946228027,59.0298843383789,59.1947288513184,59.3109397888184,59.4293937683105,59.5351600646973,59.6956062316895,59.8232421875,59.8374977111816,59.8244094848633,59.7999992370605,59.6808624267578,59.6525344848633,59.517578125,59.3714790344238,59.3044891357422,59.0687522888184,58.8957023620605,58.6904296875,58.5341835021973,58.4742202758789,58.3487281799316,58.2660140991211,58.208984375,58.2316398620605,58.2450180053711,58.1694374084473,58.1029319763184,57.9471664428711,57.86181640625,57.8436508178711,57.8021507263184,57.7412109375,57.7141609191895,57.7151412963867,57.7608413696289,57.7639694213867,57.7903327941895,57.8116226196289,57.7384796142578,57.6757850646973,57.4279289245605,57.1765632629395,56.9572257995605,56.8259735107422,56.6550788879395,56.55078125,56.3834953308105,56.2703132629395,55.9976577758789,55.6138648986816,55.4791030883789,55.25537109375,55.2381858825684,55.2814445495605,55.2956047058105,55.2751960754395,55.1737327575684,55.0641593933105,54.7718734741211,54.6646461486816,54.5665054321289,54.376953125,54.0681648254395,53.9543914794922,53.775390625,53.60986328125,53.2977561950684,53.0856437683105,53.0249977111816,52.96435546875,52.9037132263184,52.8429679870605,52.8005867004395,52.7292022705078,52.6859397888184,52.6417007446289,52.5973625183105,52.5531234741211,52.5088844299316,52.4645500183105,52.4203109741211,52.3760719299316,52.3317375183105,52.2874984741211,52.2432632446289,52.1990242004395,52.1546897888184,52.1104507446289,52.0662117004395,52.0218772888184,51.9776382446289,52.1185531616211,52.2906227111816,52.4626960754395,52.6346664428711,52.8067359924316,52.9787101745605,53.1507797241211,53.3228530883789,53.4948234558105,53.6668930053711,53.8388671875,54.0109367370605,54.1830062866211,54.35498046875,54.5270500183105,54.6990242004395,54.87109375,54.9773445129395,55.021484375,55.0582008361816,55.0947265625,55.1314468383789,55.1680679321289,55.20458984375,55.2412109375,55.2779312133789,55.314453125,55.35107421875,55.3876953125,55.42431640625,55.4609375,55.49755859375,55.5341796875,55.57080078125,55.607421875,55.6410140991211,55.5777359008789,55.4927749633789,55.40380859375,55.3201179504395,55.25927734375,55.1858367919922,55.1940422058105,55.1921882629395,55.1999015808105,55.2702140808105,55.3532218933105,55.4138679504395,55.46630859375,55.5084953308105,55.5316390991211,55.5193367004395,55.4917984008789,55.4684562683105,55.5478515625,55.6965827941895,55.7791023254395,55.8941383361816,55.9851531982422,55.9921875,55.9663124084473,55.9286155700684,55.7997093200684,55.7608413696289,55.8056640625,55.8040046691895,55.7868156433105,55.7681617736816,55.7775382995605,55.8039054870605,55.80419921875,55.7915992736816,55.7957038879395,55.8228530883789,55.8707008361816,55.9158210754395,55.9630851745605,56.0005836486816,56.0167007446289,56.00634765625,55.9796867370605,55.9703140258789,56.0083999633789,56.0638656616211,56.1065444946289,56.1544914245605,56.2046890258789,56.2678718566895,56.3135757446289,56.3529281616211,56.3879890441895],[56.2105484008789,56.2165031433105,56.2342758178711,56.27734375,56.2877922058105,56.2818336486816,56.240234375,56.2105484008789],[56.2978515625,56.2785148620605,56.24951171875,56.18359375,56.1446304321289,56.1519508361816,56.1541023254395,56.1725578308105,56.16748046875,56.1165046691895,56.0804710388184,56.1644554138184,56.197265625,56.2284202575684,56.3055648803711,56.3464851379395,56.3787117004395,56.4130859375,56.4297866821289,56.4177742004395,56.4164047241211,56.3736305236816,56.3292961120605,56.3072242736816,56.2978515625],[76.7668914794922,76.8127899169922,76.8822250366211,76.927734375,76.9789123535156,77.0044937133789,77.0486297607422,77.0306625366211,77.0008850097656,76.8917007446289,76.7829132080078,76.7575225830078,76.7490234375,76.6962890625,76.5944290161133,76.5099639892578,76.4567413330078,76.1724624633789,76.041015625,75.9382858276367,75.8621139526367,75.7091751098633,75.6055603027344,75.4525375366211,75.2640609741211,75.1875,75.1184539794922,74.9518508911133,74.7887649536133,74.5941390991211,74.4979553222656,74.3003921508789,74.1719741821289,74.0558624267578,73.9612274169922,73.8831024169922,73.85009765625,73.8121109008789,73.7945327758789,73.8099594116211,73.9246063232422,73.9723663330078,73.9794921875,73.9382781982422,73.9039001464844,73.9040985107422,73.92236328125,73.9499053955078,74.1126022338867,74.208984375,74.2464828491211,74.2508773803711,74.2156295776367,74.0784149169922,74.0009765625,73.9764709472656,73.9775390625,74.0040054321289,74.0697250366211,74.1312561035156,74.1500015258789,74.142578125,74.1177673339844,74.0503921508789,73.9942398071289,73.9898376464844,74.0038070678711,74.0491180419922,74.1262664794922,74.2220687866211,74.2835922241211,74.3036117553711,74.32275390625,74.3299789428711,74.3054733276367,74.3545837402344,74.4833984375,74.5882797241211,74.6324234008789,74.6632843017578,74.6433639526367,74.6578140258789,74.6857452392578,74.7888717651367,74.9873046875,75.1041030883789,75.2336959838867,75.3026351928711,75.33349609375,75.32470703125,75.2541046142578,75.1387710571289,75.0714797973633,74.7394561767578,74.6357421875,74.5555648803711,74.5259780883789,74.5099563598633,74.5818328857422,74.5939407348633,74.5349578857422,74.5176773071289,74.5397491455078,74.6103515625,74.6257781982422,74.6328125,74.509765625,74.38037109375,74.33935546875,74.2156295776367,74.0089874267578,73.8993148803711,73.8916015625,73.8827133178711,73.9246063232422,73.9333953857422,73.8865203857422,73.8091812133789,73.6580123901367,73.4674835205078,73.3816375732422,73.3172836303711,73.2578125,73.2311477661133,73.1283187866211,72.94873046875,72.9033203125,72.6255874633789,72.3418960571289,72.2919921875,72.23388671875,72.17919921875,72.1285171508789,71.9480438232422,71.8888702392578,71.8703079223633,71.7166976928711,71.54296875,71.2901306152344,71.1847610473633,70.8749008178711,70.7979507446289,70.7374038696289,70.6916046142578,70.6491241455078,70.6291046142578,70.5692443847656,70.4885787963867,70.4037094116211,70.3184585571289,70.2443389892578,70.1939392089844,70.14453125,70.0498046875,69.8962936401367,69.7248077392578,69.6613311767578,69.62158203125,69.5679702758789,69.5370178222656,69.4945297241211,69.4700164794922,69.4812469482422,69.5069351196289,69.6005859375,69.7359390258789,69.9114227294922,70.0593719482422,70.1146545410156,70.1476516723633,70.1568374633789,70.1492156982422,70.1326141357422,70.0777359008789,70.07861328125,70.1001892089844,70.2646484375,70.3251953125,70.4485397338867,70.505859375,70.5695266723633,70.6148452758789,70.6484375,70.6572265625,70.6520538330078,70.7025375366211,70.8004913330078,70.8777313232422,70.9508743286133,71.0207061767578,71.0478515625,71.0023422241211,70.9763717651367,70.9698257446289,70.9793014526367,70.9732437133789,71.0062484741211,71.0453109741211,71.0440444946289,70.9828109741211,70.9281234741211,70.88623046875,70.8050842285156,70.7672805786133,70.71630859375,70.6594772338867,70.5792922973633,70.5558624267578,70.5650405883789,70.5467758178711,70.4892578125,70.2890625,70.0982437133789,70.0651397705078,70.0210876464844,69.9337844848633,69.80517578125,69.7162094116211,69.6341781616211,69.5591812133789,69.4434585571289,69.2350616455078,69.1195373535156,69.0515594482422,68.9845657348633,68.9007797241211,68.8634796142578,68.8283233642578,68.7999954223633,68.7811508178711,68.7589797973633,68.7396469116211,68.7281265258789,68.72412109375,68.5866241455078,68.4886703491211,68.3812484741211,68.2825241088867,68.2341766357422,68.1650390625,68.1488265991211,68.1155319213867,68.0677719116211,68.0370101928711,68.00146484375,67.9509735107422,67.8599548339844,67.8190460205078,67.66845703125,67.6495132446289,67.6458053588867,67.5630874633789,67.5036163330078,67.4768524169922,67.4539031982422,67.4276351928711,67.365234375,67.3093795776367,67.3042984008789,67.2886734008789,67.1714782714844,67.1005859375,66.7030258178711,66.6822280883789,66.7098617553711,66.6986312866211,66.5699234008789,66.5338821411133,66.4286193847656,66.32421875,66.2190475463867,66.1623077392578,66.1311569213867,66.3564453125,66.4071273803711,66.4676818847656,66.4029312133789,66.3283233642578,66.2346649169922,65.8835983276367,65.6796875,65.40625,65.0613327026367,64.7766571044922,64.6589813232422,64.5940399169922,64.5437545776367,64.1520538330078,64.1249008178711,64.0593719482422,63.9873008728027,63.9355506896973,63.7208976745605,63.5566368103027,63.4957046508789,63.4914016723633,63.2857398986816,63.1700210571289,63.0150375366211,62.6647453308105,62.5724639892578,62.4447250366211,62.3912086486816,62.3153343200684,62.2487335205078,62.1986312866211,62.1521453857422,62.0894508361816,61.9079132080078,61.74365234375,61.56689453125,61.587890625,61.6154289245605,61.64013671875,61.6713905334473,61.6618156433105,61.6686515808105,61.7376976013184,61.7543983459473,61.78076171875,61.8099594116211,61.8423843383789,61.8698272705078,62.0890617370605,62.1259727478027,62.2393531799316,62.2496070861816,62.2596664428711,62.3123092651367,62.3850555419922,62.4392585754395,62.6364288330078,62.7515640258789,62.7866249084473,63.0929718017578,63.1578102111816,63.1680679321289,63.1861343383789,63.2416038513184,63.2503929138184,63.2314453125,63.2420883178711,63.3051795959473,63.3015632629395,63.2562446594238,63.1960945129395,63.1667976379395,62.9154281616211,62.8116226196289,62.7629890441895,62.7527313232422,62.7625007629395,62.7642555236816,62.8008766174316,62.81201171875,62.7823257446289,62.73974609375,62.7625007629395,62.7580032348633,62.7494125366211,62.7175750732422,62.5645523071289,62.4338836669922,62.35302734375,62.1305618286133,62.0330085754395,61.8898468017578,61.7580108642578,61.623046875,61.5687522888184,61.5085906982422,61.337890625,61.3394546508789,61.318359375,61.1521453857422,61.0341796875,60.8433609008789,61.2244148254395,61.5214805603027,62.0009727478027,62.3734359741211,62.4765586853027,63.5675811767578,63.9709968566895,64.0987319946289,64.1179733276367,64.1721649169922,64.26611328125,64.3937530517578,64.5210952758789,64.7035140991211,64.8273391723633,64.9189453125,65.0955047607422,65.1804656982422,65.4709930419922,65.6662139892578,65.9616241455078,66.1770553588867,66.2312545776367,66.2869110107422,66.3133773803711,66.2471694946289,66.2384796142578,66.2818374633789,66.3054656982422,66.3009719848633,66.2869110107422,66.3468704223633,66.3971710205078,66.4973602294922,66.5667953491211,66.5958023071289,66.6242218017578,66.7313461303711,66.8292922973633,66.92431640625,67.0277328491211,67.1159210205078,67.2873001098633,67.4528274536133,67.5963897705078,67.6615219116211,67.7378921508789,67.7334976196289,67.6470642089844,67.5975570678711,67.5782165527344,67.6267547607422,67.7398452758789,68.0171890258789,68.1301727294922,68.1610336303711,68.2139663696289,68.31982421875,68.4432601928711,68.5207061767578,68.59765625,68.6732406616211,68.7136688232422,68.7823257446289,68.8689498901367,68.9734344482422,69.0831069946289,69.1869125366211,69.279296875,69.2565383911133,69.2414016723633,69.2899398803711,69.3594741821289,69.4053726196289,69.4045944213867,69.453125,69.5015563964844,69.5677719116211,69.7037124633789,69.9201202392578,70.0902328491211,70.2611312866211,70.2841796875,70.2197265625,70.1341781616211,70.056640625,69.8680648803711,69.8896484375,69.9947280883789,70.2536087036133,70.32568359375,70.4157257080078,70.6540069580078,70.8484420776367,71.0515594482422,71.09130859375,71.0890579223633,71.0923843383789,71.095703125,71.02294921875,70.9789123535156,70.9656219482422,71.0163116455078,71.0656280517578,71.11328125,71.2257766723633,71.2941436767578,71.3581085205078,71.455078125,71.51708984375,71.5455093383789,71.6016616821289,71.6205062866211,71.6052703857422,71.5772476196289,71.5455093383789,71.5455093383789,71.5719757080078,71.6005859375,71.58740234375,71.5719757080078,71.51904296875,71.4835891723633,71.4275360107422,71.3975601196289,71.3428726196289,71.22021484375,71.18505859375,71.23291015625,71.3125991821289,71.4632797241211,71.5458984375,71.6205062866211,71.7164077758789,71.7726593017578,71.822265625,71.9207000732422,72.0956039428711,72.15673828125,72.2498016357422,72.3269577026367,72.43115234375,72.5313491821289,72.6228561401367,72.7662124633789,72.9937438964844,73.1167984008789,73.4111328125,73.7318344116211,73.7691421508789,73.9078140258789,74.0018539428711,74.0388717651367,74.1947250366211,74.4310607910156,74.5414047241211,74.6005859375,74.6921920776367,74.7660140991211,74.8412094116211,74.8892593383789,74.9491195678711,75.0539093017578,75.1452178955078,75.3466796875,75.3768539428711,75.4242248535156,75.4602508544922,75.57373046875,75.6671829223633,75.7721633911133,75.8402328491211,75.8849563598633,75.9330062866211,75.9518508911133,75.9744110107422,75.9686508178711,75.93408203125,75.9048843383789,75.9123077392578,75.9451217651367,76.0104522705078,76.0708999633789,76.1033172607422,76.1478500366211,76.1778335571289,76.2516555786133,76.3857421875,76.5020523071289,76.55126953125,76.5634765625,76.6318359375,76.7275390625,76.7668914794922],[-81.603271484375,-81.6581115722656,-81.7701187133789,-81.85205078125,-81.8585968017578,-81.85693359375,-81.8121566772461,-81.7522964477539,-81.728759765625,-81.6714324951172,-81.71044921875,-81.6947250366211,-81.6134262084961,-81.603271484375],[-79.0654296875,-79.1103515625,-79.1275329589844,-79.0962905883789,-79.0853042602539,-79.0654296875],[-78.8983459472656,-78.9181137084961,-78.9649429321289,-78.9574203491211,-78.9605941772461,-78.916015625,-78.8832550048828,-78.8561553955078,-78.8391571044922,-78.8532257080078,-78.8983459472656],[-82.2334976196289,-82.2444305419922,-82.3217315673828,-82.2757797241211,-82.2594223022461,-82.2334976196289],[-77.3742218017578,-77.39306640625,-77.4483413696289,-77.478515625,-77.4072799682617,-77.3858871459961,-77.3455047607422,-77.2826156616211,-77.2123031616211,-77.1959915161133,-77.2159652709961,-77.282958984375,-77.3456115722656,-77.3627471923828,-77.3507766723633,-77.5382843017578,-77.5865707397461,-77.6186065673828,-77.6585922241211,-77.7063446044922,-77.7320327758789,-77.7469253540039,-77.7619171142578,-77.7687530517578,-77.743896484375,-77.7646942138672,-77.8283157348633,-77.9011688232422,-77.9297866821289,-78.1701126098633,-78.3782272338867,-78.4215850830078,-78.3676300048828,-78.3154296875,-78.287353515625,-78.2548828125,-78.2812042236328,-78.1800308227539,-78.1418914794922,-78.1138687133789,-78.0477523803711,-77.95166015625,-77.8336410522461,-77.7605514526367,-77.8529281616211,-78.0125045776367,-78.0571746826172,-78.0994644165039,-78.1618194580078,-78.1907730102539,-78.2230453491211,-78.2511215209961,-78.256103515625,-78.3501434326172,-78.3743133544922,-78.3992233276367,-78.3878860473633,-78.369384765625,-78.3792953491211,-78.4098587036133,-78.43603515625,-78.4694366455078,-78.5140609741211,-78.6209030151367,-78.6698226928711,-78.710205078125,-78.7696762084961,-78.8482437133789,-78.9551773071289,-79.0863800048828,-79.2466812133789,-79.4415054321289,-79.507080078125,-79.5516586303711,-79.5723648071289,-79.6874465942383,-79.7310485839844,-79.758544921875,-79.81591796875,-79.7504348754883,-80.1257858276367,-80.2000961303711,-80.3686981201172,-80.4075698852539,-80.458984375,-80.4658737182617,-80.4581069946289,-80.4091339111328,-80.3655776977539,-80.2609405517578,-80.0751953125,-80.0400390625,-80.01123046875,-80.0672836303711,-80.110595703125,-80.2873001098633,-80.3482437133789,-80.3729476928711,-80.4388656616211,-80.6667022705078,-80.8455581665039,-80.9012222290039,-80.9146499633789,-81.0351028442383,-81.0638656616211,-81.0939483642578,-81.1578140258789,-81.1793899536133,-81.1954574584961,-81.2190399169922,-81.2684020996094,-81.3695831298828,-81.504150390625,-81.6756820678711,-81.6942901611328,-81.7276382446289,-81.8602523803711,-81.9732971191406,-82.0967254638672,-82.1598587036133,-82.2243194580078,-82.2354507446289,-82.3648376464844,-82.5309600830078,-82.6795425415039,-82.7811584472656,-82.8661193847656,-82.8543472290039,-82.8793487548828,-82.88330078125,-82.9128875732422,-82.9484405517578,-83.0233840942383,-83.02734375,-82.99755859375,-82.8616180419922,-82.8447799682617,-82.8426284790039,-82.855712890625,-82.9170379638672,-82.8819808959961,-82.8119125366211,-82.739990234375,-82.7278289794922,-82.7411651611328,-82.7830505371094,-82.88134765625,-82.9403305053711,-82.9428253173828,-82.9398422241211,-82.925048828125,-82.8889617919922,-82.8601608276367,-82.843994140625,-82.801025390625,-82.723388671875,-82.6440963745117,-82.6112823486328,-82.5865249633789,-82.5692443847656,-82.5635757446289,-82.5003356933594,-82.3707962036133,-82.3631820678711,-82.3753967285156,-82.3397445678711,-82.2724609375,-82.2048873901367,-82.1881408691406,-82.20068359375,-82.2354507446289,-82.2441940307617,-82.13330078125,-82.077880859375,-81.8941421508789,-81.826416015625,-81.7802276611328,-81.8314895629883,-81.900146484375,-81.8944778442383,-81.8423767089844,-81.8025894165039,-81.7122116088867,-81.5456085205078,-81.3547897338867,-81.2037582397461,-81.0630798339844,-80.8386688232422,-80.6764678955078,-80.546875,-80.1270980834961,-79.9779739379883,-79.9150848388672,-79.8550796508789,-79.7230987548828,-79.6522445678711,-79.5773010253906,-79.35546875,-79.2116241455078,-79.1122589111328,-79.0166931152344,-78.9749984741211,-78.931640625,-78.6969223022461,-78.5043411254883,-78.082763671875,-77.830810546875,-77.6972198486328,-77.3742218017578],[-128.290084838867,-128.299987792969,-128.320648193359,-128.342193603516,-128.350189208984,-128.330123901367,-128.303619384766,-128.290817260742,-128.290084838867],[-69.9659194946289,-69.9720230102539,-70.0039520263672,-70.0533218383789,-70.1288070678711,-70.1839828491211,-70.2391586303711,-70.31689453125,-70.3436508178711,-70.4046401977539,-70.5306625366211,-70.6345748901367,-70.7215805053711,-70.7995147705078,-70.8660125732422,-70.9156265258789,-70.9736862182617,-71.1442413330078,-71.2350082397461,-71.3167953491211,-71.4382781982422,-71.5213317871094,-71.6683654785156,-71.8447265625,-71.9431610107422,-71.982421875,-72.08251953125,-72.2567901611328,-72.3528289794922,-72.468994140625,-72.6083526611328,-72.69873046875,-72.8319320678711,-72.8870620727539,-72.907470703125,-72.8958053588867,-72.9182586669922,-72.9589309692383,-72.97021484375,-72.9798812866211,-73.0680694580078,-73.1628875732422,-73.2093734741211,-73.2355499267578,-73.2064971923828,-73.167724609375,-73.1353530883789,-73.1263198852539,-73.1373519897461,-73.1774444580078,-73.2403335571289,-73.3254852294922,-73.4999084472656,-73.6945343017578,-73.7581100463867,-73.7762680053711,-73.8046417236328,-73.7930145263672,-73.7582015991211,-73.7233428955078,-73.7204132080078,-73.7494659423828,-73.8046417236328,-73.85400390625,-73.8917465209961,-73.929443359375,-73.9643096923828,-73.9643096923828,-73.9526824951172,-73.95849609375,-73.9817428588867,-74.0020523071289,-73.9817428588867,-73.9468688964844,-73.8946304321289,-73.8220748901367,-73.7668991088867,-73.7204132080078,-73.714599609375,-73.7320251464844,-73.772705078125,-73.7755813598633,-73.7204132080078,-73.6826705932617,-73.6449203491211,-73.610107421875,-73.610107421875,-73.5723648071289,-73.5491256713867,-73.5491256713867,-73.4881362915039,-73.4358901977539,-73.3981399536133,-73.3603973388672,-73.3517074584961,-73.3567352294922,-73.3024444580078,-73.203125,-73.12255859375,-73.0705108642578,-72.9740219116211,-72.9703598022461,-73.08984375,-73.2094268798828,-73.0137710571289,-72.8142547607422,-72.60546875,-72.4647521972656,-72.3790512084961,-72.3180618286133,-72.2890167236328,-72.2658157348633,-72.2599563598633,-72.1728515625,-72.1791000366211,-72.1815872192383,-72.1429748535156,-71.887451171875,-71.6080093383789,-71.3394088745117,-71.2379379272461,-71.1152801513672,-71.041748046875,-70.9707565307617,-70.884521484375,-70.8162612915039,-70.7584991455078,-70.6724624633789,-70.6369171142578,-70.60791015625,-70.5411148071289,-70.5701599121094,-70.5922393798828,-70.5991744995117,-70.5672302246094,-70.5937957763672,-70.6369171142578,-70.6375961303711,-70.6385269165039,-70.6393508911133,-70.6403274536133,-70.6415557861328,-70.642333984375,-70.5965347290039,-70.5332565307617,-70.4508819580078,-70.3922882080078,-70.3419952392578,-70.2903747558594,-70.2200698852539,-70.0663070678711,-69.9603500366211,-69.8397979736328,-69.6740188598633,-69.57861328125,-69.45361328125,-69.3620147705078,-69.2577133178711,-69.1737289428711,-69.0461883544922,-68.93603515625,-68.8186950683594,-68.6852569580078,-68.7281265258789,-68.7628860473633,-68.7590866088867,-68.8118133544922,-68.86767578125,-68.9337387084961,-68.9786148071289,-68.9805221557617,-68.9722671508789,-68.9834518432617,-69.0175323486328,-69.0528335571289,-69.0741195678711,-69.0230484008789,-68.9742660522461,-68.9374542236328,-68.8917007446289,-68.8709030151367,-68.8803176879883,-68.9717788696289,-69.0044937133789,-69.0131301879883,-69.0527801513672,-69.1197280883789,-69.1626968383789,-69.1992721557617,-69.2349090576172,-69.2523498535156,-69.2760238647461,-69.3594741821289,-69.3737335205078,-69.3747100830078,-69.3307113647461,-69.1871109008789,-69.1724624633789,-69.2543029785156,-69.3019027709961,-69.4185028076172,-69.4208984375,-69.3918914794922,-69.2175827026367,-69.18798828125,-69.1341781616211,-69.0462417602539,-68.9134750366211,-68.8488235473633,-68.8427734375,-68.8578109741211,-68.9280319213867,-69.0062484741211,-69.0329132080078,-69.0383758544922,-69.0206985473633,-69.0545349121094,-69.1325149536133,-69.1997985839844,-69.2672348022461,-69.3815383911133,-69.4210968017578,-69.4383316040039,-69.5033187866211,-69.6248550415039,-69.6457061767578,-69.6258773803711,-69.5638198852539,-69.5219192504883,-69.510986328125,-69.5110778808594,-69.5109405517578,-69.58642578125,-69.6847686767578,-69.8061065673828,-69.8520965576172,-69.8415069580078,-69.8024444580078,-69.8025894165039,-69.8396987915039,-69.9263687133789,-70.05908203125,-70.1837844848633,-70.2822799682617,-70.3774871826172,-70.4182586669922,-70.4916000366211,-70.8174743652344,-70.9416961669922,-71.0565872192383,-71.3369598388672,-71.3649444580078,-71.3994140625,-71.4358901977539,-71.5322265625,-71.7744598388672,-71.8683547973633,-71.9668960571289,-72.1112823486328,-72.2686080932617,-72.3625030517578,-72.4676818847656,-72.7939453125,-72.9577178955078,-73.2637710571289,-73.4000473022461,-73.7276840209961,-73.824951171875,-74.1470718383789,-74.3729019165039,-74.5548858642578,-75.104248046875,-75.1905288696289,-75.2745590209961,-75.3965835571289,-75.5336380004883,-75.7376937866211,-75.9338836669922,-76.0062942504883,-76.136474609375,-76.1751480102539,-76.2890090942383,-76.2970275878906,-76.37646484375,-76.3194808959961,-76.2592239379883,-76.1839370727539,-76.2236328125,-76.4273452758789,-76.5021514892578,-76.5552291870117,-76.6371078491211,-76.7580108642578,-76.8321304321289,-76.9940948486328,-77.0381317138672,-77.0626983642578,-77.1527328491211,-77.1576156616211,-77.2203140258789,-77.3099136352539,-77.6332015991211,-77.6385726928711,-77.664306640625,-77.736083984375,-78.095458984375,-78.1856002807617,-78.2755889892578,-78.3564987182617,-78.4456481933594,-78.5801239013672,-78.6645965576172,-78.7545852661133,-78.7622604370117,-78.9253921508789,-79.0122604370117,-79.1644058227539,-79.3128433227539,-79.3772506713867,-79.5888671875,-79.6177291870117,-79.761962890625,-79.9046859741211,-79.9949722290039,-80.1102600097656,-80.8116226196289,-81.0584411621094,-81.1420364379883,-81.1805191040039,-81.164306640625,-81.0918426513672,-80.9916534423828,-80.9306640625,-80.8827209472656,-80.8819351196289,-80.943115234375,-81.1676712036133,-81.1507339477539,-81.1084976196289,-81.195068359375,-81.2893981933594,-81.3366165161133,-81.283203125,-81.2320327758789,-80.8919448852539,-80.798583984375,-80.6527328491211,-80.503662109375,-80.3246536254883,-80.29833984375,-80.2735366821289,-80.2718734741211,-80.2652359008789,-80.2454147338867,-80.2437438964844,-80.2206039428711,-80.2189483642578,-80.21728515625,-80.2288589477539,-80.2175827026367,-80.1792526245117,-80.1941375732422,-80.2305145263672,-80.2668914794922,-80.3032684326172,-80.3578567504883,-80.4372024536133,-80.4901351928711,-80.510009765625,-80.4934616088867,-80.4884796142578,-80.4537582397461,-80.3528823852539,-80.44384765625,-80.4884796142578,-80.4785614013672,-80.4241714477539,-80.3834991455078,-80.2933578491211,-80.232177734375,-80.1974563598633,-80.1395492553711,-80.0635223388672,-79.962890625,-79.8451156616211,-79.7972640991211,-79.7109832763672,-79.6385269165039,-79.5776901245117,-79.5161590576172,-79.5019073486328,-79.4557647705078,-79.3994140625,-79.3309631347656,-79.2681121826172,-79.1866760253906,-79.0762710571289,-79.0333023071289,-78.9952621459961,-78.9753875732422,-78.919189453125,-78.9142074584961,-78.92578125,-78.9076156616211,-78.8615188598633,-78.7430648803711,-78.68603515625,-78.6744613647461,-78.6529235839844,-78.6612319946289,-78.6851577758789,-78.6793975830078,-78.6479949951172,-78.6033706665039,-78.5651397705078,-78.5504379272461,-78.5090866088867,-78.4934539794922,-78.4710464477539,-78.4197769165039,-78.4214401245117,-78.3999481201172,-78.3980484008789,-78.3472595214844,-78.3453674316406,-78.3230514526367,-78.2841796875,-78.250732421875,-78.2403869628906,-78.226318359375,-78.1948776245117,-78.1584930419922,-78.1609878540039,-78.1874465942383,-78.1946258544922,-78.1833038330078,-78.1282272338867,-78.0679244995117,-77.9384765625,-77.860595703125,-77.6589813232422,-77.5064926147461,-77.3600616455078,-77.1614761352539,-76.8807601928711,-76.6791000366211,-76.4993667602539,-76.3601531982422,-76.2409210205078,-76.0897979736328,-75.8854522705078,-75.7445297241211,-75.6416473388672,-75.570556640625,-75.5138702392578,-75.4491729736328,-75.42041015625,-75.4080581665039,-75.380126953125,-75.3481979370117,-75.3091812133789,-75.2724151611328,-75.2496109008789,-75.2835922241211,-75.2787094116211,-75.2593765258789,-75.2632369995117,-75.3252487182617,-75.4247055053711,-75.4659652709961,-75.4910583496094,-75.5605926513672,-75.6320266723633,-75.6262741088867,-75.583740234375,-75.4759750366211,-75.3983917236328,-75.3404769897461,-75.2844696044922,-75.224609375,-75.18408203125,-75.1383819580078,-75.0546875,-75.0049743652344,-74.9453125,-74.8888168334961,-74.8375015258789,-74.8017578125,-74.7804718017578,-74.75537109375,-74.691650390625,-74.6163635253906,-74.5550765991211,-74.5138702392578,-74.4651870727539,-74.4178695678711,-74.3749008178711,-74.3531265258789,-74.32861328125,-74.3344268798828,-74.2838897705078,-74.2463912963867,-74.1807556152344,-74.0543899536133,-73.98681640625,-73.9269561767578,-73.8631820678711,-73.8071746826172,-73.7357406616211,-73.664306640625,-73.6102523803711,-73.5754852294922,-73.5213851928711,-73.4943389892578,-73.5252456665039,-73.4962921142578,-73.4402847290039,-73.3495101928711,-73.2664566040039,-73.2239685058594,-73.1969757080078,-73.1814956665039,-73.1452102661133,-73.1265106201172,-73.1602020263672,-73.1726531982422,-73.1544952392578,-73.0681610107422,-72.9896469116211,-72.9411163330078,-72.8871536254883,-72.8112335205078,-72.7141571044922,-72.66015625,-72.6253433227539,-72.5867233276367,-72.5006790161133,-72.3956069946289,-72.3007354736328,-72.2184600830078,-72.1368179321289,-72.0538101196289,-71.9842758178711,-71.9324722290039,-71.8672866821289,-71.802734375,-71.7525405883789,-71.6714859008789,-71.5594787597656,-71.49609375,-71.4474639892578,-71.39697265625,-71.3001022338867,-71.1963882446289,-71.1133804321289,-71.0272979736328,-70.9685516357422,-70.91455078125,-70.7053756713867,-70.6480026245117,-70.5758819580078,-70.5167999267578,-70.418212890625,-70.3641586303711,-70.2946243286133,-70.2444305419922,-70.1647415161133,-70.0958480834961,-70.0647430419922,-70.064453125,-70.0740280151367,-70.1470642089844,-70.2901382446289,-70.4189910888672,-70.6216735839844,-70.735107421875,-70.7061996459961,-70.5296859741211,-70.48583984375,-70.4210968017578,-70.3791961669922,-70.3395004272461,-70.2984390258789,-70.2402877807617,-70.1983871459961,-70.1675338745117,-70.0947723388672,-70.0171813964844,-69.9659194946289],[120.250381469727,120.223243713379,120.191604614258,120.149993896484,120.118354797363,120.1005859375,120.013282775879,119.958106994629,119.877540588379,119.821487426758,119.827346801758,119.982711791992,120.079681396484,120.165237426758,120.2080078125,120.229385375977,120.250381469727],[121.159370422363,121.2138671875,121.282524108887,121.391502380371,121.414642333984,121.411041259766,121.29443359375,121.218170166016,121.083015441895,121.018547058105,120.930656433105,120.876373291016,120.898239135742,121.037696838379,121.159370422363],[122.092872619629,121.991401672363,121.959175109863,121.879875183105,121.872459411621,121.808692932129,121.832038879395,121.914939880371,122.058296203613,122.2880859375,122.323539733887,122.251754760742,122.200973510742,122.092872619629],[125.784568786621,125.768943786621,125.707511901855,125.683006286621,125.714454650879,125.783401489258,125.784568786621],[122.937110900879,122.948051452637,122.943649291992,122.839546203613,122.8046875,122.796585083008,122.822166442871,122.871192932129,122.914840698242,122.937110900879],[117.079879760742,117.028327941895,116.969528198242,116.975784301758,116.993560791016,117.077049255371,117.079879760742],[117.355270385742,117.287010192871,117.272270202637,117.280860900879,117.32958984375,117.353713989258,117.355270385742],[124.806640625,124.777923583984,124.665817260742,124.639068603516,124.653312683105,124.708106994629,124.736824035645,124.790229797363,124.806640625],[123.697662353516,123.706253051758,123.614456176758,123.540718078613,123.493453979492,123.493553161621,123.53515625,123.626068115234,123.654876708984,123.697662353516],[125.970504760742,125.952445983887,125.922065734863,125.948539733887,125.9677734375,125.992973327637,125.970504760742],[126.005950927734,126.087600708008,126.193359375,126.191993713379,126.209083557129,126.304595947266,126.319526672363,126.262985229492,126.22021484375,126.1416015625,126.139549255371,126.173049926758,126.282325744629,126.365333557129,126.379783630371,126.458686828613,126.456634521484,126.425300598145,126.435356140137,126.494430541992,126.54443359375,126.570114135742,126.593360900879,126.589263916016,126.58154296875,126.546684265137,126.439071655273,126.294052124023,126.216896057129,126.192085266113,126.240226745605,126.221183776855,126.189353942871,126.142471313477,126.109771728516,126.080070495605,126.043067932129,125.984954833984,125.961624145508,125.901176452637,125.824417114258,125.773628234863,125.689262390137,125.670219421387,125.660247802734,125.640731811523,125.542182922363,125.464744567871,125.400978088379,125.380661010742,125.432914733887,125.486625671387,125.564544677734,125.588478088379,125.670700073242,125.66796875,125.607810974121,125.455863952637,125.346481323242,125.287895202637,125.241012573242,125.233200073242,125.264945983887,125.268455505371,125.231544494629,125.191017150879,125.174018859863,125.076171875,125.035354614258,124.975196838379,124.927345275879,124.636322021484,124.398818969727,124.212799072266,124.078132629395,124.049705505371,124.048149108887,123.987892150879,123.980857849121,123.985252380371,124.045112609863,124.117584228516,124.158210754395,124.190719604492,124.212890625,124.206634521484,124.182418823242,124.067970275879,123.968452453613,123.764747619629,123.717391967773,123.665817260742,123.60888671875,123.553215026855,123.493072509766,123.477447509766,123.481636047363,123.476364135742,123.390922546387,123.282035827637,123.178230285645,123.150688171387,123.138778686523,123.121192932129,123.096672058105,123.048934936523,122.989547729492,122.916893005371,122.842971801758,122.818756103516,122.791801452637,122.713966369629,122.6162109375,122.497955322266,122.474418640137,122.448623657227,122.319725036621,122.25146484375,122.176170349121,122.142486572266,122.09814453125,122.027641296387,121.964256286621,121.904205322266,121.924613952637,121.991111755371,122.047164916992,122.114845275879,122.119918823242,122.131843566895,122.243362426758,122.337104797363,122.38671875,122.589454650879,122.672943115234,122.804397583008,122.911140441895,122.996284484863,123.002731323242,122.998825073242,123.017578125,123.050582885742,123.095893859863,123.147163391113,123.292861938477,123.341209411621,123.380180358887,123.434562683105,123.498931884766,123.563667297363,123.680084228516,123.783393859863,123.849227905273,123.860549926758,123.877449035645,123.853416442871,123.753120422363,123.799415588379,123.931144714355,123.996879577637,124.159370422363,124.197662353516,124.225784301758,124.283203125,124.3251953125,124.35791015625,124.404884338379,124.451263427734,124.621772766113,124.731155395508,124.761726379395,124.786819458008,124.806144714355,124.868942260742,124.94384765625,125.046394348145,125.141021728516,125.176170349121,125.209663391113,125.247856140137,125.375579833984,125.498725891113,125.533401489258,125.510162353516,125.413963317871,125.471282958984,125.520896911621,125.642478942871,125.876655578613,125.954681396484,126.005950927734],[126.059379577637,126.046783447266,125.9912109375,125.998634338379,126.073829650879,126.129493713379,126.12890625,126.120796203613,126.172554016113,126.136917114258,126.059379577637],[125.28076171875,125.287689208984,125.158981323242,125.133010864258,125.175880432129,125.23095703125,125.28076171875],[124.593849182129,124.584274291992,124.505661010742,124.477531433105,124.403411865234,124.35986328125,124.122459411621,123.935638427734,123.871681213379,123.829978942871,123.817192077637,123.863868713379,123.908889770508,124.059768676758,124.093841552734,124.1728515625,124.335739135742,124.351554870605,124.373237609863,124.405860900879,124.486328125,124.5771484375,124.555084228516,124.582229614258,124.593849182129],[125.690231323242,125.672554016113,125.648635864258,125.590530395508,125.534477233887,125.494819641113,125.521980285645,125.524612426758,125.580169677734,125.605850219727,125.647956848145,125.666793823242,125.684562683105,125.646682739258,125.703323364258,125.684371948242,125.692474365234,125.690231323242],[119.916213989258,119.793159484863,119.764457702637,119.85205078125,119.9501953125,120.008407592773,119.981155395508,119.916213989258],[124.316207885742,124.288475036621,124.334671020508,124.37109375,124.38232421875,124.38134765625,124.316207885742],[122.649513244629,122.621879577637,122.59716796875,122.538375854492,122.516693115234,122.537498474121,122.625785827637,122.6484375,122.672561645508,122.701271057129,122.729194641113,122.737197875977,122.681251525879,122.649513244629],[123.130851745605,123.064651489258,122.994728088379,122.947860717773,122.866607666016,122.7724609375,122.664558410645,122.6103515625,122.5625,122.410934448242,122.399520874023,122.425582885742,122.471481323242,122.523239135742,122.648246765137,122.712982177734,122.855567932129,122.865814208984,122.866500854492,122.852348327637,122.816993713379,122.855567932129,122.905868530273,122.958404541016,122.968742370605,122.9697265625,122.983306884766,123.0244140625,123.221778869629,123.256637573242,123.510635375977,123.562492370605,123.567573547363,123.527725219727,123.492866516113,123.406929016113,123.343559265137,123.296089172363,123.266212463379,123.186614990234,123.162010192871,123.162696838379,123.1494140625,123.149803161621,123.308395385742,123.321876525879,123.320510864258,123.293357849121,123.228713989258,123.192474365234,123.130851745605],[123.370307922363,123.331741333008,123.316017150879,123.327056884766,123.403701782227,123.38623046875,123.514350891113,123.592872619629,123.711418151855,123.726455688477,123.831550598145,123.929885864258,123.924606323242,123.950096130371,123.964065551758,123.967178344727,124.038871765137,124.057914733887,124.036521911621,124.039840698242,124.052528381348,124.053321838379,124.027534484863,124.051277160645,124.004981994629,123.952140808105,123.873924255371,123.788665771484,123.700492858887,123.643363952637,123.633979797363,123.493553161621,123.370307922363],[123.757034301758,123.815620422363,123.736717224121,123.707618713379,123.741401672363,123.757034301758],[117.311134338379,117.218559265137,117.228515625,117.255867004395,117.349906921387,117.417770385742,117.529975891113,117.59326171875,117.744918823242,117.884765625,117.931549072266,117.983009338379,118.023826599121,118.114845275879,118.343940734863,118.533401489258,118.7275390625,118.820114135742,118.845115661621,119.023834228516,119.079879760742,119.143058776855,119.18603515625,119.223831176758,119.287010192871,119.312698364258,119.296684265137,119.261123657227,119.305656433105,119.340728759766,119.465339660645,119.501266479492,119.553321838379,119.560249328613,119.534568786621,119.532615661621,119.561912536621,119.526657104492,119.616119384766,119.684379577637,119.686904907227,119.595222473145,119.540519714355,119.422454833984,119.369331359863,119.284759521484,119.23193359375,119.218559265137,119.191497802734,118.948631286621,118.834663391113,118.782127380371,118.754981994629,118.773834228516,118.569625854492,118.504493713379,118.434959411621,118.349609375,118.229293823242,118.134078979492,118.069435119629,117.989547729492,117.888572692871,117.77978515625,117.679885864258,117.572174072266,117.539649963379,117.5166015625,117.469146728516,117.412498474121,117.311134338379],[119.861434936523,119.882904052734,119.854888916016,119.830665588379,119.798637390137,119.72998046875,119.725578308105,119.761421203613,119.826759338379,119.861434936523],[124.574607849121,124.644340515137,124.724319458008,124.821090698242,124.929977416992,124.993949890137,125.026557922363,125.044326782227,125.039741516113,125.01318359375,125.033782958984,125.083892822266,125.127532958984,125.164161682129,125.187698364258,125.197158813477,125.260055541992,125.268455505371,125.253318786621,125.1484375,125.140037536621,125.142578125,125.105361938477,125.0439453125,124.987503051758,125.0048828125,125.023536682129,125.026557922363,124.929107666016,124.812789916992,124.780769348145,124.791694641113,124.78955078125,124.737693786621,124.798637390137,124.797172546387,124.786720275879,124.738662719727,124.662696838379,124.616119384766,124.502830505371,124.445510864258,124.411720275879,124.366012573242,124.330955505371,124.308197021484,124.330657958984,124.374122619629,124.435935974121,124.510932922363,124.548240661621,124.574607849121],[124.6083984375,124.483489990234,124.428810119629,124.3603515625,124.437400817871,124.510932922363,124.564949035645,124.622268676758,124.6083984375],[124.854393005371,124.8359375,124.806640625,124.781051635742,124.74365234375,124.730865478516,124.788375854492,124.821487426758,124.854393005371],[122.496192932129,122.612701416016,122.726272583008,122.838081359863,122.931243896484,122.900787353516,122.89453125,123.102737426758,123.158294677734,123.156455993652,123.144142150879,123.119529724121,123.075485229492,123.016502380371,122.938774108887,122.8466796875,122.802932739258,122.789451599121,122.791122436523,122.769920349121,122.673149108887,122.522064208984,122.197654724121,122.108589172363,122.051750183105,121.988380432129,121.953994750977,121.938278198242,121.933792114258,121.980072021484,121.972358703613,121.950294494629,121.96435546875,122.020698547363,122.050880432129,122.059677124023,122.103507995605,122.101364135742,122.066993713379,121.940818786621,121.891212463379,121.916023254395,121.963668823242,122.029197692871,122.086814880371,122.290725708008,122.399215698242,122.496192932129],[120.038772583008,119.963874816895,119.944923400879,119.931739807129,119.932807922363,119.860939025879,119.916015625,119.956535339355,119.997856140137,120.035942077637,120.070709228516,120.062400817871,120.073143005371,120.038772583008],[120.099998474121,120.154693603516,120.193748474121,120.228225708008,120.260551452637,120.341400146484,120.314544677734,120.243461608887,120.173629760742,120.100090026855,120.010551452637,119.95703125,119.896087646484,119.865913391113,119.86962890625,119.891799926758,119.880081176758,119.885749816895,119.896675109863,119.916404724121,119.963874816895,120.077537536621,120.099998474121],[122.654487609863,122.603317260742,122.499320983887,122.438873291016,122.422943115234,122.471878051758,122.603614807129,122.673629760742,122.683296203613,122.654487609863],[125.239547729492,125.310356140137,125.327537536621,125.320213317871,125.352249145508,125.408790588379,125.481246948242,125.535636901855,125.503318786621,125.513374328613,125.45654296875,125.464263916016,125.496871948242,125.5,125.491806030273,125.505767822266,125.592971801758,125.608978271484,125.582328796387,125.573532104492,125.627342224121,125.704002380371,125.749122619629,125.735641479492,125.674415588379,125.628120422363,125.431831359863,125.311531066895,125.233406066895,125.155853271484,125.087890625,125.034278869629,124.945304870605,124.916984558105,124.978912353516,124.998237609863,124.995018005371,124.935646057129,124.884269714355,124.821090698242,124.795806884766,124.749801635742,124.6767578125,124.571868896484,124.529098510742,124.445701599121,124.384864807129,124.325782775879,124.294731140137,124.565818786621,124.840133666992,125.150199890137,125.239547729492],[123.716598510742,123.908302307129,124.040336608887,124.0556640625,124.045509338379,123.982711791992,123.847747802734,123.753997802734,123.725204467773,123.736038208008,123.674812316895,123.667579650879,123.612014770508,123.531059265137,123.473731994629,123.418838500977,123.292671203613,123.157821655273,123.155868530273,123.210548400879,123.245315551758,123.267181396484,123.239845275879,123.236419677734,123.337013244629,123.462989807129,123.558982849121,123.574798583984,123.716598510742],[122.310836791992,122.27978515625,122.260932922363,122.247848510742,122.278022766113,122.287498474121,122.310836791992],[122.094039916992,122.013969421387,121.960159301758,121.98193359375,121.935638427734,121.923248291016,121.941017150879,121.989456176758,122.001564025879,122.103813171387,122.145027160645,122.130271911621,122.131637573242,122.094039916992],[123.775390625,123.779106140137,123.741500854492,123.62060546875,123.587203979492,123.621475219727,123.708686828613,123.775390625],[123.281837463379,123.3671875,123.274215698242,123.166412353516,123.054206848145,122.973442077637,122.949020385742,122.95751953125,123.01708984375,123.043548583984,123.206245422363,123.281837463379],[120.704399108887,120.75537109375,120.915328979492,120.980766296387,121.024711608887,121.079299926758,121.122459411621,121.202728271484,121.284378051758,121.356834411621,121.442184448242,121.522750854492,121.538665771484,121.48974609375,121.474807739258,121.479682922363,121.540618896484,121.519241333008,121.458000183105,121.412307739258,121.418167114258,121.400093078613,121.394332885742,121.356834411621,121.322372436523,121.288871765137,121.236717224121,121.155464172363,121.116981506348,121.107612609863,121.083396911621,121.048530578613,120.962501525879,120.922172546387,120.921478271484,120.8994140625,120.854789733887,120.795989990234,120.7763671875,120.768753051758,120.763671875,120.680274963379,120.651374816895,120.573135375977,120.508308410645,120.480659484863,120.455467224121,120.438095092773,120.387496948242,120.338478088379,120.352729797363,120.401260375977,120.468360900879,120.653327941895,120.704399108887],[121.914848327637,121.9765625,121.995704650879,122.114547729492,122.107322692871,122.122360229492,122.0546875,122.042381286621,122.0048828125,121.875877380371,121.829193115234,121.815040588379,121.866218566895,121.914848327637],[120.271286010742,120.272857666016,120.104202270508,120.099411010742,120.103408813477,120.120704650879,120.21142578125,120.271286010742],[124.353607177734,124.327049255371,124.294525146484,124.248245239258,124.175392150879,124.057029724121,124.038871765137,124.123733520508,124.122856140137,124.153717041016,124.186225891113,124.224899291992,124.308296203613,124.336723327637,124.417182922363,124.396293640137,124.403999328613,124.353607177734],[122.175392150879,122.172264099121,121.956253051758,121.946388244629,121.945991516113,121.959175109863,122.175392150879],[122.033500671387,122.051559448242,122.03173828125,122.017288208008,121.970306396484,122.021682739258,121.989654541016,121.932998657227,121.910636901855,121.922172546387,121.9345703125,121.923042297363,121.889259338379,121.862312316895,121.820320129395,121.839836120605,121.971672058105,122.033500671387],[121.1015625,121.254493713379,121.592964172363,121.716796875,121.845603942871,121.947563171387,122.038475036621,122.076957702637,122.146675109863,122.22119140625,122.265525817871,122.299812316895,122.315040588379,122.293853759766,122.222846984863,122.179489135742,122.150978088379,122.15234375,122.175201416016,122.236824035645,122.269050598145,122.3623046875,122.387496948242,122.392868041992,122.407524108887,122.467872619629,122.519142150879,122.5,122.467971801758,122.425788879395,122.225883483887,122.214157104492,122.135162353516,121.974708557129,121.788665771484,121.685157775879,121.595314025879,121.560935974121,121.590423583984,121.609184265137,121.607040405273,121.579200744629,121.489845275879,121.452049255371,121.411911010742,121.392280578613,121.398918151855,121.434959411621,121.543937683105,121.660545349121,121.685638427734,121.695404052734,121.626556396484,121.627922058105,121.648536682129,121.751853942871,121.7666015625,121.800483703613,121.853317260742,121.911720275879,122.07958984375,122.144340515137,122.211715698242,122.228424072266,122.287498474121,122.2744140625,122.202537536621,122.19970703125,122.237503051758,122.282608032227,122.383697509766,122.490821838379,122.627151489258,122.761039733887,122.856063842773,122.934181213379,123.014549255371,123.070991516113,123.070709228516,123.056938171387,123.059951782227,123.101943969727,123.2314453125,123.296974182129,123.305374145508,123.25927734375,123.280471801758,123.320304870605,123.377433776855,123.432327270508,123.632820129395,123.68408203125,123.725982666016,123.815727233887,123.857612609863,123.806251525879,123.607131958008,123.549613952637,123.608108520508,123.703620910645,123.764846801758,123.819236755371,123.816604614258,123.78515625,123.872749328613,123.955177307129,124.069145202637,124.104591369629,124.142768859863,124.137306213379,124.059768676758,123.961715698242,123.877830505371,123.894920349121,123.948532104492,123.917961120605,123.863868713379,123.802345275879,123.736038208008,123.626754760742,123.402336120605,123.310935974121,123.290435791016,123.295501708984,123.205955505371,123.191596984863,123.163284301758,122.896194458008,122.863479614258,122.781349182129,122.59521484375,122.543060302734,122.486328125,122.467971801758,122.493743896484,122.504196166992,122.500190734863,122.508010864258,122.596183776855,122.609382629395,122.667877197266,122.675102233887,122.599899291992,122.515235900879,122.512504577637,122.497955322266,122.406936645508,122.376564025879,122.205268859863,122.072746276855,121.777931213379,121.742874145508,121.691703796387,121.643455505371,121.501075744629,121.450775146484,121.446296691895,121.344146728516,121.203506469727,121.095512390137,121.006156921387,120.932327270508,120.840721130371,120.729095458984,120.637107849121,120.617385864258,120.616798400879,120.642669677734,120.688278198242,120.922065734863,120.951560974121,120.941314697266,120.888092041016,120.804489135742,120.707908630371,120.638282775879,120.583694458008,120.546775817871,120.582717895508,120.588668823242,120.5556640625,120.495697021484,120.438774108887,120.396102905273,120.365242004395,120.283889770508,120.250778198242,120.2138671875,120.137985229492,120.082130432129,120.044532775879,120.036628723145,120.004974365234,119.959381103516,119.932807922363,119.891593933105,119.881446838379,119.859672546387,119.808197021484,119.768951416016,119.761817932129,119.772560119629,119.790237426758,119.830764770508,119.886131286621,119.930374145508,119.985153198242,120.033401489258,120.1240234375,120.159759521484,120.271286010742,120.337020874023,120.368751525879,120.388763427734,120.389259338379,120.324996948242,120.305274963379,120.304397583008,120.321189880371,120.408882141113,120.420112609863,120.411712646484,120.427146911621,120.424514770508,120.3720703125,120.3583984375,120.505081176758,120.550979614258,120.58447265625,120.599708557129,120.709373474121,120.813766479492,120.867782592773,120.925003051758,121.051361083984,121.1015625],[121.921676635742,121.858200073242,121.8251953125,121.860740661621,121.859771728516,121.888862609863,121.943359375,121.987892150879,121.921676635742],[121.252250671387,121.246673583984,121.196090698242,121.184860229492,121.189949035645,121.213180541992,121.244720458984,121.252250671387],[121.520896911621,121.53125,121.472076416016,121.382911682129,121.374603271484,121.375968933105,121.3916015625,121.520896911621],[121.960060119629,121.941307067871,121.914070129395,121.941207885742,121.9912109375,122.031150817871,121.960060119629],[121.878128051758,121.82958984375,121.790618896484,121.796478271484,121.847854614258,121.866996765137,121.878128051758],[131.17236328125,131.149597167969,131.134963989258,131.13671875,131.151550292969,131.17236328125,131.187896728516,131.186325073242,131.17236328125],[134.595413208008,134.53466796875,134.506240844727,134.515731811523,134.555953979492,134.599716186523,134.608688354492,134.651168823242,134.659576416016,134.632720947266,134.598251342773,134.595413208008],[153.5361328125,153.703216552734,153.759872436523,153.699508666992,153.5537109375,153.519241333008,153.378997802734,153.357040405273,153.286819458008,153.322372436523,153.234481811523,153.20703125,153.20361328125,153.306732177734,153.5361328125],[154.28076171875,154.266006469727,154.229598999023,154.121200561523,154.064071655273,154.031143188477,154.0234375,154.11767578125,154.101760864258,154.237884521484,154.28076171875],[150.898727416992,150.884674072266,150.802337646484,150.785736083984,150.79931640625,150.8720703125,150.898727416992],[151.080963134766,151.123443603516,151.1943359375,151.255661010742,151.296493530273,151.230865478516,151.175491333008,150.959167480469,150.952438354492,150.896087646484,150.861328125,150.789642333984,150.776077270508,150.816696166992,150.8623046875,151.051467895508,151.044143676758,151.080963134766],[150.528427124023,150.669036865234,150.746475219727,150.788665771484,150.879104614258,150.884078979492,150.898635864258,150.89404296875,150.844039916992,150.848236083984,150.809967041016,150.678314208984,150.576278686523,150.436233520508,150.495315551758,150.508483886719,150.434677124023,150.431442260742,150.437301635742,150.498931884766,150.528427124023],[150.345413208008,150.331344604492,150.272842407227,150.109771728516,150.134963989258,150.208297729492,150.320114135742,150.357025146484,150.3681640625,150.345413208008],[152.630966186523,152.689254760742,152.81005859375,152.849792480469,152.905075073242,152.952926635742,152.995315551758,152.995025634766,152.984954833984,152.959274291992,152.966888427734,152.922760009766,152.867477416992,152.759475708008,152.720123291016,152.708206176758,152.638092041016,152.51513671875,152.577056884766,152.630966186523],[143.59033203125,143.608200073242,143.462799072266,143.324111938477,143.253799438477,143.206848144531,143.293060302734,143.443359375,143.59033203125],[151.106826782227,151.124130249023,151.046188354492,151.080764770508,151.086822509766,151.082809448242,151.004974365234,151.046279907227,151.090133666992,151.117584228516,151.116409301758,151.138580322266,151.106826782227],[143.586822509766,143.543167114258,143.366897583008,143.321868896484,143.528228759766,143.581451416016,143.592590332031,143.586822509766],[148.025787353516,147.985443115234,147.967971801758,147.87451171875,147.781066894531,147.782516479492,147.794631958008,147.846481323242,148.054794311523,148.076065063477,148.060455322266,148.025787353516],[155.957611083984,155.933212280273,155.914947509766,155.891891479492,155.804977416992,155.763473510742,155.719345092773,155.617370605469,155.520889282227,155.427352905273,155.344039916992,155.260543823242,155.208587646484,155.234481811523,155.2021484375,155.044631958008,155.010162353516,154.940231323242,154.870315551758,154.781936645508,154.75927734375,154.721099853516,154.708984375,154.741119384766,154.729309082031,154.772659301758,154.818466186523,154.870513916016,154.9970703125,155.093841552734,155.186721801758,155.197860717773,155.2275390625,155.323043823242,155.37255859375,155.466995239258,155.519332885742,155.5810546875,155.638488769531,155.734176635742,155.822555541992,155.882217407227,155.927627563477,155.957611083984],[147.17626953125,147.120208740234,147.029006958008,147.005859375,147.014739990234,147.131057739258,147.206359863281,147.221878051758,147.17626953125],[154.647277832031,154.627334594727,154.583892822266,154.576171875,154.562789916992,154.5400390625,154.605560302734,154.632614135742,154.682037353516,154.689147949219,154.727142333984,154.698440551758,154.647277832031],[146.019332885742,145.952346801758,145.904006958008,145.883590698242,145.900192260742,145.958786010742,145.995803833008,146.037414550781,146.053421020508,146.019332885742],[151.915618896484,151.967590332031,152.1171875,152.197265625,152.299407958984,152.405654907227,152.363571166992,152.376068115234,152.403518676758,152.400009155273,152.351165771484,152.257614135742,152.215728759766,152.166595458984,152.013275146484,151.983688354492,151.993942260742,152.076858520508,152.142974853516,152.077041625977,151.968460083008,151.865432739258,151.694915771484,151.51513671875,151.4814453125,151.48046875,151.455184936523,151.422470092773,151.331253051758,151.229293823242,151.090042114258,151.043151855469,150.919921875,150.808975219727,150.759567260742,150.705764770508,150.588088989258,150.473541259766,150.428314208984,150.190826416016,149.850982666016,149.750305175781,149.652542114258,149.598434448242,149.483001708984,149.38232421875,149.272644042969,149.126571655273,149.099029541016,148.807510375977,148.719146728516,148.624801635742,148.509765625,148.401168823242,148.337203979492,148.3447265625,148.432037353516,148.56494140625,148.615814208984,148.665817260742,148.724319458008,148.783493041992,148.999221801758,149.1240234375,149.245315551758,149.35888671875,149.475387573242,149.631744384766,149.681060791016,149.831436157227,149.962783813477,150.011901855469,150.045303344727,150.090026855469,150.122268676758,150.170318603516,150.108688354492,150.08154296875,150.072463989258,150.106246948242,150.18310546875,150.298736572266,150.404403686523,150.519439697266,150.625778198242,150.734481811523,150.784378051758,150.842590332031,150.900299072266,150.952926635742,151.022277832031,151.06884765625,151.137802124023,151.326553344727,151.380950927734,151.439834594727,151.572555541992,151.671188354492,151.678909301758,151.664657592773,151.551956176758,151.544235229492,151.560546875,151.593063354492,151.703704833984,151.8193359375,151.86474609375,151.915618896484],[153.659271240234,153.650085449219,153.591506958008,153.639755249023,153.662994384766,153.659271240234],[152.670608520508,152.646194458008,152.585052490234,152.543258666992,152.569931030273,152.638778686523,152.670608520508],[152.099212646484,152.088470458984,152.057312011719,151.971099853516,151.95458984375,151.974700927734,152.074615478516,152.099212646484],[151.957229614258,151.933410644531,151.929779052734,151.946380615234,152.001953125,152.011322021484,151.957229614258],[140.976165771484,140.975982666016,140.975982666016,140.975875854492,140.975784301758,140.975677490234,140.9755859375,140.9755859375,140.975494384766,140.975387573242,140.975296020508,140.975204467773,140.975204467773,140.919540405273,140.8623046875,140.874603271484,140.944046020508,140.974990844727,140.974990844727,140.974899291992,140.974807739258,140.974700927734,140.974609375,140.974609375,140.974517822266,140.974411010742,140.974319458008,140.974212646484,140.974212646484,140.974029541016,140.974029541016,140.973922729492,140.973831176758,140.973724365234,140.9736328125,140.973526000977,140.973434448242,141.003326416016,141.104782104492,141.185638427734,141.686813354492,141.836532592773,141.887496948242,141.937789916992,141.985733032227,142.211517333984,142.549026489258,142.905181884766,143.015625,143.129974365234,143.378326416016,143.508987426758,143.700576782227,143.797180175781,143.8876953125,144.015823364258,144.06640625,144.121978759766,144.247955322266,144.374404907227,144.426559448242,144.477737426758,144.524505615234,144.548233032227,144.626663208008,144.737899780273,144.843460083008,144.9384765625,145.008392333984,145.087799072266,145.2080078125,145.334564208984,145.766983032227,145.7880859375,145.792877197266,145.745208740234,145.852828979492,145.999420166016,146.205383300781,146.403427124023,147.034271240234,147.120895385742,147.248229980469,147.376663208008,147.422744750977,147.5185546875,147.566696166992,147.653015136719,147.730072021484,147.762893676758,147.802062988281,147.824508666992,147.8544921875,147.845504760742,147.810440063477,147.709579467773,147.355758666992,147.119140625,146.95361328125,146.94921875,146.960739135742,147.104888916016,147.190048217773,147.260162353516,147.365341186523,147.458984375,147.545120239258,147.724319458008,147.821884155273,147.936126708984,148.126770019531,148.151947021484,148.206451416016,148.22998046875,148.233581542969,148.246871948242,148.414443969727,148.451171875,148.52587890625,148.583099365234,148.679489135742,148.791793823242,149.097457885742,149.141708374023,149.198333740234,149.247650146484,149.264053344727,149.216201782227,149.203018188477,149.26318359375,149.418746948242,149.475784301758,149.755767822266,149.865615844727,149.973541259766,150.011032104492,149.984664916992,149.92822265625,149.864349365234,149.76123046875,149.763092041016,149.8212890625,149.874420166016,149.919143676758,149.967575073242,150.088577270508,150.206253051758,150.283889770508,150.364059448242,150.538879394531,150.6669921875,150.849502563477,150.691314697266,150.636810302734,150.446090698242,150.410263061523,150.488861083984,150.60546875,150.647171020508,150.617965698242,150.482421875,150.42578125,150.319931030273,150.142379760742,150.016799926758,149.981536865234,149.948043823242,149.834762573242,149.754089355469,149.6513671875,149.544342041016,149.352630615234,148.936813354492,148.837692260742,148.712890625,148.654205322266,148.591217041016,148.430572509766,148.383407592773,148.268753051758,148.150497436523,148.101272583008,148.051376342773,147.89013671875,147.768661499023,147.668853759766,147.614349365234,147.553115844727,147.496490478516,147.408294677734,147.298934936523,147.064453125,147.017181396484,146.925384521484,146.930374145508,146.963775634766,146.913284301758,146.856246948242,146.696578979492,146.630859375,146.524124145508,146.455856323242,146.296493530273,146.250595092773,146.18408203125,146.142959594727,146.108779907227,146.078506469727,146.033203125,145.810928344727,145.771789550781,145.728713989258,145.563385009766,145.4677734375,145.287490844727,145.1943359375,145.082336425781,144.973831176758,144.9208984375,144.885345458984,144.8642578125,144.7734375,144.684371948242,144.597946166992,144.509857177734,144.44970703125,144.431243896484,144.403427124023,144.351852416992,144.326171875,144.270217895508,144.225387573242,144.142868041992,143.9736328125,143.898254394531,143.834182739258,143.779098510742,143.723342895508,143.654891967773,143.742095947266,143.942291259766,143.892181396484,143.840621948242,143.887985229492,143.833404541016,143.779296875,143.6650390625,143.551559448242,143.518157958984,143.542190551758,143.58203125,143.61376953125,143.449996948242,143.282028198242,143.094924926758,142.905471801758,142.808303833008,142.708587646484,142.615036010742,142.524124145508,142.447555541992,142.399215698242,142.37646484375,142.347473144531,142.27587890625,142.206832885742,142.325088500977,142.360549926758,142.391021728516,142.474807739258,142.575973510742,142.797943115234,143.013671875,143.064834594727,143.11181640625,143.222946166992,143.306732177734,143.377243041992,143.392181396484,143.387496948242,143.3662109375,143.226852416992,143.078231811523,142.859191894531,142.647155761719,142.535751342773,142.435241699219,142.396286010742,142.292770385742,142.229598999023,141.978912353516,141.727340698242,141.62158203125,141.518753051758,141.405670166016,141.293655395508,141.216995239258,141.133209228516,140.976165771484],[152.9658203125,152.891693115234,152.845611572266,152.786529541016,152.739944458008,152.6806640625,152.677734375,152.693359375,152.69677734375,152.668167114258,152.598434448242,152.355758666992,152.279388427734,152.192184448242,152.136337280273,152.023239135742,151.972946166992,151.879791259766,151.793167114258,151.578506469727,151.465042114258,151.405075073242,151.066787719727,150.968063354492,150.847854614258,150.74609375,150.826461791992,150.842971801758,150.825378417969,150.995300292969,151.174606323242,151.226470947266,151.314743041992,151.475387573242,151.585754394531,151.689834594727,151.80712890625,152.032913208008,152.065032958984,152.179397583008,152.329483032227,152.380462646484,153.016799926758,153.124221801758,153.132507324219,153.111511230469,153.044342041016,153.045608520508,153.023239135742,152.9658203125],[150.436630249023,150.237503051758,150.165725708008,150.1015625,150.04345703125,149.985168457031,149.961624145508,150.1025390625,150.227142333984,150.429489135742,150.449996948242,150.451568603516,150.446090698242,150.436630249023],[147.876953125,147.844528198242,147.768936157227,147.735549926758,147.790237426758,147.812210083008,147.835830688477,147.876953125],[147.067581176758,147.400588989258,147.422561645508,147.418746948242,147.424407958984,147.444137573242,147.438095092773,147.385452270508,147.336517333984,147.301361083984,147.206359863281,147.142196655273,147.063858032227,146.926376342773,146.747863769531,146.699111938477,146.635452270508,146.572463989258,146.546493530273,146.531356811523,146.532424926758,146.607025146484,146.595901489258,146.65625,146.760055541992,146.857131958008,147.067581176758],[149.765441894531,149.76318359375,149.69091796875,149.671096801758,149.5458984375,149.5478515625,149.580947875977,149.633010864258,149.725280761719,149.765441894531],[19.6043949127197,19.6442394256592,19.92431640625,20.2082023620605,20.6647453308105,21.1405277252197,21.6341800689697,22.1684589385986,22.7318363189697,22.7662105560303,22.8237285614014,22.8939456939697,22.9767589569092,23.0155277252197,23.0319347381592,23.0421886444092,23.0874996185303,23.1703128814697,23.2823238372803,23.3701171875,23.45361328125,23.4813461303711,23.4830074310303,23.4776363372803,23.4846687316895,23.5989246368408,23.7892589569092,23.8591785430908,23.8871097564697,23.9093761444092,23.9163093566895,23.9154300689697,23.9012699127197,23.8447265625,23.4795894622803,23.4109382629395,23.3033218383789,23.2041015625,23.1812496185303,23.1750984191895,23.1969718933105,23.3271484375,23.4583988189697,23.5011711120605,23.5979480743408,23.63330078125,23.6524410247803,23.6510734558105,23.607421875,23.6256847381592,23.5813484191895,23.5448226928711,23.5396480560303,23.6052742004395,23.6588878631592,23.6796875,23.6576175689697,23.6644515991211,23.7122077941895,23.8634777069092,23.9380874633789,23.9857425689697,24.0958003997803,24.1057605743408,24.0616207122803,24.0259780883789,23.9970703125,23.9784164428711,24.00732421875,24.0462894439697,24.0947265625,24.0899410247803,24.0526351928711,24.0049800872803,23.97265625,23.7117195129395,23.6490230560303,23.5061531066895,23.4085941314697,23.2644538879395,23.0363273620605,22.9522457122803,22.8907241821289,22.7061519622803,22.6494140625,22.66064453125,22.7199230194092,22.732421875,22.7219734191895,22.7023429870605,22.7056636810303,22.7601566314697,22.8470706939697,22.85205078125,22.8397464752197,22.8097648620605,22.7012691497803,22.5799808502197,22.5386714935303,22.4730472564697,22.2025394439697,22.0201168060303,22.0021476745605,21.9676761627197,21.89013671875,21.7121086120605,21.6396484375,21.3504886627197,21.2249984741211,21.1361331939697,21.0793952941895,21.0011711120605,20.947265625,20.8684577941895,20.7995128631592,20.72900390625,20.6161136627197,20.5345706939697,20.4745121002197,20.4226570129395,20.4046878814697,20.3629875183105,20.3025398254395,20.2365245819092,20.1636714935303,20.1076183319092,20.0576171875,19.9161128997803,19.8689460754395,19.80224609375,19.7566413879395,19.7673816680908,19.7879886627197,19.7870121002197,19.77392578125,19.7300777435303,19.6641597747803,19.6302719116211,19.6266593933105,19.5930671691895,19.5347671508789,19.4796867370605,19.4416027069092,19.38623046875,19.3023433685303,19.2501945495605,19.1494140625,18.9683589935303,18.9572257995605,18.9381828308105,18.8322257995605,18.8292961120605,18.8069324493408,18.5946273803711,18.56884765625,18.5771484375,18.5624008178711,18.5162105560303,18.3484382629395,18.3052749633789,18.2663097381592,18.2052726745605,18.0992183685303,18.0876941680908,18.0495128631592,18.0283203125,18.0146484375,17.9837894439697,17.8748054504395,17.8312511444092,17.7916984558105,17.74658203125,17.6810550689697,17.6270503997803,17.5962886810303,17.58935546875,17.7092761993408,17.7354488372803,17.7201156616211,17.7022476196289,17.6546878814697,17.5545902252197,17.4623031616211,17.4152336120605,17.1519546508789,16.9807624816895,16.8800773620605,16.869140625,16.9147472381592,16.9933586120605,16.9896488189697,16.8953132629395,16.841796875,16.7786140441895,16.7252922058105,16.6791019439697,16.63916015625,16.5966796875,16.4875965118408,16.3504867553711,16.3341808319092,16.2913093566895,16.2307605743408,16.2103500366211,16.24072265625,16.2825202941895,16.3566398620605,16.3791007995605,16.3922843933105,16.4197273254395,16.4125003814697,16.3599605560303,16.2822265625,16.06640625,16.0072269439697,15.9738283157349,15.9485340118408,15.8939456939697,15.8192386627197,15.7305660247803,15.6439456939697,15.4639644622803,15.3946285247803,15.3543949127197,15.3125972747803,15.2770519256592,15.2585935592651,15.1259765625,14.993748664856,14.9844722747803,14.9899415969849,14.9829092025757,14.8958005905151,14.8093748092651,14.8142585754395,14.9174795150757,14.9638671875,15.0166015625,14.953125,14.935546875,14.9059572219849,14.7247076034546,14.7109384536743,14.7386722564697,14.7249021530151,14.6813478469849,14.6239252090454,14.6016607284546,14.6749019622803,14.6929683685303,14.7248048782349,14.7481441497803,14.7525396347046,14.70458984375,14.6923828125,14.7053718566895,14.6798830032349,14.6156244277954,14.5739259719849,14.5545902252197,14.5697259902954,14.6194343566895,14.5140628814697,14.2537107467651,14.1286134719849,14.1388673782349,14.1936521530151,14.2931642532349,14.3685541152954,14.4109373092651,14.4123048782349,14.41455078125,14.2987308502197,14.2798824310303,14.26611328125,14.2588863372803,14.4875974655151,14.5834951400757,14.5715818405151,14.5521478652954,14.56494140625,14.5583982467651,14.3508796691895,14.2136716842651,14.1982421875,14.21142578125,14.2493162155151,14.3841800689697,14.7157230377197,15.2883796691895,15.9000005722046,16.0427742004395,16.1863269805908,16.2393550872803,16.2922859191895,16.3755855560303,16.5597648620605,16.8854484558105,17.0070304870605,17.2619152069092,17.8429698944092,18.0856437683105,18.3234367370605,18.53515625,18.75927734375,18.7996082305908,18.6783199310303,18.5015621185303,18.4362316131592,18.5871105194092,18.6696281433105,18.83642578125,18.9762706756592,19.4071292877197,19.5601558685303,19.6043949127197],[-67.8724670410156,-67.8818359375,-67.8911666870117,-67.8954620361328,-67.9019012451172,-67.9303741455078,-67.9370574951172,-67.9306182861328,-67.9189453125,-67.861083984375,-67.8551788330078,-67.8436508178711,-67.8433609008789,-67.8490676879883,-67.8591766357422,-67.8634262084961,-67.8667984008789,-67.8681106567383,-67.8724670410156],[-65.4255828857422,-65.5040054321289,-65.5550765991211,-65.5722122192383,-65.4771499633789,-65.3662109375,-65.3026885986328,-65.2948760986328,-65.4255828857422],[-66.12939453125,-66.0984878540039,-66.0684051513672,-66.0926742553711,-66.0704040527344,-65.8787612915039,-65.7555694580078,-65.6288070678711,-65.620849609375,-65.7184066772461,-65.7822265625,-65.8341293334961,-65.9708023071289,-66.135498046875,-66.2450180053711,-66.285888671875,-66.3257827758789,-66.4085464477539,-66.5107879638672,-66.5984344482422,-66.7724151611328,-66.8375930786133,-66.9000015258789,-66.9612274169922,-67.0133285522461,-67.1423873901367,-67.1968765258789,-67.17431640625,-67.1724548339844,-67.2041473388672,-67.2389602661133,-67.2640686035156,-67.21337890625,-67.1717758178711,-67.1586380004883,-67.113037109375,-67.0596160888672,-66.8128890991211,-66.1885757446289,-66.153076171875,-66.12939453125],[124.905265808105,124.848930358887,124.846092224121,124.889541625977,124.9345703125,124.905265808105],[130.526947021484,130.554107666016,130.617965698242,130.651565551758,130.658004760742,130.687316894531,130.636520385742,130.569244384766,130.45751953125,130.314743041992,130.235733032227,130.179885864258,130.068267822266,130.00732421875,129.92822265625,129.876358032227,129.75634765625,129.686325073242,129.682418823242,129.758972167969,129.765823364258,129.712112426758,129.742004394531,129.708694458008,129.341110229492,129.2451171875,129.109771728516,128.945220947266,128.842971801758,128.701370239258,128.610748291016,128.51123046875,128.392959594727,128.304489135742,128.106338500977,127.966598510742,127.867080688477,127.568168640137,127.527442932129,127.547264099121,127.548919677734,127.522857666016,127.457420349121,127.422264099121,127.383491516113,127.39453125,127.496971130371,127.580955505371,127.698921203613,127.7861328125,127.9716796875,128.123046875,128.162506103516,128.249404907227,128.329483032227,128.374603271484,128.339462280273,128.279296875,128.22314453125,128.168655395508,128.106246948242,128.038955688477,127.905281066895,127.78466796875,127.745506286621,127.579490661621,127.53271484375,127.294044494629,127.169532775879,127.090324401855,127.009658813477,126.940032958984,126.87890625,126.754295349121,126.666801452637,126.66650390625,126.664543151855,126.633895874023,126.623237609863,126.572746276855,126.369918823242,126.203125,126.161041259766,126.11669921875,126.05029296875,125.941696166992,125.769134521484,125.695014953613,125.676177978516,125.581535339355,125.449317932129,125.406646728516,125.357810974121,125.364845275879,125.310745239258,125.101951599121,125.026763916016,124.988777160645,125.193161010742,125.246681213379,125.206741333008,125.162605285645,124.995018005371,124.907028198242,124.779495239258,124.690925598145,124.874519348145,124.882713317871,124.880569458008,124.973731994629,125.067390441895,125.309669494629,125.415328979492,125.491806030273,125.554489135742,125.488677978516,125.424224853516,125.298919677734,125.168846130371,125.157325744629,125.409660339355,125.413185119629,125.373626708984,125.36083984375,125.180076599121,125.100090026855,124.867866516113,124.775291442871,124.738868713379,124.732231140137,124.699211120605,124.638275146484,124.607620239258,124.557418823242,124.40380859375,124.3486328125,124.375091552734,124.362106323242,124.386627197266,124.481056213379,124.712394714355,124.77197265625,124.889350891113,124.942283630371,124.996871948242,125.013374328613,125.025970458984,125.072952270508,125.185935974121,125.314453125,125.416893005371,125.542579650879,125.593849182129,125.645111083984,125.659187316895,125.688278198242,125.728317260742,125.783981323242,125.874908447266,125.989067077637,126.066795349121,126.093162536621,126.144538879395,126.253616333008,126.328704833984,126.411819458008,126.451461791992,126.490432739258,126.513572692871,126.540138244629,126.578323364258,126.601272583008,126.696975708008,126.721580505371,126.743064880371,126.787696838379,126.847267150879,126.903518676758,126.954788208008,127.006927490234,127.061325073242,127.085357666016,127.128410339355,127.13671875,127.179695129395,127.270797729492,127.420310974121,127.516998291016,127.572166442871,127.687698364258,127.918647766113,128.013092041016,128.052734375,128.111236572266,128.1494140625,128.200286865234,128.2548828125,128.290924072266,128.289260864258,128.2578125,128.181732177734,128.131927490234,128.084167480469,128.056060791016,128.032913208008,128.028717041016,128.045211791992,128.16015625,128.307815551758,128.42724609375,128.626770019531,128.7490234375,128.83984375,128.923446655273,128.960632324219,129.077239990234,129.133682250977,129.195510864258,129.205383300781,129.2177734375,129.252532958984,129.313674926758,129.365814208984,129.423629760742,129.48486328125,129.523727416992,129.567581176758,129.603912353516,129.6279296875,129.697845458984,129.7197265625,129.746475219727,129.7734375,129.779190063477,129.841506958008,129.861038208008,129.898239135742,129.941207885742,129.976959228516,130.022262573242,130.082626342773,130.124801635742,130.151275634766,130.240325927734,130.248825073242,130.246673583984,130.295608520508,130.360748291016,130.450286865234,130.498229980469,130.526947021484],[-17.1908683776855,-17.0544929504395,-16.92919921875,-16.7739753723145,-16.6932621002197,-16.7652835845947,-16.83740234375,-17.0182609558105,-17.1711902618408,-17.2260265350342,-17.2410163879395,-17.1908683776855],[-25.02734375,-25.0315437316895,-25.08837890625,-25.159912109375,-25.1983871459961,-25.1635246276855,-25.0829105377197,-25.0443363189697,-25.02734375],[-25.6489734649658,-25.5854969024658,-25.2666015625,-25.1819343566895,-25.1907234191895,-25.2511234283447,-25.4390144348145,-25.7344722747803,-25.8336906433105,-25.8478527069092,-25.8458976745605,-25.7837409973145,-25.6489734649658],[-28.1472663879395,-28.0647945404053,-28.1897468566895,-28.2311534881592,-28.3324222564697,-28.4544925689697,-28.5311527252197,-28.548828125,-28.51025390625,-28.4021472930908,-28.1472663879395],[-28.6413097381592,-28.7438468933105,-28.842041015625,-28.69775390625,-28.6554203033447,-28.6242179870605,-28.6058101654053,-28.6413097381592],[-27.7784671783447,-27.8258781433105,-28.0923347473145,-28.187255859375,-28.3106441497803,-27.962646484375,-27.7784671783447],[-27.0752429962158,-27.0953121185303,-27.3028316497803,-27.3619136810303,-27.3859367370605,-27.3510265350342,-27.2596683502197,-27.1270008087158,-27.0419425964355,-27.0419921875,-27.0752429962158],[-31.1371078491211,-31.1813468933105,-31.2576160430908,-31.282958984375,-31.2608394622803,-31.1998538970947,-31.1386241912842,-31.1371078491211],[-7.40615272521973,-7.49360275268555,-7.83413124084473,-7.939697265625,-8.13676738739014,-8.48432636260986,-8.59765625,-8.73911190032959,-8.84843730926514,-8.93535137176514,-8.997802734375,-8.92626953125,-8.81416034698486,-8.81855487823486,-8.79184532165527,-8.82265567779541,-8.87895488739014,-8.80224704742432,-8.81093692779541,-8.881103515625,-8.66831111907959,-8.73398399353027,-8.79887676239014,-8.86162185668945,-8.914794921875,-9.09599590301514,-9.18671894073486,-9.21328163146973,-9.203369140625,-9.25039100646973,-9.17783164978027,-9.09331130981445,-9.021484375,-8.97705078125,-9.00048732757568,-8.93808650970459,-8.79160118103027,-8.86748027801514,-8.95429706573486,-9.09101581573486,-9.13579082489014,-9.25229454040527,-9.35673809051514,-9.41020488739014,-9.47412109375,-9.479736328125,-9.47475624084473,-9.43144512176514,-9.41435527801514,-9.35283184051514,-9.35722637176514,-9.374755859375,-9.31962966918945,-9.25141620635986,-9.14829158782959,-9.00405311584473,-8.83784198760986,-8.851318359375,-8.88662052154541,-8.87265586853027,-8.77241134643555,-8.73159217834473,-8.68461894989014,-8.67397499084473,-8.65556621551514,-8.65981388092041,-8.67460918426514,-8.73837852478027,-8.8056640625,-8.81083965301514,-8.75542068481445,-8.84638690948486,-8.88759708404541,-8.87822341918945,-8.77714824676514,-8.68295955657959,-8.58964824676514,-8.5380859375,-8.32255935668945,-8.26606369018555,-8.21308517456055,-8.20419979095459,-8.173583984375,-8.13930702209473,-8.12998104095459,-8.21333026885986,-8.22475624084473,-8.18125057220459,-8.17353534698486,-8.15249061584473,-8.09443378448486,-7.990966796875,-7.92084980010986,-7.89638662338257,-7.69306659698486,-7.64467811584473,-7.61259746551514,-7.51259756088257,-7.40361356735229,-7.26855516433716,-7.20961904525757,-7.19833993911743,-7.19536161422729,-7.17792940139771,-7.14711904525757,-7.09912157058716,-7.03046846389771,-6.86552762985229,-6.83320283889771,-6.77729511260986,-6.70361280441284,-6.61826229095459,-6.57534170150757,-6.55751991271973,-6.55258750915527,-6.55898427963257,-6.54218721389771,-6.48466825485229,-6.39169979095459,-6.30805683135986,-6.24311542510986,-6.22168016433716,-6.21250009536743,-6.24433565139771,-6.28935575485229,-6.40312480926514,-6.56591796875,-6.69013643264771,-6.77578115463257,-6.8828125,-6.91552734375,-6.92846727371216,-6.85771465301514,-6.83588838577271,-6.818359375,-6.829833984375,-6.835693359375,-6.85205078125,-6.84794950485229,-6.82177782058716,-6.81015634536743,-6.85888719558716,-6.94843721389771,-7.01469707489014,-7.03261661529541,-7.02783203125,-6.91640615463257,-6.89609384536743,-6.91118144989014,-6.97539043426514,-7.03671884536743,-7.04741239547729,-7.11767625808716,-7.45410203933716,-7.53569316864014,-7.52421855926514,-7.44511699676514,-7.36269569396973,-7.33544921875,-7.30576181411743,-7.17241239547729,-7.04296827316284,-6.99794864654541,-7.00624990463257,-7.04604482650757,-7.12548828125,-7.21992254257202,-7.28154277801514,-7.28637647628784,-7.30595731735229,-7.33579158782959,-7.343017578125,-7.10639667510986,-6.97480487823486,-6.95756864547729,-6.98110342025757,-7.02285146713257,-7.072509765625,-7.18544960021973,-7.292236328125,-7.37890625,-7.44394588470459,-7.50351524353027,-7.49604463577271,-7.46718788146973,-7.40615272521973],[-58.1597633361816,-58.1377906799316,-58.1246109008789,-58.0915031433105,-58.0584449768066,-58.0253944396973,-58.0022468566895,-58.0088386535645,-57.9956092834473,-57.9790496826172,-57.9625015258789,-57.9151344299316,-57.8914070129395,-57.9084968566895,-57.9019050598145,-57.8848152160645,-57.9004898071289,-57.8922386169434,-57.8600082397461,-57.8302268981934,-57.8269577026367,-57.8600082397461,-57.886474609375,-57.89306640625,-57.8732452392578,-57.9062957763672,-57.9459915161133,-57.9360809326172,-57.929443359375,-57.9261703491211,-57.9162139892578,-57.929443359375,-57.9426727294922,-57.9493179321289,-57.9327621459961,-57.9625015258789,-57.9790496826172,-57.9856910705566,-57.9559097290039,-57.8798370361328,-57.8203125,-57.7640647888184,-57.7210960388184,-57.6417007446289,-57.5689468383789,-57.4763679504395,-57.3936500549316,-57.3308601379395,-57.2382316589355,-57.142333984375,-57.0298843383789,-56.937255859375,-56.8446769714355,-56.7751960754395,-56.7024421691895,-56.6330070495605,-56.580078125,-56.55029296875,-56.5238265991211,-56.4477996826172,-56.3948745727539,-56.3518524169922,-56.2757797241211,-56.2460441589355,-56.1898422241211,-56.0674781799316,-55.9914054870605,-55.9053726196289,-55.8491706848145,-55.799560546875,-55.7532691955566,-55.7466316223145,-55.7036628723145,-55.6474113464355,-55.61767578125,-55.6275863647461,-55.654052734375,-55.6507301330566,-55.6209983825684,-55.6209983825684,-55.6011238098145,-55.5614280700684,-55.5481948852539,-55.5548324584961,-55.5283699035645,-55.5184593200684,-55.5349617004395,-55.5416030883789,-55.5382804870605,-55.5184593200684,-55.4588851928711,-55.4423828125,-55.4423828125,-55.4159164428711,-55.3663101196289,-55.2869148254395,-55.1943359375,-55.0818862915039,-54.982666015625,-54.9264640808105,-54.8172836303711,-54.7213859558105,-54.6717796325684,-54.62548828125,-54.5295906066895,-54.4402351379395,-54.3708038330078,-54.2417984008789,-54.2668952941895,-54.3182601928711,-54.3172836303711,-54.281005859375,-54.3129386901855,-54.4129905700684,-54.4541015625,-54.4362297058105,-54.47314453125,-54.6105461120605,-54.6158676147461,-54.6319351196289,-54.677734375,-54.7550773620605,-54.8254890441895,-54.888916015625,-54.9344749450684,-54.962158203125,-55.0136222839355,-55.0888671875,-55.129638671875,-55.1359367370605,-55.2080078125,-55.3458023071289,-55.4266624450684,-55.4506340026855,-55.4967269897461,-55.5648918151855,-55.5972633361816,-55.5937995910645,-55.6329078674316,-55.7146492004395,-55.7899894714355,-55.8590316772461,-55.9514656066895,-56.0673332214355,-56.1640625,-56.24169921875,-56.310546875,-56.3705062866211,-56.4371566772461,-56.5105476379395,-56.6033706665039,-56.7157211303711,-56.80517578125,-56.8717269897461,-56.9739761352539,-57.11181640625,-57.3912582397461,-57.8122100830078,-58.1682624816895,-58.6048316955566,-58.6417503356934,-58.6186027526855,-58.5477066040039,-58.5032196044922,-58.4852561950684,-58.4363250732422,-58.3564453125,-58.3225631713867,-58.3346633911133,-58.3176765441895,-58.2716751098633,-58.2455596923828,-58.2393569946289,-58.2220687866211,-58.1913070678711,-58.1879386901855,-58.2051773071289,-58.2030258178711,-58.1814918518066,-58.1546897888184,-58.1356468200684,-58.1180648803711,-58.1111335754395,-58.0824203491211,-57.9431190490723,-57.8906288146973,-57.88623046875,-57.865234375,-57.782470703125,-57.7570762634277,-57.7547836303711,-57.7254867553711,-57.6258277893066,-57.5716781616211,-57.5631332397461,-57.5871543884277,-57.6438941955566,-57.8216781616211,-57.9598159790039,-58.136474609375,-58.2527847290039,-58.3086891174316,-58.3653793334961,-58.4228019714355,-58.5196266174316,-58.7240257263184,-59.187255859375,-59.3729476928711,-59.4353981018066,-59.6085891723633,-59.8924789428711,-60.1103019714355,-60.26220703125,-60.5053749084473,-60.8398399353027,-61.0329093933105,-61.084716796875,-61.2083969116211,-61.4039535522461,-61.5055198669434,-61.5130386352539,-61.5709915161133,-61.6794891357422,-61.7985343933105,-61.9280281066895,-62.0665969848633,-62.2141571044922,-62.3725090026855,-62.5415496826172,-62.6259765625,-62.6256828308105,-62.6509742736816,-62.6285171508789,-62.5669441223145,-62.4778289794922,-62.3854522705078,-62.2766609191895,-62.2765121459961,-62.2763175964355,-62.1216278076172,-62.0118141174316,-61.9169425964355,-61.8208961486816,-61.7568359375,-61.5118179321289,-61.0959930419922,-60.8887672424316,-60.4516143798828,-60.0073699951172,-59.5408706665039,-59.0905265808105,-58.7411155700684,-58.4742202758789,-58.1801719665527,-58.1600570678711,-58.1399421691895,-58.1597633361816],[34.2453117370605,34.2124977111816,34.1981468200684,34.3873062133789,34.4773445129395,34.5241203308105,34.5255851745605,34.3501968383789,34.3483390808105,34.2453117370605],[35.4505882263184,35.40869140625,35.2766609191895,35.1011734008789,34.9078140258789,34.8804664611816,34.8727531433105,34.92919921875,34.9509773254395,35.03466796875,35.1534194946289,35.2037086486816,35.1980476379395,35.1271476745605,35.05322265625,34.9830055236816,34.9611320495605,34.9538078308105,34.9783210754395,34.98974609375,34.9788093566895,34.9713897705078,34.9559555053711,34.99951171875,35.0105438232422,35.0650405883789,35.1932601928711,35.3038101196289,35.3621101379395,35.38671875,35.4026374816895,35.484375,35.5514640808105,35.5720672607422,35.5347671508789,35.5314483642578,35.5589866638184,35.4994125366211,35.4654312133789,35.4505882263184],[-138.505859375,-138.534866333008,-138.524017333984,-138.54638671875,-138.568359375,-138.54638671875,-138.514938354492,-138.505859375],[-136.293899536133,-136.314056396484,-136.316009521484,-136.344039916992,-136.382904052734,-136.435699462891,-136.464263916016,-136.478515625,-136.458694458008,-136.426116943359,-136.38037109375,-136.32763671875,-136.293899536133],[-140.685348510742,-140.671875,-140.696044921875,-140.773239135742,-140.78173828125,-140.685348510742],[-136.971725463867,-136.971343994141,-137.067581176758,-137.029632568359,-136.971725463867],[-140.829879760742,-140.822708129883,-140.860061645508,-140.895462036133,-140.958648681641,-140.973526000977,-140.925155639648,-140.893264770508,-140.829879760742],[-140.809371948242,-140.804443359375,-140.8408203125,-140.8515625,-140.850723266602,-140.824264526367,-140.803619384766,-140.761428833008,-140.686187744141,-140.649810791016,-140.638229370117,-140.652053833008,-140.776321411133,-140.815185546875,-140.83251953125,-140.829254150391,-140.809371948242],[-149.321533203125,-149.177688598633,-149.15087890625,-149.181777954102,-149.254501342773,-149.290481567383,-149.341110229492,-149.481689453125,-149.578903198242,-149.6328125,-149.635009765625,-149.611434936523,-149.508102416992,-149.379196166992,-149.330078125,-149.321533203125],[-149.813674926758,-149.844924926758,-149.886581420898,-149.905120849609,-149.911819458008,-149.902145385742,-149.808792114258,-149.782424926758,-149.813674926758],[-151.409820556641,-151.449462890625,-151.485488891602,-151.476409912109,-151.466751098633,-151.411178588867,-151.364501953125,-151.409820556641],[-143.440567016602,-143.386169433594,-143.458541870117,-143.550689697266,-143.515579223633,-143.464691162109,-143.440567016602],[-143.571136474609,-143.610641479492,-143.707427978516,-143.670211791992,-143.614791870117,-143.571136474609],[-151.466598510742,-151.484909057617,-151.504150390625,-151.512405395508,-151.505767822266,-151.457427978516,-151.438079833984,-151.466598510742],[-145.486679077148,-145.482223510742,-145.502746582031,-145.539840698242,-145.553115844727,-145.576705932617,-145.609130859375,-145.61279296875,-145.613815307617,-145.577087402344,-145.542327880859,-145.516983032227,-145.486679077148],[-142.511810302734,-142.529586791992,-142.5068359375,-142.481201171875,-142.511810302734],[-145.051361083984,-145.057678222656,-145.076416015625,-145.137939453125,-145.160736083984,-145.133544921875,-145.051361083984],[-138.651123046875,-138.687744140625,-138.690383911133,-138.642929077148,-138.624465942383,-138.632369995117,-138.651123046875],[-139.059707641602,-139.133987426758,-139.134231567383,-139.107467651367,-139.083160400391,-139.059707641602],[-139.024322509766,-138.874465942383,-138.827331542969,-138.874954223633,-139.024261474609,-139.073669433594,-139.134078979492,-139.166458129883,-139.024322509766],[-140.075637817383,-140.097351074219,-140.138031005859,-140.144393920898,-140.070953369141,-140.031097412109,-140.075637817383],[-139.556198120117,-139.620986938477,-139.631790161133,-139.611770629883,-139.583984375,-139.534576416016,-139.508346557617,-139.509902954102,-139.556198120117],[-140.072601318359,-140.170562744141,-140.217437744141,-140.252685546875,-140.240036010742,-140.224411010742,-140.057662963867,-140.043701171875,-140.046142578125,-140.072601318359],[51.2679710388184,51.1780281066895,51.0933609008789,51.0227546691895,50.9660148620605,50.9283180236816,50.8556632995605,50.8043937683105,50.8359375,50.8467750549316,50.77734375,50.7545890808105,50.7628898620605,50.8026351928711,50.86865234375,50.90380859375,51.0031280517578,51.1081047058105,51.2623062133789,51.3890609741211,51.5430679321289,51.572265625,51.5269546508789,51.4853515625,51.51025390625,51.51953125,51.5614242553711,51.6019515991211,51.60888671875,51.5869140625,51.5333976745605,51.4279289245605,51.396484375,51.2679710388184],[28.2125015258789,28.3176765441895,28.4512691497803,28.7607421875,28.7882804870605,28.7914066314697,28.7698249816895,28.7666015625,28.78173828125,28.8243160247803,28.8943367004395,29.0274391174316,29.2235355377197,29.4037113189697,29.5676765441895,29.6519508361816,29.7058582305908,29.6890640258789,29.6786136627197,29.6353511810303,29.60546875,29.5575199127197,29.0482406616211,29.0810527801514,29.0691413879395,29.0477523803711,29.0953121185303,28.9806632995605,28.9305667877197,28.8915061950684,28.9261722564697,28.8704109191895,28.8490219116211,28.8464851379395,28.8135738372803,28.8070297241211,28.88818359375,28.8517570495605,28.69921875,28.6454105377197,28.6585922241211,28.5907230377197,28.5853500366211,28.4234371185303,28.3751945495605,28.2219734191895,28.0500011444092,27.9489250183105,27.88427734375,27.7385749816895,27.7107410430908,27.6708984375,27.56103515625,27.4253902435303,27.1207027435303,27.0869140625,26.8477535247803,26.4892578125,26.2158203125,25.9333992004395,25.81884765625,25.6861343383789,25.4970703125,25.15966796875,24.8082027435303,24.4305648803711,24.2267570495605,23.9507808685303,23.5345706939697,23.224609375,22.9190425872803,22.86767578125,22.8564453125,22.8682613372803,22.9113273620605,22.9853496551514,23.0244140625,23.0285148620605,22.9454116821289,22.7751960754395,22.705078125,22.6878910064697,22.6833000183105,22.64794921875,22.5818367004395,22.5306644439697,22.4945316314697,22.5023441314697,22.5540027618408,22.6201171875,22.7007808685303,22.734375,22.7208995819092,22.64208984375,22.4976558685303,22.3506832122803,22.2009773254395,22.0930671691895,22.0269527435303,21.9092788696289,21.740234375,21.6361312866211,21.5970706939697,21.5231456756592,21.3600597381592,21.35791015625,21.3843746185303,21.4421863555908,21.5199222564697,21.5323238372803,21.533203125,21.4719734191895,21.4099597930908,21.3777332305908,21.3570308685303,21.3529300689697,21.37109375,21.3958988189697,21.4207019805908,21.4344730377197,21.4678707122803,21.4917964935303,21.490234375,21.4654293060303,21.4314441680908,21.3817386627197,21.2264633178711,21.1478519439697,21.0999031066895,21.0238285064697,20.9417972564697,20.8708000183105,20.7940425872803,20.7742176055908,20.7724609375,20.7865238189697,20.7860355377197,20.7658214569092,20.779296875,20.7757816314697,20.7749996185303,20.7601566314697,20.7468757629395,20.7278327941895,20.7092761993408,20.6527328491211,20.5811519622803,20.5326175689697,20.43798828125,20.3585929870605,20.3013687133789,20.2417964935303,20.2809581756592,20.5081062316895,20.6136722564697,20.6610355377197,20.7074222564697,20.7327156066895,20.7374019622803,20.76025390625,20.8370113372803,21.0398445129395,21.1216793060303,21.1519527435303,21.17041015625,21.1917972564697,21.2645511627197,21.2632827758789,21.2522468566895,21.2945308685303,21.3202152252197,21.361328125,21.4110336303711,21.4970703125,21.47705078125,21.4944343566895,21.5841789245605,21.6526374816895,21.6514644622803,21.6614265441895,21.7217769622803,21.7854499816895,21.8693370819092,21.8992195129395,21.9542980194092,21.9953136444092,21.9997062683105,22.0379886627197,22.1119136810303,22.18505859375,22.24462890625,22.2906246185303,22.3514652252197,22.41748046875,22.4914073944092,22.5628910064697,22.6083984375,22.6767578125,22.8517570495605,22.8766593933105,22.9128913879395,23.0547847747803,23.0908203125,23.1394538879395,23.20263671875,23.408203125,23.6287097930908,23.6690425872803,23.6820316314697,23.708984375,24.0018558502197,24.04736328125,24.0597667694092,24.177734375,24.2819347381592,24.3809566497803,24.4840812683105,24.5789070129395,24.6509761810303,24.837890625,24.8933582305908,24.9791011810303,25.0738277435303,25.1696300506592,25.4642581939697,25.6892585754395,25.90869140625,26.1626949310303,26.2362308502197,26.2769527435303,26.3056640625,26.4423828125,26.5724601745605,26.6189460754395,26.7137699127197,26.7873058319092,26.9009761810303,26.9807624816895,27.01220703125,27.0803718566895,27.1520519256592,27.2308597564697,27.2481441497803,27.2779293060303,27.3369140625,27.44921875,27.46484375,27.5158195495605,27.6140613555908,27.6961917877197,27.7679691314697,27.8023433685303,27.8538093566895,27.9742183685303,28.07177734375,28.1499996185303,28.2046871185303,28.2394523620605,28.22265625,28.2443370819092,28.1996097564697,28.119140625,28.0997066497803,28.1135749816895,28.1155281066895,28.1349620819092,28.15625,28.1597652435303,28.130859375,28.09033203125,28.07470703125,28.1119136810303,28.1625003814697,28.2125015258789],[146.045608520508,146.032333374023,146.028030395508,146.048919677734,146.088577270508,146.100784301758,146.086318969727,146.069915771484,146.045608520508],[145.881546020508,145.895599365234,145.913864135742,145.93115234375,145.941116333008,145.943542480469,145.93115234375,145.9072265625,145.893951416016,145.886535644531,145.881546020508,145.869140625,145.869140625,145.881546020508],[146.358779907227,146.332321166992,146.288192749023,146.273834228516,146.28369140625,146.310150146484,146.333312988281,146.349807739258,146.358779907227],[146.713958740234,146.683013916016,146.608596801758,146.613479614258,146.621978759766,146.824615478516,146.884078979492,146.899017333984,146.713958740234],[146.207611083984,146.35595703125,146.567779541016,146.516204833984,146.4365234375,146.296188354492,146.172943115234,146.1123046875,145.9140625,145.887313842773,145.766983032227,145.586822509766,145.555847167969,145.439254760742,145.426177978516,145.461715698242,145.666305541992,145.748336791992,145.773345947266,145.851959228516,145.890228271484,145.9404296875,146.112106323242,146.207611083984],[148.599502563477,148.414642333984,148.262298583984,148.005264282227,147.913772583008,147.784072875977,147.657821655273,147.621871948242,147.609573364258,147.563079833984,147.310165405273,147.207412719727,147.098434448242,146.8974609375,146.933502197266,146.974227905273,147.140914916992,147.15478515625,147.24658203125,147.430465698242,147.557815551758,147.657913208008,147.769439697266,147.885543823242,147.872650146484,147.924026489258,147.964553833008,148.056060791016,148.130081176758,148.32421875,148.6123046875,148.706649780273,148.772659301758,148.812210083008,148.826171875,148.825393676758,148.803024291992,148.837097167969,148.790725708008,148.599502563477],[47.9830093383789,47.9676780700684,47.9203147888184,47.9175796508789,47.9471664428711,47.9871101379395,47.9830093383789],[149.687698364258,149.538864135742,149.447067260742,149.665924072266,149.796295166016,149.962310791016,150.308792114258,150.3486328125,150.553115844727,150.234573364258,150.195007324219,150.056640625,149.9541015625,149.883392333984,149.687698364258],[152.002044677734,151.815628051758,151.754104614258,151.723434448242,151.71533203125,151.864349365234,152.039840698242,152.165817260742,152.234664916992,152.288864135742,152.002044677734],[153.10107421875,153.053802490234,153.004104614258,152.984268188477,153.049118041992,153.079193115234,153.10107421875],[154.081253051758,154.04296875,154.000686645508,153.992279052734,154.091705322266,154.126358032227,154.199020385742,154.228424072266,154.204696655273,154.081253051758],[154.810440063477,154.71484375,154.610931396484,154.612991333008,154.824890136719,154.899612426758,154.88330078125,154.802337646484,154.829879760742,154.810440063477],[155.921081542969,155.792373657227,155.607513427734,155.516403198242,155.448928833008,155.397171020508,155.288665771484,155.243072509766,155.243072509766,155.195114135742,155.218353271484,155.326751708984,155.433883666992,155.68017578125,155.772750854492,155.884765625,156.001663208008,156.096862792969,156.122848510742,156.1005859375,156.04443359375,155.921081542969],[156.405075073242,156.365432739258,156.325775146484,156.1962890625,156.16796875,156.213073730469,156.37646484375,156.455871582031,156.487503051758,156.483108520508,156.405075073242],[155.644821166992,155.553512573242,155.512786865234,155.483489990234,155.467391967773,155.568557739258,155.6396484375,155.653625488281,155.644821166992],[142.761032104492,142.976165771484,142.985946655273,142.967086791992,142.926559448242,142.911422729492,142.936416625977,142.91796875,143.095504760742,143.2236328125,143.259963989258,143.287887573242,143.32470703125,143.332611083984,143.323638916016,143.295120239258,143.264251708984,143.200988769531,143.172271728516,143.155563354492,143.190628051758,143.250579833984,143.294738769531,143.299514770508,143.320510864258,143.417770385742,143.455474853516,143.467391967773,143.472946166992,143.48876953125,143.5341796875,143.736038208008,143.816009521484,144.047958374023,144.141311645508,144.199615478516,144.239929199219,144.272064208984,144.341217041016,144.431747436523,144.606842041016,144.685546875,144.706634521484,144.713775634766,144.672653198242,144.620986938477,144.536331176758,144.411819458008,144.283782958984,144.12548828125,144.048736572266,143.9677734375,143.819137573242,143.732330322266,143.382232666016,143.236328125,143.10498046875,143.02685546875,142.9716796875,142.650970458984,142.57421875,142.5458984375,142.556930541992,142.579010009766,142.670104980469,142.745407104492,142.80078125,142.863967895508,142.905471801758,142.940322875977,143.005569458008,143.089248657227,143.177932739258,143.217666625977,143.318649291992,143.384368896484,143.447265625,143.485656738281,143.540328979492,143.578704833984,143.580657958984,143.508605957031,143.490615844727,143.482330322266,143.463485717773,143.431640625,143.418640136719,143.370315551758,143.352142333984,143.282333374023,143.0478515625,142.829299926758,142.795516967773,142.747360229492,142.69189453125,142.6357421875,142.578033447266,142.478805541992,142.406433105469,142.349990844727,142.304000854492,142.208602905273,142.149703979492,142.0771484375,142.015625,141.961608886719,141.929977416992,141.916305541992,141.830368041992,141.866500854492,142.011032104492,142.038681030273,142.016906738281,141.984176635742,141.962509155273,141.964065551758,142.015625,142.075973510742,142.149215698242,142.181732177734,142.135345458984,142.028717041016,141.897262573242,141.873046875,141.866302490234,141.979598999023,142.020126342773,142.066497802734,142.108703613281,142.142288208008,142.153121948242,142.14306640625,142.071090698242,142.066024780273,142.100494384766,142.147262573242,142.207916259766,142.206741333008,142.090713500977,142.005950927734,141.872940063477,141.771865844727,141.722366333008,141.771865844727,141.80810546875,141.721008300781,141.66845703125,141.660842895508,141.682418823242,141.74755859375,141.803314208984,141.855575561523,141.873626708984,141.8388671875,141.823532104492,141.852447509766,141.964462280273,142.141983032227,142.179885864258,142.318939208984,142.370513916016,142.424026489258,142.526168823242,142.58349609375,142.509185791016,142.552536010742,142.679595947266,142.688873291016,142.642868041992,142.683013916016,142.705963134766,142.670227050781,142.466598510742,142.3349609375,142.551666259766,142.615615844727,142.666198730469,142.692779541016,142.761032104492],[168.0390625,168.081344604492,167.677352905273,167.488082885742,167.441497802734,167.51171875,167.592468261719,167.710647583008,167.882614135742,168.0390625],[137.178619384766,137.055267333984,136.969436645508,136.902740478516,136.76513671875,136.714645385742,136.795318603516,136.995697021484,137.077545166016,137.156051635742,137.178619384766],[137.940536499023,138.03125,138.172073364258,138.206146240234,138.096481323242,138.0166015625,137.9912109375,137.95947265625,137.8701171875,137.790222167969,137.721496582031,137.6611328125,137.525588989258,137.462692260742,137.276077270508,137.23291015625,137.275192260742,137.384384155273,137.435546875,137.543655395508,137.577346801758,137.910446166992,137.940536499023],[21.2357425689697,21.2975578308105,21.3892574310303,21.4470710754395,21.5546875,21.6827144622803,21.8739261627197,22.0723628997803,22.1378917694092,22.3463859558105,22.5672855377197,22.62744140625,22.7365226745605,22.82470703125,22.8312492370605,22.7096691131592,22.6844711303711,22.6798820495605,22.7243156433105,22.7662105560303,22.7318363189697,22.1684589385986,21.6341800689697,21.1405277252197,20.6647453308105,20.2082023620605,19.92431640625,19.6442394256592,19.6043949127197,19.7584953308105,19.85888671875,19.9441394805908,19.9532222747803,19.9745121002197,20.1076183319092,20.3966789245605,20.5203132629395,20.6789073944092,20.845703125,20.8998050689697,20.9578113555908,20.859375,20.5948238372803,20.677734375,20.7740230560303,20.8875007629395,20.9958992004395,21.1888675689697,21.2228527069092,21.2357425689697],[166.650283813477,166.645111083984,166.521286010742,166.463684082031,166.381744384766,166.324798583984,166.229873657227,166.119720458984,166.082321166992,166.066314697266,165.991882324219,165.751068115234,165.830459594727,165.931243896484,166.2119140625,166.275772094727,166.22998046875,166.248046875,166.404296875,166.4794921875,166.577346801758,166.650283813477],[150.589950561523,150.511123657227,150.471771240234,150.47021484375,150.592468261719,150.666213989258,150.712707519531,150.727737426758,150.589950561523],[163.635162353516,163.471389770508,163.447265625,163.431838989258,163.42724609375,163.576766967773,163.7265625,163.784469604492,163.7666015625,163.760940551758,164.2021484375,164.517379760742,164.572662353516,164.629287719727,164.66162109375,164.61572265625,164.27880859375,163.960052490234,163.635162353516],[35.8161125183105,35.8484382629395,35.8583984375,35.8273429870605,35.84228515625,35.7787094116211,35.6800765991211,35.6213874816895,35.55859375,35.5289039611816,35.5857429504395,35.6086921691895,35.7291030883789,35.8161125183105],[70.0206985473633,69.8447265625,69.6513671875,69.4693374633789,69.5027313232422,69.6164093017578,69.8003921508789,69.9175796508789,70.07666015625,70.0576171875,70.0572280883789,70.1100616455078,70.0591812133789,70.0206985473633],[42.7136688232422,42.6755867004395,42.4773406982422,42.4600601196289,42.4685554504395,42.5474624633789,42.6314468383789,42.6907196044922,42.7136688232422],[50.2652320861816,50.2830085754395,50.2206039428711,50.1644554138184,50.1409187316895,50.0939445495605,49.9207992553711,49.83984375,49.6262702941895,49.1804695129395,48.9103546142578,48.6669921875,48.4390640258789,48.31591796875,48.29443359375,48.27880859375,48.2802734375,48.2962875366211,48.3199195861816,48.4138679504395,48.63134765625,48.8449249267578,48.9533233642578,49.2251930236816,49.9962882995605,50.1672859191895,50.2652320861816],[67.3449172973633,67.2639617919922,67.0978546142578,67.0472640991211,67.02587890625,67.2161102294922,67.3289031982422,67.3449172973633],[161.467086791992,161.422470092773,161.456253051758,161.461135864258,161.364059448242,161.182525634766,161.136520385742,161.12548828125,161.16455078125,161.082809448242,161.110748291016,161.323348999023,161.409759521484,161.505172729492,161.520706176758,161.617782592773,161.609283447266,161.540328979492,161.374420166016,161.350891113281,161.372665405273,161.377532958984,161.394241333008,161.494720458984,161.516998291016,161.506729125977,161.467086791992],[169.200775146484,168.915725708008,168.348037719727,168.144332885742,167.99267578125,167.8212890625,167.788864135742,167.81396484375,167.86474609375,168.0595703125,168.1962890625,168.35791015625,169.374801635742,169.420700073242,169.43359375,169.418167114258,169.332427978516,169.299118041992,169.263381958008,169.245803833008,169.200775146484],[60.4504852294922,60.4806632995605,60.4772491455078,60.4402351379395,60.3271484375,60.2159194946289,60.0261764526367,59.9195289611816,59.8127937316895,59.724609375,59.6370124816895,59.5782241821289,59.5812492370605,59.5026359558105,59.3815422058105,59.2683601379395,59.1442413330078,59.0825233459473,59.0040016174316,58.9527359008789,58.6800804138184,58.6341819763184,58.6055641174316,58.5680656433105,58.4730453491211,58.5199203491211,58.6153297424316,58.6780242919922,58.7942428588867,59.0052719116211,59.0480422973633,59.0882797241211,59.3098640441895,59.4259796142578,59.5291023254395,59.6363296508789,59.9558563232422,60.1722640991211,60.3925819396973,60.4504852294922],[66.5609359741211,66.5685501098633,66.5158157348633,66.4486389160156,66.4076232910156,66.3948211669922,66.4181671142578,66.4402313232422,66.462890625,66.4577102661133,66.5609359741211],[160.718948364258,160.6513671875,160.504791259766,160.436904907227,160.4404296875,160.448532104492,160.565826416016,160.644912719727,160.718948364258],[52.9033203125,52.994140625,53.0740242004395,53.1414070129395,53.1925773620605,53.2051773071289,53.0714836120605,53.0481414794922,53.1057624816895,53.1209983825684,53.0226554870605,53.0044898986816,52.9496116638184,52.8353500366211,52.7889633178711,52.7383804321289,52.5465812683105,52.4254875183105,52.2894515991211,52.2496070861816,52.2398452758789,52.2965812683105,52.5125961303711,52.6173820495605,52.7296905517578,52.7203140258789,52.7322273254395,52.7767601013184,52.9033203125],[178.861526489258,178.792572021484,178.648239135742,178.628326416016,178.683883666992,178.829010009766,178.89111328125,179.235061645508,179.547653198242,179.715927124023,179.886428833008,180.000045776367,180.155136108398,180.308990478516,180.45361328125,180.597946166992,180.743515014648,180.888427734375,181.005950927734,181.12353515625,181.56103515625,181.646438598633,181.785308837891,181.866104125977,181.943359375,182.025192260742,182.182998657227,182.415863037109,182.467819213867,182.501510620117,182.476409912109,182.17822265625,181.937301635742,181.472015380859,180.843109130859,180.584320068359,180.493301391602,180.265975952148,180.000045776367,179.88134765625,179.647659301758,179.152542114258,178.861526489258],[137.959869384766,137.711822509766,137.612884521484,137.511810302734,137.457809448242,137.403213500977,137.34423828125,137.265533447266,137.078720092773,137.064056396484,137.081848144531,137.129486083984,137.168167114258,137.281829833984,137.816802978516,137.857620239258,137.933792114258,137.959869384766],[77.6325149536133,77.1456069946289,76.9059524536133,76.87109375,76.9031295776367,77.1495132446289,77.2604522705078,77.3778305053711,77.5787124633789,77.74853515625,78.2791061401367,78.3529281616211,78.3651351928711,78.1544952392578,78.0072250366211,77.7808609008789,77.6325149536133],[79.50146484375,79.4306640625,78.8805618286133,78.6902313232422,78.6332015991211,78.6568374633789,79.1642532348633,79.3565444946289,79.4124984741211,79.5413131713867,79.5378875732422,79.50146484375],[74.6605453491211,74.6383819580078,74.5880813598633,74.4347686767578,74.1806640625,74.1001968383789,74.1423873901367,74.1985397338867,74.4087905883789,74.5998992919922,74.7252960205078,74.9615249633789,74.7425765991211,74.6472702026367,74.66015625,74.6971664428711,74.6605453491211],[120.261329650879,120.007911682129,119.792091369629,119.640434265137,119.761917114258,119.964462280273,120.078506469727,120.23681640625,120.261329650879],[55.31982421875,55.7873039245605,56.1376953125,56.3504867553711,56.4295883178711,56.3974609375,56.3346710205078,56.18896484375,56.1669921875,56.19287109375,56.1705093383789,56.1216812133789,56.0837898254395,55.8197288513184,55.7234344482422,55.7184562683105,55.7009773254395,55.6164054870605,55.4413070678711,55.4033203125,55.4168930053711,55.35595703125,55.3595695495605,55.3904304504395,55.3991203308105,55.51806640625,55.4949226379395,55.4033203125,55.375,55.2978515625,55.4710960388184,55.5466804504395,55.6136703491211,55.8193359375,56.0431632995605,56.4543952941895,56.8948249816895,57.0656242370605,57.4835968017578,57.5564460754395,57.6253890991211,57.4471702575684,57.263671875,57.2469711303711,57.1458969116211,56.6488265991211,56.6216812133789,56.5686531066895,56.5100593566895,56.3857421875,56.2600593566895,56.3347663879395,56.4171867370605,56.5613288879395,56.4997062683105,56.4345703125,56.1424827575684,56.11474609375,56.0871086120605,55.9416007995605,55.9072265625,55.796875,55.7067375183105,55.7064476013184,55.6873054504395,55.2369155883789,55.0516586303711,54.8670921325684,54.6451148986816,54.6082038879395,54.6011695861816,54.5173797607422,54.3326187133789,54.1994171142578,53.7223663330078,53.3835945129395,53.4677734375,53.6135749816895,53.6156272888184,53.5925788879395,53.5877914428711,53.6705055236816,53.8570327758789,53.8342781066895,53.9222679138184,54.0939445495605,54.1556663513184,53.8861312866211,53.5908203125,53.6221694946289,53.5152359008789,53.4099617004395,53.3190422058105,53.33251953125,53.41162109375,53.3638687133789,52.9089851379395,52.6787109375,52.4188461303711,52.1799812316895,51.9378929138184,51.8125953674316,51.6916007995605,51.5904312133789,51.5113296508789,51.4386749267578,51.4286117553711,51.4435539245605,51.4822273254395,51.58251953125,51.6531257629395,51.8054695129395,51.8854484558105,52.0686531066895,52.2520484924316,52.3323249816895,52.40673828125,52.4619140625,52.5861320495605,52.6220703125,52.6619110107422,52.7057609558105,52.7138671875,52.7487297058105,52.8636741638184,52.8232421875,52.8390617370605,52.9165992736816,52.68310546875,52.60498046875,52.5285148620605,52.5505867004395,52.5792999267578,52.8122062683105,52.9131851196289,53.0242195129395,53.1349601745605,53.2535171508789,53.3698234558105,53.2472686767578,53.2371101379395,53.18896484375,53.1979484558105,53.2511711120605,53.3576202392578,53.51220703125,53.6336898803711,53.7532234191895,53.8656234741211,54.0910186767578,54.2023429870605,54.32763671875,54.6760749816895,54.8039093017578,54.9406242370605,55.1213836669922,55.31982421875],[70.6739273071289,70.38037109375,70.29833984375,70.11865234375,70.0407180786133,69.9201202392578,69.9303741455078,69.9856414794922,70.0187454223633,69.9959030151367,70.1496047973633,70.3499984741211,70.9402389526367,71.0232467651367,71.1412048339844,71.2316436767578,71.3511734008789,71.4449157714844,71.5895462036133,71.6304702758789,71.6261749267578,71.3556594848633,70.88671875,70.6739273071289],[76.7560577392578,76.6593704223633,76.2344741821289,76.0830993652344,76.1395492553711,76.2506790161133,76.7560577392578],[75.5037078857422,75.3443374633789,75.375,75.5697250366211,75.93017578125,76.0394515991211,76.0515594482422,75.9009780883789,75.8271484375,75.5037078857422],[142.184860229492,142.43505859375,142.63916015625,143.34375,143.410736083984,143.463973999023,143.491302490234,143.451477050781,143.193267822266,142.841598510742,142.5869140625,142.342193603516,142.126373291016,141.5966796875,141.182708740234,140.753997802734,140.662780761719,140.392471313477,140.026947021484,139.925094604492,139.785537719727,139.685546875,139.920120239258,140.155181884766,140.380661010742,140.593551635742,140.697463989258,140.8837890625,140.983581542969,141.084762573242,141.18994140625,141.311920166016,141.681930541992,141.931838989258,142.184860229492],[124.54296875,124.481742858887,124.366409301758,124.335739135742,124.336517333984,124.429679870605,124.547653198242,124.636909484863,124.652931213379,124.54296875],[83.5490264892578,83.4957962036133,83.4500045776367,83.41064453125,83.1589889526367,82.8177719116211,82.9029312133789,83.1498107910156,83.5134735107422,83.6183624267578,83.5490264892578],[82.7099609375,82.61279296875,82.4781265258789,82.3815383911133,82.3293914794922,82.3824234008789,82.5255813598633,82.6110382080078,82.68896484375,82.7099609375],[136.197448730469,136.121688842773,136.051467895508,135.714553833008,135.633407592773,135.448638916016,135.402435302734,135.387008666992,135.628326416016,136.036819458008,136.259170532227,136.197448730469],[141.01025390625,140.507217407227,140.409469604492,140.183197021484,140.1015625,140.193557739258,140.30029296875,140.407424926758,140.849227905273,140.9443359375,141.03857421875,141.079498291016,141.097457885742,141.046875,141.01025390625],[84.7589874267578,84.71044921875,84.4289016723633,84.3894500732422,84.5403289794922,84.6798858642578,84.8728561401367,84.7589874267578],[113.38720703125,113.353126525879,113.299217224121,113.258888244629,113.190238952637,112.977638244629,112.811325073242,112.782417297363,112.19580078125,112.105079650879,111.912109375,111.642967224121,111.50341796875,111.570114135742,111.637504577637,111.879783630371,111.94921875,111.982810974121,111.989349365234,112.007614135742,112.08447265625,112.951759338379,113.286224365234,113.38720703125],[86.6531295776367,86.7371139526367,87.0005874633789,87.0521469116211,87.1243133544922,87.01171875,86.9271469116211,86.6919937133789,86.3905258178711,86.2585983276367,86.3306579589844,86.5044860839844,86.60546875,86.6531295776367],[82.17236328125,82.2087936401367,82.2215881347656,82.1792984008789,82.0501022338867,81.978515625,81.9050827026367,81.8605499267578,81.6976547241211,81.65478515625,81.5792922973633,81.5321350097656,81.5005798339844,81.7121047973633,81.8421859741211,81.9265594482422,81.9097671508789,81.9127883911133,82.0218734741211,82.1656265258789,82.17236328125],[146.795211791992,147.060348510742,147.443557739258,147.496978759766,148.432434082031,148.508880615234,148.518844604492,148.489166259766,148.474990844727,148.590133666992,148.892181396484,149.083206176758,149.645309448242,150.103912353516,150.280670166016,150.417190551758,150.530563354492,150.612899780273,150.690338134766,150.756927490234,150.822372436523,150.646286010742,150.580276489258,150.331253051758,149.838073730469,149.596878051758,149.050186157227,148.296875,148.092376708984,147.971862792969,147.740905761719,147.626861572266,147.257034301758,147.14404296875,146.924896240234,146.703323364258,146.148529052734,146.186126708984,146.257614135742,146.342956542969,146.4384765625,146.537506103516,146.7509765625,146.748245239258,146.795211791992],[135.948638916016,135.745895385742,135.451950073242,135.473052978516,135.5234375,135.592666625977,135.561233520508,135.578414916992,135.613876342773,135.698638916016,135.788284301758,135.849227905273,135.90478515625,136.127349853516,136.1689453125,135.9833984375,135.965133666992,136.0205078125,135.948638916016],[152.885940551758,152.786331176758,152.55859375,152.642776489258,152.799423217773,152.835052490234,152.86376953125,152.885940551758],[140.048721313477,140.152145385742,140.2744140625,140.389053344727,140.496292114258,140.546676635742,140.602142333984,140.65673828125,140.81591796875,140.889266967773,140.944137573242,140.9404296875,140.926574707031,140.92578125,140.950302124023,140.9853515625,141.032608032227,141.29931640625,141.485443115234,141.742279052734,142.00146484375,142.460357666016,142.669540405273,142.9267578125,143.185165405273,143.311141967773,143.559967041016,143.685836791992,145.255264282227,145.309768676758,145.359970092773,145.0234375,144.803131103516,144.726760864258,144.814254760742,144.883499145508,144.407806396484,144.216018676758,144.019729614258,143.625869750977,143.396087646484,143.170318603516,142.922058105469,142.820114135742,142.7294921875,142.699600219727,142.734481811523,142.867568969727,142.986038208008,143.00244140625,142.941802978516,142.551559448242,142.307907104492,142.086227416992,142.151077270508,142.198822021484,142.264739990234,142.616790771484,142.696960449219,142.9296875,143.1279296875,142.778213500977,142.626068115234,142.472763061523,142.37841796875,142.287399291992,142.184188842773,142.099990844727,141.9873046875,141.748428344727,141.529968261719,141.310455322266,140.660736083984,140.4638671875,140.267868041992,140.011032104492,139.758193969727,139.681243896484,139.605850219727,139.548049926758,139.512313842773,139.430068969727,139.325592041016,139.21533203125,139.09912109375,138.981735229492,138.865631103516,138.09228515625,138.001358032227,137.9150390625,137.683013916016,137.568069458008,137.446975708008,137.217971801758,137.006240844727,136.962295532227,136.947662353516,136.982421875,137.166015625,137.289749145508,137.215240478516,137.268859863281,137.358489990234,137.70654296875,137.593566894531,137.501174926758,137.560546875,137.625381469727,137.7744140625,137.97705078125,138.038665771484,138.096008300781,138.207626342773,138.4306640625,138.81396484375,138.919525146484,139.017578125,139.109176635742,139.211334228516,139.528518676758,139.743362426758,140.048721313477],[96.5324249267578,96.6139602661133,96.5896453857422,96.4867248535156,96.3507843017578,96.3534164428711,96.3005828857422,96.1087875366211,95.8445358276367,95.6786117553711,95.3111267089844,95.3220748901367,95.3798828125,95.5944366455078,95.7862319946289,96.1509780883789,96.2707061767578,96.5324249267578],[96.8539047241211,96.7978515625,96.7544937133789,96.7393569946289,96.740234375,96.8329086303711,96.8352508544922,96.8779296875,96.990234375,97.0453186035156,97.0530242919922,96.9742202758789,96.8539047241211],[112.47802734375,112.632514953613,112.660842895508,112.614158630371,112.586524963379,112.574798583984,112.531639099121,112.394828796387,112.296875,112.15380859375,112.002731323242,111.968940734863,112.011131286621,112.281448364258,112.394142150879,112.47802734375],[97.58837890625,97.5352478027344,97.4303741455078,97.3416976928711,97.3103561401367,97.3816375732422,97.58837890625],[149.150192260742,148.398635864258,148.448150634766,148.719619750977,149.406433105469,149.268356323242,149.204788208008,149.150192260742],[67.7653350830078,67.365234375,67.126953125,66.8931655883789,66.6574249267578,66.2824249267578,65.619140625,65.2015609741211,64.7445297241211,64.2625961303711,63.779296875,63.6594734191895,63.3166999816895,63.0459938049316,62.0661125183105,61.6162071228027,61.4865226745605,61.35595703125,61.2488250732422,61.1472702026367,60.9356422424316,60.8292007446289,60.7192420959473,60.6553726196289,60.5337867736816,60.4755859375,60.27685546875,60.2411155700684,60.4548835754395,60.5013656616211,60.4391632080078,60.30078125,60.2224617004395,60.080078125,59.9823265075684,59.7472648620605,59.7346687316895,59.771484375,59.7527351379395,59.6740226745605,59.5959968566895,59.2401351928711,59.1820335388184,59.1570320129395,59.1460914611816,59.1009788513184,59.0404281616211,58.92822265625,58.53466796875,58.5021476745605,58.5620155334473,58.6457061767578,58.6650428771973,58.6178703308105,58.44140625,57.7673835754395,57.7784194946289,57.8534164428711,57.8722686767578,57.8449211120605,57.7559585571289,57.6574249267578,57.6037101745605,57.4485359191895,57.3130874633789,57.2909164428711,57.4642601013184,57.5425834655762,57.4597663879395,57.1343765258789,56.9638671875,56.6341819763184,56.4303703308105,56.2283210754395,56.0345726013184,55.5492172241211,55.2801780700684,55.0068359375,54.7686538696289,54.5658226013184,54.2999000549316,54.1315460205078,54.20458984375,53.8386726379395,53.7628898620605,53.8513679504395,53.9634780883789,54.1740226745605,54.3863258361816,54.6056632995605,54.6426734924316,54.7333984375,54.8312492370605,54.9203109741211,55.0228500366211,55.3409194946289,55.4164047241211,56.0783195495605,56.1371078491211,55.9474601745605,55.7517585754395,55.6615219116211,55.6103515625,55.5822257995605,55.65966796875,55.9136695861816,56.2178688049316,56.4987297058105,56.4285163879395,56.3400382995605,55.998046875,55.8631858825684,55.8211936950684,55.81005859375,55.9207000732422,56.0355491638184,56.1622085571289,56.2886734008789,56.3890609741211,56.4852561950684,56.5703125,56.8762702941895,56.8292999267578,56.8094749450684,56.8444328308105,56.9894523620605,57.0875015258789,57.3017578125,57.6068344116211,57.6315460205078,57.7081985473633,57.7834014892578,58.0936546325684,58.0725555419922,58.0583000183105,58.4183616638184,58.6527328491211,58.8812522888184,58.9947280883789,59.1104507446289,59.3465843200684,59.7819328308105,60.0361328125,60.1181640625,60.2792930603027,60.6061553955078,60.7305641174316,60.8011703491211,60.9421844482422,60.9977569580078,61.0539054870605,61.0369148254395,61.0343742370605,61.1569328308105,61.2016563415527,61.5694351196289,61.7871055603027,62.2373046875,62.4710960388184,62.7820320129395,62.9714813232422,63.5261688232422,64.4634780883789,64.7076110839844,64.9499969482422,65.0728530883789,65.1971664428711,65.3097610473633,65.5284194946289,65.6369171142578,65.7551727294922,65.8628921508789,65.9588851928711,66.06298828125,66.34521484375,66.8288116455078,67.263671875,67.5349655151367,67.65185546875,68.0172882080078,68.4857406616211,68.6991195678711,68.8733367919922,68.9117202758789,68.9416961669922,68.8905258178711,68.8580093383789,68.8998031616211,68.55859375,68.2223587036133,68.1654281616211,67.7653350830078],[96.2854461669922,96.2535095214844,96.2098617553711,96.0914077758789,95.8546905517578,95.7658233642578,95.6808624267578,95.3640670776367,95.2703094482422,95.4207000732422,95.8541030883789,96.5284194946289,96.5619125366211,96.5613250732422,96.42431640625,96.2854461669922],[89.5142593383789,89.2995147705078,89.1792907714844,89.1417007446289,89.2004852294922,89.2815399169922,89.6162109375,89.6795883178711,89.6658248901367,89.5142593383789],[107.414741516113,107.30224609375,107.26953125,107.366409301758,107.486427307129,107.593650817871,107.629295349121,107.66455078125,107.679489135742,107.414741516113],[130.687316894531,130.658004760742,130.651565551758,130.617965698242,130.554107666016,130.526947021484,130.58447265625,130.576568603516,130.520614624023,130.439270019531,130.419921875,130.4248046875,130.452728271484,130.492965698242,130.577255249023,130.722457885742,130.803314208984,130.868560791016,130.94287109375,131.005569458008,131.068557739258,131.08349609375,131.086135864258,131.108978271484,131.135543823242,131.175582885742,131.2119140625,131.239349365234,131.25732421875,131.261810302734,131.243942260742,131.209182739258,131.182418823242,131.180084228516,131.18359375,131.174224853516,131.213272094727,131.255264282227,131.125778198242,131.0869140625,131.060653686523,131.00390625,130.9677734375,130.981643676758,131.033020019531,131.082321166992,131.227935791016,131.268264770508,131.446884155273,131.487487792969,131.578720092773,131.613967895508,131.654006958008,131.742095947266,131.794921875,131.851852416992,131.909271240234,131.9775390625,132.0673828125,132.181350708008,132.362991333008,132.549026489258,132.665618896484,132.72314453125,132.838668823242,132.888778686523,132.93603515625,133.01171875,133.113479614258,133.096878051758,133.113372802734,133.18603515625,133.266983032227,133.3095703125,133.35546875,133.436416625977,133.465621948242,133.449127197266,133.475769042969,133.484664916992,133.513076782227,133.551162719727,133.608001708984,133.647857666016,133.685729980469,133.711135864258,133.70068359375,133.750198364258,133.832809448242,133.861328125,133.874801635742,133.880279541016,133.902725219727,133.88671875,133.866592407227,133.95751953125,134.022659301758,134.03857421875,134.046005249023,134.071380615234,134.08642578125,134.136917114258,134.2021484375,134.189254760742,134.162994384766,134.167678833008,134.225204467773,134.260055541992,134.290817260742,134.339447021484,134.382522583008,134.483505249023,134.541900634766,134.59619140625,134.69580078125,134.728134155273,134.752334594727,134.698638916016,134.650299072266,134.59130859375,134.566009521484,134.605361938477,134.647262573242,134.669342041016,134.680847167969,134.665237426758,134.563583374023,134.456146240234,134.3349609375,134.293365478516,134.205856323242,133.842193603516,133.671768188477,133.5732421875,133.468353271484,133.301162719727,133.14404296875,133.020126342773,132.877151489258,132.772857666016,132.707229614258,132.636917114258,132.561904907227,132.476272583008,132.380172729492,132.149810791016,131.785247802734,131.556732177734,131.464263916016,131.3193359375,131.121871948242,131.002731323242,130.9619140625,130.932815551758,130.915420532227,130.8486328125,130.732620239258,130.712097167969,130.787200927734,130.804290771484,130.763473510742,130.746871948242,130.6591796875,130.597259521484,130.552154541016,130.565628051758,130.6171875,130.553131103516,130.355270385742,130.195999145508,130.037109375,129.792587280273,129.671096801758,129.591400146484,129.53369140625,129.498153686523,129.440719604492,129.384674072266,129.35009765625,129.309860229492,129.248428344727,129.185165405273,129.1201171875,129.065139770508,129.020309448242,128.938278198242,128.8193359375,128.770309448242,128.791015625,128.76904296875,128.703994750977,128.526763916016,128.237121582031,127.999603271484,127.814262390137,127.711135864258,127.690139770508,127.636711120605,127.55078125,127.502449035645,127.491790771484,127.512306213379,127.590225219727,127.395309448242,127.337211608887,127.351173400879,127.3408203125,127.306053161621,127.308197021484,127.34716796875,127.346870422363,127.307037353516,127.1982421875,127.020416259766,126.9248046875,126.911529541016,126.887687683105,126.854400634766,126.833786010742,126.847763061523,126.827339172363,126.801750183105,126.805465698242,126.774513244629,126.709175109863,126.688667297363,126.700790405273,126.653717041016,126.510551452637,126.468070983887,126.455574035645,126.394821166992,126.391502380371,126.383499145508,126.346290588379,126.324211120605,126.341697692871,126.312889099121,126.237594604492,126.202934265137,126.194435119629,126.156639099121,126.0458984375,126.016021728516,126.023246765137,126.047073364258,126.060157775879,126.056060791016,126.048149108887,126.004302978516,125.941596984863,125.871879577637,125.782814025879,125.728126525879,125.680770874023,125.6953125,125.691696166992,125.649017333984,125.595993041992,125.545997619629,125.422462463379,125.225593566895,125.074996948242,124.970893859863,124.906646728516,124.882125854492,124.812309265137,124.639846801758,124.465919494629,124.369140625,124.291412353516,124.219924926758,124.154304504395,123.994720458984,123.740913391113,123.607818603516,123.559761047363,123.534759521484,123.489448547363,123.424018859863,123.3095703125,123.154106140137,122.957611083984,122.744728088379,122.515823364258,122.380172729492,122.337799072266,122.0888671875,121.743942260742,121.405464172363,120.985450744629,120.7041015625,120.421287536621,120.218162536621,120.094535827637,120.044334411621,120.067581176758,120.172752380371,120.360061645508,120.521095275879,120.656150817871,120.699226379395,120.650390625,120.665428161621,120.744537353516,120.749809265137,120.681449890137,120.510543823242,120.237007141113,120.06689453125,119.966987609863,119.813186645508,119.756645202637,119.746002197266,119.684951782227,119.573432922363,119.512298583984,119.501754760742,119.445709228516,119.344047546387,119.280662536621,119.255851745605,119.216705322266,119.163673400879,119.19189453125,119.301559448242,119.346282958984,119.326065063477,119.259857177734,119.1474609375,118.9794921875,118.755950927734,118.451560974121,118.186614990234,117.873435974121,117.812591552734,117.698432922363,117.477149963379,117.24560546875,117.021682739258,116.888961791992,116.683303833008,116.631546020508,116.551170349121,116.351165771484,116.216804504395,116.134567260742,115.925971984863,115.795219421387,115.7177734375,115.587989807129,115.42919921875,115.365036010742,115.274513244629,115.098045349121,115.003326416016,114.879585266113,114.7431640625,114.674903869629,114.554008483887,114.386322021484,114.297065734863,114.221778869629,114.070701599121,113.881149291992,113.732421875,113.57421875,113.445510864258,113.319038391113,113.164161682129,113.092094421387,113.055564880371,112.914840698242,112.806449890137,112.697364807129,112.494926452637,112.375198364258,112.079689025879,111.934471130371,111.833396911621,111.735549926758,111.574798583984,111.511917114258,111.429298400879,111.336616516113,111.204200744629,110.827926635742,110.709762573242,110.631050109863,110.529594421387,110.427833557129,110.321388244629,110.199905395508,109.994537353516,109.750389099121,109.528709411621,109.453712463379,109.236717224121,108.919921875,108.733001708984,108.613677978516,108.5224609375,108.406929016113,108.213081359863,108.098045349121,108.033790588379,108.009574890137,107.965431213379,107.936721801758,107.938774108887,107.934860229492,107.947853088379,107.916603088379,107.786819458008,107.630950927734,107.347068786621,107.233299255371,107.14306640625,107.040229797363,106.941307067871,106.853713989258,106.711135864258,106.574409484863,106.368453979492,106.217872619629,106.08251953125,105.996482849121,105.875190734863,105.692581176758,105.541603088379,105.383598327637,105.266700744629,105.185935974121,105.0947265625,104.976951599121,104.685356140137,104.596382141113,104.46630859375,104.353904724121,104.259963989258,104.1796875,104.078712463379,103.95849609375,103.856155395508,103.802635192871,103.723236083984,103.632911682129,103.496292114258,103.421195983887,103.304389953613,103.233787536621,103.161712646484,103.039451599121,102.859672546387,102.765426635742,102.683303833008,102.546287536621,102.469436645508,102.406837463379,102.33642578125,102.288383483887,102.285743713379,102.303314208984,102.316604614258,102.276565551758,102.235054016113,102.215034484863,102.226173400879,102.210258483887,102.194534301758,102.151954650879,102.142387390137,102.160064697266,102.155662536621,102.111526489258,101.979194641113,101.821189880371,101.570899963379,101.46435546875,101.381248474121,101.304489135742,101.223236083984,101.085350036621,100.903610229492,100.710746765137,100.536231994629,100.468948364258,100.230369567871,100.034568786621,99.9216766357422,99.7878875732422,99.71923828125,99.6128921508789,99.5323257446289,99.4070281982422,99.1761703491211,99.0914077758789,99.0342788696289,98.9580993652344,98.8931579589844,98.8486328125,98.8025360107422,98.7601547241211,98.6405258178711,98.3527374267578,98.3031234741211,98.27685546875,98.2374954223633,98.2199249267578,98.1846694946289,98.1031265258789,98.03759765625,97.9891586303711,97.9468765258789,97.9232406616211,97.9273452758789,97.9178695678711,97.9108352661133,97.8357391357422,97.8252944946289,97.8561553955078,97.9198226928711,97.953125,97.9641571044922,97.9619140625,98.0011749267578,98.02978515625,98.0789108276367,98.14501953125,98.2205123901367,98.2794876098633,98.2926788330078,98.27734375,98.2502899169922,98.1999969482422,98.1701202392578,98.1219711303711,98.1034240722656,98.00390625,97.9366226196289,97.8539047241211,97.7855453491211,97.720703125,97.6509780883789,97.58935546875,97.5408172607422,97.4183578491211,97.3597640991211,97.2085876464844,97.1369171142578,97.09765625,97.0491180419922,96.9857406616211,96.7117233276367,96.6402359008789,96.5984344482422,96.5432586669922,96.5057678222656,96.4664077758789,96.3811569213867,96.3150405883789,96.2296905517578,96.1117172241211,96.0655212402344,96.0185546875,95.9895553588867,95.9357452392578,95.8994140625,95.8519515991211,95.7893524169922,95.7078170776367,95.5671844482422,95.5226516723633,95.4417953491211,95.3856506347656,95.3294906616211,95.1662139892578,95.1114273071289,95.0443344116211,95.0128936767578,94.9302749633789,94.8112258911133,94.7180633544922,94.6754837036133,94.61474609375,94.5646438598633,94.4968795776367,94.45849609375,94.4001922607422,94.3546829223633,94.3468780517578,94.3193359375,94.2870101928711,94.2510681152344,94.0757827758789,93.9898452758789,93.79541015625,93.6620101928711,93.6255798339844,93.5010757446289,93.3868103027344,93.2705078125,93.2225646972656,93.1031265258789,93.0098648071289,92.970703125,92.9635772705078,92.9413070678711,92.8564453125,92.779296875,92.7386703491211,92.6813507080078,92.6266632080078,92.5789031982422,92.4864273071289,92.4263687133789,92.3547821044922,92.2957992553711,92.2790069580078,92.2653274536133,92.1923828125,92.10400390625,91.95654296875,91.8042984008789,91.7063446044922,91.6341781616211,91.5968704223633,91.5216827392578,91.4464874267578,91.4150390625,91.3408203125,91.3005828857422,91.2337875366211,91.0627899169922,91.0215835571289,90.9171905517578,90.8380889892578,90.7607421875,90.71435546875,90.6550827026367,90.5168991088867,90.3648452758789,90.3113327026367,90.2245101928711,90.1037139892578,90.0537109375,90.0049819946289,89.9773483276367,89.8780288696289,89.7442398071289,89.6438522338867,89.63427734375,89.6695327758789,89.6540985107422,89.5792007446289,89.4749984741211,89.3956069946289,89.2992172241211,89.2439498901367,89.2029342651367,89.1799850463867,89.1094741821289,89.0083923339844,88.9706039428711,88.9454040527344,88.9001922607422,88.8638687133789,88.8603515625,88.8316421508789,88.7478485107422,88.6827087402344,88.6332015991211,88.5443420410156,88.4524383544922,88.3933563232422,88.3377914428711,88.1925735473633,88.1355438232422,88.13427734375,88.11572265625,88.0285186767578,87.9880905151367,87.9347686767578,87.8182601928711,87.8142623901367,87.7625045776367,87.6683578491211,87.5765609741211,87.5158233642578,87.4761734008789,87.4167022705078,87.3228454589844,87.296875,87.2336959838867,87.1480407714844,87.0706024169922,87.0009765625,86.9529342651367,86.8121109008789,86.71435546875,86.62646484375,86.6142578125,86.6653366088867,86.7306671142578,86.7287139892578,86.6754913330078,86.6101608276367,86.5222702026367,86.41796875,86.2923812866211,86.2421875,86.1808624267578,86.0929718017578,86.0295944213867,85.9744110107422,85.93359375,85.8804702758789,85.4984359741211,85.37158203125,85.2918930053711,85.2326202392578,85.2101516723633,85.1365280151367,85.0764694213867,85.0007858276367,84.9751968383789,84.9997100830078,84.9894485473633,84.9240264892578,84.8389663696289,84.6073226928711,84.4990234375,84.4009780883789,84.3232421875,84.2578125,84.1945343017578,84.1759719848633,84.0993118286133,84.0023422241211,83.9451141357422,83.8597640991211,83.7177734375,83.5814514160156,83.3573226928711,83.2737274169922,83.1602478027344,83.0927734375,83.0192337036133,82.9190444946289,82.7608413696289,82.7185592651367,82.6929702758789,82.6117172241211,82.4939422607422,82.3263702392578,82.2119140625,82.0980453491211,81.9336929321289,81.7520523071289,81.6338882446289,81.4659194946289,81.4314498901367,81.4515609741211,81.4376983642578,81.41015625,81.3882827758789,81.3191375732422,81.1246109008789,81.0714797973633,81.0775451660156,81.1124038696289,81.1410140991211,81.1272430419922,81.0267562866211,80.9656219482422,80.93408203125,80.8773422241211,80.8130874633789,80.7352523803711,80.6504898071289,80.60546875,80.5506820678711,80.4910125732422,80.4480438232422,80.4214782714844,80.43359375,80.4522476196289,80.4236297607422,80.34521484375,80.2704086303711,80.22021484375,80.1272430419922,80.0863342285156,80.0720672607422,80.06591796875,79.9862289428711,79.8596725463867,79.7164077758789,79.5542907714844,79.4688491821289,79.1487274169922,78.9920883178711,78.7214813232422,78.4754867553711,78.1980514526367,78.0335006713867,77.8599624633789,77.7994155883789,77.7043914794922,77.46923828125,77.1324234008789,76.8207015991211,76.57568359375,76.5130844116211,76.4847640991211,76.4585952758789,76.4220733642578,76.4216842651367,76.6545944213867,76.7030258178711,76.7889633178711,76.8373031616211,76.7593765258789,76.6155319213867,76.5391616821289,76.4964828491211,76.2666015625,76.1405258178711,75.8806610107422,75.69287109375,75.6568374633789,75.4372024536133,75.3981475830078,75.3923797607422,75.3770523071289,75.22021484375,75.0521469116211,74.9889678955078,74.8868179321289,74.8341827392578,74.6814422607422,74.4519500732422,74.4304733276367,74.4292984008789,74.4027328491211,74.3515625,74.27734375,74.2099609375,74.0686569213867,73.8589782714844,73.7311553955078,73.6429672241211,73.4699172973633,73.4069290161133,73.3718719482422,73.3619155883789,73.3268585205078,73.2857437133789,73.3056640625,73.3994140625,73.55419921875,73.6789016723633,73.7155303955078,73.71240234375,73.6664047241211,73.6179733276367,73.5899429321289,73.5056610107422,73.3806610107422,73.2765655517578,73.2298812866211,73.1193313598633,72.9140625,72.7410125732422,72.6222686767578,72.5827102661133,72.5642547607422,72.5755920410156,72.5992202758789,72.5859375,72.5302734375,72.44677734375,72.404296875,72.3830032348633,72.3873062133789,72.3294906616211,72.2691421508789,72.18603515625,72.1053695678711,72.0656280517578,72.0044860839844,71.8873977661133,71.6771469116211,71.33642578125,71.0931625366211,71.052734375,71.1521530151367,71.1597671508789,71.1591796875,71.185546875,71.1262741088867,70.9917984008789,70.91015625,70.7903366088867,70.7380828857422,70.486328125,70.4171905517578,70.3714904785156,70.2933578491211,70.1824188232422,70.08740234375,69.9817352294922,69.8702087402344,69.740234375,69.4932632446289,69.2469711303711,68.9772491455078,68.8429641723633,68.712890625,68.5248031616211,68.4384765625,68.3019561767578,68.2062454223633,68.2252883911133,68.2440414428711,68.2093734741211,68.1558609008789,68.0738296508789,67.93994140625,67.8298873901367,67.693359375,67.4846649169922,67.25732421875,67.0983352661133,66.7544937133789,66.5553741455078,66.22265625,65.9546890258789,65.9142532348633,65.7078094482422,65.4769515991211,65.4343719482422,65.3781280517578,65.31591796875,65.2374038696289,65.1921920776367,65.1578140258789,65.08837890625,64.9954071044922,64.9267578125,64.8092727661133,64.64990234375,64.5251007080078,64.4612274169922,64.1994171142578,64.0628890991211,64.0373992919922,64.00390625,63.8470726013184,63.72119140625,63.7013702392578,63.58203125,63.4136695861816,63.2926788330078,63.1913032531738,63.1265602111816,63.0739288330078,62.6327133178711,62.5882797241211,62.4990234375,62.0402336120605,62.0023460388184,61.9856452941895,61.9287109375,61.59814453125,61.3336906433105,61.2310562133789,61.1437492370605,61.1131858825684,61.1131858825684,61.1131858825684,61.0735359191895,60.9855499267578,60.9794921875,61.0985336303711,61.2479515075684,61.3361282348633,61.4099578857422,61.47412109375,61.5191383361816,61.5349655151367,61.5265617370605,61.4985389709473,61.4009742736816,61.3116226196289,61.2289047241211,61.1859397888184,61.1627922058105,61.19921875,61.3109397888184,61.4368133544922,61.576171875,61.6598587036133,61.7662124633789,62.0146484375,62.0810585021973,62.0827178955078,62.0371131896973,61.9742202758789,61.8885765075684,61.7193374633789,61.5335922241211,61.4007835388184,61.2069358825684,61.0474586486816,61.0065460205078,60.9447250366211,60.8932609558105,60.8023452758789,60.7744140625,60.8212890625,60.9794921875,60.9945335388184,60.9375953674316,60.8284149169922,60.6703109741211,60.4993171691895,60.4254875183105,60.2336883544922,60.0655288696289,60.0302734375,60.0674858093262,60.2803726196289,60.3874969482422,60.4183578491211,60.4647483825684,60.6303749084473,60.9735374450684,60.9933586120605,61.0148429870605,61.3630905151367,61.4113311767578,61.5546875,61.5850563049316,61.51220703125,61.4650344848633,61.3894538879395,61.2268524169922,60.9422874450684,60.6379890441895,60.5084953308105,60.4248046875,60.2880859375,60.1867179870605,60.1121139526367,60.0585899353027,60.0052757263184,59.9551734924316,59.8877944946289,59.8124046325684,59.7511711120605,59.5230484008789,59.4978523254395,59.5239219665527,59.4951133728027,59.4523429870605,59.1708984375,59.0643539428711,58.98486328125,58.8836898803711,58.8140640258789,58.66455078125,58.5474586486816,58.3591766357422,58.1884765625,58.1747093200684,58.0451202392578,57.8388671875,57.8289031982422,57.7648429870605,57.7169914245605,57.6538124084473,57.5578117370605,57.4421844482422,57.3125,57.1790046691895,57.01171875,56.849609375,56.7903327941895,56.6202163696289,56.56689453125,56.4914054870605,56.3255844116211,56.1439476013184,56.1044921875,56.0497055053711,55.92919921875,55.7976570129395,55.6862335205078,55.5422859191895,55.3611335754395,55.1952171325684,55.0148429870605,54.8679695129395,54.7271461486816,54.6416015625,54.5729484558105,54.5460929870605,54.5656242370605,54.6062507629395,54.6378898620605,54.6500015258789,54.6361351013184,54.59619140625,54.5552711486816,54.5173797607422,54.4714813232422,54.443359375,54.4214859008789,54.2978515625,54.1911125183105,54.1397476196289,54.04150390625,53.9568328857422,53.7764625549316,53.6880874633789,53.53466796875,53.4486312866211,53.3380851745605,53.2472686767578,53.2273445129395,53.0383796691895,52.9026374816895,52.8205070495605,52.7350578308105,52.7281265258789,52.6351547241211,52.6177711486816,52.5711898803711,52.4961929321289,52.4230461120605,52.3310546875,52.2191429138184,52.0071296691895,51.775390625,51.6090850830078,51.4734344482422,51.39599609375,51.3445281982422,51.3010711669922,51.2907218933105,51.2699203491211,51.1634750366211,51.0178718566895,50.8824234008789,50.7939453125,50.7561531066895,50.6439437866211,50.5163078308105,50.3537101745605,50.3092765808105,50.2468757629395,50.1048851013184,49.9323234558105,49.822265625,49.6663093566895,49.498046875,49.4246063232422,49.3794898986816,49.3234367370605,49.0586891174316,48.9137687683105,48.8083992004395,48.7347640991211,48.6551780700684,48.6250953674316,48.666015625,48.7004890441895,48.7494125366211,48.7847671508789,48.8179664611816,48.84326171875,48.8102531433105,48.7589874267578,48.5999984741211,48.4342765808105,48.3349609375,48.2248039245605,48.1813468933105,48.0607414245605,47.849609375,47.7057609558105,47.599609375,47.5036125183105,47.42919921875,47.3763656616211,47.3264656066895,47.2947273254395,47.2976570129395,47.2952156066895,47.2483406066895,47.1295890808105,46.9919929504395,46.8895530700684,46.8231468200684,46.8020515441895,46.8529281616211,46.9534187316895,47.0181655883789,47.0313491821289,47.0142593383789,46.9622077941895,46.8529281616211,46.70263671875,46.6091804504395,46.6609344482422,46.8531227111816,47.0042953491211,47.0646476745605,47.1190452575684,47.1115226745605,47.09326171875,47.1307601928711,47.2020492553711,47.2923851013184,47.3873062133789,47.48193359375,47.6001968383789,47.9346656799316,48.1099586486816,48.1669921875,48.2756843566895,48.4130859375,48.5525398254395,48.6006813049316,48.71435546875,48.8318367004395,48.9593734741211,48.9502906799316,48.8835906982422,48.7763671875,48.6935539245605,48.6470680236816,48.6052703857422,48.5583953857422,48.5185546875,48.5023460388184,48.5091781616211,48.5412101745605,48.5860366821289,48.6101570129395,48.7743148803711,48.958984375,49.1842765808105,49.2322273254395,49.2458992004395,49.12548828125,49.1106452941895,49.07958984375,48.8099632263184,48.7425804138184,48.6836929321289,48.6873054504395,48.7034187316895,48.7496070861816,48.7295875549316,48.6896476745605,48.6374015808105,48.5890617370605,48.5373039245605,48.4870109558105,48.2576141357422,48.1591796875,48.0528335571289,47.8301773071289,47.7639617919922,47.7010765075684,47.6498031616211,47.63330078125,47.5740242004395,47.5083999633789,47.4793930053711,47.4632835388184,47.5242156982422,47.5294952392578,47.5145492553711,47.4886741638184,47.4544906616211,47.4130859375,47.39111328125,47.3512687683105,47.2961921691895,47.2214813232422,47.1615257263184,47.11474609375,47.0837898254395,47.0392570495605,47.0029296875,46.9836921691895,46.9574203491211,46.8412132263184,46.7552757263184,46.7161140441895,46.7072257995605,46.7208976745605,46.7530288696289,46.9157218933105,47.0236320495605,47.1226577758789,47.2298851013184,47.3070297241211,47.3615226745605,47.42919921875,47.4627914428711,47.5625953674316,47.646484375,47.6278305053711,47.5679664611816,47.5089874267578,47.4898452758789,47.5116233825684,47.5128898620605,47.4631843566895,47.4888687133789,47.5290069580078,47.6348609924316,47.7090835571289,47.7277336120605,47.7697257995605,47.8223648071289,48.0801734924316,48.2286148071289,48.3030242919922,48.3837890625,48.4263648986816,48.4767570495605,48.5728530883789,48.5186500549316,48.4306640625,48.3914070129395,48.2981452941895,48.1422843933105,48.0560531616211,47.9636726379395,47.8611335754395,47.791015625,47.591796875,47.5206031799316,47.3176765441895,47.2611312866211,47.2052764892578,47.142578125,47.06396484375,47.0101585388184,46.98779296875,46.9308586120605,46.8255844116211,46.7493133544922,46.6903343200684,46.6160163879395,46.5712890625,46.5521469116211,46.5376968383789,46.4298858642578,46.4115257263184,46.2677726745605,46.2126960754395,46.1597633361816,46.0484390258789,45.9540061950684,45.9103546142578,45.8459968566895,45.7265625,45.6385765075684,45.63427734375,45.6883773803711,45.7275390625,45.7052726745605,45.6555671691895,45.5628890991211,45.34375,45.2082023620605,45.1602554321289,45.0715827941895,44.943359375,44.8709945678711,44.8504867553711,44.7710952758789,44.6917953491211,44.6443367004395,44.5764656066895,44.505859375,44.3294906616211,44.19970703125,44.1027336120605,44.0046882629395,43.9574203491211,43.8259773254395,43.7598648071289,43.7383804321289,43.7499008178711,43.79541015625,43.7987327575684,43.7826194763184,43.623046875,43.5578117370605,43.3479499816895,43.0891609191895,43.0001983642578,42.9916000366211,42.8900413513184,42.7606468200684,42.6602554321289,42.5660171508789,42.4190444946289,42.2796859741211,42.1222648620605,42.0877914428711,42.0500030517578,41.58056640625,41.4607391357422,41.3582038879395,41.0831069946289,40.9419898986816,40.8016624450684,40.6480484008789,40.5189476013184,40.34228515625,40.1501960754395,40.0845718383789,40.0237312316895,39.9783210754395,39.8736343383789,39.5166969299316,39.3293952941895,38.71728515625,38.6358375549316,38.3118133544922,38.1812477111816,37.8514633178711,37.7048835754395,37.5724601745605,37.4951171875,37.4113273620605,37.3523445129395,37.2840843200684,37.2047843933105,36.9444351196289,36.6507797241211,36.6276359558105,36.619140625,36.873046875,36.9412117004395,36.81103515625,36.7616233825684,36.7204132080078,36.7937507629395,36.8659172058105,36.9778327941895,37.103515625,37.2135734558105,37.2642593383789,37.6471710205078,37.6729507446289,37.671875,37.6343765258789,37.6099624633789,37.6124038696289,37.6692390441895,37.8409194946289,37.93310546875,38.0142593383789,38.0738258361816,38.0697250366211,38.07958984375,38.1328125,38.18359375,38.3118133544922,38.400390625,38.4922866821289,38.3152351379395,38.0777359008789,37.9775390625,37.9138679504395,37.8095703125,37.7665023803711,37.8673820495605,37.9679679870605,38.1594696044922,38.22998046875,38.3434562683105,38.5009765625,38.4879913330078,38.4386711120605,38.6307601928711,38.8010749816895,39.1267585754395,39.2707023620605,39.2890625,39.29345703125,39.2445335388184,39.1957015991211,39.0237274169922,38.9283218383789,38.6681632995605,38.5524444580078,38.6443367004395,38.7360343933105,38.7619132995605,38.5772476196289,38.4847640991211,38.21435546875,38.2058601379395,38.2013664245605,38.22119140625,38.2653312683105,38.28076171875,38.28076171875,38.2410163879395,38.2080078125,38.2013664245605,38.21240234375,38.2432594299316,38.2565422058105,38.2587890625,38.2873992919922,38.3688468933105,38.5109405517578,38.640625,38.7189445495605,38.822265625,38.9002952575684,39.0578117370605,39.1584968566895,39.3910140991211,39.6584968566895,39.7359352111816,39.7787094116211,39.7757797241211,39.81396484375,39.8850555419922,39.9610366821289,39.9579124450684,39.9181632995605,39.8663063049316,39.8474617004395,39.8499031066895,39.8898468017578,39.8826179504395,39.8575210571289,39.8356437683105,39.7654304504395,39.6447257995605,39.67041015625,39.70458984375,39.755859375,39.7928695678711,39.9041023254395,39.9844741821289,40.0036125183105,39.9891586303711,39.86376953125,39.7533187866211,39.7056655883789,39.6865234375,39.7594718933105,39.8897476196289,39.9763679504395,40.0700187683105,40.1087913513184,40.1283226013184,40.1261711120605,40.0578117370605,40.0578117370605,40.0949211120605,40.0806617736816,40.0306625366211,39.95849609375,39.8768539428711,39.7805671691895,39.6265602111816,39.4627952575684,39.3684539794922,39.3029289245605,39.2459983825684,39.2118148803711,39.1748046875,39.1149406433105,39.0277366638184,38.9183616638184,38.7766609191895,38.6477546691895,38.5519561767578,38.451171875,38.2585945129395,38.2086944580078,38.1775398254395,38.1626968383789,38.1467781066895,38.1124992370605,38.046875,37.9502944946289,37.7041969299316,37.6050796508789,37.5823249816895,37.5013694763184,37.4228515625,37.3431663513184,37.2548828125,37.1710968017578,37.1312522888184,36.9884757995605,36.7590827941895,36.6963882446289,36.6194343566895,36.5596656799316,36.4998016357422,36.3688468933105,36.3060531616211,36.2433624267578,36.189453125,36.1164054870605,36.0078125,35.8902320861816,35.7961921691895,35.6737289428711,35.5911102294922,35.5455055236816,35.4884796142578,35.41162109375,35.3917007446289,35.41162109375,35.4401359558105,35.4401359558105,35.4173812866211,35.3832054138184,35.3460922241211,35.3147468566895,35.30908203125,35.3347663879395,35.3119125366211,35.2691383361816,35.1980476379395,35.1581039428711,35.1153297424316,35.0925788879395,35.0640640258789,34.990234375,34.8685531616211,34.7603530883789,34.7123069763184,34.6167945861816,34.4910163879395,34.2341766357422,34.2138671875,34.2284164428711,34.2806663513184,34.2749977111816,34.2298812866211,34.20654296875,34.2092742919922,34.2008781433105,34.1467742919922,34.1154289245605,34.12109375,34.2391624450684,34.3792953491211,34.4027328491211,34.3978500366211,34.1130867004395,34.0153312683105,33.9220695495605,33.81884765625,33.7352523803711,33.6133804321289,33.4518547058105,33.287109375,33.1484375,32.8997077941895,32.8064460754395,32.6454124450684,32.5079078674316,32.4354476928711,32.3913078308105,32.3629875183105,32.2828140258789,32.216796875,32.1222686767578,32.0416030883789,31.9738292694092,31.8755874633789,31.7824211120605,31.7633781433105,31.7585926055908,31.6906261444092,31.6499004364014,31.6015605926514,31.5773448944092,31.5765628814697,31.5855464935303,31.6159172058105,31.5261726379395,31.5194339752197,31.5634746551514,31.5648441314697,31.5351581573486,31.4427719116211,31.35302734375,31.2951183319092,31.2587871551514,31.3029308319092,31.3645496368408,31.3883800506592,31.4178695678711,31.5629863739014,31.6682605743408,31.7474594116211,31.7774410247803,31.8497085571289,32.0554695129395,32.1419906616211,32.2506866455078,32.42626953125,32.4693374633789,32.5780296325684,32.6444358825684,32.7042999267578,32.7102546691895,32.7064476013184,32.6857452392578,32.4696311950684,32.4423828125,32.4251976013184,32.4509773254395,32.4501953125,32.2003898620605,31.9921855926514,31.8207988739014,31.7541980743408,31.7830085754395,31.8252944946289,31.8377933502197,31.8259773254395,31.7919940948486,31.6284160614014,31.4036140441895,31.2991199493408,31.2455062866211,31.1847648620605,31.0748062133789,31.0819339752197,31.1548824310303,31.1521472930908,31.1212882995605,30.9841804504395,30.798828125,30.791015625,30.8044910430908,30.8298816680908,30.8667964935303,30.9777355194092,30.9777355194092,30.9588871002197,30.87744140625,30.814453125,30.810546875,30.8209972381592,30.86181640625,30.9005851745605,30.9087886810303,30.9068355560303,30.8822250366211,30.85595703125,30.8007831573486,30.7216777801514,30.6623039245605,30.6255855560303,30.5867176055908,30.4753913879395,30.4562511444092,30.2335929870605,30.0426750183105,29.9370136260986,29.8816413879395,29.8239250183105,29.7441425323486,29.6845684051514,29.6300773620605,29.4822254180908,29.4129867553711,29.3534183502197,29.3731422424316,29.3979511260986,29.3960933685303,29.3750019073486,29.2830085754395,29.08740234375,29.0317401885986,28.9474620819092,28.7947254180908,28.7408218383789,28.6908206939697,28.6369132995605,28.5639629364014,28.4070320129395,28.39208984375,28.3163070678711,28.2842769622803,28.14794921875,28.17333984375,28.2020511627197,28.1916999816895,28.1692390441895,28.11083984375,28.1031246185303,28.0075206756592,27.9916000366211,27.94140625,27.89208984375,27.8815422058105,27.8486328125,27.8060550689697,27.6556625366211,27.6394538879395,27.7111320495605,27.7173824310303,27.7627925872803,27.8145503997803,27.8302726745605,27.8382816314697,27.82861328125,27.796875,27.6727542877197,27.5386714935303,27.5111331939697,27.4697265625,27.3519535064697,27.3542957305908,27.3717784881592,27.3999996185303,27.4919929504395,27.5147457122803,27.5420894622803,27.7528324127197,27.7769527435303,27.7785167694092,27.7687511444092,27.7219715118408,27.6734371185303,27.6441402435303,27.5710926055908,27.50244140625,27.48779296875,27.5055656433105,27.5300788879395,27.5313472747803,27.4270515441895,27.4341793060303,27.4644527435303,27.5130863189697,27.6217765808105,27.7576179504395,27.8495101928711,27.8976573944092,27.9381828308105,28.0164070129395,28.0460929870605,28.0613288879395,28.1283206939697,28.1510734558105,28.1330070495605,28.0658206939697,28.0124988555908,28.06396484375,28.0462875366211,28.0139636993408,28.0580081939697,28.1311511993408,28.2125015258789,28.3345699310303,28.4237289428711,28.4539051055908,28.5181636810303,28.6039066314697,28.7476539611816,28.8668937683105,28.9472637176514,28.9815444946289,29.0133800506592,29.0791034698486,29.1472644805908,29.6697273254395,30.12255859375,30.1568355560303,30.1726570129395,30.0599594116211,29.9767570495605,29.8722667694092,29.7211933135986,29.5693378448486,29.3704109191895,29.0691413879395,28.8126945495605,28.6431636810303,28.5222663879395,28.4916019439697,28.6224613189697,28.6403312683105,28.6505870819092,28.5778331756592,28.5127944946289,28.1792964935303,27.7976551055908,28.1519527435303,28.4074211120605,28.455078125,28.5681648254395,28.6628894805908,28.7390632629395,28.9929676055908,29.2516613006592,29.4923820495605,29.5793952941895,29.6901359558105,29.9332008361816,30.0099620819092,30.3064441680908,30.4796886444092,30.5656261444092,30.9357433319092,31.1867179870605,31.28564453125,31.3824214935303,31.4373054504395,31.5339832305908,31.5365238189697,31.50927734375,31.43701171875,31.3367195129395,31.2474594116211,31.1808586120605,30.9748058319092,30.6552715301514,30.4185543060303,30.0553703308105,29.9915046691895,30.0041027069092,30.2102546691895,30.4153308868408,30.5039043426514,30.5260753631592,30.5279293060303,30.5137710571289,30.4878921508789,30.3906269073486,30.1081047058105,30.0418930053711,29.9866199493408,29.9855480194092,30.1201171875,30.1261711120605,30.1102542877197,30.0728511810303,29.783203125,29.7016620635986,29.6374988555908,29.6041984558105,29.6008796691895,29.6224613189697,29.7200183868408,29.8108386993408,29.8269538879395,29.826171875,29.8105449676514,29.6296882629395,29.6124019622803,29.6080055236816,29.6171894073486,29.71484375,29.7280254364014,29.8194351196289,29.7159194946289,29.7239246368408,29.8826179504395,30.0290050506592,30.0953121185303,30.1027336120605,30.0874996185303,29.9366207122803,29.9034194946289,29.8035163879395,29.7207050323486,29.6708984375,29.5907230377197,29.5443344116211,29.4643535614014,29.3711929321289,29.2932605743408,29.0930671691895,29.0662117004395,29.0690422058105,29.0870132446289,29.2433586120605,29.3876953125,29.5722675323486,29.7505855560303,29.9412097930908,29.9880847930908,29.9792003631592,29.8215827941895,29.5242195129395,29.3438491821289,29.06298828125,28.6851558685303,28.5601558685303,28.470703125,28.4792976379395,28.7520503997803,28.7776355743408,28.7728519439697,28.7448234558105,28.7059574127197,28.4535160064697,28.4140625,28.5660171508789,28.6921863555908,28.89892578125,28.9658222198486,29.1185550689697,29.1708984375,29.2099590301514,29.35302734375,29.3882827758789,29.8327159881592,29.9940452575684,30.0873050689697,30.1318378448486,30.1637687683105,30.1867179870605,30.1964855194092,30.1597652435303,30.1801738739014,30.2275390625,30.3796882629395,30.6154289245605,30.7888679504395,30.8607406616211,30.8966789245605,30.9224624633789,30.9241218566895,30.8697261810303,31.0495109558105,31.4527339935303,31.5469741821289,31.6662120819092,31.78857421875,31.8793964385986,31.9979515075684,32.0305671691895,31.9693355560303,31.9845695495605,32.3916015625,32.5654296875,32.9416999816895,33.0078125,33.0125961303711,32.99462890625,32.9150390625,32.7542953491211,32.1767578125,32.0915031433105,32.1613273620605,32.33056640625,32.3777313232422,32.6368141174316,32.8837890625,32.9998016357422,33.02099609375,32.9416046142578,32.9789047241211,33.255859375,33.3848609924316,33.4542961120605,33.4636726379395,33.41796875,33.4129867553711,33.3277320861816,33.1963844299316,33.1412124633789,33.3334007263184,33.4356422424316,33.6270484924316,33.6843757629395,34.2293930053711,34.3527336120605,34.8639678955078,35.0095672607422,35.1758766174316,35.2332038879395,35.2898406982422,35.85791015625,36.6182632446289,37.7305679321289,38.3576202392578,38.43017578125,38.6568336486816,38.70556640625,38.83154296875,39.5689468383789,39.8233375549316,39.7897453308105,39.7462882995605,39.8092765808105,39.8956069946289,40.0357398986816,40.2066383361816,40.3806610107422,40.5257797241211,40.6565437316895,40.7663078308105,40.9664039611816,41.0609397888184,41.1338844299316,41.1338844299316,41.26171875,41.3587913513184,41.3542938232422,41.2755889892578,41.18896484375,40.5215835571289,40.1033172607422,39.2890625,38.6539039611816,38.3975563049316,37.9006805419922,37.6282234191895,37.2948226928711,36.9836921691895,36.7699203491211,36.3734397888184,35.5134773254395,35.3639640808105,34.8246116638184,34.6102561950684,34.4826164245605,34.3960952758789,34.4308624267578,34.4515647888184,34.1460914611816,33.8936538696289,33.7595710754395,33.5954093933105,33.52294921875,33.4820327758789,33.1501960754395,33.001953125,32.8475570678711,32.88525390625,32.9304695129395,32.39990234375,31.8953132629395,31.9830055236816,32.2015609741211,32.3406257629395,32.5009765625,32.4636688232422,32.6864280700684,32.8573265075684,32.8624038696289,32.9287109375,33.1805686950684,33.2244148254395,33.1829109191895,33.2173805236816,33.4052734375,33.5176734924316,33.6559562683105,33.59326171875,33.4769515991211,33.3605461120605,33.4158210754395,33.5666999816895,34.1126976013184,34.3998031616211,34.6917991638184,34.7863273620605,34.7931632995605,34.7769546508789,34.7347679138184,34.7155265808105,34.61572265625,34.5441398620605,34.4064445495605,34.5359344482422,34.6710929870605,34.8035163879395,34.8271484375,34.8326148986816,34.9522476196289,34.9054718017578,34.8582992553711,34.8695335388184,35.0353546142578,35.2840805053711,35.4320297241211,35.6470718383789,35.8020515441895,36.146484375,36.3019523620605,36.3649406433105,36.7137680053711,36.9751930236816,37.3727531433105,37.4421882629395,37.6353530883789,37.9679679870605,38.07080078125,38.0622062683105,37.9771499633789,37.9537124633789,37.8435554504395,37.7406234741211,37.4295921325684,37.28955078125,37.1836891174316,37.0404281616211,36.7693367004395,36.6242179870605,36.5787086486816,36.5282249450684,36.5345687866211,36.6529312133789,36.7859382629395,36.8828125,37.0501937866211,37.1408195495605,37.5281257629395,38.0093727111816,38.11572265625,38.2282257080078,38.412109375,38.4419898986816,38.5409164428711,38.6130828857422,39.0535163879395,39.5673828125,39.7580070495605,39.8330078125,39.8486328125,40.0578117370605,40.2037124633789,40.4078140258789,40.4449195861816,40.3753890991211,40.28125,40.1426734924316,39.896484375,39.7980461120605,39.7491226196289,39.7811546325684,39.8165016174316,40.3278312683105,40.5127906799316,40.6916046142578,40.7744140625,41.0760765075684,41.4757804870605,41.7808570861816,42.0835914611816,42.2105445861816,42.3136711120605,42.4507789611816,42.6021461486816,42.8065452575684,43.0059585571289,43.2332038879395,43.5508766174316,43.6033210754395,43.6531257629395,43.5503883361816,43.5418968200684,43.6239280700684,43.7370109558105,43.84375,43.9441375732422,44.0167007446289,44.1043968200684,44.1324195861816,44.1453132629395,44.09716796875,44.220703125,44.31640625,44.4886741638184,44.4371070861816,44.4292984008789,44.4039077758789,44.2917976379395,44.0744132995605,43.8553695678711,43.7824211120605,43.7957038879395,43.8563461303711,44.0364265441895,44.2253913879395,44.2315444946289,44.2138671875,44.2264671325684,44.2046890258789,44.1691398620605,43.4040031433105,43.3580055236816,43.3332023620605,43.4132804870605,43.4719734191895,44.0480461120605,44.17529296875,45.078125,45.5194320678711,45.8919944763184,46.1583976745605,46.4296875,46.68359375,46.6904296875,46.4289093017578,46.1742210388184,45.5287132263184,45.3741226196289,44.939453125,44.9021492004395,44.939453125,45.1388664245605,45.5622062683105,45.7525367736816,45.8853492736816,45.9860343933105,46.083984375,46.2977523803711,46.4485359191895,46.4923820495605,46.5523414611816,46.6908226013184,47.4964828491211,47.6558570861816,47.7090835571289,47.76806640625,47.8392562866211,47.908203125,47.8826179504395,47.8747100830078,48.2787094116211,48.65380859375,48.8332023620605,48.8779296875,48.7626953125,48.6957054138184,48.7542953491211,48.8406219482422,48.9539070129395,49.1552734375,49.9312515258789,50.2332038879395,50.4140625,50.6994132995605,50.8388671875,51.0785140991211,51.3361320495605,51.61669921875,51.9947280883789,52.0556640625,52.1288108825684,52.2853507995605,52.2274398803711,52.1834945678711,52.2591819763184,52.322265625,52.3966789245605,52.4750022888184,52.6697273254395,52.72265625,52.6476593017578,52.5500984191895,52.43505859375,52.3440437316895,52.68359375,53.4128875732422,53.8019523620605,54.1858406066895,54.4912109375,54.3762702941895,53.8744163513184,53.7976570129395,53.7982406616211,53.9195327758789,53.9706039428711,53.9292945861816,53.8912086486816,53.8338890075684,53.7588844299316,53.9176788330078,53.9308586120605,53.8294944763184,53.6900405883789,53.5666961669922,53.3425788879395,53.2933616638184,53.2605476379395,53.4031257629395,53.51513671875,53.9136695861816,53.9678726196289,54.0992202758789,54.23291015625,54.3939476013184,54.4761734008789,54.5612297058105,54.7179679870605,54.861328125,54.9230461120605,55.15087890625,55.4180641174316,55.67529296875,55.9246101379395,56.0436515808105,56.2756843566895,56.6202163696289,56.9093742370605,57.1268539428711,57.4443359375,58.1730499267578,58.2370147705078,58.3539047241211,58.9189453125,59.0573272705078,59.0598602294922,59.1101570129395,59.2204093933105,59.3705101013184,59.2983436584473,59.2225570678711,59.1123008728027,59.0990257263184,59.3107414245605,59.6042938232422,59.7256813049316,59.8275375366211,59.8587875366211,59.8973655700684,59.9228553771973,59.94140625,59.8651390075684,59.8959999084473,60.1602516174316,60.4891586303711,60.6376953125,60.8151359558105,60.93359375,60.8585891723633,60.66455078125,60.3373031616211,60.1706047058105,60.2764663696289,60.5586891174316,60.8129844665527,60.9090805053711,61.0159187316895,61.7705078125,62.6312484741211,63.3614234924316,64.1904296875,64.5921936035156,64.9285125732422,64.8962936401367,65.0315399169922,65.3267593383789,65.5279312133789,65.7357406616211,65.8126983642578,66.0847625732422,66.3656234741211,66.4161071777344,66.7564468383789,67.00244140625,67.1492233276367,67.6396484375,67.7307662963867,68.1569366455078,68.3711929321289,68.5042037963867,68.8294906616211,69.0243148803711,69.1405334472656,68.9244155883789,68.7628860473633,68.6595687866211,68.5427703857422,68.3550796508789,68.1173782348633,68.0730438232422,68.005859375,67.7743148803711,67.6241149902344,67.064453125,66.9640655517578,66.9341812133789,66.8966827392578,66.8402328491211,66.8040008544922,66.8029327392578,66.8322296142578,66.9263687133789,67.0690460205078,67.1444320678711,67.2392578125,67.1974639892578,67.146484375,67.1568374633789,67.2468719482422,67.2847671508789,67.2115173339844,67.1433639526367,66.8224563598633,66.7022476196289,66.6751937866211,66.6661148071289,66.7587890625,66.8470687866211,66.6925735473633,66.6396484375,66.76806640625,66.9175796508789,67.2742233276367,67.5417938232422,67.9593811035156,68.2692337036133,68.4694366455078,68.607421875,68.8296890258789,69.0390625,69.3914108276367,69.61181640625,69.6943359375,69.708984375,69.6587905883789,69.6451187133789,69.73828125,69.8874969482422,70.1721649169922,70.6553726196289,71.5001983642578,71.6169891357422,71.9295883178711,72.1009826660156,72.4463882446289,72.6337890625,72.8121109008789,72.7873992919922,72.7529296875,72.6244125366211,72.5741195678711,72.375,72.2794952392578,72.1296844482422,71.9120101928711,71.8843765258789,71.8672866821289,72.0792922973633,72.5813446044922,72.7044982910156,72.7316360473633,72.7000045776367,72.6533203125,72.5619125366211,72.4694366455078,72.5296936035156,72.5994110107422,72.6156311035156,72.5573272705078,72.5270538330078,72.52734375,72.5767593383789,72.6783218383789,72.8119125366211,73.1907196044922,73.5480422973633,73.5734405517578,73.5916976928711,73.4652328491211,73.2664108276367,73.1394577026367,73.12939453125,73.1730422973633,73.1521453857422,73.0667953491211,72.94873046875,72.5943374633789,71.8474655151367,71.6681671142578,71.365234375,71.4489288330078,71.5511703491211,71.53955078125,71.3419952392578,71.0656280517578,70.939453125,70.7248992919922,70.5614242553711,70.3828125,70.4088897705078,70.4425735473633,70.5679702758789,70.6907196044922,70.6307601928711,70.5791015625,70.4439468383789,70.2834014892578,70.09375,69.9486312866211,69.8771438598633,69.7404327392578,69.2177734375,69.0787124633789,69.0134735107422,69.0511703491211,69.0911102294922,69.1439437866211,69.1943359375,69.4120101928711,69.7009735107422,69.982421875,70.3394546508789,71.1455078125,71.3580017089844,71.5656204223633,71.9169921875,72.0675811767578,72.3215789794922,72.3839797973633,72.4173812866211,73.3415985107422,73.5135726928711,73.7920837402344,73.88330078125,73.9862289428711,74.0745162963867,74.6760787963867,74.76953125,74.7873077392578,74.7782211303711,74.74267578125,74.6324234008789,74.51123046875,74.3914108276367,74.48095703125,74.57958984375,75.1246109008789,75.5895538330078,76.1075210571289,76.3160095214844,76.4591827392578,76.6057662963867,76.7350616455078,77.1117172241211,77.2384796142578,77.2610397338867,77.2484359741211,77.1744155883789,77.3250961303711,77.3956069946289,77.5792007446289,77.6750946044922,77.7715835571289,77.9855499267578,78.5895462036133,78.9224624633789,78.8875961303711,78.8390655517578,78.55908203125,78.1612319946289,77.5882797241211,77.5201187133789,77.5359420776367,77.6648483276367,77.7568359375,77.8682632446289,77.9951171875,77.9586944580078,77.9068374633789,77.7852554321289,77.6506881713867,77.46630859375,77.3283233642578,76.6449203491211,76.0009765625,75.5611343383789,75.4200210571289,75.0535125732422,74.8148422241211,74.3626022338867,73.9774475097656,73.8360290527344,73.7756805419922,73.8909225463867,73.8327102661133,73.6632843017578,73.5601501464844,73.578125,73.8301773071289,73.9374084472656,74.2067413330078,74.3433609008789,74.3109359741211,73.7315444946289,73.5765609741211,73.5072250366211,73.365234375,73.1504821777344,73.0862274169922,73.6717758178711,73.939453125,74.3112258911133,74.4890594482422,74.8041000366211,74.9921875,75.05322265625,75.0899353027344,75.0970687866211,75.0603485107422,75.0080108642578,74.8968734741211,74.7868118286133,74.8649368286133,74.9421844482422,75.1524429321289,75.3693313598633,75.4748992919922,75.603515625,75.6031265258789,75.5914077758789,75.6443405151367,75.6911087036133,75.7414093017578,75.6944274902344,75.6443405151367,75.5502014160156,75.39453125,75.2738265991211,75.2474670410156,75.5032196044922,75.4685592651367,75.4171829223633,75.2802734375,75.2980499267578,75.33203125,75.7335968017578,76.1104507446289,76.7419891357422,76.9290008544922,76.9952163696289,77.5896530151367,78.0682601928711,78.3206024169922,78.5257797241211,78.9421844482422,79.0154342651367,79.0838851928711,78.888671875,78.8035125732422,78.7239303588867,78.5876922607422,78.4914093017578,78.3865203857422,78.2126007080078,77.9083938598633,77.7066421508789,77.4810485839844,77.1136703491211,76.8711929321289,76.4333953857422,76.3121109008789,76.2157211303711,76.1036148071289,76.0324249267578,76.1240234375,76.4216842651367,76.8713912963867,77.0613327026367,77.55078125,77.7775344848633,78.1869201660156,78.232421875,78.1408233642578,78.0164108276367,77.7806625366211,77.4928741455078,77.4108352661133,77.4397506713867,77.4715881347656,77.6252899169922,77.7332000732422,77.9681701660156,78.2253952026367,78.4826202392578,79.4220657348633,79.9539108276367,80.4740295410156,80.69921875,80.7624969482422,80.8147430419922,80.8560562133789,81.51123046875,81.66162109375,82.0798873901367,82.5472640991211,82.7578125,82.9861373901367,83.1066436767578,83.2335968017578,83.16552734375,83.1056671142578,82.97705078125,82.91796875,82.4931640625,82.3228530883789,82.2769470214844,82.2542953491211,82.2391586303711,82.3160095214844,82.3359375,82.2706985473633,82.1631851196289,82.1824188232422,82.22119140625,82.23583984375,82.2314453125,82.2583999633789,82.45166015625,82.5924758911133,82.73779296875,82.869140625,83.0101547241211,83.0510787963867,83.0584030151367,83.0301742553711,82.9198226928711,82.7424850463867,82.6823196411133,82.7672805786133,82.8565368652344,82.9610366821289,83.0807647705078,83.1095733642578,83.1320266723633,83.0941390991211,83.0738296508789,83.29345703125,83.4970703125,83.6598663330078,83.7004928588867,83.7359390258789,83.6512680053711,83.5789108276367,83.3338851928711,83.1512680053711,83.2660140991211,83.4576187133789,83.5310516357422,83.5504837036133,83.5712890625,83.5535125732422,83.5343780517578,83.3404312133789,83.2002868652344,82.7550811767578,82.6454086303711,82.3191375732422,82.2806625366211,82.2092742919922,82.18359375,82.0936584472656,81.7928771972656,81.5862350463867,81.28271484375,81.09814453125,80.8270492553711,80.7977600097656,80.7196273803711,80.65625,80.6753845214844,80.7737274169922,80.8415985107422,80.7574234008789,80.638671875,80.5096664428711,80.4554672241211,80.4245147705078,80.4189453125,80.3980484008789,80.4583053588867,80.5958938598633,80.5619201660156,80.5831985473633,81.4688491821289,81.8169937133789,83.5447311401367,83.6669921875,84.4173812866211,84.7378845214844,85.0774459838867,85.2005844116211,85.4483337402344,85.6114273071289,85.9792938232422,86.5914001464844,86.8929672241211,86.9613265991211,87.0294952392578,86.6976547241211,86.3659210205078,86.0941390991211,85.8270492553711,85.8004913330078,85.7925796508789,85.8024444580078,85.8181610107422,86.09814453125,86.3079071044922,86.5143508911133,86.6770553588867,86.7150344848633,86.1216812133789,85.9708023071289,85.9100570678711,85.93896484375,85.9989242553711,86.0923843383789,86.1550750732422,86.3762741088867,87.1201171875,87.29443359375,87.3695297241211,87.5711975097656,87.5032196044922,87.3375015258789,87.2096633911133,86.6970748901367,86.5710906982422,86.1778259277344,86.0013656616211,86.1829071044922,86.3957977294922,86.5384750366211,86.6647491455078,86.89794921875,87.2296829223633,87.1061553955078,86.8942413330078,86.7000961303711,86.4256820678711,86.1161117553711,85.791015625,85.8807601928711,86.0588912963867,86.1195297241211,86.2012710571289,86.6514587402344,86.8628921508789,87.0418014526367,87.4193420410156,87.4675827026367,87.2873992919922,87.1407241821289,86.9390563964844,86.9216766357422,87.0059585571289,87.1707992553711,87.67138671875,88.5037155151367,88.7331085205078,89.3102569580078,89.5951156616211,90.1849670410156,91.0046920776367,91.4794921875,91.8454132080078,92.4075241088867,92.6025390625,93.5498046875,94.0751953125,94.1563491821289,93.68701171875,93.5740203857422,93.4754867553711,93.4060516357422,93.1781234741211,93.1163101196289,93.0686492919922,92.9866180419922,92.8904266357422,92.8585968017578,92.9715881347656,93.1048812866211,93.25927734375,93.3595657348633,93.6484375,93.8428726196289,94.1023406982422,94.3882827758789,94.5067367553711,94.5755844116211,95.0384750366211,95.3592758178711,95.5787124633789,95.919921875,96.0754928588867,95.9860382080078,95.6533203125,95.7438507080078,95.9347686767578,96.5085906982422,96.6005859375,96.5376968383789,96.4970703125,96.8791961669922,97.2054672241211,97.3506851196289,97.4992218017578,97.6376953125,97.6698303222656,97.9183578491211,98.02001953125,98.1946334838867,98.3419952392578,98.6620178222656,98.7712936401367,98.9846649169922,99.1873016357422,99.5626983642578,99.6156234741211,99.6631851196289,99.7704086303711,99.6892623901367,99.6023483276367,99.4421844482422,99.5407257080078,99.609375,99.7375030517578,99.8513717651367,99.8253936767578,99.6167984008789,99.4606399536133,99.0938491821289,98.9695281982422,98.8056640625,98.8694305419922,99.5762710571289,99.9357452392578,100.322364807129,100.84375,101.060745239258,101.310737609863,101.597755432129,101.683792114258,101.212989807129,101.002639770508,100.928024291992,101.006248474121,101.099319458008,101.008201599121,100.92041015625,100.905860900879,100.989944458008,101.185745239258,101.292869567871,101.517677307129,102.610160827637,103.131446838379,103.331245422363,103.560745239258,104.014556884766,104.184867858887,104.814262390137,104.965232849121,105.308982849121,105.710250854492,105.89453125,105.9833984375,106.0595703125,105.734176635742,105.384574890137,104.911911010742,104.323631286621,104.202438354492,105.320213317871,105.645896911621,105.712013244629,105.822166442871,106.145408630371,106.338668823242,106.705078125,106.78369140625,106.941604614258,107.278907775879,107.429786682129,107.190238952637,106.94091796875,106.638763427734,106.545509338379,106.384666442871,106.41357421875,106.683204650879,106.825386047363,107.15771484375,107.624221801758,107.72216796875,107.902244567871,107.949905395508,108.027931213379,108.181640625,108.35205078125,108.638381958008,109.369331359863,109.981155395508,110.471488952637,111.114845275879,111.392486572266,111.6005859375,111.7861328125,111.938674926758,112.093940734863,112.016799926758,111.942672729492,112.142768859863,112.297073364258,112.413284301758,112.619529724121,112.684181213379,112.742576599121,112.798439025879,112.721878051758,112.65625,112.818946838379,113.046676635742,113.094039916992,113.150390625,113.066017150879,112.987991333008,113.086036682129,113.272659301758,113.365524291992,113.427734375,113.563865661621,113.857223510742,113.870994567871,113.748725891113,113.619926452637,113.567573547363,113.485939025879,113.517189025879,113.469047546387,113.3916015625,113.126365661621,112.629196166992,112.496681213379,112.466110229492,112.453025817871,112.729591369629,112.955665588379,113.161521911621,113.242965698242,113.356254577637,113.558891296387,113.726173400879,113.613571166992,112.924903869629,112.191993713379,111.868263244629,111.299026489258,110.892768859863,110.37353515625,110.225883483887,109.84033203125,109.866409301758,109.911331176758,109.863868713379,109.810256958008,109.510841369629,109.074996948242,108.199516296387,107.765426635742,107.271095275879,107.1669921875,106.794235229492,106.679389953613,106.188667297363,105.677154541016,105.392776489258,105.143943786621,105.402732849121,105.708198547363,106.066696166992,106.159568786621,106.208786010742,106.315032958984,106.477928161621,107.108787536621,107.368751525879,107.750396728516,108.001266479492,108.150978088379,108.285354614258,108.351463317871,108.575393676758,109.089942932129,109.165618896484,109.3310546875,109.637107849121,109.855270385742,110.4287109375,110.773338317871,110.8681640625,110.799217224121,110.722366333008,110.388282775879,110.091209411621,109.752731323242,109.706733703613,109.665626525879,109.774124145508,109.869140625,110.083885192871,110.261421203613,110.920112609863,111.056251525879,111.130859375,111.341407775879,111.550590515137,111.4599609375,111.228126525879,111.299507141113,111.400390625,111.8037109375,112.147270202637,112.399993896484,112.79541015625,112.85595703125,112.939651489258,112.8359375,112.934959411621,113.032814025879,113.181541442871,113.326850891113,113.416213989258,113.364456176758,113.156929016113,113.276954650879,113.490913391113,113.487594604492,113.474609375,113.369338989258,113.247367858887,113.127830505371,113.158203125,113.186134338379,113.312210083008,113.66455078125,113.7119140625,113.630081176758,113.391403198242,113.298149108887,113.215530395508,113.3115234375,113.41748046875,113.542778015137,113.581443786621,113.558891296387,113.63916015625,113.765235900879,113.829299926758,113.88623046875,113.795120239258,113.711326599121,113.539451599121,113.510353088379,113.85693359375,114.060546875,114.816009521484,115.337692260742,116.495498657227,117.30859375,118.4501953125,118.870903015137,118.911231994629,118.936416625977,118.754493713379,118.457038879395,118.376564025879,118.430274963379,118.960350036621,119.425300598145,119.750389099121,119.921676635742,120.597946166992,120.997169494629,121.354293823242,121.747848510742,121.886039733887,122.02978515625,122.260154724121,122.537498474121,122.692092895508,122.751953125,122.730865478516,122.501953125,122.526763916016,122.615226745605,122.999313354492,123.160346984863,123.301170349121,123.404594421387,123.461624145508,123.521881103516,123.572463989258,123.622268676758,123.500984191895,123.383880615234,123.355278015137,123.322654724121,123.305076599121,123.416206359863,123.491119384766,123.796875,123.933891296387,124.019035339355,124.388084411621,124.541206359863,124.796287536621,125.617095947266,125.598533630371,125.79443359375,125.887886047363,126.107421875,126.254486083984,126.295989990234,126.344924926758,126.308883666992,126.257423400879,126.29248046875,126.335456848145,126.552543640137,126.838470458984,126.955169677734,127.031341552734,127.740325927734,127.955085754395,127.996871948242,128.025680541992,128.141693115234,128.281448364258,128.26416015625,128.2578125,128.587005615234,128.73046875,128.888671875,128.871688842773,128.913375854492,129.059188842773,129.1005859375,129.0537109375,128.853515625,128.735260009766,128.599014282227,128.674026489258,129.017288208008,129.229110717773,129.250381469727,129.117584228516,128.815338134766,128.633407592773,128.508499145508,128.418258666992,128.549423217773,129.116592407227,129.281341552734,129.411712646484,129.41064453125,129.283508300781,128.934951782227,128.475189208984,128.196960449219,127.803413391113,127.726081848145,127.841400146484,128.026565551758,128.358795166016,128.911422729492,129.040145874023,129.116592407227,129.154205322266,129.12158203125,129.210250854492,129.291793823242,129.460845947266,129.234176635742,128.949020385742,128.84326171875,128.922668457031,129.13427734375,129.224517822266,129.389846801758,129.761917114258,130.025970458984,130.28125,130.537109375,130.66845703125,130.757125854492,130.831924438477,130.898040771484,131.021575927734,131.157424926758,131.268264770508,131.432312011719,131.56201171875,131.76904296875,131.906448364258,132.035354614258,131.990829467773,132.003707885742,132.098815917969,132.227645874023,132.325775146484,132.562301635742,132.653915405273,132.7158203125,132.7685546875,132.803619384766,132.839263916016,133.130859375,133.426177978516,133.688858032227,134.102828979492,134.702743530273,134.813858032227,135.022369384766,135.359375,135.559173583984,135.884765625,136.09033203125,136.406158447266,137.115814208984,137.3154296875,137.41748046875,137.650588989258,137.7978515625,137.939651489258,137.99169921875,137.973724365234,137.901962280273,137.844039916992,138.0126953125,138.032516479492,138.090621948242,138.314056396484,138.09716796875,138.022171020508,137.918365478516,137.927337646484,137.995712280273,138.04833984375,138.118453979492,138.234176635742,138.318069458008,138.525192260742,138.670028686523,138.780181884766,139.0048828125,139.209381103516,139.320220947266,139.632125854492,139.984176635742,139.938766479492,139.695114135742,139.722946166992,139.552352905273,139.359268188477,139.640243530273,139.847076416016,140.014053344727,140.187698364258,140.134368896484,139.616989135742,139.505279541016,139.430465698242,139.176361083984,139.14501953125,139.140823364258,139.4736328125,139.601165771484,140.450592041016,140.705078125,141.079284667969,140.983200073242,140.972854614258,140.65234375,140.675979614258,140.708099365234,140.808197021484,141.309768676758,141.518371582031,142.061431884766,143.515823364258,143.680953979492,144.303909301758,144.568664550781,145.199310302734,145.485748291016,145.714157104492,146.083312988281,146.2529296875,146.234756469727,145.467086791992,145.212890625,144.8974609375,144.7763671875,144.587585449219,144.360931396484,144.169235229492,144.294921875,144.470703125,145.039169311523,146.594131469727,146.831848144531,146.807037353516,146.599212646484,146.401657104492,146.11328125,146.005859375,146.230285644531,146.137313842773,146.051467895508,145.799423217773,145.758605957031,145.709671020508,145.710159301758,145.6640625,145.756744384766,145.756744384766,145.4072265625,145.271194458008,145.125778198242,145.06396484375,145.046875,145.077743530273,145.07373046875,145.017868041992,144.989639282227,145.075576782227,145.188583374023,145.804779052734,146.0732421875,146.367965698242,146.894714355469,147.127059936523,147.261810302734,147.433990478516,148.402053833008,148.96484375,149.501556396484,149.766220092773,149.963088989258,149.998138427734,150.016891479492,149.881057739258,149.279678344727,149.048736572266,148.96533203125,148.954879760742,148.92333984375,148.968154907227,149.237884521484,149.498046875,149.857116699219,149.912689208984,150.026473999023,150.060836791992,150.599807739258,150.634857177734,150.667770385742,150.525100708008,150.384765625,150.09765625,150.242980957031,150.821685791016,150.9677734375,151.145309448242,151.582412719727,151.759765625,152.0927734375,151.999801635742,151.762008666992,152.5087890625,152.79833984375,153.460647583008,153.794143676758,154.413970947266,155.029495239258,155.595901489258,155.895217895508,156.6845703125,157.447357177734,158.037017822266,158.7021484375,159.350677490234,159.727935791016,159.8046875,159.911819458008,159.958587646484,160.006439208984,159.9833984375,159.8896484375,159.831436157227,159.839157104492,159.729400634766,159.83251953125,160.119140625,160.739456176758,160.910751342773,160.928894042969,160.982025146484,161.035552978516,161.140808105469,161.309860229492,161.340621948242,161.128997802734,160.996688842773,160.856048583984,161.1044921875,161.230178833008,161.365127563477,161.495315551758,161.565628051758,161.565628051758,161.480087280273,161.480087280273,161.536911010742,161.945114135742,162.166015625,162.375686645508,162.944625854492,163.201370239258,163.498046875,163.705276489258,163.945999145508,164.159561157227,164.513275146484,165.7607421875,165.98046875,166.8203125,166.884368896484,167.073150634766,167.628128051758,167.856826782227,167.950103759766,168.047653198242,168.149993896484,168.22998046875,168.303039550781,168.423049926758,168.587585449219,168.946197509766,169.310653686523,169.414642333984,169.60986328125,170.065628051758,170.53759765625,170.995407104492,170.996688842773,170.8837890625,170.714157104492,170.582229614258,170.160934448242,170.201171875,170.359573364258,170.503112792969,170.525390625,170.48681640625,170.867980957031,171.246673583984,171.970504760742,172.5595703125,172.869232177734,173.056350708008,173.277450561523,173.353317260742,173.438659667969,173.7333984375,173.948043823242,174.319427490234,174.785552978516,175.295608520508,175.751174926758,175.921478271484,176.107528686523,176.410446166992,176.924407958984,177.39453125,177.933700561523,178.442779541016,178.848342895508,178.906921386719,178.925003051758,178.95068359375,179.272659301758,179.868270874023,180.000045776367,180.201461791992,180.404586791992,180.485488891602,180.529159545898,180.64404296875,180.720703125,181.126129150391,181.310684204102,181.461471557617,181.386322021484,181.24853515625,181.263473510742,181.307373046875,181.526077270508,181.755508422852,181.902542114258,181.951324462891,181.981292724609,181.944198608398,182.077590942383,182.203216552734,182.13818359375,181.715469360352,181.626953125,181.750152587891,182.316787719727,182.472747802734,182.406784057617,182.360641479492,182.410781860352,182.479110717773,182.592483520508,182.702590942383,182.828170776367,183.092727661133,184.65478515625,184.690124511719,184.734069824219,184.760452270508,184.767471313477,184.625289916992,184.844924926758,184.877197265625,184.934371948242,184.997314453125,185.081939697266,185.150146484375,185.130081176758,185.069580078125,185.061874389648,185.114944458008,185.171295166016,185.216354370117,185.228820800781,185.1298828125,185.075088500977,185.1357421875,185.325332641602,185.387542724609,185.496246337891,185.522262573242,185.546234130859,185.581298828125,185.605911254883,185.633941650391,185.743026733398,185.794006347656,185.915237426758,185.982284545898,185.934967041016,185.974563598633,186.005523681641,186.044525146484,186.100051879883,186.16796875,186.226028442383,186.157470703125,186.079055786133,185.898147583008,185.803665161133,185.768417358398,185.860397338867,185.939407348633,185.994476318359,185.981140136719,185.958984375,185.91357421875,185.845657348633,185.716445922852,185.658157348633,185.569091796875,185.481048583984,185.445510864258,185.449905395508,185.552383422852,186.115966796875,186.3203125,186.413436889648,186.506011962891,186.842193603516,186.832382202148,186.775833129883,186.676467895508,186.656936645508,186.652633666992,186.741058349609,186.824600219727,186.783843994141,186.771728515625,186.806991577148,186.853164672852,186.941497802734,187.037414550781,187.359420776367,187.450637817383,187.479888916016,187.417037963867,186.998092651367,186.992477416992,187.378952026367,187.552688598633,187.72607421875,187.968505859375,188.204437255859,188.430419921875,188.639495849609,188.850723266602,189.073333740234,189.4443359375,189.490478515625,189.526901245117,189.457183837891,189.395553588867,189.516693115234,189.638870239258,189.698776245117,189.753021240234,189.788375854492,189.808059692383,189.756057739258,190.111190795898,190.222122192383,190.270843505859,190.168319702148,190.108306884766,190.050689697266,189.996185302734,189.840576171875,189.598968505859,189.459320068359,189.436965942383,189.458587646484,189.439010620117,189.333694458008,189.103134155273,188.99853515625,188.880996704102,188.767974853516,188.623153686523,188.578460693359,188.548828125,188.598281860352,188.69677734375,188.865570068359,188.945755004883,188.894134521484,188.830032348633,188.783981323242,188.63623046875,188.533737182617,188.209625244141,188.092880249023,188.052841186523,188.042831420898,187.868499755859,187.76611328125,187.717727661133,187.677734375,187.564300537109,187.392288208008,187.280807495117,187.216705322266,187.443466186523,187.646041870117,187.608001708984,187.582229614258,187.694274902344,187.767181396484,187.788421630859,187.730133056641,187.690734863281,187.338088989258,187.426849365234,187.517929077148,187.621292114258,187.713958740234,187.776306152344,187.786819458008,187.695663452148,187.601272583008,187.407180786133,187.20751953125,187.102630615234,187.000885009766,186.933792114258,186.914199829102,187.001953125,187.103134155273,187.198928833008,187.188430786133,187.097412109375,187.075973510742,187.110946655273,187.09912109375,187.145843505859,187.253112792969,187.383895874023,187.512603759766,187.563369750977,187.606155395508,187.621246337891,187.598541259766,187.305313110352,187.260833740234,187.244049072266,187.20849609375,187.096832275391,187.050979614258,187.084121704102,187.039932250977,186.990859985352,186.842590332031,186.724517822266,186.624313354492,186.624465942383,186.690780639648,186.690673828125,186.672515869141,186.604339599609,186.525054931641,186.396377563477,186.334030151367,186.270263671875,186.102157592773,185.998626708984,185.795166015625,185.681991577148,185.429443359375,185.169525146484,184.963973999023,184.854156494141,184.744094848633,184.604873657227,184.557861328125,184.516799926758,184.479339599609,184.284133911133,184.146148681641,184.140533447266,184.169784545898,184.143859863281,184.077056884766,183.90673828125,183.452545166016,183.077880859375,182.943740844727,182.824752807617,182.51123046875,182.301376342773,181.689788818359,181.587509155273,181.495361328125,181.474075317383,181.500686645508,181.497665405273,181.473785400391,181.441467285156,181.320861816406,181.208938598633,181.120651245117,181.060943603516,181.141754150391,181.253280639648,181.269439697266,181.306198120117,181.383743286133,181.413467407227,181.465866088867,181.473434448242,181.384231567383,181.247207641602,181.179153442383,181.131896972656,181.08447265625,180.973876953125,180.894927978516,180.893127441406,180.856597900391,180.821640014648,180.807327270508,180.706832885742,180.659866333008,180.683792114258,180.672805786133,180.577346801758,180.383834838867,180.316696166992,180.259124755859,180.216354370117,180.210311889648,180.271682739258,180.359375,180.550933837891,180.634033203125,180.655624389648,180.647903442383,180.54833984375,180.480651855469,180.364852905273,180.29541015625,180,179.827331542969,179.6513671875,179.4482421875,179.150009155273,178.698440551758,178.51953125,178.285354614258,177.748641967773,177.581634521484,177.337005615234,177.251846313477,177.17919921875,176.880859375,176.624801635742,176.4130859375,176.341018676758,176.4521484375,176.6455078125,176.940032958984,177.037307739258,177.123443603516,177.222854614258,177.148239135742,177.068740844727,176.8310546875,176.556640625,176.429489135742,176.061126708984,175.781143188477,175.396484375,175.097763061523,174.548828125,174.698623657227,175.097076416016,175.330673217773,175.677932739258,175.858581542969,175.945892333984,176.056533813477,176.169235229492,176.246963500977,176.300888061523,176.350982666016,176.283203125,176.219421386719,176.140914916992,176.507614135742,176.73095703125,176.842880249023,177.0498046875,177.387496948242,177.427444458008,177.467193603516,177.409866333008,177.432907104492,177.6875,177.953323364258,178.044723510742,178.130569458008,178.163970947266,178.2294921875,178.31298828125,178.381439208984,178.477142333984,178.474807739258,178.451370239258,178.536041259766,178.650283813477,178.692474365234,178.7314453125,178.681335449219,178.6259765625,178.4404296875,178.466110229492,178.653701782227,178.706451416016,178.668838500977,178.6787109375,178.744049072266,178.786712646484,178.775390625,178.79296875,178.918548583984,178.921493530273,179.028121948242,179.332321166992,179.388580322266,179.405075073242,179.329010009766,179.259567260742,179.302154541016,179.381057739258,179.510940551758,179.570510864258,179.570510864258,179.477249145508,179.288665771484,179.176956176758,179.133880615234,179.120697021484,179.044631958008,178.9638671875,178.019241333008,177.6630859375,177.351272583008,177.292572021484,177.2958984375,177.315826416016,177.359664916992,177.338958740234,177.29833984375,177.258697509766,177.172668457031,177.091201782227,177.023529052734,176.990036010742,176.963485717773,176.964752197266,177.008010864258,177.189651489258,177.159469604492,176.907424926758,176.702545166016,176.4365234375,176.328430175781,175.613861083984,175.441986083984,175.365814208984,175.267868041992,175.1923828125,174.797561645508,174.715042114258,174.610549926758,174.514358520508,174.284973144531,174.138870239258,173.822357177734,173.623443603516,173.390716552734,173.1318359375,173.054595947266,172.856536865234,172.806838989258,172.837890625,172.908004760742,172.867767333984,172.7890625,172.730667114258,172.690032958984,172.696975708008,172.584762573242,172.4970703125,172.396087646484,172.362411499023,172.392776489258,172.213272094727,172.067291259766,171.997665405273,171.91796875,171.83056640625,171.7294921875,171.48974609375,170.949325561523,170.79931640625,170.608215332031,170.589752197266,170.588577270508,170.512298583984,170.423431396484,170.396484375,170.350982666016,170.154098510742,169.982620239258,169.92724609375,169.897552490234,169.887008666992,169.854293823242,169.814743041992,169.618362426758,169.275680541992,169.226760864258,168.788284301758,168.670303344727,168.462783813477,168.137496948242,167.745986938477,167.626083374023,167.226760864258,166.964050292969,166.452545166016,166.331832885742,166.273040771484,166.1865234375,166.14892578125,166.136032104492,166.168350219727,166.229782104492,166.29248046875,166.30810546875,166.352157592773,166.18017578125,165.941986083984,165.5830078125,165.415817260742,165.285247802734,165.192581176758,165.084564208984,165.073638916016,165.018936157227,164.953720092773,164.854293823242,164.779388427734,164.669723510742,164.525299072266,164.440048217773,164.376846313477,164.251556396484,164.11328125,164.135055541992,164.017578125,163.912887573242,163.780075073242,163.743850708008,163.690032958984,163.574310302734,163.493743896484,163.409973144531,163.364852905273,163.321182250977,163.26904296875,163.272857666016,163.084869384766,163.010162353516,162.974899291992,162.940032958984,163.004287719727,162.969833374023,162.9345703125,162.847274780273,162.643356323242,162.453033447266,162.1416015625,162.049209594727,161.960052490234,162.001953125,162.039657592773,162.097946166992,162.197463989258,162.411422729492,162.392181396484,162.391403198242,162.466995239258,162.52197265625,162.654296875,162.718353271484,163.14501953125,163.225784301758,163.2138671875,163.187896728516,163.108779907227,162.95703125,162.779296875,162.762298583984,162.761520385742,162.80810546875,162.814834594727,162.791107177734,162.802642822266,162.849899291992,162.922073364258,163.04638671875,163.165435791016,163.256546020508,163.243255615234,163.294052124023,163.335540771484,163.261322021484,163.189254760742,163.04736328125,162.9716796875,162.84033203125,162.628128051758,162.713180541992,162.893264770508,162.975204467773,163.038375854492,162.944137573242,162.877639770508,162.671493530273,162.589065551758,162.488677978516,162.528213500977,162.461135864258,162.334075927734,162.146102905273,162.0849609375,161.924011230469,161.775588989258,161.723922729492,161.729400634766,161.784957885742,161.82421875,161.99609375,162.080276489258,162.105560302734,161.966903686523,161.725677490234,161.624801635742,161.294036865234,161.1298828125,160.935546875,160.772659301758,160.517196655273,160.288864135742,160.074417114258,160.010162353516,159.921783447266,159.84375,159.870910644531,159.914260864258,159.955856323242,159.899124145508,159.897659301758,160.002151489258,160.025100708008,159.947463989258,159.771575927734,159.5859375,159.136123657227,158.952056884766,158.745407104492,158.683700561523,158.639541625977,158.564651489258,158.472061157227,158.432327270508,158.560150146484,158.608795166016,158.53369140625,158.480758666992,158.500381469727,158.4931640625,158.463470458984,158.331634521484,158.103515625,157.8232421875,157.62890625,157.530944824219,157.489852905273,157.202239990234,156.847473144531,156.747756958008,156.724319458008,156.713470458984,156.670791625977,156.54345703125,156.521194458008,156.500381469727,156.489837646484,156.377349853516,156.36474609375,156.228607177734,156.154388427734,156.1103515625,156.098815917969,155.9501953125,155.904891967773,155.706436157227,155.620315551758,155.563873291016,155.554885864258,155.643447875977,155.716598510742,155.982513427734,156.025390625,156.067489624023,156.529296875,156.728424072266,156.848831176758,156.976760864258,156.963577270508,156.9482421875,156.89990234375,156.791595458984,156.829879760742,156.871978759766,156.985733032227,157.216796875,157.450393676758,157.666412353516,157.974609375,158.21044921875,158.275192260742,158.321090698242,158.449417114258,158.68701171875,159.036911010742,159.210647583008,159.308395385742,159.45263671875,159.591506958008,159.847366333008,160.350387573242,160.547454833984,160.71142578125,160.855270385742,161.218948364258,161.449325561523,161.753509521484,161.845993041992,162.003616333008,162.068161010742,162.266311645508,162.713180541992,162.97314453125,163.352340698242,163.466415405273,163.585159301758,163.7099609375,163.553512573242,163.589263916016,163.61962890625,163.893356323242,164.005462646484,163.992095947266,163.972747802734,163.804397583008,163.837112426758,163.882705688477,164.01953125,164.067977905273,164.07421875,164.207229614258,164.287490844727,164.598342895508,164.670700073242,164.8876953125,165.124130249023,165.208114624023,165.225677490234,165.2138671875,165.280380249023,165.417388916016,165.396575927734,165.044052124023,164.792388916016,164.566986083984,164.418365478516,164.255661010742,163.331741333008,163.287109375,163.244232177734,163.302139282227,163.258010864258,163.213287353516,163.163482666016,163.118453979492,163.131042480469,163.017684936523,163.00927734375,163.207611083984,163.2578125,163.197845458984,163.138870239258,163.085266113281,163.047271728516,162.993942260742,162.921676635742,162.85595703125,162.752349853516,162.717864990234,162.699020385742,162.607604980469,162.506439208984,162.392578125,162.188369750977,161.037109375,160.9150390625,160.7666015625,160.482025146484,160.3681640625,160.287307739258,160.173629760742,160.177352905273,160.201080322266,160.225784301758,160.37890625,160.28125,160.184280395508,160.003997802734,159.883102416992,159.790435791016,159.834564208984,159.94921875,159.913955688477,159.883102416992,159.930862426758,160.162689208984,160.246871948242,160.3173828125,160.321472167969,160.309371948242,160.23779296875,160.182525634766,159.72216796875,159.552337646484,159.496276855469,159.423049926758,159.295028686523,159.189254760742,159.07666015625,158.824310302734,158.547180175781,158.333694458008,158.151565551758,158.070114135742,157.79931640625,157.469345092773,157.370697021484,157.084167480469,156.891799926758,156.790618896484,156.680282592773,156.629699707031,156.482604980469,156.344131469727,156.055953979492,155.853317260742,155.716110229492,155.427825927734,154.970809936523,154.578216552734,154.440719604492,154.389846801758,154.293075561523,154.2666015625,154.268844604492,154.209182739258,154.149795532227,154.212890625,154.272171020508,154.357620239258,154.58251953125,154.971282958984,155.166687011719,155.153030395508,155.160446166992,155.016708374023,154.82373046875,154.703506469727,154.4580078125,154.3759765625,154.246673583984,154.010940551758,153.891708374023,153.695205688477,153.361129760742,153.27294921875,153.196090698242,153.077728271484,152.882232666016,152.81787109375,152.575576782227,152.400680541992,152.319625854492,152.165237426758,152.087890625,151.70458984375,151.326766967773,151.12109375,151.504974365234,151.733505249023,151.990036010742,152.260635375977,152.169525146484,152.1044921875,151.9423828125,151.798049926758,151.485748291016,151.348236083984,151.170303344727,151.033599853516,150.982513427734,150.911911010742,150.86328125,150.823425292969,150.7294921875,150.615234375,150.483581542969,150.539840698242,150.667282104492,150.457229614258,150.325592041016,150.202545166016,149.642578125,149.424514770508,149.290435791016,149.065231323242,149.127731323242,149.175384521484,149.204971313477,149.133010864258,148.924987792969,148.797058105469,148.708892822266,148.744140625,148.8896484375,148.964645385742,148.9140625,148.726654052734,148.4912109375,148.257431030273,147.874603271484,147.687896728516,147.514450073242,147.0400390625,146.8037109375,146.537200927734,146.4443359375,146.2734375,146.049514770508,145.931640625,145.8291015625,145.756439208984,145.554595947266,144.4833984375,144.123443603516,143.868759155273,143.523834228516,143.192184448242,142.580261230469,142.330368041992,142.025390625,141.754684448242,141.602935791016,141.347076416016,140.987701416016,140.790237426758,140.684951782227,140.4951171875,140.446868896484,140.002349853516,139.861526489258,139.803314208984,139.619247436523,139.506637573242,139.44384765625,139.181640625,138.965713500977,138.662109375,138.2177734375,138.180084228516,138.140625,138.073837280273,137.691497802734,137.572952270508,137.384078979492,137.189834594727,137.012100219727,136.793548583984,136.460250854492,136.351181030273,136.175186157227,135.750778198242,135.540618896484,135.262496948242,135.234771728516,135.211517333984,135.257720947266,135.325393676758,135.437789916992,135.8515625,136.237991333008,136.580276489258,136.714553833008,136.797271728516,136.82373046875,136.820404052734,136.770416259766,136.729385375977,136.683013916016,136.718841552734,136.802642822266,136.886428833008,137.018737792969,137.155380249023,137.258010864258,137.172470092773,137.09619140625,137.1416015625,137.377746582031,137.525100708008,137.666015625,137.51318359375,137.451278686523,137.403427124023,137.339263916016,137.476455688477,137.622756958008,137.834762573242,137.7861328125,137.644821166992,137.516998291016,137.313674926758,137.221481323242,137.253723144531,137.328323364258,137.738189697266,137.950485229492,138.2529296875,138.37890625,138.493560791016,138.527923583984,138.568161010742,138.569137573242,138.407028198242,138.292190551758,138.249710083008,138.3203125,138.45068359375,138.510940551758,138.660736083984,138.699401855469,138.7216796875,138.704681396484,138.715927124023,138.6572265625,138.695709228516,139.105087280273,139.319732666016,139.707427978516,139.795501708984,139.8583984375,140.1787109375,140.24169921875,140.347061157227,140.687591552734,141.005661010742,141.015029907227,141.217666625977,141.373733520508,141.402038574219,141.327926635742,141.181259155273,140.887313842773,140.839645385742,140.87451171875,141.086822509766,141.255859375,141.265914916992,141.245025634766,141.132415771484,141.169815063477,141.329681396484,141.409088134766,141.485244750977,141.385543823242,141.366897583008,141.258392333984,141.12939453125,140.9326171875,140.838562011719,140.687683105469,140.670700073242,140.645599365234,140.520889282227,140.476364135742,140.535446166992,140.564056396484,140.62451171875,140.61328125,140.584564208984,140.462692260742,140.464553833008,140.511337280273,140.517181396484,140.431045532227,140.399108886719,140.364349365234,140.3486328125,140.325576782227,140.308990478516,140.333694458008,140.378326416016,140.224227905273,140.170608520508,140.11328125,139.998443603516,139.7607421875,139.67626953125,139.5205078125,139.372665405273,139.1669921875,139.001373291016,138.586822509766,138.529693603516,138.50048828125,138.391784667969,138.3369140625,138.210159301758,138.106338500977,137.769134521484,137.685455322266,137.425201416016,137.14697265625,136.803512573242,136.737197875977,136.604095458984,136.46044921875,136.251174926758,136.208694458008,136.142288208008,135.987014770508,135.874603271484,135.533203125,135.489074707031,135.4833984375,135.260162353516,135.131057739258,134.9169921875,134.691787719727,134.156448364258,134.010437011719,133.709381103516,133.586730957031,133.329483032227,133.159957885742,133.059387207031,132.99658203125,132.923919677734,132.863571166992,132.708984375,132.576461791992,132.481338500977,132.303817749023,132.334365844727,132.3095703125,132.233200073242,132.028701782227,131.947265625,131.866592407227,131.898345947266,132.013092041016,131.976272583008,131.93896484375,131.794723510742,131.722076416016,131.516418457031,131.393264770508,131.29248046875,131.245315551758,131.158309936523,131.024795532227,130.945709228516,130.756164550781,130.709365844727,130.834182739258,130.729888916016,130.687316894531],[107.695503234863,107.606246948242,107.481636047363,107.343849182129,107.001663208008,106.41552734375,106.583297729492,107.50830078125,107.5732421875,107.695503234863],[106.270408630371,106.151077270508,106.023635864258,106.058395385742,106.3505859375,106.456840515137,106.640426635742,106.691207885742,106.719627380371,106.718948364258,106.679100036621,106.504692077637,106.472457885742,106.270408630371],[102.884765625,102.787307739258,102.745796203613,102.447853088379,102.412307739258,102.587303161621,102.747657775879,102.844825744629,102.950386047363,103.07568359375,103.199127197266,103.433403015137,103.6728515625,103.80078125,103.925682067871,104.003997802734,104.091110229492,104.404205322266,104.44921875,104.476951599121,104.452049255371,104.633201599121,104.881050109863,105.0146484375,105.14599609375,105.20458984375,105.256057739258,105.310157775879,105.342674255371,105.312599182129,104.832618713379,104.741798400879,104.519432067871,104.297462463379,103.719337463379,103.003128051758,102.796684265137,102.734375,102.673149108887,102.6171875,102.180473327637,101.6923828125,101.2041015625,101.039939880371,100.541213989258,100.082229614258,99.8450164794922,99.5002899169922,99.3917007446289,99.287109375,99.4386749267578,99.5456085205078,99.6779327392578,100.018943786621,100.057518005371,100.12353515625,100.162986755371,100.215042114258,100.257225036621,100.2626953125,100.283988952637,100.416404724121,100.515625,100.61962890625,100.875579833984,100.955764770508,100.89794921875,100.856246948242,100.864555358887,100.9013671875,100.965423583984,101.030860900879,101.068168640137,101.05224609375,101.148834228516,101.196098327637,101.310447692871,101.543060302734,101.555274963379,101.590629577637,101.643356323242,101.761329650879,101.82421875,101.912109375,102.005279541016,102.128509521484,102.251274108887,102.17724609375,102.1806640625,102.22509765625,102.282424926758,102.404884338379,102.789848327637,103.041603088379,103.097946166992,103.052436828613,102.939643859863,102.884765625],[76.2489242553711,76.37255859375,76.4673843383789,77.3601531982422,77.54931640625,77.5889663696289,76.8101577758789,76.6495132446289,76.6365280151367,76.4576187133789,76.1537170410156,76.0718765258789,76.0515594482422,76.1484375,76.2489242553711],[100.135932922363,99.9154281616211,99.9422836303711,99.9557647705078,100.068359375,100.141502380371,100.30029296875,100.135932922363],[92.6834945678711,92.4406280517578,92.1537094116211,91.68359375,91.3762741088867,91.1260757446289,91.0703125,91.2293014526367,91.4259719848633,91.751953125,92.1734390258789,92.5927734375,93.4815444946289,93.8031234741211,93.603515625,93.3820343017578,93.1550750732422,92.92626953125,92.6834945678711],[51.4092750549316,51.4351577758789,51.4312477111816,51.0762710571289,50.4541015625,50.0914039611816,50.47265625,50.67578125,50.9363288879395,51.25439453125,51.2378883361816,51.2427749633789,51.3269538879395,51.4092750549316],[59.6888656616211,59.3306617736816,59.20263671875,59.1692390441895,59.1003913879395,58.9192390441895,58.9460983276367,59.0014610290527,59.5445289611816,59.8016586303711,59.9110374450684,59.6888656616211],[97.6745147705078,97.9036102294922,98.0177688598633,97.90673828125,97.8079147338867,97.7599563598633,97.626953125,97.59130859375,97.6516571044922,97.7245101928711,97.8707046508789,98.0645523071289,98.2732391357422,98.3531265258789,98.4990234375,98.4718780517578,98.5318374633789,98.5964889526367,98.8659133911133,99.294921875,99.3707046508789,99.4730453491211,99.5361328125,99.7265625,99.818359375,99.9465789794922,100.061233520508,99.9158172607422,99.8392562866211,99.8054733276367,99.7816390991211,99.7710952758789,99.7488250732422,99.72119140625,99.7062454223633,99.7215805053711,99.6806640625,99.5373001098633,99.3877868652344,99.1670913696289,99.1043930053711,99.0418014526367,99.3173828125,99.5172805786133,99.7507781982422,99.8146514892578,99.8996124267578,99.9292907714844,99.5408172607422,99.4395523071289,98.8195343017578,98.4111328125,98.2825241088867,98.05419921875,97.9051742553711,97.6885757446289,97.5554656982422,97.2481460571289,96.9329071044922,96.8711929321289,96.8078079223633,96.4299850463867,96.3473663330078,95.7964859008789,95.7028350830078,95.5310592651367,95.4369125366211,95.1332015991211,95.0204086303711,94.791015625,94.65234375,94.6316452026367,94.6197204589844,94.4821319580078,94.3137741088867,94.21875,93.7585906982422,93.4787139892578,93.2722625732422,93.07080078125,93.4046859741211,93.8472671508789,94.0381774902344,94.2571258544922,94.3472671508789,94.7194290161133,94.8150405883789,94.94677734375,94.9873046875,95.2813491821289,95.3379898071289,95.3907241821289,95.49755859375,95.8578109741211,96.1624984741211,96.27734375,96.4166030883789,97.1205062866211,97.5868225097656,97.6745147705078],[50.0517578125,49.9709014892578,49.5882797241211,49.5560531616211,49.8836936950684,50.2509765625,50.3099594116211,50.3191413879395,50.072265625,50.0517578125],[55.4796867370605,55.1951179504395,55.0484390258789,54.9796867370605,55.0916023254395,55.2400398254395,55.3532218933105,55.4347648620605,55.4796867370605],[57.0787124633789,57.1226539611816,57.1189460754395,57.07275390625,57.0801734924316,56.9869155883789,56.2005844116211,55.8116188049316,55.7240219116211,55.9422836303711,56.01220703125,55.9898452758789,56.0244140625,56.6550788879395,56.7072257995605,56.9445304870605,57.0787124633789],[53.5213851928711,52.8563461303711,52.6359367370605,52.6070327758789,52.5504875183105,52.3435554504395,52.21337890625,52.2702140808105,52.57666015625,52.6805648803711,52.7160148620605,52.8539047241211,53.1856460571289,53.3291969299316,53.3458976745605,53.4861335754395,53.8516616821289,53.7779273986816,53.6529273986816,53.5213851928711],[57.9562530517578,57.8000984191895,57.3922843933105,57.3323287963867,57.2814445495605,57.2140617370605,57.2117156982422,57.1862297058105,57.0833969116211,57.0111312866211,57.0750007629395,57.5219764709473,58.48046875,58.9716835021973,59.1159172058105,59.2554702758789,58.3979454040527,58.2838897705078,58.2857398986816,58.2554702758789,58.1631851196289,57.9562530517578],[54.4153327941895,54.27587890625,53.8119163513184,53.8499984741211,53.9001960754395,53.9015617370605,53.85888671875,53.8772468566895,54.1767578125,54.2053718566895,54.4071273803711,54.4373054504395,54.4153327941895],[47.4419937133789,47.8995132446289,48.2432632446289,48.34521484375,48.4457015991211,48.54736328125,48.6865234375,48.68359375,48.62548828125,48.0443344116211,47.77734375,47.7052726745605,47.60009765625,47.5123062133789,47.4141578674316,47.3039054870605,47.1982421875,47.1449241638184,47.0110359191895,46.6775398254395,46.6239280700684,46.513671875,46.3781242370605,46.1414070129395,46.0598640441895,46.0236320495605,45.9690399169922,45.6408195495605,45.3892555236816,45.1492156982422,44.9049797058105,45.12451171875,46.3274421691895,46.7991218566895,47.0206069946289,47.3523406982422,47.4419937133789],[62.1677742004395,62.2277374267578,62.1917953491211,62.1145515441895,62.0757789611816,61.7691421508789,61.6812515258789,61.5974617004395,61.28515625,61.05126953125,60.7222671508789,60.2783164978027,59.9001922607422,59.6498031616211,59.3463859558105,59.3043937683105,59.2881851196289,59.3062515258789,59.3865203857422,59.4951133728027,59.5494155883789,59.5922889709473,59.7158164978027,60.0945320129395,60.2349586486816,60.2780227661133,60.4815444946289,60.8202133178711,61.3131790161133,61.5974617004395,61.8505859375,62.1029281616211,62.1677742004395],[50.2781257629395,50.4314460754395,50.8010749816895,50.9176788330078,51.4547843933105,51.5910148620605,51.70361328125,51.1461906433105,50.9608421325684,50.2796859741211,49.8459968566895,49.7498054504395,49.7941398620605,49.5859375,48.8960952758789,48.81103515625,48.6770515441895,48.68896484375,48.9219703674316,48.9595718383789,48.9908180236816,49.0107421875,48.9775390625,48.8918952941895,48.79736328125,48.5817375183105,48.5545883178711,48.5326156616211,48.466796875,48.38623046875,48.1671867370605,48.0958976745605,48.0257835388184,47.93994140625,47.7373046875,47.6324195861816,47.72314453125,47.9775390625,47.8929672241211,47.6423835754395,47.4443359375,47.3430671691895,47.2486343383789,46.9910163879395,46.8458976745605,46.7381820678711,46.6444358825684,47.4029312133789,47.6560516357422,47.8958015441895,48.2082023620605,48.30615234375,48.4026374816895,48.4647445678711,48.6250953674316,49.0877952575684,49.1852531433105,49.1926765441895,49.1474609375,49.2443351745605,49.5078125,50.1243171691895,50.2781257629395],[80.0266647338867,79.0985336303711,79.0068359375,78.9776382446289,79.10986328125,79.2173843383789,79.806640625,80.2795944213867,80.4279327392578,80.3733444213867,80.3448257446289,80.0266647338867],[61.1408195495605,60.8267555236816,60.3210945129395,60.0582008361816,60.0783233642578,60.1475601196289,60.5866203308105,61.4574203491211,61.5673789978027,61.4719772338867,61.1408195495605],[54.7189445495605,55.470703125,56.1701164245605,56.4722671508789,56.90966796875,57.5677719116211,57.6941452026367,57.5803718566895,56.8147468566895,56.3155288696289,55.8833999633789,55.7125015258789,55.5406265258789,55.1171875,54.6681632995605,54.6233367919922,54.5328102111816,54.3760757446289,54.0666007995605,54.04541015625,54.2405281066895,54.3672828674316,54.4167976379395,54.6339836120605,54.7189445495605],[58.6223678588867,58.7615242004395,58.8153343200684,58.9025421142578,58.9305648803711,58.8599624633789,58.6418952941895,58.28564453125,57.9378929138184,57.7498016357422,57.4051780700684,57.2109375,57.4102516174316,57.65625,58.0495109558105,58.1023445129395,58.18994140625,58.5076179504395,58.6223678588867],[50.7537117004395,50.6160163879395,50.5181617736816,50.4119148254395,50.37744140625,50.3684577941895,50.4649429321289,50.5059585571289,50.5215797424316,50.591796875,50.7159156799316,50.8786125183105,50.9461898803711,50.7887725830078,50.7537117004395],[63.3738288879395,63.1875953674316,63.0021514892578,62.7604484558105,62.5203132629395,62.5925788879395,62.8193321228027,63.1158180236816,63.6147422790527,63.85595703125,64.095703125,64.1659164428711,64.21044921875,64.255859375,64.3101577758789,64.5753860473633,64.8020477294922,65.0277328491211,65.1719741821289,65.3097610473633,65.3820343017578,65.3600616455078,65.3720703125,65.4374008178711,64.9974594116211,64.54833984375,63.3738288879395],[91.5671920776367,91.2228546142578,89.9757843017578,89.91943359375,89.9011688232422,90.0699234008789,91.1089859008789,91.4778289794922,91.5671920776367],[96.5265655517578,96.5630798339844,96.6932601928711,96.7549743652344,97.4136734008789,97.7030258178711,97.8318405151367,97.8699264526367,97.8564453125,97.7471618652344,97.6654281616211,97.2213821411133,97.1130828857422,97.025390625,97.0725631713867,97.1150360107422,97.2501907348633,97.2868194580078,97.4169921875,97.2984390258789,97.1752014160156,95.8557662963867,94.9613265991211,94.6612319946289,94.5650405883789,94.3284149169922,93.8723678588867,93.6546859741211,93.0023422241211,92.2015609741211,92.0921859741211,91.8916015625,91.6374053955078,91.5238265991211,91.6877899169922,91.8966751098633,92.2466735839844,92.5779266357422,92.8267593383789,92.9810485839844,93.2624969482422,92.77294921875,92.5925750732422,92.6101608276367,92.7103500366211,92.7646484375,92.9386672973633,93.0651397705078,93.3586959838867,93.4973602294922,93.63671875,93.8888626098633,94.14013671875,94.37548828125,94.6116256713867,94.837890625,95.0609436035156,95.1595687866211,95.8006820678711,95.9019546508789,95.9839859008789,96.0751953125,96.1869125366211,96.4710922241211,96.5265655517578],[59.3130836486816,59.0969772338867,58.7190399169922,58.6101531982422,58.6344680786133,58.8805694580078,59.0750045776367,59.2808570861816,59.3746109008789,59.3130836486816],[57.8102569580078,57.8626976013184,58.0166053771973,58.4360389709473,58.5638656616211,58.3718757629395,57.8586883544922,57.9119186401367,58.0153312683105,57.9128952026367,57.7697257995605,57.4509735107422,57.1594734191895,56.8218727111816,56.6692390441895,56.5125007629395,56.3639640808105,56.1919937133789,55.7166976928711,55.5726547241211,55.4660148620605,55.7819328308105,56.1568336486816,56.4046897888184,56.71875,56.9730453491211,57.0915069580078,57.3650360107422,57.4564476013184,57.7166023254395,57.8102569580078],[63.6509780883789,63.5285186767578,62.8849639892578,62.5730514526367,62.53125,62.5152320861816,62.1064453125,62.2839813232422,62.794921875,63.7095718383789,63.7673835754395,63.7824249267578,63.6509780883789],[58.29541015625,57.96484375,57.9206047058105,57.9092788696289,57.9451179504395,57.9849586486816,58.1345672607422,59.2618141174316,59.4084930419922,59.3568382263184,59.3564491271973,58.29541015625],[30.5099601745605,30.4770526885986,30.47021484375,30.5081062316895,30.6319332122803,30.7107429504395,30.7622051239014,30.8125991821289,30.8275394439697,30.8067378997803,30.8191413879395,30.8646488189697,30.8765640258789,30.85498046875,30.8287105560303,30.7976570129395,30.7624988555908,30.7148456573486,30.6566410064697,30.5933589935303,30.5536117553711,30.5289058685303,30.4822254180908,30.4084968566895,30.27099609375,30.2337875366211,30.1833000183105,30.1422843933105,30.1172847747803,30.0918941497803,29.9734382629395,29.93017578125,29.9124031066895,29.892578125,29.8681640625,29.7833976745605,29.6980476379395,29.6513671875,29.4636707305908,29.3902359008789,29.3498039245605,29.2970714569092,29.1975593566895,29.1020488739014,29.0631828308105,29.0286140441895,29.0143566131592,28.9217777252197,28.8939437866211,28.8914070129395,28.8576183319092,28.8763656616211,28.9126968383789,28.9895496368408,29.1064453125,29.1315422058105,29.1480464935303,29.1406269073486,29.12939453125,29.1432628631592,29.1965827941895,29.2681636810303,29.3516578674316,29.4019527435303,29.4679698944092,29.5377922058105,29.5769519805908,29.6096687316895,29.8253898620605,29.8468742370605,29.8816413879395,29.8999996185303,29.9300765991211,29.9905281066895,30.1015625,30.1499977111816,30.20703125,30.2798824310303,30.3205070495605,30.3602523803711,30.4123058319092,30.4699211120605,30.5099601745605],[-8.683349609375,-8.68310546875,-8.682861328125,-8.6826171875,-8.68232345581055,-8.68212890625,-8.68212890625,-8.68222618103027,-8.88564395904541,-9.07192420959473,-9.25820350646973,-9.44453144073486,-9.630859375,-9.81718826293945,-10.0035152435303,-10.1897945404053,-10.3761224746704,-10.5624513626099,-10.748779296875,-10.9351072311401,-11.1213865280151,-11.3077144622803,-11.4940433502197,-11.6803703308105,-11.8666505813599,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0163078308105,-12.0234375,-12.0833501815796,-12.2261714935303,-12.3729009628296,-12.5593748092651,-12.6204099655151,-12.7395992279053,-12.89599609375,-13.031494140625,-13.1208982467651,-13.1532707214355,-13.16650390625,-13.1559572219849,-13.1073246002197,-13.0943355560303,-13.0867671966553,-13.0784664154053,-13.069580078125,-13.0606441497803,-13.0512199401855,-13.041748046875,-13.0322265625,-13.0250968933105,-13.0162115097046,-13.1674318313599,-13.396728515625,-13.6260261535645,-13.8553705215454,-14.0846681594849,-14.31396484375,-14.5432624816895,-14.7726078033447,-15.0019044876099,-15.2312002182007,-15.4605464935303,-15.6897945404053,-15.9191408157349,-16.1484375,-16.3777351379395,-16.6070308685303,-16.8363265991211,-16.9645500183105,-17.0059070587158,-17.0423831939697,-17.06396484375,-17.0480461120605,-17.0987796783447,-17.0096187591553,-17.0030765533447,-17.0029792785645,-16.9511222839355,-16.73095703125,-16.5810070037842,-16.1908702850342,-16.041015625,-15.9208498001099,-15.7509288787842,-15.6107902526855,-15.4609375,-15.2909660339355,-15.15087890625,-14.9711427688599,-14.8408212661743,-14.7509765625,-14.6708498001099,-14.64111328125,-14.6107902526855,-14.62109375,-14.630859375,-14.5810060501099,-14.52099609375,-14.4608888626099,-14.4409189224243,-14.3808097839355,-14.31103515625,-14.27099609375,-14.22119140625,-14.2109375,-14.1908693313599,-14.1908693313599,-14.1708984375,-14.1410636901855,-14.12109375,-14.10107421875,-14.0409669876099,-14.02099609375,-13.9809083938599,-13.9311027526855,-13.89111328125,-13.8407716751099,-13.7709474563599,-13.6610832214355,-13.5810546875,-13.48095703125,-13.39111328125,-13.3109865188599,-13.28076171875,-13.23095703125,-13.1611328125,-13.12109375,-13.06103515625,-12.9911623001099,-12.9478511810303,-12.9111328125,-12.8207521438599,-12.7109375,-12.6308097839355,-12.56103515625,-12.5009765625,-12.43115234375,-12.40087890625,-12.3608388900757,-12.3109865188599,-12.2709474563599,-12.23095703125,-12.2011232376099,-12.1708498001099,-12.1308603286743,-12.1010255813599,-12.0810537338257,-12.0810537338257,-12.0609865188599,-12.0567874908447,-12.03076171875,-11.9608888626099,-11.8808584213257,-11.7548828125,-11.7182130813599,-11.6992197036743,-11.6845216751099,-11.63720703125,-11.583984375,-11.5531740188599,-11.5116701126099,-11.470703125,-11.39990234375,-11.337890625,-11.3168458938599,-11.3168458938599,-11.3612794876099,-11.392578125,-11.2636232376099,-11.1503419876099,-11.0468263626099,-10.9228029251099,-10.830078125,-10.7577638626099,-10.6542482376099,-10.55126953125,-10.4789543151855,-10.3549308776855,-10.2514638900757,-10.189453125,-10.123046875,-10.0668458938599,-10.03271484375,-9.98090839385986,-9.90034198760986,-9.81787109375,-9.7353515625,-9.67333984375,-9.56982421875,-9.4873046875,-9.41303730010986,-9.35297870635986,-9.28559589385986,-9.20844745635986,-9.08442401885986,-9.00190448760986,-8.88906288146973,-8.79487228393555,-8.75385761260986,-8.75385761260986,-8.77436542510986,-8.79682636260986,-8.80268573760986,-8.78896427154541,-8.77436542510986,-8.78457069396973,-8.81391620635986,-8.81777381896973,-8.81782245635986,-8.683349609375,-8.683349609375,-8.683349609375],[41.9876976013184,42.0650367736816,42.033203125,42.0263671875,42.0599594116211,42.17041015625,42.1671867370605,42.1578140258789,42.1277351379395,42.1083984375,42.1023445129395,42.07177734375,41.9641609191895,41.8972663879395,41.8015632629395,41.7760734558105,41.8161125183105,41.8582038879395,41.8849601745605,41.8604507446289,41.9172859191895,41.9479484558105,41.9539070129395,41.9625015258789,41.9466819763184,41.9876976013184],[36.9016609191895,36.8751945495605,36.8050804138184,36.7638702392578,36.7220726013184,36.5302734375,36.5042991638184,36.5335922241211,36.5541000366211,36.5887718200684,36.74755859375,36.9244117736816,36.9547882080078,36.9016609191895],[36.5955085754395,36.5861320495605,36.5439453125,36.5464859008789,36.5827140808105,36.5798835754395,36.5956077575684,36.5955085754395],[46.5314445495605,46.7248039245605,46.9822273254395,47.1387672424316,47.4332008361816,47.5212898254395,47.55322265625,47.5831069946289,47.6712875366211,47.8719711303711,48.0496101379395,48.2687492370605,48.4424819946289,48.49853515625,48.5230484008789,48.6263656616211,48.7737312316895,48.8089828491211,48.8328132629395,48.8072280883789,48.7971687316895,48.9064445495605,49.0869140625,49.1575202941895,49.2374992370605,49.1750984191895,49.2815437316895,49.4052734375,49.5376968383789,49.7165031433105,49.9861297607422,50.1498031616211,50.1346664428711,50.0866203308105,50.0263671875,50.0081024169922,50.0113296508789,50.02734375,50.1107406616211,50.1849632263184,50.2138671875,50.1554679870605,50.13525390625,50.0959968566895,50.0539093017578,50.0316429138184,50.0810546875,50.1302719116211,50.1896476745605,50.2389640808105,50.28125,50.4551734924316,50.5084991455078,50.5579109191895,50.6668930053711,50.7255859375,50.8043937683105,50.8556632995605,50.9283180236816,50.9660148620605,51.0227546691895,51.0933609008789,51.1780281066895,51.2679710388184,51.3384780883789,51.4112281799316,51.4183616638184,51.3699226379395,51.3098640441895,51.3952140808105,51.4767570495605,51.5347633361816,51.568359375,51.568359375,51.5721664428711,51.5925788879395,51.6292991638184,51.6843757629395,51.7393569946289,51.7943344116211,51.8494148254395,51.9043922424316,51.95947265625,52.0144500732422,52.0694351196289,52.12451171875,52.1794929504395,52.2345733642578,52.28955078125,52.3445320129395,52.3996086120605,52.45458984375,52.5095672607422,52.5550765991211,52.63916015625,52.6659202575684,52.7416000366211,52.8592796325684,53.0119132995605,53.1923828125,53.39404296875,53.6095695495605,53.8321304321289,54.0545883178711,54.2701187133789,54.4716796875,54.6522483825684,54.8048820495605,54.9224586486816,54.9982452392578,55.0250015258789,55.1042976379395,55.1194343566895,55.1858367919922,55.25927734375,55.3201179504395,55.40380859375,55.4927749633789,55.5777359008789,55.6410140991211,55.607421875,55.57080078125,55.5341796875,55.49755859375,55.4609375,55.42431640625,55.3876953125,55.35107421875,55.314453125,55.2779312133789,55.2412109375,55.20458984375,55.1680679321289,55.1314468383789,55.0947265625,55.0582008361816,55.021484375,54.9773445129395,54.87109375,54.6990242004395,54.5270500183105,54.35498046875,54.1830062866211,54.0109367370605,53.8388671875,53.6668930053711,53.4948234558105,53.3228530883789,53.1507797241211,52.9787101745605,52.8067359924316,52.6346664428711,52.4626960754395,52.2906227111816,52.1185531616211,51.9776382446289,51.7429695129395,51.5149383544922,51.2583961486816,50.9500007629395,50.7082023620605,50.3552742004395,50.0389671325684,49.7420883178711,49.4451141357422,49.1923828125,49.0419921875,48.8648414611816,48.5929679870605,48.3158187866211,48.1721649169922,48.0216789245605,47.9455070495605,47.8078117370605,47.7037124633789,47.57958984375,47.525390625,47.4417991638184,47.36962890625,47.2512702941895,47.1435546875,46.9756851196289,46.8799781799316,46.7785148620605,46.7276344299316,46.6820297241211,46.5134773254395,46.3103523254395,46.07080078125,45.79443359375,45.5353507995605,45.4065437316895,45.2366180419922,45.1927719116211,45.1480445861816,44.9464836120605,44.7467765808105,44.5464820861816,44.3546867370605,44.1559562683105,44.0859375,44.0082015991211,43.9596672058105,43.9169921875,43.8664054870605,43.8042984008789,43.7129898071289,43.6534194946289,43.5972633361816,43.5392570495605,43.4742202758789,43.41796875,43.3460960388184,43.3021507263184,43.19091796875,43.1863288879395,43.2369155883789,43.2213859558105,43.1559562683105,43.1359367370605,43.1261711120605,43.1165046691895,43.1456031799316,43.1844711303711,43.1863288879395,43.1650390625,43.1047859191895,43.0607414245605,43.0335922241211,42.986328125,42.79931640625,42.7898445129395,42.7306632995605,42.7263679504395,42.6988296508789,42.6474609375,42.5529289245605,42.5441436767578,42.4749984741211,42.38330078125,42.3324241638184,42.2939453125,42.05224609375,41.75,41.6580085754395,41.5076179504395,41.4317398071289,41.2294921875,41.2207984924316,41.1908226013184,41.1441421508789,41.1160125732422,40.9132804870605,40.8478507995605,40.7915992736816,40.7770538330078,40.7591819763184,40.6159172058105,40.4822273254395,40.0806617736816,39.8840827941895,39.7283210754395,39.6136741638184,39.4912109375,39.2760734558105,39.0935554504395,39.1506805419922,39.1470680236816,39.0910186767578,39.02978515625,38.9878921508789,39.0211906433105,39.0339851379395,39.0699234008789,39.0958976745605,39.06201171875,39.0013656616211,39.0074234008789,38.9387702941895,38.8829116821289,38.9411163330078,38.8355484008789,38.796875,38.7570304870605,38.7060546875,38.5422859191895,38.4641609191895,38.2888641357422,38.0986328125,37.9778327941895,37.9197273254395,37.8209991455078,37.71337890625,37.63818359375,37.5430679321289,37.4309577941895,37.3384742736816,37.1808586120605,37.2204093933105,37.2663078308105,37.2434577941895,37.2183609008789,37.1488304138184,36.9207038879395,36.8601570129395,36.7626953125,36.7025413513184,36.6751976013184,36.5187492370605,36.2496109008789,36.09375,36.0320320129395,35.8516578674316,35.7629890441895,35.5813484191895,35.423828125,35.1804656982422,35.0783233642578,34.8275375366211,34.7220687866211,34.625,34.6162109375,34.6833000183105,34.7798843383789,34.7991218566895,34.9507827758789,35.1637725830078,35.3391609191895,35.5953140258789,35.8603515625,36.0154304504395,36.0684547424316,36.2828102111816,36.47607421875,36.591796875,36.7039070129395,36.7552719116211,36.9270515441895,37.1994132995605,37.46923828125,37.49072265625,37.5536117553711,37.6335945129395,37.64990234375,37.6697273254395,37.8628921508789,37.9800758361816,37.81298828125,37.6554679870605,37.47900390625,37.3294944763184,37.1052742004395,36.9585952758789,37.2156257629395,37.4933586120605,37.7738265991211,38.1114234924316,38.37548828125,38.7696304321289,38.9623069763184,38.9970703125,39.1454086303711,39.36865234375,39.7041015625,40.02783203125,40.3693351745605,40.4789085388184,40.8083992004395,41.0224609375,41.2724609375,41.5850563049316,41.7997055053711,42.0744132995605,42.28857421875,42.5597648620605,42.8577156066895,43.1031227111816,43.4408187866211,43.7737312316895,44.099609375,44.3607444763184,44.6908187866211,44.7165031433105,45.05029296875,45.4989242553711,45.94970703125,46.3564453125,46.5314445495605],[36.8713874816895,36.8826179504395,36.9269523620605,37.0811500549316,37.2117156982422,37.2585945129395,37.26318359375,37.2572288513184,37.2174797058105,37.1505889892578,37.14111328125,37.1568374633789,37.1726570129395,37.2275390625,37.1878890991211,37.1931648254395,37.2625999450684,37.2484359741211,37.3615226745605,37.4712867736816,37.5316390991211,37.5994110107422,37.7297821044922,37.921875,38.0740203857422,38.1281242370605,38.2017555236816,38.2521514892578,38.2831039428711,38.3329086303711,38.5740242004395,38.6094703674316,38.5228500366211,38.4224624633789,38.3971672058105,38.3855476379395,38.3737297058105,38.3473625183105,38.2898406982422,38.2672843933105,38.2535171508789,38.2190437316895,38.1815414428711,38.1485366821289,38.0989265441895,38.0252914428711,37.9500999450684,37.9225578308105,37.8629913330078,37.8033218383789,37.7824211120605,37.7259750366211,37.65673828125,37.5759735107422,37.5474624633789,37.5101547241211,37.4529304504395,37.4110374450684,37.3404273986816,37.2488288879395,37.1695327758789,37.0615234375,37.0089836120605,36.9952125549316,36.9757804870605,36.9787139892578,36.9357414245605,36.8877906799316,36.9054679870605,36.9137725830078,36.8258819580078,36.8134765625,36.7245140075684,36.67919921875,36.5660171508789,36.5217781066895,36.4267578125,36.4481430053711,36.4707984924316,36.4922866821289,36.5243186950684,36.4439468383789,36.4470710754395,36.390625,36.3462867736816,36.3068351745605,36.2735366821289,36.2122077941895,36.16015625,36.1371116638184,36.1353530883789,36.1251945495605,36.1075172424316,35.9875984191895,35.8206062316895,35.7305679321289,35.6702156066895,35.5960960388184,35.4496116638184,35.3727531433105,35.25244140625,35.1123046875,35.0827140808105,35.0596694946289,35.0079078674316,34.9607429504395,34.9691429138184,34.9249000549316,34.9314460754395,34.88232421875,34.8162078857422,34.7712860107422,34.6749992370605,34.6017570495605,34.5718765258789,34.5080070495605,34.4314460754395,34.3439483642578,34.2756805419922,34.3148422241211,34.3112297058105,34.29150390625,34.1852531433105,34.1590805053711,34.1203117370605,34.0792961120605,34.078125,34.0767555236816,33.8909187316895,33.8878898620605,33.8848609924316,33.8821296691895,33.8788070678711,33.8714866638184,33.8677711486816,33.8740234375,33.8949241638184,33.9591827392578,33.9625015258789,33.9632797241211,33.9632797241211,33.9573249816895,33.9499015808105,33.9461898803711,33.9573249816895,33.9584007263184,33.9568367004395,33.9518547058105,33.9070281982422,33.8921890258789,33.4590835571289,33.3798828125,33.3713874816895,33.3607406616211,33.1408195495605,33.1300773620605,33.1314430236816,33.1382827758789,33.1647453308105,33.16845703125,33.1721687316895,33.0730476379395,33.0733413696289,33.0778312683105,33.08154296875,33.0946311950684,33.1060523986816,33.119140625,33.1224594116211,33.1361351013184,33.1350593566895,33.1930656433105,33.1993179321289,32.7218742370605,32.7189445495605,32.7185554504395,32.7198219299316,32.7205047607422,32.7201156616211,32.71630859375,32.71533203125,32.7158203125,32.7230453491211,32.7356452941895,32.7376937866211,32.73828125,32.7367172241211,32.072265625,32.3353500366211,32.3384780883789,32.3430671691895,32.3449211120605,32.3499031066895,32.3357429504395,32.3388671875,32.3541984558105,32.4253921508789,32.4207992553711,32.4041023254395,31.9330101013184,31.919921875,31.8542976379395,31.7919940948486,31.7642593383789,31.6548843383789,31.2249031066895,31.1544933319092,30.9403343200684,30.8270511627197,30.8141593933105,30.794921875,30.7831058502197,30.7691421508789,30.75537109375,30.7393550872803,30.474609375,30.0030269622803,29.9579105377197,29.6910171508789,29.6359386444092,29.60546875,29.6039085388184,29.5574207305908,29.47314453125,29.2423820495605,29.1223621368408,28.9996109008789,28.9795913696289,28.9795913696289,28.9323234558105,28.8394527435303,28.8293952941895,28.8445301055908,28.0489273071289,27.9962902069092,27.8858413696289,27.880859375,27.7998046875,27.07421875,26.9705085754395,26.76318359375,26.65869140625,26.5513668060303,26.1695308685303,26.0870113372803,26.0570316314697,26.0005855560303,25.9191398620605,25.8915042877197,25.8828125,25.88525390625,25.8582038879395,25.7980461120605,25.28515625,25.2117195129395,25.10400390625,25.0669918060303,25.0236320495605,25.0148429870605,25.0162124633789,25.0029296875,24.9638671875,24.8176765441895,24.7852535247803,24.7921886444092,24.7826175689697,24.7603511810303,24.6966800689697,24.6736335754395,24.6628913879395,24.6593742370605,24.6480464935303,24.5682621002197,24.5494136810303,24.5448246002197,24.5319347381592,24.3001956939697,24.2135734558105,24.1604480743408,24.1473636627197,24.0481452941895,23.9219722747803,23.6792984008789,23.5832042694092,23.5373039245605,23.5518550872803,23.5280284881592,23.4890632629395,23.4627933502197,23.46826171875,23.5960941314697,23.6226558685303,23.6427726745605,23.65625,23.6462898254395,23.5450191497803,23.4566402435303,23.3123054504395,23.255859375,22.9643535614014,22.9307613372803,22.8600597381592,22.8948230743408,22.9376964569092,22.9427738189697,22.9226570129395,22.8490238189697,22.7833995819092,22.7540035247803,22.6973648071289,22.6410140991211,22.5911140441895,22.5563488006592,22.5809574127197,22.5643558502197,22.4898433685303,22.4724617004395,22.4754886627197,22.4352531433105,22.3902339935303,22.4144515991211,22.3523445129395,22.2333984375,22.1211910247803,22.0006828308105,21.9281234741211,21.8781261444092,21.8433589935303,21.8252925872803,21.841796875,21.90771484375,21.990234375,22.1580085754395,22.20263671875,22.2281246185303,22.2326183319092,22.2213878631592,22.2023448944092,22.1529293060303,22.1076164245605,22.1064453125,22.1282234191895,22.1731452941895,22.2621097564697,22.2834949493408,22.33935546875,22.38818359375,22.5099601745605,22.53857421875,22.5282230377197,22.4983406066895,22.4493179321289,22.4393558502197,22.4249992370605,22.3997077941895,22.3815422058105,22.4162120819092,22.4677734375,22.5320301055908,22.6318359375,22.6708984375,22.6824207305908,22.67919921875,22.7149410247803,22.7632827758789,22.8021488189697,22.8671875,22.9323234558105,22.9613285064697,22.9695320129395,22.9338874816895,23.0091800689697,23.1051750183105,23.2434558868408,23.4580078125,23.60400390625,23.7082023620605,23.9459953308105,23.9652347564697,23.9708023071289,23.9833984375,23.9833011627197,23.98291015625,23.9825191497803,23.9822273254395,23.9818363189697,23.9814453125,23.9810543060303,23.9806652069092,23.9802742004395,23.9802742004395,23.9802742004395,23.9802742004395,23.9802742004395,24.2269515991211,24.4736328125,24.7204093933105,24.9669914245605,24.97021484375,24.9732437133789,24.9763679504395,24.9794921875,24.9796867370605,24.9798831939697,24.9800777435303,24.9802722930908,25.3623046875,25.7443351745605,26.1263675689697,26.5083980560303,26.8904285430908,27.2724609375,27.6544914245605,28.0364265441895,28.4185543060303,28.8005847930908,29.1825199127197,29.5645484924316,29.9466800689697,30.3286151885986,30.7106437683105,31.0926780700684,31.2091808319092,31.2606449127197,31.3584938049316,31.4002914428711,31.4642562866211,31.4861335754395,31.4664039611816,31.4344730377197,31.6216793060303,31.9498043060303,32.27783203125,32.6060562133789,32.93408203125,33.26220703125,33.59033203125,33.91845703125,34.2464828491211,34.5746078491211,34.9027328491211,35.2308616638184,35.5589866638184,35.8870124816895,36.2152328491211,36.5432624816895,36.8713874816895],[34.078125,34.0771484375,34.0845718383789,34.0910148620605,34.1015625,34.1017570495605,34.0945320129395,34.07275390625,34.0197257995605,33.9533195495605,33.7850608825684,33.6448249816895,33.5453147888184,33.4093742370605,33.2810516357422,33.2342758178711,33.1652336120605,33.0652351379395,33.0125961303711,32.9989242553711,33.0146484375,33.0807609558105,33.2259788513184,33.3922843933105,33.5163078308105,33.6009750366211,33.6661109924316,33.9024429321289,33.9779319763184,34.0204124450684,34.0301742553711,34.0642585754395,34.2003898620605,34.279296875,34.484375,34.5627937316895,34.6387672424316,34.7106475830078,34.7492179870605,34.8380851745605,34.8978500366211,34.958984375,34.9835929870605,35.0319328308105,35.0819358825684,35.1644554138184,35.25244140625,35.2683563232422,35.08447265625,34.8783187866211,34.6398429870605,34.3801727294922,34.1768569946289,33.97607421875,33.7416038513184,33.5684585571289,33.53955078125,33.4893569946289,33.3243141174316,33.1540985107422,32.9972648620605,32.8380851745605,32.7371101379395,32.6769523620605,32.5347671508789,32.3357429504395,32.2455062866211,32.1966781616211,32.15625,32.1359367370605,32.0994148254395,32.0482406616211,31.9417953491211,31.8882808685303,31.8386707305908,31.7980480194092,31.62890625,31.5471687316895,31.47998046875,31.357421875,31.2219734191895,31.1523418426514,31.0480461120605,30.9293956756592,30.8681659698486,30.8385734558105,30.81689453125,30.7969703674316,30.7572269439697,30.6999034881592,30.6476573944092,30.5867176055908,30.5593757629395,30.5535163879395,30.5369148254395,30.50830078125,30.4207019805908,30.1949214935303,30.0213871002197,29.9339828491211,29.8702144622803,29.7798843383789,29.6768550872803,29.5520496368408,29.4696292877197,29.3848628997803,29.2249011993408,29.1514663696289,29.0574207305908,28.9393539428711,28.72705078125,28.6395492553711,28.5248050689697,28.4275398254395,28.3671875,28.31103515625,28.2472648620605,28.1920890808105,28.07861328125,28.0198249816895,27.9806632995605,27.9166011810303,27.8416004180908,27.7880859375,27.7614250183105,27.71923828125,27.6641597747803,27.4910163879395,27.4392566680908,27.4033203125,27.3324203491211,27.2567386627197,27.2325191497803,27.2291011810303,27.21337890625,27.1812496185303,27.1439456939697,27.0833988189697,26.9422855377197,26.7964859008789,26.7263660430908,26.5936527252197,26.5142574310303,26.4474601745605,26.4205074310303,26.3533210754395,26.3246097564697,26.30859375,26.36181640625,26.2845706939697,26.1693363189697,26.0869140625,26.0365238189697,25.8889636993408,25.5666027069092,25.3806629180908,25.2789058685303,25.1901378631592,25.1813488006592,25.2386722564697,25.2473640441895,25.2003917694092,25.0072269439697,24.8533191680908,24.7367191314697,24.4560546875,24.37548828125,24.2914047241211,24.2083988189697,24.1799793243408,24.2208995819092,24.19482421875,24.1473636627197,24.1604480743408,24.2135734558105,24.3001956939697,24.5319347381592,24.5448246002197,24.5494136810303,24.5682621002197,24.6480464935303,24.6593742370605,24.6628913879395,24.6736335754395,24.6966800689697,24.7603511810303,24.7826175689697,24.7921886444092,24.7852535247803,24.8176765441895,24.9638671875,25.0029296875,25.0162124633789,25.0148429870605,25.0236320495605,25.0669918060303,25.10400390625,25.2117195129395,25.28515625,25.7980461120605,25.8582038879395,25.88525390625,25.8828125,25.8915042877197,25.9191398620605,26.0005855560303,26.0570316314697,26.0870113372803,26.1695308685303,26.5513668060303,26.65869140625,26.76318359375,26.9705085754395,27.07421875,27.7998046875,27.880859375,27.8858413696289,27.9962902069092,28.0489273071289,28.8445301055908,28.8293952941895,28.8394527435303,28.9323234558105,28.9795913696289,28.9795913696289,28.9996109008789,29.1223621368408,29.2423820495605,29.47314453125,29.5574207305908,29.6039085388184,29.60546875,29.6359386444092,29.6910171508789,29.9579105377197,30.0030269622803,30.474609375,30.7393550872803,30.75537109375,30.7691421508789,30.7831058502197,30.794921875,30.8141593933105,30.8270511627197,30.9403343200684,31.1544933319092,31.2249031066895,31.6548843383789,31.7642593383789,31.7919940948486,31.8542976379395,31.919921875,31.9330101013184,32.4041023254395,32.4207992553711,32.4253921508789,32.3541984558105,32.3388671875,32.3357429504395,32.3499031066895,32.3449211120605,32.3430671691895,32.3384780883789,32.3353500366211,32.072265625,32.7367172241211,32.73828125,32.7376937866211,32.7356452941895,32.7230453491211,32.7158203125,32.71533203125,32.71630859375,32.7201156616211,32.7205047607422,32.7198219299316,32.7185554504395,32.7189445495605,32.7218742370605,33.1993179321289,33.1930656433105,33.1350593566895,33.1361351013184,33.1224594116211,33.119140625,33.1060523986816,33.0946311950684,33.08154296875,33.0778312683105,33.0733413696289,33.0730476379395,33.1721687316895,33.16845703125,33.1647453308105,33.1382827758789,33.1314430236816,33.1300773620605,33.1408195495605,33.3607406616211,33.3713874816895,33.3798828125,33.4590835571289,33.8921890258789,33.9070281982422,33.9518547058105,33.9568367004395,33.9584007263184,33.9573249816895,33.9461898803711,33.9499015808105,33.9573249816895,33.9632797241211,33.9632797241211,33.9625015258789,33.9591827392578,33.8949241638184,33.8740234375,33.8677711486816,33.8714866638184,33.8788070678711,33.8821296691895,33.8848609924316,33.8878898620605,33.8909187316895,34.0767555236816,34.078125],[-12.2806148529053,-12.1865234375,-12.2068367004395,-12.2284183502197,-12.1752443313599,-12.1128902435303,-12.068359375,-12.0191898345947,-12.0111818313599,-12.0201168060303,-11.9880857467651,-11.9608888626099,-11.9663572311401,-11.9841794967651,-12.0441408157349,-12.05419921875,-11.9570798873901,-11.8945808410645,-11.8952150344849,-11.8777828216553,-11.8316898345947,-11.8033685684204,-11.772216796875,-11.7582521438599,-11.6744632720947,-11.6349611282349,-11.5813484191895,-11.5616703033447,-11.5487794876099,-11.4928216934204,-11.4441404342651,-11.4339351654053,-11.390380859375,-11.4174308776855,-11.4143552780151,-11.444091796875,-11.4505863189697,-11.4487791061401,-11.3824224472046,-11.3894033432007,-11.4567384719849,-11.5736808776855,-11.80810546875,-11.8885746002197,-12.0423822402954,-12.1519536972046,-12.2912101745605,-12.3990726470947,-12.4573736190796,-12.5342283248901,-12.6208009719849,-12.7130374908447,-12.7973136901855,-12.88818359375,-12.9307126998901,-12.9605464935303,-12.9856443405151,-13.0119142532349,-13.061279296875,-13.079833984375,-13.0644044876099,-13.0597658157349,-13.0829105377197,-13.1384763717651,-13.2280759811401,-13.37255859375,-13.40576171875,-13.729248046875,-14.0648431777954,-14.3492183685303,-14.7081546783447,-14.9605951309204,-15.1960935592651,-15.3779296875,-15.5748052597046,-15.8395500183105,-16.1441879272461,-16.2415027618408,-16.34228515625,-16.4163093566895,-16.5213375091553,-16.6569328308105,-16.7118167877197,-16.745849609375,-16.7848644256592,-16.7603015899658,-16.6776371002197,-16.55322265625,-16.4880867004395,-16.449951171875,-16.44287109375,-16.4550285339355,-16.548828125,-16.59765625,-16.6378421783447,-16.6725578308105,-16.701416015625,-16.743896484375,-16.7679691314697,-16.7784175872803,-16.7589836120605,-16.7689456939697,-16.7573719024658,-16.7633304595947,-16.7045402526855,-16.6487789154053,-16.4308605194092,-16.2283191680908,-16.0330562591553,-15.8342771530151,-15.8144044876099,-15.7515621185303,-15.6573238372803,-15.4818363189697,-15.2862300872803,-15.2445316314697,-15.2121086120605,-15.1916027069092,-15.151123046875,-15.0963859558105,-15.0246095657349,-14.9502935409546,-14.8650388717651,-14.8082513809204,-14.6719245910645,-14.4385738372803,-14.2467775344849,-14.014892578125,-13.8475103378296,-13.8267087936401,-13.8528318405151,-13.9773921966553,-14.1469717025757,-14.1990232467651,-14.2780275344849,-14.3255367279053,-14.4054679870605,-14.5069818496704,-14.5708494186401,-14.66015625,-14.7660150527954,-14.9357919692993,-15.0244626998901,-15.1083498001099,-15.26953125,-15.4268550872803,-15.5096673965454,-15.6671867370605,-16.0016117095947,-16.3087406158447,-16.5623054504395,-16.5877914428711,-16.6478519439697,-16.7454109191895,-16.7669429779053,-16.73388671875,-16.6395988464355,-16.6181144714355,-16.66748046875,-16.7421398162842,-16.791748046875,-16.7977542877197,-16.8805179595947,-16.9738273620605,-17.0793952941895,-17.1680660247803,-17.2606449127197,-17.3458023071289,-17.41845703125,-17.4450206756592,-17.53564453125,-17.4118175506592,-17.1471672058105,-16.8434085845947,-16.5707530975342,-16.5352554321289,-16.5020503997803,-16.4800777435303,-16.4410171508789,-16.4043445587158,-16.3581047058105,-16.3022956848145,-16.239013671875,-16.1683597564697,-16.11328125,-16.0740242004395,-15.958984375,-15.7682123184204,-15.6208009719849,-15.5166988372803,-15.3799810409546,-15.2105464935303,-15.1214361190796,-15.1126470565796,-15.0905752182007,-15.0552244186401,-15.0219240188599,-14.9906244277954,-14.9595212936401,-14.9286127090454,-14.7867193222046,-14.5337409973145,-14.3000984191895,-14.0856447219849,-13.9750490188599,-13.9681634902954,-13.9326171875,-13.8684577941895,-13.809814453125,-13.7566404342651,-13.7149410247803,-13.6846685409546,-13.6235342025757,-13.5555171966553,-13.5069828033447,-13.4981441497803,-13.4869632720947,-13.4541015625,-13.40966796875,-13.3475580215454,-13.2970218658447,-13.2580080032349,-13.2064447402954,-13.1423826217651,-13.1052732467651,-13.097900390625,-13.0792961120605,-13.0485353469849,-12.9943361282349,-12.9308595657349,-12.8626947402954,-12.8519039154053,-12.8626461029053,-12.8584966659546,-12.8131837844849,-12.7703123092651,-12.7352542877197,-12.6596193313599,-12.5435543060303,-12.4598636627197,-12.40869140625,-12.3025388717651,-12.2806148529053],[103.9697265625,103.819923400879,103.650199890137,103.705276489258,103.817970275879,103.908988952637,103.960838317871,103.996383666992,103.9697265625],[-26.2641105651855,-26.2598648071289,-26.4153308868408,-26.4510269165039,-26.4012203216553,-26.303466796875,-26.2793960571289,-26.2641105651855],[-37.1033172607422,-37.0060539245605,-36.9289054870605,-36.8489265441895,-36.80517578125,-36.7600593566895,-36.7038078308105,-36.6068840026855,-36.6474113464355,-36.541015625,-36.4486312866211,-36.40673828125,-36.3858375549316,-36.3264617919922,-36.2852554321289,-36.2356452941895,-36.172607421875,-36.1168975830078,-36.0731468200684,-36.0331077575684,-35.9646492004395,-35.8953094482422,-35.9215316772461,-35.9132804870605,-35.798583984375,-35.866943359375,-35.9389152526855,-36.0854988098145,-36.1236343383789,-36.251708984375,-36.3114738464355,-36.4457511901855,-36.4720687866211,-36.5065422058105,-36.6281242370605,-36.7349586486816,-36.8238754272461,-36.8517074584961,-36.885986328125,-37.0067367553711,-37.0828132629395,-37.1581039428711,-37.4976577758789,-37.6309051513672,-37.6922874450684,-37.6890144348145,-37.6188468933105,-37.9127922058105,-38.0174293518066,-37.9455070495605,-37.5358390808105,-37.3822250366211,-37.3687515258789,-37.2328109741211,-37.1033172607422],[-5.692138671875,-5.78251934051514,-5.77504920959473,-5.70786094665527,-5.66249990463257,-5.65971660614014,-5.692138671875],[-14.3643560409546,-14.3986806869507,-14.4086923599243,-14.4149417877197,-14.3985834121704,-14.3836421966553,-14.3604011535645,-14.328857421875,-14.3025388717651,-14.3167963027954,-14.3643560409546],[166.92919921875,166.8408203125,166.805953979492,166.747451782227,166.790924072266,166.85546875,166.875106811523,166.92919921875],[160.576278686523,160.506546020508,160.443069458008,160.39453125,160.355087280273,160.270217895508,160.149505615234,160.100006103516,160.087112426758,160.003524780273,159.979293823242,159.986328125,160,160.077346801758,160.44873046875,160.537109375,160.576278686523],[166.133209228516,166.053314208984,166.027923583984,165.968154907227,165.903991699219,165.856536865234,165.8193359375,165.791015625,165.790420532227,165.8359375,165.85986328125,165.9091796875,166.023818969727,166.125671386719,166.162109375,166.1298828125,166.133209228516],[161.71533203125,161.841110229492,161.914367675781,162.022857666016,162.105377197266,162.156829833984,162.287200927734,162.287979125977,162.373336791992,162.30126953125,162.201263427734,162.123641967773,162.042678833008,161.905868530273,161.786819458008,161.537887573242,161.539260864258,161.499130249023,161.487014770508,161.39794921875,161.2939453125,161.285552978516,161.304779052734,161.38232421875,161.475677490234,161.65380859375,161.697952270508,161.71533203125],[161.5478515625,161.558883666992,161.553802490234,161.477920532227,161.442474365234,161.409759521484,161.412017822266,161.4169921875,161.402252197266,161.364166259766,161.406829833984,161.5478515625],[159.750396728516,159.970611572266,160.065338134766,160.354598999023,160.525192260742,160.62548828125,160.681838989258,160.75146484375,160.794326782227,160.818954467773,160.801651000977,160.713073730469,160.649215698242,160.481643676758,160.321090698242,160.002349853516,159.853698730469,159.802734375,159.755477905273,159.680465698242,159.621871948242,159.6123046875,159.607421875,159.625595092773,159.686325073242,159.750396728516],[160.168167114258,160.225677490234,160.253524780273,160.3193359375,160.407531738281,160.371490478516,160.300003051758,160.275970458984,160.268157958984,160.253128051758,160.175186157227,160.105377197266,160.096282958984,160.168167114258],[159.188568115234,159.175094604492,159.128128051758,159.071090698242,159.036331176758,159.07763671875,159.129791259766,159.153701782227,159.176071166992,159.228408813477,159.233993530273,159.188568115234],[157.645416259766,157.643173217773,157.585830688477,157.457901000977,157.453506469727,157.5263671875,157.579299926758,157.623245239258,157.645416259766],[158.200790405273,158.178802490234,158.155364990234,158.2099609375,158.236328125,158.25341796875,158.200790405273],[158.10791015625,158.009475708008,157.937591552734,157.879302978516,157.8984375,157.909271240234,157.938278198242,157.966995239258,157.998428344727,158.10546875,158.132232666016,158.068359375,158.089645385742,158.103515625,158.10791015625],[157.388961791992,157.389053344727,157.333892822266,157.212310791016,157.233779907227,157.345108032227,157.379501342773,157.410949707031,157.383499145508,157.347061157227,157.332229614258,157.388961791992],[159.687698364258,159.640045166016,159.569244384766,159.538482666016,159.55322265625,159.594619750977,159.6416015625,159.646286010742,159.687698364258],[160.749420166016,160.997650146484,160.98779296875,160.9541015625,160.9443359375,160.9755859375,161.04345703125,161.15869140625,161.204696655273,161.208786010742,161.256637573242,161.258499145508,161.367965698242,161.377532958984,161.367385864258,161.321868896484,161.191009521484,161.04150390625,161.0244140625,160.873443603516,160.772064208984,160.66259765625,160.714065551758,160.590423583984,160.596282958984,160.648529052734,160.684768676758,160.7021484375,160.749420166016],[157.763473510742,157.826278686523,157.8984375,157.885452270508,157.833694458008,157.8193359375,157.749221801758,157.655960083008,157.587890625,157.564559936523,157.558013916016,157.504196166992,157.351364135742,157.302444458008,157.232421875,157.217575073242,157.228515625,157.321578979492,157.340621948242,157.433395385742,157.490631103516,157.598831176758,157.6123046875,157.651275634766,157.763473510742],[156.603897094727,156.591705322266,156.539642333984,156.542282104492,156.55126953125,156.5703125,156.612396240234,156.603897094727],[157.171875,157.150009155273,157.041213989258,156.958312988281,156.958984375,157.024124145508,157.102737426758,157.145797729492,157.186141967773,157.200592041016,157.191497802734,157.171875],[156.687881469727,156.668746948242,156.635345458984,156.611038208008,156.611709594727,156.510940551758,156.50244140625,156.560943603516,156.6396484375,156.717681884766,156.80908203125,156.790237426758,156.7080078125,156.687881469727],[159.879104614258,159.880859375,159.746490478516,159.64453125,159.354110717773,159.291702270508,159.2392578125,159.090225219727,158.944046020508,158.854583740234,158.831848144531,158.778030395508,158.686233520508,158.596984863281,158.5654296875,158.478805541992,158.457427978516,158.734375,158.86279296875,158.972457885742,159.010543823242,159.109375,159.198043823242,159.286819458008,159.36767578125,159.431457519531,159.843063354492,159.7939453125,159.8486328125,159.879104614258],[155.83984375,155.739349365234,155.677536010742,155.704986572266,155.738967895508,155.864639282227,155.83984375],[157.486724853516,157.518661499023,157.441314697266,157.339248657227,157.317291259766,157.314651489258,157.243453979492,157.1015625,156.904296875,156.69580078125,156.494918823242,156.457427978516,156.452545166016,156.479385375977,156.604202270508,156.765441894531,157.0302734375,157.1025390625,157.1484375,157.193359375,157.336135864258,157.41162109375,157.451553344727,157.486724853516],[-12.5260744094849,-12.5406255722046,-12.607177734375,-12.9516115188599,-12.8543949127197,-12.615234375,-12.544189453125,-12.5124998092651,-12.5006351470947,-12.5260744094849],[-10.283203125,-10.2857427597046,-10.3146486282349,-10.3598136901855,-10.3895511627197,-10.5167474746704,-10.5708494186401,-10.6175785064697,-10.6474609375,-10.6913089752197,-10.8780755996704,-11.000244140625,-11.0854005813599,-11.1661128997803,-11.2676763534546,-11.3766603469849,-11.4545412063599,-11.5075187683105,-11.5475101470947,-11.7334470748901,-11.9291982650757,-12.3466310501099,-12.4856443405151,-12.4806642532349,-12.4327154159546,-12.5104494094849,-12.4802732467651,-12.5104494094849,-12.5702152252197,-12.6976079940796,-12.7819337844849,-12.8508787155151,-12.8809576034546,-12.9251461029053,-12.9569339752197,-13.0208005905151,-13.1489744186401,-13.2017574310303,-13.2727537155151,-13.26123046875,-13.2033195495605,-13.157958984375,-13.0850095748901,-12.9942388534546,-12.9129400253296,-12.8940916061401,-12.9040040969849,-12.953369140625,-13.0882320404053,-13.1216306686401,-13.1818351745605,-13.2284183502197,-13.2261724472046,-13.2069339752197,-13.071044921875,-13.0594730377197,-13.1537103652954,-13.2716312408447,-13.2926759719849,-13.2342281341553,-13.1783695220947,-13.1298828125,-13.0772943496704,-13.0280275344849,-12.9986324310303,-12.9587888717651,-12.8311033248901,-12.755859375,-12.6844234466553,-12.6516599655151,-12.6221685409546,-12.6036128997803,-12.5898447036743,-12.557861328125,-12.5243654251099,-12.5014638900757,-12.427978515625,-12.2777347564697,-12.142333984375,-11.9227533340454,-11.9110841751099,-11.7100591659546,-11.471923828125,-11.2736330032349,-11.2056636810303,-11.1808595657349,-11.1156740188599,-11.0474615097046,-10.9630861282349,-10.8647947311401,-10.7585935592651,-10.6905279159546,-10.6827144622803,-10.6876459121704,-10.7212400436401,-10.7499513626099,-10.7470216751099,-10.7268552780151,-10.615966796875,-10.6057615280151,-10.6056156158447,-10.5517578125,-10.5005369186401,-10.5031251907349,-10.6284666061401,-10.6773433685303,-10.7021484375,-10.7121095657349,-10.6869630813599,-10.6526365280151,-10.60400390625,-10.5577154159546,-10.4964361190796,-10.3944339752197,-10.3600578308105,-10.283203125],[-89.3625946044922,-89.3372573852539,-89.1701126098633,-89.1205062866211,-89.05712890625,-89.02685546875,-89.0001907348633,-88.8683090209961,-88.845947265625,-88.7473678588867,-88.7076110839844,-88.6656188964844,-88.5831527709961,-88.5125503540039,-88.5043487548828,-88.4976577758789,-88.482666015625,-88.4491195678711,-88.4084930419922,-88.2762222290039,-88.1510238647461,-88.0804672241211,-88.0387191772461,-87.9910202026367,-87.8919906616211,-87.80224609375,-87.7314453125,-87.71533203125,-87.758544921875,-87.7742233276367,-87.7818832397461,-87.7564392089844,-87.7316436767578,-87.7370071411133,-87.814208984375,-87.83837890625,-87.8207015991211,-87.8780746459961,-87.9308547973633,-88.0234375,-88.1806640625,-88.4171371459961,-88.591552734375,-88.6856384277344,-88.6558609008789,-88.58154296875,-88.48388671875,-88.5120086669922,-88.8670349121094,-89.2776336669922,-89.5232391357422,-89.80419921875,-89.970458984375,-90.09521484375,-90.1059036254883,-90.104736328125,-90.0481414794922,-89.9426803588867,-89.8727111816406,-89.8399429321289,-89.793701171875,-89.7493591308594,-89.7111358642578,-89.6712875366211,-89.5702590942383,-89.5471649169922,-89.5550308227539,-89.5769577026367,-89.5736312866211,-89.54052734375,-89.5008773803711,-89.4188461303711,-89.3832473754883,-89.3625946044922],[12.3968753814697,12.4411134719849,12.5037107467651,12.5146484375,12.4852533340454,12.4263668060303,12.3968753814697],[48.9385757446289,48.9384765625,48.9384765625,48.9383773803711,48.9382820129395,48.9382820129395,48.9380874633789,48.9380874633789,48.7935523986816,48.6166000366211,48.4286117553711,48.2727546691895,48.1267547607422,47.9782218933105,47.6376953125,47.3056640625,46.9782218933105,46.9195289611816,46.6447296142578,46.2959938049316,45.86328125,45.5554695129395,45.2269515991211,44.8935546875,44.6320304870605,44.3062477111816,44.0228500366211,43.9837875366211,43.8267593383789,43.6205062866211,43.5810546875,43.4825210571289,43.3943367004395,43.3031272888184,43.2184562683105,43.181640625,43.0689468383789,43.0147438049316,42.9125022888184,42.8415985107422,42.81640625,42.78369140625,42.7251968383789,42.6692352294922,42.6564445495605,42.6595726013184,42.7630882263184,42.8097648620605,42.8628883361816,42.9061508178711,42.9227523803711,43.0486335754395,43.1593742370605,43.2459945678711,43.4412117004395,43.6311531066895,43.8527336120605,44.158203125,44.279296875,44.3865242004395,44.9429702758789,45.3376960754395,45.6958999633789,45.8166999816895,46.0245132446289,46.25390625,46.4602546691895,46.5650405883789,46.9734382629395,47.2300796508789,47.4049797058105,47.4738273620605,47.7125015258789,48.0192375183105,48.4388694763184,48.5725593566895,48.6744155883789,48.9031257629395,48.9385757446289],[41.53271484375,41.53759765625,41.5218734741211,41.4269523620605,41.2498054504395,41.1158180236816,40.9787101745605,40.9782218933105,40.9765625,40.9732437133789,40.9700202941895,40.9666976928711,40.9650382995605,40.9644508361816,40.9787101745605,41.1349601745605,41.341796875,41.6134757995605,41.7609367370605,41.8839836120605,41.9153289794922,42.0241203308105,42.2284164428711,42.3551750183105,42.7916030883789,42.8566398620605,42.8947257995605,42.9309577941895,43.0160140991211,43.1256828308105,43.333984375,43.5382843017578,43.58349609375,43.8291969299316,43.8894538879395,43.9888687133789,44.0281219482422,44.3695297241211,44.6366233825684,44.91162109375,44.9405288696289,45.1328125,45.4384765625,45.6335945129395,45.9349632263184,46.1667938232422,46.4229469299316,46.6717796325684,46.97119140625,47.1597633361816,47.4528312683105,47.7316398620605,47.9782218933105,48.1267547607422,48.2727546691895,48.4286117553711,48.6166000366211,48.7935523986816,48.9380874633789,48.9380874633789,48.9382820129395,48.9382820129395,48.9383773803711,48.9384765625,48.9384765625,48.9385757446289,49.0621109008789,49.3882827758789,49.64208984375,50.1100578308105,50.4662132263184,50.5283203125,50.6359405517578,50.7922859191895,51.1913070678711,51.2548828125,51.2318344116211,51.2181663513184,51.1363296508789,51.0842781066895,51.1222648620605,51.140625,51.1312484741211,51.1048812866211,51.0938491821289,51.05078125,51.0318374633789,51.0631828308105,51.1882820129395,51.185546875,51.1929664611816,51.2957038879395,51.369140625,51.3902359008789,51.3845710754395,51.2681655883789,51.2087898254395,51.0359382629395,50.9300765991211,50.8984375,50.8737335205078,50.8328132629395,50.8249969482422,50.6851577758789,50.6379890441895,50.4297866821289,50.3211898803711,50.2857398986816,50.1028327941895,49.85205078125,49.76123046875,49.6711921691895,49.5700187683105,49.3485374450684,49.2349624633789,49.0926742553711,49.04931640625,48.6490211486816,48.2339820861816,47.9752922058105,47.5114250183105,46.8788070678711,46.0511703491211,45.8262710571289,44.9202156066895,44.3327140808105,44.03271484375,43.7175788879395,43.4676742553711,42.7121086120605,42.6341819763184,42.5607414245605,42.4656257629395,42.3994140625,42.2189445495605,42.1062507629395,41.9798812866211,41.92626953125,41.8882827758789,41.84619140625,41.7322273254395,41.6320304870605,41.53271484375],[-56.1507339477539,-56.1716804504395,-56.2432594299316,-56.2091789245605,-56.18505859375,-56.1526374816895,-56.1373519897461,-56.1392593383789,-56.1507339477539],[-56.26708984375,-56.3542022705078,-56.384765625,-56.3772468566895,-56.3325691223145,-56.3339347839355,-56.3869132995605,-56.3779296875,-56.3646469116211,-56.287353515625,-56.2783699035645,-56.3148918151855,-56.289794921875,-56.2554702758789,-56.26708984375],[20.2417964935303,20.3013687133789,20.3585929870605,20.43798828125,20.5326175689697,20.5811519622803,20.6527328491211,20.7092761993408,20.7278327941895,20.7468757629395,20.7601566314697,20.7749996185303,20.7757816314697,20.779296875,20.7658214569092,20.7860355377197,20.7865238189697,20.7724609375,20.7742176055908,20.7940425872803,20.8708000183105,20.9417972564697,21.0238285064697,21.0999031066895,21.1478519439697,21.2264633178711,21.3817386627197,21.4314441680908,21.4654293060303,21.490234375,21.4917964935303,21.4678707122803,21.4344730377197,21.4207019805908,21.3958988189697,21.37109375,21.3529300689697,21.3570308685303,21.3777332305908,21.4099597930908,21.4719734191895,21.533203125,21.5323238372803,21.5199222564697,21.4421863555908,21.3843746185303,21.35791015625,21.3600597381592,21.5231456756592,21.5970706939697,21.6361312866211,21.740234375,21.9092788696289,22.0269527435303,22.0930671691895,22.2009773254395,22.3506832122803,22.4976558685303,22.64208984375,22.7208995819092,22.734375,22.7007808685303,22.6201171875,22.5540027618408,22.5023441314697,22.4945316314697,22.5306644439697,22.5818367004395,22.64794921875,22.6833000183105,22.6878910064697,22.705078125,22.66748046875,22.6265621185303,22.6034183502197,22.5974597930908,22.4690437316895,22.4208011627197,22.3990230560303,22.3654308319092,22.36962890625,22.3869152069092,22.3948249816895,22.4363269805908,22.47412109375,22.4991207122803,22.5545883178711,22.6969738006592,22.767578125,22.8197269439697,22.8595714569092,22.9768562316895,22.9679698944092,22.9422836303711,22.9152355194092,22.8568344116211,22.7999019622803,22.7061519622803,22.55810546875,22.5227527618408,22.466796875,22.4392585754395,22.4656238555908,22.4632816314697,22.4362297058105,22.4720706939697,22.5242195129395,22.5324211120605,22.5235366821289,22.4457015991211,22.4220695495605,22.3440437316895,22.3173828125,22.2770519256592,22.23974609375,22.1466808319092,22.0520515441895,21.9775390625,21.9041004180908,21.85302734375,21.8146495819092,21.7392578125,21.6182632446289,21.5625,21.5416030883789,21.5189437866211,21.5299797058105,21.60986328125,21.6190433502197,21.7306632995605,21.7521476745605,21.7529296875,21.7238273620605,21.6625003814697,21.390625,21.4030284881592,21.3231449127197,21.2371082305908,21.22265625,21.1270503997803,21.0570316314697,20.9676742553711,20.8907222747803,20.8444347381592,20.8238277435303,20.8238277435303,20.8005867004395,20.7633781433105,20.7005863189697,20.6231441497803,20.6096668243408,20.6375980377197,20.6576175689697,20.6485347747803,20.6240234375,20.47509765625,20.4583988189697,20.48681640625,20.4688472747803,20.3443374633789,20.34765625,20.3399410247803,20.2684574127197,20.1678695678711,19.9440422058105,19.8580074310303,19.7811508178711,19.6709957122803,19.6144542694092,19.5515632629395,19.4146480560303,19.2982425689697,19.21875,19.1964836120605,19.1916027069092,19.1943359375,19.2544918060303,19.30078125,19.3603515625,19.3996086120605,19.4512691497803,19.47998046875,19.4951171875,19.4881839752197,19.3640632629395,19.2572269439697,19.2450199127197,19.26806640625,19.3052730560303,19.34521484375,19.4494152069092,19.5495128631592,19.5836906433105,19.5837898254395,19.5471687316895,19.43017578125,19.3388671875,19.2315425872803,19.15185546875,19.1283206939697,19.1184577941895,19.1273441314697,19.1324214935303,19.1513671875,19.22314453125,19.2918949127197,19.33447265625,19.3568363189697,19.3486328125,19.3126945495605,19.23681640625,19.1315422058105,19.0420894622803,19.0071296691895,18.9955081939697,19.0095710754395,19.03759765625,19.060546875,19.0852546691895,19.1000003814697,19.0628910064697,19.1296882629395,19.1307621002197,19.1369152069092,19.2059574127197,19.3030261993408,19.3880863189697,19.4009761810303,19.3999996185303,19.3823223114014,19.3522472381592,19.3302745819092,19.2728519439697,19.0930671691895,19.0046863555908,19.0076179504395,19.0333003997803,19.0642585754395,19.0550785064697,18.9537105560303,18.9178714752197,18.947265625,18.89453125,18.8390617370605,18.8935546875,18.9010753631592,18.9053707122803,18.9278316497803,19.0157222747803,19.0476551055908,19.0662117004395,19.0873050689697,19.1462898254395,19.2083988189697,19.2781257629395,19.3302745819092,19.3928699493408,19.4212894439697,19.45751953125,19.53076171875,19.6134757995605,19.7245121002197,19.8444328308105,19.93408203125,20.1614265441895,20.2101554870605,20.2417964935303],[6.65996122360229,6.55683612823486,6.51972627639771,6.49697256088257,6.46816396713257,6.47753858566284,6.52431631088257,6.62587881088257,6.68691444396973,6.74980497360229,6.74999952316284,6.65996122360229],[7.423828125,7.38662099838257,7.34238338470459,7.33066368103027,7.38759756088257,7.41445302963257,7.43701124191284,7.45039081573486,7.45234441757202,7.423828125],[-54.1559600830078,-54.2401847839355,-54.3316421508789,-54.4521980285645,-54.4468765258789,-54.4733390808105,-54.4796867370605,-54.4711456298828,-54.4402351379395,-54.4260749816895,-54.4496078491211,-54.440673828125,-54.416015625,-54.396240234375,-54.3983879089355,-54.369140625,-54.3421401977539,-54.3507308959961,-54.2555198669434,-54.1974639892578,-54.11279296875,-54.0819816589355,-54.0342292785645,-54.0059089660645,-53.990478515625,-54.0059585571289,-54.0095710754395,-54.0631828308105,-54.1880378723145,-54.203125,-54.1707000732422,-54.1880836486816,-54.1955070495605,-54.2567367553711,-54.4020042419434,-54.4855461120605,-54.5359382629395,-54.5684089660645,-54.604736328125,-54.6162605285645,-54.661865234375,-54.6974105834961,-54.7029304504395,-54.7222137451172,-54.766845703125,-54.8516616821289,-54.8760757446289,-54.9265594482422,-54.9684104919434,-54.9786605834961,-55.0058135986328,-55.0703125,-55.1141128540039,-55.1488304138184,-55.1876945495605,-55.2860336303711,-55.343994140625,-55.3853530883789,-55.658935546875,-55.7305679321289,-55.8937492370605,-55.9359359741211,-55.9574699401855,-55.9755859375,-55.9935035705566,-56.0203628540039,-56.0451164245605,-56.0877914428711,-56.12939453125,-56.1376953125,-56.0736351013184,-56.0200691223145,-55.9619636535645,-55.9153327941895,-55.921630859375,-55.9296379089355,-55.9633293151855,-56.0199203491211,-56.2271499633789,-56.3858413696289,-56.4528312683105,-56.4828109741211,-56.5223617553711,-56.5626945495605,-56.627197265625,-56.704345703125,-56.7611312866211,-56.81982421875,-56.8405265808105,-56.8864250183105,-56.9314956665039,-56.9452133178711,-56.9792976379395,-56.9971199035645,-57.0234870910645,-57.0289535522461,-57.0419425964355,-57.0604476928711,-57.0968742370605,-57.1051292419434,-57.1211433410645,-57.1636238098145,-57.1973648071289,-57.2098159790039,-57.2069320678711,-57.2249984741211,-57.2305679321289,-57.2316398620605,-57.2489738464355,-57.2779273986816,-57.2829132080078,-57.2899436950684,-57.3036613464355,-57.4255867004395,-57.4378890991211,-57.4905776977539,-57.5496101379395,-57.6027336120605,-57.646728515625,-57.6561050415039,-57.6494598388672,-57.7203636169434,-57.8326683044434,-57.8665504455566,-57.90771484375,-58.0322227478027,-58.0542984008789,-58.0544891357422,-58.0107421875,-57.9497566223145,-57.9247093200684,-57.90625,-57.8747062683105,-57.8460006713867,-57.8678703308105,-57.9048805236816,-57.9170379638672,-57.8811073303223,-57.8449211120605,-57.8041038513184,-57.7520027160645,-57.7110824584961,-57.6488265991211,-57.5708999633789,-57.4121589660645,-57.3310012817383,-57.3057594299316,-57.3095703125,-57.269287109375,-57.2268562316895,-57.2098159790039,-57.2073249816895,-57.2184600830078,-57.2353019714355,-57.2796401977539,-57.3185501098633,-57.2918930053711,-57.2575187683105,-57.2478981018066,-57.1947746276855,-57.18212890625,-57.1408195495605,-57.1360359191895,-57.1045875549316,-57.056640625,-56.9698257446289,-56.4660186767578,-56.235595703125,-55.9395484924316,-55.8976058959961,-55.8955078125,-55.909912109375,-55.8281745910645,-55.6483421325684,-55.3792953491211,-55.1482925415039,-54.8336906433105,-54.3561515808105,-54.142333984375,-54.05419921875,-54.0374031066895,-54.0459480285645,-54.0804672241211,-54.1559600830078],[22.5386714935303,22.5241222381592,22.4832038879395,22.4320316314697,22.3894519805908,22.3326168060303,22.2952136993408,22.1428699493408,22.1318359375,22.111328125,21.7669906616211,21.7214832305908,21.6746101379395,21.6486320495605,21.6325187683105,21.6026363372803,21.5631847381592,21.5046863555908,21.4513664245605,21.3824214935303,21.1963882446289,21.0672855377197,20.9811534881592,20.8666019439697,20.6431636810303,20.4900398254395,20.4750003814697,20.3337898254395,20.1286144256592,19.9503898620605,19.8986320495605,19.81005859375,19.7091808319092,19.6253910064697,19.5642566680908,19.4974613189697,19.4669914245605,19.26513671875,18.9141597747803,18.7918949127197,18.7500972747803,18.7483406066895,18.7780265808105,18.7406253814697,18.7242183685303,18.4762687683105,18.1456050872803,17.9479484558105,17.7619132995605,17.63525390625,17.4806632995605,17.3172855377197,17.3015632629395,17.2772445678711,17.1746101379395,17.1473636627197,17.0859394073486,17.06787109375,16.97265625,16.8654289245605,16.8626956939697,16.9044933319092,16.943359375,16.9488277435303,16.953125,16.9852523803711,17.0632801055908,17.1356449127197,17.1884765625,17.296875,17.4826164245605,17.6253910064697,17.7584972381592,17.8308601379395,17.8926753997803,17.9132804870605,17.9407234191895,18.0508785247803,18.0859375,18.1003913879395,18.1099605560303,18.1326179504395,18.1609363555908,18.3648433685303,18.3831062316895,18.4158210754395,18.47607421875,18.5345706939697,18.5964851379395,18.6761722564697,18.7497081756592,18.8070316314697,18.8322257995605,18.9381828308105,18.9572257995605,18.9683589935303,19.1494140625,19.2501945495605,19.3023433685303,19.38623046875,19.4416027069092,19.4796867370605,19.5347671508789,19.5930671691895,19.6266593933105,19.6302719116211,19.6641597747803,19.7300777435303,19.77392578125,19.7870121002197,19.7879886627197,19.7673816680908,19.7566413879395,19.80224609375,19.8689460754395,19.9161128997803,20.0576171875,20.1076183319092,20.1636714935303,20.2365245819092,20.3025398254395,20.3629875183105,20.4046878814697,20.4226570129395,20.4745121002197,20.5345706939697,20.6161136627197,20.72900390625,20.7995128631592,20.8684577941895,20.947265625,21.0011711120605,21.0793952941895,21.1361331939697,21.2249984741211,21.3504886627197,21.6396484375,21.7121086120605,21.89013671875,21.9676761627197,22.0021476745605,22.0201168060303,22.2025394439697,22.4730472564697,22.5386714935303],[16.5162105560303,16.4276371002197,16.3211917877197,16.3011722564697,16.2583980560303,16.2367191314697,16.2533187866211,16.2274417877197,16.1064453125,16.0665035247803,16.0006828308105,15.9333009719849,15.8475589752197,15.7842769622803,15.7041997909546,15.6359367370605,15.6089849472046,15.5925788879395,15.5968751907349,15.6662101745605,15.6755867004395,15.6680660247803,15.6521482467651,15.6248054504395,15.4541025161743,15.2770519256592,15.2729482650757,15.3537101745605,15.3569345474243,15.2901363372803,15.2835931777954,15.2912111282349,15.3266592025757,15.3394527435303,15.2420892715454,15.1104497909546,14.95458984375,14.8999996185303,14.8470706939697,14.7930660247803,14.7335939407349,14.6495113372803,14.6085929870605,14.5917978286743,14.5688486099243,14.5339851379395,14.5051755905151,14.4273433685303,14.3699226379395,14.2830076217651,14.1612310409546,14.0855464935303,13.9927740097046,13.9703130722046,13.9701175689697,13.9356441497803,13.8787107467651,13.615234375,13.5779294967651,13.6373052597046,13.7198247909546,13.7759771347046,13.8447256088257,13.8747072219849,13.8311529159546,13.7216796875,13.6634769439697,13.5833988189697,13.5696287155151,13.6139650344849,13.6005859375,13.5091791152954,13.4876947402954,13.4802742004395,13.4864253997803,13.5480470657349,13.6166019439697,13.6349601745605,13.6325197219849,13.5447263717651,13.4917974472046,13.4498052597046,13.4209966659546,13.3996095657349,13.3782224655151,13.3995122909546,13.478515625,13.5632810592651,13.6371097564697,13.6796875,13.6999998092651,13.7439460754395,13.8313484191895,13.9288082122803,14.0196285247803,14.0995121002197,14.2672853469849,14.4199209213257,14.4659185409546,14.5035152435303,14.5498056411743,14.5771484375,14.5969734191895,14.6801748275757,14.7567377090454,14.8105459213257,14.8406248092651,14.8932619094849,14.9494142532349,15.0006837844849,15.2169923782349,15.4392585754395,15.5453128814697,15.6326179504395,15.7602529525757,15.7668943405151,15.9576168060303,15.9722661972046,15.98046875,15.9768562316895,16.0372066497803,16.0930671691895,16.2835922241211,16.3084964752197,16.3184566497803,16.33544921875,16.3671875,16.3845710754395,16.3812503814697,16.41845703125,16.5056648254395,16.5162105560303],[16.5285167694092,16.4771480560303,16.431640625,16.4012699127197,16.3941402435303,16.4123039245605,16.63037109375,16.7277336120605,16.8646488189697,16.9015617370605,16.9609375,16.9959964752197,17.025390625,17.0892581939697,17.11767578125,17.0503902435303,17.0582027435303,17.0535163879395,16.8836917877197,16.8382816314697,16.7780265808105,16.5285167694092],[19.0764636993408,18.9937496185303,18.9451179504395,18.8781261444092,18.8138675689697,18.7909183502197,18.9079093933105,18.8436527252197,18.7848625183105,18.7428703308105,18.6999015808105,18.5384750366211,18.4773445129395,18.38720703125,18.3402347564697,18.2489242553711,18.1463871002197,18.20654296875,18.2853527069092,18.2095718383789,18.1639652252197,18.1050777435303,18.1519527435303,18.12890625,18.1365242004395,18.2048816680908,18.283203125,18.4051761627197,18.5374031066895,18.7218742370605,18.80517578125,18.8411121368408,18.9005870819092,18.9564456939697,19.0764636993408],[19.1563472747803,19.1383800506592,19.0865230560303,19.0392589569092,19.1348628997803,19.2811527252197,19.3314456939697,19.1563472747803],[18.4162120819092,18.3718757629395,18.3499031066895,18.3772449493408,18.3975582122803,18.4649410247803,18.4855461120605,18.4162120819092],[18.5954093933105,18.5703125,18.5451183319092,18.55517578125,18.5723628997803,18.6208992004395,18.6984367370605,18.6979503631592,18.6238269805908,18.5954093933105],[24.1554698944092,23.8905258178711,23.69140625,23.5920906066895,23.4183597564697,23.2210941314697,23.1545906066895,23.1023426055908,22.9193344116211,22.74658203125,22.6203136444092,22.53857421875,22.4651355743408,22.4009761810303,22.3663082122803,22.3359375,22.28759765625,22.2749996185303,22.2666015625,22.2540054321289,22.0862312316895,22.0962886810303,22.1328125,22.1475601196289,22.0867176055908,21.9201183319092,21.9031257629395,21.9499988555908,21.9134769439697,21.8795890808105,21.6806640625,21.5655288696289,21.5326175689697,21.5234375,21.5452156066895,21.5959968566895,21.6126956939697,21.6091804504395,21.56689453125,21.4468746185303,21.4103507995605,21.4377937316895,21.50634765625,21.5459957122803,21.5806636810303,21.5739269256592,21.4249038696289,21.2937507629395,21.1958980560303,21.13818359375,21.2049808502197,21.279296875,21.33154296875,21.3938484191895,21.5196285247803,21.4943351745605,21.4650382995605,21.2557621002197,21.0184574127197,20.7626934051514,20.6776351928711,20.4537105560303,20.3713855743408,20.2046890258789,19.9136734008789,19.7816390991211,19.7220706939697,19.65576171875,19.5900402069092,19.5023441314697,19.4909172058105,19.49462890625,19.3542976379395,19.2880859375,19.236328125,19.0343742370605,18.8166999816895,18.7922859191895,18.8501949310303,18.8589839935303,18.8194332122803,18.7595710754395,18.6671867370605,18.6064453125,18.57763671875,18.5306644439697,18.40771484375,18.34423828125,18.3128910064697,18.5020503997803,18.4869136810303,18.4826164245605,18.4630851745605,18.248046875,18.2149410247803,18.1700191497803,18.0744132995605,18.0779304504395,18.0935554504395,17.9510746002197,17.9066410064697,17.8795890808105,17.8956069946289,17.9329109191895,17.9744148254395,17.9407234191895,17.9030265808105,17.9304695129395,18.0065422058105,18.0373039245605,17.9470691680908,17.83447265625,17.7177734375,17.6463871002197,17.5706062316895,17.5089855194092,17.4102535247803,17.37841796875,17.3733386993408,17.4290027618408,17.5352535247803,17.6336898803711,17.5628910064697,17.5101566314697,17.4465827941895,17.4120121002197,17.37451171875,17.3982429504395,17.4172840118408,17.4654293060303,17.3345718383789,17.1963863372803,17.2156257629395,17.1307621002197,17.1465816497803,17.1642570495605,17.1379871368408,17.1779289245605,17.1996097564697,17.1638679504395,17.1797847747803,17.1857433319092,17.212890625,17.2029285430908,17.2789058685303,17.26123046875,17.2509765625,17.35986328125,17.45703125,17.5554676055908,17.5930671691895,17.6307621002197,17.6611328125,17.7421875,17.87158203125,17.9557609558105,18.0113277435303,18.1625003814697,18.25048828125,18.3999996185303,18.5575199127197,18.5354499816895,18.6011714935303,18.7870121002197,18.8527336120605,18.88427734375,18.9332027435303,18.9904289245605,18.9705066680908,18.8956050872803,18.71875,18.6399421691895,18.578125,18.4024429321289,18.3380870819092,18.2764644622803,18.2168941497803,18.16357421875,17.9642581939697,17.9797840118408,18.1326179504395,18.2105464935303,18.2705078125,18.3360347747803,18.3958015441895,18.4591789245605,18.5088863372803,18.5602531433105,18.6175785064697,18.4986324310303,18.4142589569092,18.373046875,18.3219718933105,18.2853527069092,18.09814453125,17.974609375,17.8290023803711,17.7654304504395,17.6696300506592,17.4567375183105,17.34765625,17.1028308868408,16.9781265258789,16.6393547058105,16.3158187866211,16.2142581939697,16.3180656433105,16.3908195495605,16.47802734375,16.6830081939697,16.7884769439697,16.923828125,16.8243160247803,16.6519527435303,16.7166004180908,16.7699203491211,16.7000980377197,16.6949214935303,16.5969715118408,16.5553703308105,16.5862293243408,16.5837898254395,16.6042003631592,16.6522445678711,16.6308574676514,16.4759769439697,16.4794921875,16.50732421875,16.5279293060303,16.45751953125,16.4078121185303,16.3487300872803,16.2165050506592,16.1506843566895,15.9966802597046,15.9203119277954,15.82666015625,15.7222652435303,15.6265630722046,15.5096673965454,15.3265619277954,15.0511722564697,14.7820320129395,14.7139644622803,14.7547845840454,14.6555662155151,14.5585947036743,14.4732427597046,14.4019527435303,14.2619142532349,14.2150382995605,14.2029304504395,14.2764644622803,14.3416996002197,14.1737298965454,14.0799808502197,13.8063468933105,13.3213863372803,12.8858404159546,12.9406251907349,12.9387693405151,12.96337890625,12.97802734375,12.9739255905151,12.9419927597046,12.8345699310303,12.5925779342651,12.52099609375,12.47119140625,12.5070304870605,12.7063474655151,12.7528324127197,12.8016605377197,12.7421875,12.6911134719849,12.6564455032349,12.7731447219849,12.857421875,12.9195318222046,12.8836908340454,12.7931632995605,12.7175779342651,12.5726566314697,12.4214849472046,12.15185546875,12.05322265625,11.9615230560303,11.9169921875,11.8850584030151,11.8787117004395,11.7349605560303,11.7291011810303,11.7032222747803,11.5490236282349,11.4493160247803,11.4315423965454,11.3299798965454,11.2482423782349,11.2520503997803,11.2715816497803,11.2238283157349,11.2079105377197,11.1691408157349,11.1471672058105,11.1668949127197,11.19580078125,11.2953128814697,11.3882808685303,11.470703125,11.5435552597046,11.6427736282349,11.7122068405151,11.7518548965454,11.7981443405151,11.7433595657349,11.6848630905151,11.6807613372803,11.8342771530151,11.8812494277954,11.93212890625,11.98828125,12.0718755722046,12.1692390441895,12.2919921875,12.4020509719849,12.4861326217651,12.5146484375,12.5158205032349,12.5528326034546,12.5886716842651,12.5538082122803,12.4453125,12.3146486282349,12.2941408157349,12.3537111282349,12.4675779342651,12.6830081939697,12.7060546875,12.7278318405151,12.7763662338257,12.8282222747803,12.8636713027954,12.8807621002197,12.7575197219849,12.5960941314697,12.48681640625,12.2920904159546,12.1553707122803,12.2336912155151,12.2919921875,12.3013668060303,12.3035163879395,12.1145515441895,12.1218748092651,12.1398429870605,12.1196298599243,12.1085939407349,12.1410160064697,12.2181644439697,12.1446285247803,12.138671875,11.9999027252197,12.2121095657349,12.1751956939697,12.3019533157349,12.53271484375,12.6625003814697,12.6900386810303,12.7927732467651,12.9875974655151,13.2035150527954,13.2996091842651,13.6707029342651,13.9605474472046,14.0027351379395,14.0632810592651,14.1412105560303,14.1480474472046,14.1199216842651,14.07763671875,13.87353515625,13.6502933502197,13.9248046875,14.1151361465454,14.3524417877197,14.42626953125,14.4796867370605,14.5495119094849,14.5958003997803,14.6345701217651,14.6351556777954,14.6099605560303,14.5432624816895,14.9179677963257,15.0400390625,15.1533193588257,15.3749027252197,15.4837894439697,15.4229497909546,15.5570316314697,15.8841791152954,16.2376956939697,16.4035148620605,16.4207038879395,16.4342765808105,16.3606433868408,16.2815437316895,16.12744140625,16.1935558319092,16.30712890625,16.4571285247803,16.5741214752197,16.5855464935303,16.7835941314697,17.1705074310303,17.3246097564697,17.5647468566895,17.9167003631592,18.0732421875,18.125,18.1766605377197,18.1559581756592,18.1470699310303,18.16259765625,18.3030261993408,18.3786125183105,18.7698249816895,18.8682613372803,19.0526371002197,19.2589836120605,19.6912097930908,19.8700199127197,19.9698238372803,20.0559577941895,20.2400398254395,19.9688472747803,20.1474609375,20.2400398254395,20.3194332122803,20.3480472564697,20.3371105194092,20.2823238372803,20.11669921875,20.4919910430908,20.6221675872803,20.8951168060303,20.9070320129395,20.9089851379395,20.9185543060303,21.1833972930908,21.259765625,21.42236328125,21.4654293060303,21.6160144805908,21.7240219116211,21.8501949310303,21.9974613189697,22.1951160430908,22.3621101379395,22.7824230194092,22.8541011810303,22.9753894805908,23.0978527069092,23.1825199127197,23.3185539245605,23.35546875,23.4742202758789,23.6388664245605,23.6329097747803,23.5018558502197,23.48779296875,23.5001945495605,23.5413093566895,23.5370121002197,23.5044918060303,23.4654293060303,23.4514656066895,23.4548835754395,23.4680671691895,23.537109375,23.6608390808105,23.7335948944092,23.77490234375,23.7609367370605,23.6566410064697,23.6260738372803,23.623046875,23.6415042877197,23.6773433685303,23.7589836120605,23.8693351745605,23.9417972564697,23.97607421875,23.9885730743408,23.9388675689697,23.8941421508789,23.8858394622803,23.8655281066895,23.7683601379395,23.701171875,23.6820316314697,23.673828125,23.6935539245605,23.7002944946289,23.7209968566895,23.75146484375,23.9073238372803,23.99462890625,24.0490226745605,24.1554698944092],[31.9482440948486,31.9684581756592,32.060546875,32.06884765625,32.0599632263184,32.0414085388184,32.04833984375,32.0779304504395,32.10595703125,32.1128883361816,32.0816421508789,32.0248031616211,31.9947261810303,31.9671878814697,31.9460926055908,31.9583969116211,31.7425765991211,31.4695301055908,31.2740230560303,31.0633792877197,30.9380855560303,30.88330078125,30.8067378997803,30.7943344116211,30.7875003814697,30.7890625,30.8033199310303,30.9452133178711,31.0333023071289,31.0880870819092,31.2073249816895,31.3351554870605,31.3826198577881,31.4151382446289,31.6404285430908,31.8714828491211,31.9216785430908,31.9482440948486],[-63.1230430603027,-63.0111808776855,-63.0123062133789,-63.0230445861816,-63.0904312133789,-63.1247024536133,-63.1230430603027],[55.5403327941895,55.54296875,55.4947280883789,55.4812507629395,55.4167976379395,55.3833999633789,55.4557609558105,55.5403327941895],[42.3589859008789,42.3590812683105,42.35009765625,42.2373046875,42.083984375,41.9740219116211,41.78857421875,41.6501960754395,41.4167938232422,41.3541984558105,41.2959976196289,41.2618141174316,41.2517585754395,41.24560546875,41.3001937866211,41.3526344299316,41.359375,41.3540992736816,41.3033218383789,41.2483406066895,41.2164039611816,41.1996078491211,41.19921875,41.1947250366211,41.0990257263184,40.9870109558105,40.93505859375,40.689453125,40.4214820861816,40.1219711303711,39.8500022888184,39.564453125,39.2683601379395,39.0567398071289,38.7735366821289,38.515625,38.2542991638184,38.0557632446289,37.7541046142578,37.5774383544922,37.3175773620605,37.0889663696289,36.818359375,36.4791984558105,36.3720703125,36.2842750549316,36.2197265625,36.0594711303711,35.9564437866211,35.8947257995605,35.7873039245605,35.8014640808105,35.8568382263184,35.9134750366211,35.8820304870605,35.8717765808105,35.8680648803711,35.9066390991211,35.8884773254395,35.8587913513184,35.8370132446289,35.8371086120605,35.8515625,35.869140625,35.9147453308105,35.9265632629395,35.9675788879395,36.0222625732422,36.0344734191895,36.0266609191895,35.9716796875,35.9423828125,35.9684562683105,35.9861335754395,36.0188484191895,36.0921897888184,36.1498031616211,36.1994132995605,36.2833976745605,36.3485336303711,36.3650398254395,36.36279296875,36.2822265625,36.27783203125,36.2978515625,36.3548851013184,36.4228515625,36.45751953125,36.53515625,36.5849609375,36.50439453125,36.45556640625,36.37646484375,36.3298835754395,36.3262672424316,36.388671875,36.4330062866211,36.3838882446289,36.2962875366211,36.2635726928711,36.1510734558105,35.9762687683105,35.8993186950684,35.8878898620605,35.8899421691895,35.9430694580078,35.9180679321289,35.916015625,35.9024429321289,35.7644500732422,35.8396492004395,35.8926734924316,35.9675788879395,36.1273422241211,36.1536102294922,36.2019538879395,36.2488288879395,36.3475570678711,36.3753929138184,36.4214859008789,36.47705078125,36.5624008178711,36.63671875,36.6414070129395,36.5374984741211,36.5466804504395,36.5968742370605,36.62841796875,36.6585960388184,36.7765617370605,36.9417991638184,36.9853515625,37.0662117004395,37.1874008178711,37.3270530700684,37.4363288879395,37.5235366821289,37.7203140258789,37.8179702758789,37.9066429138184,38.1916999816895,38.3058586120605,38.3839836120605,38.4437522888184,38.5780296325684,38.6888694763184,38.7666015625,38.9064445495605,39.1083984375,39.3566398620605,39.50146484375,39.6865234375,40.0164031982422,40.4503898620605,40.7056617736816,40.8156242370605,40.9588851928711,41.1021499633789,41.2646484375,41.3395500183105,41.5155258178711,41.7435531616211,41.8868179321289,42.0598640441895,42.1678695678711,42.2027359008789,42.24755859375,42.2685546875,42.3128929138184,42.3589859008789],[-72.3328094482422,-72.2186508178711,-72.1498031616211,-72.1443405151367,-72.1815414428711,-72.190673828125,-72.3008804321289,-72.33544921875,-72.3423843383789,-72.3328094482422],[-71.6614227294922,-71.6653823852539,-71.7217712402344,-71.8304214477539,-71.84765625,-71.80615234375,-71.6683654785156,-71.6369171142578,-71.6614227294922],[-71.8799362182617,-71.8974609375,-71.9554672241211,-71.9637680053711,-71.9845275878906,-72.01904296875,-72.0106430053711,-71.9315414428711,-71.8996124267578,-71.8799362182617],[23.9802742004395,23.9806652069092,23.9810543060303,23.9814453125,23.9818363189697,23.9822273254395,23.9825191497803,23.98291015625,23.9833011627197,23.9833984375,23.9708023071289,23.9652347564697,23.9459953308105,23.7082023620605,23.60400390625,23.4580078125,23.2434558868408,23.1051750183105,23.0091800689697,22.9338874816895,22.9695320129395,22.9613285064697,22.9323234558105,22.8671875,22.8021488189697,22.7632827758789,22.7149410247803,22.67919921875,22.6824207305908,22.6708984375,22.6318359375,22.5320301055908,22.4677734375,22.4162120819092,22.3815422058105,22.3997077941895,22.4249992370605,22.4393558502197,22.4493179321289,22.4983406066895,22.5282230377197,22.53857421875,22.5099601745605,22.38818359375,22.33935546875,22.2834949493408,22.2621097564697,22.1731452941895,22.1282234191895,22.1064453125,22.1076164245605,22.1529293060303,22.2023448944092,22.2213878631592,22.2326183319092,22.2281246185303,22.20263671875,22.1580085754395,21.990234375,21.90771484375,21.841796875,21.8252925872803,21.8433589935303,21.8781261444092,21.9281234741211,22.0006828308105,22.1211910247803,22.2333984375,22.3523445129395,22.4144515991211,22.3902339935303,22.4352531433105,22.4754886627197,22.4724617004395,22.4898433685303,22.5643558502197,22.5809574127197,22.5563488006592,22.5911140441895,22.6410140991211,22.6973648071289,22.7540035247803,22.7833995819092,22.8490238189697,22.9226570129395,22.9427738189697,22.9376964569092,22.8948230743408,22.8600597381592,22.8173828125,22.7301769256592,22.6240234375,22.4938488006592,22.3698234558105,22.2359371185303,22.1936531066895,22.15625,22.09716796875,22.0431632995605,22.0137691497803,21.96484375,21.771484375,21.7306632995605,21.70654296875,21.70654296875,21.7261714935303,21.7257823944092,21.6827144622803,21.6327152252197,21.5757808685303,21.5280265808105,21.4968738555908,21.39599609375,21.3524417877197,21.2638664245605,21.0094718933105,20.9841804504395,20.8910160064697,20.7732410430908,20.6681652069092,20.65966796875,20.6314449310303,20.56689453125,20.3420886993408,20.0726547241211,19.9535160064697,19.8376960754395,19.6683597564697,19.6174812316895,19.4002933502197,19.1455078125,19.0478515625,18.9562511444092,18.8882827758789,18.8783206939697,18.8885746002197,18.8860340118408,19.0641593933105,19.1086921691895,19.0638675689697,19.0423831939697,19.0398445129395,19.0108394622803,18.9064445495605,18.7474613189697,18.6662101745605,18.6335945129395,18.5916004180908,18.5641613006592,18.455078125,18.2388668060303,17.9401378631592,17.7608394622803,17.6494140625,17.49267578125,17.4364261627197,17.4021492004395,17.2469730377197,17.1179695129395,17.0719718933105,16.8903312683105,16.8181648254395,16.7847652435303,16.6683597564697,16.5889644622803,16.5501956939697,16.5453128814697,16.5232410430908,16.4593753814697,16.4043941497803,16.37890625,16.1911144256592,16.0306644439697,15.9576168060303,15.8450193405151,15.7012691497803,15.5892581939697,15.4800777435303,15.5324220657349,15.5526361465454,15.5578117370605,15.5498046875,15.4844722747803,15.4429693222046,15.3490228652954,15.2523431777954,15.1162109375,14.9679689407349,14.8607416152954,14.8262691497803,14.7712898254395,14.7328119277954,14.5361328125,14.3323240280151,14.2800779342651,14.1779289245605,14.0641603469849,14.0049800872803,13.9772462844849,14.0559568405151,14.1397457122803,14.2432613372803,14.3772459030151,14.5979499816895,14.8358402252197,15.0715827941895,15.1327142715454,15.1931648254395,15.3200187683105,15.540919303894,15.6548824310303,15.5319337844849,15.3999032974243,15.2760753631592,15.2009773254395,15.1322259902954,15.0686521530151,15.0298824310303,15.0357427597046,15.0554685592651,15.1219720840454,15.0780267715454,15.0876951217651,15.0812501907349,15.0598640441895,14.9738292694092,14.9567384719849,14.8806629180908,14.8470706939697,14.76123046875,14.6232423782349,14.5447273254395,14.5162115097046,14.4617185592651,14.2448244094849,14.06396484375,13.9323244094849,13.7634763717651,13.6063480377197,13.5057621002197,13.4482421875,13.513671875,13.6423835754395,13.80712890625,14.17822265625,14.3679685592651,14.7466793060303,15.2121086120605,15.4743156433105,15.5166988372803,15.5615243911743,15.5955076217651,15.637598991394,15.6729497909546,15.6986331939697,15.7350587844849,15.7662115097046,15.9488277435303,15.9631834030151,15.9292974472046,15.66845703125,15.5871095657349,15.5403318405151,15.6073236465454,15.2936525344849,15.2158203125,15.1818351745605,15.1778316497803,15.1722660064697,15.0889654159546,14.9790048599243,15.3474607467651,15.6271486282349,15.9840822219849,16.3150386810303,16.7941417694092,17.2732429504395,17.7522449493408,18.2313480377197,18.71044921875,19.189453125,19.6685543060303,20.1476554870605,20.6267585754395,21.1058597564697,21.5849609375,22.0640621185303,22.5430660247803,23.0221691131592,23.5012683868408,23.9802742004395],[0.900488495826721,0.874804556369781,0.821874737739563,0.787500083446503,0.763379216194153,0.779980599880219,0.792187511920929,0.958300530910492,1.17617177963257,1.33007800579071,1.34287095069885,1.34511697292328,1.34707021713257,1.37890601158142,1.38574194908142,1.42431628704071,1.56630849838257,1.60019552707672,1.60380852222443,1.60664069652557,1.62460923194885,1.62460923194885,1.62460923194885,1.62470722198486,1.62470722198486,1.53095698356628,1.58203101158142,1.59082019329071,1.60292959213257,1.57753896713257,1.59853506088257,1.63925802707672,1.74316394329071,1.77792978286743,1.61093723773956,1.62265598773956,1.31064438819885,1.18720698356628,1.18505835533142,1.13964855670929,1.08447277545929,1.04990220069885,1.00214838981628,0.984960854053497,0.912207305431366,0.822460830211639,0.736914217472076,0.707226455211639,0.715429484844208,0.702246248722076,0.672753870487213,0.595703065395355,0.548046767711639,0.525586187839508,0.533398509025574,0.523046851158142,0.538085877895355,0.579492330551147,0.592480182647705,0.59619128704071,0.619531214237213,0.634765625,0.591015696525574,0.537304937839508,0.509570300579071,0.498925894498825,0.500000238418579,0.605175793170929,0.583593606948853,0.599218726158142,0.647070467472076,0.688086211681366,0.68632835149765,0.616210997104645,0.483300775289536,0.415331959724426,0.378613352775574,0.372558355331421,0.453124821186066,0.488769680261612,0.493261635303497,0.460351556539536,0.466113179922104,0.497168093919754,0.529003918170929,0.525683641433716,0.447558522224426,0.405273675918579,0.370995879173279,0.28935569524765,0.259960681200027,0.241503983736038,0.233398362994194,0.261913895606995,0.251562267541885,0.275488436222076,0.327343970537186,0.342578321695328,0.272753715515137,0.264550656080246,0.26953136920929,0.289648503065109,0.311718851327896,0.323925703763962,0.334570109844208,0.343066513538361,0.351855665445328,0.362695455551147,0.378613352775574,0.380859375,0.331835955381393,0.21601539850235,0.148241952061653,0.0892576575279236,0.0394533015787601,0,-0.0577146112918854,-0.0863281190395355,-0.0901852771639824,-0.0605954453349113,-0.0138672972097993,0,0.00942401029169559,0,-0.004736069124192,-0.0686036720871925,0,0.159277111291885,0.484179675579071,0.490722894668579,0.492675513029099,0.549121022224426,0.642968654632568,0.900488495826721],[99.6630859375,99.64404296875,99.6066436767578,99.6540069580078,99.7013626098633,99.6630859375],[99.0784149169922,99.1043930053711,99.06787109375,99.0376968383789,99.0380859375,99.0451202392578,99.0784149169922],[98.5919876098633,98.5799865722656,98.5293960571289,98.6043014526367,98.5919876098633],[98.4090881347656,98.3984375,98.357421875,98.3156280517578,98.2962951660156,98.2623062133789,98.3013687133789,98.3220672607422,98.3509750366211,98.4349594116211,98.3988265991211,98.4090881347656],[98.3075180053711,98.2507858276367,98.2583999633789,98.2736358642578,98.3011779785156,98.3125,98.3075180053711],[100.070701599121,100.075286865234,100.0537109375,99.96240234375,99.9312515258789,99.9395523071289,99.95361328125,100.04296875,100.070701599121],[100.074119567871,100.064453125,100.025680541992,99.998046875,99.9833984375,100.04345703125,100.073051452637,100.074119567871],[102.6064453125,102.589942932129,102.532814025879,102.546485900879,102.568946838379,102.6064453125],[102.4267578125,102.429985046387,102.378120422363,102.359962463379,102.301948547363,102.273338317871,102.277435302734,102.31884765625,102.378120422363,102.408401489258,102.4267578125],[100.122459411621,100.114936828613,100.139747619629,100.174125671387,100.218070983887,100.266014099121,100.317962646484,100.373146057129,100.431549072266,100.491600036621,100.51953125,100.539939880371,100.543067932129,100.514549255371,100.466209411621,100.397651672363,100.420120239258,100.513572692871,100.62548828125,100.743942260742,100.806838989258,100.858200073242,100.906051635742,100.966506958008,101.154685974121,101.2119140625,101.220802307129,101.197555541992,101.2265625,101.279884338379,101.286331176758,101.220504760742,101.16552734375,101.106346130371,101.060447692871,101.046974182129,101.050582885742,101.0927734375,101.137496948242,101.148727416992,101.143951416016,101.11328125,100.9990234375,100.908500671387,100.955856323242,101.045700073242,101.105171203613,101.16748046875,101.299713134766,101.413673400879,101.555076599121,101.563667297363,101.6875,101.744140625,101.774803161621,101.818656921387,101.87548828125,101.947456359863,102.034568786621,102.101463317871,102.148246765137,102.231643676758,102.351852416992,102.458786010742,102.552536010742,102.598236083984,102.596092224121,102.616798400879,102.66064453125,102.680076599121,102.675193786621,102.717575073242,102.807418823242,102.898628234863,102.991401672363,103.051368713379,103.091209411621,103.148536682129,103.19970703125,103.26318359375,103.279594421387,103.248924255371,103.251754760742,103.288276672363,103.366989135742,103.487983703613,103.629684448242,103.792282104492,103.898826599121,103.949615478516,104.048728942871,104.196189880371,104.322654724121,104.428123474121,104.539253234863,104.655860900879,104.739646911621,104.816017150879,104.758979797363,104.743553161621,104.750587463379,104.8193359375,104.949897766113,105.025779724121,105.047172546387,105.148735046387,105.330665588379,105.40625,105.375587463379,105.373245239258,105.39892578125,105.462013244629,105.562400817871,105.6220703125,105.641014099121,105.638870239258,105.615623474121,105.57373046875,105.51318359375,105.505859375,105.490425109863,105.490425109863,105.533393859863,105.546684265137,105.523048400879,105.500198364258,105.497360229492,105.4755859375,105.422660827637,105.342185974121,105.24365234375,105.183303833008,105.169143676758,105.1259765625,105.074119567871,105.03369140625,105.00341796875,104.982421875,104.9697265625,104.878814697266,104.778999328613,104.575782775879,104.41162109375,104.227737426758,104.054298400879,103.981834411621,103.898628234863,103.818359375,103.741889953613,103.600387573242,103.54638671875,103.432426452637,103.3134765625,103.199409484863,103.031051635742,102.909278869629,102.873245239258,102.812797546387,102.728904724121,102.620407104492,102.544723510742,102.565528869629,102.546875,102.428512573242,102.336326599121,102.319725036621,102.330764770508,102.362983703613,102.422653198242,102.461715698242,102.49072265625,102.499610900879,102.629684448242,102.703323364258,102.755661010742,102.737403869629,102.706253051758,102.736618041992,102.918067932129,102.933891296387,102.912307739258,102.883689880371,102.791603088379,102.762992858887,102.654884338379,102.594139099121,102.574806213379,102.540237426758,102.43408203125,102.343162536621,102.259078979492,102.248435974121,102.229591369629,102.134178161621,102.034370422363,101.944526672363,101.889060974121,101.835746765137,101.7236328125,101.444923400879,101.090232849121,100.953712463379,100.897750854492,100.86328125,100.896385192871,100.903907775879,100.946090698242,100.92626953125,100.946975708008,100.962692260742,100.906547546387,100.656051635742,100.602928161621,100.536422729492,100.235641479492,100.122360229492,100.017478942871,99.9905242919922,100.051078796387,100.089942932129,99.9820327758789,99.9639663696289,100.005661010742,99.9890670776367,99.9302673339844,99.8371047973633,99.7987289428711,99.7254867553711,99.6273422241211,99.5613250732422,99.5143508911133,99.4869155883789,99.2847671508789,99.2373046875,99.1650390625,99.1903305053711,99.1946258544922,99.1693344116211,99.1607437133789,99.1913070678711,99.2882843017578,99.2650375366211,99.25390625,99.33544921875,99.3938522338867,99.7238311767578,99.8355484008789,99.8775405883789,99.9046859741211,99.9606475830078,99.9895477294922,100.056243896484,100.129295349121,100.154106140137,100.158882141113,100.163475036621,100.228706359863,100.279396057129,100.453521728516,100.503715515137,100.545211791992,100.439353942871,100.410743713379,100.38037109375,100.342971801758,100.283790588379,100.279983520508,100.324317932129,100.3173828125,100.256637573242,100.158203125,100.160743713379,100.204887390137,100.371383666992,100.423530578613,100.48974609375,100.586227416992,100.70166015625,100.792572021484,101.017868041992,101.154388427734,101.301948547363,101.40087890625,101.497947692871,101.6142578125,101.799217224121,102.10107421875,102.068359375,102.05517578125,101.936134338379,101.917190551758,101.873634338379,101.790725708008,101.719528198242,101.678413391113,101.649993896484,101.601364135742,101.576759338379,101.556053161621,101.404197692871,101.257026672363,101.229782104492,101.190628051758,101.147659301758,101.113960266113,101.081741333008,101.025199890137,100.981643676758,100.992774963379,101.075584411621,101.086524963379,101.075973510742,101.053512573242,101.029388427734,100.98876953125,100.873924255371,100.816505432129,100.793746948242,100.754493713379,100.715629577637,100.629493713379,100.563865661621,100.345413208008,100.261428833008,100.216598510742,100.1767578125,100.161224365234,100.137992858887,100.119140625,99.86865234375,99.6959991455078,99.7203140258789,99.6677703857422,99.6024398803711,99.5530242919922,99.5969696044922,99.5290985107422,99.4351577758789,99.3585891723633,99.3003921508789,99.263671875,99.1833953857422,99.07763671875,99.0426712036133,99.0510787963867,98.9739303588867,98.8724594116211,98.7886734008789,98.7035140991211,98.6363296508789,98.5792007446289,98.4998016357422,98.4740295410156,98.4209976196289,98.3607406616211,98.3054733276367,98.2381820678711,98.2269515991211,98.2417907714844,98.3259735107422,98.3713912963867,98.4431610107422,98.4929656982422,98.5619125366211,98.7025375366211,98.7184600830078,98.7468719482422,98.7683639526367,98.775390625,98.7572250366211,98.7572250366211,98.7869110107422,98.8871078491211,99.025390625,99.1901397705078,99.3587875366211,99.4426803588867,99.4779281616211,99.5152359008789,99.5728530883789,99.6124954223633,99.61474609375,99.52294921875,99.462890625,99.4324188232422,99.4163055419922,99.3942413330078,99.4050750732422,99.3719711303711,99.29736328125,99.2198257446289,99.1735305786133,99.1735305786133,99.1239242553711,99.107421875,99.1371078491211,99.1761703491211,99.1716842651367,99.1560592651367,99.1368179321289,99.0862274169922,99.0146484375,98.93359375,98.72119140625,98.5700149536133,98.4950256347656,98.4001922607422,98.3321304321289,98.2459945678711,98.2021484375,98.1779327392578,98.1910171508789,98.2322235107422,98.2861328125,98.3293991088867,98.4521484375,98.5373001098633,98.5569305419922,98.5652389526367,98.5544967651367,98.5582046508789,98.5740203857422,98.5923767089844,98.8179702758789,98.8655242919922,98.8884735107422,98.8882751464844,98.8693389892578,98.83544921875,98.6892623901367,98.6607437133789,98.5936508178711,98.5647430419922,98.5231399536133,98.4781265258789,98.47119140625,98.4388656616211,98.2565383911133,98.1746139526367,98.0630874633789,97.9292984008789,97.79296875,97.7290954589844,97.7064437866211,97.6985321044922,97.7399368286133,97.7197265625,97.6515655517578,97.6224594116211,97.6322250366211,97.5993194580078,97.5238342285156,97.4507827758789,97.3806610107422,97.3739242553711,97.3970718383789,97.4849624633789,97.51513671875,97.5773468017578,97.6715850830078,97.7277297973633,97.7459030151367,97.7540054321289,97.7060546875,97.7141647338867,97.8039016723633,97.7935562133789,97.8167953491211,97.9164047241211,97.9912109375,98.0150375366211,98.0490264892578,98.1110382080078,98.2390594482422,98.2936553955078,98.3712844848633,98.4549789428711,98.4938507080078,98.7606430053711,98.8195343017578,98.8757781982422,98.9166946411133,98.9580078125,98.9874038696289,99.0206985473633,99.0397415161133,99.07421875,99.1307601928711,99.1968765258789,99.28369140625,99.337890625,99.3992156982422,99.4515609741211,99.4859390258789,99.5016632080078,99.4874954223633,99.4479522705078,99.4588851928711,99.5316390991211,99.638671875,99.7201232910156,99.7733383178711,99.8251953125,99.8903350830078,99.9542999267578,100.003616333008,100.122459411621],[70.70166015625,70.6121063232422,70.5595703125,70.5186538696289,70.4892578125,70.4828109741211,70.4977569580078,70.5670928955078,70.6641616821289,70.6982421875,70.70166015625],[70.6525421142578,70.6227569580078,70.5687484741211,70.5500030517578,70.5720672607422,70.6183547973633,70.6492156982422,70.6525421142578],[70.9580078125,70.9609375,70.9463806152344,70.7385787963867,70.6443405151367,70.6241226196289,70.5992202758789,70.5568389892578,70.51513671875,70.4513626098633,70.37890625,70.2744140625,70.0714874267578,69.966796875,69.7652359008789,69.5302734375,69.49365234375,69.46875,69.4709930419922,69.4878921508789,69.4762649536133,69.4319305419922,69.3654251098633,69.3072280883789,69.27880859375,69.2447204589844,69.2291030883789,69.2802734375,69.2976608276367,69.3915023803711,69.4632797241211,69.5988311767578,69.6669921875,69.7720718383789,69.9559555053711,70.1016616821289,70.1368179321289,70.1710968017578,70.2092742919922,70.2448272705078,70.39208984375,70.5011749267578,70.5679702758789,70.6078109741211,70.6786117553711,70.7331008911133,70.79931640625,71.0048828125,71.0650405883789,71.1180648803711,71.2027359008789,71.2728500366211,71.3285140991211,71.404296875,71.4703140258789,71.5030288696289,71.5173797607422,71.505859375,71.5033187866211,71.5462875366211,71.6726608276367,71.7322235107422,71.7353515625,71.7256774902344,71.7786178588867,71.8059539794922,71.9910125732422,72.0427703857422,72.0841751098633,72.1473617553711,72.22998046875,72.2498016357422,72.2872009277344,72.3577117919922,72.490234375,72.5633773803711,72.6399459838867,72.8724594116211,72.9494094848633,73.1092758178711,73.2349624633789,73.3361282348633,73.3874053955078,73.4704132080078,73.5755844116211,73.6316375732422,73.6363296508789,73.6231384277344,73.6073226928711,73.6904296875,73.7437515258789,73.7956008911133,73.8052749633789,73.7945327758789,73.72998046875,73.7068405151367,73.6960983276367,73.716796875,73.7541046142578,73.8016662597656,73.869140625,73.9700164794922,74.0255813598633,74.0653381347656,74.13134765625,74.1873016357422,74.2774429321289,74.5140686035156,74.7450180053711,74.8123016357422,74.8359375,74.7720718383789,74.7750930786133,74.7896499633789,74.8424835205078,74.8908157348633,74.9002914428711,74.9212875366211,74.9382781982422,74.9123077392578,74.8942413330078,74.9158248901367,74.9864273071289,75.0974578857422,75.1187515258789,75.0790023803711,75.0083999633789,74.9181671142578,74.8913116455078,74.8753890991211,74.8304672241211,74.7305679321289,74.6593780517578,74.5242233276367,74.4449234008789,74.3490219116211,74.2596664428711,74.2035140991211,74.1670913696289,74.0777282714844,73.9488220214844,73.7496109008789,73.6535186767578,73.6275405883789,73.6488265991211,73.71728515625,73.7337875366211,73.7206115722656,73.6571273803711,73.6326141357422,73.6046829223633,73.4813461303711,73.3829116821289,73.2111282348633,72.8955078125,72.7570343017578,72.6574249267578,72.3587951660156,72.1535186767578,71.9419937133789,71.8020477294922,71.7337875366211,71.6656265258789,71.5974578857422,71.5308532714844,71.4718780517578,71.4329147338867,71.4547805786133,71.4796829223633,71.5050811767578,71.5461883544922,71.5803680419922,71.5822296142578,71.5519561767578,71.48779296875,71.3896484375,71.3199234008789,71.2785186767578,71.2828140258789,71.3327102661133,71.255859375,71.0521469116211,70.87890625,70.7359390258789,70.6158218383789,70.5185546875,70.4177780151367,70.3132781982422,70.23876953125,70.2146453857422,70.1994171142578,70.2549743652344,70.25146484375,70.1886672973633,70.1198196411133,70.0447235107422,69.9849624633789,69.9406280517578,69.8208999633789,69.6257781982422,69.4920959472656,69.4201202392578,69.3992156982422,69.4296875,69.4144515991211,69.3538055419922,69.3039093017578,69.2648468017578,69.18017578125,69.0500030517578,68.96044921875,68.9118118286133,68.88525390625,68.8553695678711,68.8384780883789,68.82373046875,68.7820358276367,68.7232360839844,68.6691360473633,68.6370162963867,68.5464859008789,68.3869171142578,68.2995147705078,68.2847671508789,68.2609405517578,68.2121124267578,68.0677719116211,67.9580078125,67.83447265625,67.7660140991211,67.7589874267578,67.7980499267578,67.8143539428711,67.8635711669922,68.0109405517578,68.0875930786133,68.1740264892578,68.2365188598633,68.2940444946289,68.3412094116211,68.3544921875,68.3502883911133,68.3331069946289,68.2513656616211,68.1441421508789,68.0872039794922,68.0559539794922,68.0478515625,68.1485366821289,68.1325149536133,68.103515625,68.0443420410156,67.9595718383789,67.8756790161133,67.7685546875,67.6944351196289,67.6765594482422,67.6672821044922,67.6483383178711,67.6165008544922,67.400390625,67.3576202392578,67.349609375,67.4261703491211,67.4595718383789,67.49169921875,67.54248046875,67.7190475463867,67.9085922241211,68.0771484375,68.2449188232422,68.3030242919922,68.3990249633789,68.4632797241211,68.5069351196289,68.5861282348633,68.6103515625,68.6389617919922,68.6869125366211,68.7352523803711,68.7582015991211,68.7679672241211,68.77783203125,68.7976531982422,68.8324203491211,68.8522491455078,68.8687515258789,68.8638687133789,68.8244171142578,68.7894515991211,68.7927703857422,68.8046875,68.9084930419922,68.9556655883789,68.9720687866211,68.9660186767578,68.9268569946289,68.7845687866211,68.6397476196289,68.6224594116211,68.6306610107422,68.6525421142578,68.9517593383789,69.1103515625,69.2283172607422,69.27490234375,69.2195358276367,69.29443359375,69.30419921875,69.2062530517578,69.2599639892578,69.31396484375,69.3093719482422,69.3572235107422,69.4138641357422,69.4982376098633,69.62841796875,69.6707992553711,69.712890625,69.7732467651367,70.0056610107422,70.1363296508789,70.2920913696289,70.3189392089844,70.3726577758789,70.4019546508789,70.4415054321289,70.5782241821289,70.6573257446289,70.6573257446289,70.634765625,70.63916015625,70.7509765625,70.7510757446289,70.7255859375,70.7120132446289,70.6983413696289,70.548828125,70.3826217651367,70.3771514892578,70.3697280883789,70.37158203125,70.3982391357422,70.4699249267578,70.5335922241211,70.5658187866211,70.6027374267578,70.6531219482422,70.8994140625,70.9580078125],[53.1095695495605,53.1001968383789,53.0458984375,53.0185546875,53.0533218383789,53.0921859741211,53.05517578125,53.1095695495605],[66.5222625732422,66.4718704223633,66.3502960205078,66.1083984375,65.9006805419922,65.7650375366211,65.7438507080078,65.6830062866211,65.6412124633789,65.6080093383789,65.5549774169922,65.3036193847656,65.0896453857422,64.9515609741211,64.8163070678711,64.7824249267578,64.7531204223633,64.67431640625,64.6025390625,64.5658187866211,64.5110321044922,64.3580093383789,64.1843719482422,64.0921859741211,64.0513687133789,64.0423812866211,64.0096740722656,63.9380836486816,63.8624992370605,63.6965827941895,63.5169906616211,63.3016586303711,63.1789093017578,63.1299819946289,63.1085929870605,63.1299819946289,63.1507797241211,63.1697273254395,63.1193351745605,63.0841789245605,63.056640625,62.9802742004395,62.8580055236816,62.7226600646973,62.6880836486816,62.6105461120605,62.5331077575684,62.4628944396973,62.3866195678711,62.3078117370605,62.2711906433105,62.2528305053711,62.2130851745605,62.0896492004395,61.9838829040527,61.9380874633789,61.8410186767578,61.7197265625,61.6209983825684,61.5427703857422,61.4217796325684,61.3777313232422,61.3447303771973,61.2620086669922,61.2388687133789,61.2355499267578,61.2586898803711,61.2521476745605,61.2058563232422,61.1529312133789,61.1594734191895,61.1826133728027,61.21240234375,61.2120132446289,61.1750946044922,61.1603469848633,61.1699256896973,61.1196250915527,60.7079086303711,60.3413047790527,60.3207054138184,60.1783180236816,60.0627899169922,59.9486312866211,59.6872062683105,59.5622062683105,59.4549789428711,59.3673820495605,59.3447265625,59.3269538879395,59.3017578125,59.2741241455078,59.2408180236816,58.9372024536133,58.8154296875,58.7007827758789,58.6501922607422,58.5504837036133,58.4357452392578,58.38671875,58.3181648254395,58.2616233825684,58.1087913513184,57.9805679321289,57.88818359375,57.7105484008789,57.5209999084473,57.4238243103027,57.3537139892578,57.3357429504395,57.3367195129395,57.3314476013184,57.3081092834473,57.2601585388184,57.1935539245605,57.0790023803711,56.9066390991211,56.7746086120605,56.669921875,56.5440406799316,56.4406280517578,56.3668937683105,56.3241195678711,56.2969741821289,56.2720718383789,56.2288093566895,56.1711921691895,56.05029296875,55.84130859375,55.5784187316895,55.380859375,55.2247047424316,55.0755844116211,54.9000968933105,54.8486328125,54.7452163696289,54.6994132995605,54.6396484375,54.5789070129395,54.4586906433105,54.2998046875,54.1916007995605,53.9141578674316,53.8978538513184,53.8478507995605,53.8235321044922,53.8251953125,53.8541030883789,53.8518562316895,53.8400382995605,53.8515625,53.8737335205078,53.8853492736816,53.86865234375,53.81494140625,53.72412109375,53.7097663879395,53.70458984375,53.6175804138184,53.5394515991211,53.4749984741211,53.3363304138184,53.2667961120605,53.2033195495605,53.1566390991211,53.1240234375,53.1248054504395,53.2356452941895,53.3049812316895,53.3896484375,53.4973640441895,53.6031265258789,53.5824203491211,53.5333023071289,53.4722671508789,53.4504890441895,53.4582977294922,53.4873046875,53.4542961120605,53.4042015075684,53.28857421875,53.1385765075684,52.9874992370605,52.9521484375,53.0355491638184,52.96484375,52.8982391357422,52.8046875,52.7444343566895,52.7336921691895,52.7847671508789,52.8499031066895,52.8892593383789,52.9434547424316,52.9976577758789,53.0595703125,53.1452140808105,53.1919937133789,53.3329124450684,53.4236335754395,53.5203132629395,53.615234375,53.6937484741211,53.7637710571289,53.8700180053711,54.0888671875,54.1929664611816,54.2833023071289,54.3298835754395,54.3773460388184,54.3362312316895,54.3194313049316,54.3744163513184,54.5470695495605,54.6570320129395,54.68505859375,54.7100563049316,54.7232437133789,54.7179679870605,54.7037124633789,54.6714859008789,54.5921897888184,54.2845726013184,54.1810569763184,54.0948257446289,54.0398445129395,53.9952125549316,53.9538078308105,53.8464813232422,53.8046875,53.7523422241211,53.6249008178711,53.4958992004395,53.2849578857422,53.1641616821289,53.1082992553711,52.9700202941895,52.9052734375,52.8148460388184,52.8834991455078,52.8822288513184,52.8301734924316,52.86181640625,52.8503913879395,52.8255844116211,52.7472648620605,52.609375,52.4938507080078,52.6968727111816,52.8705062866211,53.0125007629395,53.0558586120605,53.2500991821289,53.5007820129395,53.6853523254395,53.9263687133789,54.0051765441895,54.1209945678711,54.2149391174316,54.2718734741211,54.4728507995605,54.6779289245605,54.8538093566895,54.9037094116211,54.931640625,54.9523468017578,55.1018562316895,55.1623039245605,55.2496070861816,55.3197288513184,55.3883781433105,55.4343757629395,55.4870109558105,55.5452156066895,55.6786117553711,55.8390655517578,55.9349594116211,55.9774398803711,56.2419891357422,56.4798812866211,56.7736320495605,56.86083984375,56.9658203125,57.0179672241211,57.0642547607422,57.0948257446289,57.1188468933105,57.1138687133789,57.07666015625,57.0181617736816,56.98486328125,56.9640617370605,57.03369140625,57.1135749816895,57.2288093566895,57.2906265258789,57.3817367553711,57.6861305236816,57.8142585754395,57.85595703125,57.9234390258789,57.9457054138184,57.9834976196289,58.0289039611816,58.0754852294922,58.1656227111816,58.2340850830078,58.2829132080078,58.3272476196289,58.3705101013184,58.3771476745605,58.3970718383789,58.4314498901367,58.4570350646973,58.4744110107422,58.4858360290527,58.4769515991211,58.4181632995605,58.2886695861816,58.2040977478027,58.1620101928711,58.1515617370605,58.2064476013184,58.2596702575684,58.3531265258789,58.4771499633789,58.5323219299316,58.5890617370605,58.7299842834473,58.876953125,58.9308586120605,59.0358390808105,59.1231422424316,59.1595687866211,59.1991195678711,59.2765617370605,59.3542976379395,59.4510726928711,59.7625961303711,59.8583030700684,59.9365234375,59.9851608276367,60.0060539245605,60.0007820129395,59.9816436767578,59.9791984558105,59.97412109375,59.9493141174316,59.9417953491211,59.9626007080078,60.10693359375,60.1555671691895,60.1920852661133,60.2007827758789,60.1763648986816,60.1085929870605,60.0755844116211,60.0755844116211,60.1240234375,60.1379852294922,60.1060523986816,60.0687484741211,60.0673828125,60.0896492004395,60.2000045776367,60.4549827575684,60.5135765075684,60.7548828125,60.8671913146973,60.9332046508789,61.1199188232422,61.1792945861816,61.2423858642578,61.3289031982422,61.3875007629395,61.4173812866211,61.4436492919922,61.4969749450684,61.6445350646973,61.7999000549316,61.90283203125,61.9535179138184,62.017578125,62.0950202941895,62.1884765625,62.2980461120605,62.375,62.4416046142578,62.4832038879395,62.5254898071289,62.6506805419922,62.9068336486816,63.0581016540527,63.2918930053711,63.5060539245605,63.7208023071289,63.7636756896973,63.9525375366211,64.1627883911133,64.3099594116211,64.5316390991211,64.6218719482422,64.6599578857422,64.8207015991211,65.07666015625,65.3996047973633,65.6128845214844,65.6708984375,65.728515625,65.7902297973633,65.8571243286133,65.97119140625,66.0948257446289,66.1731491088867,66.263671875,66.3353500366211,66.3897476196289,66.5745086669922,66.6062469482422,66.6263656616211,66.6292953491211,66.5255813598633,66.5113296508789,66.5106506347656,66.5222625732422],[124.036331176758,124.198135375977,124.444435119629,124.438285827637,124.412986755371,124.375679016113,124.3193359375,124.282325744629,124.134574890137,124.115531921387,124.090141296387,124.052436828613,124.036331176758],[125.068168640137,125.033592224121,124.996971130371,124.968254089355,124.958595275879,124.960151672363,124.977546691895,125.100486755371,125.149421691895,125.149024963379,125.124404907227,125.100395202637,124.973243713379,124.936805725098,124.915031433105,124.922264099121,125.026954650879,125.11572265625,125.178024291992,125.323135375977,125.3818359375,125.804298400879,125.905082702637,126.1728515625,126.531059265137,126.619728088379,126.734573364258,126.845695495605,126.904685974121,126.966400146484,127.058502197266,127.21484375,127.257026672363,127.296096801758,127.114547729492,126.915229797363,126.792472839355,126.665435791016,126.568557739258,126.486907958984,126.382514953613,126.264747619629,126.164260864258,126.073051452637,125.946090698242,125.894729614258,125.84033203125,125.735160827637,125.408012390137,125.210250854492,125.068168640137],[125.646095275879,125.579490661621,125.507133483887,125.584075927734,125.621101379395,125.646095275879],[-174.913146972656,-174.918655395508,-174.967529296875,-174.972946166992,-174.923477172852,-174.913146972656],[-175.161911010742,-175.147659301758,-175.131942749023,-175.078170776367,-175.084091186523,-175.156600952148,-175.202346801758,-175.33544921875,-175.362350463867,-175.318054199219,-175.322616577148,-175.300445556641,-175.225387573242,-175.158004760742,-175.199768066406,-175.161911010742],[-173.953506469727,-173.991302490234,-174.009323120117,-174.053115844727,-174.069152832031,-174.00244140625,-173.968063354492,-173.921875,-173.923980712891,-173.953506469727],[-61.0121040344238,-61.1742668151855,-61.5966796875,-61.7716789245605,-61.9061012268066,-61.6614723205566,-61.6327171325684,-61.5288581848145,-61.4993171691895,-61.4647407531738,-61.478271484375,-61.4988250732422,-61.5409202575684,-61.6353073120117,-61.6511726379395,-61.5918464660645,-61.46484375,-61.3700218200684,-61.1737289428711,-61.0785140991211,-60.9176254272461,-60.9967308044434,-61.0337371826172,-61.0193405151367,-61.0375022888184,-61.0164108276367,-60.9684524536133,-60.9996070861816,-61.0041007995605,-61.0121040344238],[-60.7563018798828,-60.8106422424316,-60.8042984008789,-60.7089347839355,-60.5627937316895,-60.5254898071289,-60.5464859008789,-60.7563018798828],[10.9576168060303,10.9313478469849,10.8830080032349,10.857421875,10.7847652435303,10.7570314407349,10.7220706939697,10.73388671875,10.7452154159546,10.9219732284546,11.0178709030151,11.0335931777954,11.03759765625,10.9930667877197,10.9576168060303],[11.2780275344849,11.1236333847046,11.1530265808105,11.2548828125,11.2810544967651,11.2780275344849],[11.5045900344849,11.50244140625,11.4671878814697,11.4591798782349,11.4539060592651,11.4539060592651,11.5337886810303,11.5359373092651,11.5049810409546,11.3580074310303,11.1682615280151,11.0051755905151,10.8263673782349,10.7715826034546,10.6830072402954,10.60888671875,10.5955076217651,10.5436515808105,10.4757814407349,10.3060541152954,10.2746095657349,10.1959962844849,10.1598634719849,10.1149415969849,10.1726570129395,10.2433595657349,10.2570314407349,10.2560548782349,10.2164068222046,10.1259765625,10.0597658157349,9.93251991271973,9.89501953125,9.80742168426514,9.63798904418945,9.51874923706055,9.4580078125,9.40605449676514,9.36328220367432,9.28789043426514,9.22402381896973,9.16025352478027,9.10234355926514,9.04404258728027,9.01894569396973,8.84404277801514,8.68290996551514,8.51513576507568,8.33339881896973,8.30419921875,8.2109375,8.11250019073486,8.07558536529541,7.87724590301514,7.7626953125,7.73134708404541,7.70917987823486,7.62753868103027,7.53437471389771,7.50019502639771,7.49560499191284,7.51386737823486,7.55449247360229,7.74853563308716,7.83828163146973,7.94941377639771,8.04560565948486,8.12343788146973,8.19277381896973,8.24560546875,8.25468730926514,8.27685546875,8.31210994720459,8.39423847198486,8.35986328125,8.31640625,8.32900428771973,8.31806659698486,8.28290939331055,8.2470703125,8.24570274353027,8.2802734375,8.30673789978027,8.34873104095459,8.333984375,8.302734375,8.20878982543945,8.20761680603027,8.23076152801514,8.36962890625,8.44423770904541,8.50673866271973,8.60126876831055,8.59765625,8.57656288146973,8.82353591918945,9.05888652801514,9.14199256896973,9.68798828125,9.75888633728027,9.83847713470459,9.81552696228027,9.78398418426514,9.83027362823486,9.89638614654541,9.87939453125,9.87558650970459,9.98808574676514,10.08740234375,10.1963863372803,10.1887702941895,10.3340826034546,10.2932615280151,10.4123048782349,10.5181636810303,10.5712890625,10.7662115097046,10.9513673782349,11.0539064407349,11.0770502090454,11.1266603469849,11.0565433502197,10.9671878814697,10.7981452941895,10.6423826217651,10.5256834030151,10.4879884719849,10.4765625,10.5057611465454,10.5908193588257,10.68896484375,10.78369140625,11.0042972564697,11.0006837844849,11.0315437316895,11.0432615280151,11.1201171875,10.9558591842651,10.8662109375,10.69091796875,10.5348634719849,10.2003908157349,10.1183595657349,10.0648441314697,10.0400390625,10.0490236282349,10.1589841842651,10.3052730560303,10.4542970657349,10.7131843566895,10.7042961120605,10.7227544784546,10.828125,10.8984375,10.9580078125,11.0845708847046,11.1502923965454,11.2574224472046,11.2699213027954,11.2321290969849,11.2026376724243,11.2342767715454,11.3380861282349,11.4005851745605,11.5045900344849],[25.9700202941895,25.740234375,25.6689453125,25.7409191131592,25.8748054504395,25.9183597564697,25.9770526885986,25.9700202941895],[41.5100593566895,41.5765647888184,41.7017593383789,41.7793922424316,41.8235359191895,41.92578125,42.0777359008789,42.2111320495605,42.2799797058105,42.3643531799316,42.4664077758789,42.5079116821289,42.5673828125,42.5904312133789,42.6068344116211,42.6824226379395,42.7541007995605,42.7878913879395,42.8216781616211,42.90673828125,43.05712890625,43.1490249633789,43.1712913513184,43.1410140991211,43.15283203125,43.2054672241211,43.279296875,43.3589820861816,43.40234375,43.4333992004395,43.4416007995605,43.439453125,43.4552726745605,43.5174827575684,43.5916976928711,43.6316413879395,43.6964836120605,43.72265625,43.712890625,43.6678733825684,43.5693359375,43.59375,43.6158180236816,43.6083984375,43.6781234741211,43.7098655700684,43.6833000183105,43.6662139892578,43.7916984558105,43.9419898986816,44.00537109375,44.1780242919922,44.2892570495605,44.3996124267578,44.5604476928711,44.7337875366211,44.7682647705078,44.7833976745605,44.8171882629395,44.7821273803711,44.7250022888184,44.5871086120605,44.5167007446289,44.4559555053711,44.3893547058105,44.33544921875,44.2404289245605,44.1240234375,44.0439453125,44.0232429504395,44.0337905883789,44.0575218200684,44.0743141174316,44.0791015625,44.1212882995605,44.1780242919922,44.1805648803711,44.171875,44.1587867736816,44.14453125,44.1708030700684,44.232421875,44.2716827392578,44.2570304870605,44.2801742553711,44.2978515625,44.2908210754395,44.2985343933105,44.3196296691895,44.3757781982422,44.4308586120605,44.4499015808105,44.4496078491211,44.380859375,44.3727531433105,44.3489265441895,44.3293952941895,44.2679672241211,44.2289085388184,44.2113304138184,44.2229461669922,44.3362312316895,44.3977508544922,44.5612297058105,44.5899391174316,44.5453109741211,44.5460929870605,44.5671844482422,44.5771484375,44.5731468200684,44.5740242004395,44.6041030883789,44.7151374816895,44.7941398620605,44.7967758178711,44.75830078125,44.7667007446289,44.76513671875,44.73095703125,44.6693344116211,44.60595703125,44.5660133361816,44.4959945678711,44.4019546508789,44.3255844116211,44.2818374633789,44.2457008361816,44.2174797058105,44.20166015625,44.2084007263184,44.1917991638184,44.15625,44.1144523620605,44.0646476745605,44.01318359375,43.9400367736816,43.83642578125,43.67578125,43.5679702758789,43.5158233642578,43.3067398071289,43.2630844116211,43.1851539611816,43.0924797058105,42.9366226196289,42.869140625,42.7746086120605,42.7411155700684,42.6354484558105,42.4558601379395,42.3589859008789,42.3128929138184,42.2685546875,42.24755859375,42.2027359008789,42.1678695678711,42.0598640441895,41.8868179321289,41.7435531616211,41.5155258178711,41.3395500183105,41.2646484375,41.1021499633789,40.9588851928711,40.8156242370605,40.7056617736816,40.4503898620605,40.0164031982422,39.6865234375,39.50146484375,39.3566398620605,39.1083984375,38.9064445495605,38.7666015625,38.6888694763184,38.5780296325684,38.4437522888184,38.3839836120605,38.3058586120605,38.1916999816895,37.9066429138184,37.8179702758789,37.7203140258789,37.5235366821289,37.4363288879395,37.3270530700684,37.1874008178711,37.0662117004395,36.9853515625,36.9417991638184,36.7765617370605,36.6585960388184,36.62841796875,36.5968742370605,36.5466804504395,36.5374984741211,36.6414070129395,36.63671875,36.5624008178711,36.47705078125,36.4214859008789,36.3753929138184,36.3475570678711,36.2488288879395,36.2019538879395,36.1536102294922,36.1273422241211,35.9675788879395,35.8926734924316,35.9569320678711,35.8871078491211,35.8109397888184,35.8828125,36.03173828125,36.1884765625,36.1881828308105,36.1800804138184,36.1351585388184,36.0489273071289,35.9045906066895,35.8015632629395,35.7342758178711,35.6611328125,35.6255874633789,35.5374031066895,35.3931655883789,35.1761703491211,34.9431610107422,34.8112297058105,34.70361328125,34.6013679504395,34.2996101379395,34.0234375,33.9548835754395,33.6947250366211,33.5227546691895,33.4417991638184,33.0995140075684,32.9294929504395,32.7948265075684,32.5337905883789,32.3777313232422,32.2837867736816,32.1305656433105,32.02197265625,31.7779293060303,31.3525390625,31.2406253814697,30.9502925872803,30.64404296875,30.58203125,30.5584945678711,30.5060558319092,30.4835948944092,30.4460945129395,30.3873023986816,30.29541015625,30.2316398620605,30.0832023620605,29.7892589569092,29.6890640258789,29.3483390808105,29.2236328125,29.1432628631592,29.1161136627197,29.0655288696289,29.0581035614014,29.0382823944092,28.9696292877197,28.8958969116211,28.8168964385986,28.7176780700684,28.4835929870605,28.3037109375,28.1956043243408,28.1115226745605,28.0194339752197,28.01416015625,28.083984375,27.8038082122803,27.6558589935303,27.5404300689697,27.4539070129395,27.4668941497803,27.5546875,27.630859375,27.9344730377197,28.00537109375,28.0830078125,28.2244148254395,28.2423839569092,28.1336917877197,27.6683578491211,27.3489265441895,27.31103515625,27.2629890441895,27.2497081756592,27.3001937866211,27.3681640625,27.5350589752197,27.5201187133789,27.4005870819092,27.3762683868408,27.28955078125,27.21923828125,27.2039051055908,27.14794921875,27.0679702758789,27.0778331756592,27.2244129180908,27.2547855377197,27.232421875,27.15869140625,26.94384765625,26.8786125183105,26.8074226379395,26.6828117370605,26.62109375,26.5824222564697,26.5247058868408,26.4279289245605,26.3329105377197,26.2907238006592,26.3436527252197,26.4164066314697,26.4296894073486,26.3722648620605,26.3778324127197,26.4413089752197,26.5135746002197,26.5865230560303,26.6103515625,26.5950202941895,26.6413078308105,26.6742172241211,26.6963863372803,26.7273426055908,26.7699222564697,26.8614273071289,27.0986347198486,27.1442394256592,26.9704113006592,26.9068355560303,26.8377914428711,26.7953128814697,26.7876949310303,26.763671875,26.7901363372803,26.9091796875,27.013671875,26.9701175689697,26.9203128814697,26.8662109375,26.81494140625,26.8083019256592,26.8493156433105,26.8536128997803,26.7193355560303,26.6818370819092,26.7107429504395,26.8132801055908,26.9109363555908,26.8992176055908,26.8270492553711,26.4840831756592,26.3507804870605,26.1130847930908,26.0959968566895,26.1013679504395,26.1546878814697,26.1498050689697,26.1813488006592,26.3133792877197,26.4753913879395,26.7380867004395,27.0121097564697,27.1216812133789,27.2845706939697,27.3141613006592,27.3326168060303,27.4755859375,27.72802734375,27.7893562316895,27.8485355377197,27.7318344116211,27.7691402435303,27.8749027252197,27.9895515441895,27.9948253631592,27.96435546875,27.9289073944092,27.9625988006592,28.2890625,28.6302738189697,28.7388687133789,29.0071296691895,29.0551776885986,28.9740238189697,28.8946304321289,28.8412113189697,28.7878913879395,28.9580078125,29.0541019439697,29.5076160430908,29.8449211120605,29.8492183685303,29.8005867004395,29.36474609375,29.259765625,29.1138648986816,29.0822277069092,29.0455093383789,29.0673828125,29.0943355560303,29.1481437683105,29.322265625,29.9193363189697,30.3449211120605,30.81005859375,31.2548847198486,31.3466777801514,31.4580078125,32.08642578125,32.3064460754395,32.5421867370605,32.9466819763184,33.2847633361816,33.38134765625,34.1929702758789,34.75048828125,35.0064430236816,35.1548805236816,35.1410140991211,35.1140594482422,35.1220703125,35.2091827392578,35.2977561950684,35.5580101013184,35.9198265075684,35.9781227111816,36.0517578125,36.17919921875,36.2784156799316,36.4053726196289,36.5096664428711,36.5871086120605,36.6470680236816,36.7777328491211,36.9919929504395,37.0662117004395,37.4309577941895,37.765625,37.9100608825684,38.3810539245605,38.5569343566895,38.8521461486816,39.4263687133789,39.8079109191895,39.9111328125,40.0001983642578,40.12841796875,40.2652359008789,40.6875,40.8195343017578,40.95947265625,41.0835952758789,41.4143562316895,41.5100593566895],[28.0144519805908,27.9873046875,28.05029296875,28.1978511810303,28.3463859558105,28.9467792510986,29.0572261810303,29.0321292877197,28.9959964752197,28.9562492370605,28.7803707122803,28.294921875,28.1721687316895,28.0855464935303,27.9251956939697,27.7473640441895,27.4994144439697,27.43017578125,27.2580070495605,26.974609375,26.7720699310303,26.4679679870605,26.3299789428711,26.2717761993408,26.2027359008789,26.2259769439697,26.2601566314697,26.2523422241211,26.2538070678711,26.3552742004395,26.4474601745605,26.7203121185303,26.7920894622803,26.578125,26.3609371185303,26.2242202758789,26.10546875,26.0677738189697,26.0389633178711,26.0697250366211,26.1091804504395,26.1789054870605,26.2412109375,26.3310546875,26.3541030883789,26.3541030883789,26.3326187133789,26.3284168243408,26.32568359375,26.3306636810303,26.5364246368408,26.6023445129395,26.6249027252197,26.6097640991211,26.5813484191895,26.5445308685303,26.4950199127197,26.4624996185303,26.4105472564697,26.3208980560303,26.3179683685303,26.3272457122803,26.3603515625,26.5114250183105,26.529296875,26.5497074127197,26.5796871185303,26.6153335571289,26.67919921875,26.8003902435303,26.8848628997803,26.96875,27.01171875,27.193359375,27.2443351745605,27.294921875,27.3628902435303,27.4748039245605,27.5348625183105,27.5798816680908,27.6611328125,27.7388668060303,27.8016605377197,27.8319339752197,27.8791999816895,28.0144519805908],[118.407424926758,118.451179504395,118.432723999023,118.295120239258,118.287307739258,118.33935546875,118.407424926758],[121.008796691895,120.946868896484,120.897369384766,120.877342224121,120.87841796875,120.8642578125,120.83984375,120.742774963379,120.690139770508,120.678024291992,120.607620239258,120.581253051758,120.479782104492,120.387603759766,120.316215515137,120.325584411621,120.272857666016,120.232818603516,120.150100708008,120.12158203125,120.083396911621,120.072456359863,120.085548400879,120.121185302734,120.142974853516,120.125389099121,120.132125854492,120.158981323242,120.629692077637,120.757415771484,120.8359375,120.901557922363,120.964057922363,121.040626525879,121.095413208008,121.365425109863,121.449615478516,121.517082214355,121.593650817871,121.643058776855,121.687110900879,121.733299255371,121.852828979492,121.905174255371,121.929000854492,121.856246948242,121.820114135742,121.813377380371,121.826370239258,121.828018188477,121.737014770508,121.639358520508,121.613090515137,121.583404541016,121.526069641113,121.477157592773,121.3974609375,121.352241516113,121.2958984375,121.161231994629,121.008796691895],[39.7113265991211,39.6572265625,39.6361351013184,39.6029281616211,39.66064453125,39.7166023254395,39.8465843200684,39.8909187316895,39.9071273803711,39.8977546691895,39.8244132995605,39.7618179321289,39.7113265991211],[39.4964828491211,39.5730476379395,39.5631828308105,39.5091781616211,39.48095703125,39.4473648071289,39.4236335754395,39.3826179504395,39.3126945495605,39.2434577941895,39.1823234558105,39.2062492370605,39.1923828125,39.2669944763184,39.3089866638184,39.3572273254395,39.3682594299316,39.4333000183105,39.4878921508789,39.4964828491211],[39.8650360107422,39.8709983825684,39.8556632995605,39.8589820861816,39.85302734375,39.7958984375,39.7493171691895,39.7076187133789,39.6734390258789,39.6467742919922,39.7010765075684,39.6734390258789,39.78076171875,39.8650360107422],[33.9032211303711,33.9793968200684,34.0515632629395,34.1316413879395,34.3447265625,34.5579071044922,34.7710914611816,34.9842758178711,35.1974601745605,35.4105453491211,35.6237297058105,35.8369140625,36.0500030517578,36.2630844116211,36.4763641357422,36.689453125,36.9026374816895,37.1158218383789,37.3290023803711,37.5421867370605,37.6438484191895,37.6591796875,37.6768569946289,37.68798828125,37.6818351745605,37.6253929138184,37.6086921691895,37.6082038879395,37.6220703125,37.6701164245605,37.7110366821289,37.7261695861816,37.7574234008789,37.7972679138184,37.8873062133789,38.0408210754395,38.1943359375,38.3478507995605,38.5013656616211,38.6548805236816,38.8083953857422,38.9619140625,39.1154327392578,39.1901359558105,39.2217788696289,39.2018585205078,39.1232414245605,39.1187515258789,39.0879859924316,39.0583000183105,38.9782218933105,38.9110336303711,38.8192367553711,38.8046875,38.8552742004395,38.8740234375,38.9814453125,39.0673828125,39.12548828125,39.2284164428711,39.2873077392578,39.4723625183105,39.5460968017578,39.5192375183105,39.4333953857422,39.3531265258789,39.2884788513184,39.2870140075684,39.3304710388184,39.4284172058105,39.4410171508789,39.3400382995605,39.3089866638184,39.3040008544922,39.3773460388184,39.4883804321289,39.4800796508789,39.4512672424316,39.6413078308105,39.62548828125,39.6966819763184,39.7279281616211,39.7748031616211,39.7837905883789,39.7251968383789,39.86376953125,39.9452133178711,39.9835929870605,40.0836944580078,40.1378898620605,40.2160148620605,40.3887710571289,40.435546875,40.4525413513184,40.4635734558105,40.3474617004395,40.1662101745605,39.9886703491211,39.8170890808105,39.6944351196289,39.5634765625,39.4391632080078,39.3215827941895,39.1709938049316,38.9874992370605,38.7947273254395,38.6033210754395,38.4917984008789,38.3151397705078,38.1765632629395,38.0172843933105,37.9202117919922,37.8853530883789,37.8550758361816,37.8292961120605,37.7248039245605,37.5416984558105,37.3728523254395,37.2183609008789,37.1138687133789,37.0591773986816,36.9789085388184,36.8726577758789,36.7710952758789,36.673828125,36.5186500549316,36.3056640625,36.1913070678711,36.1754875183105,36.0822257995605,35.9113273620605,35.7854461669922,35.7046890258789,35.6309547424316,35.5643539428711,35.50439453125,35.4513664245605,35.4182586669922,35.1826171875,34.95947265625,34.95263671875,34.93701171875,34.890625,34.8505859375,34.8008804321289,34.7738265991211,34.7521476745605,34.7264633178711,34.6884765625,34.6380844116211,34.60791015625,34.59765625,34.6056671142578,34.65234375,34.6670913696289,34.6618156433105,34.6365242004395,34.5835952758789,34.5895500183105,34.5715827941895,34.5697250366211,34.5799789428711,34.5699234008789,34.5242195129395,34.5242195129395,34.4759788513184,34.3278312683105,34.3208961486816,34.0885772705078,33.99560546875,33.9621086120605,33.9496078491211,33.9593772888184,33.9537086486816,33.9439468383789,33.8888664245605,33.8541984558105,33.7662124633789,33.6976585388184,33.5275421142578,33.4677734375,33.4208984375,33.3308601379395,33.2252960205078,33.1304664611816,32.9740219116211,32.9373016357422,32.919921875,32.86328125,32.7566413879395,32.6083984375,32.4871101379395,32.4332046508789,32.3193359375,32.2208976745605,32.1297836303711,32.0353546142578,31.9425792694092,31.9218769073486,31.9186515808105,31.8861331939697,31.8180675506592,31.7447280883789,31.7000007629395,31.6736335754395,31.61279296875,31.5562496185303,31.5348644256592,31.4492206573486,31.3505859375,31.0763664245605,31.0333976745605,30.9683609008789,30.8919925689697,30.8306636810303,30.7767581939697,30.7511711120605,30.7208995819092,30.6538066864014,30.5588874816895,30.4856452941895,30.40673828125,30.3745098114014,30.3131828308105,30.2126960754395,30.1618156433105,30.1062507629395,29.9618167877197,29.7981452941895,29.7096672058105,29.5906238555908,29.5408210754395,29.5062503814697,29.4800758361816,29.4908199310303,29.5963878631592,29.6070327758789,29.5941410064697,29.5423812866211,29.5037117004395,29.4764633178711,29.4201183319092,29.3427734375,29.3234367370605,29.32568359375,29.3675765991211,29.4041996002197,29.4032230377197,29.7177734375,29.7695293426514,29.947265625,30.1471672058105,30.1871109008789,30.2685527801514,30.3484363555908,30.3791007995605,30.4000015258789,30.4249992370605,30.5298843383789,30.6319332122803,30.6246070861816,30.6109371185303,30.6260738372803,30.6818351745605,30.7902336120605,30.8114280700684,30.8111343383789,30.7935562133789,30.796875,30.7802753448486,30.70947265625,30.6042976379395,30.5150375366211,30.4555683135986,30.4334964752197,30.4240226745605,30.4413089752197,30.4504890441895,30.4733390808105,30.4343738555908,30.4242210388184,30.4419918060303,30.53369140625,30.5536117553711,30.5933589935303,30.6566410064697,30.7148456573486,30.7624988555908,30.7976570129395,30.8287105560303,30.85498046875,30.8765640258789,30.8646488189697,30.8191413879395,30.8067378997803,30.8275394439697,30.8125991821289,30.7622051239014,30.7107429504395,30.6319332122803,30.5081062316895,30.47021484375,30.4770526885986,30.5099601745605,30.5199222564697,30.5987319946289,30.6727542877197,30.7419910430908,30.8091793060303,30.8236331939697,30.8447284698486,30.94970703125,31.1275386810303,31.3052730560303,31.4831047058105,31.6608390808105,31.8385753631592,32.0164070129395,32.1941413879395,32.3718757629395,32.5497055053711,32.7274436950684,32.9051780700684,33.0830078125,33.2607421875,33.4384765625,33.6163101196289,33.7940444946289,33.9032211303711],[33.9032211303711,33.7940444946289,33.6163101196289,33.4384765625,33.2607421875,33.0830078125,32.9051780700684,32.7274436950684,32.5497055053711,32.3718757629395,32.1941413879395,32.0164070129395,31.8385753631592,31.6608390808105,31.4831047058105,31.3052730560303,31.1275386810303,30.94970703125,30.8447284698486,30.8236331939697,30.8091793060303,30.7419910430908,30.6727542877197,30.5987319946289,30.5199222564697,30.5099601745605,30.4699211120605,30.4123058319092,30.3602523803711,30.3205070495605,30.2798824310303,30.20703125,30.1499977111816,30.1015625,29.9905281066895,29.9300765991211,29.8999996185303,29.8816413879395,29.8468742370605,29.8253898620605,29.6096687316895,29.5769519805908,29.5799808502197,29.5640640258789,29.5619125366211,29.5900382995605,29.6064453125,29.6082038879395,29.6478538513184,29.6332035064697,29.6843738555908,29.6978511810303,29.7176742553711,29.7497081756592,29.77783203125,29.8146495819092,29.8854484558105,29.9344711303711,29.9238262176514,29.931640625,29.9428691864014,30.0473651885986,30.1829090118408,30.2401371002197,30.3210945129395,30.4778308868408,30.4781246185303,30.9425773620605,31.1587905883789,31.2527351379395,31.2560558319092,31.2740230560303,31.2363262176514,31.1914081573486,31.1763668060303,31.1375961303711,31.0821285247803,31.0453128814697,31.0036125183105,30.9619140625,30.8300762176514,30.7286148071289,30.7298812866211,30.76953125,30.8466777801514,30.8507823944092,30.8399391174316,30.8213863372803,30.7865238189697,30.7540016174316,30.7792987823486,30.8278331756592,30.8675785064697,30.9064445495605,30.8953113555908,30.8385734558105,30.8681659698486,30.9293956756592,31.0480461120605,31.1523418426514,31.2219734191895,31.357421875,31.47998046875,31.5471687316895,31.62890625,31.7980480194092,31.8386707305908,31.8882808685303,31.9417953491211,32.0482406616211,32.0994148254395,32.1359367370605,32.15625,32.1966781616211,32.2455062866211,32.3357429504395,32.5347671508789,32.6769523620605,32.7371101379395,32.8380851745605,32.9972648620605,33.1540985107422,33.3243141174316,33.4893569946289,33.53955078125,33.5684585571289,33.7416038513184,33.97607421875,34.1320343017578,34.1857414245605,34.17822265625,34.1650390625,34.26708984375,34.3928718566895,34.4376945495605,34.4417953491211,34.3994140625,34.4072265625,34.4478492736816,34.5225601196289,34.5891609191895,34.7232398986816,34.7424812316895,34.7734375,34.814453125,34.8466796875,34.8662109375,34.90576171875,34.8830070495605,34.9139633178711,34.9640617370605,34.9775390625,34.9782218933105,34.9764671325684,34.9652328491211,34.9412117004395,34.8983383178711,34.8509750366211,34.8095703125,34.7835960388184,34.8038101196289,34.7986335754395,34.78759765625,34.7267570495605,34.6491241455078,34.6019515991211,34.5352516174316,34.4817390441895,34.4108390808105,34.2925758361816,34.2725563049316,34.1609382629395,34.1117210388184,34.08056640625,34.0372047424316,33.9431648254395,33.9214859008789,33.9244117736816,33.9000015258789,33.9032211303711],[32.01220703125,32.1500968933105,32.0093727111816,31.7001953125,31.5638675689697,31.5287094116211,31.5087890625,31.5848636627197,31.6384773254395,32.01220703125],[38.21435546875,38.1783180236816,37.8287124633789,37.5433616638184,37.33984375,37.2185554504395,37.0475578308105,36.9320297241211,36.7948226928711,36.6886711120605,36.5587882995605,36.4320297241211,36.2794914245605,36.1946296691895,36.02490234375,35.8271484375,35.4001960754395,35.2566413879395,35.2043914794922,35.13232421875,35.0552749633789,35.0145530700684,35.2177734375,35.2801742553711,35.2909164428711,35.2919921875,35.2303733825684,35.0640640258789,34.9695320129395,34.849609375,34.84375,34.8573226928711,34.9066429138184,35.0228500366211,35.2601547241211,35.3739242553711,35.45751953125,35.5580101013184,35.7509765625,35.83349609375,36.0128898620605,36.0771484375,36.1705055236816,36.2903327941895,36.4270515441895,36.5750007629395,36.5142555236816,36.4507789611816,36.4284172058105,36.3933601379395,36.2298812866211,36.0547866821289,35.8701171875,35.8036117553711,35.7594718933105,35.6775398254395,35.5695304870605,35.4725608825684,35.3578147888184,35.15478515625,35.0876960754395,34.8877944946289,34.7168960571289,34.4699211120605,34.28173828125,34.0744132995605,33.9099578857422,33.7556648254395,33.6558609008789,33.45068359375,33.4626960754395,33.4913101196289,33.5300788879395,33.6122093200684,33.6011695861816,33.55517578125,33.3924789428711,33.2615203857422,33.1869125366211,32.9186515808105,32.7726593017578,32.611328125,32.5518569946289,32.5080070495605,32.8280258178711,33.1422843933105,33.2800788879395,33.4662132263184,33.6648445129395,33.63671875,33.5941429138184,33.4988250732422,33.4298820495605,33.2634773254395,33.2022476196289,32.9417991638184,32.796875,32.4767570495605,32.3298835754395,32.0357437133789,31.9251956939697,31.8312492370605,31.7799816131592,31.8428707122803,31.9159164428711,31.9916973114014,32.0130882263184,32.0084953308105,31.8557624816895,31.7136707305908,31.6236324310303,31.5548839569092,31.7160148620605,31.8779296875,32.1314468383789,32.361328125,32.4189453125,32.5525398254395,32.5780296325684,32.3540992736816,32.1272468566895,32.0443344116211,31.9743175506592,31.9449214935303,31.9640636444092,31.9395484924316,31.8647441864014,31.8381824493408,31.7591800689697,31.8369159698486,31.8659191131592,31.9126968383789,31.9016590118408,31.8728523254395,31.7795886993408,31.6570320129395,31.5321292877197,31.5633792877197,31.4968738555908,31.4029293060303,31.3203144073486,31.1368179321289,30.7962875366211,30.7728500366211,30.7216777801514,30.6722660064697,30.6567363739014,30.5115242004395,30.4929695129395,30.2190418243408,30.1841812133789,30.0066394805908,29.9016590118408,29.8211898803711,29.6850566864014,29.62841796875,29.6016616821289,29.6011714935303,29.6703109741211,29.7269535064697,29.7058582305908,29.6519508361816,29.5676765441895,29.4037113189697,29.2235355377197,29.0274391174316,28.8943367004395,28.8243160247803,28.78173828125,28.7666015625,28.7698249816895,28.7914066314697,28.7882804870605,28.7607421875,28.4512691497803,28.3176765441895,28.2125015258789,28.2648448944092,28.3103523254395,28.4713878631592,28.4990234375,28.5017585754395,28.5137691497803,28.5094718933105,28.4916019439697,28.5623035430908,28.6675777435303,28.7292957305908,28.73876953125,28.8495140075684,28.9477558135986,28.9718761444092,29.0062484741211,28.9437503814697,28.9305667877197,28.9274425506592,28.9583969116211,29.0499019622803,29.1462879180908,29.1862316131592,29.2007789611816,29.2045917510986,29.2238292694092,29.2545890808105,29.3048820495605,29.3395519256592,29.3928718566895,29.4328117370605,29.4587898254395,29.4910144805908,29.5550785064697,29.6149425506592,29.6645488739014,29.7068347930908,29.751953125,29.837890625,29.8780269622803,30.07568359375,30.1075191497803,30.1310558319092,29.92431640625,29.9347648620605,29.9424800872803,29.9180660247803,29.8778324127197,29.7197265625,29.5977535247803,29.5719738006592,29.5686531066895,29.5634765625,29.5150394439697,29.5109386444092,29.5417976379395,29.5493183135986,29.5391616821289,29.5106449127197,29.4556655883789,29.3833980560303,29.3337879180908,29.2005844116211,29.1597652435303,29.1348609924316,29.1229496002197,29.15087890625,29.1860332489014,29.2107410430908,29.2111339569092,29.19482421875,29.1253910064697,29.0929679870605,29.0369129180908,28.9733409881592,28.9231433868408,28.8658199310303,28.7738285064697,28.6016597747803,28.5304679870605,28.4630851745605,28.4419918060303,28.4230461120605,28.3875007629395,28.3405284881592,28.3269519805908,28.34716796875,28.291015625,28.1587886810303,28.0884761810303,28.080078125,28.0384769439697,27.96337890625,27.890625,27.8200187683105,27.7144546508789,27.57373046875,27.5622081756592,27.5492172241211,27.4583988189697,27.40380859375,27.3369140625,27.228515625,27.0084953308105,26.9005851745605,26.8470687866211,26.6404285430908,26.6189460754395,26.5724601745605,26.4423828125,26.3056640625,26.2769527435303,26.2362308502197,26.1626949310303,25.90869140625,25.6892585754395,25.4642581939697,25.1696300506592,25.0738277435303,24.9791011810303,24.8933582305908,24.837890625,24.6509761810303,24.5789070129395,24.4840812683105,24.3809566497803,24.2819347381592,24.177734375,24.0597667694092,24.04736328125,24.0018558502197,23.708984375,23.6820316314697,23.6690425872803,23.6287097930908,23.408203125,23.20263671875,23.1394538879395,23.0908203125,23.0547847747803,22.9128913879395,22.8766593933105,22.8560543060303,22.8464832305908,22.8572254180908,22.8362312316895,22.7822265625,22.7691402435303,22.7015628814697,22.68310546875,22.6763668060303,22.5824222564697,22.5201168060303,22.423828125,22.3501949310303,22.3166980743408,22.2951164245605,22.2721672058105,22.2694339752197,22.2537117004395,22.2311515808105,22.2271480560303,22.1318359375,22.1428699493408,22.2952136993408,22.3326168060303,22.3894519805908,22.4320316314697,22.4832038879395,22.5241222381592,22.5386714935303,22.5799808502197,22.7012691497803,22.8097648620605,22.8397464752197,22.85205078125,22.8470706939697,22.7601566314697,22.7056636810303,22.7023429870605,22.7219734191895,22.732421875,22.7199230194092,22.66064453125,22.6494140625,22.7061519622803,22.8907241821289,22.9522457122803,23.0363273620605,23.2644538879395,23.4085941314697,23.5061531066895,23.6490230560303,23.7117195129395,23.97265625,24.0049800872803,24.0526351928711,24.0899410247803,24.0947265625,24.0462894439697,24.00732421875,23.9784164428711,23.9970703125,24.0259780883789,24.0616207122803,24.1057605743408,24.0958003997803,23.9857425689697,23.9380874633789,23.8634777069092,23.7122077941895,23.6644515991211,23.6576175689697,23.6796875,23.6588878631592,23.6052742004395,23.6137714385986,23.6085929870605,23.6466808319092,23.7068347930908,23.7916984558105,23.8642578125,23.951171875,23.9783191680908,24.1268558502197,24.2800769805908,24.32373046875,24.3619136810303,24.4952144622803,24.611328125,24.6851558685303,24.8664073944092,24.9738292694092,25.0666980743408,25.2671871185303,25.5802745819092,25.7857418060303,25.92529296875,26.26708984375,26.3943347930908,26.4534187316895,26.56689453125,26.7734375,26.9528312683105,27.0741214752197,27.1419925689697,27.2701168060303,27.2962894439697,27.34765625,27.4523429870605,27.6013679504395,27.6897449493408,27.6767578125,27.6999988555908,27.7413082122803,27.7888679504395,27.8288078308105,27.8585948944092,28.0107421875,28.0802745819092,28.1444339752197,28.1837882995605,28.2916011810303,28.4246082305908,28.5320301055908,28.5990238189697,28.6477527618408,28.6902332305908,28.7312488555908,28.7932605743408,28.8495140075684,28.9275379180908,28.9777336120605,29.0130863189697,29.0607414245605,29.1020488739014,29.1356430053711,29.1742172241211,29.2304668426514,29.298828125,29.3464851379395,29.4696292877197,29.5531253814697,29.7060527801514,29.9087905883789,30.0637702941895,30.1607418060303,30.2195301055908,30.3089847564697,30.3333988189697,30.4495124816895,30.5445308685303,30.5769538879395,30.6325206756592,30.6117191314697,30.6023445129395,30.5607414245605,30.5330085754395,30.5838871002197,30.6394538879395,30.6672859191895,30.7552738189697,30.8457050323486,30.9806613922119,31.0792980194092,31.1684589385986,31.2179698944092,31.3459949493408,31.57373046875,31.7633781433105,31.7824211120605,31.8755874633789,31.9738292694092,32.0416030883789,32.1222686767578,32.216796875,32.2828140258789,32.3629875183105,32.3913078308105,32.4354476928711,32.5079078674316,32.6454124450684,32.8064460754395,32.8997077941895,33.1484375,33.287109375,33.4518547058105,33.6133804321289,33.7352523803711,33.81884765625,33.9220695495605,34.0153312683105,34.1130867004395,34.3978500366211,34.4027328491211,34.3792953491211,34.2391624450684,34.12109375,34.1154289245605,34.1467742919922,34.2008781433105,34.2092742919922,34.20654296875,34.2298812866211,34.2749977111816,34.2806663513184,34.2284164428711,34.2138671875,34.2341766357422,34.4910163879395,34.6167945861816,34.7123069763184,34.7603530883789,34.8685531616211,34.990234375,35.0640640258789,35.0925788879395,35.1153297424316,35.1581039428711,35.1980476379395,35.2691383361816,35.3119125366211,35.3347663879395,35.30908203125,35.3147468566895,35.3460922241211,35.3832054138184,35.4173812866211,35.4401359558105,35.4401359558105,35.41162109375,35.3917007446289,35.41162109375,35.4884796142578,35.5455055236816,35.5911102294922,35.6737289428711,35.7961921691895,35.8902320861816,36.0078125,36.1164054870605,36.189453125,36.2433624267578,36.3060531616211,36.3688468933105,36.4998016357422,36.5596656799316,36.6194343566895,36.6963882446289,36.7590827941895,36.9884757995605,37.1312522888184,37.1710968017578,37.2548828125,37.3431663513184,37.4228515625,37.5013694763184,37.5823249816895,37.6050796508789,37.7041969299316,37.9502944946289,38.046875,38.1124992370605,38.1467781066895,38.1626968383789,38.1775398254395,38.2086944580078,38.2585945129395,38.451171875,38.5519561767578,38.6477546691895,38.7766609191895,38.9183616638184,39.0277366638184,39.1149406433105,39.1748046875,39.2118148803711,39.2459983825684,39.3029289245605,39.3684539794922,39.4627952575684,39.6265602111816,39.7805671691895,39.8768539428711,39.95849609375,40.0306625366211,40.0806617736816,40.0949211120605,40.0578117370605,40.0578117370605,40.1261711120605,40.1283226013184,40.1087913513184,40.0700187683105,39.9763679504395,39.8897476196289,39.7594718933105,39.6865234375,39.7056655883789,39.7533187866211,39.86376953125,39.9891586303711,40.0036125183105,39.9844741821289,39.9041023254395,39.7928695678711,39.755859375,39.70458984375,39.67041015625,39.6447257995605,39.7654304504395,39.8356437683105,39.8575210571289,39.8826179504395,39.8898468017578,39.8499031066895,39.8474617004395,39.8663063049316,39.9181632995605,39.9579124450684,39.9610366821289,39.8850555419922,39.81396484375,39.7757797241211,39.7787094116211,39.7359352111816,39.6584968566895,39.3910140991211,39.1584968566895,39.0578117370605,38.9002952575684,38.822265625,38.7189445495605,38.640625,38.5109405517578,38.3688468933105,38.2873992919922,38.2587890625,38.2565422058105,38.2432594299316,38.21240234375,38.2013664245605,38.2080078125,38.2410163879395,38.28076171875,38.28076171875,38.2653312683105,38.22119140625,38.2013664245605,38.2058601379395,38.21435546875],[-53.37060546875,-53.4195785522461,-53.4724617004395,-53.5345191955566,-53.742919921875,-53.7853012084961,-54.01025390625,-54.1685523986816,-54.2721214294434,-54.3653297424316,-54.9022941589355,-55.0951156616211,-55.2378921508789,-55.37060546875,-55.6731452941895,-55.8629379272461,-56.117919921875,-56.1946296691895,-56.2499504089355,-56.3878440856934,-56.4630889892578,-56.8551750183105,-57.1707038879395,-57.54345703125,-57.8291015625,-57.8732452392578,-57.9021492004395,-57.9612312316895,-58.2070274353027,-58.4001922607422,-58.4381370544434,-58.4113273620605,-58.3533668518066,-58.3635215759277,-58.2921867370605,-58.2215843200684,-58.1535606384277,-58.0926780700684,-58.0823249816895,-58.1295890808105,-58.1622085571289,-58.2011680603027,-58.123046875,-58.1197242736816,-58.1647987365723,-58.177001953125,-58.1563491821289,-58.160400390625,-58.1890106201172,-58.1674842834473,-58.0958480834961,-58.0423316955566,-58.0069808959961,-57.9888648986816,-57.9879837036133,-58.0096702575684,-58.0538558959961,-58.0333976745605,-57.9483413696289,-57.8933601379395,-57.868408203125,-57.8700675964355,-57.8982925415039,-57.8863296508789,-57.8340797424316,-57.8105964660645,-57.8185539245605,-57.8725051879883,-57.8311996459961,-57.7126960754395,-57.6508827209473,-57.645751953125,-57.6088905334473,-57.5522956848145,-57.3838424682617,-57.2144508361816,-57.1869125366211,-57.1205062866211,-57.03271484375,-56.937255859375,-56.8327178955078,-56.7216796875,-56.4072265625,-56.1761703491211,-56.1058578491211,-56.0448226928711,-55.9989776611328,-56.0184555053711,-56.0155258178711,-56.0046844482422,-55.9519996643066,-55.8736839294434,-55.8077659606934,-55.75634765625,-55.7059555053711,-55.6652374267578,-55.6504898071289,-55.6271476745605,-55.60302734375,-55.5573272705078,-55.4495582580566,-55.3660621643066,-55.3455085754395,-55.3132820129395,-55.2789535522461,-55.254638671875,-55.1735343933105,-55.0911636352539,-55.0360336303711,-54.89599609375,-54.587646484375,-54.5309104919434,-54.4776840209961,-54.3699226379395,-54.2205581665039,-54.1004409790039,-53.9851570129395,-53.9206047058105,-53.8765144348145,-53.8061027526855,-53.76171875,-53.74658203125,-53.7011222839355,-53.6536102294922,-53.6017074584961,-53.4894027709961,-53.3627433776855,-53.2312507629395,-53.1572761535645,-53.1255874633789,-53.2140617370605,-53.3101081848145,-53.3952140808105,-53.4828605651855,-53.5118637084961,-53.5313491821289,-53.5303726196289,-53.5376472473145,-53.5313491821289,-53.5188484191895,-53.4635772705078,-53.3975601196289,-53.37060546875],[-155.581344604492,-155.625640869141,-155.680755615234,-155.881301879883,-155.905609130859,-155.890716552734,-155.9658203125,-156.048675537109,-155.988433837891,-155.908889770508,-155.8203125,-155.892776489258,-155.874267578125,-155.831634521484,-155.6220703125,-155.198776245117,-155.086090087891,-155.06591796875,-154.989013671875,-154.952575683594,-154.841354370117,-154.80419921875,-154.850296020508,-155.053466796875,-155.309616088867,-155.535247802734,-155.581344604492],[-156.849609375,-156.908889770508,-156.973388671875,-156.988433837891,-157.050582885742,-156.941787719727,-156.880569458008,-156.848281860352,-156.809371948242,-156.849609375],[-156.48681640625,-156.460830688477,-156.354400634766,-156.277542114258,-156.148345947266,-156.103515625,-156.018646240234,-155.989837646484,-156.013565063477,-156.107131958008,-156.234771728516,-156.309951782227,-156.408798217773,-156.438232421875,-156.448867797852,-156.480087280273,-156.543853759766,-156.615432739258,-156.689697265625,-156.69775390625,-156.656890869141,-156.585403442383,-156.532318115234,-156.48681640625],[-157.213623046875,-157.002288818359,-156.952331542969,-156.917190551758,-156.7421875,-156.712158203125,-156.747894287109,-156.85986328125,-157.020904541016,-157.290328979492,-157.279495239258,-157.253814697266,-157.249954223633,-157.213623046875],[-157.799362182617,-157.764999389648,-157.720901489258,-157.705520629883,-157.654159545898,-157.635391235352,-157.690872192383,-157.798782348633,-157.849319458008,-157.901763916016,-157.958450317383,-157.968307495117,-157.978424072266,-158.017288208008,-157.98095703125,-158.079147338867,-158.1103515625,-158.137832641602,-158.239120483398,-158.238662719727,-158.273147583008,-158.123107910156,-158.020355224609,-157.962493896484,-157.851516723633,-157.854339599609,-157.82958984375,-157.799362182617],[-160.180023193359,-160.200241088867,-160.234725952148,-160.243453979492,-160.220901489258,-160.163864135742,-160.100631713867,-160.048736572266,-160.076705932617,-160.080032348633,-160.153411865234,-160.180023193359],[-159.372756958008,-159.460693359375,-159.511871337891,-159.608840942383,-159.646377563477,-159.748001098633,-159.789154052734,-159.726608276367,-159.579208374023,-159.35205078125,-159.304794311523,-159.300674438477,-159.330169677734,-159.34375,-159.372756958008],[-81.7838363647461,-81.8092269897461,-81.8114242553711,-81.7676773071289,-81.7386703491211,-81.73974609375,-81.7838363647461],[-81.5666961669922,-81.6315002441406,-81.5792465209961,-81.5623092651367,-81.5316390991211,-81.5322265625,-81.5666961669922],[-81.3348159790039,-81.3647918701172,-81.3790512084961,-81.3790512084961,-81.4216842651367,-81.4200668334961,-81.3223190307617,-81.31982421875,-81.3348159790039],[-81.044189453125,-81.0899963378906,-81.1373519897461,-81.0852508544922,-80.9304656982422,-80.9889144897461,-81.044189453125],[-80.8293914794922,-80.8483428955078,-80.8388671875,-80.7994155883789,-80.7852020263672,-80.7867660522461,-80.8293914794922],[-80.6382751464844,-80.6651382446289,-80.6256866455078,-80.6146011352539,-80.6382751464844],[-80.3818359375,-80.58056640625,-80.5585403442383,-80.4810562133789,-80.4560089111328,-80.4036636352539,-80.3549270629883,-80.3512649536133,-80.2804641723633,-80.257080078125,-80.3818359375],[-82.0372009277344,-82.0728530883789,-82.1449737548828,-82.1843719482422,-82.2013702392578,-82.1385726928711,-82.1160659790039,-82.0372009277344],[-82.0837860107422,-82.085205078125,-82.1355895996094,-82.1691436767578,-82.1211395263672,-82.0837860107422],[-97.1707077026367,-97.1845245361328,-97.267333984375,-97.402099609375,-97.4071807861328,-97.385986328125,-97.3512191772461,-97.2022476196289,-97.1707077026367],[-97.3536148071289,-97.3848190307617,-97.376220703125,-97.2950210571289,-97.1300277709961,-97.060546875,-97.2508773803711,-97.3536148071289],[-80.186767578125,-80.1705093383789,-80.262451171875,-80.3760757446289,-80.4369125366211,-80.395751953125,-80.3555145263672,-80.186767578125],[-97.0143508911133,-97.0360336303711,-96.9876480102539,-96.9786605834961,-96.8993148803711,-96.857421875,-96.8397445678711,-96.9213409423828,-97.0143508911133],[-96.764404296875,-96.8011245727539,-96.755615234375,-96.681640625,-96.5193328857422,-96.453125,-96.4186553955078,-96.403564453125,-96.413330078125,-96.5438995361328,-96.764404296875],[-95.0396957397461,-95.0896453857422,-94.8716812133789,-94.8259735107422,-94.7676239013672,-94.8649444580078,-95.0396957397461],[-91.793701171875,-91.8308639526367,-91.9962387084961,-92.0066452026367,-91.925048828125,-91.875244140625,-91.7964859008789,-91.7676773071289,-91.7542953491211,-91.7619094848633,-91.793701171875],[-84.9079132080078,-85.0082550048828,-85.1167449951172,-85.04931640625,-85.0005340576172,-84.8769989013672,-84.8122100830078,-84.7371597290039,-84.9079132080078],[-88.8893051147461,-88.943603515625,-88.9411087036133,-88.9011764526367,-88.8726577758789,-88.8893051147461],[-88.8274459838867,-88.8556671142578,-88.8279724121094,-88.8668975830078,-88.8258819580078,-88.8125915527344,-88.8274459838867],[-89.2239761352539,-89.220458984375,-89.2694320678711,-89.3419876098633,-89.31005859375,-89.2876510620117,-89.2764663696289,-89.1846694946289,-89.210693359375,-89.2239761352539],[-88.55810546875,-88.5706558227539,-88.6592330932617,-88.7130889892578,-88.7228546142578,-88.573974609375,-88.55810546875],[-88.0713348388672,-88.1593246459961,-88.2897415161133,-88.3162612915039,-88.263916015625,-88.109375,-88.0713348388672],[-81.4189987182617,-81.4634780883789,-81.4827194213867,-81.484619140625,-81.450927734375,-81.4189987182617],[-118.350387573242,-118.408599853516,-118.473197937012,-118.528900146484,-118.590187072754,-118.557083129883,-118.507469177246,-118.383201599121,-118.350387573242],[-119.438041687012,-119.482521057129,-119.543647766113,-119.5751953125,-119.52513885498,-119.478805541992,-119.442047119141,-119.438041687012],[-118.347946166992,-118.297454833984,-118.370208740234,-118.4462890625,-118.469329833984,-118.492034912109,-118.507316589355,-118.559417724609,-118.563323974609,-118.569435119629,-118.554832458496,-118.391700744629,-118.347946166992],[-120.043556213379,-120.113914489746,-120.167137145996,-120.251899719238,-120.071830749512,-119.99439239502,-119.983940124512,-120.043556213379],[-120.306594848633,-120.359718322754,-120.441551208496,-120.412940979004,-120.367729187012,-120.353317260742,-120.306594848633],[-119.882377624512,-119.678863525391,-119.569137573242,-119.549263000488,-119.562202453613,-119.809562683105,-119.88550567627,-119.892425537109,-119.918067932129,-119.882377624512],[-76.5462341308594,-76.5685043334961,-76.6078109741211,-76.6619644165039,-76.6739273071289,-76.6222610473633,-76.5462341308594],[-76.503662109375,-76.528564453125,-76.43701171875,-76.2562026977539,-76.2073745727539,-76.3577194213867,-76.503662109375],[-75.7819290161133,-75.9636688232422,-75.9841842651367,-75.8649368286133,-75.7819290161133],[-75.5441360473633,-75.6782684326172,-75.6900863647461,-75.536376953125,-75.4878921508789,-75.4812469482422,-75.5042953491211,-75.5035171508789,-75.478515625,-75.4564437866211,-75.4647445678711,-75.5093231201172,-75.5441360473633],[-75.6356887817383,-75.6507797241211,-75.7171859741211,-75.6488800048828,-75.6366653442383,-75.6356887817383],[-75.3330612182617,-75.3785171508789,-75.2259750366211,-75.1374053955078,-75.097900390625,-75.13623046875,-75.2032241821289,-75.3330612182617],[-74.1332015991211,-74.25048828125,-74.253173828125,-74.1067428588867,-74.1332015991211],[-74.1881332397461,-74.2358932495117,-74.1881866455078,-74.1004867553711,-74.0687484741211,-74.0673828125,-74.0796890258789,-74.1385269165039,-74.1881332397461],[-72.509765625,-72.5808639526367,-72.5166015625,-72.4613265991211,-72.4089889526367,-72.2874526977539,-72.1838836669922,-72.1512680053711,-72.1018981933594,-72.0039596557617,-71.9032287597656,-72.3389663696289,-72.4280776977539,-72.5555648803711,-72.6760711669922,-72.7628402709961,-73.1942901611328,-73.228515625,-73.2655258178711,-73.6209030151367,-73.7667465209961,-73.8995590209961,-73.8013229370117,-73.7991714477539,-73.8226547241211,-73.8751983642578,-73.9290008544922,-74.014892578125,-74.0320281982422,-74.0033721923828,-73.9645462036133,-73.8792495727539,-73.7572250366211,-73.6952133178711,-73.6522445678711,-73.642822265625,-73.6097717285156,-73.5738296508789,-73.4874038696289,-73.4408721923828,-73.4072265625,-73.3727035522461,-73.2781753540039,-73.1858367919922,-73.1112823486328,-73.0337905883789,-72.8288116455078,-72.6250915527344,-72.5436477661133,-72.37255859375,-72.2741165161133,-72.4273910522461,-72.509765625],[-69.9779281616211,-70.0550765991211,-70.2330627441406,-70.0866165161133,-70.0626907348633,-70.0436019897461,-70.0412139892578,-69.985595703125,-69.9779281616211],[-70.5099105834961,-70.7853012084961,-70.8292007446289,-70.760498046875,-70.6737289428711,-70.6160125732422,-70.5253448486328,-70.5099105834961],[-71.3653335571289,-71.39306640625,-71.4034194946289,-71.3839874267578,-71.3643112182617,-71.3544921875,-71.3653335571289],[-71.2414093017578,-71.2909164428711,-71.3462371826172,-71.3181610107422,-71.3074722290039,-71.2801742553711,-71.2644500732422,-71.2320327758789,-71.2414093017578],[-68.6231918334961,-68.6611785888672,-68.7017135620117,-68.7030258178711,-68.6907653808594,-68.6767578125,-68.6559524536133,-68.6231918334961],[-68.187255859375,-68.2454605102539,-68.3092727661133,-68.3079605102539,-68.3150863647461,-68.3857955932617,-68.4117202758789,-68.4094772338867,-68.3470153808594,-68.2994155883789,-68.238037109375,-68.19091796875,-68.187255859375],[-122.853080749512,-122.862602233887,-122.876754760742,-122.907958984375,-122.911918640137,-122.885101318359,-122.849174499512,-122.853080749512],[-122.394134521484,-122.398727416992,-122.437110900879,-122.456985473633,-122.458198547363,-122.468559265137,-122.509902954102,-122.5068359375,-122.486480712891,-122.468597412109,-122.442092895508,-122.394134521484],[-122.497268676758,-122.502639770508,-122.557815551758,-122.575927734375,-122.57373046875,-122.560111999512,-122.549751281738,-122.517242431641,-122.507858276367,-122.497268676758],[-122.572761535645,-122.523826599121,-122.502830505371,-122.366744995117,-122.366592407227,-122.383148193359,-122.411430358887,-122.437599182129,-122.461616516113,-122.492286682129,-122.557510375977,-122.591361999512,-122.603179931641,-122.606292724609,-122.622657775879,-122.657272338867,-122.690383911133,-122.741500854492,-122.748733520508,-122.724510192871,-122.668998718262,-122.628616333008,-122.603523254395,-122.572456359863,-122.535545349121,-122.542434692383,-122.692138671875,-122.69702911377,-122.624404907227,-122.597602844238,-122.572761535645],[-122.820899963379,-122.836570739746,-122.890037536621,-122.921630859375,-122.932273864746,-122.912208557129,-122.885498046875,-122.86890411377,-122.861923217773,-122.814598083496,-122.820899963379],[-123.013137817383,-122.986763000488,-123.094436645508,-123.139938354492,-123.153419494629,-123.169578552246,-123.162162780762,-123.114158630371,-123.02416229248,-123.013137817383],[-122.782127380371,-122.768852233887,-122.808990478516,-122.837593078613,-122.883110046387,-122.903022766113,-122.887016296387,-122.892532348633,-122.985641479492,-123.002830505371,-122.976654052734,-122.918022155762,-122.89771270752,-122.782127380371],[-74.7088928222656,-74.6632308959961,-74.4303741455078,-74.0142593383789,-73.59814453125,-73.1820373535156,-72.7659225463867,-72.3497543334961,-71.9336395263672,-71.5175323486328,-71.4190368652344,-71.3272933959961,-71.2016143798828,-71.1346664428711,-71.0602569580078,-70.9999084472656,-70.9601516723633,-70.9262237548828,-70.8979949951172,-70.8650360107422,-70.8368148803711,-70.8377914428711,-70.7991714477539,-70.7533187866211,-70.7109375,-70.6897964477539,-70.692138671875,-70.7074203491211,-70.7022476196289,-70.5963821411133,-70.4666061401367,-70.4210968017578,-70.4078674316406,-70.3334503173828,-70.2962417602539,-70.2871627807617,-70.3064422607422,-70.3044891357422,-70.2789001464844,-70.248291015625,-70.1796875,-70.0671844482422,-70.0382385253906,-70.0077209472656,-69.8717269897461,-69.717529296875,-69.6297836303711,-69.4714889526367,-69.35888671875,-69.3021469116211,-69.2428741455078,-69.1462860107422,-69.0501937866211,-69.0642547607422,-69.048583984375,-69.0031280517578,-68.9372100830078,-68.8874053955078,-68.8287124633789,-68.6685562133789,-68.4803695678711,-68.3768997192383,-68.3580093383789,-68.3108901977539,-68.2354965209961,-68.0967712402344,-67.9348678588867,-67.8067855834961,-67.8028335571289,-67.8003463745117,-67.7977066040039,-67.7957992553711,-67.7925338745117,-67.7899398803711,-67.7864685058594,-67.78466796875,-67.7670364379883,-67.7776336669922,-67.7822799682617,-67.7811508178711,-67.7741241455078,-67.7752914428711,-67.7916946411133,-67.7999038696289,-67.80224609375,-67.78466796875,-67.7553176879883,-67.7306594848633,-67.698974609375,-67.6579055786133,-67.5957489013672,-67.5312042236328,-67.4866180419922,-67.4326705932617,-67.4138641357422,-67.4244155883789,-67.4549331665039,-67.48779296875,-67.49365234375,-67.4772491455078,-67.4537582397461,-67.4279251098633,-67.4385299682617,-67.4619598388672,-67.4725570678711,-67.4525833129883,-67.3998107910156,-67.366943359375,-67.3152847290039,-67.2906723022461,-67.2707061767578,-67.2496109008789,-67.2132339477539,-67.1709976196289,-67.1248550415039,-67.13037109375,-67.1022415161133,-67.0804672241211,-67.1139144897461,-67.1067352294922,-67.0140151977539,-66.991455078125,-66.9870147705078,-67.1912612915039,-67.3640670776367,-67.4578094482422,-67.5560073852539,-67.5990753173828,-67.6529769897461,-67.726806640625,-67.7904815673828,-67.8390655517578,-67.9070281982422,-67.9626998901367,-67.98486328125,-68.0139617919922,-68.056640625,-68.0937042236328,-68.1172866821289,-68.1520538330078,-68.1982421875,-68.2457580566406,-68.2774429321289,-68.3167495727539,-68.3737335205078,-68.4168472290039,-68.4505844116211,-68.4794387817383,-68.5214309692383,-68.5144577026367,-68.5325164794922,-68.5723571777344,-68.6120071411133,-68.7232971191406,-68.8119125366211,-68.7938995361328,-68.7101058959961,-68.7358856201172,-68.7770004272461,-68.794921875,-68.7655258178711,-68.7626953125,-68.8001937866211,-68.8473663330078,-68.9614791870117,-68.9561538696289,-69.0635757446289,-69.068359375,-69.1372604370117,-69.22607421875,-69.3445358276367,-69.4349594116211,-69.4808578491211,-69.520751953125,-69.5415496826172,-69.5566864013672,-69.5899887084961,-69.6239242553711,-69.6367721557617,-69.6528854370117,-69.6991195678711,-69.7298278808594,-69.7620086669922,-69.7722702026367,-69.7953109741211,-69.80322265625,-69.7916030883789,-69.808349609375,-69.84033203125,-69.8725051879883,-69.9255828857422,-69.9743194580078,-69.9745101928711,-69.9652328491211,-70.0623550415039,-70.1788101196289,-70.2692413330078,-70.2378845214844,-70.2025909423828,-70.3596725463867,-70.5207061767578,-70.642333984375,-70.691162109375,-70.7331008911133,-70.7776336669922,-70.8290557861328,-70.80029296875,-70.7813491821289,-70.7356948852539,-70.6968765258789,-70.6548385620117,-70.6239700317383,-70.6041488647461,-70.6129379272461,-70.6614303588867,-70.7518539428711,-70.8311538696289,-70.8708953857422,-70.9304656982422,-71.0461959838867,-70.9967269897461,-70.8179702758789,-70.73828125,-70.61767578125,-70.6452102661133,-70.6561584472656,-70.5489273071289,-70.5146942138672,-70.4266586303711,-70.2954635620117,-70.135009765625,-70.0014114379883,-70.006103515625,-70.0900421142578,-70.1102523803711,-70.1725540161133,-70.1962356567383,-70.2365188598633,-70.2410659790039,-70.2035140991211,-70.1598587036133,-70.1089324951172,-69.9778823852539,-69.9416046142578,-69.933837890625,-69.9486312866211,-69.9867706298828,-70.0595169067383,-70.4046859741211,-70.4813537597656,-70.6571350097656,-70.6680679321289,-70.6553726196289,-70.6664581298828,-70.7011260986328,-70.9742202758789,-71.0797805786133,-71.1685562133789,-71.1884231567383,-71.2042999267578,-71.1487274169922,-71.1783218383789,-71.2710876464844,-71.3107376098633,-71.3306198120117,-71.3591766357422,-71.39013671875,-71.3636779785156,-71.4265594482422,-71.4438018798828,-71.5228500366211,-71.769287109375,-71.929931640625,-72.0738754272461,-72.2652893066406,-72.3710479736328,-72.4793930053711,-72.84716796875,-72.9247055053711,-73.0237350463867,-73.1822738647461,-73.5830078125,-73.67138671875,-73.7790069580078,-73.8512725830078,-73.9106903076172,-73.9472122192383,-73.9871063232422,-73.9485855102539,-73.90673828125,-73.8719711303711,-73.8822250366211,-73.9253463745117,-73.9699249267578,-73.9176712036133,-73.9092330932617,-73.9271926879883,-74.0254898071289,-74.0673370361328,-74.1162643432617,-74.1531219482422,-74.1871566772461,-74.2267150878906,-74.2642059326172,-74.2415084838867,-74.0498504638672,-73.9984436035156,-73.9722671508789,-73.9576187133789,-73.9719696044922,-74.0040054321289,-74.0283203125,-74.0489273071289,-74.0799331665039,-74.083984375,-74.0645980834961,-74.0959930419922,-74.1176300048828,-74.1761245727539,-74.2565460205078,-74.3306121826172,-74.4070281982422,-74.3898468017578,-74.4108428955078,-74.4288101196289,-74.474365234375,-74.5171890258789,-74.5787124633789,-74.6029815673828,-74.6047821044922,-74.6459503173828,-74.7944869995117,-74.9234390258789,-74.9542999267578,-74.9203109741211,-74.8970184326172,-74.9752960205078,-75.0501937866211,-75.1361312866211,-75.2310562133789,-75.3534164428711,-75.5242156982422,-75.5192337036133,-75.5235290527344,-75.4716262817383,-75.421875,-75.3531723022461,-75.15380859375,-75.1038131713867,-75.0741653442383,-75.1729431152344,-75.3208999633789,-75.400634765625,-75.4644012451172,-75.5021438598633,-75.5876007080078,-75.5815887451172,-75.5672836303711,-75.5738754272461,-75.5198211669922,-75.4126434326172,-75.3921890258789,-75.3104019165039,-75.18505859375,-75.0886764526367,-75.083984375,-75.1284713745117,-75.1871109008789,-75.11083984375,-75.0728530883789,-75.035888671875,-75.0387649536133,-75.05126953125,-75.0743637084961,-75.0733871459961,-75.0897445678711,-75.1167449951172,-75.1342239379883,-75.1415100097656,-75.1600112915039,-75.2254409790039,-75.2918014526367,-75.353515625,-75.5963821411133,-75.5871047973633,-75.6315460205078,-75.6988296508789,-75.7668914794922,-75.8120574951172,-75.85400390625,-75.9343719482422,-75.9845199584961,-75.9973602294922,-75.9750518798828,-75.8881378173828,-75.7923812866211,-75.7193374633789,-75.6592788696289,-75.7351608276367,-75.850830078125,-75.8290557861328,-75.7953109741211,-75.8556137084961,-75.8913116455078,-75.9280776977539,-75.8849563598633,-75.8639144897461,-75.8767547607422,-75.8586883544922,-75.8888168334961,-75.937255859375,-75.9673843383789,-75.9857482910156,-76.0066833496094,-76.0203094482422,-76.0512161254883,-76.1165008544922,-76.211669921875,-76.2646484375,-76.2948760986328,-76.26416015625,-76.1983871459961,-76.1129379272461,-76.0009307861328,-76.0169448852539,-76.0569381713867,-76.1750030517578,-76.2129821777344,-76.2783203125,-76.30810546875,-76.3411636352539,-76.3003463745117,-76.2469711303711,-76.1681671142578,-76.1910705566406,-76.2408218383789,-76.3306655883789,-76.32958984375,-76.312744140625,-76.2450180053711,-76.1856918334961,-76.1352005004883,-76.1329574584961,-76.2168426513672,-76.2356948852539,-76.1531295776367,-76.0743637084961,-75.9759750366211,-75.8759765625,-75.938720703125,-76.0031280517578,-75.9547348022461,-75.9134750366211,-75.8729476928711,-75.9704132080078,-75.9589385986328,-76.0062942504883,-76.06298828125,-76.0850524902344,-76.0807113647461,-76.0972671508789,-76.141357421875,-76.2158203125,-76.2230453491211,-76.2476577758789,-76.2568359375,-76.2763671875,-76.330810546875,-76.34716796875,-76.3450698852539,-76.3589782714844,-76.4056625366211,-76.4027862548828,-76.4208984375,-76.5704116821289,-76.5739288330078,-76.4893569946289,-76.4275894165039,-76.4200668334961,-76.4730911254883,-76.5462341308594,-76.5585479736328,-76.518798828125,-76.4937438964844,-76.51953125,-76.5155258178711,-76.5210952758789,-76.536865234375,-76.5013122558594,-76.45849609375,-76.4164123535156,-76.3940963745117,-76.4387741088867,-76.5099182128906,-76.5724105834961,-76.6468734741211,-76.6591796875,-76.6773452758789,-76.6685562133789,-76.6419906616211,-76.4087829589844,-76.36572265625,-76.3329086303711,-76.3411636352539,-76.4019546508789,-76.4543914794922,-76.5936050415039,-76.7691421508789,-76.8681182861328,-76.8677749633789,-76.8897476196289,-76.9502410888672,-76.9883728027344,-77.0011672973633,-77.0767135620117,-77.1559066772461,-77.2325210571289,-77.2416000366211,-77.2208938598633,-77.1349105834961,-77.0539016723633,-77.0181655883789,-77.0303649902344,-77.0456085205078,-77.0918960571289,-77.1646499633789,-77.2603988647461,-77.2837905883789,-77.3136672973633,-77.2732391357422,-77.23193359375,-77.1099090576172,-77.0467758178711,-76.9063491821289,-76.6448745727539,-76.5495071411133,-76.4717712402344,-76.3549270629883,-76.2642593383789,-76.2618179321289,-76.293212890625,-76.3056182861328,-76.3441390991211,-76.4366226196289,-76.4924850463867,-76.7927703857422,-76.82861328125,-76.9399871826172,-77.0706558227539,-77.111083984375,-76.9250946044922,-76.8491668701172,-76.7154312133789,-76.6198272705078,-76.5494613647461,-76.4840850830078,-76.3055725097656,-76.3676300048828,-76.2685546875,-76.25439453125,-76.2634735107422,-76.4009780883789,-76.4054641723633,-76.3931655883789,-76.4539031982422,-76.5383758544922,-76.7577133178711,-76.755859375,-76.7380828857422,-76.6108932495117,-76.4973602294922,-76.401123046875,-76.3269577026367,-76.30078125,-76.2833023071289,-76.3382797241211,-76.40087890625,-76.4620132446289,-76.5068359375,-76.602294921875,-76.6309127807617,-76.7035140991211,-77.0069808959961,-77.2508773803711,-77.22705078125,-77.1961898803711,-77.001953125,-76.9251937866211,-76.7654342651367,-76.671875,-76.6339340209961,-76.504638671875,-76.4878387451172,-76.3995590209961,-76.2442398071289,-76.1439971923828,-75.9994125366211,-75.9663543701172,-75.9415512084961,-75.8904266357422,-75.7578659057617,-75.5586929321289,-75.5341796875,-75.5804672241211,-75.7282180786133,-75.8097686767578,-75.8935546875,-75.9178771972656,-75.9464874267578,-75.96533203125,-75.9734344482422,-75.9597625732422,-75.9927749633789,-75.9784622192383,-75.9248504638672,-75.8666000366211,-75.820068359375,-75.8830032348633,-75.9501953125,-76.0547332763672,-76.1478500366211,-76.1410675048828,-76.1499938964844,-76.2217788696289,-76.2706069946289,-76.2273941040039,-76.3211898803711,-76.3836898803711,-76.42431640625,-76.4788131713867,-76.5593719482422,-76.6789093017578,-76.7176284790039,-76.733642578125,-76.7400360107422,-76.71875,-76.7262191772461,-76.6111297607422,-76.5035171508789,-76.3582992553711,-76.2635803222656,-76.20654296875,-76.0697784423828,-76.06005859375,-76.07568359375,-76.0835952758789,-76.0457000732422,-76.0011749267578,-75.9789047241211,-75.8539123535156,-75.81201171875,-75.772216796875,-75.7588424682617,-75.7447280883789,-75.77392578125,-75.9659652709961,-76.103515625,-76.173828125,-76.2752456665039,-76.3902359008789,-76.4466323852539,-76.489501953125,-76.515625,-76.532470703125,-76.5772018432617,-76.6110382080078,-76.6341247558594,-76.7414016723633,-76.8872528076172,-77.0399932861328,-76.9744644165039,-76.595458984375,-76.5527801513672,-76.512939453125,-76.5659713745117,-76.6075210571289,-76.6133804321289,-76.6280288696289,-76.7791519165039,-76.8610305786133,-77.0702667236328,-76.9749450683594,-76.8986358642578,-76.7449722290039,-76.4567413330078,-76.3622055053711,-76.4397964477539,-76.5168914794922,-76.6180191040039,-76.7070846557617,-76.7332000732422,-76.7966766357422,-76.8958969116211,-77.0495147705078,-77.1338882446289,-77.2517547607422,-77.2962417602539,-77.3583984375,-77.3844757080078,-77.4122543334961,-77.4129409790039,-77.4020538330078,-77.3797836303711,-77.5176773071289,-77.649658203125,-77.6969757080078,-77.750732421875,-77.86083984375,-77.8880386352539,-77.9278335571289,-77.932861328125,-77.926025390625,-77.9532699584961,-77.9705581665039,-78.0133285522461,-78.4058609008789,-78.5776824951172,-78.8414535522461,-78.9203186035156,-79.13818359375,-79.1937942504883,-79.2383804321289,-79.2273483276367,-79.2264633178711,-79.2813491821289,-79.229248046875,-79.2760238647461,-79.419921875,-79.4986801147461,-79.5871124267578,-79.6149368286133,-79.7350082397461,-79.8049850463867,-79.93310546875,-79.8936538696289,-79.9407272338867,-80.0217819213867,-80.12255859375,-80.1803207397461,-80.2296905517578,-80.2683639526367,-80.3628463745117,-80.4609909057617,-80.5722198486328,-80.6341781616211,-80.530029296875,-80.4742660522461,-80.4857406616211,-80.5136184692383,-80.579345703125,-80.6082077026367,-80.6258239746094,-80.647216796875,-80.6777877807617,-80.6830596923828,-80.7093276977539,-80.8025360107422,-80.7978973388672,-80.7653350830078,-80.7338409423828,-80.7020492553711,-80.6942367553711,-80.7580032348633,-80.7908172607422,-80.8492126464844,-80.882080078125,-80.8723602294922,-80.9234390258789,-81.0455551147461,-81.0828628540039,-81.11328125,-81.0955047607422,-81.0650329589844,-81.0661163330078,-81.098388671875,-81.162109375,-81.1979064941406,-81.1865692138672,-81.16552734375,-81.169921875,-81.2423782348633,-81.2593765258789,-81.223388671875,-81.1957015991211,-81.1754379272461,-81.2189025878906,-81.2579116821289,-81.2949676513672,-81.3809585571289,-81.3777389526367,-81.3291549682617,-81.2884750366211,-81.3648910522461,-81.41259765625,-81.4417495727539,-81.4603500366211,-81.4532241821289,-81.4713821411133,-81.5005798339844,-81.5204086303711,-81.5162124633789,-81.5039596557617,-81.4571762084961,-81.3857421875,-81.3371124267578,-81.24951171875,-81.1045379638672,-80.9000015258789,-80.5643081665039,-80.5241165161133,-80.5678253173828,-80.5811538696289,-80.5849609375,-80.5728530883789,-80.5331573486328,-80.4568862915039,-80.4995574951172,-80.6100082397461,-80.6228485107422,-80.60693359375,-80.6328659057617,-80.6539077758789,-80.6654739379883,-80.6935043334961,-80.7317352294922,-80.7290496826172,-80.6884765625,-80.7002410888672,-80.7659149169922,-80.7798843383789,-80.77099609375,-80.8086929321289,-80.8381881713867,-80.8184127807617,-80.7872009277344,-80.7486343383789,-80.6863784790039,-80.6500930786133,-80.2261276245117,-80.1257858276367,-80.0886764526367,-80.050048828125,-80.0413055419922,-80.110595703125,-80.1263732910156,-80.1362838745117,-80.1429214477539,-80.158935546875,-80.2190933227539,-80.3008346557617,-80.3277359008789,-80.366943359375,-80.4846649169922,-80.5576171875,-80.7365264892578,-80.8622055053711,-81.011962890625,-81.1104965209961,-81.1673889160156,-81.15869140625,-81.1360321044922,-81.09765625,-80.9653854370117,-80.9404296875,-80.9803695678711,-81.0568313598633,-81.1133346557617,-81.2271499633789,-81.3450698852539,-81.3649444580078,-81.5682601928711,-81.7154846191406,-81.8114700317383,-81.8665542602539,-81.9314956665039,-81.9589385986328,-81.8955078125,-81.8286666870117,-81.8815460205078,-81.9205551147461,-81.9701690673828,-82.0064010620117,-82.0396041870117,-82.077880859375,-82.0669403076172,-82.0132827758789,-82.095703125,-82.1810989379883,-82.1686019897461,-82.1806640625,-82.2428665161133,-82.2900390625,-82.3540496826172,-82.4413604736328,-82.6204605102539,-82.6553726196289,-82.714599609375,-82.6867218017578,-82.6358413696289,-82.5208511352539,-82.4305191040039,-82.4005355834961,-82.40576171875,-82.4457015991211,-82.4981460571289,-82.5206069946289,-82.57958984375,-82.6359405517578,-82.6752014160156,-82.6337890625,-82.5965805053711,-82.6109848022461,-82.6260299682617,-82.660888671875,-82.71533203125,-82.7428741455078,-82.7752914428711,-82.8075637817383,-82.843505859375,-82.74853515625,-82.66064453125,-82.6505813598633,-82.64404296875,-82.6514663696289,-82.7693405151367,-83.2904739379883,-83.6943817138672,-84.0442428588867,-84.3096694946289,-84.3556137084961,-84.3753433227539,-84.3586883544922,-84.3828125,-84.4540557861328,-84.5499954223633,-84.800537109375,-84.888916015625,-84.9691925048828,-85.029296875,-85.18603515625,-85.3189468383789,-85.3763656616211,-85.413818359375,-85.413818359375,-85.3834533691406,-85.33642578125,-85.3148956298828,-85.3068389892578,-85.3536148071289,-85.5042953491211,-85.67578125,-85.6234817504883,-85.6102523803711,-85.6634292602539,-85.6409683227539,-85.603515625,-85.6758804321289,-85.7408218383789,-85.7429733276367,-85.7558135986328,-85.790771484375,-85.8556594848633,-86.1751480102539,-86.4544448852539,-86.2400894165039,-86.1238327026367,-86.1376953125,-86.1656723022461,-86.2573699951172,-86.3741683959961,-86.4479522705078,-86.5233917236328,-86.6060562133789,-86.6796340942383,-86.9676208496094,-87.201171875,-87.1637191772461,-87.123779296875,-86.9857940673828,-86.9651412963867,-86.99755859375,-87.0338821411133,-87.072021484375,-87.1187973022461,-87.1706008911133,-87.1846694946289,-87.2510757446289,-87.2810516357422,-87.4757843017578,-87.500732421875,-87.4437484741211,-87.4482879638672,-87.5132827758789,-87.6222610473633,-88.0059585571289,-87.9850082397461,-87.9039993286133,-87.790283203125,-87.8132858276367,-87.8571243286133,-87.8976058959961,-87.92431640625,-87.9229965209961,-87.9488754272461,-88.0113220214844,-88.0324249267578,-88.078369140625,-88.1165542602539,-88.1354522705078,-88.2492218017578,-88.3499069213867,-88.6920928955078,-88.8199234008789,-88.8729476928711,-88.9052276611328,-89.0540542602539,-89.2236328125,-89.2635726928711,-89.320556640625,-89.4435043334961,-89.5884780883789,-89.9542465209961,-90.0452117919922,-90.1259765625,-90.2252960205078,-90.3319854736328,-90.4130401611328,-90.2849578857422,-90.1753387451172,-89.9941864013672,-89.89404296875,-89.812255859375,-89.7731475830078,-89.7374496459961,-89.6675262451172,-89.6650390625,-89.7146987915039,-89.7772521972656,-89.815185546875,-89.7437973022461,-89.6316909790039,-89.5895004272461,-89.5633773803711,-89.4944305419922,-89.4007339477539,-89.4140625,-89.4009246826172,-89.3578643798828,-89.36279296875,-89.3544387817383,-89.4554214477539,-89.5306625366211,-89.5908660888672,-89.559326171875,-89.6206512451172,-89.662109375,-89.6829605102539,-89.689208984375,-89.7209014892578,-89.6748046875,-89.580322265625,-89.513671875,-89.2457046508789,-89.1807632446289,-89.1168441772461,-89.0653305053711,-89.0157241821289,-89.0213851928711,-89.1095199584961,-89.1333465576172,-89.155517578125,-89.1952590942383,-89.236083984375,-89.33056640625,-89.3761291503906,-89.353515625,-89.3892059326172,-89.4431610107422,-89.5217819213867,-89.5771484375,-89.6202621459961,-89.6724624633789,-89.7169952392578,-89.7923889160156,-89.79736328125,-89.8182678222656,-89.8772430419922,-90.1590881347656,-90.1607894897461,-90.1412582397461,-90.1007843017578,-90.0523452758789,-90.0527877807617,-90.07373046875,-90.0827178955078,-90.1013717651367,-90.1358413696289,-90.2127914428711,-90.2467269897461,-90.3016128540039,-90.3792037963867,-90.5024948120117,-90.5862274169922,-90.677490234375,-90.7510223388672,-91.0027389526367,-91.2901382446289,-91.28271484375,-91.2375030517578,-91.1507797241211,-91.1553726196289,-91.2439880371094,-91.26025390625,-91.2488250732422,-91.2777328491211,-91.3309631347656,-91.5142059326172,-91.5647888183594,-91.6724624633789,-91.8244171142578,-91.8931655883789,-92.017333984375,-92.0802230834961,-92.135498046875,-92.1139678955078,-92.0588912963867,-92.0840377807617,-92.2608413696289,-92.6712875366211,-92.7913131713867,-92.952392578125,-93.1756820678711,-93.283203125,-93.3884811401367,-93.69482421875,-93.7659149169922,-93.8264694213867,-93.86572265625,-93.8838882446289,-93.8483428955078,-93.8087921142578,-93.7730941772461,-93.76904296875,-93.7939987182617,-93.8414611816406,-93.9462890625,-93.8863830566406,-93.8904800415039,-94.0996627807617,-94.574462890625,-94.7596206665039,-94.7501449584961,-94.5262680053711,-94.6053161621094,-94.7326202392578,-94.7782745361328,-94.724365234375,-94.741943359375,-94.8323287963867,-94.889892578125,-94.9298858642578,-94.9822769165039,-95.0228500366211,-94.9928207397461,-94.9358901977539,-94.8882827758789,-95.018310546875,-95.1390609741211,-95.1521530151367,-95.2734909057617,-95.3876495361328,-95.6558532714844,-95.7323760986328,-95.8534164428711,-96.0204086303711,-96.1805191040039,-96.2345275878906,-96.1322784423828,-96.0110321044922,-96.1150360107422,-96.2753448486328,-96.3734359741211,-96.3741226196289,-96.44873046875,-96.5260238647461,-96.5597152709961,-96.57568359375,-96.6084976196289,-96.6400375366211,-96.524658203125,-96.4754867553711,-96.4210891723633,-96.4888153076172,-96.5617141723633,-96.6763687133789,-96.7735366821289,-96.7945785522461,-96.806884765625,-96.8395004272461,-96.8916015625,-96.9198684692383,-96.9333038330078,-96.9666519165039,-97.0154800415039,-97.0960464477539,-97.156494140625,-97.1550750732422,-97.1412582397461,-97.0343246459961,-97.0730972290039,-97.1714324951172,-97.2515563964844,-97.3741226196289,-97.4043884277344,-97.4314956665039,-97.2887191772461,-97.3804626464844,-97.4391098022461,-97.4797821044922,-97.5238800048828,-97.68212890625,-97.7684555053711,-97.6923828125,-97.485107421875,-97.4745101928711,-97.4756851196289,-97.5165023803711,-97.5546875,-97.5265121459961,-97.4937973022461,-97.4658203125,-97.43505859375,-97.40234375,-97.2139205932617,-97.150390625,-97.1401901245117,-97.146240234375,-97.2817916870117,-97.3386688232422,-97.3497543334961,-97.358154296875,-97.3756332397461,-97.4402847290039,-97.5872573852539,-97.8014144897461,-98.0828170776367,-98.2750473022461,-98.3781280517578,-98.4858932495117,-98.5982894897461,-98.69140625,-98.7652282714844,-98.8731994628906,-99.0152816772461,-99.1077651977539,-99.1720733642578,-99.17236328125,-99.2299270629883,-99.3024368286133,-99.4435501098633,-99.4564971923828,-99.45654296875,-99.4577178955078,-99.4402313232422,-99.4551315307617,-99.4998016357422,-99.5100555419922,-99.48583984375,-99.4842834472656,-99.5053253173828,-99.5953140258789,-99.7542495727539,-99.8896484375,-100.001411437988,-100.111961364746,-100.221290588379,-100.296051025391,-100.336280822754,-100.34814453125,-100.331733703613,-100.39892578125,-100.549705505371,-100.636329650879,-100.658645629883,-100.754592895508,-100.924118041992,-101.016304016113,-101.038619995117,-101.038970947266,-101.303512573242,-101.38037109375,-101.440376281738,-101.50927734375,-101.544624328613,-101.54638671875,-101.568702697754,-101.611625671387,-101.752349853516,-101.990921020508,-102.1630859375,-102.268951416016,-102.343070983887,-102.385643005371,-102.476272583008,-102.614936828613,-102.734176635742,-102.833984375,-102.877830505371,-102.865676879883,-102.891990661621,-102.956840515137,-103.022850036621,-103.089988708496,-103.168312072754,-103.257713317871,-103.422950744629,-103.663963317871,-103.852928161621,-103.98974609375,-104.110595703125,-104.215522766113,-104.312210083008,-104.400634765625,-104.504005432129,-104.622215270996,-104.681350708008,-104.681350708008,-104.835891723633,-104.917869567871,-104.978805541992,-105.09814453125,-105.275833129883,-105.514015197754,-105.812698364258,-106.024070739746,-106.148048400879,-106.255714416504,-106.346977233887,-106.43603515625,-106.445411682129,-106.453224182129,-106.673049926758,-106.892868041992,-107.112693786621,-107.33251953125,-107.552345275879,-107.772216796875,-107.992042541504,-108.211814880371,-108.212493896484,-108.213188171387,-108.213813781738,-108.214447021484,-108.56787109375,-108.921340942383,-109.274757385254,-109.628227233887,-109.981636047363,-110.335105895996,-110.688522338867,-111.0419921875,-111.516212463379,-111.990478515625,-112.464744567871,-112.93896484375,-113.413177490234,-113.887451171875,-114.361717224121,-114.835945129395,-114.787994384766,-114.724761962891,-114.839057922363,-115.125198364258,-115.41136932373,-115.69750213623,-115.983688354492,-116.269821166992,-116.555961608887,-116.842094421387,-117.128280639648,-117.130462646484,-117.137405395508,-117.183738708496,-117.243461608887,-117.270698547363,-117.255752563477,-117.262985229492,-117.318855285645,-117.467437744141,-117.788528442383,-117.952095031738,-118.080513000488,-118.161918640137,-118.26439666748,-118.294189453125,-118.410453796387,-118.392967224121,-118.506202697754,-118.598831176758,-118.832023620605,-119.143745422363,-119.23583984375,-119.267669677734,-119.413673400879,-119.606048583984,-119.713188171387,-119.853317260742,-120.052978515625,-120.169525146484,-120.396484375,-120.481201171875,-120.55982208252,-120.644676208496,-120.62671661377,-120.637603759766,-120.624900817871,-120.663040161133,-120.633598327637,-120.659072875977,-120.707038879395,-120.857368469238,-120.884864807129,-120.860305786133,-120.899604797363,-121.022850036621,-121.137939453125,-121.283836364746,-121.343841552734,-121.433738708496,-121.464996337891,-121.664360046387,-121.877388000488,-121.910163879395,-121.918655395508,-121.835151672363,-121.789993286133,-121.794525146484,-121.807418823242,-121.880668640137,-122.164199829102,-122.394927978516,-122.408447265625,-122.499221801758,-122.500442504883,-122.514205932617,-122.445610046387,-122.384078979492,-122.390281677246,-122.369720458984,-122.297607421875,-122.228668212891,-122.166023254395,-122.119041442871,-122.070503234863,-122.096527099609,-122.124122619629,-122.158058166504,-122.222221374512,-122.295997619629,-122.333442687988,-122.365478515625,-122.385444641113,-122.314262390137,-122.217041015625,-122.086723327637,-121.716842651367,-121.638084411621,-121.572998046875,-121.525344848633,-121.62574005127,-121.682228088379,-121.748626708984,-121.880767822266,-121.934173583984,-121.993118286133,-122.031494140625,-122.153755187988,-122.208297729492,-122.337104797363,-122.393356323242,-122.48388671875,-122.494918823242,-122.466888427734,-122.521339416504,-122.584175109863,-122.680717468262,-122.760398864746,-122.872947692871,-122.931983947754,-122.99878692627,-123.00146484375,-122.968154907227,-122.977584838867,-122.876808166504,-122.908149719238,-122.986518859863,-123.046188354492,-123.121139526367,-123.289749145508,-123.424812316895,-123.701118469238,-123.719535827637,-123.820304870605,-123.777778625488,-123.783493041992,-123.832916259766,-123.884468078613,-124.108497619629,-124.324020385742,-124.356544494629,-124.371681213379,-124.324508666992,-124.28369140625,-124.253898620605,-124.242332458496,-124.250579833984,-124.220024108887,-124.208450317383,-124.190231323242,-124.222511291504,-124.219192504883,-124.199897766113,-124.133102416992,-124.140037536621,-124.068504333496,-124.071929931641,-124.11767578125,-124.163238525391,-124.244621276855,-124.208740234375,-124.21167755127,-124.355270385742,-124.410011291504,-124.420501708984,-124.406158447266,-124.443794250488,-124.539649963379,-124.498580932617,-124.454444885254,-124.346588134766,-124.320602416992,-124.275482177734,-124.196922302246,-124.23314666748,-124.287979125977,-124.239212036133,-124.184371948242,-124.148727416992,-124.130668640137,-124.099166870117,-124.047462463379,-124.0654296875,-124.044525146484,-124.059181213379,-123.948593139648,-123.963081359863,-123.929344177246,-123.961227416992,-123.947113037109,-123.975242614746,-123.989311218262,-123.962936401367,-123.911666870117,-123.673637390137,-123.521636962891,-123.466354370117,-123.402297973633,-123.321578979492,-123.220603942871,-123.251319885254,-123.298683166504,-123.404731750488,-123.464851379395,-123.650344848633,-123.688377380371,-123.895698547363,-123.959762573242,-124.072746276855,-124.045112609863,-124.050193786621,-124.044334411621,-124.016403198242,-123.946144104004,-123.912406921387,-123.889152526855,-123.957717895508,-124.071685791016,-124.11254119873,-123.842872619629,-123.986030578613,-124.04222869873,-124.111724853516,-124.116790771484,-124.139259338379,-124.16357421875,-124.170501708984,-124.198837280273,-124.309280395508,-124.376022338867,-124.460060119629,-124.62109375,-124.6630859375,-124.70166015625,-124.679977416992,-124.709968566895,-124.632621765137,-124.429054260254,-124.175491333008,-124.098785400391,-123.975791931152,-123.29443359375,-123.249900817871,-123.161865234375,-123.124420166016,-123.024223327637,-122.97388458252,-122.908882141113,-122.860885620117,-122.778617858887,-122.767524719238,-122.769088745117,-122.73974609375,-122.679489135742,-122.656646728516,-122.778411865234,-122.8017578125,-122.805366516113,-122.821388244629,-123.050636291504,-123.131057739258,-123.139060974121,-123.136329650879,-123.104202270508,-123.030914306641,-122.922164916992,-122.916893005371,-123.018211364746,-123.066802978516,-123.060157775879,-123.048629760742,-122.982475280762,-122.912887573242,-122.814071655273,-122.757133483887,-122.717872619629,-122.608154296875,-122.587890625,-122.592674255371,-122.585746765137,-122.532814025879,-122.510795593262,-122.52392578125,-122.618408203125,-122.630172729492,-122.613632202148,-122.628273010254,-122.664306640625,-122.675483703613,-122.585845947266,-122.557418823242,-122.553565979004,-122.577880859375,-122.603904724121,-122.648628234863,-122.707710266113,-122.720901489258,-122.767776489258,-122.783302307129,-122.812553405762,-122.828468322754,-122.919525146484,-122.956207275391,-122.987648010254,-123.02759552002,-122.914161682129,-122.811965942383,-122.729881286621,-122.701957702637,-122.627044677734,-122.604148864746,-122.542182922363,-122.511077880859,-122.46484375,-122.420120239258,-122.353813171387,-122.351119995117,-122.375244140625,-122.368354797363,-122.380760192871,-122.410499572754,-122.406784057617,-122.383644104004,-122.381980895996,-122.401809692383,-122.392868041992,-122.33032989502,-122.318458557129,-122.241989135742,-122.261276245117,-122.317489624023,-122.352973937988,-122.388671875,-122.415817260742,-122.424705505371,-122.386619567871,-122.394775390625,-122.494049072266,-122.516998291016,-122.529151916504,-122.520317077637,-122.467041015625,-122.403366088867,-122.408546447754,-122.488433837891,-122.541656494141,-122.582565307617,-122.637794494629,-122.662490844727,-122.668998718262,-122.657272338867,-122.627975463867,-122.542678833008,-122.496780395508,-122.501068115234,-122.514793395996,-122.512748718262,-122.545120239258,-122.562004089355,-122.580177307129,-122.599411010742,-122.653022766113,-122.685935974121,-122.722465515137,-122.788772583008,-122.686370849609,-122.26000213623,-121.833595275879,-121.4072265625,-120.980857849121,-120.554489135742,-120.128067016602,-119.701713562012,-119.275344848633,-118.848922729492,-118.422554016113,-117.996192932129,-117.569778442383,-117.143409729004,-116.71704864502,-116.290626525879,-115.8642578125,-115.437889099121,-115.011520385742,-114.585105895996,-114.158744812012,-113.732376098633,-113.305953979492,-112.879592895508,-112.453224182129,-112.026809692383,-111.600440979004,-111.174072265625,-110.747657775879,-110.3212890625,-109.894920349121,-109.468551635742,-109.042137145996,-108.615768432617,-108.189407348633,-107.762985229492,-107.336624145508,-106.910255432129,-106.483840942383,-106.057472229004,-105.631103515625,-105.204689025879,-104.7783203125,-104.351951599121,-103.925582885742,-103.499168395996,-103.072799682617,-102.646430969238,-102.220016479492,-101.793647766113,-101.367286682129,-100.940872192383,-100.514503479004,-100.088134765625,-99.6617202758789,-99.2353515625,-98.8089828491211,-98.3826217651367,-97.9561996459961,-97.5298385620117,-97.1034698486328,-96.6770553588867,-96.2506866455078,-95.8243179321289,-95.3979034423828,-95.1620635986328,-95.1582489013672,-95.1552734375,-94.9393539428711,-94.8748016357422,-94.8543472290039,-94.8603973388672,-94.8425827026367,-94.803466796875,-94.7127914428711,-94.7125473022461,-94.705078125,-94.6753387451172,-94.6208953857422,-94.4141616821289,-94.05517578125,-93.8516159057617,-93.8035659790039,-93.7077178955078,-93.5642547607422,-93.463623046875,-93.3778762817383,-93.2579650878906,-93.1552200317383,-93.0517120361328,-92.9962463378906,-92.8367156982422,-92.732666015625,-92.583251953125,-92.5005798339844,-92.4608840942383,-92.4145965576172,-92.3484420776367,-92.2986831665039,-92.1717758178711,-92.0051727294922,-91.8583984375,-91.6473159790039,-91.518310546875,-91.38720703125,-91.2206497192383,-91.04345703125,-90.9160690307617,-90.84033203125,-90.7973175048828,-90.744384765625,-90.6070785522461,-90.3201217651367,-90.091796875,-90.0399398803711,-89.99365234375,-89.9010238647461,-89.775390625,-89.5505828857422,-89.4556655883789,-89.273193359375,-89.1856460571289,-89.0625991821289,-88.898681640625,-88.6117630004883,-88.378173828125,-88.16064453125,-87.9874496459961,-87.9205093383789,-87.743896484375,-87.4942398071289,-87.2080078125,-86.9218292236328,-86.6721725463867,-86.4955520629883,-86.4285659790039,-86.2344741821289,-86.0403747558594,-85.8463363647461,-85.6522445678711,-85.4581985473633,-85.2641143798828,-85.070068359375,-84.8759765625,-84.8270492553711,-84.7793960571289,-84.665771484375,-84.561767578125,-84.5015640258789,-84.4404830932617,-84.4017028808594,-84.3367156982422,-84.1921920776367,-84.1494598388672,-84.1251907348633,-84.1231918334961,-84.1281204223633,-84.1504821777344,-84.1151885986328,-84.1077651977539,-84.08837890625,-84.0291976928711,-83.977783203125,-83.9130325317383,-83.76318359375,-83.6692810058594,-83.615966796875,-83.5247573852539,-83.4801330566406,-83.469482421875,-83.5926742553711,-83.3973159790039,-83.1792907714844,-82.9193344116211,-82.7603988647461,-82.5510711669922,-82.5152359008789,-82.4850616455078,-82.4465866088867,-82.4073715209961,-82.3682632446289,-82.3268051147461,-82.28125,-82.2407684326172,-82.1965789794922,-82.1378402709961,-82.1903762817383,-82.3047866821289,-82.408203125,-82.417236328125,-82.4883346557617,-82.5453186035156,-82.6451187133789,-82.7441864013672,-82.8677749633789,-83.0037078857422,-83.0731430053711,-83.1095199584961,-83.149658203125,-83.1419448852539,-83.0299758911133,-82.8662109375,-82.6900405883789,-82.4390640258789,-82.2133255004883,-81.9741668701172,-81.7609329223633,-81.50732421875,-81.2776412963867,-81.0282211303711,-80.6826171875,-80.24755859375,-80.0357437133789,-79.7620162963867,-79.4462432861328,-79.1737289428711,-79.0367126464844,-78.9392547607422,-78.9150924682617,-78.9208450317383,-78.9459991455078,-78.9807662963867,-79.0116729736328,-79.0261688232422,-79.029052734375,-79.0479965209961,-79.0660629272461,-79.0592269897461,-79.0830612182617,-79.171875,-79.0024871826172,-78.8455581665039,-78.7204132080078,-78.458251953125,-78.2147979736328,-77.8792495727539,-77.5965347290039,-77.2667007446289,-77.0733337402344,-76.8199691772461,-76.6964874267578,-76.5861358642578,-76.464599609375,-76.24853515625,-76.185791015625,-76.1511688232422,-76.0202178955078,-75.8759307861328,-75.8193359375,-75.7919464111328,-75.4012756347656,-75.1793899536133,-74.9961395263672,-74.8566360473633,-74.762451171875,-74.7088928222656],[179.451568603516,179.278121948242,178.925872802734,178.7470703125,178.64794921875,178.692184448242,178.908004760742,179.084274291992,179.181732177734,179.294342041016,179.41552734375,179.451568603516],[-176.286712646484,-176.349655151367,-176.396102905273,-176.413711547852,-176.378570556641,-176.280227661133,-176.286712646484],[-176.008987426758,-176.093353271484,-176.204437255859,-176.193649291992,-176.071640014648,-176.008987426758],[-177.879058837891,-177.901260375977,-177.925354003906,-178.058898925781,-178.078460693359,-178.000045776367,-177.977249145508,-177.986389160156,-178.045120239258,-178.153457641602,-178.194519042969,-178.168258666992,-178.116607666016,-177.953796386719,-177.865875244141,-177.799606323242,-177.644470214844,-177.724945068359,-177.770660400391,-177.826950073242,-177.879058837891],[-177.148193359375,-177.177001953125,-177.229888916016,-177.382369995117,-177.474655151367,-177.577575683594,-177.654891967773,-177.670211791992,-177.667633056641,-177.334716796875,-177.257278442383,-177.209762573242,-177.166412353516,-177.131484985352,-177.110061645508,-177.063034057617,-177.079544067383,-177.121383666992,-177.135116577148,-177.148193359375],[-176.593307495117,-176.587936401367,-176.473388671875,-176.437454223633,-176.437347412109,-176.452346801758,-176.469787597656,-176.510986328125,-176.557525634766,-176.770950317383,-176.837112426758,-176.961624145508,-176.874420166016,-176.773635864258,-176.736434936523,-176.7451171875,-176.698348999023,-176.596817016602,-176.549896240234,-176.551605224609,-176.593307495117],[178.575485229492,178.511810302734,178.477737426758,178.475006103516,178.509368896484,178.570617675781,178.607330322266,178.575485229492],[179.727737426758,179.645217895508,179.549606323242,179.497665405273,179.50390625,179.627151489258,179.779983520508,179.727737426758],[-176.021530151367,-176.045059204102,-176.142868041992,-176.177551269531,-176.184524536133,-176.155654907227,-176.077392578125,-176.031204223633,-175.988082885742,-175.975296020508,-176.021530151367],[177.415435791016,177.328506469727,177.260650634766,177.250289916992,177.380661010742,177.478424072266,177.5205078125,177.563766479492,177.636520385742,177.669631958008,177.653030395508,177.595993041992,177.594146728516,177.415435791016],[-173.553314208984,-173.357223510742,-173.11328125,-173.024322509766,-173.022903442383,-173.178863525391,-173.232223510742,-173.368408203125,-173.460983276367,-173.672561645508,-173.835784912109,-173.878952026367,-173.930221557617,-173.989608764648,-173.992477416992,-173.938919067383,-173.794097900391,-173.779006958008,-173.656829833984,-173.553314208984],[-172.464782714844,-172.539123535156,-172.619827270508,-172.582168579102,-172.543655395508,-172.470397949219,-172.383102416992,-172.313629150391,-172.464782714844],[-174.677383422852,-175.2138671875,-175.295562744141,-175.214157104492,-175.11767578125,-174.915908813477,-174.667770385742,-174.474273681641,-174.30615234375,-174.258834838867,-174.406494140625,-174.435546875,-174.365432739258,-174.306884765625,-174.168899536133,-174.045608520508,-174.018356323242,-174.030075073242,-174.054885864258,-174.163223266602,-174.179397583008,-174.120666503906,-174.343551635742,-174.677383422852],[173.722763061523,173.657821655273,173.6162109375,173.40234375,173.424514770508,173.516510009766,173.657608032227,173.776077270508,173.744720458984,173.722763061523],[-170.7333984375,-170.79736328125,-170.816070556641,-170.827041625977,-170.791168212891,-170.682083129883,-170.608062744141,-170.584609985352,-170.586624145508,-170.614013671875,-170.649276733398,-170.692291259766,-170.7333984375],[-169.691940307617,-169.708114624023,-169.722763061523,-169.877334594727,-169.980575561523,-169.991851806641,-169.982559204102,-169.820663452148,-169.7548828125,-169.710983276367,-169.691940307617],[172.811813354492,172.983978271484,173.102142333984,173.251647949219,173.43603515625,173.394714355469,173.348251342773,173.302536010742,173.15869140625,173.080276489258,172.935165405273,172.775588989258,172.721786499023,172.595123291016,172.494827270508,172.677932739258,172.811813354492],[-167.96435546875,-168.270706176758,-168.3701171875,-168.445999145508,-168.505615234375,-168.549026489258,-168.597412109375,-168.698547363281,-168.741012573242,-169.06591796875,-169.088912963867,-169.073089599609,-168.973876953125,-168.9091796875,-168.836090087891,-168.795837402344,-168.783004760742,-168.777786254883,-168.759613037109,-168.689834594727,-168.639007568359,-168.572174072266,-168.436614990234,-168.380416870117,-168.362991333008,-168.397262573242,-168.405319213867,-168.396423339844,-168.357223510742,-168.287689208984,-168.193069458008,-168.073287963867,-167.985687255859,-167.828079223633,-167.8046875,-167.843109130859,-167.865142822266,-167.96435546875],[-166.209777832031,-166.223831176758,-166.249420166016,-166.250732421875,-166.234375,-166.187744140625,-166.154541015625,-166.113723754883,-166.102691650391,-166.138626098633,-166.183731079102,-166.209777832031],[-166.615341186523,-166.572174072266,-166.497451782227,-166.442779541016,-166.400039672852,-166.372314453125,-166.335632324219,-166.230865478516,-166.319000244141,-166.48876953125,-166.545608520508,-166.549224853516,-166.384719848633,-166.338760375977,-166.309478759766,-166.354537963867,-166.444183349609,-166.522018432617,-166.702194213867,-166.770401000977,-166.850982666016,-166.960739135742,-167.153656005859,-167.270797729492,-167.300445556641,-167.337265014648,-167.381301879883,-167.428802490234,-167.479827880859,-167.5224609375,-167.592193603516,-167.628616333008,-167.66943359375,-167.780868530273,-167.808792114258,-167.710098266602,-167.638717651367,-167.530166625977,-167.423538208008,-167.2041015625,-167.136077880859,-167.092330932617,-167.042419433594,-167.015716552734,-166.894149780273,-166.838333129883,-166.818756103516,-166.808990478516,-166.803649902344,-166.741256713867,-166.777252197266,-166.889587402344,-166.972946166992,-167.027252197266,-167.071487426758,-167.105621337891,-167.121139526367,-167.1181640625,-167.090484619141,-167.0380859375,-166.978073120117,-166.848678588867,-166.734024047852,-166.673294067383,-166.627395629883,-166.615341186523],[-165.841552734375,-165.87939453125,-165.909851074219,-165.932922363281,-166.036422729492,-166.056640625,-166.102844238281,-166.105819702148,-166.087738037109,-166.041259765625,-165.966415405273,-165.892883300781,-165.764450073242,-165.704238891602,-165.69287109375,-165.737884521484,-165.841552734375],[-165.561141967773,-165.604827880859,-165.615386962891,-165.620498657227,-165.654144287109,-165.59033203125,-165.550628662109,-165.533782958984,-165.487701416016,-165.441741943359,-165.407867431641,-165.467575073242,-165.561141967773],[-162.554397583008,-162.64111328125,-162.733108520508,-162.811721801758,-162.820556640625,-162.645416259766,-162.607955932617,-162.554397583008],[-162.298141479492,-162.321914672852,-162.390762329102,-162.415771484375,-162.433898925781,-162.293655395508,-162.264602661133,-162.238388061523,-162.233734130859,-162.272567749023,-162.298141479492],[-163.476028442383,-163.378952026367,-163.3369140625,-163.274505615234,-163.187118530273,-163.135055541992,-163.089263916016,-163.083251953125,-163.358093261719,-163.530868530273,-163.5830078125,-164.073287963867,-164.171279907227,-164.234619140625,-164.3466796875,-164.403518676758,-164.463485717773,-164.5908203125,-164.743789672852,-164.823440551758,-164.866165161133,-164.903961181641,-164.903717041016,-164.887649536133,-164.75146484375,-164.706207275391,-164.52978515625,-164.478622436523,-164.42431640625,-164.273681640625,-164.145080566406,-163.867965698242,-163.80712890625,-163.607467651367,-163.553039550781,-163.510894775391,-163.476028442383],[-159.362014770508,-159.394485473633,-159.421340942383,-159.45849609375,-159.4619140625,-159.390426635742,-159.363189697266,-159.362014770508],[-159.51513671875,-159.520401000977,-159.534957885742,-159.561462402344,-159.617721557617,-159.648483276367,-159.635406494141,-159.6396484375,-159.597946166992,-159.588043212891,-159.595260620117,-159.574768066406,-159.545059204102,-159.51513671875],[-131.339736938477,-131.237457275391,-131.232025146484,-131.329544067383,-131.406204223633,-131.445709228516,-131.456100463867,-131.431350708008,-131.481735229492,-131.5400390625,-131.592239379883,-131.595123291016,-131.555999755859,-131.577835083008,-131.578475952148,-131.5654296875,-131.512649536133,-131.404647827148,-131.339736938477],[-159.873001098633,-159.933929443359,-159.953079223633,-159.999404907227,-160.038421630859,-160.169570922852,-160.22705078125,-160.16357421875,-160.153625488281,-160.152389526367,-160.172073364258,-160.133743286133,-160.102203369141,-160.038757324219,-159.981628417969,-159.920471191406,-159.887344360352,-159.871047973633,-159.898239135742,-159.839401245117,-159.854095458984,-159.873001098633],[-132.862258911133,-132.837738037109,-132.812896728516,-132.772323608398,-132.700637817383,-132.648864746094,-132.617233276367,-132.634033203125,-132.64697265625,-132.676651000977,-132.705810546875,-132.807281494141,-132.889587402344,-133.008926391602,-133.075393676758,-133.08056640625,-133.122711181641,-133.204650878906,-133.251174926758,-133.324859619141,-133.41796875,-133.453811645508,-133.429061889648,-133.296585083008,-133.097412109375,-133.067077636719,-132.995742797852,-132.982177734375,-132.945999145508,-132.862258911133],[-160.329299926758,-160.343307495117,-160.480758666992,-160.517471313477,-160.492919921875,-160.3623046875,-160.329299926758],[-160.684906005859,-160.669723510742,-160.638824462891,-160.573974609375,-160.552795410156,-160.552490234375,-160.583160400391,-160.531204223633,-160.482666015625,-160.487548828125,-160.609085083008,-160.701812744141,-160.750625610352,-160.795074462891,-160.825485229492,-160.846527099609,-160.839645385742,-160.789215087891,-160.723922729492,-160.695648193359,-160.672164916992,-160.666366577148,-160.684906005859],[-133.305068969727,-133.283203125,-133.281692504883,-133.426467895508,-133.429092407227,-133.463088989258,-133.493453979492,-133.54736328125,-133.650192260742,-133.635009765625,-133.737106323242,-133.634216308594,-133.566696166992,-133.454772949219,-133.345550537109,-133.305068969727],[-155.566009521484,-155.604873657227,-155.680618286133,-155.723190307617,-155.737350463867,-155.62060546875,-155.593948364258,-155.5732421875,-155.563903808594,-155.566009521484],[-130.979156494141,-131.013916015625,-131.082763671875,-131.187896728516,-131.261871337891,-131.316314697266,-131.366851806641,-131.420715332031,-131.450927734375,-131.42236328125,-131.447570800781,-131.474517822266,-131.521820068359,-131.641311645508,-131.723678588867,-131.762496948242,-131.810989379883,-131.841995239258,-131.846084594727,-131.759475708008,-131.647552490234,-131.624954223633,-131.269241333008,-131.236190795898,-131.120651245117,-130.997802734375,-130.965972900391,-130.965042114258,-130.979156494141],[-133.566116333008,-133.376617431641,-133.202972412109,-133.143707275391,-133.1044921875,-133.081741333008,-133.075439453125,-133.080123901367,-133.101226806641,-133.096633911133,-132.757568359375,-132.597610473633,-132.533782958984,-132.496963500977,-132.43017578125,-132.288864135742,-132.214752197266,-132.172698974609,-132.196350097656,-132.2958984375,-132.511291503906,-132.528854370117,-132.54833984375,-132.581741333008,-132.631301879883,-132.591613769531,-132.417877197266,-132.272018432617,-132.215270996094,-132.160247802734,-132.158401489258,-132.1904296875,-132.214889526367,-132.206680297852,-132.165969848633,-132.005065917969,-131.976409912109,-132.000381469727,-131.977584838867,-131.977935791016,-131.99658203125,-131.997222900391,-131.982711791992,-131.980865478516,-132.021682739258,-132.064743041992,-132.134323120117,-132.189254760742,-132.266311645508,-132.34130859375,-132.370208740234,-132.468643188477,-132.486480712891,-132.549362182617,-132.593841552734,-132.588485717773,-132.626953125,-132.622161865234,-132.665328979492,-132.701766967773,-132.682861328125,-132.704162597656,-132.782318115234,-132.91259765625,-133.060592651367,-133.118560791016,-133.10302734375,-133.030029296875,-132.970809936523,-132.958877563477,-133.082458496094,-133.078414916992,-133.033401489258,-133.089645385742,-133.243759155273,-133.298248291016,-133.342819213867,-133.368988037109,-133.502746582031,-133.553268432617,-133.640472412109,-133.68017578125,-133.664398193359,-133.584091186523,-133.537155151367,-133.446975708008,-133.411727905273,-133.322128295898,-133.308486938477,-133.241500854492,-133.252151489258,-133.289215087891,-133.371246337891,-133.538619995117,-133.684234619141,-133.742538452148,-133.755172729492,-133.599227905273,-133.530853271484,-133.544097900391,-133.594436645508,-133.5986328125,-133.566116333008],[-132.779891967773,-132.830947875977,-132.891448974609,-133.035003662109,-133.037643432617,-133.01708984375,-132.935501098633,-132.902053833008,-132.7060546875,-132.643356323242,-132.629104614258,-132.632278442383,-132.65283203125,-132.657577514648,-132.646575927734,-132.669387817383,-132.779891967773],[-132.112350463867,-132.132965087891,-132.172607421875,-132.210311889648,-132.287307739258,-132.368606567383,-132.406585693359,-132.420608520508,-132.406051635742,-132.451171875,-132.602966308594,-132.659912109375,-132.691360473633,-132.699020385742,-132.675201416016,-132.598724365234,-132.539016723633,-132.505950927734,-132.379837036133,-132.316497802734,-132.205612182617,-132.06689453125,-132.112350463867],[-154.682815551758,-154.751220703125,-154.77392578125,-154.777160644531,-154.760940551758,-154.729339599609,-154.623733520508,-154.517532348633,-154.46337890625,-154.444885253906,-154.511184692383,-154.682815551758],[-154.208633422852,-154.2578125,-154.332122802734,-154.322219848633,-154.216751098633,-154.110412597656,-154.102249145508,-154.107177734375,-154.115966796875,-154.149795532227,-154.208633422852],[-169.755233764648,-169.623916625977,-169.550491333008,-169.485687255859,-169.474319458008,-169.586868286133,-169.632614135742,-169.766174316406,-169.755233764648],[-132.746871948242,-132.757629394531,-132.884719848633,-132.930801391602,-132.948043823242,-132.936233520508,-132.906539916992,-132.870651245117,-132.842529296875,-132.655868530273,-132.598678588867,-132.567977905273,-132.634231567383,-132.714462280273,-132.746871948242],[-133.989593505859,-133.9248046875,-133.830856323242,-133.778121948242,-133.738388061523,-133.767288208008,-133.809036254883,-133.855270385742,-133.883590698242,-133.870468139648,-133.884613037109,-133.938522338867,-133.94970703125,-133.970794677734,-133.993988037109,-134.024017333984,-134.067474365234,-134.122421264648,-134.189605712891,-134.245071411133,-134.195465087891,-134.084365844727,-134.150482177734,-134.290237426758,-134.278366088867,-134.384429931641,-134.390625,-134.373687744141,-134.2744140625,-134.143264770508,-134.051803588867,-134.000579833984,-133.989593505859],[-133.3662109375,-133.299713134766,-133.263519287109,-133.195999145508,-133.07080078125,-132.996246337891,-132.954147338867,-132.950592041016,-132.963333129883,-132.954010009766,-132.959182739258,-132.975875854492,-133.004089355469,-133.034912109375,-133.132369995117,-133.244003295898,-133.328948974609,-133.332427978516,-133.30908203125,-133.239700317383,-133.227249145508,-133.178466796875,-133.156646728516,-133.144241333008,-133.144729614258,-133.158157348633,-133.180801391602,-133.212646484375,-133.382766723633,-133.484176635742,-133.602783203125,-133.63134765625,-133.649261474609,-133.658309936523,-133.688186645508,-133.680953979492,-133.757522583008,-133.823043823242,-133.917282104492,-133.979446411133,-133.962356567383,-133.865966796875,-133.707717895508,-133.3662109375],[-153.007080078125,-153.134231567383,-153.156845092773,-153.235397338867,-153.29541015625,-153.374603271484,-153.354339599609,-153.285202026367,-152.935455322266,-152.908401489258,-152.907760620117,-152.933441162109,-153.007080078125],[-170.160537719727,-170.264007568359,-170.358001708984,-170.385879516602,-170.386627197266,-170.116165161133,-170.160537719727],[-134.969772338867,-134.884857177734,-134.823196411133,-134.768508911133,-134.676849365234,-134.634078979492,-134.620712280273,-134.610549926758,-134.624313354492,-134.651718139648,-134.657089233398,-134.631683349609,-134.630020141602,-134.654006958008,-134.681884765625,-134.750289916992,-134.806442260742,-134.848007202148,-134.950134277344,-134.980560302734,-134.982421875,-134.966659545898,-134.933197021484,-134.875106811523,-134.883453369141,-134.927581787109,-135.017822265625,-135.09716796875,-135.159027099609,-135.146575927734,-135.163131713867,-135.284820556641,-135.330612182617,-135.340621948242,-135.33837890625,-135.315124511719,-135.199600219727,-135.211227416992,-135.267395019531,-135.341354370117,-135.375289916992,-135.454925537109,-135.501953125,-135.608932495117,-135.661865234375,-135.812301635742,-135.781646728516,-135.767730712891,-135.821136474609,-135.82275390625,-135.787109375,-135.680908203125,-135.62451171875,-135.58056640625,-135.569625854492,-135.4873046875,-135.448669433594,-135.346298217773,-135.130661010742,-135.065231323242,-134.969772338867],[-153.240615844727,-153.2685546875,-153.294982910156,-153.350830078125,-153.465026855469,-153.51708984375,-153.520065307617,-153.481063842773,-153.346969604492,-153.2900390625,-153.240615844727],[-152.898040771484,-152.890823364258,-152.850158691406,-152.696243286133,-152.616012573242,-152.511566162109,-152.428756713867,-152.411911010742,-152.419143676758,-152.485397338867,-152.482620239258,-152.411483764648,-152.236526489258,-152.215286254883,-152.216217041016,-152.336669921875,-152.380859375,-152.412200927734,-152.630966186523,-152.831146240234,-152.912155151367,-152.940780639648,-152.997467041016,-152.956848144531,-152.781341552734,-152.719528198242,-152.692520141602,-152.679061889648,-152.714065551758,-152.789108276367,-152.879058837891,-152.990280151367,-153.051620483398,-153.274368286133,-153.443695068359,-153.503570556641,-153.5244140625,-153.588272094727,-153.732574462891,-153.646545410156,-153.633056640625,-153.631439208984,-153.643310546875,-153.757232666016,-153.972702026367,-154.02734375,-154.05078125,-154.070022583008,-154.070846557617,-153.793212890625,-153.80419921875,-153.879730224609,-153.999359130859,-154.083801269531,-154.102981567383,-154.080474853516,-154.025436401367,-154.035049438477,-154.065322875977,-154.134857177734,-154.243743896484,-154.324401855469,-154.376815795898,-154.381103515625,-154.26953125,-154.239212036133,-154.209121704102,-154.190826416016,-154.184326171875,-154.207717895508,-154.260940551758,-154.338973999023,-154.498779296875,-154.5693359375,-154.705963134766,-154.712219238281,-154.673202514648,-154.535293579102,-154.387054443359,-154.281448364258,-154.179351806641,-154.116149902344,-154.029830932617,-153.995010375977,-154.015869140625,-154.007904052734,-153.947372436523,-153.881881713867,-153.805419921875,-153.754592895508,-153.687698364258,-153.756927490234,-153.797805786133,-153.818359375,-153.838134765625,-153.799468994141,-153.690124511719,-153.693161010742,-153.808502197266,-153.879455566406,-153.906097412109,-153.904449462891,-153.841552734375,-153.805801391602,-153.768997192383,-153.695602416992,-153.662643432617,-153.568557739258,-153.524459838867,-153.487945556641,-153.454055786133,-153.422698974609,-153.390441894531,-153.357131958008,-153.252395629883,-153.217483520508,-153.200302124023,-153.201019287109,-153.175201416016,-153.168853759766,-153.225921630859,-153.160446166992,-152.943267822266,-152.850387573242,-152.898040771484],[-135.730361938477,-135.587509155273,-135.586273193359,-135.615386962891,-135.693115234375,-135.671142578125,-135.613235473633,-135.572021484375,-135.421188354492,-135.374710083008,-135.346633911133,-135.162841796875,-135.002105712891,-134.954681396484,-134.927978515625,-134.970657348633,-135.102584838867,-135.164749145508,-135.231201171875,-135.338470458984,-135.249557495117,-134.978851318359,-134.896636962891,-134.873092651367,-134.931488037109,-135.084854125977,-135.22021484375,-135.497848510742,-135.564208984375,-135.608535766602,-135.620666503906,-135.617813110352,-135.691940307617,-135.910797119141,-135.996673583984,-136.076614379883,-136.378219604492,-136.459915161133,-136.568603515625,-136.525100708008,-136.512298583984,-136.454391479492,-136.369537353516,-136.321975708008,-136.245697021484,-136.143753051758,-136.142333984375,-136.094390869141,-135.994384765625,-135.947418212891,-135.881744384766,-135.787063598633,-135.730361938477],[-134.312744140625,-134.319869995117,-134.456253051758,-134.593994140625,-134.661575317383,-134.647994995117,-134.519973754883,-134.398880004883,-134.312744140625],[-134.680282592773,-134.426132202148,-134.240081787109,-134.070159912109,-133.965530395508,-133.904098510742,-133.869293212891,-133.82275390625,-133.826904296875,-133.925003051758,-133.995559692383,-134.031646728516,-134.067245483398,-134.104736328125,-134.177536010742,-134.180267333984,-134.212600708008,-134.249954223633,-134.292343139648,-134.306884765625,-134.300384521484,-134.26708984375,-134.083694458008,-133.961135864258,-133.93701171875,-133.920837402344,-133.973739624023,-133.908843994141,-133.9111328125,-133.92529296875,-134.100051879883,-134.260147094727,-134.435302734375,-134.516006469727,-134.554779052734,-134.591506958008,-134.613082885742,-134.619537353516,-134.575866699219,-134.489212036133,-134.486770629883,-134.594818115234,-134.659851074219,-134.695114135742,-134.754104614258,-134.781494140625,-134.820114135742,-134.869964599609,-134.907669067383,-134.93310546875,-134.923492431641,-134.836959838867,-134.733200073242,-134.680282592773],[-152.416946411133,-152.380752563477,-152.343017578125,-152.316268920898,-152.197952270508,-152.125244140625,-152.078521728516,-152.03662109375,-151.997756958008,-151.974365234375,-151.982513427734,-152.068908691406,-152.109085083008,-152.165481567383,-152.1865234375,-152.223571777344,-152.251663208008,-152.268356323242,-152.334381103516,-152.332672119141,-152.305221557617,-152.309234619141,-152.381149291992,-152.451614379883,-152.537643432617,-152.558197021484,-152.571334838867,-152.598236083984,-152.638778686523,-152.683059692383,-152.763854980469,-152.781539916992,-152.840728759766,-152.928421020508,-152.982559204102,-153.305465698242,-153.38134765625,-153.115814208984,-152.976135253906,-152.895355224609,-152.814559936523,-152.771881103516,-152.768692016602,-152.843948364258,-152.841110229492,-152.674652099609,-152.6123046875,-152.543563842773,-152.478469848633,-152.416946411133],[-152.486083984375,-152.515533447266,-152.588623046875,-152.636627197266,-152.604873657227,-152.463180541992,-152.3955078125,-152.367919921875,-152.356842041016,-152.362258911133,-152.392822265625,-152.486083984375],[-160.918991088867,-160.992385864258,-161.070251464844,-161.131500244141,-161.084579467773,-160.986236572266,-160.768615722656,-160.715133666992,-160.918991088867],[-144.565628051758,-144.613571166992,-144.541549682617,-144.444915771484,-144.353958129883,-144.235748291016,-144.248977661133,-144.403213500977,-144.565628051758],[-148.021789550781,-148.074172973633,-148.271881103516,-148.230667114258,-148.07958984375,-147.914215087891,-148.021789550781],[-147.735885620117,-147.846343994141,-147.872467041016,-147.814346313477,-147.76806640625,-147.733642578125,-147.606689453125,-147.4658203125,-147.336517333984,-147.205215454102,-147.180862426758,-147.120025634766,-147.019882202148,-146.957855224609,-146.986724853516,-147.318450927734,-147.346328735352,-147.376510620117,-147.40380859375,-147.447555541992,-147.479400634766,-147.499313354492,-147.540222167969,-147.60205078125,-147.644927978516,-147.668746948242,-147.735885620117],[-166.135452270508,-166.043640136719,-165.994918823242,-165.840911865234,-165.784469604492,-165.729690551758,-165.69580078125,-165.689346313477,-165.714401245117,-165.706939697266,-165.712356567383,-165.630569458008,-165.605026245117,-165.591796875,-165.769287109375,-165.946731567383,-166.099853515625,-166.131195068359,-166.106689453125,-166.148742675781,-166.187545776367,-166.261627197266,-166.342971801758,-166.627639770508,-166.985046386719,-167.138870239258,-167.295120239258,-167.436431884766,-167.344329833984,-167.251708984375,-166.836334228516,-166.784378051758,-166.73095703125,-166.598968505859,-166.475677490234,-166.420364379883,-166.363876342773,-166.246963500977,-166.184967041016,-166.135452270508],[-145.118499755859,-145.150497436523,-145.237640380859,-145.284271240234,-145.128112792969,-145.102432250977,-145.118499755859],[-146.393951416016,-146.371673583984,-146.179550170898,-146.124267578125,-146.102233886719,-146.128280639648,-146.202392578125,-146.419189453125,-146.595306396484,-146.618316650391,-146.650451660156,-146.683013916016,-146.702880859375,-146.702545166016,-146.670272827148,-146.605911254883,-146.560302734375,-146.393951416016],[-147.658248901367,-147.65869140625,-147.690048217773,-147.659957885742,-147.712005615234,-147.732131958008,-147.759918212891,-147.787841796875,-147.815826416016,-147.821685791016,-147.805282592773,-147.871337890625,-147.891448974609,-147.854873657227,-147.841705322266,-147.837600708008,-147.794525146484,-147.779144287109,-147.774169921875,-147.760208129883,-147.7373046875,-147.702972412109,-147.688583374023,-147.658248901367],[-152.020751953125,-152.069046020508,-152.004501342773,-151.959716796875,-151.8994140625,-151.887298583984,-151.986907958984,-152.020751953125],[-172.742233276367,-172.526077270508,-172.387496948242,-172.277526855469,-172.232086181641,-172.397171020508,-172.6357421875,-172.958389282227,-173.074035644531,-173.047653198242,-172.923873901367,-172.860198974609,-172.742233276367],[-147.930709838867,-148.057418823242,-148.115432739258,-148.123779296875,-148.099700927734,-148.101654052734,-148.037734985352,-147.964416503906,-147.943115234375,-147.930709838867],[-171.463027954102,-171.447860717773,-171.343353271484,-171.196929931641,-171.034866333008,-170.874618530273,-170.672500610352,-170.551864624023,-170.430419921875,-170.299362182617,-170.171295166016,-170.121826171875,-170.082412719727,-170.056304931641,-170.017379760742,-169.777435302734,-169.624114990234,-169.587203979492,-169.554534912109,-169.427581787109,-169.295059204102,-169.221099853516,-168.996047973633,-168.716018676758,-168.761322021484,-168.852401733398,-169.109039306641,-169.364700317383,-169.470840454102,-169.559280395508,-169.5712890625,-169.622863769531,-169.676361083984,-169.719818115234,-169.777786254883,-169.818603515625,-169.863433837891,-169.988479614258,-170.115386962891,-170.189605712891,-170.243118286133,-170.272705078125,-170.323532104492,-170.424179077148,-170.527099609375,-170.848388671875,-170.954040527344,-171.061233520508,-171.176025390625,-171.287307739258,-171.401168823242,-171.519134521484,-171.6318359375,-171.737838745117,-171.790969848633,-171.819381713867,-171.817916870117,-171.803512573242,-171.746383666992,-171.646484375,-171.463027954102],[-166.10986328125,-166.148620605469,-166.146484375,-166.032516479492,-165.822219848633,-165.829879760742,-165.942291259766,-166.10986328125],[-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-141.002151489258,-140.762741088867,-140.525451660156,-140.452819824219,-140.196929931641,-139.973297119141,-139.830657958984,-139.676315307617,-139.467971801758,-139.234756469727,-139.079254150391,-139.079254150391,-139.136962890625,-139.185150146484,-139.04345703125,-138.868759155273,-138.705474853516,-138.632278442383,-138.45361328125,-138.317626953125,-138.187438964844,-138.001113891602,-137.870544433594,-137.696624755859,-137.593307495117,-137.543701171875,-137.484176635742,-137.520904541016,-137.438568115234,-137.277542114258,-137.126220703125,-136.939315795898,-136.813278198242,-136.578750610352,-136.466735839844,-136.466354370117,-136.347854614258,-136.277984619141,-136.247131347656,-136.321823120117,-136.09716796875,-135.934677124023,-135.702590942383,-135.475921630859,-135.367874145508,-135.260787963867,-135.051025390625,-135.036666870117,-135.050827026367,-135.0712890625,-134.943740844727,-134.9072265625,-134.802383422852,-134.67724609375,-134.621978759766,-134.440765380859,-134.410202026367,-134.39306640625,-134.363525390625,-134.329650878906,-134.296966552734,-134.218505859375,-134.069198608398,-133.965713500977,-133.820755004883,-133.673919677734,-133.54638671875,-133.401123046875,-133.422561645508,-133.275299072266,-133.120407104492,-133.001419067383,-132.916854858398,-132.815521240234,-132.691513061523,-132.550491333008,-132.442474365234,-132.301666259766,-132.232177734375,-132.279403686523,-132.337982177734,-132.157028198242,-132.031539916992,-132.062896728516,-132.104293823242,-131.962509155273,-131.866165161133,-131.885986328125,-131.833099365234,-131.824264526367,-131.651519775391,-131.575103759766,-131.471878051758,-131.335800170898,-131.199417114258,-131.082916259766,-130.930221557617,-130.74169921875,-130.649078369141,-130.477096557617,-130.413131713867,-130.214706420898,-130.097854614258,-130.055953979492,-130.022903442383,-130.014053344727,-130.025100708008,-130.074661254883,-130.111968994141,-130.137054443359,-130.146530151367,-130.140426635742,-130.120407104492,-130.059478759766,-130.039260864258,-130.036575317383,-130.171813964844,-130.218505859375,-130.214065551758,-130.312545776367,-130.493270874023,-130.575332641602,-130.615829467773,-130.849609375,-130.934616088867,-130.979675292969,-131.0478515625,-131.0458984375,-130.983932495117,-130.750396728516,-130.748199462891,-130.835067749023,-130.85595703125,-130.879791259766,-130.873397827148,-130.879638671875,-130.918548583984,-130.977005004883,-131.127685546875,-131.140380859375,-131.074020385742,-131.032760620117,-131.28759765625,-131.63525390625,-131.7841796875,-131.815475463867,-131.826171875,-131.799072265625,-131.803268432617,-131.833587646484,-131.869430541992,-131.945022583008,-131.9833984375,-132.119003295898,-132.155410766602,-132.223434448242,-132.20751953125,-132.157958984375,-132.090667724609,-132.005706787109,-131.843856811523,-131.738037109375,-131.551376342773,-131.84423828125,-131.887893676758,-131.927291870117,-131.962295532227,-132.021926879883,-132.133239746094,-132.182022094727,-132.255569458008,-132.304992675781,-132.33203125,-132.336669921875,-132.357666015625,-132.434417724609,-132.475936889648,-132.487106323242,-132.639495849609,-132.701950073242,-132.802200317383,-132.829879760742,-132.838821411133,-132.814254760742,-132.824615478516,-132.913436889648,-133.465866088867,-133.436660766602,-133.538970947266,-133.648727416992,-133.626953125,-133.603363037109,-133.55419921875,-133.342346191406,-133.142822265625,-133.117034912109,-133.435729980469,-133.515472412109,-133.535217285156,-133.536422729492,-133.511123657227,-133.212066650391,-133.1943359375,-133.497406005859,-133.559371948242,-133.625732421875,-133.657287597656,-133.722320556641,-133.744140625,-133.821380615234,-133.894485473633,-134.031097412109,-134.056732177734,-134.063339233398,-134.045257568359,-133.933639526367,-133.888519287109,-133.876754760742,-133.9111328125,-133.94384765625,-134.0361328125,-134.131195068359,-134.208831787109,-134.257614135742,-134.331451416016,-134.485458374023,-134.663619995117,-134.776123046875,-134.942535400391,-134.964797973633,-134.986129760742,-135.076477050781,-135.1318359375,-135.217376708984,-135.330322265625,-135.358444213867,-135.348922729492,-135.363662719727,-135.402542114258,-135.412750244141,-135.484085083008,-135.416946411133,-135.400146484375,-135.433746337891,-135.502349853516,-135.386138916016,-135.334075927734,-135.257034301758,-135.207077026367,-135.1845703125,-135.151901245117,-135.06201171875,-135.049713134766,-135.060501098633,-135.090240478516,-135.141555786133,-135.302536010742,-135.363128662109,-135.449951171875,-135.57177734375,-135.873428344727,-135.897552490234,-135.896347045898,-135.861724853516,-135.889556884766,-136.045516967773,-136.043121337891,-135.826370239258,-135.931686401367,-136.0166015625,-136.049362182617,-136.100631713867,-136.133682250977,-136.150054931641,-136.159484863281,-136.12353515625,-136.118408203125,-136.124160766602,-136.146835327148,-136.186325073242,-136.225830078125,-136.299026489258,-136.380279541016,-136.451171875,-136.477584838867,-136.511184692383,-136.566223144531,-136.830947875977,-136.989013671875,-137.002151489258,-136.952835083008,-136.948059082031,-136.987884521484,-137.059036254883,-137.038375854492,-136.963043212891,-136.879104614258,-136.740142822266,-136.613922119141,-136.568206787109,-136.54931640625,-136.533508300781,-136.410110473633,-136.404205322266,-136.483734130859,-136.319885253906,-136.224609375,-136.102890014648,-136.061477661133,-136.055953979492,-136.081253051758,-136.129638671875,-136.46240234375,-136.582626342773,-136.607421875,-136.698928833008,-136.864990234375,-137.071929931641,-137.544006347656,-137.556930541992,-137.564605712891,-137.597076416016,-137.661087036133,-137.75,-137.863723754883,-137.933990478516,-137.960891723633,-138.026901245117,-138.24072265625,-138.352493286133,-138.451324462891,-138.537170410156,-138.560302734375,-138.520706176758,-138.514892578125,-138.704208374023,-138.884323120117,-139.340972900391,-139.576797485352,-139.714462280273,-139.773284912109,-139.799118041992,-139.766067504883,-139.674118041992,-139.611618041992,-139.513031005859,-139.505569458008,-139.558502197266,-139.582183837891,-139.581161499023,-139.569137573242,-139.554092407227,-139.512313842773,-139.483001708984,-139.446884155273,-139.330947875977,-139.314651489258,-139.320007324219,-139.286727905273,-139.276275634766,-139.265625,-139.258743286133,-139.245697021484,-139.220794677734,-139.178863525391,-139.048294067383,-138.988082885742,-139.242477416992,-139.402496337891,-139.431442260742,-139.518951416016,-139.611679077148,-139.850204467773,-139.916885375977,-140.216751098633,-140.419830322266,-140.648391723633,-140.843154907227,-141.331924438477,-141.408309936523,-141.294631958008,-141.289932250977,-141.329544067383,-141.362152099609,-141.408737182617,-141.421676635742,-141.422164916992,-141.409729003906,-141.447067260742,-141.530181884766,-141.670166015625,-142.104110717773,-142.548583984375,-142.945648193359,-143.506103515625,-143.805084228516,-143.9794921875,-144.147216796875,-144.160934448242,-144.084274291992,-144.088516235352,-144.185501098633,-144.332626342773,-144.529983520508,-144.642974853516,-144.671569824219,-144.741409301758,-144.852432250977,-144.901321411133,-144.862457275391,-144.824401855469,-144.786575317383,-144.691116333008,-144.724411010742,-144.863082885742,-144.984024047852,-145.095993041992,-145.162689208984,-145.248291015625,-145.381790161133,-145.563140869141,-145.718444824219,-145.847747802734,-145.898880004883,-145.810653686523,-145.759811401367,-145.690231323242,-145.674896240234,-146.149032592773,-146.166397094727,-146.167083740234,-146.182312011719,-146.251022338867,-146.34716796875,-146.502975463867,-146.570465087891,-146.54638671875,-146.495513916016,-146.391998291016,-146.531921386719,-146.603561401367,-146.638427734375,-146.636032104492,-146.59912109375,-146.284912109375,-146.384368896484,-146.582717895508,-146.715911865234,-146.8740234375,-146.980178833008,-147.034332275391,-147.10595703125,-147.195022583008,-147.2548828125,-147.285598754883,-147.321090698242,-147.361373901367,-147.390579223633,-147.433410644531,-147.523284912109,-147.567291259766,-147.592575073242,-147.623291015625,-147.655654907227,-147.8076171875,-147.89111328125,-147.990768432617,-148.005126953125,-147.97119140625,-147.751861572266,-147.773773193359,-147.844833374023,-147.986373901367,-148.049423217773,-148.157913208008,-148.208694458008,-148.27001953125,-148.341903686523,-148.388763427734,-148.410751342773,-148.395843505859,-148.287399291992,-148.225875854492,-148.208694458008,-148.293167114258,-148.344421386719,-148.393310546875,-148.471038818359,-148.55615234375,-148.557373046875,-148.398681640625,-148.341262817383,-148.267868041992,-148.256744384766,-148.284225463867,-148.304977416992,-148.338424682617,-148.4677734375,-148.509567260742,-148.596618652344,-148.64013671875,-148.624267578125,-148.549118041992,-148.439849853516,-148.29638671875,-148.189453125,-148.119201660156,-148.050689697266,-147.984039306641,-147.964111328125,-147.990966796875,-148.045989990234,-148.129196166992,-148.181686401367,-148.203567504883,-148.215866088867,-148.218658447266,-148.197601318359,-148.213760375977,-148.245025634766,-148.291351318359,-148.333099365234,-148.430709838867,-148.465087890625,-148.506057739258,-148.542373657227,-148.574066162109,-148.643600463867,-148.750885009766,-148.842727661133,-149.004257202148,-149.070114135742,-149.12158203125,-149.2666015625,-149.304931640625,-149.395263671875,-149.414840698242,-149.432235717773,-149.459716796875,-149.549163818359,-149.598037719727,-149.612884521484,-149.629638671875,-149.684677124023,-149.7138671875,-149.794784545898,-149.803649902344,-149.782470703125,-149.80126953125,-149.964981079102,-150.005325317383,-150.015960693359,-149.960144042969,-149.966506958008,-150.198043823242,-150.258499145508,-150.296478271484,-150.338134765625,-150.4853515625,-150.525970458984,-150.58154296875,-150.607376098633,-150.621154785156,-150.622894287109,-150.677444458008,-150.852783203125,-150.899307250977,-150.934524536133,-150.960739135742,-151.063583374023,-151.182769775391,-151.19921875,-151.163040161133,-151.170700073242,-151.222259521484,-151.287506103516,-151.366363525391,-151.477005004883,-151.619384765625,-151.738189697266,-151.903854370117,-151.949508666992,-151.964065551758,-151.931686401367,-151.884628295898,-151.849960327148,-151.692565917969,-151.5126953125,-151.399612426758,-151.262100219727,-151.189407348633,-151.046478271484,-151.057327270508,-151.089447021484,-151.403656005859,-151.450103759766,-151.512603759766,-151.763824462891,-151.816940307617,-151.853225708008,-151.783447265625,-151.734527587891,-151.611862182617,-151.451461791992,-151.39599609375,-151.312698364258,-151.317520141602,-151.355026245117,-151.3564453125,-151.32177734375,-150.953765869141,-150.779479980469,-150.441253662109,-150.34912109375,-150.281494140625,-150.202789306641,-150.113037109375,-149.99755859375,-149.856246948242,-149.632461547852,-149.1728515625,-149.075088500977,-149.0712890625,-149.142242431641,-149.459121704102,-149.592483520508,-149.967727661133,-150.053268432617,-150.0185546875,-149.9267578125,-149.895324707031,-149.882019042969,-149.829208374023,-149.736907958984,-149.595993041992,-149.329055786133,-149.433547973633,-149.625442504883,-149.695266723633,-149.82373046875,-149.873672485352,-149.945220947266,-149.975677490234,-150.108932495117,-150.471771240234,-150.533203125,-150.567230224609,-150.612258911133,-150.945510864258,-151.064987182617,-151.150146484375,-151.281890869141,-151.460113525391,-151.593505859375,-151.733978271484,-151.781631469727,-151.784423828125,-151.75048828125,-151.785110473633,-151.866149902344,-151.996246337891,-152.270706176758,-152.306594848633,-152.305084228516,-152.260299682617,-152.29150390625,-152.368850708008,-152.540924072266,-152.653945922852,-152.727294921875,-152.797897338867,-152.923385620117,-153.025009155273,-153.03125,-152.892929077148,-152.752380371094,-152.664733886719,-152.630126953125,-152.628570556641,-152.660110473633,-152.759475708008,-152.85693359375,-153.106048583984,-153.186370849609,-153.211227416992,-153.040084838867,-153.024612426758,-153.048141479492,-153.093612670898,-153.236175537109,-153.364013671875,-153.383499145508,-153.359619140625,-153.366455078125,-153.414413452148,-153.482620239258,-153.652542114258,-153.670700073242,-153.609375,-153.622268676758,-153.71435546875,-153.752578735352,-153.814163208008,-154.088333129883,-154.067489624023,-154.138824462891,-154.178329467773,-154.129837036133,-153.899551391602,-153.787933349609,-153.656402587891,-153.418258666992,-153.338958740234,-153.327056884766,-153.334426879883,-153.362930297852,-153.437606811523,-153.617340087891,-153.698593139648,-153.821487426758,-153.861968994141,-154.019882202148,-154.062454223633,-154.055709838867,-154.085891723633,-154.289016723633,-154.281784057617,-154.208068847656,-154.235107421875,-154.247024536133,-154.282287597656,-154.409225463867,-154.570602416992,-154.581939697266,-154.584915161133,-155.006881713867,-155.099273681641,-155.147369384766,-155.312744140625,-155.413955688477,-155.529632568359,-155.590240478516,-155.595855712891,-155.628707885742,-155.728958129883,-155.777984619141,-155.813674926758,-156.000198364258,-156.037353515625,-156.055374145508,-156.089889526367,-156.156005859375,-156.2421875,-156.435897827148,-156.478424072266,-156.473678588867,-156.443557739258,-156.397659301758,-156.400482177734,-156.475143432617,-156.501312255859,-156.592041015625,-156.629013061523,-156.712646484375,-156.779891967773,-156.823867797852,-156.871734619141,-156.923431396484,-156.988433837891,-157.066696166992,-157.13916015625,-157.205764770508,-157.270553588867,-157.333602905273,-157.390228271484,-157.440567016602,-157.489654541016,-157.528717041016,-157.578369140625,-157.609771728516,-157.673873901367,-157.770706176758,-157.869079589844,-158.027893066406,-158.078323364258,-157.978271484375,-157.9287109375,-157.929977416992,-157.982177734375,-158.070938110352,-158.124374389648,-158.189407348633,-158.352478027344,-158.414413452148,-158.537399291992,-158.552139282227,-158.536376953125,-158.467330932617,-158.386123657227,-158.343994140625,-158.317001342773,-158.291412353516,-158.275634765625,-158.431838989258,-158.476119995117,-158.504684448242,-158.523330688477,-158.542678833008,-158.554443359375,-158.591156005859,-158.626754760742,-158.704879760742,-158.789855957031,-159.429443359375,-159.523254394531,-159.541305541992,-159.567626953125,-159.610061645508,-159.65966796875,-159.670257568359,-159.665328979492,-159.678512573242,-159.743011474609,-159.771392822266,-159.810409545898,-159.874359130859,-159.913528442383,-159.962310791016,-160.045654296875,-160.243804931641,-160.373199462891,-160.407409667969,-160.462692260742,-160.499328613281,-160.553512573242,-160.625244140625,-160.682907104492,-160.726501464844,-160.770843505859,-160.896728515625,-160.952194213867,-161.024215698242,-161.099502563477,-161.178024291992,-161.381927490234,-161.4638671875,-161.480514526367,-161.476715087891,-161.443801879883,-161.413330078125,-161.372695922852,-161.313278198242,-161.202087402344,-161.214706420898,-161.255126953125,-161.357467651367,-161.458786010742,-161.516937255859,-161.598785400391,-161.654296875,-161.683547973633,-161.720352172852,-161.741561889648,-161.980331420898,-162.073974609375,-162.166595458984,-162.211471557617,-162.274658203125,-162.332916259766,-162.386383056641,-162.427932739258,-162.457473754883,-162.452392578125,-162.412551879883,-162.426803588867,-162.495269775391,-162.541885375977,-162.63037109375,-162.644134521484,-162.614303588867,-162.618896484375,-162.674362182617,-162.819580078125,-162.865036010742,-162.995895385742,-163.11962890625,-163.127822875977,-163.100189208984,-163.131103515625,-163.220550537109,-163.288619995117,-163.335296630859,-163.337890625,-163.296340942383,-163.285705566406,-163.305953979492,-163.303665161133,-163.27880859375,-163.114501953125,-163.045364379883,-163.008255004883,-162.961959838867,-162.906600952148,-162.87158203125,-162.857131958008,-162.786239624023,-162.658981323242,-162.513381958008,-162.349365234375,-162.157135009766,-161.936630249023,-161.697311401367,-161.215621948242,-161.178619384766,-161.222564697266,-161.192535400391,-161.145172119141,-160.968658447266,-160.898635864258,-160.877838134766,-160.902389526367,-161.008407592773,-161.00537109375,-160.851318359375,-160.802825927734,-160.762603759766,-160.745513916016,-160.758392333984,-160.706344604492,-160.599716186523,-160.530227661133,-160.497909545898,-160.436904907227,-160.347305297852,-160.291702270508,-160.270111083984,-160.308502197266,-160.479888916016,-160.527435302734,-160.5390625,-160.514694213867,-160.460845947266,-160.377487182617,-160.302047729492,-160.149261474609,-160.046234130859,-159.785064697266,-159.283096313477,-159.159042358398,-158.990386962891,-158.918014526367,-158.918014526367,-158.894866943359,-158.782073974609,-158.708847045898,-158.675140380859,-158.665924072266,-158.681060791016,-158.684814453125,-158.67724609375,-158.660781860352,-158.585601806641,-158.473739624023,-158.320938110352,-158.224517822266,-158.133544921875,-158.045700073242,-157.894332885742,-157.845748901367,-157.737213134766,-157.697418212891,-157.674026489258,-157.645553588867,-157.535308837891,-157.4619140625,-157.473876953125,-157.533493041992,-157.571624755859,-157.607574462891,-157.6806640625,-157.697204589844,-157.683990478516,-157.621185302734,-157.610885620117,-157.555038452148,-157.442672729492,-157.193710327148,-157.339401245117,-157.393615722656,-157.488388061523,-157.5244140625,-157.523635864258,-157.460891723633,-157.228851318359,-156.974655151367,-157.009033203125,-157.040481567383,-156.923248291016,-156.808883666992,-156.96337890625,-157.142044067383,-157.665725708008,-158.021911621094,-158.19091796875,-158.302581787109,-158.3896484375,-158.439315795898,-158.503173828125,-158.476272583008,-158.425643920898,-158.314498901367,-158.189208984375,-158.080520629883,-158.220611572266,-158.422805786133,-158.514404296875,-158.58447265625,-158.678268432617,-158.760589599609,-158.809463500977,-158.775527954102,-158.837738037109,-158.861373901367,-158.772125244141,-158.788619995117,-158.95068359375,-159.082656860352,-159.358200073242,-159.454193115234,-159.670257568359,-159.741455078125,-159.832229614258,-159.920211791992,-160.152587890625,-160.260787963867,-160.363128662109,-160.519927978516,-160.656631469727,-160.817092895508,-160.924270629883,-161.215911865234,-161.246826171875,-161.287902832031,-161.328125,-161.361328125,-161.755462646484,-162.144927978516,-162.008682250977,-161.856491088867,-161.724365234375,-161.780517578125,-161.790283203125,-161.788665771484,-161.644393920898,-161.794479370117,-161.890762329102,-161.981048583984,-162.023284912109,-161.920104980469,-161.872222900391,-161.831695556641,-161.828704833984,-161.908630371094,-162.138137817383,-162.242477416992,-162.421340942383,-162.287796020508,-162.138870239258,-161.946578979492,-161.962005615234,-162.068252563477,-162.138031005859,-162.199905395508,-162.265029907227,-162.468704223633,-162.599700927734,-162.684951782227,-162.547714233398,-162.526947021484,-162.50048828125,-162.53564453125,-162.570755004883,-162.732620239258,-162.877822875977,-163.219390869141,-163.680374145508,-163.906890869141,-164.142822265625,-164.14111328125,-164.131546020508,-164.470504760742,-164.662261962891,-164.799957275391,-164.919723510742,-165.061141967773,-165.048736572266,-165.026504516602,-165.11328125,-165.224517822266,-165.353805541992,-165.016021728516,-164.899810791016,-164.80517578125,-164.682373046875,-164.512939453125,-164.370071411133,-164.318511962891,-164.265686035156,-164.321380615234,-164.372375488281,-164.309661865234,-164.1318359375,-163.999557495117,-163.936141967773,-163.894927978516,-163.821395874023,-163.72998046875,-163.528717041016,-163.420944213867,-163.511856079102,-163.623046875,-163.906555175781,-163.837310791016,-163.655410766602,-163.5869140625,-163.658935546875,-163.7490234375,-163.99462890625,-164.441558837891,-164.753952026367,-165.065612792969,-165.114837646484,-165.175491333008,-164.999893188477,-164.875595092773,-164.868988037109,-164.941207885742,-165.077087402344,-165.1376953125,-165.127792358398,-165.150054931641,-165.203765869141,-165.27978515625,-165.344879150391,-165.310791015625,-165.243942260742,-165.273635864258,-165.333694458008,-165.392044067383,-165.379302978516,-165.380767822266,-165.48046875,-165.565872192383,-165.627593994141,-165.691360473633,-165.863952636719,-165.906295776367,-165.797119140625,-165.845306396484,-165.961334228516,-166.093994140625,-166.152740478516,-166.163528442383,-166.168121337891,-166.131149291992,-166.100494384766,-165.834579467773,-165.808929443359,-166.019912719727,-166.078811645508,-165.991409301758,-165.833831787109,-165.61279296875,-165.705810546875,-165.725234985352,-165.743942260742,-165.707275390625,-165.447662353516,-165.194534301758,-165.115631103516,-164.999710083008,-164.891845703125,-164.779205322266,-164.757858276367,-164.796096801758,-164.844390869141,-164.68798828125,-164.596282958984,-164.589462280273,-164.68896484375,-164.792663574219,-164.818649291992,-164.845413208008,-164.799667358398,-164.764068603516,-164.677444458008,-164.428131103516,-164.384231567383,-164.375106811523,-164.525207519531,-164.463272094727,-164.409027099609,-164.107620239258,-163.94287109375,-163.736236572266,-163.616302490234,-163.633743286133,-163.66357421875,-163.725723266602,-163.748977661133,-163.737838745117,-163.649368286133,-163.504348754883,-163.423187255859,-163.358840942383,-163.287841796875,-163.062255859375,-162.947692871094,-162.807769775391,-162.621490478516,-162.359817504883,-162.282806396484,-162.193298339844,-162.112503051758,-162.056243896484,-161.973968505859,-161.505416870117,-161.266006469727,-161.099716186523,-160.926712036133,-160.826522827148,-160.778564453125,-160.840484619141,-160.903961181641,-160.987548828125,-161.220108032227,-161.385696411133,-161.49072265625,-161.414596557617,-161.193069458008,-161.048782348633,-160.931930541992,-160.893692016602,-160.836044311523,-160.855911254883,-160.886962890625,-160.967483520508,-161.063232421875,-161.130172729492,-161.186920166016,-161.466354370117,-161.633987426758,-161.759368896484,-161.868316650391,-162.172271728516,-162.334625244141,-162.6357421875,-162.711090087891,-162.807022094727,-162.876419067383,-163.203903198242,-163.302825927734,-163.248291015625,-163.174072265625,-163.0517578125,-163.1044921875,-163.144348144531,-163.267044067383,-163.486175537109,-163.713088989258,-164.303955078125,-164.691848754883,-164.727478027344,-164.764938354492,-164.829544067383,-164.857269287109,-164.899505615234,-164.978759765625,-165.138122558594,-165.446197509766,-166.142761230469,-166.325088500977,-166.481399536133,-166.478134155273,-166.40869140625,-166.415237426758,-166.550888061523,-166.826950073242,-166.928421020508,-166.906402587891,-166.856781005859,-166.762557983398,-166.531005859375,-166.45166015625,-166.279678344727,-166.121490478516,-166.157028198242,-166.197402954102,-166.609375,-166.665390014648,-167.404006958008,-167.987258911133,-168.035018920898,-168.08837890625,-168.009658813477,-167.930572509766,-167.927001953125,-167.914352416992,-167.580032348633,-167.405334472656,-167.07421875,-166.997207641602,-166.894424438477,-166.747665405273,-166.540130615234,-166.398742675781,-166.214599609375,-166.057418823242,-166.008941650391,-165.723678588867,-165.629928588867,-165.589981079102,-165.560211181641,-165.840225219727,-165.811874389648,-165.776184082031,-165.449417114258,-165.198287963867,-165.06396484375,-164.674133300781,-164.460494995117,-164.058242797852,-163.727676391602,-163.638229370117,-163.815719604492,-163.893936157227,-163.838226318359,-163.775497436523,-163.793701171875,-163.902877807617,-163.893936157227,-163.964981079102,-164.033737182617,-163.695358276367,-163.171432495117,-162.886474609375,-162.721771240234,-162.586868286133,-162.214263916016,-161.933685302734,-161.816299438477,-161.556838989258,-161.455413818359,-161.345062255859,-161.201065063477,-161.109237670898,-161.034271240234,-161.069534301758,-161.120315551758,-161.54443359375,-161.828170776367,-161.916885375977,-161.887603759766,-162.191162109375,-162.317718505859,-162.467422485352,-162.543655395508,-162.607421875,-162.478317260742,-162.361618041992,-162.253555297852,-162.131393432617,-162.017639160156,-162.050735473633,-161.909561157227,-161.591018676758,-161.3359375,-161.155807495117,-161.048141479492,-160.784469604492,-160.650543212891,-160.231689453125,-160.227340698242,-160.262542724609,-160.360885620117,-160.643798828125,-160.864013671875,-161.051467895508,-161.398056030273,-161.571731567383,-161.680908203125,-161.856689453125,-161.878768920898,-161.731292724609,-161.622222900391,-161.719924926758,-161.965423583984,-162.391555786133,-162.411575317383,-162.409423828125,-162.583099365234,-162.761428833008,-163.001708984375,-163.531829833984,-163.720565795898,-163.7998046875,-163.942687988281,-164.125198364258,-165.386032104492,-165.959564208984,-166.235931396484,-166.409133911133,-166.574462890625,-166.786285400391,-166.643890380859,-166.5458984375,-166.647842407227,-166.570404052734,-166.447021484375,-166.380508422852,-166.282958984375,-166.182022094727,-166.209091186523,-165.509475708008,-165.0439453125,-164.889694213867,-164.302337646484,-164.150192260742,-163.867919921875,-163.535690307617,-163.250534057617,-163.205169677734,-163.187118530273,-163.161468505859,-163.131011962891,-163.093551635742,-162.952102661133,-162.350387573242,-162.071136474609,-161.977981567383,-161.880966186523,-161.812591552734,-161.779922485352,-161.761077880859,-161.818405151367,-161.911956787109,-162.042388916016,-162.073867797852,-161.997421264648,-161.768173217773,-161.639007568359,-160.996292114258,-160.647659301758,-160.634124755859,-160.117126464844,-160.045608520508,-159.963134765625,-160.106399536133,-160.005569458008,-160.095077514648,-159.907562255859,-159.865676879883,-159.855224609375,-159.857513427734,-159.842620849609,-159.814987182617,-159.683303833008,-159.386764526367,-159.746185302734,-159.961807250977,-160.081588745117,-159.680908203125,-159.314498901367,-159.231735229492,-159.191757202148,-159.183166503906,-159.26220703125,-159.33984375,-159.304153442383,-159.251174926758,-159.075057983398,-158.996292114258,-158.620956420898,-158.510833740234,-158.484375,-157.998489379883,-157.909362792969,-157.605621337891,-157.324768066406,-157.1953125,-156.973327636719,-156.783309936523,-156.47021484375,-156.395263671875,-156.496688842773,-156.567245483398,-156.469970703125,-155.811141967773,-155.645599365234,-155.579437255859,-155.634582519531,-155.804351806641,-156.146575927734,-156.041946411133,-155.973526000977,-155.872207641602,-155.708053588867,-155.579391479492,-155.313385009766,-155.229736328125,-155.166839599609,-154.943801879883,-154.817535400391,-154.673675537109,-154.726318359375,-154.785202026367,-154.5986328125,-154.392196655273,-154.195205688477,-153.918212890625,-153.701370239258,-153.497711181641,-153.23291015625,-152.784912109375,-152.670852661133,-152.4912109375,-152.300384521484,-152.23291015625,-152.437255859375,-152.470596313477,-152.399230957031,-152.269683837891,-152.253372192383,-152.172958374023,-151.76904296875,-151.799896240234,-151.819625854492,-151.944671630859,-151.224807739258,-151.128036499023,-150.979049682617,-150.662643432617,-150.543502807617,-150.403228759766,-150.273635864258,-150.152481079102,-149.8701171875,-149.544036865234,-149.410598754883,-149.269439697266,-148.844787597656,-148.688385009766,-148.479202270508,-148.371154785156,-148.248779296875,-148.142715454102,-148.0390625,-147.869522094727,-147.790588378906,-147.705368041992,-147.062927246094,-146.744873046875,-146.28125,-146.057662963867,-145.823150634766,-145.440093994141,-145.23681640625,-145.197357177734,-144.619186401367,-144.416885375977,-144.064117431641,-143.746444702148,-143.56640625,-143.357040405273,-143.276473999023,-143.218322753906,-142.707855224609,-142.422119140625,-142.296981811523,-141.69921875,-141.5263671875,-141.407913208008,-141.338623046875,-141.289642333984,-141.080810546875,-141.002151489258],[71.7796859741211,71.7529296875,71.7058563232422,71.6689453125,71.6812515258789,71.7365188598633,71.7653274536133,71.7899398803711,71.7796859741211],[71.2061462402344,71.15625,71.0642623901367,71.01171875,71.0436553955078,71.0448303222656,71.041015625,71.0144500732422,70.9744186401367,70.9606475830078,70.9762649536133,71.0054702758789,71.0241165161133,71.0803680419922,71.1302719116211,71.2287063598633,71.1792984008789,71.2156219482422,71.2061462402344],[70.94677734375,70.97900390625,71.0360336303711,71.1299819946289,71.2126922607422,71.2323226928711,71.228515625,71.0322265625,70.9103546142578,70.8566436767578,70.8418960571289,70.8033218383789,70.7277374267578,70.630859375,70.5628890991211,70.4549789428711,70.1809616088867,70.1769561767578,70.2008743286133,70.2900390625,70.4078063964844,70.4713821411133,70.6458969116211,70.6888656616211,70.734375,70.7824172973633,70.8604431152344,70.9626007080078,71.0259780883789,71.1107406616211,71.2234344482422,71.298828125,71.39306640625,71.4084014892578,71.4208984375,71.5,71.5456008911133,71.5855484008789,71.6062469482422,71.61962890625,71.6022491455078,71.6374969482422,71.6649398803711,71.6851577758789,71.697265625,71.70068359375,71.7577133178711,71.79248046875,71.8257827758789,71.8580093383789,71.8786163330078,71.95849609375,72.0524444580078,72.1154327392578,72.1642532348633,72.1809539794922,72.1806640625,72.1873092651367,72.2130813598633,72.294921875,72.3640670776367,72.4273452758789,72.5059585571289,72.6204071044922,72.6583023071289,72.8309555053711,72.8666000366211,72.9259796142578,72.9900360107422,73.1321258544922,73.1369171142578,73.1128921508789,72.7738265991211,72.7488327026367,72.6795959472656,72.6040954589844,72.5674819946289,72.4020462036133,72.3826141357422,72.3690414428711,72.3697280883789,72.4059524536133,72.3892517089844,72.3577117919922,72.2540054321289,72.2346649169922,72.2328186035156,72.19287109375,72.1312484741211,72.0125961303711,71.9710998535156,71.9556655883789,71.9027328491211,71.8454055786133,71.7726593017578,71.6924819946289,71.6668014526367,71.65087890625,71.6298828125,71.5804748535156,71.5204162597656,71.4574279785156,71.3761749267578,71.3046875,71.0945281982422,70.9906234741211,70.9580078125,70.8994140625,70.6531219482422,70.6027374267578,70.5658187866211,70.5335922241211,70.4699249267578,70.3982391357422,70.37158203125,70.3697280883789,70.3771514892578,70.3826217651367,70.548828125,70.6983413696289,70.7120132446289,70.7255859375,70.7510757446289,70.7509765625,70.63916015625,70.634765625,70.6573257446289,70.6573257446289,70.5782241821289,70.4415054321289,70.4019546508789,70.3726577758789,70.3189392089844,70.2920913696289,70.1363296508789,70.0056610107422,69.7732467651367,69.712890625,69.6707992553711,69.62841796875,69.4982376098633,69.4138641357422,69.3572235107422,69.3093719482422,69.31396484375,69.2599639892578,69.2062530517578,69.30419921875,69.29443359375,69.2195358276367,69.27490234375,69.2283172607422,69.1103515625,68.9517593383789,68.6525421142578,68.6306610107422,68.6224594116211,68.6397476196289,68.7845687866211,68.9268569946289,68.9660186767578,68.9720687866211,68.9556655883789,68.9084930419922,68.8046875,68.7927703857422,68.7894515991211,68.8244171142578,68.8638687133789,68.8687515258789,68.8522491455078,68.8324203491211,68.7976531982422,68.77783203125,68.7679672241211,68.7582015991211,68.7352523803711,68.6869125366211,68.6389617919922,68.6103515625,68.5861282348633,68.5069351196289,68.4632797241211,68.3990249633789,68.3030242919922,68.2449188232422,68.0771484375,67.9085922241211,67.7190475463867,67.54248046875,67.49169921875,67.4595718383789,67.4261703491211,67.349609375,67.3576202392578,67.400390625,67.6165008544922,67.6483383178711,67.6672821044922,67.6765594482422,67.6944351196289,67.7685546875,67.8756790161133,67.9595718383789,68.0443420410156,68.103515625,68.1325149536133,68.1485366821289,68.0478515625,68.0559539794922,68.0872039794922,68.1441421508789,68.2513656616211,68.3331069946289,68.3502883911133,68.3544921875,68.3412094116211,68.2940444946289,68.2365188598633,68.1740264892578,68.0875930786133,68.0109405517578,67.8635711669922,67.8143539428711,67.7980499267578,67.7589874267578,67.7529296875,67.7000045776367,67.607421875,67.5464859008789,67.5172882080078,67.4416961669922,67.3197250366211,67.1955108642578,67.06884765625,66.8277282714844,66.5222625732422,66.5106506347656,66.5113296508789,66.5255813598633,66.6292953491211,66.6263656616211,66.6062469482422,66.5745086669922,66.3897476196289,66.3353500366211,66.263671875,66.1731491088867,66.0948257446289,65.97119140625,65.8571243286133,65.7902297973633,65.728515625,65.6708984375,65.6128845214844,65.3996047973633,65.07666015625,64.8207015991211,64.6599578857422,64.6218719482422,64.5316390991211,64.3099594116211,64.1627883911133,63.9525375366211,63.7636756896973,63.7208023071289,63.5060539245605,63.2918930053711,63.0581016540527,62.9068336486816,62.6506805419922,62.5254898071289,62.4832038879395,62.4416046142578,62.375,62.2980461120605,62.1884765625,62.0950202941895,62.017578125,61.9535179138184,61.90283203125,61.7999000549316,61.6445350646973,61.4969749450684,61.4436492919922,61.4173812866211,61.3875007629395,61.3289031982422,61.2423858642578,61.1792945861816,61.1199188232422,60.9332046508789,60.8671913146973,60.7548828125,60.5135765075684,60.4549827575684,60.2000045776367,60.0896492004395,60.0673828125,60.0687484741211,60.1060523986816,60.1379852294922,60.1240234375,60.0755844116211,60.0755844116211,60.1085929870605,60.1763648986816,60.2007827758789,60.1920852661133,60.1555671691895,60.10693359375,59.9626007080078,59.9417953491211,59.9493141174316,59.97412109375,59.9791984558105,59.9816436767578,60.0007820129395,60.0060539245605,59.9851608276367,59.9365234375,59.8583030700684,59.7625961303711,59.4510726928711,59.3542976379395,59.2765617370605,59.1991195678711,59.1595687866211,59.1231422424316,59.0358390808105,58.9308586120605,58.876953125,58.7299842834473,58.5890617370605,58.5323219299316,58.4771499633789,58.3531265258789,58.2596702575684,58.2064476013184,58.1515617370605,58.1620101928711,58.2040977478027,58.2886695861816,58.4181632995605,58.4769515991211,58.4858360290527,58.4744110107422,58.4570350646973,58.4314498901367,58.3970718383789,58.3771476745605,58.3705101013184,58.3272476196289,58.2829132080078,58.2340850830078,58.1656227111816,58.0754852294922,58.0289039611816,57.9834976196289,57.9457054138184,57.9234390258789,57.85595703125,57.8142585754395,57.6861305236816,57.3817367553711,57.2906265258789,57.2288093566895,57.1135749816895,57.03369140625,56.9640617370605,56.98486328125,57.0181617736816,57.07666015625,57.1138687133789,57.1188468933105,57.0948257446289,57.0642547607422,57.0179672241211,56.9658203125,56.86083984375,56.7736320495605,56.4798812866211,56.2419891357422,55.9774398803711,55.9773445129395,55.9771499633789,55.97705078125,55.9769515991211,55.9768562316895,55.9767570495605,55.9766578674316,55.9765625,55.9764633178711,55.9763679504395,55.9762687683105,55.9761734008789,55.97607421875,55.9759750366211,55.9757804870605,55.9756851196289,56.1004905700684,56.2579078674316,56.4091796875,56.5887680053711,56.7918968200684,56.9650382995605,57.1716766357422,57.3292961120605,57.4773483276367,57.6666984558105,57.9610328674316,58.1251983642578,58.2911148071289,58.4494132995605,58.5552749633789,58.6689491271973,58.8070335388184,58.9451217651367,59.0833969116211,59.2214813232422,59.3595695495605,59.4978523254395,59.6359405517578,59.7740249633789,59.9122047424316,60.05029296875,60.1884727478027,60.3266639709473,60.4647483825684,60.6029281616211,60.7411117553711,60.8791961669922,61.0079116821289,61.0653343200684,61.0970726013184,61.1607437133789,61.2714805603027,61.3850631713867,61.52587890625,61.6236343383789,61.7232437133789,61.8875999450684,61.9902381896973,62.0719718933105,62.2378921508789,62.4593734741211,62.6344757080078,62.8461875915527,63.0476608276367,63.2070274353027,63.4448204040527,63.6796836853027,63.84814453125,64.0132827758789,64.2087936401367,64.3181610107422,64.4431610107422,64.49609375,64.6041030883789,64.7060546875,64.8118133544922,64.9054718017578,65.0031204223633,65.0848693847656,65.1708984375,65.2705078125,65.3665008544922,65.4961929321289,65.5707015991211,65.6702117919922,65.7356491088867,65.8030242919922,65.9010696411133,66.0056610107422,66.1002960205078,66.0888671875,66.0785140991211,66.0626983642578,66.0498046875,66.0155258178711,66.01318359375,66.01123046875,66.0095672607422,66.1931686401367,66.3288040161133,66.4986343383789,66.5150375366211,66.5378952026367,66.5725555419922,66.6016616821289,66.6453094482422,66.6686477661133,66.7096633911133,66.7498092651367,66.8142547607422,67.0386657714844,67.2250061035156,67.37158203125,67.5280227661133,67.7350616455078,67.8050765991211,67.86572265625,67.9357376098633,67.9914016723633,68.0197219848633,68.0593719482422,68.1130828857422,68.09033203125,68.0570297241211,68.0476608276367,68.1123046875,68.1602554321289,68.2918930053711,68.4150390625,68.4957046508789,68.5726547241211,68.6006851196289,68.5936508178711,68.5565414428711,68.5592803955078,68.5840835571289,68.6627883911133,68.7371139526367,68.8511657714844,68.9869155883789,69.04345703125,69.06494140625,69.1536178588867,69.2493133544922,69.3683547973633,69.4009780883789,69.5651321411133,69.6638641357422,69.7880859375,69.9599609375,70.0956115722656,70.2258834838867,70.3289108276367,70.416015625,70.4890594482422,70.5401382446289,70.5842742919922,70.61328125,70.6624984741211,70.7152328491211,70.7645492553711,70.8603515625,70.94677734375],[70.6525421142578,70.6492156982422,70.6183547973633,70.5720672607422,70.5500030517578,70.5687484741211,70.6227569580078,70.6525421142578],[12.4391603469849,12.4305658340454,12.4275388717651,12.4305658340454,12.4383792877197,12.4391603469849],[-61.3343734741211,-61.3445320129395,-61.3535118103027,-61.3511238098145,-61.3397445678711,-61.3344230651855,-61.3363265991211,-61.3358345031738,-61.3268051147461,-61.3198738098145,-61.3167953491211,-61.3148422241211,-61.3201217651367,-61.3259239196777,-61.3288078308105,-61.3343734741211],[-61.2262191772461,-61.2422370910645,-61.2347145080566,-61.2422370910645,-61.2472610473633,-61.2552261352539,-61.2653350830078,-61.2769546508789,-61.2769546508789,-61.2620620727539,-61.2393531799316,-61.2392578125,-61.2459449768066,-61.2490234375,-61.2406234741211,-61.2128372192383,-61.201171875,-61.1998062133789,-61.1993141174316,-61.1979484558105,-61.1937484741211,-61.2080574035645,-61.2262191772461],[-61.1745147705078,-61.2039031982422,-61.2772941589355,-61.2684555053711,-61.2240753173828,-61.18212890625,-61.1389617919922,-61.1240272521973,-61.1345252990723,-61.1745147705078],[-60.9978981018066,-61.0599594116211,-61.0691909790039,-61.0504913330078,-60.9447746276855,-60.9158210754395,-60.8945770263672,-60.8998985290527,-60.8491706848145,-60.8614234924316,-60.9164085388184,-60.9978981018066],[-60.8211898803711,-60.94140625,-60.939453125,-60.9072761535645,-60.8448753356934,-60.8213882446289,-60.7815933227539,-60.7588844299316,-60.73583984375,-60.7903785705566,-60.8211898803711],[-65.2125015258789,-65.2664031982422,-65.365234375,-65.4146499633789,-65.3832015991211,-65.3023452758789,-65.2265625,-65.2125015258789],[-63.849365234375,-63.8172836303711,-63.8270988464355,-63.9176292419434,-63.9935531616211,-64.0546875,-64.1011657714844,-64.160888671875,-64.2189483642578,-64.3624954223633,-64.40234375,-64.3486328125,-64.249755859375,-64.2136764526367,-64.184814453125,-64.11279296875,-64.0283203125,-64.00732421875,-63.8931121826172,-63.849365234375],[-60.0175285339355,-59.8316383361816,-59.8289070129395,-59.8490753173828,-59.9648475646973,-59.9907188415527,-60.0324249267578,-60.1781730651855,-60.2789039611816,-60.3467788696289,-60.380615234375,-60.5136260986328,-60.5563468933105,-60.610107421875,-60.6494636535645,-60.7186546325684,-60.7192420959473,-60.6237258911133,-60.6065444946289,-60.6361808776855,-60.6333045959473,-60.5832023620605,-60.523193359375,-60.4649391174316,-60.3923835754395,-60.3450698852539,-60.3254890441895,-60.3220710754395,-60.3520965576172,-60.39501953125,-60.5860862731934,-60.6710472106934,-60.7179222106934,-60.8208465576172,-60.8733367919922,-60.91357421875,-60.9379920959473,-61.007080078125,-61.1047859191895,-61.1456069946289,-61.17724609375,-61.2036094665527,-61.1815910339355,-61.1510238647461,-61.1522941589355,-61.1287117004395,-61.1594734191895,-61.2249488830566,-61.3031272888184,-61.3908195495605,-61.3768043518066,-61.1671867370605,-60.9540023803711,-60.7421340942383,-60.7119598388672,-60.6719779968262,-60.635009765625,-60.6044921875,-60.6038589477539,-60.6275863647461,-60.6791496276855,-60.7417449951172,-60.8334007263184,-60.90625,-60.9664039611816,-61.0028305053711,-61.0362777709961,-61.1024436950684,-61.2094230651855,-61.2800827026367,-61.3675270080566,-61.4793930053711,-61.5542449951172,-61.8208465576172,-62.0815887451172,-62.1531219482422,-62.4106407165527,-62.4725570678711,-62.5439453125,-62.6097640991211,-62.6653327941895,-62.7121124267578,-62.7399406433105,-62.7646026611328,-62.8569793701172,-62.9686546325684,-63.0453109741211,-63.1362342834473,-63.2947273254395,-63.3386726379395,-63.3797836303711,-63.5268058776855,-63.5966300964355,-63.6529312133789,-63.7469749450684,-63.9146499633789,-64.021484375,-64.0733871459961,-64.1217269897461,-64.154296875,-64.1924819946289,-64.2556686401367,-64.5255355834961,-64.5763626098633,-64.6136703491211,-64.66552734375,-64.7222671508789,-64.7886734008789,-64.81787109375,-64.7025833129883,-64.6689987182617,-64.5679244995117,-64.2752914428711,-64.2210922241211,-64.22705078125,-64.228759765625,-64.2188491821289,-64.1435546875,-64.0377883911133,-64.009033203125,-64.0287170410156,-64.048828125,-64.0465850830078,-64.02490234375,-63.9241714477539,-63.7125511169434,-63.5846214294434,-63.3892593383789,-63.3748512268066,-63.3939437866211,-63.4325218200684,-63.4639167785645,-63.5702629089355,-63.68212890625,-63.844482421875,-63.9371604919434,-63.9757804870605,-64.0084991455078,-64.0354461669922,-64.0670394897461,-64.1148452758789,-64.2050247192383,-64.30419921875,-64.4051284790039,-64.4860382080078,-64.5262756347656,-64.5843734741211,-64.6674346923828,-64.7315368652344,-64.8179702758789,-64.9101104736328,-65.0265655517578,-65.103759765625,-65.1696319580078,-65.2639694213867,-65.36083984375,-65.4072265625,-65.473388671875,-65.5560531616211,-65.5626907348633,-65.5229949951172,-65.5660171508789,-65.6446762084961,-65.6814498901367,-65.7181167602539,-65.8113250732422,-65.9258804321289,-65.996337890625,-66.06005859375,-66.1912155151367,-66.3016586303711,-66.3471221923828,-66.4292449951172,-66.6190414428711,-66.8760299682617,-66.8955078125,-66.8844680786133,-66.9311065673828,-66.9583511352539,-66.9815444946289,-66.9881286621094,-67.0438919067383,-67.0895538330078,-67.1314468383789,-67.1138229370117,-67.1314468383789,-67.1654815673828,-67.2152328491211,-67.1976089477539,-67.2108383178711,-67.2527313232422,-67.312255859375,-67.3916015625,-67.4864273071289,-67.5349655151367,-67.5680160522461,-67.5966796875,-67.6187057495117,-67.667236328125,-67.7664489746094,-67.8590850830078,-67.8612289428711,-67.8347702026367,-67.5148391723633,-67.3536148071289,-67.3362808227539,-67.3221664428711,-67.3111343383789,-67.3477096557617,-67.4986801147461,-67.5511245727539,-67.6025390625,-67.66162109375,-67.7323226928711,-67.783203125,-67.7986297607422,-67.79541015625,-67.8143081665039,-67.8552703857422,-67.8552703857422,-67.8143081665039,-67.80419921875,-67.8249053955078,-67.7884216308594,-67.6946258544922,-67.6422882080078,-67.63134765625,-67.5751953125,-67.473876953125,-67.4393539428711,-67.4715805053711,-67.4819793701172,-67.5680694580078,-67.7271423339844,-67.8591766357422,-67.9388656616211,-68.14306640625,-68.4717788696289,-68.7364730834961,-68.9372100830078,-69.0899353027344,-69.1945266723633,-69.2681655883789,-69.3108367919922,-69.3570785522461,-69.4271545410156,-69.4392547607422,-69.5948257446289,-69.7389602661133,-69.9041976928711,-70.0950241088867,-70.1291961669922,-70.1881408691406,-70.26611328125,-70.3874969482422,-70.4706573486328,-70.5355453491211,-70.6550750732422,-70.7371520996094,-70.8106918334961,-71.0132751464844,-71.1286163330078,-71.2178192138672,-71.4571304321289,-71.6208953857422,-71.811279296875,-71.8926696777344,-72.0066375732422,-72.0842819213867,-72.1566848754883,-72.2077178955078,-72.2963333129883,-72.3946304321289,-72.4429702758789,-72.4719772338867,-72.4789581298828,-72.4688949584961,-72.4595718383789,-72.446044921875,-72.3917007446289,-72.3576202392578,-72.3641586303711,-72.3903350830078,-72.4165573120117,-72.5257339477539,-72.6654281616211,-72.7255401611328,-72.79638671875,-72.8521041870117,-72.9044418334961,-72.9601593017578,-73.00927734375,-73.0584030151367,-73.13671875,-73.1931610107422,-73.3367156982422,-73.3662109375,-73.3563461303711,-73.295654296875,-73.2242660522461,-73.1412582397461,-73.0640640258789,-73.0065383911133,-72.9673843383789,-72.9403839111328,-72.8693313598633,-72.7391586303711,-72.6900863647461,-72.572265625,-72.5180206298828,-72.4460983276367,-72.2484817504883,-72.0123062133789,-71.9581069946289,-71.719482421875,-71.5360794067383,-71.4001922607422,-71.3555679321289,-71.3197250366211,-71.3494110107422,-71.41455078125,-71.4883804321289,-71.86865234375,-71.9075241088867,-71.9569320678711,-71.9572219848633,-71.9469757080078,-71.8351058959961,-71.7914581298828,-71.6416015625,-71.6756820678711,-71.7309036254883,-71.6904296875,-71.5984344482422,-71.5943374633789,-71.6648483276367,-71.7935028076172,-71.884765625,-71.9557113647461,-72.1128463745117,-71.9932632446289,-71.9762649536133,-71.873046875,-71.8056640625,-71.7607421875,-71.7813491821289,-71.7401351928711,-71.6867218017578,-71.6195297241211,-71.53662109375,-71.2979507446289,-71.2414093017578,-71.2053680419922,-71.0858459472656,-71.0784149169922,-71.0526885986328,-71.0817413330078,-71.2072296142578,-71.26220703125,-71.3866271972656,-71.4627914428711,-71.4942398071289,-71.5178756713867,-71.5446243286133,-71.4611358642578,-71.4695281982422,-71.2643585205078,-70.8205108642578,-70.5456085205078,-70.2325210571289,-70.1599655151367,-70.0971145629883,-70.0485382080078,-69.8853454589844,-69.8047866821289,-69.7729034423828,-69.8173294067383,-69.9109344482422,-70.1925735473633,-70.2201156616211,-70.2200241088867,-70.2865295410156,-70.2451171875,-70.2027816772461,-70.1220245361328,-70.0039520263672,-69.9143524169922,-69.860107421875,-69.8306198120117,-69.810546875,-69.7624053955078,-69.7119140625,-69.631591796875,-69.56982421875,-69.5257339477539,-69.2325668334961,-69.0545959472656,-68.8279800415039,-68.6162109375,-68.3986358642578,-68.3431701660156,-68.3248062133789,-68.2720718383789,-68.32470703125,-68.2962875366211,-68.2340850830078,-68.1399383544922,-67.8716278076172,-67.5813446044922,-67.13330078125,-66.9890594482422,-66.2472152709961,-66.1058578491211,-66.0921401977539,-66.0904846191406,-65.8517608642578,-65.6558609008789,-65.4893569946289,-65.3173828125,-65.1291046142578,-65.0232925415039,-64.9440383911133,-64.8504867553711,-64.1883316040039,-63.8336906433105,-63.7790489196777,-63.7318878173828,-63.8626937866211,-64.1579055786133,-64.2475128173828,-64.2981948852539,-64.2019577026367,-63.8734359741211,-63.4967765808105,-63.1898918151855,-63.0354957580566,-62.9467315673828,-62.7023468017578,-62.2422828674316,-61.8794937133789,-61.92138671875,-62.0404319763184,-62.2329139709473,-62.3799819946289,-62.6935539245605,-62.91357421875,-62.8429718017578,-62.8430213928223,-62.8129386901855,-62.7812461853027,-62.706298828125,-62.6858367919922,-62.6616249084473,-62.6946754455566,-62.7405738830566,-62.6511726379395,-62.6004867553711,-62.60791015625,-62.6004867553711,-62.5503387451172,-62.51513671875,-62.4009284973145,-62.3204078674316,-62.2998085021973,-62.2806625366211,-62.2567367553711,-62.2211456298828,-62.1904296875,-62.1719741821289,-62.1533699035645,-62.1703147888184,-62.1474609375,-62.1553230285645,-62.11962890625,-62.0770950317383,-62.0165061950684,-61.9085960388184,-61.8372535705566,-61.8311538696289,-61.8053703308105,-61.7587394714355,-61.7359352111816,-61.7317428588867,-61.7591781616211,-61.7659187316895,-61.6253890991211,-61.5888671875,-61.5123062133789,-61.3093757629395,-61.2344207763672,-61.0133781433105,-60.8740730285645,-60.79248046875,-60.8409690856934,-60.9710426330566,-61.0231437683105,-61.0530738830566,-61.0535659790039,-61.0929679870605,-61.0988273620605,-61.1223640441895,-61.1758804321289,-61.2472610473633,-61.6187019348145,-61.5269050598145,-61.4425773620605,-61.3040008544922,-61.1937484741211,-61.0359840393066,-60.8652381896973,-60.8009796142578,-60.4814949035645,-60.4044914245605,-60.3402328491211,-60.1674766540527,-60.0175285339355],[-64.5936508178711,-64.6718215942383,-64.6951141357422,-64.6509780883789,-64.5691375732422,-64.545166015625,-64.5936508178711],[-64.3952178955078,-64.421142578125,-64.4380416870117,-64.4437484741211,-64.4260711669922,-64.3246612548828,-64.3952178955078],[-64.2878952026367,-64.2735824584961,-64.2823257446289,-64.3394546508789,-64.3833999633789,-64.4014663696289,-64.4114227294922,-64.3230972290039,-64.2878952026367],[-64.765625,-64.6818389892578,-64.5804672241211,-64.686279296875,-64.8891143798828,-64.8847122192383,-64.765625],[-64.6598129272461,-64.7259750366211,-64.7705993652344,-64.7876968383789,-64.75244140625,-64.6598129272461],[-64.8450164794922,-64.9199676513672,-65.0236358642578,-64.9420394897461,-64.8891143798828,-64.8450164794922],[106.617477416992,106.589256286621,106.567970275879,106.658592224121,106.649513244629,106.652534484863,106.617477416992],[104.06396484375,104.0830078125,104.075782775879,104.036811828613,104.04833984375,104.018463134766,103.9521484375,103.867973327637,103.849510192871,103.8984375,103.98583984375,104.027732849121,104.06396484375],[107.167678833008,107.083786010742,107.077926635742,107.15087890625,107.176559448242,107.194923400879,107.167678833008],[107.031349182129,106.990036010742,106.91064453125,106.953422546387,107.043746948242,107.064453125,107.06396484375,107.042289733887,107.031349182129],[106.865623474121,106.854103088379,106.803123474121,106.769432067871,106.795310974121,106.855079650879,106.865623474121],[107.521286010742,107.465530395508,107.399223327637,107.478614807129,107.518943786621,107.55126953125,107.551078796387,107.521286010742],[107.602729797363,107.458686828613,107.403518676758,107.452537536621,107.476272583008,107.562690734863,107.602729797363],[107.97265625,107.92578125,107.80908203125,107.707229614258,107.63671875,107.526954650879,107.409965515137,107.376174926758,107.373336791992,107.354293823242,107.164749145508,107.111717224121,107.0751953125,107.019233703613,106.9814453125,106.936424255371,106.88623046875,106.820610046387,106.76025390625,106.725196838379,106.683395385742,106.675483703613,106.7373046875,106.75341796875,106.55078125,106.572845458984,106.517967224121,106.3955078125,106.165718078613,106.062210083008,105.984077453613,105.81396484375,105.812110900879,105.785354614258,105.791114807129,105.716407775879,105.639060974121,105.621780395508,105.732032775879,105.744239807129,105.808204650879,105.839256286621,105.888282775879,106.065628051758,106.14453125,106.239547729492,106.411918640137,106.4990234375,106.459373474121,106.478904724121,106.355857849121,106.370513916016,106.516792297363,106.735740661621,106.926177978516,107.119918823242,107.180374145508,107.355079650879,107.54931640625,107.540824890137,107.593460083008,107.72412109375,107.803123474121,107.833793640137,107.882034301758,107.936325073242,107.99072265625,108.029396057129,108.087989807129,108.169723510742,108.208984375,108.240821838379,108.267387390137,108.274024963379,108.286041259766,108.395317077637,108.447463989258,108.577827453613,108.674217224121,108.742774963379,108.8212890625,108.898239135742,108.93994140625,109.0224609375,109.084861755371,109.087013244629,109.137306213379,109.19140625,109.207321166992,109.223922729492,109.24462890625,109.303321838379,109.2880859375,109.2470703125,109.252052307129,109.2880859375,109.271873474121,109.3095703125,109.376754760742,109.423927307129,109.420021057129,109.444923400879,109.381439208984,109.335540771484,109.274024963379,109.218948364258,109.3046875,109.206832885742,109.215721130371,109.256256103516,109.259185791016,109.247261047363,109.22021484375,109.214546203613,109.199119567871,109.199905395508,109.167282104492,109.157524108887,109.192680358887,109.198631286621,109.173248291016,109.132514953613,109.039649963379,109.018463134766,108.986717224121,108.82080078125,108.700294494629,108.55126953125,108.418556213379,108.271682739258,108.176170349121,108.094924926758,108.001365661621,107.845115661621,107.564453125,107.470314025879,107.384475708008,107.261520385742,107.235054016113,107.194145202637,107.087791442871,107.035743713379,107.020706176758,107.006637573242,106.983688354492,106.966110229492,106.947463989258,106.902053833008,106.812698364258,106.727340698242,106.605857849121,106.64306640625,106.698432922363,106.7412109375,106.777534484863,106.776275634766,106.757423400879,106.6435546875,106.49169921875,106.464065551758,106.602447509766,106.72900390625,106.785255432129,106.785255432129,106.714157104492,106.6591796875,106.658103942871,106.656837463379,106.595603942871,106.557327270508,106.449119567871,106.136428833008,106.18359375,106.507423400879,106.564361572266,106.572456359863,106.539161682129,106.484085083008,106.378021240234,106.204002380371,105.925682067871,105.830963134766,106.112503051758,106.158592224121,106.206146240234,106.192581176758,106.168357849121,105.5009765625,105.4013671875,105.322265625,105.191207885742,105.114356994629,104.891891479492,104.770408630371,104.896293640137,104.818557739258,104.814651489258,104.84521484375,104.903221130371,104.987113952637,105.092575073242,105.094917297363,105.08447265625,105.02783203125,104.9658203125,104.873245239258,104.801948547363,104.747657775879,104.663475036621,104.612693786621,104.594039916992,104.51611328125,104.426368713379,104.466995239258,104.514068603516,104.564262390137,104.689651489258,104.8154296875,104.8505859375,104.901268005371,104.98388671875,105.04638671875,105.061126708984,105.0361328125,105.022262573242,105.045700073242,105.159469604492,105.284278869629,105.314643859863,105.386528015137,105.40576171875,105.452728271484,105.576560974121,105.69775390625,105.755073547363,105.810745239258,105.853324890137,105.875190734863,105.938179016113,105.990135192871,106.098823547363,106.163963317871,106.131546020508,106.16796875,106.160942077637,106.099510192871,105.8916015625,105.856056213379,105.860939025879,105.85400390625,105.835350036621,105.838470458984,105.851463317871,105.889846801758,105.926559448242,105.956253051758,106.006050109863,106.102928161621,106.239158630371,106.33984375,106.399215698242,106.412506103516,106.410743713379,106.417778015137,106.413871765137,106.499610900879,106.630958557129,106.700096130371,106.7646484375,106.9306640625,107.050682067871,107.158981323242,107.212104797363,107.279685974121,107.330078125,107.393356323242,107.445991516113,107.506446838379,107.5380859375,107.555465698242,107.543556213379,107.511520385742,107.481544494629,107.475387573242,107.545509338379,107.60546875,107.593948364258,107.528610229492,107.462303161621,107.389450073242,107.362113952637,107.342575073242,107.331443786621,107.3603515625,107.364456176758,107.448440551758,107.4931640625,107.535255432129,107.519432067871,107.513771057129,107.524513244629,107.504684448242,107.480369567871,107.496292114258,107.555274963379,107.589645385742,107.633689880371,107.653121948242,107.621681213379,107.564254760742,107.459571838379,107.338775634766,107.279396057129,107.232620239258,107.189552307129,107.165924072266,107.188873291016,107.360641479492,107.391990661621,107.41015625,107.396385192871,107.35009765625,107.296485900879,107.217384338379,107.069725036621,107.001953125,106.9306640625,106.892776489258,106.85107421875,106.832420349121,106.791595458984,106.739555358887,106.696090698242,106.656440734863,106.637504577637,106.593650817871,106.546188354492,106.53369140625,106.525978088379,106.502243041992,106.46533203125,106.425979614258,106.333396911621,106.26953125,106.006248474121,105.973533630371,105.902732849121,105.779495239258,105.69140625,105.627243041992,105.59765625,105.588478088379,105.5185546875,105.458206176758,105.400001525879,105.33349609375,105.273239135742,105.163284301758,105.114547729492,105.085838317871,105.087005615234,105.113479614258,105.145408630371,105.146484375,105.115135192871,104.9931640625,104.716506958008,104.61328125,104.517967224121,104.44580078125,104.108589172363,104.00634765625,103.918357849121,103.8916015625,103.896385192871,103.932029724121,104.027534484863,104.062889099121,104.051559448242,104.013473510742,104.032028198242,104.062797546387,104.127151489258,104.259864807129,104.546287536621,104.587890625,104.7431640625,104.8017578125,104.815132141113,104.845802307129,104.927925109863,104.92919921875,104.888671875,104.847846984863,104.812698364258,104.69873046875,104.676948547363,104.661911010742,104.656448364258,104.618850708008,104.496192932129,104.392189025879,104.367774963379,104.407814025879,104.478607177734,104.53271484375,104.5751953125,104.583198547363,104.530372619629,104.46142578125,104.349609375,104.1953125,104.101364135742,104.052047729492,103.882034301758,103.79052734375,103.714454650879,103.635063171387,103.5546875,103.463577270508,103.210739135742,103.1044921875,102.8837890625,102.851173400879,102.872268676758,102.887496948242,102.909568786621,102.948631286621,102.959182739258,102.949607849121,102.917671203613,102.876174926758,102.84521484375,102.81591796875,102.798240661621,102.771095275879,102.738571166992,102.6953125,102.662010192871,102.640815734863,102.631248474121,102.609664916992,102.58251953125,102.487503051758,102.442672729492,102.301361083984,102.183013916016,102.12744140625,102.175979614258,102.237007141113,102.30224609375,102.375778198242,102.406448364258,102.427925109863,102.470893859863,102.517181396484,102.598533630371,102.721000671387,102.830078125,102.874221801758,102.935157775879,102.98193359375,103.00537109375,103.075881958008,103.136329650879,103.137596130371,103.193359375,103.266304016113,103.300590515137,103.32666015625,103.356056213379,103.470993041992,103.492965698242,103.525390625,103.570701599121,103.620208740234,103.637298583984,103.9150390625,103.941505432129,103.971389770508,103.990821838379,104.0126953125,104.053909301758,104.14306640625,104.212501525879,104.23828125,104.29833984375,104.371780395508,104.52685546875,104.577537536621,104.631736755371,104.687309265137,104.740036010742,104.795700073242,104.814743041992,104.826560974121,104.86474609375,104.91015625,104.995704650879,105.189064025879,105.23876953125,105.275390625,105.350494384766,105.440139770508,105.494529724121,105.530860900879,105.548141479492,105.691207885742,105.782325744629,105.842964172363,105.902633666992,105.962303161621,106.0009765625,106.068458557129,106.1484375,106.183982849121,106.249412536621,106.279006958008,106.338088989258,106.450881958008,106.541801452637,106.6240234375,106.7802734375,106.736328125,106.701560974121,106.633102416992,106.582427978516,106.550392150879,106.536331176758,106.553611755371,106.593162536621,106.636528015137,106.654197692871,106.660057067871,106.65771484375,106.66357421875,106.697654724121,106.7294921875,106.794143676758,106.87451171875,106.925201416016,106.970993041992,107.006439208984,107.019821166992,107.061622619629,107.178512573242,107.272071838379,107.351165771484,107.433494567871,107.471389770508,107.641014099121,107.75927734375,107.802047729492,107.908393859863,107.97265625],[169.896286010742,169.861129760742,169.807022094727,169.737503051758,169.750671386719,169.829498291016,169.852340698242,169.896286010742],[169.491302490234,169.4384765625,169.347259521484,169.261901855469,169.217483520508,169.247451782227,169.291122436523,169.336715698242,169.359970092773,169.491302490234],[169.334381103516,169.288284301758,169.248046875,168.986907958984,168.997848510742,168.987106323242,169.015823364258,169.087890625,169.143844604492,169.178024291992,169.255767822266,169.201171875,169.296203613281,169.334381103516],[168.44580078125,168.54541015625,168.5849609375,168.524597167969,168.3994140625,168.251663208008,168.305862426758,168.27783203125,168.233200073242,168.182022094727,168.158203125,168.19091796875,168.273147583008,168.297454833984,168.319534301758,168.341018676758,168.44580078125],[168.44677734375,168.4765625,168.460159301758,168.32275390625,168.212295532227,168.181442260742,168.148529052734,168.124313354492,168.135345458984,168.181838989258,168.19921875,168.233795166016,168.265426635742,168.296096801758,168.366027832031,168.44677734375],[168.296676635742,168.182418823242,168.021682739258,167.95703125,167.929000854492,167.984573364258,168.064270019531,168.163864135742,168.198333740234,168.235458374023,168.275680541992,168.297958374023,168.296676635742],[167.412506103516,167.458587646484,167.483703613281,167.498733520508,167.641998291016,167.681350708008,167.714447021484,167.775970458984,167.792572021484,167.836624145508,167.759765625,167.611419677734,167.5263671875,167.449310302734,167.436126708984,167.446868896484,167.400970458984,167.380279541016,167.349227905273,167.315612792969,167.24609375,167.218063354492,167.151458740234,167.182998657227,167.199508666992,167.253707885742,167.335739135742,167.412506103516],[167.218948364258,167.200775146484,167.0947265625,167.119049072266,167.234375,167.218948364258],[168.212890625,168.196197509766,168.179290771484,168.122848510742,168.159973144531,168.183486938477,168.267776489258,168.25634765625,168.212890625],[167.911315917969,167.84423828125,167.72021484375,167.674224853516,167.826278686523,168.002548217773,167.911315917969],[168.189163208008,168.171875,168.130477905273,168.104187011719,168.114944458008,168.136428833008,168.186920166016,168.189163208008],[166.745803833008,166.810165405273,166.885147094727,166.923446655273,166.967575073242,166.9873046875,167.026565551758,167.075576782227,167.054290771484,167.068557739258,167.1064453125,167.131637573242,167.182037353516,167.200775146484,167.199615478516,167.093933105469,166.936630249023,166.825790405273,166.75830078125,166.758987426758,166.698928833008,166.631057739258,166.647842407227,166.527252197266,166.526077270508,166.5673828125,166.607803344727,166.66259765625,166.745803833008],[167.584869384766,167.543258666992,167.430267333984,167.403503417969,167.410736083984,167.439071655273,167.506439208984,167.598922729492,167.584869384766],[167.488861083984,167.474212646484,167.451080322266,167.391799926758,167.406829833984,167.481063842773,167.547271728516,167.553512573242,167.553024291992,167.542877197266,167.498626708984,167.488861083984],[-178.046676635742,-178.103317260742,-178.158599853516,-178.194396972656,-178.178024291992,-178.142227172852,-178.105026245117,-178.043640136719,-178.046676635742],[-176.160598754883,-176.176895141602,-176.195358276367,-176.171188354492,-176.14794921875,-176.128082275391,-176.160598754883],[-171.4541015625,-171.728225708008,-171.86376953125,-171.911911010742,-172.028076171875,-172.0458984375,-171.98486328125,-171.858154296875,-171.603897094727,-171.5654296875,-171.506881713867,-171.461380004883,-171.449569702148,-171.4541015625],[-172.33349609375,-172.221542358398,-172.176849365234,-172.224960327148,-172.330856323242,-172.484512329102,-172.535690307617,-172.658798217773,-172.744094848633,-172.778518676758,-172.669631958008,-172.510894775391,-172.33349609375],[53.76318359375,53.8248023986816,53.9185562133789,54.1874008178711,54.5111312866211,54.4500007629395,54.4137725830078,54.2712898254395,54.1294898986816,53.7188453674316,53.5983390808105,53.4994125366211,53.3158226013184,53.3884773254395,53.4039077758789,53.4309577941895,53.5349617004395,53.6384773254395,53.76318359375],[42.755859375,42.6897468566895,42.7349586486816,42.78125,42.7941398620605,42.755859375],[42.7874031066895,42.7742195129395,42.7560539245605,42.6940422058105,42.7621116638184,42.79833984375,42.7874031066895],[42.5902366638184,42.5586891174316,42.5490226745605,42.5697288513184,42.6023445129395,42.62451171875,42.6104507446289,42.5902366638184],[53.0856437683105,52.5814476013184,52.4484405517578,52.3277359008789,52.2373046875,52.1740226745605,52.2220687866211,52.2174797058105,52.0873069763184,51.9658203125,51.8307609558105,51.7486343383789,51.6815414428711,51.6037101745605,51.3224601745605,51.01513671875,50.5270500183105,50.3385734558105,50.1668968200684,49.9063491821289,49.5486335754395,49.3499031066895,49.1029281616211,49.0480461120605,49.0046882629395,48.9287071228027,48.7799835205078,48.6683578491211,48.59375,48.4490242004395,48.27783203125,47.9899406433105,47.916015625,47.8550796508789,47.6333961486816,47.40771484375,47.2425765991211,46.9756851196289,46.7888679504395,46.6634750366211,46.501953125,46.203125,45.9197273254395,45.6573257446289,45.5339851379395,45.3935546875,45.1638679504395,45.1097640991211,45.0386695861816,44.8898429870605,44.7552719116211,44.6177749633789,44.3584976196289,44.2603492736816,44.1115226745605,44.005859375,43.9297866821289,43.8353500366211,43.6343765258789,43.4875946044922,43.4752960205078,43.23193359375,43.2826194763184,43.2824211120605,43.2340812683105,43.0890617370605,43.0933609008789,43.0448265075684,43.0062484741211,43.0187530517578,43.0210952758789,42.9469718933105,42.9221687316895,42.9129905700684,42.9373054504395,42.9364280700684,42.8970718383789,42.8556632995605,42.6578140258789,42.6978530883789,42.7364234924316,42.7884788513184,42.7990226745605,42.7999000549316,42.7171897888184,42.8396492004395,42.79931640625,42.986328125,43.0335922241211,43.0607414245605,43.1047859191895,43.1650390625,43.1863288879395,43.1844711303711,43.1456031799316,43.1165046691895,43.1261711120605,43.1359367370605,43.1559562683105,43.2213859558105,43.2369155883789,43.1863288879395,43.19091796875,43.3021507263184,43.3460960388184,43.41796875,43.4742202758789,43.5392570495605,43.5972633361816,43.6534194946289,43.7129898071289,43.8042984008789,43.8664054870605,43.9169921875,43.9596672058105,44.0082015991211,44.0859375,44.1559562683105,44.3546867370605,44.5464820861816,44.7467765808105,44.9464836120605,45.1480445861816,45.1927719116211,45.2366180419922,45.4065437316895,45.5353507995605,45.79443359375,46.07080078125,46.3103523254395,46.5134773254395,46.6820297241211,46.7276344299316,46.7785148620605,46.8799781799316,46.9756851196289,47.1435546875,47.2512702941895,47.36962890625,47.4417991638184,47.525390625,47.57958984375,47.7037124633789,47.8078117370605,47.9455070495605,48.0216789245605,48.1721649169922,48.3158187866211,48.5929679870605,48.8648414611816,49.0419921875,49.1923828125,49.4451141357422,49.7420883178711,50.0389671325684,50.3552742004395,50.7082023620605,50.9500007629395,51.2583961486816,51.5149383544922,51.7429695129395,51.9776382446289,52.0218772888184,52.0662117004395,52.1104507446289,52.1546897888184,52.1990242004395,52.2432632446289,52.2874984741211,52.3317375183105,52.3760719299316,52.4203109741211,52.4645500183105,52.5088844299316,52.5531234741211,52.5973625183105,52.6417007446289,52.6859397888184,52.7292022705078,52.8005867004395,52.8429679870605,52.9037132263184,52.96435546875,53.0249977111816,53.0856437683105],[37.85693359375,37.81396484375,37.61181640625,37.5900382995605,37.6497077941895,37.6848602294922,37.78955078125,37.8728523254395,37.8876953125,37.85693359375],[31.2878913879395,31.2931632995605,31.3001956939697,31.3480472564697,31.4193344116211,31.4666976928711,31.53173828125,31.5296859741211,31.5456047058105,31.6041011810303,31.6755867004395,31.7000007629395,31.7240219116211,31.7996082305908,31.8583011627197,31.9080066680908,31.9505863189697,31.9666004180908,31.98583984375,31.9832038879395,31.984375,31.9857425689697,31.9870128631592,31.9793930053711,31.9845695495605,31.9203128814697,31.9283199310303,31.9482440948486,31.9216785430908,31.8714828491211,31.6404285430908,31.4151382446289,31.3826198577881,31.3351554870605,31.2073249816895,31.0880870819092,31.0333023071289,30.9452133178711,30.8033199310303,30.7890625,30.7875003814697,30.7943344116211,30.8067378997803,30.88330078125,30.9380855560303,31.0633792877197,31.2740230560303,31.4695301055908,31.7425765991211,31.9583969116211,31.9460926055908,31.9671878814697,31.9947261810303,32.0248031616211,32.0816421508789,32.1128883361816,32.1996078491211,32.353515625,32.4777336120605,32.5887680053711,32.7765617370605,32.8861312866211,32.84912109375,32.7058601379395,32.6570281982422,32.5347671508789,32.3751945495605,32.2857398986816,32.0272445678711,31.9553718566895,31.8915061950684,31.7782230377197,31.3351554870605,31.1699199676514,31.0233402252197,30.8776359558105,30.66357421875,30.4722671508789,30.2886714935303,29.97119140625,29.8302745819092,29.7351570129395,29.4829120635986,29.1278343200684,28.85595703125,28.4494152069092,28.2140636444092,27.8606433868408,27.7621097564697,27.36376953125,27.0774421691895,26.6136722564697,26.4294929504395,25.9895515441895,25.8058605194092,25.6524429321289,25.63818359375,25.57421875,25.4772453308105,25.1697273254395,25.0029296875,24.9055652618408,24.8271484375,24.5955085754395,24.1830062866211,23.6978511810303,23.5855464935303,23.3503894805908,23.2681655883789,22.9255867004395,22.7355461120605,22.5538082122803,22.4144515991211,22.2455081939697,21.7889652252197,21.55322265625,21.3498058319092,21.2489242553711,21.0601558685303,20.9898433685303,20.8824214935303,20.7748050689697,20.5298824310303,20.4346675872803,20.0206069946289,19.92626953125,19.8500003814697,19.6349620819092,19.3915042877197,19.2982425689697,19.3232421875,19.3307628631592,19.2793941497803,19.24462890625,19.1491222381592,19.0983390808105,18.9521484375,18.9015636444092,18.8313465118408,18.8250980377197,18.8306636810303,18.8263664245605,18.8087882995605,18.7521476745605,18.7086906433105,18.6051750183105,18.5338878631592,18.5003910064697,18.4621086120605,18.4616203308105,18.4103507995605,18.35205078125,18.3333988189697,18.3543949127197,18.4650382995605,18.4564456939697,18.4330062866211,18.3094730377197,18.26123046875,18.1563491821289,18.0748043060303,17.9925785064697,17.9583969116211,17.8782234191895,17.85107421875,17.8953132629395,17.9652347564697,18.0365238189697,18.125,18.2508792877197,18.3252925872803,18.3298835754395,18.3107414245605,18.2108402252197,18.1636714935303,17.9385738372803,17.6774406433105,17.3470706939697,17.1890621185303,16.9500007629395,16.7394523620605,16.4807605743408,16.4475574493408,16.4871082305908,16.6261711120605,16.689453125,16.7230472564697,16.7557621002197,16.7875003814697,16.7945327758789,16.8101558685303,16.8412113189697,16.8752918243408,16.9333000183105,17.0562496185303,17.1494140625,17.1884765625,17.20458984375,17.2458000183105,17.31201171875,17.3586921691895,17.3857421875,17.3802738189697,17.3425769805908,17.3478527069092,17.3958969116211,17.4157218933105,17.4479484558105,17.6167964935303,17.6993160247803,17.8416023254395,17.97607421875,18.1027355194092,18.3108386993408,18.6003913879395,18.8387699127197,19.0260734558105,19.1617183685303,19.2458019256592,19.2822265625,19.27099609375,19.3126945495605,19.4072265625,19.48291015625,19.5398445129395,19.6714839935303,19.8778324127197,19.98046875,19.98046875,19.98046875,19.98046875,19.98046875,19.98046875,19.98046875,19.98046875,19.98046875,19.98046875,20.0286121368408,20.34521484375,20.4306659698486,20.47314453125,20.6092777252197,20.7107429504395,20.7931632995605,20.7994155883789,20.81103515625,20.8226566314697,20.8150386810303,20.7570323944092,20.6978511810303,20.6267585754395,20.6199226379395,20.6414051055908,20.68505859375,20.7398433685303,20.8708972930908,20.9539070129395,21.0709972381592,21.4549808502197,21.5013675689697,21.6462879180908,21.6947269439697,21.7380867004395,21.7882804870605,21.8332042694092,21.91455078125,22.0109367370605,22.0909175872803,22.2175769805908,22.4708995819092,22.5486335754395,22.5976543426514,22.6402339935303,22.72900390625,22.7960948944092,22.8189449310303,22.8788089752197,22.9512691497803,23.0220718383789,23.0575199127197,23.1487312316895,23.2660160064697,23.3892574310303,23.521484375,23.6707038879395,23.8234367370605,23.8937511444092,23.9695320129395,24.1044921875,24.1929683685303,24.33056640625,24.4001941680908,24.5558605194092,24.7481460571289,24.8692378997803,24.9989261627197,25.0924816131592,25.21337890625,25.34619140625,25.4436531066895,25.5181636810303,25.5837879180908,25.6591796875,25.70263671875,25.7699203491211,25.8524417877197,25.8818359375,25.912109375,26.0318374633789,26.130859375,26.3971691131592,26.4517593383789,26.5015621185303,26.6177730560303,26.7611331939697,26.8350582122803,26.9706058502197,26.9870109558105,27.0855464935303,27.1463871002197,27.185546875,27.2412109375,27.3133792877197,27.3992176055908,27.4987316131592,27.5631847381592,27.5926742553711,27.6438484191895,27.716796875,27.75830078125,27.7685546875,27.8125972747803,27.8905277252197,27.9313488006592,27.93505859375,28.0279312133789,28.2101573944092,28.3817386627197,28.5428695678711,28.6955070495605,28.8398456573486,28.9458026885986,29.0134754180908,29.1298828125,29.3648452758789,29.3774433135986,29.6630840301514,29.9023418426514,30.1904296875,30.4601573944092,30.7116203308105,30.9161128997803,31.0734367370605,31.197265625,31.2878913879395],[28.7369136810303,28.9010753631592,28.9752922058105,29.0290031433105,29.0980472564697,29.1219730377197,29.1421871185303,29.1951179504395,29.2492198944092,29.2935543060303,29.3488273620605,29.38671875,29.3907241821289,29.3708992004395,29.3359394073486,29.3013687133789,29.259765625,29.1780281066895,29.0580081939697,28.9537105560303,28.8562507629395,28.8162097930908,28.7217788696289,28.68115234375,28.6526374816895,28.6257801055908,28.5833969116211,28.4718742370605,28.2326183319092,28.0843753814697,27.9598617553711,27.8303699493408,27.7355480194092,27.6604499816895,27.5902347564697,27.5271472930908,27.4910163879395,27.4580078125,27.4249019622803,27.3568363189697,27.2945327758789,27.2074222564697,27.09521484375,27.0569343566895,27.0517578125,27.091796875,27.1304683685303,27.1935539245605,27.23974609375,27.3126945495605,27.3553695678711,27.3497066497803,27.3640632629395,27.3884754180908,27.4085922241211,27.4314441680908,27.4919929504395,27.5065422058105,27.5490226745605,27.5896472930908,27.6666011810303,27.7531242370605,27.90185546875,28.0181636810303,28.0568351745605,28.0963859558105,28.1287117004395,28.1390628814697,28.1761703491211,28.3154296875,28.39208984375,28.4390621185303,28.4996089935303,28.57666015625,28.6343746185303,28.6468753814697,28.7369136810303],[32.919921875,32.92333984375,32.9510726928711,32.9798812866211,32.9821281433105,32.9959945678711,33.0377922058105,33.0724639892578,33.1044921875,33.1480445861816,33.1957015991211,33.2126960754395,33.25,33.3104515075684,33.3509788513184,33.3371086120605,33.3115234375,33.3935546875,33.5000953674316,33.5289077758789,33.53759765625,33.5537109375,33.6261711120605,33.6615257263184,33.6590843200684,33.4647483825684,33.4031257629395,33.3449211120605,33.2927742004395,33.2613258361816,33.2727546691895,33.2932624816895,33.3386726379395,33.3797836303711,33.3455047607422,33.2683601379395,33.2327156066895,33.2263679504395,33.25,33.2882843017578,33.3039054870605,33.3050765991211,33.3009757995605,33.2523422241211,33.3401374816895,33.3700180053711,33.4914054870605,33.5123062133789,33.4832038879395,33.4306640625,33.39794921875,33.2434577941895,33.0215797424316,32.9751930236816,32.9456062316895,32.9705085754395,33,32.9904289245605,32.9710922241211,32.9776344299316,32.9675788879395,32.9385719299316,32.8997077941895,32.8518524169922,32.8140640258789,32.7583999633789,32.67041015625,32.6720695495605,32.7717781066895,32.7974624633789,32.8067359924316,32.7853507995605,32.76513671875,32.81103515625,32.8671875,32.9203147888184,32.9675788879395,32.9812469482422,32.9920883178711,33.00927734375,33.0423812866211,33.1036148071289,33.1480445861816,33.2017593383789,32.9871101379395,32.87451171875,32.55322265625,32.2728538513184,32.1999015808105,32.0544891357422,31.9821281433105,31.7289066314697,31.6230487823486,31.5378894805908,31.3285179138184,31.1308574676514,30.9151363372803,30.6733417510986,30.5376930236816,30.4460945129395,30.2318344116211,30.2217769622803,30.2250003814697,30.2521476745605,30.3056659698486,30.3505840301514,30.3798828125,30.3960933685303,30.2506828308105,29.9949226379395,29.7295894622803,29.4873027801514,29.2878913879395,29.0505847930908,28.9730472564697,28.9130859375,28.8755836486816,28.8567371368408,28.8567371368408,28.8327140808105,28.7605476379395,28.7606468200684,28.3998050689697,28.1637687683105,27.9322261810303,27.7565441131592,27.63671875,27.4378910064697,27.2357425689697,27.0207996368408,26.7798824310303,26.5775375366211,26.3333988189697,26.1395511627197,25.9958972930908,25.86328125,25.7416019439697,25.6396465301514,25.55712890625,25.4517574310303,25.2587890625,25.0921878814697,25.0017566680908,24.9324226379395,24.73291015625,24.27490234375,24.2271480560303,24.0369148254395,23.7992191314697,23.5949211120605,23.3806648254395,23.181640625,22.9558601379395,22.7219734191895,22.5459957122803,22.4594745635986,22.3050765991211,22.1939449310303,22.1506843566895,22.0402355194092,21.9797840118408,21.9797840118408,21.9796867370605,21.9795894622803,21.9794921875,21.9793949127197,21.9792976379395,21.9791011810303,21.9791011810303,21.97900390625,21.9789066314697,22.2095699310303,22.4709968566895,22.7443351745605,23.04150390625,23.3386726379395,23.6358413696289,23.8431644439697,23.8974609375,23.9629878997803,23.9680652618408,23.8824214935303,23.8865242004395,23.9093761444092,23.9447269439697,23.9913101196289,23.9964847564697,23.9588871002197,23.9623050689697,23.9734382629395,23.98388671875,23.9709968566895,23.98681640625,24.0146484375,24.029296875,24.0466804504395,24.0414066314697,24.0255870819092,24.0100574493408,23.98828125,23.9665031433105,24.0027351379395,24.0784187316895,24.1151371002197,24.1365222930908,24.1872081756592,24.3199214935303,24.36572265625,24.3962879180908,24.3779296875,24.3351554870605,24.3779296875,24.4666023254395,24.5185546875,24.6682605743408,24.7281246185303,24.8063468933105,24.8768558502197,25.0759754180908,25.1848640441895,25.2459964752197,25.2887687683105,25.3193359375,25.2917976379395,25.2826156616211,25.3207035064697,25.3494129180908,25.4133777618408,25.4599609375,25.5119152069092,25.6188468933105,25.8548812866211,25.9265632629395,26.0259761810303,26.0963878631592,26.3396472930908,26.4296894073486,26.5963859558105,26.7296867370605,26.8240242004395,26.8904285430908,26.9308586120605,26.9496097564697,26.9768543243408,27.0266609191895,27.0460948944092,27.0954093933105,27.1591796875,27.1963863372803,27.2380867004395,27.4236335754395,27.4870128631592,27.5333995819092,27.5738296508789,27.6443347930908,27.7568378448486,27.857421875,28.06884765625,28.2373046875,28.3577136993408,28.4128894805908,28.4514656066895,28.4744148254395,28.51123046875,28.5508785247803,28.6154289245605,28.6729488372803,28.7300777435303,28.7731456756592,28.8587894439697,28.9216804504395,28.9422836303711,29.0142574310303,29.1116218566895,29.2018566131592,29.2537097930908,29.3818340301514,29.4814453125,29.5541973114014,29.5971660614014,29.6302719116211,29.6476554870605,29.6517562866211,29.7226581573486,29.7751960754395,29.7953128814697,29.7964859008789,29.7962913513184,29.7960929870605,29.7958011627197,29.7956047058105,29.7955093383789,29.7953128814697,29.7951183319092,29.7496089935303,29.6919937133789,29.5597648620605,29.5082035064697,29.4919910430908,29.5022468566895,29.5048847198486,29.4855480194092,29.4275379180908,29.34375,29.1912117004395,29.0643577575684,28.9734363555908,28.8499984741211,28.7694320678711,28.5746097564697,28.5416011810303,28.4825210571289,28.4318351745605,28.4070320129395,28.3833999633789,28.3572254180908,28.4041996002197,28.4703121185303,28.5179691314697,28.5442390441895,28.6388664245605,28.6455078125,28.607421875,28.6171875,28.62353515625,28.62890625,28.6300792694092,28.6041984558105,28.54052734375,28.4001960754395,28.4006824493408,28.4842777252197,28.6165027618408,28.6812515258789,28.7587890625,28.7935543060303,28.8695297241211,28.9177742004395,28.9344749450684,28.8981437683105,28.9722652435303,29.2156257629395,29.4837894439697,29.7662105560303,30.0513668060303,30.3275394439697,30.5779304504395,30.7511711120605,30.7767581939697,30.8306636810303,30.8919925689697,30.9683609008789,31.0333976745605,31.0763664245605,31.3505859375,31.4492206573486,31.5348644256592,31.5562496185303,31.61279296875,31.6736335754395,31.7000007629395,31.7447280883789,31.8180675506592,31.8861331939697,31.9186515808105,31.9218769073486,31.9425792694092,32.0353546142578,32.1297836303711,32.2208976745605,32.3193359375,32.4332046508789,32.4871101379395,32.6083984375,32.7566413879395,32.86328125,32.919921875],[31.2878913879395,31.197265625,31.0734367370605,30.9161128997803,30.7116203308105,30.4601573944092,30.1904296875,29.9023418426514,29.6630840301514,29.3774433135986,29.3648452758789,29.3152332305908,29.2372093200684,29.1068382263184,29.0714836120605,29.0423812866211,29.0233383178711,29.0158214569092,29.0373058319092,29.0255851745605,28.9907207489014,28.9193344116211,28.7477531433105,28.5320301055908,28.181640625,28.0456047058105,28.0140609741211,27.974609375,27.9074230194092,27.8441410064697,27.6934585571289,27.66943359375,27.6769523620605,27.6880855560303,27.7042980194092,27.6969738006592,27.69482421875,27.6996097564697,27.6792964935303,27.6246089935303,27.4689464569092,27.28076171875,27.2746105194092,27.2567386627197,27.2214851379395,27.17822265625,27.091796875,26.9167003631592,26.67822265625,26.474609375,26.2410144805908,26.1680660247803,26.0819339752197,25.95068359375,25.9591808319092,25.9393558502197,25.8119144439697,25.78369140625,25.76123046875,25.5583019256592,25.4892559051514,25.4367198944092,25.3843746185303,25.3402347564697,25.2824211120605,25.2422847747803,25.2240238189697,25.2390613555908,25.2587890625,25.4517574310303,25.55712890625,25.6396465301514,25.7416019439697,25.86328125,25.9958972930908,26.1395511627197,26.3333988189697,26.5775375366211,26.7798824310303,27.0207996368408,27.2357425689697,27.4378910064697,27.63671875,27.7565441131592,27.9322261810303,28.1637687683105,28.3998050689697,28.7606468200684,28.7605476379395,28.8327140808105,28.8567371368408,28.8567371368408,28.8755836486816,28.9130859375,28.9730472564697,29.0505847930908,29.2878913879395,29.4873027801514,29.7295894622803,29.9949226379395,30.2506828308105,30.3960933685303,30.3981456756592,30.4093742370605,30.4377918243408,30.6301746368408,30.9387702941895,31.2362308502197,31.4261722564697,31.4898433685303,31.6875972747803,31.9398422241211,32.2432594299316,32.4519538879395,32.6358375549316,32.7417945861816,32.8102531433105,32.9029312133789,32.9480476379395,32.9378890991211,32.8762702941895,32.8843765258789,32.9693374633789,32.9807586669922,32.9546852111816,32.95556640625,32.9646492004395,32.978515625,32.9963874816895,32.9930648803711,32.9424819946289,32.9016609191895,32.9002952575684,32.8845710754395,32.8544921875,32.7219696044922,32.69921875,32.69970703125,32.7165031433105,32.7662086486816,32.826171875,32.8498039245605,32.8499984741211,32.8309555053711,32.7776374816895,32.8307609558105,32.8904304504395,32.97265625,33.0067405700684,33.0048828125,32.9927749633789,32.86962890625,32.7808609008789,32.6725578308105,32.529296875,32.4923820495605,32.4776344299316,32.4828109741211,32.4761734008789,32.3536109924316,32.4297866821289,32.4124031066895,32.37109375,32.1947250366211,32.0163116455078,31.8859348297119,31.7376956939697,31.5714836120605,31.4294910430908,31.2878913879395],[5.79511737823486,5.87001943588257,5.8671875,5.83544874191284,5.65341806411743,5.619140625,5.61308574676514,5.61796855926514,5.62529277801514,5.6669921875,5.65683555603027,5.54726600646973,5.45498037338257,5.42041015625,5.4130859375,5.39824199676514,5.37724637985229,5.26894474029541,5.15234375,5.06806659698486,5.09404325485229,5.11757802963257,5.16484403610229,5.24746084213257,5.24169921875,5.13798809051514,5.06591796875,5.05996131896973,5.06982421875,5.06904268264771,5.06230449676514,5.03623008728027,5.01406240463257,5.03994131088257,5.31601619720459,5.49531269073486,5.79511737823486],[5.49912118911743,5.42441415786743,5.24238300323486,5.18388652801514,5.16855478286743,5.376953125,5.53925800323486,5.65810585021973,5.73554706573486,5.79746055603027,5.82949209213257,5.83417940139771,5.80087900161743,5.74101591110229,5.62060546875,5.54804706573486,5.49912118911743],[12.4391603469849,12.4383792877197,12.4305658340454,12.4275388717651,12.4305658340454,12.4391603469849]],"ys":[[12.4520015716553,12.4229984283447,12.4385251998901,12.50048828125,12.5469722747803,12.5970697402954,12.6141109466553,12.567626953125,12.48046875,12.4520015716553],[37.2316398620605,37.2250518798828,37.2491722106934,37.28564453125,37.2907218933105,37.2667007446289,37.2366218566895,37.15771484375,37.1373519897461,37.0572242736816,37.0306625366211,37.0221710205078,36.9836921691895,36.8968772888184,36.8257331848145,36.8230972290039,36.8529319763184,36.8884773254395,36.8877906799316,36.8816909790039,36.8685569763184,36.8516120910645,36.8350105285645,36.82958984375,36.802001953125,36.7658195495605,36.7423858642578,36.7347183227539,36.7008819580078,36.6337394714355,36.5341796875,36.486083984375,36.4318351745605,36.4265632629395,36.4364738464355,36.377685546875,36.2932624816895,36.1711883544922,36.1217765808105,36.0420875549316,36.0006828308105,35.9385261535645,35.8801765441895,35.833740234375,35.714599609375,35.5975074768066,35.5468254089355,35.4608383178711,35.4079093933105,35.3704109191895,35.3285140991211,35.2888641357422,35.2479972839355,35.2117652893066,35.1830062866211,35.1506843566895,35.1014137268066,35.0511245727539,34.9669418334961,34.9096183776855,34.8677253723145,34.779541015625,34.6815948486328,34.599609375,34.5546379089355,34.5303688049316,34.4862823486328,34.43115234375,34.3694343566895,34.2732429504395,34.2040557861328,34.1202659606934,34.0497093200684,33.9818840026855,33.9522933959961,33.950439453125,33.9611320495605,33.9759750366211,34.0518074035645,34.0072746276855,33.8976554870605,33.7198715209961,33.6207542419434,33.4546852111816,33.3690414428711,33.2890129089355,33.1980972290039,33.1124992370605,33.0947265625,33.0641593933105,33.0200691223145,32.8328132629395,32.7642555236816,32.6827163696289,32.59033203125,32.5305671691895,32.4335441589355,32.2494621276855,31.9368171691895,31.8380870819092,31.7384757995605,31.6673831939697,31.6342296600342,31.6464347839355,31.7080574035645,31.7597179412842,31.8029804229736,31.7941417694092,31.7544918060303,31.7676734924316,31.8073749542236,31.8029804229736,31.7632827758789,31.6779804229736,31.5481929779053,31.5387706756592,31.5064945220947,31.4533214569092,31.4099597930908,31.3792476654053,31.3439464569092,31.31298828125,31.2776870727539,31.234619140625,31.2178230285645,31.2429218292236,31.3002433776855,31.3056144714355,31.263671875,31.1945323944092,31.0460433959961,31.0199699401855,30.9965839385986,30.9645500183105,30.9122066497803,30.8027839660645,30.6079120635986,30.5029792785645,30.3211421966553,30.1934566497803,30.109619140625,30.0435066223145,29.9685554504395,29.9200191497803,29.86572265625,29.8355941772461,29.7789077758789,29.7013206481934,29.6515617370605,29.5776348114014,29.5594749450684,29.5527839660645,29.5641593933105,29.5671367645264,29.5645008087158,29.5443344116211,29.5069332122803,29.4603519439697,29.4142570495605,29.3919429779053,29.4300804138184,29.4979991912842,29.4083499908447,29.4253902435303,29.5304183959961,29.6656742095947,29.7494144439697,29.8586902618408,30.12841796875,30.3637218475342,30.5993633270264,30.8319358825684,30.9132823944092,31.0725593566895,31.2853031158447,31.3824214935303,31.4216289520264,31.4511222839355,31.4832515716553,31.4951667785645,31.6605968475342,31.7344722747803,31.877197265625,31.9871120452881,32.16796875,32.2494125366211,32.6000022888184,32.7943840026855,32.994873046875,33.0587882995605,33.1378402709961,33.3235359191895,33.3638191223145,33.4562492370605,33.5052223205566,33.5389633178711,33.5586891174316,33.5604019165039,33.5883293151855,33.638916015625,33.7119140625,33.8419914245605,34.0947761535645,34.2196311950684,34.3071746826172,34.3194313049316,34.4180183410645,34.4752426147461,34.4917945861816,34.5182647705078,34.5447273254395,34.5546379089355,34.5876960754395,34.6339836120605,34.6538543701172,34.7100601196289,34.749755859375,34.7993659973145,34.8556175231934,34.9217300415039,35.0011215209961,35.0507316589355,35.09375,35.1565437316895,35.20947265625,35.2723121643066,35.2888641357422,35.31201171875,35.3616218566895,35.4244613647461,35.4740715026855,35.5137710571289,35.5534210205078,35.6195793151855,35.6294898986816,35.5931167602539,35.5457992553711,35.4578628540039,35.4323234558105,35.41943359375,35.4314918518066,35.4478988647461,35.4437026977539,35.3796882629395,35.2899398803711,35.250244140625,35.1891098022461,35.1707992553711,35.2312507629395,35.2513694763184,35.2398948669434,35.233154296875,35.2553215026855,35.2713394165039,35.3496589660645,35.4091796875,35.44580078125,35.5680656433105,35.6375503540039,35.6781234741211,35.728271484375,35.7667465209961,35.8186988830566,35.84619140625,35.8584480285645,35.8583984375,35.9131355285645,35.9678192138672,36.0123519897461,36.0197257995605,36.0121116638184,36.0250968933105,36.067626953125,36.1126976013184,36.14892578125,36.22607421875,36.3406753540039,36.4275894165039,36.5545425415039,36.7501945495605,36.9647941589355,37.0592765808105,37.132080078125,37.1935577392578,37.2379379272461,37.2467765808105,37.2511711120605,37.368408203125,37.4678230285645,37.5191421508789,37.5608406066895,37.5691413879395,37.5081062316895,37.4147453308105,37.3681640625,37.3447265625,37.3484840393066,37.3712882995605,37.3348159790039,37.2352066040039,37.2095680236816,37.2580108642578,37.2666511535645,37.2356452941895,37.2225074768066,37.2272453308105,37.1998062133789,37.1722145080566,37.14013671875,37.064208984375,36.9720230102539,36.9498023986816,37.0215339660645,37.0130844116211,37.0363273620605,37.0884284973145,37.1375007629395,37.1834487915039,37.2244644165039,37.2583999633789,37.2680206298828,37.2580108642578,37.2707023620605,37.3028335571289,37.3168449401855,37.3280754089355,37.3339385986328,37.3250465393066,37.2665023803711,37.1583023071289,37.1083984375,37.116943359375,37.1500473022461,37.207763671875,37.2908706665039,37.3993148803711,37.4867172241211,37.5530776977539,37.5940437316895,37.6095695495605,37.6002922058105,37.566162109375,37.5472183227539,37.5435028076172,37.5824699401855,37.6641578674316,37.765380859375,37.8860359191895,37.9244117736816,37.9412078857422,37.9848136901855,38.075439453125,38.1919937133789,38.3344230651855,38.4225616455078,38.4563980102539,38.4178695678711,38.3069839477539,38.1702613830566,38.0079078674316,37.9184112548828,37.90185546875,37.9062995910645,37.931884765625,37.9331512451172,37.9101066589355,37.8642578125,37.795654296875,37.6029319763184,37.43603515625,37.2718276977539,37.1275405883789,37.0150871276855,36.8451156616211,36.73291015625,36.6969223022461,36.6840324401855,36.6942863464355,36.7664527893066,36.9005393981934,36.98291015625,37.029052734375,37.1727066040039,37.2675285339355,37.4084968566895,37.4622573852539,37.4716796875,37.446044921875,37.4372062683105,37.4304695129395,37.4187507629395,37.3757820129395,37.3294410705566,37.2912101745605,37.2615699768066,37.2393569946289,37.2317886352539,37.2831535339355,37.3162117004395,37.3294410705566,37.3724594116211,37.4154319763184,37.4187507629395,37.3956069946289,37.3823738098145,37.3944816589355,37.3570289611816,37.2859382629395,37.2419929504395,37.2316398620605],[-10.8717775344849,-11.0028324127197,-11.1847658157349,-11.3156251907349,-11.3741207122803,-11.4053707122803,-11.4391593933105,-11.5176753997803,-11.5872068405151,-11.6358404159546,-11.7249994277954,-11.8529300689697,-11.9878902435303,-12.1177730560303,-12.3506832122803,-12.4221677780151,-12.5437498092651,-12.6361322402954,-12.7432613372803,-12.7990236282349,-12.9569339752197,-12.9884767532349,-12.9982414245605,-13.0009765625,-13.0009765625,-13.0009765625,-13.0009765625,-13.0009765625,-13.0009765625,-13.0009765625,-13.0009765625,-13.1568365097046,-13.4777336120605,-13.7987308502197,-14.1196279525757,-14.4405279159546,-14.7614250183105,-15.0823249816895,-15.4032220840454,-15.7241220474243,-15.95556640625,-16.2627925872803,-16.59716796875,-16.6281242370605,-16.6895503997803,-16.8151359558105,-16.9102535247803,-17.0752944946289,-17.2857418060303,-17.4744148254395,-17.640625,-17.6988277435303,-17.7816410064697,-17.8374996185303,-17.9051761627197,-17.94775390625,-18.0006828308105,-17.99951171875,-17.9629878997803,-17.9557609558105,-18.0060558319092,-18.0197257995605,-17.9966793060303,-17.9525394439697,-17.8874034881592,-17.8636722564697,-17.88134765625,-17.86865234375,-17.8254871368408,-17.8084964752197,-17.8176746368408,-17.8035163879395,-17.7663097381592,-17.7032222747803,-17.5700187683105,-17.4427738189697,-17.4246082305908,-17.4051742553711,-17.3994140625,-17.39599609375,-17.3927726745605,-17.392578125,-17.3919925689697,-17.3914070129395,-17.3908195495605,-17.3902359008789,-17.3896484375,-17.38916015625,-17.3885746002197,-17.3879890441895,-17.3876953125,-17.3977527618408,-17.4088859558105,-17.4041996002197,-17.3887710571289,-17.3607425689697,-17.2883777618408,-17.2334957122803,-17.1412124633789,-17.0400390625,-17.0078125,-16.9895515441895,-16.9716796875,-16.9676761627197,-17.0154304504395,-17.0625972747803,-17.1082038879395,-17.1605472564697,-17.2126941680908,-17.2058601379395,-17.21337890625,-17.2099609375,-17.16455078125,-17.1685543060303,-17.2265625,-17.2492198944092,-16.8712902069092,-16.7041015625,-16.5042972564697,-15.9864263534546,-15.9153327941895,-15.831934928894,-15.7683591842651,-15.7198238372803,-15.6339836120605,-15.513671875,-15.2482423782349,-14.6374998092651,-14.0390615463257,-13.7554693222046,-13.4377927780151,-13.0277347564697,-12.7756834030151,-12.6521482467651,-12.5204105377197,-12.2861328125,-12.1238279342651,-11.8127927780151,-11.4879884719849,-11.0543947219849,-10.9296875,-10.7571287155151,-10.6335935592651,-10.5123052597046,-10.4207038879395,-10.2571287155151,-9.99892520904541,-9.82675838470459,-9.70322322845459,-9.55068302154541,-9.3896484375,-9.23037147521973,-9.04804706573486,-8.99101638793945,-8.92226600646973,-8.89970684051514,-8.93427753448486,-8.97519588470459,-9.0068359375,-8.68720722198486,-8.62470722198486,-8.55478572845459,-8.46923828125,-8.36972618103027,-7.78017520904541,-7.23183584213257,-6.95478487014771,-6.59033155441284,-6.35341739654541,-6.18730497360229,-6.12431621551514,-6.09257841110229,-6.08427715301514,-6.04589796066284,-6.00390625,-5.90761709213257,-5.86484336853027,-5.85624980926514,-5.8818359375,-5.86337900161743,-5.86181640625,-5.86171865463257,-5.85517597198486,-5.85722637176514,-5.86513662338257,-5.8759765625,-5.89267587661743,-5.88886690139771,-5.88007783889771,-5.87451171875,-5.86884737014771,-5.86386728286743,-5.86494159698486,-5.86562538146973,-5.90019464492798,-5.96582078933716,-6.02529287338257,-6.05156230926514,-6.11455059051514,-6.16425752639771,-6.24140644073486,-6.34599637985229,-6.4716796875,-6.61845731735229,-6.77255916595459,-6.93398475646973,-7.06210947036743,-7.15703105926514,-7.25742197036743,-7.36308574676514,-7.41904306411743,-7.46132755279541,-7.62333965301514,-7.88193321228027,-8.07587909698486,-8.09902381896973,-8.09072303771973,-8.07138633728027,-8.06767559051514,-8.10761737823486,-8.10078144073486,-8.02382755279541,-8.00029277801514,-7.96855449676514,-7.93593788146973,-7.93603515625,-7.99814462661743,-8.00146484375,-8.00146484375,-7.96660184860229,-7.70654344558716,-7.65507793426514,-7.55732440948486,-7.47216796875,-7.39072227478027,-7.27949285507202,-7.14443397521973,-7.037109375,-6.986328125,-6.97646474838257,-6.94628858566284,-6.91582012176514,-6.91992139816284,-6.93515634536743,-7.12177705764771,-7.18281316757202,-7.24443340301514,-7.27773475646973,-7.28144502639771,-7.28496122360229,-7.29667997360229,-7.30546855926514,-7.31464862823486,-7.32861280441284,-7.42099618911743,-7.60165977478027,-7.86542987823486,-8.11191463470459,-8.34111309051514,-8.693359375,-8.90351486206055,-9.16845703125,-9.46875,-9.59423828125,-9.7255859375,-9.86279296875,-10.0406255722046,-10.2590827941895,-10.3966789245605,-10.4533205032349,-10.5515632629395,-10.6913089752197,-10.7839841842651,-10.8294916152954,-10.8922853469849,-11.0126943588257,-11.1219730377197,-11.1636714935303,-11.1941404342651,-11.1986322402954,-11.1594734191895,-11.0867185592651,-11.0558595657349,-11.0597648620605,-11.0802736282349,-11.087890625,-11.0748052597046,-10.9764652252197,-10.9693355560303,-10.9786128997803,-11.0076169967651,-11.0136728286743,-10.9832029342651,-10.9434576034546,-10.8915033340454,-10.8717775344849],[-5.75869178771973,-5.73193311691284,-5.63271474838257,-5.53867149353027,-5.46826171875,-5.32480430603027,-5.19716787338257,-5.03515625,-5.00429677963257,-4.95214891433716,-4.83769512176514,-4.77861356735229,-4.76552724838257,-4.72412109375,-4.65771484375,-4.619140625,-4.58750009536743,-4.53115272521973,-4.46972703933716,-4.43056631088257,-4.42890644073486,-4.44511699676514,-4.5517578125,-4.61923837661743,-4.634765625,-4.65107393264771,-4.6953125,-4.73662090301514,-4.90537118911743,-4.97841835021973,-4.99658203125,-5.03691339492798,-5.07148408889771,-5.09062528610229,-5.11269521713257,-5.14892578125,-5.42460918426514,-5.69580125808716,-5.71875,-5.72773408889771,-5.74648380279541,-5.75869178771973],[18.2217769622803,18.17138671875,18.2002925872803,18.2697257995605,18.2647953033447,18.2217769622803],[42.5472679138184,42.524658203125,42.4258766174316,42.3389663696289,42.3087921142578,42.2749519348145,42.223388671875,42.1714820861816,42.0130844116211,41.9174308776855,41.8736801147461,41.8623504638672,41.7064933776855,41.6270484924316,41.5747566223145,41.5541038513184,41.5212898254395,41.3914070129395,41.3360824584961,41.2726058959961,41.1278305053711,41.0830574035645,41.0616683959961,40.9283676147461,40.9052734375,40.9179229736328,40.9031257629395,40.8715362548828,40.8499031066895,40.7752952575684,40.7177734375,40.6586418151855,40.6224594116211,40.5633811950684,40.494384765625,40.4679183959961,40.4454612731934,40.3918952941895,40.3349151611328,40.2926750183105,40.2463874816895,40.1517601013184,40.1173820495605,40.0826644897461,40.0685043334961,40.0655746459961,40.0494613647461,40.0171852111816,39.9910659790039,39.9794425964355,39.9507789611816,39.890625,39.841796875,39.8026390075684,39.791748046875,39.7966804504395,39.7822265625,39.7385749816895,39.701171875,39.6783676147461,39.6535186767578,39.66162109375,39.6991233825684,39.710693359375,39.7094268798828,39.801025390625,39.8722686767578,40.0435562133789,40.2099609375,40.2848625183105,40.3477058410645,40.4070816040039,40.4087409973145,40.3936996459961,40.3756828308105,40.4053688049316,40.4702644348145,40.6220703125,40.663818359375,40.7907218933105,40.9333000183105,41.1060562133789,41.2363739013672,41.3209953308105,41.4247550964355,41.5626945495605,41.5968246459961,41.6404304504395,41.7874984741211,41.8561553955078,41.8690910339355,41.9188499450684,41.9977531433105,42.0240211486816,42.069091796875,42.1292953491211,42.1725578308105,42.249267578125,42.3418922424316,42.4153823852539,42.491943359375,42.5654296875,42.6285667419434,42.64794921875,42.634521484375,42.60693359375,42.525146484375,42.4969253540039,42.4761695861816,42.486328125,42.5066871643066,42.5499038696289,42.5472679138184],[60.0406761169434,60.0169410705566,60.0116691589355,60.0327644348145,60.0301284790039,60.0406761169434,60.0617179870605,60.0749015808105,60.0696296691895,60.0406761169434],[60.1871566772461,60.1647453308105,60.1703605651855,60.1626968383789,60.1350631713867,60.1449737548828,60.1845664978027,60.2438468933105,60.24609375,60.1871566772461],[60.3511695861816,60.3508758544922,60.3593254089355,60.3534164428711,60.314697265625,60.2937469482422,60.2830123901367,60.2612762451172,60.1935539245605,60.1922836303711,60.2008781433105,60.1934585571289,60.1806678771973,60.1524925231934,60.0935554504395,60.0817375183105,60.0989723205566,60.2330055236816,60.2676277160645,60.2823753356934,60.2855453491211,60.2133750915527,60.2205543518066,60.2681121826172,60.3016090393066,60.3185081481934,60.3315925598145,60.3540496826172,60.3901824951172,60.4058113098145,60.3575210571289,60.3511695861816],[42.5033226013184,42.4966773986816,42.4559555053711,42.4416999816895,42.4344711303711,42.4374504089355,42.4613304138184,42.4978485107422,42.5308074951172,42.5483856201172,42.5958976745605,42.6216773986816,42.6427268981934,42.635009765625,42.6044425964355,42.575927734375,42.5567359924316,42.525634765625,42.5033226013184],[24.1771984100342,24.1433601379395,24.1531257629395,24.1355476379395,24.1453132629395,24.1697750091553,24.2107906341553,24.2589340209961,24.2151355743408,24.1771984100342],[24.28857421875,24.2812976837158,24.3352527618408,24.3523445129395,24.3767566680908,24.3326168060303,24.28857421875],[24.2585926055908,24.2529296875,24.2909183502197,24.3416004180908,24.3644523620605,24.4110355377197,24.3711910247803,24.3079109191895,24.2808589935303,24.2585926055908],[24.4427719116211,24.42333984375,24.4250984191895,24.4427719116211,24.4710445404053,24.5045890808105,24.50634765625,24.4710445404053,24.4427719116211],[25.6506824493408,25.5693855285645,25.018310546875,24.9791984558105,24.9732913970947,24.9312992095947,24.86669921875,24.8333015441895,24.7955074310303,24.7486820220947,24.73876953125,24.7982406616211,24.8589344024658,24.8720703125,24.8764171600342,24.90771484375,24.9532222747803,24.9702625274658,24.9717769622803,24.951416015625,24.9112796783447,24.8681163787842,24.7812995910645,24.68359375,24.63623046875,24.5773448944092,24.4906253814697,24.4235363006592,24.383544921875,24.3498039245605,24.24267578125,24.22265625,24.2151355743408,24.1426277160645,24.0929679870605,24.0633773803711,24.0414066314697,24.01708984375,24.0241222381592,23.9913578033447,23.9411125183105,23.90966796875,23.885498046875,23.8190441131592,23.724609375,23.6329097747803,23.5187492370605,23.387451171875,23.18994140625,23.0347652435303,22.9229507446289,22.8500003814697,22.7041015625,22.6239261627197,22.6214847564697,22.6311531066895,22.6343746185303,22.6436538696289,22.6580085754395,22.6766605377197,22.69873046875,22.7233390808105,22.7496585845947,22.7768058776855,22.8040046691895,22.830322265625,22.8549308776855,22.8770008087158,22.8956050872803,22.9100093841553,22.9192867279053,22.9225101470947,22.9328117370605,22.9869632720947,23.0524425506592,23.1179695129395,23.1834964752197,23.2489738464355,23.314453125,23.3799800872803,23.4454593658447,23.510986328125,23.5764656066895,23.6419925689697,23.70751953125,23.7729969024658,23.8384761810303,23.9040031433105,23.9695320129395,24.0350112915039,24.078857421875,24.1283187866211,24.2579097747803,24.2861824035645,24.3384284973145,24.3010730743408,24.2504405975342,24.2627925872803,24.25439453125,24.0747547149658,24.0108871459961,23.9853515625,23.9710941314697,23.9952144622803,24.1125011444092,24.1546382904053,24.1473140716553,24.0984382629395,24.0694828033447,24.0770511627197,24.1711902618408,24.2542972564697,24.2781734466553,24.3582534790039,24.4626941680908,24.5309581756592,24.5635261535645,24.6212882995605,24.7155284881592,24.8104496002197,25.0416011810303,25.23681640625,25.2998046875,25.3944835662842,25.4981441497803,25.7939949035645,25.916015625,26.0527839660645,26.0626468658447,26.0681629180908,26.0474605560303,25.9451656341553,25.8484859466553,25.74609375,25.6905269622803,25.6449222564697,25.6253910064697,25.6277332305908,25.6506824493408],[25.2132816314697,25.2088375091553,25.2355461120605,25.2786140441895,25.3008785247803,25.3038082122803,25.2666988372803,25.2132816314697],[-54.7162132263184,-54.7393569946289,-54.7219734191895,-54.7216796875,-54.7298812866211,-54.7423820495605,-54.7229499816895,-54.72509765625,-54.7679672241211,-54.8106460571289,-54.7925758361816,-54.7964820861816,-54.84033203125,-54.8399391174316,-54.9025382995605,-54.8629875183105,-54.8265647888184,-54.7747039794922,-54.7736320495605,-54.7527313232422,-54.7162132263184],[-54.8536148071289,-54.6278343200684,-54.3240203857422,-54.0529289245605,-53.7888679504395,-53.5154304504395,-53.2418937683105,-52.9495124816895,-52.6526374816895,-52.6949234008789,-52.9000968933105,-52.9839820861816,-53.0818367004395,-53.0196304321289,-53.0552749633789,-53.11376953125,-53.17724609375,-53.2218742370605,-53.2609367370605,-53.294921875,-53.3064422607422,-53.3190422058105,-53.5640602111816,-53.6187515258789,-53.6622085571289,-53.787109375,-53.9219741821289,-54.0498046875,-54.1480445861816,-54.2225608825684,-54.3135757446289,-54.4410171508789,-54.5334968566895,-54.5989265441895,-54.6534194946289,-54.6321296691895,-54.6380844116211,-54.6781272888184,-54.7888679504395,-54.8779296875,-54.9146499633789,-54.9281234741211,-54.9263648986816,-54.9099617004395,-54.9193344116211,-54.9567375183105,-54.9752960205078,-54.9777336120605,-55.0093727111816,-55.0321273803711,-55.0132789611816,-54.9249000549316,-54.90380859375,-54.86865234375,-54.8484382629395,-54.8175773620605,-54.8163070678711,-54.8362312316895,-54.8337898254395,-54.8536148071289],[-39.171875,-39.23486328125,-39.2274436950684,-39.1668930053711,-39.1101531982422,-39.0862274169922,-39.1122093200684,-39.1356468200684,-39.171875],[-22.2336921691895,-22.2615222930908,-22.2904300689697,-22.349609375,-22.4391613006592,-22.6124019622803,-22.8694343566895,-23.0592765808105,-23.1820316314697,-23.2687511444092,-23.3194332122803,-23.3604488372803,-23.3919925689697,-23.45751953125,-23.5570316314697,-23.6564464569092,-23.7556648254395,-23.8581047058105,-23.9635734558105,-24.0139656066895,-24.0091800689697,-24.0935554504395,-24.2667961120605,-24.3870105743408,-24.4539051055908,-24.5623054504395,-24.78662109375,-24.8428707122803,-24.8941402435303,-24.9592761993408,-24.9791011810303,-24.9538097381592,-24.9771480560303,-25.0492191314697,-25.1364269256592,-25.3284187316895,-25.4050788879395,-25.4737300872803,-25.5341815948486,-25.5987319946289,-25.6671867370605,-25.6970710754395,-25.7259750366211,-25.78369140625,-25.9069328308105,-25.9642562866211,-26.0065422058105,-26.0529308319092,-26.1385746002197,-26.18017578125,-26.2249031066895,-26.25146484375,-26.2625980377197,-26.3074207305908,-26.3814449310303,-26.4765625,-26.5925788879395,-26.6299800872803,-26.6499996185303,-26.6768550872803,-26.7310543060303,-26.7707023620605,-26.7958984375,-26.8249015808105,-26.8576164245605,-26.8900394439697,-26.9219722747803,-26.9684581756592,-27.0294914245605,-27.083984375,-27.1321296691895,-27.1960945129395,-27.3143558502197,-27.2734375,-27.3166027069092,-27.4304676055908,-27.4701175689697,-27.4357414245605,-27.4406242370605,-27.4846668243408,-27.4937496185303,-27.4678707122803,-27.4878921508789,-27.5538082122803,-27.5374011993408,-27.4387683868408,-27.3667964935303,-27.3214836120605,-27.3077144622803,-27.32568359375,-27.3619136810303,-27.4164066314697,-27.4148445129395,-27.3571300506592,-27.2880859375,-27.2076168060303,-27.1499996185303,-27.1153316497803,-27.068359375,-27.00927734375,-26.97314453125,-26.9601573944092,-26.93115234375,-26.8860359191895,-26.8445320129395,-26.806640625,-26.7593746185303,-26.7025394439697,-26.6667976379395,-26.6522464752197,-26.5329093933105,-26.3087882995605,-26.0057621002197,-25.5760746002197,-25.5764656066895,-25.6082992553711,-25.625,-25.5886726379395,-25.5718746185303,-25.5704097747803,-25.5295886993408,-25.5230484008789,-25.5452156066895,-25.5718746185303,-25.5779304504395,-25.6476554870605,-25.6688480377197,-25.7488269805908,-25.9595699310303,-26.0836925506592,-26.22509765625,-26.2881832122803,-26.3518543243408,-26.4431648254395,-26.66650390625,-26.7486324310303,-26.8046875,-26.8828125,-26.9783191680908,-27.12109375,-27.1595706939697,-27.1611328125,-27.2437496185303,-27.2747077941895,-27.2538089752197,-27.2896480560303,-27.3820323944092,-27.4235363006592,-27.4464855194092,-27.4573249816895,-27.4541015625,-27.4771499633789,-27.5265617370605,-27.544921875,-27.5325202941895,-27.5505847930908,-27.5992183685303,-27.6519527435303,-27.7085933685303,-27.7471694946289,-27.7677745819092,-27.7962875366211,-27.8359375,-27.8667984008789,-27.8988285064697,-27.9559574127197,-28.0377941131592,-28.08935546875,-28.1209945678711,-28.2041015625,-28.2554683685303,-28.3028316497803,-28.3449211120605,-28.3816413879395,-28.3866214752197,-28.3597660064697,-28.3541984558105,-28.3700180053711,-28.3996086120605,-28.4432621002197,-28.4728527069092,-28.4885730743408,-28.5246105194092,-28.5808601379395,-28.6517581939697,-28.7372074127197,-28.8524398803711,-28.9972648620605,-29.0924797058105,-29.1380863189697,-29.2030277252197,-29.2873039245605,-29.4178714752197,-29.5948238372803,-29.7162094116211,-29.7821273803711,-29.8565425872803,-29.9394550323486,-30.0338859558105,-30.1399402618408,-30.1877937316895,-30.2269535064697,-30.2950210571289,-30.3844718933105,-30.4952125549316,-30.5910129547119,-30.7120113372803,-30.8585929870605,-30.91748046875,-30.9374027252197,-30.9751949310303,-31.0310554504395,-31.1043949127197,-31.1953105926514,-31.2994136810303,-31.4166030883789,-31.4949207305908,-31.5343761444092,-31.5761737823486,-31.6206073760986,-31.6849613189697,-31.7692375183105,-31.8318367004395,-31.8726577758789,-31.9242191314697,-31.9865245819092,-32.0515632629395,-32.1190414428711,-32.1848640441895,-32.2489242553711,-32.3218765258789,-32.4716796875,-32.56396484375,-32.9592742919922,-33.0146484375,-33.0783195495605,-33.0829086303711,-33.0718765258789,-33.1115226745605,-33.2859382629395,-33.6634750366211,-33.7530250549316,-33.8983383178711,-33.9909172058105,-34.0607414245605,-34.1929702758789,-34.2525367736816,-34.2629890441895,-34.2961883544922,-34.4574241638184,-34.5316390991211,-34.6834945678711,-34.89453125,-35.0189476013184,-35.1884765625,-35.3625030517578,-35.5059547424316,-35.7203102111816,-35.9002952575684,-36.0267601013184,-36.1441421508789,-36.2967758178711,-36.3525390625,-36.3464813232422,-36.3890609741211,-36.4264640808105,-36.7352523803711,-36.8512725830078,-36.9577178955078,-37.4463844299316,-37.74462890625,-37.9092788696289,-38.0856437683105,-38.1696281433105,-38.4358406066895,-38.67333984375,-38.7966804504395,-38.8381843566895,-38.9739265441895,-38.9929695129395,-38.9808578491211,-38.9988288879395,-38.9618186950684,-38.9191436767578,-38.8132820129395,-38.8000984191895,-38.8529281616211,-38.9880867004395,-39.1505851745605,-39.2432632446289,-39.2618141174316,-39.3097686767578,-39.3738288879395,-39.3804702758789,-39.4315414428711,-39.4615249633789,-39.568359375,-39.8253936767578,-39.8804702758789,-39.8953132629395,-39.95068359375,-40.1965827941895,-40.35595703125,-40.4587898254395,-40.6746101379395,-40.8146476745605,-40.8908195495605,-41.0471687316895,-41.1096687316895,-41.1524429321289,-41.1597633361816,-41.1500015258789,-41.0078125,-40.9224586486816,-40.8544921875,-40.8137664794922,-40.7932586669922,-40.7565460205078,-40.73583984375,-40.7313499450684,-40.8052749633789,-40.8806648254395,-40.9469757080078,-41.1056632995605,-41.23876953125,-41.56689453125,-41.7451171875,-41.9699211120605,-42.10205078125,-42.1618156433105,-42.2208023071289,-42.2610321044922,-42.2545890808105,-42.2702140808105,-42.2992172241211,-42.35595703125,-42.416015625,-42.4337882995605,-42.4216804504395,-42.3951187133789,-42.3534202575684,-42.26611328125,-42.2507820129395,-42.21826171875,-42.1828117370605,-42.1246070861816,-42.1138687133789,-42.1529273986816,-42.1886711120605,-42.28271484375,-42.4065437316895,-42.5555648803711,-42.69580078125,-42.7457046508789,-42.8052749633789,-42.8812484741211,-42.8614234924316,-42.7555656433105,-42.6460914611816,-42.572265625,-42.5134773254395,-42.5314445495605,-42.6332054138184,-42.6663093566895,-42.7588882446289,-42.9089851379395,-42.95068359375,-42.94921875,-42.9689445495605,-43.0246086120605,-43.0591773986816,-43.1355476379395,-43.1888656616211,-43.2935523986816,-43.5220718383789,-43.5718727111816,-43.6299819946289,-43.7874984741211,-44.0487289428711,-44.158203125,-44.2796859741211,-44.3607444763184,-44.4773445129395,-44.6614265441895,-44.7961921691895,-44.8755874633789,-44.9450187683105,-45.0078125,-45.0071296691895,-44.9647483825684,-45.0335922241211,-45.1175804138184,-45.1578140258789,-45.1829109191895,-45.2276382446289,-45.25732421875,-45.5772476196289,-45.7755851745605,-45.9701156616211,-46.0525398254395,-46.1667938232422,-46.26953125,-46.3454093933105,-46.4427719116211,-46.5538063049316,-47.005859375,-47.0453109741211,-47.09375,-47.15673828125,-47.2567405700684,-47.3449211120605,-47.568359375,-47.63818359375,-47.7015647888184,-47.7832984924316,-47.8267555236816,-47.8576164245605,-47.8532257080078,-47.8267555236816,-47.8532257080078,-47.9411125183105,-47.9767570495605,-48.0193367004395,-48.0842781066895,-48.3423805236816,-48.4195327758789,-48.52294921875,-48.6277351379395,-48.6878890991211,-48.8142585754395,-48.9517593383789,-49.2466773986816,-49.3040046691895,-49.3421897888184,-49.85888671875,-49.9196281433105,-49.9844703674316,-50.0914039611816,-50.1045913696289,-50.0426750183105,-49.9779281616211,-49.8669929504395,-49.7525405883789,-49.79345703125,-49.8629875183105,-49.9357414245605,-49.96875,-50.0030250549316,-49.9876937866211,-50.0094718933105,-50.0361328125,-50.0914039611816,-50.1579093933105,-50.1947250366211,-50.2251968383789,-50.2811508178711,-50.38232421875,-50.4991188049316,-50.5831069946289,-50.7525405883789,-50.8644523620605,-50.9505844116211,-51.0281257629395,-51.0458030700684,-51.0061531066895,-50.99365234375,-51.0969734191895,-51.3035163879395,-51.4464836120605,-51.4889640808105,-51.5471687316895,-51.5612297058105,-51.5594711303711,-51.58447265625,-51.6102561950684,-51.60107421875,-51.6623039245605,-51.63623046875,-51.6771507263184,-51.7146492004395,-52.0130844116211,-52.1975593566895,-52.3070297241211,-52.3566398620605,-52.2904281616211,-52.2733383178711,-52.2554702758789,-52.2081031799316,-52.1361312866211,-52.1361312866211,-52.0753898620605,-52.0082015991211,-52.0022468566895,-51.9981460571289,-51.9939460754395,-51.9913063049316,-51.9895515441895,-51.9641609191895,-51.88037109375,-51.8186492919922,-51.7440452575684,-51.6911125183105,-51.6203117370605,-51.5408210754395,-51.4703102111816,-51.2989234924316,-51.2233390808105,-51.17041015625,-51.0954093933105,-51.0601539611816,-51.0333976745605,-50.9102516174316,-50.78955078125,-50.6818389892578,-50.63427734375,-50.6117172241211,-50.6075172424316,-50.6476554870605,-50.6376953125,-50.6531219482422,-50.6964836120605,-50.7603492736816,-50.73828125,-50.6700172424316,-50.6107406616211,-50.5584945678711,-50.4725570678711,-50.3619155883789,-50.2311515808105,-50.1252937316895,-50.0302734375,-49.9109382629395,-49.7945327758789,-49.6980476379395,-49.5829124450684,-49.4638671875,-49.3976554870605,-49.3138656616211,-49.3006858825684,-49.18798828125,-49.0968742370605,-49.0143547058105,-48.9767608642578,-48.9439430236816,-48.8962898254395,-48.8415985107422,-48.7928695678711,-48.7296867370605,-48.6624984741211,-48.5193328857422,-48.4753875732422,-48.4173851013184,-48.3658180236816,-48.2291030883789,-48.1100578308105,-48.0159187316895,-47.9733390808105,-47.8763694763184,-47.7841796875,-47.685546875,-47.5720710754395,-47.49267578125,-47.4462890625,-47.3427734375,-47.2414054870605,-47.2138671875,-47.20166015625,-47.1443367004395,-47.0874977111816,-47.0160179138184,-46.9368171691895,-46.8312492370605,-46.7915992736816,-46.7058563232422,-46.6513671875,-46.5784187316895,-46.4278335571289,-46.31982421875,-46.2799835205078,-46.2067375183105,-46.1605453491211,-46.1027336120605,-46.0418968200684,-45.9537124633789,-45.8787117004395,-45.8390617370605,-45.7244148254395,-45.5789070129395,-45.5347633361816,-45.5126953125,-45.4376983642578,-45.3319358825684,-45.23046875,-45.1682624816895,-45.06787109375,-44.9791984558105,-44.9306640625,-44.9041976928711,-44.8204116821289,-44.7718772888184,-44.79150390625,-44.7744140625,-44.7704124450684,-44.7620086669922,-44.7498054504395,-44.78515625,-44.7630844116211,-44.6307601928711,-44.5602531433105,-44.4940452575684,-44.4412117004395,-44.4249038696289,-44.3831062316895,-44.3301734924316,-44.2414054870605,-44.1507797241211,-44.1060562133789,-44.0666999816895,-43.9844703674316,-43.9295921325684,-43.8583984375,-43.7532234191895,-43.7046852111816,-43.6467781066895,-43.5901374816895,-43.5271492004395,-43.4401359558105,-43.3475608825684,-43.3229522705078,-43.2946281433105,-43.2373046875,-43.1667938232422,-43.1453094482422,-43.1019515991211,-43.0656242370605,-42.9900398254395,-42.7767562866211,-42.6482391357422,-42.5771484375,-42.5224609375,-42.4732398986816,-42.3584976196289,-42.29833984375,-42.2518539428711,-42.2053718566895,-42.14794921875,-42.13427734375,-42.1670875549316,-42.1478500366211,-42.1014633178711,-42.0467758178711,-41.9685554504395,-41.77197265625,-41.650390625,-41.6066398620605,-41.560546875,-41.3933601379395,-41.2923851013184,-40.99462890625,-40.8929672241211,-40.7891578674316,-40.6916999816895,-40.62060546875,-40.5244140625,-40.4391593933105,-40.40087890625,-40.3817367553711,-40.3352546691895,-40.2997093200684,-40.2443351745605,-40.1766624450684,-40.1247062683105,-40.0946273803711,-40.0949211120605,-40.0208015441895,-39.92919921875,-39.8868141174316,-39.8333015441895,-39.70703125,-39.63525390625,-39.6051750183105,-39.59423828125,-39.6111335754395,-39.6024398803711,-39.5641593933105,-39.5231475830078,-39.4952125549316,-39.40234375,-39.2872085571289,-39.2059593200684,-38.9856452941895,-38.93505859375,-38.8888664245605,-38.8454093933105,-38.8093757629395,-38.7575187683105,-38.7384757995605,-38.6810569763184,-38.6044921875,-38.5416030883789,-38.4978523254395,-38.4458999633789,-38.3148460388184,-38.1939468383789,-38.0412101745605,-37.9099617004395,-37.7623062133789,-37.6310539245605,-37.5591812133789,-37.4451141357422,-37.3932609558105,-37.30029296875,-37.2274398803711,-37.1143569946289,-37.0569305419922,-36.9202156066895,-36.8436508178711,-36.7616233825684,-36.68505859375,-36.64404296875,-36.5780258178711,-36.5237312316895,-36.4873046875,-36.419921875,-36.4117164611816,-36.4117164611816,-36.392578125,-36.3406257629395,-36.283203125,-36.2119140625,-36.1463851928711,-36.1327133178711,-36.0617179870605,-35.9705085754395,-35.8785171508789,-35.7718772888184,-35.6091804504395,-35.5230445861816,-35.4519538879395,-35.3753890991211,-35.326171875,-35.3079109191895,-35.2468719482422,-35.216796875,-35.1936531066895,-35.1468772888184,-34.9217758178711,-34.85498046875,-34.7745132446289,-34.7328147888184,-34.6726570129395,-34.5812530517578,-34.4928703308105,-34.4320335388184,-34.3500022888184,-34.30078125,-34.2762680053711,-34.2699203491211,-34.25439453125,-34.2243156433105,-34.1804695129395,-34.0835914611816,-33.9297828674316,-33.7313461303711,-33.6009750366211,-33.4697265625,-33.3986320495605,-33.3439445495605,-33.2837867736816,-33.2509765625,-33.2793922424316,-33.271484375,-33.2017593383789,-33.1279296875,-33.0267562866211,-32.9636726379395,-32.8845710754395,-32.8599624633789,-32.8074188232422,-32.6260757446289,-32.4716796875,-32.4306640625,-32.3099594116211,-32.2666969299316,-32.1764640808105,-32.08349609375,-32.0423851013184,-32.0310516357422,-31.9577140808105,-31.9165992736816,-31.8810558319092,-31.8837909698486,-31.8418941497803,-31.6664066314697,-31.5694351196289,-31.4279308319092,-31.3173847198486,-31.2228507995605,-31.1484394073486,-31.11279296875,-31.1292972564697,-31.12109375,-31.0604496002197,-31.0226535797119,-30.9925785064697,-30.9597644805908,-30.90234375,-30.833984375,-30.6772441864014,-30.5046863555908,-30.4402332305908,-30.3855476379395,-30.3609390258789,-30.3882808685303,-30.3582038879395,-30.2816410064697,-30.2132797241211,-30.1750011444092,-30.1203117370605,-30.1039047241211,-30.0783195495605,-30.0164070129395,-29.8740253448486,-29.7691402435303,-29.54541015625,-29.3240242004395,-29.2500019073486,-29.1488304138184,-29.1032238006592,-29.0455093383789,-28.7838859558105,-28.64111328125,-28.56201171875,-28.41357421875,-28.28564453125,-28.2008800506592,-28.1926765441895,-28.1653308868408,-28.07080078125,-27.9736328125,-27.9247074127197,-27.84814453125,-27.7435550689697,-27.5700187683105,-27.4490242004395,-27.4051761627197,-27.2466793060303,-27.1537113189697,-27.1154289245605,-27.1044921875,-27.1483402252197,-27.1400375366211,-27.0853500366211,-27.0481452941895,-27.0279293060303,-26.9732418060303,-26.8775386810303,-26.8064460754395,-26.6703128814697,-26.5183601379395,-26.4704093933105,-26.4180660247803,-26.3519535064697,-26.2769527435303,-26.1537113189697,-26.0654296875,-25.7410163879395,-25.6515617370605,-25.4856452941895,-25.4200191497803,-25.2367191314697,-25.1629886627197,-25.1493167877197,-25.1247081756592,-25.0918960571289,-25.0509777069092,-24.9989261627197,-24.9251937866211,-24.8992195129395,-24.8376960754395,-24.7473640441895,-24.6297855377197,-24.5969734191895,-24.5451164245605,-24.4972648620605,-24.4603519439697,-24.3919925689697,-24.3083000183105,-24.2433586120605,-24.1189441680908,-24.0337886810303,-23.9748039245605,-23.9346675872803,-23.6339836120605,-23.2451171875,-23.0013675689697,-22.8216800689697,-22.7738285064697,-22.65087890625,-22.55224609375,-22.5098628997803,-22.40966796875,-22.3430671691895,-22.2693347930908,-22.21630859375,-22.2053718566895,-22.1583995819092,-22.11376953125,-22.0531253814697,-21.9474601745605,-21.8304691314697,-21.8025398254395,-21.8056640625,-21.8350582122803,-21.8794918060303,-22.0197277069092,-22.099609375,-22.1102542877197,-22.0945320129395,-22.09814453125,-22.1027355194092,-22.1096668243408,-22.1439456939697,-22.1712894439697,-22.185546875,-22.2288074493408,-22.37158203125,-22.4853515625,-22.5853519439697,-22.7610359191895,-22.82763671875,-22.7953128814697,-22.6033210754395,-22.4913101196289,-22.3658199310303,-22.0725593566895,-22.0286140441895,-22.0072269439697,-22.0054683685303,-22.0272464752197,-22.0275402069092,-22.0042953491211,-22.0005855560303,-21.9972667694092,-21.9991207122803,-22.0496101379395,-22.1598644256592,-22.2179679870605,-22.2336921691895],[40.6160659790039,40.5995597839355,40.6069793701172,40.6483421325684,40.6648445129395,40.6640129089355,40.649169921875,40.6160659790039],[38.9066886901855,38.9126472473145,38.8690414428711,38.8777809143066,38.9548797607422,39.0175323486328,39.1781272888184,39.243896484375,39.2819328308105,39.3501968383789,39.3784637451172,39.417236328125,39.4881324768066,39.5456085205078,39.5629386901855,39.5640602111816,39.5498046875,39.4944839477539,39.5298843383789,39.5655746459961,39.595458984375,39.5706062316895,39.5826683044434,39.6565895080566,39.6963386535645,39.7428207397461,39.76513671875,39.7191429138184,39.7035140991211,39.7464866638184,39.8875961303711,39.9957504272461,40.0403823852539,40.0357398986816,40.0141105651855,40.0231437683105,40.0702629089355,40.1263694763184,40.149658203125,40.16650390625,40.2393074035645,40.3565940856934,40.393310546875,40.4440422058105,40.4823722839355,40.5740737915039,40.6477546691895,40.7195320129395,40.7941398620605,40.9290046691895,40.9682121276855,41.0048332214355,41.064697265625,41.1071281433105,41.1155281066895,41.1166496276855,41.131103515625,41.1589813232422,41.1825180053711,41.203369140625,41.2133293151855,41.1910171508789,41.2082023620605,41.2113761901855,41.2201690673828,41.2485847473145,41.2593765258789,41.2774887084961,41.2909660339355,41.2457008361816,41.1954612731934,41.1751480102539,41.1474113464355,41.1263694763184,41.100830078125,41.088134765625,41.0755844116211,41.0693359375,41.0062484741211,41.0048828125,40.9856948852539,40.947998046875,40.896728515625,40.846923828125,40.8297386169434,40.8044929504395,40.7071266174316,40.673583984375,40.6380844116211,40.5323753356934,40.4168434143066,40.3291015625,40.2337875366211,40.1748046875,40.1046867370605,40.0570793151855,40.0248527526855,40.0112800598145,40.0142097473145,40.0028343200684,39.989013671875,39.9775390625,39.9561996459961,39.881103515625,39.808349609375,39.7765617370605,39.7185554504395,39.6644515991211,39.594482421875,39.617431640625,39.55517578125,39.5128440856934,39.47509765625,39.4338836669922,39.4167976379395,39.4024887084961,39.3822746276855,39.3485336303711,39.2985343933105,39.2236824035645,39.201416015625,39.1981925964355,39.2073707580566,39.1921882629395,39.1676750183105,39.1108894348145,39.0694351196289,38.9974594116211,38.9066886901855],[41.0272445678711,41.0158195495605,41.0272445678711,41.0538558959961,41.0779800415039,41.0855941772461,41.0792465209961,41.0526351928711,41.0272445678711],[-14.3511714935303,-14.3597660064697,-14.3121089935303,-14.2759761810303,-14.2574214935303,-14.2667970657349,-14.2822265625,-14.3511714935303],[-83.1187515258789,-83.1417922973633,-83.0967788696289,-83.0596694946289,-83.0215835571289,-82.9876937866211,-82.9685516357422,-82.9273452758789,-82.9022445678711,-82.8792953491211,-82.8568344116211,-82.8426742553711,-82.8648452758789,-82.8992233276367,-82.9227523803711,-83.0269546508789,-83.0425796508789,-83.1187515258789],[-82.10498046875,-82.1120147705078,-82.0584945678711,-81.9488296508789,-81.87548828125,-81.8186492919922,-81.779296875,-81.77978515625,-81.9004898071289,-81.947265625,-82.03076171875,-82.10498046875],[-81.5894546508789,-81.5978546142578,-81.482421875,-81.44482421875,-81.4236373901367,-81.4040985107422,-81.3909149169922,-81.3521499633789,-81.3240203857422,-81.2814407348633,-81.3132781982422,-81.3966827392578,-81.4634780883789,-81.5220718383789,-81.5669937133789,-81.5894546508789],[-80.0333023071289,-80.0360336303711,-80.0260772705078,-80.00048828125,-79.96630859375,-79.8998031616211,-79.8464813232422,-79.82958984375,-79.7655258178711,-79.8506851196289,-79.935546875,-79.9876937866211,-80.0333023071289],[-80.3441467285156,-80.55810546875,-80.6565399169922,-80.7332992553711,-80.7743148803711,-80.8407287597656,-80.88134765625,-80.9481430053711,-80.8890609741211,-80.8342742919922,-80.7658157348633,-80.7345733642578,-80.62744140625,-80.5948257446289,-80.6169891357422,-80.650390625,-80.6678771972656,-80.7339859008789,-80.74853515625,-80.607421875,-80.4419937133789,-80.3576202392578,-80.3185577392578,-80.2938461303711,-80.2604446411133,-80.2385787963867,-80.2244186401367,-80.2222671508789,-80.2347640991211,-80.2582092285156,-80.34619140625,-80.3612289428711,-80.3844757080078,-80.3733444213867,-80.36865234375,-80.3441467285156,-80.3065414428711,-80.2566452026367,-80.2438507080078,-80.2058563232422,-80.1343765258789,-80.0693359375,-80.0197296142578,-79.9958038330078,-79.9783172607422,-79.9505844116211,-79.8868179321289,-79.8622055053711,-79.8088912963867,-79.7410202026367,-79.7769546508789,-79.8752899169922,-79.93798828125,-80.0010757446289,-80.1009750366211,-80.1150360107422,-80.15087890625,-80.1961975097656,-80.1976547241211,-80.2080078125,-80.2554702758789,-80.3153305053711,-80.3441467285156],[-79.7984390258789,-79.8184585571289,-79.7712860107422,-79.7258834838867,-79.6473617553711,-79.7331085205078,-79.8232421875,-79.9095764160156,-79.9298858642578,-79.9259796142578,-79.9361343383789,-80.0108337402344,-79.9384765625,-79.8876953125,-79.8495178222656,-79.732421875,-79.63427734375,-79.6447219848633,-79.7984390258789],[-79.6736297607422,-79.6828079223633,-79.6583023071289,-79.59228515625,-79.53466796875,-79.5298843383789,-79.56787109375,-79.6038055419922,-79.6070327758789,-79.6736297607422],[-80.0778350830078,-80.0814514160156,-80.0750961303711,-80.0542984008789,-79.9733428955078,-79.9088897705078,-79.8726577758789,-79.7933578491211,-79.7618179321289,-79.6202087402344,-79.5894546508789,-79.5458984375,-79.5284194946289,-79.5603485107422,-79.5686492919922,-79.5822296142578,-79.6080093383789,-79.6120071411133,-79.6244125366211,-79.7377014160156,-79.7708053588867,-79.8369140625,-79.9542999267578,-80.04052734375,-80.0540008544922,-80.0778350830078],[-79.45263671875,-79.5011749267578,-79.478515625,-79.4639663696289,-79.4374008178711,-79.3331985473633,-79.2982406616211,-79.2849578857422,-79.2271499633789,-79.2140579223633,-79.2229461669922,-79.2684555053711,-79.3115234375,-79.32763671875,-79.39404296875,-79.45263671875],[-79.3204040527344,-79.3571243286133,-79.2940444946289,-79.28271484375,-79.2087860107422,-79.1957015991211,-79.18115234375,-79.14892578125,-79.0918960571289,-79.0900421142578,-79.2229461669922,-79.2785186767578,-79.3053741455078,-79.3204040527344],[-79.8074188232422,-79.8445281982422,-79.81201171875,-79.7035140991211,-79.5047836303711,-79.4419937133789,-79.3887710571289,-79.32080078125,-79.2455062866211,-79.1376953125,-79.05078125,-78.9953079223633,-78.9288024902344,-78.8557662963867,-78.7798767089844,-78.7220687866211,-78.7191390991211,-78.7251968383789,-78.7417984008789,-78.7601547241211,-78.79345703125,-78.9009780883789,-79.0070343017578,-79.1316452026367,-79.271484375,-79.3243179321289,-79.4024429321289,-79.5081100463867,-79.5452194213867,-79.5910110473633,-79.6373062133789,-79.6745147705078,-79.6945343017578,-79.7351608276367,-79.7804718017578,-79.8074188232422],[-79.6798782348633,-79.6830062866211,-79.6744155883789,-79.6402359008789,-79.6236343383789,-79.5680618286133,-79.53857421875,-79.5005874633789,-79.4443359375,-79.2457046508789,-79.12890625,-79.0596618652344,-78.9015579223633,-78.8836898803711,-78.8093719482422,-78.7691421508789,-78.6861343383789,-78.3625030517578,-78.3157196044922,-78.3497085571289,-78.3836898803711,-78.4216766357422,-78.51416015625,-78.5695343017578,-78.6824188232422,-78.8708953857422,-79.01318359375,-79.2103500366211,-79.27978515625,-79.443359375,-79.517578125,-79.6183624267578,-79.666015625,-79.6798782348633],[-78.1414108276367,-78.2160110473633,-78.2490234375,-78.2224578857422,-78.2842712402344,-78.3064422607422,-78.2746124267578,-78.21337890625,-78.1312484741211,-78.1019515991211,-78.0896453857422,-78.1480484008789,-78.1979446411133,-78.1963806152344,-78.1299819946289,-78.0457992553711,-78.0064468383789,-77.9923782348633,-78.0222702026367,-78.1117172241211,-78.1414108276367],[-78.8107452392578,-78.8142547607422,-78.8042984008789,-78.806640625,-78.8184585571289,-78.8460922241211,-78.9019546508789,-78.9896469116211,-79.0847625732422,-79.1630859375,-79.2999954223633,-79.3500061035156,-79.47705078125,-79.5791015625,-79.80126953125,-79.8914031982422,-79.9786148071289,-79.9693374633789,-79.9739303588867,-79.9901351928711,-80.0029296875,-80.0205078125,-80.0955123901367,-80.123046875,-80.1550750732422,-80.1780242919922,-80.19140625,-80.6427764892578,-80.6669006347656,-80.7123031616211,-80.7532272338867,-80.7841796875,-80.8701171875,-80.8638687133789,-80.8469772338867,-80.7603530883789,-80.6236343383789,-80.5694274902344,-80.5165023803711,-80.4875030517578,-80.28369140625,-80.18896484375,-80.1087875366211,-80.1144485473633,-80.1609344482422,-80.1750030517578,-80.1559600830078,-80.0990219116211,-80.0665969848633,-80.0778350830078,-80.1260757446289,-80.1412124633789,-79.9898376464844,-79.8197250366211,-79.6267547607422,-79.51171875,-79.4794921875,-79.4296875,-79.3814468383789,-79.3381881713867,-79.3212890625,-79.3132858276367,-79.28271484375,-79.2328109741211,-79.1722640991211,-79.1043930053711,-79.0217742919922,-78.9798812866211,-78.9499053955078,-78.9228515625,-78.8822250366211,-78.8332977294922,-78.8182601928711,-78.7804641723633,-78.6959991455078,-78.6052703857422,-78.5567398071289,-78.4622116088867,-78.2224578857422,-78.0938491821289,-78.0474548339844,-77.8401336669922,-77.8190460205078,-77.79052734375,-77.7852554321289,-77.8048858642578,-77.8268585205078,-77.8814468383789,-77.9883804321289,-78.03515625,-78.0928726196289,-78.1672897338867,-78.2584991455078,-78.2865219116211,-78.3363265991211,-78.3850631713867,-78.4326171875,-78.5298843383789,-78.5975646972656,-78.6614227294922,-78.68701171875,-78.7398376464844,-78.7912139892578,-78.8107452392578],[-77.3363265991211,-77.3868179321289,-77.37060546875,-77.3009796142578,-77.2747039794922,-77.2799835205078,-77.3150329589844,-77.3363265991211],[-77.369140625,-77.3737258911133,-77.2962875366211,-77.2733383178711,-77.2264633178711,-77.2200241088867,-77.3350524902344,-77.3490219116211,-77.369140625],[-77.3216857910156,-77.3943328857422,-77.3861312866211,-77.4546890258789,-77.5247116088867,-77.560546875,-77.6533203125,-77.6812515258789,-77.6825180053711,-77.6441421508789,-77.6486282348633,-77.70263671875,-77.7564468383789,-77.8509750366211,-77.7740249633789,-77.7003860473633,-77.5474624633789,-77.5246047973633,-77.4940414428711,-77.4437484741211,-77.3767623901367,-77.3355484008789,-77.2888717651367,-77.251953125,-77.1893539428711,-77.1617202758789,-77.1865234375,-77.2706069946289,-77.3074188232422,-77.3216857910156],[-77.0068359375,-77.0191421508789,-76.9930648803711,-76.9158172607422,-76.9001998901367,-76.9356460571289,-76.95458984375,-76.9771499633789,-76.9977569580078,-77.0068359375],[-77.1205062866211,-77.1366195678711,-77.1285171508789,-77.1141586303711,-77.0994110107422,-77.07568359375,-77.0042953491211,-76.9816436767578,-76.9484405517578,-76.9260787963867,-76.8987274169922,-76.875,-76.8892593383789,-76.9270477294922,-77.0016632080078,-77.0492172241211,-77.0789031982422,-77.0931625366211,-77.1205062866211],[-76.9613265991211,-77.0056610107422,-76.9927749633789,-76.9296875,-76.8865280151367,-76.8662109375,-76.8373031616211,-76.8371047973633,-76.8831024169922,-76.9485321044922,-76.9613265991211],[-76.7173843383789,-76.7300796508789,-76.7205123901367,-76.7444381713867,-76.7716827392578,-76.794921875,-76.8020477294922,-76.8407211303711,-76.8453140258789,-76.8177719116211,-76.7571258544922,-76.7173843383789],[-76.7764663696289,-76.7889633178711,-76.7615280151367,-76.7367172241211,-76.7141571044922,-76.6913070678711,-76.7188491821289,-76.7351531982422,-76.7764663696289],[-76.6498031616211,-76.6627960205078,-76.6534194946289,-76.59716796875,-76.5771484375,-76.5580062866211,-76.5768585205078,-76.61083984375,-76.6498031616211],[-76.6331024169922,-76.7140655517578,-76.6708984375,-76.6188507080078,-76.5525436401367,-76.5315475463867,-76.5549774169922,-76.5632781982422,-76.6331024169922],[-76.2463836669922,-76.2609405517578,-76.1974639892578,-76.1466751098633,-76.1045913696289,-76.0902328491211,-76.0734405517578,-76.0627975463867,-76.09814453125,-76.2463836669922],[-75.5670852661133,-75.6961898803711,-75.69189453125,-75.6685562133789,-75.5966796875,-75.5662078857422,-75.5573196411133,-75.5670852661133],[-75.7112350463867,-75.7161102294922,-75.6926727294922,-75.6119155883789,-75.5735397338867,-75.5333938598633,-75.5047836303711,-75.5138702392578,-75.5879898071289,-75.6414108276367,-75.6828155517578,-75.7112350463867],[-74.8327102661133,-74.8327102661133,-74.8028335571289,-74.7337951660156,-74.7115249633789,-74.62890625,-74.6077117919922,-74.73193359375,-74.7527313232422,-74.8327102661133],[-74.44189453125,-74.4984359741211,-74.4617156982422,-74.4215850830078,-74.3866195678711,-74.4099578857422,-74.44189453125],[-74.5837860107422,-74.6047821044922,-74.5537109375,-74.5425796508789,-74.5143585205078,-74.4886703491211,-74.4638671875,-74.42578125,-74.34912109375,-74.3238296508789,-74.3296890258789,-74.3672866821289,-74.41357421875,-74.4140625,-74.4562454223633,-74.5150451660156,-74.5837860107422],[-74.6226577758789,-74.6405258178711,-74.5746078491211,-74.5425796508789,-74.4894561767578,-74.4782180786133,-74.4662094116211,-74.3274459838867,-74.3122024536133,-74.3318328857422,-74.4052734375,-74.4251937866211,-74.4801712036133,-74.5784225463867,-74.6226577758789],[-74.1650390625,-74.1927719116211,-74.1609344482422,-74.1224594116211,-74.0828170776367,-73.8655242919922,-73.8956069946289,-73.9104461669922,-74.0176773071289,-74.0555648803711,-74.0685501098633,-74.0832977294922,-74.0955123901367,-74.1256866455078,-74.1650390625],[-74.1102600097656,-74.1220703125,-74.11962890625,-74.0972671508789,-74.08154296875,-74.0291976928711,-74.0061492919922,-73.9893569946289,-73.8146438598633,-73.8092803955078,-73.77490234375,-73.7776336669922,-73.8094787597656,-73.8342742919922,-73.8780212402344,-73.9669952392578,-73.9976577758789,-74.076171875,-74.1102600097656],[-73.7560577392578,-73.8558578491211,-73.9219665527344,-73.9891586303711,-74.1570281982422,-74.1734390258789,-74.2599639892578,-74.279296875,-74.2928695678711,-74.3263626098633,-74.3373031616211,-74.4031219482422,-74.3426742553711,-74.3020477294922,-74.2403335571289,-74.22705078125,-74.2186508178711,-74.1904296875,-74.1412124633789,-74.0990219116211,-73.99365234375,-73.9655227661133,-73.8666076660156,-73.8441390991211,-73.8375930786133,-73.8493194580078,-73.8430633544922,-73.8030319213867,-73.7386779785156,-73.6822204589844,-73.6729507446289,-73.6796875,-73.681640625,-73.7118148803711,-73.7328109741211,-73.751953125,-73.7560577392578],[-73.8866195678711,-74.10498046875,-74.1505889892578,-74.1968765258789,-74.3173828125,-74.4085006713867,-74.443359375,-74.49267578125,-74.48095703125,-74.4547882080078,-74.4377899169922,-74.22509765625,-74.1760711669922,-74.1326217651367,-74.091796875,-74.1065444946289,-74.056640625,-73.9893569946289,-73.9398422241211,-73.8800811767578,-73.7904281616211,-73.6766662597656,-73.6251983642578,-73.6192321777344,-73.7118148803711,-73.7978515625,-73.8866195678711],[-73.60498046875,-73.6252899169922,-73.5615234375,-73.5394515991211,-73.4180679321289,-73.3790969848633,-73.3460922241211,-73.3204040527344,-73.32421875,-73.4586944580078,-73.5143585205078,-73.5680618286133,-73.60498046875],[-73.2862319946289,-73.3148422241211,-73.3568420410156,-73.3882751464844,-73.4056625366211,-73.4532241821289,-73.5364227294922,-73.5625,-73.6178665161133,-73.6664047241211,-73.6905288696289,-73.7107467651367,-73.7027359008789,-73.7183609008789,-73.7486343383789,-73.7801742553711,-73.8019561767578,-73.8221664428711,-73.8201141357422,-73.7955093383789,-73.80078125,-73.8259811401367,-73.8297805786133,-73.7496109008789,-73.7352523803711,-73.73974609375,-73.833984375,-73.88037109375,-73.9068374633789,-73.9442367553711,-73.9710922241211,-74.0080108642578,-74.0570297241211,-74.0762710571289,-74.1173782348633,-74.1682586669922,-74.2256774902344,-74.2561492919922,-74.2560577392578,-74.2255859375,-74.2082977294922,-74.1824264526367,-74.0699234008789,-74.0626983642578,-74.0354461669922,-73.95458984375,-73.8909225463867,-73.8121109008789,-73.7462921142578,-73.7001953125,-73.68017578125,-73.669921875,-73.6536102294922,-73.6573257446289,-73.6767578125,-73.7257766723633,-73.7341766357422,-73.7244110107422,-73.7134780883789,-73.5855484008789,-73.5674819946289,-73.5163116455078,-73.4468765258789,-73.3822250366211,-73.3040008544922,-73.2943344116211,-73.3080139160156,-73.2908172607422,-73.2789077758789,-73.2862319946289],[-73.3568420410156,-73.3760757446289,-73.3655242919922,-73.3154296875,-73.2766571044922,-73.2493209838867,-73.22021484375,-73.11328125,-73.1003952026367,-73.1238250732422,-73.2249984741211,-73.2962875366211,-73.3568420410156],[-73.1663131713867,-73.2121047973633,-73.2005844116211,-73.1259765625,-73.0263671875,-72.9915084838867,-72.9659194946289,-72.9410171508789,-73.1663131713867],[-73.0984420776367,-73.2291946411133,-73.2439422607422,-73.2752914428711,-73.3114242553711,-73.3277359008789,-73.369140625,-73.4271469116211,-73.4642562866211,-73.5654296875,-73.611328125,-73.3326187133789,-73.2879943847656,-73.2546844482422,-73.2028350830078,-73.1504821777344,-73.1087875366211,-73.08544921875,-73.0563430786133,-73.0542984008789,-73.10888671875,-73.10107421875,-73.0515670776367,-73.0503921508789,-73.0093765258789,-72.9942398071289,-72.95263671875,-72.9110336303711,-72.8793029785156,-72.8204116821289,-72.8937530517578,-72.9189453125,-72.9512710571289,-72.9970703125,-73.0984420776367],[-72.9197311401367,-72.9202117919922,-72.8838806152344,-72.8600616455078,-72.8230438232422,-72.8197250366211,-72.8820343017578,-72.9076156616211,-72.9197311401367],[-73.1822280883789,-73.2071304321289,-73.1955108642578,-72.9678726196289,-72.9094772338867,-72.8678741455078,-72.7536087036133,-72.6237258911133,-72.5938491821289,-72.54736328125,-72.5563507080078,-72.6106414794922,-72.6810531616211,-72.7317428588867,-72.8236312866211,-72.85400390625,-72.9166030883789,-72.9929656982422,-73.083984375,-73.1365280151367,-73.1822280883789],[-72.6650390625,-72.6698226928711,-72.6468734741211,-72.5994110107422,-72.5171890258789,-72.4680709838867,-72.4757766723633,-72.49169921875,-72.5891571044922,-72.6125946044922,-72.6650390625],[-72.6791000366211,-72.7028274536133,-72.7021484375,-72.65234375,-72.5822296142578,-72.5308609008789,-72.4735336303711,-72.4585952758789,-72.47802734375,-72.5999984741211,-72.6791000366211],[-72.3001022338867,-72.3001937866211,-72.2604522705078,-72.1901321411133,-72.1031265258789,-72.0891647338867,-72.1654281616211,-72.2287139892578,-72.2759780883789,-72.3001022338867],[-72.17333984375,-72.1960983276367,-72.1876983642578,-72.1371078491211,-72.0989227294922,-72.064453125,-72.0146484375,-72.0006866455078,-72.0065460205078,-72.0712890625,-72.1400451660156,-72.17333984375],[-71.9177703857422,-72.0466766357422,-72.0441360473633,-71.96826171875,-71.9219741821289,-71.8939437866211,-71.9078140258789,-71.9177703857422],[-71.9124984741211,-72.0184555053711,-72.123046875,-72.1166000366211,-71.9188461303711,-71.8826217651367,-72.0002975463867,-72.0951156616211,-72.1492233276367,-72.1882858276367,-72.1890640258789,-72.1318359375,-72.0910110473633,-72.04541015625,-71.9440460205078,-71.8509750366211,-71.8363265991211,-71.8955078125,-72.0451126098633,-72.1316375732422,-72.2218704223633,-72.2469711303711,-72.2594757080078,-72.2554702758789,-72.2076187133789,-72.1219787597656,-72.056640625,-72.0684509277344,-72.1749954223633,-72.2487335205078,-72.4099655151367,-72.43896484375,-72.4538040161133,-72.5247039794922,-72.5542984008789,-72.5772476196289,-72.5476531982422,-72.5580062866211,-72.5783233642578,-72.5738296508789,-72.5208969116211,-72.5217742919922,-72.5476531982422,-72.5570297241211,-72.5560531616211,-72.5476531982422,-72.48974609375,-72.4732437133789,-72.4719772338867,-72.4066390991211,-72.3798828125,-72.3124008178711,-72.2870101928711,-72.2726593017578,-72.2781295776367,-72.1756820678711,-72.177734375,-72.1903305053711,-72.1849670410156,-72.13525390625,-72.0810546875,-72.0321350097656,-72.00927734375,-71.9854431152344,-71.86572265625,-71.8329086303711,-71.8369140625,-71.939453125,-72.0460891723633,-72.0443344116211,-72.0330047607422,-71.9449234008789,-71.97216796875,-71.9325180053711,-71.8542938232422,-71.7637634277344,-71.7815475463867,-71.8200225830078,-71.9124984741211],[-71.2137756347656,-71.2366180419922,-71.2302703857422,-71.2008743286133,-71.1429672241211,-71.1198196411133,-71.0811538696289,-71.0621109008789,-71.0517578125,-71.0759811401367,-71.0948257446289,-71.2137756347656],[-71.0529251098633,-71.0586929321289,-71.0411148071289,-71.0074234008789,-70.9673767089844,-70.9343795776367,-70.9140625,-70.9201126098633,-70.9625015258789,-70.99951171875,-71.0409240722656,-71.0529251098633],[-70.7677764892578,-70.8003921508789,-70.7960891723633,-70.8208999633789,-70.8553695678711,-70.9015655517578,-70.9979553222656,-71.04443359375,-71.14111328125,-71.16748046875,-71.1126937866211,-71.0567398071289,-71.0147476196289,-70.9822311401367,-70.9403305053711,-70.88330078125,-70.8514633178711,-70.7359390258789,-70.703125,-70.6832962036133,-70.6744155883789,-70.6941375732422,-70.7677764892578],[-70.6351547241211,-70.7234420776367,-70.7943344116211,-70.9241256713867,-70.9734344482422,-71.0122985839844,-71.1324234008789,-71.1328125,-71.1167984008789,-71.0904312133789,-70.9908218383789,-70.94140625,-70.8941421508789,-70.86376953125,-70.8359375,-70.7725524902344,-70.7517547607422,-70.70556640625,-70.6505889892578,-70.60888671875,-70.5902404785156,-70.5905227661133,-70.6309585571289,-70.7919921875,-70.7697296142578,-70.7266616821289,-70.5867233276367,-70.5758819580078,-70.6146469116211,-70.6553726196289,-70.5767517089844,-70.5529327392578,-70.5609359741211,-70.578125,-70.6351547241211],[-70.7105407714844,-70.7105407714844,-70.6896514892578,-70.6290054321289,-70.59912109375,-70.5325241088867,-70.517578125,-70.5087890625,-70.5442352294922,-70.6033172607422,-70.6466827392578,-70.7105407714844],[-70.55224609375,-70.6115188598633,-70.5855484008789,-70.5502014160156,-70.45263671875,-70.4457015991211,-70.4046859741211,-70.4214859008789,-70.4322280883789,-70.55224609375],[-70.5973587036133,-70.62353515625,-70.5933609008789,-70.5345687866211,-70.5003967285156,-70.3816452026367,-70.3926773071289,-70.4026412963867,-70.4488220214844,-70.5191421508789,-70.5539093017578,-70.5973587036133],[-70.5337829589844,-70.5354537963867,-70.5080108642578,-70.4883804321289,-70.3440399169922,-70.3073272705078,-70.2796859741211,-70.2884826660156,-70.3202133178711,-70.3761749267578,-70.4622039794922,-70.5070343017578,-70.5337829589844],[-70.26513671875,-70.29541015625,-70.2672882080078,-70.2978515625,-70.3174819946289,-70.3915023803711,-70.4388656616211,-70.4422912597656,-70.31884765625,-70.2883758544922,-70.26513671875],[-70.6326217651367,-70.6330108642578,-70.6219711303711,-70.5491180419922,-70.4974594116211,-70.4435501098633,-70.3363265991211,-70.2843780517578,-70.2642593383789,-70.3122100830078,-70.36767578125,-70.4055709838867,-70.4568405151367,-70.5009765625,-70.5245132446289,-70.5746078491211,-70.6090850830078,-70.6326217651367],[-70.4787139892578,-70.5026321411133,-70.4512710571289,-70.4169921875,-70.3252944946289,-70.2902374267578,-70.2667922973633,-70.2408218383789,-70.2513656616211,-70.2942352294922,-70.36865234375,-70.4325180053711,-70.4787139892578],[-70.3811492919922,-70.4193420410156,-70.4123992919922,-70.4479522705078,-70.4342803955078,-70.3729476928711,-70.2945327758789,-70.2614288330078,-70.1999053955078,-70.0724639892578,-70.060546875,-70.0782241821289,-70.1144561767578,-70.18603515625,-70.3299789428711,-70.3811492919922],[-70.2551727294922,-70.38134765625,-70.3781280517578,-70.30419921875,-70.2243194580078,-70.1689453125,-70.0940475463867,-70.0498046875,-70.0227508544922,-70.0407180786133,-70.1356506347656,-70.2551727294922],[-69.9757766723633,-69.9779281616211,-69.9498062133789,-69.9325180053711,-69.8932647705078,-69.8563461303711,-69.8831100463867,-69.9552764892578,-69.9757766723633],[-69.7278289794922,-69.75244140625,-69.8609390258789,-69.9168930053711,-69.9496078491211,-69.9716796875,-69.998046875,-70.1317367553711,-70.1792984008789,-70.1494140625,-70.0960922241211,-70.0850524902344,-70.0381774902344,-69.9839859008789,-69.9161148071289,-69.8816375732422,-69.8402328491211,-69.8168029785156,-69.7493209838867,-69.7351608276367,-69.7406234741211,-69.7278289794922],[-70.0076141357422,-70.0719757080078,-69.9820327758789,-69.955078125,-69.9390640258789,-69.884765625,-69.86279296875,-69.8280258178711,-69.7732467651367,-69.7284164428711,-69.7049865722656,-69.7232437133789,-69.7502975463867,-69.8444366455078,-70.0076141357422],[-69.6984405517578,-69.7401351928711,-69.70703125,-69.64501953125,-69.5290985107422,-69.4688491821289,-69.43310546875,-69.4130859375,-69.4518585205078,-69.4917984008789,-69.6984405517578],[-69.7218780517578,-69.7294921875,-69.6366195678711,-69.4949188232422,-69.2881851196289,-69.1804656982422,-69.1545944213867,-69.14599609375,-69.17578125,-69.2147445678711,-69.3003921508789,-69.3761749267578,-69.44189453125,-69.5146484375,-69.5333938598633,-69.5876007080078,-69.69140625,-69.7218780517578],[-69.1890640258789,-69.3109359741211,-69.2672805786133,-69.2765579223633,-69.3208999633789,-69.5831985473633,-69.6663131713867,-69.9090805053711,-70.09033203125,-70.4081039428711,-70.5814437866211,-70.6829071044922,-70.81787109375,-70.8560562133789,-70.9117202758789,-71.0970687866211,-71.3134765625,-71.7251968383789,-71.8221740722656,-71.9751968383789,-72.0853500366211,-72.1576156616211,-72.2097625732422,-72.4265594482422,-72.5341796875,-72.6261749267578,-72.6644592285156,-72.6229476928711,-72.6130828857422,-72.626953125,-72.6393585205078,-72.6697311401367,-72.6583023071289,-72.6172866821289,-72.5895462036133,-72.5958938598633,-72.58056640625,-72.5466766357422,-72.4840850830078,-72.4475555419922,-72.4078140258789,-72.30419921875,-72.2805633544922,-72.2750930786133,-72.2962875366211,-72.3313446044922,-72.3589859008789,-72.3664093017578,-72.3563461303711,-72.3225555419922,-72.2277374267578,-72.1910171508789,-72.1677703857422,-72.1633834838867,-72.1696319580078,-72.2291030883789,-72.2640609741211,-72.2844696044922,-72.2498016357422,-72.15283203125,-72.1208038330078,-72.0470657348633,-72.0345687866211,-71.9874038696289,-71.94580078125,-71.9065475463867,-71.83642578125,-71.837890625,-71.8505859375,-71.8218765258789,-71.7396469116211,-71.6412124633789,-71.6322250366211,-71.6623001098633,-71.9216766357422,-71.9236373901367,-71.9045944213867,-71.8531265258789,-71.8349609375,-71.8489227294922,-71.8702163696289,-71.9293899536133,-71.98095703125,-72.0223617553711,-72.15234375,-72.1698226928711,-72.1587905883789,-72.1422882080078,-72.0726547241211,-72.0555648803711,-72.0699234008789,-72.0635757446289,-72.0333023071289,-71.9884796142578,-71.9639663696289,-71.9139633178711,-71.87841796875,-71.8279266357422,-71.7802734375,-71.7523498535156,-71.7255859375,-71.6796875,-71.6452178955078,-71.6149444580078,-71.5553741455078,-71.5433578491211,-71.6174850463867,-71.6415023803711,-71.6432647705078,-71.63818359375,-71.6179656982422,-71.5793914794922,-71.5072250366211,-71.4569320678711,-71.4149398803711,-71.39990234375,-71.3883819580078,-71.3830108642578,-71.4104461669922,-71.4381866455078,-71.5169906616211,-71.5730438232422,-71.5787124633789,-71.5588912963867,-71.5279235839844,-71.4480438232422,-71.3965835571289,-71.3507766723633,-71.3248977661133,-71.3211898803711,-71.36865234375,-71.3835906982422,-71.3883819580078,-71.3350524902344,-71.2750015258789,-71.22314453125,-71.1865234375,-71.126953125,-71.0729446411133,-71.0748062133789,-71.1451187133789,-71.1115264892578,-71.0108413696289,-70.9847640991211,-70.9925765991211,-70.9463806152344,-70.951171875,-70.9647445678711,-71.1337890625,-71.1256866455078,-71.0925750732422,-71.03369140625,-70.9734344482422,-70.9136734008789,-70.8759765625,-70.88037109375,-70.8970718383789,-70.8826141357422,-70.8505859375,-70.8361282348633,-70.81787109375,-70.7858428955078,-70.7621154785156,-70.7129898071289,-70.6595687866211,-70.537109375,-70.4040985107422,-70.3612289428711,-70.3506851196289,-70.3601531982422,-70.4147491455078,-70.4123077392578,-70.3980484008789,-70.30517578125,-70.2341766357422,-70.1804656982422,-70.15966796875,-70.1394500732422,-70.1923828125,-70.2013702392578,-70.1964874267578,-70.0677719116211,-70.0537109375,-70.0051803588867,-69.9693374633789,-69.9089889526367,-69.84716796875,-69.8070297241211,-69.6494140625,-69.5240249633789,-69.4227523803711,-69.3667984008789,-69.3287048339844,-69.2671890258789,-69.2253875732422,-69.1765670776367,-69.1145477294922,-69.06005859375,-69.0009765625,-68.9708023071289,-68.9410095214844,-68.87353515625,-68.7889633178711,-68.8322296142578,-68.9229507446289,-68.9593734741211,-69.1395492553711,-69.1890640258789],[-68.7977600097656,-68.79833984375,-68.7981491088867,-68.7986297607422,-68.8001937866211,-68.8018569946289,-68.80322265625,-68.80322265625,-68.8013687133789,-68.7969741821289,-68.7882843017578,-68.7796936035156,-68.7708969116211,-68.7595672607422,-68.7544937133789,-68.7472686767578,-68.7378921508789,-68.7297821044922,-68.7213897705078,-68.7164077758789,-68.7138671875,-68.7121047973633,-68.7123031616211,-68.7129898071289,-68.7139663696289,-68.7150344848633,-68.7183609008789,-68.7284164428711,-68.7384796142578,-68.7429733276367,-68.7488250732422,-68.755859375,-68.7642593383789,-68.7712860107422,-68.783203125,-68.794921875,-68.7972640991211,-68.7977600097656],[-68.767578125,-68.7950210571289,-68.7784194946289,-68.7588882446289,-68.7097625732422,-68.6806640625,-68.6876983642578,-68.7220687866211,-68.767578125],[-67.7662124633789,-67.7852554321289,-67.7634735107422,-67.6876907348633,-67.6794891357422,-67.6612319946289,-67.6044921875,-67.5906219482422,-67.5987319946289,-67.62451171875,-67.6501922607422,-67.7119140625,-67.7372055053711,-67.7662124633789],[-67.5404281616211,-67.56884765625,-67.56005859375,-67.5000991821289,-67.4078140258789,-67.2888641357422,-67.2593688964844,-67.3260726928711,-67.3636703491211,-67.4185562133789,-67.4474639892578,-67.50390625,-67.5404281616211],[-66.89453125,-66.9019546508789,-66.8972625732422,-66.8755874633789,-66.8036117553711,-66.7562484741211,-66.7369079589844,-66.7535171508789,-66.8152313232422,-66.8409194946289,-66.89453125],[-66.9533157348633,-66.9796905517578,-66.96533203125,-66.9508819580078,-66.8542938232422,-66.7759780883789,-66.7233428955078,-66.728515625,-66.7746047973633,-66.8941421508789,-66.9533157348633],[-66.7841796875,-66.8073196411133,-66.7978515625,-66.7738265991211,-66.7500991821289,-66.7242202758789,-66.7038040161133,-66.7009735107422,-66.7196273803711,-66.7311553955078,-66.767578125,-66.7783203125,-66.7841796875],[-66.8211898803711,-66.8819351196289,-66.8679656982422,-66.8191375732422,-66.7005844116211,-66.6884765625,-66.7087936401367,-66.767578125,-66.7984390258789,-66.8211898803711],[-66.7674865722656,-66.7919921875,-66.78759765625,-66.73291015625,-66.6966781616211,-66.6748046875,-66.6869125366211,-66.7057571411133,-66.7181625366211,-66.7674865722656],[-67.4744110107422,-67.5386734008789,-67.5581970214844,-67.5396499633789,-67.5324249267578,-67.5553741455078,-67.65625,-67.7071304321289,-67.7328109741211,-67.7225646972656,-67.7228469848633,-67.7457046508789,-67.75341796875,-67.7442398071289,-67.6799850463867,-67.6027297973633,-67.5779342651367,-67.5152282714844,-67.45263671875,-67.4031295776367,-67.2335968017578,-67.1572265625,-67.0704116821289,-66.9925765991211,-66.8533172607422,-66.8020477294922,-66.6568374633789,-66.6243133544922,-66.6330108642578,-66.708984375,-66.7461929321289,-66.8445358276367,-66.9821319580078,-67.0322265625,-67.0448226928711,-67.0624008178711,-67.0819320678711,-67.1229476928711,-67.1473617553711,-67.2191390991211,-67.25537109375,-67.2999954223633,-67.3441390991211,-67.3719787597656,-67.3824234008789,-67.41796875,-67.4502868652344,-67.4744110107422],[-66.6119079589844,-66.6434631347656,-66.6371078491211,-66.6042938232422,-66.5837936401367,-66.5185546875,-66.5215835571289,-66.5560531616211,-66.6119079589844],[-66.4698257446289,-66.4816360473633,-66.4533233642578,-66.39990234375,-66.3826141357422,-66.3692321777344,-66.4205093383789,-66.4698257446289],[-66.4773406982422,-66.5251007080078,-66.5201110839844,-66.3997039794922,-66.3474578857422,-66.3037109375,-66.2646484375,-66.2512741088867,-66.4326171875,-66.4773406982422],[-66.2165985107422,-66.2294921875,-66.2027359008789,-66.1880874633789,-66.1310501098633,-66.1123962402344,-66.1799774169922,-66.2165985107422],[-66.20068359375,-66.3126983642578,-66.3054656982422,-66.2938537597656,-66.2748107910156,-66.2335968017578,-66.11083984375,-66.0667953491211,-66.0824203491211,-66.1338882446289,-66.1786117553711,-66.20068359375],[-66.0358428955078,-66.0608367919922,-66.0584030151367,-66.0967788696289,-66.1394500732422,-66.1639709472656,-66.2007827758789,-66.2249984741211,-66.1858367919922,-66.0801773071289,-66.0458984375,-66.0358428955078],[-65.8082962036133,-65.8216781616211,-65.8072280883789,-65.7600555419922,-65.7399444580078,-65.7067413330078,-65.7021484375,-65.7306671142578,-65.7604446411133,-65.7748031616211,-65.8082962036133],[-65.8424758911133,-65.880859375,-65.8665084838867,-65.8263626098633,-65.7737350463867,-65.7447204589844,-65.6661148071289,-65.6328125,-65.5709991455078,-65.5272445678711,-65.5477523803711,-65.6261749267578,-65.6529312133789,-65.67431640625,-65.6866226196289,-65.7384719848633,-65.8137664794922,-65.8424758911133],[-65.6775436401367,-65.7012710571289,-65.6753921508789,-65.6729507446289,-65.6512680053711,-65.6033172607422,-65.5206985473633,-65.4656219482422,-65.4089889526367,-65.3963851928711,-65.3781280517578,-65.4025344848633,-65.4722671508789,-65.5276412963867,-65.5645523071289,-65.6775436401367],[-65.4453125,-65.4685516357422,-65.4546890258789,-65.43505859375,-65.33837890625,-65.31201171875,-65.2853546142578,-65.2359390258789,-65.1678695678711,-65.1363296508789,-65.1296844482422,-65.1906280517578,-65.2371139526367,-65.3077163696289,-65.33056640625,-65.3773422241211,-65.4264602661133,-65.4453125],[-64.8611297607422,-64.9065399169922,-64.9059524536133,-64.7962875366211,-64.7920913696289,-64.7297821044922,-64.73876953125,-64.7908172607422,-64.8611297607422],[-64.5667953491211,-64.5707015991211,-64.5402297973633,-64.4884719848633,-64.4598617553711,-64.4387741088867,-64.4353485107422,-64.3523406982422,-64.3330078125,-64.3817367553711,-64.42724609375,-64.4679718017578,-64.5667953491211],[-64.4695358276367,-64.5733413696289,-64.5723648071289,-64.5349578857422,-64.5193328857422,-64.5718765258789,-64.6109390258789,-64.6802749633789,-64.7173843383789,-64.73388671875,-64.7273483276367,-64.7341766357422,-64.8030242919922,-64.8342819213867,-64.8084030151367,-64.79150390625,-64.7685546875,-64.7327117919922,-64.7095642089844,-64.6975631713867,-64.6353530883789,-64.5819320678711,-64.509765625,-64.4871063232422,-64.4572296142578,-64.42138671875,-64.3839874267578,-64.3427734375,-64.3141632080078,-64.27294921875,-64.2605438232422,-64.2662124633789,-64.3236312866211,-64.3806686401367,-64.4695358276367],[-64.4244155883789,-64.4646530151367,-64.4716796875,-64.4541015625,-64.5142593383789,-64.4959945678711,-64.47900390625,-64.4716796875,-64.4445266723633,-64.3916015625,-64.25341796875,-64.2106399536133,-64.1396484375,-64.1163101196289,-64.0755844116211,-64.0457000732422,-64.0124053955078,-64.0134735107422,-64.0399398803711,-64.0900421142578,-64.1380844116211,-64.2345657348633,-64.2959976196289,-64.3688507080078,-64.4013671875,-64.4244155883789],[-64.0771484375,-64.0803680419922,-64.02734375,-63.9902305603027,-63.9665985107422,-64.0269546508789,-64.0544891357422,-64.0771484375],[-64.0539016723633,-64.0675811767578,-64.0615234375,-64.0478515625,-64.01513671875,-63.9670906066895,-63.9616203308105,-64.0106430053711,-64.0970687866211,-64.1662139892578,-64.2213897705078,-64.23779296875,-64.2958984375,-64.3182601928711,-64.3669891357422,-64.37890625,-64.3503875732422,-64.3572235107422,-64.3109359741211,-64.2932586669922,-64.3020477294922,-64.4009780883789,-64.4100570678711,-64.39404296875,-64.3312530517578,-64.3204116821289,-64.3215866088867,-64.3685531616211,-64.3697280883789,-64.3145523071289,-64.2419967651367,-64.2061538696289,-64.1607437133789,-64.0973663330078,-64.1068344116211,-64.1306610107422,-64.134765625,-64.1134796142578,-64.0677719116211,-63.9943389892578,-63.9162139892578,-63.8776359558105,-63.8474617004395,-63.8346672058105,-63.8060569763184,-63.8038101196289,-63.8682594299316,-63.9068374633789,-63.94921875,-64.0539016723633],[-63.8072242736816,-63.8248062133789,-63.8157196044922,-63.8412094116211,-63.8755874633789,-63.8683624267578,-63.8785171508789,-63.8536148071289,-63.8126945495605,-63.7914047241211,-63.8072242736816],[-63.8666000366211,-63.9021492004395,-63.8910179138184,-63.8490257263184,-63.8366203308105,-63.7166976928711,-63.6688461303711,-63.6958999633789,-63.7589836120605,-63.8079071044922,-63.85009765625,-63.8666000366211],[-63.5356407165527,-63.5799827575684,-63.5132789611816,-63.4688491821289,-63.4369125366211,-63.4073219299316,-63.4220695495605,-63.4920921325684,-63.5356407165527],[-84.3515625,-84.3375015258789,-84.3523406982422,-84.3952178955078,-84.3993148803711,-84.4183578491211,-84.4752960205078,-84.5103530883789,-84.4797821044922,-84.4654312133789,-84.4626922607422,-84.5423812866211,-84.68359375,-84.7286148071289,-84.8336868286133,-84.8192367553711,-84.90087890625,-84.9011688232422,-84.9866256713867,-85.1856384277344,-85.1921844482422,-85.18603515625,-85.0793991088867,-84.9822235107422,-84.9123992919922,-84.8913116455078,-84.8892593383789,-84.8396530151367,-84.8111343383789,-84.7780303955078,-84.5287170410156,-84.5130844116211,-84.4927749633789,-84.4704132080078,-84.4454116821289,-84.4313507080078,-84.4098587036133,-84.3812484741211,-84.3695297241211,-84.3746109008789,-84.3526382446289,-84.32421875,-84.3054656982422,-84.29052734375,-84.2740249633789,-84.1910171508789,-84.1545867919922,-84.0966796875,-84.0715789794922,-84.0535202026367,-84.015625,-83.9460906982422,-83.8058624267578,-83.7566452026367,-83.7902297973633,-83.7919921875,-83.8108367919922,-83.7908172607422,-83.7354507446289,-83.6368103027344,-83.611328125,-83.529296875,-83.4498062133789,-83.2564468383789,-83.2037124633789,-82.9000930786133,-82.8477554321289,-82.79345703125,-82.9158172607422,-82.9167938232422,-82.8841781616211,-82.89306640625,-82.8624038696289,-82.8474655151367,-82.9429702758789,-83.0960006713867,-83.1502914428711,-83.1878890991211,-83.2015609741211,-83.2288055419922,-83.2132797241211,-83.2265625,-83.2173843383789,-83.2007827758789,-83.2155303955078,-83.2900390625,-83.4124984741211,-83.4676742553711,-83.4247055053711,-83.373046875,-83.3291015625,-83.3470687866211,-83.41064453125,-83.5189437866211,-83.4895477294922,-83.4947280883789,-83.5431671142578,-83.3812484741211,-83.3463897705078,-83.234375,-83.1984405517578,-83.1874008178711,-83.1294937133789,-83.1066436767578,-83.0751953125,-83.02685546875,-82.9807586669922,-82.8583984375,-82.6693344116211,-82.5862274169922,-82.4496078491211,-82.1765594482422,-81.9407196044922,-81.8675765991211,-81.810546875,-81.7654342651367,-81.7001953125,-81.6232376098633,-81.5662078857422,-81.5103530883789,-81.376953125,-81.3191375732422,-81.23095703125,-81.1623077392578,-81.1333999633789,-81.0290069580078,-80.9007797241211,-80.8357391357422,-80.7600631713867,-80.7415008544922,-80.7186508178711,-80.67041015625,-80.6041946411133,-80.5930633544922,-80.5862350463867,-80.5104522705078,-80.48046875,-80.4094772338867,-80.3537139892578,-80.2110366821289,-80.1500015258789,-80.11767578125,-80.10595703125,-80.1081085205078,-80.0905227661133,-80.0709991455078,-80.0027389526367,-79.9712905883789,-79.9294891357422,-79.9065399169922,-79.90771484375,-79.8567428588867,-79.77587890625,-79.7314453125,-79.6569290161133,-79.5456085205078,-79.4596710205078,-79.3933639526367,-79.3176727294922,-79.2805633544922,-79.2416000366211,-79.1927719116211,-79.1159210205078,-79.1027374267578,-79.1348648071289,-79.1172866821289,-79.0465850830078,-78.96484375,-78.74462890625,-78.6353530883789,-78.5586929321289,-78.5103530883789,-78.3981475830078,-78.35888671875,-78.2590866088867,-78.2169876098633,-78.2208023071289,-78.1919937133789,-78.1861343383789,-78.1998062133789,-78.0739288330078,-77.9507827758789,-77.8877944946289,-77.7783203125,-77.6148376464844,-77.4976577758789,-77.3542938232422,-77.1571273803711,-77.0912094116211,-77.0792007446289,-77.2312469482422,-77.2420883178711,-77.2129898071289,-77.134765625,-77.1056671142578,-77.0980453491211,-77.13330078125,-77.1195297241211,-77.126953125,-77.2269515991211,-77.2742156982422,-77.3101501464844,-77.3630905151367,-77.416015625,-77.4424819946289,-77.41259765625,-77.4258804321289,-77.5735321044922,-77.7314453125,-77.77099609375,-77.7974624633789,-77.7742156982422,-77.71484375,-77.6426773071289,-77.5511703491211,-77.4623031616211,-77.41259765625,-77.361328125,-77.34326171875,-77.28369140625,-77.2023468017578,-77.125,-77.1050796508789,-77.2113265991211,-77.3097610473633,-77.3252944946289,-77.3207092285156,-77.2858352661133,-77.2598648071289,-77.4724578857422,-77.4867172241211,-77.4880828857422,-77.4552764892578,-77.3983383178711,-77.33837890625,-77.3299789428711,-77.2732391357422,-77.2212905883789,-77.19921875,-77.1617202758789,-77.1033248901367,-77.0941390991211,-77.0687484741211,-77.0289993286133,-77.0121078491211,-76.9537124633789,-76.8844757080078,-76.7966766357422,-76.7490234375,-76.6576156616211,-76.5070343017578,-76.4383850097656,-76.4932632446289,-76.4580078125,-76.4189453125,-76.3653335571289,-76.3112335205078,-76.2717819213867,-76.23828125,-76.16796875,-76.1179733276367,-76.1044921875,-76.130859375,-76.3180694580078,-76.3377914428711,-76.42431640625,-76.4288024902344,-76.4091796875,-76.32568359375,-76.2666015625,-76.0997085571289,-76.0203094482422,-75.8887710571289,-75.87890625,-75.8321304321289,-75.7314453125,-75.5635681152344,-75.54345703125,-75.4909210205078,-75.52978515625,-75.6904296875,-75.7460021972656,-75.7507781982422,-75.7458953857422,-75.5458984375,-75.52001953125,-75.4976501464844,-75.447265625,-75.4058609008789,-75.2127914428711,-75.16015625,-75.0755920410156,-75.1526412963867,-75.1617202758789,-75.1394500732422,-75.0358352661133,-74.8360366821289,-74.7653350830078,-74.6904296875,-74.6941452026367,-74.7761764526367,-74.8296890258789,-74.8545913696289,-74.8518600463867,-74.80615234375,-74.7893524169922,-74.7657241821289,-74.8109359741211,-74.8259811401367,-74.890625,-74.8914031982422,-74.8289108276367,-74.8202133178711,-74.71923828125,-74.6978530883789,-74.7425765991211,-74.7146453857422,-74.7357482910156,-74.7730484008789,-74.75,-74.6545867919922,-74.62158203125,-74.5178680419922,-74.4222640991211,-74.3927764892578,-74.3815460205078,-74.4029312133789,-74.4732437133789,-74.4470672607422,-74.4873962402344,-74.455078125,-74.2750015258789,-73.9885787963867,-73.9029235839844,-73.9250030517578,-74.0888671875,-74.1583938598633,-74.2079086303711,-74.2277374267578,-74.36669921875,-74.4063491821289,-74.3942337036133,-74.4542007446289,-74.5587921142578,-74.6181640625,-74.6444320678711,-74.8429641723633,-74.9090805053711,-74.9818344116211,-74.9818344116211,-74.9521484375,-74.9436492919922,-74.8917007446289,-74.8322296142578,-74.8011703491211,-74.7921905517578,-74.7508773803711,-74.6534194946289,-74.5716781616211,-74.5040969848633,-74.3866195678711,-74.2697296142578,-74.1814498901367,-74.2007827758789,-74.1880874633789,-74.23046875,-74.2689437866211,-74.2888641357422,-74.36669921875,-74.5363311767578,-74.7106475830078,-74.8363265991211,-74.9512710571289,-75.1333999633789,-75.2199249267578,-75.1908187866211,-75.1991195678711,-75.18505859375,-75.2066421508789,-75.2525405883789,-75.3215866088867,-75.33447265625,-75.3093719482422,-75.3439483642578,-75.1976547241211,-75.1151351928711,-75.15625,-75.1207046508789,-75.1525344848633,-75.1012725830078,-75.09521484375,-75.1169967651367,-75.1273422241211,-75.2217712402344,-75.3658218383789,-75.421875,-75.3981475830078,-75.3534164428711,-75.3704147338867,-75.3089828491211,-75.3274459838867,-75.3170928955078,-75.2771453857422,-75.1897506713867,-75.1408233642578,-75.0785217285156,-74.9488296508789,-74.9216766357422,-74.9378890991211,-74.9143524169922,-74.8723602294922,-74.8229446411133,-74.662109375,-74.5150451660156,-74.4841842651367,-74.4888687133789,-74.5411148071289,-74.5049819946289,-74.4857406616211,-74.35009765625,-74.0963821411133,-74.0237350463867,-73.9577102661133,-73.92578125,-73.8837890625,-73.7835998535156,-73.6456985473633,-73.6164093017578,-73.6305694580078,-73.6554718017578,-73.6667938232422,-73.6952133178711,-73.73486328125,-73.7572250366211,-73.7578125,-73.7201156616211,-73.6941452026367,-73.6451110839844,-73.6341781616211,-73.6408157348633,-73.6111373901367,-73.5709915161133,-73.4951171875,-73.4025421142578,-73.3531265258789,-73.31787109375,-73.32958984375,-73.3112258911133,-73.3208999633789,-73.28515625,-73.1845703125,-72.9453125,-72.81884765625,-72.7720718383789,-72.72119140625,-72.7162094116211,-72.7356491088867,-72.7601547241211,-72.8349609375,-72.9114227294922,-72.951171875,-72.9874038696289,-72.9981460571289,-73.0208969116211,-73.0298843383789,-72.9954147338867,-72.9811553955078,-73.0155258178711,-73.0413055419922,-72.9999008178711,-73.0222625732422,-73.033203125,-73.1017608642578,-73.1444320678711,-73.1262664794922,-73.2064437866211,-73.2685546875,-73.3011703491211,-73.3092727661133,-73.2938461303711,-73.2414093017578,-73.2201232910156,-73.2389678955078,-73.24951171875,-73.31298828125,-73.2867202758789,-73.2150344848633,-73.1646499633789,-73.1784133911133,-73.3070373535156,-73.3191452026367,-73.2432632446289,-73.11865234375,-72.9779281616211,-72.9445266723633,-72.9601593017578,-72.8626022338867,-72.8708953857422,-72.8895492553711,-72.8257751464844,-72.6931610107422,-72.6830062866211,-72.7023468017578,-72.7875061035156,-72.8585891723633,-72.9343719482422,-73.1207046508789,-73.22900390625,-73.2195358276367,-73.2409210205078,-73.1945343017578,-73.1919937133789,-73.3539123535156,-73.3636703491211,-73.3537139892578,-73.20849609375,-73.1920852661133,-73.2589797973633,-73.4132843017578,-73.5020523071289,-73.5567398071289,-73.57275390625,-73.6451110839844,-73.7059555053711,-73.7072296142578,-73.7323226928711,-73.8568344116211,-73.7957000732422,-73.73828125,-73.6324234008789,-73.4737319946289,-73.31494140625,-73.2488250732422,-73.2355499267578,-73.4141616821289,-73.30810546875,-73.2249984741211,-73.0833969116211,-72.9776306152344,-72.9445266723633,-73.0000991821289,-73.0281219482422,-73.0895538330078,-73.3124008178711,-73.5067443847656,-73.5557632446289,-73.5470657348633,-73.5150375366211,-73.4879837036133,-73.4957962036133,-73.46044921875,-73.5663070678711,-73.7184600830078,-73.8176803588867,-73.8441390991211,-73.8205032348633,-73.7894515991211,-73.8053741455078,-73.7364273071289,-73.7112274169922,-73.6387710571289,-73.6451110839844,-73.6580123901367,-73.7152328491211,-73.6838912963867,-73.6955108642578,-73.6998062133789,-73.4479446411133,-73.4523468017578,-73.4383773803711,-73.3792037963867,-73.35302734375,-73.3544921875,-73.2627944946289,-73.2740249633789,-73.2264633178711,-73.1696319580078,-73.10546875,-72.935546875,-72.8345718383789,-72.6111297607422,-72.3875961303711,-72.0904235839844,-71.8972702026367,-71.8122024536133,-71.7189483642578,-71.5267562866211,-71.2845764160156,-71.0578079223633,-70.8446273803711,-70.6861343383789,-70.4216842651367,-70.2499008178711,-70.0197219848633,-69.8091812133789,-69.6438446044922,-69.5263671875,-69.4322280883789,-69.4126892089844,-69.3839874267578,-69.3475570678711,-69.4123001098633,-69.3175735473633,-69.248046875,-69.1610336303711,-69.0287094116211,-68.9744110107422,-68.8612289428711,-68.7706985473633,-68.7706985473633,-68.6714859008789,-68.5748062133789,-68.453125,-68.2976531982422,-68.2404251098633,-68.1467742919922,-68.0246047973633,-67.9300765991211,-67.8314514160156,-67.6925735473633,-67.5933609008789,-67.5602569580078,-67.5271530151367,-67.4916000366211,-67.4850540161133,-67.5469741821289,-67.53466796875,-67.5029296875,-67.4351577758789,-67.2692337036133,-67.11279296875,-67.0907211303711,-67.07080078125,-66.9517517089844,-66.9451141357422,-66.9847640991211,-67.1435546875,-67.1799774169922,-67.2559585571289,-67.2824249267578,-67.2325134277344,-67.2085952758789,-67.2625961303711,-67.2890625,-67.2427749633789,-67.1142578125,-67.0625,-66.9792938232422,-66.9401397705078,-66.8751983642578,-66.74072265625,-66.6898422241211,-66.60888671875,-66.5919952392578,-66.5924835205078,-66.6456069946289,-66.6498031616211,-66.6249008178711,-66.5732421875,-66.4027404785156,-66.3425750732422,-66.2879867553711,-66.2544937133789,-66.13525390625,-66.1292953491211,-66.1398468017578,-66.1167984008789,-66.0684509277344,-65.9942398071289,-65.9579086303711,-65.9462890625,-65.9927749633789,-66.01904296875,-65.9595718383789,-65.9000015258789,-65.8668975830078,-65.8140640258789,-65.7478485107422,-65.7809600830078,-65.7683563232422,-65.70849609375,-65.6798858642578,-65.640625,-65.6329116821289,-65.6173782348633,-65.5705032348633,-65.5537109375,-65.5559539794922,-65.5315399169922,-65.4803695678711,-65.4673843383789,-65.4171905517578,-65.2782211303711,-65.1790084838867,-65.0930709838867,-65.0334930419922,-65.0849609375,-65.0731430053711,-65.1260757446289,-65.1393585205078,-65.0797805786133,-65.0279312133789,-64.9424819946289,-64.8416976928711,-64.8575134277344,-64.8333969116211,-64.7556610107422,-64.6564407348633,-64.6432571411133,-64.7291946411133,-64.7468719482422,-64.7267608642578,-64.6253890991211,-64.60986328125,-64.6046905517578,-64.5456085205078,-64.4755859375,-64.4271469116211,-64.3624954223633,-64.3147506713867,-64.1497039794922,-64.10791015625,-64.0734405517578,-63.9239234924316,-63.9095687866211,-63.8207015991211,-63.7138710021973,-63.6703147888184,-63.5518569946289,-63.5343704223633,-63.4512672424316,-63.3187484741211,-63.2262687683105,-63.2347640991211,-63.2625045776367,-63.3728523254395,-63.5055694580078,-63.5716781616211,-63.6312522888184,-63.6246109008789,-63.6377944946289,-63.5716781616211,-63.5235328674316,-63.4791030883789,-63.4906234741211,-63.5135726928711,-63.5465850830078,-63.6166000366211,-63.6568374633789,-63.7633819580078,-63.9154281616211,-64.0774383544922,-64.1868209838867,-64.1949157714844,-64.2344741821289,-64.2659225463867,-64.279296875,-64.2926712036133,-64.3389587402344,-64.3521499633789,-64.3888702392578,-64.44482421875,-64.5242156982422,-64.5374069213867,-64.4513626098633,-64.4435501098633,-64.4035110473633,-64.3456039428711,-64.3587875366211,-64.4401397705078,-64.53076171875,-64.5836944580078,-64.5588836669922,-64.4513626098633,-64.43359375,-64.4313430786133,-64.546875,-64.5505905151367,-64.609375,-64.6765670776367,-64.7291946411133,-64.9068374633789,-64.9812545776367,-65.0238342285156,-65.0176773071289,-64.9997100830078,-64.98779296875,-64.9872055053711,-65.0334930419922,-65.1856460571289,-65.2385711669922,-65.2353515625,-65.1922836303711,-65.2325210571289,-65.2732391357422,-65.3317337036133,-65.4568405151367,-65.5134735107422,-65.52294921875,-65.5692367553711,-65.5891571044922,-65.6988296508789,-65.7750015258789,-65.8404312133789,-65.9164047241211,-66.0313491821289,-66.1128921508789,-66.1195297241211,-66.0947265625,-66.0714874267578,-66.0587921142578,-65.97998046875,-65.9745101928711,-65.9886703491211,-65.9919891357422,-65.9402389526367,-65.9208984375,-65.93408203125,-65.93310546875,-65.9793930053711,-66.0323257446289,-66.1050796508789,-66.0653305053711,-66.0719680786133,-66.1105499267578,-66.1910171508789,-66.2637710571289,-66.3365249633789,-66.2902374267578,-66.2117156982422,-66.16455078125,-66.1447296142578,-66.2256851196289,-66.24951171875,-66.3431625366211,-66.42919921875,-66.4027404785156,-66.2960891723633,-66.208984375,-66.1970748901367,-66.2193298339844,-66.2174835205078,-66.2373046875,-66.3101501464844,-66.3636779785156,-66.4357376098633,-66.4896545410156,-66.5111312866211,-66.5560531616211,-66.6209945678711,-66.70703125,-66.7061538696289,-66.6800765991211,-66.4528274536133,-66.3525390625,-66.2637710571289,-66.2437515258789,-66.24169921875,-66.2777404785156,-66.31982421875,-66.3829116821289,-66.4089889526367,-66.5059585571289,-66.5887680053711,-66.6066360473633,-66.6540985107422,-66.7609405517578,-66.80322265625,-66.8729476928711,-66.9120101928711,-66.92724609375,-66.8533172607422,-66.8517608642578,-66.7996139526367,-66.8062515258789,-66.8941421508789,-66.9719696044922,-67.0245132446289,-67.1047821044922,-67.1237335205078,-67.1560516357422,-67.1832046508789,-67.2140579223633,-67.2427749633789,-67.2691421508789,-67.3073272705078,-67.3353500366211,-67.3419876098633,-67.3109436035156,-67.326171875,-67.3772506713867,-67.4446258544922,-67.5282211303711,-67.587890625,-67.6101608276367,-67.6595687866211,-67.7883758544922,-67.8163070678711,-67.8756790161133,-67.9299774169922,-68.0094680786133,-68.04833984375,-68.1305694580078,-68.1466827392578,-68.150390625,-68.1400375366211,-68.0675735473633,-68.0563430786133,-68.0831069946289,-68.12744140625,-68.1683578491211,-68.2874984741211,-68.3367156982422,-68.3641662597656,-68.3702163696289,-68.4078140258789,-68.4493179321289,-68.4892578125,-68.5832061767578,-68.6179656982422,-68.67333984375,-68.74609375,-68.7711944580078,-68.6869125366211,-68.58251953125,-68.4976577758789,-68.4697265625,-68.4188461303711,-68.4207000732422,-68.4425811767578,-68.486328125,-68.4706039428711,-68.4994125366211,-68.5921859741211,-68.6318359375,-68.70458984375,-68.76416015625,-68.8104476928711,-68.951171875,-69.0418930053711,-69.1410140991211,-69.2530288696289,-69.3289031982422,-69.3718719482422,-69.4772415161133,-69.5843734741211,-69.8272399902344,-70.0279312133789,-70.1201171875,-70.1995086669922,-70.2789001464844,-70.2332000732422,-70.2789001464844,-70.3648376464844,-70.4244155883789,-70.4971694946289,-70.4905319213867,-70.5699234008789,-70.61669921875,-70.67578125,-70.7087860107422,-70.7286071777344,-70.8013687133789,-70.9005889892578,-70.8567352294922,-70.8511734008789,-70.8675765991211,-71.0022506713867,-71.1668930053711,-71.24462890625,-71.3193359375,-71.3418960571289,-71.4005813598633,-71.4523468017578,-71.4791030883789,-71.6160125732422,-71.630859375,-71.6578063964844,-71.6725540161133,-71.67529296875,-71.5640640258789,-71.5885696411133,-71.6613235473633,-71.7472610473633,-71.8200225830078,-71.8628921508789,-71.9036178588867,-72.017578125,-72.0709915161133,-72.052734375,-72.0726547241211,-72.1126937866211,-72.0915069580078,-72.0501937866211,-72.0515594482422,-72.0726547241211,-72.1441421508789,-72.2698211669922,-72.3625030517578,-72.4259719848633,-72.4705047607422,-72.46826171875,-72.6007766723633,-72.69970703125,-72.6468734741211,-72.67333984375,-72.8321304321289,-73.00732421875,-73.0172805786133,-72.9378890991211,-72.9378890991211,-73.0305709838867,-73.1892623901367,-73.2752914428711,-73.240234375,-73.21142578125,-73.27099609375,-73.3204040527344,-73.3282241821289,-73.2502975463867,-73.19140625,-73.1607437133789,-73.1476593017578,-73.2157211303711,-73.2548828125,-73.37548828125,-73.5001983642578,-73.4670944213867,-73.5386734008789,-73.6120071411133,-73.7118148803711,-73.87060546875,-73.9294891357422,-73.9566421508789,-73.89599609375,-73.89599609375,-73.923828125,-73.9961929321289,-74.0320281982422,-74.0359344482422,-74.0557556152344,-74.1219711303711,-74.2079086303711,-74.1947250366211,-74.2281265258789,-74.2896423339844,-74.32861328125,-74.3069305419922,-74.2410125732422,-74.30712890625,-74.3729476928711,-74.4783248901367,-74.5118179321289,-74.5135726928711,-74.4757843017578,-74.4528350830078,-74.4413070678711,-74.5055694580078,-74.5499954223633,-74.7130813598633,-74.7767639160156,-74.86279296875,-74.9263687133789,-74.9521484375,-74.8958053588867,-74.7371139526367,-74.6908264160156,-74.6775360107422,-74.6841812133789,-74.7645492553711,-74.8495101928711,-74.9090805053711,-74.8783187866211,-74.9056701660156,-74.9523468017578,-75.0044937133789,-75.0302734375,-75.0347671508789,-75.11474609375,-75.15380859375,-75.1714859008789,-75.2061538696289,-75.2928695678711,-75.3293914794922,-75.3279266357422,-75.3363342285156,-75.3522415161133,-75.3985366821289,-75.4514694213867,-75.57958984375,-75.7381820678711,-75.7874984741211,-75.8151321411133,-75.95166015625,-76.0133819580078,-76.1097640991211,-76.3507843017578,-76.52197265625,-76.6544876098633,-76.6741180419922,-76.7180633544922,-76.7393569946289,-76.7527313232422,-76.6890640258789,-76.6754913330078,-76.69677734375,-76.5814437866211,-76.5867233276367,-76.6082000732422,-76.6082000732422,-76.5920867919922,-76.5853576660156,-76.6297836303711,-76.70166015625,-76.8338851928711,-76.9934539794922,-77.27490234375,-77.33447265625,-77.3984375,-77.4742202758789,-77.47802734375,-77.5355529785156,-77.5902328491211,-77.69384765625,-77.8942337036133,-77.9708023071289,-78.109375,-78.1778335571289,-78.1578140258789,-78.0441360473633,-77.9403305053711,-77.8425827026367,-77.7965774536133,-77.751953125,-77.7976608276367,-77.841796875,-77.8460922241211,-77.8856430053711,-78.154296875,-78.3509750366211,-78.4014663696289,-78.4346694946289,-78.5603485107422,-78.6595764160156,-78.7455139160156,-78.7520523071289,-78.8207015991211,-78.82275390625,-78.7542953491211,-78.55908203125,-78.4123992919922,-78.2466812133789,-78.1146469116211,-77.9835968017578,-78.0663070678711,-78.1480484008789,-78.248046875,-78.3570327758789,-78.4015655517578,-78.4076156616211,-78.3552780151367,-78.4041061401367,-78.4611358642578,-78.5373077392578,-78.6110382080078,-78.7742156982422,-78.8167037963867,-78.9163055419922,-79.0998077392578,-79.1628875732422,-79.2978515625,-79.400390625,-79.5018615722656,-79.5171890258789,-79.5127944946289,-79.4261703491211,-79.3209915161133,-79.2945251464844,-79.26806640625,-79.3043975830078,-79.32568359375,-79.38720703125,-79.4651412963867,-79.6270523071289,-79.8208999633789,-79.9035110473633,-79.9552764892578,-79.994140625,-80.0095748901367,-79.9954071044922,-79.9968795776367,-80.0896530151367,-80.1668014526367,-80.1529312133789,-80.1312561035156,-80.0949249267578,-80.1600570678711,-80.2950210571289,-80.3381805419922,-80.3827133178711,-80.5308609008789,-80.6174774169922,-80.7189407348633,-80.8026351928711,-80.8600616455078,-80.8865203857422,-80.8379898071289,-80.81591796875,-80.8936538696289,-80.9172821044922,-80.8531188964844,-80.7638626098633,-80.6822280883789,-80.6467742919922,-80.6190414428711,-80.6262741088867,-80.6465835571289,-80.7354507446289,-80.8566436767578,-80.9177703857422,-80.9615249633789,-80.9658203125,-81.0048828125,-80.9679641723633,-81.0041046142578,-81.0738296508789,-81.13037109375,-81.1482391357422,-81.4605484008789,-81.5216827392578,-81.55322265625,-81.5567398071289,-81.57666015625,-81.6361389160156,-81.6783218383789,-81.6839828491211,-81.6671829223633,-81.6760711669922,-81.6597671508789,-81.6716766357422,-81.65234375,-81.6964874267578,-81.7293014526367,-81.7750015258789,-81.78564453125,-81.802734375,-81.7948226928711,-81.8693389892578,-81.8874969482422,-81.9022445678711,-81.9138641357422,-81.9534149169922,-81.9710006713867,-81.9932556152344,-82.0455093383789,-82.1832046508789,-82.2791976928711,-82.2943344116211,-82.2803726196289,-82.3182601928711,-82.3705139160156,-82.3744125366211,-82.3042984008789,-82.3068313598633,-82.2630844116211,-82.2710952758789,-82.1867141723633,-82.1885757446289,-82.1998977661133,-82.2757797241211,-82.4666061401367,-82.5033264160156,-82.52734375,-82.6206970214844,-82.7181701660156,-82.8223648071289,-82.9766616821289,-83.0099563598633,-82.9391555786133,-82.9917984008789,-83.0979461669922,-83.1841735839844,-83.232421875,-83.2793960571289,-83.3412094116211,-83.3956069946289,-83.4275360107422,-83.4407272338867,-83.4423828125,-83.4583969116211,-83.1207046508789,-82.9585952758789,-82.8902359008789,-82.8402328491211,-82.6333999633789,-82.5702133178711,-82.4784240722656,-82.46484375,-82.3162078857422,-82.2005920410156,-82.1784210205078,-82.16943359375,-82.14453125,-82.1536102294922,-82.1348648071289,-82.0615234375,-82.0152282714844,-81.9754867553711,-81.9676742553711,-81.8922882080078,-81.9251937866211,-82.0040054321289,-82.0032272338867,-81.9792022705078,-81.9469680786133,-82.03955078125,-82.15869140625,-82.2710952758789,-82.33984375,-82.45458984375,-82.5116195678711,-82.4949188232422,-82.43798828125,-82.3659210205078,-82.3177719116211,-82.3314514160156,-82.2701187133789,-82.0171813964844,-81.7616195678711,-81.5978546142578,-81.40771484375,-81.2977523803711,-81.21484375,-81.17236328125,-81.1651382446289,-81.0320358276367,-80.88232421875,-80.9542999267578,-81.0638656616211,-80.9747085571289,-80.9595718383789,-80.9205093383789,-80.8909225463867,-80.8126983642578,-80.7457046508789,-80.6506805419922,-80.6034164428711,-80.5404281616211,-80.5186538696289,-80.5316390991211,-80.5138702392578,-80.4607391357422,-80.4446258544922,-80.4501037597656,-80.30810546875,-80.2796859741211,-80.2233352661133,-80.1818389892578,-80.1720733642578,-80.0619125366211,-80.0089874267578,-79.96484375,-79.8589782714844,-79.8148422241211,-79.7708053588867,-79.7579116821289,-79.7746047973633,-79.7624969482422,-79.5990219116211,-79.5853500366211,-79.4852523803711,-79.3042984008789,-79.1630859375,-79.1241226196289,-79.1279296875,-79.1452102661133,-79.22216796875,-79.2288055419922,-79.1102600097656,-78.9775390625,-78.9330062866211,-78.8435592651367,-78.7742156982422,-78.6155242919922,-78.4683609008789,-78.0412063598633,-77.8371124267578,-77.8206100463867,-77.728515625,-77.5218734741211,-77.4253921508789,-77.3112335205078,-77.2816467285156,-77.1408157348633,-77.13623046875,-77.1598587036133,-77.0331039428711,-76.7623062133789,-76.6603469848633,-76.5979461669922,-76.3703155517578,-76.2578125,-76.2263641357422,-76.1573257446289,-76.0546875,-75.9572296142578,-75.7669982910156,-75.7176742553711,-75.6610412597656,-75.6941452026367,-75.68310546875,-75.6343765258789,-75.5939483642578,-75.4919891357422,-75.5399398803711,-75.47021484375,-75.4626007080078,-75.4313430786133,-75.396484375,-75.3899383544922,-75.3423843383789,-75.2420883178711,-75.1153335571289,-75.0519561767578,-74.9745101928711,-74.8629837036133,-74.69921875,-74.3790969848633,-74.3338851928711,-74.31982421875,-74.32763671875,-74.3239288330078,-74.4073257446289,-74.3755798339844,-74.2808609008789,-74.1632766723633,-73.9888610839844,-73.9375,-73.8517532348633,-73.8888702392578,-73.9456100463867,-73.9157180786133,-73.8669967651367,-73.8302764892578,-73.81591796875,-73.7576141357422,-73.7090759277344,-73.6813507080078,-73.64404296875,-73.5559616088867,-73.4256820678711,-73.3884735107422,-73.33447265625,-73.1521453857422,-73.0967788696289,-73.0474624633789,-73.123046875,-73.1024398803711,-73.0005874633789,-72.84326171875,-72.7845687866211,-72.7330093383789,-72.7562484741211,-72.7920913696289,-72.7850570678711,-72.62890625,-72.4981384277344,-72.4440383911133,-72.4128952026367,-72.2816390991211,-72.0315475463867,-71.9019546508789,-71.8223648071289,-71.7579116821289,-71.77685546875,-71.7855453491211,-71.7193374633789,-71.6136703491211,-71.4951171875,-71.3897476196289,-71.3322296142578,-71.296875,-71.2886734008789,-71.3312530517578,-71.4397506713867,-71.4811553955078,-71.56005859375,-71.5533218383789,-71.4426803588867,-71.2955093383789,-71.2502899169922,-71.2009811401367,-71.1306610107422,-71.0609359741211,-71.0240249633789,-70.982421875,-70.9357452392578,-70.9263610839844,-71.02734375,-71.0953140258789,-71.1175765991211,-71.1740188598633,-71.361328125,-71.6727523803711,-71.6748046875,-71.6470718383789,-71.6353530883789,-71.5464859008789,-71.3243179321289,-71.2237319946289,-71.1214828491211,-71.0171890258789,-70.9403305053711,-70.8845672607422,-70.8427734375,-70.8290100097656,-70.7869110107422,-70.8351516723633,-70.8445358276367,-70.81689453125,-70.7557601928711,-70.7126922607422,-70.7453079223633,-70.8567352294922,-70.96826171875,-71.0518569946289,-71.1541061401367,-71.265625,-71.3259811401367,-71.3415985107422,-71.3277359008789,-71.3384780883789,-71.3388671875,-71.3746109008789,-71.3604507446289,-71.3209991455078,-71.32080078125,-71.3571319580078,-71.4333953857422,-71.4123077392578,-71.3868179321289,-71.2841796875,-71.265625,-71.3492202758789,-71.5397491455078,-71.6302795410156,-71.7126998901367,-71.6418914794922,-71.5588912963867,-71.4711990356445,-71.39794921875,-71.2742156982422,-71.2023468017578,-71.0802764892578,-71.0036163330078,-70.9000930786133,-70.8444366455078,-70.6556625366211,-70.6363296508789,-70.58642578125,-70.5352554321289,-70.4944305419922,-70.3563537597656,-70.4616165161133,-70.4738311767578,-70.3908233642578,-70.1836929321289,-70.26904296875,-70.4029312133789,-70.5079116821289,-70.6876983642578,-70.7287139892578,-70.7666015625,-70.7365264892578,-70.6165084838867,-70.4432601928711,-70.3701171875,-70.3086929321289,-70.21337890625,-70.1623077392578,-70.1436538696289,-70.1173858642578,-70.0656204223633,-70.0536117553711,-70.2295913696289,-70.2875061035156,-70.3431625366211,-70.2996139526367,-70.2947311401367,-70.3307647705078,-70.3240203857422,-70.1934585571289,-70.1451110839844,-70.2038040161133,-70.3972625732422,-70.4508819580078,-70.5403289794922,-70.5182571411133,-70.41552734375,-70.2899398803711,-70.2694396972656,-70.2013702392578,-70.2121124267578,-70.2931671142578,-70.4918975830078,-70.6740264892578,-70.8208999633789,-70.90234375,-70.9169921875,-70.9206085205078,-70.91015625,-70.9175796508789,-70.8434600830078,-70.6805648803711,-70.4951171875,-70.2584915161133,-70.2767562866211,-70.3003921508789,-70.4172821044922,-70.47509765625,-70.5613250732422,-70.6426696777344,-70.6956100463867,-70.73974609375,-70.8103561401367,-70.7962875366211,-70.7232437133789,-70.5749969482422,-70.4045944213867,-70.4133758544922,-70.4486312866211,-70.5369110107422,-70.7043991088867,-70.8204040527344,-70.89208984375,-70.9709930419922,-70.9906234741211,-71.0373992919922,-71.01953125,-70.96728515625,-70.9537124633789,-70.9109344482422,-70.8132858276367,-70.7724609375,-70.6820297241211,-70.5831069946289,-70.40625,-70.3000030517578,-70.2462921142578,-70.2247085571289,-70.2257766723633,-70.0998001098633,-70.0259780883789,-70.0005874633789,-69.9093780517578,-69.7336883544922,-69.6242141723633,-69.5169906616211,-69.3787155151367,-69.2548828125,-69.07421875,-68.8689498901367,-68.7831039428711,-68.6891632080078,-68.6707000732422,-68.6830062866211,-68.7024383544922,-68.7906265258789,-68.8853530883789,-68.9792938232422,-69.02001953125,-69.0993194580078,-69.1105422973633,-69.0945281982422,-69.1676712036133,-69.4869155883789,-69.6373062133789,-69.6813430786133,-69.6600646972656,-69.6618194580078,-69.6393508911133,-69.6378936767578,-69.6522445678711,-69.7255859375,-69.8104476928711,-69.7478485107422,-69.7183532714844,-69.7256851196289,-69.82421875,-70.05615234375,-70.171875,-70.0978469848633,-70.0060577392578,-69.9242172241211,-69.7859344482422,-69.6080093383789,-69.4255905151367,-69.17333984375,-68.9669952392578,-68.8677749633789,-68.8048782348633,-68.7388687133789,-68.7236328125,-68.5750961303711,-68.5149383544922,-68.4326171875,-68.3518524169922,-68.1232376098633,-68.0953140258789,-68.0597686767578,-68.0456008911133,-67.9724578857422,-67.9613265991211,-67.904296875,-67.7692413330078,-67.7311553955078,-67.7364273071289,-67.6597671508789,-67.6570358276367,-67.6175765991211,-67.5334014892578,-67.4765625,-67.4019546508789,-67.3036117553711,-67.2681655883789,-67.2748031616211,-67.3572311401367,-67.3619155883789,-67.4091796875,-67.46826171875,-67.5726623535156,-67.6649398803711,-67.7279281616211,-67.7162094116211,-67.6600570678711,-67.6993179321289,-67.7852554321289,-67.91748046875,-67.9880828857422,-67.9263687133789,-67.7940368652344,-67.6251983642578,-67.5206069946289,-67.3524398803711,-67.2268524169922,-67.19970703125,-67.2168960571289,-67.1712875366211,-67.04345703125,-66.9382781982422,-66.9415969848633,-67.0309600830078,-67.1751937866211,-67.1721649169922,-67.1943359375,-67.1501922607422,-66.9385757446289,-66.8200225830078,-66.7533264160156,-66.6034164428711,-66.4446258544922,-66.3564453125,-66.3154296875,-66.0721664428711,-66.02001953125,-65.9691467285156,-65.9455108642578,-65.8586883544922,-65.9163131713867,-65.9542007446289,-66.0026397705078,-66.0799865722656,-66.2093734741211,-66.3727493286133,-66.4234390258789,-66.4748001098633,-66.61328125,-66.7043914794922,-66.7126998901367,-66.6592788696289,-66.6034164428711,-66.6260757446289,-66.7210922241211,-66.77978515625,-66.8591766357422,-66.9738311767578,-67.1993179321289,-67.2645492553711,-67.2124938964844,-67.1159210205078,-67.0733413696289,-67.0562515258789,-67.0526351928711,-67.0140609741211,-67.0413131713867,-67.1034164428711,-67.1630859375,-67.2295913696289,-67.4849624633789,-67.4585952758789,-67.4031295776367,-67.3851547241211,-67.49951171875,-67.5402374267578,-67.5754852294922,-67.6475601196289,-67.5618133544922,-67.52685546875,-67.50830078125,-67.5260772705078,-67.6204071044922,-67.7164077758789,-67.7655334472656,-67.7679672241211,-67.8101577758789,-67.8541030883789,-67.8895568847656,-67.8621063232422,-67.8248062133789,-67.7429733276367,-67.76318359375,-67.8645477294922,-68.041015625,-68.1608428955078,-68.2794952392578,-68.3794860839844,-68.4642562866211,-68.5353546142578,-68.5985336303711,-68.7369155883789,-68.8566360473633,-68.9322280883789,-69.0240173339844,-69.1537094116211,-69.2316360473633,-69.2937545776367,-69.3314437866211,-69.33740234375,-69.3727569580078,-69.4697265625,-69.5402297973633,-69.5779266357422,-69.6548843383789,-69.7696304321289,-69.8666000366211,-69.9118118286133,-69.92138671875,-69.9021530151367,-69.8373031616211,-69.8944320678711,-69.9527359008789,-70.087890625,-70.1771469116211,-70.2731475830078,-70.3259735107422,-70.4228515625,-70.4124984741211,-70.3699264526367,-70.3251953125,-70.333984375,-70.4310531616211,-70.5852508544922,-70.70458984375,-71.03515625,-71.0907211303711,-71.1814422607422,-71.2515640258789,-71.2865219116211,-71.3910140991211,-71.5797882080078,-71.7367172241211,-72.0029296875,-72.2906265258789,-72.46142578125,-72.64111328125,-72.9486312866211,-72.9998016357422,-73.12548828125,-73.2091751098633,-73.2168960571289,-73.2364196777344,-73.30029296875,-73.1681594848633,-73.0856475830078,-72.9181671142578,-72.7505874633789,-72.6506881713867,-72.5150375366211,-72.4186553955078,-72.4087905883789,-72.37451171875,-72.2536163330078,-72.1329116821289,-72.0553741455078,-71.9309539794922,-71.8420867919922,-71.822265625,-71.7367172241211,-71.6715774536133,-71.6239242553711,-71.5138702392578,-71.3091735839844,-71.1545867919922,-70.94921875,-70.80126953125,-70.7066345214844,-70.65673828125,-70.5986328125,-70.4720687866211,-70.3956985473633,-70.2391586303711,-70.0958938598633,-70.0096664428711,-69.8489227294922,-69.8257827758789,-69.7431640625,-69.8003921508789,-69.8795928955078,-69.85546875,-69.89306640625,-69.8489227294922,-69.7254867553711,-69.5755844116211,-69.4874038696289,-69.490234375,-69.3396530151367,-69.2059555053711,-69.1744155883789,-69.0689392089844,-68.8918991088867,-68.7561569213867,-68.6257781982422,-68.3937530517578,-68.27783203125,-68.1753921508789,-68.1193313598633,-67.9468688964844,-67.8312530517578,-67.6900329589844,-67.6916961669922,-67.6130905151367,-67.6105499267578,-67.60302734375,-67.4412155151367,-67.2919921875,-67.244140625,-67.1144485473633,-67.1022415161133,-67.1255798339844,-67.1609344482422,-67.1613235473633,-67.0549774169922,-67.037109375,-66.9855422973633,-66.9401397705078,-66.7884826660156,-66.8174819946289,-66.7919921875,-66.7994155883789,-66.8181610107422,-66.8230438232422,-66.7696304321289,-66.7342758178711,-66.6028366088867,-66.5720748901367,-66.5374984741211,-66.5079116821289,-66.55859375,-66.6042938232422,-66.6144561767578,-66.6244125366211,-66.5710906982422,-66.58544921875,-66.6429672241211,-66.6896438598633,-66.6888656616211,-66.6471710205078,-66.5435562133789,-66.5013656616211,-66.5274429321289,-66.5711898803711,-66.6309509277344,-66.6211929321289,-66.599609375,-66.5505828857422,-66.4994125366211,-66.57861328125,-66.6073226928711,-66.4674835205078,-66.49853515625,-66.5347671508789,-66.5531234741211,-66.6707992553711,-66.6482391357422,-66.5486297607422,-66.4739303588867,-66.4251937866211,-66.3580093383789,-66.1004867553711,-66.0208969116211,-65.9730529785156,-65.951171875,-65.9542007446289,-65.9326171875,-65.8651428222656,-65.9168930053711,-65.9989242553711,-65.98974609375,-65.9880828857422,-66.0391616821289,-66.1368179321289,-66.1640625,-66.41064453125,-66.4704132080078,-66.5523452758789,-66.5803680419922,-66.6640625,-66.6720733642578,-66.6390609741211,-66.7658233642578,-66.8619155883789,-66.90869140625,-66.8336944580078,-66.62109375,-66.5240173339844,-66.3123016357422,-66.07666015625,-65.9609375,-65.9000930786133,-65.8479461669922,-65.7999038696289,-65.8487243652344,-65.8863296508789,-65.9295883178711,-66.0604476928711,-66.3601608276367,-66.4679641723633,-66.4769515991211,-66.4929733276367,-66.5608444213867,-66.7711944580078,-66.9580078125,-67.0279312133789,-67.1084899902344,-67.1722640991211,-67.2119140625,-67.36767578125,-67.4412155151367,-67.4056625366211,-67.3878860473633,-67.3565444946289,-67.3078155517578,-67.2380828857422,-67.2019500732422,-67.1427764892578,-67.10791015625,-67.0471649169922,-67.0554733276367,-67.1143569946289,-67.1092758178711,-67.1285095214844,-67.0853500366211,-67.0824203491211,-67.1150360107422,-67.1609344482422,-67.1716766357422,-67.1448211669922,-67.07080078125,-66.9915084838867,-66.9662170410156,-66.9666061401367,-66.9837875366211,-67.0758819580078,-67.19921875,-67.3197250366211,-67.3707046508789,-67.2360305786133,-67.1357421875,-67.0907211303711,-67.0570297241211,-66.9017562866211,-66.8594741821289,-66.8048858642578,-66.7451171875,-66.6768493652344,-66.6081085205078,-66.6007766723633,-66.6522445678711,-66.7082061767578,-66.6945266723633,-66.64111328125,-66.5158157348633,-66.4244155883789,-66.3933563232422,-66.3644561767578,-66.3955078125,-66.46240234375,-66.49755859375,-66.7593765258789,-66.9896469116211,-67.0510711669922,-67.0279312133789,-67.119140625,-67.1071243286133,-67.0803756713867,-67.0982437133789,-67.0415954589844,-66.7529296875,-66.4685516357422,-66.3448257446289,-66.29150390625,-66.2684555053711,-66.2085952758789,-66.19140625,-66.2155303955078,-66.23583984375,-66.1654281616211,-66.1780319213867,-66.0948257446289,-66.0814437866211,-66.1536102294922,-66.2042999267578,-66.2770462036133,-66.34765625,-66.4767532348633,-66.4798812866211,-66.3533172607422,-66.3301696777344,-66.1271438598633,-66.1800765991211,-66.2667922973633,-66.2921905517578,-66.43896484375,-66.40771484375,-66.3396453857422,-66.3464813232422,-66.4064483642578,-66.4569320678711,-66.5439453125,-66.564453125,-66.5404281616211,-66.5740280151367,-66.6376037597656,-66.7151336669922,-66.751953125,-66.8318405151367,-66.7940444946289,-66.8067321777344,-66.8736343383789,-66.9483413696289,-67.0127944946289,-67.0000991821289,-66.9486312866211,-66.8767623901367,-66.8767623901367,-66.9385757446289,-67.0907211303711,-67.0876922607422,-67.0179672241211,-67.0355453491211,-67.1414108276367,-67.2825164794922,-67.4787139892578,-67.6441345214844,-67.7940368652344,-67.8645477294922,-67.8998031616211,-67.7942352294922,-67.7208938598633,-67.6259765625,-67.5909194946289,-67.6242218017578,-67.7508773803711,-67.9646530151367,-68.041015625,-68.1203079223633,-68.1912078857422,-68.2736358642578,-68.36865234375,-68.38427734375,-68.3750991821289,-68.4669876098633,-68.43115234375,-68.4313507080078,-68.4177703857422,-68.419921875,-68.4357452392578,-68.4029312133789,-68.3584899902344,-68.3849639892578,-68.623046875,-68.76416015625,-68.8170928955078,-68.76416015625,-68.6936492919922,-68.7255859375,-68.7295913696289,-68.7676773071289,-68.8568344116211,-68.8179702758789,-68.7643585205078,-68.7289047241211,-68.640625,-68.4933624267578,-68.349609375,-68.3231506347656,-68.349609375,-68.4178695678711,-68.63427734375,-68.7743148803711,-68.84130859375,-68.8946304321289,-69.0244140625,-69.0778350830078,-69.1830062866211,-69.17626953125,-69.3086853027344,-69.2046890258789,-69.1807632446289,-69.2088851928711,-69.2994155883789,-69.3201141357422,-69.4683609008789,-69.5218734741211,-69.6305694580078,-69.7342758178711,-69.8401412963867,-69.9749069213867,-70.08056640625,-70.1820297241211,-70.3171844482422,-70.8267593383789,-70.9161148071289,-70.9073257446289,-71.03955078125,-71.0218811035156,-70.9690475463867,-70.625,-70.4398422241211,-70.333984375,-70.3045883178711,-70.5013732910156,-70.6208953857422,-70.6422805786133,-70.6365280151367,-70.5104446411133,-70.5565414428711,-70.57080078125,-70.6453094482422,-70.6328125,-70.6642532348633,-70.7712936401367,-70.8102493286133,-70.8543930053711,-70.9249038696289,-71.0130844116211,-71.0924835205078,-71.1831970214844,-71.1973648071289,-71.2748031616211,-71.5113296508789,-71.5806655883789,-71.6304702758789,-71.5687484741211,-71.4439392089844,-71.4187545776367,-71.60400390625,-71.7451171875,-71.8685531616211,-71.9685592651367,-71.9479522705078,-71.9479522705078,-72.1155242919922,-72.40283203125,-72.3977508544922,-72.3712921142578,-72.4771423339844,-72.5653305053711,-72.6005859375,-72.5337905883789,-72.48681640625,-72.46875,-72.3844757080078,-72.3791046142578,-72.3833999633789,-72.47265625,-72.5524444580078,-72.6212921142578,-72.7288131713867,-72.7946243286133,-72.876953125,-73.0503921508789,-73.2003860473633,-73.0912170410156,-73.06591796875,-73.1297836303711,-73.1224594116211,-73.1472625732422,-73.01123046875,-72.93603515625,-72.9974594116211,-73.2243118286133,-73.2757797241211,-73.3368148803711,-73.3942413330078,-73.447265625,-73.4400405883789,-73.5443344116211,-73.5269546508789,-73.5337905883789,-73.57666015625,-73.5926818847656,-73.6307678222656,-73.7351531982422,-73.8228530883789,-73.86669921875,-73.8460922241211,-73.87939453125,-73.7824249267578,-73.5711898803711,-73.3826141357422,-73.37451171875,-73.3967742919922,-73.5587844848633,-73.8376998901367,-73.9258728027344,-74.0029296875,-74.0285110473633,-74.2634811401367,-74.4261703491211,-74.4791946411133,-74.55859375,-74.59375,-74.5626983642578,-74.5783233642578,-74.568359375,-74.5334014892578,-74.5232467651367,-74.5673828125,-74.5637664794922,-74.4173812866211,-74.3821258544922,-74.42626953125,-74.6019592285156,-74.6560592651367,-74.7361373901367,-74.8231430053711,-75.1670913696289,-75.2376022338867,-75.2345657348633,-75.2616195678711,-75.23388671875,-75.2177734375,-75.3346710205078,-75.3958969116211,-75.3861312866211,-75.4042053222656,-75.4668960571289,-75.6216812133789,-75.6865234375,-75.7580032348633,-75.7932662963867,-75.84619140625,-75.9519577026367,-76.0490188598633,-76.1548843383789,-76.2077102661133,-76.2253875732422,-76.4635696411133,-76.5693359375,-76.6575241088867,-76.7457046508789,-76.8287124633789,-76.8692398071289,-76.9556655883789,-77.0067367553711,-77.0235366821289,-77.0323181152344,-77.12646484375,-77.2693328857422,-77.3877944946289,-77.5823211669922,-77.6999053955078,-77.7746124267578,-77.85302734375,-77.8770523071289,-77.8834915161133,-77.9540023803711,-78.0421829223633,-78.1467742919922,-78.2238311767578,-78.2362289428711,-78.3156280517578,-78.22607421875,-78.1286163330078,-78.0421829223633,-78.0635757446289,-78.15380859375,-78.3056640625,-78.45166015625,-78.4973602294922,-78.5215835571289,-78.5184555053711,-78.6061553955078,-78.68603515625,-78.6798858642578,-78.6949157714844,-78.6278305053711,-78.5710983276367,-78.6032257080078,-78.6300811767578,-78.7170867919922,-78.7585983276367,-78.8448181152344,-78.8977584838867,-78.6942367553711,-78.544921875,-78.5361328125,-78.5713882446289,-78.67724609375,-78.9074172973633,-78.96826171875,-79.0299835205078,-79.0609359741211,-79.0599594116211,-79.0150375366211,-78.9787139892578,-79.0497055053711,-79.2014694213867,-79.3023452758789,-79.35888671875,-79.4268569946289,-79.5543975830078,-79.5612335205078,-79.5856475830078,-79.6325225830078,-79.6355438232422,-79.6915054321289,-79.7366180419922,-79.7898483276367,-79.8589782714844,-79.8924865722656,-79.8794937133789,-79.9295883178711,-80.0105514526367,-80.0544891357422,-80.0880889892578,-80.2933578491211,-80.3489227294922,-80.4234390258789,-80.4425811767578,-80.4250030517578,-80.4499053955078,-80.5076141357422,-80.5833969116211,-80.6365203857422,-80.6740188598633,-80.7298889160156,-80.7531204223633,-80.779296875,-80.7867202758789,-80.8467788696289,-80.9011764526367,-80.9074249267578,-81.1128845214844,-81.1996078491211,-81.24169921875,-81.2701187133789,-81.3406295776367,-81.3902359008789,-81.6097640991211,-81.6104507446289,-81.6530227661133,-81.7649383544922,-81.83203125,-81.8662109375,-81.9207000732422,-81.9689407348633,-82.12060546875,-82.1873016357422,-82.314453125,-82.4078063964844,-82.4899444580078,-82.4815444946289,-82.5349578857422,-82.5189437866211,-82.4632797241211,-82.3967819213867,-82.3544006347656,-82.3849639892578,-82.6296920776367,-82.72216796875,-82.7570343017578,-82.7646484375,-82.80126953125,-82.87939453125,-82.9523468017578,-82.98583984375,-82.9988250732422,-83.0474624633789,-83.03076171875,-82.9747085571289,-82.9872055053711,-83.0653305053711,-83.1267547607422,-83.1550827026367,-83.2107391357422,-83.2298812866211,-83.2431640625,-83.2429733276367,-83.2314453125,-83.2721710205078,-83.3162078857422,-83.3620147705078,-83.3990173339844,-83.4788131713867,-83.4357376098633,-83.4484405517578,-83.4749984741211,-83.5554656982422,-83.5813446044922,-83.64404296875,-83.6753921508789,-83.6731414794922,-83.7587890625,-83.7609405517578,-83.8101577758789,-83.8390655517578,-83.87744140625,-83.9400405883789,-83.9675827026367,-83.97314453125,-84.0749053955078,-84.1298828125,-84.1266632080078,-84.1357421875,-84.1814498901367,-84.2061538696289,-84.2683639526367,-84.3515625],[-63.1735343933105,-63.1996116638184,-63.1986351013184,-63.2047882080078,-63.2493133544922,-63.3243179321289,-63.3531265258789,-63.3358383178711,-63.2966804504395,-63.2984352111816,-63.3414993286133,-63.3826179504395,-63.4373092651367,-63.4180679321289,-63.3576202392578,-63.3342742919922,-63.3182640075684,-63.3019523620605,-63.2835006713867,-63.2340812683105,-63.1571273803711,-63.1283187866211,-63.1568336486816,-63.1735343933105],[-63.0785179138184,-63.1731452941895,-63.1688461303711,-63.0983390808105,-63.0616188049316,-63.0451202392578,-62.9822273254395,-63.0052719116211,-63.0126953125,-63.0185546875,-63.0546875,-63.0785179138184],[-62.9673843383789,-62.9775390625,-62.9690437316895,-62.9866180419922,-63.0089874267578,-63.0179672241211,-62.9957008361816,-62.9486351013184,-62.9055633544922,-62.8952178955078,-62.9673843383789],[-63.0693397521973,-63.0738258361816,-63.0319366455078,-62.923828125,-62.8741188049316,-62.9177780151367,-62.9715843200684,-63.0693397521973],[-62.56005859375,-62.5726547241211,-62.5487289428711,-62.6184577941895,-62.6149406433105,-62.7454109191895,-62.7075233459473,-62.67919921875,-62.6165046691895,-62.6333999633789,-62.6207046508789,-62.6623077392578,-62.6791038513184,-62.6789054870605,-62.6341819763184,-62.5890617370605,-62.5916976928711,-62.5336952209473,-62.4751930236816,-62.4910125732422,-62.56005859375],[-62.4443359375,-62.4514656066895,-62.39501953125,-62.3543014526367,-62.3521499633789,-62.3672828674316,-62.4128875732422,-62.4443359375],[-62.3025398254395,-62.3477554321289,-62.3017539978027,-62.2831077575684,-62.2390594482422,-62.2492179870605,-62.2638664245605,-62.2678718566895,-62.3025398254395],[-61.9119148254395,-61.9399375915527,-61.9211921691895,-61.9420890808105,-61.9982414245605,-62.0204124450684,-62.0119171142578,-62.0775413513184,-62.0634765625,-62.1177711486816,-62.1458015441895,-62.1700210571289,-62.1194381713867,-62.13720703125,-62.2256889343262,-62.2439460754395,-62.2477569580078,-62.2251968383789,-62.2178726196289,-62.2060585021973,-62.1712913513184,-62.2097663879395,-62.1642570495605,-62.0447273254395,-62.0082015991211,-61.9382781982422,-61.9533195495605,-61.9119148254395],[-61.2991256713867,-61.3084945678711,-61.2697257995605,-61.2465782165527,-61.2017593383789,-61.14208984375,-61.13525390625,-61.25537109375,-61.2991256713867],[-61.2204132080078,-61.2485313415527,-61.2116241455078,-61.1463890075684,-61.1061515808105,-61.0726547241211,-61.1169891357422,-61.1397476196289,-61.1686553955078,-61.2204132080078],[-60.5209007263184,-60.5464859008789,-60.5827140808105,-60.6238288879395,-60.6397438049316,-60.6481437683105,-60.671875,-60.69873046875,-60.7330093383789,-60.6497116088867,-60.6454086303711,-60.5860328674316,-60.6199226379395,-60.5974578857422,-60.5683631896973,-60.5265617370605,-60.5434608459473,-60.5209007263184],[-12.4256830215454,-12.4359369277954,-12.4341802597046,-12.4239253997803,-12.4256830215454],[-49.0588836669922,-49.0667991638184,-49.0342750549316,-48.9912109375,-48.95703125,-48.9190444946289,-48.8829078674316,-48.8788070678711,-48.8904304504395,-48.951171875,-49.0342750549316,-49.0588836669922],[-49.1095695495605,-49.1154289245605,-49.1062507629395,-48.9743194580078,-48.9709930419922,-49.0719718933105,-49.1173820495605,-49.1290054321289,-49.1240234375,-49.1817359924316,-49.2556648254395,-49.2658195495605,-49.2649421691895,-49.2481422424316,-49.2215805053711,-49.1598625183105,-49.1360359191895,-49.1349601745605,-49.0764656066895,-49.05859375,-49.0611343383789,-49.0838890075684,-49.1369132995605,-49.2014656066895,-49.2655258178711,-49.32763671875,-49.3656272888184,-49.4109382629395,-49.4339866638184,-49.4352531433105,-49.4248046875,-49.37158203125,-49.3429718017578,-49.3449249267578,-49.3485374450684,-49.3892593383789,-49.4205055236816,-49.4376945495605,-49.43017578125,-49.4475593566895,-49.4901351928711,-49.5440444946289,-49.5816421508789,-49.58935546875,-49.5177726745605,-49.5093727111816,-49.5306625366211,-49.58349609375,-49.6007843017578,-49.6288070678711,-49.6650390625,-49.7043914794922,-49.7085952758789,-49.6893577575684,-49.6449241638184,-49.6135749816895,-49.6017608642578,-49.6421890258789,-49.6509780883789,-49.6173820495605,-49.5631828308105,-49.5427742004395,-49.5296859741211,-49.6529312133789,-49.7049789428711,-49.7098655700684,-49.6996078491211,-49.6512680053711,-49.599609375,-49.5501937866211,-49.4996109008789,-49.4443359375,-49.3921890258789,-49.3539047241211,-49.2853507995605,-49.2316398620605,-49.1920928955078,-49.1649398803711,-49.1413078308105,-49.1350593566895,-49.1037139892578,-49.06591796875,-48.9947242736816,-48.9261703491211,-48.8487319946289,-48.7755851745605,-48.69384765625,-48.6612319946289,-48.6564445495605,-48.6792984008789,-48.7239265441895,-48.7528305053711,-48.7660179138184,-48.8610343933105,-48.89990234375,-48.9375991821289,-49.017578125,-49.0819358825684,-49.1095695495605],[-46.43994140625,-46.44873046875,-46.4281272888184,-46.3736343383789,-46.3268547058105,-46.35888671875,-46.3947257995605,-46.43994140625],[17.0370121002197,16.9971675872803,17.0133304595947,17.0631332397461,17.09814453125,17.1689453125,17.1384754180908,17.1050777435303,17.0984382629395,17.06982421875,17.0489253997803,17.0370121002197],[17.574951171875,17.5486812591553,17.5961418151855,17.6854496002197,17.7042961120605,17.7140617370605,17.6968746185303,17.6904792785645,17.6613292694092,17.574951171875],[-54.7097625732422,-54.7492179870605,-54.7040023803711,-54.5060539245605,-54.4723625183105,-54.5750007629395,-54.7097625732422],[-43.39697265625,-43.5007820129395,-43.4831047058105,-43.5001945495605,-43.43115234375,-43.4128913879395,-43.4302749633789,-43.4078140258789,-43.3791961669922,-43.3713874816895,-43.3304672241211,-43.2789077758789,-43.2802734375,-43.3462905883789,-43.39697265625],[-43.24072265625,-43.2408218383789,-43.232421875,-43.1833000183105,-43.1617164611816,-43.1453094482422,-43.1146469116211,-43.0802726745605,-43.1182594299316,-43.24072265625],[-42.71044921875,-42.71923828125,-42.71484375,-42.6633796691895,-42.6404304504395,-42.5931663513184,-42.6159172058105,-42.6517601013184,-42.6805686950684,-42.71044921875],[-40.7867164611816,-40.7906227111816,-40.76513671875,-40.7699203491211,-40.8263664245605,-40.8582000732422,-40.8523445129395,-40.8639640808105,-40.9041023254395,-40.9390602111816,-40.9620094299316,-40.9971694946289,-41.0246086120605,-41.1180648803711,-41.1634788513184,-41.1423835754395,-41.1162109375,-41.0780296325684,-41.1136703491211,-41.1680679321289,-41.109375,-41.0583000183105,-41.0177726745605,-40.9923858642578,-40.9942398071289,-40.9833984375,-40.9597663879395,-40.9564437866211,-40.9855461120605,-41.0016593933105,-40.9641571044922,-40.8755836486816,-40.8447265625,-40.8716812133789,-40.87255859375,-40.7795906066895,-40.7809562683105,-40.8548812866211,-40.9470710754395,-41.1153297424316,-41.1746101379395,-41.2331047058105,-41.3497047424316,-41.4650421142578,-41.5549812316895,-41.6461906433105,-41.8157234191895,-41.927734375,-42.0041999816895,-42.0399398803711,-42.07373046875,-42.1111335754395,-42.1591796875,-42.21533203125,-42.2616195678711,-42.2549781799316,-42.2194328308105,-42.1734352111816,-42.1364250183105,-42.1026344299316,-42.0647430419922,-42.0419921875,-42.0218734741211,-41.9700202941895,-42.0123062133789,-42.06982421875,-42.0882797241211,-42.1037101745605,-42.1703109741211,-42.2594718933105,-42.3451156616211,-42.4359397888184,-42.505859375,-42.5724601745605,-42.6584968566895,-42.81640625,-42.96044921875,-43.1570320129395,-43.1818351745605,-43.1951179504395,-43.2200202941895,-43.12255859375,-43.0206069946289,-42.9798851013184,-43.00341796875,-42.9802742004395,-42.9541015625,-42.9281234741211,-42.8719749450684,-42.845703125,-42.8780288696289,-42.9745101928711,-42.9964866638184,-43.0333976745605,-42.8938484191895,-42.7909164428711,-42.8405265808105,-42.9265594482422,-42.9644546508789,-43.0134773254395,-43.03173828125,-43.0710945129395,-43.12646484375,-43.2159194946289,-43.255859375,-43.1563491821289,-43.1898422241211,-43.21875,-43.27587890625,-43.3190422058105,-43.36962890625,-43.50244140625,-43.6124992370605,-43.6193351745605,-43.6019554138184,-43.5088844299316,-43.51953125,-43.5127906799316,-43.5471687316895,-43.44482421875,-43.4084014892578,-43.3760719299316,-43.3543930053711,-43.3552742004395,-43.3162078857422,-43.3017578125,-43.3112297058105,-43.2771492004395,-43.2923812866211,-43.2440414428711,-43.0759773254395,-42.9982452392578,-42.9679679870605,-42.9513664245605,-42.9266586303711,-42.5443382263184,-42.45556640625,-42.2308578491211,-42.3384780883789,-42.4065437316895,-42.4928703308105,-42.38818359375,-42.3544921875,-42.2275390625,-42.1907234191895,-42.1470680236816,-42.1910133361816,-42.1969718933105,-42.1073226928711,-42.0196266174316,-41.8267555236816,-41.64404296875,-41.4188461303711,-41.3900375366211,-41.3415031433105,-41.1907234191895,-41.0789070129395,-40.9808578491211,-40.7829093933105,-40.6722679138184,-40.7216796875,-40.7867164611816],[-40.51513671875,-40.5925788879395,-40.5439453125,-40.521484375,-40.5031242370605,-40.5050773620605,-40.51513671875],[-40.5067367553711,-40.5894546508789,-40.4852523803711,-40.47021484375,-40.4348640441895,-40.4403305053711,-40.5067367553711],[-40.3069343566895,-40.3671875,-40.4324226379395,-40.4865226745605,-40.4972648620605,-40.4345703125,-40.45751953125,-40.45166015625,-40.4041976928711,-40.3805656433105,-40.3568344116211,-40.35791015625,-40.3069343566895],[-39.7576141357422,-39.9384765625,-39.9666976928711,-39.9857406616211,-40.0654296875,-40.0995101928711,-40.1444320678711,-40.1735343933105,-40.1724624633789,-40.2336921691895,-40.2621116638184,-40.2408218383789,-40.2127952575684,-40.1719741821289,-40.0145492553711,-39.9713859558105,-39.9054679870605,-39.9104499816895,-39.8703155517578,-39.83154296875,-39.7259750366211,-39.7576141357422],[-40.1161117553711,-40.1202125549316,-40.06396484375,-39.9835929870605,-39.9041023254395,-39.82421875,-39.7379875183105,-39.7000007629395,-39.6581077575684,-39.5836906433105,-39.5801734924316,-39.6380882263184,-39.7852554321289,-39.8740234375,-39.9538078308105,-40.0220718383789,-40.0782203674316,-40.1161117553711],[-38.4908218383789,-38.5381851196289,-38.5570335388184,-38.5197257995605,-38.5276336669922,-38.4585914611816,-38.47216796875,-38.4908218383789],[-38.3548812866211,-38.4209938049316,-38.390625,-38.3410148620605,-38.3189468383789,-38.3141593933105,-38.3548812866211],[-35.7386703491211,-35.7621078491211,-35.72607421875,-35.7551765441895,-35.8523406982422,-35.9005889892578,-35.9076156616211,-35.8677711486816,-35.89794921875,-35.9380836486816,-36.0271492004395,-36.0748062133789,-36.0208969116211,-35.982421875,-36.0390625,-36.02392578125,-36.0466766357422,-36.0331077575684,-35.9353485107422,-35.89013671875,-35.8086891174316,-35.7488288879395,-35.6638679504395,-35.5924797058105,-35.6050796508789,-35.6202163696289,-35.6564445495605,-35.7222633361816,-35.7386703491211],[-27.4364261627197,-27.7117176055908,-27.7064456939697,-27.6650390625,-27.5056629180908,-27.4053707122803,-27.4224624633789,-27.4364261627197],[-27.3160152435303,-27.3309574127197,-27.2353515625,-27.1388664245605,-27.0494136810303,-27.0298824310303,-27.0380859375,-27.2014636993408,-27.3160152435303],[-26.0531253814697,-26.0945320129395,-25.7831058502197,-25.56982421875,-25.5315418243408,-25.5202140808105,-25.8150405883789,-25.8826179504395,-25.9519519805908,-25.9738273620605,-26.0531253814697],[-25.7507820129395,-25.7783203125,-25.7289066314697,-25.5513668060303,-25.4484386444092,-25.3542976379395,-25.30224609375,-25.1931648254395,-25.0705089569092,-25.0057621002197,-24.9225597381592,-24.8326187133789,-24.8148441314697,-24.7648429870605,-24.7395515441895,-24.7289066314697,-24.73828125,-24.9152336120605,-24.9777336120605,-25.0630855560303,-25.5127944946289,-25.6825199127197,-25.7507820129395],[-23.4908199310303,-23.5162124633789,-23.5130863189697,-23.5296878814697,-23.5949211120605,-23.66845703125,-23.7203121185303,-23.7623043060303,-23.7757816314697,-23.74072265625,-23.5301761627197,-23.4605464935303,-23.4908199310303],[-22.3225593566895,-22.32470703125,-22.3077163696289,-22.2678718566895,-22.2107429504395,-22.2283210754395,-22.3069324493408,-22.3225593566895],[-22.1930656433105,-22.2232437133789,-22.150390625,-22.0740242004395,-22.0487308502197,-22.1493167877197,-22.1930656433105],[-20.7877941131592,-20.8660163879395,-20.8505859375,-20.8111324310303,-20.7462902069092,-20.66796875,-20.71630859375,-20.7877941131592],[-20.29150390625,-20.3025398254395,-20.3017578125,-20.28369140625,-20.1535148620605,-20.2214832305908,-20.2775382995605,-20.29150390625],[-20.14990234375,-20.154296875,-20.1435546875,-20.1019535064697,-20.0689449310303,-20.0443363189697,-20.1346683502197,-20.14990234375],[-18.2312507629395,-18.3260746002197,-18.4000968933105,-18.4486331939697,-18.4847660064697,-18.4507808685303,-18.3628902435303,-18.2923812866211,-18.2517585754395,-18.2551765441895,-18.2414073944092,-18.2258796691895,-18.2312507629395],[-17.1145496368408,-17.1316413879395,-17.0906257629395,-17.0491218566895,-16.9904308319092,-17.0419921875,-17.0944328308105,-17.1145496368408],[-16.5730476379395,-16.6610355377197,-16.6486320495605,-16.6965827941895,-16.7194347381592,-16.7186527252197,-16.74169921875,-16.7138671875,-16.6258792877197,-16.5275382995605,-16.46728515625,-16.4384765625,-16.3952159881592,-16.4032211303711,-16.5149421691895,-16.5294914245605,-16.5730476379395],[-15.7781248092651,-15.8244142532349,-15.7757797241211,-15.7259759902954,-15.7117176055908,-15.6657218933105,-15.6524419784546,-15.5948247909546,-15.6628904342651,-15.7380857467651,-15.7781248092651],[-15.6282224655151,-15.6324214935303,-15.6273431777954,-15.5831060409546,-15.5431642532349,-15.5336904525757,-15.5441408157349,-15.6282224655151],[-15.6199216842651,-15.6273431777954,-15.5440435409546,-15.5025386810303,-15.5888671875,-15.6199216842651],[-15.4019536972046,-15.43017578125,-15.4214849472046,-15.3824224472046,-15.34033203125,-15.2924795150757,-15.2674808502197,-15.2703123092651,-15.3108406066895,-15.3565435409546,-15.4019536972046],[-14.5794916152954,-14.6416988372803,-14.5916986465454,-14.4919910430908,-14.4560546875,-14.4748048782349,-14.5526371002197,-14.5794916152954],[-13.8039064407349,-13.8454103469849,-13.8424806594849,-13.7509765625,-13.763671875,-13.78662109375,-13.8269529342651,-13.8965816497803,-13.9073247909546,-13.94580078125,-14.0726566314697,-14.115234375,-14.1578121185303,-14.197265625,-14.1790037155151,-14.1842765808105,-14.2459964752197,-14.2930660247803,-14.2734375,-14.2804689407349,-14.2345705032349,-14.2289066314697,-14.2118158340454,-14.1754884719849,-14.12646484375,-14.0111322402954,-13.8648433685303,-13.7937498092651,-13.7210931777954,-13.6758794784546,-13.681640625,-13.7261714935303,-13.8039064407349],[-13.8245124816895,-13.8359375,-13.8166017532349,-13.7805662155151,-13.7531242370605,-13.7367181777954,-13.6647462844849,-13.7066411972046,-13.7911128997803,-13.8245124816895],[-11.6023445129395,-11.6767578125,-11.5764656066895,-11.4871091842651,-11.4659175872803,-11.5092763900757,-11.5832033157349,-11.6023445129395],[-11.6792964935303,-11.7031240463257,-11.7371091842651,-11.7732419967651,-11.8166007995605,-11.8356447219849,-11.7717771530151,-11.8245115280151,-11.8254880905151,-11.7873048782349,-11.6807613372803,-11.6970701217651,-11.658203125,-11.5412101745605,-11.4775390625,-11.3605470657349,-11.3368158340454,-11.3370113372803,-11.4201173782349,-11.5098638534546,-11.5921878814697,-11.6792964935303],[-11.3760747909546,-11.3843746185303,-11.3092775344849,-11.33984375,-11.3343744277954,-11.2630863189697,-11.2425775527954,-11.1898441314697,-11.2468748092651,-11.3131837844849,-11.3825197219849,-11.4152336120605,-11.4369144439697,-11.5095701217651,-11.587890625,-11.58251953125,-11.7109375,-11.9264650344849,-11.7423830032349,-11.6178712844849,-11.44580078125,-11.3049812316895,-11.2149410247803,-11.1921873092651,-11.1804685592651,-11.18310546875,-11.2794923782349,-11.3059577941895,-11.3760747909546],[-11.37890625,-11.4388675689697,-11.3938474655151,-11.35791015625,-11.2116203308105,-11.1776361465454,-11.1583976745605,-11.1047849655151,-11.0246095657349,-11.0124998092651,-11.1946287155151,-11.37890625],[-11.3028316497803,-11.318359375,-11.1636714935303,-11.1160154342651,-11.0373048782349,-11.0284175872803,-10.9688472747803,-10.9976568222046,-11.1064453125,-11.1691408157349,-11.3028316497803],[-11.9544925689697,-12.0758790969849,-12.1696290969849,-12.2259769439697,-12.3035163879395,-12.3612298965454,-12.3976564407349,-12.4988279342651,-12.6399412155151,-12.7361326217651,-12.8557615280151,-13.0945310592651,-13.3038082122803,-13.4436521530151,-13.7410154342651,-13.86279296875,-13.9636716842651,-14.16455078125,-14.3488273620605,-14.4010744094849,-14.4628915786743,-14.39453125,-14.3019533157349,-14.2793951034546,-14.2318363189697,-14.3546876907349,-14.4924793243408,-14.67431640625,-14.791015625,-14.85693359375,-14.9431648254395,-15.0293941497803,-15.0974607467651,-15.2039051055908,-15.3272466659546,-15.4766607284546,-15.7015619277954,-15.8810539245605,-16.0564460754395,-16.2369136810303,-16.3049793243408,-16.4061527252197,-16.5321273803711,-16.6250972747803,-16.72607421875,-16.8794918060303,-16.9103507995605,-16.9125003814697,-17.0702152252197,-17.3810539245605,-17.63525390625,-17.9773426055908,-18.1757831573486,-18.2728519439697,-18.5098648071289,-18.5537109375,-18.6666984558105,-18.8412113189697,-18.97705078125,-19.0787105560303,-19.1394538879395,-19.1874027252197,-19.2357425689697,-19.2560558319092,-19.3326168060303,-19.3931636810303,-19.4141597747803,-19.4029293060303,-19.3781242370605,-19.4193363189697,-19.47412109375,-19.6227550506592,-19.7701168060303,-19.7947273254395,-19.8692398071289,-19.8895492553711,-19.8986320495605,-19.9558601379395,-20.0874996185303,-20.10888671875,-20.1452140808105,-20.28955078125,-20.3664073944092,-20.4808597564697,-20.49169921875,-20.4677734375,-20.5801753997803,-20.7356452941895,-20.84521484375,-20.9611339569092,-21.1250972747803,-21.2501964569092,-21.2995109558105,-21.47607421875,-21.5787105560303,-21.7654285430908,-22.0236320495605,-22.2576179504395,-22.3283195495605,-22.4405269622803,-22.42626953125,-22.3898429870605,-22.5013675689697,-22.5506839752197,-22.5215816497803,-22.30810546875,-22.1842765808105,-22.1683597564697,-22.1644515991211,-22.2654304504395,-22.3729496002197,-22.4689445495605,-22.55908203125,-22.5557613372803,-22.4861316680908,-22.3839855194092,-22.3672847747803,-22.4181652069092,-22.576171875,-22.9029293060303,-23.1765613555908,-23.4580078125,-23.5319328308105,-23.6017570495605,-23.6960945129395,-23.7840805053711,-23.8249988555908,-24.0124034881592,-24.0335941314697,-24.0383796691895,-24.1229496002197,-24.2009754180908,-24.4944324493408,-24.5975589752197,-24.6993160247803,-24.7325191497803,-24.8024425506592,-24.9040031433105,-24.9639644622803,-25.0720710754395,-25.2019538879395,-25.2741203308105,-25.43212890625,-25.6885738372803,-25.8162117004395,-25.8703136444092,-25.9226570129395,-25.9641590118408,-26.3038082122803,-26.9827156066895,-27.1944332122803,-27.4046878814697,-27.7685546875,-27.8976573944092,-28.04833984375,-28.2405261993408,-28.5333995819092,-28.6730461120605,-28.8544921875,-29.0501937866211,-29.2904300689697,-29.49658203125,-29.8924808502197,-29.9986305236816,-30.1638679504395,-30.5633792877197,-30.7201175689697,-30.9071292877197,-31.0866222381592,-31.2087879180908,-31.4348621368408,-31.7863273620605,-32.0457038879395,-32.2430648803711,-32.3301734924316,-32.4390602111816,-32.5575218200684,-32.6086921691895,-32.6781234741211,-32.6781234741211,-32.6999015808105,-32.7216796875,-32.7574234008789,-32.8203125,-32.9010734558105,-33.0986328125,-33.2018585205078,-33.3009757995605,-33.3474578857422,-33.3973655700684,-33.5215797424316,-33.5439453125,-33.5809555053711,-33.6993179321289,-33.8348655700684,-33.9266586303711,-33.9850578308105,-33.9640617370605,-33.9734344482422,-34.0052719116211,-34.0152320861816,-34.0296859741211,-34.1624984741211,-34.2970695495605,-34.3866233825684,-34.4991226196289,-34.7492179870605,-34.8921890258789,-34.9938507080078,-35.0128936767578,-35.0204086303711,-35.0071258544922,-35.0418968200684,-35.07666015625,-35.1197280883789,-35.1345710754395,-35.1551742553711,-35.177734375,-35.1776351928711,-35.2142562866211,-35.5841789245605,-35.6823272705078,-35.8335952758789,-35.9706039428711,-36.1204109191895,-36.3726577758789,-36.5503921508789,-36.7227554321289,-36.8455085754395,-37.0802726745605,-37.2583999633789,-37.35302734375,-37.44384765625,-37.5285148620605,-37.5478515625,-37.6169929504395,-37.72998046875,-37.7711906433105,-37.8021507263184,-37.7884750366211,-37.8306655883789,-37.8560523986816,-37.9341812133789,-38.0556640625,-38.2191390991211,-38.6634788513184,-38.7118186950684,-38.7118186950684,-38.6998023986816,-38.7274436950684,-38.78271484375,-38.8402328491211,-38.8942375183105,-38.8196296691895,-38.84033203125,-38.9779281616211,-39.0650367736816,-39.1123046875,-39.1455078125,-39.1238288879395,-39.07666015625,-38.9644546508789,-38.86572265625,-38.8340835571289,-38.8670883178711,-38.9017601013184,-38.7759780883789,-38.6669921875,-38.6556625366211,-38.6568336486816,-38.6096687316895,-38.5353507995605,-38.4773445129395,-38.4163093566895,-38.3938484191895,-38.3114242553711,-38.2437477111816,-38.2256813049316,-38.2375984191895,-38.2912101745605,-38.3835945129395,-38.5007820129395,-38.4363288879395,-38.34033203125,-38.3473625183105,-38.3440437316895,-38.2583999633789,-38.2048835754395,-38.09130859375,-38.0109367370605,-37.9522476196289,-37.8998031616211,-38.0771484375,-38.1025390625,-38.1369132995605,-38.1664085388184,-38.1576156616211,-38.2099609375,-38.2840805053711,-38.3037109375,-38.3482437133789,-38.4623031616211,-38.6988258361816,-38.7668952941895,-38.8208999633789,-38.7578125,-38.7431640625,-38.6458969116211,-38.5808601379395,-38.45166015625,-38.3863296508789,-38.3721656799316,-38.3994140625,-38.2837867736816,-38.2713890075684,-38.3877944946289,-38.3797874450684,-38.3634757995605,-38.1719741821289,-38.0769538879395,-38.0284156799316,-37.8966789245605,-37.6421852111816,-37.35205078125,-37.2458000183105,-37.1416969299316,-37.0595703125,-36.9026374816895,-36.748046875,-36.662109375,-36.3713874816895,-36.0966796875,-36.0103530883789,-35.8273429870605,-35.6892585754395,-35.6175804138184,-35.5807609558105,-35.5984382629395,-35.5422859191895,-35.5230445861816,-35.59765625,-35.611328125,-35.4859352111816,-35.4266624450684,-35.3994140625,-35.3753890991211,-35.3472671508789,-35.3895492553711,-35.4432601928711,-35.4888687133789,-35.5368156433105,-35.5383796691895,-35.55078125,-35.6423835754395,-35.6447257995605,-35.6126937866211,-35.4865226745605,-35.4117202758789,-35.3257789611816,-35.0244140625,-34.7635726928711,-34.65625,-34.4403305053711,-34.3072280883789,-34.1698226928711,-34.2498054504395,-34.3340835571289,-34.4560546875,-34.7274398803711,-35.1429710388184,-35.1480484008789,-35.13134765625,-35.1787109375,-35.2364273071289,-35.2365226745605,-35.2548828125,-35.23974609375,-34.9158210754395,-34.9247055053711,-34.9115257263184,-34.9169921875,-34.9132804870605,-34.7644538879395,-34.5977516174316,-34.490234375,-34.37890625,-34.2521476745605,-34.1611328125,-33.8590812683105,-33.703125,-33.5791015625,-33.4613304138184,-33.3140640258789,-33.2007827758789,-33.1651382446289,-33.09423828125,-32.7707061767578,-32.6737289428711,-32.578125,-32.7019500732422,-32.8232421875,-32.97802734375,-33.0891571044922,-33.1935539245605,-33.43017578125,-33.6294898986816,-33.7030296325684,-33.7195320129395,-33.7501945495605,-33.8296890258789,-33.8965835571289,-33.9841804504395,-34.0299797058105,-34.4287109375,-34.5619163513184,-34.61572265625,-34.6609382629395,-34.7238273620605,-34.7667961120605,-34.9437484741211,-34.9818344116211,-34.9619140625,-34.86328125,-34.8992156982422,-34.9396476745605,-34.7582054138184,-34.7155265808105,-34.6426734924316,-34.5797843933105,-34.5726547241211,-34.5857429504395,-34.5365219116211,-34.49658203125,-34.4873046875,-34.5456047058105,-34.59765625,-34.6019515991211,-34.5810546875,-34.3755874633789,-34.1955108642578,-34.1422843933105,-33.9597663879395,-33.90673828125,-33.7777328491211,-33.6263656616211,-33.4446296691895,-33.3283195495605,-33.2551765441895,-33.1901359558105,-33.1650390625,-32.9790992736816,-32.7486343383789,-32.7333984375,-32.7305679321289,-32.65869140625,-32.5485343933105,-32.4117202758789,-32.2688446044922,-32.2072257995605,-32.1829071044922,-32.1884765625,-32.1837882995605,-31.9562492370605,-31.9493160247803,-32.02001953125,-32.0071258544922,-31.6962890625,-31.5485343933105,-31.52099609375,-31.4957027435303,-31.5318355560303,-31.5658226013184,-31.6040019989014,-31.5791034698486,-31.6272449493408,-31.6599617004395,-31.70263671875,-31.8876953125,-32.0665054321289,-32.1512680053711,-32.2640609741211,-32.296875,-32.3109359741211,-32.2568359375,-32.2969741821289,-32.505859375,-32.5565452575684,-32.6144523620605,-32.8827171325684,-32.9401359558105,-32.9583969116211,-33.0152359008789,-33.12939453125,-33.4462890625,-33.5963859558105,-33.8363265991211,-33.9162101745605,-33.9053688049316,-33.98828125,-33.9005851745605,-33.8837890625,-33.8908233642578,-33.9917945861816,-33.8744163513184,-33.8567390441895,-33.8624992370605,-33.8267593383789,-33.8712921142578,-33.9197273254395,-33.9630851745605,-33.9354515075684,-33.9747047424316,-34.04150390625,-34.1011695861816,-34.3682632446289,-34.4564476013184,-34.4593734741211,-34.4798851013184,-34.7371101379395,-34.9866180419922,-35.0132827758789,-35.0549812316895,-35.0749015808105,-35.0977516174316,-35.03369140625,-35.0265617370605,-34.9878883361816,-34.8658218383789,-34.7950172424316,-34.5260734558105,-34.42578125,-34.3039093017578,-34.3084983825684,-34.341796875,-34.255859375,-34.1451148986816,-34.0511703491211,-33.80419921875,-33.5153312683105,-33.5802726745605,-33.6434555053711,-33.6399421691895,-33.5313491821289,-33.3722648620605,-33.19287109375,-33.0021476745605,-32.6669921875,-32.5965843200684,-32.5679664611816,-32.4010734558105,-31.8878898620605,-31.6945323944092,-31.3025379180908,-30.9618186950684,-30.8080081939697,-30.5604476928711,-30.2162113189697,-30.0422859191895,-29.7215824127197,-29.5397453308105,-29.43359375,-29.1429691314697,-28.8717765808105,-28.7716808319092,-28.6662120819092,-28.5428695678711,-28.294921875,-28.0806636810303,-27.9764652252197,-27.5442390441895,-27.3472671508789,-26.8477535247803,-26.4173812866211,-26.2414054870605,-26.1822280883789,-26.1742191314697,-26.197265625,-26.240234375,-26.2438468933105,-26.2083015441895,-26.1260738372803,-26.0804691314697,-26.1055679321289,-26.1980476379395,-26.4367179870605,-26.55810546875,-26.5951175689697,-26.5632820129395,-26.5005855560303,-26.3321304321289,-26.2559566497803,-26.2236328125,-26.0986328125,-25.8983383178711,-25.7132816314697,-25.6471672058105,-25.59912109375,-25.6251945495605,-25.7316398620605,-25.8307609558105,-26.0041999816895,-26.0516605377197,-26.0916996002197,-26.1297855377197,-26.1597671508789,-26.1158199310303,-26.0144519805908,-26.0276355743408,-26.2586917877197,-26.3214855194092,-26.3936519622803,-26.3374996185303,-26.2894535064697,-26.1263675689697,-25.96875,-25.8515625,-25.5448246002197,-25.1657218933105,-24.97705078125,-24.6929702758789,-24.5946292877197,-24.4356441497803,-24.2540035247803,-24.13232421875,-23.86962890625,-23.7328128814697,-23.4181632995605,-23.2825202941895,-23.1804676055908,-23.0236339569092,-22.91455078125,-22.8128890991211,-22.6377925872803,-22.3321285247803,-21.9391593933105,-21.8814449310303,-21.82861328125,-21.9097671508789,-22.1813488006592,-22.3233394622803,-22.4831066131592,-22.4558582305908,-22.4253902435303,-22.3415050506592,-22.2610340118408,-21.9421882629395,-21.8234386444092,-21.7359390258789,-21.6305675506592,-21.49169921875,-21.3581066131592,-21.2422866821289,-21.11669921875,-21.0303707122803,-20.71337890625,-20.65380859375,-20.6470718383789,-20.6576175689697,-20.6409187316895,-20.7130870819092,-20.72119140625,-20.6427726745605,-20.5725593566895,-20.4190425872803,-20.3751945495605,-20.32666015625,-20.2619132995605,-19.9953117370605,-20.0123043060303,-20.0382804870605,-19.9583988189697,-19.9094715118408,-19.8419914245605,-19.6650390625,-19.6043949127197,-19.4779281616211,-19.3199234008789,-19.1064453125,-18.9151363372803,-18.8166007995605,-18.6599597930908,-18.5359382629395,-18.47705078125,-18.3936519622803,-18.1590824127197,-18.1119136810303,-18.0369148254395,-17.9949226379395,-17.9685535430908,-17.7203121185303,-17.5490226745605,-17.4284172058105,-17.3136730194092,-17.1357421875,-17.0593738555908,-16.9704093933105,-16.94287109375,-16.8649406433105,-16.7876949310303,-16.7101573944092,-16.5524406433105,-16.4326171875,-16.4368152618408,-16.71533203125,-16.8630867004395,-17.0368156433105,-17.2927742004395,-17.4099617004395,-17.4857425689697,-17.5208988189697,-17.4722652435303,-17.4154300689697,-17.2199211120605,-17.0827140808105,-17.0303707122803,-17.00830078125,-17.0232429504395,-17.0998039245605,-17.1271476745605,-17.1208000183105,-16.9968738555908,-16.9186515808105,-16.86474609375,-16.8677730560303,-16.8009757995605,-16.7236328125,-16.6680660247803,-16.5407218933105,-16.49072265625,-16.4675769805908,-16.4708995819092,-16.4163093566895,-16.3430671691895,-16.2240238189697,-16.1798839569092,-16.1924800872803,-16.38232421875,-16.3635749816895,-16.2869148254395,-16.2649402618408,-16.27880859375,-16.3335933685303,-16.3882808685303,-16.3820304870605,-16.3952159881592,-16.3861331939697,-16.4026355743408,-16.3733406066895,-16.3387680053711,-16.3318367004395,-16.3352527618408,-16.2989273071289,-16.2030277252197,-16.1334953308105,-16.1038093566895,-16.1163101196289,-16.1136703491211,-16.0201168060303,-15.9375,-15.8702144622803,-15.8054695129395,-15.8226556777954,-15.9724607467651,-15.8505859375,-15.7582025527954,-15.6258792877197,-15.4935550689697,-15.4753904342651,-15.4962892532349,-15.4188480377197,-15.3596677780151,-15.31103515625,-15.2736330032349,-15.2852544784546,-15.4042959213257,-15.4665040969849,-15.4422855377197,-15.37451171875,-15.3067388534546,-15.3169927597046,-15.31005859375,-15.2719736099243,-15.2405271530151,-15.1607418060303,-15.1099615097046,-15.1066398620605,-15.0718746185303,-15.0244131088257,-15.0041017532349,-15.0323238372803,-15.0454092025757,-15.1068363189697,-15.1198244094849,-15.0868167877197,-15.0156240463257,-14.9445304870605,-14.8746099472046,-14.7940425872803,-14.7147464752197,-14.6484365463257,-14.5840826034546,-14.5579099655151,-14.5568351745605,-14.5022449493408,-14.4832029342651,-14.3616199493408,-14.2781248092651,-14.2566413879395,-14.2914056777954,-14.3879880905151,-14.4801750183105,-14.5294923782349,-14.525390625,-14.5048837661743,-14.4443359375,-14.4691410064697,-14.5338859558105,-14.5972652435303,-14.6179695129395,-14.5204105377197,-14.4945306777954,-14.3712882995605,-14.2830076217651,-14.2166986465454,-14.1843748092651,-14.1140623092651,-14.0655269622803,-13.9772462844849,-13.9577150344849,-14.0020503997803,-14.1133785247803,-14.16357421875,-14.13623046875,-14.0621099472046,-14.0189456939697,-14.0789070129395,-14.1609373092651,-14.08935546875,-13.9551763534546,-13.8730478286743,-13.7884759902954,-13.7441415786743,-13.7767581939697,-13.8673830032349,-13.9347658157349,-14.0314455032349,-14.0946292877197,-14.1951169967651,-14.2994146347046,-14.4851560592651,-14.7116212844849,-14.7517576217651,-14.8273439407349,-14.9241209030151,-15.0879888534546,-15.3292970657349,-15.31201171875,-15.2255859375,-15.2433605194092,-15.2985363006592,-15.24560546875,-15.2135734558105,-15.1022462844849,-15.0431642532349,-14.9957027435303,-14.9388666152954,-14.9016590118408,-14.869140625,-14.8289060592651,-14.7879877090454,-14.7745122909546,-14.7809581756592,-14.8843755722046,-14.9875984191895,-15.1150398254395,-15.1602535247803,-15.0801763534546,-14.9060544967651,-14.871485710144,-14.8984375,-14.9332027435303,-15.0473623275757,-15.1033210754395,-15.1397457122803,-15.0868167877197,-15.0118169784546,-14.9258794784546,-14.8509759902954,-14.8450202941895,-14.8289060592651,-14.7997064590454,-14.78955078125,-14.7208986282349,-14.6470708847046,-14.5752925872803,-14.5574216842651,-14.5596685409546,-14.48974609375,-14.3924808502197,-14.2134761810303,-14.0383787155151,-13.97998046875,-13.9208984375,-13.8119144439697,-13.7199220657349,-13.6484375,-13.5729494094849,-13.5016603469849,-13.4761714935303,-13.4483404159546,-13.3826169967651,-13.30224609375,-13.1455078125,-13.0591793060303,-12.9574213027954,-12.8829097747803,-12.6878910064697,-12.6585931777954,-12.6643562316895,-12.6468744277954,-12.4913082122803,-12.4310541152954,-12.4069337844849,-12.427734375,-12.4953117370605,-12.5578126907349,-12.5236330032349,-12.455078125,-12.3671875,-12.3482418060303,-12.3428707122803,-12.2710943222046,-12.2138671875,-12.1896486282349,-12.1779289245605,-12.1190423965454,-12.06787109375,-12.0958976745605,-12.2100582122803,-12.2769527435303,-12.2781248092651,-12.23193359375,-12.25927734375,-12.28076171875,-12.2269525527954,-12.18603515625,-12.2391605377197,-12.2951173782349,-12.1763668060303,-12.1348628997803,-12.1102542877197,-12.1300783157349,-12.1234369277954,-12.03515625,-11.9546880722046,-11.8358402252197,-11.7271480560303,-11.6490230560303,-11.4915037155151,-11.4676761627197,-11.5006837844849,-11.4747066497803,-11.3485355377197,-11.3024415969849,-11.2713861465454,-11.1808595657349,-11.1963863372803,-11.2811517715454,-11.3111333847046,-11.3049812316895,-11.23876953125,-11.2040042877197,-11.2235345840454,-11.3668947219849,-11.5055665969849,-11.4689455032349,-11.39111328125,-11.4073247909546,-11.4528322219849,-11.6217775344849,-11.7056646347046,-11.7282228469849,-11.7603521347046,-11.8162107467651,-11.8113269805908,-11.83203125,-11.9401369094849,-12.0077152252197,-12.0257816314697,-12.052734375,-12.0608406066895,-11.984375,-12.0546875,-12.1025381088257,-12.1937503814697,-12.2216796875,-12.1291990280151,-12.0606451034546,-11.9561519622803,-11.9070310592651,-11.8216791152954,-11.8257808685303,-11.9054689407349,-11.95068359375,-11.9695310592651,-11.9927730560303,-12.0547857284546,-12.1515626907349,-12.2098636627197,-12.24169921875,-12.2275400161743,-12.1785154342651,-12.1521492004395,-12.19140625,-12.3308601379395,-12.4224605560303,-12.4351558685303,-12.4337892532349,-12.3055667877197,-12.1963872909546,-12.1730470657349,-12.1316404342651,-11.9514646530151,-11.9576177597046,-12.1335935592651,-12.2264642715454,-12.2191410064697,-12.2435550689697,-12.3499021530151,-12.7842769622803,-12.8328123092651,-12.91162109375,-13.0038080215454,-13.2251949310303,-13.2361326217651,-13.1763668060303,-13.1379880905151,-13.1649408340454,-13.1810541152954,-13.3042974472046,-13.62158203125,-13.8101568222046,-13.9348630905151,-14.1531248092651,-14.2341794967651,-14.28662109375,-14.5849609375,-14.656641960144,-14.7582035064697,-14.8556652069092,-14.9231452941895,-15.0003910064697,-15.16015625,-15.2702150344849,-15.4034185409546,-15.4952144622803,-15.5701179504395,-15.6552734375,-15.70654296875,-15.6933584213257,-15.6755867004395,-15.6753902435303,-15.6852531433105,-15.751953125,-15.7884759902954,-15.8349618911743,-15.8942375183105,-15.8923835754395,-15.8783197402954,-15.9413080215454,-15.9821290969849,-16.0663089752197,-16.1670894622803,-16.2330074310303,-16.4765625,-16.6169929504395,-16.7183589935303,-16.78955078125,-16.77783203125,-16.8606452941895,-16.8993148803711,-17.0140628814697,-17.10107421875,-17.1677722930908,-17.32861328125,-17.3805675506592,-17.5407238006592,-17.611328125,-17.6536140441895,-17.70263671875,-17.7043952941895,-17.62451171875,-17.5437507629395,-17.4144535064697,-17.1925773620605,-17.0145511627197,-16.6461925506592,-16.4634761810303,-16.2210922241211,-16.0695304870605,-15.9046878814697,-15.6052732467651,-15.1954107284546,-15.0566396713257,-14.8527345657349,-14.4701175689697,-14.337890625,-14.15283203125,-14.0186529159546,-13.9267578125,-13.7975587844849,-13.5538082122803,-13.4250984191895,-13.2590827941895,-12.9434566497803,-12.83349609375,-12.7787113189697,-12.7782230377197,-12.8029298782349,-12.7398443222046,-12.6813478469849,-12.61328125,-12.5787115097046,-12.5666007995605,-12.5293951034546,-12.4914064407349,-12.35107421875,-12.080078125,-11.9755859375,-12.0192384719849,-12.0542964935303,-11.9762697219849,-11.8961915969849,-11.6317377090454,-11.2732419967651,-10.9465818405151,-10.8841800689697,-10.80224609375,-10.7073240280151,-10.7073240280151,-10.7482414245605,-10.8194341659546,-10.8744144439697,-11.0104494094849,-11.1153316497803,-11.2139654159546,-11.3069334030151,-11.4322261810303,-11.63232421875,-11.8213863372803,-11.8807611465454,-11.9190425872803,-11.9241209030151,-11.9544925689697],[-10.7047853469849,-10.7620115280151,-10.73193359375,-10.66845703125,-10.640625,-10.5919923782349,-10.7047853469849],[-10.1921873092651,-10.2541999816895,-10.2356443405151,-10.1993169784546,-10.1494131088257,-10.1404304504395,-10.1921873092651],[-10.1541013717651,-10.1812496185303,-10.1217775344849,-10.0517578125,-10.0852537155151,-10.1541013717651],[48.5988273620605,48.5885772705078,48.5509262084961,48.5035171508789,48.44140625,48.3869132995605,48.1980934143066,48.083251953125,48.03955078125,48.0059585571289,47.963623046875,47.90087890625,47.8729476928711,47.8371086120605,47.8045425415039,47.7637672424316,47.7075691223145,47.6953125,47.697265625,47.6939964294434,47.6786613464355,47.686279296875,47.739013671875,47.7505340576172,47.7473640441895,47.7244606018066,47.695068359375,47.6744651794434,47.6562995910645,47.60888671875,47.5360374450684,47.4766120910645,47.4475593566895,47.4246559143066,47.404541015625,47.3995132446289,47.367431640625,47.2731437683105,47.2527351379395,47.2234344482422,47.1459007263184,47.140380859375,47.1226577758789,47.0912590026855,47.057861328125,47.0224609375,47.0067863464355,46.9969711303711,47.002197265625,46.971923828125,46.86328125,46.8448257446289,46.8013687133789,46.7058563232422,46.6972160339355,46.6776351928711,46.7112770080566,46.7107429504395,46.6984367370605,46.6546401977539,46.629638671875,46.6429672241211,46.6259765625,46.6132354736328,46.6059074401855,46.5804710388184,46.5445785522461,46.4991226196289,46.4634284973145,46.4360809326172,46.4129371643066,46.3997077941895,46.4170417785645,46.4161148071289,46.4279327392578,46.4407234191895,46.4619140625,46.482177734375,46.4981918334961,46.51123046875,46.5143051147461,46.520263671875,46.5555648803711,46.5579109191895,46.5726547241211,46.6258811950684,46.6474609375,46.6541023254395,46.6725082397461,46.70263671875,46.7598114013672,46.8358879089355,46.9352531433105,46.9847640991211,47.0281753540039,47.0608863830566,47.0750007629395,47.0821304321289,47.0396957397461,46.986083984375,46.9846649169922,46.99658203125,46.9974098205566,46.9830589294434,46.9756851196289,46.9361839294434,46.859130859375,46.7969741821289,46.7770004272461,46.7694816589355,46.7752418518066,46.7933120727539,46.8463859558105,46.8537101745605,46.8551292419434,46.8649406433105,46.8994140625,46.9644050598145,46.9847640991211,46.8623542785645,46.8515129089355,46.8853530883789,46.9376945495605,46.9759750366211,47.0073738098145,47.037109375,47.0574684143066,47.057373046875,47.0758285522461,47.1071281433105,47.132080078125,47.1579093933105,47.1854972839355,47.2122535705566,47.234130859375,47.254638671875,47.270751953125,47.3917961120605,47.467041015625,47.5111351013184,47.5242156982422,47.5340309143066,47.52587890625,47.55078125,47.5755348205566,47.5522956848145,47.5053215026855,47.4735832214355,47.4490699768066,47.4285163879395,47.3933563232422,47.3795928955078,47.374267578125,47.3634262084961,47.3171882629395,47.27880859375,47.2841339111328,47.3134269714355,47.3660659790039,47.4169921875,47.5410652160645,47.5515632629395,47.5417976379395,47.5472183227539,47.5241203308105,47.5202140808105,47.5007820129395,47.470458984375,47.4267082214355,47.3981437683105,47.3931159973145,47.4088859558105,47.4251976013184,47.4136238098145,47.4249038696289,47.4602508544922,47.4871559143066,47.506103515625,47.5497550964355,47.58349609375,47.6195335388184,47.646728515625,47.7090835571289,47.71826171875,47.7027359008789,47.6881828308105,47.6661148071289,47.6373062133789,47.6361351013184,47.6562995910645,47.6693344116211,47.639404296875,47.6070289611816,47.5904273986816,47.5641593933105,47.5421867370605,47.5064468383789,47.4756813049316,47.4780769348145,47.5080108642578,47.5791473388672,47.6551284790039,47.69873046875,47.7094230651855,47.7128448486328,47.7218742370605,47.7458038330078,47.8077621459961,47.890625,47.9848136901855,48.0759773254395,48.1069831848145,48.1608390808105,48.2037124633789,48.2750968933105,48.2899436950684,48.3019027709961,48.3312492370605,48.3613777160645,48.3941421508789,48.5645523071289,48.5718231201172,48.5818328857422,48.5230484008789,48.5327644348145,48.5423851013184,48.5874519348145,48.6216773986816,48.6864242553711,48.7475090026855,48.7669410705566,48.7598648071289,48.72802734375,48.6924324035645,48.6024894714355,48.5785675048828,48.5762214660645,48.6162567138672,48.6255378723145,48.6133270263672,48.5992164611816,48.6719245910645,48.7473640441895,48.7740211486816,48.7713851928711,48.8277359008789,48.9839363098145,49.0011215209961,48.9978523254395,48.9693336486816,48.9462890625,48.9481430053711,48.9638671875,48.9740257263184,48.9573707580566,48.8863754272461,48.8604469299316,48.8654289245605,48.8644523620605,48.7547836303711,48.7394027709961,48.7389678955078,48.7720680236816,48.8000984191895,48.7962417602539,48.7818832397461,48.7342300415039,48.7220191955566,48.7143058776855,48.7037124633789,48.6208992004395,48.5988273620605],[38.8777809143066,38.9079093933105,38.9728050231934,39.0062522888184,39.0958976745605,39.13916015625,39.1946792602539,39.2156219482422,39.2542953491211,39.3115730285645,39.3628883361816,39.4235343933105,39.6291007995605,39.6504402160645,39.6846694946289,39.7035140991211,39.7191429138184,39.76513671875,39.7428207397461,39.6963386535645,39.6565895080566,39.5826683044434,39.5706062316895,39.595458984375,39.5655746459961,39.5298843383789,39.4944839477539,39.5498046875,39.5640602111816,39.5629386901855,39.5456085205078,39.4881324768066,39.417236328125,39.3784637451172,39.3501968383789,39.2819328308105,39.243896484375,39.1781272888184,39.0175323486328,38.9548797607422,38.8777809143066],[41.0272445678711,41.0526351928711,41.0792465209961,41.0855941772461,41.0779800415039,41.0538558959961,41.0272445678711,41.0158195495605,41.0272445678711],[41.844482421875,41.78662109375,41.6295890808105,41.3739738464355,41.3017082214355,41.2177734375,41.1161117553711,41.0262222290039,40.7998542785645,40.71630859375,40.6081047058105,40.583984375,40.5771980285645,40.5768089294434,40.5345230102539,40.5047836303711,40.4617691040039,40.4122085571289,40.2794952392578,40.3232421875,40.31640625,40.2878913879395,40.2490234375,40.1941413879395,40.0872535705566,39.83984375,39.6083488464355,39.501220703125,39.3983879089355,39.3495597839355,39.3289070129395,39.28515625,39.0726585388184,39.0302734375,39.00390625,39.029052734375,39.084716796875,39.1339836120605,39.0787582397461,38.9617691040039,38.8388175964355,38.8153343200684,38.4355010986328,38.437255859375,38.3987274169922,38.4110832214355,38.5862312316895,38.6056175231934,38.6134757995605,38.6422882080078,38.689208984375,38.72412109375,38.8190422058105,38.853759765625,38.88427734375,38.9118156433105,38.9350090026855,38.9586448669434,38.9789543151855,38.9936027526855,39.0188484191895,39.0592765808105,39.09912109375,39.171630859375,39.2028350830078,39.2411117553711,39.2811050415039,39.3123512268066,39.35498046875,39.3990707397461,39.4483375549316,39.560546875,39.6839370727539,39.68505859375,39.6485824584961,39.5433578491211,39.4983367919922,39.4238739013672,39.3409690856934,39.2528839111328,39.18017578125,39.1484375,39.08740234375,38.9043960571289,38.9066886901855,38.9974594116211,39.0694351196289,39.1108894348145,39.1676750183105,39.1921882629395,39.2073707580566,39.1981925964355,39.201416015625,39.2236824035645,39.2985343933105,39.3485336303711,39.3822746276855,39.4024887084961,39.4167976379395,39.4338836669922,39.47509765625,39.5128440856934,39.55517578125,39.617431640625,39.594482421875,39.6644515991211,39.7185554504395,39.7765617370605,39.808349609375,39.881103515625,39.9561996459961,39.9775390625,39.989013671875,40.0028343200684,40.0142097473145,40.0112800598145,40.0248527526855,40.0570793151855,40.1046867370605,40.1748046875,40.2337875366211,40.3291015625,40.4168434143066,40.5323753356934,40.6380844116211,40.673583984375,40.7071266174316,40.8044929504395,40.8297386169434,40.846923828125,40.896728515625,40.947998046875,40.9856948852539,41.0048828125,41.0062484741211,41.0693359375,41.0755844116211,41.088134765625,41.100830078125,41.1263694763184,41.1474113464355,41.1751480102539,41.1954612731934,41.2457008361816,41.2909660339355,41.4231910705566,41.4495582580566,41.42529296875,41.337646484375,41.2890129089355,41.2616195678711,41.2244110107422,41.1867218017578,41.1672859191895,41.183837890625,41.1978492736816,41.1544456481934,41.0993194580078,41.0770530700684,41.0702133178711,41.0885734558105,41.15966796875,41.2455062866211,41.2868156433105,41.34375,41.4055671691895,41.4598617553711,41.5077171325684,41.6021499633789,41.6125946044922,41.6248512268066,41.6570816040039,41.7021484375,41.7368659973145,41.7517585754395,41.757080078125,41.7901840209961,41.8550796508789,41.8909683227539,41.8704109191895,41.8123054504395,41.8000984191895,41.8069305419922,41.8313484191895,41.8125991821289,41.743408203125,41.67041015625,41.6213874816895,41.5874977111816,41.5546875,41.5160636901855,41.4556159973145,41.3150863647461,41.2824211120605,41.2290534973145,41.2181167602539,41.1992683410645,41.2127418518066,41.333984375,41.4586906433105,41.4847640991211,41.5450172424316,41.6019020080566,41.663330078125,41.7793464660645,41.844482421875],[40.6160659790039,40.649169921875,40.6640129089355,40.6648445129395,40.6483421325684,40.6069793701172,40.5995597839355,40.6160659790039],[-2.40009784698486,-2.42626953125,-2.61347651481628,-2.64160132408142,-2.65888667106628,-2.69433569908142,-2.75322246551514,-2.76904320716858,-2.82402372360229,-2.87451148033142,-2.89316415786743,-2.91757774353027,-2.93525409698486,-2.97724604606628,-2.98486304283142,-3.01513671875,-3.06933617591858,-3.11640644073486,-3.20058584213257,-3.27460956573486,-3.30937504768372,-3.34736323356628,-3.36640596389771,-3.388671875,-3.41865253448486,-3.49248027801514,-3.58886742591858,-3.65390610694885,-3.73076200485229,-3.77978539466858,-3.85048842430115,-3.99287128448486,-4.08535194396973,-4.30732440948486,-4.41806697845459,-4.45585966110229,-4.44931650161743,-4.29970693588257,-4.09540987014771,-3.91083955764771,-3.83378911018372,-3.68496084213257,-3.47568345069885,-3.36328125,-3.28125023841858,-3.138671875,-3.05351543426514,-2.95527315139771,-2.85078144073486,-2.79960942268372,-2.75830078125,-2.72021508216858,-2.66455054283142,-2.60253953933716,-2.59570336341858,-2.62031245231628,-2.67304682731628,-2.79150366783142,-2.80859375,-2.80839848518372,-2.79277348518372,-2.79472661018372,-2.76640629768372,-2.71640610694885,-2.66464853286743,-2.54863262176514,-2.33955049514771,-2.33710908889771,-2.41152310371399,-2.41660141944885,-2.41396450996399,-2.37705063819885,-2.34706997871399,-2.34785151481628,-2.31298828125,-2.37607431411743,-2.39560532569885,-2.40009784698486],[50.7504386901855,50.7322273254395,50.6792488098145,50.6372566223145,50.5966796875,50.5453605651855,50.5225105285645,50.4991188049316,50.4854965209961,50.4517593383789,50.4002418518066,50.316162109375,50.232666015625,50.1393547058105,50.1209983825684,50.123779296875,50.1545906066895,50.154296875,50.1671867370605,50.0828132629395,50.0126953125,49.9612312316895,49.9196281433105,49.8756370544434,49.857177734375,49.8333511352539,49.8083000183105,49.7588882446289,49.732177734375,49.644775390625,49.6128425598145,49.5783195495605,49.5538063049316,49.5382804870605,49.5392074584961,49.5282211303711,49.5110321044922,49.5108871459961,49.5544891357422,49.6198234558105,49.6509780883789,49.6779289245605,49.6892585754395,49.7214851379395,49.7565422058105,49.7783699035645,49.7892570495605,49.7881355285645,49.8471183776855,49.9145050048828,49.9595680236816,50.1358909606934,50.1531753540039,50.1390647888184,50.0970726013184,50.046875,50.00244140625,49.9602546691895,49.9449691772461,49.9602546691895,49.9715843200684,49.9844703674316,50,50.0238800048828,50.0528335571289,50.0941390991211,50.1298828125,50.143798828125,50.1784172058105,50.2217750549316,50.2464828491211,50.3213386535645,50.3359375,50.3385734558105,50.3469734191895,50.343505859375,50.3216781616211,50.3060531616211,50.3248062133789,50.4573249816895,50.4773445129395,50.4994659423828,50.5073738098145,50.5315437316895,50.5911636352539,50.6629371643066,50.731689453125,50.7489280700684,50.7794456481934,50.7668952941895,50.72705078125,50.7160148620605,50.7117691040039,50.7506332397461,50.8114280700684,50.8759269714355,50.9117698669434,50.9552726745605,50.9885749816895,51.0495109558105,51.0971183776855,51.2654304504395,51.3516120910645,51.377685546875,51.2911148071289,51.2636222839355,51.2457504272461,51.2422370910645,51.2636222839355,51.2861824035645,51.2756843566895,51.2548332214355,51.2332038879395,51.2125968933105,51.2076683044434,51.2470703125,51.3070793151855,51.3487319946289,51.386474609375,51.3615226745605,51.3560028076172,51.3670883178711,51.4275894165039,51.4598121643066,51.4747047424316,51.4485855102539,51.4219245910645,51.4217262268066,51.4911117553711,51.4773941040039,51.4527320861816,51.4328117370605,51.4120597839355,51.4032745361328,51.4077644348145,51.4453620910645,51.4690933227539,51.453125,51.4068336486816,51.3464851379395,51.2789573669434,51.2597160339355,51.2729988098145,51.2850608825684,51.2749977111816,51.2393074035645,51.1984405517578,51.1694831848145,51.153076171875,51.1256370544434,51.08642578125,50.98876953125,50.9599113464355,50.9502449035645,50.93212890625,50.8666496276855,50.8436050415039,50.8059539794922,50.7747535705566,50.7595710754395,50.7545433044434,50.752555847168,50.7504386901855],[11.6962890625,11.6318845748901,11.4992189407349,11.3954095840454,11.1768560409546,11.1545906066895,11.1203126907349,11.07958984375,10.971923828125,10.8504390716553,10.7687492370605,10.6537599563599,10.607421875,10.4358882904053,10.4176263809204,10.4126949310303,10.4277830123901,10.4089841842651,10.3506841659546,10.29248046875,10.2683591842651,10.16015625,10.0045413970947,9.90732383728027,9.85190391540527,9.83862400054932,9.81279373168945,9.77846717834473,9.66704082489014,9.56562519073486,9.49467754364014,9.45161151885986,9.32060527801514,9.18828105926514,9.08383846282959,9.06137752532959,9.04853439331055,8.78251934051514,8.614013671875,8.44189453125,8.371826171875,8.27299785614014,8.0498046875,7.87373018264771,7.82661151885986,7.72309541702271,7.61625957489014,7.54189395904541,7.47685623168945,7.44340801239014,7.42250919342041,7.39506816864014,7.14321231842041,7.06791973114014,7.01982450485229,6.98027324676514,6.85283231735229,6.77163076400757,6.71171855926514,6.66176795959473,6.595703125,6.42768573760986,6.36923837661743,6.32807588577271,6.26064443588257,6.216796875,6.25083017349243,6.29462909698486,6.42626905441284,6.58154249191284,6.61020517349243,6.68740224838257,6.73808574676514,6.77226543426514,6.87700223922729,6.99243116378784,6.997314453125,7.36918926239014,7.72587871551514,8.03022480010986,8.27099609375,8.55927753448486,8.77099609375,9.050048828125,9.13725662231445,9.28500938415527,9.36166954040527,9.46298789978027,9.56752967834473,9.75019454956055,9.96294021606445,9.99697303771973,10.098388671875,10.2420415878296,10.3515625,10.3595695495605,10.3866701126099,10.7102537155151,10.7525873184204,10.8857421875,10.9932613372803,10.9928226470947,11.0277833938599,11.0790042877197,11.068115234375,11.0582027435303,11.0763673782349,11.1160154342651,11.1563472747803,11.1743650436401,11.2104005813599,11.2519044876099,11.2627449035645,11.2610349655151,11.2739753723145,11.2952642440796,11.37890625,11.4080085754395,11.4287109375,11.4471197128296,11.45556640625,11.4491205215454,11.4006338119507,11.443359375,11.4184083938599,11.629150390625,11.6912593841553,11.840087890625,11.8970699310303,11.9993162155151,12.1884269714355,12.221923828125,12.2627925872803,12.2943363189697,12.2967767715454,12.3127927780151,12.3536138534546,12.3838386535645,12.3736820220947,12.3677253723145,12.1180658340454,11.9918947219849,11.9271488189697,11.8804683685303,11.8519525527954,11.7874507904053,11.6962890625],[14.9114742279053,14.8650388717651,14.7828121185303,14.6529293060303,14.4972162246704,14.3964357376099,14.2880373001099,14.2458009719849,14.1390132904053,14.0763664245605,13.9721193313599,13.8397455215454,13.7034177780151,13.6854009628296,13.6745119094849,13.6500492095947,13.6264162063599,13.6109371185303,13.5811519622803,13.5519533157349,13.4678707122803,13.412353515625,13.3575201034546,13.32958984375,13.3407716751099,13.3648433685303,13.3245115280151,13.1703615188599,13.0418949127197,13.0248041152954,13.0011224746704,12.8342771530151,12.6764650344849,12.6353998184204,12.6198234558105,12.61328125,12.6278810501099,12.7074222564697,12.7162103652954,12.7139654159546,12.7012691497803,12.6364259719849,12.5384273529053,12.466064453125,12.42724609375,12.41259765625,12.3938474655151,12.379150390625,12.3579587936401,12.3093748092651,12.2779788970947,12.136474609375,11.9459953308105,11.8970699310303,11.840087890625,11.6912593841553,11.629150390625,11.4184083938599,11.443359375,11.4006338119507,11.4491205215454,11.45556640625,11.4471197128296,11.4287109375,11.4080085754395,11.37890625,11.2952642440796,11.2739753723145,11.2610349655151,11.2627449035645,11.2519044876099,11.2104005813599,11.1743650436401,11.1563472747803,11.1160154342651,11.0763673782349,11.0582027435303,11.068115234375,11.0790042877197,11.0277833938599,10.9928226470947,10.9932613372803,10.9830570220947,10.9554204940796,10.9549808502197,10.9781742095947,10.9919929504395,11.0696287155151,11.1017780303955,11.1156244277954,11.1668949127197,11.118896484375,11.0879392623901,11.085693359375,11.09326171875,11.0562982559204,11.0076169967651,10.9836912155151,10.9536619186401,10.9273929595947,10.9267578125,10.9889650344849,10.9952630996704,10.9847173690796,11.001708984375,11.0100584030151,10.9972171783447,11.0226554870605,11.0088872909546,10.9976568222046,10.9946775436401,10.9914054870605,10.9887208938599,10.9863767623901,10.9969720840454,10.9983882904053,10.9774904251099,10.7279787063599,10.5923337936401,10.5079593658447,10.4546384811401,10.4324216842651,10.4019041061401,10.3629388809204,10.3228511810303,10.2815923690796,10.2381839752197,10.1925783157349,10.0831050872803,9.90966796875,9.79721641540527,9.745849609375,9.65805625915527,9.533935546875,9.48134803771973,9.45712947845459,9.42470741271973,9.42582988739014,9.50092792510986,9.53461933135986,9.61074256896973,9.68735313415527,9.72089862823486,9.75209999084473,9.84916973114014,9.89545917510986,9.88222694396973,9.90029239654541,9.92431735992432,9.91718673706055,9.89492225646973,9.85962009429932,9.78173828125,9.74326229095459,9.64570331573486,9.64799785614014,9.67924785614014,9.72348594665527,9.71357440948486,9.75654315948486,9.84116172790527,9.86894512176514,9.93007850646973,10.0464839935303,10.1283197402954,10.2416009902954,10.2926273345947,10.3196773529053,10.3140134811401,10.3595695495605,10.426025390625,10.4834470748901,10.5650873184204,10.6439456939697,10.7713871002197,10.9310550689697,11.0423831939697,11.0887203216553,11.1302738189697,11.2059574127197,11.3757820129395,11.5224609375,11.5767574310303,11.619873046875,11.6833009719849,11.7604494094849,11.8279294967651,11.8902826309204,11.9423828125,11.967529296875,11.9933109283447,12.0321292877197,12.076171875,12.1202154159546,12.155029296875,12.2264642715454,12.2817869186401,12.3375978469849,12.4930667877197,12.5815916061401,12.63037109375,12.6722173690796,12.793701171875,12.8794918060303,12.975341796875,13.052490234375,13.1190423965454,13.1973142623901,13.2561521530151,13.3062009811401,13.3824224472046,13.4021968841553,13.37353515625,13.1941900253296,13.1827154159546,13.1963863372803,13.2437009811401,13.28076171875,13.4007816314697,13.5774421691895,13.6583499908447,13.6728515625,13.6391115188599,13.6371097564697,13.6484375,13.6794929504395,13.7363767623901,13.7867679595947,13.9507322311401,14.07373046875,14.2275876998901,14.25830078125,14.2741212844849,14.16845703125,14.1946287155151,14.45654296875,14.4814929962158,14.4860353469849,14.5084962844849,14.5268068313599,14.6260747909546,14.7615242004395,14.8195314407349,14.8413572311401,14.9374017715454,15.0477533340454,15.0697755813599,15.0778818130493,15.0596675872803,15.0285167694092,15.0124998092651,15.0594234466553,14.987060546875,14.9848155975342,14.9114742279053],[21.5080547332764,21.5033206939697,21.532958984375,21.5744152069092,21.7087898254395,21.7229480743408,21.72216796875,21.6825695037842,21.6097660064697,21.5080547332764],[21.8321285247803,21.750244140625,21.809814453125,21.8853511810303,21.9270496368408,21.9266586303711,21.8836421966553,21.8321285247803],[22.1751956939697,22.1051769256592,22.5197257995605,22.3654308319092,22.2830066680908,22.1751956939697],[22.3822250366211,22.352783203125,22.37841796875,22.4756832122803,22.5988292694092,22.6165027618408,22.5765609741211,22.49072265625,22.4253902435303,22.3822250366211],[22.0893058776855,22.05419921875,22.0651359558105,22.3274898529053,22.4449710845947,22.5406742095947,22.6176261901855,22.6725578308105,22.7475090026855,22.8353500366211,22.863525390625,22.8131828308105,22.7853031158447,22.7135257720947,22.6387195587158,22.48486328125,22.3905754089355,22.1599617004395,22.0893058776855],[22.9629878997803,22.9200191497803,22.945556640625,22.9754390716553,23.014892578125,23.0354480743408,22.9629878997803],[21.9780769348145,21.9403324127197,21.6090335845947,21.4673347473145,21.3507328033447,21.3062019348145,21.2701644897461,21.2633304595947,21.31982421875,21.3629875183105,21.4090328216553,21.4397945404053,21.4275875091553,21.3578605651855,21.2931156158447,21.2022457122803,21.1126956939697,21.0614738464355,21.0046882629395,20.9315910339355,20.8644523620605,20.7918453216553,20.7904300689697,20.8835945129395,20.9842777252197,21.1748046875,21.5162601470947,21.6847667694092,21.883056640625,22.1573734283447,22.2286624908447,22.3173332214355,22.3504886627197,22.3431167602539,22.2974624633789,22.4066905975342,22.5047855377197,22.7076663970947,22.7974128723145,22.8848152160645,22.7970218658447,22.7351551055908,22.6422367095947,22.6140632629395,22.5970211029053,22.7213878631592,23.0254878997803,23.09423828125,23.2730464935303,23.4423351287842,23.5316410064697,23.5913581848145,23.578125,23.537109375,23.4742679595947,23.4215335845947,23.431884765625,23.4558582305908,23.366943359375,23.3461437225342,23.2664051055908,23.2041511535645,23.1339359283447,23.0849628448486,23.0539073944092,22.9867668151855,22.9048824310303,22.8817882537842,22.8281745910645,22.7519054412842,22.6846675872803,22.6348152160645,22.5887222290039,22.539306640625,22.4358386993408,22.3621597290039,22.2584476470947,22.2181644439697,22.1789073944092,22.0482425689697,21.8994140625,21.8297843933105,21.8168449401855,21.847412109375,21.8872566223145,21.9599113464355,22.0174808502197,22.1565933227539,22.1378917694092,22.0981922149658,22.0228519439697,22.1161632537842,22.2025890350342,22.3083992004395,22.4664058685303,22.3875980377197,22.2889652252197,22.1730461120605,22.0909175872803,21.9834957122803,21.9190425872803,21.877685546875,21.8141613006592,21.7674312591553,21.860595703125,21.9836921691895,22.2755374908447,22.2129402160645,22.0318851470947,21.9143562316895,21.8210926055908,21.7210941314697,21.7069816589355,21.7223625183105,21.7842769622803,21.8727531433105,22.0149402618408,22.0931644439697,22.1862297058105,22.2746086120605,22.5109367370605,22.6320323944092,22.671142578125,22.6875495910645,22.843505859375,22.9388675689697,23.04052734375,23.1866207122803,23.2103996276855,23.2296867370605,23.2549800872803,23.2928218841553,23.4366207122803,23.4930171966553,23.5500011444092,23.57275390625,23.6021976470947,23.6744136810303,23.8263664245605,24.0025405883789,24.0696296691895,24.1862316131592,24.2309093475342,24.27490234375,24.3259773254395,24.3466320037842,24.3892574310303,24.453857421875,24.479736328125,24.4606437683105,24.4857902526855,24.5499019622803,24.6278324127197,24.6644535064697,24.7130374908447,24.9146480560303,24.9206066131592,24.8819332122803,24.8818359375,24.9615230560303,25.1884288787842,25.1878910064697,25.1804695129395,25.1689453125,25.1762199401855,25.1943855285645,25.2229976654053,25.25927734375,25.290771484375,25.3335456848145,25.3655281066895,25.4562492370605,25.490478515625,25.4953136444092,25.5370121002197,25.5744132995605,25.6981945037842,25.789794921875,25.8114242553711,25.8411140441895,25.8882312774658,25.9563484191895,26.0182113647461,26.087158203125,26.1780757904053,26.2575206756592,26.31201171875,26.3694820404053,26.4010257720947,26.4371089935303,26.4715328216553,26.4825687408447,26.5047855377197,26.5641117095947,26.571533203125,26.5177745819092,26.4306640625,26.3529777526855,26.2917003631592,26.2818355560303,26.2793960571289,26.252197265625,26.260498046875,26.245361328125,26.2508792877197,26.2861328125,26.3379878997803,26.3750972747803,26.412109375,26.4195327758789,26.4102554321289,26.3769054412842,26.308349609375,26.2022457122803,26.10595703125,26.03759765625,26.006103515625,25.9835453033447,26.0052719116211,26.0724124908447,26.13232421875,26.18603515625,26.2156734466553,26.2138195037842,26.1712417602539,25.94140625,25.839599609375,25.5601558685303,25.3758296966553,25.3361320495605,25.3053703308105,25.2927722930908,25.2931632995605,25.2583484649658,25.219970703125,25.1849613189697,25.15771484375,25.1666011810303,25.167724609375,25.1594734191895,25.174072265625,25.177978515625,25.151611328125,25.1421375274658,25.16064453125,25.1694831848145,25.1109371185303,25.01513671875,24.9441413879395,24.9033203125,24.8685054779053,24.8494129180908,24.8487796783447,24.8950672149658,24.88134765625,24.7862300872803,24.77099609375,24.6857414245605,24.4939441680908,24.4080562591553,24.3861808776855,24.3743648529053,24.3708992004395,24.3567371368408,24.3255367279053,24.2606945037842,24.1953125,24.17529296875,24.2106437683105,24.205078125,24.1900882720947,24.15283203125,24.1065921783447,24.0907726287842,24.10009765625,24.093505859375,24.0604972839355,24.018798828125,23.9204597473145,23.7628898620605,23.66064453125,23.5810565948486,23.3736324310303,23.1043949127197,23.0770034790039,23.068359375,23.0745601654053,23.1304683685303,23.197998046875,23.2138671875,23.1999015808105,23.1412601470947,23.03369140625,22.9915523529053,22.9796886444092,23.0048332214355,23.0535163879395,23.1061038970947,23.2098140716553,23.2873039245605,23.3610343933105,23.4710445404053,23.5046863555908,23.5982418060303,23.6859855651855,23.6919918060303,23.6777839660645,23.7209968566895,23.7218761444092,23.675537109375,23.68359375,23.4924793243408,23.3238277435303,23.2423820495605,23.06982421875,22.9290027618408,22.8970203399658,22.8218250274658,22.7344245910645,22.6854019165039,22.5256843566895,22.4103031158447,22.0480461120605,21.9780769348145],[43.7422370910645,43.5013160705566,43.3893051147461,43.4268531799316,43.3956069946289,43.2682609558105,43.2305183410645,43.1861305236816,43.0207061767578,42.7497062683105,42.7166481018066,42.70654296875,42.4680671691895,42.4009780883789,42.3499526977539,42.2080078125,42.0474128723145,41.9690437316895,41.9866218566895,41.9813003540039,41.95654296875,41.9615249633789,41.9613304138184,41.9329109191895,41.9208030700684,41.9468765258789,42.0250473022461,42.0795402526855,42.09326171875,42.0770988464355,42.0586433410645,42.02685546875,41.9918441772461,41.9751472473145,41.9633293151855,41.9648933410645,41.9479484558105,41.896728515625,41.8466796875,41.8263664245605,41.8015632629395,41.7728004455566,41.7446784973145,41.716552734375,41.7437973022461,41.7256851196289,41.7041511535645,41.6732406616211,41.6401863098145,41.6082038879395,41.5215339660645,41.4348640441895,41.3857421875,41.3506813049316,41.3119125366211,41.3304214477539,41.3150405883789,41.3101081848145,41.2998046875,41.2643547058105,41.2435531616211,41.3157730102539,41.364990234375,41.3942375183105,41.3729019165039,41.3561019897461,41.4199714660645,41.4427261352539,41.4673843383789,41.5525398254395,41.5552253723145,41.5235328674316,41.5250473022461,41.5308113098145,41.5272445678711,41.4690895080566,41.4600563049316,41.4522933959961,41.4559555053711,41.4129867553711,41.3867645263672,41.3860359191895,41.3987312316895,41.3896484375,41.3849601745605,41.3220710754395,41.3256378173828,41.3362770080566,41.3561019897461,41.6056175231934,41.7169456481934,41.7398414611816,41.7571792602539,41.7750968933105,41.835205078125,41.9936065673828,42.0256843566895,42.0591316223145,42.1048355102539,42.1650886535645,42.31396484375,42.328857421875,42.359130859375,42.4409675598145,42.481201171875,42.50390625,42.5433120727539,42.6291007995605,42.70947265625,42.7507820129395,42.7916526794434,42.8424797058105,42.8703117370605,42.8784675598145,42.8839378356934,42.9857406616211,43.0182609558105,43.0759773254395,43.0970687866211,43.1420402526855,43.18798828125,43.2523422241211,43.3007316589355,43.3541526794434,43.3910636901855,43.4544944763184,43.5188484191895,43.6022491455078,43.6654777526855,43.7066383361816,43.7401351928711,43.7812957763672,43.8621101379395,43.9695320129395,44.0074195861816,44.0180168151855,44.0752944946289,44.1485824584961,44.194091796875,44.22021484375,44.23779296875,44.1952133178711,44.1272926330566,44.0779762268066,44.0472183227539,44.0169944763184,43.9872055053711,43.9478988647461,43.8990211486816,43.8645515441895,43.83447265625,43.8738784790039,43.8535652160645,43.7866706848145,43.7634773254395,43.7943840026855,43.7384262084961,43.6863288879395,43.6708030700684,43.7117652893066,43.766845703125,43.8705558776855,44.0072746276855,44.083984375,44.1461906433105,44.1673851013184,44.1461410522461,44.0205078125,44.0200653076172,43.997802734375,43.964599609375,43.956298828125,43.9873542785645,43.9186058044434,43.8224105834961,43.7728500366211,43.7447776794434,43.740478515625,43.7422370910645],[25.8831062316895,25.8067874908447,25.83349609375,25.9655284881592,26.0584468841553,26.1908187866211,26.2289562225342,26.2464370727539,26.24072265625,26.1982917785645,26.1244621276855,26.0023441314697,25.8831062316895],[21.1923828125,21.1190414428711,20.9791488647461,20.9438953399658,20.9374027252197,20.9755859375,21.0091304779053,21.0615711212158,21.0822277069092,21.1033210754395,21.1259288787842,21.1908206939697,21.2017574310303,21.1561527252197,21.1544933319092,21.2047843933105,21.3133792877197,21.2995109558105,21.1923828125],[21.5066890716553,21.4576168060303,21.5153312683105,21.5616207122803,21.5066890716553],[22.4290523529053,22.4145984649658,22.4156265258789,22.3855953216553,22.3443851470947,22.327392578125,22.2906246185303,22.3602523803711,22.3692378997803,22.3675785064697,22.3807125091553,22.455322265625,22.4290523529053],[22.2137699127197,22.1836910247803,22.2355461120605,22.3233890533447,22.400634765625,22.427978515625,22.44921875,22.4777336120605,22.5274410247803,22.5680179595947,22.6350593566895,22.6822757720947,22.7155265808105,22.7310562133789,22.6807613372803,22.5384273529053,22.3611812591553,22.3062496185303,22.2137699127197],[22.7234859466553,22.70556640625,22.6654300689697,22.715087890625,22.711669921875,22.7644538879395,22.8035640716553,22.839599609375,22.8115711212158,22.7599105834961,22.7234859466553],[22.8943367004395,22.8687019348145,23.0685539245605,23.1170902252197,23.1653327941895,23.1927261352539,23.2046394348145,23.2679214477539,23.3363761901855,23.474609375,23.5682621002197,23.58984375,23.6683597564697,23.5467758178711,23.4386711120605,23.3328132629395,23.1501960754395,23.088134765625,22.9999027252197,22.8943367004395],[23.4501457214355,23.4442386627197,23.4706535339355,23.59228515625,23.602783203125,23.67138671875,23.6474132537842,23.5425300598145,23.489990234375,23.4501457214355],[24.0680675506592,23.959716796875,23.9689464569092,24.1050777435303,24.1266593933105,24.12548828125,24.0680675506592],[24.2494640350342,24.2265625,24.1634769439697,24.1184558868408,24.2163562774658,24.1368160247803,23.9876461029053,23.961669921875,23.9394054412842,23.9108390808105,23.8694324493408,23.7391605377197,23.7525386810303,23.8623542785645,23.8835468292236,24.0403804779053,24.0909175872803,24.2198257446289,24.2530765533447,24.2419929504395,24.2575187683105,24.2699222564697,24.2875480651855,24.2494640350342],[24.2000007629395,24.149169921875,24.1594715118408,24.1395988464355,24.1390628814697,24.1738777160645,24.220947265625,24.2657718658447,24.3304195404053,24.4912605285645,24.5293941497803,24.58984375,24.6546878814697,24.6893539428711,24.697509765625,24.6808586120605,24.4495124816895,24.4273433685303,24.2000007629395],[25.0138664245605,24.9931144714355,25.0135746002197,25.030029296875,25.0576648712158,25.0807132720947,25.0830078125,25.0557613372803,25.0438480377197,25.0138664245605],[24.7074222564697,24.5863285064697,24.4957504272461,24.4634761810303,24.4029293060303,24.3690929412842,24.3349609375,24.2874507904053,24.3646488189697,24.412353515625,24.4934558868408,24.466064453125,24.4827632904053,24.544189453125,24.6275882720947,24.6420402526855,24.5902347564697,24.65380859375,24.6873035430908,24.6916007995605,24.75390625,24.9170894622803,25.0223636627197,25.1912612915039,25.2023448944092,25.1431140899658,25.0848140716553,25.0047855377197,24.9428234100342,24.7943840026855,24.7074222564697],[25.4874019622803,25.3746089935303,25.33203125,25.1908206939697,25.1405277252197,25.0947265625,24.8856449127197,24.759765625,24.6494140625,24.6820812225342,24.7543449401855,24.7958984375,24.8176765441895,24.8224620819092,24.9362297058105,25.0259761810303,25.1193351745605,25.2221202850342,25.3125972747803,25.341552734375,25.431640625,25.4427242279053,25.4268569946289,25.4805679321289,25.5516109466553,25.5648918151855,25.4874019622803],[26.7290515899658,26.6979484558105,26.7021980285645,26.72265625,26.7143058776855,26.7442378997803,26.6911144256592,26.6633777618408,26.6373538970947,26.5593757629395,26.5065441131592,26.5006828308105,26.5284671783447,26.6895027160645,26.6734371185303,26.5824222564697,26.5990238189697,26.6591796875,26.7046394348145,26.7261714935303,26.7979507446289,26.7290515899658],[25.9041996002197,25.8954582214355,25.99560546875,26.0247058868408,26.0955085754395,26.1563472747803,26.2890625,26.3617668151855,26.4247074127197,26.4889640808105,26.5611324310303,26.6183586120605,26.8459949493408,26.9012699127197,26.903564453125,26.9400882720947,26.9356441497803,26.9139156341553,26.9034175872803,26.83642578125,26.7476081848145,26.711669921875,26.6888179779053,26.6630363464355,26.6388187408447,26.5972652435303,26.5301761627197,26.3334484100342,26.2403316497803,25.9554691314697,25.9041996002197],[44.8691902160645,44.871337890625,44.8996124267578,44.9142570495605,44.8974609375,44.8809051513672,44.8585433959961,44.7806625366211,44.69677734375,44.6095695495605,44.52734375,44.4837875366211,44.41455078125,44.3599624633789,44.3302726745605,44.3025398254395,44.2805671691895,44.225830078125,44.1544914245605,44.073486328125,44.04345703125,44.0110855102539,43.9871063232422,43.97802734375,43.985107421875,43.9933586120605,43.9834480285645,43.9650382995605,43.9433097839355,43.8447723388672,43.7035636901855,43.6428718566895,43.5951652526855,43.5620613098145,43.5675773620605,43.5934562683105,43.591796875,43.5843772888184,43.5332984924316,43.5354499816895,43.5277328491211,43.5177230834961,43.5325164794922,43.5423316955566,43.5266609191895,43.4967269897461,43.4423828125,43.3573226928711,43.2924308776855,43.285400390625,43.3394508361816,43.3481903076172,43.3463363647461,43.2835426330566,43.2308120727539,43.1939468383789,43.1536598205566,43.1246070861816,43.0276832580566,43.0121574401855,42.9978981018066,42.9684562683105,42.8440933227539,42.7772445678711,42.6741714477539,42.6416015625,42.6201171875,42.5791969299316,42.5645027160645,42.5597190856934,42.586669921875,42.5994148254395,42.6905746459961,42.7412605285645,42.8074226379395,42.8450698852539,42.9022445678711,42.9154777526855,42.8971176147461,42.9383811950684,42.9597663879395,42.9800796508789,43.006591796875,43.0427742004395,43.1989250183105,43.3056144714355,43.3438491821289,43.4457511901855,43.47021484375,43.5165519714355,43.6490249633789,43.77880859375,43.8150367736816,43.9131851196289,44.0025901794434,44.0596160888672,44.12451171875,44.2151374816895,44.3520011901855,44.4737319946289,44.52099609375,44.53759765625,44.6819343566895,44.7658195495605,44.8563957214355,45.0075187683105,45.178955078125,45.2027816772461,45.2157211303711,45.2107925415039,45.1896018981934,45.0722160339355,45.026611328125,45.0088348388672,45.058349609375,45.1620140075684,45.2166976928711,45.1968765258789,45.2765617370605,45.1717758178711,45.1560554504395,45.1705589294434,45.1639633178711,45.13330078125,45.120361328125,45.12255859375,45.1634750366211,45.1584014892578,45.078125,45.0772476196289,45.1118659973145,45.1417961120605,45.119384765625,45.1329116821289,45.13427734375,45.1205558776855,45.1020050048828,45.0858383178711,45.0774421691895,45.0265121459961,44.9772453308105,44.9472160339355,44.9148941040039,44.8832511901855,44.8651885986328,44.8691902160645],[17.87646484375,17.8751964569092,17.8836421966553,17.8985843658447,17.91357421875,17.9222660064697,17.9182624816895,17.90869140625,17.8976573944092,17.8854503631592,17.87646484375],[56.1429176330566,56.055908203125,56.0525398254395,56.0867195129395,56.0890159606934,56.0919914245605,56.061767578125,56.0026359558105,55.9553718566895,55.9425773620605,56.0021018981934,56.0217781066895,56.0211448669434,55.9678726196289,55.938720703125,55.9122047424316,55.8810577392578,55.834716796875,55.7843742370605,55.724853515625,55.6845703125,55.7511749267578,55.7697257995605,55.7704086303711,55.7951202392578,55.8323249816895,55.8452606201172,55.83642578125,55.84521484375,55.7868156433105,55.768798828125,55.7002906799316,55.666259765625,55.6554679870605,55.6221199035645,55.6011238098145,55.6075210571289,55.5963859558105,55.5700187683105,55.5253448486328,55.3974151611328,55.3604011535645,55.3302726745605,55.3069839477539,55.2787132263184,55.2234382629395,55.1375961303711,55.0877914428711,55.0504875183105,54.9407234191895,54.9149894714355,54.8609352111816,54.8060035705566,54.7832527160645,54.6958999633789,54.6484870910645,54.6253433227539,54.6109390258789,54.51708984375,54.4917945861816,54.4529800415039,54.3916511535645,54.2916984558105,54.1959457397461,54.1111793518066,54.055908203125,54.0307121276855,54.0007820129395,53.9350090026855,53.85498046875,53.8104515075684,53.7919425964355,53.796875,53.78125,53.6929206848145,53.6533203125,53.6172866821289,53.5792465209961,53.5469703674316,53.4481468200684,53.41943359375,53.3714370727539,53.3363304138184,53.3289031982422,53.3124046325684,53.2703132629395,53.2105941772461,53.1283721923828,53.0911636352539,53.0894508361816,53.106201171875,53.1468772888184,53.1841812133789,53.200927734375,53.2024917602539,53.196044921875,53.184814453125,53.1389656066895,53.0608863830566,53.0167007446289,52.9897956848145,52.9334487915039,52.86181640625,52.7982444763184,52.7592277526855,52.7314453125,52.69873046875,52.6330070495605,52.5461921691895,52.532470703125,52.426025390625,52.3123054504395,52.2848129272461,52.26220703125,52.2206535339355,52.1258316040039,52.10107421875,52.1081047058105,52.1053733825684,52.0502471923828,52.0629386901855,52.0769538879395,52.0461921691895,51.9530754089355,51.8951644897461,51.8141136169434,51.7700691223145,51.68896484375,51.5963401794434,51.531494140625,51.4712409973145,51.4063453674316,51.3554191589355,51.318359375,51.2650413513184,51.2743148803711,51.3255386352539,51.3996086120605,51.4512214660645,51.4778785705566,51.4820327758789,51.4580078125,51.4395523071289,51.4345703125,51.4083480834961,51.382568359375,51.4130363464355,51.4970207214355,51.5806159973145,51.6172866821289,51.6275405883789,51.6254386901855,51.5989265441895,51.57177734375,51.5621566772461,51.5401840209961,51.5103492736816,51.4333992004395,51.4388694763184,51.45654296875,51.5426254272461,51.5624504089355,51.5636215209961,51.5818328857422,51.6078605651855,51.6016578674316,51.5650405883789,51.5597648620605,51.5923843383789,51.5774421691895,51.5291481018066,51.4825668334961,51.4779777526855,51.4899406433105,51.5724105834961,51.6016120910645,51.6061019897461,51.5941390991211,51.597412109375,51.6135749816895,51.7520484924316,51.7608413696289,51.7540016174316,51.7707023620605,51.8019027709961,51.8134307861328,51.8444366455078,51.8550300598145,51.9135246276855,51.923828125,51.9247550964355,51.937744140625,51.9305152893066,51.9111328125,51.8991203308105,51.8882827758789,51.8895034790039,51.883056640625,51.8675270080566,51.8384284973145,51.7747077941895,51.6646461486816,51.59130859375,51.5850601196289,51.6239738464355,51.6371078491211,51.6413078308105,51.6288566589355,51.6104965209961,51.525390625,51.5179214477539,51.618896484375,51.7102546691895,51.7624015808105,51.8093223571777,51.8797836303711,51.9729995727539,52.0403785705566,52.069580078125,52.1030769348145,52.140380859375,52.1695327758789,52.2084465026855,52.2569351196289,52.28662109375,52.3069801330566,52.337890625,52.4283676147461,52.5162124633789,52.5515594482422,52.6642074584961,52.70361328125,52.770263671875,52.8187522888184,52.9048843383789,53.0275382995605,53.1121063232422,53.2709465026855,53.5992202758789,53.9397926330566,53.9198265075684,53.9122543334961,53.9356918334961,53.93896484375,53.931640625,53.9450187683105,53.950439453125,53.9199714660645,53.8929672241211,53.9318351745605,53.9798355102539,53.9746589660645,53.9982414245605,54.1189918518066,54.1451683044434,54.133056640625,54.1549301147461,54.2142562866211,54.2512702941895,54.251220703125,54.2927742004395,54.2649421691895,54.2151336669922,54.1752433776855,54.1596183776855,54.139892578125,54.1404762268066,54.156982421875,54.1797866821289,54.22119140625,54.2596664428711,54.29296875,54.3101043701172,54.3106918334961,54.3318367004395,54.3770523071289,54.4603996276855,54.5357933044434,54.5642585754395,54.5903816223145,54.6360359191895,54.7178726196289,54.833251953125,54.9192886352539,54.9471664428711,54.9623031616211,55.0032691955566,55.0503883361816,55.0901374816895,55.12451171875,55.1395988464355,55.1301765441895,55.2041969299316,55.2249031066895,55.2467765808105,55.2730941772461,55.2933578491211,55.3064460754395,55.3180160522461,55.3424835205078,55.3719253540039,55.4481468200684,55.5464820861816,55.6226577758789,55.6675300598145,55.6796379089355,55.6939926147461,55.709228515625,55.8129386901855,55.83056640625,55.8039054870605,55.8059577941895,55.8035163879395,55.7987785339355,55.8091773986816,55.9117202758789,55.9415512084961,56.076171875,56.13330078125,56.1458015441895,56.1429176330566],[17.4228515625,17.2830085754395,17.3229503631592,17.4264640808105,17.4627933502197,17.5010738372803,17.5462894439697,17.5242195129395,17.4795894622803,17.4228515625],[17.9249515533447,17.9063472747803,17.9640140533447,18.0010738372803,18.1549320220947,18.154052734375,18.140380859375,17.9249515533447],[15.890625,15.8898439407349,15.9006834030151,15.8886709213257,15.8944339752197,16.142822265625,16.5271492004395,16.8089847564697,17.0846672058105,17.2912120819092,17.572265625,17.8148441314697,17.9019527435303,17.9708023071289,17.9997062683105,17.9396495819092,17.91455078125,17.9288082122803,17.9655265808105,18.0716304779053,18.29052734375,18.4458999633789,18.4767570495605,18.4823246002197,18.472412109375,18.3588390350342,18.3440914154053,18.3546867370605,18.3507328033447,18.2261219024658,18.1216316223145,17.8460941314697,17.7513656616211,17.60986328125,17.5166015625,17.392578125,17.3126964569092,17.192138671875,16.9630374908447,16.6327648162842,16.4886226654053,16.4337882995605,16.2904300689697,16.2476558685303,16.0166492462158,15.9560070037842,15.890625],[32.29345703125,32.2596168518066,32.2623023986816,32.2738761901855,32.3077125549316,32.3869132995605,32.3819351196289,32.29345703125],[-20.1646480560303,-19.9988288879395,-19.8548831939697,-19.81787109375,-19.6460933685303,-19.490234375,-19.2862300872803,-19.2917957305908,-19.2975597381592,-19.3887691497803,-19.478515625,-19.52099609375,-19.6064453125,-19.6453132629395,-19.8094730377197,-20.0553722381592,-20.1990222930908,-20.3499031066895,-20.5625,-20.82080078125,-21.0660152435303,-21.4117183685303,-21.7052745819092,-21.9886722564697,-22.1839847564697,-22.2336921691895,-22.2179679870605,-22.1598644256592,-22.0496101379395,-21.9991207122803,-21.9972667694092,-22.0005855560303,-22.0042953491211,-22.0275402069092,-22.0272464752197,-22.0054683685303,-22.0072269439697,-22.0286140441895,-22.0725593566895,-22.3658199310303,-22.4913101196289,-22.6033210754395,-22.7953128814697,-22.82763671875,-22.7610359191895,-22.5853519439697,-22.4853515625,-22.37158203125,-22.2288074493408,-22.185546875,-22.1712894439697,-22.1439456939697,-22.1096668243408,-22.1027355194092,-22.09814453125,-22.0945320129395,-22.1102542877197,-22.099609375,-22.0197277069092,-21.8794918060303,-21.8350582122803,-21.8056640625,-21.8025398254395,-21.8304691314697,-21.9474601745605,-22.0531253814697,-22.11376953125,-22.1583995819092,-22.2053718566895,-22.21630859375,-22.2693347930908,-22.3430671691895,-22.40966796875,-22.5098628997803,-22.55224609375,-22.65087890625,-22.7738285064697,-22.8216800689697,-22.8551750183105,-22.8916988372803,-22.88916015625,-22.8794937133789,-22.8577156066895,-22.8229484558105,-22.7841796875,-22.7292003631592,-22.6305656433105,-22.4933605194092,-22.3336906433105,-22.2822265625,-22.2040042877197,-22.05712890625,-21.9828109741211,-21.8606433868408,-21.7530269622803,-21.6185550689697,-21.447265625,-21.30029296875,-21.1296863555908,-20.9482421875,-20.9236316680908,-20.9019527435303,-20.8498039245605,-20.7691402435303,-20.7201175689697,-20.6407222747803,-20.62841796875,-20.6120109558105,-20.4929676055908,-20.4585933685303,-20.4162101745605,-20.3780269622803,-20.3389644622803,-20.3100566864014,-20.2251949310303,-20.1484355926514,-20.1155261993408,-20.0908203125,-20.0696277618408,-20.044921875,-19.9670906066895,-19.90234375,-19.8565425872803,-19.74072265625,-19.7210941314697,-19.5601558685303,-19.4543952941895,-19.4328117370605,-19.4099617004395,-19.3819332122803,-19.3411140441895,-19.2966785430908,-19.2423820495605,-19.1622066497803,-19.0933609008789,-19.0251941680908,-18.9679679870605,-18.9096698760986,-18.81298828125,-18.65625,-18.5500965118408,-18.4330062866211,-18.3566398620605,-18.2824211120605,-18.2024402618408,-18.14404296875,-18.1027355194092,-18.0707035064697,-18.0504875183105,-17.96484375,-17.9431648254395,-17.7716789245605,-17.6195316314697,-17.5060558319092,-17.5048828125,-17.4603500366211,-17.3889656066895,-17.3329105377197,-17.29443359375,-17.24853515625,-17.2001953125,-17.1047840118408,-17.08837890625,-17.0400390625,-17.0013675689697,-16.8609371185303,-16.7684574127197,-16.7130851745605,-16.67431640625,-16.6421871185303,-16.5426750183105,-16.4759769439697,-16.4336910247803,-16.3890628814697,-16.3547859191895,-16.337890625,-16.3127937316895,-16.2619132995605,-16.2176761627197,-16.2219734191895,-16.1828136444092,-16.1491222381592,-15.7369146347046,-15.6406240463257,-15.6034183502197,-15.3994140625,-15.3329105377197,-15.2366199493408,-15.1987295150757,-15.0389652252197,-14.9629878997803,-14.8875007629395,-14.7953128814697,-14.7458982467651,-14.6710939407349,-14.5970706939697,-14.5725584030151,-14.5309572219849,-14.4703121185303,-14.4175786972046,-14.3772459030151,-14.2650384902954,-14.234375,-14.1988277435303,-14.1697263717651,-14.0943365097046,-14.0146484375,-13.9759769439697,-13.7802724838257,-13.6828126907349,-13.6439456939697,-13.5944337844849,-13.4963874816895,-13.38232421875,-12.9625978469849,-12.8800783157349,-12.8220701217651,-12.7551755905151,-12.7295904159546,-12.6872072219849,-12.6077146530151,-12.5607423782349,-12.5019521713257,-12.2704105377197,-12.0667972564697,-11.8756828308105,-11.654296875,-11.5085935592651,-11.3275394439697,-11.1687498092651,-10.9517574310303,-10.9481439590454,-10.9556646347046,-10.9943361282349,-11.0111322402954,-11.0446290969849,-11.09765625,-11.1224613189697,-11.11279296875,-11.1091794967651,-11.0547847747803,-11.0187501907349,-10.9751949310303,-10.9331064224243,-10.7850580215454,-10.703125,-10.6744146347046,-10.6627931594849,-10.68603515625,-10.6831045150757,-10.5989255905151,-10.5059566497803,-10.3898439407349,-10.35791015625,-10.3172845840454,-10.3114252090454,-10.2689456939697,-9.97548866271973,-9.89990234375,-9.88613319396973,-9.8681640625,-9.82607364654541,-9.78544902801514,-9.76845741271973,-9.80908203125,-9.79746055603027,-9.73173809051514,-9.71044921875,-9.71240234375,-9.79023456573486,-9.87265682220459,-9.93554782867432,-10.0269536972046,-10.1467771530151,-10.2530279159546,-10.3922843933105,-10.4490232467651,-10.5074224472046,-10.5862302780151,-10.7147464752197,-10.8927736282349,-11.0248050689697,-11.1103515625,-11.1842765808105,-11.2462892532349,-11.2899417877197,-11.3150386810303,-11.36474609375,-11.4391593933105,-11.5110349655151,-11.58056640625,-11.6468753814697,-11.7100591659546,-11.74951171875,-11.76513671875,-11.7523431777954,-11.7350578308105,-11.7412109375,-11.8293943405151,-11.8473634719849,-11.9200201034546,-11.9751958847046,-12.0059566497803,-12.0302734375,-12.0593748092651,-12.146484375,-12.2039060592651,-12.2509765625,-12.326171875,-12.4397459030151,-12.4833011627197,-12.5050783157349,-12.5296878814697,-12.4694328308105,-12.47802734375,-12.5189456939697,-12.5466794967651,-12.6051759719849,-12.6800775527954,-12.7079105377197,-12.6662111282349,-12.6516599655151,-12.6691408157349,-12.7503900527954,-12.8055667877197,-12.8470697402954,-12.9537105560303,-12.9972658157349,-12.9943361282349,-13.0642576217651,-13.1324214935303,-13.1436519622803,-13.1336917877197,-13.1597652435303,-13.2419919967651,-13.40625,-13.4704103469849,-13.5255861282349,-13.5248041152954,-13.5412111282349,-13.5265617370605,-13.49853515625,-13.48974609375,-13.5614261627197,-13.6643552780151,-13.7453126907349,-13.7898435592651,-13.8624029159546,-13.9379892349243,-14.0192384719849,-14.1000003814697,-14.1324224472046,-14.1847648620605,-14.2630863189697,-14.3328123092651,-14.4187498092651,-14.5705080032349,-14.6185550689697,-15.0887689590454,-15.0927734375,-15.0983400344849,-15.1431646347046,-15.3182611465454,-15.4795904159546,-15.7386713027954,-15.9019527435303,-16.1321277618408,-16.2693367004395,-16.28173828125,-16.2959957122803,-16.3131847381592,-16.3282222747803,-16.32666015625,-16.3079090118408,-16.2835922241211,-16.2843742370605,-16.3399410247803,-16.4102535247803,-16.4908199310303,-16.6501941680908,-16.70068359375,-16.9107418060303,-17.08056640625,-17.2342758178711,-17.2821292877197,-17.3630847930908,-17.5128898620605,-17.5323238372803,-17.5121097564697,-17.5730476379395,-17.6717777252197,-17.9473628997803,-18.1222667694092,-18.18310546875,-18.2146492004395,-18.2373046875,-18.2464847564697,-18.279296875,-18.4750003814697,-18.7332019805908,-18.9142570495605,-18.9171867370605,-18.9673824310303,-19.0440425872803,-19.0535163879395,-19.0809574127197,-19.2294921875,-19.4242172241211,-19.6252937316895,-19.7445316314697,-19.8327140808105,-19.9795913696289,-20.0204105377197,-20.0407218933105,-20.05517578125,-20.1103515625,-20.1510753631592,-20.1646480560303],[-27.7669925689697,-27.8122081756592,-27.5748043060303,-27.4955062866211,-27.4363288879395,-27.3996086120605,-27.4514636993408,-27.5663089752197,-27.70703125,-27.7669925689697],[-26.4015617370605,-26.4137687683105,-26.2896480560303,-26.1703128814697,-26.21875,-26.3131847381592,-26.3796882629395,-26.4015617370605],[-23.88916015625,-23.9413089752197,-23.9147453308105,-23.9349594116211,-23.8956050872803,-23.7275390625,-23.751953125,-23.7826156616211,-23.8253898620605,-23.85302734375,-23.88916015625],[-23.1418952941895,-23.1693344116211,-23.1666011810303,-23.1908187866211,-23.2123031616211,-23.1720714569092,-23.1162109375,-23.0741214752197,-23.0829086303711,-23.11328125,-23.1418952941895],[-13.4734373092651,-13.5323238372803,-13.5235347747803,-13.4840822219849,-13.4456052780151,-13.41552734375,-13.3984375,-13.4010744094849,-13.4734373092651],[-13.0970706939697,-13.11865234375,-13.0550785064697,-12.9749021530151,-12.8801765441895,-12.9240236282349,-12.9724617004395,-12.9925775527954,-13.0970706939697],[-2.93964815139771,-3.03759765625,-2.92392563819885,-2.84560513496399,-2.78496098518372,-2.72626948356628,-2.71757841110229,-2.78974580764771,-2.81191420555115,-2.93964815139771],[-1.31787109375,-1.36601567268372,-1.39082014560699,-1.37236344814301,-1.34472680091858,-1.34755837917328,-1.26728522777557,-1.27685534954071,-1.31787109375],[-1.43378901481628,-1.45263648033142,-1.20253932476044,-1.08613264560699,-0.855078220367432,-0.649609267711639,-0.565917909145355,-0.541406154632568,-0.666699409484863,-1.02177739143372,-1.02382802963257,-1.21113312244415,-1.34189462661743,-1.43378901481628],[-0.229199171066284,-0.233593747019768,-0.214648321270943,-0.167870908975601,-0.158691555261612,-0.163574293255806,-0.215527251362801,-0.231640532612801,-0.248241990804672,-0.271874785423279,-0.297363370656967,-0.352831989526749,-0.441504091024399,-0.534765541553497,-0.664941608905792,-0.691406488418579,-0.684472560882568,-0.800976634025574,-0.847558736801147,-0.892871081829071,-0.986914098262787,-1.10664045810699,-1.13173830509186,-1.17333960533142,-1.22656226158142,-1.27656257152557,-1.32695305347443,-1.39003908634186,-1.48232436180115,-1.50468742847443,-1.51406276226044,-1.50507819652557,-1.41259765625,-1.48496103286743,-1.55898439884186,-1.59951198101044,-1.59521484375,-1.55556666851044,-1.51162135601044,-1.63046884536743,-1.71240234375,-1.73808586597443,-1.75537133216858,-1.79023408889771,-1.76298820972443,-1.70849597454071,-1.70380866527557,-1.74785125255585,-1.75595688819885,-1.80068373680115,-1.78798818588257,-1.69775414466858,-1.63769555091858,-1.51601529121399,-1.37148451805115,-1.24023413658142,-1.12675762176514,-1.13056659698486,-1.1474609375,-1.13945329189301,-1.10312485694885,-1.07294940948486,-1.07773411273956,-1.01035141944885,-0.906250178813934,-0.689843475818634,-0.645410001277924,-0.583398222923279,-0.528515756130219,-0.470214992761612,-0.364453285932541,-0.272851675748825,-0.157421872019768,-0.116406112909317,-0.229199171066284],[-0.131640493869781,-0.327343970537186,-0.263086199760437,-0.225878864526749,-0.188378781080246,-0.172363460063934,-0.105273544788361,-0.05019511282444,-0.0408690795302391,-0.0580079630017281,-0.105859108269215,-0.131640493869781],[-0.11240229010582,-0.143749952316284,-0.093896321952343,-0.0518555454909801,0.0151369711384177,0.0626953318715096,0.0836913734674454,0.0572264529764652,0.00107429595664144,-0.0554686151444912,-0.11240229010582],[0.268164306879044,0.215966701507568,0.00688466103747487,-0.0231933314353228,-0.0292970519512892,0.0330080986022949,0.0285642370581627,0.0433591529726982,0.134472623467445,0.231738492846489,0.22651369869709,0.304540812969208,0.268164306879044],[0.139257967472076,-0.00766617525368929,0.0543943233788013,0.204785421490669,0.246923610568047,0.326904505491257,0.42495134472847,0.558251678943634,0.581396341323853,0.590869069099426,0.581738352775574,0.381591618061066,0.25903308391571,0.139257967472076],[0.393017530441284,0.359179794788361,0.390820235013962,0.516503691673279,0.585449397563934,0.604736328125,0.62500011920929,0.638037204742432,0.594824135303497,0.522802770137787,0.393017530441284],[1.93852531909943,1.89287102222443,1.91049802303314,2.02954077720642,2.12861323356628,2.16147470474243,2.15444326400757,2.14174842834473,1.97958993911743,1.93852531909943],[1.94213843345642,1.93232417106628,1.92387712001801,1.88535141944885,1.84223628044128,1.85707998275757,1.88749992847443,1.97666037082672,2.03955101966858,2.09511709213257,2.15815424919128,2.23676753044128,2.25903344154358,2.29951167106628,2.34130835533142,2.36440420150757,2.39277315139771,2.49750995635986,2.51596665382385,2.52045917510986,2.51660132408142,2.48950219154358,2.40615224838257,2.41874980926514,2.44062495231628,2.48876929283142,2.49965834617615,2.54750967025757,2.55078101158142,2.53920912742615,2.54833984375,2.59296870231628,2.59765601158142,2.54833984375,2.49736332893372,2.45039033889771,2.43955063819885,2.45473623275757,2.441650390625,2.39794921875,2.35981440544128,2.32753920555115,2.32675743103027,2.31376957893372,2.29306650161743,2.24545884132385,2.20751929283142,2.15424799919128,2.15332055091858,2.13706040382385,2.12104511260986,2.15048837661743,2.23256826400757,2.27827143669128,2.31293964385986,2.34599614143372,2.35483431816101,2.33500981330872,2.30854511260986,2.29291987419128,2.26191425323486,2.25312519073486,2.27944326400757,2.32421875,2.33974599838257,2.29521512985229,2.23227572441101,2.20488262176514,2.21132826805115,2.20170903205872,2.18173861503601,2.18354511260986,2.21152377128601,2.26665043830872,2.31718730926514,2.36367201805115,2.42573261260986,2.52890610694885,2.57314419746399,2.64765620231628,2.86416053771973,2.90385723114014,2.97221684455872,3.05156230926514,3.11772465705872,3.18173837661743,3.23710942268372,3.27167963981628,3.36469769477844,3.45229482650757,3.64687538146973,3.70200181007385,3.735107421875,3.77695322036743,3.82856488227844,3.86958003044128,3.92993211746216,3.99267578125,4.03969764709473,4.06127977371216,4.23378944396973,4.31088829040527,4.31376934051514,4.22475576400757,4.09360361099243,3.67167949676514,3.28183579444885,3.07753920555115,2.65185546875,2.57304692268372,2.47778344154358,2.37675786018372,2.21035146713257,2.17944312095642,2.13403344154358,2.13095736503601,2.10410141944885,1.99858403205872,1.92724609375,1.82958996295929,1.79765629768372,1.78598630428314,1.730712890625,1.65986323356628,1.41992175579071,1.26904308795929,1.16298806667328,1.12143540382385,1.05195343494415,1.01508808135986,0.835741996765137,0.75102561712265,0.637304961681366,0.420508027076721,0.222558572888374,0.172558844089508,0.160986244678497,0.130273416638374,-0.0312502644956112,-0.0852051302790642,-0.17880867421627,-0.392676055431366,-0.509472906589508,-0.549121022224426,-0.762304902076721,-0.855468988418579,-1.01845705509186,-1.11777353286743,-1.18085908889771,-1.3203125,-1.36796879768372,-1.39902365207672,-1.36249995231628,-1.51406276226044,-1.55175757408142,-1.55957055091858,-1.64013695716858,-1.58671891689301,-1.39433574676514,-1.35410153865814,-1.22353529930115,-1.13652360439301,-1.03212893009186,-0.98632800579071,-0.937597870826721,-0.999609053134918,-1.03886699676514,-1.11523425579071,-1.16445302963257,-1.22626948356628,-1.31142592430115,-1.37626922130585,-1.48994123935699,-1.64384746551514,-1.69472658634186,-1.76171863079071,-1.81708991527557,-1.84990251064301,-2.01552724838257,-1.92294943332672,-1.89619112014771,-1.85751950740814,-1.83183598518372,-1.87060523033142,-1.92636716365814,-1.86718761920929,-1.73173797130585,-1.97158217430115,-2.19150400161743,-2.2802734375,-2.51992177963257,-2.58388662338257,-2.65693354606628,-2.63144493103027,-2.59687495231628,-2.50458979606628,-2.34433579444885,-1.91650366783142,-1.87851548194885,-1.82978546619415,-1.48769521713257,-1.48876953125,-1.56748068332672,-1.61396479606628,-1.52040994167328,-1.48212897777557,-1.43583965301514,-1.39384758472443,-1.32382810115814,-1.22919905185699,-1.14550769329071,-1.03984391689301,-0.960547089576721,-0.895117044448853,-0.827929735183716,-0.795215010643005,-0.737499833106995,-0.713671624660492,-0.705077886581421,-0.769628942012787,-0.693359136581421,-0.663476526737213,-0.67675769329071,-0.71044933795929,-0.724804759025574,-0.718749761581421,-0.669921696186066,-0.748534977436066,-0.765917956829071,-0.721875250339508,-0.680956900119781,-0.64765602350235,-0.626659989356995,-0.645410001277924,-0.680468738079071,-0.745410084724426,-0.750195443630219,-0.743359446525574,-0.779882609844208,-0.779687285423279,-0.836523532867432,-0.916406452655792,-0.970605432987213,-0.996874988079071,-1.03007781505585,-1.03916037082672,-1.03125011920929,-1.09980452060699,-1.11835920810699,-1.10302746295929,-1.18740236759186,-1.25078117847443,-1.34785175323486,-1.33066403865814,-1.35625004768372,-1.56738269329071,-1.71728515625,-1.69658184051514,-1.62949204444885,-1.50703144073486,-1.46640634536743,-1.51347661018372,-1.58886694908142,-1.67167937755585,-1.72480463981628,-1.73349583148956,-1.79882788658142,-1.79228544235229,-1.74580073356628,-1.84179675579071,-1.94628882408142,-2.05273413658142,-2.11386704444885,-2.15214872360229,-2.2275390625,-2.24111366271973,-2.26552724838257,-2.32041025161743,-2.37324213981628,-2.23046851158142,-2.19033241271973,-2.16806650161743,-2.26962924003601,-2.36552739143372,-2.40546894073486,-2.48124980926514,-2.52421903610229,-2.57343769073486,-2.67685532569885,-2.76249980926514,-3.14228510856628,-3.20478487014771,-3.13789057731628,-2.94443345069885,-2.73837900161743,-2.53515601158142,-2.47128891944885,-2.47119164466858,-2.49345684051514,-2.56005835533142,-2.59853506088257,-2.69960927963257,-2.75498056411743,-2.80957007408142,-2.64218735694885,-2.58349609375,-2.59540987014771,-2.51816368103027,-2.50205039978027,-2.41367197036743,-2.37607431411743,-2.38603520393372,-2.46503925323486,-2.52958989143372,-2.58964824676514,-2.66103482246399,-2.79199242591858,-2.80605459213257,-2.74658203125,-2.80888676643372,-2.87861323356628,-2.91650414466858,-2.93623042106628,-2.88613271713257,-2.86962890625,-2.79560542106628,-2.81318378448486,-2.86152386665344,-2.98584008216858,-3.05625033378601,-3.12558603286743,-3.19736361503601,-3.39023423194885,-3.50175762176514,-3.65371108055115,-3.71748018264771,-3.87646436691284,-3.94804692268372,-4.21640586853027,-4.404296875,-4.59208965301514,-4.71308612823486,-4.91240262985229,-4.93671894073486,-4.96660137176514,-5.05068349838257,-5.09755849838257,-5.08427762985229,-5.09375,-5.05439472198486,-5.12939453125,-5.166015625,-5.25087928771973,-5.56669950485229,-5.91718721389771,-6.18535137176514,-6.39374971389771,-6.78505849838257,-6.908203125,-7.0029296875,-7.02441358566284,-7.28837871551514,-7.39482402801514,-7.53330087661743,-7.59502029418945,-7.63427734375,-7.69209003448486,-7.74746036529541,-7.87177658081055,-7.97148466110229,-8.09218788146973,-8.40761661529541,-8.93056583404541,-9.23066425323486,-9.54062461853027,-9.70253944396973,-9.71904373168945,-9.68701171875,-9.7724609375,-9.84765625,-10.0757818222046,-10.22509765625,-10.4840812683105,-10.4899415969849,-10.5899419784546,-10.6716794967651,-10.8204097747803,-11.0547847747803,-11.0849618911743,-11.0684576034546,-11.1875,-11.3759775161743,-11.4039058685303,-11.3504877090454,-11.3098630905151,-11.2666015625,-11.2151365280151,-11.2525396347046,-11.3937501907349,-11.4972658157349,-11.6536130905151,-12.1000003814697,-12.4754886627197,-12.59130859375,-12.84423828125,-12.9662113189697,-12.9670896530151,-12.9566411972046,-12.7623052597046,-12.6446294784546,-12.6239261627197,-12.74853515625,-12.78271484375,-12.7901372909546,-12.8444337844849,-12.9072265625,-13.0329103469849,-13.1471681594849,-13.2730464935303,-13.3651361465454,-13.48046875,-13.5881834030151,-13.5587882995605,-13.5814456939697,-13.6150388717651,-13.66455078125,-13.7581052780151,-13.9910154342651,-14.0439453125,-14.1011724472046,-14.00341796875,-14.0306644439697,-14.65478515625,-14.9356451034546,-15.2538089752197,-15.5643548965454,-15.8419923782349,-15.8642587661743,-16.1865234375,-16.50439453125,-16.7635746002197,-17.0435562133789,-17.1781253814697,-17.3158206939697,-17.64208984375,-17.7039070129395,-17.8494129180908,-17.9200191497803,-17.9901351928711,-18.2523441314697,-18.6398448944092,-18.8459968566895,-19.27783203125,-19.4539070129395,-19.57177734375,-19.6491203308105,-19.7419929504395,-19.96826171875,-20.2060546875,-20.2926769256592,-20.42578125,-20.5694332122803,-20.7837886810303,-20.84619140625,-20.9060554504395,-21.0313472747803,-21.2378902435303,-21.5056648254395,-21.5968761444092,-21.61083984375,-21.9203128814697,-21.9990234375,-22.0843753814697,-22.24365234375,-22.3096675872803,-22.5806655883789,-22.6446285247803,-22.73583984375,-22.7882804870605,-22.8451156616211,-22.9470710754395,-22.9408206939697,-22.9410152435303,-22.9733390808105,-22.9670906066895,-22.9425773620605,-22.9025402069092,-22.85009765625,-22.7707023620605,-22.7233390808105,-22.7251949310303,-22.7476558685303,-22.7951183319092,-22.8288078308105,-22.8781242370605,-22.9385738372803,-22.9912109375,-22.998046875,-23.04638671875,-23.0666007995605,-23.1014652252197,-23.0573234558105,-23.0352535247803,-23.0459957122803,-23.0094738006592,-22.96630859375,-22.9105472564697,-22.9447269439697,-23.0110359191895,-23.0049800872803,-23.0554695129395,-23.10693359375,-23.2066402435303,-23.228515625,-23.2740230560303,-23.31640625,-23.3351573944092,-23.3814449310303,-23.5755863189697,-23.5997066497803,-23.6853504180908,-23.7584953308105,-23.8025398254395,-23.8047847747803,-23.7648448944092,-23.763671875,-23.7955074310303,-24.1103515625,-24.236328125,-24.4931640625,-24.7810554504395,-24.9529304504395,-24.9974613189697,-24.9999027252197,-25.0357418060303,-25.0654296875,-25.0681629180908,-25.1682624816895,-25.2367191314697,-25.41650390625,-25.4033203125,-25.3098640441895,-25.3063468933105,-25.2720699310303,-25.3107433319092,-25.4033203125,-25.4429683685303,-25.4474620819092,-25.4365234375,-25.3687496185303,-25.4915046691895,-25.5212879180908,-25.5501956939697,-25.5973625183105,-25.81591796875,-25.8443355560303,-25.8751945495605,-25.875,-25.9354496002197,-26.1793937683105,-26.2257823944092,-26.2269535064697,-26.2686519622803,-26.3483390808105,-26.4064445495605,-26.5191402435303,-26.6124019622803,-26.7029304504395,-26.8781242370605,-27.0580081939697,-27.1234378814697,-27.1959972381592,-27.2638683319092,-27.3727531433105,-27.5579109191895,-27.8251953125,-28.0755863189697,-28.2072257995605,-28.3101558685303,-28.4426746368408,-28.5752925872803,-28.6986312866211,-28.8711929321289,-29.0753917694092,-29.3631839752197,-29.8009757995605,-30.42578125,-30.8976554870605,-31.0680656433105,-31.2583980560303,-31.4803733825684,-31.7024421691895,-31.9002933502197,-31.9895496368408,-32.1148414611816,-32.0630874633789,-31.9775390625,-31.9134750366211,-31.8303718566895,-31.8150386810303,-31.8677749633789,-31.83203125,-31.7966785430908,-31.7746105194092,-31.5573253631592,-31.4769535064697,-31.3397464752197,-31.2667961120605,-31.1188468933105,-31.0813465118408,-31.09423828125,-31.0521469116211,-31.0054702758789,-30.9037094116211,-30.8133811950684,-30.7042007446289,-30.4259757995605,-30.4134769439697,-30.4568347930908,-30.4388675689697,-30.31689453125,-30.2536125183105,-30.23681640625,-30.3743152618408,-30.3686542510986,-30.2606449127197,-30.2110366821289,-30.1213874816895,-30.0599594116211,-30.0348625183105,-30.1410160064697,-30.2441387176514,-30.3642578125,-30.4119129180908,-30.4675788879395,-30.5912113189697,-30.7515640258789,-30.7027359008789,-30.6745128631592,-30.8468761444092,-30.9127941131592,-30.9775371551514,-31.0526351928711,-31.1044940948486,-31.2437515258789,-31.3388690948486,-31.3837909698486,-31.4899425506592,-31.5990238189697,-31.6949234008789,-31.8855457305908,-31.9675769805908,-32.0884780883789,-32.1677742004395,-32.2207984924316,-32.32373046875,-32.4397468566895,-32.8752937316895,-33.1377944946289,-33.2664070129395,-33.4019546508789,-33.7421875,-33.7373046875,-33.7098617553711,-33.67724609375,-33.6554718017578,-33.6228523254395,-33.5002899169922,-33.1708984375,-33.1086921691895,-33.0685539245605,-33.0103530883789,-32.9270515441895,-32.8210945129395,-32.7367172241211,-32.6800765991211,-32.6253929138184,-32.5811538696289,-32.5032234191895,-32.4030303955078,-32.2987327575684,-32.1863288879395,-32.0974578857422,-32.0568389892578,-32.0399436950684,-31.9945316314697,-31.9523429870605,-31.9281253814697,-31.9015636444092,-31.8551750183105,-31.7450180053711,-31.6227550506592,-31.5419902801514,-31.4851570129395,-31.3912105560303,-31.2790050506592,-31.31396484375,-31.2795886993408,-31.2255859375,-31.1841773986816,-31.1417007446289,-31.0929679870605,-31.0461902618408,-30.9644546508789,-30.8759765625,-30.8507823944092,-30.8581047058105,-30.8920879364014,-30.9249038696289,-30.9465827941895,-30.9871101379395,-31.0367183685303,-31.0696296691895,-31.0808582305908,-31.0791988372803,-31.0596675872803,-30.9918937683105,-30.8372077941895,-30.7776374816895,-30.7137699127197,-30.6284198760986,-30.4474601745605,-30.1869125366211,-30.1072273254395,-30.10107421875,-30.1099605560303,-30.1444358825684,-30.2648448944092,-30.2833995819092,-30.2806644439697,-30.2612323760986,-30.1877937316895,-30.1399402618408,-30.0338859558105,-29.9394550323486,-29.8565425872803,-29.7821273803711,-29.7162094116211,-29.5948238372803,-29.4178714752197,-29.2873039245605,-29.2030277252197,-29.1380863189697,-29.0924797058105,-28.9972648620605,-28.8524398803711,-28.7372074127197,-28.6517581939697,-28.5808601379395,-28.5246105194092,-28.4885730743408,-28.4728527069092,-28.4432621002197,-28.3996086120605,-28.3700180053711,-28.3541984558105,-28.3597660064697,-28.3866214752197,-28.3816413879395,-28.3449211120605,-28.3028316497803,-28.2554683685303,-28.2041015625,-28.1209945678711,-28.08935546875,-28.0377941131592,-27.9559574127197,-27.8988285064697,-27.8667984008789,-27.8359375,-27.7962875366211,-27.7677745819092,-27.7471694946289,-27.7085933685303,-27.6519527435303,-27.5992183685303,-27.5505847930908,-27.5325202941895,-27.544921875,-27.5265617370605,-27.4771499633789,-27.4541015625,-27.4573249816895,-27.4464855194092,-27.4235363006592,-27.3820323944092,-27.2896480560303,-27.2538089752197,-27.2747077941895,-27.2437496185303,-27.1611328125,-27.1595706939697,-27.12109375,-26.9783191680908,-26.8828125,-26.8046875,-26.7486324310303,-26.66650390625,-26.4431648254395,-26.3518543243408,-26.2881832122803,-26.22509765625,-26.0836925506592,-25.9595699310303,-25.7488269805908,-25.6688480377197,-25.6476554870605,-25.5779304504395,-25.5718746185303,-25.5452156066895,-25.5230484008789,-25.5295886993408,-25.5704097747803,-25.5718746185303,-25.5886726379395,-25.625,-25.6082992553711,-25.5764656066895,-25.5760746002197,-25.4327144622803,-25.22021484375,-25.1212882995605,-25.0652351379395,-24.8674812316895,-24.5281238555908,-24.3060550689697,-24.2012691497803,-24.1281242370605,-24.0658206939697,-24.0472660064697,-23.97119140625,-23.9017562866211,-23.8521480560303,-23.8125,-23.8290042877197,-23.8521480560303,-23.8884773254395,-23.9513683319092,-23.9745121002197,-23.9976558685303,-24.0174808502197,-24.0042972564697,-23.9910163879395,-23.9513683319092,-23.8653316497803,-23.7925777435303,-23.6867179870605,-23.6272468566895,-23.5809574127197,-23.5247058868408,-23.4619140625,-23.4156246185303,-23.359375,-23.3196277618408,-23.2501964569092,-23.154296875,-23.0947265625,-23.0252933502197,-22.9558601379395,-22.8864250183105,-22.8103523254395,-22.7409191131592,-22.6714839935303,-22.6218757629395,-22.5920906066895,-22.5126953125,-22.41015625,-22.3539066314697,-22.3076171875,-22.3076171875,-22.2811527252197,-22.2844715118408,-22.2811527252197,-22.2646484375,-22.2282218933105,-22.1786136627197,-22.0926761627197,-22.076171875,-22.1025390625,-22.1356449127197,-22.1819343566895,-22.23486328125,-22.2315425872803,-22.2613277435303,-22.2646484375,-22.2712879180908,-22.2448253631592,-22.2150402069092,-22.1952152252197,-22.2150402069092,-22.1984367370605,-22.1885738372803,-22.1819343566895,-22.1290035247803,-22.0992183685303,-22.1091785430908,-22.1422843933105,-22.1356449127197,-22.1091785430908,-22.04638671875,-22.0066413879395,-21.9669914245605,-21.9107418060303,-21.8511714935303,-21.79833984375,-21.751953125,-21.6991214752197,-21.6495132446289,-21.5965824127197,-21.5469722747803,-21.4940433502197,-21.41796875,-21.3550777435303,-21.30224609375,-21.2658195495605,-21.2062492370605,-21.1335945129395,-20.9979496002197,-20.9185543060303,-20.8970699310303,-20.873046875,-20.8416996002197,-20.8093757629395,-20.7763671875,-20.7474613189697,-20.6903324127197,-20.673828125,-20.6573257446289,-20.5944347381592,-20.5216789245605,-20.4654293060303,-20.4158191680908,-20.3861312866211,-20.3332042694092,-20.29345703125,-20.2373046875,-20.1646480560303,-20.1510753631592,-20.1103515625,-20.05517578125,-20.0407218933105,-20.0204105377197,-19.9795913696289,-19.8327140808105,-19.7445316314697,-19.6252937316895,-19.4242172241211,-19.2294921875,-19.0809574127197,-19.0535163879395,-19.0440425872803,-18.9673824310303,-18.9171867370605,-18.9142570495605,-18.7332019805908,-18.4750003814697,-18.279296875,-18.2464847564697,-18.2373046875,-18.2146492004395,-18.18310546875,-18.1222667694092,-17.9473628997803,-17.6717777252197,-17.5730476379395,-17.5121097564697,-17.5323238372803,-17.5128898620605,-17.3630847930908,-17.2821292877197,-17.2342758178711,-17.08056640625,-16.9107418060303,-16.70068359375,-16.6501941680908,-16.4908199310303,-16.4102535247803,-16.3399410247803,-16.2843742370605,-16.2835922241211,-16.3079090118408,-16.32666015625,-16.3282222747803,-16.3131847381592,-16.2959957122803,-16.28173828125,-16.2693367004395,-16.1321277618408,-15.9019527435303,-15.7386713027954,-15.4795904159546,-15.3182611465454,-15.1431646347046,-15.0983400344849,-15.0927734375,-15.0887689590454,-14.6185550689697,-14.5705080032349,-14.4187498092651,-14.3328123092651,-14.2630863189697,-14.1847648620605,-14.1324224472046,-14.1000003814697,-14.0192384719849,-13.9379892349243,-13.8624029159546,-13.7898435592651,-13.7453126907349,-13.6643552780151,-13.5614261627197,-13.48974609375,-13.49853515625,-13.5265617370605,-13.5412111282349,-13.5248041152954,-13.5255861282349,-13.4704103469849,-13.40625,-13.2419919967651,-13.1597652435303,-13.1336917877197,-13.1436519622803,-13.1324214935303,-13.0642576217651,-12.9943361282349,-12.9972658157349,-12.9537105560303,-12.8470697402954,-12.8055667877197,-12.7503900527954,-12.6691408157349,-12.6516599655151,-12.6662111282349,-12.7079105377197,-12.6800775527954,-12.6051759719849,-12.5466794967651,-12.5189456939697,-12.47802734375,-12.4694328308105,-12.5296878814697,-12.5050783157349,-12.4833011627197,-12.4397459030151,-12.326171875,-12.2509765625,-12.2039060592651,-12.146484375,-12.0593748092651,-12.0302734375,-12.0059566497803,-11.9751958847046,-11.9200201034546,-11.8473634719849,-11.8293943405151,-11.7412109375,-11.7350578308105,-11.7523431777954,-11.76513671875,-11.74951171875,-11.7100591659546,-11.6468753814697,-11.58056640625,-11.5110349655151,-11.4391593933105,-11.36474609375,-11.3150386810303,-11.2899417877197,-11.2462892532349,-11.1842765808105,-11.1103515625,-11.0248050689697,-10.8927736282349,-10.7147464752197,-10.5862302780151,-10.5074224472046,-10.4490232467651,-10.3922843933105,-10.2530279159546,-10.1467771530151,-10.0269536972046,-9.93554782867432,-9.87265682220459,-9.79023456573486,-9.71240234375,-9.71044921875,-9.73173809051514,-9.79746055603027,-9.80908203125,-9.76845741271973,-9.78544902801514,-9.82607364654541,-9.8681640625,-9.88613319396973,-9.89990234375,-9.97548866271973,-10.2689456939697,-10.3114252090454,-10.3172845840454,-10.35791015625,-10.3898439407349,-10.5059566497803,-10.5989255905151,-10.6831045150757,-10.68603515625,-10.6627931594849,-10.6744146347046,-10.703125,-10.7850580215454,-10.9331064224243,-10.9751949310303,-11.0187501907349,-11.0547847747803,-11.1091794967651,-11.11279296875,-11.1224613189697,-11.09765625,-11.0446290969849,-11.0111322402954,-10.9943361282349,-10.9556646347046,-10.9481439590454,-10.9517574310303,-10.9541015625,-10.9333982467651,-10.9298830032349,-10.982421875,-11.0476570129395,-11.0642576217651,-11.0666990280151,-11.05859375,-11.0248050689697,-10.9468746185303,-10.9768562316895,-11.01025390625,-10.8408203125,-10.5860347747803,-10.361328125,-10.1815423965454,-9.97177696228027,-9.82373046875,-9.76748085021973,-9.70458984375,-9.62050724029541,-9.54345703125,-9.48984336853027,-9.4375,-9.46367168426514,-9.47822284698486,-9.51796817779541,-9.57168006896973,-9.62529277801514,-9.66904258728027,-9.76572227478027,-9.81875038146973,-9.85244083404541,-9.96601581573486,-9.98857498168945,-10.0060548782349,-10.0055665969849,-10.0051765441895,-10.0037107467651,-9.91015625,-9.84404277801514,-9.77431678771973,-9.6884765625,-9.62919902801514,-9.556640625,-9.51015663146973,-9.4921875,-9.45205116271973,-9.41035175323486,-9.40742206573486,-9.41142654418945,-9.26572322845459,-9.1201171875,-8.9931640625,-8.8828125,-8.81406211853027,-8.71933555603027,-8.65400409698486,-8.56699180603027,-8.51416015625,-8.47929763793945,-8.458984375,-8.42705059051514,-8.39218807220459,-8.34580039978027,-8.29931640625,-8.24990177154541,-8.19189453125,-8.14541053771973,-8.07285118103027,-8.02060508728027,-7.98574209213257,-7.93642520904541,-7.89570283889771,-7.87539100646973,-7.82900428771973,-7.78251981735229,-7.75351572036743,-7.73896455764771,-7.65478515625,-7.61123085021973,-7.58505916595459,-7.55605459213257,-7.53574180603027,-7.50663995742798,-7.46025371551514,-7.41669940948486,-7.37890625,-7.36728525161743,-7.37314462661743,-7.34990262985229,-7.34121131896973,-7.33535146713257,-7.30927753448486,-7.26279306411743,-7.17275381088257,-7.13505840301514,-7.07988309860229,-6.97353506088257,-6.90576171875,-6.83378887176514,-6.67949199676514,-6.57470703125,-6.56406259536743,-6.52519512176514,-6.46581983566284,-6.40087890625,-6.34433603286743,-6.26064443588257,-6.15644502639771,-6.09843730926514,-6.02871131896973,-5.93339872360229,-5.78955030441284,-5.63486337661743,-5.58964824676514,-5.49521446228027,-5.30253934860229,-5.1982421875,-5.15771484375,-5.12275362014771,-5.09375,-5.06718778610229,-5.00957059860229,-4.90126943588257,-4.78603506088257,-4.74892616271973,-4.64228487014771,-4.57460975646973,-4.55332040786743,-4.50439500808716,-4.4873046875,-4.46972703933716,-4.43759775161743,-4.42431640625,-4.38818359375,-4.38720703125,-4.35048818588257,-4.29531240463257,-4.22958993911743,-4.17333984375,-4.15888690948486,-4.16865205764771,-4.16757822036743,-4.15009784698486,-4.19365262985229,-4.24697256088257,-4.30117177963257,-4.29814434051514,-4.28662109375,-4.33310556411743,-4.32724571228027,-4.30117177963257,-4.23593759536743,-4.20058631896973,-3.99658226966858,-3.65986299514771,-3.35458993911743,-3.01669907569885,-2.66767597198486,-2.31425786018372,-2.02421855926514,-1.77490234375,-1.62197244167328,-1.42167961597443,-1.24570298194885,-1.19492185115814,-1.15224623680115,-1.09160149097443,-1.06494116783142,-1.02958965301514,-0.998730778694153,-0.965722680091858,-0.945800960063934,-0.917187392711639,-0.877441346645355,-0.837793171405792,-0.795898497104645,-0.762793004512787,-0.720898330211639,-0.681250274181366,-0.639355599880219,-0.59960949420929,-0.553320229053497,-0.509277522563934,-0.482421875,-0.452539265155792,-0.38134753704071,-0.317480504512787,-0.196191638708115,-0.138867199420929,0.0185546651482582,0.189355686306953,0.447363138198853,0.578613460063934,0.585839629173279,0.589404582977295,0.598486542701721,0.607470452785492,0.626367211341858,0.649804592132568,0.665087699890137,0.65966808795929,0.680371344089508,0.700195074081421,0.716406404972076,0.729931592941284,0.698388576507568,0.666894793510437,0.651562452316284,0.655175566673279,0.652441382408142,0.627246081829071,0.625439584255219,0.629931569099426,0.635351777076721,0.642529189586639,0.658789157867432,0.686669886112213,0.712841987609863,0.753320336341858,0.801953017711639,0.86406272649765,0.898291230201721,0.963134586811066,1.01538097858429,1.05048835277557,1.06401360034943,1.04238271713257,1.038818359375,1.05859398841858,1.05947291851044,1.06577146053314,1.0732421875,1.05908203125,1.07661128044128,1.07841765880585,1.05952155590057,1.30878913402557,1.54389667510986,1.70873999595642,1.70517575740814,1.73486351966858,1.73945295810699,1.77075183391571,1.77324199676514,1.75791013240814,1.72578096389771,1.72124016284943,1.72128903865814,1.72138690948486,1.72148442268372,1.72158169746399,1.7216796875,1.71982443332672,1.77456033229828,1.84550797939301,1.90136730670929,1.95761740207672,1.98701190948486,1.95576167106628,1.860107421875,1.78803682327271,1.75253927707672,1.74848616123199,1.76059556007385,1.79008793830872,1.922119140625,2.03505849838257,2.072998046875,2.10790991783142,2.12114262580872,2.11669921875,2.08583974838257,2.03208017349243,1.84482443332672,1.70361316204071,1.61557602882385,1.40058624744415,1.21000957489014,1.18540036678314,1.17836904525757,1.22304654121399,0.992138862609863,0.821679949760437,0.767187595367432,0.75195324420929,0.763281226158142,0.785351514816284,0.809765815734863,0.863134562969208,0.937255859375,0.97802746295929,0.983447134494781,0.970361351966858,0.926073968410492,0.843408167362213,0.747509956359863,0.68798828125,0.691259801387787,0.790478348731995,0.868652164936066,0.93188488483429,1.02221691608429,1.10810554027557,1.15844738483429,1.21972632408142,1.25712895393372,1.25332045555115,1.29384756088257,1.36987328529358,1.43100559711456,1.45278310775757,1.44687485694885,1.45527327060699,1.52949190139771,1.61928701400757,1.7705078125,1.90444350242615,1.93159198760986,1.95302748680115,1.96699225902557,1.97670900821686,2.04814481735229,2.12050747871399,2.13603544235229,2.15556621551514,2.22250962257385,2.34042930603027,2.41191387176514,2.43393564224243,2.43403315544128,2.45244145393372,2.48188471794128,2.50239253044128,2.52509760856628,2.57607436180115,2.67187523841858,2.801513671875,3.0048828125,3.20468759536743,3.343994140625,3.49121117591858,3.58740234375,3.66269540786743,3.89980459213257,4.01181650161743,4.08930683135986,4.23227548599243,4.27602529525757,4.2744140625,4.23710966110229,4.15771436691284,4.13989305496216,4.13999032974243,4.14033222198486,4.12685585021973,4.10014677047729,4.06699180603027,3.97441411018372,3.92910170555115,3.9306640625,3.93256855010986,3.94082021713257,3.91503882408142,3.89370131492615,3.94287109375,3.94389629364014,3.92226552963257,3.75644516944885,3.68647456169128,3.59394526481628,3.59345722198486,3.67294955253601,3.94033217430115,4.01792001724243,4.03964900970459,4.04228544235229,4.08432626724243,4.13852500915527,4.15673828125,4.098388671875,4.12631845474243,4.197021484375,4.28779268264771,4.40224599838257,4.43300771713257,4.51689481735229,4.508056640625,4.50468778610229,4.51933574676514,4.53525352478027,4.57470703125,4.68681669235229,4.72919940948486,4.77412080764771,4.82709980010986,4.89252901077271,4.94936513900757,4.99458026885986,5.08198261260986,5.16435575485229,5.19155311584473,5.20205068588257,5.22114276885986,5.19248056411743,5.18808555603027,5.21015644073486,5.19931650161743,5.25795888900757,5.23881816864014,5.23881816864014,5.19423866271973,5.14399385452271,5.08286142349243,4.98984384536743,4.90751886367798,4.81269550323486,4.74052715301514,4.66665029525757,4.59765625,4.56962871551514,4.53325176239014,4.51118135452271,4.50458955764771,4.501708984375,4.48032236099243,4.47592782974243,4.41665029525757,4.381103515625,4.353515625,4.28764677047729,4.22675800323486,4.18818330764771,4.160400390625,4.02314472198486,3.97539091110229,3.96000957489014,3.93354487419128,3.88344717025757,3.81967735290527,3.75273442268372,3.69980454444885,3.66655278205872,3.58750009536743,3.462158203125,3.39858388900757,3.34921836853027,3.28310537338257,3.08784198760986,2.99047875404358,2.76542949676514,2.68999028205872,2.58837866783142,2.36293911933899,2.32705092430115,2.27412104606628,2.12163066864014,1.96240210533142,1.90063464641571,1.87417018413544,1.86147451400757,1.84233379364014,1.79521489143372,1.74628889560699,1.71801745891571,1.70000016689301,1.63242208957672,1.52734398841858,1.50820314884186,1.46459996700287,1.37602519989014,1.34365248680115,1.30458998680115,1.24887669086456,1.20361304283142,1.20122087001801,1.20849633216858,1.24750959873199,1.28105461597443,1.27915060520172,1.28466820716858,1.31225621700287,1.34775376319885,1.43867182731628,1.46625971794128,1.48173809051514,1.53022468090057,1.55668962001801,1.58754849433899,1.59194314479828,1.57431614398956,1.56328094005585,1.54785180091858,1.51699233055115,1.51435554027557,1.52026379108429,1.53994154930115,1.57431614398956,1.64843738079071,1.65058600902557,1.66728544235229,1.70000016689301,1.70478487014771,1.70410132408142,1.72607433795929,1.77382802963257,1.90893542766571,1.94013655185699,1.96347665786743,1.95922863483429,1.98159182071686,2.01396489143372,2.00581049919128,1.93647480010986,1.92124044895172,1.91640639305115,1.88125014305115,1.89218735694885,1.91430652141571,1.92265617847443,1.90722632408142,1.92724609375,1.94213843345642],[13.0819826126099,13.0622062683105,13.1021003723145,13.1502933502197,13.3031253814697,13.3176755905151,13.1968259811401,13.1527833938599,13.0819826126099],[4.89970684051514,4.89975595474243,4.86669921875,4.75058603286743,4.63398408889771,4.45634746551514,4.38076210021973,4.36528253555298,4.35258817672729,4.34721708297729,4.36420917510986,4.39042949676514,4.58266592025757,4.69135713577271,4.82114219665527,4.89970684051514],[4.89970684051514,4.85624980926514,4.80175733566284,4.75483417510986,4.71806621551514,4.66650390625,4.55302762985229,4.46391582489014,4.42876005172729,4.39321327209473,4.3544921875,4.28076171875,4.26650381088257,4.16879892349243,4.09653329849243,4.03764677047729,4.02397489547729,4.049072265625,4.11357402801514,4.20356416702271,4.255859375,4.26279258728027,4.30419969558716,4.354736328125,4.41425800323486,4.47788047790527,4.52695322036743,4.56523418426514,4.59267568588257,4.59096717834473,4.607177734375,4.660400390625,4.72456073760986,4.79814481735229,4.88100576400757,4.94638633728027,5.02236366271973,5.016357421875,4.96245098114014,4.89970684051514],[27.7599620819092,27.7373046875,27.677001953125,27.6114253997803,27.5576667785645,27.4936027526855,27.4425277709961,27.4386234283447,27.4583015441895,27.4501953125,27.3646965026855,27.2906246185303,27.2143058776855,27.1473636627197,27.0999031066895,27.0792980194092,27.0408191680908,26.9751949310303,26.9148445129395,26.8748531341553,26.85498046875,26.86083984375,26.8600578308105,26.8529796600342,26.8307628631592,26.802001953125,26.8073253631592,26.8668937683105,26.8670902252197,26.7899417877197,26.8034172058105,26.7777347564697,26.7716789245605,26.7802238464355,26.7965831756592,26.8507823944092,26.8903312683105,26.8541507720947,26.8475093841553,26.7545909881592,26.7239265441895,26.7015628814697,26.7139167785645,26.7194328308105,26.7411117553711,26.76220703125,26.7789554595947,26.7962417602539,26.8034172058105,26.8265609741211,26.8486328125,26.816162109375,26.8650398254395,26.9322261810303,26.9614753723145,27.0655765533447,27.0990238189697,27.1342296600342,27.1755867004395,27.2181167602539,27.2974605560303,27.3160648345947,27.4640121459961,27.5178699493408,27.5925769805908,27.7112789154053,27.8331546783447,27.9581546783447,28.0599613189697,28.107421875,28.1583003997803,28.1881847381592,28.2562999725342,28.2941398620605,28.3111820220947,28.3020496368408,28.2777347564697,28.2439460754395,28.2165031433105,28.1681652069092,28.119140625,28.093994140625,28.0802249908447,28.0708503723145,28.0785636901855,28.0717296600342,28.0265140533447,27.9945812225342,27.9700679779053,27.9744625091553,28.0267581939697,28.0712413787842,28.078369140625,28.0640144348145,28.0216312408447,27.9817867279053,27.9517097473145,27.9232425689697,27.8008785247803,27.7599620819092],[-17.7935543060303,-17.8430671691895,-17.9152336120605,-17.9690418243408,-18.0412101745605,-18.1044921875,-18.1419906616211,-18.2349605560303,-18.3512706756592,-18.4417972564697,-18.6492176055908,-18.7235355377197,-18.7970695495605,-18.9386711120605,-18.9856452941895,-19.0817394256592,-19.3699207305908,-19.5382823944092,-19.5693359375,-19.7486324310303,-19.8927745819092,-19.9901371002197,-20.05419921875,-20.1009769439697,-20.1457996368408,-20.2320308685303,-20.3818359375,-20.4787101745605,-20.4748058319092,-20.4835929870605,-20.5030269622803,-20.5306644439697,-20.5945320129395,-20.6897468566895,-20.7664070129395,-20.8483390808105,-20.94482421875,-21.0642566680908,-21.1110363006592,-21.2615242004395,-21.3590831756592,-21.5067367553711,-21.55419921875,-21.5730457305908,-21.58935546875,-21.6512699127197,-21.7076168060303,-21.7660160064697,-21.7814464569092,-21.796875,-21.8113288879395,-21.93994140625,-21.9812488555908,-22.0183601379395,-22.0474605560303,-22.0657215118408,-22.0794925689697,-22.15771484375,-22.1939449310303,-22.2132816314697,-22.2784194946289,-22.3951187133789,-22.4808597564697,-22.5354499816895,-22.5729484558105,-22.5933589935303,-22.6936511993408,-22.8737297058105,-22.9870109558105,-23.0335941314697,-23.0739269256592,-23.1080074310303,-23.14892578125,-23.19677734375,-23.2196292877197,-23.2176761627197,-23.2526359558105,-23.3246097564697,-23.3683586120605,-23.3835926055908,-23.4242191314697,-23.4900379180908,-23.5234375,-23.5244140625,-23.5779285430908,-23.70458984375,-23.7634754180908,-24.2408218383789,-24.2971687316895,-24.3955078125,-24.5132808685303,-24.5827140808105,-24.6135730743408,-24.6714839935303,-24.7024421691895,-24.7474613189697,-24.7879867553711,-24.9352531433105,-25.146484375,-25.3023433685303,-25.4378910064697,-25.6062488555908,-25.6627922058105,-25.7144527435303,-25.7399425506592,-25.7562503814697,-25.75146484375,-25.7540035247803,-25.8134765625,-25.8173828125,-25.7831058502197,-25.7498054504395,-25.7428703308105,-25.6329097747803,-25.6348628997803,-25.6260738372803,-25.6008796691895,-25.5446300506592,-25.4339847564697,-25.3444328308105,-25.2914066314697,-25.2666015625,-25.2886714935303,-25.3123054504395,-25.3241214752197,-25.3703117370605,-25.4579105377197,-25.5951175689697,-25.6791019439697,-25.8573246002197,-26.0711898803711,-26.1327152252197,-26.1784172058105,-26.2190437316895,-26.3888683319092,-26.5801753997803,-26.6358394622803,-26.6619148254395,-26.6783218383789,-26.7100601196289,-26.8068370819092,-26.8409175872803,-26.8541984558105,-26.8426761627197,-26.8328132629395,-26.8517589569092,-26.8210945129395,-26.8087902069092,-26.8488273620605,-26.8224620819092,-26.7421875,-26.5808601379395,-26.44384765625,-26.3401355743408,-26.26416015625,-26.1649417877197,-26.12060546875,-26.08056640625,-25.9990234375,-25.9156246185303,-25.7332038879395,-25.4912109375,-25.2212905883789,-25.1470699310303,-25.0298824310303,-24.8070316314697,-24.7767562866211,-24.751953125,-24.5357418060303,-24.2490234375,-23.96240234375,-23.67578125,-23.38916015625,-23.1025390625,-22.81591796875,-22.529296875,-22.2425785064697,-22.0001945495605,-22.0001945495605,-22.0001945495605,-22.0001945495605,-22.0001945495605,-21.9619140625,-21.7840824127197,-21.3760738372803,-20.9681625366211,-20.5602550506592,-20.15234375,-19.7443351745605,-19.33642578125,-18.9285144805908,-18.5205078125,-18.31884765625,-18.3068370819092,-18.265625,-18.1986331939697,-18.11572265625,-18.0671882629395,-18.0095691680908,-17.9997062683105,-18.0075206756592,-18.02734375,-18.2310543060303,-18.3864250183105,-18.4529285430908,-18.4599609375,-18.4494132995605,-18.42431640625,-18.26953125,-18.2292003631592,-18.1541023254395,-18.0775394439697,-18.0234375,-17.9782218933105,-17.9894523620605,-18.0285167694092,-18.052734375,-17.8646488189697,-17.8213882446289,-17.7875995635986,-17.7935543060303],[10.919677734375,10.7953128814697,10.7518072128296,10.4578123092651,10.3879404067993,10.1742677688599,10.0300779342651,9.82290077209473,9.71035194396973,9.61391639709473,9.34062480926514,9.26191425323486,9.11474609375,9.04848575592041,8.99331092834473,8.97060585021973,8.94321346282959,8.81581974029541,8.76582050323486,8.73247146606445,8.709716796875,8.69130802154541,8.66562461853027,8.65336894989014,8.60825252532959,8.46113300323486,8.369140625,8.29140663146973,8.25844764709473,8.23945331573486,8.19155216217041,8.13754844665527,7.96484422683716,7.80791044235229,7.72456073760986,7.64897441864014,7.55722713470459,7.51933574676514,7.427490234375,7.33339834213257,7.22871112823486,7.06494140625,6.95522451400757,6.87211942672729,6.78173828125,6.69902372360229,6.63530302047729,6.455322265625,6.39623975753784,6.34492206573486,6.274169921875,6.18300724029541,6.06923818588257,6.01752948760986,5.99824237823486,5.94550800323486,5.85493135452271,5.77685594558716,5.72294950485229,5.67514657974243,5.61879873275757,5.5625,5.44077157974243,5.28964853286743,5.18632793426514,5.10917997360229,5.19785118103027,5.19975614547729,5.18437480926514,5.07568359375,5.06240272521973,5.07192373275757,5.085205078125,5.17114210128784,5.25371074676514,5.28369140625,5.31210947036743,5.25590801239014,5.06269502639771,5.02456092834473,4.96743202209473,4.98295879364014,4.93007850646973,5.00996065139771,4.994140625,4.953857421875,4.86626005172729,4.81635713577271,4.77080059051514,4.70126962661743,4.66313505172729,4.66352510452271,4.70297861099243,4.73691415786743,4.74384784698486,4.723876953125,4.64667987823486,4.59174776077271,4.445556640625,4.20766639709473,4.15976572036743,4.15512704849243,4.13496065139771,4.25390625,4.28139686584473,4.24482440948486,4.31137657165527,4.32309579849243,4.30219745635986,4.33217811584473,4.41313457489014,4.44731473922729,4.43564414978027,4.46269512176514,4.54155302047729,4.68618154525757,4.82963848114014,4.94472646713257,5.03129863739014,5.08930683135986,5.12138700485229,5.12749052047729,5.07075214385986,4.89140605926514,4.5234375,4.38261747360229,4.34624004364014,4.257568359375,4.11660146713257,3.95429706573486,3.6787109375,3.47841811180115,3.51020550727844,3.60410165786743,3.62299776077271,3.58081030845642,3.54267597198486,3.50541996955872,3.49980425834656,3.55107450485229,3.56030297279358,3.55083036422729,3.5517578125,3.55839848518372,3.55385708808899,3.58417987823486,3.66162085533142,3.68730497360229,3.68461942672729,3.6171875,3.59843730926514,3.55668926239014,3.53627943992615,3.53520512580872,3.50537133216858,3.46308588981628,3.36953091621399,3.20883798599243,3.16513705253601,3.10097646713257,2.99321269989014,2.89653325080872,2.83173847198486,2.70102572441101,2.54277348518372,2.40678715705872,2.27006840705872,2.36376953125,2.47348618507385,2.59921908378601,2.63266611099243,2.67001938819885,2.67817378044128,2.77299833297729,2.839111328125,2.90859389305115,2.97666001319885,3.02871108055115,3.07578134536743,3.09584951400757,3.10307645797729,3.12719750404358,3.22968745231628,3.32929682731628,3.45683598518372,3.56713843345642,3.70214867591858,3.826904296875,3.94721674919128,4.016357421875,4.02294921875,4.02446317672729,4.03691434860229,4.06914091110229,4.16396522521973,4.28486347198486,4.35854482650757,4.47187471389771,4.55810546875,4.60239219665527,4.66557645797729,5.06552743911743,5.17905235290527,5.22880840301514,5.251708984375,5.27993154525757,5.35107421875,5.41474628448486,5.43964910507202,5.49550771713257,5.86513662338257,5.88398408889771,5.916015625,5.91357421875,5.91689443588257,5.970703125,6.03872108459473,6.08671855926514,6.12680673599243,6.16191387176514,6.19121122360229,6.250244140625,6.27978515625,6.31635713577271,6.36572265625,6.55571269989014,6.74531269073486,6.78442335128784,6.90991258621216,7.06357479095459,7.13491201400757,7.20615196228027,7.26357412338257,7.358154296875,7.52377939224243,7.51503944396973,7.48842811584473,7.47529268264771,7.50756883621216,7.57211923599243,7.62343692779541,7.68354511260986,7.77236318588257,7.81899404525757,7.85996150970459,7.865478515625,7.83588838577271,7.74336004257202,7.65175771713257,7.55097627639771,7.55732440948486,7.63369178771973,7.68081045150757,7.70190382003784,7.81298828125,7.88457059860229,7.89091825485229,7.90981483459473,7.98359394073486,7.97382736206055,7.98544883728027,8.02036190032959,8.03203105926514,8.0458984375,8.060791015625,8.167724609375,8.19770526885986,8.24379825592041,8.40507793426514,8.54121017456055,8.5869140625,8.59028339385986,8.59882831573486,8.65615272521973,8.71543025970459,8.83603572845459,8.85249042510986,8.87319374084473,8.88974666595459,8.93886661529541,8.99501895904541,9.01596641540527,9.01162052154541,9.02358436584473,9.02089786529541,9.04936504364014,9.07514667510986,9.13320350646973,9.12709999084473,9.27495098114014,9.30136680603027,9.32451152801514,9.34711933135986,9.40566444396973,9.52714824676514,9.63627910614014,9.71323204040527,9.974609375,9.96914100646973,10.0013675689697,10.1756830215454,10.2078123092651,10.2185544967651,10.23828125,10.2898435592651,10.3665533065796,10.4616203308105,10.5378904342651,10.5748043060303,10.6086912155151,10.642822265625,10.7366695404053,10.7820320129395,10.8227052688599,10.830078125,10.8260746002197,10.8513669967651,10.8941402435303,10.9515132904053,10.9962396621704,10.9773435592651,10.9540529251099,10.9271974563599,10.919677734375],[43.9396018981934,43.9039077758789,43.9066429138184,43.9391136169434,43.953369140625,43.9396018981934,43.9471702575684,44.0028305053711,43.9396018981934],[44.2922859191895,44.25732421875,44.2916030883789,44.3790054321289,44.3920440673828,44.2922859191895],[44.6817855834961,44.62890625,44.763916015625,44.8053703308105,44.7914085388184,44.7098121643066,44.6817855834961],[45.4899406433105,45.4817390441895,45.4960441589355,45.5373039245605,45.5578117370605,45.5773429870605,45.5672874450684,45.5155258178711,45.4899406433105],[45.5854988098145,45.5648918151855,45.5735816955566,45.6718254089355,45.6944808959961,45.5854988098145],[45.4690933227539,45.4491195678711,45.4676284790039,45.4419403076172,45.44140625,45.5157203674316,45.5614242553711,45.701171875,45.7047386169434,45.5464363098145,45.4898452758789,45.4690933227539],[46.8729476928711,46.8648452758789,46.8995628356934,46.96142578125,46.99609375,46.995361328125,46.9195289611816,46.8729476928711],[45.9447250366211,45.9371109008789,45.9855461120605,45.9834976196289,45.9486312866211,45.9106941223145,45.8822250366211,45.8558120727539,45.8379859924316,45.7950172424316,45.7483901977539,45.703369140625,45.7477073669434,45.7480964660645,45.7380867004395,45.7514190673828,45.7733383178711,45.946533203125,45.9687004089355,45.9329109191895,45.9565925598145,46.0614280700684,46.1166534423828,46.203857421875,46.255615234375,46.2845687866211,46.3112297058105,46.2701148986816,46.195556640625,46.2060089111328,46.1909675598145,46.1595230102539,46.1414070129395,46.1129379272461,46.0616226196289,46.0194358825684,45.9651374816895,45.9415512084961,45.8804664611816,45.8188972473145,45.7430152893066,45.6546363830566,45.5908203125,45.5908203125,45.6106910705566,45.6061515808105,45.5823745727539,45.5850105285645,45.572509765625,45.5738754272461,45.5984878540039,45.6690940856934,45.7162132263184,45.9414558410645,46.0597648620605,46.1703605651855,46.2438468933105,46.3025398254395,46.6504898071289,46.7294425964355,46.7967796325684,46.8633804321289,46.9757804870605,46.9988288879395,47.0097160339355,47.0035133361816,46.9629402160645,46.9231948852539,46.767822265625,46.7370109558105,46.6133308410645,46.4135246276855,46.3033676147461,46.270263671875,46.2145500183105,46.1721687316895,46.0926742553711,46.0741195678711,46.0445785522461,45.9447250366211],[46.4687004089355,46.4546394348145,46.4805183410645,46.5619163513184,46.5406265258789,46.50390625,46.5120086669922,46.5082550048828,46.460205078125,46.4222183227539,46.427734375,46.4502944946289,46.4594230651855,46.478271484375,46.4872055053711,46.4657211303711,46.4457015991211,46.4215812683105,46.3553695678711,46.2783203125,46.202880859375,46.1659164428711,46.0979499816895,46.0286636352539,46.02294921875,45.9997062683105,45.977294921875,45.9668922424316,45.9731941223145,46.0013694763184,46.0682640075684,46.0584487915039,46.0666007995605,46.1235847473145,46.1951637268066,46.18994140625,46.2239265441895,46.2698249816895,46.2921409606934,46.3163566589355,46.2953605651855,46.2528305053711,46.2367210388184,46.2000007629395,46.1883316040039,46.1598625183105,46.1532707214355,46.209228515625,46.23046875,46.2890625,46.3673324584961,46.370361328125,46.3843765258789,46.3976097106934,46.4081535339355,46.4048347473145,46.4254417419434,46.5621109008789,46.5997085571289,46.6314468383789,46.640869140625,46.6916007995605,46.7692375183105,46.8357391357422,46.9012680053711,46.9548835754395,47.0615692138672,46.9817390441895,46.9129867553711,46.7754402160645,46.6391105651855,46.6089859008789,46.5723609924316,46.5386734008789,46.5087890625,46.493896484375,46.4687004089355],[47.2845230102539,47.2655258178711,47.267578125,47.2598152160645,47.2226066589355,47.218994140625,47.2342796325684,47.4251441955566,47.4690895080566,47.5938491821289,47.6317863464355,47.6467781066895,47.6376457214355,47.56396484375,47.5600090026855,47.4987297058105,47.4308128356934,47.3920402526855,47.3446273803711,47.2845230102539],[47.4413566589355,47.4065437316895,47.4081039428711,47.4385261535645,47.4976577758789,47.5399932861328,47.5655250549316,47.5851097106934,47.6070823669434,47.6468276977539,47.5730972290039,47.4413566589355],[47.88671875,47.8137702941895,47.7519035339355,47.7476081848145,47.7536125183105,47.7935562133789,47.863037109375,47.8724594116211,47.8662605285645,47.88671875],[47.9588851928711,47.9072265625,47.9849624633789,48.0050811767578,48.0137672424316,48.0069351196289,47.9588851928711],[48.7544441223145,48.728759765625,48.732177734375,48.7501487731934,48.92578125,48.9220695495605,48.8673858642578,48.845703125,48.7933578491211,48.7560539245605,48.7544441223145],[48.8861312866211,48.8751945495605,48.9459495544434,49.0386238098145,49.0951156616211,48.9546890258789,48.9082527160645,48.8861312866211],[45.0038566589355,45.0415992736816,45.1882820129395,45.2414093017578,45.3954582214355,45.4250984191895,45.458984375,45.5867691040039,45.63232421875,45.7578125,45.8636703491211,46.0100593566895,46.1036148071289,46.1818351745605,46.3526878356934,46.40478515625,46.4420890808105,46.5115203857422,46.5512199401855,46.6319351196289,46.6537590026855,46.7207527160645,46.7562522888184,46.819091796875,46.8521995544434,47.0325202941895,47.116943359375,47.2898445129395,47.3777847290039,47.471435546875,47.6234397888184,47.8236846923828,47.96728515625,48.0470199584961,48.2750015258789,48.3660125732422,48.3764152526855,48.4573249816895,48.5416984558105,48.6264152526855,48.7309112548828,48.85595703125,48.9641609191895,49.1263694763184,49.213134765625,49.2256813049316,49.2661628723145,49.2620582580566,49.1917495727539,49.1047859191895,48.921875,48.8736343383789,48.8062019348145,48.8389625549316,48.8411102294922,48.8036155700684,48.6911125183105,48.5503921508789,48.4231948852539,48.3605003356934,48.3105964660645,48.2280731201172,48.1964836120605,48.1598625183105,48.1062469482422,48.021240234375,48.0111351013184,48.031494140625,48.1116714477539,48.1888656616211,48.1466789245605,48.1026878356934,48.1173324584961,48.097900390625,48.11962890625,48.0224609375,48.0110816955566,48.0669441223145,48.0606422424316,47.9885749816895,47.9110336303711,47.8597679138184,47.6961441040039,47.6700210571289,47.68701171875,47.7679176330566,47.811279296875,47.8468284606934,47.7930183410645,47.7972183227539,47.724853515625,47.6734886169434,47.5698738098145,47.36865234375,47.2337913513184,47.1012191772461,47.0692367553711,47.049560546875,47.0888214111328,47.086181640625,47.060791015625,46.9578132629395,46.887939453125,46.8228530883789,46.6986846923828,46.6714363098145,46.5123023986816,46.4255828857422,46.35595703125,46.3114242553711,46.2403335571289,46.22021484375,46.19287109375,46.1658172607422,46.1461906433105,46.107177734375,46.0213356018066,45.959228515625,45.8580055236816,45.8779296875,45.8747062683105,45.811279296875,45.7798843383789,45.751953125,45.7579574584961,45.7824249267578,45.7763671875,45.7405738830566,45.6859855651855,45.6482391357422,45.6606941223145,45.621826171875,45.6405258178711,45.6646461486816,45.7308578491211,45.8681640625,45.8511734008789,45.7991180419922,45.7142105102539,45.6556129455566,45.6421852111816,45.68701171875,45.6482925415039,45.5736808776855,45.4760246276855,45.4410629272461,45.4105949401855,45.36669921875,45.3486328125,45.3301773071289,45.291748046875,45.2528305053711,45.2334480285645,45.256103515625,45.2355003356934,45.18505859375,45.1570320129395,45.15380859375,45.1305160522461,45.094482421875,45.0844230651855,44.9944839477539,44.9364738464355,44.8436508178711,44.7851066589355,44.7147979736328,44.7085418701172,44.7113265991211,44.642578125,44.6519050598145,44.6399421691895,44.6550750732422,44.6832046508789,44.610595703125,44.5437507629395,44.5144538879395,44.47998046875,44.4864273071289,44.5106468200684,44.5463371276855,44.6038589477539,44.6449203491211,44.587890625,44.54541015625,44.4874496459961,44.586669921875,44.5503425598145,44.4448738098145,44.4147453308105,44.3340835571289,44.2919921875,44.3035659790039,44.1851577758789,44.1420402526855,44.0213394165039,43.9293441772461,43.8678703308105,43.7271957397461,43.7313957214355,43.7267570495605,43.6681137084961,43.5496101379395,43.5652809143066,43.5614280700684,43.5242195129395,43.51806640625,43.5532722473145,43.5340347290039,43.5607452392578,43.734375,43.7952156066895,43.8148422241211,43.7781257629395,43.7421875,43.8138198852539,44.0796890258789,44.1438484191895,44.3674812316895,44.5687980651855,44.5755348205566,44.4359359741211,44.4697265625,44.5049324035645,44.5617179870605,44.6150894165039,44.6461906433105,44.6509284973145,44.680419921875,44.732666015625,44.7604026794434,44.728515625,44.6971206665039,44.73828125,44.7603034973145,45.1208000183105,45.1802253723145,45.2560539245605,45.3057136535645,45.3374519348145,45.309326171875,45.2682113647461,45.2382316589355,45.187255859375,45.1382293701172,45.1143074035645,45.0230445861816,45.1470222473145,45.2170867919922,45.3108863830566,45.3210945129395,45.3647956848145,45.3941421508789,45.378173828125,45.410888671875,45.3895492553711,45.4100570678711,45.3829612731934,45.3243637084961,45.3502426147461,45.3545913696289,45.3748054504395,45.4755363464355,45.62548828125,45.755859375,45.783203125,45.835693359375,45.826904296875,45.8063468933105,45.8666038513184,45.9466323852539,45.913330078125,45.8136711120605,45.638427734375,45.6259765625,45.5442390441895,45.4730949401855,45.3373031616211,45.222900390625,45.2224617004395,45.3166007995605,45.3594703674316,45.4175796508789,45.4008293151855,45.3756332397461,45.335205078125,45.2569313049316,45.2275848388672,45.1890144348145,45.1332054138184,45.0958976745605,45.1433601379395,45.0833969116211,45.0672836303711,45.09765625,45.1456069946289,45.1571769714355,45.1439437866211,45.16943359375,45.1819801330566,45.1925315856934,45.2007827758789,45.1867179870605,45.1679229736328,45.15380859375,45.1737785339355,45.2101554870605,45.2476577758789,45.27587890625,45.3086929321289,45.3403816223145,45.3779296875,45.4212379455566,45.4458999633789,45.4740715026855,45.5010261535645,45.5139656066895,45.5304183959961,45.5655746459961,45.6031227111816,45.618408203125,45.612548828125,45.6207542419434,45.6441917419434,45.6711921691895,45.6864776611328,45.6864776611328,45.7017059326172,45.7275390625,45.769775390625,45.7955589294434,45.81787109375,45.842529296875,45.8601570129395,45.8741683959961,45.8917999267578,45.927001953125,45.9527816772461,46.0421371459961,46.2093276977539,46.33740234375,46.4983863830566,46.6156234741211,46.7798843383789,46.9357414245605,47.0828132629395,47.1676254272461,47.2748527526855,47.345947265625,47.3544921875,47.3445320129395,47.316162109375,47.2857933044434,47.2534675598145,47.2033195495605,47.2028312683105,47.2112312316895,47.2364273071289,47.2736320495605,47.338134765625,47.4266090393066,47.4447746276855,47.4629898071289,47.4020004272461,47.3506355285645,47.2386703491211,47.0813484191895,46.994873046875,46.8429183959961,46.7089347839355,46.5714378356934,46.4410667419434,46.3418464660645,46.2508773803711,46.1500015258789,46.057373046875,45.9798316955566,45.9391593933105,45.9061050415039,45.8680686950684,45.8019027709961,45.7382316589355,45.7068367004395,45.6439933776855,45.5513687133789,45.4989242553711,45.4553718566895,45.4283180236816,45.4094696044922,45.4106941223145,45.40478515625,45.3661613464355,45.3106918334961,45.2707023620605,45.262451171875,45.2907218933105,45.3331069946289,45.3372573852539,45.3091316223145,45.2628402709961,45.2603492736816,45.2900848388672,45.2003402709961,45.007568359375,45.007080078125,45.006591796875,45.006103515625,45.005615234375,45.0051765441895,45.0046882629395,45.0041999816895,45.00390625,45.0038566589355],[49.35400390625,49.2636222839355,49.2781257629395,49.295654296875,49.3390617370605,49.3797836303711,49.3650398254395,49.35400390625],[49.5888671875,49.5306625366211,49.5077629089355,49.4961433410645,49.5144538879395,49.5760726928711,49.5965805053711,49.599365234375,49.5912094116211,49.5720672607422,49.5623512268066,49.5621566772461,49.6220703125,49.6314926147461,49.6199722290039,49.5888671875],[49.7444343566895,49.6794929504395,49.6619644165039,49.5916519165039,49.5870132446289,49.5953636169434,49.6608390808105,49.7114753723145,49.718994140625,49.6885261535645,49.7511749267578,49.7444343566895],[49.5311508178711,49.5103492736816,49.5881843566895,49.6342315673828,49.66748046875,49.6863288879395,49.7356910705566,49.7583465576172,49.7751007080078,49.7649421691895,49.727783203125,49.6672859191895,49.5311508178711],[49.6058082580566,49.6013679504395,49.6134757995605,49.64208984375,49.7184562683105,49.7356910705566,49.75048828125,49.7627906799316,49.7829093933105,49.8377456665039,49.8723640441895,49.8436508178711,49.8084945678711,49.7458000183105,49.6751480102539,49.6268043518066,49.6058082580566],[49.0938987731934,49.0791015625,49.140869140625,49.1707038879395,49.2249526977539,49.3993148803711,49.4599151611328,49.5343246459961,49.6020011901855,49.65771484375,49.8277320861816,49.886962890625,49.9259262084961,49.9443855285645,49.941650390625,49.875244140625,49.8168449401855,49.772705078125,49.7054672241211,49.6239280700684,49.4070777893066,49.3897933959961,49.2835426330566,49.2037620544434,49.1390151977539,49.1057624816895,49.0938987731934],[50.0295867919922,50.020751953125,50.1340827941895,50.1751937866211,50.1958503723145,50.2171401977539,50.1659202575684,50.1315422058105,50.083984375,50.0712890625,50.0295867919922],[50.0971183776855,50.0443344116211,50.1300315856934,50.3115234375,50.3539543151855,50.4140625,50.4178237915039,50.3897438049316,50.3396949768066,50.3202629089355,50.2677726745605,50.2206535339355,50.1634788513184,50.0971183776855],[50.7196769714355,50.7086906433105,50.7090301513672,50.7207984924316,50.7401847839355,50.7807159423828,50.8012237548828,50.79638671875,50.7759284973145,50.7421379089355,50.7196769714355],[50.640380859375,50.5155296325684,50.453857421875,50.3808097839355,50.3585433959961,50.3424797058105,50.3167991638184,50.254638671875,50.106689453125,50.01220703125,49.8481941223145,49.7316436767578,49.6853523254395,49.6704559326172,49.6431655883789,49.5300750732422,49.4286613464355,49.3802719116211,49.3005867004395,49.2240219116211,49.1708030700684,49.1191902160645,49.08349609375,48.9512214660645,48.8240242004395,48.5820808410645,48.602294921875,48.6744155883789,48.6904792785645,48.6981925964355,48.6702117919922,48.6064453125,48.4551734924316,48.4110336303711,48.406494140625,48.42724609375,48.4000968933105,48.344970703125,48.3228034973145,48.3335418701172,48.3865737915039,48.4364280700684,48.5152320861816,48.5973167419434,48.6536140441895,48.7114753723145,48.7607917785645,48.8026390075684,48.8224105834961,48.9563484191895,49.0282745361328,49.0833015441895,49.1415519714355,49.212646484375,49.2071266174316,49.1390609741211,49.0785140991211,49.031005859375,49.0142097473145,48.9910163879395,48.9982414245605,48.9410629272461,48.9337882995605,48.9528350830078,49.0291519165039,49.0918464660645,49.1072273254395,49.1392097473145,49.185791015625,49.1932106018066,49.1903800964355,49.1998519897461,49.2602043151855,49.2766609191895,49.24951171875,49.248046875,49.2878913879395,49.3797836303711,49.4014625549316,49.3680152893066,49.3790016174316,49.4087905883789,49.4212875366211,49.4151840209961,49.4426727294922,49.431884765625,49.3921890258789,49.3820304870605,49.4490242004395,49.4511222839355,49.3999519348145,49.3967781066895,49.4189453125,49.5432624816895,49.57861328125,49.5904808044434,49.6192855834961,49.650146484375,49.6723136901855,49.6608390808105,49.677734375,49.72021484375,49.7195816040039,49.7333984375,49.7641143798828,49.87646484375,49.9049301147461,49.9228019714355,49.9441413879395,49.9347190856934,49.9026870727539,49.8828125,49.8715362548828,49.8797378540039,49.9104499816895,49.9491691589355,49.992431640625,50.05029296875,50.0731430053711,50.0999031066895,50.1214866638184,50.1379890441895,50.1293487548828,50.0955581665039,50.0708503723145,50.0519523620605,50.0850105285645,50.130859375,50.1634254455566,50.163330078125,50.1211433410645,50.1177253723145,50.1277351379395,50.1500968933105,50.21142578125,50.293212890625,50.313720703125,50.3262214660645,50.3459968566895,50.4452171325684,50.4639663696289,50.4710426330566,50.4790992736816,50.4649429321289,50.4046363830566,50.4273452758789,50.4957542419434,50.5367660522461,50.5831069946289,50.5966796875,50.6073760986328,50.5777359008789,50.5357437133789,50.4988784790039,50.4926261901855,50.4984855651855,50.5205574035645,50.6092758178711,50.6965827941895,50.7442398071289,50.7941398620605,50.8281745910645,50.8577613830566,50.8605461120605,50.8207550048828,50.640380859375],[51.5365219116211,51.4369621276855,51.3885726928711,51.3728981018066,51.3586921691895,51.343017578125,51.328369140625,51.2618637084961,51.2269020080566,51.1992683410645,51.19140625,51.2079086303711,51.205078125,51.191162109375,51.1661643981934,51.1314468383789,51.0870590209961,51.0333023071289,50.9073753356934,50.8376960754395,50.7809562683105,50.75927734375,50.7337875366211,50.6509742736816,50.5847663879395,50.4169921875,50.3800277709961,50.3504867553711,50.2708473205566,50.2067413330078,50.0596694946289,50.0077133178711,49.966552734375,49.9084968566895,49.8829116821289,49.833740234375,49.7877464294434,49.7653312683105,49.724609375,49.7103996276855,49.67333984375,49.6134757995605,49.651611328125,49.7876930236816,49.86962890625,49.8974113464355,49.9336929321289,50.0137710571289,50.090087890625,50.114990234375,50.100341796875,50.0628395080566,50.01513671875,49.957275390625,49.9401359558105,49.9826164245605,50.0177726745605,50.0131340026855,49.96044921875,49.966552734375,49.9971694946289,49.983154296875,49.936767578125,49.8923835754395,49.8290023803711,49.7046852111816,49.619140625,49.6217269897461,49.6583976745605,49.6781234741211,49.6808586120605,49.670166015625,49.6459465026855,49.5802764892578,49.4519538879395,49.4568367004395,49.5152816772461,49.4346160888672,49.4624977111816,49.48974609375,49.472900390625,49.4376945495605,49.380859375,49.3729019165039,49.3919410705566,49.4127426147461,49.5138168334961,49.52392578125,49.5081558227539,49.4820327758789,49.4084968566895,49.3346672058105,49.2669906616211,49.1681137084961,49.1255874633789,49.0794448852539,49.077880859375,49.1208953857422,49.1385726928711,49.1796417236328,49.1998062133789,49.2444343566895,49.29736328125,49.3053703308105,49.2930183410645,49.2603530883789,49.2681159973145,49.3162612915039,49.3454093933105,49.35546875,49.3885765075684,49.4445304870605,49.4908180236816,49.52734375,49.52978515625,49.4693336486816,49.4005393981934,49.3417472839355,49.3294448852539,49.3921356201172,49.4150390625,49.4241180419922,49.4192886352539,49.4418487548828,49.4263191223145,49.3853034973145,49.3216285705566,49.26416015625,49.1916999816895,49.1412086486816,49.0775375366211,49.035400390625,48.993408203125,48.9513664245605,48.9254417419434,48.88916015625,48.7876968383789,48.7847671508789,48.8067855834961,48.8113288879395,48.7966766357422,48.767822265625,48.7388687133789,48.7249031066895,48.7066917419434,48.6846656799316,48.6954116821289,48.6798324584961,48.655517578125,48.5763206481934,48.5263671875,48.4845695495605,48.4188499450684,48.3936042785645,48.3883781433105,48.4366226196289,48.4488258361816,48.44921875,48.4958000183105,48.51123046875,48.4817848205566,48.5621566772461,48.5726051330566,48.5633316040039,48.577880859375,48.632568359375,48.6590309143066,48.6566429138184,48.6347160339355,48.5716285705566,48.515869140625,48.4803237915039,48.40185546875,48.3743629455566,48.364013671875,48.3681640625,48.35595703125,48.2943344116211,48.2318878173828,48.2077140808105,48.173828125,48.1084480285645,48.0880851745605,48.0679206848145,48.0568351745605,48.0423812866211,48.0196800231934,48.0097160339355,48.0257339477539,48.0146484375,47.9686508178711,47.9211921691895,47.7992706298828,47.7870140075684,47.7272453308105,47.6820335388184,47.6500968933105,47.6482429504395,47.6623077392578,47.7438468933105,47.9978485107422,48.0685043334961,48.1470680236816,48.1311531066895,48.1129913330078,48.0939483642578,48.029296875,47.9759292602539,47.8119163513184,47.7345695495605,47.6529808044434,47.5121116638184,47.48779296875,47.4551277160645,47.4831047058105,47.5528335571289,47.6194343566895,47.7278785705566,47.7694358825684,47.7689476013184,47.7453155517578,47.6930198669434,47.6217765808105,47.5494155883789,47.4698257446289,47.4263191223145,47.1032218933105,47.0458526611328,47.0110855102539,46.97412109375,46.8194351196289,46.7227516174316,46.6812477111816,46.6558113098145,46.646484375,46.6604995727539,46.6977081298828,46.717041015625,46.7183609008789,46.71142578125,46.6325187683105,46.6282691955566,46.6388664245605,46.6802711486816,46.8884773254395,46.957275390625,47.0103530883789,47.0994148254395,47.1332511901855,47.14599609375,47.1376953125,47.0929222106934,47.0118179321289,46.939453125,46.839599609375,46.8199691772461,46.8249015808105,46.8385772705078,46.88037109375,46.9171867370605,46.9674835205078,47.0862312316895,47.261962890625,47.3870124816895,47.4403305053711,47.4635734558105,47.5093269348145,47.6446762084961,47.7562026977539,47.8056144714355,47.8598136901855,47.8467292785645,47.7716789245605,47.555908203125,47.4623031616211,47.4275856018066,47.4038581848145,47.3751945495605,47.4250984191895,47.5362319946289,47.5470695495605,47.457763671875,47.408203125,47.3954582214355,47.3986320495605,47.385009765625,47.1739273071289,47.1035614013672,47.0459480285645,46.9417495727539,46.9057121276855,46.8992691040039,46.9172859191895,46.9140167236328,46.8814468383789,46.8672332763672,46.8738288879395,46.88720703125,46.9276847839355,46.9563980102539,46.9732437133789,47.0168952941895,47.0716323852539,47.0920867919922,47.11962890625,47.16064453125,47.2214813232422,47.2585945129395,47.448974609375,47.5161628723145,47.5708999633789,47.6403312683105,47.6647453308105,47.6294898986816,47.620849609375,47.6338844299316,47.6575698852539,47.6500511169434,47.6610832214355,47.6428718566895,47.5503921508789,47.5012664794922,47.4847679138184,47.4750518798828,47.4652366638184,47.498291015625,47.516357421875,47.5300788879395,47.4999504089355,47.5028305053711,47.5245094299316,47.5923347473145,47.787841796875,47.8191909790039,47.7918968200684,47.7637214660645,47.7718772888184,47.7891616821289,47.7745132446289,47.67138671875,47.658447265625,47.6544914245605,47.616943359375,47.59228515625,47.5649909973145,47.574462890625,47.631103515625,47.6253890991211,47.6600074768066,47.6749000549316,47.6688461303711,47.6768531799316,47.7198715209961,47.7308578491211,47.6833992004395,47.652587890625,47.626220703125,47.5804672241211,47.5707015991211,47.6025390625,47.6341781616211,47.7369117736816,47.8656768798828,47.8889656066895,47.9336433410645,47.9955596923828,48.1593742370605,48.3250503540039,48.4113273620605,48.4420394897461,48.513671875,48.5221176147461,48.5130386352539,48.5328636169434,48.5407218933105,48.5217781066895,48.5217781066895,48.5584945678711,48.627685546875,48.7464332580566,48.746826171875,48.6915512084961,48.6501960754395,48.6227073669434,48.6053237915039,48.5980453491211,48.6220703125,48.7494125366211,48.8968734741211,49.0032234191895,49.0843238830566,49.0965347290039,49.0813484191895,49.0619163513184,48.987548828125,48.9812507629395,48.9879417419434,49.009765625,49.0447273254395,49.0774383544922,49.1799812316895,49.20947265625,49.2296371459961,49.2300758361816,49.2587394714355,49.3051261901855,49.3866691589355,49.4353981018066,49.4997062683105,49.54248046875,49.5315399169922,49.4738273620605,49.489990234375,49.508544921875,49.6003913879395,49.6684074401855,49.7008323669434,50.02490234375,50.1987800598145,50.4636726379395,50.5058097839355,50.5839347839355,50.6051750183105,50.6053695678711,50.6148452758789,50.6493682861328,50.6733894348145,50.69873046875,50.7252960205078,50.7449226379395,50.7874031066895,50.8573265075684,50.9396514892578,50.9677238464355,50.9956550598145,51.0108413696289,51.0279769897461,51.125732421875,51.1444854736328,51.27490234375,51.332763671875,51.3624534606934,51.3993148803711,51.4886245727539,51.568359375,51.5639152526855,51.50830078125,51.4713401794434,51.5110359191895,51.5594253540039,51.5789070129395,51.5963859558105,51.58984375,51.5623054504395,51.5365219116211],[51.473876953125,51.4571762084961,51.4572296142578,51.4740219116211,51.5111312866211,51.626708984375,51.6465797424316,51.6667938232422,51.7083969116211,51.7038078308105,51.673583984375,51.60546875,51.58544921875,51.5061988830566,51.473876953125],[51.8896484375,51.8888168334961,51.9000511169434,51.9384765625,51.9828605651855,51.9951667785645,51.9299812316895,51.8896484375],[51.9519538879395,51.9448738098145,51.9529304504395,51.97802734375,52.0100593566895,52.09814453125,52.0868186950684,52.0710945129395,52.02392578125,51.9922828674316,51.9683113098145,51.9519538879395],[51.9616203308105,51.959716796875,51.9804191589355,52.0138664245605,52.1010246276855,52.1365737915039,52.150634765625,52.0952644348145,51.9616203308105],[52.40087890625,52.3875007629395,52.4411125183105,52.4728546142578,52.5027351379395,52.5603523254395,52.6963882446289,52.7818832397461,52.7843742370605,52.7412109375,52.5482406616211,52.458984375,52.4260749816895,52.40087890625],[52.510009765625,52.4642562866211,52.5743675231934,52.6053199768066,52.72216796875,52.7723655700684,52.80078125,52.8112297058105,52.8038558959961,52.791259765625,52.6617164611816,52.6007308959961,52.510009765625],[52.9921875,52.9842300415039,52.9976081848145,53.0237312316895,53.0977554321289,53.12890625,53.1793937683105,53.1883277893066,53.1830101013184,53.1747055053711,53.0527839660645,52.9921875],[52.7472648620605,52.7339859008789,52.7606430053711,52.7798843383789,52.8520011901855,52.9579086303711,53.010498046875,53.0499000549316,53.1322250366211,53.1655731201172,53.186279296875,53.2242660522461,53.205810546875,53.0371589660645,52.9232444763184,52.8315925598145,52.7873992919922,52.7472648620605],[53.195556640625,53.1029777526855,53.0200691223145,52.9221687316895,52.8850593566895,52.9146461486816,52.9091300964355,52.8798332214355,52.86669921875,52.8186988830566,52.7563934326172,52.7452125549316,52.7017097473145,52.6233406066895,52.5782241821289,52.4533195495605,52.422119140625,52.4157218933105,52.4258308410645,52.4159164428711,52.3900413513184,52.3175277709961,52.3030738830566,52.2916526794434,52.2911109924316,52.2190895080566,52.1536140441895,52.2379875183105,52.3220710754395,52.3999519348145,52.4439964294434,52.5416984558105,52.7527847290039,52.7833023071289,52.8667984008789,52.906982421875,52.9338874816895,52.9480934143066,52.9574737548828,52.9993171691895,53.0718727111816,53.0867195129395,53.1375007629395,53.1449241638184,53.1360855102539,53.1604995727539,53.1791496276855,53.2019500732422,53.2314453125,53.229736328125,53.195556640625],[52.9397430419922,52.6207046508789,52.5186042785645,52.4518051147461,52.3398933410645,52.2896461486816,52.3565406799316,52.4677238464355,52.5560569763184,52.598388671875,52.7633781433105,52.751220703125,52.6788063049316,52.673828125,52.7559585571289,52.8224601745605,52.891845703125,52.9649391174316,52.9906730651855,53.0179176330566,53.0906753540039,53.1396980285645,53.2406234741211,53.2799339294434,53.2743644714355,53.2285652160645,53.1788558959961,53.1124992370605,52.9397430419922],[53.1178741455078,53.1107406616211,53.1109352111816,53.1211433410645,53.1421394348145,53.1739768981934,53.2123031616211,53.2854995727539,53.3166999816895,53.330078125,53.293212890625,53.2591323852539,53.1178741455078],[53.3041534423828,53.2742652893066,53.2800788879395,53.2971687316895,53.3240737915039,53.3442878723145,53.3644523620605,53.3522453308105,53.3041534423828],[53.1679191589355,53.1645011901855,53.1766586303711,53.345703125,53.4073715209961,53.4908218383789,53.5442352294922,53.6311492919922,53.6204109191895,53.5496597290039,53.4811019897461,53.4363784790039,53.2447738647461,53.21728515625,53.1679191589355],[53.9585456848145,53.922607421875,53.8662605285645,53.8439445495605,53.8555183410645,53.8617668151855,53.9178733825684,53.9402809143066,53.94140625,53.921142578125,53.89404296875,53.8600082397461,53.8445281982422,53.8475570678711,53.8634757995605,53.8922386169434,53.9214859008789,53.9512672424316,53.9912605285645,54.0741691589355,54.0890121459961,54.085693359375,54.0469245910645,54.03564453125,53.9585456848145],[54.1274909973145,54.0686531066895,54.1060523986816,54.098876953125,54.0763168334961,54.0284194946289,53.9552268981934,53.8751945495605,53.8465347290039,53.8147468566895,53.6876449584961,53.6753883361816,53.6639671325684,53.6517105102539,53.6533203125,53.684814453125,53.7068367004395,53.8069839477539,53.8601570129395,53.9002952575684,53.995849609375,54.0342788696289,54.0419921875,54.0773429870605,54.1431655883789,54.141357421875,54.0228042602539,53.9863777160645,53.8415031433105,53.7139625549316,53.587890625,53.3791999816895,53.3086891174316,53.2651863098145,53.189208984375,53.1940422058105,53.26318359375,53.3104972839355,53.3678703308105,53.3705558776855,53.3592796325684,53.337890625,53.3369636535645,53.3504409790039,53.4585914611816,53.5077171325684,53.5626945495605,53.6053695678711,53.6291999816895,53.7781257629395,53.8370132446289,53.9202613830566,54.005615234375,54.14404296875,54.158935546875,54.1578140258789,54.1407699584961,54.1274909973145],[54.4790534973145,54.477783203125,54.4986801147461,54.5418472290039,54.6148910522461,54.6317863464355,54.618896484375,54.61376953125,54.5997047424316,54.5767097473145,54.5438003540039,54.4790534973145],[54.7696800231934,54.8341789245605,54.9459495544434,55.02587890625,54.947021484375,54.8145484924316,54.7487297058105,54.7696800231934],[56.039306640625,56.01513671875,56.0325698852539,56.0478515625,56.0639190673828,56.0889625549316,56.1183624267578,56.1408233642578,56.1292495727539,56.0988311767578,56.0804176330566,56.039306640625],[56.1453132629395,56.1314430236816,56.1328125,56.1663551330566,56.2320785522461,56.2898445129395,56.3396492004395,56.38330078125,56.4207038879395,56.439208984375,56.4386215209961,56.3173828125,56.260498046875,56.212890625,56.1745109558105,56.1453132629395],[56.20703125,56.1994132995605,56.2138671875,56.287353515625,56.3179168701172,56.3484344482422,56.3671875,56.3765144348145,56.3765144348145,56.4039535522461,56.4588394165039,56.4663581848145,56.3865242004395,56.3268089294434,56.20703125],[56.2660636901855,56.1649894714355,56.06787109375,55.8850593566895,55.8785171508789,55.9224624633789,56.1364250183105,56.1602554321289,56.1807136535645,56.212158203125,56.1759757995605,55.9348640441895,55.8961906433105,55.874755859375,55.8706550598145,55.8756828308105,55.8067893981934,56.0931625366211,56.114990234375,56.1283721923828,56.1209449768066,55.9405784606934,55.8710479736328,55.8921394348145,55.9110336303711,55.9320793151855,56.1141586303711,56.2444839477539,56.3265151977539,56.40380859375,56.4603500366211,56.5226097106934,56.5397453308105,56.5365715026855,56.5130348205566,56.4474601745605,56.3128395080566,56.24658203125,56.1919937133789,56.1800765991211,56.2110824584961,56.2764663696289,56.3763198852539,56.4630889892578,56.6004371643066,56.5956535339355,56.5682601928711,56.5489273071289,56.53759765625,56.5199699401855,56.4739265441895,56.45361328125,56.4364242553711,56.4217262268066,56.3714370727539,56.3449211120605,56.3279266357422,56.3204116821289,56.2660636901855],[56.6566886901855,56.6438484191895,56.6449203491211,56.7334938049316,56.7648468017578,56.7809562683105,56.7957038879395,56.7987289428711,56.7714347839355,56.74267578125,56.7144050598145,56.6865234375,56.6672859191895,56.6566886901855],[56.7745628356934,56.7571296691895,56.7767562866211,56.826904296875,56.8652839660645,56.863525390625,56.8431167602539,56.8160171508789,56.7745628356934],[57.5545883178711,57.5249519348145,57.4160652160645,57.4224586486816,57.4954109191895,57.5484886169434,57.5576133728027,57.5667457580566,57.5541038513184,57.5731468200684,57.5793495178223,57.5545883178711],[57.5155258178711,57.5075225830078,57.5144538879395,57.4485855102539,57.4424324035645,57.4830093383789,57.5177268981934,57.5415992736816,57.5592727661133,57.5987319946289,57.6079597473145,57.6045913696289,57.5550270080566,57.5155258178711],[59.0402374267578,58.9675788879395,58.9766120910645,58.9616203308105,58.9607391357422,59.0289535522461,59.0748062133789,59.1212425231934,59.1448745727539,59.1461448669434,59.0927734375,59.0727081298828,59.0635757446289,59.0402374267578],[59.6241226196289,59.6210441589355,59.6332015991211,59.6741676330566,59.6791534423828,59.724609375,59.7088851928711,59.6834983825684,59.6449203491211,59.6241226196289],[59.7708015441895,59.7638702392578,59.8231964111328,59.8518562316895,59.8701667785645,59.8769569396973,59.8531265258789,59.8099098205566,59.7708015441895],[60.3670883178711,60.2978515625,60.3232421875,60.3756332397461,60.410400390625,60.4484367370605,60.5010223388672,60.5096168518066,60.5146026611328,60.4414024353027,60.4302215576172,60.3670883178711],[60.2409172058105,60.23291015625,60.2540512084961,60.3147430419922,60.360595703125,60.4555702209473,60.56201171875,60.5878410339355,60.5704116821289,60.5398445129395,60.4888191223145,60.4495086669922,60.3916511535645,60.3398933410645,60.3046417236328,60.2409172058105],[60.7285652160645,60.7168922424316,60.7313499450684,60.7723121643066,60.8081092834473,60.8186531066895,60.7838897705078,60.7563972473145,60.7285652160645],[61.3660659790039,61.3544425964355,61.3570785522461,61.410400390625,61.4320335388184,61.4529800415039,61.5628433227539,61.6110382080078,61.6495132446289,61.6682624816895,61.6856918334961,61.6851043701172,61.6622085571289,61.6376495361328,61.5930213928223,61.5393562316895,61.4714851379395,61.4384269714355,61.4132843017578,61.3660659790039],[61.8440895080566,61.8417015075684,61.8927230834961,61.9185523986816,61.9350090026855,61.8896980285645,61.8674774169922,61.8440895080566],[61.8790550231934,61.8702659606934,61.8806114196777,61.8716773986816,61.8433609008789,61.82373046875,61.7981452941895,61.7798805236816,61.7615203857422,61.7332992553711,61.7325210571289,61.7976531982422,61.864013671875,61.8977088928223,61.9283676147461,61.9432106018066,61.9422340393066,61.926025390625,61.8790550231934],[62.4117202758789,62.384521484375,62.293701171875,62.2476539611816,62.1859817504883,62.103515625,62.0260734558105,61.9677734375,61.8940963745117,61.8080062866211,61.7096176147461,61.6444320678711,61.612548828125,61.5959510803223,61.5946311950684,61.630126953125,61.7025337219238,61.7468299865723,61.7772445678711,61.8182106018066,61.8585929870605,61.9894981384277,62.0546379089355,62.1090354919434,62.1526870727539,62.2127914428711,62.3429679870605,62.3928680419922,62.4043426513672,62.39501953125,62.3982925415039,62.4132308959961,62.4117202758789],[62.5587425231934,62.5479965209961,62.5518074035645,62.5359344482422,62.4874000549316,62.4178695678711,62.3914070129395,62.3835945129395,62.40625,62.4210433959961,62.4583511352539,62.4850082397461,62.494140625,62.5049819946289,62.5254364013672,62.5433120727539,62.5587425231934],[62.618408203125,62.6096649169922,62.6219749450684,62.6687965393066,62.7127418518066,62.726318359375,62.7332992553711,62.7265090942383,62.69580078125,62.6803245544434,62.6626968383789,62.6360855102539,62.618408203125],[62.5487289428711,62.5448226928711,62.5523414611816,62.5731925964355,62.5968742370605,62.6480941772461,62.704345703125,62.7877922058105,62.81591796875,62.8739280700684,62.8779754638672,62.8653335571289,62.8400917053223,62.8070335388184,62.7337913513184,62.6658172607422,62.6159629821777,62.5780754089355,62.5615768432617,62.5487289428711],[62.9542007446289,62.9262199401855,62.8840370178223,62.82763671875,62.7763175964355,62.7300758361816,62.6522445678711,62.5191383361816,62.4465827941895,62.4032249450684,62.2099075317383,62.2003898620605,62.2041053771973,62.2329597473145,62.2381324768066,62.1602554321289,62.1735877990723,62.2571754455566,62.3035163879395,62.4024925231934,62.425537109375,62.4541511535645,62.4764633178711,62.56884765625,62.9049301147461,62.9215812683105,62.8841323852539,62.8720703125,62.8739280700684,62.9445304870605,62.9361839294434,62.9774398803711,62.9776840209961,62.9705619812012,62.9542007446289],[63.4705581665039,63.4278335571289,63.4308586120605,63.3959999084473,63.2870559692383,63.2689437866211,63.2336463928223,63.1884307861328,63.1646003723145,63.1295890808105,63.1144027709961,63.1388702392578,63.2398414611816,63.3579063415527,63.4237289428711,63.4511260986328,63.469970703125,63.4895515441895,63.4789505004883,63.4705581665039],[63.3939437866211,63.384033203125,63.4063453674316,63.4497528076172,63.5883293151855,63.6820335388184,63.5735855102539,63.5035667419434,63.3939437866211],[63.9918937683105,63.9457054138184,63.9620094299316,63.9760246276855,63.9929161071777,64.0147933959961,64.03564453125,64.0372085571289,64.0304183959961,64.0220642089844,63.9918937683105],[65.2610855102539,65.2489776611328,65.2559051513672,65.3052749633789,65.4473190307617,65.4606399536133,65.4584503173828,65.3672409057617,65.3163070678711,65.2454605102539,65.2178192138672,65.1812438964844,65.1689910888672,65.1317901611328,65.1039047241211,64.9679641723633,64.9596710205078,64.9041061401367,64.7803192138672,64.7619171142578,64.7211456298828,64.690673828125,64.644287109375,64.5594253540039,64.4259796142578,64.212646484375,64.1705093383789,64.1455612182617,64.118896484375,64.03125,64.0164031982422,64.0218734741211,64.0757827758789,64.037109375,64.0310592651367,64.0333023071289,64.1004867553711,64.0899429321289,64.0247573852539,63.9720726013184,63.9319381713867,63.9092254638672,63.9014663696289,63.8629379272461,63.8019523620605,63.76220703125,63.6737823486328,63.5963897705078,63.4802780151367,63.4625473022461,63.4615707397461,63.4832000732422,63.5380859375,63.6644515991211,63.6911582946777,63.706787109375,63.7365264892578,63.9269523620605,63.960693359375,64.0004348754883,64.0232391357422,64.0614242553711,64.1269989013672,64.1590347290039,64.1575164794922,64.143798828125,64.0992202758789,64.05810546875,64.013427734375,63.9178237915039,63.8726081848145,63.8133773803711,63.6598625183105,63.6137199401855,63.6004867553711,63.5857887268066,63.5290985107422,63.3900375366211,63.3499984741211,63.3092269897461,63.2469215393066,63.1972198486328,63.1393051147461,63.1196784973145,63.1391105651855,63.2708969116211,63.657958984375,63.6841316223145,63.7003402709961,63.7065391540527,63.6567878723145,63.6623077392578,63.5752906799316,63.5689926147461,63.5717811584473,63.5856475830078,63.5951194763184,63.6328163146973,63.6722679138184,63.7148933410645,63.8304214477539,63.9016571044922,63.9237327575684,64.0515594482422,64.0936584472656,64.1368637084961,64.1812515258789,64.238037109375,64.3765106201172,64.5029830932617,64.5658187866211,64.662353515625,64.8963394165039,65.0102996826172,65.4172821044922,65.5338363647461,65.6402816772461,65.7042465209961,65.8319320678711,65.8831481933594,65.9186553955078,65.91455078125,65.8997039794922,65.8455581665039,65.7955093383789,65.7468719482422,65.6929244995117,65.6227035522461,65.5920867919922,65.5457534790039,65.5262222290039,65.5103073120117,65.4374008178711,65.2610855102539],[65.5749969482422,65.563720703125,65.570068359375,65.5989761352539,65.6505355834961,65.6891632080078,65.7373504638672,65.7561950683594,65.8208541870117,65.8853530883789,65.9437484741211,65.972412109375,66.015380859375,66.0254821777344,66.008544921875,65.9970245361328,65.9720764160156,65.9414978027344,65.8589401245117,65.7931594848633,65.6573715209961,65.6314926147461,65.6040573120117,65.5749969482422],[65.7967300415039,65.7574691772461,65.7351989746094,65.7232894897461,65.7150421142578,65.6965866088867,65.6678237915039,65.6310577392578,65.6299819946289,65.6559600830078,65.669189453125,65.6986312866211,65.7013626098633,65.6915054321289,65.6785202026367,65.662353515625,65.6577606201172,65.6688919067383,65.6783218383789,65.7100143432617,65.7561950683594,65.7704086303711,65.7874984741211,65.7584457397461,65.7515106201172,65.7717742919922,65.87744140625,65.9159698486328,65.942138671875,65.9697799682617,65.9906234741211,66.0118179321289,66.0644073486328,66.0882797241211,66.1062545776367,66.1310043334961,66.0778350830078,66.0274963378906,65.9657669067383,65.9201126098633,65.890380859375,65.8607406616211,65.8311538696289,65.7967300415039],[66.2828140258789,66.2706527709961,66.2719268798828,66.25732421875,66.2084426879883,66.1992645263672,66.2342300415039,66.2770462036133,66.302978515625,66.33154296875,66.3364791870117,66.2828140258789],[67.0051727294922,66.9972610473633,66.9985809326172,66.9213409423828,66.8718719482422,66.8578109741211,66.8848571777344,66.9468765258789,67.0051727294922],[67.0562973022461,67.02880859375,67.0625915527344,67.0721130371094,67.112548828125,67.1482391357422,67.1769561767578,67.1900405883789,67.1884765625,67.1783218383789,67.1342315673828,67.0562973022461],[67.4018096923828,67.3182144165039,67.3260269165039,67.3659133911133,67.3850631713867,67.429443359375,67.5248107910156,67.5681686401367,67.6285629272461,67.6648941040039,67.6221160888672,67.5493621826172,67.5135726928711,67.4764633178711,67.467041015625,67.4372100830078,67.4018096923828],[67.9823760986328,67.9718780517578,67.979736328125,67.9394073486328,67.8985366821289,67.8844680786133,67.8788070678711,67.9240264892578,67.9517059326172,67.9749984741211,67.9823760986328],[67.7838363647461,67.7825164794922,67.7895965576172,67.8048782348633,67.8286590576172,67.9055633544922,67.9848175048828,68.0184631347656,68.0487823486328,68.0670928955078,68.0934524536133,68.0605926513672,68.0219192504883,68.0153274536133,68.0006332397461,67.9898986816406,67.9700164794922,67.8787155151367,67.8299331665039,67.7930603027344,67.7838363647461],[67.9908752441406,67.9876022338867,68.0470199584961,68.0979995727539,68.0458450317383,68.0133285522461,67.9908752441406],[67.7359390258789,67.73486328125,67.7501525878906,67.8105010986328,67.8365707397461,67.8670425415039,67.9019470214844,67.9380874633789,68.01025390625,68.0675735473633,68.1002426147461,68.1186981201172,68.1628875732422,68.1905288696289,68.2296905517578,68.3056106567383,68.2876968383789,68.2254867553711,68.1834487915039,68.1387176513672,68.0739288330078,67.9889221191406,67.9272994995117,67.8889694213867,67.8169860839844,67.7835998535156,67.752197265625,67.7359390258789],[68.322509765625,68.2340316772461,68.201904296875,68.1731414794922,68.1412124633789,68.0753860473633,68.0490264892578,67.9852523803711,67.9652328491211,67.7514114379883,67.634765625,67.5373077392578,67.4591827392578,67.4004364013672,67.36669921875,67.2835464477539,67.2620086669922,67.2581024169922,67.23583984375,67.240478515625,67.2502975463867,67.2669372558594,67.3196258544922,67.4083557128906,67.5082015991211,67.6851043701172,67.7061004638672,67.85009765625,67.9470672607422,68.0909652709961,68.2312469482422,68.25439453125,68.2789535522461,68.3187026977539,68.3087921142578,68.3138198852539,68.3323287963867,68.3368148803711,68.322509765625],[68.1928176879883,68.1817932128906,68.2349624633789,68.2644577026367,68.3352584838867,68.35302734375,68.310302734375,68.2681655883789,68.1928176879883],[68.4059066772461,68.4021987915039,68.4182662963867,68.4539489746094,68.4917526245117,68.5315399169922,68.5590362548828,68.57421875,68.581787109375,68.5767135620117,68.5615234375,68.5035171508789,68.470703125,68.4495162963867,68.43115234375,68.4059066772461],[68.3486785888672,68.3422393798828,68.4041519165039,68.4744644165039,68.5254821777344,68.5501480102539,68.5888214111328,68.6360855102539,68.687744140625,68.6960983276367,68.6847152709961,68.6476058959961,68.5446319580078,68.4944381713867,68.4579620361328,68.4407196044922,68.4237747192383,68.413330078125,68.3940887451172,68.365966796875,68.3486785888672],[68.5863265991211,68.5849609375,68.6028289794922,68.6636734008789,68.6819839477539,68.7075729370117,68.740478515625,68.7740249633789,68.8253936767578,68.7989730834961,68.7746124267578,68.75341796875,68.72412109375,68.6521530151367,68.6368637084961,68.5863265991211],[68.8066940307617,68.7750473022461,68.7660675048828,68.7288055419922,68.7238235473633,68.74755859375,68.7861785888672,68.792236328125,68.7906723022461,68.7660675048828,68.7664031982422,68.7829055786133,68.81591796875,68.8652801513672,68.9413528442383,68.9690933227539,68.9901809692383,69.0094223022461,69.0268096923828,69.0350570678711,69.028076171875,68.99755859375,68.9576644897461,68.9261703491211,68.9039077758789,68.8066940307617],[69.0135192871094,68.9540023803711,68.9699249267578,68.991455078125,69.0403823852539,69.0527801513672,69.0714797973633,69.114013671875,69.1294937133789,69.1114730834961,69.0865707397461,69.047119140625,69.0135192871094],[69.2210922241211,69.2094268798828,69.2509231567383,69.2594680786133,69.2871627807617,69.2925796508789,69.3359909057617,69.3573303222656,69.3741683959961,69.3678283691406,69.3246078491211,69.2904281616211,69.2466278076172,69.2210922241211],[68.845458984375,68.8387222290039,68.857666015625,68.8901824951172,68.9230499267578,68.93994140625,68.9559097290039,68.9923324584961,69.0492630004883,69.0874557495117,69.1229019165039,69.1353988647461,69.235107421875,69.252197265625,69.2623519897461,69.2751922607422,69.2997512817383,69.3250961303711,69.3512191772461,69.37060546875,69.3895034790039,69.3860321044922,69.3787155151367,69.3612365722656,69.3145980834961,69.3040008544922,69.2626953125,69.1991653442383,69.1460418701172,69.128662109375,69.1062469482422,69.0790481567383,69.013671875,68.9504852294922,68.9156723022461,68.8829116821289,68.845458984375],[69.4190902709961,69.3570785522461,69.2728958129883,69.2578125,69.252197265625,69.2625961303711,69.3287048339844,69.369873046875,69.3904800415039,69.4162063598633,69.4287109375,69.4179153442383,69.43603515625,69.4190902709961],[69.1437454223633,69.1321258544922,69.1380844116211,69.1616668701172,69.193603515625,69.2740249633789,69.3115234375,69.378662109375,69.4038619995117,69.4163055419922,69.4400863647461,69.4374008178711,69.4117736816406,69.4040069580078,69.3804168701172,69.3661651611328,69.3485870361328,69.3276824951172,69.2667541503906,69.224853515625,69.1746597290039,69.1437454223633],[69.3970718383789,69.3884811401367,69.3905792236328,69.4178237915039,69.4314956665039,69.4412612915039,69.4629440307617,69.4798355102539,69.4928207397461,69.5174331665039,69.5397033691406,69.5592269897461,69.5760803222656,69.5735397338867,69.540771484375,69.4954605102539,69.4619140625,69.4569320678711,69.443359375,69.4147033691406,69.3970718383789],[69.5736312866211,69.506591796875,69.4740753173828,69.4197769165039,69.3784637451172,69.35009765625,69.3358383178711,69.335693359375,69.3475570678711,69.3895492553711,69.4020004272461,69.43896484375,69.5002899169922,69.5380401611328,69.5596160888672,69.5605010986328,69.5405731201172,69.4998016357422,69.447021484375,69.3822326660156,69.3517608642578,69.3671417236328,69.3918914794922,69.4327087402344,69.4696731567383,69.5088424682617,69.5670394897461,69.6060104370117,69.625732421875,69.6243209838867,69.601806640625,69.5736312866211],[69.576904296875,69.539306640625,69.5785598754883,69.5978546142578,69.6496124267578,69.6476593017578,69.616943359375,69.5896987915039,69.576904296875],[69.5409698486328,69.5348663330078,69.5804214477539,69.6167526245117,69.6570281982422,69.6787643432617,69.6818389892578,69.6749954223633,69.6314392089844,69.5917434692383,69.5409698486328],[69.7148971557617,69.6648864746094,69.6389617919922,69.6083984375,69.5518112182617,69.5025329589844,69.4915542602539,69.5026397705078,69.479736328125,69.4828109741211,69.5231399536133,69.6388168334961,69.650634765625,69.6748046875,69.6671371459961,69.6871566772461,69.7168502807617,69.739501953125,69.7392120361328,69.7148971557617],[69.7877960205078,69.7304153442383,69.7123565673828,69.6851577758789,69.630859375,69.6086883544922,69.6343231201172,69.5563430786133,69.5234832763672,69.5096664428711,69.5138702392578,69.5359344482422,69.5624008178711,69.5931625366211,69.6000061035156,69.5828628540039,69.5867691040039,69.6326141357422,69.6497039794922,69.677001953125,69.6892623901367,69.7103500366211,69.7404327392578,69.7505874633789,69.7371139526367,69.7447738647461,69.78271484375,69.797607421875,69.7937927246094,69.8019485473633,69.7824249267578,69.7372589111328,69.7455062866211,69.7389602661133,69.7555236816406,69.7957000732422,69.8104934692383,69.7877960205078],[69.6426773071289,69.6307601928711,69.640869140625,69.6734924316406,69.6796340942383,69.6734924316406,69.614990234375,69.5536117553711,69.510009765625,69.4710922241211,69.3443832397461,69.2586898803711,69.1254425048828,69.0237350463867,68.9535675048828,68.8976516723633,68.8351135253906,68.880615234375,68.8921356201172,68.8731994628906,68.8260726928711,68.8050308227539,68.7783737182617,68.7472686767578,68.7450180053711,68.73583984375,68.6864700317383,68.6272506713867,68.6072769165039,68.5079116821289,68.470703125,68.4608383178711,68.5386734008789,68.5277328491211,68.543701171875,68.6259307861328,68.6724624633789,68.7393569946289,68.749267578125,68.7718734741211,68.8071746826172,68.8307571411133,68.8427276611328,68.8417053222656,68.818359375,68.7982406616211,68.8027801513672,68.8167419433594,68.838623046875,68.8643569946289,68.893798828125,68.9164581298828,68.9324188232422,68.932861328125,68.9176712036133,68.8988800048828,68.8765640258789,68.8633346557617,68.8631820678711,68.8762664794922,68.9176712036133,68.9595718383789,69.0341262817383,69.0542984008789,69.099609375,69.1312026977539,69.1497573852539,69.1675796508789,69.2191390991211,69.3082962036133,69.3346710205078,69.3540573120117,69.375,69.4263153076172,69.46142578125,69.47802734375,69.4795379638672,69.4993667602539,69.5274429321289,69.5449752807617,69.5728988647461,69.5790557861328,69.5650405883789,69.4845199584961,69.4688491821289,69.4566345214844,69.51220703125,69.6290054321289,69.6692886352539,69.6916961669922,69.7544403076172,69.780029296875,69.7969741821289,69.8330535888672,69.8582534790039,69.8616180419922,69.8412628173828,69.8022003173828,69.7384796142578,69.71240234375,69.7002410888672,69.6827163696289,69.6667938232422,69.6426773071289],[70.1132278442383,70.1052703857422,70.1150436401367,70.105712890625,70.0772476196289,70.0476531982422,70.0170440673828,69.9953155517578,69.976318359375,69.9857482910156,70.0111312866211,69.9998550415039,70.0185546875,70.0439453125,70.0801239013672,70.1022491455078,70.1272430419922,70.1466751098633,70.1132278442383],[70.4958038330078,70.4876937866211,70.5250015258789,70.5469284057617,70.5631332397461,70.5962448120117,70.6461868286133,70.6703109741211,70.6686096191406,70.6494140625,70.5946273803711,70.578369140625,70.5424346923828,70.5083999633789,70.4958038330078],[45.0038566589355,44.9990730285645,45.00390625,44.9701156616211,44.8993644714355,44.7722663879395,44.4970703125,44.468017578125,44.4169921875,44.3625984191895,44.303955078125,44.2422370910645,44.214111328125,44.0576171875,43.92431640625,43.7848167419434,43.6288108825684,43.6268577575684,43.6274909973145,43.6286125183105,43.6295394897461,43.6306648254395,43.6314926147461,43.6249504089355,43.5833473205566,43.5271492004395,43.466552734375,43.3313980102539,43.278076171875,43.1061019897461,43.0873069763184,43.061767578125,43.017333984375,42.9970207214355,42.9806137084961,42.9613304138184,42.9352035522461,42.9091300964355,42.8637199401855,42.8023452758789,42.74853515625,42.6514663696289,42.5389633178711,42.4414520263672,42.3660125732422,42.2997550964355,42.2471694946289,42.2091789245605,42.1034698486328,41.98681640625,41.8887214660645,41.7787094116211,41.6748542785645,41.6751937866211,41.7530288696289,41.8329582214355,41.9758796691895,42.1419410705566,42.2506828308105,42.30029296875,42.3317375183105,42.3852043151855,42.4934577941895,42.5580558776855,42.6247062683105,42.739501953125,43.0173835754395,43.0726547241211,43.2632293701172,43.4740715026855,43.5708999633789,43.8222160339355,44.0153350830078,44.1922340393066,44.3915557861328,44.572998046875,44.7439460754395,44.91552734375,45.083740234375,45.2043952941895,45.3473625183105,45.4477081298828,45.5179672241211,45.6327629089355,45.7290534973145,45.817138671875,45.9946784973145,46.0237312316895,46.0586891174316,46.1168479919434,46.1227531433105,46.1090812683105,46.0729026794434,46.0849113464355,46.1470222473145,46.2265167236328,46.2886238098145,46.3708000183105,46.4447746276855,46.4835929870605,46.5029296875,46.5272445678711,46.5427742004395,46.549560546875,46.5185050964355,46.515625,46.4981460571289,46.4618682861328,46.4573707580566,46.5432624816895,46.6373023986816,46.766845703125,46.89990234375,46.9799346923828,47.0599632263184,47.1399421691895,47.219970703125,47.2999992370605,47.3800277709961,47.4600563049316,47.5400886535645,47.5666007995605,47.6364288330078,47.7352066040039,47.8484878540039,47.9617652893066,48.060546875,48.13037109375,48.1568832397461,48.2253913879395,48.3030776977539,48.2640113830566,48.1557121276855,48.0937995910645,48.0474128723145,48.0199699401855,47.9962425231934,47.9999008178711,48.0153312683105,47.9954605102539,48.0153312683105,48.0781745910645,48.1181144714355,48.0991706848145,48.1125984191895,48.1045875549316,48.1310539245605,48.2005348205566,48.2091331481934,48.1937026977539,48.1045875549316,48.0585479736328,48.0583000183105,48.1045875549316,48.1975593566895,48.3018531799316,48.33837890625,48.3289070129395,48.276611328125,48.276611328125,48.3658714294434,48.4353523254395,48.465087890625,48.5318374633789,48.5677757263184,48.61181640625,48.619873046875,48.6253433227539,48.6288566589355,48.6165504455566,48.561279296875,48.5369148254395,48.5254402160645,48.5489273071289,48.6072769165039,48.6590309143066,48.7041015625,48.7426261901855,48.7744140625,48.8084983825684,48.8629875183105,48.8634300231934,49.0029296875,49.1191902160645,49.2585945129395,49.3045921325684,49.3190422058105,49.3494148254395,49.3696746826172,49.2030792236328,48.9917488098145,48.9931640625,48.9931640625,48.9931640625,48.9931640625,48.9931640625,48.9931640625,48.9931640625,48.9931640625,48.9931640625,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930191040039,48.9930191040039,48.9930191040039,48.9930191040039,48.9930191040039,48.9930191040039,48.9930191040039,48.9930191040039,48.9930191040039,49.0284194946289,49.0746574401855,49.0746078491211,49.0608901977539,49.0385246276855,48.9930191040039,48.9777374267578,48.980224609375,48.9930191040039,49.0563468933105,49.0846176147461,49.1183586120605,49.130615234375,49.1210441589355,49.1294937133789,49.147705078125,49.2195320129395,49.260498046875,49.2777366638184,49.2915534973145,49.2932624816895,49.3231925964355,49.39892578125,49.329345703125,49.3221702575684,49.3481903076172,49.3439483642578,49.3594703674316,49.3749542236328,49.3904762268066,49.4430198669434,49.5904808044434,49.644287109375,49.6735343933105,49.6803245544434,49.5776863098145,49.5451164245605,49.5169944763184,49.4591789245605,49.44189453125,49.4513168334961,49.4024391174316,49.3973159790039,49.4828643798828,49.4947280883789,49.5347175598145,49.6028785705566,49.6617164611816,49.7113304138184,49.7361793518066,49.7361793518066,49.717529296875,49.6366691589355,49.5865707397461,49.5935554504395,49.6575698852539,49.6812477111816,49.6569328308105,49.6584968566895,49.6851539611816,49.73681640625,49.7954597473145,49.9811515808105,50.0170402526855,50.043701171875,50.0879859924316,50.1067390441895,50.1442375183105,50.1736335754395,50.1882820129395,50.1839370727539,50.1025848388672,50.0720672607422,49.9927749633789,49.9695320129395,49.8920440673828,49.8755836486816,49.8536643981934,49.7926750183105,49.7721176147461,49.7781257629395,49.8082008361816,49.9576683044434,50.0201187133789,50.0728034973145,50.258056640625,50.2978973388672,50.3556137084961,50.36376953125,50.4186515808105,50.5374031066895,50.6373062133789,50.6686515808105,50.7173347473145,50.8256340026855,50.8724136352539,50.8105964660645,50.7646980285645,50.7184066772461,50.6656723022461,50.5919418334961,50.5138664245605,50.476318359375,50.4971656799316,50.5072746276855,50.5341300964355,50.6490211486816,50.6348648071289,50.5736351013184,50.4860382080078,50.4662132263184,50.4645538330078,50.4785652160645,50.5082054138184,50.5106468200684,50.4873542785645,50.4967308044434,50.4976081848145,50.5232925415039,50.5298843383789,50.5497093200684,50.5877456665039,50.6069831848145,50.6238288879395,50.664306640625,50.6843757629395,50.7049293518066,50.7113800048828,50.666748046875,50.672119140625,50.6793975830078,50.7244606018066,50.7672882080078,50.8070793151855,50.8373527526855,50.8501930236816,50.8418464660645,50.8660659790039,50.9604988098145,51.0568351745605,50.9654769897461,50.9151344299316,50.8936996459961,50.8667945861816,50.8675308227539,50.9160652160645,50.945556640625,50.9894027709961,51.0875473022461,51.1511688232422,51.2686538696289,51.3434562683105,51.4272956848145,51.6080551147461,51.6423835754395,51.6541023254395,51.669921875,51.692626953125,51.7034187316895,51.7167015075684,51.7073745727539,51.678955078125,51.5629386901855,51.5140609741211,51.4784660339355,51.4775886535645,51.4901885986328,51.5055160522461,51.5435562133789,51.6039047241211,51.6731948852539,51.7752456665039,51.82080078125,51.8790016174316,51.9932136535645,51.9902839660645,52.0864753723145,52.1910171508789,52.2529296875,52.297607421875,52.3561515808105,52.3951187133789,52.3148422241211,52.2906761169434,52.2545394897461,52.1883316040039,52.1251449584961,52.06494140625,52.0606918334961,52.1123504638672,52.2254867553711,52.2653312683105,52.30859375,52.3709449768066,52.3948745727539,52.4576644897461,52.4982414245605,52.5376968383789,52.6579093933105,52.7212409973145,52.7510261535645,52.78466796875,52.8425788879395,52.8424797058105,52.7545928955078,52.719970703125,52.6817359924316,52.6526870727539,52.6328125,52.3592758178711,52.3432121276855,52.3185043334961,52.2893562316895,52.2509765625,52.15087890625,51.9505386352539,51.7884254455566,51.998291015625,52.1588859558105,52.3181648254395,52.3424835205078,52.4275398254395,52.4533195495605,52.4906730651855,52.545166015625,52.5507354736328,52.5311508178711,52.4079132080078,52.3682632446289,52.3629913330078,52.4354972839355,52.623291015625,52.8058128356934,52.8580551147461,52.9106903076172,52.9068832397461,52.8257827758789,52.8766098022461,53.1406745910645,53.2438468933105,53.328125,53.3672866821289,53.4422874450684,53.5335960388184,53.64111328125,53.692138671875,53.7151870727539,53.7045402526855,53.6651840209961,53.559326171875,53.5494155883789,53.420654296875,53.4103050231934,53.4598159790039,53.4578628540039,53.4177703857422,53.3694343566895,53.2747077941895,53.329833984375,53.4459457397461,53.4832038879395,53.4903793334961,53.4708976745605,53.4765625,53.506103515625,53.5545921325684,53.6608390808105,53.710205078125,53.7468757629395,53.7801742553711,53.8100090026855,53.8316421508789,53.8450698852539,53.8581047058105,53.9188499450684,53.9186019897461,53.8297882080078,53.8228034973145,53.8400382995605,53.8414573669434,53.7974624633789,53.7777862548828,53.6416015625,53.576416015625,53.4790534973145,53.41796875,53.3931617736816,53.3465805053711,53.25146484375,53.3335456848145,53.4127464294434,53.5513687133789,53.5756340026855,53.6541519165039,53.7239265441895,53.867431640625,53.9757804870605,54.1056632995605,54.133544921875,54.165771484375,54.2302742004395,54.2361297607422,54.2263679504395,54.181396484375,54.2703590393066,54.3516616821289,54.4209976196289,54.4796371459961,54.5393524169922,54.6200180053711,54.6553230285645,54.7002944946289,54.7302703857422,54.82275390625,54.8872566223145,55.0810546875,55.1646461486816,55.2804679870605,55.4625511169434,55.4522476196289,55.4366722106934,55.4385719299316,55.4580078125,55.4982414245605,55.5595703125,55.5326156616211,55.4175758361816,55.319091796875,55.2506332397461,55.1114730834961,55.0572738647461,55.1077613830566,55.1947746276855,55.2640609741211,55.3588371276855,55.471923828125,55.5628890991211,55.6318359375,55.6947746276855,55.751708984375,55.813720703125,55.8807640075684,55.8882293701172,55.9505348205566,56.0144996643066,56.0652351379395,56.1092758178711,56.0828132629395,56.1225090026855,56.2305679321289,56.263671875,56.3408203125,56.3786163330078,56.4048309326172,56.44921875,56.501220703125,56.5567398071289,56.5988273620605,56.5960960388184,56.5899887084961,56.684814453125,56.7421379089355,56.7928199768066,56.8187026977539,56.8567848205566,56.953369140625,57.0265655517578,57.0481910705566,57.0794448852539,57.1453628540039,57.1985359191895,57.2763175964355,57.4067344665527,57.4999008178711,57.6451148986816,57.772705078125,57.8770027160645,57.9489784240723,58.0777320861816,58.2228546142578,58.3370628356934,58.410888671875,58.5034675598145,58.59716796875,58.7050323486328,58.7578620910645,58.7955055236816,58.8498992919922,58.8984832763672,58.939697265625,58.9687538146973,59.0091819763184,59.0562477111816,59.0853500366211,59.1553192138672,59.1992645263672,59.25,59.2711944580078,59.2882804870605,59.4414558410645,59.4960441589355,59.5506858825684,59.5786590576172,59.6950187683105,59.7433090209961,59.7932586669922,59.728759765625,59.6626434326172,59.6383781433105,59.6048355102539,59.5329132080078,59.4803199768066,59.4560508728027,59.4590835571289,59.2799301147461,59.1522445678711,59.1500473022461,59.1061019897461,59.0409698486328,58.9881858825684,58.9031295776367,58.9153823852539,58.9912109375,59.1194343566895,59.2262725830078,59.2811508178711,59.3735847473145,59.4429168701172,59.5419425964355,59.6111335754395,59.6833953857422,59.7782745361328,59.9013175964355,59.9457511901855,59.9932594299316,60.0835952758789,60.1727027893066,60.2794456481934,60.3437042236328,60.3397483825684,60.3336906433105,60.3283233642578,60.2528839111328,60.1831550598145,60.2374992370605,60.2997093200684,60.2183570861816,60.2591323852539,60.3002433776855,60.5924301147461,60.8846664428711,61.1768569946289,61.4690437316895,61.7612838745117,62.0534629821777,62.3456993103027,62.6378898620605,62.9300765991211,63.2222671508789,63.5144538879395,63.8066902160645,64.098876953125,64.39111328125,64.6832962036133,64.9755325317383,65.2677230834961,65.5599136352539,65.8521499633789,66.1443405151367,66.4365234375,66.728759765625,67.0209426879883,67.3131332397461,67.6053695678711,67.8975601196289,68.1897430419922,68.4819869995117,68.774169921875,69.0663604736328,69.3585968017578,69.6507797241211,69.63525390625,69.6024932861328,69.6217269897461,69.5155258178711,69.3167953491211,69.2190399169922,69.1519546508789,69.0928192138672,68.964111328125,68.9508743286133,68.88916015625,68.8973159790039,68.8822250366211,68.8326187133789,68.6964340209961,68.684326171875,68.6942825317383,68.8289566040039,68.8419952392578,68.8922348022461,68.9169921875,68.9267120361328,68.9741668701172,68.9926300048828,69.0010223388672,69.0006332397461,69.0082473754883,69.0269470214844,69.0313034057617,69.0494155883789,69.0813903808594,69.1114730834961,69.3111801147461,69.291015625,69.337158203125,69.307861328125,69.3238296508789,69.4252014160156,69.4496078491211,69.4678268432617,69.4858932495117,69.4679183959961,69.4776306152344,69.5453109741211,69.5719223022461,69.6328125,69.6654739379883,69.6817855834961,69.6688461303711,69.6388168334961,69.5872573852539,69.557861328125,69.5282211303711,69.5077133178711,69.4294967651367,69.3884811401367,69.2805709838867,69.2528305053711,69.3013229370117,69.368408203125,69.4053726196289,69.4121627807617,69.4338836669922,69.4706497192383,69.5082550048828,69.629638671875,69.6506805419922,69.6432571411133,69.6469268798828,69.6587448120117,69.674072265625,69.69287109375,69.6981430053711,69.7066879272461,69.7265625,69.7380828857422,69.7518005371094,69.7081527709961,69.7049865722656,69.7534637451172,69.8821334838867,69.9179229736328,69.9241714477539,69.9007873535156,69.9068832397461,69.9794921875,70.0181121826172,70.0516052246094,70.1270446777344,70.1431655883789,70.1292495727539,70.0979995727539,70.0858917236328,70.0950698852539,70.0909194946289,70.1061553955078,70.1920928955078,70.1929626464844,70.1676254272461,70.1051788330078,70.0739212036133,69.9977493286133,69.7799758911133,69.6859893798828,69.6514587402344,69.6320266723633,69.6157760620117,69.5966262817383,69.5794906616211,69.5493621826172,69.5347137451172,69.5176239013672,69.40234375,69.3646926879883,69.3079605102539,69.2731475830078,69.2598571777344,69.2057647705078,69.1014099121094,69.0287628173828,68.9671401977539,68.8866729736328,68.8442840576172,68.7884750366211,68.7698745727539,68.7398452758789,68.74658203125,68.7765121459961,68.8197326660156,68.8352508544922,68.847412109375,68.8148956298828,68.8478012084961,68.8756332397461,68.8899459838867,68.8958969116211,68.9224624633789,68.9724578857422,69.0121536254883,69.0416030883789,69.0792007446289,69.140625,69.1669387817383,69.20166015625,69.2344741821289,69.29052734375,69.3359909057617,69.3712844848633,69.3888702392578,69.401611328125,69.4319839477539,69.4590835571289,69.4613800048828,69.4354019165039,69.4150924682617,69.3611755371094,69.3637161254883,69.4321823120117,69.4549789428711,69.45947265625,69.45068359375,69.4287109375,69.3628921508789,69.2532653808594,69.2090835571289,69.2848663330078,69.3200225830078,69.4813003540039,69.5696792602539,69.6558151245117,69.7200698852539,69.8267135620117,69.8554153442383,69.8819351196289,69.9049758911133,69.9334487915039,69.9661636352539,69.9634780883789,69.8948669433594,69.875,69.83251953125,69.8001022338867,69.7500457763672,69.717041015625,69.7010726928711,69.71240234375,69.7510223388672,69.8101577758789,69.9601593017578,69.9875946044922,70.1081085205078,70.1613311767578,70.2218780517578,70.2603530883789,70.2929229736328,70.3153305053711,70.3287582397461,70.3631362915039,70.3973693847656,70.41845703125,70.4797897338867,70.5238265991211,70.56640625,70.5738296508789,70.549072265625,70.5171432495117,70.3687515258789,70.296142578125,70.2393569946289,70.0617141723633,69.9590835571289,69.8533706665039,69.777099609375,69.7303237915039,69.5452651977539,69.4670867919922,69.4185562133789,69.3799819946289,69.3515625,69.3492202758789,69.427978515625,69.4797821044922,69.566162109375,69.6259765625,69.6624526977539,69.7323760986328,69.7567367553711,69.8288040161133,69.8150405883789,69.8178253173828,69.8442840576172,69.8943862915039,69.935791015625,69.968505859375,69.9900436401367,70.0055160522461,70.0055694580078,70.0125961303711,70.026611328125,70.041748046875,70.0801773071289,70.1169967651367,70.1414566040039,70.1512222290039,70.14111328125,70.110595703125,70.0619125366211,69.9825668334961,69.9185104370117,69.7674331665039,69.7345275878906,69.6899948120117,69.6531753540039,69.4938507080078,69.454833984375,69.4251403808594,69.4000473022461,69.3794403076172,69.3648452758789,69.35888671875,69.3728485107422,69.37744140625,69.3893508911133,69.4200210571289,69.4966278076172,69.52001953125,69.54150390625,69.6324768066406,69.7381362915039,69.782470703125,69.8100128173828,69.81884765625,69.8084487915039,69.8173828125,69.8084487915039,69.816162109375,69.7975082397461,69.7757797241211,69.7415542602539,69.660400390625,69.6168441772461,69.4205551147461,69.3805618286133,69.3423309326172,69.2571792602539,69.2342758178711,69.1448745727539,69.0927276611328,69.0429229736328,68.9999008178711,68.9349136352539,68.9134292602539,68.9071273803711,68.9159698486328,68.8788070678711,68.880615234375,68.8736343383789,68.8468246459961,68.8370132446289,68.8554153442383,68.9579086303711,68.9740676879883,68.975341796875,68.9581069946289,68.9873046875,68.9866180419922,68.9725646972656,68.94091796875,68.891845703125,68.8500518798828,68.74609375,68.6595687866211,68.5520553588867,68.4773483276367,68.4354019165039,68.4146499633789,68.4149932861328,68.3990707397461,68.3064956665039,68.2833938598633,68.2667999267578,68.2478485107422,68.2702102661133,68.1952667236328,68.1320343017578,68.1043930053711,68.0841827392578,68.0185546875,67.9984436035156,67.9235305786133,67.90234375,67.8716812133789,67.8191909790039,67.806396484375,67.8135757446289,67.7952117919922,67.751220703125,67.7311553955078,67.7350082397461,67.7269058227539,67.7068862915039,67.699951171875,67.7017593383789,67.6866683959961,67.6798782348633,67.6819381713867,67.6847610473633,67.7195739746094,67.7311096191406,67.7317352294922,67.75732421875,67.7568359375,67.7761688232422,67.8152313232422,67.8225555419922,67.7982406616211,67.7876434326172,67.7908248901367,67.8323211669922,67.9542007446289,67.9540023803711,67.9922409057617,67.992919921875,67.9771957397461,67.8878936767578,67.87353515625,67.8658218383789,67.8201141357422,67.7517623901367,67.7327194213867,67.7297897338867,67.7107391357422,67.691162109375,67.6371078491211,67.5323715209961,67.4939498901367,67.4380874633789,67.4219741821289,67.4375,67.5828094482422,67.606201171875,67.5980453491211,67.5908660888672,67.4832992553711,67.4034194946289,67.2563934326172,67.2024917602539,67.1625442504883,67.1268081665039,67.0951690673828,67.0697708129883,67.0505905151367,67.0575180053711,67.0904312133789,67.09228515625,67.06298828125,66.9412536621094,66.8926239013672,66.8603515625,66.8443374633789,66.8180160522461,66.7813034057617,66.6836929321289,66.6371078491211,66.4917984008789,66.4346694946289,66.4018096923828,66.3985366821289,66.4249038696289,66.6184997558594,66.7400360107422,66.7691879272461,66.8137741088867,66.9614715576172,66.984130859375,67.0031280517578,66.9361801147461,66.9267578125,66.9307098388672,66.947998046875,66.9319839477539,66.882568359375,66.8817367553711,66.9763717651367,67.0225524902344,67.0547866821289,67.103271484375,67.1277847290039,67.1991271972656,67.2730484008789,67.384765625,67.42822265625,67.4742126464844,67.5112762451172,67.5868682861328,67.6392059326172,67.7000045776367,67.7320251464844,67.818603515625,67.8563461303711,67.9068298339844,67.9588394165039,68.0125045776367,68.0369110107422,68.0321807861328,68.0591354370117,68.0496597290039,68.0611801147461,68.0937957763672,68.1084442138672,68.1063003540039,68.1141586303711,68.1286163330078,68.1448211669922,68.206787109375,68.2160186767578,68.2092742919922,68.1956558227539,68.2005844116211,68.2884750366211,68.3193359375,68.3832015991211,68.389892578125,68.4073257446289,68.443115234375,68.4751434326172,68.5265655517578,68.59228515625,68.6111373901367,68.6365203857422,68.6233444213867,68.5765609741211,68.5164566040039,68.4605941772461,68.3889617919922,68.3573760986328,68.3873062133789,68.3868179321289,68.3743667602539,68.3468246459961,68.30419921875,68.2964324951172,68.3235397338867,68.3310546875,68.2857437133789,68.2520523071289,68.2029266357422,68.1737289428711,68.1629409790039,68.1692886352539,68.14990234375,68.1540985107422,68.1775436401367,68.27734375,68.2974624633789,68.3785095214844,68.5978012084961,68.6107940673828,68.64892578125,68.6888732910156,68.8094711303711,68.8194808959961,68.8994674682617,68.9198760986328,68.9060592651367,68.8647918701172,68.8281707763672,68.7824249267578,68.7186508178711,68.5780792236328,68.458251953125,68.413818359375,68.3303756713867,68.2979965209961,68.2878875732422,68.3074264526367,68.310546875,68.3030242919922,68.25048828125,68.2452621459961,68.2517547607422,68.2300796508789,68.2139205932617,68.1487808227539,68.1214828491211,68.0631866455078,68.0412139892578,68.0311965942383,68.0410690307617,68.069091796875,68.1150360107422,68.0638198852539,67.9402313232422,67.8527297973633,67.8115692138672,67.76220703125,67.7356414794922,67.7327194213867,67.7533187866211,67.7453155517578,67.7086410522461,67.6931610107422,67.7623519897461,67.7656707763672,67.7989807128906,67.80908203125,67.8082504272461,67.8184127807617,67.8394546508789,67.8385696411133,67.8148422241211,67.7840805053711,67.7453155517578,67.7236328125,67.7188491821289,67.7257766723633,67.7444305419922,67.7797393798828,67.7978973388672,67.8071823120117,67.8264617919922,67.8558120727539,67.9114227294922,67.9657211303711,68.0001983642578,68.0419921875,68.0661163330078,68.0725631713867,68.046630859375,67.9884262084961,67.7696762084961,67.7386169433594,67.7107925415039,67.6310577392578,67.6169967651367,67.666259765625,67.6969223022461,67.7264175415039,67.7548370361328,67.7962417602539,67.8219299316406,67.8550796508789,67.9013671875,67.9607467651367,67.9781723022461,67.9535675048828,67.9030303955078,67.9230041503906,68.064697265625,68.11767578125,68.1322784423828,68.1153335571289,68.1324691772461,68.2007827758789,68.2236328125,68.3311538696289,68.363525390625,68.370849609375,68.3833999633789,68.3821258544922,68.3174285888672,68.3463363647461,68.3875961303711,68.4495162963867,68.5104522705078,68.523681640625,68.5327682495117,68.4819869995117,68.4749450683594,68.4951629638672,68.4965286254883,68.4791488647461,68.4529266357422,68.3779830932617,68.3328628540039,68.2649383544922,68.2554168701172,68.2502975463867,68.3105926513672,68.2900924682617,68.2428207397461,68.0612335205078,68.0387725830078,68.0484390258789,68.0631332397461,68.0849609375,68.1358413696289,68.15087890625,68.2365188598633,68.2491226196289,68.1577606201172,67.8316879272461,67.7178192138672,67.67919921875,67.5538558959961,67.509765625,67.404296875,67.3755874633789,67.2889709472656,67.2718276977539,67.2708969116211,67.2984924316406,67.3167953491211,67.2987289428711,67.1937942504883,67.1846160888672,67.2115783691406,67.2152862548828,67.2091751098633,67.1555709838867,67.1155776977539,67.0561065673828,67.0132293701172,66.9894485473633,66.9798812866211,66.9727478027344,66.9756774902344,66.9666976928711,66.9781723022461,67.0108871459961,67.0187530517578,66.9935531616211,66.9977111816406,67.0700225830078,67.063232421875,67.0517578125,67.0361862182617,66.9894027709961,66.7413558959961,66.6901397705078,66.6165466308594,66.6167984008789,66.6904296875,66.72607421875,66.8704605102539,66.9231414794922,66.9375,66.9522476196289,66.916259765625,66.9241180419922,66.949462890625,66.980712890625,67.1524963378906,67.2625503540039,67.3610305786133,67.517822265625,67.6102066040039,67.703857421875,67.7374496459961,68.0213928222656,68.0452575683594,68.0555648803711,68.0597152709961,68.0832977294922,68.05029296875,68.0416488647461,68.0708999633789,68.1900863647461,68.2269973754883,68.2968215942383,68.3994140625,68.4738311767578,68.5431137084961,68.598876953125,68.618896484375,68.6236801147461,68.6331100463867,68.6859893798828,68.7837448120117,68.8381881713867,68.8872604370117,68.9310531616211,68.9695816040039,68.99267578125,69.0003433227539,68.996826171875,68.982177734375,68.8890609741211,68.8206024169922,68.7847671508789,68.7605438232422,68.7427749633789,68.7755355834961,68.80322265625,68.9116668701172,68.9581604003906,69.0497589111328,69.1231002807617,69.1358337402344,69.1363754272461,69.1514663696289,69.2416000366211,69.2752380371094,69.3137741088867,69.3417434692383,69.3763656616211,69.4169921875,69.40283203125,69.2809143066406,69.2526321411133,69.2261276245117,69.296875,69.3551788330078,69.3750534057617,69.38232421875,69.4064407348633,69.45068359375,69.4809036254883,69.51904296875,69.4978485107422,69.4576721191406,69.4467239379883,69.4459457397461,69.4551315307617,69.4742660522461,69.5170440673828,69.5834503173828,69.649658203125,69.6568832397461,69.6494140625,69.58544921875,69.5777816772461,69.6673812866211,69.7176284790039,69.7557144165039,69.7782211303711,69.7851104736328,69.8027877807617,69.8311538696289,69.8718719482422,69.9249496459961,69.9917984008789,70.1249008178711,70.2103042602539,70.2430191040039,70.3272476196289,70.4701614379883,70.5113754272461,70.5417022705078,70.5612258911133,70.5670928955078,70.5489730834961,70.5932159423828,70.6168441772461,70.6942901611328,70.69775390625,70.6382827758789,70.6422805786133,70.6786575317383,70.8087387084961,70.8897476196289,71.0023422241211,71.0697250366211,71.1270523071289,71.1431579589844,71.1592254638672,71.1764678955078,71.2398910522461,71.2736282348633,71.339111328125,71.3963851928711,71.4138717651367,71.41064453125,71.39306640625,71.361083984375,71.3281707763672,71.3188018798828,71.3367691040039,71.4600601196289,71.4916534423828,71.50537109375,71.5040588378906,71.5142593383789,71.5260772705078,71.5731430053711,71.5982437133789,71.6854019165039,71.7768020629883,71.9037094116211,71.96337890625,71.9829635620117,71.9868698120117,71.9789581298828,71.91552734375,71.8485870361328,71.764892578125,71.7584457397461,71.7711410522461,71.7662582397461,71.7428283691406,71.7166519165039,71.67431640625,71.6380386352539,71.5687026977539,71.5206985473633,71.4608383178711,71.335693359375,71.3003463745117,71.2621078491211,71.1223602294922,71.0693359375,70.9160614013672,70.8871078491211,70.8522415161133,70.838134765625,70.7981414794922,70.7188034057617,70.6932144165039,70.6504364013672,70.63427734375,70.6075210571289,70.492919921875,70.3896484375,70.3673782348633,70.3187484741211,70.3033218383789,70.2855529785156,70.2947769165039,70.3311538696289,70.3416519165039,70.3262710571289,70.2992172241211,70.2329559326172,70.1782760620117,70.1615753173828,70.1478500366211,70.1326675415039,70.1432113647461,70.169921875,70.197509765625,70.2398452758789,70.2353515625,70.2008285522461,70.1504440307617,70.1038589477539,70.0831604003906,70.0845260620117,70.0714340209961,70.0386734008789,69.9839859008789,69.8921356201172,69.7139205932617,69.668212890625,69.6548843383789,69.6514587402344,69.6592788696289,69.6832046508789,69.6729049682617,69.6533660888672,69.6343231201172,69.6033248901367,69.53125,69.5456085205078,69.6150360107422,69.6494598388672,69.644775390625,69.6371612548828,69.6203079223633,69.581298828125,69.5379409790039,69.5256805419922,69.543212890625,69.5154800415039,69.5085983276367,69.5155258178711,69.5044937133789,69.4754409790039,69.4569778442383,69.4451141357422,69.4453125,69.427734375,69.3924789428711,69.3467254638672,69.2904815673828,69.2672805786133,69.277099609375,69.2930221557617,69.3152313232422,69.3184051513672,69.3025894165039,69.3021011352539,69.2855453491211,69.1059036254883,68.9468765258789,68.8811569213867,68.86376953125,68.819580078125,68.7859878540039,68.6888732910156,68.6112823486328,68.4747085571289,68.4322280883789,68.3947296142578,68.3467330932617,68.330322265625,68.2916564941406,68.2674331665039,68.2574691772461,68.2702102661133,68.3055191040039,68.3385772705078,68.3980484008789,68.4907760620117,68.5215301513672,68.5943908691406,68.6255874633789,68.7359390258789,68.8124465942383,68.9315948486328,69.0146026611328,69.0849151611328,69.2269973754883,69.2554702758789,69.2694854736328,69.26611328125,69.2204055786133,69.1358947753906,69.058837890625,68.9544448852539,68.9150390625,68.8117141723633,68.7092819213867,68.564697265625,68.4776382446289,68.4480972290039,68.4041519165039,68.345703125,68.2999496459961,68.2668914794922,68.2481460571289,68.2420425415039,68.2511672973633,68.2660140991211,68.3348617553711,68.3390579223633,68.2882766723633,68.2598648071289,68.165771484375,67.98876953125,67.9503402709961,67.7658157348633,67.6256790161133,67.3553237915039,67.3246078491211,67.2141647338867,67.1910629272461,67.17724609375,67.1728515625,67.183837890625,67.2677764892578,67.3562545776367,67.4023895263672,67.4061050415039,67.4223098754883,67.4508285522461,67.4821319580078,67.5161590576172,67.6494674682617,67.713134765625,67.8000946044922,67.8248062133789,68.0453643798828,68.0724563598633,68.3280334472656,68.4450149536133,68.5154800415039,68.5782699584961,68.630126953125,68.6709442138672,68.69970703125,68.7288055419922,68.7698211669922,68.7739715576172,68.7742691040039,68.7462921142578,68.7413635253906,68.7733383178711,68.7903823852539,68.8440475463867,68.8709487915039,68.9079055786133,68.94921875,68.9622573852539,68.988525390625,69.02099609375,69.073974609375,69.0927734375,69.1658706665039,69.1627426147461,69.1723175048828,69.2318801879883,69.3184051513672,69.3538589477539,69.410888671875,69.4267578125,69.4524917602539,69.4882354736328,69.5199203491211,69.5477523803711,69.5806655883789,69.6187515258789,69.6515197753906,69.7481460571289,69.7777862548828,69.8190383911133,69.8350601196289,69.8452606201172,69.8495101928711,69.8361282348633,69.8051300048828,69.8047866821289,69.8350601196289,69.8497085571289,69.8437042236328,69.8350067138672,69.745361328125,69.69970703125,69.7039489746094,69.6858825683594,69.6951217651367,69.6910629272461,69.6417999267578,69.6008758544922,69.5322265625,69.5181198120117,69.494384765625,69.4583969116211,69.4100112915039,69.3325653076172,69.2970199584961,69.2649917602539,69.2488784790039,69.2488784790039,69.2760772705078,69.2581024169922,69.1981430053711,69.1856460571289,69.138916015625,69.1199264526367,69.0030212402344,68.9567337036133,68.9090805053711,68.8836364746094,68.8789520263672,68.8655700683594,68.8500518798828,68.8279800415039,68.7806091308594,68.7431640625,68.6924285888672,68.6572265625,68.5559539794922,68.5243682861328,68.4868698120117,68.4587860107422,68.462646484375,68.49853515625,68.5062484741211,68.4775848388672,68.4786148071289,68.4686050415039,68.4464874267578,68.3824234008789,68.357177734375,68.3383331298828,68.3065872192383,68.2965850830078,68.2852554321289,68.145263671875,68.1344223022461,68.1396942138672,68.1796875,68.1959457397461,68.1939010620117,68.1733856201172,68.0514602661133,68.0188980102539,67.9898376464844,67.9281692504883,67.8620147705078,67.802490234375,67.7223663330078,67.6369094848633,67.5953598022461,67.4974136352539,67.4599151611328,67.3569793701172,67.1885681152344,67.0928726196289,67.0698776245117,67.0020065307617,66.986083984375,66.9879379272461,66.9747009277344,66.92041015625,66.8250961303711,66.7646484375,66.7391128540039,66.7094268798828,66.6213836669922,66.5875015258789,66.5508270263672,66.4314956665039,66.3921356201172,66.3712463378906,66.3687515258789,66.3878402709961,66.460693359375,66.484619140625,66.5343780517578,66.6790542602539,66.728515625,66.739501953125,66.731689453125,66.736328125,66.7817840576172,66.8111343383789,66.8225631713867,66.8392028808594,66.8627471923828,66.9274368286133,66.9613342285156,66.9728088378906,67.0166015625,67.0287094116211,66.9560546875,66.9069366455078,66.8909149169922,66.8720703125,66.8812561035156,66.9265670776367,66.940673828125,66.93359375,66.90234375,66.8751449584961,66.8566436767578,66.7118148803711,66.6824645996094,66.6478500366211,66.5902328491211,66.5262222290039,66.4205551147461,66.2899932861328,66.2384796142578,66.2135772705078,66.2117614746094,66.231201171875,66.2917938232422,66.2906723022461,66.2587432861328,66.1862335205078,66.1792907714844,66.2077178955078,66.2713394165039,66.3253479003906,66.3696823120117,66.4403305053711,66.537353515625,66.5682601928711,66.532958984375,66.5203628540039,66.5313491821289,66.5230407714844,66.5108871459961,66.4574737548828,66.4427719116211,66.432861328125,66.4170837402344,66.3604049682617,66.3219223022461,66.2699203491211,66.2252960205078,66.1868133544922,66.1544494628906,66.1190414428711,66.0484924316406,66.0225601196289,65.6705551147461,65.5282745361328,65.4408187866211,65.383056640625,65.3548355102539,65.3389587402344,65.3353500366211,65.3489303588867,65.3945770263672,65.5162124633789,65.587646484375,65.611572265625,65.6787567138672,65.691650390625,65.7030258178711,65.7389678955078,65.8607864379883,65.9093246459961,65.93603515625,65.9408187866211,65.93359375,65.8722610473633,65.8685531616211,65.882568359375,65.8824234008789,65.9263687133789,65.9205017089844,65.9293441772461,65.953857421875,65.9657211303711,65.9645538330078,65.9593734741211,65.9478988647461,65.8944320678711,65.829833984375,65.8855514526367,65.89990234375,65.9192428588867,65.8848190307617,65.8126983642578,65.8056182861328,65.7802734375,65.7367172241211,65.6477584838867,65.4463882446289,65.3956069946289,65.3482894897461,65.2798843383789,65.2803268432617,65.2605438232422,65.2248001098633,65.1980972290039,65.1085968017578,65.0636215209961,64.9268112182617,64.826171875,64.4004364013672,64.302490234375,64.2439498901367,64.1833038330078,64.0892562866211,64.0093765258789,63.9922332763672,64.01123046875,64.034423828125,64.11376953125,64.1054229736328,63.9811019897461,63.968505859375,63.9841346740723,64.0399932861328,64.0296936035156,64.0145034790039,64.0147933959961,64.0306091308594,64.0769577026367,64.0995101928711,64.1682586669922,64.1805725097656,64.140869140625,64.1277389526367,64.1001968383789,64.0806198120117,63.9788055419922,63.9569816589355,63.9435539245605,63.9819831848145,63.9787559509277,63.8774948120117,63.829345703125,63.8042945861816,63.6896476745605,63.6418914794922,63.6244125366211,63.6361808776855,63.6654281616211,63.6612777709961,63.623779296875,63.596923828125,63.5809059143066,63.5878372192383,63.6178245544434,63.7255821228027,63.7422332763672,63.7570762634277,63.7723159790039,63.7868156433105,63.8005867004395,63.8124504089355,63.8224143981934,63.8130378723145,63.784423828125,63.7759742736816,63.7876472473145,63.8295402526855,63.9376449584961,64.02880859375,64.1471710205078,64.0405807495117,64.0044937133789,63.9728012084961,63.9414024353027,63.8652801513672,63.8379859924316,63.8408699035645,63.8533210754395,63.8722686767578,63.9000473022461,63.9412117004395,63.948486328125,63.9269027709961,63.9017601013184,63.76123046875,63.7349128723145,63.7078094482422,63.6916999816895,63.6567878723145,63.5694313049316,63.5550804138184,63.5629920959473,63.6399955749512,63.6756324768066,63.6975593566895,63.6597213745117,63.5622062683105,63.5068321228027,63.4758796691895,63.4427719116211,63.3515625,63.3040504455566,63.1105461120605,63.0638656616211,63.0174827575684,62.9716339111328,62.9459457397461,62.9404335021973,62.9215812683105,62.8188972473145,62.8040542602539,62.8347129821777,62.8634300231934,62.8617210388184,62.8390655517578,62.8288078308105,62.8193893432617,62.8008766174316,62.7724113464355,62.7338409423828,62.711669921875,62.6836433410645,62.665283203125,62.644775390625,62.5923881530762,62.5754890441895,62.5405235290527,62.5440902709961,62.5853500366211,62.5869636535645,62.5646018981934,62.5572242736816,62.5648460388184,62.5467262268066,62.5028839111328,62.4700698852539,62.4182090759277,62.3799819946289,62.3499488830566,62.3282203674316,62.279052734375,62.2022933959961,62.1684074401855,62.1773414611816,62.207763671875,62.2597160339355,62.3062019348145,62.3668479919434,62.3649406433105,62.3495597839355,62.2859344482422,62.2449684143066,62.2151336669922,62.1497573852539,62.1278305053711,62.1086463928223,62.0926742553711,62.060546875,62.0336418151855,62.02978515625,62.0145530700684,61.9815902709961,61.9610862731934,61.9329109191895,61.928955078125,61.9420433044434,61.8716316223145,61.8469276428223,61.8121109008789,61.77978515625,61.7672843933105,61.7395477294922,61.7058067321777,61.6025428771973,61.4814414978027,61.4436492919922,61.3640670776367,61.3440475463867,61.3080101013184,61.3178253173828,61.3036575317383,61.2661628723145,61.2112808227539,61.1388664245605,61.0254402160645,60.8709945678711,60.7307167053223,60.6045417785645,60.5419921875,60.5376930236816,60.4982452392578,60.4775428771973,60.4533195495605,60.4164047241211,60.3010749816895,60.1073722839355,59.9533157348633,59.4781265258789,59.2678718566895,59.1513175964355,59.0879859924316,59.06884765625,59.0503387451172,59.0206069946289,58.9754409790039,58.9033203125,58.8701171875,58.875732421875,58.8684577941895,58.8483924865723,58.7455062866211,58.7160148620605,58.658935546875,58.339111328125,58.2973670959473,58.3780288696289,58.6263694763184,58.7367172241211,58.760009765625,58.7725639343262,58.7444839477539,58.7410125732422,58.7563934326172,58.7256317138672,58.694580078125,58.5644035339355,58.4898452758789,58.2245140075684,58.0758743286133,57.8440437316895,57.7777862548828,57.4686050415039,57.3848648071289,57.3203125,57.2750511169434,57.2052726745605,57.1109390258789,57.0390129089355,56.9895515441895,56.95263671875,56.9283218383789,56.9219741821289,56.9335441589355,56.9583015441895,57.0223159790039,57.0367164611816,57.0458488464355,57.0227546691895,56.9751472473145,56.9706039428711,57.0089836120605,57.0637664794922,57.2412109375,57.2569313049316,57.2244606018066,57.1490745544434,57.0519027709961,56.9813499450684,56.9154319763184,56.8838386535645,56.851318359375,56.8142585754395,56.7250480651855,56.6086883544922,56.5356903076172,56.46728515625,56.3416023254395,56.0563468933105,56.0212898254395,55.9746551513672,55.91455078125,55.7732429504395,55.7178726196289,55.6958999633789,55.6569328308105,55.60107421875,55.5401840209961,55.4742660522461,55.43115234375,55.38330078125,55.3489723205566,55.2974624633789,55.095458984375,55.0792961120605,55.224365234375,55.2662124633789,55.28564453125,55.2833480834961,55.2592277526855,55.2588882446289,55.2825202941895,55.2931137084961,55.2908210754395,55.2978019714355,55.3141593933105,55.3146476745605,55.2645034790039,55.2618179321289,55.214599609375,55.2313957214355,55.2222175598145,55.1606941223145,55.1559066772461,55.16552734375,55.1487312316895,55.0678215026855,54.9981460571289,54.8559112548828,54.8134765625,54.4834976196289,54.3558082580566,54.2445793151855,54.1804695129395,54.072998046875,54.0448265075684,53.8856964111328,53.817626953125,53.7395477294922,53.6109352111816,53.5128402709961,53.3646011352539,53.26416015625,53.2114715576172,53.1598129272461,53.0661125183105,53.0307121276855,52.9611320495605,52.9216804504395,52.8773918151855,52.8116188049316,52.6514129638672,52.5636215209961,52.4326171875,52.3672866821289,52.3240699768066,52.2938957214355,52.2536125183105,52.2242202758789,52.2171897888184,52.2390632629395,52.2367668151855,52.2044906616211,52.1422348022461,52.0892066955566,52.04541015625,51.9722175598145,51.79833984375,51.7583503723145,51.667236328125,51.5250968933105,51.4322242736816,51.3885726928711,51.3446769714355,51.2647476196289,51.125,51.1318359375,51.1908683776855,51.3073272705078,51.3298835754395,51.3163604736328,51.2828598022461,51.2351531982422,51.17333984375,51.0078125,50.8755874633789,50.7626457214355,50.8345184326172,50.9172859191895,51.0490264892578,51.1175765991211,51.1504859924316,51.186279296875,51.2516593933105,51.3465805053711,51.4135246276855,51.4524383544922,51.4938468933105,51.5376930236816,51.5699234008789,51.628173828125,51.622802734375,51.552001953125,51.5373039245605,51.5262184143066,51.501708984375,51.4637680053711,51.4253425598145,51.3863792419434,51.2591323852539,51.2002944946289,51.2716789245605,51.3839378356934,51.4299774169922,51.4974594116211,51.526611328125,51.5657691955566,51.7337875366211,51.7745628356934,51.798828125,51.8451194763184,51.9134254455566,51.9629859924316,52.03271484375,52.1396980285645,52.2132835388184,52.2520980834961,52.2613754272461,52.2911109924316,52.3106956481934,52.399169921875,52.4918975830078,52.5351066589355,52.6277351379395,52.6553688049316,52.7600593566895,52.8124008178711,52.8564453125,52.8989753723145,52.97607421875,53.0433578491211,53.2061996459961,53.4103507995605,53.5605010986328,53.6566429138184,53.7171859741211,53.7422866821289,53.8179702758789,53.8365707397461,53.8315925598145,53.8402328491211,53.8810539245605,53.932373046875,53.951416015625,54.0024909973145,54.0239753723145,54.0519523620605,54.098876953125,54.116943359375,54.1572265625,54.1692352294922,54.1856956481934,54.2168464660645,54.2633819580078,54.3366203308105,54.394775390625,54.4915542602539,54.6016578674316,54.6291046142578,54.6468276977539,54.6549797058105,54.6718254089355,54.6974639892578,54.8814964294434,54.9080085754395,55.0110359191895,55.0685539245605,55.1513175964355,55.2364273071289,55.291259765625,55.3441886901855,55.5555191040039,55.6635246276855,55.7562980651855,55.8672370910645,55.9964332580566,56.1072273254395,56.1995620727539,56.3587913513184,56.4999504089355,56.7069816589355,56.8917961120605,57.1811981201172,57.2722663879395,57.3805694580078,57.5985832214355,57.657958984375,57.7581100463867,58.0188980102539,58.1953163146973,58.2396011352539,58.2913551330566,58.3507308959961,58.399169921875,58.5806617736816,58.6024436950684,58.6491203308105,58.6823768615723,58.7690925598145,58.8291015625,58.8732948303223,58.9017562866211,59.0350608825684,59.1417465209961,59.2001914978027,59.2455062866211,59.3050308227539,59.3800315856934,59.410400390625,59.4435043334961,59.4757843017578,59.5581550598145,59.5809555053711,59.6585006713867,59.6758804321289,59.6805191040039,59.5692367553711,59.5789527893066,59.6096229553223,59.6845703125,59.7156753540039,59.7966346740723,59.8334007263184,59.8843727111816,59.9250946044922,60.0220222473145,60.0423812866211,60.0611305236816,60.0881843566895,60.1009788513184,60.1334991455078,60.1457977294922,60.3624992370605,60.4271011352539,60.5067405700684,60.5427207946777,60.5631866455078,60.56689453125,60.5777816772461,60.6398468017578,60.6790504455566,60.6969757080078,60.7895050048828,60.80859375,60.8251991271973,60.7858428955078,60.8182106018066,60.8096199035645,60.8191375732422,60.8521957397461,61.0026397705078,61.0840339660645,61.1575202941895,61.2063980102539,61.2306632995605,61.3930206298828,61.4373550415039,61.4786643981934,61.5563011169434,61.6264114379883,61.6947746276855,61.7287101745605,61.7618637084961,61.8320770263672,61.9233894348145,62.1073760986328,62.2086944580078,62.2822761535645,62.3181114196777,62.3554191589355,62.4265632629395,62.5313987731934,62.572509765625,62.5499496459961,62.525390625,62.4656715393066,62.3158645629883,62.24951171875,62.1934089660645,62.1795883178711,62.2864265441895,62.3070831298828,62.3121109008789,62.270751953125,62.2644538879395,62.2300300598145,62.1156730651855,62.1251945495605,62.1834487915039,62.2111320495605,62.2718238830566,62.3213844299316,62.3700218200684,62.4343757629395,62.4687538146973,62.47314453125,62.4541969299316,62.3688468933105,62.3250465393066,62.2791481018066,62.1982421875,62.1804237365723,62.1253929138184,62.131103515625,62.1245574951172,62.1138687133789,62.0766105651855,62.0527801513672,62.0272445678711,61.955322265625,61.8404312133789,61.8386192321777,61.8632278442383,61.9071273803711,61.9226608276367,61.8877906799316,61.8315925598145,61.8018074035645,61.7532234191895,61.728271484375,61.6802749633789,61.6646957397461,61.6414070129395,61.60205078125,61.5872573852539,61.6119613647461,61.6362762451172,61.6769485473633,61.6885261535645,61.6171836853027,61.5923843383789,61.5728988647461,61.5509262084961,61.5267601013184,61.4660148620605,61.4397926330566,61.4211921691895,61.4131355285645,61.3987274169922,61.3720703125,61.3372573852539,61.2132835388184,61.158935546875,61.1489753723145,61.1465301513672,61.1255340576172,61.05517578125,61.04248046875,61.0639686584473,61.0686531066895,61.04052734375,61.0206565856934,60.9811019897461,60.9218254089355,60.8802757263184,60.8564949035645,60.860107421875,60.9066886901855,60.9146461486816,60.9495620727539,61.01416015625,61.0495147705078,61.0596694946289,61.0404319763184,61.0109367370605,60.9224586486816,60.8828582763672,60.8467750549316,60.8142547607422,60.7795944213867,60.7427215576172,60.6897964477539,60.5674324035645,60.4874534606934,60.4402351379395,60.3885231018066,60.332275390625,60.2859420776367,60.2203636169434,60.1985816955566,60.1454582214355,60.1221199035645,60.0758781433105,60.0297317504883,60.017822265625,60.0151863098145,60.0262222290039,59.9842796325684,59.9708518981934,59.9713897705078,59.9448738098145,59.9180183410645,59.8707542419434,59.8218307495117,59.7223129272461,59.6750946044922,59.6227035522461,59.5650901794434,59.4884262084961,59.3925323486328,59.3417472839355,59.3377952575684,59.3030776977539,59.2771987915039,59.1800308227539,59.1524429321289,59.0868682861328,59.0682144165039,59.0491714477539,59,58.920654296875,58.8692359924316,58.8294906616211,58.8208045959473,58.8313484191895,58.9396018981934,58.9557151794434,58.945556640625,58.9287605285645,58.9053688049316,58.8811569213867,58.8561515808105,58.8163566589355,58.7894058227539,58.7605972290039,58.7451667785645,58.6969718933105,58.6893081665039,58.7282676696777,58.8507347106934,58.8839340209961,58.8966331481934,58.8982429504395,58.888916015625,58.904541015625,58.8928680419922,58.8659172058105,58.823486328125,58.7827110290527,58.7435073852539,58.5954055786133,58.556640625,58.5281715393066,58.4845657348633,58.3992156982422,58.2269058227539,58.1632347106934,58.0763206481934,58.0368690490723,57.9998550415039,57.9687995910645,57.9260292053223,57.9024925231934,57.9758262634277,58.0116729736328,58.0517578125,58.0907249450684,58.1776885986328,58.4025917053223,58.4733390808105,58.4853057861328,58.4612274169922,58.3293914794922,58.2957534790039,58.2672386169434,58.1389656066895,58.1520538330078,58.2726058959961,58.3102531433105,58.3654747009277,58.4045906066895,58.3854484558105,58.2437973022461,58.1402359008789,58.1070289611816,58.0087432861328,57.9911117553711,58.0761222839355,58.1403312683105,58.1861343383789,58.2134780883789,58.3000030517578,58.370361328125,58.4329109191895,58.4627914428711,58.4910163879395,58.5489273071289,58.6366195678711,58.6973152160645,58.7309112548828,58.7911605834961,58.7945289611816,58.7722702026367,58.7271003723145,58.6590309143066,58.6056175231934,58.5667953491211,58.4308128356934,58.4312019348145,58.5350570678711,58.5719718933105,58.6109390258789,58.6498527526855,58.7347640991211,58.7878875732422,58.8206520080566,58.8392105102539,58.8466300964355,58.8605003356934,58.8956069946289,58.9146499633789,58.97705078125,58.9804649353027,58.9706039428711,59.0025901794434,59.0237770080566,59.0320358276367,59.0118637084961,59.0362281799316,59.0384254455566,59.0602073669434,59.0913047790527,59.1107444763184,59.1277313232422,59.1527862548828,59.2133293151855,59.2293930053711,59.2296867370605,59.213134765625,59.2449722290039,59.3197288513184,59.3503875732422,59.3149871826172,59.3302268981934,59.44775390625,59.4703140258789,59.4788093566895,59.4641609191895,59.3780250549316,59.3878936767578,59.4114723205566,59.4622535705566,59.4954643249512,59.5110359191895,59.5393524169922,59.6167945861816,59.6486854553223,59.6902313232422,59.7765121459961,59.7952117919922,59.8090858459473,59.8180656433105,59.8095207214355,59.7527847290039,59.7707061767578,59.8016128540039,59.8301277160645,59.8666496276855,59.9080047607422,59.9934120178223,60.0622024536133,60.2520027160645,60.2865219116211,60.3083038330078,60.3310546875,60.3361358642578,60.2682647705078,60.2281227111816,60.1713829040527,60.0945320129395,60.0371589660645,60.0121116638184,59.9975547790527,60.0434074401855,60.0647926330566,60.0640602111816,59.9729461669922,59.8465347290039,59.7412109375,59.7936019897461,59.8225593566895,59.7537117004395,59.6976089477539,59.6449203491211,59.5744132995605,59.5125999450684,59.4477996826172,59.4090805053711,59.3801765441895,59.3492698669434,59.3186492919922,59.2771492004395,59.2773399353027,59.3414535522461,59.3328628540039,59.1943893432617,59.1151847839355,59.0789031982422,59.0634727478027,59.0655784606934,59.0538101196289,59.0273895263672,59.0270004272461,59.0470199584961,59.0796394348145,59.0815925598145,59.0683097839355,59.0571746826172,59.034423828125,59.0264663696289,59.0031776428223,58.9279747009277,58.9110374450684,58.8673858642578,58.8577613830566,58.8781700134277,58.8554191589355,58.7650413513184,58.6724586486816,58.5457534790039,58.51953125,58.4525413513184,58.3988265991211,58.3299331665039,58.3306617736816,58.4412078857422,58.4669456481934,58.46044921875,58.4417495727539,58.4108428955078,58.414794921875,58.4793968200684,58.4921913146973,58.4963912963867,58.4740219116211,58.3191871643066,58.2003898620605,58.1271018981934,58.0841827392578,58.0146980285645,58.0021514892578,58.0933151245117,58.1292457580566,58.1581039428711,58.154052734375,57.9722671508789,57.9546394348145,57.9641151428223,57.9117660522461,57.8613243103027,57.8250503540039,57.8033180236816,57.7694320678711,57.6685562133789,57.6119155883789,57.5619125366211,57.5365715026855,57.5287590026855,57.4845237731934,57.4779815673828,57.4892120361328,57.4619636535645,57.4481964111328,57.440673828125,57.45458984375,57.4528312683105,57.4207992553711,57.3813018798828,57.3704147338867,57.3478546142578,57.2743682861328,57.2479476928711,57.2281265258789,57.1975593566895,57.1862297058105,57.1961936950684,57.1831512451172,57.0105934143066,56.9215812683105,56.8529777526855,56.7758331298828,56.6808090209961,56.6545867919922,56.6990737915039,56.7669944763184,56.7876968383789,56.8328132629395,56.836181640625,56.8184585571289,56.8017082214355,56.7300262451172,56.6668472290039,56.5908203125,56.5842781066895,56.5705108642578,56.5260238647461,56.5107421875,56.5054206848145,56.453857421875,56.423583984375,56.3970718383789,56.3903312683105,56.3606452941895,56.3275871276855,56.2887191772461,56.23095703125,56.2078132629395,56.2218284606934,56.2160148620605,56.0762214660645,56.0471687316895,56.0223617553711,55.9957046508789,55.9736785888672,55.9553718566895,55.9302749633789,55.8885726928711,55.8663558959961,55.8623542785645,55.9142112731934,55.9578628540039,55.9414558410645,55.886962890625,55.8250007629395,55.8148460388184,55.7270011901855,55.8051300048828,55.78857421875,55.78466796875,55.7090835571289,55.6495590209961,55.6123542785645,55.5569839477539,55.4809112548828,55.4443855285645,55.3663101196289,55.2427711486816,55.199951171875,55.1290054321289,55.0602035522461,55.0674819946289,55.1939468383789,55.2364273071289,55.2594223022461,55.2948722839355,55.3095703125,55.2691383361816,55.1963348388672,55.17333984375,55.1973648071289,55.1759262084961,55.1301765441895,54.9425773620605,54.8672370910645,54.81396484375,54.8870124816895,55.0555191040039,55.0807113647461,55.15283203125,55.199951171875,55.1832542419434,55.1494636535645,55.0550765991211,54.9522476196289,54.83837890625,54.7831077575684,54.7741203308105,54.8126945495605,54.8659210205078,54.8822250366211,54.875732421875,54.7731437683105,54.7186546325684,54.6737327575684,54.6503410339355,54.6402854919434,54.5908699035645,54.5704116821289,54.5174789428711,54.4404296875,54.3865737915039,54.3840827941895,54.3504371643066,54.3199729919434,54.2864723205566,54.2533187866211,54.2281265258789,54.1029777526855,54.049560546875,54.0394058227539,54.0444831848145,54.0331077575684,54.01025390625,53.9762687683105,53.963623046875,53.9291000366211,53.8341827392578,53.8312530517578,53.84228515625,53.8344230651855,53.8077621459961,53.7615737915039,53.7333526611328,53.7010231018066,53.6342277526855,53.610107421875,53.5961418151855,53.6100578308105,53.6533203125,53.6074714660645,53.5299835205078,53.4869613647461,53.4498062133789,53.3914566040039,53.360107421875,53.3435554504395,53.3170890808105,53.2890129089355,53.2774429321289,53.26611328125,53.3065414428711,53.392822265625,53.4800758361816,53.5045394897461,53.5368156433105,53.5747566223145,53.6437492370605,53.7439460754395,53.8752937316895,53.9778823852539,54.0518035888672,54.0895004272461,54.0911636352539,54.1035652160645,54.1267585754395,54.1312980651855,54.1144523620605,54.1544952392578,54.1719245910645,54.20166015625,54.2281761169434,54.1911125183105,54.1627426147461,53.9243621826172,53.8477058410645,53.7918434143066,53.7568855285645,53.7154808044434,53.6331024169922,53.6114234924316,53.5999031066895,53.583251953125,53.560546875,53.4690895080566,53.5285186767578,53.672607421875,53.7394523620605,53.7576675415039,53.7664566040039,53.7650413513184,53.7183113098145,53.6244659423828,53.60009765625,53.5875968933105,53.4711418151855,53.3908195495605,53.3438987731934,53.312255859375,53.2858390808105,53.2457504272461,53.2119598388672,53.1346664428711,53.0004425048828,52.87841796875,52.8233871459961,52.7356910705566,52.6771469116211,52.6431655883789,52.6233406066895,52.5747566223145,52.5737800598145,52.5445327758789,52.5359878540039,52.5374031066895,52.5076141357422,52.4745597839355,52.4282722473145,52.3915023803711,52.3642578125,52.3695793151855,52.3944854736328,52.3704109191895,52.3104019165039,52.2799301147461,52.2416000366211,52.1901397705078,52.1377906799316,51.9292945861816,51.7970695495605,51.6810035705566,51.4576644897461,51.44677734375,51.4425277709961,51.478271484375,51.4690933227539,51.4259300231934,51.3995132446289,51.3220710754395,51.3109855651855,51.2952156066895,51.305908203125,51.2950706481934,51.2571258544922,51.237060546875,51.1716766357422,50.8791007995605,50.7798805236816,50.6754417419434,50.4920883178711,50.4182586669922,50.31640625,50.2545890808105,50.2388648986816,50.2211418151855,50.2498054504395,50.2054214477539,50.1915054321289,50.2019500732422,50.1040496826172,50.1969718933105,50.2328605651855,50.2389183044434,50.2772941589355,50.2845230102539,50.3016586303711,50.3014640808105,50.2913551330566,50.2937965393066,50.2425765991211,50.2582015991211,50.3046379089355,50.3143577575684,50.303955078125,50.2694358825684,50.3089370727539,50.2754898071289,50.2978973388672,50.3200187683105,50.25927734375,50.2941436767578,50.2010269165039,50.2203636169434,50.2066421508789,50.2242660522461,50.2118644714355,50.1611824035645,50.1554222106934,50.0655250549316,49.9937019348145,49.6017608642578,49.451171875,49.3484382629395,49.3346214294434,49.332275390625,49.2567863464355,49.1971664428711,49.149658203125,49.1143569946289,49.0995101928711,49.05615234375,49.0071792602539,48.9394989013672,48.8289527893066,48.5736312866211,48.3864250183105,48.2507820129395,48.1991691589355,48.191162109375,48.2073745727539,48.2709465026855,48.2779808044434,48.3665008544922,48.4556159973145,48.4223136901855,48.3673820495605,48.3532218933105,48.3543434143066,48.2435569763184,48.2057609558105,48.1722640991211,48.0980949401855,47.9525871276855,47.8322296142578,47.7398948669434,47.5030250549316,47.4234390258789,47.1397933959961,47.0066909790039,46.9249496459961,46.7959480285645,46.6983909606934,46.673583984375,46.6868171691895,46.607421875,46.5588874816895,46.4850578308105,46.2873039245605,46.2624015808105,46.209716796875,46.1202659606934,46.0663108825684,46.0250015258789,45.8998565673828,45.7382316589355,45.711181640625,45.6549301147461,45.5641593933105,45.5018577575684,45.5310554504395,45.4928703308105,45.433349609375,45.3451156616211,45.3240203857422,45.2063941955566,45.0038566589355],[72.9366226196289,72.9267120361328,72.9969253540039,73.0624008178711,73.0984878540039,73.1301727294922,73.1572723388672,73.1888198852539,73.1654815673828,73.1373062133789,73.1262664794922,73.1019058227539,73.0738296508789,73.0415496826172,72.992431640625,72.9609375,72.9469757080078,72.9366226196289],[72.5929183959961,72.5803680419922,72.5907745361328,72.6240692138672,72.6514663696289,72.6728057861328,72.6468276977539,72.652099609375,72.6944351196289,72.7222671508789,72.753662109375,72.8220672607422,72.8632354736328,72.9497985839844,72.9810104370117,72.9952621459961,72.9860382080078,72.9366226196289,72.8880386352539,72.7659225463867,72.7137222290039,72.6685562133789,72.5721206665039,72.4356002807617,72.3863296508789,72.3561019897461,72.3352584838867,72.3001480102539,72.3509292602539,72.40771484375,72.454833984375,72.4590301513672,72.4490737915039,72.4248504638672,72.401123046875,72.3779754638672,72.3638687133789,72.3566436767578,72.3653335571289,72.4319839477539,72.5338821411133,72.5997085571289,72.6333541870117,72.6612777709961,72.6810531616211,72.7588806152344,72.7920379638672,72.8614273071289,72.9445266723633,73.0082092285156,72.9989242553711,72.9921417236328,72.983642578125,72.8756790161133,72.8084487915039,72.7750473022461,72.7264099121094,72.6868667602539,72.6707992553711,72.6540985107422,72.6368179321289,72.5960006713867,72.5828628540039,72.5675277709961,72.5510787963867,72.499267578125,72.4120101928711,72.3173294067383,72.1387710571289,71.900390625,71.7511749267578,71.7237854003906,71.7049331665039,71.677490234375,71.638671875,71.6261749267578,71.6296920776367,71.6630325317383,71.7161102294922,71.8192443847656,71.8352508544922,71.8589859008789,71.8751449584961,71.8860778808594,71.8946838378906,72.0253448486328,72.1493148803711,72.3026351928711,72.3474578857422,72.4427795410156,72.4678192138672,72.4907684326172,72.5204086303711,72.556640625,72.5877456665039,72.6526870727539,73.1058120727539,73.14990234375,73.1830596923828,73.2020568847656,73.2067413330078,73.2171401977539,73.2331008911133,73.2813949584961,73.3037109375,73.3487319946289,73.3290557861328,73.2883834838867,73.192138671875,73.1974105834961,73.2453079223633,73.2760238647461,73.2659149169922,73.1961898803711,73.0719223022461,73.0106430053711,72.927490234375,72.8489761352539,72.788330078125,72.7646484375,72.7403793334961,72.708740234375,72.6697311401367,72.6348114013672,72.5604476928711,72.4635772705078,72.4150848388672,71.97998046875,71.9031753540039,71.8675765991211,71.69921875,71.5769500732422,71.4951171875,71.4716796875,71.4339828491211,71.382080078125,71.3375473022461,71.30029296875,71.2479019165039,71.1324234008789,71.1040496826172,71.0642623901367,70.9272003173828,70.7626419067383,70.7337875366211,70.630859375,70.5724563598633,70.5473175048828,70.51025390625,70.5088348388672,70.5158233642578,70.5259323120117,70.5409622192383,70.6190948486328,70.6497039794922,70.6550750732422,70.5218734741211,70.4688415527344,70.4132308959961,70.2850570678711,70.2745590209961,70.286376953125,70.2782669067383,70.2655792236328,70.2483367919922,70.1724166870117,70.135009765625,70.1509704589844,70.1476058959961,70.1356964111328,70.1107940673828,70.0294952392578,69.8691864013672,69.8117218017578,69.71533203125,69.6798858642578,69.668701171875,69.6796340942383,69.7102584838867,69.749267578125,69.8338928222656,69.8501968383789,69.8331527709961,69.7556610107422,69.7212905883789,69.6985321044922,69.7041473388672,69.7380828857422,69.8246078491211,69.845947265625,69.8422393798828,69.81298828125,69.7582015991211,69.7179183959961,69.6921920776367,69.6598129272461,69.6208038330078,69.5920867919922,69.5735855102539,69.551513671875,69.5477523803711,69.5646514892578,69.5946731567383,69.67431640625,69.6853561401367,69.6676788330078,69.6444778442383,69.6114196777344,69.5684509277344,69.4972610473633,69.4833526611328,69.4849166870117,69.4717788696289,69.4334945678711,69.3675765991211,69.2359848022461,69.20458984375,69.2120132446289,69.34130859375,69.3775863647461,69.4344711303711,69.476318359375,69.4876937866211,69.46484375,69.4250946044922,69.4070281982422,69.3604507446289,69.3371124267578,69.3076171875,69.2811508178711,69.2576675415039,69.2360382080078,69.2455062866211,69.2399444580078,69.2170867919922,69.181640625,69.1322708129883,69.0239715576172,68.988525390625,68.9228515625,68.888916015625,68.864990234375,68.833251953125,68.8236389160156,68.8287124633789,68.8085403442383,68.8479995727539,68.8655700683594,68.9281692504883,68.9123992919922,68.8721160888672,68.92041015625,68.9553680419922,68.9781188964844,69.052490234375,69.0680694580078,69.0812530517578,69.093994140625,69.133544921875,69.1531753540039,69.1476058959961,69.1620101928711,69.194580078125,69.224365234375,69.251220703125,69.2806167602539,69.3396530151367,69.3810577392578,69.4137725830078,69.4430694580078,69.4395980834961,69.4071273803711,69.3473129272461,69.1807632446289,69.1522979736328,69.0316925048828,69.0021514892578,68.954345703125,68.9347610473633,68.8974075317383,68.8274383544922,68.7598114013672,68.6767120361328,68.6302719116211,68.6100082397461,68.5784225463867,68.5941925048828,68.5883331298828,68.5420379638672,68.5330581665039,68.5162124633789,68.4852523803711,68.4770965576172,68.4813461303711,68.494140625,68.535400390625,68.5987777709961,68.767578125,68.8384780883789,68.9598617553711,69.0301742553711,69.1819839477539,69.1950225830078,69.2513122558594,69.2691421508789,69.2727584838867,69.2647399902344,69.282958984375,69.3035659790039,69.337158203125,69.4246139526367,69.4335479736328,69.4626922607422,69.5120086669922,69.5762176513672,69.7193908691406,69.8042449951172,69.8258743286133,69.8881378173828,69.9910659790039,70.0540542602539,70.0924835205078,70.1001434326172,70.175048828125,70.2571258544922,70.3124542236328,70.3074645996094,70.2815399169922,70.2696762084961,70.2638168334961,70.2252426147461,70.2285614013672,70.2546920776367,70.2755813598633,70.2729034423828,70.2857437133789,70.308837890625,70.35205078125,70.4468765258789,70.6163558959961,70.6523895263672,70.6907196044922,70.6961975097656,70.6742706298828,70.6752471923828,70.6422348022461,70.6213912963867,70.6011734008789,70.5862808227539,70.5906753540039,70.6164016723633,70.6237335205078,70.6036682128906,70.6295394897461,70.8883285522461,70.9677200317383,71.0000457763672,71.0347213745117,71.1346664428711,71.158447265625,71.2121124267578,71.2769546508789,71.3379821777344,71.3591766357422,71.3616714477539,71.3817901611328,71.4010772705078,71.423095703125,71.4542999267578,71.4692916870117,71.485107421875,71.4658203125,71.472412109375,71.4936981201172,71.5108947753906,71.54638671875,71.4441909790039,71.4346160888672,71.3906707763672,71.39208984375,71.4359359741211,71.4490737915039,71.4670867919922,71.5257339477539,71.5608444213867,71.6593246459961,71.6610336303711,71.6399383544922,71.6490249633789,71.686767578125,71.7317428588867,71.7642593383789,71.8030242919922,71.9130859375,71.9722137451172,71.9855499267578,72.16748046875,72.2054672241211,72.2628860473633,72.2869110107422,72.31103515625,72.3695297241211,72.3992233276367,72.4276885986328,72.4725112915039,72.5338821411133,72.6328125,72.8311080932617,72.9144058227539,72.9593276977539,73.054931640625,73.2134704589844,73.3726577758789,73.3307113647461,73.2977981567383,73.2698211669922,73.230712890625,73.1802749633789,73.0709991455078,73.0146026611328,72.9580612182617,72.9068298339844,72.8609848022461,72.8050765991211,72.7390594482422,72.6258773803711,72.5929183959961],[72.919921875,72.9148941040039,72.93798828125,72.9893035888672,73.0933074951172,73.1963882446289,73.2581024169922,73.3041000366211,73.4133834838867,73.4577178955078,73.4798278808594,73.5103530883789,73.5990753173828,73.669921875,73.6956024169922,73.7185974121094,73.7657699584961,73.7671356201172,73.7444305419922,73.6472625732422,73.6362762451172,73.6144027709961,73.5780792236328,73.5411148071289,73.465576171875,73.3539123535156,73.3111343383789,73.1676330566406,73.0886764526367,73.0375518798828,72.9970169067383,72.9473114013672,72.919921875],[73.6544952392578,73.641357421875,73.6658248901367,73.6476516723633,73.5366744995117,73.4995651245117,73.4504852294922,73.373046875,73.3560562133789,73.3100128173828,73.2542037963867,73.225341796875,73.1592788696289,73.121826171875,73.1004867553711,73.0810012817383,72.9979019165039,72.959228515625,72.9123992919922,72.8811950683594,72.8430633544922,72.8206558227539,72.843994140625,72.8968276977539,72.8818359375,72.8577194213867,72.7716293334961,72.7577133178711,72.7559509277344,72.8263168334961,72.8636245727539,72.8924789428711,72.9769973754883,73.0782241821289,73.1613235473633,73.2246551513672,73.24560546875,73.2707977294922,73.3054733276367,73.3341751098633,73.3806610107422,73.428955078125,73.4715347290039,73.4819793701172,73.4831008911133,73.53466796875,73.5914077758789,73.6705551147461,73.7212371826172,73.7434539794922,73.7577667236328,73.767333984375,73.7654266357422,73.7070770263672,73.7015151977539,73.6544952392578],[71.0107955932617,70.98876953125,71.0167999267578,71.0489730834961,71.125732421875,71.1524429321289,71.1519546508789,71.137451171875,71.0915985107422,71.07861328125,71.0586929321289,71.03173828125,71.0018005371094,71.0286102294922,71.09326171875,71.3586883544922,71.4387741088867,71.5146026611328,71.5861358642578,71.6314468383789,71.6586456298828,71.654052734375,71.6612319946289,71.67529296875,71.697265625,71.72705078125,71.8165512084961,71.8664016723633,71.9564437866211,71.9865264892578,72.02197265625,72.0627899169922,72.1015625,72.21484375,72.233154296875,72.2181625366211,72.1294937133789,72.0526351928711,72.0444793701172,72.0584487915039,72.0944366455078,72.1895523071289,72.2587890625,72.3081512451172,72.3295364379883,72.3514175415039,72.3765640258789,72.40625,72.4056091308594,72.3843765258789,72.3829116821289,72.4215316772461,72.4439926147461,72.5105895996094,72.568603515625,72.6046371459961,72.6332015991211,72.72216796875,72.7744598388672,72.8191909790039,72.8563995361328,72.9251480102539,72.9450225830078,72.9540023803711,72.9198684692383,72.7967300415039,72.83642578125,73.0026397705078,73.04541015625,73.10546875,73.3354949951172,73.3895492553711,73.4564895629883,73.4593734741211,73.4169006347656,73.428466796875,73.5083999633789,73.5283203125,73.5758743286133,73.6316833496094,73.676025390625,73.6991195678711,73.7154312133789,73.7295913696289,73.7364730834961,73.7298355102539,73.6959991455078,73.634521484375,73.5977554321289,73.4795455932617,73.3140106201172,73.2452163696289,73.2071762084961,73.1658172607422,73.1211929321289,73.06494140625,72.9971694946289,72.9276885986328,72.8565902709961,72.8162536621094,72.7701644897461,72.7455596923828,72.7174758911133,72.6789093017578,72.5586395263672,72.4262161254883,72.3117218017578,72.2779312133789,72.4571838378906,72.4508285522461,72.42578125,72.3382797241211,72.26025390625,72.2101516723633,72.1912612915039,72.1034622192383,72.0890121459961,72.09619140625,72.0882797241211,72.0723114013672,72.0482406616211,72.0143508911133,71.970703125,71.9380874633789,71.9165496826172,71.9076614379883,71.9114303588867,71.92919921875,71.9881286621094,72.1487808227539,72.2087860107422,72.2140579223633,72.1749496459961,72.1772003173828,72.3008804321289,72.3321762084961,72.3783187866211,72.3942413330078,72.4281692504883,72.4462890625,72.438720703125,72.3759307861328,72.3321762084961,72.3146514892578,72.3373031616211,72.3908157348633,72.355712890625,72.2720184326172,72.188232421875,72.1043472290039,72.0429229736328,71.9303665161133,71.8810043334961,71.880615234375,71.8975067138672,71.9349594116211,71.972412109375,72.0302734375,72.100830078125,72.2654266357422,72.3514175415039,72.329345703125,72.27978515625,72.275146484375,72.2803192138672,72.1799774169922,72.1777877807617,72.21875,72.2384338378906,72.2938537597656,72.3598175048828,72.4352111816406,72.4706115722656,72.5087432861328,72.5423355102539,72.571533203125,72.6001968383789,72.6875991821289,72.7247543334961,72.7368621826172,72.7358856201172,72.7206573486328,72.695068359375,72.6333541870117,72.5722122192383,72.5613250732422,72.562744140625,72.5765075683594,72.571533203125,72.4808578491211,72.4342346191406,72.3777313232422,72.3228530883789,72.2695846557617,72.2263717651367,72.039794921875,72.0079574584961,71.9371566772461,71.8030776977539,71.7312927246094,71.7172393798828,71.7137222290039,71.7458953857422,71.8385772705078,71.91845703125,71.984375,72.06298828125,72.1004867553711,72.096923828125,72.0856475830078,72.0665588378906,72.0505828857422,72.0376968383789,71.9786682128906,71.9386749267578,71.8936462402344,71.8426742553711,71.7862777709961,71.7419891357422,71.7255401611328,71.7091293334961,71.6915969848633,71.66748046875,71.6755905151367,71.6469192504883,71.5708923339844,71.5047378540039,71.4505920410156,71.4065856933594,71.3140563964844,71.2305221557617,71.2181167602539,71.338134765625,71.4694366455078,71.5848617553711,71.6483917236328,71.6725082397461,71.6822280883789,71.7496109008789,71.7710494995117,71.7714385986328,71.746337890625,71.7198791503906,71.599365234375,71.4041519165039,71.4264678955078,71.4728546142578,71.5191421508789,71.5875930786133,71.5255355834961,71.479248046875,71.3734359741211,71.3224639892578,71.2828598022461,71.349853515625,71.4842758178711,71.5379867553711,71.5648880004883,71.6777801513672,71.64013671875,71.6067886352539,71.6156234741211,71.6109848022461,71.5927734375,71.5612335205078,71.5162582397461,71.4637222290039,71.4034652709961,71.36181640625,71.3387680053711,71.2787170410156,71.2388153076172,71.1468734741211,71.1051254272461,71.0863723754883,71.1047821044922,71.0942840576172,71.0653305053711,70.9388122558594,70.8840789794922,70.8492202758789,70.8307647705078,70.83251953125,70.8701629638672,70.940673828125,71.013427734375,71.046875,70.9751434326172,70.97802734375,71.0499954223633,71.0990219116211,71.1087341308594,71.1033248901367,71.0522003173828,71.006591796875,70.9024353027344,70.8708953857422,70.792236328125,70.6741180419922,70.6297836303711,70.60595703125,70.5658645629883,70.5335464477539,70.4876937866211,70.45703125,70.4315414428711,70.3941879272461,70.3952178955078,70.4175796508789,70.5056686401367,70.5440444946289,70.5521011352539,70.5484390258789,70.5311508178711,70.5002899169922,70.4251937866211,70.1277847290039,70.128662109375,70.2093276977539,70.51904296875,70.5810546875,70.6436004638672,70.73828125,70.787841796875,70.8295364379883,70.8450164794922,70.8345718383789,70.785888671875,70.7771453857422,70.7892532348633,70.783447265625,70.76416015625,70.7280807495117,70.6871109008789,70.6102600097656,70.5940933227539,70.5649948120117,70.5228958129883,70.4812469482422,70.4399871826172,70.4148483276367,70.4056625366211,70.3832015991211,70.3244094848633,70.314453125,70.2891616821289,70.2768096923828,70.2531280517578,70.1893081665039,70.0708465576172,70.0426254272461,70.029052734375,70.0469207763672,70.1287612915039,70.1600570678711,70.1851043701172,70.2069854736328,70.2035675048828,70.1991653442383,70.1798324584961,70.1456527709961,70.1010284423828,70.0799331665039,69.9789581298828,69.9527359008789,69.94140625,69.9684600830078,70.0304718017578,70.0648422241211,70.0716247558594,70.08740234375,70.1122055053711,70.12841796875,70.1605987548828,70.18017578125,70.228271484375,70.281494140625,70.3145980834961,70.3172378540039,70.2817840576172,70.2198257446289,70.034423828125,69.9984359741211,69.8606948852539,69.7994689941406,69.7568359375,69.730712890625,69.7212905883789,69.7209930419922,69.7773895263672,69.7700653076172,69.7542953491211,69.7306137084961,69.7007827758789,69.6646957397461,69.6443862915039,69.6436538696289,69.62353515625,69.5745162963867,69.5474090576172,69.5119171142578,69.5181198120117,69.564208984375,69.5772933959961,69.4758758544922,69.4601058959961,69.4747009277344,69.479248046875,69.4725112915039,69.4601058959961,69.4211883544922,69.336669921875,69.3118667602539,69.2857360839844,69.2584457397461,69.1910629272461,69.168212890625,69.1527328491211,69.170654296875,69.1847152709961,69.1669921875,69.1731948852539,69.2002487182617,69.2481460571289,69.2026824951172,69.2322235107422,69.2060012817383,69.0979995727539,69.079345703125,69.1720733642578,69.1664123535156,69.1326217651367,69.0659637451172,69.0386734008789,68.933837890625,68.8633346557617,68.7839889526367,68.794677734375,68.8440475463867,68.850830078125,68.8427734375,68.8113250732422,68.8102111816406,68.872802734375,68.8757858276367,68.869384765625,68.8569793701172,68.7599563598633,68.7493667602539,68.7325668334961,68.7029800415039,68.6810531616211,68.6161117553711,68.5507354736328,68.524169921875,68.52294921875,68.5470199584961,68.5507354736328,68.5339813232422,68.4978942871094,68.48779296875,68.4658737182617,68.4614715576172,68.4716262817383,68.457763671875,68.4457015991211,68.4246597290039,68.3741683959961,68.3260803222656,68.2635269165039,68.2156295776367,68.1798858642578,68.0984878540039,68.0657272338867,68.0630874633789,68.1290054321289,68.1205596923828,68.0292510986328,68.034423828125,68.1100082397461,68.2106475830078,68.2503433227539,68.2804183959961,68.1227111816406,68.040771484375,67.904296875,67.8602981567383,67.8511734008789,67.8338394165039,67.8319320678711,67.853271484375,67.958740234375,68.0685501098633,68.0709457397461,68.0311965942383,67.9574279785156,67.9228515625,67.9570846557617,67.9866714477539,67.9823226928711,67.96826171875,67.9356918334961,67.7993698120117,67.765625,67.6748504638672,67.6802749633789,67.7240676879883,67.8323211669922,67.8792495727539,67.9394989013672,68.0262222290039,68.0434112548828,68.0316390991211,67.989990234375,67.9651336669922,67.9391098022461,67.8920364379883,67.8238296508789,67.7875518798828,67.7842788696289,67.8402328491211,67.8126907348633,67.7399368286133,67.6230010986328,67.6548843383789,67.5660629272461,67.49560546875,67.3473129272461,67.3534622192383,67.3418502807617,67.3551712036133,67.3505325317383,67.3155288696289,67.3010787963867,67.2561569213867,67.2572784423828,67.2659149169922,67.2641143798828,67.315673828125,67.3451156616211,67.3775405883789,67.3583526611328,67.3363265991211,67.2350082397461,67.1743621826172,67.1170349121094,67.0685043334961,67.024658203125,66.9944839477539,66.8223648071289,66.82080078125,66.8624038696289,66.92431640625,66.9492721557617,66.9327087402344,66.9319839477539,66.9541015625,66.9286117553711,66.9053726196289,67.0467300415039,67.01904296875,66.9317321777344,66.7784652709961,66.689208984375,66.6487350463867,66.6088409423828,66.5665969848633,66.55810546875,66.6377944946289,66.6781234741211,66.6737823486328,66.64306640625,66.6259307861328,66.5031280517578,66.4124984741211,66.3729019165039,66.3128433227539,66.309326171875,66.3379898071289,66.3915023803711,66.4108352661133,66.4171905517578,66.4068374633789,66.31591796875,66.28857421875,66.2709045410156,66.2270050048828,66.14794921875,66.0675354003906,66.0353012084961,66.0113754272461,66.005859375,66.0174789428711,66.034423828125,66.0162582397461,65.9740295410156,65.9455108642578,65.90576171875,65.8683090209961,65.8333053588867,65.8044967651367,65.7236328125,65.6399383544922,65.6319808959961,65.647705078125,65.6223602294922,65.6573257446289,65.695556640625,65.85302734375,65.8353500366211,65.7558059692383,65.7085952758789,65.67431640625,65.6609878540039,65.6360321044922,65.6167449951172,65.543212890625,65.229736328125,65.1184539794922,65.021240234375,64.9280776977539,64.9891128540039,65.0513687133789,65.0833053588867,65.1092758178711,65.1214828491211,65.1219253540039,65.066162109375,65.1143035888672,65.1724166870117,65.3245544433594,65.4007797241211,65.4001922607422,65.3641586303711,65.2527313232422,65.1166000366211,65.1689453125,65.2340850830078,65.299560546875,65.3750915527344,65.4637680053711,65.5681610107422,65.5896530151367,65.6766586303711,65.7015151977539,65.7097702026367,65.7640151977539,65.8220672607422,65.8906707763672,65.93994140625,65.988525390625,66.0159149169922,66.0785598754883,66.1927261352539,66.2721633911133,66.317138671875,66.3254852294922,66.2870101928711,66.2309112548828,66.1374053955078,66.0777359008789,66.0084457397461,65.9945755004883,65.9872055053711,65.9969177246094,66.0202102661133,66.08642578125,66.2047348022461,66.2130813598633,66.1711959838867,66.1422348022461,66.12744140625,66.1327133178711,66.2063980102539,66.2291030883789,66.2544937133789,66.2797393798828,66.46044921875,66.5084991455078,66.5556640625,66.5953140258789,66.6274948120117,66.6222152709961,66.5818862915039,66.5471649169922,66.5254898071289,66.5330123901367,66.5697250366211,66.5203628540039,66.4327621459961,66.3217315673828,66.3102569580078,66.3037643432617,66.3174819946289,66.4004440307617,66.4582061767578,66.4901351928711,66.4674301147461,66.3673324584961,66.2686080932617,66.1872100830078,66.0902862548828,66.034423828125,65.95556640625,65.9297332763672,65.9217300415039,65.921630859375,65.9651870727539,66.0138168334961,66.1298370361328,66.249267578125,66.2486343383789,66.2000503540039,66.1922378540039,66.188720703125,66.1731948852539,66.078857421875,66.0389709472656,65.9945755004883,65.9386291503906,65.8709945678711,65.8277816772461,65.7972717285156,65.7932586669922,65.773681640625,65.6231002807617,65.5819320678711,65.5648880004883,65.5634765625,65.6253356933594,65.6404266357422,65.6435546875,65.626220703125,65.58837890625,65.5493698120117,65.5091781616211,65.48291015625,65.4403839111328,65.4205093383789,65.3566360473633,65.3465881347656,65.3419418334961,65.3038101196289,65.2440948486328,65.1729965209961,65.1380386352539,65.1048355102539,65.0849151611328,65.0813446044922,65.093994140625,65.0915985107422,65.0741729736328,65.0196762084961,64.8600540161133,64.815185546875,64.8136749267578,64.9738311767578,65.0003433227539,64.9719696044922,64.9096145629883,64.8541030883789,64.828125,64.7777328491211,64.7553253173828,64.722412109375,64.7349166870117,64.7912063598633,64.8465805053711,64.8857421875,64.8535614013672,64.770751953125,64.7423324584961,64.7064971923828,64.7264175415039,64.6315460205078,64.5885238647461,64.5259780883789,64.5047836303711,64.5096206665039,64.5097122192383,64.4845657348633,64.4366683959961,64.3032760620117,64.3150863647461,64.3183135986328,64.3111343383789,64.2938461303711,64.2323226928711,64.181640625,64.1298370361328,64.087158203125,64.0675277709961,64.0379867553711,64.0281753540039,64.0088348388672,64.0261764526367,64.0327606201172,64.0279769897461,64.0095672607422,63.9609375,63.9159660339355,63.9054222106934,63.9183578491211,63.8973617553711,63.7903327941895,63.7063484191895,63.6870613098145,63.6796836853027,63.5725593566895,63.4627914428711,63.3575668334961,63.2639656066895,63.2431678771973,63.2453575134277,63.2688484191895,63.5487289428711,63.5992660522461,63.6433601379395,63.7642555236816,63.7448234558105,63.6890602111816,63.6059608459473,63.4401359558105,63.3334007263184,63.2928237915039,63.2828636169434,63.2634811401367,63.2346229553223,63.1256332397461,63.0600090026855,62.9917945861816,62.94580078125,62.9219703674316,62.9023933410645,62.8871574401855,62.8798828125,62.8891563415527,62.9523429870605,62.9326171875,62.7014656066895,62.6461410522461,62.6264610290527,62.6494598388672,62.7150917053223,62.7881813049316,62.8689002990723,62.9319839477539,62.9302749633789,62.9115753173828,62.9085426330566,62.921142578125,62.968505859375,63.0006828308105,63.107177734375,63.1082496643066,63.0763206481934,63.0062484741211,62.9909706115723,62.9966773986816,63.0271949768066,63.0972671508789,63.2188987731934,63.2647438049316,63.2349090576172,63.1190414428711,63.0801315307617,63.0695304870605,63.0801773071289,63.111083984375,63.1622581481934,63.2011184692383,63.2276840209961,63.2555618286133,63.3051300048828,63.3165016174316,63.3050270080566,63.3407211303711,63.4814453125,63.6339340209961,63.7145538330078,63.7337379455566,63.6350059509277,63.4892578125,63.4227523803711,63.4197273254395,63.44921875,63.6370620727539,63.7254867553711,63.7411155700684,63.7518577575684,63.7032203674316,63.5951194763184,63.5136680603027,63.4589385986328,63.3521957397461,63.2146987915039,63.1723175048828,63.1136741638184,63.0980949401855,63.0935592651367,63.0726585388184,63.03369140625,63.0095672607422,62.9482383728027,62.9141616821289,62.8575668334961,62.843505859375,62.7008285522461,62.6780776977539,62.6317901611328,62.60205078125,62.5099601745605,62.4631309509277,62.3519020080566,62.3026847839355,62.2463912963867,62.2302780151367,62.2088851928711,62.1583023071289,62.13720703125,62.1023941040039,62.0539093017578,61.9674797058105,61.9138679504395,61.8930625915527,61.8682632446289,61.8702659606934,61.8907203674316,61.9255905151367,62.0125999450684,62.0728530883789,62.1050300598145,62.1340789794922,62.1512718200684,62.2351531982422,62.255615234375,62.2812995910645,62.3189926147461,62.4051780700684,62.4239730834961,62.5718765258789,62.7445831298828,62.7677230834961,62.7904739379883,62.7761726379395,62.7572250366211,62.7633819580078,62.7915077209473,62.8691864013672,62.9104957580566,62.978271484375,63.0022468566895,63.0196762084961,63.1207008361816,63.1192893981934,63.0425300598145,63.0661163330078,63.1264190673828,63.1872062683105,63.3552703857422,63.4161605834961,63.4298820495605,63.4354515075684,63.4302253723145,63.444091796875,63.5122566223145,63.5550270080566,63.5803260803223,63.5865745544434,63.5988273620605,63.6267585754395,63.6625938415527,63.7061500549316,63.7249488830566,63.7088890075684,63.7279815673828,63.7569808959961,63.8387184143066,63.8716812133789,63.8898887634277,63.8934059143066,63.891357421875,63.8181190490723,63.823486328125,63.9007835388184,63.9890632629395,64.02001953125,64.0304718017578,64.1171875,64.2818908691406,64.33349609375,64.3795852661133,64.3992614746094,64.4234848022461,64.5602493286133,64.58251953125,64.5741729736328,64.6025390625,64.5682678222656,64.5662078857422,64.5853500366211,64.578125,64.4658203125,64.437744140625,64.4226608276367,64.4246597290039,64.4437026977539,64.4699249267578,64.6078109741211,64.6281204223633,64.6334915161133,64.6446838378906,64.670166015625,64.7173309326172,64.7861785888672,64.8239288330078,64.8306655883789,64.8251495361328,64.8073196411133,64.7962417602539,64.7919921875,64.7655258178711,64.7168960571289,64.6473617553711,64.5570755004883,64.49658203125,64.4657211303711,64.4566879272461,64.4693832397461,64.4904251098633,64.540771484375,64.5243682861328,64.3919448852539,64.3846664428711,64.3880844116211,64.3763122558594,64.3031692504883,64.29296875,64.3016128540039,64.283935546875,64.2420425415039,64.2376480102539,64.2708511352539,64.2850570678711,64.2803726196289,64.2998962402344,64.34375,64.3634719848633,64.3601531982422,64.3670883178711,64.4610824584961,64.499267578125,64.6177215576172,64.6646423339844,64.7147445678711,64.7518081665039,64.8077163696289,64.9392623901367,64.98291015625,65.0226058959961,65.0729446411133,65.1615676879883,65.196533203125,65.2197723388672,65.3281707763672,65.3559112548828,65.3721160888672,65.4354553222656,65.453125,65.462890625,65.4308624267578,65.4180145263672,65.4138717651367,65.3697280883789,65.2854461669922,65.22705078125,65.1408157348633,65.0560073852539,65.0130844116211,64.947021484375,64.9276809692383,64.905029296875,64.8792037963867,64.8415985107422,64.8558578491211,64.9007797241211,64.9385223388672,64.9690475463867,65.0087432861328,65.0575637817383,65.0997085571289,65.1351547241211,65.2570343017578,65.2975082397461,65.3157196044922,65.2748031616211,65.283935546875,65.3314437866211,65.3639678955078,65.3814468383789,65.3890609741211,65.366943359375,65.3636703491211,65.3716812133789,65.3975601196289,65.48388671875,65.5034713745117,65.5169906616211,65.5188522338867,65.4843292236328,65.4852523803711,65.5429153442383,65.6532211303711,65.7666931152344,65.80517578125,65.8770523071289,66.0127487182617,66.0969696044922,66.1390151977539,66.1670913696289,66.2081527709961,66.3580551147461,66.5069351196289,66.5831527709961,66.6362838745117,66.6749572753906,66.6991653442383,66.7281723022461,66.765380859375,66.8285140991211,66.8832550048828,67.0306167602539,67.0704574584961,67.0980987548828,67.1333999633789,67.2542953491211,67.284423828125,67.3072738647461,67.3418960571289,67.6586456298828,67.8116226196289,67.9447784423828,68.1069869995117,68.2667541503906,68.3089828491211,68.3569869995117,68.3678207397461,68.2977523803711,68.2945251464844,68.3249969482422,68.3629379272461,68.4293975830078,68.4641571044922,68.4970703125,68.5780258178711,68.6192855834961,68.6586380004883,68.6859893798828,68.7013702392578,68.7109832763672,68.7149429321289,68.700927734375,68.5787582397461,68.5486297607422,68.5354537963867,68.5412063598633,68.5560531616211,68.5799331665039,68.70751953125,68.7555694580078,68.790283203125,68.808349609375,68.7967300415039,68.7958984375,68.8081512451172,68.8231430053711,68.89208984375,68.9133758544922,68.9361267089844,68.9407272338867,68.9610900878906,68.9828643798828,69.0206527709961,69.0455093383789,69.0642547607422,69.0658187866211,69.0246047973633,68.9405746459961,68.9093780517578,68.9482955932617,68.9612808227539,68.9527359008789,68.8877410888672,68.8401870727539,68.72802734375,68.6923294067383,68.69873046875,68.7213897705078,68.7598648071289,68.7915573120117,68.8163070678711,68.8466796875,68.9744567871094,69.0094757080078,69.0304183959961,69.0524444580078,69.0261688232422,69.0308074951172,69.060302734375,69.1029281616211,69.1588439941406,69.2125473022461,69.299560546875,69.3186569213867,69.3863754272461,69.4109878540039,69.421630859375,69.4411163330078,69.4694290161133,69.5166015625,69.5486831665039,69.5909194946289,69.6199645996094,69.6534729003906,69.662109375,69.6868133544922,69.6839294433594,69.65625,69.5912628173828,69.5728988647461,69.6111831665039,69.6168441772461,69.6351089477539,69.6527328491211,69.6707534790039,69.6890640258789,69.7451629638672,69.775390625,69.8248519897461,69.8361282348633,69.8545913696289,69.8362350463867,69.8456039428711,69.9004440307617,69.9657287597656,70.04150390625,70.1707992553711,70.238525390625,70.2470703125,70.2191390991211,70.2187957763672,70.2291488647461,70.3155746459961,70.34619140625,70.3534240722656,70.4453125,70.4631805419922,70.508544921875,70.5813522338867,70.6035614013672,70.5752487182617,70.5347137451172,70.4818801879883,70.437255859375,70.4007339477539,70.372314453125,70.3251953125,70.293701171875,70.2418975830078,70.1785583496094,70.1036148071289,70.0476531982422,70.0104522705078,69.9774856567383,69.9253387451172,69.8948211669922,69.8875961303711,69.8947296142578,69.9959945678711,69.9967727661133,70.0104522705078,70.0521011352539,70.056640625,70.0911636352539,70.1112289428711,70.0946350097656,70.0480499267578,70.0246124267578,70.0243606567383,69.9828186035156,69.9000015258789,69.8505859375,69.7916488647461,69.7713851928711,69.7309112548828,69.9427185058594,69.8687515258789,69.8412094116211,69.8369140625,69.865966796875,69.9681625366211,70.00390625,70.0090866088867,69.9647979736328,69.9627380371094,70.0052261352539,70.0336456298828,70.0633316040039,70.0782241821289,70.0782241821289,70.1113815307617,70.0366744995117,70.1051330566406,70.1454086303711,70.1730499267578,70.28857421875,70.3503875732422,70.40625,70.4445266723633,70.46533203125,70.4012680053711,70.3907241821289,70.3882827758789,70.4119644165039,70.3998489379883,70.37744140625,70.3448181152344,70.3250961303711,70.3097152709961,70.32568359375,70.3187484741211,70.309814453125,70.2582473754883,70.24658203125,70.2519073486328,70.3686065673828,70.4424819946289,70.4708557128906,70.4944839477539,70.5228958129883,70.7597122192383,70.8106918334961,70.9961395263672,71.0356979370117,71.0617218017578,71.0671920776367,71.0446319580078,71.0456085205078,71.0305633544922,70.9843292236328,70.9513168334961,70.9443817138672,70.9565963745117,70.987548828125,71.0116195678711,71.0528335571289,71.1075668334961,71.1785125732422,71.2085494995117,71.2272491455078,71.2402801513672,71.2879409790039,71.3521957397461,71.4234848022461,71.4623031616211,71.4922866821289,71.5857467651367,71.7427215576172,71.8480453491211,71.9018020629883,71.9487838745117,72.0490188598633,72.157958984375,72.175048828125,72.1801300048828,72.2078094482422,72.2483444213867,72.3126449584961,72.3671875,72.4119186401367,72.4677276611328,72.5680694580078,72.6898422241211,72.8041458129883,72.841552734375,72.9429702758789,73.0169448852539,73.0689926147461,73.1080551147461,73.1821746826172,73.2524871826172,73.3124008178711,73.3345718383789,73.3652801513672,73.38818359375,73.4032745361328,73.5953140258789,73.67333984375,73.722900390625,73.7594223022461,73.833984375,73.8547897338867,73.8501434326172,73.8081512451172,73.7786102294922,73.7637176513672,73.7216339111328,73.6947784423828,73.6035690307617,73.5276794433594,73.4614791870117,73.3125457763672,73.26025390625,72.9602508544922,72.9105453491211,72.8708038330078,72.7625503540039,72.7240219116211,72.6611328125,72.524658203125,72.4608383178711,72.4021530151367,72.2622528076172,72.1913070678711,72.1231918334961,72.0257797241211,71.8991165161133,71.7709426879883,71.641357421875,71.555419921875,71.4921340942383,71.3984909057617,71.3532257080078,71.3034210205078,71.2267532348633,71.1939468383789,71.1626510620117,71.0958938598633,71.0426254272461,71.0107955932617],[73.9458923339844,73.7315902709961,73.749267578125,73.7605438232422,73.7920837402344,73.8182144165039,73.8657684326172,73.8793411254883,73.8877410888672,73.8875427246094,73.8618621826172,73.8437957763672,73.8248519897461,73.7903289794922,73.7061462402344,73.6749038696289,73.66650390625,73.61474609375,73.5921859741211,73.5707550048828,73.564208984375,73.526611328125,73.53662109375,73.5022964477539,73.4813461303711,73.4711456298828,73.4735794067383,73.4882354736328,73.48095703125,73.4584426879883,73.4212875366211,73.3868179321289,73.3392105102539,73.2853012084961,73.1157760620117,73.044677734375,73.0225067138672,73.000244140625,72.9580612182617,72.9410171508789,72.9341354370117,72.9930648803711,73.0205078125,73.0355987548828,73.0366668701172,73.0276412963867,72.9922866821289,72.9378433227539,72.9180221557617,72.8981475830078,72.8781280517578,72.8649444580078,72.8374481201172,72.7628402709961,72.7175750732422,72.6727523803711,72.6427688598633,72.6275863647461,72.6368179321289,72.68701171875,72.7131805419922,72.7102508544922,72.69873046875,72.6298828125,72.5524444580078,72.4873046875,72.4343719482422,72.3931121826172,72.342041015625,72.3226089477539,72.3224105834961,72.3137664794922,72.271240234375,72.2372589111328,72.2044906616211,72.1728515625,72.1459045410156,72.0459518432617,72.0316848754883,72.025146484375,71.9675827026367,71.8338394165039,71.7918930053711,71.7607421875,71.7108383178711,71.6734848022461,71.6342239379883,71.6296920776367,71.6624526977539,71.6814956665039,71.7155227661133,71.7645034790039,71.8030776977539,71.8311080932617,71.8475570678711,71.8523406982422,71.8242645263672,71.773193359375,71.7164993286133,71.5589370727539,71.4912109375,71.4624557495117,71.4408721923828,71.3488235473633,71.317626953125,71.3021011352539,71.3136672973633,71.3523406982422,71.3694763183594,71.3671875,71.3871078491211,71.4242172241211,71.5571823120117,71.6515579223633,71.7572326660156,71.9115219116211,72.0038528442383,72.15234375,72.1859436035156,72.1994171142578,72.2100601196289,72.2285690307617,72.279052734375,72.3169937133789,72.32177734375,72.3128433227539,72.2778778076172,72.3148956298828,72.3409194946289,72.3850631713867,72.4092788696289,72.4310531616211,72.4861373901367,72.5947265625,72.7194290161133,72.7645034790039,72.7829132080078,72.8428192138672,72.9107971191406,72.978271484375,73.0059127807617,73.0641098022461,73.0772933959961,73.0699234008789,73.0569839477539,73.0180130004883,72.9730987548828,72.9428176879883,72.9097137451172,72.883056640625,72.821044921875,72.7462921142578,72.7216796875,72.7132797241211,72.7259292602539,72.77294921875,72.77880859375,72.8068389892578,72.9769973754883,72.9777374267578,72.89892578125,72.8902816772461,72.9066925048828,72.9449691772461,72.963134765625,73.0553207397461,73.0954055786133,73.1203079223633,73.1284637451172,73.1205596923828,73.1382827758789,73.1637191772461,73.1978530883789,73.2339401245117,73.2545928955078,73.2651824951172,73.2546844482422,73.2110824584961,73.201416015625,73.2138671875,73.239501953125,73.3402328491211,73.3590316772461,73.3158187866211,73.299560546875,73.2784729003906,73.2753448486328,73.4309539794922,73.4458541870117,73.4863739013672,73.5050277709961,73.5338363647461,73.5719680786133,73.5958480834961,73.5997543334961,73.5712890625,73.4942932128906,73.4493103027344,73.4654769897461,73.5097198486328,73.5753936767578,73.5933609008789,73.6129379272461,73.6580581665039,73.69140625,73.7272033691406,73.7653350830078,73.7914047241211,73.8053741455078,73.843505859375,73.80126953125,73.795166015625,73.8470230102539,73.8571319580078,73.8437957763672,73.8440933227539,73.8725128173828,73.8891067504883,73.9288558959961,73.9458923339844],[73.8684997558594,73.847412109375,73.8564987182617,73.828857421875,73.817138671875,73.8120574951172,73.8619613647461,73.8792953491211,73.8954086303711,73.910888671875,73.9257354736328,73.9482955932617,73.9649353027344,73.9881820678711,74.0068817138672,74.02099609375,74.0345230102539,74.1046905517578,74.1146469116211,74.1086959838867,74.0901336669922,74.0716323852539,74.0530242919922,74.0055160522461,73.9684524536133,73.8888168334961,73.8684997558594],[74.1609878540039,74.1137237548828,74.0827178955078,74.0620651245117,73.9923782348633,73.9723663330078,74.0127868652344,74.0277862548828,74.00927734375,73.9517059326172,73.9084014892578,73.86865234375,73.8247528076172,73.7538604736328,73.6864242553711,73.5806198120117,73.5276794433594,73.5022964477539,73.4670944213867,73.41552734375,73.3040008544922,73.284912109375,73.236083984375,73.2142105102539,73.1948776245117,73.1453628540039,73.02587890625,72.9153823852539,72.8493118286133,72.7538070678711,72.726806640625,72.7184524536133,72.8018493652344,72.800537109375,72.7569274902344,72.7356491088867,72.703369140625,72.668212890625,72.5586395263672,72.5313034057617,72.4994583129883,72.43701171875,72.421142578125,72.2526397705078,72.1299819946289,72.0287551879883,72.0008316040039,72.0436096191406,72.0423355102539,72.0127944946289,72.0274353027344,72.1800308227539,72.3447799682617,72.501953125,72.7815475463867,72.8311538696289,72.8844680786133,72.9415969848633,72.9990768432617,73.1152801513672,73.1741714477539,73.3277359008789,73.5574722290039,73.6385269165039,73.6707992553711,73.6954574584961,73.7281799316406,73.7516555786133,73.755126953125,73.6857376098633,73.6625518798828,73.66357421875,73.6714401245117,73.6860809326172,73.7160186767578,73.8050765991211,73.8812484741211,73.9063949584961,73.9327621459961,73.9603042602539,73.9850616455078,74.0238723754883,74.0414123535156,74.0859909057617,74.1131362915039,74.131591796875,74.1183547973633,74.1671371459961,74.1787567138672,74.1609878540039],[74.1126480102539,74.1084442138672,74.2012252807617,74.2062454223633,74.197998046875,74.1861801147461,74.1678695678711,74.1275863647461,74.0278778076172,74.0155258178711,74.0211944580078,74.0447311401367,74.1920928955078,74.2325210571289,74.24462890625,74.2667541503906,74.2660675048828,74.2523422241211,74.2317352294922,74.171142578125,74.1014175415039,74.0271453857422,73.7479476928711,73.66552734375,73.6187515258789,73.5846710205078,73.5418930053711,73.501953125,73.4647979736328,73.4388732910156,73.416748046875,73.3232421875,73.2945861816406,73.2532272338867,73.1072769165039,73.0377502441406,72.9021911621094,72.6841354370117,72.6403350830078,72.6088409423828,72.3603973388672,72.3026885986328,72.2438430786133,72.2291488647461,72.212646484375,72.1267547607422,71.9840774536133,71.8880310058594,71.6308135986328,71.6050796508789,71.5574188232422,71.5057601928711,71.4462356567383,71.4149856567383,71.3890151977539,71.4067840576172,71.4476089477539,71.451171875,71.4447784423828,71.2659225463867,71.1935577392578,71.128173828125,71.0974578857422,71.0879898071289,71.0938034057617,71.1234359741211,71.169189453125,71.2188491821289,71.4231948852539,71.4931106567383,71.5280303955078,71.6524887084961,71.6774368286133,71.8351516723633,71.9236373901367,71.9547882080078,71.9730529785156,71.9608383178711,71.9656219482422,71.9786682128906,72.0250015258789,72.0542449951172,72.0829162597656,72.129150390625,72.1374969482422,72.1830596923828,72.1925277709961,72.2103042602539,72.2365264892578,72.2548370361328,72.2652893066406,72.3077163696289,72.382080078125,72.423828125,72.4507293701172,72.5226058959961,72.5516128540039,72.5879821777344,72.6043930053711,72.6169967651367,72.644775390625,72.7314453125,72.7760772705078,72.8133239746094,72.8433151245117,72.8631820678711,72.9259262084961,72.9441375732422,72.9650421142578,72.9887161254883,73.0053253173828,73.0189437866211,73.022705078125,73.0587921142578,73.0762710571289,73.1256866455078,73.2044448852539,73.2433090209961,73.418701171875,73.5273895263672,73.6442413330078,73.7681655883789,73.7853012084961,73.8275909423828,73.8568878173828,73.9020080566406,73.9532699584961,74.2481460571289,74.27001953125,74.3043441772461,74.3269958496094,74.3481903076172,74.4361343383789,74.4641647338867,74.5406265258789,74.5451126098633,74.5299758911133,74.490234375,74.4207534790039,74.3529281616211,74.2537155151367,74.2328109741211,74.1536636352539,74.1299285888672,74.1126480102539],[74.5263214111328,74.4656753540039,74.4892120361328,74.5105438232422,74.6024856567383,74.6265640258789,74.5979995727539,74.5763702392578,74.5596694946289,74.5263214111328],[74.5054168701172,74.5003890991211,74.506103515625,74.5507278442383,74.5690383911133,74.5824661254883,74.5986862182617,74.615966796875,74.6369094848633,74.6367721557617,74.585693359375,74.53955078125,74.5191879272461,74.5054168701172],[75.0363311767578,75.0309600830078,75.061279296875,75.1197280883789,75.1477584838867,75.1604995727539,75.1730499267578,75.2110366821289,75.3207015991211,75.3497619628906,75.4130401611328,75.429931640625,75.4245071411133,75.391845703125,75.3708038330078,75.3455123901367,75.2890625,75.25244140625,75.210693359375,75.1865692138672,75.1629409790039,75.1390609741211,75.114990234375,75.0797348022461,75.0363311767578],[75.0279235839844,74.9519577026367,74.9213333129883,74.8564987182617,74.77197265625,74.7564926147461,74.7493133544922,74.7275390625,74.6910629272461,74.6688537597656,74.660888671875,74.6441879272461,74.6474075317383,74.63671875,74.6421890258789,74.6601104736328,74.699951171875,74.7940902709961,74.79736328125,74.8304214477539,74.9325180053711,74.9507827758789,74.9203109741211,74.9271926879883,74.9477081298828,74.98193359375,74.9994659423828,74.9903793334961,75.0018539428711,75.0317840576172,75.057861328125,75.0987319946289,75.2113800048828,75.2192916870117,75.2400894165039,75.3009262084961,75.3583068847656,75.4437942504883,75.4690399169922,75.5286636352539,75.621826171875,75.6300277709961,75.623046875,75.5933609008789,75.5440902709961,75.4225082397461,75.3490219116211,75.2735366821289,75.230224609375,75.1368637084961,75.100341796875,75.0511703491211,75.0279235839844],[75.5101013183594,75.4772415161133,75.4748077392578,75.5059585571289,75.4942398071289,75.4312973022461,75.3941879272461,75.3807601928711,75.369140625,75.3796920776367,75.4126434326172,75.4680633544922,75.5098190307617,75.5379409790039,75.5836486816406,75.6063461303711,75.6307144165039,75.6468276977539,75.6546401977539,75.613525390625,75.5857925415039,75.5543441772461,75.5418472290039,75.5101013183594],[75.7452621459961,75.7406234741211,75.7518539428711,75.7774963378906,75.8475112915039,75.8675308227539,75.8838424682617,75.9029769897461,75.9274444580078,75.9375,75.927978515625,75.9066925048828,75.8445281982422,75.814453125,75.7887725830078,75.7657241821289,75.7452621459961],[75.7493209838867,75.7488708496094,75.7696762084961,75.7914047241211,75.8211898803711,75.8589859008789,75.8892059326172,75.911865234375,75.9307632446289,75.9459457397461,75.9544448852539,75.9572296142578,75.9964370727539,75.9921875,75.971435546875,75.9170913696289,75.8848648071289,75.8259735107422,75.7880859375,75.7763214111328,75.7659149169922,75.7493209838867],[75.5796890258789,75.5154342651367,75.5221176147461,75.5693359375,75.5853500366211,75.6010284423828,75.6173324584961,75.6625442504883,75.6986312866211,75.7695846557617,75.90625,75.9579620361328,75.9944839477539,76.0760726928711,76.1124496459961,76.1150894165039,76.0994110107422,76.0771942138672,75.9659652709961,75.921142578125,75.8054656982422,75.6111831665039,75.5796890258789],[75.9258804321289,75.8669967651367,75.8696823120117,75.8311538696289,75.8256301879883,75.8429183959961,75.8832550048828,75.9583511352539,76.0108413696289,76.0923843383789,76.1458969116211,76.1346740722656,76.1063003540039,76.0254440307617,75.9851608276367,75.9258804321289],[76.014892578125,75.9535140991211,75.9393997192383,75.927734375,75.8951644897461,75.8691864013672,75.8084030151367,75.7802276611328,75.763427734375,75.7642059326172,75.8229446411133,75.9188537597656,75.95849609375,75.8923797607422,75.93310546875,76.0370101928711,76.0465316772461,76.0469741821289,76.1084976196289,76.1823272705078,76.2229995727539,76.2582015991211,76.3114776611328,76.3070297241211,76.2816390991211,76.1964340209961,76.0950698852539,76.08642578125,76.014892578125],[76.5831069946289,76.5775909423828,76.5975112915039,76.60107421875,76.5634307861328,76.5388641357422,76.4774475097656,76.4498596191406,76.4314956665039,76.4051742553711,76.370849609375,76.3475570678711,76.3290481567383,76.3262710571289,76.3346176147461,76.3651351928711,76.4789505004883,76.5401840209961,76.5827178955078,76.6064910888672,76.63037109375,76.6661148071289,76.6661148071289,76.656982421875,76.6387634277344,76.6141662597656,76.5831069946289],[76.4664993286133,76.4218292236328,76.3873977661133,76.3631362915039,76.3352508544922,76.3037109375,76.2242202758789,76.1815414428711,76.1387176513672,76.1094284057617,76.0526428222656,75.9791488647461,75.940185546875,75.8793411254883,75.85107421875,75.8025894165039,75.7603454589844,75.7380905151367,75.6845703125,75.6725158691406,75.5520935058594,75.4198226928711,75.417236328125,75.4586410522461,75.5077590942383,75.4161148071289,75.2603073120117,75.1908187866211,75.1511688232422,75.1272964477539,75.1166534423828,75.121826171875,75.1532669067383,75.2008361816406,75.1991653442383,75.1762161254883,75.1529846191406,75.1106948852539,75.0601577758789,75.0457992553711,75.03271484375,75.0321731567383,75.0093307495117,75.0058135986328,75.0181655883789,75.0210876464844,75.0157241821289,75.0257797241211,75.0494155883789,75.0437545776367,74.9837417602539,75.0028305053711,75.0077133178711,75.0277328491211,75.0667495727539,75.1884307861328,75.2190933227539,75.2356491088867,75.2461471557617,75.28955078125,75.3214340209961,75.3465347290039,75.3943328857422,75.4063491821289,75.4609909057617,75.5685043334961,75.6122589111328,75.6333999633789,75.6553726196289,75.6686096191406,75.6983871459961,75.6812515258789,75.6204147338867,75.5904312133789,75.60791015625,75.5136184692383,75.513671875,75.5436096191406,75.5996551513672,75.6387252807617,75.7128448486328,75.7777328491211,75.8127899169922,75.8750457763672,75.8838424682617,75.8326644897461,75.7819290161133,75.7582015991211,75.7628860473633,75.7984313964844,75.8023986816406,75.7836380004883,75.7891082763672,75.8458480834961,75.8819351196289,75.9180679321289,75.943603515625,75.9919967651367,76.0079116821289,76.0413589477539,76.0830993652344,76.1012725830078,76.1501007080078,76.21728515625,76.23583984375,76.234375,76.2431106567383,76.2848587036133,76.3311996459961,76.3990249633789,76.4390182495117,76.4512710571289,76.4510269165039,76.4249038696289,76.4104995727539,76.3451614379883,76.307861328125,76.2719268798828,76.2455596923828,76.2070770263672,76.0076599121094,75.96044921875,75.9395523071289,75.9242172241211,75.9273986816406,75.9414596557617,75.959716796875,76.0294952392578,76.0666046142578,76.1172332763672,76.133544921875,76.1392059326172,76.1326141357422,76.1462860107422,76.1675796508789,76.1958465576172,76.1972122192383,76.2425231933594,76.2566909790039,76.2711410522461,76.291259765625,76.2998962402344,76.3124542236328,76.3427734375,76.3592758178711,76.3959426879883,76.43701171875,76.4565887451172,76.4754867553711,76.5238723754883,76.5846176147461,76.6135787963867,76.634765625,76.6322326660156,76.6241226196289,76.5212860107422,76.4536590576172,76.465576171875,76.5365753173828,76.6145553588867,76.6431655883789,76.69384765625,76.6673812866211,76.5987243652344,76.5753402709961,76.5329132080078,76.518798828125,76.49609375,76.4664993286133],[76.579345703125,76.5749969482422,76.5870132446289,76.6045913696289,76.6277389526367,76.6654281616211,76.7341766357422,76.74267578125,76.7524871826172,76.7503433227539,76.734130859375,76.649169921875,76.579345703125],[76.05712890625,76.0542984008789,76.0652313232422,76.0577163696289,76.0353088378906,75.9954071044922,75.9740219116211,75.9556198120117,75.9402847290039,75.839599609375,75.8047866821289,75.7962875366211,75.8021469116211,75.8775863647461,75.9011764526367,75.9065933227539,75.8915481567383,75.8785629272461,75.8631820678711,75.8454132080078,75.7730941772461,75.6796340942383,75.6892623901367,75.7416458129883,75.7824249267578,75.8099670410156,75.8190383911133,75.8415985107422,75.8724136352539,75.9300765991211,75.9515609741211,75.9746627807617,76.0237274169922,76.0530319213867,76.0601043701172,76.0089797973633,75.9669876098633,75.9453582763672,75.929931640625,75.8806610107422,75.7456588745117,75.7022476196289,75.6323699951172,75.5013656616211,75.4124984741211,75.1915512084961,75.1314926147461,75.0894546508789,75.0154266357422,74.9400939941406,74.9281692504883,74.9271469116211,74.9521484375,75.0000457763672,74.9864730834961,74.951904296875,74.942626953125,74.9472122192383,74.9595718383789,74.991943359375,75.0067367553711,75.0232925415039,75.0403289794922,75.0648956298828,75.0103073120117,74.8827667236328,74.8399887084961,74.81396484375,74.7803726196289,74.752685546875,74.6876983642578,74.6387252807617,74.5851516723633,74.501953125,74.4168472290039,74.4019088745117,74.4300842285156,74.4530334472656,74.4889602661133,74.57373046875,74.6043472290039,74.6708526611328,74.715087890625,74.76611328125,74.8125534057617,74.8752899169922,74.9755859375,74.9944305419922,75.009765625,75.0003890991211,75.0056686401367,75.0194320678711,75.0556182861328,75.1277389526367,75.1952133178711,75.2267608642578,75.2563018798828,75.2604446411133,75.1911163330078,75.1677780151367,75.1661605834961,75.1424255371094,75.1329040527344,75.1336898803711,75.1999969482422,75.211669921875,75.2046890258789,75.1871566772461,75.13818359375,75.12060546875,75.1078109741211,75.09326171875,75.068603515625,75.0838394165039,75.1122055053711,75.1294937133789,75.187744140625,75.2109375,75.2593765258789,75.2963409423828,75.3217315673828,75.3966827392578,75.4161148071289,75.412109375,75.3754425048828,75.38818359375,75.4300765991211,75.4342727661133,75.4168930053711,75.3923797607422,75.2912139892578,75.239501953125,75.2499465942383,75.2811508178711,75.285400390625,75.2754898071289,75.2580108642578,75.1712875366211,75.1409683227539,75.087890625,74.999755859375,74.9761734008789,74.9852981567383,75.0094757080078,75.048828125,75.1015625,75.1133804321289,75.114990234375,75.1041030883789,75.0807113647461,75.0558624267578,75.0095672607422,74.9741668701172,74.9681167602539,75.0415573120117,75.1717758178711,75.1515121459961,75.1561050415039,75.203857421875,75.2333526611328,75.2716751098633,75.2925262451172,75.3140640258789,75.3567886352539,75.4214859008789,75.4423370361328,75.4595184326172,75.4729995727539,75.4805145263672,75.4829635620117,75.4929656982422,75.6180725097656,75.6385726928711,75.6785202026367,75.6950149536133,75.705810546875,75.606689453125,75.5853500366211,75.6015167236328,75.6171417236328,75.6448745727539,75.7184143066406,75.7457504272461,75.7715835571289,75.8082046508789,75.840576171875,75.84130859375,75.8669967651367,75.8963394165039,75.8948211669922,75.8810501098633,75.890625,75.9292907714844,75.9575729370117,75.9915542602539,76.016845703125,76.07373046875,76.0958023071289,76.1432189941406,76.1944351196289,76.2017059326172,76.1842269897461,76.1661148071289,76.172607421875,76.1948699951172,76.2114791870117,76.2398452758789,76.2525405883789,76.27001953125,76.2957992553711,76.329833984375,76.3647003173828,76.4375,76.4974594116211,76.5057144165039,76.5017623901367,76.4748077392578,76.4514617919922,76.4226608276367,76.3958511352539,76.3494567871094,76.3311996459961,76.3007354736328,76.2291564941406,76.2068405151367,76.2484359741211,76.2577667236328,76.2446746826172,76.2017059326172,76.0718765258789,75.9393081665039,75.9107437133789,75.8664093017578,75.847412109375,75.834228515625,75.8255386352539,75.8320770263672,75.8221206665039,75.8106918334961,75.7621612548828,75.6764678955078,75.6125030517578,75.5702133178711,75.5485382080078,75.5469284057617,75.5595245361328,75.5553207397461,75.5064926147461,75.5149917602539,75.5417938232422,75.5869674682617,75.6240768432617,75.6747512817383,75.698974609375,75.863037109375,75.9290542602539,76.021240234375,76.0425338745117,76.0718231201172,76.109130859375,76.1446762084961,76.2124557495117,76.22265625,76.2894592285156,76.3063507080078,76.3329544067383,76.369384765625,76.3974151611328,76.4169921875,76.4847640991211,76.5223693847656,76.691650390625,76.7599639892578,76.7919921875,76.8118667602539,76.8211441040039,76.758056640625,76.7541961669922,76.7376022338867,76.708251953125,76.6802749633789,76.6297378540039,76.6085433959961,76.5867233276367,76.5363235473633,76.5031280517578,76.4471664428711,76.4389114379883,76.3916473388672,76.330078125,76.2334518432617,76.2000503540039,76.154052734375,76.11572265625,76.0850524902344,76.0665512084961,76.05712890625],[76.5074234008789,76.4938507080078,76.5008773803711,76.487548828125,76.4951171875,76.5235824584961,76.5794906616211,76.6627960205078,76.7345733642578,76.7542953491211,76.787841796875,76.8101577758789,76.8369674682617,76.8362350463867,76.7820281982422,76.763427734375,76.7411651611328,76.7198257446289,76.701171875,76.6596221923828,76.630615234375,76.6021957397461,76.5610885620117,76.533935546875,76.5208053588867,76.5074234008789],[76.7432632446289,76.7105407714844,76.7588882446289,76.7740707397461,76.794677734375,76.8510284423828,76.8753433227539,76.8948669433594,76.8729553222656,76.8473129272461,76.8250427246094,76.7832489013672,76.7432632446289],[76.9124526977539,76.9037551879883,76.9170913696289,76.9141616821289,76.8738250732422,76.8122024536133,76.7843246459961,76.770263671875,76.7546844482422,76.7080078125,76.6863784790039,76.6690902709961,76.62646484375,76.5736846923828,76.5271453857422,76.4920425415039,76.4477081298828,76.47412109375,76.6204147338867,76.6029739379883,76.6160125732422,76.67578125,76.6851119995117,76.6807632446289,76.6619110107422,76.5813446044922,76.54296875,76.5157775878906,76.4957580566406,76.4646987915039,76.4835891723633,76.500244140625,76.5105972290039,76.509765625,76.49853515625,76.4558639526367,76.4464797973633,76.4373016357422,76.3016128540039,76.2582015991211,76.239013671875,76.2177276611328,76.1891632080078,76.1580123901367,76.1856002807617,76.22998046875,76.2200698852539,76.159912109375,76.1415557861328,76.10595703125,76.076171875,76.053466796875,76.0302734375,75.9708938598633,75.96630859375,75.9248504638672,75.8536148071289,75.8440933227539,75.85693359375,75.7950668334961,75.7620086669922,75.7372589111328,75.698486328125,75.6457977294922,75.583740234375,75.5650405883789,75.5723648071289,75.5641098022461,75.4539566040039,75.4519500732422,75.4634780883789,75.5024871826172,75.53857421875,75.588623046875,75.6249008178711,75.6474609375,75.6586380004883,75.658447265625,75.6451187133789,75.5120086669922,75.5756301879883,75.5470657348633,75.49365234375,75.48486328125,75.59130859375,75.6177291870117,75.4913558959961,75.46337890625,75.436279296875,75.4063491821289,75.39501953125,75.4419479370117,75.5022430419922,75.5287094116211,75.5797882080078,75.5726089477539,75.6449203491211,75.6546859741211,75.6534729003906,75.7626419067383,75.7799301147461,75.8189468383789,75.8128356933594,75.7508239746094,75.7564392089844,75.8182601928711,75.8333511352539,75.8310546875,75.794921875,75.7560043334961,75.7355499267578,75.6843795776367,75.6692428588867,75.6581573486328,75.6431121826172,75.6421356201172,75.6290969848633,75.5811538696289,75.5620651245117,75.5421371459961,75.4903793334961,75.4794464111328,75.4674301147461,75.4614715576172,75.4495162963867,75.3848648071289,75.2953643798828,75.2598114013672,75.1993179321289,75.1186065673828,75.0515594482422,75.0341796875,75.0021514892578,74.9880905151367,74.9909210205078,75.0214385986328,75.0208511352539,74.9897003173828,74.9587936401367,74.9176254272461,74.880126953125,74.8336410522461,74.8948287963867,74.9083023071289,74.9029769897461,74.8761749267578,74.8276901245117,74.7957000732422,74.7801742553711,74.7494583129883,74.70361328125,74.6570358276367,74.58447265625,74.5815963745117,74.566650390625,74.5535125732422,74.5023422241211,74.4766082763672,74.4727096557617,74.4820785522461,74.5352020263672,74.5251998901367,74.5302734375,74.5655746459961,74.5834426879883,74.6297836303711,74.693115234375,74.7321319580078,74.7883758544922,74.8165512084961,74.8167419433594,74.8284149169922,74.8848114013672,74.9014663696289,74.8922882080078,74.8481903076172,74.8341369628906,74.8019027709961,74.7646026611328,74.670166015625,74.6549758911133,74.585693359375,74.56591796875,74.5643997192383,74.5151824951172,74.5081024169922,74.5195770263672,74.5419921875,74.5676727294922,74.6041946411133,74.60693359375,74.5276870727539,74.5174331665039,74.5186538696289,74.5433120727539,74.6005859375,74.600341796875,74.5669937133789,74.5451126098633,74.5347671508789,74.4989776611328,74.498779296875,74.5397491455078,74.5355911254883,74.5134811401367,74.555419921875,74.5570373535156,74.4891128540039,74.4786148071289,74.4803237915039,74.502197265625,74.4703598022461,74.4893569946289,74.494140625,74.5097122192383,74.5414505004883,74.5697250366211,74.6087875366211,74.6669006347656,74.744140625,74.7848587036133,74.8037109375,74.8289108276367,74.8317337036133,74.802001953125,74.7151870727539,74.6899871826172,74.7110824584961,74.7638168334961,74.7894973754883,74.7880859375,74.7740249633789,74.7472686767578,74.7375946044922,74.744873046875,74.7317810058594,74.6982421875,74.6668395996094,74.6375503540039,74.6091842651367,74.5679244995117,74.5547409057617,74.548583984375,74.5608825683594,74.6104507446289,74.6127471923828,74.6958999633789,74.715087890625,74.7451629638672,74.8010787963867,74.8177795410156,74.7362747192383,74.7102584838867,74.6498565673828,74.6455078125,74.667236328125,74.6506805419922,74.6555633544922,74.6991729736328,74.7435073852539,74.793212890625,74.9483871459961,75.0510711669922,75.072021484375,75.1009750366211,75.12353515625,75.1812515258789,75.2297897338867,75.2633285522461,75.2972640991211,75.3463821411133,75.3827133178711,75.40625,75.4794464111328,75.6106414794922,75.6344757080078,75.6579132080078,75.727294921875,75.7968292236328,75.8465347290039,75.9151382446289,75.9864730834961,76.1144561767578,76.2139587402344,76.35400390625,76.3660125732422,76.359619140625,76.3114318847656,76.2885284423828,76.2731475830078,76.2696762084961,76.2823257446289,76.2971649169922,76.2932586669922,76.2577133178711,76.264404296875,76.363037109375,76.4161605834961,76.4459991455078,76.4867172241211,76.5133285522461,76.5372085571289,76.5634307861328,76.5846710205078,76.56640625,76.5696334838867,76.7029266357422,76.7264175415039,76.7383270263672,76.7540054321289,76.7734832763672,76.8027877807617,76.7657699584961,76.7630310058594,76.7740707397461,76.7972183227539,76.8106918334961,76.8551712036133,76.888916015625,76.9134750366211,76.9482421875,76.9717788696289,76.9850082397461,76.9879379272461,77.0045928955078,77.050048828125,77.0662155151367,77.0637664794922,77.017333984375,76.9583511352539,76.9124526977539],[77.36328125,77.3243179321289,77.3086471557617,77.2921447753906,77.2659225463867,77.17822265625,77.1370620727539,77.1016616821289,77.0299758911133,76.96923828125,76.9391098022461,76.908447265625,76.918212890625,76.9155731201172,76.9014129638672,76.8743209838867,76.7845230102539,76.7362365722656,76.7112808227539,76.6869125366211,76.6535110473633,76.611083984375,76.5771484375,76.5315933227539,76.49609375,76.4690933227539,76.4035110473633,76.3731002807617,76.3219223022461,76.2980041503906,76.2815399169922,76.2725601196289,76.272705078125,76.3167495727539,76.3448257446289,76.4058074951172,76.446533203125,76.4966812133789,76.5740203857422,76.653076171875,76.7339401245117,76.7842788696289,76.8041076660156,76.8050842285156,76.7723617553711,76.760498046875,76.7366714477539,76.7009735107422,76.6623001098633,76.5879898071289,76.54736328125,76.5251922607422,76.5255813598633,76.5129852294922,76.48583984375,76.4637680053711,76.4175262451172,76.3658676147461,76.3346633911133,76.277099609375,76.2578125,76.1676712036133,76.1448745727539,76.1240692138672,76.1265182495117,76.1594696044922,76.2217788696289,76.275390625,76.3203125,76.3402786254883,76.3265151977539,76.2798767089844,76.2436981201172,76.1898956298828,76.1566925048828,76.1177215576172,76.0999526977539,76.0520553588867,76.0305633544922,75.9972229003906,75.982177734375,75.9845733642578,75.9459915161133,75.9154281616211,75.8588409423828,75.8519592285156,75.8247604370117,75.8256301879883,75.8701629638672,75.9583511352539,76.0084457397461,76.0340347290039,76.1340866088867,76.1663055419922,76.1792984008789,76.1826629638672,76.1634292602539,76.020263671875,75.9836883544922,75.9770050048828,75.9811019897461,76.0203094482422,76.0347671508789,76.0182189941406,75.9598083496094,75.9442367553711,75.950927734375,75.9729995727539,76.0090866088867,76.031005859375,76.0386734008789,76.0805206298828,76.0973129272461,76.1214370727539,76.1402816772461,76.1529769897461,76.1620559692383,76.16748046875,76.1623992919922,76.1347198486328,76.164794921875,76.2276840209961,76.3531723022461,76.3900833129883,76.4012222290039,76.4414520263672,76.4534683227539,76.6221694946289,76.6607437133789,76.6914520263672,76.793212890625,76.8164520263672,76.8869171142578,76.9045867919922,76.9313507080078,77.0738754272461,77.1769027709961,77.2406768798828,77.3050765991211,77.3327102661133,77.3812026977539,77.3173828125,77.3133773803711,77.3314437866211,77.36083984375,77.3484878540039,77.3395538330078,77.3465881347656,77.3799285888672,77.3982391357422,77.431884765625,77.448974609375,77.4651336669922,77.5038604736328,77.5288543701172,77.547607421875,77.5428237915039,77.5160140991211,77.4606399536133,77.36328125],[77.2676315307617,77.2103958129883,77.2124557495117,77.3294906616211,77.3873062133789,77.42626953125,77.4815444946289,77.5571823120117,77.6080551147461,77.625732421875,77.6438980102539,77.6549835205078,77.6486282348633,77.6283721923828,77.5946807861328,77.4914093017578,77.442138671875,77.3781280517578,77.3389663696289,77.3104019165039,77.2925796508789,77.2676315307617],[77.1417465209961,77.1239776611328,77.1646041870117,77.1820831298828,77.2542495727539,77.3526382446289,77.4613800048828,77.525390625,77.5634307861328,77.6265106201172,77.7253875732422,77.7398376464844,77.75439453125,77.7359848022461,77.700927734375,77.64208984375,77.5482940673828,77.5067367553711,77.4496612548828,77.418701171875,77.4132308959961,77.3377456665039,77.3085479736328,77.2491149902344,77.2208023071289,77.162353515625,77.1417465209961],[77.7919921875,77.7538070678711,77.7740707397461,77.7762145996094,77.7599105834961,77.770751953125,77.7643051147461,77.7397918701172,77.710205078125,77.66015625,77.6296920776367,77.4744110107422,77.4666519165039,77.4645462036133,77.4522476196289,77.4742202758789,77.484130859375,77.5034713745117,77.5945281982422,77.6305618286133,77.6725540161133,77.7005386352539,77.71435546875,77.7822799682617,77.7919921875],[77.6965866088867,77.6873550415039,77.6921844482422,77.7281265258789,77.7701644897461,77.836669921875,77.8734893798828,77.880615234375,77.8893508911133,77.8996124267578,77.89208984375,77.8541488647461,77.8297882080078,77.8125991821289,77.77783203125,77.762451171875,77.735107421875,77.7290573120117,77.7183151245117,77.6965866088867],[77.754638671875,77.720703125,77.7214889526367,77.7693328857422,77.9154281616211,77.967529296875,77.9769058227539,77.992919921875,78.0155258178711,78.0403366088867,78.0775375366211,78.0631790161133,78.03271484375,78.0042953491211,77.9982376098633,77.9779281616211,77.9155731201172,77.903564453125,77.889892578125,77.8689422607422,77.8324203491211,77.8134765625,77.7757797241211,77.754638671875],[78.1032180786133,78.0792465209961,78.0747604370117,78.0568389892578,77.9993209838867,77.9574203491211,77.9048385620117,77.8572235107422,77.8341293334961,77.8203582763672,77.8031768798828,77.7814407348633,77.786376953125,77.7770004272461,77.762939453125,77.7423782348633,77.715576171875,77.6247024536133,77.5721206665039,77.5245132446289,77.4906234741211,77.4458999633789,77.4259796142578,77.4331512451172,77.4285125732422,77.3441848754883,77.34375,77.3641128540039,77.4437026977539,77.4749526977539,77.5107421875,77.5302734375,77.558837890625,77.5801773071289,77.5997543334961,77.6175308227539,77.6326217651367,77.6764602661133,77.7183151245117,77.7784194946289,77.8130416870117,77.8356475830078,77.8600616455078,77.9035186767578,77.912353515625,77.9191360473633,77.9416046142578,78.0067825317383,78.088134765625,78.0806121826172,78.0965805053711,78.1032180786133],[78.1464309692383,78.1263656616211,78.1381378173828,78.150634765625,78.165771484375,78.1834945678711,78.245849609375,78.2672348022461,78.271240234375,78.2500457763672,78.2181625366211,78.1464309692383],[78.650390625,78.5920944213867,78.59326171875,78.5824737548828,78.5671844482422,78.4928665161133,78.4560089111328,78.408447265625,78.36865234375,78.336669921875,78.31640625,78.3037643432617,78.3228073120117,78.2981948852539,78.2949676513672,78.3107452392578,78.3223114013672,78.3676300048828,78.3862762451172,78.3763198852539,78.3365707397461,78.287353515625,78.2747039794922,78.282958984375,78.3660659790039,78.3415069580078,78.2929229736328,78.2837905883789,78.2978973388672,78.3343734741211,78.352783203125,78.4084014892578,78.4668426513672,78.4998016357422,78.5478057861328,78.57470703125,78.6032257080078,78.64404296875,78.7083969116211,78.7350616455078,78.7578125,78.7566452026367,78.7044372558594,78.678466796875,78.650390625],[78.5313034057617,78.505126953125,78.5166015625,78.49755859375,78.4302673339844,78.3905258178711,78.3604965209961,78.34521484375,78.3125991821289,78.2626419067383,78.2250518798828,78.1780776977539,78.1362762451172,78.1063995361328,78.0756301879883,77.9926223754883,77.9681701660156,77.9708023071289,77.9632339477539,77.9244613647461,77.8873977661133,77.8721694946289,77.8493118286133,77.8119125366211,77.8060073852539,77.8274459838867,77.8590850830078,77.8809585571289,77.9081039428711,77.9334945678711,77.9822769165039,78.0502471923828,78.0716323852539,78.0906219482422,78.1032180786133,78.116943359375,78.1390075683594,78.15185546875,78.1574249267578,78.2032241821289,78.2306137084961,78.2623596191406,78.325927734375,78.3863220214844,78.4030227661133,78.4292449951172,78.4378890991211,78.4768524169922,78.4981384277344,78.5174789428711,78.558349609375,78.586669921875,78.6923828125,78.751220703125,78.7735366821289,78.8045425415039,78.8052291870117,78.7957992553711,78.7829132080078,78.7576675415039,78.7202682495117,78.6871109008789,78.6651840209961,78.5953598022461,78.5731887817383,78.5511245727539,78.5313034057617],[79.3156280517578,79.2953109741211,79.2311019897461,79.0950164794922,79.0775909423828,79.0549774169922,79.0271453857422,78.9910125732422,78.9693374633789,78.9429702758789,78.9301223754883,78.9009246826172,78.87939453125,78.9006881713867,78.9332046508789,78.9541015625,79.0102996826172,79.0383834838867,79.0792007446289,79.08837890625,79.0789031982422,78.982177734375,78.972900390625,78.9615249633789,78.9390182495117,78.9146957397461,78.8582992553711,78.823974609375,78.8016586303711,78.7829132080078,78.8203125,78.7286148071289,78.61962890625,78.5830535888672,78.5632781982422,78.544677734375,78.4935073852539,78.4553680419922,78.438232421875,78.3929672241211,78.3645477294922,78.3250961303711,78.302978515625,78.2793426513672,78.1175308227539,78.0723648071289,78.0159606933594,77.9968719482422,77.9656295776367,77.8771438598633,77.85693359375,77.8396530151367,77.8240737915039,77.7938003540039,77.8327178955078,77.8917999267578,77.9306640625,77.9776916503906,77.9960479736328,78.0716323852539,78.0877456665039,78.1302261352539,78.19384765625,78.1993713378906,78.2641143798828,78.279541015625,78.2750015258789,78.2489318847656,78.2559051513672,78.2752456665039,78.306884765625,78.3301773071289,78.3710479736328,78.319580078125,78.260009765625,78.2694778442383,78.2946319580078,78.3516616821289,78.4012680053711,78.468017578125,78.5185089111328,78.5526428222656,78.5728988647461,78.5793914794922,78.5397415161133,78.51953125,78.5398483276367,78.5939483642578,78.6230010986328,78.6349105834961,78.6633758544922,78.6926803588867,78.736328125,78.7516098022461,78.7691421508789,78.7640151977539,78.7813034057617,78.7956008911133,78.81396484375,78.9026870727539,78.9188003540039,78.9478530883789,78.985595703125,78.9898910522461,78.9561538696289,78.825927734375,78.8070755004883,78.8081588745117,78.8291549682617,78.8564910888672,78.8901901245117,78.9911117553711,79.027099609375,79.0511169433594,79.033203125,79.0325241088867,79.0609817504883,79.1142120361328,79.1642074584961,79.2424850463867,79.30224609375,79.3235855102539,79.3109893798828,79.34814453125,79.35205078125,79.3156280517578],[79.9811019897461,79.8508834838867,79.785400390625,79.737060546875,79.7256851196289,79.7240753173828,79.7618637084961,79.7840805053711,79.8028793334961,79.8395462036133,79.8871536254883,79.8794937133789,79.884033203125,79.8982391357422,79.9186477661133,80.0012664794922,80.0304183959961,80.0811080932617,80.0933609008789,80.14013671875,80.1440963745117,80.1264190673828,80.1242218017578,80.1111297607422,80.081787109375,80.037353515625,79.9811019897461],[81.1328582763672,81.04931640625,80.85009765625,80.7776794433594,80.6876983642578,80.6553192138672,80.6417007446289,80.5936965942383,80.575927734375,80.5482406616211,80.4984359741211,80.5012741088867,80.53076171875,80.538818359375,80.53173828125,80.5106430053711,80.4796371459961,80.4574203491211,80.4402389526367,80.4069366455078,80.39404296875,80.3785095214844,80.3603515625,80.2892532348633,80.26318359375,80.198486328125,80.1662139892578,80.1311492919922,80.1336898803711,80.1114730834961,80.1251983642578,80.16650390625,80.2251968383789,80.25537109375,80.2897491455078,80.3482894897461,80.3868637084961,80.4180221557617,80.4280700683594,80.4294967651367,80.4156265258789,80.3721237182617,80.3484420776367,80.3016128540039,80.2074737548828,80.1872100830078,80.1338882446289,80.0977020263672,80.0875015258789,80.0794448852539,80.0465316772461,80.043212890625,79.9669494628906,79.8942413330078,79.805419921875,79.6626510620117,79.6299285888672,79.5801696777344,79.5711441040039,79.5909652709961,79.5977096557617,79.646240234375,79.6349563598633,79.6224136352539,79.6054153442383,79.5512237548828,79.4794387817383,79.4859848022461,79.5730514526367,79.5945281982422,79.6152801513672,79.6114273071289,79.5303192138672,79.3872528076172,79.3281707763672,79.2845687866211,79.2337417602539,79.2083511352539,79.0999984741211,79.0386657714844,78.9913101196289,78.9754867553711,78.9828109741211,78.9748992919922,78.898681640625,78.8661117553711,78.8134765625,78.7181625366211,78.6763229370117,78.7068405151367,78.7513656616211,78.8516082763672,78.8931198120117,78.9150390625,78.9505844116211,79.0363235473633,79.0453186035156,79.0381851196289,78.9953079223633,78.9728088378906,78.9334945678711,78.867431640625,78.7455139160156,78.6963882446289,78.6719665527344,78.6530303955078,78.626953125,78.6155242919922,78.5947265625,78.5647506713867,78.5370635986328,78.4944305419922,78.4770965576172,78.4965362548828,78.6019058227539,78.5960922241211,78.5840301513672,78.5464401245117,78.4621124267578,78.3919906616211,78.333740234375,78.24169921875,78.1924285888672,78.1858825683594,78.1844253540039,78.209228515625,78.3702087402344,78.4388656616211,78.5730514526367,78.6006851196289,78.6068344116211,78.5491714477539,78.4957962036133,78.3702087402344,78.2789077758789,78.2032699584961,78.1719284057617,78.1596145629883,78.1666030883789,78.1930160522461,78.2375946044922,78.262451171875,78.291259765625,78.3130874633789,78.3280258178711,78.3309097290039,78.3217315673828,78.3077163696289,78.2685546875,78.2466812133789,78.21875,78.1847686767578,78.1632766723633,78.1498565673828,78.1584014892578,78.18798828125,78.2368698120117,78.3128967285156,78.3891143798828,78.4297409057617,78.4601135253906,78.4866638183594,78.5207977294922,78.5426712036133,78.5617141723633,78.6050262451172,78.6129379272461,78.6015625,78.6082992553711,78.6426773071289,78.7078170776367,78.7509231567383,78.7677688598633,78.77734375,78.7691879272461,78.775634765625,78.8080520629883,78.8722152709961,78.9289093017578,78.9510269165039,78.9728088378906,78.9941940307617,79.0374069213867,79.1395034790039,79.1553726196289,79.1563949584961,79.185791015625,79.2826156616211,79.3174285888672,79.3608932495117,79.3727035522461,79.36474609375,79.3734359741211,79.4392623901367,79.450439453125,79.4498977661133,79.4292449951172,79.3681640625,79.3539581298828,79.2907257080078,79.2952194213867,79.302734375,79.3150863647461,79.3570327758789,79.3856887817383,79.3955078125,79.4015655517578,79.4004364013672,79.3905258178711,79.3350601196289,79.2935485839844,79.2898864746094,79.354736328125,79.390380859375,79.418212890625,79.52734375,79.5497589111328,79.5680618286133,79.6671447753906,79.6861877441406,79.736328125,79.7256393432617,79.6771926879883,79.653076171875,79.6532211303711,79.66015625,79.6737823486328,79.7046890258789,79.8475036621094,79.9166488647461,79.9776840209961,80.024169921875,80.1357879638672,80.06640625,80.0532760620117,80.0487289428711,80.0555191040039,80.0736312866211,80.1029739379883,80.1255874633789,80.1414031982422,80.181640625,80.1948699951172,80.2015151977539,80.1343765258789,80.135009765625,80.23095703125,80.214111328125,80.2217254638672,80.2458953857422,80.2669906616211,80.2930679321289,80.3150405883789,80.3328628540039,80.352783203125,80.3804168701172,80.383056640625,80.3652801513672,80.3666000366211,80.396240234375,80.4708480834961,80.553466796875,80.69140625,80.7206497192383,80.7254409790039,80.6905746459961,80.685791015625,80.6464309692383,80.5708999633789,80.5723571777344,80.5580596923828,80.5591812133789,80.586181640625,80.6097183227539,80.640625,80.7512741088867,80.8083038330078,80.838134765625,80.8632278442383,81.0007858276367,81.0496597290039,81.0311965942383,81.0571746826172,81.105712890625,81.083251953125,81.0853500366211,81.1002960205078,81.1288604736328,81.1551284790039,81.1792526245117,81.2090835571289,81.2132797241211,81.2249984741211,81.2409210205078,81.2649383544922,81.2896957397461,81.3151397705078,81.3307647705078,81.3392639160156,81.3493118286133,81.3505859375,81.3644027709961,81.3462905883789,81.2782745361328,81.2436065673828,81.1855010986328,81.1328582763672],[83.0167999267578,82.998779296875,83.0052719116211,82.961181640625,82.9560089111328,82.9644012451172,82.9539031982422,82.9440383911133,82.9268569946289,82.9061508178711,82.8612289428711,82.8179244995117,82.6768035888672,82.6533660888672,82.6459426879883,82.6524429321289,82.6681137084961,82.716064453125,82.7188491821289,82.7420959472656,82.80712890625,82.8424301147461,82.8269500732422,82.8023910522461,82.7996139526367,82.8185043334961,82.8701171875,82.888916015625,82.9022979736328,82.9008255004883,82.87646484375,82.818603515625,82.7784194946289,82.7777328491211,82.8231964111328,82.8291015625,82.8125915527344,82.7925720214844,82.771240234375,82.7487335205078,82.7293014526367,82.7129898071289,82.6940460205078,82.6534652709961,82.5652313232422,82.5328063964844,82.4668426513672,82.4501953125,82.5195770263672,82.4886245727539,82.4674301147461,82.44189453125,82.3997573852539,82.341064453125,82.2798309326172,82.1844253540039,82.1102523803711,82.0434036254883,82.0067901611328,81.8455047607422,81.7936553955078,81.7426223754883,81.7337417602539,81.7435073852539,81.7153778076172,81.6680679321289,81.6455535888672,81.6294403076172,81.6164016723633,81.5630416870117,81.5268020629883,81.4986877441406,81.485107421875,81.2933044433594,81.26123046875,81.2479934692383,81.26123046875,81.4942398071289,81.5096664428711,81.4928741455078,81.4386215209961,81.2847595214844,81.1461410522461,81.0409164428711,80.8594284057617,80.6787109375,80.5868682861328,80.4228515625,80.3832550048828,80.366943359375,80.373779296875,80.3976593017578,80.458984375,80.5275421142578,80.5395965576172,80.5055694580078,80.2777328491211,80.2335968017578,80.1870651245117,80.1458969116211,80.1359405517578,80.1433563232422,80.13916015625,80.1232452392578,80.1055679321289,80.086181640625,80.071044921875,80.122314453125,80.11865234375,80.0937042236328,80.0709991455078,79.9982452392578,79.9112777709961,79.9063491821289,79.8755416870117,79.8478012084961,79.7825622558594,79.7617645263672,79.7010726928711,79.6868209838867,79.6943893432617,79.8270950317383,79.8462905883789,79.8797836303711,79.8740692138672,79.8351593017578,79.8155746459961,79.7782211303711,79.77099609375,79.7564468383789,79.732177734375,79.6439971923828,79.5965805053711,79.552490234375,79.5215835571289,79.5040054321289,79.4951629638672,79.5014114379883,79.4905242919922,79.4647445678711,79.4535598754883,79.4586944580078,79.4210433959961,79.4141616821289,79.43115234375,79.4731903076172,79.4944305419922,79.5122985839844,79.4882354736328,79.4780731201172,79.4136276245117,79.3261260986328,79.3113327026367,79.2395553588867,79.2283248901367,79.2039031982422,79.2353515625,79.2294921875,79.052734375,79.0355453491211,79.0355453491211,79.0612258911133,79.0877456665039,79.1177749633789,79.1003952026367,79.1041488647461,79.0865249633789,79.087158203125,79.0572738647461,79.0569305419922,79.0762176513672,79.0821762084961,79.0749969482422,79.0545883178711,79.0483856201172,79.01513671875,78.9639129638672,78.9423828125,78.9545440673828,78.9784698486328,79.0178680419922,79.0242156982422,79.0068359375,78.9851531982422,78.9590377807617,78.8897399902344,78.8813018798828,78.8582992553711,78.7577133178711,78.7500991821289,78.72412109375,78.6592788696289,78.6203155517578,78.5448226928711,78.5228500366211,78.5298309326172,78.5210876464844,78.5115203857422,78.49169921875,78.4035186767578,78.355712890625,78.3277359008789,78.2210922241211,78.0098190307617,77.9931182861328,77.9873046875,77.9910125732422,77.9378890991211,77.92724609375,77.9471740722656,77.946044921875,77.9117202758789,77.8460922241211,77.7473602294922,77.615478515625,77.51904296875,77.4581069946289,77.4130859375,77.369384765625,77.3421401977539,77.33251953125,77.3310089111328,77.299560546875,77.3014678955078,77.3147964477539,77.3585891723633,77.4821319580078,77.5095672607422,77.5254440307617,77.4988327026367,77.4297866821289,77.3852005004883,77.3651809692383,77.3440475463867,77.3108444213867,77.2959442138672,77.2965316772461,77.2836456298828,77.2478561401367,77.214111328125,77.2040023803711,77.2144546508789,77.257080078125,77.2696304321289,77.2594757080078,77.2442932128906,77.1509246826172,77.1465835571289,77.193603515625,77.1960983276367,77.1584014892578,77.0851516723633,77.0257797241211,76.9803695678711,76.93603515625,76.8928680419922,76.8835906982422,76.9080047607422,76.9672393798828,76.9812545776367,76.9779815673828,76.9349060058594,76.8519515991211,76.7549819946289,76.64404296875,76.5712432861328,76.451171875,76.4039611816406,76.3547897338867,76.3105010986328,76.2512664794922,76.2401809692383,76.1764678955078,76.173583984375,76.1839370727539,76.2149963378906,76.2701644897461,76.321533203125,76.369140625,76.4086456298828,76.4700698852539,76.4984893798828,76.5127410888672,76.5044937133789,76.4876480102539,76.4844207763672,76.4949722290039,76.5208511352539,76.62939453125,76.6432113647461,76.6398010253906,76.6553726196289,76.6898880004883,76.7232894897461,76.6978073120117,76.6360321044922,76.57470703125,76.5137634277344,76.4658203125,76.4392547607422,76.453125,76.4950180053711,76.6753387451172,76.3565444946289,76.3045883178711,76.3133850097656,76.3490219116211,76.4349136352539,76.4918518066406,76.5486373901367,76.5796356201172,76.5848617553711,76.5165023803711,76.3766098022461,76.4127426147461,76.4480438232422,76.5858383178711,76.3862838745117,76.4127426147461,76.4052734375,76.580078125,76.7728500366211,76.65087890625,76.5472106933594,76.4208984375,76.4568405151367,76.4744644165039,76.491943359375,76.65966796875,76.8268051147461,76.9933624267578,77.0722198486328,77.1039581298828,77.1240234375,77.136474609375,77.1268615722656,77.13623046875,77.1658706665039,77.1744155883789,77.1849136352539,77.2002944946289,77.3077163696289,77.3321228027344,77.343017578125,77.3478012084961,77.3948287963867,77.4363784790039,77.4928207397461,77.5998077392578,77.7191925048828,77.7847137451172,77.8362274169922,77.8719253540039,77.8917999267578,77.8922348022461,77.8637161254883,77.80859375,77.7461471557617,77.6139221191406,77.5086441040039,77.4611358642578,77.3749542236328,77.3610382080078,77.3679733276367,77.3905258178711,77.4142074584961,77.4422378540039,77.4825668334961,77.5136260986328,77.5848159790039,77.7327194213867,77.8495101928711,77.8888168334961,77.9363327026367,77.9921417236328,77.96240234375,77.6737289428711,77.6212921142578,77.5326156616211,77.518310546875,77.522705078125,77.5619583129883,77.49951171875,77.515380859375,77.5590286254883,77.7638702392578,77.9276809692383,78.0105972290039,78.06201171875,78.1957015991211,78.1970672607422,78.176025390625,78.2063522338867,78.2513656616211,78.2396926879883,78.527587890625,78.3124008178711,78.1995086669922,78.1424331665039,78.1095733642578,78.0811996459961,78.1869659423828,78.3428726196289,78.2846145629883,78.197021484375,78.1510238647461,78.1268081665039,78.1326675415039,78.1766128540039,78.284423828125,78.4171905517578,78.4787063598633,78.5576171875,78.6639175415039,78.7743682861328,78.8236312866211,78.8437042236328,78.9020004272461,78.9122543334961,78.8845748901367,78.8391571044922,78.8044891357422,78.7793502807617,78.7703094482422,78.807861328125,78.8441390991211,78.8404312133789,78.8470687866211,78.8641128540039,78.8984832763672,78.9503402709961,78.9757843017578,78.974853515625,78.9618606567383,78.924072265625,78.9036636352539,78.9075241088867,78.9394989013672,78.9452667236328,78.9598159790039,78.9752883911133,78.99658203125,79.028564453125,79.0712890625,79.1012649536133,79.1185531616211,79.1221694946289,79.0986785888672,79.058837890625,79.0536575317383,79.0900421142578,79.1631393432617,79.22509765625,79.30126953125,79.3766098022461,79.4947280883789,79.6121597290039,79.6641082763672,79.6898422241211,79.721923828125,79.7428207397461,79.84521484375,80.0181655883789,80.12353515625,80.2582550048828,80.3193359375,80.2717742919922,80.2789077758789,80.261962890625,80.2289505004883,80.14697265625,80.0545883178711,79.9927749633789,79.9082565307617,79.7827606201172,79.7225646972656,79.685791015625,79.6541519165039,79.6142044067383,79.6010284423828,79.6062545776367,79.6352005004883,79.6694869995117,79.678955078125,79.6749496459961,79.693115234375,79.7334518432617,79.7877883911133,79.8902359008789,79.9571762084961,80.0663604736328,80.1749038696289,80.2778854370117,80.3226089477539,80.353759765625,80.3755340576172,80.4452667236328,80.528564453125,80.625244140625,80.6478500366211,80.7843780517578,80.8347625732422,80.8429183959961,80.8647918701172,80.878173828125,80.8964309692383,80.9054183959961,80.9048309326172,80.9214401245117,80.95166015625,81.0010681152344,81.04345703125,81.1143569946289,81.1676254272461,81.3210906982422,81.3856964111328,81.4302749633789,81.330810546875,81.2589416503906,81.1510314941406,81.1192398071289,81.1276321411133,81.1175842285156,81.0890655517578,81.036865234375,80.9609832763672,80.9093246459961,80.8817901611328,80.8419418334961,80.763916015625,80.6548843383789,80.627197265625,80.622802734375,80.5613327026367,80.5588836669922,80.5775375366211,80.6306610107422,80.7286605834961,80.7723159790039,80.7627944946289,80.7360305786133,80.7139663696289,80.674072265625,80.6017608642578,80.5562515258789,80.5377960205078,80.5267562866211,80.5211410522461,80.5259780883789,80.5811538696289,80.5621109008789,80.5657730102539,80.604736328125,80.6300277709961,80.6640167236328,80.72802734375,80.78955078125,80.9246063232422,80.9878921508789,81.0423812866211,81.1033172607422,81.14794921875,81.0980911254883,81.0350570678711,81.0119094848633,80.9500961303711,80.7262725830078,80.6697692871094,80.65625,80.6753845214844,80.7038040161133,80.7700653076172,80.8056182861328,80.8295364379883,80.8536605834961,80.8819351196289,80.914306640625,80.9413070678711,80.999755859375,80.9883804321289,81.0004348754883,81.0357437133789,81.1235809326172,81.2468795776367,81.2862319946289,81.2948684692383,81.2853012084961,81.2412109375,81.1226577758789,81.080810546875,81.0584945678711,81.0253372192383,81.032470703125,81.0648422241211,81.1247024536133,81.1726608276367,81.2264709472656,81.2390594482422,81.2500991821289,81.3020477294922,81.32861328125,81.387451171875,81.47412109375,81.5014190673828,81.518798828125,81.5093231201172,81.5258255004883,81.5586395263672,81.5646514892578,81.54150390625,81.401123046875,81.4053726196289,81.4295425415039,81.4642028808594,81.6116714477539,81.6348648071289,81.6315460205078,81.6385269165039,81.656005859375,81.6404800415039,81.5919952392578,81.5712432861328,81.5782241821289,81.6356887817383,81.683837890625,81.7442398071289,81.7877426147461,81.8274383544922,81.8772506713867,81.89404296875,81.89453125,81.916748046875,81.9554214477539,82.0180206298828,82.0610809326172,82.0964813232422,82.0850601196289,82.05419921875,82.0000991821289,81.958740234375,81.9921417236328,82.0333480834961,82.051025390625,82.0451202392578,82.0255355834961,81.9756851196289,81.9532775878906,81.9546432495117,81.9822311401367,81.9828186035156,81.9945297241211,82.0233840942383,82.0439910888672,82.1872100830078,82.2185516357422,82.2479476928711,82.2830581665039,82.2916030883789,82.3663101196289,82.4052200317383,82.4494094848633,82.4373474121094,82.3983383178711,82.3739242553711,82.3506851196289,82.3264617919922,82.1872100830078,82.1417007446289,82.0949172973633,82.0773010253906,82.0660171508789,82.0924835205078,82.1583023071289,82.1964340209961,82.2287063598633,82.2457504272461,82.2472686767578,82.2184600830078,82.1205596923828,82.0045928955078,81.9776382446289,81.9362335205078,81.8858871459961,81.8511199951172,81.8544387817383,81.9323272705078,82.0283660888672,82.1923828125,82.2782669067383,82.3363265991211,82.39501953125,82.4271011352539,82.4646530151367,82.494384765625,82.5062561035156,82.5186538696289,82.563232421875,82.6017608642578,82.628662109375,82.6492156982422,82.6430130004883,82.594482421875,82.571533203125,82.5863800048828,82.715576171875,82.7446746826172,82.76171875,82.779052734375,82.7690963745117,82.7061996459961,82.6746597290039,82.6793899536133,82.6939010620117,82.7327651977539,82.7849578857422,82.8164978027344,82.8589859008789,82.8942337036133,82.9111328125,82.9385223388672,82.9332046508789,82.89111328125,82.9063491821289,82.8958511352539,82.8831558227539,82.8372116088867,82.6708984375,82.6444320678711,82.6041030883789,82.5498504638672,82.53515625,82.5724105834961,82.6085433959961,82.6435089111328,82.7236328125,82.7579116821289,82.8158187866211,82.91943359375,82.967529296875,83.008544921875,83.0471725463867,83.0131378173828,82.989013671875,82.9553756713867,82.9041976928711,82.8518524169922,82.7715759277344,82.7216262817383,82.7556610107422,82.84423828125,82.9048385620117,82.9458541870117,82.9771499633789,82.998779296875,83.0811996459961,83.1060562133789,83.1014175415039,82.974853515625,82.9230422973633,82.9022445678711,82.9112777709961,82.9695816040039,83.0012741088867,83.0211410522461,83.0826644897461,83.09814453125,83.1161117553711,83.109619140625,83.092529296875,83.02490234375,83.0167999267578],[47.5242156982422,47.5111351013184,47.467041015625,47.3917961120605,47.270751953125,47.1726570129395,47.0975074768066,47.062255859375,47.062744140625,47.057373046875,47.0574684143066,47.037109375,47.0073738098145,46.9759750366211,46.9376945495605,46.8853530883789,46.8515129089355,46.8623542785645,46.9847640991211,46.9644050598145,46.8994140625,46.8649406433105,46.73486328125,46.6650390625,46.6188468933105,46.5828590393066,46.550048828125,46.5470695495605,46.5648460388184,46.62109375,46.6143531799316,46.5999031066895,46.5467758178711,46.4832038879395,46.4478988647461,46.4207534790039,46.3628425598145,46.2879867553711,46.2535133361816,46.2382316589355,46.2279777526855,46.2380867004395,46.3276824951172,46.36181640625,46.3677749633789,46.3460426330566,46.2958984375,46.2960929870605,46.3061981201172,46.3487777709961,46.4308090209961,46.4823226928711,46.4806632995605,46.4955558776855,46.4751968383789,46.3912582397461,46.2867660522461,46.21923828125,46.1024398803711,46.0514640808105,46.014892578125,45.9831047058105,45.9281234741211,45.8755874633789,45.845703125,45.8300285339355,45.8619651794434,45.9186973571777,45.9961929321289,46.06103515625,46.0771484375,46.1107902526855,46.1598167419434,46.2458992004395,46.2828598022461,46.4027824401855,46.431884765625,46.446044921875,46.4451179504395,46.4034156799316,46.3412132263184,46.2710456848145,46.2560043334961,46.1875953674316,46.1609382629395,46.0519065856934,46.0159149169922,45.9474601745605,45.9218292236328,45.9722175598145,45.9781723022461,45.9444313049316,45.912353515625,45.8804206848145,45.90380859375,45.92578125,45.9588356018066,46.0171394348145,46.0517578125,46.0894050598145,46.1306648254395,46.1651344299316,46.2751922607422,46.31396484375,46.3691902160645,46.4066429138184,46.415771484375,46.4373512268066,46.4305191040039,46.3936996459961,46.3326148986816,46.3194351196289,46.3084487915039,46.2522430419922,46.1930694580078,46.1470222473145,46.142333984375,46.1512184143066,46.2146949768066,46.2380867004395,46.2793960571289,46.337646484375,46.3786125183105,46.4281730651855,46.4585418701172,46.5160636901855,46.5669937133789,46.6110343933105,46.6830558776855,46.7554206848145,46.832275390625,46.9258804321289,46.9483413696289,47.0043487548828,47.0265159606934,47.0582504272461,47.1631851196289,47.2671852111816,47.3020515441895,47.322509765625,47.3394546508789,47.3525390625,47.3612327575684,47.3942375183105,47.4532241821289,47.4893569946289,47.4898452758789,47.4732437133789,47.4537124633789,47.4327163696289,47.42578125,47.43310546875,47.4551734924316,47.5076637268066,47.54736328125,47.5927238464355,47.5698738098145,47.5638656616211,47.576171875,47.60693359375,47.60693359375,47.589599609375,47.5921401977539,47.59619140625,47.6126937866211,47.6240234375,47.6377944946289,47.6519050598145,47.6591300964355,47.6518058776855,47.6626968383789,47.6877937316895,47.7313461303711,47.7668952941895,47.775634765625,47.76611328125,47.7000503540039,47.6980438232422,47.7099113464355,47.716552734375,47.70361328125,47.6626968383789,47.6563949584961,47.6707038879395,47.6707038879395,47.5989265441895,47.5242156982422],[-55.8896484375,-55.8917007446289,-55.8731422424316,-55.8648452758789,-55.8572273254395,-55.8425788879395,-55.8275375366211,-55.8161163330078,-55.8259773254395,-55.8328094482422,-55.8443374633789,-55.87744140625,-55.8896484375],[-55.77685546875,-55.7847633361816,-55.7660140991211,-55.7527351379395,-55.7248039245605,-55.7078132629395,-55.6836891174316,-55.6620101928711,-55.640625,-55.5851554870605,-55.58935546875,-55.6121101379395,-55.6886711120605,-55.7437477111816,-55.7627944946289,-55.77685546875],[-55.2290992736816,-55.2728500366211,-55.2699203491211,-55.2541046142578,-55.234375,-55.2130889892578,-55.1936531066895,-55.16943359375,-55.16552734375,-55.1897468566895,-55.2290992736816],[-55.15380859375,-55.1920928955078,-55.2425765991211,-55.2818374633789,-55.2925758361816,-55.2722625732422,-55.2365226745605,-55.201171875,-55.1817359924316,-55.1774406433105,-55.1785163879395,-55.1919937133789,-55.2429695129395,-55.2564468383789,-55.2595710754395,-55.2210922241211,-55.2068328857422,-55.1726570129395,-55.0712890625,-54.9806632995605,-54.9293937683105,-54.9296875,-54.9689445495605,-54.9776382446289,-55.0635757446289,-55.115234375,-55.15380859375],[-54.9190444946289,-55.0177764892578,-54.9579124450684,-54.9596672058105,-55.0419921875,-55.1283226013184,-55.1555671691895,-55.177734375,-55.1916046142578,-55.2194328308105,-55.2551765441895,-55.3082046508789,-55.3327140808105,-55.3566398620605,-55.4369125366211,-55.4783210754395,-55.5179710388184,-55.5875015258789,-55.6431655883789,-55.6505851745605,-55.6336898803711,-55.6015625,-55.5213851928711,-55.5052719116211,-55.4890594482422,-55.4500007629395,-55.4522476196289,-55.4356460571289,-55.4501953125,-55.4238243103027,-55.37060546875,-55.3473625183105,-55.2632827758789,-55.2412109375,-55.2385749816895,-55.2557640075684,-55.2443351745605,-55.1833992004395,-55.171875,-55.1658210754395,-55.27392578125,-55.3006858825684,-55.3693351745605,-55.4748039245605,-55.4767570495605,-55.4442367553711,-55.4240226745605,-55.3708992004395,-55.3399429321289,-55.3208999633789,-55.2984390258789,-55.22900390625,-55.2189445495605,-55.2365226745605,-55.2198257446289,-55.1906280517578,-55.1741218566895,-55.1474609375,-55.1307640075684,-55.1110343933105,-55.0611343383789,-54.8820304870605,-54.9190444946289],[-54.8679695129395,-54.9313507080078,-54.9429702758789,-54.9676742553711,-54.9527359008789,-54.9456024169922,-54.9212875366211,-54.9088897705078,-55.06591796875,-55.11376953125,-55.1656227111816,-55.1770515441895,-55.1613273620605,-55.1349601745605,-55.1119155883789,-55.08203125,-55.0848655700684,-55.10693359375,-55.1042022705078,-55.0798835754395,-55.0619163513184,-55.0396499633789,-55.0068359375,-54.9904289245605,-54.9377937316895,-54.8929710388184,-54.8869171142578,-54.8922843933105,-54.9137687683105,-54.9342765808105,-54.9308624267578,-54.9137687683105,-54.8892593383789,-54.83935546875,-54.8345718383789,-54.8444366455078,-54.8674812316895,-54.8679695129395],[-54.0328140258789,-54.1125984191895,-54.11181640625,-54.1617202758789,-54.2466812133789,-54.2811508178711,-54.3163070678711,-54.3663063049316,-54.3740234375,-54.3135757446289,-54.2311515808105,-54.24560546875,-54.2253913879395,-54.2297859191895,-54.2764625549316,-54.3008766174316,-54.2072257995605,-54.1187515258789,-54.0477561950684,-53.9388694763184,-53.9212913513184,-53.8848609924316,-53.92333984375,-53.9560546875,-54.0328140258789],[-53.4816398620605,-53.5627937316895,-53.5783195495605,-53.5653305053711,-53.5579109191895,-53.5880851745605,-53.5988273620605,-53.6875,-53.7253913879395,-53.8074226379395,-53.8621101379395,-53.9408226013184,-53.9807624816895,-54.0038108825684,-54.0277366638184,-54.07373046875,-54.0789070129395,-54.1031227111816,-54.1250953674316,-54.1265602111816,-54.1146507263184,-54.0920906066895,-54.06591796875,-54.0415992736816,-53.9547843933105,-53.8648414611816,-53.8485336303711,-53.86083984375,-53.8605461120605,-53.8328132629395,-53.8752937316895,-53.9159202575684,-53.97802734375,-53.998046875,-54.0093765258789,-53.98583984375,-53.9439468383789,-53.9196319580078,-53.8358421325684,-53.7920913696289,-53.7291984558105,-53.72265625,-53.7240257263184,-53.7361335754395,-53.6554679870605,-53.5703125,-53.5457992553711,-53.4268569946289,-53.4100570678711,-53.47021484375,-53.5119132995605,-53.4480476379395,-53.4251976013184,-53.3967781066895,-53.3944320678711,-53.41455078125,-53.4230461120605,-53.4424819946289,-53.4816398620605],[-52.92236328125,-52.9314460754395,-52.9292984008789,-52.9455108642578,-52.96533203125,-53.01220703125,-53.0560569763184,-53.06982421875,-53.1256828308105,-53.1400413513184,-53.1443367004395,-53.2476577758789,-53.2594718933105,-53.3409194946289,-53.3539047241211,-53.3583984375,-53.3205070495605,-53.3184585571289,-53.3068351745605,-53.3001937866211,-53.2529296875,-53.2296905517578,-53.1207046508789,-53.0968780517578,-53.0757827758789,-53.0905265808105,-53.0764656066895,-53.08154296875,-52.9949226379395,-52.921875,-52.8347625732422,-52.7681617736816,-52.7487297058105,-52.73388671875,-52.7712898254395,-52.8356437683105,-52.8600616455078,-52.92236328125],[-52.6526374816895,-52.9495124816895,-53.2418937683105,-53.5154304504395,-53.7888679504395,-54.0529289245605,-54.3240203857422,-54.6278343200684,-54.8536148071289,-54.8536148071289,-54.8767585754395,-54.9098663330078,-54.85888671875,-54.8127937316895,-54.7121086120605,-54.7391624450684,-54.7818374633789,-54.8155250549316,-54.8192367553711,-54.7775382995605,-54.75634765625,-54.7517585754395,-54.8095703125,-54.7505836486816,-54.71435546875,-54.6941413879395,-54.61962890625,-54.6261711120605,-54.6015625,-54.5287094116211,-54.4948234558105,-54.4744110107422,-54.4339866638184,-54.4436531066895,-54.4971656799316,-54.4953117370605,-54.4449234008789,-54.4001960754395,-54.3954124450684,-54.4505844116211,-54.4442367553711,-54.4195327758789,-54.3980484008789,-54.3600578308105,-54.337890625,-54.3272476196289,-54.3488311767578,-54.4147491455078,-54.4854507446289,-54.50439453125,-54.5022468566895,-54.5285148620605,-54.4855461120605,-54.3732414245605,-54.3034172058105,-54.2623062133789,-54.2413101196289,-54.1104507446289,-53.9958000183105,-53.8841781616211,-53.8228492736816,-53.7274436950684,-53.6550750732422,-53.6273422241211,-53.8934555053711,-53.9867172241211,-54.0056648254395,-54.0055694580078,-54.1361351013184,-54.1806640625,-54.2774391174316,-54.34765625,-54.3792953491211,-54.38134765625,-54.3674812316895,-54.32080078125,-54.3058624267578,-54.3640632629395,-54.4071273803711,-54.4375991821289,-54.4881820678711,-54.5426788330078,-54.5714836120605,-54.5574188232422,-54.4833030700684,-54.4576187133789,-54.4450187683105,-54.4284172058105,-54.40673828125,-54.3543968200684,-54.1090812683105,-54.0111312866211,-53.8880844116211,-53.7611312866211,-53.7217788696289,-53.6715812683105,-53.6008796691895,-53.4994125366211,-53.47998046875,-53.4163055419922,-53.3734359741211,-53.3419914245605,-53.3340835571289,-53.3372077941895,-53.3504867553711,-53.4181632995605,-53.4139633178711,-53.3776397705078,-53.3047828674316,-53.2062530517578,-53.1433601379395,-53.0855484008789,-53.0264625549316,-53.0006828308105,-53.0041007995605,-52.990234375,-52.9699211120605,-52.9427719116211,-52.9193344116211,-52.8990211486816,-52.8572273254395,-52.8169937133789,-52.751953125,-52.7337913513184,-52.7236328125,-52.7685546875,-52.8212890625,-52.8210945129395,-52.7990226745605,-52.7313499450684,-52.6462898254395,-52.54931640625,-52.4914054870605,-52.4862289428711,-52.6675796508789,-52.67431640625,-52.5767555236816,-52.58203125,-52.6315422058105,-52.6526374816895],[-51.9310569763184,-51.9420890808105,-51.9187507629395,-51.89794921875,-51.8451194763184,-51.7906265258789,-51.7256851196289,-51.7249031066895,-51.7506828308105,-51.7702140808105,-51.8116226196289,-51.8708992004395,-51.8884773254395,-51.9111328125,-51.9310569763184],[-51.6301765441895,-51.82470703125,-51.8518562316895,-51.8662109375,-51.9651374816895,-51.9919891357422,-52.1600570678711,-52.2792015075684,-52.2707023620605,-52.1522483825684,-52.0378913879395,-51.9039039611816,-51.7888679504395,-51.7237319946289,-51.73828125,-51.6500015258789,-51.6301765441895],[-51.2770500183105,-51.36083984375,-51.3874969482422,-51.3957023620605,-51.3702163696289,-51.3673820495605,-51.4117202758789,-51.4341812133789,-51.4283180236816,-51.3983421325684,-51.5243186950684,-51.5666999816895,-51.6253890991211,-51.5564460754395,-51.4535140991211,-51.38330078125,-51.27880859375,-51.3181648254395,-51.2794914245605,-51.2076187133789,-51.2071304321289,-51.2454109191895,-51.2770500183105],[-50.6799812316895,-50.7723655700684,-50.7643547058105,-50.7411117553711,-50.6825218200684,-50.6541976928711,-50.5955085754395,-50.5303726196289,-50.4805641174316,-50.4839859008789,-50.4967765808105,-50.5104522705078,-50.5543975830078,-50.5806655883789,-50.5968742370605,-50.6799812316895],[-50.2960929870605,-50.3762702941895,-50.3430671691895,-50.3433609008789,-50.2566413879395,-50.1926765441895,-50.16796875,-50.1126937866211,-50.0118179321289,-50.04541015625,-50.0552749633789,-50.0886726379395,-50.1099586486816,-50.197265625,-50.2373046875,-50.2960929870605],[-48.8365211486816,-48.916015625,-49.0689468383789,-49.1591796875,-49.2306632995605,-49.22998046875,-49.1954116821289,-49.1388664245605,-49.0824241638184,-49.0095710754395,-48.9884757995605,-48.9807624816895,-48.9425773620605,-48.8859367370605,-48.85888671875,-48.8381805419922,-48.8504905700684,-48.8106460571289,-48.7786140441895,-48.77294921875,-48.8365211486816],[-49.1478538513184,-49.2945289611816,-49.44189453125,-49.6229515075684,-49.6595687866211,-49.6685562133789,-49.6911163330078,-49.7862281799316,-49.8594741821289,-49.9191398620605,-49.99072265625,-50.0066375732422,-50.0192375183105,-50.0114288330078,-49.9296875,-49.8794937133789,-49.8138656616211,-49.7258758544922,-49.6921882629395,-49.6341781616211,-49.6052742004395,-49.5160179138184,-49.4892578125,-49.4523429870605,-49.43701171875,-49.423828125,-49.4224586486816,-49.5330047607422,-49.5641593933105,-49.6056632995605,-49.7517585754395,-49.8362312316895,-49.8523445129395,-49.85595703125,-49.8474617004395,-49.7699203491211,-49.7913055419922,-49.6970710754395,-49.6216812133789,-49.6282234191895,-49.4940452575684,-49.4624977111816,-49.4084014892578,-49.35888671875,-49.3220710754395,-49.2686500549316,-49.2628898620605,-49.2927703857422,-49.2702140808105,-49.1853523254395,-49.1480484008789,-49.0835952758789,-49.0220718383789,-48.9601554870605,-48.8894538879395,-48.81884765625,-48.7913093566895,-48.7332000732422,-48.705078125,-48.7088851928711,-48.7499008178711,-48.7547836303711,-48.7668952941895,-48.8125991821289,-49.1478538513184],[-48.7634773254395,-48.7646484375,-48.5863304138184,-48.3288116455078,-48.2306632995605,-48.15673828125,-48.0958976745605,-48.0708999633789,-48.0197257995605,-48.0740242004395,-48.2184562683105,-48.4251937866211,-48.6226539611816,-48.67138671875,-48.72119140625,-48.7634773254395],[-48.5919914245605,-48.61572265625,-48.6011695861816,-48.62646484375,-48.5357398986816,-48.3914070129395,-48.3615226745605,-48.279296875,-48.2252922058105,-48.1417007446289,-48.0534172058105,-48.0267601013184,-47.974609375,-47.9228515625,-47.83935546875,-47.8503875732422,-48.0207977294922,-48.0782241821289,-48.1258773803711,-48.1455078125,-48.2058601379395,-48.29931640625,-48.3430671691895,-48.3703155517578,-48.39306640625,-48.4251937866211,-48.5919914245605],[-47.8376960754395,-47.8506813049316,-47.8180656433105,-47.7638664245605,-47.72802734375,-47.6906242370605,-47.6947250366211,-47.72314453125,-47.7566413879395,-47.8004875183105,-47.8245086669922,-47.8376960754395],[-45.6915016174316,-45.73583984375,-45.7572250366211,-45.7224617004395,-45.7385749816895,-45.66259765625,-45.5999984741211,-45.5255889892578,-45.4258766174316,-45.28515625,-45.2529296875,-45.2032241821289,-45.1726531982422,-45.2772445678711,-45.4640617370605,-45.5745124816895,-45.611328125,-45.6536140441895,-45.6915016174316],[-44.89013671875,-44.9065437316895,-44.9017601013184,-44.8699188232422,-44.8156242370605,-44.7951164245605,-44.7951164245605,-44.8239250183105,-44.8705062866211,-44.89013671875],[-44.8214836120605,-44.8329124450684,-44.8311538696289,-44.796875,-44.7516593933105,-44.68408203125,-44.6521492004395,-44.6139640808105,-44.5591812133789,-44.5442390441895,-44.5462875366211,-44.61083984375,-44.6807632446289,-44.7529296875,-44.8214836120605],[-44.7800788879395,-44.8599624633789,-44.8332061767578,-44.7743148803711,-44.7248077392578,-44.6410179138184,-44.5960960388184,-44.5313491821289,-44.4895515441895,-44.4402313232422,-44.3941383361816,-44.3502922058105,-44.3349609375,-44.3840827941895,-44.4577140808105,-44.5085945129395,-44.5490226745605,-44.6384773254395,-44.7120132446289,-44.7800788879395],[-44.39453125,-44.4375,-44.4451179504395,-44.4948234558105,-44.5379867553711,-44.5909156799316,-44.6546859741211,-44.7288093566895,-44.83984375,-44.94580078125,-44.9786109924316,-45.0335922241211,-45.1190452575684,-45.1576194763184,-45.1958961486816,-45.2667999267578,-45.2765617370605,-45.2834968566895,-45.3265609741211,-45.3406257629395,-45.3449211120605,-45.3253898620605,-45.1957015991211,-45.1448249816895,-45.0589828491211,-44.9108390808105,-44.865234375,-44.7481460571289,-44.64794921875,-44.5845718383789,-44.4735336303711,-44.4354476928711,-44.3957023620605,-44.4269523620605,-44.4159164428711,-44.3893547058105,-44.27587890625,-44.1864280700684,-44.1402359008789,-44.1348609924316,-44.1853523254395,-44.2349624633789,-44.2741241455078,-44.3253898620605,-44.39453125],[-43.8272476196289,-43.87646484375,-43.8832054138184,-43.8753890991211,-43.9142570495605,-43.9219703674316,-43.8875007629395,-43.8721656799316,-43.8209953308105,-43.8165054321289,-43.7837905883789,-43.7889633178711,-43.8272476196289],[-43.6078147888184,-43.6253929138184,-43.5955085754395,-43.5703125,-43.5494117736816,-43.5359382629395,-43.5530281066895,-43.5774383544922,-43.599609375,-43.6078147888184],[-43.3458976745605,-43.3667984008789,-43.3719749450684,-43.3566398620605,-43.35791015625,-43.31884765625,-43.2635765075684,-43.2316398620605,-43.1857414245605,-43.0794906616211,-42.8787117004395,-42.5905265808105,-42.4813499450684,-42.43603515625,-42.3815422058105,-42.3254852294922,-42.2689476013184,-42.2164039611816,-42.1058578491211,-42.0562515258789,-42.0023422241211,-41.8909187316895,-41.85400390625,-41.82275390625,-41.7955093383789,-41.8772468566895,-41.8962898254395,-41.9808578491211,-42.0471687316895,-42.1659164428711,-42.19287109375,-42.27783203125,-42.314453125,-42.392578125,-42.46630859375,-42.4925765991211,-42.5082015991211,-42.5287132263184,-42.5447273254395,-42.5857429504395,-42.6218757629395,-42.7043952941895,-42.7616233825684,-42.84716796875,-42.9365234375,-42.9932594299316,-43.07373046875,-43.1271514892578,-43.1590805053711,-43.2914047241211,-43.3458976745605],[-33.646484375,-33.6677742004395,-33.6617202758789,-33.6441421508789,-33.6135749816895,-33.5763664245605,-33.5751953125,-33.578125,-33.5850563049316,-33.6101570129395,-33.6273422241211,-33.6416015625,-33.646484375],[-27.1404285430908,-27.1712894439697,-27.1162109375,-27.068359375,-27.0958976745605,-27.10107421875,-27.1404285430908],[-22.8216800689697,-23.0013675689697,-23.2451171875,-23.6339836120605,-23.9346675872803,-23.9748039245605,-24.0337886810303,-24.1189441680908,-24.2433586120605,-24.3083000183105,-24.3919925689697,-24.4603519439697,-24.4972648620605,-24.5451164245605,-24.5969734191895,-24.6297855377197,-24.7473640441895,-24.8376960754395,-24.8992195129395,-24.9251937866211,-24.9989261627197,-25.0509777069092,-25.0918960571289,-25.1247081756592,-25.1493167877197,-25.1629886627197,-25.2367191314697,-25.4200191497803,-25.4856452941895,-25.6515617370605,-25.7410163879395,-26.0654296875,-26.1537113189697,-26.2769527435303,-26.3519535064697,-26.4180660247803,-26.4704093933105,-26.5183601379395,-26.6703128814697,-26.8064460754395,-26.8775386810303,-26.9732418060303,-27.0279293060303,-27.0481452941895,-27.0853500366211,-27.1400375366211,-27.1483402252197,-27.1044921875,-27.1154289245605,-27.1537113189697,-27.2466793060303,-27.4051761627197,-27.4490242004395,-27.5700187683105,-27.7435550689697,-27.84814453125,-27.9247074127197,-27.9736328125,-28.07080078125,-28.1653308868408,-28.1926765441895,-28.2008800506592,-28.28564453125,-28.41357421875,-28.56201171875,-28.64111328125,-28.7838859558105,-29.0455093383789,-29.1032238006592,-29.1488304138184,-29.2500019073486,-29.3240242004395,-29.54541015625,-29.7691402435303,-29.8740253448486,-30.0164070129395,-30.0783195495605,-30.1039047241211,-30.1203117370605,-30.1750011444092,-30.2132797241211,-30.2816410064697,-30.3582038879395,-30.3882808685303,-30.3609390258789,-30.3855476379395,-30.4402332305908,-30.5046863555908,-30.6772441864014,-30.833984375,-30.90234375,-30.9597644805908,-30.9925785064697,-31.0226535797119,-31.0604496002197,-31.12109375,-31.1292972564697,-31.11279296875,-31.1484394073486,-31.2228507995605,-31.3173847198486,-31.4279308319092,-31.5694351196289,-31.6664066314697,-31.8418941497803,-31.8837909698486,-31.8810558319092,-31.9165992736816,-31.9577140808105,-32.0310516357422,-32.0423851013184,-32.08349609375,-32.1764640808105,-32.2666969299316,-32.3099594116211,-32.4306640625,-32.4716796875,-32.6260757446289,-32.8074188232422,-32.8599624633789,-32.8845710754395,-32.9636726379395,-33.0267562866211,-33.1279296875,-33.2017593383789,-33.271484375,-33.2793922424316,-33.2509765625,-33.2837867736816,-33.3439445495605,-33.3986320495605,-33.4697265625,-33.6009750366211,-33.7313461303711,-33.9297828674316,-34.0835914611816,-34.1804695129395,-34.2243156433105,-34.25439453125,-34.2699203491211,-34.2762680053711,-34.30078125,-34.3500022888184,-34.4320335388184,-34.4928703308105,-34.5812530517578,-34.6726570129395,-34.7328147888184,-34.7745132446289,-34.85498046875,-34.9217758178711,-35.1468772888184,-35.1936531066895,-35.216796875,-35.2468719482422,-35.3079109191895,-35.326171875,-35.3753890991211,-35.4519538879395,-35.5230445861816,-35.6091804504395,-35.7718772888184,-35.8785171508789,-35.9705085754395,-36.0617179870605,-36.1327133178711,-36.1463851928711,-36.2119140625,-36.283203125,-36.3406257629395,-36.392578125,-36.4117164611816,-36.4117164611816,-36.419921875,-36.4873046875,-36.5237312316895,-36.5780258178711,-36.64404296875,-36.68505859375,-36.7616233825684,-36.8436508178711,-36.9202156066895,-37.0569305419922,-37.1143569946289,-37.2274398803711,-37.30029296875,-37.3932609558105,-37.4451141357422,-37.5591812133789,-37.6310539245605,-37.7623062133789,-37.9099617004395,-38.0412101745605,-38.1939468383789,-38.3148460388184,-38.4458999633789,-38.4978523254395,-38.5416030883789,-38.6044921875,-38.6810569763184,-38.7384757995605,-38.7575187683105,-38.8093757629395,-38.8454093933105,-38.8888664245605,-38.93505859375,-38.9856452941895,-39.2059593200684,-39.2872085571289,-39.40234375,-39.4952125549316,-39.5231475830078,-39.5641593933105,-39.6024398803711,-39.6111335754395,-39.59423828125,-39.6051750183105,-39.63525390625,-39.70703125,-39.8333015441895,-39.8868141174316,-39.92919921875,-40.0208015441895,-40.0949211120605,-40.0946273803711,-40.1247062683105,-40.1766624450684,-40.2443351745605,-40.2997093200684,-40.3352546691895,-40.3817367553711,-40.40087890625,-40.4391593933105,-40.5244140625,-40.62060546875,-40.6916999816895,-40.7891578674316,-40.8929672241211,-40.99462890625,-41.2923851013184,-41.3933601379395,-41.560546875,-41.6066398620605,-41.650390625,-41.77197265625,-41.9685554504395,-42.0467758178711,-42.1014633178711,-42.1478500366211,-42.1670875549316,-42.13427734375,-42.14794921875,-42.2053718566895,-42.2518539428711,-42.29833984375,-42.3584976196289,-42.4732398986816,-42.5224609375,-42.5771484375,-42.6482391357422,-42.7767562866211,-42.9900398254395,-43.0656242370605,-43.1019515991211,-43.1453094482422,-43.1667938232422,-43.2373046875,-43.2946281433105,-43.3229522705078,-43.3475608825684,-43.4401359558105,-43.5271492004395,-43.5901374816895,-43.6467781066895,-43.7046852111816,-43.7532234191895,-43.8583984375,-43.9295921325684,-43.9844703674316,-44.0666999816895,-44.1060562133789,-44.1507797241211,-44.2414054870605,-44.3301734924316,-44.3831062316895,-44.4249038696289,-44.4412117004395,-44.4940452575684,-44.5602531433105,-44.6307601928711,-44.7630844116211,-44.78515625,-44.7498054504395,-44.7620086669922,-44.7704124450684,-44.7744140625,-44.79150390625,-44.7718772888184,-44.8204116821289,-44.9041976928711,-44.9306640625,-44.9791984558105,-45.06787109375,-45.1682624816895,-45.23046875,-45.3319358825684,-45.4376983642578,-45.5126953125,-45.5347633361816,-45.5789070129395,-45.7244148254395,-45.8390617370605,-45.8787117004395,-45.9537124633789,-46.0418968200684,-46.1027336120605,-46.1605453491211,-46.2067375183105,-46.2799835205078,-46.31982421875,-46.4278335571289,-46.5784187316895,-46.6513671875,-46.7058563232422,-46.7915992736816,-46.8312492370605,-46.9368171691895,-47.0160179138184,-47.0874977111816,-47.1443367004395,-47.20166015625,-47.2138671875,-47.2414054870605,-47.3427734375,-47.4462890625,-47.49267578125,-47.5720710754395,-47.685546875,-47.7841796875,-47.8763694763184,-47.9733390808105,-48.0159187316895,-48.1100578308105,-48.2291030883789,-48.3658180236816,-48.4173851013184,-48.4753875732422,-48.5193328857422,-48.6624984741211,-48.7296867370605,-48.7928695678711,-48.8415985107422,-48.8962898254395,-48.9439430236816,-48.9767608642578,-49.0143547058105,-49.0968742370605,-49.18798828125,-49.3006858825684,-49.3138656616211,-49.3976554870605,-49.4638671875,-49.5829124450684,-49.6980476379395,-49.7945327758789,-49.9109382629395,-50.0302734375,-50.1252937316895,-50.2311515808105,-50.3619155883789,-50.4725570678711,-50.5584945678711,-50.6107406616211,-50.6700172424316,-50.73828125,-50.7603492736816,-50.6964836120605,-50.6531219482422,-50.6376953125,-50.6476554870605,-50.6075172424316,-50.6117172241211,-50.63427734375,-50.6818389892578,-50.78955078125,-50.9102516174316,-51.0333976745605,-51.0601539611816,-51.0954093933105,-51.17041015625,-51.2233390808105,-51.2989234924316,-51.4703102111816,-51.5408210754395,-51.6203117370605,-51.6911125183105,-51.7440452575684,-51.8186492919922,-51.88037109375,-51.9641609191895,-51.9895515441895,-51.9913063049316,-51.9939460754395,-51.9981460571289,-52.0022468566895,-52.0082015991211,-52.0753898620605,-52.1361312866211,-52.1361312866211,-52.2081031799316,-52.2554702758789,-52.2733383178711,-52.2904281616211,-52.3566398620605,-52.2626953125,-52.21142578125,-52.2054672241211,-52.2694320678711,-52.4215812683105,-52.4647445678711,-52.5055656433105,-52.5135765075684,-52.6608390808105,-52.6734390258789,-52.7125015258789,-52.7687492370605,-52.8895492553711,-52.9630851745605,-53.2269554138184,-53.3736305236816,-53.4483413696289,-53.5704116821289,-53.779296875,-53.8249969482422,-53.8833961486816,-53.8409194946289,-53.8031234741211,-53.72265625,-53.6658210754395,-53.63232421875,-53.47119140625,-53.4177703857422,-53.3501930236816,-53.2537117004395,-53.2466812133789,-53.2496109008789,-53.2340812683105,-53.2857398986816,-53.3983421325684,-53.4583969116211,-53.4955062866211,-53.5235328674316,-53.4845733642578,-53.2326164245605,-53.1070327758789,-53.03369140625,-52.9205055236816,-52.8880844116211,-52.8456039428711,-52.8106422424316,-52.7642555236816,-53.0017585754395,-53.0643539428711,-53.13232421875,-53.2544937133789,-53.2906265258789,-53.3716773986816,-53.4607391357422,-53.4200210571289,-53.2907257080078,-53.2434539794922,-53.17724609375,-53.1219749450684,-52.9365234375,-52.87158203125,-52.8195304870605,-52.7623062133789,-52.7490234375,-52.7738304138184,-52.8175811767578,-52.814453125,-52.6500015258789,-52.6460952758789,-52.6828117370605,-52.6607398986816,-52.6439476013184,-52.6053733825684,-52.56005859375,-52.5370140075684,-52.52099609375,-52.53857421875,-52.6257820129395,-52.60400390625,-52.56005859375,-52.5291023254395,-52.5355491638184,-52.5774421691895,-52.642578125,-52.71240234375,-52.7816390991211,-52.8917961120605,-52.9774398803711,-53.0220680236816,-53.0456047058105,-53.0739250183105,-53.0546875,-52.96484375,-52.9035186767578,-52.8370132446289,-52.7542991638184,-52.7071304321289,-52.6019554138184,-52.5350608825684,-52.4879875183105,-52.4878883361816,-52.5626945495605,-52.6240234375,-52.5951156616211,-52.6857452392578,-52.6615219116211,-52.6881866455078,-52.6393547058105,-52.5772476196289,-52.5125961303711,-52.4029273986816,-52.3762702941895,-52.3825187683105,-52.3171844482422,-52.2023429870605,-52.1712913513184,-52.1178703308105,-52.1048851013184,-52.1202125549316,-52.15478515625,-52.1591796875,-52.2339859008789,-52.2160148620605,-52.1988296508789,-52.13671875,-52.0777320861816,-52.0777320861816,-52.1531257629395,-52.14599609375,-52.1659164428711,-52.1578140258789,-52.1296882629395,-52.046875,-51.9611320495605,-51.9495086669922,-51.9605484008789,-51.9851570129395,-52.0447273254395,-52.0998992919922,-52.1451148986816,-52.2000961303711,-52.2541999816895,-52.3302726745605,-52.3567390441895,-52.3846702575684,-52.37158203125,-52.333984375,-52.2823257446289,-52.2554702758789,-52.2170906066895,-52.1703147888184,-52.0370101928711,-52.0069351196289,-51.9464836120605,-51.8909187316895,-51.8475608825684,-51.763671875,-51.7061500549316,-51.5732421875,-51.43994140625,-51.4539031982422,-51.47802734375,-51.49560546875,-51.5044937133789,-51.6142578125,-51.6779289245605,-51.6950225830078,-51.7373046875,-51.7991180419922,-51.85986328125,-51.9906234741211,-52.0700187683105,-52.041015625,-51.9603538513184,-51.8562507629395,-51.7955055236816,-51.8011741638184,-51.7899398803711,-51.7578125,-51.7844734191895,-51.7121086120605,-51.6805648803711,-51.5787086486816,-51.6178703308105,-51.3314476013184,-51.2663078308105,-51.1954078674316,-51.20458984375,-51.1950187683105,-51.1624984741211,-51.1496086120605,-51.1306648254395,-51.0865211486816,-51.0628890991211,-50.8810539245605,-50.7855453491211,-50.6812515258789,-50.6789093017578,-50.6620140075684,-50.6184577941895,-50.5353507995605,-50.4699211120605,-50.4084968566895,-50.3609390258789,-50.3820304870605,-50.4878921508789,-50.5595703125,-50.7780265808105,-50.7974624633789,-50.8177757263184,-50.9400405883789,-50.9383811950684,-50.8358421325684,-50.6966781616211,-50.6506843566895,-50.6511726379395,-50.6279296875,-50.5700187683105,-50.49267578125,-50.490234375,-50.5398445129395,-50.78271484375,-50.8270492553711,-50.7170867919922,-50.6378936767578,-50.6097640991211,-50.4853515625,-50.5105476379395,-50.4698257446289,-50.3980484008789,-50.3629875183105,-50.3501930236816,-50.265625,-50.1940422058105,-50.0652351379395,-49.974609375,-50.0227546691895,-49.9947280883789,-49.9285163879395,-49.9485359191895,-49.9073257446289,-49.7833976745605,-49.7201156616211,-49.6041030883789,-49.5792999267578,-49.5553703308105,-49.5930671691895,-49.6237297058105,-49.609375,-49.5234375,-49.4909172058105,-49.4296875,-49.36181640625,-49.3056640625,-49.244140625,-49.1580085754395,-49.0909194946289,-49.0599594116211,-49.0460929870605,-49.0209007263184,-49.0261726379395,-49.1110343933105,-49.1883773803711,-49.25048828125,-49.3205108642578,-49.4043922424316,-49.5005874633789,-49.4639625549316,-49.42626953125,-49.4004859924316,-49.3513679504395,-49.0478515625,-48.7936515808105,-48.595703125,-48.5169906616211,-48.494140625,-48.5041999816895,-48.5036125183105,-48.4749984741211,-48.4274406433105,-48.45458984375,-48.4925765991211,-48.4639663696289,-48.3623046875,-48.2744140625,-48.1619148254395,-47.9990234375,-48.0130844116211,-48.044921875,-48.0421867370605,-48.1982421875,-48.1773452758789,-48.1458969116211,-48.1066398620605,-48.0193367004395,-47.9939460754395,-47.9415054321289,-47.88037109375,-47.6554679870605,-47.6613273620605,-47.7384757995605,-47.8669929504395,-47.9293937683105,-47.9546852111816,-47.9689445495605,-47.9443359375,-47.8912086486816,-47.8296890258789,-47.7996101379395,-47.77294921875,-47.7580070495605,-47.7022476196289,-47.6179695129395,-47.5676727294922,-47.57763671875,-47.6003913879395,-47.6666984558105,-47.6792984008789,-47.6266593933105,-47.5908203125,-47.568359375,-47.5596694946289,-47.5314483642578,-47.4304695129395,-47.3275375366211,-47.2095718383789,-47.1825180053711,-47.0831069946289,-46.9744148254395,-46.884765625,-46.7881813049316,-46.7667961120605,-46.7950210571289,-46.8345680236816,-46.8643569946289,-46.8858413696289,-46.8851585388184,-46.8639678955078,-46.7997093200684,-46.7411117553711,-46.6953125,-46.6280288696289,-46.5121078491211,-46.5105438232422,-46.60009765625,-46.6470718383789,-46.6623992919922,-46.69873046875,-46.7287101745605,-46.7463836669922,-46.7507820129395,-46.8626976013184,-46.9056663513184,-46.9345703125,-46.9401359558105,-46.86279296875,-46.7750015258789,-46.7052726745605,-46.6103515625,-46.4830055236816,-46.4291038513184,-46.3693351745605,-46.2345733642578,-46.15966796875,-46.09765625,-46.0044937133789,-45.8749008178711,-45.8236351013184,-45.8447265625,-45.8407211303711,-45.8096694946289,-45.8030281066895,-45.7671890258789,-45.716796875,-45.6783218383789,-45.6447257995605,-45.6034164428711,-45.4961891174316,-45.4603538513184,-45.4176750183105,-45.4043960571289,-45.40771484375,-45.4468727111816,-45.50244140625,-45.5693359375,-45.8352546691895,-45.8953132629395,-45.9473609924316,-46.0558586120605,-46.1318359375,-46.2126960754395,-46.2173843383789,-46.2462882995605,-46.2394523620605,-46.2223625183105,-46.1541023254395,-46.0499038696289,-45.8468780517578,-45.8181648254395,-45.8117179870605,-45.8595695495605,-45.9666976928711,-46.0703125,-46.3773422241211,-46.5006828308105,-46.5331039428711,-46.5715827941895,-46.5660171508789,-46.4998054504395,-46.4152336120605,-46.2974624633789,-46.2121124267578,-46.1592788696289,-45.9865226745605,-45.8991203308105,-45.77685546875,-45.7307624816895,-45.7028312683105,-45.6279296875,-45.47998046875,-45.4837913513184,-45.3828125,-45.34619140625,-45.3538093566895,-45.451171875,-45.4523429870605,-45.4172859191895,-45.392578125,-45.3597640991211,-45.2551765441895,-45.2381820678711,-45.1023445129395,-44.9782218933105,-44.9610366821289,-44.9202156066895,-44.7341804504395,-44.5939483642578,-44.4364242553711,-44.3954086303711,-44.2923851013184,-44.2374992370605,-44.1686515808105,-44.0657234191895,-43.89794921875,-43.8620147705078,-43.6315422058105,-43.4551773071289,-43.3236351013184,-43.2113265991211,-43.1335945129395,-43.0481452941895,-43.0394554138184,-42.9929656982422,-42.908203125,-42.8080101013184,-42.6691398620605,-42.5051765441895,-42.5166015625,-42.5096664428711,-42.4104499816895,-42.30126953125,-42.2577133178711,-42.2205085754395,-42.1998062133789,-42.2557601928711,-42.4338836669922,-42.38818359375,-42.2066421508789,-41.9808578491211,-42.0105476379395,-41.99462890625,-41.9595680236816,-41.9087867736816,-41.8467750549316,-41.8005867004395,-41.7424774169922,-41.7220687866211,-41.6491203308105,-41.4990234375,-41.5138664245605,-41.6459007263184,-41.6906242370605,-41.68408203125,-41.6593780517578,-41.5443344116211,-41.517578125,-41.5147476196289,-41.5438461303711,-41.74658203125,-41.7808570861816,-41.7970695495605,-41.7736320495605,-41.7424774169922,-41.6924819946289,-41.6392593383789,-41.6119117736816,-41.5813484191895,-41.5736351013184,-41.5174789428711,-41.4463882446289,-41.3193359375,-41.1182594299316,-40.9743156433105,-40.87158203125,-40.4684562683105,-40.2629890441895,-40.0823249816895,-39.9631843566895,-39.8542976379395,-39.7891616821289,-39.42236328125,-39.2244148254395,-38.6240234375,-38.5093765258789,-38.3667984008789,-38.1300773620605,-38.0403327941895,-37.9105453491211,-37.6985359191895,-37.5904312133789,-37.4791030883789,-37.3410148620605,-37.2554664611816,-37.1884765625,-37.2243156433105,-37.2074203491211,-37.1668930053711,-37.0535163879395,-36.8761711120605,-36.7999000549316,-36.6883773803711,-36.6434555053711,-36.5377922058105,-36.3904266357422,-35.978515625,-35.876953125,-35.7596664428711,-35.5857429504395,-35.50537109375,-35.4469718933105,-35.3408203125,-35.2404289245605,-35.09619140625,-34.9202117919922,-34.6158180236816,-34.4205055236816,-34.2884750366211,-34.1653289794922,-34.015625,-33.8895492553711,-33.8195304870605,-33.6526374816895,-33.5192375183105,-33.4290046691895,-33.2890625,-33.0951194763184,-33.0225563049316,-32.9695320129395,-32.6595687866211,-32.5378913879395,-32.3868179321289,-32.2079124450684,-31.8058586120605,-31.4963855743408,-31.1695308685303,-30.9866218566895,-30.7592792510986,-30.6280269622803,-30.3303699493408,-30.1429710388184,-29.9332008361816,-29.6497077941895,-29.4431629180908,-29.3503913879395,-29.1982421875,-28.9264659881592,-28.8552742004395,-28.7787113189697,-28.6724605560303,-28.5075187683105,-28.3778324127197,-28.0640621185303,-27.814453125,-27.7273445129395,-27.6178722381592,-27.5886726379395,-27.5051746368408,-27.3079090118408,-27.1875,-26.9505863189697,-26.8409175872803,-26.5969715118408,-26.4217777252197,-26.3293933868408,-26.2253913879395,-25.99267578125,-25.8611335754395,-25.7841796875,-25.5456066131592,-25.4874992370605,-25.37646484375,-25.2518539428711,-25.1726551055908,-24.7785167694092,-24.6443347930908,-24.3316402435303,-24.1296863555908,-23.9685535430908,-23.8857421875,-23.78173828125,-23.6555652618408,-23.56591796875,-23.5285167694092,-23.4828109741211,-23.3683586120605,-23.2554683685303,-23.17333984375,-23.0570316314697,-23.0341796875,-22.9696292877197,-22.8486328125,-22.5560550689697,-22.1931648254395,-21.974609375,-21.8666019439697,-21.6408214569092,-21.4930667877197,-21.3568344116211,-21.2532234191895,-20.7253913879395,-20.5314445495605,-20.2297859191895,-19.8050785064697,-19.7058582305908,-19.6129894256592,-19.4869136810303,-19.267578125,-18.8275394439697,-18.59521484375,-18.3980464935303,-18.3456058502197,-18.3335933685303,-18.3253898620605,-18.3251934051514,-18.2834968566895,-18.2060546875,-18.0934581756592,-17.990234375,-17.8999996185303,-17.7851543426514,-17.7038097381592,-17.6649417877197,-17.6498050689697,-17.5732421875,-17.5060558319092,-17.6195316314697,-17.7716789245605,-17.9431648254395,-17.96484375,-18.0504875183105,-18.0707035064697,-18.1027355194092,-18.14404296875,-18.2024402618408,-18.2824211120605,-18.3566398620605,-18.4330062866211,-18.5500965118408,-18.65625,-18.81298828125,-18.9096698760986,-18.9679679870605,-19.0251941680908,-19.0933609008789,-19.1622066497803,-19.2423820495605,-19.2966785430908,-19.3411140441895,-19.3819332122803,-19.4099617004395,-19.4328117370605,-19.4543952941895,-19.5601558685303,-19.7210941314697,-19.74072265625,-19.8565425872803,-19.90234375,-19.9670906066895,-20.044921875,-20.0696277618408,-20.0908203125,-20.1155261993408,-20.1484355926514,-20.2251949310303,-20.3100566864014,-20.3389644622803,-20.3780269622803,-20.4162101745605,-20.4585933685303,-20.4929676055908,-20.6120109558105,-20.62841796875,-20.6407222747803,-20.7201175689697,-20.7691402435303,-20.8498039245605,-20.9019527435303,-20.9236316680908,-20.9482421875,-21.1296863555908,-21.30029296875,-21.447265625,-21.6185550689697,-21.7530269622803,-21.8606433868408,-21.9828109741211,-22.05712890625,-22.2040042877197,-22.2822265625,-22.3336906433105,-22.4933605194092,-22.6305656433105,-22.7292003631592,-22.7841796875,-22.8229484558105,-22.8577156066895,-22.8794937133789,-22.88916015625,-22.8916988372803,-22.8551750183105,-22.8216800689697],[19.991943359375,19.9475593566895,19.88330078125,19.7646980285645,19.6554698944092,19.5860824584961,19.5579109191895,19.2912120819092,19.20703125,19.171875,19.1351566314697,19.0985355377197,18.97021484375,18.8125991821289,18.7479496002197,18.6983394622803,18.6732921600342,18.6695308685303,18.65576171875,18.56982421875,18.5052242279053,18.4756336212158,18.4475593566895,18.416259765625,18.4220695495605,18.3966789245605,18.3482913970947,18.2591304779053,18.2471179962158,18.226318359375,18.21826171875,18.2811031341553,18.2996101379395,18.3251457214355,18.3677730560303,18.4161128997803,18.5352535247803,18.750244140625,18.8663101196289,18.90771484375,19.2650394439697,19.3041019439697,19.3382797241211,19.4181652069092,19.4813480377197,19.6135749816895,19.6741199493408,19.7611331939697,19.7574710845947,19.7684555053711,19.8428211212158,19.8826675415039,19.9043941497803,19.8888168334961,19.9042472839355,19.9703121185303,19.984375,19.9627437591553,19.992919921875,20.0537109375,20.0560550689697,20.038818359375,19.9755840301514,20.0180168151855,20.0592288970947,20.0547370910645,19.9763660430908,20.0724601745605,20.0976085662842,20.1377429962158,20.1370601654053,20.0594711303711,20.014404296875,19.991943359375],[21.0931644439697,21.05859375,21.0831050872803,21.0390129089355,21.0184574127197,20.9677238464355,21.0068836212158,20.9977531433105,21.0011711120605,21.0251960754395,21.0747547149658,21.0931644439697],[21.6396484375,21.6185073852539,21.623046875,21.6747550964355,21.7102527618408,21.6396484375],[21.6018543243408,21.5818347930908,21.6180667877197,21.669921875,21.6979484558105,21.7332515716553,21.7722663879395,21.7645015716553,21.7526359558105,21.712158203125,21.6948738098145,21.6018543243408],[22.80419921875,22.7579097747803,22.8283214569092,22.8323726654053,22.8585948944092,22.90283203125,22.904541015625,22.85205078125,22.80419921875],[24.4962882995605,24.4361324310303,24.4461441040039,24.4888668060303,24.5014171600342,24.5412120819092,24.5523452758789,24.5185050964355,24.4962882995605],[25.4569816589355,25.4106922149658,25.4327144622803,25.4947261810303,25.5505847930908,25.5908699035645,25.6388187408447,25.6531753540039,25.6232433319092,25.6073722839355,25.591064453125,25.5596675872803,25.5078125,25.4795894622803,25.4569816589355],[28.08642578125,28.0625,28.0625972747803,28.13525390625,28.2043952941895,28.1812992095947,28.1452159881592,28.08642578125],[29.6790027618408,29.6602535247803,29.725341796875,29.7359390258789,29.7727527618408,29.7822265625,29.7007808685303,29.6790027618408],[29.8923835754395,29.8460922241211,29.8526840209961,29.9349594116211,29.9552249908447,29.9502429962158,29.8923835754395],[29.9634284973145,29.9438457489014,30.0012683868408,30.0133304595947,30.0638179779053,30.1431140899658,30.1397457122803,30.0680198669434,30.0313949584961,29.9634284973145],[31.4922847747803,31.4637680053711,31.5496101379395,31.6437511444092,31.7581043243408,31.8053703308105,31.7973613739014,31.7564449310303,31.6936511993408,31.6739273071289,31.6373023986816,31.5521469116211,31.5263671875,31.4922847747803],[42.535400390625,42.5705070495605,42.5816917419434,42.630859375,42.6849594116211,42.7448234558105,42.8726081848145,42.8917999267578,42.91796875,42.9560050964355,42.9741706848145,42.9625968933105,42.974853515625,42.9956550598145,42.9981460571289,42.965087890625,42.8942375183105,42.7765617370605,42.7054672241211,42.6038055419922,42.4749984741211,42.4481430053711,42.4442863464355,42.4358901977539,42.39208984375,42.3846664428711,42.4103050231934,42.4358901977539,42.439208984375,42.41357421875,42.3578605651855,42.3126945495605,42.2705535888672,42.2184562683105,42.1685066223145,42.1423835754395,42.0687980651855,42.0382347106934,42.037841796875,42.0406761169434,42.0208511352539,42.0107421875,42.025634765625,42.0116195678711,41.9874992370605,41.9516143798828,41.8984870910645,41.86376953125,41.840576171875,41.7691421508789,41.7000503540039,41.6553688049316,41.607421875,41.5627937316895,41.5065422058105,41.4330062866211,41.3877449035645,41.3892555236816,41.4156227111816,41.4486808776855,41.4611320495605,41.4399909973145,41.4547386169434,41.4817352294922,41.4837875366211,41.5198249816895,41.5313491821289,41.5545387268066,41.607421875,41.643798828125,41.6873512268066,41.7420425415039,41.7694816589355,41.7810554504395,41.7479972839355,41.7182121276855,41.724853515625,41.716552734375,41.69189453125,41.6409645080566,41.5943374633789,41.4955558776855,41.3939933776855,41.3580551147461,41.3518524169922,41.3213348388672,41.2256813049316,41.1377906799316,41.0782699584961,41.023681640625,40.9740715026855,40.9046363830566,40.8922348022461,40.8720245361328,40.86669921875,40.8386726379395,40.7958984375,40.7789535522461,40.7789535522461,40.7425765991211,40.659912109375,40.6446304321289,40.5894050598145,40.5474624633789,40.5238761901855,40.4978523254395,40.4647445678711,40.4581527709961,40.4598121643066,40.3837394714355,40.3192367553711,40.181640625,40.104248046875,40.0040512084961,40.0115737915039,39.9241714477539,39.8410148620605,39.8224143981934,39.881591796875,39.8408203125,39.7861328125,39.7678718566895,39.762939453125,39.7269058227539,39.6866226196289,39.6735343933105,39.6199226379395,39.600830078125,39.3661117553711,39.267333984375,39.1519050598145,39.0937995910645,39.0531730651855,39.0365219116211,38.9964828491211,38.9914054870605,39.0096702575684,39.00341796875,38.954833984375,38.8917961120605,38.8650894165039,38.8307609558105,38.8082008361816,38.7669448852539,38.7435073852539,38.7316398620605,38.8132820129395,38.9208030700684,38.9466781616211,38.9602546691895,39.1086883544922,39.2201652526855,39.2687492370605,39.3475570678711,39.3865203857422,39.4008293151855,39.3748512268066,39.3768043518066,39.384765625,39.4521980285645,39.5194320678711,39.544677734375,39.6212425231934,39.64013671875,39.6389656066895,39.6852531433105,39.7548828125,39.8448219299316,39.9505386352539,40.04638671875,40.1358413696289,40.3582534790039,40.3960456848145,40.5001945495605,40.5418434143066,40.6027374267578,40.6881866455078,40.8420906066895,40.9742660522461,40.968505859375,40.8758773803711,40.8461418151855,40.8434066772461,40.87841796875,40.9012680053711,40.8416023254395,40.7491188049316,40.68310546875,40.6492195129395,40.5890617370605,40.23095703125,40.203857421875,39.9874534606934,39.9026374816895,39.7524909973145,39.66162109375,39.5608901977539,39.4080581665039,39.2223663330078,39.1825675964355,39.1664047241211,39.172119140625,39.1604957580566,39.1768569946289,39.1180152893066,39.0670890808105,39.195068359375,39.2267570495605,39.1912612915039,39.1344757080078,38.8528823852539,38.6914558410645,38.6251449584961,38.4242210388184,38.3116683959961,38.1833992004395,38.0949211120605,38.1263694763184,38.1266593933105,38.0427742004395,37.9040031433105,37.8091773986816,37.7765121459961,37.7485847473145,37.7007331848145,37.6610336303711,37.641357421875,37.4940910339355,37.3311538696289,37.277099609375,37.201171875,37.1382827758789,37.124755859375,37.1550788879395,37.2533683776855,37.2957992553711,37.350830078125,37.4950218200684,37.6227035522461,37.656494140625,37.6790046691895,37.6920890808105,37.7010269165039,37.8339347839355,37.7251930236816,37.6001472473145,37.5789566040039,37.5150375366211,37.4603500366211,37.4566421508789,37.4453125,37.4957504272461,37.5289039611816,37.5223121643066,37.4561538696289,37.4052734375,37.407958984375,37.4264183044434,37.40283203125,37.3179206848145,37.1811027526855,37.1378440856934,37.068115234375,37.0222663879395,37.0026359558105,36.9468269348145,36.9151382446289,36.8322257995605,36.8338356018066,36.849853515625,36.8795433044434,36.9272003173828,36.9586410522461,36.9707527160645,36.95947265625,36.8363800048828,36.7383804321289,36.6604499816895,36.6113777160645,36.5979499816895,36.6351585388184,36.6328125,36.6072235107422,36.5389175415039,36.4853019714355,36.4441413879395,36.4260749816895,36.456298828125,36.4132804870605,36.3407211303711,36.1683578491211,36.1299324035645,36.1086883544922,36.0538558959961,36.0792007446289,36.110107421875,36.189453125,36.2281723022461,36.2261734008789,36.2024421691895,36.1502914428711,36.118896484375,36.0174789428711,36.0072288513184,35.9844245910645,35.9349136352539,35.8611335754395,35.7993659973145,35.740234375,35.6932144165039,35.6436538696289,35.6177253723145,35.5887222290039,35.4698715209961,35.3585968017578,35.3014144897461,35.1138191223145,35.0117683410645,34.8488273620605,34.7484359741211,34.7494125366211,34.7141609191895,34.5822296142578,34.4961929321289,34.4478034973145,34.32568359375,34.2740211486816,34.1689949035645,33.8663063049316,33.7164535522461,33.63818359375,33.4905281066895,33.2366218566895,33.0165023803711,32.8432121276855,32.7641105651855,32.661376953125,32.5670433044434,32.4573249816895,32.425048828125,32.3719253540039,32.2062492370605,32.1533203125,32.12109375,32.051025390625,31.9928722381592,31.8997573852539,31.8164558410645,31.7035617828369,31.6995124816895,31.712158203125,31.8587894439697,31.8626956939697,31.8423347473145,31.8693866729736,32.03173828125,32.0810546875,32.1058616638184,31.9661636352539,31.9759788513184,31.9602546691895,31.9362773895264,31.9063472747803,31.9520988464355,32.0198249816895,31.9837398529053,31.9228534698486,31.8197746276855,31.7501964569092,31.7194328308105,31.6280746459961,31.4853515625,31.3197269439697,31.1628913879395,31.0616207122803,30.9169921875,30.870361328125,30.86376953125,30.8409671783447,30.7897968292236,30.6997051239014,30.5582523345947,30.4697265625,30.3926277160645,30.3546371459961,30.390869140625,30.3878421783447,30.2835464477539,30.2413082122803,30.2495632171631,30.2630386352539,30.2474117279053,30.3030776977539,30.1331558227539,30.1606426239014,30.3017578125,30.3041019439697,30.2823715209961,30.2266597747803,29.9791011810303,29.9521484375,29.8940925598145,29.8876953125,29.870361328125,29.7796859741211,29.5837898254395,29.5370121002197,29.4845714569092,29.510986328125,29.6046390533447,29.6277828216553,29.6059093475342,29.4906253814697,29.1350078582764,29.12890625,29.2256832122803,29.25634765625,29.2361335754395,29.2367172241211,29.1931648254395,29.13134765625,29.1184558868408,29.0105972290039,28.953125,28.9159164428711,28.931884765625,28.8514156341553,28.7679195404053,28.7348136901855,28.7136707305908,28.6414070129395,28.5210933685303,28.3666019439697,28.2921390533447,28.3242683410645,28.2298812866211,28.2221202850342,28.34619140625,28.32666015625,28.29052734375,28.1572761535645,28.0370121002197,28.00390625,28.0133781433105,28.0099601745605,27.9774417877197,27.9377918243408,27.8914546966553,27.7445793151855,27.6878890991211,27.6394538879395,27.5807609558105,27.4821281433105,27.4124031066895,27.318359375,27.2562484741211,27.155517578125,27.0970687866211,26.8861331939697,26.7806644439697,26.6715812683105,26.6338386535645,26.5863761901855,26.6104488372803,26.6830081939697,26.6893081665039,26.7369136810303,26.797607421875,26.8463859558105,26.8314952850342,26.7747077941895,26.7286624908447,26.7472648620605,26.7849617004395,26.73046875,26.6758785247803,26.6211910247803,26.6094226837158,26.5466289520264,26.4501953125,26.4141597747803,26.3709468841553,26.3341808319092,26.3001480102539,26.2364253997803,26.1273441314697,26.0546875,26.0540523529053,26.0625495910645,26.1043930053711,26.1217765808105,25.9748039245605,25.94873046875,25.954345703125,26.0091800689697,26.0035648345947,25.918701171875,25.8228988647461,25.6986827850342,25.5912590026855,25.4374523162842,25.3911609649658,25.3680171966553,25.4086418151855,25.4596195220947,25.4462890625,25.468017578125,25.4498062133789,25.414306640625,25.355712890625,25.3070316314697,25.2322273254395,25.2059574127197,25.2234382629395,25.2092761993408,25.1268062591553,25.0047855377197,24.9289054870605,24.9111328125,24.8498039245605,24.8355464935303,24.80908203125,24.7823238372803,24.7461433410645,24.6214351654053,24.5803718566895,24.6007328033447,24.57275390625,24.6258296966553,24.6270027160645,24.5599117279053,24.4819831848145,24.4742183685303,24.4798336029053,24.4743156433105,24.4324703216553,24.3958988189697,24.3796405792236,24.3271484375,24.24609375,24.1064453125,24.0123043060303,24.0147953033447,23.9392585754395,23.8367195129395,23.8569812774658,23.840576171875,23.7916984558105,23.771484375,23.7362308502197,23.6209945678711,23.588623046875,23.6357421875,23.7087879180908,23.71435546875,23.6470222473145,23.5987796783447,23.5787582397461,23.6234378814697,23.6466808319092,23.4530773162842,23.3825187683105,23.3604984283447,23.3538570404053,23.327392578125,23.2777824401855,23.2281742095947,23.21826171875,23.1796875,23.006591796875,22.9458999633789,22.9410648345947,22.9813480377197,22.9495601654053,22.9186515808105,22.887451171875,22.8791007995605,22.8015613555908,22.8239250183105,22.8534183502197,22.82470703125,22.7651863098145,22.7188472747803,22.7188472747803,22.7759761810303,22.8172855377197,22.7816886901855,22.7089366912842,22.6846179962158,22.6395015716553,22.6167964935303,22.6263179779053,22.7387199401855,22.7552738189697,22.6984386444092,22.654052734375,22.62060546875,22.5289058685303,22.5270519256592,22.583251953125,22.5932121276855,22.5409660339355,22.553955078125,22.5649890899658,22.5649890899658,22.55126953125,22.54296875,22.5144538879395,22.5119132995605,22.5310535430908,22.6072254180908,22.733642578125,22.8016586303711,22.8614253997803,22.9688968658447,23.02001953125,23.0769538879395,23.1274909973145,23.1020984649658,23.0550785064697,22.9957027435303,22.9405765533447,22.9120121002197,22.8888168334961,22.8645992279053,22.7894039154053,22.7261219024658,22.6923828125,22.5940418243408,22.4041500091553,22.3504886627197,22.2972660064697,22.2251949310303,22.22412109375,22.2459487915039,22.2415523529053,22.2174797058105,22.195556640625,22.1944332122803,22.1783695220947,22.1648426055908,22.1454105377197,22.0887699127197,22.0750007629395,22.2079582214355,22.1193351745605,21.938232421875,21.9073238372803,21.8814449310303,21.9446296691895,21.90234375,21.8594722747803,21.8198719024658,21.77685546875,21.8183097839355,21.880615234375,21.9273433685303,21.9581050872803,21.9813461303711,21.97802734375,21.91748046875,21.8012218475342,21.767578125,21.74169921875,21.7631340026855,21.8064937591553,21.843017578125,21.8496589660645,21.7762680053711,21.7171382904053,21.7097644805908,21.71923828125,21.6552238464355,21.6084957122803,21.55908203125,21.5351066589355,21.4861316680908,21.493896484375,21.4822254180908,21.4847164154053,21.510986328125,21.51171875,21.4302730560303,21.3959465026855,21.3865222930908,21.2791004180908,21.2140617370605,21.2074222564697,21.2305679321289,21.326904296875,21.338134765625,21.2477035522461,21.17236328125,21.13134765625,21.0376472473145,20.9446296691895,20.8585929870605,20.8375968933105,20.79052734375,20.7520503997803,20.7199211120605,20.6716804504395,20.5182628631592,20.4600086212158,20.4268550872803,20.3554210662842,20.2948246002197,20.2637195587158,20.2951164245605,20.3640613555908,20.4131355285645,20.3988780975342,20.4032726287842,20.4481449127197,20.4743671417236,20.5143051147461,20.6218738555908,20.7114753723145,20.7807121276855,20.8387680053711,20.8736343383789,20.9168949127197,21.052734375,21.1316413879395,21.2283687591553,21.3374519348145,21.37646484375,21.4805660247803,21.4835929870605,21.56005859375,21.5279788970947,21.5246086120605,21.6719741821289,21.6905746459961,21.6934089660645,21.5379390716553,21.4794921875,21.4539546966553,21.4434070587158,21.425537109375,21.4402828216553,21.4873542785645,21.5436038970947,21.5904788970947,21.6265125274658,21.6244125366211,21.6344718933105,21.6304683685303,21.6512699127197,21.7246589660645,21.7704601287842,21.8159675598145,21.868896484375,21.9010257720947,21.9046382904053,21.8288078308105,21.7394046783447,21.67138671875,21.6334476470947,21.6073246002197,21.67919921875,21.6969242095947,21.6935062408447,21.6219234466553,21.5583972930908,21.565185546875,21.5259761810303,21.5079574584961,21.5604000091553,21.6451663970947,21.655029296875,21.6139163970947,21.5983390808105,21.6422843933105,21.60888671875,21.7106437683105,21.7170906066895,21.794189453125,21.8348636627197,21.8934078216553,21.9239253997803,21.9201183319092,21.9512691497803,21.9819812774658,22.0003414154053,21.9861812591553,21.9789066314697,22.0182132720947,22.136474609375,22.241455078125,22.2886238098145,22.3245105743408,22.3416996002197,22.3954105377197,22.5013675689697,22.5732421875,22.5860347747803,22.6377429962158,22.7109375,22.7789058685303,22.874267578125,22.9083499908447,22.8938961029053,22.8634757995605,22.8574714660645,22.8694343566895,22.9551277160645,22.9700679779053,22.9755363464355,22.9747543334961,22.9374504089355,22.9249515533447,22.9228038787842,22.9693355560303,23.0299320220947,23.0726547241211,23.1219730377197,23.1808586120605,23.2353515625,23.3076648712158,23.34521484375,23.3221206665039,23.2810554504395,23.1943359375,23.1605472564697,23.1363754272461,23.1001949310303,23.0107917785645,22.9111328125,22.8604984283447,22.8222160339355,22.8182125091553,22.8200206756592,22.8041019439697,22.7040519714355,22.7120113372803,22.7685050964355,22.8094234466553,22.8001461029053,22.7522945404053,22.6663589477539,22.5861320495605,22.5504875183105,22.5400886535645,22.5382328033447,22.77001953125,22.7820320129395,22.7344245910645,22.611572265625,22.5879878997803,22.5974102020264,22.7546882629395,22.769775390625,22.764404296875,22.7135257720947,22.6385250091553,22.5929679870605,22.542236328125,22.4975090026855,22.4529781341553,22.4482440948486,22.4661617279053,22.525390625,22.587158203125,22.6484870910645,22.7003917694092,22.7410163879395,22.7509288787842,22.7328128814697,22.7080078125,22.6466312408447,22.5459957122803,22.4660148620605,22.4146480560303,22.3791980743408,22.4122562408447,22.439208984375,22.4394054412842,22.3884773254395,22.4903316497803,22.4952640533447,22.486572265625,22.4623050689697,22.4054203033447,22.3274402618408,22.2763671875,22.253662109375,22.2098636627197,22.1624011993408,22.1208972930908,22.0552730560303,21.9896965026855,21.8824710845947,21.8265132904053,21.7779769897461,21.6057605743408,21.5338382720947,21.39501953125,21.31494140625,21.2789058685303,21.2359867095947,21.2125968933105,21.2041492462158,21.1563968658447,21.150146484375,21.1696281433105,21.1844234466553,21.2035636901855,21.2342777252197,21.2308120727539,21.1841297149658,21.1973152160645,21.2237300872803,21.2782230377197,21.3424320220947,21.38330078125,21.4075183868408,21.5220718383789,21.5674800872803,21.5816402435303,21.7051277160645,21.7355480194092,21.74609375,21.755859375,21.7363777160645,21.6551761627197,21.5049304962158,21.4717769622803,21.4581050872803,21.4840812683105,21.5010261535645,21.4629878997803,21.4805183410645,21.5111827850342,21.5579090118408,21.6170406341553,21.6606464385986,21.6827640533447,21.7016105651855,21.7587413787842,21.82080078125,21.901611328125,21.9883308410645,22.0280284881592,22.0497074127197,22.0891590118408,22.1107902526855,22.1006336212158,22.1101551055908,22.1259765625,22.1533203125,22.1924800872803,22.2825679779053,22.370361328125,22.498046875,22.5865230560303,22.6886730194092,22.8250980377197,22.9272937774658,22.9591312408447,23.0045890808105,23.0462398529053,23.0692386627197,23.0958995819092,23.1033191680908,23.1305675506592,23.1912593841553,23.3074703216553,23.3803215026855,23.4400882720947,23.4825191497803,23.5204105377197,23.6243667602539,23.7378425598145,23.7831058502197,23.841796875,23.9050788879395,23.9640617370605,24.090576171875,24.1211929321289,24.1187000274658,24.1160640716553,24.06982421875,24.0988292694092,24.1156730651855,24.1190414428711,24.1106452941895,24.0654296875,23.9862785339355,23.931884765625,23.8980960845947,23.8871574401855,23.9110355377197,23.9884757995605,24.1308116912842,24.228759765625,24.312744140625,24.3799819946289,24.4229507446289,24.44384765625,24.49169921875,24.6312007904053,24.7748031616211,24.8201160430908,24.8419914245605,24.869873046875,24.9703598022461,25.0343265533447,25.1580562591553,25.2518539428711,25.2361335754395,25.2593269348145,25.2925281524658,25.3489742279053,25.4157238006592,25.5710945129395,25.5945301055908,25.56884765625,25.5867671966553,25.677978515625,25.7888660430908,25.8232421875,25.8267097473145,25.8635749816895,25.9177722930908,26.0037097930908,26.0724124908447,26.1140613555908,26.1394538879395,26.1893539428711,26.2985343933105,26.4296894073486,26.5833988189697,26.6981430053711,26.7857418060303,26.8773918151855,27.044921875,27.1906242370605,27.2453117370605,27.4219245910645,27.5724601745605,27.5988273620605,27.6476554870605,27.6572265625,27.6394538879395,27.5870609283447,27.5380859375,27.5500965118408,27.5990715026855,27.6631832122803,27.9675769805908,28.0552234649658,28.1422843933105,28.1858882904053,28.2115230560303,28.3138179779053,28.3564949035645,28.3635749816895,28.3563480377197,28.3561515808105,28.4071292877197,28.4693374633789,28.5,28.5170402526855,28.5102043151855,28.4563465118408,28.4256362915039,28.3539066314697,28.2544937133789,28.2179679870605,28.23681640625,28.3403339385986,28.3689441680908,28.3376941680908,28.3624019622803,28.3670406341553,28.4497566223145,28.4599132537842,28.406005859375,28.367919921875,28.3672847747803,28.3865242004395,28.4120597839355,28.4281730651855,28.4685554504395,28.496826171875,28.525390625,28.6065425872803,28.763671875,28.82958984375,28.9593257904053,29.0222663879395,29.0506839752197,29.0274391174316,28.9097175598145,28.922607421875,28.9634761810303,29.0820808410645,29.1176776885986,29.1612300872803,29.2098159790039,29.2490730285645,29.2609844207764,29.2458019256592,29.2724590301514,29.3813972473145,29.4241199493408,29.4471683502197,29.3909187316895,29.3138198852539,29.2063465118408,29.1511726379395,29.1370124816895,29.102294921875,29.0542964935303,29.0374011993408,29.035888671875,29.0495586395264,29.1040534973145,29.1491718292236,29.1440410614014,29.1758804321289,29.2012691497803,29.2516098022461,29.2970199584961,29.3124027252197,29.2162132263184,29.1446285247803,29.0599098205566,28.9758796691895,28.9595203399658,28.9075183868408,28.8607902526855,28.80322265625,28.7297840118408,28.6902332305908,28.6540546417236,28.6294918060303,28.5908203125,28.4927253723145,28.4022941589355,28.32763671875,28.2281246185303,28.1471195220947,28.0933589935303,28.0615234375,28.0498542785645,28.0257320404053,27.9889640808105,27.9489250183105,27.8791999816895,27.845947265625,27.8246097564697,27.8207511901855,27.8302230834961,27.8415031433105,27.8269519805908,27.812255859375,27.8076171875,27.7303714752197,27.7296867370605,27.7464370727539,27.7598152160645,27.7599620819092,27.8008785247803,27.9232425689697,27.9517097473145,27.9817867279053,28.0216312408447,28.0640144348145,28.078369140625,28.0712413787842,28.0267581939697,27.9744625091553,27.9700679779053,27.9945812225342,28.0265140533447,28.0717296600342,28.0785636901855,28.0708503723145,28.0802249908447,28.093994140625,28.119140625,28.1681652069092,28.2165031433105,28.2439460754395,28.2777347564697,28.3020496368408,28.3111820220947,28.2941398620605,28.2562999725342,28.1881847381592,28.1583003997803,28.107421875,28.0599613189697,27.9581546783447,27.8331546783447,27.7112789154053,27.5925769805908,27.5178699493408,27.4640121459961,27.3160648345947,27.3628406524658,27.4298820495605,27.5218753814697,27.7673835754395,27.86865234375,27.9072761535645,28.0069332122803,28.0396976470947,28.0918464660645,28.0933589935303,28.057373046875,28.0344715118408,28.0116691589355,27.9688472747803,27.9489250183105,27.9330081939697,27.904541015625,27.87060546875,27.8833980560303,27.8908214569092,27.8860855102539,27.8213882446289,27.815185546875,27.8218269348145,27.8238277435303,27.8219242095947,27.8383293151855,27.9286613464355,27.9684562683105,27.9991703033447,28.0220699310303,28.0706558227539,28.0949211120605,28.10302734375,28.085205078125,27.9635257720947,27.9395503997803,27.9286613464355,27.9595203399658,28.0220699310303,28.0916996002197,28.1143550872803,28.0835933685303,27.9945812225342,27.9347171783447,27.910400390625,27.92822265625,27.9896965026855,28.1353511810303,28.2206554412842,28.2774410247803,28.2760257720947,28.2926254272461,28.3159656524658,28.3722667694092,28.4842777252197,28.5718746185303,28.5922355651855,28.6026363372803,28.6096687316895,28.5536136627197,28.5602054595947,28.5792484283447,28.5955562591553,28.6215343475342,28.6595687866211,28.7529315948486,28.8039054870605,28.8681163787842,28.9117679595947,29.036376953125,29.1562976837158,29.2199687957764,29.2538566589355,29.2794914245605,29.2274417877197,29.1875972747803,29.1835956573486,29.3063468933105,29.4391593933105,29.5545902252197,29.6126480102539,29.6180667877197,29.6833972930908,29.8311996459961,29.9415035247803,30.0638656616211,30.1151847839355,30.1589832305908,30.2450675964355,30.3267574310303,30.3624038696289,30.3875007629395,30.3375968933105,30.0933113098145,30.0398921966553,30.0368156433105,30.0989742279053,30.1645030975342,30.2371082305908,30.2905769348145,30.3603992462158,30.4148426055908,30.4488773345947,30.4635238647461,30.5094718933105,30.5613288879395,30.5684089660645,30.6053237915039,30.6837406158447,30.7592277526855,30.7898445129395,30.7819347381592,30.8887691497803,30.8941879272461,30.9246101379395,30.96826171875,30.9652328491211,30.9490718841553,30.9937019348145,31.064208984375,31.0799331665039,31.105712890625,31.2417488098145,31.4026355743408,31.4262199401855,31.4141101837158,31.32861328125,31.3372077941895,31.3313484191895,31.301513671875,31.2936515808105,31.302490234375,31.3237800598145,31.4365711212158,31.4718246459961,31.55029296875,31.6180648803711,31.6683597564697,31.7403812408447,31.8055171966553,31.8876457214355,31.9579601287842,31.9837875366211,32.0230484008789,32.2157707214355,32.2362327575684,32.3003387451172,32.3973617553711,32.4667015075684,32.5198745727539,32.5447273254395,32.5577163696289,32.57080078125,32.5789527893066,32.5970191955566,32.5583992004395,32.499267578125,32.4680671691895,32.4119644165039,32.3582038879395,32.3651390075684,32.38818359375,32.4757804870605,32.4972152709961,32.5010757446289,32.507568359375,32.5640144348145,32.7030754089355,32.7587890625,32.8090324401855,32.8648452758789,32.946044921875,33.00146484375,33.0226554870605,33.0525398254395,33.1081047058105,33.1719245910645,33.2189445495605,33.2262687683105,33.2503890991211,33.2914543151855,33.3465347290039,33.3867683410645,33.4311027526855,33.4997062683105,33.6503410339355,33.8087921142578,33.8875961303711,34.0133781433105,34.0555648803711,34.0876960754395,34.18896484375,34.2282218933105,34.2581062316895,34.3026390075684,34.3519554138184,34.3903312683105,34.4529304504395,34.5181655883789,34.5579605102539,34.6063957214355,34.6539077758789,34.7698211669922,34.9464836120605,35.1349143981934,35.2510261535645,35.3069343566895,35.4494132995605,35.4797859191895,35.490234375,35.4716300964355,35.4490242004395,35.4607925415039,35.4845199584961,35.4927749633789,35.4958992004395,35.4805641174316,35.4718246459961,35.4734382629395,35.4755859375,35.5081558227539,35.5520515441895,35.61328125,35.6617202758789,35.6786613464355,35.7293968200684,35.7729988098145,35.8870620727539,35.8782234191895,35.837158203125,35.8109397888184,35.810546875,35.8290023803711,35.94921875,35.9830093383789,35.996337890625,36.017578125,36.0489730834961,36.0884742736816,36.1339340209961,36.1688461303711,36.3824234008789,36.4581069946289,36.5215797424316,36.6007347106934,36.6497039794922,36.6949234008789,36.7419929504395,36.7593269348145,36.7250480651855,36.7382316589355,36.8836898803711,36.9134788513184,36.9732398986816,36.9871597290039,36.9683570861816,36.9524383544922,36.9791030883789,37.0127449035645,37.0357398986816,37.0366706848145,37.0221710205078,37.0306625366211,37.0572242736816,37.1373519897461,37.15771484375,37.2366218566895,37.2667007446289,37.2907218933105,37.28564453125,37.2491722106934,37.2250518798828,37.2316398620605,37.25,37.2935523986816,37.3440437316895,37.3856925964355,37.4512710571289,37.5303688049316,37.5728034973145,37.6014175415039,37.6873054504395,37.7725105285645,37.8049812316895,37.8327140808105,37.92578125,38.0380859375,38.1036148071289,38.19189453125,38.2747535705566,38.404296875,38.4603042602539,38.510009765625,38.6000022888184,38.6597671508789,38.6575164794922,38.6611824035645,38.6084938049316,38.5398445129395,38.53369140625,38.5628890991211,38.6068878173828,38.6989250183105,38.8172340393066,38.8542976379395,38.88623046875,38.9146957397461,38.9413108825684,38.9686546325684,39.0021476745605,39.0445289611816,39.1045417785645,39.2291984558105,39.2978515625,39.3966789245605,39.4488792419434,39.4622535705566,39.4889640808105,39.5333023071289,39.5785140991211,39.6064949035645,39.7145500183105,39.7628402709961,39.8001480102539,39.8286628723145,39.8779296875,39.9788093566895,40.0431137084961,40.0593719482422,40.0743179321289,40.092041015625,40.13720703125,40.2721672058105,40.3105964660645,40.3298835754395,40.3285179138184,40.344970703125,40.4285163879395,40.4587898254395,40.4826164245605,40.4935073852539,40.4495124816895,40.4541015625,40.4802742004395,40.6275405883789,40.6251983642578,40.6053199768066,40.5166015625,40.3292465209961,40.3058090209961,40.30322265625,40.3714332580566,40.3875465393066,40.37646484375,40.4084014892578,40.4307632446289,40.3522453308105,40.3897933959961,40.4495124816895,40.51123046875,40.577880859375,40.662353515625,40.7422370910645,40.7796363830566,40.818115234375,40.9823226928711,41.024169921875,41.0391616821289,41.0107421875,41.0143547058105,40.9927749633789,41.0243148803711,41.0556144714355,41.0506820678711,41.0756340026855,41.2814445495605,41.3251953125,41.3716316223145,41.4175300598145,41.4595680236816,41.56005859375,41.7191390991211,41.7828102111816,41.7810554504395,41.8209953308105,41.8988761901855,41.9957542419434,42.0149917602539,42.0324211120605,42.04345703125,42.059814453125,42.1298332214355,42.1900367736816,42.2078132629395,42.2354011535645,42.274169921875,42.3994140625,42.5183563232422,42.6255378723145,42.66552734375,42.7344703674316,42.7972640991211,42.8557624816895,42.8734855651855,42.9117202758789,42.935546875,42.9737777709961,42.99560546875,43.0204124450684,43.0431137084961,43.0857887268066,43.1282730102539,43.1024894714355,43.1189460754395,43.1615753173828,43.204345703125,43.2742691040039,43.31005859375,43.3529815673828,43.4270515441895,43.5641593933105,43.6851081848145,43.89208984375,43.9517593383789,44.0471687316895,44.0972671508789,44.1712875366211,44.2232894897461,44.3265151977539,44.4383773803711,44.552001953125,44.6268081665039,44.6554222106934,44.6769065856934,44.68408203125,44.7146492004395,44.74609375,44.7703132629395,44.80810546875,44.8037605285645,44.7972145080566,44.8251953125,44.86083984375,44.8837890625,44.944091796875,45.0064468383789,45.0339851379395,45.0750961303711,45.10498046875,45.1265144348145,45.1355476379395,45.1292953491211,45.1691398620605,45.2461929321289,45.3108406066895,45.349365234375,45.3108406066895,45.2260246276855,45.1820793151855,45.1608390808105,45.161865234375,45.1948738098145,45.2190933227539,45.2058601379395,45.1624488830566,45.1235847473145,45.12548828125,45.1554183959961,45.2159690856934,45.2931137084961,45.3744163513184,45.4242668151855,45.4425773620605,45.4719734191895,45.5199203491211,45.563720703125,45.5949211120605,45.6715316772461,45.8119125366211,46.0058097839355,46.15869140625,46.3866691589355,46.6244621276855,46.9660148620605,47.0334968566895,47.1414566040039,47.1859397888184,47.2093772888184,47.1865730285645,47.108642578125,47.043212890625,47.0210456848145,46.9978523254395,46.9705085754395,46.9786148071289,46.9947280883789,46.9961395263672,46.9757804870605,46.9749488830566,46.9723625183105,46.9393539428711,46.8643569946289,46.8307151794434,46.8431663513184,46.9092292785645,46.9612312316895,47.036376953125,47.0467300415039,47.0635261535645,47.1007804870605,47.1884765625,47.254638671875,47.33837890625,47.3974113464355,47.49365234375,47.5584945678711,47.7464866638184,47.9156227111816,48.0518569946289,48.2040061950684,48.2505378723145,48.3118171691895,48.3850593566895,48.4080581665039,48.4237289428711,48.4545402526855,48.4862289428711,48.50537109375,48.5286140441895,48.6355476379395,48.6971702575684,48.8607406616211,48.9393577575684,49.0088386535645,49.0497055053711,49.090576171875,49.0975570678711,49.1099090576172,49.1058616638184,49.0857887268066,49.0766105651855,49.0914535522461,49.1224136352539,49.1323699951172,49.147216796875,49.1658172607422,49.1623039245605,49.1163063049316,49.0802726745605,49.0319328308105,49.0001449584961,48.9655265808105,48.9455070495605,48.9185562133789,48.8816413879395,48.8357429504395,48.7916526794434,48.7652816772461,48.735595703125,48.7071800231934,48.675048828125,48.6404304504395,48.6033210754395,48.5810546875,48.5551300048828,48.537841796875,48.5090827941895,48.4720687866211,48.4034156799316,48.3844718933105,48.3174324035645,48.2201652526855,48.1705589294434,48.1017112731934,48.0890121459961,48.0499496459961,48.0025405883789,47.9876976013184,47.9809074401855,48.0248527526855,48.029052734375,48.0039558410645,47.9090805053711,47.879150390625,47.8524894714355,47.8270034790039,47.8232917785645,47.8443336486816,47.8863258361816,47.877685546875,47.8504867553711,47.8035659790039,47.7454109191895,47.7020988464355,47.6761703491211,47.6551742553711,47.5969734191895,47.556640625,47.5041007995605,47.4081535339355,47.3288078308105,47.28515625,47.2140121459961,47.1002922058105,47.0038566589355,46.9851570129395,46.9544944763184,46.8832511901855,46.7490234375,46.6610832214355,46.5957527160645,46.5660629272461,46.5290031433105,46.3879890441895,46.3242683410645,46.2706565856934,46.1772918701172,46.10498046875,46.0357933044434,45.9850578308105,45.921630859375,45.8854026794434,45.853515625,45.7308120727539,45.5951652526855,45.5252418518066,45.4746589660645,45.4189453125,45.3706550598145,45.2628936767578,45.1960906982422,45.1939430236816,45.2159156799316,45.2174301147461,45.1939430236816,45.14453125,45.1181144714355,45.124755859375,45.0982437133789,45.0765113830566,45.0689468383789,45.0693359375,45.0685043334961,45.0352554321289,45.0089340209961,45.010986328125,45.0357437133789,45.0201683044434,44.983154296875,44.9444808959961,44.9009780883789,44.8319358825684,44.7242202758789,44.6749534606934,44.6451644897461,44.5194854736328,44.4725074768066,44.350830078125,44.3033218383789,44.2594261169434,44.278076171875,44.2615242004395,44.1954078674316,44.1048812866211,44.0393562316895,44.0059547424316,43.9861831665039,43.9539527893066,43.8922386169434,43.8536109924316,43.6640625,43.3836898803711,43.2759742736816,43.2064933776855,43.0961456298828,43.0145034790039,42.9287109375,42.8493156433105,42.7467765808105,42.7203598022461,42.7438468933105,42.76025390625,42.789794921875,42.7362785339355,42.6845245361328,42.6387176513672,42.6162109375,42.5682106018066,42.6294441223145,42.6773452758789,42.6707534790039,42.6167984008789,42.5877952575684,42.5513153076172,42.5378913879395,42.5387687683105,42.5235366821289,42.5000495910645,42.4658203125,42.2923316955566,42.2158699035645,42.1581077575684,42.0920906066895,42.052001953125,42.0059585571289,41.9365730285645,41.8558616638184,41.7513160705566,41.7969741821289,41.8461456298828,41.8770027160645,41.65869140625,41.6411628723145,41.6437492370605,41.5955085754395,41.6159210205078,41.6632804870605,41.738037109375,41.7709007263184,41.854736328125,41.8750991821289,41.9939956665039,42.1405754089355,42.2115745544434,42.2273445129395,42.2635765075684,42.2887191772461,42.3088874816895,42.321533203125,42.3492660522461,42.4009780883789,42.4058570861816,42.4271965026855,42.4473152160645,42.436767578125,42.4292945861816,42.4161148071289,42.4264640808105,42.4405746459961,42.4383811950684,42.4559555053711,42.5105476379395,42.5538101196289,42.6062469482422,42.6605949401855,42.7100105285645,42.7427253723145,42.773681640625,42.8135719299316,42.8441390991211,42.895263671875,42.9905281066895,43.0738792419434,43.1107902526855,43.194091796875,43.2568855285645,43.3414077758789,43.3687515258789,43.3919944763184,43.4749031066895,43.4927749633789,43.4962921142578,43.5631828308105,43.6211433410645,43.6646003723145,43.68017578125,43.71142578125,43.75244140625,43.81494140625,43.87890625,43.9346694946289,44.0411148071289,44.1071281433105,44.19189453125,44.2716293334961,44.3223648071289,44.3672866821289,44.419189453125,44.5115699768066,44.56982421875,44.6729011535645,44.8271484375,44.899169921875,44.9695320129395,45.0640602111816,45.0816383361816,45.0629386901855,45.0630378723145,45.0582046508789,45.0109367370605,44.9176788330078,44.8834457397461,44.8103523254395,44.7948265075684,44.7916488647461,44.7674331665039,44.7623558044434,44.7457046508789,44.7634735107422,44.825927734375,44.8961944580078,44.9123039245605,44.9425773620605,44.9711418151855,45.0498542785645,45.1108894348145,45.2025871276855,45.271728515625,45.3163070678711,45.3646011352539,45.3899917602539,45.4132804870605,45.4196281433105,45.3782691955566,45.3902320861816,45.3961944580078,45.4199676513672,45.4395027160645,45.458251953125,45.5348167419434,45.6261711120605,45.6769523620605,45.6867179870605,45.7393569946289,45.7959938049316,45.8457527160645,45.8869132995605,45.9630355834961,46.0965805053711,46.1587905883789,46.2090797424316,46.289794921875,46.3130836486816,46.3219757080078,46.3766593933105,46.387939453125,46.3617668151855,46.3550758361816,46.3522453308105,46.3620109558105,46.3913078308105,46.4366683959961,46.53759765625,46.5708961486816,46.5862312316895,46.5882797241211,46.552001953125,46.5220718383789,46.5181655883789,46.5376968383789,46.6193351745605,46.6666030883789,46.6785659790039,46.717041015625,46.7031745910645,46.69189453125,46.70166015625,46.69189453125,46.7470703125,46.7602043151855,46.73486328125,46.6921882629395,46.638671875,46.6138191223145,46.6266593933105,46.6039543151855,46.6060028076172,46.627197265625,46.6721687316895,46.7328605651855,46.7914543151855,46.8578109741211,46.9065895080566,46.9788093566895,47.0270042419434,47.0900382995605,47.1500015258789,47.2224578857422,47.2559051513672,47.380859375,47.41015625,47.4307136535645,47.47265625,47.4925804138184,47.5251922607422,47.5584945678711,47.6162605285645,47.6541519165039,47.6853523254395,47.7029304504395,47.72509765625,47.7576179504395,47.822265625,47.9432640075684,47.9839859008789,47.99951171875,48.0289077758789,48.0189437866211,47.9996070861816,47.9998550415039,47.9878921508789,47.9082984924316,47.8046875,47.7413597106934,47.6757316589355,47.6521987915039,47.6663551330566,47.7402839660645,47.806396484375,47.8365707397461,47.8530769348145,47.8697738647461,47.864501953125,47.8395500183105,47.8440437316895,47.85986328125,47.8582038879395,47.78955078125,47.7113265991211,47.6869163513184,47.7382316589355,47.7989234924316,47.8748054504395,47.9450187683105,48.130859375,48.1862297058105,48.2482414245605,48.3463401794434,48.4557113647461,48.5772476196289,48.6893539428711,48.7822761535645,48.8400382995605,48.9361343383789,49.0374526977539,49.1703605651855,49.4062004089355,49.684814453125,49.8237762451172,49.73779296875,49.6929702758789,49.6248512268066,49.6094245910645,49.5358428955078,49.5135269165039,49.5134773254395,49.6927757263184,49.844482421875,49.9628410339355,49.9788589477539,50.0133781433105,50.06640625,50.1549339294434,50.2789535522461,50.3539047241211,50.3798332214355,50.406005859375,50.4325180053711,50.4841804504395,50.5609893798828,50.6338844299316,50.7028350830078,50.7792510986328,50.8631362915039,50.94677734375,51.0301246643066,51.1077117919922,51.1794929504395,51.2670402526855,51.422119140625,51.6006813049316,51.7229957580566,51.8485374450684,51.9730453491211,52.0965347290039,52.2054672241211,52.2999038696289,52.3958969116211,52.4936027526855,52.566650390625,52.6150398254395,52.6269989013672,52.6024894714355,52.6329116821289,52.7182159423828,52.7872047424316,52.8398933410645,52.9680671691895,53.1718292236328,53.2845687866211,53.3170394897461,53.3835945129395,53.4514656066895,53.4850082397461,53.4625015258789,53.4569816589355,53.468505859375,53.4977073669434,53.5445823669434,53.5556144714355,53.53076171875,53.5294418334961,53.5264663696289,53.5266609191895,53.5465354919434,53.510986328125,53.4056129455566,53.3586921691895,53.3701171875,53.3408660888672,53.2709465026855,53.2296371459961,53.2106437683105,53.1338386535645,53.1297378540039,53.1726570129395,53.1973152160645,53.2036628723145,53.1658210754395,53.083740234375,53.047607421875,53.0574684143066,53.0422859191895,53.0037117004395,52.956298828125,52.9308090209961,52.8907203674316,52.8907203674316,52.8715324401855,52.8006858825684,52.7678680419922,52.7394523620605,52.7158660888672,52.6919937133789,52.6734848022461,52.6430168151855,52.6102066040039,52.5733375549316,52.546630859375,52.5191421508789,52.4838409423828,52.44482421875,52.3997573852539,52.3620109558105,52.3316383361816,52.3062515258789,52.2865219116211,52.2145004272461,52.1729965209961,52.12646484375,52.0312995910645,51.9258308410645,51.7812995910645,51.7030258178711,51.6099128723145,51.5663108825684,51.5450668334961,51.5056648254395,51.4480476379395,51.4122543334961,51.3741683959961,51.3148918151855,51.2613754272461,51.2301292419434,51.1723136901855,51.1001472473145,50.9858894348145,50.8294448852539,50.7079582214355,50.621337890625,50.5500984191895,50.4941902160645,50.4535140991211,50.4280738830566,50.3936042785645,50.3501472473145,50.298583984375,50.208984375,50.0716781616211,49.9750480651855,49.8734359741211,49.8018035888672,49.7602043151855,49.7167472839355,49.6715316772461,49.6221199035645,49.568603515625,49.5592765808105,49.59423828125,49.6001472473145,49.5769538879395,49.5418434143066,49.4947280883789,49.4637680053711,49.4489288330078,49.4192390441895,49.3746604919434,49.362060546875,49.3813972473145,49.378662109375,49.3538589477539,49.3623542785645,49.3894538879395,49.3894538879395,49.3888168334961,49.3234367370605,49.2866706848145,49.2785186767578,49.1988754272461,48.9722633361816,48.8916511535645,48.8663558959961,48.8611831665039,48.773193359375,48.6801261901855,48.6024894714355,48.5746574401855,48.4833984375,48.4303703308105,48.388427734375,48.3415031433105,48.2545890808105,48.1276359558105,48.0192375183105,47.929443359375,47.8429222106934,47.7598152160645,47.7093238830566,47.6914558410645,47.6976547241211,47.7278327941895,47.7226066589355,47.6820335388184,47.6805191040039,47.7179679870605,47.7294921875,47.7149925231934,47.7685050964355,47.8900871276855,47.947265625,47.9400863647461,47.9790992736816,48.0644035339355,48.1056632995605,48.1015167236328,48.09716796875,48.1330108642578,48.2077140808105,48.2737312316895,48.3599128723145,48.3734397888184,48.3688468933105,48.3553199768066,48.3217277526855,48.25390625,48.21044921875,48.1533203125,48.1201629638672,48.0829086303711,48.0225105285645,47.9751930236816,47.874267578125,47.8014183044434,47.7154273986816,47.6844749450684,47.6248550415039,47.5238761901855,47.4851570129395,47.4473648071289,47.438232421875,47.4294929504395,47.41357421875,47.3777351379395,47.3526344299316,47.3022003173828,47.2587432861328,47.1942367553711,47.1280746459961,47.0689964294434,46.9781265258789,46.9507827758789,46.8819808959961,46.858154296875,46.7131843566895,46.6142578125,46.4991226196289,46.4305686950684,46.366943359375,46.3360328674316,46.30908203125,46.2477531433105,46.2242698669434,46.1859359741211,46.1397438049316,46.0696296691895,46.0089378356934,45.9552230834961,45.9203109741211,45.8978004455566,45.8788070678711,45.8104476928711,45.7576675415039,45.705078125,45.6512222290039,45.6046867370605,45.5722160339355,45.5530776977539,45.5452613830566,45.4948234558105,45.3214378356934,45.220458984375,45.1307144165039,45.0745582580566,45.0299301147461,45.0460433959961,45.0611305236816,45.08056640625,45.0937004089355,45.122802734375,45.1599617004395,45.2032699584961,45.2259788513184,45.2439918518066,45.2737312316895,45.3268547058105,45.3052749633789,45.2426261901855,45.2053718566895,45.1365699768066,45.0836448669434,45.0131340026855,44.9840316772461,44.9361305236816,44.920166015625,44.9100112915039,44.8888664245605,44.8443336486816,44.7999534606934,44.7532234191895,44.65966796875,44.5956535339355,44.4691886901855,44.0715789794922,44.0029296875,43.7047386169434,43.65087890625,43.5670890808105,43.5055656433105,43.4904289245605,43.4690399169922,43.4330558776855,43.3780784606934,43.337646484375,43.2577629089355,43.1421852111816,43.0976104736328,43.0624504089355,43.0380859375,42.956298828125,42.9022445678711,42.8831024169922,42.8517570495605,42.8633308410645,42.8568344116211,42.8358383178711,42.8116226196289,42.7790985107422,42.7554206848145,42.72705078125,42.6998519897461,42.685546875,42.67431640625,42.6232414245605,42.5673332214355,42.535400390625],[5.12831974029541,5.08867216110229,5.11406278610229,5.147216796875,5.12831974029541],[10.426025390625,10.3595695495605,10.3140134811401,10.3196773529053,10.2926273345947,10.2416009902954,10.1283197402954,10.0464839935303,9.93007850646973,9.86894512176514,9.84116172790527,9.75654315948486,9.71357440948486,9.72348594665527,9.67924785614014,9.64799785614014,9.64570331573486,9.74326229095459,9.78173828125,9.85962009429932,9.89492225646973,9.91718673706055,9.92431735992432,9.90029239654541,9.88222694396973,9.89545917510986,9.84916973114014,9.75209999084473,9.72089862823486,9.68735313415527,9.61074256896973,9.53461933135986,9.50092792510986,9.42582988739014,9.42470741271973,9.45712947845459,9.48134803771973,9.43173789978027,9.35136699676514,9.30166053771973,9.28261756896973,9.21860313415527,9.10962009429932,9.04511737823486,9.02509784698486,8.95659160614014,8.83960056304932,8.80043888092041,8.7763671875,8.49301719665527,8.208740234375,8.171630859375,8.16079139709473,8.14755821228027,8.12109279632568,8.08222579956055,8.04667949676514,8.022216796875,7.93193387985229,7.89599561691284,7.81904268264771,7.77207040786743,7.68500947952271,7.45454120635986,7.26362371444702,7.20488309860229,7.16376972198486,7.10458946228027,6.94096660614014,6.80722713470459,6.74912071228027,6.69077157974243,6.64868211746216,6.53564500808716,6.44106435775757,6.34824228286743,6.08564424514771,5.92627000808716,5.79775333404541,5.71132802963257,5.67626905441284,5.64301776885986,5.61918973922729,5.60009765625,5.43251943588257,5.35693359375,5.32822275161743,5.26411104202271,5.18452167510986,5.15302705764771,5.14902353286743,5.11884784698486,5.13081073760986,5.15053701400757,5.15771484375,5.20302772521973,5.34829139709473,5.3544921875,5.33540010452271,5.16079139709473,5.13066387176514,5.220703125,5.29316377639771,5.30971670150757,5.30141544342041,5.27988243103027,5.23588895797729,5.26162099838257,5.25664091110229,5.23012733459473,5.17255878448486,5.13833045959473,5.14775371551514,5.20361280441284,5.21025419235229,5.19199228286743,5.15078115463257,5.15971708297729,5.16215848922729,5.13066387176514,5.08945274353027,5.01093769073486,4.95283222198486,4.76176738739014,4.67148399353027,4.63832998275757,4.54472637176514,4.48598623275757,4.37602519989014,4.351318359375,4.38642549514771,4.57231426239014,4.821533203125,4.916748046875,5.00644493103027,5.08066415786743,5.10849618911743,5.13979482650757,5.23642539978027,5.32451152801514,5.47788095474243,5.50991249084473,5.55058622360229,5.65131807327271,5.84130907058716,5.85371112823486,5.84550809860229,5.842041015625,5.90771484375,5.91904306411743,5.97509765625,6.03891611099243,6.07636690139771,6.15014696121216,6.23486328125,6.2861328125,6.29838848114014,6.28754901885986,6.29072237014771,6.31904363632202,6.35127019882202,6.41318368911743,6.46250009536743,6.45639705657959,6.46806669235229,6.49052715301514,6.50781297683716,6.70512676239014,6.80156278610229,6.86039972305298,6.92001962661743,6.98095750808716,7.07402372360229,7.41181659698486,7.51640653610229,7.54702091217041,7.55849552154541,7.60185527801514,7.590576171875,7.55673789978027,7.59023475646973,7.76074171066284,7.82402372360229,7.86772489547729,7.98442363739014,8.02973651885986,8.07851505279541,8.14492225646973,8.16972637176514,8.16513729095459,8.18144607543945,8.21967792510986,8.25371074676514,8.40791034698486,8.45566368103027,8.48325157165527,8.49067401885986,8.49531269073486,8.47773456573486,8.46753025054932,8.46767616271973,8.42197227478027,8.375244140625,8.37558650970459,8.41035175323486,8.5625,8.64301681518555,8.72060585021973,8.78681659698486,8.87944316864014,8.97978591918945,9.01708984375,9.08085918426514,9.11503887176514,9.151611328125,9.18852519989014,9.30869197845459,9.41586875915527,9.40385723114014,9.39765644073486,9.4306640625,9.49570274353027,9.6748046875,9.88173866271973,9.97319316864014,10.0220699310303,10.0670890808105,10.1252927780151,10.1625003814697,10.1634769439697,10.1857423782349,10.236572265625,10.34228515625,10.4274415969849,10.4212398529053,10.4368171691895,10.4397945404053,10.3839349746704,10.34130859375,10.3401365280151,10.2593746185303,10.2256832122803,10.2035150527954,10.144775390625,10.1432609558105,10.1556644439697,10.1762208938599,10.19873046875,10.2518548965454,10.299560546875,10.3423337936401,10.3450679779053,10.3569822311401,10.3571290969849,10.3494625091553,10.3921871185303,10.5120124816895,10.5780277252197,10.6337890625,10.6564455032349,10.58642578125,10.5612306594849,10.5591306686401,10.5723638534546,10.6487302780151,10.6717767715454,10.6851081848145,10.6928215026855,10.7240724563599,10.7179203033447,10.5975093841553,10.55810546875,10.4762697219849,10.4002933502197,10.3694334030151,10.3223628997803,10.2791986465454,10.2616205215454,10.2321290969849,10.201904296875,10.19482421875,10.2391119003296,10.2750978469849,10.3072271347046,10.3547353744507,10.3895511627197,10.4332036972046,10.43994140625,10.426025390625],[7.52377939224243,7.358154296875,7.26357412338257,7.20615196228027,7.13491201400757,7.06357479095459,6.90991258621216,6.78442335128784,6.74531269073486,6.55571269989014,6.36572265625,6.31635713577271,6.27978515625,6.250244140625,6.19121122360229,6.16191387176514,6.12680673599243,6.08671855926514,6.03872108459473,5.970703125,5.91689443588257,5.91357421875,5.916015625,5.88398408889771,5.86513662338257,5.49550771713257,5.43964910507202,5.41474628448486,5.35107421875,5.27993154525757,5.251708984375,5.22880840301514,5.17905235290527,5.06552743911743,4.66557645797729,4.60239219665527,4.55810546875,4.47187471389771,4.35854482650757,4.28486347198486,4.16396522521973,4.06914091110229,4.03691434860229,4.02446317672729,4.02294921875,4.016357421875,3.94721674919128,3.826904296875,3.70214867591858,3.56713843345642,3.45683598518372,3.32929682731628,3.22968745231628,3.12719750404358,3.10307645797729,3.09584951400757,3.07578134536743,3.02871108055115,2.97666001319885,2.90859389305115,2.839111328125,2.77299833297729,2.67817378044128,2.67001938819885,2.63266611099243,2.59921908378601,2.47348618507385,2.36376953125,2.27006840705872,2.262451171875,2.20478510856628,2.16782236099243,2.10678696632385,2.02167987823486,1.91806614398956,1.79594707489014,1.72421848773956,1.71411156654358,1.69125962257385,1.67622077465057,1.76001012325287,1.81660175323486,1.91498994827271,1.95039081573486,1.95673859119415,1.94472646713257,1.98173844814301,2.02446269989014,2.03559589385986,2.00234365463257,2.00087881088257,2.01376962661743,2.01230454444885,2.06933617591858,2.08046889305115,2.0751953125,2.12241220474243,2.11713862419128,2.13208031654358,2.19912099838257,2.15473628044128,2.16035175323486,2.15888667106628,2.15742206573486,2.15952134132385,2.16157221794128,2.22421860694885,2.25644540786743,2.25942349433899,2.24677729606628,2.25678730010986,2.26503896713257,2.28134775161743,2.29599595069885,2.28437542915344,2.28750014305115,2.28515625,2.30219721794128,2.29970693588257,2.26142621040344,2.23383808135986,2.16743183135986,2.16748046875,2.16757845878601,2.16762709617615,2.16772437095642,2.16777300834656,2.21328163146973,2.24238276481628,2.27548813819885,2.29780268669128,2.30444312095642,2.53925800323486,2.73496079444885,2.91655278205872,3.07905292510986,3.23964834213257,3.30976581573486,3.53759789466858,3.62382817268372,3.61176776885986,3.69648432731628,3.79804658889771,3.81430673599243,3.87001943588257,3.85292983055115,3.88012671470642,3.96533203125,4.00834989547729,4.056396484375,4.07665967941284,4.02690458297729,4.02841806411743,4.06064462661743,4.06611299514771,4.00073194503784,3.94252943992615,3.92231416702271,3.92573285102844,3.94038105010986,3.97294902801514,3.99785137176514,4.04106426239014,4.09160184860229,4.23042011260986,4.29023456573486,4.35781288146973,4.43515634536743,4.55375957489014,4.57275390625,4.57924842834473,4.57343769073486,4.58002901077271,4.64570331573486,4.67099618911743,4.55024385452271,4.516357421875,4.52622032165527,4.57187509536743,4.60585975646973,4.75209951400757,4.75522470474243,4.83281278610229,4.92700242996216,5.04687547683716,5.19746112823486,5.46376991271973,5.62968730926514,5.781005859375,5.917724609375,6.00908231735229,6.18613290786743,6.31962871551514,6.37338876724243,6.41865253448486,6.47041034698486,6.531982421875,6.65000009536743,6.76015567779541,6.783935546875,6.80327177047729,6.92138671875,6.99643611907959,6.95917987823486,6.91279315948486,6.8916015625,6.87675809860229,6.87773466110229,6.89125967025757,6.93046855926514,7.03745079040527,7.05771493911743,7.06308603286743,6.98828172683716,6.88178682327271,6.77656269073486,6.73911094665527,6.69789981842041,6.50551748275757,6.45771503448486,6.43793964385986,6.45053672790527,6.48466825485229,6.533935546875,6.597412109375,6.65502882003784,6.69726610183716,6.85463809967041,6.88886690139771,6.95156240463257,7.05620098114014,7.11640596389771,7.13798809051514,7.20195293426514,7.27226543426514,7.34506845474243,7.40073299407959,7.58974647521973,7.65200185775757,7.72778272628784,7.94248056411743,8.22739219665527,8.28232383728027,8.41972637176514,8.59555625915527,8.62412166595459,8.66777420043945,8.74565410614014,8.81787109375,8.88662052154541,9.01943397521973,9.17075252532959,9.30351638793945,9.42626953125,9.48832988739014,9.53964805603027,9.56377029418945,9.64516544342041,9.81401348114014,9.91591835021973,9.96005821228027,10.0361814498901,10.1714353561401,10.3832521438599,10.6050786972046,10.8731441497803,11.1400880813599,11.2118644714355,11.2450189590454,11.24853515625,11.2681636810303,11.4011716842651,11.4461421966553,11.4922847747803,11.5324220657349,11.5911626815796,11.7287111282349,11.829833984375,11.9866209030151,12.1086912155151,12.1509761810303,12.2094240188599,12.2220706939697,12.2982425689697,12.3441410064697,12.3564939498901,12.3837890625,12.4472169876099,12.4840822219849,12.5240726470947,12.61279296875,13.0785150527954,13.0773439407349,13.0217771530151,12.979736328125,12.8202142715454,12.7299318313599,12.6556148529053,12.5020990371704,12.2693843841553,12.13037109375,12.1083498001099,11.9071283340454,11.8455076217651,11.724365234375,11.642578125,11.5412588119507,11.3685541152954,11.2625007629395,11.1136722564697,10.8510732650757,10.6484870910645,10.4845209121704,10.3573732376099,10.2168951034546,10.0884761810303,10.0078125,9.96030235290527,9.95429706573486,9.98149394989014,9.98286151885986,9.96596622467041,9.94169902801514,9.95307636260986,9.98505878448486,9.979736328125,9.90180587768555,9.78437519073486,9.69155216217041,9.58872032165527,9.53173828125,9.406494140625,9.28505802154541,9.20351600646973,9.02524471282959,8.86567306518555,8.83915996551514,8.810302734375,8.79863262176514,8.707275390625,8.55732345581055,8.32236385345459,8.08383750915527,7.85185575485229,7.812744140625,7.78788995742798,7.73803758621216,7.66450166702271,7.60439443588257,7.52377939224243],[5.10917997360229,5.03920888900757,4.96757793426514,4.84599637985229,4.7783203125,4.70322275161743,4.64467763900757,4.59775400161743,4.56791973114014,4.53208017349243,4.47939443588257,4.4248046875,4.35024404525757,4.34853553771973,4.33803701400757,4.31865215301514,4.32416963577271,4.37285184860229,4.45449209213257,4.50498008728027,4.487060546875,4.44594717025757,4.38818359375,4.39189434051514,4.49838876724243,4.61181640625,4.63603544235229,4.5869140625,4.48095655441284,4.3271484375,4.26850557327271,4.17763710021973,3.98193359375,3.88388681411743,3.835693359375,3.78720664978027,3.72294902801514,3.65278291702271,3.62421870231628,3.63413095474243,3.64409160614014,3.62421870231628,3.57314467430115,3.53334927558899,3.49072241783142,3.46367192268372,3.40893578529358,3.34213829040527,3.28261733055115,3.16337895393372,3.04179668426514,3.00136709213257,2.96757817268372,2.93349647521973,2.89365243911743,2.84702157974243,2.677978515625,2.53027319908142,2.45537114143372,2.40043926239014,2.40327143669128,2.36938500404358,2.31552767753601,2.2880859375,2.28886723518372,2.27006840705872,2.23227572441101,2.19135737419128,2.14628887176514,2.08847641944885,2.04457974433899,1.92202115058899,1.68281257152557,1.23906242847443,1.23881840705872,1.18530297279358,1.10278332233429,0.973486244678497,0.863525331020355,0.819238543510437,0.792871057987213,0.673925578594208,0.49902331829071,0.418945580720901,0.263623058795929,0.166357159614563,0.147216930985451,0.0983396098017693,-0.0602052547037601,-0.113573983311653,-0.441699475049973,-0.535253763198853,-0.691308498382568,-0.783105492591858,-0.887109398841858,-0.977343916893005,-1.12138664722443,-1.35673832893372,-1.38789057731628,-1.40976560115814,-1.46806621551514,-1.50742161273956,-1.51757788658142,-1.62158226966858,-1.71992194652557,-1.81601560115814,-1.86025404930115,-1.98457062244415,-2.13183617591858,-2.19511723518372,-2.23320341110229,-2.31279277801514,-2.37031245231628,-2.40029335021973,-2.44667983055115,-2.55556654930115,-2.63505864143372,-2.68203163146973,-2.72021508216858,-2.75830078125,-2.79960942268372,-2.85078144073486,-2.95527315139771,-3.05351543426514,-3.138671875,-3.28125023841858,-3.36328125,-3.47568345069885,-3.68496084213257,-3.83378911018372,-3.91083955764771,-4.09540987014771,-4.29970693588257,-4.44931650161743,-4.49667978286743,-4.66884756088257,-4.83564472198486,-4.89882802963257,-4.98310565948486,-5.17617177963257,-5.31660127639771,-5.40097618103027,-5.49980449676514,-5.65078163146973,-5.72265625,-5.77597665786743,-5.96542978286743,-6.02500009536743,-6.17207050323486,-6.31386709213257,-6.39443397521973,-6.61689472198486,-6.69189453125,-6.80312490463257,-6.9150390625,-6.97304677963257,-7.03789043426514,-7.20371103286743,-7.33867216110229,-7.46064472198486,-7.62714862823486,-7.78193330764771,-7.97089815139771,-8.10439395904541,-8.19365215301514,-8.22002029418945,-8.25820255279541,-8.30029296875,-8.34375,-8.38691425323486,-8.42783164978027,-8.46494102478027,-8.48544979095459,-8.59023380279541,-8.70058631896973,-8.78584003448486,-8.89101600646973,-8.9326171875,-9.0146484375,-9.07226657867432,-9.16943454742432,-9.22480487823486,-9.27499961853027,-9.51005840301514,-9.67880821228027,-9.83124923706055,-9.91875076293945,-10.0988283157349,-10.31298828125,-10.3973627090454,-10.5501956939697,-10.6692390441895,-10.8023433685303,-10.9332036972046,-11.1095695495605,-11.3543949127197,-11.4830083847046,-11.5666990280151,-11.6228513717651,-11.6983394622803,-11.8121099472046,-11.8791990280151,-11.9081048965454,-12.05126953125,-12.1205081939697,-12.2578125,-12.3488283157349,-12.3702154159546,-12.4047842025757,-12.4312496185303,-12.41845703125,-12.3861331939697,-12.3175773620605,-12.2668943405151,-12.2282228469849,-12.2024412155151,-12.1983394622803,-12.1640625,-12.1554679870605,-12.30615234375,-12.4505863189697,-12.6258783340454,-12.8270511627197,-12.9920892715454,-13.16748046875,-13.3697261810303,-13.3927736282349,-13.4380855560303,-13.4538087844849,-13.4143562316895,-13.3729486465454,-13.2985343933105,-13.2605466842651,-13.2489261627197,-13.2679681777954,-13.3228521347046,-13.3708009719849,-13.3983402252197,-13.3951168060303,-13.3688478469849,-13.30712890625,-13.2146482467651,-13.1194334030151,-12.9819326400757,-12.9254884719849,-12.861328125,-12.8541011810303,-12.8361320495605,-12.7421875,-12.6233396530151,-12.5774412155151,-12.5180654525757,-12.4820308685303,-12.4345703125,-12.3681650161743,-12.2848625183105,-12.2808589935303,-12.2667970657349,-12.22705078125,-12.1953134536743,-12.0796871185303,-11.9445314407349,-11.7834959030151,-11.6050777435303,-11.5791988372803,-11.59375,-11.6159172058105,-11.6637697219849,-11.8246097564697,-11.8988285064697,-11.9193353652954,-11.9435548782349,-11.9652347564697,-11.9759769439697,-11.9720706939697,-11.9478521347046,-11.9298820495605,-11.9032220840454,-11.89013671875,-11.8552742004395,-11.8201169967651,-11.744140625,-11.75341796875,-11.6998043060303,-11.6735353469849,-11.623046875,-11.5535163879395,-11.4049806594849,-11.3254880905151,-11.2369146347046,-11.21240234375,-11.21240234375,-11.2429685592651,-11.2600584030151,-11.2991209030151,-11.3211917877197,-11.3377933502197,-11.3529291152954,-11.4384765625,-11.4476556777954,-11.4170894622803,-11.3712892532349,-11.3193359375,-11.2551755905151,-11.1298828125,-11.07177734375,-11.0299816131592,-11.0259761810303,-10.9556646347046,-10.8915033340454,-10.8791017532349,-10.8717775344849,-10.8915033340454,-10.9434576034546,-10.9832029342651,-11.0136728286743,-11.0076169967651,-10.9786128997803,-10.9693355560303,-10.9764652252197,-11.0748052597046,-11.087890625,-11.0802736282349,-11.0597648620605,-11.0558595657349,-11.0867185592651,-11.1594734191895,-11.1986322402954,-11.1941404342651,-11.1636714935303,-11.1219730377197,-11.0126943588257,-10.8922853469849,-10.8294916152954,-10.7839841842651,-10.6913089752197,-10.5515632629395,-10.4533205032349,-10.3966789245605,-10.2590827941895,-10.0406255722046,-9.86279296875,-9.7255859375,-9.59423828125,-9.46875,-9.16845703125,-8.90351486206055,-8.693359375,-8.34111309051514,-8.11191463470459,-7.86542987823486,-7.60165977478027,-7.42099618911743,-7.32861280441284,-7.31464862823486,-7.30546855926514,-7.29667997360229,-7.28496122360229,-7.28144502639771,-7.27773475646973,-7.24443340301514,-7.18281316757202,-7.12177705764771,-6.93515634536743,-6.91992139816284,-6.91582012176514,-6.94628858566284,-6.97646474838257,-6.986328125,-7.037109375,-7.14443397521973,-7.27949285507202,-7.39072227478027,-7.47216796875,-7.55732440948486,-7.65507793426514,-7.70654344558716,-7.96660184860229,-8.00146484375,-8.00146484375,-7.99814462661743,-7.93603515625,-7.93593788146973,-7.96855449676514,-8.00029277801514,-8.02382755279541,-8.10078144073486,-8.10761737823486,-8.06767559051514,-8.07138633728027,-8.09072303771973,-8.09902381896973,-8.07587909698486,-7.88193321228027,-7.62333965301514,-7.46132755279541,-7.41904306411743,-7.36308574676514,-7.25742197036743,-7.15703105926514,-7.06210947036743,-6.93398475646973,-6.77255916595459,-6.61845731735229,-6.4716796875,-6.34599637985229,-6.24140644073486,-6.16425752639771,-6.11455059051514,-6.05156230926514,-6.02529287338257,-5.96582078933716,-5.90019464492798,-5.86562538146973,-5.86494159698486,-5.86386728286743,-5.86884737014771,-5.87451171875,-5.88007783889771,-5.88886690139771,-5.89267587661743,-5.8759765625,-5.86513662338257,-5.85722637176514,-5.85517597198486,-5.86171865463257,-5.86181640625,-5.86337900161743,-5.8818359375,-5.85624980926514,-5.86484336853027,-5.83613300323486,-5.85410165786743,-5.87773466110229,-5.96083974838257,-6.00419950485229,-6.00048875808716,-5.986328125,-5.89531230926514,-5.80732440948486,-5.75869178771973,-5.74648380279541,-5.72773408889771,-5.71875,-5.69580125808716,-5.42460918426514,-5.14892578125,-5.11269521713257,-5.09062528610229,-5.07148408889771,-5.03691339492798,-4.99658203125,-4.97841835021973,-4.90537118911743,-4.73662090301514,-4.6953125,-4.65107393264771,-4.634765625,-4.60195302963257,-4.60429668426514,-4.62031269073486,-4.65585947036743,-4.70585966110229,-4.76523447036743,-4.82939481735229,-4.83740234375,-4.80498027801514,-4.75673818588257,-4.72148418426514,-4.68867158889771,-4.61835956573486,-4.54326200485229,-4.45449209213257,-4.44248056411743,-4.43388652801514,-4.45888662338257,-4.48466825485229,-4.48466825485229,-4.46123027801514,-4.41748046875,-4.40000009536743,-4.35810565948486,-4.30410146713257,-4.29941415786743,-4.36972665786743,-4.41904306411743,-4.44951200485229,-4.50810527801514,-4.58554697036743,-4.681640625,-4.77499961853027,-4.83125019073486,-4.85410165786743,-4.86494159698486,-4.85166025161743,-4.85576152801514,-4.88505887985229,-4.88173866271973,-4.845703125,-4.70556640625,-4.46103525161743,-4.3076171875,-4.24492168426514,-4.17177772521973,-4.08798837661743,-4.03095674514771,-3.98554730415344,-3.93427729606628,-3.76621079444885,-3.46416020393372,-3.19443345069885,-3.03027367591858,-2.46474599838257,-2.27910184860229,-2.17783212661743,-2.10820341110229,-1.96083974838257,-1.84013640880585,-1.69892585277557,-1.37636721134186,-1.27246069908142,-1.22587871551514,-1.13994133472443,-1.064453125,-0.999609053134918,-0.774999916553497,-0.549023628234863,-0.277539014816284,-0.0523924082517624,0.234130620956421,0.537304937839508,0.856884717941284,1.11806643009186,1.42211902141571,1.53486335277557,1.71938502788544,2.01328158378601,2.41494154930115,2.65541982650757,2.92441368103027,3.08701181411743,3.30405282974243,3.47841811180115,3.6787109375,3.95429706573486,4.11660146713257,4.257568359375,4.34624004364014,4.38261747360229,4.5234375,4.89140605926514,5.07075214385986,5.12749052047729,5.12138700485229,5.08930683135986,5.03129863739014,4.94472646713257,4.82963848114014,4.68618154525757,4.54155302047729,4.46269512176514,4.43564414978027,4.44731473922729,4.41313457489014,4.33217811584473,4.30219745635986,4.32309579849243,4.31137657165527,4.24482440948486,4.28139686584473,4.25390625,4.13496065139771,4.15512704849243,4.15976572036743,4.20766639709473,4.445556640625,4.59174776077271,4.64667987823486,4.723876953125,4.74384784698486,4.73691415786743,4.70297861099243,4.66352510452271,4.66313505172729,4.70126962661743,4.77080059051514,4.81635713577271,4.86626005172729,4.953857421875,4.994140625,5.00996065139771,4.93007850646973,4.98295879364014,4.96743202209473,5.02456092834473,5.06269502639771,5.25590801239014,5.31210947036743,5.28369140625,5.25371074676514,5.17114210128784,5.085205078125,5.07192373275757,5.06240272521973,5.07568359375,5.18437480926514,5.19975614547729,5.19785118103027,5.10917997360229],[3.47841811180115,3.30405282974243,3.08701181411743,2.92441368103027,2.65541982650757,2.41494154930115,2.01328158378601,1.71938502788544,1.53486335277557,1.42211902141571,1.11806643009186,0.856884717941284,0.537304937839508,0.234130620956421,-0.0523924082517624,-0.277539014816284,-0.549023628234863,-0.774999916553497,-0.999609053134918,-1.064453125,-1.13994133472443,-1.22587871551514,-1.27246069908142,-1.37636721134186,-1.69892585277557,-1.84013640880585,-1.96083974838257,-2.10820341110229,-2.17783212661743,-2.27910184860229,-2.46474599838257,-3.03027367591858,-3.19443345069885,-3.46416020393372,-3.76621079444885,-3.93427729606628,-3.98554730415344,-4.03095674514771,-4.08798837661743,-4.17177772521973,-4.24492168426514,-4.3076171875,-4.46103525161743,-4.70556640625,-4.845703125,-4.88173866271973,-4.88505887985229,-4.85576152801514,-4.85166025161743,-4.86494159698486,-4.85410165786743,-4.83125019073486,-4.77499961853027,-4.681640625,-4.58554697036743,-4.50810527801514,-4.44951200485229,-4.41904306411743,-4.36972665786743,-4.29941415786743,-4.30410146713257,-4.35810565948486,-4.40000009536743,-4.41748046875,-4.46123027801514,-4.48466825485229,-4.48466825485229,-4.45888662338257,-4.43388652801514,-4.44248056411743,-4.45449209213257,-4.54326200485229,-4.61835956573486,-4.68867158889771,-4.72148418426514,-4.75673818588257,-4.80498027801514,-4.83740234375,-4.82939481735229,-4.76523447036743,-4.70585966110229,-4.65585947036743,-4.62031269073486,-4.60429668426514,-4.60195302963257,-4.634765625,-4.61923837661743,-4.5517578125,-4.44511699676514,-4.42890644073486,-4.43056631088257,-4.46972703933716,-4.53115272521973,-4.58750009536743,-4.619140625,-4.65771484375,-4.72412109375,-4.76552724838257,-4.77861356735229,-4.83769512176514,-4.95214891433716,-5.00429677963257,-4.98203134536743,-4.95439481735229,-4.86572265625,-4.75546884536743,-4.70517587661743,-4.67656230926514,-4.56582069396973,-4.43427753448486,-4.20029258728027,-4.13056612014771,-3.91630887985229,-3.76201176643372,-3.69082045555115,-3.64111375808716,-3.52031230926514,-3.52500033378601,-3.68203115463257,-3.69453096389771,-3.69023418426514,-3.69667935371399,-3.66591787338257,-3.62539052963257,-3.580078125,-3.53144526481628,-3.47861361503601,-3.42021489143372,-3.35097646713257,-3.31855487823486,-3.28320288658142,-3.22910118103027,-3.17695307731628,-3.126953125,-3.06308555603027,-3.01123070716858,-2.98310542106628,-2.93652319908142,-2.88662075996399,-2.85537123680115,-2.83671855926514,-2.76962876319885,-2.67099618911743,-2.59540987014771,-2.39707016944885,-2.36093735694885,-2.34257817268372,-2.36455106735229,-2.39472651481628,-2.35146474838257,-2.34482407569885,-2.38281226158142,-2.41259741783142,-2.32998061180115,-2.24560546875,-2.16923809051514,-2.11201190948486,-2.07529330253601,-2.04755854606628,-1.99033224582672,-1.92890644073486,-1.90000021457672,-1.82685530185699,-1.82958996295929,-1.86943340301514,-1.93183600902557,-2.06328129768372,-2.17626976966858,-2.31337881088257,-2.36914038658142,-2.40478515625,-2.39540982246399,-2.27861309051514,-2.18750023841858,-2.13847684860229,-2.16376924514771,-2.28369116783142,-2.33017587661743,-2.37451195716858,-2.42988324165344,-2.46543002128601,-2.49062538146973,-2.46689438819885,-2.41796898841858,-2.35419940948486,-2.30058598518372,-2.26552724838257,-2.21757817268372,-2.17988252639771,-2.07675766944885,-2.00146484375,-1.95351564884186,-1.92021489143372,-1.89003884792328,-1.71152353286743,-1.64697289466858,-1.59335958957672,-1.52509784698486,-1.41318368911743,-1.22978520393372,-1.10390627384186,-0.972070515155792,-0.798828065395355,-0.618359565734863,-0.573437333106995,-0.518652260303497,-0.468554556369781,-0.427344024181366,-0.361913949251175,-0.292382657527924,-0.270116925239563,-0.24257829785347,-0.203320384025574,-0.0908201113343239,0.0752929598093033,0.190820157527924,0.283984243869781,0.35380831360817,0.427734196186066,0.514990508556366,0.536572098731995,0.551122963428497,0.587451338768005,0.624218642711639,0.673828125,0.755712985992432,0.81147438287735,0.849121153354645,0.901464879512787,1.00444316864014,1.09023439884186,1.12084984779358,1.32255840301514,1.37021481990814,1.39589822292328,1.41875004768372,1.38227558135986,1.31459951400757,1.26777338981628,1.24101567268372,1.24843728542328,1.27924799919128,1.30542016029358,1.36669921875,1.45458972454071,1.53505873680115,1.64809596538544,1.78857421875,1.92041015625,2.09169936180115,2.16157221794128,2.15952134132385,2.15742206573486,2.15888667106628,2.16035175323486,2.15473628044128,2.19912099838257,2.13208031654358,2.11713862419128,2.12241220474243,2.0751953125,2.08046889305115,2.06933617591858,2.01230454444885,2.01376962661743,2.00087881088257,2.00234365463257,2.03559589385986,2.02446269989014,1.98173844814301,1.94472646713257,1.95673859119415,1.95039081573486,1.91498994827271,1.81660175323486,1.76001012325287,1.67622077465057,1.69125962257385,1.71411156654358,1.72421848773956,1.79594707489014,1.91806614398956,2.02167987823486,2.10678696632385,2.16782236099243,2.20478510856628,2.262451171875,2.27006840705872,2.40678715705872,2.54277348518372,2.70102572441101,2.83173847198486,2.89653325080872,2.99321269989014,3.10097646713257,3.16513705253601,3.20883798599243,3.36953091621399,3.46308588981628,3.50537133216858,3.53520512580872,3.53627943992615,3.55668926239014,3.59843730926514,3.6171875,3.68461942672729,3.68730497360229,3.66162085533142,3.58417987823486,3.55385708808899,3.55839848518372,3.5517578125,3.55083036422729,3.56030297279358,3.55107450485229,3.49980425834656,3.50541996955872,3.54267597198486,3.58081030845642,3.62299776077271,3.60410165786743,3.51020550727844,3.47841811180115],[-21.2492179870605,-21.24951171875,-21.2420902252197,-21.2380847930908,-21.2291030883789,-21.2004890441895,-21.1864261627197,-21.1884765625,-21.2081069946289,-21.2406253814697,-21.2492179870605],[2.541748046875,2.51967740058899,2.55927753448486,2.60917997360229,2.64633798599243,2.63417983055115,2.60361313819885,2.541748046875],[11.8619146347046,11.8497562408447,11.8235359191895,11.7740726470947,11.7268562316895,11.6664066314697,11.6019535064697,11.1964349746704,11.1142578125,11.0539064407349,10.9771480560303,10.8358402252197,10.7271966934204,10.4912605285645,10.1957521438599,10.0297355651855,9.78916072845459,9.668212890625,9.55463790893555,9.443603515625,9.322021484375,9.22685527801514,9.19414043426514,9.16792011260986,9.19414043426514,9.22280311584473,9.25957012176514,9.23994159698486,9.13515567779541,9.1220703125,9.13515567779541,9.10898399353027,8.84829044342041,8.62758827209473,8.48969745635986,8.38198280334473,8.28706073760986,8.15278339385986,8.08730411529541,8.04770565032959,7.96611356735229,7.80986356735229,7.75795936584473,7.61323213577271,7.52426719665527,7.45488262176514,7.41508817672729,7.39453172683716,7.37026405334473,7.24970722198486,7.09687519073486,7.03261661529541,6.99033212661743,7.00581026077271,7.03291034698486,7.0263671875,6.98520517349243,6.98671913146973,6.99443387985229,7.07758760452271,7.09003925323486,7.08276414871216,7.04052686691284,7.00712919235229,6.97260761260986,6.94794940948486,6.95205068588257,6.95361375808716,6.93793964385986,6.70024394989014,6.49438524246216,6.32148456573486,6.13491201400757,6.12397432327271,6.14799785614014,6.13759756088257,6.09970664978027,6.11533212661743,6.18437528610229,6.19819307327271,6.15678739547729,6.15654325485229,6.197509765625,6.24194288253784,6.28989315032959,6.28496074676514,6.24179697036743,6.18027353286743,6.11977529525757,6.02553701400757,5.92998027801514,5.83310556411743,5.70937490463257,5.55878925323486,5.447509765625,5.37548828125,5.27045869827271,5.13251924514771,4.93081045150757,4.66547870635986,4.50688505172729,4.455078125,4.38071250915527,4.28388690948486,4.1982421875,4.08652353286743,3.86425757408142,3.768798828125,3.73383784294128,3.69111323356628,3.46376919746399,3.41586899757385,3.37397503852844,3.34262704849243,3.32265615463257,3.187255859375,2.89282202720642,2.85532188415527,2.79360342025757,2.83330106735229,2.80019545555115,2.79360342025757,2.76933574676514,2.68994116783142,2.67675805091858,2.64365243911743,2.55991172790527,2.47167992591858,2.42944359779358,2.39013695716858,2.33276391029358,2.27548813819885,2.142578125,2.10126948356628,2.05058598518372,1.99985361099243,1.94033193588257,1.82319355010986,1.68017590045929,1.60078108310699,1.56420910358429,1.45800805091858,1.35825204849243,1.28989243507385,1.22304654121399,1.17836904525757,1.18540036678314,1.21000957489014,1.40058624744415,1.61557602882385,1.70361316204071,1.84482443332672,2.03208017349243,2.08583974838257,2.11669921875,2.12114262580872,2.10790991783142,2.072998046875,2.03505849838257,1.922119140625,1.79008793830872,1.76059556007385,1.74848616123199,1.75253927707672,1.78803682327271,1.860107421875,1.95576167106628,1.98701190948486,1.95761740207672,1.90136730670929,1.84550797939301,1.77456033229828,1.71982443332672,1.7216796875,1.72158169746399,1.72148442268372,1.72138690948486,1.72128903865814,1.72124016284943,1.72578096389771,1.75791013240814,1.77324199676514,1.77075183391571,1.73945295810699,1.73486351966858,1.70517575740814,1.70873999595642,1.54389667510986,1.30878913402557,1.05952155590057,1.07841765880585,1.07661128044128,1.05908203125,1.0732421875,1.06577146053314,1.05947291851044,1.05859398841858,1.038818359375,1.04238271713257,1.06401360034943,1.05048835277557,1.01538097858429,0.963134586811066,0.898291230201721,0.86406272649765,0.801953017711639,0.753320336341858,0.712841987609863,0.686669886112213,0.658789157867432,0.642529189586639,0.635351777076721,0.629931569099426,0.625439584255219,0.627246081829071,0.652441382408142,0.655175566673279,0.651562452316284,0.666894793510437,0.698388576507568,0.729931592941284,0.716406404972076,0.700195074081421,0.680371344089508,0.65966808795929,0.665087699890137,0.649804592132568,0.626367211341858,0.607470452785492,0.598486542701721,0.589404582977295,0.585839629173279,0.578613460063934,0.447363138198853,0.189355686306953,0.0185546651482582,-0.138867199420929,-0.196191638708115,-0.317480504512787,-0.38134753704071,-0.452539265155792,-0.482421875,-0.509277522563934,-0.553320229053497,-0.59960949420929,-0.639355599880219,-0.681250274181366,-0.720898330211639,-0.762793004512787,-0.795898497104645,-0.837793171405792,-0.877441346645355,-0.917187392711639,-0.945800960063934,-0.965722680091858,-0.998730778694153,-1.02958965301514,-1.06494116783142,-1.09160149097443,-1.15224623680115,-1.19492185115814,-1.24570298194885,-1.42167961597443,-1.62197244167328,-1.77490234375,-2.02421855926514,-2.31425786018372,-2.66767597198486,-3.01669907569885,-3.35458993911743,-3.65986299514771,-3.99658226966858,-4.20058631896973,-4.23593759536743,-4.16201162338257,-4.09218788146973,-4.05019521713257,-3.9951171875,-3.88271498680115,-3.84423828125,-3.81435537338257,-3.81874990463257,-3.849609375,-3.86933588981628,-3.86640620231628,-3.78896450996399,-3.78154325485229,-3.60458970069885,-3.28828096389771,-3.08730459213257,-2.86406254768372,-2.75019502639771,-2.73076152801514,-2.70165967941284,-2.65820336341858,-2.63984394073486,-2.60654330253601,-2.55253911018372,-2.529296875,-2.49072265625,-2.45312523841858,-2.41826176643372,-2.40576148033142,-2.34199190139771,-2.21855449676514,-2.20683574676514,-2.22578120231628,-2.24541020393372,-2.31308627128601,-2.33486318588257,-2.33408236503601,-2.29375004768372,-2.27919912338257,-2.22421860694885,-2.18212866783142,-2.15273404121399,-2.16630864143372,-2.22773432731628,-2.28867197036743,-2.32656264305115,-2.32460951805115,-2.38066434860229,-2.40048813819885,-2.40927720069885,-2.42890620231628,-2.39501976966858,-2.36513662338257,-2.35166001319885,-2.36103534698486,-2.39218759536743,-2.40546894073486,-2.40849637985229,-2.39404273033142,-2.33974599838257,-2.31201195716858,-2.27822279930115,-2.20839834213257,-2.15634799003601,-2.08105492591858,-2.00332045555115,-1.88037133216858,-1.83027362823486,-1.78769540786743,-1.77226555347443,-1.78388690948486,-1.73740231990814,-1.69306623935699,-1.63886725902557,-1.53662133216858,-1.44970703125,-1.40136706829071,-1.31640601158142,-1.24882793426514,-1.21416008472443,-1.21796846389771,-1.19667971134186,-1.12519514560699,-1.09814465045929,-1.02861332893372,-0.997753798961639,-0.970605432987213,-0.927831768989563,-0.850878953933716,-0.808398187160492,-0.766601502895355,-0.691406488418579,-0.580664098262787,-0.517675936222076,-0.470117002725601,-0.429882794618607,-0.37001958489418,-0.335253655910492,-0.29863303899765,-0.244531527161598,-0.200097486376762,-0.203320384025574,-0.199413955211639,-0.188183397054672,-0.155859425663948,-0.11669946461916,-0.050488468259573,-0.0417479984462261,-0.0417479984462261,-0.106543220579624,-0.0384277030825615,0.0628907158970833,0.0892576575279236,0.150976672768593,0.247753828763962,0.313085913658142,0.34555658698082,0.439404189586639,0.448486179113388,0.404980331659317,0.378857463598251,0.303905993700027,0.261230379343033,0.235448971390724,0.240966573357582,0.268164306879044,0.272118836641312,0.25083002448082,0.241650119423866,0.247753828763962,0.268505781888962,0.296240389347076,0.355077981948853,0.347753882408142,0.360400199890137,0.393896460533142,0.424853384494781,0.636523485183716,0.651171684265137,0.660351574420929,0.689502000808716,0.723633110523224,0.782226622104645,0.837841928005219,0.825390338897705,0.89873069524765,0.968554735183716,1.04609370231628,1.19882833957672,1.23666977882385,1.28344714641571,1.35869157314301,1.43466794490814,1.45537126064301,1.52407228946686,1.62368166446686,1.75219714641571,1.84873032569885,1.77377927303314,1.92363309860229,2.05624985694885,2.30678701400757,2.35664057731628,2.42368125915527,2.47006845474243,2.48349595069885,2.46054673194885,2.510498046875,2.48818397521973,2.50913071632385,2.54306650161743,2.56899452209473,2.62924814224243,2.69882822036743,2.72587871551514,2.71635746955872,2.74638652801514,2.78730440139771,2.87885737419128,2.91933584213257,3.007568359375,3.03994154930115,3.05117201805115,3.07597661018372,3.16025400161743,3.23378944396973,3.34179711341858,3.34858417510986,3.47475576400757,3.58535146713257,3.91328144073486,3.90605497360229,3.86225605010986,3.86743187904358,3.89321255683899,4.04096698760986,4.05849647521973,3.94472670555115,4.06044912338257,4.13095712661743,4.20078086853027,4.24775409698486,4.21279335021973,4.25629901885986,4.301025390625,4.34760713577271,4.39829111099243,4.47499990463257,4.59384775161743,4.72172832489014,4.78466796875,4.83852577209473,5.07656240463257,5.21518564224243,5.32397508621216,5.41616201400757,5.537109375,5.67563438415527,5.78017568588257,5.99536180496216,6.1767578125,6.28564405441284,6.271728515625,6.27499961853027,6.50449180603027,6.57558584213257,6.69033193588257,6.693115234375,6.83730506896973,6.86962890625,6.96040010452271,7.13725566864014,7.22934627532959,7.44282197952271,7.48369121551514,7.53696250915527,7.66806650161743,7.69882774353027,7.71186494827271,7.71093797683716,7.69121026992798,7.63461875915527,7.56455039978027,7.54306602478027,7.56625986099243,7.70585966110229,7.74907255172729,7.83652353286743,7.90815448760986,7.93251943588257,7.97246074676514,8.03388595581055,8.18706035614014,8.26953125,8.35165977478027,8.42724609375,8.49843788146973,8.56586933135986,8.64467716217041,8.65830135345459,8.63671875,8.49370098114014,8.40058517456055,8.25034141540527,8.14682579040527,8.12797927856445,8.09047889709473,8.06269550323486,8.03339767456055,7.97319412231445,7.93945264816284,7.91796875,7.93159151077271,8.00214862823486,8.310546875,8.46469783782959,8.51274394989014,8.57373046875,8.619873046875,8.64067459106445,8.69472694396973,8.98911094665527,9.265625,9.36577129364014,9.430908203125,9.41562461853027,9.450439453125,9.53847599029541,9.65781211853027,9.72978591918945,9.83427715301514,9.99272441864014,10.1258296966553,10.2051763534546,10.2364253997803,10.1721677780151,10.1434078216553,10.1963376998901,10.3277339935303,10.5276365280151,10.6108884811401,10.7271966934204,10.7832517623901,10.8704099655151,11.0575685501099,11.1097164154053,10.9890623092651,10.9966793060303,10.9746580123901,10.9671878814697,10.9344730377197,10.8625001907349,10.7870607376099,10.7652339935303,10.813720703125,10.9522457122803,11.1053228378296,11.2657222747803,11.3208494186401,11.3406248092651,11.3088865280151,11.2756834030151,11.2714834213257,11.2957515716553,11.712158203125,11.8017091751099,11.8892574310303,12.0602054595947,12.1885738372803,12.2384281158447,12.23828125,12.26953125,12.30908203125,12.4199705123901,12.4343748092651,12.4322757720947,12.3353033065796,12.1641607284546,12.0463380813599,11.9183101654053,11.8619146347046],[-12.3070316314697,-12.3682622909546,-12.3428707122803,-12.2876958847046,-12.2470703125,-12.2559566497803,-12.3070316314697],[-12.08154296875,-12.2195310592651,-12.3235349655151,-12.3565425872803,-12.3351564407349,-12.2522459030151,-12.17138671875,-12.1647462844849,-12.1730470657349,-12.1656246185303,-12.1201171875,-12.0929689407349,-12.0713872909546,-12.08154296875],[-11.9012699127197,-11.91455078125,-11.8575201034546,-11.8440427780151,-11.7518548965454,-11.43212890625,-11.3912115097046,-11.37451171875,-11.3684568405151,-11.4085941314697,-11.6141595840454,-11.7525396347046,-11.8621091842651,-11.9012699127197],[14.8562984466553,14.8182134628296,14.8348140716553,14.8742189407349,14.9312496185303,14.9802732467651,15.0382823944092,15.0194816589355,14.9295415878296,14.8562984466553],[15.1367683410645,15.1331052780151,15.1781253814697,15.2405271530151,15.2569828033447,15.3235349655151,15.3177251815796,15.2684078216553,15.1666498184204,15.1367683410645],[15.0079593658447,14.9161138534546,14.9234867095947,14.9613275527954,15.0769052505493,15.1661138534546,15.2435550689697,15.310791015625,15.3285150527954,15.3168935775757,15.2716302871704,15.160888671875,15.1392574310303,15.0079593658447],[16.2372570037842,16.218994140625,16.2255382537842,16.2215328216553,16.1690444946289,16.11328125,16.0433597564697,15.9860353469849,15.9927244186401,16.0451164245605,16.1484375,16.2372570037842],[16.6225109100342,16.5930671691895,16.572021484375,16.56103515625,16.5994129180908,16.575927734375,16.4931163787842,16.618408203125,16.6644535064697,16.6777839660645,16.6448745727539,16.6225109100342],[16.6590824127197,16.60791015625,16.6830558776855,16.7008781433105,16.808837890625,16.8410148620605,16.84375,16.7321300506592,16.6590824127197],[16.818115234375,16.794189453125,16.7972164154053,16.83251953125,16.8707046508789,16.9132328033447,16.922119140625,16.8464832305908,16.818115234375],[16.9464855194092,16.9259281158447,16.9358406066895,17.015380859375,17.0677242279053,17.0910167694092,17.1936531066895,17.1764659881592,17.0947265625,17.04931640625,16.9464855194092],[10.917236328125,10.8774900436401,10.8149909973145,10.7347164154053,10.4659185409546,10.3153810501099,10.0416011810303,9.99125957489014,9.77094745635986,9.73456954956055,9.66953086853027,9.61601543426514,9.57666015625,9.55820369720459,9.538818359375,9.51923847198486,9.505859375,9.54609394073486,9.591796875,9.57080078125,9.511474609375,9.48100566864014,9.46904277801514,9.44917011260986,9.24887657165527,9.06010723114014,9.05585956573486,8.99028301239014,8.95170879364014,8.91606426239014,8.89858436584473,8.857421875,8.80532264709473,8.74033164978027,8.63530254364014,8.56396484375,8.48935508728027,8.45351600646973,8.36777305603027,8.33774471282959,8.31601524353027,8.2568359375,8.19960975646973,8.13056564331055,8.07065391540527,8.18173885345459,8.28774356842041,8.35307598114014,8.50546836853027,8.58818340301514,8.66435527801514,8.71772480010986,8.70683574676514,8.61923885345459,8.50688457489014,8.46381855010986,8.406005859375,8.41489219665527,8.4384765625,8.44584941864014,8.48032283782959,8.61445331573486,8.72890567779541,8.80405330657959,8.95981407165527,9.03535175323486,9.15029239654541,9.27641582489014,9.37944412231445,9.46249961853027,9.52617168426514,9.56836032867432,9.64667892456055,9.70288181304932,9.78940486907959,9.8994140625,10.11572265625,10.1953125,10.2420902252197,10.2566404342651,10.1073732376099,10.0174312591553,9.93344688415527,9.88457012176514,9.82094669342041,9.69926738739014,9.66831016540527,9.60195350646973,9.581787109375,9.62006855010986,9.81093788146973,9.90244197845459,9.95859336853027,10.1328611373901,10.2920408248901,10.3981437683105,10.5634765625,10.6354494094849,10.6797847747803,10.7450199127197,10.7905759811401,10.8499507904053,10.8975591659546,10.9212884902954,10.9852533340454,11.04296875,11.0621099472046,11.0662593841553,11.08154296875,11.0974607467651,11.153076171875,11.1844720840454,11.189453125,11.1663084030151,11.1064453125,11.0399408340454,10.9453125,11.0059080123901,11.0521974563599,11.0456056594849,10.9916505813599,10.9744625091553,10.9798831939697,10.9007329940796,10.84130859375,10.8017091751099,10.7803707122803,10.7756834030151,10.7353515625,10.7432613372803,10.785888671875,10.8368654251099,10.917236328125],[21.5716800689697,21.5186519622803,21.4438953399658,21.4413089752197,21.4693851470947,21.5318851470947,21.5934562683105,21.623046875,21.5736827850342,21.5494136810303,21.5655765533447,21.59228515625,21.7914066314697,21.83349609375,21.9427242279053,21.9095230102539,21.8902835845947,21.8211421966553,21.7668933868408,21.621826171875,21.5716800689697],[21.9519538879395,21.9213390350342,21.965576171875,21.9704093933105,21.9879398345947,22.0371570587158,22.0880851745605,22.0919437408447,22.0829601287842,22.0540046691895,21.9964847564697,21.9519538879395],[22.1275386810303,22.1247081756592,22.1664047241211,22.2012691497803,22.2479972839355,22.2987308502197,22.3020992279053,22.2406730651855,22.2145500183105,22.2010746002197,22.1489753723145,22.1275386810303],[22.28515625,22.2685070037842,22.3057613372803,22.3219738006592,22.3799800872803,22.4022445678711,22.423583984375,22.4376468658447,22.4314937591553,22.38720703125,22.305908203125,22.28515625],[22.55224609375,22.5310535430908,22.5437488555908,22.5474605560303,22.5386238098145,22.4554672241211,22.4451160430908,22.4451160430908,22.4601078033447,22.4640140533447,22.4881839752197,22.5088367462158,22.5339851379395,22.55224609375],[22.6639175415039,22.6376953125,22.7111320495605,22.7876472473145,22.8052253723145,22.8067378997803,22.6813468933105,22.6639175415039],[23.1630382537842,23.1165046691895,23.1296882629395,23.1568355560303,23.1286144256592,23.0596675872803,23.054931640625,23.08984375,23.1030750274658,23.083740234375,23.0166015625,22.9750003814697,22.9434070587158,22.9349613189697,22.9493656158447,22.9423332214355,22.8769035339355,22.869140625,22.8870105743408,22.8271980285645,22.7430667877197,22.5777835845947,22.5098628997803,22.4489269256592,22.4076175689697,22.3878898620605,22.39599609375,22.3909168243408,22.3673343658447,22.3580551147461,22.3668460845947,22.1094226837158,21.9719715118408,21.9005851745605,21.79736328125,21.7746086120605,21.788330078125,21.81103515625,21.8683109283447,21.8892574310303,21.8716297149658,21.7552719116211,21.7122554779053,21.6724128723145,21.6436023712158,21.59765625,21.59375,21.6126461029053,21.5378913879395,21.4834957122803,21.4788570404053,21.5386219024658,21.5890140533447,21.458984375,21.3995113372803,21.3647956848145,21.3304195404053,21.3624019622803,21.35888671875,21.3404293060303,21.2845230102539,21.2721195220947,21.2736339569092,21.2273921966553,21.1334476470947,21.1142578125,21.1110363006592,21.0613288879395,20.9946784973145,20.9474601745605,20.8981456756592,20.837646484375,20.8119640350342,20.7755374908447,20.7361812591553,20.7145500183105,20.7334957122803,20.7166500091553,20.7016124725342,20.7138671875,20.6726570129395,20.650634765625,20.5731945037842,20.5221195220947,20.3845691680908,20.3304691314697,20.3173828125,20.326416015625,20.3114757537842,20.2921886444092,20.23193359375,20.1685543060303,20.1171398162842,20.0796871185303,20.0753421783447,20.0581531524658,20.0022945404053,19.9579105377197,19.9285640716553,19.9014167785645,19.9246597290039,19.9539051055908,20.0083484649658,19.9593753814697,19.9236335754395,19.8931159973145,19.89111328125,19.9322261810303,19.9604015350342,19.98974609375,19.9871578216553,19.9566898345947,19.940185546875,19.9213371276855,19.892822265625,19.8937511444092,19.8613777160645,19.85546875,20.0821285247803,20.3003902435303,20.3472652435303,20.4075183868408,20.4529304504395,20.4916496276855,20.5599613189697,20.61083984375,20.6437492370605,20.67236328125,20.6895027160645,20.6900882720947,20.713623046875,20.7153816223145,20.7618656158447,20.927490234375,20.973876953125,21.010986328125,21.0537109375,21.2968273162842,21.413818359375,21.5155277252197,21.5927238464355,21.6189460754395,21.5528335571289,21.5626468658447,21.5851573944092,21.7425785064697,21.8292484283447,21.8721675872803,21.9333992004395,22.0337390899658,22.1234378814697,22.087158203125,22.0635261535645,22.0528812408447,22.0735836029053,22.0979480743408,22.1342277526855,22.2069339752197,22.2679691314697,22.2029304504395,22.1429195404053,22.1094226837158,22.1041030883789,22.1837882995605,22.2001953125,22.2136726379395,22.2908687591553,22.3876953125,22.4218254089355,22.4667472839355,22.4966793060303,22.5348148345947,22.5918941497803,22.6329097747803,22.6570320129395,22.6724605560303,22.6790046691895,22.6892585754395,22.6583499908447,22.5951175689697,22.5140151977539,22.4298839569092,22.386474609375,22.3554191589355,22.30322265625,22.2229995727539,22.1871089935303,22.2089347839355,22.208740234375,22.18896484375,22.179931640625,22.1701183319092,22.149658203125,22.0920906066895,21.9801273345947,21.943115234375,21.9290027618408,21.8983402252197,21.7916507720947,21.7761707305908,21.8542976379395,21.9302749633789,21.9330081939697,21.9203624725342,21.8990726470947,21.84228515625,21.8279285430908,21.8569831848145,21.8941402435303,22.0311527252197,22.0416011810303,22.0312995910645,22.0359363555908,22.0743160247803,22.2555675506592,22.37890625,22.4742183685303,22.6185531616211,22.666015625,22.9675788879395,22.9830074310303,23.0435543060303,23.0645503997803,23.1539554595947,23.1904296875,23.1630382537842],[12.0597658157349,12.0454587936401,12.141845703125,12.2984380722046,12.3802728652954,12.3732423782349,12.342041015625,12.2313470840454,12.1585454940796,12.0597658157349],[19.348876953125,19.3294925689697,19.3413581848145,19.362548828125,19.3467769622803,19.30517578125,19.3041019439697,19.2773933410645,19.2718753814697,19.2784175872803,19.374755859375,19.3849124908447,19.348876953125],[19.7082042694092,19.7025394439697,19.7068347930908,19.6659183502197,19.6683597564697,19.6826648712158,19.6960945129395,19.7061023712158,19.7099609375,19.71826171875,19.7192878723145,19.7099609375,19.7082042694092],[19.7119140625,19.6966800689697,19.7025394439697,19.7440910339355,19.7581043243408,19.765625,19.7657222747803,19.7571277618408,19.7508792877197,19.7119140625],[35.0652351379395,35.0567855834961,35.08544921875,35.0936050415039,35.0671882629395,35.0481948852539,35.0397453308105,35.0373077392578,35.0178718566895,35.0227546691895,35.0386734008789,35.0003395080566,35.0049324035645,35.1014137268066,35.1409187316895,35.1626968383789,35.1536140441895,35.1569328308105,35.1731452941895,35.1461906433105,35.1164054870605,35.0878410339355,35.0894050598145,35.1157722473145,35.1453590393066,35.1710433959961,35.1595687866211,35.1805648803711,35.278076171875,35.3904304504395,35.3582038879395,35.3415031433105,35.3358879089355,35.3541526794434,35.4739723205566,35.5457000732422,35.5699691772461,35.6292953491211,35.6620597839355,35.593505859375,35.2920417785645,35.202392578125,35.140380859375,35.0652351379395],[35.1710433959961,35.1453590393066,35.1157722473145,35.0894050598145,35.0878410339355,35.1164054870605,35.1461906433105,35.1731452941895,35.1569328308105,35.1536140441895,35.1626968383789,35.1409187316895,35.1014137268066,35.0049324035645,35.0003395080566,35.0386734008789,35.0227546691895,35.0178718566895,35.0373077392578,35.0397453308105,35.0481948852539,35.0671882629395,35.0936050415039,35.08544921875,35.0567855834961,35.0652351379395,35.0455589294434,34.9883766174316,34.9714851379395,34.9659156799316,34.9732437133789,34.9698715209961,34.8064460754395,34.7508773803711,34.7177238464355,34.6980476379395,34.695556640625,34.6748046875,34.6369132995605,34.5999984741211,34.569580078125,34.5758781433105,34.635498046875,34.6611328125,34.6478042602539,34.6493644714355,34.7062492370605,34.7294425964355,34.7780265808105,34.9533195495605,35.0829582214355,35.0498046875,35.0899925231934,35.15576171875,35.1826667785645,35.1710433959961],[50.8589859008789,50.8613777160645,50.8865737915039,50.92724609375,51.00341796875,51.0143547058105,50.9928703308105,50.9585456848145,50.8830070495605,50.8457527160645,50.811767578125,50.7962875366211,50.7938499450684,50.7488746643066,50.7396965026855,50.7086906433105,50.6769027709961,50.6702651977539,50.6354484558105,50.6116180419922,50.6299324035645,50.6556167602539,50.6213874816895,50.5851554870605,50.5736351013184,50.5416526794434,50.5168952941895,50.50048828125,50.4830055236816,50.4546852111816,50.4237327575684,50.3940925598145,50.3718757629395,50.3668937683105,50.34521484375,50.2483901977539,50.1219215393066,50.1021499633789,50.0974617004395,50.1160659790039,50.1570281982422,50.1867179870605,50.2019500732422,50.2369155883789,50.2597198486328,50.34521484375,50.4145011901855,50.4270515441895,50.4161109924316,50.3783226013184,50.2547874450684,50.2547874450684,50.2640609741211,50.2842292785645,50.3071784973145,50.2986335754395,50.2307624816895,50.1935539245605,50.157470703125,50.1394996643066,50.1164054870605,50.1007804870605,50.0567855834961,50.006591796875,49.9832992553711,49.9722671508789,49.9990730285645,50.020263671875,50.0352554321289,50.0319366455078,50.0072746276855,49.9927749633789,49.9647445678711,49.9302749633789,49.9140625,49.9298324584961,49.9023933410645,49.8793449401855,49.8411140441895,49.8179206848145,49.7578125,49.6137237548828,49.5401382446289,49.5107917785645,49.5092277526855,49.4939956665039,49.4884757995605,49.491455078125,49.4646987915039,49.4210929870605,49.3909187316895,49.3639144897461,49.3362274169922,49.2573738098145,49.2245597839355,49.1797866821289,49.1193351745605,49.0651359558105,49.0365219116211,49.011962890625,48.9987335205078,48.9711418151855,48.9286117553711,48.8881340026855,48.8418464660645,48.8277816772461,48.8428230285645,48.8609352111816,48.841064453125,48.78076171875,48.6769027709961,48.5988273620605,48.6208992004395,48.7037124633789,48.7143058776855,48.7220191955566,48.7342300415039,48.7818832397461,48.7962417602539,48.8000984191895,48.7720680236816,48.7389678955078,48.7394027709961,48.7547836303711,48.8644523620605,48.8654289245605,48.8604469299316,48.8863754272461,48.9573707580566,48.9740257263184,48.9638671875,48.9481430053711,48.9462890625,48.9693336486816,48.9978523254395,49.0011215209961,48.9839363098145,48.8277359008789,48.7713851928711,48.7740211486816,48.7473640441895,48.6719245910645,48.5992164611816,48.6133270263672,48.6255378723145,48.6162567138672,48.5762214660645,48.5785675048828,48.6024894714355,48.6924324035645,48.72802734375,48.7598648071289,48.7669410705566,48.8159675598145,48.876708984375,48.9596672058105,48.95556640625,48.9775886535645,49.0081062316895,49.060791015625,49.0974617004395,49.1116676330566,49.1583518981934,49.2601051330566,49.3304672241211,49.329345703125,49.3662109375,49.4145011901855,49.4612312316895,49.5748558044434,49.6396980285645,49.6797828674316,49.7131843566895,49.7396469116211,49.8001441955566,49.830078125,49.8530769348145,49.87744140625,49.8958015441895,49.9555168151855,49.9985847473145,50.0423355102539,50.0975074768066,50.1480484008789,50.1758308410645,50.2134246826172,50.2685546875,50.3017578125,50.3109855651855,50.3109359741211,50.2883758544922,50.244873046875,50.1814460754395,50.2057113647461,50.2732429504395,50.3498039245605,50.3934097290039,50.3970680236816,50.4091300964355,50.4309577941895,50.4222145080566,50.4064445495605,50.4162101745605,50.4560546875,50.4903793334961,50.510498046875,50.5767593383789,50.576416015625,50.5863265991211,50.6114234924316,50.6217269897461,50.6093254089355,50.60107421875,50.616943359375,50.6928253173828,50.7046394348145,50.7165031433105,50.7612800598145,50.8011245727539,50.82275390625,50.8612289428711,50.8987312316895,50.9140625,50.9525871276855,50.9769058227539,51.0018539428711,51.0294914245605,51.0377922058105,51.0262718200684,51.0098648071289,50.9939460754395,50.9549331665039,50.9186019897461,50.9147453308105,50.8555679321289,50.8326187133789,50.814697265625,50.8183097839355,50.8423347473145,50.8589859008789],[53.8707504272461,53.8743667602539,53.8630867004395,53.8790512084961,53.93896484375,53.9966316223145,54.0362815856934,54.0595703125,54.0928230285645,54.1272430419922,54.0345687866211,53.9503440856934,53.9190444946289,53.8707504272461],[54.41796875,54.416015625,54.4560050964355,54.4661636352539,54.5154800415039,54.5333976745605,54.5012702941895,54.4383811950684,54.41796875],[54.3827133178711,54.3154296875,54.2811508178711,54.3381843566895,54.33740234375,54.2495613098145,54.245849609375,54.3256340026855,54.3645515441895,54.3969230651855,54.5089836120605,54.5442390441895,54.582763671875,54.638427734375,54.6971168518066,54.6993179321289,54.6496124267578,54.6153793334961,54.5770034790039,54.5595703125,54.5354499816895,54.4881820678711,54.4639663696289,54.4251480102539,54.3827133178711],[54.7126960754395,54.6881828308105,54.6910667419434,54.714111328125,54.7386741638184,54.7574234008789,54.7603034973145,54.7487297058105,54.7126960754395],[54.8255348205566,54.8071784973145,54.7806129455566,54.73828125,54.6739273071289,54.581298828125,54.5146484375,54.4724617004395,54.4884300231934,54.4501953125,54.408935546875,54.4383277893066,54.3162612915039,54.3756828308105,54.379150390625,54.280517578125,54.18115234375,54.0751457214355,54.0098114013672,53.9953117370605,54.0091819763184,53.9446296691895,53.9647483825684,54.113525390625,54.1454582214355,54.1683120727539,54.2258796691895,54.2837905883789,54.3470230102539,54.4673843383789,54.4457015991211,54.4226570129395,54.4110336303711,54.28271484375,54.140869140625,54.1532249450684,54.01904296875,53.8533668518066,53.8013648986816,53.7674293518066,53.7318840026855,53.7296371459961,53.7071304321289,53.624755859375,53.5564460754395,53.2834930419922,53.2167510986328,53.1990242004395,53.1055679321289,53.0267562866211,52.9823226928711,52.932861328125,52.8782196044922,52.7825202941895,52.6456069946289,52.5285148620605,52.4311027526855,52.3596649169922,52.3141593933105,52.2776374816895,52.25,52.2074699401855,52.1500473022461,52.1102027893066,52.0818367004395,52.07080078125,52.0308570861816,51.9580078125,51.9048347473145,51.8323745727539,51.7708015441895,51.6981925964355,51.6617202758789,51.6271476745605,51.544921875,51.5238761901855,51.4633293151855,51.4353523254395,51.3771476745605,51.2527351379395,51.0951194763184,51.0087394714355,50.8716316223145,50.8589859008789,50.8423347473145,50.8183097839355,50.814697265625,50.8326187133789,50.8555679321289,50.9147453308105,50.9186019897461,50.9549331665039,50.9939460754395,51.0098648071289,51.0262718200684,51.0377922058105,51.0294914245605,51.0018539428711,50.9769058227539,50.9525871276855,50.9140625,50.8987312316895,50.8612289428711,50.82275390625,50.8011245727539,50.7612800598145,50.7165031433105,50.7046394348145,50.6928253173828,50.616943359375,50.60107421875,50.6093254089355,50.6217269897461,50.6114234924316,50.5863265991211,50.576416015625,50.5767593383789,50.510498046875,50.4903793334961,50.4560546875,50.4162101745605,50.4064445495605,50.4222145080566,50.4309577941895,50.4091300964355,50.3970680236816,50.3934097290039,50.3498039245605,50.2732429504395,50.2057113647461,50.1814460754395,50.244873046875,50.2883758544922,50.3109359741211,50.3109855651855,50.3017578125,50.2685546875,50.2134246826172,50.1758308410645,50.1480484008789,50.0975074768066,50.0423355102539,49.9985847473145,49.9555168151855,49.8958015441895,49.87744140625,49.8530769348145,49.830078125,49.8001441955566,49.7396469116211,49.7131843566895,49.6797828674316,49.6396980285645,49.5748558044434,49.4612312316895,49.4145011901855,49.3662109375,49.329345703125,49.3304672241211,49.2601051330566,49.1583518981934,49.1116676330566,49.0974617004395,49.060791015625,49.0081062316895,48.9775886535645,48.95556640625,48.9596672058105,48.876708984375,48.8159675598145,48.7669410705566,48.7475090026855,48.6864242553711,48.6216773986816,48.5874519348145,48.5423851013184,48.5327644348145,48.5230484008789,48.5818328857422,48.5718231201172,48.5645523071289,48.3941421508789,48.3613777160645,48.3312492370605,48.3019027709961,48.2899436950684,48.2750968933105,48.2037124633789,48.1608390808105,48.1069831848145,48.0759773254395,47.9848136901855,47.890625,47.8077621459961,47.7458038330078,47.7218742370605,47.7128448486328,47.7094230651855,47.69873046875,47.6551284790039,47.5791473388672,47.5080108642578,47.4780769348145,47.4756813049316,47.5064468383789,47.5421867370605,47.5641593933105,47.5904273986816,47.6070289611816,47.639404296875,47.6693344116211,47.6562995910645,47.6361351013184,47.6373062133789,47.6661148071289,47.6881828308105,47.7027359008789,47.71826171875,47.7090835571289,47.646728515625,47.6195335388184,47.58349609375,47.5497550964355,47.506103515625,47.4871559143066,47.4602508544922,47.4249038696289,47.4136238098145,47.4251976013184,47.4088859558105,47.3931159973145,47.3981437683105,47.4267082214355,47.470458984375,47.5007820129395,47.5202140808105,47.5241203308105,47.5472183227539,47.5417976379395,47.5515632629395,47.5410652160645,47.4169921875,47.3660659790039,47.3134269714355,47.2841339111328,47.27880859375,47.3171882629395,47.3634262084961,47.374267578125,47.3795928955078,47.3933563232422,47.4285163879395,47.4490699768066,47.4735832214355,47.5053215026855,47.5522956848145,47.5755348205566,47.55078125,47.52587890625,47.5340309143066,47.5242156982422,47.5989265441895,47.6707038879395,47.6707038879395,47.6563949584961,47.6626968383789,47.70361328125,47.716552734375,47.7099113464355,47.6980438232422,47.7000503540039,47.76611328125,47.775634765625,47.7668952941895,47.7313461303711,47.6877937316895,47.6626968383789,47.6518058776855,47.6591300964355,47.6519050598145,47.6377944946289,47.6240234375,47.6126937866211,47.59619140625,47.5921401977539,47.589599609375,47.60693359375,47.60693359375,47.576171875,47.5638656616211,47.5698738098145,47.5927238464355,47.6065406799316,47.6738777160645,47.7736320495605,47.9056625366211,48.0025863647461,48.0643081665039,48.1567878723145,48.280029296875,48.4100112915039,48.5468254089355,48.6360359191895,48.6985359191895,48.873291015625,48.8864250183105,48.9735832214355,48.9858894348145,49.0109367370605,49.0418968200684,49.061767578125,49.0863761901855,49.1521949768066,49.153076171875,49.1295394897461,49.1136245727539,49.1275405883789,49.1248550415039,49.1126937866211,49.1234359741211,49.1798820495605,49.1946296691895,49.20751953125,49.2019538879395,49.1739273071289,49.1541481018066,49.1605987548828,49.2908706665039,49.3196792602539,49.34619140625,49.3946800231934,49.44287109375,49.4581527709961,49.4527359008789,49.5126953125,49.599609375,49.6449699401855,49.6820297241211,49.7078132629395,49.75439453125,49.7984848022461,49.8053245544434,49.837890625,49.8721694946289,49.9151344299316,49.9743156433105,50.0343742370605,50.0942344665527,50.1209983825684,50.1393547058105,50.232666015625,50.316162109375,50.4002418518066,50.4517593383789,50.4854965209961,50.4991188049316,50.5225105285645,50.5453605651855,50.5966796875,50.6372566223145,50.6792488098145,50.7322273254395,50.7504386901855,50.9048843383789,50.949951171875,50.9729499816895,50.9842262268066,51.0056648254395,51.0301246643066,51.0453147888184,51.0408210754395,51.0566902160645,51.1474151611328,51.1648445129395,51.1747093200684,51.1799812316895,51.1990242004395,51.22412109375,51.3548316955566,51.4105949401855,51.4500007629395,51.4889183044434,51.5500984191895,51.5989265441895,51.6377944946289,51.6582489013672,51.7624015808105,51.8026847839355,51.833984375,51.8539543151855,51.8704109191895,51.8807601928711,51.8507308959961,51.8246574401855,51.8300285339355,51.8583984375,51.8539543151855,51.910888671875,51.9382820129395,51.9673805236816,51.9801750183105,52.0361824035645,52.056884765625,52.0802230834961,52.0986824035645,52.1112327575684,52.1357917785645,52.2055168151855,52.2660140991211,52.3314971923828,52.3802261352539,52.4189949035645,52.444091796875,52.4402847290039,52.4422836303711,52.4640159606934,52.4992179870605,52.5301742553711,52.5496597290039,52.5735855102539,52.59765625,52.6178703308105,52.6340827941895,52.633544921875,52.6513671875,52.7447776794434,52.8870124816895,52.9662132263184,52.99951171875,53.1872062683105,53.2822761535645,53.3269538879395,53.3758316040039,53.4776344299316,53.5569801330566,53.654541015625,53.6813468933105,53.697265625,53.6907234191895,53.5434074401855,53.4676780700684,53.4324226379395,53.4453125,53.5111808776855,53.5841331481934,53.606201171875,53.5517082214355,53.5143547058105,53.3942375183105,53.556884765625,53.6707534790039,53.7811050415039,53.8384780883789,53.875,53.835693359375,53.85595703125,53.8134765625,53.6004867553711,53.5656242370605,53.5546379089355,53.6001968383789,53.859130859375,53.8912086486816,53.9009246826172,53.9262199401855,53.96533203125,54.0002899169922,54.2607917785645,54.299560546875,54.3130340576172,54.2952156066895,54.2949714660645,54.3539543151855,54.3976554870605,54.4275398254395,54.4675788879395,54.538330078125,54.5939445495605,54.6959457397461,54.7918472290039,54.9034156799316,54.9033203125,54.901123046875,54.8969230651855,54.8446769714355,54.8080062866211,54.8062973022461,54.8404273986816,54.8554229736328,54.8343734741211,54.8255348205566,54.8255348205566],[54.7869606018066,54.76708984375,54.9083023071289,55.0587387084961,55.0553703308105,55.0147438049316,54.9862823486328,54.9293937683105,54.8998565673828,54.8917465209961,54.8653793334961,54.8476066589355,54.7869606018066],[11.4998054504395,11.36572265625,11.1943359375,10.9993171691895,10.9979496002197,11.00927734375,11.0423831939697,11.0783205032349,11.0807619094849,11.0470705032349,11.0052242279053,10.9916009902954,10.9683589935303,10.9410161972046,10.955810546875,10.9804677963257,11.1877927780151,11.4128913879395,11.589111328125,11.68603515625,11.7237787246704,11.8578615188599,11.912353515625,12.1341304779053,12.3242673873901,12.4664058685303,12.494384765625,12.5213375091553,12.5136232376099,12.3765621185303,12.3803224563599,12.4228515625,12.5693359375,12.622802734375,12.6212892532349,12.6623048782349,12.7085933685303,12.6604490280151,12.4638671875,12.3670406341553,12.18994140625,12.0912590026855,12.0270013809204,11.9695310592651,11.8290529251099,11.7394046783447,11.5601081848145,11.5721673965454,11.5042972564697,11.4967775344849,11.5095701217651,11.5617189407349,11.5866212844849,11.5884761810303,11.5660152435303,11.4998054504395],[15.2490243911743,15.2272958755493,15.3998537063599,15.525146484375,15.6034660339355,15.6331062316895,15.5850591659546,15.5267086029053,15.3731441497803,15.2490243911743],[54.8916511535645,54.82958984375,54.8331565856934,54.8074226379395,54.7676773071289,54.679443359375,54.6537132263184,54.6624526977539,54.6288566589355,54.7730979919434,54.8933601379395,54.9405784606934,54.9518051147461,54.8916511535645],[54.8475608825684,54.837158203125,54.8589363098145,54.9409675598145,54.9627456665039,54.9488258361816,54.9059562683105,54.8968276977539,54.8605461120605,54.8475608825684],[54.9657745361328,54.9508781433105,54.9618148803711,54.8924827575684,54.9144020080566,54.9586906433105,54.9748077392578,54.9936027526855,55.0210952758789,55.0641098022461,55.0409202575684,55.0312004089355,55.0174827575684,54.997314453125,54.9657745361328],[54.8863754272461,54.8724594116211,54.8966293334961,54.906005859375,55.0599136352539,55.0690422058105,55.0582504272461,54.9864768981934,54.9079093933105,54.8863754272461],[54.750732421875,54.7450675964355,54.8260765075684,54.8514137268066,54.9032707214355,54.9620132446289,55.0521965026855,55.1578636169434,55.1562004089355,55.0621109008789,54.7996597290039,54.750732421875],[55.0218772888184,55.0049324035645,55.032958984375,55.1022453308105,55.238037109375,55.2967300415039,55.14453125,55.087158203125,55.0218772888184],[55.6098175048828,55.5576171875,55.4463386535645,55.3218765258789,55.269775390625,55.2030258178711,55.1333999633789,55.0524406433105,55.0487823486328,55.087890625,55.1631813049316,55.2054672241211,55.2289047241211,55.3572235107422,55.5154762268066,55.5353012084961,55.61083984375,55.5989761352539,55.5603523254395,55.5580558776855,55.6128425598145,55.6098175048828],[55.5965347290039,55.5540008544922,55.5562515258789,55.6146011352539,55.6500968933105,55.6802253723145,55.6793479919434,55.6467781066895,55.5965347290039],[55.7830581665039,55.7650871276855,55.7837905883789,55.8484878540039,55.9065895080566,55.9585456848145,55.991943359375,55.9141578674316,55.8775901794434,55.8338851928711,55.7830581665039],[55.7850570678711,55.6849632263184,55.6558113098145,55.6366195678711,55.6162605285645,55.5878410339355,55.5378913879395,55.4665031433105,55.4142570495605,55.3856468200684,55.2861824035645,55.2371101379395,55.1881332397461,55.0699234008789,54.9767608642578,54.9090347290039,54.8153305053711,54.7726097106934,54.9153327941895,54.9724617004395,55.0391616821289,55.0959968566895,55.1869125366211,55.2115249633789,55.2147483825684,55.1978530883789,55.2044448852539,55.32861328125,55.4656219482422,55.5347671508789,55.5660667419434,55.6007308959961,55.6292953491211,55.6444358825684,55.7215347290039,55.740234375,55.731201171875,55.7364730834961,55.7525405883789,55.8792991638184,55.9072265625,55.9434585571289,55.9568862915039,55.9079093933105,55.8294906616211,55.72900390625,55.70166015625,55.6976547241211,55.7718772888184,55.8079605102539,55.8280754089355,55.8958969116211,55.9373054504395,55.9681663513184,56.0521507263184,56.11865234375,56.1221160888672,56.1058578491211,56.0833969116211,56.0640602111816,56.0330085754395,55.958984375,55.91845703125,55.7850570678711],[57.2525367736816,57.2291030883789,57.2622566223145,57.30859375,57.3299293518066,57.3229026794434,57.2769050598145,57.2525367736816],[54.8255348205566,54.8255348205566,54.8343734741211,54.8554229736328,54.8404273986816,54.8062973022461,54.8080062866211,54.8446769714355,54.8969230651855,54.901123046875,54.9033203125,54.9034156799316,54.9859352111816,55.0455551147461,55.13427734375,55.1556625366211,55.3285636901855,55.418212890625,55.5103034973145,55.5998077392578,55.9011726379395,55.9823722839355,56.139892578125,56.3211898803711,56.6068840026855,56.6180686950684,56.61669921875,56.5654296875,56.560302734375,56.5144996643066,56.4956550598145,56.5442886352539,56.62744140625,56.7350578308105,56.7748031616211,56.7938499450684,56.7504386901855,56.70166015625,56.8083992004395,57.01171875,57.0436515808105,57.01611328125,56.8872566223145,56.7252922058105,56.7104034423828,56.66455078125,56.7121086120605,56.7540054321289,56.8153343200684,56.8523445129395,56.9844245910645,57.1112785339355,57.1100616455078,57.1505851745605,57.1554183959961,57.1465339660645,57.17431640625,57.2324714660645,57.4784202575684,57.5809593200684,57.6170425415039,57.7353973388672,57.7369155883789,57.648681640625,57.6145515441895,57.5622062683105,57.4485359191895,57.3793487548828,57.2432136535645,57.1722640991211,57.0213394165039,56.9991188049316,56.8229484558105,56.7809104919434,56.6205101013184,56.5548324584961,56.5205078125,56.521728515625,56.4928703308105,56.4432640075684,56.3590316772461,56.2955093383789,56.2419891357422,56.2020988464355,56.2003440856934,56.2761726379395,56.2515640258789,56.212890625,56.00537109375,55.8651847839355,55.8538055419922,55.8744621276855,55.8760719299316,55.8428192138672,55.8130836486816,55.7614250183105,55.7355461120605,55.7075691223145,55.6509780883789,55.608154296875,55.5574722290039,55.4932136535645,55.41357421875,55.3436546325684,55.2664070129395,55.2047386169434,55.1162605285645,55.03955078125,55.04052734375,55.0228004455566,55.0001449584961,54.968017578125,54.9283180236816,54.8255348205566],[18.0391597747803,18.2039546966553,18.2707996368408,18.34130859375,18.4162120819092,18.5125980377197,18.597900390625,18.6103515625,18.6141605377197,18.6455078125,18.73291015625,18.80322265625,18.8563976287842,18.9200210571289,18.9870128631592,19.0455074310303,19.1307621002197,19.1635246276855,19.1959476470947,19.2858390808105,19.324462890625,19.4219722747803,19.486572265625,19.6881847381592,19.7181644439697,19.735107421875,19.795166015625,19.8486328125,19.87744140625,19.8953609466553,19.8939952850342,19.8473644256592,19.84814453125,19.8904781341553,19.9139652252197,19.8872547149658,19.8508796691895,19.7932624816895,19.775634765625,19.7769527435303,19.771240234375,19.6760749816895,19.6380367279053,19.6361331939697,19.6729488372803,19.671875,19.5897464752197,19.4732913970947,19.3671398162842,19.2992191314697,19.3277339935303,19.2718257904053,19.2256832122803,19.2010746002197,19.2120132446289,19.2064933776855,19.1604976654053,19.1178226470947,19.1076164245605,19.0860843658447,19.0519046783447,19.0284671783447,19.01318359375,18.9884777069092,18.90478515625,18.7144527435303,18.671142578125,18.6115226745605,18.5380859375,18.417724609375,18.3790035247803,18.35546875,18.3062515258789,18.2220211029053,18.2149410247803,18.2184085845947,18.26611328125,18.3393077850342,18.4080085754395,18.3992176055908,18.4398441314697,18.4201164245605,18.4156742095947,18.4363765716553,18.4435539245605,18.417724609375,18.3736324310303,18.3456535339355,18.277099609375,18.2517585754395,18.21728515625,18.267578125,18.3362293243408,18.3456058502197,18.29248046875,18.273193359375,18.2503414154053,18.224365234375,18.1283683776855,18.0700206756592,17.849609375,17.6941413879395,17.6460933685303,17.6355953216553,17.7250003814697,17.7573738098145,17.7736320495605,17.8211421966553,17.888671875,17.9541015625,18.0054683685303,18.0391597747803],[36.9372062683105,36.8838844299316,36.8339347839355,36.7875022888184,36.7607421875,36.6325225830078,36.5452651977539,36.5189437866211,36.4951171875,36.4556159973145,36.4181671142578,36.3679695129395,36.1887702941895,36.0509796142578,35.8705558776855,35.8018074035645,35.7192878723145,35.6549339294434,35.5822257995605,35.4031257629395,35.2996101379395,35.203857421875,35.0846176147461,34.9794921875,34.8289566040039,34.7340850830078,34.6462898254395,34.5639152526855,34.5126953125,34.4687004089355,34.4103012084961,34.2544898986816,34.125,34.0805168151855,33.9765129089355,33.8324699401855,33.7179183959961,33.5486335754395,33.3623046875,33.2685050964355,33.2331047058105,33.172119140625,33.0890617370605,33.0553207397461,32.9267082214355,32.6962890625,32.5436058044434,32.4223136901855,32.3104476928711,32.2121086120605,32.1053733825684,32.0723609924316,31.8461437225342,31.621337890625,31.3736820220947,31.1253433227539,30.8329124450684,30.6667976379395,30.4653797149658,30.2293968200684,30.1792964935303,30.1152324676514,29.99365234375,29.7959461212158,29.6364269256592,29.5669898986816,29.3689460754395,29.1769523620605,29.1147937774658,28.9669933319092,28.5602054595947,28.0433101654053,27.7856922149658,27.552978515625,27.3308582305908,27.2193355560303,27.0447769165039,26.9158210754395,26.8479480743408,26.6308097839355,26.5519523620605,26.4382305145264,26.333740234375,26.2455081939697,26.1470699310303,26.067138671875,25.89013671875,25.624267578125,25.3320808410645,25.258544921875,25.051025390625,24.7902355194092,24.6762199401855,24.5910148620605,24.5302257537842,24.4855937957764,24.480224609375,24.5513668060303,24.4340343475342,24.3143558502197,24.2908191680908,24.1396961212158,23.892578125,23.69482421875,23.5178718566895,23.2125968933105,22.9072761535645,22.6020030975342,22.2967281341553,21.9914054870605,21.6861324310303,21.380859375,21.0755863189697,20.8730964660645,20.6944828033447,20.4705066680908,20.2480487823486,20.0729484558105,19.8461418151855,19.7319831848145,19.4791488647461,19.4342288970947,19.3595199584961,19.2910652160645,19.227783203125,19.1845207214355,19.1427726745605,19.083740234375,19.0416011810303,18.9961433410645,18.9884281158447,18.9866218566895,18.9883785247803,19.0132808685303,19.0729007720947,19.1031742095947,19.1500988006592,19.212158203125,19.2681636810303,19.3120613098145,19.3454093933105,19.3726081848145,19.4109382629395,19.4735851287842,19.5604000091553,19.7183113098145,19.770751953125,19.7896976470947,19.8501968383789,19.9166011810303,19.9559574127197,19.9694347381592,19.992919921875,20.0350093841553,20.0638675689697,20.2103023529053,20.247802734375,20.272705078125,20.296875,20.3315906524658,20.3783702850342,20.4588375091553,20.5243644714355,20.5555667877197,20.7135753631592,20.7672863006592,20.8174304962158,20.8913078308105,20.9819831848145,21.0625,21.1022453308105,21.19775390625,21.411865234375,21.6259746551514,21.8401355743408,21.8512420654297,22.05419921875,22.268310546875,22.4824714660645,22.696533203125,22.91064453125,23.1248035430908,23.3388671875,23.5530281066895,23.7671394348145,23.9812507629395,24.1953620910645,24.4094715118408,24.62353515625,24.8044929504395,24.99560546875,25.1354503631592,25.2745113372803,25.4237785339355,25.5164070129395,25.6270008087158,25.7375965118408,25.8481941223145,25.9587898254395,26.0694332122803,26.1799793243408,26.2905769348145,26.4011726379395,26.5117664337158,26.6223621368408,26.7329597473145,26.8435554504395,26.9541492462158,27.0647449493408,27.1753425598145,27.2859382629395,27.490234375,27.6564445495605,27.900390625,28.1120109558105,28.3236808776855,28.46923828125,28.6207523345947,28.6894035339355,28.7186031341553,28.7678718566895,28.8801784515381,28.9301738739014,28.9805164337158,29.1324234008789,29.1747570037842,29.3495101928711,29.3751964569092,29.3922367095947,29.4250011444092,29.4947280883789,29.5749034881592,29.6126480102539,29.6195793151855,29.6251945495605,29.6016101837158,29.5838375091553,29.5687999725342,29.5789546966553,29.6038570404053,29.6598644256592,29.7260246276855,29.7837886810303,29.8091316223145,29.8203601837158,29.8161144256592,29.8083019256592,29.8106918334961,29.8189468383789,29.8312511444092,29.8690414428711,29.9179706573486,29.9569339752197,30.0586414337158,30.1661624908447,30.326416015625,30.4653797149658,30.5523929595947,30.6047859191895,30.6255359649658,30.6988754272461,30.8095703125,30.9135246276855,30.92724609375,30.9444828033447,30.9640140533447,31.0009269714355,31.0657730102539,31.1113777160645,31.1354007720947,31.1618156433105,31.1666011810303,31.1978034973145,31.2554683685303,31.3088397979736,31.3618183135986,31.4371109008789,31.5123558044434,31.56640625,31.619873046875,31.6619148254395,31.6895503997803,31.7000961303711,31.686767578125,31.7045402526855,31.8342761993408,31.8742179870605,31.9639644622803,32.0425262451172,32.06884765625,32.07470703125,32.0957527160645,32.1256828308105,32.1299819946289,32.121337890625,32.1150398254395,32.1047859191895,32.0995597839355,32.0948753356934,32.0890159606934,32.1072273254395,32.16455078125,32.2711410522461,32.3376007080078,32.399169921875,32.4683113098145,32.5522956848145,32.6084938049316,32.6756820678711,32.703369140625,32.7848167419434,32.8776397705078,33.0735816955566,33.183349609375,33.3186531066895,33.5667495727539,33.7168464660645,33.7818374633789,33.8582038879395,33.9902839660645,34.1760749816895,34.367919921875,34.4332504272461,34.467041015625,34.49609375,34.5570793151855,34.6073265075684,34.6546363830566,34.7231941223145,34.7519035339355,34.8355445861816,34.9708480834961,35.02978515625,35.1041984558105,35.0850601196289,35.09423828125,35.18310546875,35.3030776977539,35.3642578125,35.4957504272461,35.578857421875,35.6684074401855,35.8615226745605,35.8631820678711,35.819091796875,35.8328132629395,35.8667793273926,35.9005355834961,36.0631370544434,36.162353515625,36.2618141174316,36.3565444946289,36.4439468383789,36.5195808410645,36.5675811767578,36.6103019714355,36.6006851196289,36.7388687133789,36.7844734191895,36.7951164245605,36.8961906433105,36.8963356018066,36.8624000549316,36.8080558776855,36.6768074035645,36.6482391357422,36.6754379272461,36.7996101379395,36.8642578125,36.9383277893066,37.0460433959961,37.0857429504395,37.0030288696289,36.91943359375,36.943359375,36.968505859375,37.0923843383789,37.0592765808105,36.999755859375,36.8802719116211,36.8563461303711,36.9103507995605,36.9372062683105],[-2.97314453125,-3.01171875,-3.00830078125,-2.99589824676514,-2.95175766944885,-2.81191420555115,-2.75312519073486,-2.69628882408142,-2.66884779930115,-2.67382788658142,-2.7255859375,-2.81953120231628,-2.83378911018372,-2.84589862823486,-2.97314453125],[-1.33994150161743,-1.34199202060699,-1.29912114143372,-1.22099602222443,-1.23984396457672,-1.26230454444885,-1.29228508472443,-1.33994150161743],[-0.911034941673279,-0.95234352350235,-0.933788895606995,-0.913476347923279,-0.888574481010437,-0.826855421066284,-0.793359160423279,-0.722265422344208,-0.680077970027924,-0.689843475818634,-0.704589664936066,-0.728417873382568,-0.785937666893005,-0.826073944568634,-0.911034941673279],[-0.771582186222076,-0.773340046405792,-0.676464915275574,-0.581445574760437,-0.517382621765137,-0.484667867422104,-0.544824421405792,-0.658789157867432,-0.741992354393005,-0.757226705551147,-0.771582186222076],[-0.460839688777924,-0.478222697973251,-0.443945467472076,-0.390820235013962,-0.310937315225601,-0.28447237610817,-0.255664080381393,-0.32246121764183,-0.420898228883743,-0.460839688777924],[-0.333984524011612,-0.364257901906967,-0.329394578933716,-0.271386623382568,-0.192187234759331,-0.160449385643005,-0.189843833446503,-0.278417974710464,-0.333984524011612],[0.0251465458422899,-0.0393066257238388,-0.223046734929085,-0.416894406080246,-0.525195419788361,-0.575585961341858,-0.595312297344208,-0.671777546405792,-0.752050638198853,-0.940527379512787,-1.01962876319885,-1.01699209213257,-0.996679544448853,-0.924609541893005,-0.860937297344208,-0.799511611461639,-0.706250071525574,-0.622851490974426,-0.559081852436066,-0.496972739696503,-0.373632669448853,-0.287207126617432,-0.023388709872961,-0.0103029264137149,-0.0466794446110725,-0.0147949168458581,0.0020998902618885,0.0622558742761612,0.105175569653511,0.125830113887787,0.0914062485098839,0.0251465458422899],[1.25278306007385,1.24536156654358,1.29321277141571,1.34892594814301,1.35976576805115,1.25278306007385],[-0.106543220579624,-0.142187505960464,-0.145995944738388,-0.157129094004631,-0.122851319611073,-0.122851319611073,-0.15761724114418,-0.200097486376762,-0.248339965939522,-0.321777105331421,-0.408886730670929,-0.506542861461639,-0.55537086725235,-0.590136766433716,-0.653906404972076,-0.707129061222076,-0.951855361461639,-0.966796934604645,-0.96806663274765,-0.966796934604645,-0.940234661102295,-0.924316167831421,-0.962206959724426,-1.07119154930115,-1.31630861759186,-1.53124976158142,-1.60732424259186,-1.72812497615814,-1.89345717430115,-2.13310527801514,-2.24394512176514,-2.33134746551514,-2.43232440948486,-2.56259775161743,-2.63593745231628,-2.73769545555115,-2.80966806411743,-2.85996079444885,-2.91240215301514,-2.98164081573486,-3.04697299003601,-3.20683622360229,-3.28388714790344,-3.35019540786743,-3.38046884536743,-3.39980506896973,-3.43212890625,-3.46513628959656,-3.48583984375,-3.48916006088257,-3.47255849838257,-3.43613290786743,-3.39902329444885,-3.38828086853027,-3.39736366271973,-3.43124985694885,-3.59482407569885,-3.67431640625,-3.70576143264771,-3.77685523033142,-3.84306621551514,-3.90205025672913,-3.95214819908142,-3.98691415786743,-4.04160118103027,-4.15732431411743,-4.24814462661743,-4.32587909698486,-4.38398408889771,-4.42509746551514,-4.45820283889771,-4.51767587661743,-4.56240224838257,-4.59267568588257,-4.6650390625,-4.71445322036743,-4.77070331573486,-4.81865215301514,-4.8583984375,-4.87324237823486,-4.90800809860229,-4.96914005279541,-4.99062490463257,-4.95820331573486,-4.95761680603027,-4.92783164978027,-4.84003925323486,-4.76621055603027,-4.67060565948486,-4.53916025161743,-4.50058603286743,-4.45488262176514,-4.46757793426514,-4.47636747360229,-4.44589853286743,-4.39033222198486,-4.32753896713257,-4.29609394073486,-4.31103515625,-4.34902334213257,-4.41679668426514,-4.46367168426514,-4.46142578125,-4.43007802963257,-4.39365243911743,-4.33583974838257,-4.20849561691284,-4.20517539978027,-4.16552734375,-4.119140625,-4.06953144073486,-4.01005887985229,-3.97861337661743,-4.00341749191284,-4.00507831573486,-3.94882822036743,-3.92402338981628,-3.90585947036743,-3.87773394584656,-3.78769516944885,-3.73886680603027,-3.71074223518372,-3.65449237823486,-3.61318373680115,-3.57675766944885,-3.52216815948486,-3.49248027801514,-3.46103525161743,-3.42460942268372,-3.40644550323486,-3.38789081573486,-3.37480449676514,-3.32431674003601,-3.27402353286743,-3.22812509536743,-3.15771460533142,-3.09013676643372,-2.77695345878601,-2.57910132408142,-2.48466801643372,-2.35654258728027,-2.16787099838257,-2.11054682731628,-2.06738305091858,-2.14570283889771,-2.42363262176514,-2.54853534698486,-2.57871127128601,-2.55673837661743,-2.35380864143372,-2.39072251319885,-2.52841806411743,-2.63056612014771,-2.66464853286743,-2.70673799514771,-2.66796875,-2.62597680091858,-2.39687490463257,-2.34902334213257,-2.26914072036743,-2.23544931411743,-2.18925786018372,-2.14121103286743,-2.07666039466858,-1.93457019329071,-1.82294952869415,-1.63242208957672,-1.38339853286743,-1.28583991527557,-1.07890653610229,-0.974707186222076,-0.89873069524765,-0.84794944524765,-0.683789014816284,-0.585449397563934,-0.625097513198853,-0.620507597923279,-0.583984375,-0.436035186052322,-0.368261724710464,-0.16582028567791,-0.113085828721523,-0.0060544447042048,0.155371263623238,0.410156399011612,0.592285394668579,0.784765422344208,0.834570288658142,0.860204994678497,0.922265529632568,0.979785323143005,0.971142888069153,1.06005847454071,1.10458981990814,1.20624971389771,1.29594755172729,1.45537126064301,1.43466794490814,1.35869157314301,1.28344714641571,1.23666977882385,1.19882833957672,1.04609370231628,0.968554735183716,0.89873069524765,0.825390338897705,0.837841928005219,0.782226622104645,0.723633110523224,0.689502000808716,0.660351574420929,0.651171684265137,0.636523485183716,0.424853384494781,0.393896460533142,0.360400199890137,0.347753882408142,0.355077981948853,0.296240389347076,0.268505781888962,0.247753828763962,0.241650119423866,0.25083002448082,0.272118836641312,0.268164306879044,0.240966573357582,0.235448971390724,0.261230379343033,0.303905993700027,0.378857463598251,0.404980331659317,0.448486179113388,0.439404189586639,0.34555658698082,0.313085913658142,0.247753828763962,0.150976672768593,0.0892576575279236,0.0628907158970833,-0.0384277030825615,-0.106543220579624],[31.3226089477539,31.2922859191895,31.2083015441895,30.9950199127197,30.8278331756592,30.5962905883789,30.5073738098145,30.4460468292236,30.1914539337158,29.9820308685303,29.8121089935303,29.5639171600342,29.4773445129395,29.43212890625,29.2706050872803,28.7579097747803,28.3573226928711,28.1064929962158,28.0160160064697,27.8889656066895,27.7643070220947,27.828857421875,28.0476570129395,28.2555675506592,28.3898429870605,28.5677242279053,28.6957054138184,28.7777824401855,28.9782733917236,29.0730457305908,29.2862281799316,29.3999996185303,29.4500007629395,29.5217781066895,29.7984390258789,29.9739742279053,29.9254398345947,29.8515129089355,29.7493152618408,29.6306648254395,29.5337886810303,29.3863296508789,29.3219242095947,29.1821784973145,28.9922370910645,28.9277324676514,28.78662109375,28.7028789520264,28.630615234375,28.5652351379395,28.4422855377197,28.2082996368408,28.0505867004395,27.9744625091553,27.8981456756592,27.7012195587158,27.6073741912842,27.4305648803711,27.3411140441895,27.2681636810303,27.1849136352539,27.0494632720947,26.6490230560303,26.5507335662842,26.0243644714355,25.691162109375,25.4425296783447,25.1397953033447,24.4751453399658,24.2699699401855,24.15478515625,24.0660152435303,23.9377937316895,23.9503421783447,23.9425773620605,23.920654296875,23.8428707122803,23.779296875,23.4425296783447,23.2710933685303,22.9461917877197,22.8487319946289,22.7856941223145,22.7396488189697,22.6288089752197,22.3941898345947,22.09765625,22.0157718658447,21.9967288970947,21.9966316223145,21.99658203125,21.9965343475342,21.9964847564697,21.9964351654053,21.9963874816895,21.996337890625,21.9962902069092,21.9962406158447,21.9961910247803,21.9961433410645,21.99609375,21.9959964752197,21.9959964752197,21.9958992004395,21.995849609375,21.995849609375,22.0846691131592,22.1478023529053,22.1915035247803,22.2024421691895,22.1886234283447,22.0022945404053,21.994873046875,21.994873046875,21.9949226379395,21.9950199127197,21.9951171875,21.9951171875,21.9952163696289,21.9952640533447,21.9953136444092,21.9953632354736,21.9954605102539,21.9955081939697,21.9955577850342,21.9956073760986,21.9956550598145,21.9957523345947,21.9958019256592,21.995849609375,22.2204093933105,22.4449710845947,22.6695308685303,22.8940925598145,23.11865234375,23.3432140350342,23.5678215026855,23.7923831939697,24.0169429779053,24.2415046691895,24.466064453125,24.6906242370605,24.9151859283447,25.1397457122803,25.3643074035645,25.5888671875,25.8134269714355,26.0379886627197,26.2625484466553,26.4871101379395,26.711669921875,26.9362316131592,27.1608390808105,27.3854007720947,27.6099605560303,27.8345222473145,28.0590839385986,28.2836418151855,28.5082035064697,28.7327651977539,28.9573249816895,29.181884765625,29.2238292694092,29.3762722015381,29.5702629089355,29.8087387084961,29.8860359191895,30.1315441131592,30.2010746002197,30.2505874633789,30.4575214385986,30.5580062866211,30.6785144805908,30.7765617370605,30.9264640808105,31.0612316131592,31.1991710662842,31.3348140716553,31.427490234375,31.5140151977539,31.5671882629395,31.6269035339355,31.6549797058105,31.5337886810303,31.5127925872803,31.6208972930908,31.5121097564697,31.4703598022461,31.3778820037842,31.2126941680908,31.1917495727539,31.1950206756592,31.0974102020264,31.0504398345947,30.9426765441895,30.8567390441895,30.8302726745605,30.8345718383789,30.866943359375,30.9274406433105,31.0115242004395,31.2274913787842,31.2654304504395,31.2556629180908,31.2583980560303,31.3168449401855,31.3570289611816,31.4027347564697,31.4576187133789,31.4729995727539,31.5668468475342,31.5223655700684,31.4169921875,31.4038581848145,31.4398937225342,31.4627914428711,31.5075664520264,31.5915508270264,31.6033191680908,31.5875968933105,31.4582538604736,31.4557609558105,31.5263195037842,31.5414066314697,31.5020999908447,31.341064453125,31.344482421875,31.4824695587158,31.4137210845947,31.2925758361816,31.2401847839355,31.2205085754395,31.1529769897461,31.0928211212158,31.1190414428711,31.2008781433105,31.2465343475342,31.2937507629395,31.294921875,31.2560558319092,31.1007328033447,31.0687503814697,31.0740242004395,31.1177234649658,31.1109371185303,31.1681652069092,31.126220703125,31.0845222473145,31.1309566497803,31.1304206848145,31.1809577941895,31.3039035797119,31.3226089477539],[15.6961431503296,15.6429195404053,15.6481447219849,15.6658687591553,15.7034673690796,15.6291990280151,15.5798835754395,15.5773439407349,15.5981435775757,15.5909175872803,15.6124515533447,15.6961431503296,15.6556148529053,15.6658687591553,15.6766109466553,15.7332515716553,15.7445316314697,15.7890634536743,15.8065910339355,15.8282718658447,15.8894033432007,15.8754892349243,15.8384761810303,15.7952632904053,15.6961431503296],[16.0824222564697,15.9857416152954,16.0226554870605,16.0426750183105,16.0809555053711,16.1044921875,16.0824222564697],[18.0050792694092,17.4271488189697,17.0855484008789,16.7291507720947,16.1937007904053,15.9210939407349,15.7866706848145,15.5321283340454,15.5225095748901,15.4525384902954,15.2136726379395,15.1248531341553,15.2012691497803,15.2453117370605,15.31884765625,15.4135732650757,15.4703121185303,15.3931150436401,15.3345212936401,15.2170896530151,15.1519536972046,15.0141115188599,14.9740238189697,14.9639644622803,14.93359375,14.8830080032349,14.7430181503296,14.6203117370605,14.243896484375,13.9830570220947,13.587646484375,13.3980960845947,13.2125978469849,13.2214841842651,13.0186033248901,12.8642578125,12.808349609375,12.8995113372803,12.8246088027954,12.7085933685303,12.6623048782349,12.6212892532349,12.622802734375,12.5693359375,12.4228515625,12.3803224563599,12.3765621185303,12.5136232376099,12.5213375091553,12.494384765625,12.4664058685303,12.5702152252197,12.6619625091553,12.7714347839355,12.8206052780151,12.88232421875,13.02587890625,13.1839351654053,13.313232421875,13.4998044967651,13.7361326217651,13.9831056594849,14.1116695404053,14.1444826126099,14.2251949310303,14.3380870819092,14.4311513900757,14.4560546875,14.4591312408447,14.4406728744507,14.4990234375,14.4990234375,14.5160646438599,14.5367193222046,14.5118656158447,14.4703121185303,14.4793949127197,14.5374994277954,14.5818853378296,14.6282234191895,14.6282234191895,14.5868663787842,14.4823246002197,14.4244136810303,14.4286136627197,14.4704093933105,14.649658203125,14.6788091659546,14.6814937591553,14.702733039856,14.7371101379395,14.8105459213257,14.8522958755493,14.70849609375,14.4572257995605,14.3225584030151,14.1490726470947,14.1438474655151,14.1563968658447,14.3724613189697,14.4537591934204,14.4459972381592,14.4060554504395,14.3339834213257,14.2892580032349,14.27197265625,14.2805662155151,14.3150396347046,14.3075685501099,14.2582035064697,14.2568359375,14.5443363189697,14.7364749908447,14.9400873184204,15.1320810317993,15.2501459121704,15.3621101379395,15.7263669967651,15.7988767623901,15.9939451217651,16.05029296875,16.2961902618408,16.4595222473145,16.6246585845947,16.7223644256592,16.8005847930908,16.8665523529053,17.0205574035645,17.0588874816895,17.061279296875,17.0414047241211,17.056884765625,17.0570812225342,17.0617179870605,17.1086921691895,17.2881336212158,17.3241214752197,17.3350105285645,17.3682613372803,17.4205093383789,17.4580078125,17.4655284881592,17.4702625274658,17.4923324584961,17.5176773071289,17.5377922058105,17.5264644622803,17.5485343933105,17.5628433227539,17.56396484375,17.5847663879395,17.61669921875,17.6370124816895,17.68359375,17.7173328399658,17.7512683868408,17.7783699035645,17.8239250183105,17.9385261535645,18.0050792694092],[27.8095703125,27.6463851928711,27.7074699401855,27.7279300689697,27.761474609375,27.7681159973145,27.8501453399658,27.8095703125],[28.1473636627197,28.0705089569092,27.9928207397461,27.8747062683105,27.8106937408447,27.7469730377197,27.7583999633789,27.7840824127197,27.8875503540039,27.9944839477539,28.0641593933105,28.154052734375,28.1369152069092,28.1542472839355,28.1593265533447,28.1473636627197],[28.02197265625,28.0135250091553,28.0382823944092,28.11767578125,28.1763172149658,28.2031745910645,28.1992683410645,28.1559581756592,28.1111316680908,28.0834484100342,28.02197265625],[28.3799304962158,28.1514148712158,28.0619144439697,28.0320796966553,28.0071792602539,28.149169921875,28.2932605743408,28.339599609375,28.3761234283447,28.3698234558105,28.4004878997803,28.4126949310303,28.5582027435303,28.5759773254395,28.5282707214355,28.3799304962158],[28.1692867279053,28.0560054779053,28.0823726654053,28.1009273529053,28.1296863555908,28.2158203125,28.4066410064697,28.617431640625,28.7066917419434,28.7414569854736,28.7446765899658,28.738037109375,28.6912097930908,28.5851554870605,28.4093265533447,28.2534675598145,28.1692867279053],[28.4932136535645,28.4856929779053,28.5645980834961,28.7582511901855,28.8445777893066,28.8467788696289,28.7865734100342,28.7244625091553,28.6885738372803,28.6160144805908,28.569091796875,28.4932136535645],[28.9112319946289,28.845458984375,28.8690929412842,29.0133285522461,29.0561027526855,29.1189956665039,29.144287109375,29.2112293243408,29.2372093200684,29.1975078582764,29.1513671875,29.0065898895264,28.9602031707764,28.9112319946289],[38.6720695495605,38.6588363647461,38.6709976196289,38.6709976196289,38.71142578125,38.7396469116211,38.7682113647461,38.7119140625,38.7014656066895,38.6720695495605],[38.918701171875,38.8572731018066,38.8790054321289,38.9038543701172,38.973388671875,38.9817390441895,39.0311546325684,39.080810546875,39.1210441589355,39.08740234375,39.038818359375,38.9325180053711,38.918701171875],[39.7900848388672,39.7566909790039,39.7867202758789,39.7772941589355,39.76123046875,39.69775390625,39.6271476745605,39.5556640625,39.47705078125,39.3866195678711,39.333251953125,39.30126953125,39.3683586120605,39.3850593566895,39.4102516174316,39.51025390625,39.5421371459961,39.5562019348145,39.5306625366211,39.4778823852539,39.5304679870605,39.5403823852539,39.5720710754395,39.6130867004395,39.8548316955566,39.9082984924316,39.9705085754395,39.9610824584961,39.9242172241211,39.90771484375,39.8898468017578,39.8613777160645,39.8365745544434,39.7900848388672],[39.8418464660645,39.8302726745605,39.9458503723145,39.958740234375,39.9763679504395,40.0364761352539,40.0630378723145,40.0750961303711,40.0323753356934,39.917236328125,39.8975067138672,39.8418464660645],[43.4073257446289,43.37255859375,43.32470703125,43.3070297241211,43.282470703125,43.2792015075684,43.2676734924316,43.2400856018066,43.1971168518066,43.1491241455078,43.10498046875,43.0711402893066,43.0517578125,43.0367660522461,43.0326156616211,43.03759765625,43.0642585754395,43.0969734191895,43.1009750366211,43.0824699401855,43.0596199035645,43.0211410522461,42.9495086669922,42.9481925964355,42.9397926330566,42.9095230102539,42.7989730834961,42.802001953125,42.79931640625,42.80810546875,42.8288078308105,42.8253402709961,42.803955078125,42.7853012084961,42.7489242553711,42.703857421875,42.6891136169434,42.6942405700684,42.7193374633789,42.6929206848145,42.6932601928711,42.7001457214355,42.686279296875,42.6896018981934,42.7006340026855,42.8004379272461,42.8357429504395,42.8451156616211,42.8380393981934,42.7789573669434,42.7420387268066,42.713134765625,42.7099609375,42.690673828125,42.5958976745605,42.5483856201172,42.5308074951172,42.4978485107422,42.4613304138184,42.4374504089355,42.4344711303711,42.4416999816895,42.4559555053711,42.4966773986816,42.5033226013184,42.4570808410645,42.4263191223145,42.3927764892578,42.3584976196289,42.353515625,42.3860855102539,42.4209442138672,42.3902816772461,42.3457984924316,42.3404808044434,42.3621101379395,42.3930168151855,42.4084968566895,42.4130363464355,42.4292488098145,42.4560546875,42.4672355651855,42.4472198486328,42.4310073852539,42.43115234375,42.3678703308105,42.3437004089355,42.2889671325684,42.2603492736816,42.2564964294434,42.1624488830566,42.135986328125,42.1111335754395,42.0822257995605,41.9442405700684,41.8610343933105,41.7674293518066,41.4665069580078,41.3207511901855,41.2873992919922,41.1956062316895,41.0975608825684,41.0620574951172,40.8916015625,40.8228530883789,40.8038101196289,40.7223625183105,40.6862297058105,40.6304702758789,40.6133308410645,40.6222152709961,40.614501953125,40.3190422058105,40.1065902709961,40.0139617919922,39.9636764526367,39.8759269714355,39.5198745727539,39.4170913696289,39.0625953674316,38.969482421875,38.8912086486816,38.8791923522949,38.8246612548828,38.7591781616211,38.69677734375,38.6166763305664,38.585693359375,38.4356422424316,38.3172874450684,38.203125,38.15185546875,37.9920425415039,37.8861351013184,37.8502464294434,37.7699203491211,37.7116241455078,37.6310539245605,37.5962409973145,37.5807647705078,37.5713386535645,37.5611343383789,37.386962890625,37.2328605651855,36.9458503723145,36.7766609191895,36.7454566955566,36.81982421875,36.8311538696289,36.8064918518066,36.74755859375,36.7147445678711,36.7431640625,36.7584953308105,36.7557601928711,36.7079086303711,36.7398452758789,36.7560539245605,36.7181167602539,36.7002449035645,36.629150390625,36.5064468383789,36.5020523071289,36.4237785339355,36.3736343383789,36.2357406616211,36.1349105834961,36.1340827941895,36.1588897705078,36.1505851745605,36.0737800598145,36.038818359375,36.0259284973145,36.0883293151855,36.1817398071289,36.1884269714355,36.3337898254395,36.4264640808105,36.5265121459961,36.5648422241211,36.5967254638672,36.6370124816895,36.7288589477539,36.84814453125,36.8989715576172,36.91357421875,36.9084930419922,36.8316421508789,36.9546394348145,37.1942405700684,37.2491722106934,37.2789077758789,37.2149429321289,37.1984367370605,37.2087860107422,37.179443359375,37.4280281066895,37.5235824584961,37.5854988098145,37.728271484375,37.786376953125,37.9064483642578,38.00634765625,38.030029296875,38.0447273254395,38.1219749450684,38.1878929138184,38.1944351196289,38.1810073852539,38.4574203491211,38.50146484375,38.5668449401855,38.6493682861328,38.7145500183105,38.7705078125,38.8269500732422,38.9070320129395,38.9852561950684,39.0564460754395,39.1070823669434,39.1352043151855,39.338134765625,39.4651374816895,39.4783210754395,39.5361824035645,39.6447257995605,39.6615715026855,39.6806640625,39.6816864013672,39.70556640625,39.7139625549316,39.7983894348145,39.9371109008789,40.0218238830566,40.0568351745605,40.1426277160645,40.1679191589355,40.2083511352539,40.2516136169434,40.3007316589355,40.3431167602539,40.3762702941895,40.4109840393066,40.4432601928711,40.483154296875,40.6190910339355,40.654052734375,40.7774887084961,40.8783187866211,41.0091323852539,41.0380363464355,41.0624046325684,41.1077156066895,41.2145004272461,41.3037109375,41.3753890991211,41.4550323486328,41.5159187316895,41.5320320129395,41.5604476928711,41.601806640625,41.6421890258789,41.6653785705566,41.6644020080566,41.6725120544434,41.7040519714355,41.78955078125,41.8741226196289,41.9130859375,41.9423828125,41.9345703125,41.95849609375,41.9641609191895,41.9452629089355,41.9506378173828,41.9642066955566,41.9811515808105,41.9716796875,41.9552268981934,41.9293937683105,41.895263671875,41.8644065856934,41.8336944580078,41.8359870910645,41.8579559326172,41.8739738464355,41.8884773254395,41.8705558776855,41.8836402893066,41.8519058227539,41.814208984375,41.8119621276855,41.8199691772461,41.8369598388672,41.8958511352539,41.9270973205566,42.0181617736816,42.0399398803711,42.0693855285645,42.1118659973145,42.1336936950684,42.1374053955078,42.1150856018066,42.0693359375,42.052734375,42.0084953308105,41.9410629272461,41.9269027709961,41.9468765258789,42.1052742004395,42.2105979919434,42.274169921875,42.2870140075684,42.2852554321289,42.33447265625,42.358154296875,42.4117164611816,42.434814453125,42.4700698852539,42.5623550415039,42.5999031066895,42.6403312683105,42.5856437683105,42.5938491821289,42.662353515625,42.7667007446289,42.798583984375,42.8140144348145,42.8652305603027,42.9109878540039,42.9769020080566,43.0357933044434,43.1740226745605,43.2142066955566,43.2389640808105,43.3344268798828,43.3166007995605,43.3370628356934,43.3858413696289,43.3968276977539,43.4394073486328,43.4969215393066,43.5396003723145,43.5798835754395,43.6290512084961,43.6943817138672,43.7069816589355,43.7645492553711,43.7273406982422,43.7399406433105,43.69580078125,43.5946273803711,43.553955078125,43.5856437683105,43.5923805236816,43.5789070129395,43.6038551330566,43.5949211120605,43.6450691223145,43.5824699401855,43.5531730651855,43.5018539428711,43.4157218933105,43.4147453308105,43.4630851745605,43.4994163513184,43.4778785705566,43.5194816589355,43.5110321044922,43.4517097473145,43.37158203125,43.439697265625,43.4544410705566,43.4127426147461,43.3280296325684,43.3219223022461,43.3450660705566,43.4008293151855,43.4073257446289],[58.6212387084961,58.5970191955566,58.6046905517578,58.6215324401855,58.605712890625,58.4834938049316,58.4508323669434,58.4359855651855,58.3984870910645,58.372314453125,58.3638648986816,58.311279296875,58.2608871459961,58.2306632995605,58.2362289428711,58.2171363830566,58.1607398986816,58.0518074035645,57.966796875,57.93603515625,57.9313507080078,57.9632797241211,57.9951629638672,58.1153335571289,58.1543464660645,58.1716842651367,58.2133827209473,58.2623519897461,58.3016624450684,58.3045921325684,58.3158683776855,58.3488311767578,58.3866691589355,58.4971694946289,58.5142593383789,58.51025390625,58.4781265258789,58.5158233642578,58.5213890075684,58.5079612731934,58.5808601379395,58.6048812866211,58.6273918151855,58.6212387084961],[58.5503425598145,58.5399856567383,58.611083984375,58.6592292785645,58.6781272888184,58.6485862731934,58.5755386352539,58.5503425598145],[58.826904296875,58.7774391174316,58.7972183227539,58.8208999633789,58.7091789245605,58.6899909973145,58.7120590209961,58.7538070678711,58.8633766174316,58.8954582214355,58.8984832763672,58.943603515625,58.9743194580078,59.0264663696289,59.0812034606934,59.0871047973633,59.0744171142578,59.0319786071777,59.0150871276855,58.9912109375,58.9198265075684,58.8339385986328,58.826904296875],[59.4842758178711,59.4531784057617,59.403076171875,59.3744125366211,59.3575668334961,59.3432655334473,59.3278274536133,59.3017120361328,59.2970199584961,59.2776336669922,59.1926727294922,59.0519981384277,58.9449691772461,58.8862762451172,58.84130859375,58.7872543334961,58.7330551147461,58.4352531433105,58.3814926147461,58.3262710571289,58.2700729370117,58.2213401794434,58.1380844116211,58.0139198303223,57.9346199035645,57.9054679870605,57.8841323852539,57.8707046508789,57.8567352294922,57.8410148620605,57.7994155883789,57.7642059326172,57.7249526977539,57.6667938232422,57.612548828125,57.55029296875,57.5281219482422,57.5254859924316,57.5383338928223,57.5787620544434,57.6091270446777,57.6087875366211,57.5887184143066,57.531005859375,57.5444793701172,57.60107421875,57.6627464294434,57.7855453491211,57.8147430419922,57.8381843566895,57.8685531616211,57.9138145446777,57.9201698303223,57.9427757263184,58.0394515991211,58.0484886169434,58.0322227478027,58.0093765258789,57.9961395263672,57.9965858459473,58.0321273803711,58.0634231567383,58.0045890808105,57.9887237548828,57.9852523803711,57.9078598022461,57.8661613464355,57.8706092834473,57.9097671508789,58.10595703125,58.2616233825684,58.2830085754395,58.3045921325684,58.354248046875,58.3860816955566,58.3813972473145,58.3280258178711,58.28955078125,58.2661094665527,58.306640625,58.36083984375,58.4330062866211,58.5056190490723,58.5758323669434,58.6585464477539,58.7162590026855,58.7541465759277,58.7871589660645,58.7819328308105,58.7898406982422,58.8195343017578,58.920654296875,58.9604949951172,58.9992218017578,59.0321769714355,59.0696754455566,59.1075706481934,59.1956520080566,59.2423362731934,59.275146484375,59.2918968200684,59.372314453125,59.3759269714355,59.47265625,59.4556617736816,59.5220718383789,59.5211448669434,59.5594749450684,59.5979957580566,59.6390113830566,59.6275367736816,59.6346702575684,59.5539093017578,59.5539093017578,59.4717750549316,59.4506340026855,59.4504852294922,59.4142112731934,59.4698219299316,59.4842758178711],[12.4664058685303,12.3242673873901,12.1341304779053,11.912353515625,11.8578615188599,11.7237787246704,11.68603515625,11.589111328125,11.4128913879395,11.1877927780151,10.9804677963257,10.955810546875,10.9410161972046,10.9683589935303,10.9916009902954,11.0052242279053,11.0470705032349,11.0807619094849,11.0783205032349,11.0423831939697,11.00927734375,10.9979496002197,10.9993171691895,10.9602546691895,10.9032220840454,10.8459959030151,10.7869138717651,10.6213865280151,10.6000003814697,10.5675783157349,10.4917488098145,10.36962890625,10.2573728561401,10.2030754089355,10.1408205032349,10.0125980377197,9.92622089385986,9.87998008728027,9.77016639709473,9.60908222198486,9.48027324676514,9.37949275970459,9.34072208404541,9.33740234375,9.15078067779541,9.00883769989014,8.98603534698486,8.89306640625,8.78608417510986,8.7001953125,8.5908203125,8.48300743103027,8.3798828125,8.23496055603027,8.1181640625,8.026123046875,7.99707078933716,7.99707078933716,7.99707078933716,7.99707078933716,7.75932645797729,7.49047803878784,7.20786094665527,7.02602577209473,6.73725605010986,6.49726581573486,6.23466730117798,5.99721670150757,5.66826152801514,5.45542001724243,5.12167978286743,4.91201210021973,4.89990234375,4.915771484375,4.93120145797729,4.95097637176514,4.95053672790527,4.93076181411743,4.91142559051514,4.85498046875,4.84033203125,4.75039052963257,4.64448261260986,4.56332969665527,4.4453125,4.361083984375,4.32421827316284,4.2919921875,4.21225595474243,4.20166015625,4.137939453125,4.03129911422729,3.97773432731628,3.97905254364014,3.96328139305115,3.94619131088257,3.94306659698486,3.94355440139771,3.96298813819885,3.99194383621216,4.05747032165527,4.19033193588257,4.27304697036743,4.17763710021973,4.08271503448486,3.94794917106628,3.85146474838257,3.75424814224243,3.57783222198486,3.46918940544128,3.45610356330872,3.47875952720642,3.50087881088257,3.52060556411743,3.55898475646973,3.60009789466858,3.60483384132385,3.61899399757385,3.64882802963257,3.74672842025757,3.86464834213257,3.98593735694885,4.11083984375,4.25454092025757,4.41147470474243,4.42734384536743,4.43012666702271,4.43725538253784,4.44472646713257,4.44970750808716,4.46811532974243,4.50380897521973,4.61982393264771,4.70263624191284,4.80800771713257,4.95048809051514,5.10556602478027,5.15693378448486,5.20810556411743,5.278564453125,5.34399366378784,5.41909170150757,5.41328144073486,5.38515615463257,5.36489248275757,5.38408231735229,5.41206073760986,5.45791006088257,5.49228525161743,5.51103496551514,5.58120155334473,5.67314434051514,5.77490234375,5.85830116271973,6.04506874084473,6.15981435775757,6.30014657974243,6.56787157058716,6.66030263900757,6.72216749191284,6.77983379364014,6.89838886260986,7.00283241271973,7.08457040786743,7.22573232650757,7.29697275161743,7.36796855926514,7.4345703125,7.509521484375,7.67099666595459,7.69043016433716,7.707763671875,7.72373056411743,7.76064443588257,7.82373046875,7.86855459213257,7.89951181411743,7.95151329040527,8.04047775268555,8.25107383728027,8.39638614654541,8.437255859375,8.44775390625,8.44340801239014,8.43256855010986,8.43110370635986,8.44350624084473,8.49209022521973,8.54526424407959,8.58222675323486,8.67636680603027,8.75185489654541,9.041259765625,9.218505859375,9.42099571228027,9.46152400970459,9.51347637176514,9.72968769073486,9.85341739654541,9.91855525970459,10.1247568130493,10.1908693313599,10.2515621185303,10.5281257629395,10.6586427688599,10.787841796875,10.8428707122803,10.8801755905151,10.8645505905151,10.804931640625,10.7461910247803,10.7591791152954,10.810546875,10.8647947311401,10.9621095657349,11.1617670059204,11.2767572402954,11.4198732376099,11.6210451126099,11.748291015625,11.8165521621704,11.9570302963257,12.1555662155151,12.3005857467651,12.5373048782349,12.6237306594849,12.6610355377197,12.6848630905151,12.706298828125,12.7264642715454,12.7570314407349,12.8053226470947,12.9111328125,13.0933103561401,13.2710933685303,13.40576171875,13.4668455123901,13.5262689590454,13.6260747909546,13.842041015625,13.9884281158447,14.2568359375,14.2582035064697,14.3075685501099,14.3150396347046,14.2805662155151,14.27197265625,14.2892580032349,14.3339834213257,14.4060554504395,14.4459972381592,14.4537591934204,14.3724613189697,14.1563968658447,14.1438474655151,14.1490726470947,14.3225584030151,14.4572257995605,14.70849609375,14.8522958755493,14.8105459213257,14.7371101379395,14.702733039856,14.6814937591553,14.6788091659546,14.649658203125,14.4704093933105,14.4286136627197,14.4244136810303,14.4823246002197,14.5868663787842,14.6282234191895,14.6282234191895,14.5818853378296,14.5374994277954,14.4793949127197,14.4703121185303,14.5118656158447,14.5367193222046,14.5160646438599,14.4990234375,14.4990234375,14.4406728744507,14.4591312408447,14.4560546875,14.4311513900757,14.3380870819092,14.2251949310303,14.1444826126099,14.1116695404053,13.9831056594849,13.7361326217651,13.4998044967651,13.313232421875,13.1839351654053,13.02587890625,12.88232421875,12.8206052780151,12.7714347839355,12.6619625091553,12.5702152252197,12.4664058685303],[60.1078109741211,60.097900390625,60.1268043518066,60.1483383178711,60.1723136901855,60.1689949035645,60.140869140625,60.1078109741211],[60.1405258178711,60.1061477661133,60.1143531799316,60.1723136901855,60.1988296508789,60.2018051147461,60.1405258178711],[60.3707504272461,60.3475570678711,60.3558616638184,60.3033714294434,60.2699699401855,60.2285652160645,60.2028312683105,60.1655807495117,60.1656265258789,60.1969718933105,60.2367706298828,60.2648887634277,60.2863311767578,60.3148956298828,60.3558616638184,60.3707504272461],[60.336669921875,60.332275390625,60.3818359375,60.4012222290039,60.4124488830566,60.4522972106934,60.4699249267578,60.4384803771973,60.4017105102539,60.3931617736816,60.355224609375,60.336669921875],[60.5295867919922,60.4830589294434,60.4882316589355,60.4797897338867,60.5259742736816,60.6038589477539,60.62060546875,60.6382827758789,60.5955581665039,60.5295867919922],[63.2413101196289,63.22265625,63.2277870178223,63.2617645263672,63.2458992004395,63.1973648071289,63.1992225646973,63.2072257995605,63.1794891357422,63.1626930236816,63.1520042419434,63.1994590759277,63.2775344848633,63.2777328491211,63.2591285705566,63.2413101196289],[64.9910202026367,64.9578094482422,64.9785690307617,65.0428771972656,65.073974609375,65.08642578125,65.0553283691406,65.0387191772461,65.0262680053711,64.9910202026367],[69.02197265625,69.0096664428711,68.9610290527344,68.92822265625,68.9041519165039,68.8722686767578,68.8655242919922,68.8564453125,68.8400421142578,68.8138198852539,68.7714309692383,68.5376510620117,68.4883728027344,68.3513717651367,68.1897964477539,68.1179656982422,68.0618667602539,67.9291000366211,67.7539978027344,67.6885757446289,67.6682586669922,67.5474624633789,67.4264221191406,67.3243637084961,67.201416015625,67.0965805053711,66.970947265625,66.9302291870117,66.8917541503906,66.8492202758789,66.6955032348633,66.6170425415039,66.5321731567383,66.439697265625,66.3568420410156,66.276123046875,66.23486328125,66.1770553588867,66.091064453125,66.02294921875,65.7865219116211,65.7262649536133,65.6816864013672,65.6707077026367,65.6636199951172,65.6343765258789,65.6245574951172,65.5687484741211,65.4734344482422,65.3369598388672,65.2653350830078,65.2486801147461,65.2347717285156,65.223876953125,65.2047348022461,65.185302734375,65.1450653076172,65.10791015625,65.080322265625,65.0394973754883,65.001953125,64.9684143066406,64.9117660522461,64.8457565307617,64.8042984008789,64.7650451660156,64.7325744628906,64.6880874633789,64.6446228027344,64.5577087402344,64.5242691040039,64.443359375,64.3661117553711,64.2824172973633,64.2365188598633,64.2000045776367,64.14111328125,64.0772933959961,64.0206069946289,63.947509765625,63.8033180236816,63.7473182678223,63.7351570129395,63.6890144348145,63.5040512084961,63.41748046875,63.3006362915039,63.2082977294922,63.1418952941895,63.0680694580078,63.0077171325684,62.955322265625,62.9216270446777,62.8854026794434,62.7761268615723,62.691650390625,62.5678253173828,62.4813957214355,62.3237838745117,62.1275863647461,62.0682106018066,61.96484375,61.7573738098145,61.7115707397461,61.5460968017578,61.4934577941895,61.4442367553711,61.2877922058105,61.1690444946289,61.0587425231934,61.0028305053711,60.960205078125,60.9196319580078,60.8969230651855,60.745849609375,60.5361289978027,60.5328598022461,60.4989738464355,60.4907722473145,60.46484375,60.5386695861816,60.5434608459473,60.525146484375,60.4714813232422,60.455078125,60.4376945495605,60.4128952026367,60.4715843200684,60.5459938049316,60.5956077575684,60.6279296875,60.6245613098145,60.5518074035645,60.466796875,60.4240760803223,60.4065895080566,60.4749031066895,60.4742202758789,60.4252891540527,60.3715858459473,60.3415031433105,60.3467788696289,60.3146018981934,60.2675247192383,60.2674293518066,60.3332061767578,60.302490234375,60.26123046875,60.2483406066895,60.1940879821777,60.157470703125,60.1583518981934,60.1142578125,60.0462913513184,60.0212898254395,60.0423355102539,60.0091819763184,59.9656715393066,59.9681663513184,59.9862327575684,59.9257774353027,59.8449211120605,59.8160171508789,59.8263664245605,59.8688011169434,59.9126968383789,59.9722175598145,60.0218238830566,60.0413093566895,60.0472679138184,60.0985336303711,60.209716796875,60.2158203125,60.1866226196289,60.1013679504395,60.0768089294434,60.0572738647461,60.0375938415527,60.0280265808105,60.0292015075684,60.072265625,60.0902824401855,60.14697265625,60.1568832397461,60.2013206481934,60.1989250183105,60.2055168151855,60.2283706665039,60.2556610107422,60.2627449035645,60.2813491821289,60.3590774536133,60.3805656433105,60.3850135803223,60.3765640258789,60.4009284973145,60.5002937316895,60.5054206848145,60.5941390991211,60.5829086303711,60.5309600830078,60.5704116821289,60.5963859558105,60.636962890625,60.6968231201172,60.7674331665039,60.8500442504883,60.9674758911133,61.0592269897461,61.1271476745605,61.1705055236816,61.2812004089355,61.4108390808105,61.4549827575684,61.4843254089355,61.4843254089355,61.509521484375,61.5232925415039,61.5519485473633,61.5671424865723,61.5682144165039,61.5778770446777,61.591552734375,61.6668472290039,61.7027320861816,61.8116683959961,61.9149398803711,61.9896507263184,62.1126480102539,62.2238311767578,62.2773933410645,62.3425788879395,62.4140625,62.5147972106934,62.6229515075684,62.6892585754395,62.7399864196777,62.7905311584473,62.9500007629395,63.0332527160645,63.039306640625,63.1137199401855,63.1555137634277,63.2042961120605,63.2376976013184,63.2102508544922,63.2441444396973,63.3104515075684,63.3456535339355,63.377197265625,63.4379348754883,63.4547882080078,63.4423828125,63.4725570678711,63.50439453125,63.4911651611328,63.5799827575684,63.6478500366211,63.6833534240723,63.8218269348145,63.8649406433105,63.8961410522461,64.0344696044922,64.0409164428711,64.1341781616211,64.2582473754883,64.2741241455078,64.385986328125,64.5152816772461,64.6801300048828,64.7386703491211,64.801025390625,64.8062973022461,64.8520965576172,64.884033203125,64.8751907348633,64.8534622192383,64.8603515625,64.9164047241211,64.9510269165039,64.9842758178711,65.0094757080078,65.0651321411133,65.0986328125,65.1432647705078,65.2432098388672,65.3527297973633,65.479248046875,65.5462875366211,65.6603469848633,65.6563949584961,65.6707077026367,65.7571258544922,65.8316955566406,65.8591766357422,65.8583526611328,65.822021484375,65.7804641723633,65.8123550415039,65.8052749633789,65.9898452758789,66.0603561401367,66.1482467651367,66.191162109375,66.2154312133789,66.2526397705078,66.3042907714844,66.3807067871094,66.4434127807617,66.4807586669922,66.505859375,66.5766143798828,66.6280212402344,66.7068862915039,66.7757339477539,66.810546875,66.8382263183594,66.8778305053711,66.9340286254883,67.0025863647461,67.068115234375,67.12939453125,67.1841278076172,67.2339401245117,67.267822265625,67.3104934692383,67.32861328125,67.4228973388672,67.4400405883789,67.4491729736328,67.449951171875,67.4602584838867,67.4792022705078,67.5178680419922,67.5621566772461,67.5903854370117,67.6143035888672,67.6961898803711,67.7965774536133,67.8751983642578,67.9332046508789,67.9543914794922,68.017333984375,68.0886764526367,68.1303253173828,68.1366195678711,68.257568359375,68.3164520263672,68.3673324584961,68.3910217285156,68.4640579223633,68.4779815673828,68.5206069946289,68.5741195678711,68.6085433959961,68.6509780883789,68.690673828125,68.724609375,68.7874526977539,68.8288040161133,68.9069290161133,68.937744140625,68.9674835205078,68.9798355102539,69.036865234375,69.0694808959961,69.0714340209961,69.041748046875,69.054443359375,69.080810546875,69.1865692138672,69.214111328125,69.2472686767578,69.273681640625,69.2774887084961,69.2735824584961,69.2707061767578,69.1544876098633,69.0411148071289,68.9927749633789,68.8558578491211,68.776611328125,68.7198715209961,68.72021484375,68.6953125,68.6743621826172,68.642578125,68.6489715576172,68.6776351928711,68.7138671875,68.7583999633789,68.805908203125,68.7984390258789,68.7608871459961,68.7115249633789,68.6886749267578,68.65283203125,68.6064910888672,68.59326171875,68.6395950317383,68.7652893066406,68.8213348388672,68.8624496459961,68.880615234375,68.8871536254883,68.9191436767578,68.9901351928711,69.0761184692383,69.2314453125,69.2826690673828,69.3665084838867,69.588623046875,69.6526336669922,69.6915588378906,69.7146987915039,69.7819290161133,69.9150390625,69.9263153076172,69.9330596923828,69.9281234741211,69.9046859741211,69.906494140625,69.918701171875,69.9600601196289,70.042236328125,70.0648422241211,70.0616683959961,69.9716796875,69.8714370727539,69.82275390625,69.7314910888672,69.6714324951172,69.4729995727539,69.3939437866211,69.36669921875,69.2879867553711,69.1769104003906,69.1189956665039,69.0605926513672,69.02197265625],[-21.6950206756592,-21.7058601379395,-21.7023429870605,-21.6800785064697,-21.66748046875,-21.6759757995605,-21.6950206756592],[-20.6677742004395,-20.6705074310303,-20.6703128814697,-20.6667957305908,-20.66015625,-20.65283203125,-20.6452140808105,-20.6457023620605,-20.65234375,-20.6597652435303,-20.6677742004395],[-19.166015625,-19.1749992370605,-19.1649417877197,-19.1513671875,-19.1371097564697,-19.1188468933105,-19.1092777252197,-19.1129875183105,-19.1187496185303,-19.1251964569092,-19.1431636810303,-19.1541004180908,-19.1570320129395,-19.166015625],[-18.9741230010986,-19.01708984375,-19.0456047058105,-19.0101566314697,-19.0037097930908,-19.0665035247803,-19.09228515625,-19.1117191314697,-19.1214847564697,-19.1516609191895,-19.12158203125,-19.10107421875,-19.0601577758789,-19.0662117004395,-19.0279312133789,-18.9696292877197,-18.95703125,-18.9344730377197,-18.9507808685303,-18.9741230010986],[-18.9403324127197,-18.9698238372803,-18.9681644439697,-18.95556640625,-18.9617176055908,-18.9708003997803,-18.9913082122803,-19.0029296875,-18.9987316131592,-18.9784183502197,-18.9641609191895,-18.9432621002197,-18.9242191314697,-18.9403324127197],[-18.23388671875,-18.25244140625,-18.22216796875,-18.2020511627197,-18.1863288879395,-18.19140625,-18.23388671875],[-18.1023445129395,-18.1104488372803,-18.0305652618408,-17.9990234375,-17.970703125,-17.9440422058105,-17.9895496368408,-18.0652332305908,-18.1023445129395],[-17.9766597747803,-17.9917964935303,-17.9883785247803,-17.9723644256592,-17.9441413879395,-17.92041015625,-17.9473628997803,-17.9766597747803],[-17.9527339935303,-17.9632816314697,-17.9208984375,-17.8757820129395,-17.8864269256592,-17.9299812316895,-17.9527339935303],[-17.72900390625,-17.7467784881592,-17.6857414245605,-17.6244144439697,-17.6188468933105,-17.6812515258789,-17.72900390625],[-17.3719730377197,-17.4162101745605,-17.4353523254395,-17.4384765625,-17.5230464935303,-17.5958003997803,-17.6514644622803,-17.6990222930908,-17.7493152618408,-17.83935546875,-17.9328117370605,-18.0808601379395,-18.1089839935303,-18.1123046875,-18.1389636993408,-18.1242198944092,-18.13525390625,-18.1839847564697,-18.2501945495605,-18.2503910064697,-18.2640628814697,-18.2548828125,-18.2195320129395,-18.1810550689697,-18.1482429504395,-18.1207027435303,-18.0775394439697,-17.9686527252197,-17.9149398803711,-17.8634777069092,-17.8460941314697,-17.8200206756592,-17.7860336303711,-17.7623043060303,-17.7537097930908,-17.7373046875,-17.68212890625,-17.6316394805908,-17.53955078125,-17.4610347747803,-17.3884773254395,-17.3951168060303,-17.3392581939697,-17.31298828125,-17.3291015625,-17.3719730377197],[-17.3667984008789,-17.3938465118408,-17.2561511993408,-17.25732421875,-17.2715816497803,-17.3062496185303,-17.3667984008789],[-17.2728519439697,-17.3070316314697,-17.294921875,-17.2375011444092,-17.2126941680908,-17.1824226379395,-17.1613292694092,-17.1483383178711,-17.1820316314697,-17.2083988189697,-17.2230472564697,-17.2486324310303,-17.2728519439697],[-17.1470699310303,-17.1638679504395,-17.0842761993408,-17.0593738555908,-17.05419921875,-17.0486335754395,-17.1048812866211,-17.1470699310303],[-16.9630851745605,-17.0002937316895,-16.9640617370605,-16.8759765625,-16.7857418060303,-16.7003898620605,-16.6882820129395,-16.7653312683105,-16.8502941131592,-16.9248046875,-16.9630851745605],[-16.5400390625,-16.5412101745605,-16.5221672058105,-16.4888668060303,-16.44140625,-16.4315433502197,-16.4791011810303,-16.5028324127197,-16.5400390625],[-16.1682624816895,-16.3016605377197,-16.3703117370605,-16.4462890625,-16.5277347564697,-16.6369152069092,-16.7474594116211,-16.6319332122803,-16.5375003814697,-16.5184574127197,-16.5194339752197,-16.5516605377197,-16.5835933685303,-16.6669921875,-16.7444343566895,-16.7369136810303,-16.7435550689697,-16.7870121002197,-16.8060550689697,-16.8065433502197,-16.7919921875,-16.7697257995605,-16.7180652618408,-16.7103519439697,-16.7126960754395,-16.8135738372803,-16.9001941680908,-16.9040050506592,-16.8860359191895,-16.9521484375,-16.9761714935303,-16.9200191497803,-16.8512706756592,-16.8005847930908,-16.7878913879395,-16.72607421875,-16.7004890441895,-16.6638660430908,-16.6218757629395,-16.6485347747803,-16.6656265258789,-16.6341800689697,-16.6314449310303,-16.5400390625,-16.4828128814697,-16.4375,-16.4051761627197,-16.3986339569092,-16.3798828125,-16.2941398620605,-16.2499027252197,-16.2232437133789,-16.2076168060303,-16.2214832305908,-16.2142581939697,-16.1529293060303,-16.1260757446289,-16.1260757446289,-16.1492195129395,-16.1682624816895],[-12.5054683685303,-12.515625,-12.50732421875,-12.4911127090454,-12.4875001907349,-12.4769535064697,-12.4823246002197,-12.4928712844849,-12.5054683685303],[-52.2316398620605,-52.2508773803711,-52.2421875,-52.2046852111816,-52.1561508178711,-52.1341781616211,-52.1414070129395,-52.1800765991211,-52.2316398620605],[-52.0110359191895,-52.0990219116211,-52.0710945129395,-52.0284194946289,-51.9994163513184,-52.0015640258789,-52.0110359191895],[-51.7857437133789,-51.7995109558105,-51.7942390441895,-51.89697265625,-51.9462890625,-51.9424819946289,-51.8752937316895,-51.8394508361816,-51.81396484375,-51.7857437133789],[-51.4619140625,-51.48095703125,-51.4105453491211,-51.3880844116211,-51.4033203125,-51.4459953308105,-51.4392585754395,-51.3957023620605,-51.4105453491211,-51.3599586486816,-51.3835945129395,-51.4275398254395,-51.478515625,-51.5109367370605,-51.55615234375,-51.5926742553711,-51.6265602111816,-51.6808586120605,-51.8077163696289,-51.9695281982422,-51.9840812683105,-51.9938468933105,-51.9827156066895,-51.9864273071289,-52.07373046875,-52.1399421691895,-52.1540031433105,-52.1602516174316,-52.1703147888184,-52.1947288513184,-52.1883773803711,-52.1477546691895,-52.0573234558105,-51.9464836120605,-51.9515647888184,-51.8771476745605,-51.8395500183105,-51.80126953125,-51.77197265625,-51.7337875366211,-51.7166023254395,-51.7183609008789,-51.7351570129395,-51.7565422058105,-51.7126922607422,-51.6963882446289,-51.6971702575684,-51.6560554504395,-51.6388664245605,-51.5804672241211,-51.5450210571289,-51.4854507446289,-51.4631843566895,-51.4278297424316,-51.3578109741211,-51.3542976379395,-51.3994140625,-51.4619140625],[-51.3959007263184,-51.39599609375,-51.3631820678711,-51.2805633544922,-51.2734375,-51.3079109191895,-51.3425750732422,-51.3959007263184],[-51.2699203491211,-51.3285179138184,-51.30810546875,-51.32421875,-51.373046875,-51.4183616638184,-51.4118156433105,-51.4239273071289,-51.4835929870605,-51.5090827941895,-51.57470703125,-51.57861328125,-51.5510749816895,-51.5060539245605,-51.45751953125,-51.4170913696289,-51.4046859741211,-51.3843727111816,-51.4035148620605,-51.5179672241211,-51.5337905883789,-51.5832023620605,-51.6045875549316,-51.6361312866211,-51.6845703125,-51.7091789245605,-51.7434577941895,-51.7654304504395,-51.8224601745605,-51.86376953125,-51.9362297058105,-51.9948234558105,-52.0230484008789,-52.0992164611816,-52.0079078674316,-52.0176773071289,-52.1730461120605,-52.2017593383789,-52.18310546875,-52.1959953308105,-52.3080062866211,-52.2364273071289,-52.1343765258789,-52.0772438049316,-51.9706039428711,-51.9253883361816,-51.7804718017578,-51.7373046875,-51.7124977111816,-51.7041015625,-51.6854476928711,-51.6501960754395,-51.5897483825684,-51.4914054870605,-51.35791015625,-51.2720718383789,-51.2699203491211],[-21.33935546875,-21.3690433502197,-21.3583011627197,-21.2736339569092,-21.2173824310303,-21.0583992004395,-21.00244140625,-20.9041004180908,-20.8651371002197,-20.8795890808105,-20.90625,-21.021484375,-21.1385746002197,-21.27783203125,-21.33935546875],[-12.9767570495605,-12.9849605560303,-12.95849609375,-12.8956050872803,-12.8350582122803,-12.7861328125,-12.7012691497803,-12.6530275344849,-12.7091798782349,-12.7129888534546,-12.7521486282349,-12.8243169784546,-12.8479499816895,-12.9202146530151,-12.9767570495605],[4.06127977371216,4.03969764709473,3.99267578125,3.92993211746216,3.86958003044128,3.82856488227844,3.77695322036743,3.735107421875,3.70200181007385,3.64687538146973,3.45229482650757,3.36469769477844,3.27167963981628,3.23710942268372,3.18173837661743,3.11772465705872,3.05156230926514,2.97221684455872,2.90385723114014,2.86416053771973,2.64765620231628,2.57314419746399,2.52890610694885,2.42573261260986,2.36367201805115,2.31718730926514,2.26665043830872,2.21152377128601,2.18354511260986,2.18173861503601,2.20170903205872,2.21132826805115,2.20488262176514,2.23227572441101,2.29521512985229,2.33974599838257,2.32421875,2.27944326400757,2.25312519073486,2.26191425323486,2.29291987419128,2.30854511260986,2.33500981330872,2.35483431816101,2.34599614143372,2.31293964385986,2.27827143669128,2.23256826400757,2.15048837661743,2.12104511260986,2.13706040382385,2.15332055091858,2.15424799919128,2.20751929283142,2.24545884132385,2.29306650161743,2.31376957893372,2.32675743103027,2.33579087257385,2.34257817268372,2.34331035614014,2.41611313819885,2.46152353286743,2.71372056007385,2.81787109375,2.87485361099243,2.99360370635986,3.13818383216858,3.17875957489014,3.35332012176514,3.44853496551514,3.53051781654358,3.58955073356628,3.62041044235229,3.62939429283142,3.70595693588257,3.76943373680115,3.83442378044128,3.90107417106628,4.05410146713257,4.14003896713257,4.17094707489014,4.20249032974243,4.24140596389771,4.337646484375,4.42802762985229,4.48501014709473,4.5830078125,4.69199228286743,4.74931669235229,4.83652353286743,4.91469717025757,4.95878934860229,5.01347637176514,5.18740224838257,5.28823232650757,5.35898447036743,5.41181612014771,5.676025390625,5.76899433135986,5.7822265625,5.5634765625,5.54326200485229,5.42504835128784,5.27348613739014,5.02133798599243,4.94218730926514,4.87612295150757,4.77089881896973,4.86279296875,4.71738290786743,4.64599609375,4.51440477371216,4.42988252639771,4.38622999191284,4.352294921875,4.39907264709473,4.43613290786743,4.52431631088257,4.63373994827271,4.63569355010986,4.57050752639771,4.28681612014771,4.22880840301514,4.13876962661743,4.09848594665527,4.06127977371216],[14.4944829940796,14.4374027252197,14.42626953125,14.4737787246704,14.4670906066895,14.5095710754395,14.5296869277954,14.6019048690796,14.6212406158447,14.6523914337158,14.8043937683105,14.8485832214355,14.8719244003296,14.8752927780151,14.826171875,14.7562494277954,14.7551755905151,14.7353506088257,14.6861820220947,14.6445322036743,14.6137208938599,14.4944829940796],[15.8899412155151,15.8860349655151,15.8946781158447,15.9548826217651,15.9962396621704,16.0062980651855,15.9599113464355,15.921238899231,15.8899412155151],[16.0069332122803,15.9620599746704,15.9759283065796,16.0620613098145,16.3009757995605,16.3404788970947,16.3552742004395,16.3259754180908,16.2921867370605,16.2708988189697,16.2271480560303,16.0477523803711,16.0069332122803],[16.2304191589355,16.2192859649658,16.22802734375,16.2996101379395,16.3602046966553,16.4337882995605,16.4776859283447,16.5066413879395,16.4683094024658,16.4134273529053,16.3631839752197,16.256103515625,16.2304191589355],[42.805419921875,42.6585960388184,42.6155738830566,42.5855941772461,42.5526390075684,42.1609382629395,42.1297378540039,41.972412109375,41.9262199401855,41.731201171875,41.6788063049316,41.6271476745605,41.4600563049316,41.3849105834961,41.4765625,41.5161590576172,41.5588874816895,41.58837890625,41.627685546875,41.6685562133789,41.70068359375,41.7371101379395,41.7614250183105,41.8040046691895,41.8704109191895,41.9251441955566,41.92236328125,41.9307136535645,41.9591293334961,41.9955558776855,42.0431175231934,42.0956077575684,42.1182098388672,42.1608390808105,42.2187995910645,42.2584495544434,42.2840347290039,42.3434104919434,42.3447265625,42.3577156066895,42.3853034973145,42.4265632629395,42.5497550964355,42.60791015625,42.6453094482422,42.6616706848145,42.7049789428711,42.73291015625,42.7291984558105,42.7124519348145,42.6946296691895,42.7131805419922,42.7668952941895,42.8140640258789,42.9438018798828,43.0173835754395,43.021484375,42.9810066223145,42.9452133178711,42.8605003356934,42.805419921875],[45.904052734375,45.8166007995605,45.8971176147461,45.9676742553711,46.032958984375,46.0503921508789,46.002685546875,45.904052734375],[49.5382804870605,49.5050773620605,49.48974609375,49.4775390625,49.4546394348145,49.4454612731934,49.4546394348145,49.4852066040039,49.4989280700684,49.4943389892578,49.4775390625,49.4527359008789,49.4581527709961,49.44287109375,49.3946800231934,49.34619140625,49.3196792602539,49.2908706665039,49.1605987548828,49.1541481018066,49.1739273071289,49.2019538879395,49.20751953125,49.1946296691895,49.1798820495605,49.1234359741211,49.1126937866211,49.1248550415039,49.1275405883789,49.1136245727539,49.1295394897461,49.153076171875,49.1521949768066,49.0863761901855,49.061767578125,49.0418968200684,49.0109367370605,48.9858894348145,48.9735832214355,48.8864250183105,48.873291015625,48.6985359191895,48.6360359191895,48.5468254089355,48.4100112915039,48.280029296875,48.1567878723145,48.0643081665039,48.0025863647461,47.9056625366211,47.7736320495605,47.6738777160645,47.6065406799316,47.5927238464355,47.54736328125,47.5076637268066,47.4551734924316,47.43310546875,47.42578125,47.4327163696289,47.4537124633789,47.4732437133789,47.4898452758789,47.4893569946289,47.4532241821289,47.3942375183105,47.3612327575684,47.3525390625,47.3394546508789,47.322509765625,47.3020515441895,47.2671852111816,47.1631851196289,47.0582504272461,47.0265159606934,47.0043487548828,46.9483413696289,46.9258804321289,46.832275390625,46.7554206848145,46.6830558776855,46.6110343933105,46.5669937133789,46.5160636901855,46.4585418701172,46.4281730651855,46.3786125183105,46.337646484375,46.2793960571289,46.2380867004395,46.2146949768066,46.1512184143066,46.142333984375,46.1470222473145,46.1930694580078,46.2522430419922,46.3084487915039,46.3194351196289,46.3326148986816,46.3936996459961,46.4305191040039,46.4373512268066,46.415771484375,46.4066429138184,46.3691902160645,46.31396484375,46.2751922607422,46.1651344299316,46.1306648254395,46.0894050598145,46.0517578125,46.0171394348145,45.9588356018066,45.92578125,45.8683624267578,45.8145523071289,45.7800788879395,45.7408676147461,45.7100105285645,45.6703605651855,45.58056640625,45.50048828125,45.4236831665039,45.4009246826172,45.3817367553711,45.3490257263184,45.2399406433105,45.2226066589355,45.215576171875,45.1356468200684,45.1453132629395,45.144287109375,45.1179695129395,45.0681648254395,45.0226097106934,44.9729995727539,44.92138671875,44.8831520080566,44.8603019714355,44.8587379455566,44.8450202941895,44.8272972106934,44.7166976928711,44.68896484375,44.6771469116211,44.6316413879395,44.5645484924316,44.5106925964355,44.4632797241211,44.4281730651855,44.3920440673828,44.3357429504395,44.280029296875,44.2017097473145,44.1379852294922,44.1273918151855,44.1683616638184,44.1648445129395,44.1160125732422,44.0831527709961,44.0336418151855,43.9654273986816,43.9110832214355,43.8648910522461,43.8229484558105,43.7671394348145,43.7504386901855,43.761474609375,43.7708969116211,43.7653312683105,43.7532234191895,43.7317390441895,43.6960945129395,43.6591300964355,43.4383316040039,43.3735847473145,43.3345680236816,43.2616691589355,43.1990699768066,43.1692886352539,43.1387214660645,43.0723648071289,43.1009750366211,43.097900390625,43.1778335571289,43.228515625,43.3449249267578,43.3524894714355,43.3489723205566,43.3666038513184,43.4062995910645,43.4445304870605,43.4269523620605,43.4269523620605,43.41162109375,43.3939437866211,43.4052238464355,43.4014167785645,43.37890625,43.373291015625,43.3871116638184,43.4472160339355,43.4563941955566,43.4796371459961,43.503662109375,43.5630378723145,43.5818367004395,43.5930671691895,43.5630874633789,43.516357421875,43.4616203308105,43.1932106018066,43.0807609558105,42.9151382446289,42.837890625,42.5908699035645,42.461181640625,42.43115234375,42.4310073852539,42.4472198486328,42.4672355651855,42.4560546875,42.4292488098145,42.4130363464355,42.4084968566895,42.3930168151855,42.3621101379395,42.3404808044434,42.3457984924316,42.3902816772461,42.4209442138672,42.3860855102539,42.353515625,42.3584976196289,42.3927764892578,42.4263191223145,42.4570808410645,42.5033226013184,42.525634765625,42.5567359924316,42.575927734375,42.6044425964355,42.635009765625,42.6427268981934,42.6216773986816,42.5958976745605,42.690673828125,42.7099609375,42.713134765625,42.7420387268066,42.7789573669434,42.8380393981934,42.8451156616211,42.8357429504395,42.8004379272461,42.7006340026855,42.6896018981934,42.686279296875,42.7001457214355,42.6932601928711,42.6929206848145,42.7193374633789,42.6942405700684,42.6891136169434,42.703857421875,42.7489242553711,42.7853012084961,42.803955078125,42.8253402709961,42.8288078308105,42.80810546875,42.79931640625,42.802001953125,42.7989730834961,42.9095230102539,42.9397926330566,42.9481925964355,42.9495086669922,43.0211410522461,43.0596199035645,43.0824699401855,43.1009750366211,43.0969734191895,43.0642585754395,43.03759765625,43.0326156616211,43.0367660522461,43.0517578125,43.0711402893066,43.10498046875,43.1491241455078,43.1971168518066,43.2400856018066,43.2676734924316,43.2792015075684,43.282470703125,43.3070297241211,43.32470703125,43.37255859375,43.4073257446289,43.4380378723145,43.5637702941895,44.0202140808105,44.5598640441895,44.6618194580078,44.6898422241211,44.7640151977539,44.7264671325684,44.6866226196289,44.6666984558105,45.1614723205566,45.3426284790039,45.5324211120605,45.4570808410645,45.3806648254395,45.3143539428711,45.0934600830078,45.047119140625,45.0005836486816,45.0513687133789,45.0901870727539,45.3846206665039,45.468017578125,45.5381813049316,45.6859359741211,45.7144508361816,45.7708969116211,45.7685050964355,45.7410621643066,45.7726554870605,45.8056640625,45.9253425598145,46.204833984375,46.252685546875,46.3113784790039,46.3245124816895,46.326904296875,46.35009765625,46.5148468017578,46.684814453125,46.810302734375,46.8650398254395,46.9205055236816,47.0376434326172,47.1116218566895,47.1263160705566,47.1629409790039,47.2239227294922,47.262939453125,47.2735824584961,47.2606430053711,47.225341796875,47.2159690856934,47.3106918334961,47.2787590026855,47.2909660339355,47.3120613098145,47.381591796875,47.4129409790039,47.4708976745605,47.5116233825684,47.5270500183105,47.5261688232422,47.5138664245605,47.5372543334961,47.601806640625,47.6255378723145,47.6144523620605,47.60107421875,47.621337890625,47.6946792602539,47.6941413879395,47.6851081848145,47.7133331298828,47.7204093933105,47.7110366821289,47.7531242370605,47.8375473022461,47.8478507995605,47.8096199035645,47.8229026794434,47.87744140625,47.9689445495605,48.0395011901855,48.0857925415039,48.0967254638672,48.1288070678711,48.1699714660645,48.2179679870605,48.229736328125,48.2469749450684,48.2900390625,48.3097152709961,48.2992706298828,48.2930679321289,48.3036613464355,48.3470687866211,48.3567390441895,48.3676261901855,48.372314453125,48.3575210571289,48.3631362915039,48.4100112915039,48.4502449035645,48.5398941040039,48.6199722290039,48.70751953125,48.6947288513184,48.7104988098145,48.7656707763672,48.8129425048828,48.8408203125,48.7906761169434,48.60107421875,48.5368156433105,48.6482925415039,48.64501953125,48.5820808410645,48.6351051330566,48.6971168518066,48.6688461303711,48.6305160522461,48.6414070129395,48.652587890625,48.6976051330566,48.8055152893066,49.202392578125,49.3131828308105,49.4901351928711,49.5951156616211,49.6313972473145,49.6837882995605,49.6809577941895,49.6676750183105,49.707275390625,49.68017578125,49.5982398986816,49.494873046875,49.44482421875,49.3878898620605,49.3931655883789,49.3597183227539,49.3545417785645,49.2967758178711,49.3302268981934,49.335636138916,49.4015159606934,49.4483871459961,49.4731941223145,49.4632797241211,49.5084495544434,49.5575218200684,49.6015625,49.7030258178711,49.8629379272461,49.9102020263672,49.9982414245605,50.0885276794434,50.205078125,50.230712890625,50.252197265625,50.2939453125,50.7392578125,50.8194847106934,50.885009765625,50.9356956481934,50.9906234741211,51.0665054321289,51.0971183776855,51.0495109558105,50.9885749816895,50.9552726745605,50.9117698669434,50.8759269714355,50.8114280700684,50.7506332397461,50.7117691040039,50.7160148620605,50.72705078125,50.7668952941895,50.7794456481934,50.7489280700684,50.731689453125,50.6629371643066,50.5911636352539,50.5315437316895,50.5073738098145,50.4994659423828,50.4773445129395,50.4573249816895,50.3248062133789,50.3060531616211,50.3216781616211,50.343505859375,50.3469734191895,50.3385734558105,50.3359375,50.3213386535645,50.2464828491211,50.2217750549316,50.1784172058105,50.143798828125,50.1298828125,50.0941390991211,50.0528335571289,50.0238800048828,50,49.9844703674316,49.9715843200684,49.9602546691895,49.9449691772461,49.9602546691895,50.00244140625,50.046875,50.0970726013184,50.1390647888184,50.1531753540039,50.1358909606934,49.9595680236816,49.9145050048828,49.8471183776855,49.7881355285645,49.7892570495605,49.7783699035645,49.7565422058105,49.7214851379395,49.6892585754395,49.6779289245605,49.6509780883789,49.6198234558105,49.5544891357422,49.5108871459961,49.5110321044922,49.5282211303711,49.5392074584961,49.5382804870605],[61.4446258544922,61.414306640625,61.4176750183105,61.4522438049316,61.5347671508789,61.6029319763184,61.6343307495117,61.6308135986328,61.6027793884277,61.5843734741211,61.5705070495605,61.536376953125,61.4959487915039,61.4446258544922],[61.8059577941895,61.768310546875,61.7686500549316,61.8153305053711,61.8407707214355,61.8622512817383,61.8991203308105,61.9037132263184,61.8953590393066,61.8617668151855,61.8267097473145,61.8059577941895],[62.1393051147461,62.1005363464355,62.0732460021973,62.0468254089355,62.0400352478027,62.046142578125,62.0748023986816,62.1402816772461,62.138671875,62.1512184143066,62.1393051147461],[62.2278785705566,62.0936050415039,62.0943374633789,62.1314926147461,62.1391143798828,62.1192855834961,62.0954093933105,62.0804214477539,61.9903793334961,61.9641609191895,61.9514617919922,61.9774436950684,62.093994140625,62.2855949401855,62.3162612915039,62.2659645080566,62.2278785705566],[62.2586441040039,62.1865234375,62.1978530883789,62.2056121826172,62.2245140075684,62.2781219482422,62.3556671142578,62.2918930053711,62.2586441040039],[5.32573270797729,5.27724599838257,5.30078125,5.31792020797729,5.33501005172729,5.32573270797729],[6.81367206573486,6.791015625,6.80126953125,6.8828125,6.90463876724243,6.94482469558716,6.97773456573486,6.95107412338257,6.89316415786743,6.85463809967041,6.81367206573486],[7.34619140625,7.33300828933716,7.33808612823486,7.34550809860229,7.35131788253784,7.35722684860229,7.37539005279541,7.37924766540527,7.38872003555298,7.39042949676514,7.37924766540527,7.36284160614014,7.34619140625],[7.43203115463257,7.42675828933716,7.43178701400757,7.45737314224243,7.46616172790527,7.46708965301514,7.46015644073486,7.45385694503784,7.43203115463257],[9.50068378448486,9.41904258728027,9.44575214385986,9.49458026885986,9.55019474029541,9.58359336853027,9.59331035614014,9.54721736907959,9.50737285614014,9.50068378448486],[2.16157221794128,2.09169936180115,1.92041015625,1.78857421875,1.64809596538544,1.53505873680115,1.45458972454071,1.36669921875,1.30542016029358,1.27924799919128,1.24843728542328,1.24101567268372,1.26777338981628,1.31459951400757,1.38227558135986,1.41875004768372,1.39589822292328,1.37021481990814,1.32255840301514,1.12084984779358,1.09023439884186,1.00444316864014,0.901464879512787,0.849121153354645,0.81147438287735,0.755712985992432,0.673828125,0.624218642711639,0.587451338768005,0.551122963428497,0.536572098731995,0.514990508556366,0.427734196186066,0.35380831360817,0.283984243869781,0.190820157527924,0.0752929598093033,-0.0908201113343239,-0.203320384025574,-0.24257829785347,-0.270116925239563,-0.292382657527924,-0.361913949251175,-0.427344024181366,-0.468554556369781,-0.518652260303497,-0.573437333106995,-0.618359565734863,-0.798828065395355,-0.972070515155792,-1.10390627384186,-1.22978520393372,-1.41318368911743,-1.52509784698486,-1.59335958957672,-1.64697289466858,-1.71152353286743,-1.89003884792328,-1.92021489143372,-1.95351564884186,-2.00146484375,-2.07675766944885,-2.17988252639771,-2.21757817268372,-2.26552724838257,-2.30058598518372,-2.35419940948486,-2.41796898841858,-2.46689438819885,-2.49062538146973,-2.46543002128601,-2.42988324165344,-2.37451195716858,-2.33017587661743,-2.28369116783142,-2.16376924514771,-2.13847684860229,-2.18750023841858,-2.27861309051514,-2.39540982246399,-2.40478515625,-2.36914038658142,-2.31337881088257,-2.17626976966858,-2.06328129768372,-1.93183600902557,-1.86943340301514,-1.82958996295929,-1.82685530185699,-1.90000021457672,-1.92890644073486,-1.99033224582672,-2.04755854606628,-2.07529330253601,-2.11201190948486,-2.16923809051514,-2.24560546875,-2.32998061180115,-2.41259741783142,-2.38281226158142,-2.34482407569885,-2.35146474838257,-2.39472651481628,-2.36455106735229,-2.34257817268372,-2.36093735694885,-2.39707016944885,-2.59540987014771,-2.67099618911743,-2.76962876319885,-2.83671855926514,-2.85537123680115,-2.88662075996399,-2.93652319908142,-2.98310542106628,-3.01123070716858,-3.06308555603027,-3.126953125,-3.17695307731628,-3.22910118103027,-3.28320288658142,-3.31855487823486,-3.35097646713257,-3.42021489143372,-3.47861361503601,-3.53144526481628,-3.580078125,-3.62539052963257,-3.66591787338257,-3.69667935371399,-3.69023418426514,-3.69453096389771,-3.68203115463257,-3.52500033378601,-3.52031230926514,-3.64111375808716,-3.69082045555115,-3.76201176643372,-3.91630887985229,-3.82646489143372,-3.66210961341858,-3.56132817268372,-3.39804697036743,-3.27802753448486,-3.01308584213257,-2.74833989143372,-2.51855444908142,-2.46757793426514,-2.47382831573486,-2.58837866783142,-2.57558560371399,-2.54990243911743,-2.48984360694885,-2.44257807731628,-2.41308569908142,-2.41562509536743,-2.36708974838257,-2.29316377639771,-2.22998046875,-2.16386723518372,-2.02763676643372,-1.97499978542328,-1.90302705764771,-1.89365243911743,-1.96230483055115,-1.93496096134186,-1.89462876319885,-1.82939469814301,-1.82509744167328,-1.77929699420929,-1.72626972198486,-1.52773416042328,-1.37910139560699,-1.30888652801514,-1.63203132152557,-1.63759756088257,-1.63457012176514,-1.59833967685699,-1.55517590045929,-1.50888669490814,-1.53017604351044,-1.53457009792328,-1.51523447036743,-1.48193359375,-1.32499980926514,-1.33291018009186,-1.36093747615814,-1.37421870231628,-1.38242161273956,-1.29834008216858,-1.07148432731628,-1.02499973773956,-0.946093678474426,-0.913574278354645,-0.591015696525574,-0.614941298961639,-0.708398699760437,-0.68876975774765,-0.634668231010437,-0.636718809604645,-0.624316573143005,-0.573339939117432,-0.351269543170929,-0.0582520477473736,0.115820556879044,0.288525491952896,0.343603372573853,0.307226836681366,0.245898574590683,0.200439542531967,0.159765854477882,0.148926064372063,0.084961049258709,0.0442380718886852,0.125586047768593,0.194970652461052,0.219873130321503,0.192480593919754,0.295947015285492,0.361913949251175,0.486718475818634,0.552099823951721,0.610840022563934,0.664843618869781,0.658691167831421,0.594189345836639,0.567724406719208,0.576513588428497,0.631640672683716,0.779443204402924,0.991308689117432,1.03198230266571,1.04667985439301,1.07470726966858,1.07998073101044,1.07470726966858,1.02568387985229,0.998730778694153,0.986230552196503,0.960107624530792,0.967138469219208,0.997705161571503,1.00400364398956,1.00356411933899,1.00307607650757,1.00214838981628,1.00126957893372,1.00039064884186,0.999707043170929,1.12075185775757,1.30761742591858,1.52836894989014,1.74018561840057,1.93588864803314,2.16743183135986,2.23383808135986,2.26142621040344,2.29970693588257,2.30219721794128,2.28515625,2.28750014305115,2.28437542915344,2.29599595069885,2.28134775161743,2.26503896713257,2.25678730010986,2.24677729606628,2.25942349433899,2.25644540786743,2.22421860694885,2.16157221794128],[50.6902351379395,50.6557121276855,50.615234375,50.5992202758789,50.5888175964355,50.5885276794434,50.6697769165039,50.6661148071289,50.7033233642578,50.7335472106934,50.7734832763672,50.7347183227539,50.6902351379395],[53.3214340209961,53.3028297424316,53.3057594299316,53.2643051147461,53.2180671691895,53.1724128723145,53.1341781616211,53.1780281066895,53.1763648986816,53.2604484558105,53.386474609375,53.4192886352539,53.417236328125,53.3214340209961],[54.0887184143066,54.0948715209961,54.0770988464355,54.0606422424316,54.0636253356934,54.0572776794434,54.0586433410645,54.0847663879395,54.1634292602539,54.1847152709961,54.1956062316895,54.21435546875,54.2686538696289,54.2940406799316,54.3291015625,54.3743171691895,54.4066886901855,54.4082527160645,54.3553695678711,54.3018074035645,54.27490234375,54.239501953125,54.214111328125,54.156005859375,54.1334457397461,54.1212425231934,54.1373062133789,54.1335945129395,54.1438484191895,54.1866722106934,54.2152824401855,54.2837905883789,54.2965812683105,54.4142608642578,54.4535140991211,54.4769554138184,54.512451171875,54.5712394714355,54.5949211120605,54.6158218383789,54.6396980285645,54.6660652160645,54.683349609375,54.6983413696289,54.7178726196289,54.7192840576172,54.71044921875,54.72802734375,54.7457046508789,54.7679672241211,54.825439453125,54.8771018981934,54.9051284790039,55.0033187866211,55.0276870727539,55.0919914245605,55.056884765625,55.0482902526855,55.0806159973145,55.1825180053711,55.1889152526855,55.1806640625,55.1934585571289,55.2410163879395,55.2417984008789,55.2168426513672,55.2173843383789,55.14453125,55.0296859741211,54.9162101745605,54.8174781799316,54.757080078125,54.7246589660645,54.6843757629395,54.6413078308105,54.6630401611328,54.6730499267578,54.6634292602539,54.61962890625,54.5540542602539,54.5001983642578,54.441650390625,54.460205078125,54.5125961303711,54.5367164611816,54.5497550964355,54.4778823852539,54.3817367553711,54.3726577758789,54.3709983825684,54.2725601196289,54.245849609375,54.23583984375,54.2009773254395,54.1560554504395,54.0890655517578,54.05126953125,54.0588874816895,54.0887184143066],[55.4488296508789,55.4480934143066,55.4567375183105,55.4810562133789,55.6183586120605,55.6669425964355,55.6907234191895,55.7091827392578,55.7169418334961,55.6909637451172,55.6667976379395,55.573974609375,55.4943389892578,55.4488296508789],[55.9305686950684,55.8021469116211,55.7225112915039,55.6953125,55.6575202941895,55.6072273254395,55.60693359375,55.619140625,55.6703147888184,55.7283706665039,55.7725105285645,55.7806129455566,55.7743644714355,55.7042503356934,55.6973152160645,55.7115707397461,55.7689933776855,55.8082542419434,55.8323745727539,55.8546409606934,55.871337890625,55.8737297058105,55.8564949035645,55.9045906066895,55.9305686950684],[55.8145523071289,55.8038101196289,55.8067893981934,55.8228988647461,55.84765625,55.8931159973145,55.9256324768066,55.9747543334961,55.9921875,56.0044441223145,56.0452651977539,56.1087875366211,56.1203117370605,56.1185569763184,56.005615234375,55.8145523071289],[56.3443374633789,56.2887191772461,56.2936515808105,56.3209457397461,56.3391609191895,56.3568840026855,56.4906234741211,56.5521507263184,56.5694351196289,56.5987777709961,56.6118659973145,56.6429672241211,56.6498527526855,56.6456565856934,56.6098136901855,56.5345191955566,56.5225563049316,56.4906730651855,56.3443374633789],[56.5850105285645,56.5794410705566,56.5936050415039,56.6612319946289,56.67236328125,56.665771484375,56.6266136169434,56.5850105285645],[56.9654312133789,56.95166015625,56.959716796875,56.9723625183105,57.0067863464355,57.0189437866211,57.0002937316895,56.9833488464355,56.9654312133789],[56.9646949768066,56.9518089294434,56.9542961120605,56.9708976745605,57.0179214477539,57.050537109375,57.0313949584961,57.009521484375,56.9852561950684,56.9646949768066],[57.1153335571289,57.1097679138184,57.1151390075684,57.1306648254395,57.192138671875,57.2293472290039,57.2984848022461,57.381103515625,57.3836898803711,57.3717803955078,57.1263656616211,57.1153335571289],[57.5049819946289,57.4607887268066,57.4088401794434,57.3536643981934,57.3142547607422,57.3017082214355,57.2835464477539,57.2632331848145,57.2689437866211,57.252685546875,57.2269020080566,57.1984405517578,57.1465339660645,57.0626487731934,57.045166015625,57.04443359375,57.0519523620605,57.2012214660645,57.18212890625,57.184326171875,57.2024879455566,57.2374992370605,57.3274917602539,57.3628921508789,57.4124526977539,57.4423828125,57.4589309692383,57.4957542419434,57.4826202392578,57.4906730651855,57.5071258544922,57.5206565856934,57.5527305603027,57.5626983642578,57.6033210754395,57.6667938232422,57.6719741821289,57.6512222290039,57.5853042602539,57.5049819946289],[57.6829643249512,57.6266593933105,57.5332984924316,57.5337448120117,57.6019554138184,57.6158676147461,57.6363258361816,57.6525421142578,57.6563911437988,57.6452102661133,57.6631355285645,57.657470703125,57.6829643249512],[58.3632850646973,58.1888694763184,58.1845703125,58.1409683227539,58.0928688049316,58.0918922424316,58.0758743286133,58.0413551330566,58.0212898254395,57.9413566589355,57.9110374450684,57.8275375366211,57.8265113830566,57.7733879089355,57.7500495910645,57.75,57.7617683410645,57.8137664794922,57.8648910522461,57.8936538696289,57.9235343933105,57.9328575134277,57.9749069213867,58.003173828125,58.0179710388184,58.0504875183105,58.0547828674316,58.0723190307617,58.0790023803711,58.0953636169434,58.1382827758789,58.1821823120117,58.2015647888184,58.2223167419434,58.2287063598633,58.2176780700684,58.1825714111328,58.1960945129395,58.1894073486328,58.1975555419922,58.2838897705078,58.3015098571777,58.3216323852539,58.3831520080566,58.4866218566895,58.5028305053711,58.4887199401855,58.4351081848145,58.3632850646973],[58.5154800415039,58.4336891174316,58.4088897705078,58.3783226013184,58.3212432861328,58.2396507263184,58.0521011352539,57.9590301513672,57.9142608642578,57.8520011901855,57.8396453857422,57.8185539245605,57.7869110107422,57.6770477294922,57.5777359008789,57.5812454223633,57.6003456115723,57.6622543334961,57.7082557678223,57.7101516723633,57.6734886169434,57.6723136901855,57.6892585754395,57.6922874450684,57.6808586120605,57.702392578125,57.6766586303711,57.6123504638672,57.4937515258789,57.4740219116211,57.4199714660645,57.3521957397461,57.2588844299316,57.2085456848145,57.1534652709961,57.1025390625,56.8633308410645,56.730712890625,56.6365699768066,56.5615730285645,56.514404296875,56.4829597473145,56.4493637084961,56.42529296875,56.3839340209961,56.3634757995605,56.3660659790039,56.3890609741211,56.3975105285645,56.3182640075684,56.25341796875,56.2021484375,56.194091796875,56.0801239013672,56.0450706481934,56.0276336669922,56.0328140258789,56.0633316040039,56.09521484375,56.0431632995605,56.0160179138184,55.9519538879395,55.9585914611816,56.0262680053711,56.0272941589355,55.9029769897461,55.8079605102539,55.6717262268066,55.6185531616211,55.5703620910645,55.4980926513672,55.259521484375,55.0264167785645,54.7738761901855,54.7037124633789,54.6544914245605,54.5414085388184,54.50390625,54.3951187133789,54.2791976928711,54.1901359558105,54.1180648803711,54.0806159973145,54.021728515625,53.941650390625,53.8651885986328,53.7536849975586,53.7428207397461,53.6092758178711,53.6294441223145,53.6405296325684,53.638370513916,53.63720703125,53.6436538696289,53.6854515075684,53.7367668151855,53.7161636352539,53.7253913879395,53.7240257263184,53.6943855285645,53.6923332214355,53.5363922119141,53.46826171875,53.3354988098145,53.1599617004395,53.0811004638672,53.030029296875,52.9715843200684,52.9056129455566,52.8086891174316,52.8116188049316,52.8251953125,52.858154296875,52.9383773803711,52.9669418334961,52.9772453308105,52.9710960388184,52.953369140625,52.958984375,52.924560546875,52.8935089111328,52.7537117004395,52.67724609375,52.5785179138184,52.468994140625,52.368896484375,52.2785186767578,52.1618156433105,52.1197776794434,52.0868644714355,51.9947738647461,51.9569358825684,51.9735374450684,51.9712409973145,51.9491195678711,51.902099609375,51.8453598022461,51.8033676147461,51.7854499816895,51.8078117370605,51.7295875549316,51.6894035339355,51.6466331481934,51.5714340209961,51.5378875732422,51.5230484008789,51.5194816589355,51.5010757446289,51.4656257629395,51.4844741821289,51.4679679870605,51.4046859741211,51.3865737915039,51.3595237731934,51.3597183227539,51.3750991821289,51.3747062683105,51.36328125,51.3108406066895,51.1820297241211,51.1554718017578,51.0472679138184,50.9716796875,50.9258804321289,50.9339828491211,50.8855476379395,50.8534202575684,50.8191909790039,50.7759780883789,50.7630386352539,50.7887725830078,50.8143577575684,50.8101577758789,50.7654304504395,50.7728004455566,50.8156242370605,50.8445816040039,50.8573265075684,50.8968772888184,50.82080078125,50.7474632263184,50.7328643798828,50.7351570129395,50.7152328491211,50.7253913879395,50.6732406616211,50.6277847290039,50.6080055236816,50.6030769348145,50.6374053955078,50.6309089660645,50.5992202758789,50.6163101196289,50.6697273254395,50.70556640625,50.722412109375,50.7165985107422,50.6324195861816,50.5479469299316,50.4281730651855,50.3218269348145,50.2399406433105,50.229248046875,50.2859382629395,50.3485336303711,50.3908195495605,50.393310546875,50.378173828125,50.3590812683105,50.3582038879395,50.3413543701172,50.2904777526855,50.2559585571289,50.1607437133789,50.1343727111816,50.038330078125,50.0213851928711,50.0829582214355,50.1044425964355,50.0833969116211,50.0506820678711,50.0772476196289,50.1318855285645,50.1969718933105,50.2461433410645,50.3737297058105,50.4515151977539,50.4952659606934,50.5231437683105,50.53369140625,50.58203125,50.7763671875,50.8209495544434,50.9006843566895,50.9774398803711,51.0271492004395,51.1885261535645,51.2013168334961,51.2309112548828,51.2285652160645,51.1969757080078,51.1941375732422,51.2050285339355,51.2485809326172,51.4056625366211,51.4748039245605,51.5372581481934,51.6085968017578,51.74072265625,51.6952171325684,51.6229972839355,51.5811042785645,51.5388641357422,51.4958000183105,51.3984832763672,51.3904304504395,51.413818359375,51.5399398803711,51.5916481018066,51.5975112915039,51.5821266174316,51.56640625,51.569091796875,51.6273422241211,51.659912109375,51.6825180053711,51.7002410888672,51.7410621643066,51.748046875,51.7376480102539,51.6836891174316,51.6262664794922,51.7058601379395,51.74072265625,51.8080558776855,51.8613777160645,51.8801765441895,51.9496574401855,51.9958992004395,52.0418434143066,52.15087890625,52.1973114013672,52.2774429321289,52.3262710571289,52.3931159973145,52.4751434326172,52.541748046875,52.5576171875,52.6078605651855,52.6588401794434,52.7040519714355,52.7607421875,52.8200187683105,52.8661613464355,52.9154815673828,52.9128456115723,52.8974113464355,52.8624496459961,52.81494140625,52.80615234375,52.8441390991211,52.89111328125,52.9582023620605,53.0138168334961,53.0560569763184,53.1051254272461,53.14453125,53.2189445495605,53.3026885986328,53.3076171875,53.2978973388672,53.310546875,53.3406715393066,53.34716796875,53.2603034973145,53.3946800231934,53.4268569946289,53.3053703308105,53.2925796508789,53.3102035522461,53.3307113647461,53.3319320678711,53.3502464294434,53.3892097473145,53.5128402709961,53.5862312316895,53.6625480651855,53.7327651977539,53.7467269897461,53.7735824584961,53.8438453674316,53.9059104919434,53.960693359375,54.0438461303711,54.1353034973145,54.17724609375,54.1705055236816,54.1534194946289,54.1263198852539,54.1279296875,54.2290992736816,54.3056144714355,54.4675788879395,54.5643577575684,54.7730979919434,54.9065933227539,54.9530754089355,54.9619636535645,54.9637680053711,54.9474143981934,54.8928718566895,54.8761253356934,54.8699226379395,54.8427734375,54.8050804138184,54.7809600830078,54.7872085571289,54.7792472839355,54.8010749816895,54.837158203125,54.8467788696289,54.835693359375,54.7870597839355,54.7583503723145,54.7890129089355,54.8461418151855,54.8252944946289,54.7722663879395,54.689453125,54.7613754272461,54.8575210571289,54.9179191589355,54.9858894348145,55.0122566223145,54.9881362915039,55.1494636535645,55.3594245910645,55.4209938049316,55.5013198852539,55.5539054870605,55.5982894897461,55.6991195678711,55.7812004089355,55.8739242553711,55.9295387268066,55.9401359558105,55.9386711120605,55.9673843383789,56.0511703491211,56.0808601379395,56.1583480834961,56.1504898071289,56.1146965026855,56.028076171875,56.0078620910645,55.9873046875,55.9446296691895,55.9334945678711,55.9286613464355,55.8888664245605,55.8863296508789,55.9292488098145,56.0003890991211,56.0658187866211,56.1169929504395,56.2333488464355,56.1974601745605,56.0899391174316,56.0192375183105,55.995361328125,55.9752426147461,55.9520530700684,55.8276863098145,55.7701148986816,55.3895988464355,55.3514175415039,55.3314437866211,55.3268547058105,55.3341331481934,55.3626441955566,55.3949737548828,55.4434547424316,55.6239738464355,55.6741218566895,55.7207527160645,55.8023910522461,55.8077125549316,55.7916984558105,55.7969741821289,55.8131332397461,56.0552749633789,56.1349601745605,56.2508277893066,56.3500480651855,56.4223136901855,56.5147933959961,56.555908203125,56.6187973022461,56.6868629455566,56.758056640625,56.7510223388672,56.5657196044922,56.531982421875,56.541015625,56.5618667602539,56.605712890625,56.6898956298828,56.692138671875,56.7066879272461,56.718017578125,56.763916015625,56.7796401977539,56.8530769348145,56.9026832580566,56.9184074401855,56.9606475830078,57.1023445129395,57.2327117919922,57.2939453125,57.3340835571289,57.3788070678711,57.4360847473145,57.468017578125,57.4992179870605,57.5235328674316,57.5467758178711,57.5716781616211,57.60107421875,57.6436538696289,57.6683120727539,57.7782249450684,57.8235359191895,57.8813438415527,57.8780746459961,57.9036102294922,57.9045867919922,57.8813438415527,57.9063987731934,58.0436019897461,58.0697288513184,58.1436996459961,58.1766624450684,58.2119140625,58.2387199401855,58.2514190673828,58.2501487731934,58.2626457214355,58.2982940673828,58.3451614379883,58.3845176696777,58.4192848205566,58.4892578125,58.5202140808105,58.5665512084961,58.5803184509277,58.58837890625,58.5729026794434,58.55419921875,58.510009765625,58.5135765075684,58.5615692138672,58.5684547424316,58.5128440856934,58.5572280883789,58.5771026611328,58.6063003540039,58.6168937683105,58.6500015258789,58.6348190307617,58.6155281066895,58.5887680053711,58.5154800415039],[58.7416000366211,58.7386245727539,58.7569351196289,58.8226547241211,58.835693359375,58.8275871276855,58.7996101379395,58.7416000366211],[58.7941856384277,58.7809562683105,58.7819328308105,58.8397445678711,58.8817825317383,58.9096183776855,58.9189949035645,58.9052734375,58.8571739196777,58.8258781433105,58.8135757446289,58.8012199401855,58.7941856384277],[59.0296363830566,59.0049781799316,59.0055656433105,58.9845237731934,58.9818878173828,58.955810546875,58.9069328308105,58.8932571411133,58.8905258178711,58.9393539428711,58.9190902709961,58.92529296875,58.9387664794922,58.9555168151855,58.9896469116211,58.9997062683105,58.9674339294434,58.9712371826172,58.9867172241211,59.0187492370605,59.0649909973145,59.1308135986328,59.1439476013184,59.1363296508789,59.0990257263184,59.0760269165039,59.0576667785645,59.0296363830566],[59.2313461303711,59.2301788330078,59.289306640625,59.3041496276855,59.2975578308105,59.2710418701172,59.2313461303711],[59.1867637634277,59.1619186401367,59.1824684143066,59.2468223571777,59.2743606567383,59.2914581298828,59.3238754272461,59.3338394165039,59.3471183776855,59.288330078125,59.2408180236816,59.2267570495605,59.219482421875,59.1867637634277],[60.5375022888184,60.4670448303223,60.4853019714355,60.4177207946777,60.4176254272461,60.4444847106934,60.381591796875,60.2069854736328,60.1773414611816,60.1242713928223,60.1139144897461,60.006591796875,59.9712409973145,59.8869132995605,59.878662109375,59.9111328125,60.0398406982422,60.1146469116211,60.1534690856934,60.1883773803711,60.1894989013672,60.1733856201172,60.1939926147461,60.2217788696289,60.2310066223145,60.2290992736816,60.2367706298828,60.2622528076172,60.2825202941895,60.2983932495117,60.2924766540527,60.3329124450684,60.4685516357422,60.4812965393066,60.4944305419922,60.5174293518066,60.5298347473145,60.5987281799316,60.6095695495605,60.6076622009277,60.5375022888184],[60.5138626098633,60.5022926330566,60.6039047241211,60.7202186584473,60.7165031433105,60.6860313415527,60.6580085754395,60.6555137634277,60.6469230651855,60.5929222106934,60.5301742553711,60.5138626098633],[60.8119621276855,60.8004875183105,60.7161598205566,60.6839370727539,60.6870079040527,60.6972694396973,60.7456550598145,60.7971649169922,60.8104515075684,60.8159141540527,60.8058128356934,60.8318862915039,60.8312492370605,60.8119621276855],[41.8909683227539,41.8550796508789,41.7901840209961,41.757080078125,41.7517585754395,41.7368659973145,41.7021484375,41.6570816040039,41.6248512268066,41.6125946044922,41.6021499633789,41.5077171325684,41.4598617553711,41.4055671691895,41.34375,41.2868156433105,41.2455062866211,41.15966796875,41.0885734558105,41.0702133178711,41.0770530700684,41.0993194580078,41.1544456481934,41.1978492736816,41.183837890625,41.1672859191895,41.1867218017578,41.2244110107422,41.2616195678711,41.2890129089355,41.337646484375,41.42529296875,41.4495582580566,41.4231910705566,41.2909660339355,41.2774887084961,41.2593765258789,41.2485847473145,41.2201690673828,41.2113761901855,41.2082023620605,41.1910171508789,41.2133293151855,41.203369140625,41.1825180053711,41.1589813232422,41.131103515625,41.1166496276855,41.1155281066895,41.1071281433105,41.1259765625,41.155517578125,41.1765632629395,41.1901359558105,41.1852035522461,41.1991691589355,41.2364273071289,41.2648429870605,41.2879409790039,41.30712890625,41.3528289794922,41.4668464660645,41.4923820495605,41.563720703125,41.5789070129395,41.5857429504395,41.5788078308105,41.5707015991211,41.5592765808105,41.4700660705566,41.4398422241211,41.4540023803711,41.4749984741211,41.4867210388184,41.4940910339355,41.4956550598145,41.432373046875,41.4405250549316,41.4715843200684,41.497314453125,41.5174789428711,41.7054176330566,41.817138671875,41.8848648071289,41.9700202941895,42.1468734741211,42.3978538513184,42.6593246459961,42.7376480102539,42.828125,42.9308586120605,43.0634765625,43.1210441589355,43.1457023620605,43.3124008178711,43.4198226928711,43.48486328125,43.5531234741211,43.5697784423828,43.542724609375,43.5120086669922,43.5338859558105,43.4799308776855,43.4180641174316,43.3744659423828,43.3333969116211,43.2763175964355,43.21923828125,43.1901359558105,43.1991233825684,43.2073249816895,43.2280769348145,43.2242164611816,43.1551246643066,43.1590805053711,43.1695785522461,43.1326179504395,43.0915031433105,43.0496559143066,42.9890632629395,42.8966789245605,42.844482421875,42.8077163696289,42.7470207214355,42.727783203125,42.7029762268066,42.6575202941895,42.6169929504395,42.5938491821289,42.571533203125,42.5665512084961,42.5956039428711,42.6163558959961,42.6536140441895,42.7035140991211,42.7486343383789,42.7484855651855,42.7347183227539,42.7096214294434,42.6167984008789,42.746826171875,42.7563972473145,42.7302742004395,42.6941413879395,42.6750030517578,42.6482391357422,42.52978515625,42.5357398986816,42.5176734924316,42.4980964660645,42.4750480651855,42.3573722839355,42.2347183227539,42.205078125,42.1588859558105,42.1099586486816,42.0707054138184,42.035400390625,42.0087394714355,41.9920387268066,41.9898910522461,41.9603500366211,41.9046363830566,41.8909683227539],[49.4945335388184,49.4287109375,49.450927734375,49.4682121276855,49.506591796875,49.5063018798828,49.4945335388184],[11.1156244277954,11.0555658340454,11.0440034866333,11.02099609375,10.9685430526733,10.891357421875,10.8005857467651,10.7155265808105,10.6730461120605,10.630615234375,10.5909690856934,10.5638666152954,10.5206050872803,10.4547853469849,10.3905267715454,10.3069343566895,10.2918453216553,10.2685546875,10.2364749908447,9.92490196228027,9.84458065032959,9.803955078125,9.68759822845459,9.67099571228027,9.67231464385986,9.66791915893555,9.64472675323486,9.62094688415527,9.60415077209473,9.58657264709473,9.57060527801514,9.53564453125,9.49560546875,9.46352577209473,9.44189453125,9.426025390625,9.43183517456055,9.48554611206055,9.49145412445068,9.48027324676514,9.39848613739014,9.35830116271973,9.22124004364014,9.11533260345459,8.97421932220459,8.89492225646973,8.85146522521973,8.81377029418945,8.75927734375,8.72202110290527,8.65273380279541,8.57529354095459,8.47963809967041,8.35488319396973,8.30424785614014,8.25346660614014,8.20956993103027,8.14580059051514,7.72822237014771,7.546875,7.49511671066284,7.43510723114014,7.39873075485229,7.38881826400757,7.35366249084473,7.22656202316284,7.09663057327271,7.03398466110229,7.00410127639771,6.97968769073486,6.93886709213257,6.88832998275757,6.85092830657959,6.802490234375,6.7421875,6.592529296875,6.58076190948486,6.54931688308716,6.51875019073486,6.45258760452271,6.38637733459473,6.32856464385986,6.32031202316284,6.2685546875,6.20263671875,6.17377948760986,6.15502977371216,6.14501905441284,6.08940410614014,6.05136680603027,5.99399375915527,5.90639686584473,5.81025362014771,5.76010704040527,5.75971651077271,5.75732374191284,5.63013315200806,5.56816434860229,5.50078105926514,5.39423847198486,5.31855487823486,5.22670888900757,5.18266630172729,5.03798818588257,4.98085975646973,4.88037061691284,4.762451171875,4.76406240463257,4.87407255172729,4.92934560775757,5.01372051239014,5.04628896713257,5.08247089385986,5.08867216110229,5.12831974029541,5.13081073760986,5.11884784698486,5.14902353286743,5.15302705764771,5.18452167510986,5.26411104202271,5.32822275161743,5.35693359375,5.43251943588257,5.60009765625,5.61918973922729,5.64301776885986,5.67626905441284,5.71132802963257,5.79775333404541,5.92627000808716,6.08564424514771,6.34824228286743,6.44106435775757,6.53564500808716,6.64868211746216,6.69077157974243,6.74912071228027,6.80722713470459,6.94096660614014,7.10458946228027,7.16376972198486,7.20488309860229,7.26362371444702,7.45454120635986,7.68500947952271,7.77207040786743,7.81904268264771,7.89599561691284,7.93193387985229,8.022216796875,8.04667949676514,8.08222579956055,8.12109279632568,8.14755821228027,8.16079139709473,8.171630859375,8.208740234375,8.49301719665527,8.7763671875,8.80043888092041,8.83960056304932,8.95659160614014,9.02509784698486,9.04511737823486,9.10962009429932,9.21860313415527,9.28261756896973,9.30166053771973,9.35136699676514,9.43173789978027,9.48134803771973,9.533935546875,9.65805625915527,9.745849609375,9.79721641540527,9.90966796875,10.0831050872803,10.1925783157349,10.2381839752197,10.2815923690796,10.3228511810303,10.3629388809204,10.4019041061401,10.4324216842651,10.4546384811401,10.5079593658447,10.5923337936401,10.7279787063599,10.9774904251099,10.9983882904053,10.9969720840454,10.9863767623901,10.9887208938599,10.9914054870605,10.9946775436401,10.9976568222046,11.0088872909546,11.0226554870605,10.9972171783447,11.0100584030151,11.001708984375,10.9847173690796,10.9952630996704,10.9889650344849,10.9267578125,10.9273929595947,10.9536619186401,10.9836912155151,11.0076169967651,11.0562982559204,11.09326171875,11.085693359375,11.0879392623901,11.118896484375,11.1668949127197,11.1156244277954],[12.4043941497803,12.377685546875,12.3192386627197,12.2471685409546,12.1986331939697,12.1669425964355,12.10400390625,12.0154294967651,12.0040521621704,12.0248537063599,12.0950193405151,12.1708011627197,12.20751953125,12.2051753997803,12.15185546875,12.0342769622803,11.92724609375,11.9164552688599,11.8987302780151,11.8994140625,11.9255380630493,11.9412107467651,11.9902839660645,12.138671875,12.1795415878296,12.1902837753296,12.212646484375,12.1774415969849,12.116455078125,12.0424795150757,12.0299320220947,12.0424795150757,12.1431150436401,12.1824703216553,12.2286624908447,12.25244140625,12.2554206848145,12.2827644348145,12.3237295150757,12.3660154342651,12.4422359466553,12.4646482467651,12.4792966842651,12.4902830123901,12.4828615188599,12.449951171875,12.40234375,12.3458986282349,12.2255859375,12.1085453033447,11.9225101470947,11.80712890625,11.6732425689697,11.6482419967651,11.6374998092651,11.6177740097046,11.5149898529053,11.485107421875,11.4780759811401,11.4122066497803,11.3862791061401,11.3665523529053,11.3394041061401,11.3047361373901,11.2807130813599,11.2359371185303,11.177001953125,11.0358400344849,11.0094728469849,10.990478515625,10.9869632720947,10.9966793060303,11.0483884811401,11.0299320220947,10.9906244277954,10.9497556686401,10.8960943222046,10.8269529342651,10.74951171875,10.6175785064697,10.4859867095947,10.43798828125,10.3218746185303,10.2784175872803,10.2295408248901,10.1625003814697,10.1252927780151,10.0670890808105,10.0220699310303,9.97319316864014,9.88173866271973,9.6748046875,9.49570274353027,9.4306640625,9.39765644073486,9.40385723114014,9.41586875915527,9.30869197845459,9.18852519989014,9.151611328125,9.11503887176514,9.08085918426514,9.01708984375,8.97978591918945,8.87944316864014,8.78681659698486,8.72060585021973,8.64301681518555,8.5625,8.41035175323486,8.37558650970459,8.375244140625,8.42197227478027,8.46767616271973,8.46753025054932,8.47773456573486,8.49531269073486,8.49067401885986,8.48325157165527,8.45566368103027,8.40791034698486,8.25371074676514,8.21967792510986,8.18144607543945,8.16513729095459,8.16972637176514,8.14492225646973,8.07851505279541,8.02973651885986,7.98442363739014,7.86772489547729,7.82402372360229,7.76074171066284,7.59023475646973,7.55673789978027,7.590576171875,7.60185527801514,7.55849552154541,7.58554649353027,7.62509775161743,7.67705059051514,7.68793964385986,7.68837928771973,7.65888643264771,7.60527324676514,7.54355478286743,7.49570322036743,7.466796875,7.39194297790527,7.32280302047729,7.2626953125,7.26616191864014,7.27461004257202,7.25888681411743,7.22548818588257,7.21591806411743,7.25058650970459,7.27841758728027,7.33330106735229,7.37773466110229,7.40869188308716,7.39492177963257,7.39843702316284,7.41586923599243,7.44252920150757,7.50996112823486,7.57187509536743,7.63955020904541,7.70380783081055,7.79462909698486,7.86669921875,7.90849685668945,7.96792030334473,8.02324199676514,8.05210018157959,8.10698318481445,8.156982421875,8.17626953125,8.260009765625,8.34609413146973,8.37861347198486,8.40234375,8.43603515625,8.47353553771973,8.48442363739014,8.482177734375,8.45888614654541,8.45395469665527,8.53456974029541,8.53769588470459,8.51918983459473,8.42988300323486,8.464599609375,8.505859375,8.51972675323486,8.48881912231445,8.48515605926514,8.49550819396973,8.48095703125,8.36210918426514,8.315673828125,8.31948184967041,8.33027362823486,8.32168006896973,8.33525371551514,8.36420917510986,8.40058517456055,8.52997970581055,8.66030216217041,8.68754863739014,8.76376914978027,8.86757850646973,8.97880840301514,9.05917930603027,9.08168888092041,9.09526348114014,9.12236309051514,9.19448184967041,9.26113319396973,9.28935527801514,9.31425857543945,9.38535213470459,9.51645469665527,9.66162109375,9.78632831573486,9.84316444396973,9.92534160614014,9.97773456573486,9.99653339385986,9.99545955657959,9.99418926239014,9.99301719665527,9.92275333404541,9.87539100646973,9.92978477478027,9.89814472198486,9.86215782165527,9.78720664978027,9.70497989654541,9.671142578125,9.63422870635986,9.60063552856445,9.56191349029541,9.48417949676514,9.37358379364014,9.30224609375,9.26333045959473,9.14692401885986,9.10356426239014,9.06962871551514,9.04755878448486,9.06088924407959,9.07011699676514,9.04921817779541,9.078369140625,9.17055702209473,9.218505859375,9.31430721282959,9.36064434051514,9.42031192779541,9.54340839385986,9.53579139709473,9.63911151885986,9.7763671875,9.85126972198486,9.92778301239014,9.92294883728027,9.87026405334473,9.88720703125,9.96870136260986,10.0478515625,10.1151371002197,10.1412601470947,10.1272459030151,10.1286134719849,10.2483396530151,10.5498542785645,10.6178226470947,10.7349119186401,10.7666997909546,10.68896484375,10.7410154342651,10.862060546875,10.931640625,10.9625482559204,10.9680662155151,10.9449214935303,10.8034181594849,10.8043460845947,10.8345699310303,10.9401369094849,10.9921875,11.0721673965454,11.4055166244507,11.48193359375,11.5084962844849,11.5116214752197,11.5562009811401,11.6297855377197,11.659912109375,11.6519536972046,11.6645994186401,11.7360343933105,11.8341312408447,11.9598627090454,12.0096683502197,12.0544919967651,12.0933113098145,12.1428709030151,12.1781730651855,12.2152338027954,12.2468748092651,12.2629880905151,12.2623529434204,12.2808103561401,12.3126945495605,12.3934078216553,12.478515625,12.5928220748901,12.6739253997803,12.6622562408447,12.6536130905151,12.6395998001099,12.6397466659546,12.6335458755493,12.6150388717651,12.5810546875,12.5362796783447,12.489990234375,12.4776363372803,12.4916505813599,12.5143556594849,12.5322751998901,12.52001953125,12.451904296875,12.4331550598145,12.3961915969849,12.3757810592651,12.3783693313599,12.3400869369507,12.3280277252197,12.3766117095947,12.3980464935303,12.4033203125,12.3873043060303,12.4263181686401,12.4175777435303,12.4043941497803],[13.0641603469849,13.1484861373901,13.341064453125,13.4253911972046,13.4750003814697,13.435302734375,13.3568353652954,13.30322265625,13.2697267532349,13.2937984466553,13.28271484375,13.326171875,13.384033203125,13.4088382720947,13.4253911972046,13.4601068496704,13.4586429595947,13.4683599472046,13.4832029342651,13.4998531341553,13.4599609375,13.4482421875,13.3433589935303,13.3531732559204,13.4579591751099,13.5873050689697,13.5968751907349,13.5927734375,13.5882816314697,13.5862302780151,13.7270021438599,13.7891111373901,13.8121089935303,13.8060054779053,13.7852048873901,13.6690921783447,13.6426267623901,13.6161623001099,13.5597171783447,13.5037107467651,13.4885740280151,13.4971675872803,13.5187501907349,13.5361328125,13.54345703125,13.4785642623901,13.4078121185303,13.3353033065796,13.2963876724243,13.23583984375,13.2688970565796,13.3517084121704,13.4111328125,13.4348630905151,13.4726076126099,13.5133304595947,13.5396480560303,13.5564947128296,13.5352544784546,13.4850587844849,13.4291009902954,13.39599609375,13.3763675689697,13.3558111190796,13.33837890625,13.3251466751099,13.1564455032349,13.1583490371704,13.1603021621704,13.1573247909546,13.1541500091553,13.1197271347046,13.0641603469849],[11.0824699401855,11.0547361373901,11.0589847564697,11.0871086120605,11.0953121185303,11.1797361373901,11.1927738189697,11.1613283157349,11.1483402252197,11.0824699401855],[11.0594234466553,11.0445804595947,11.09423828125,11.1134281158447,11.130126953125,11.1308097839355,11.1668453216553,11.1910152435303,11.1987791061401,11.1972169876099,11.1175298690796,11.0840826034546,11.0594234466553],[11.2154788970947,11.1745109558105,11.1822757720947,11.1945314407349,11.2687015533447,11.3017578125,11.2964849472046,11.2864742279053,11.2578620910645,11.2343254089355,11.2154788970947],[11.4658203125,11.4344244003296,11.4491691589355,11.4771480560303,11.5272951126099,11.5982913970947,11.589111328125,11.4658203125],[11.5370121002197,11.5137691497803,11.5334959030151,11.6176271438599,11.63232421875,11.5675287246704,11.5538578033447,11.5370121002197],[11.8820314407349,11.7597169876099,11.773193359375,11.8459968566895,11.8768072128296,11.8866701126099,11.8820314407349],[10.9401369094849,11.0110349655151,11.1419429779053,11.1400394439697,11.0342283248901,11.0309085845947,11.15625,11.160888671875,11.1520013809204,11.2172365188599,11.2662105560303,11.33447265625,11.3780755996704,11.3963375091553,11.4014644622803,11.3897457122803,11.4103021621704,11.4988765716553,11.5732908248901,11.5809574127197,11.5978031158447,11.6615724563599,11.6867666244507,11.669189453125,11.6229000091553,11.615234375,11.7237787246704,11.7783689498901,11.8428220748901,11.8717775344849,11.8709468841553,11.9073238372803,11.9139642715454,11.947021484375,11.9689950942993,11.9702634811401,11.9272947311401,11.9435548782349,11.9175777435303,11.818359375,11.7634773254395,11.78662109375,11.919677734375,11.9377927780151,11.9596195220947,11.9172849655151,11.9781246185303,12.0516119003296,12.1437501907349,12.2060546875,12.2371091842651,12.2430171966553,12.2041501998901,12.3548336029053,12.3643550872803,12.3486328125,12.36767578125,12.3995113372803,12.4433107376099,12.4574222564697,12.4378910064697,12.4903802871704,12.5889654159546,12.679931640625,12.678955078125,12.677978515625,12.6764163970947,12.6752939224243,12.6739253997803,12.5928220748901,12.478515625,12.3934078216553,12.3126945495605,12.2808103561401,12.2623529434204,12.2629880905151,12.2468748092651,12.2152338027954,12.1781730651855,12.1428709030151,12.0933113098145,12.0544919967651,12.0096683502197,11.9598627090454,11.8341312408447,11.7360343933105,11.6645994186401,11.6519536972046,11.659912109375,11.6297855377197,11.5562009811401,11.5116214752197,11.5084962844849,11.48193359375,11.4055166244507,11.0721673965454,10.9921875,10.9401369094849],[1.03198230266571,1.05444347858429,1.11479473114014,1.12065446376801,1.13925778865814,1.29638683795929,1.435302734375,1.54023432731628,1.56552743911743,1.61757791042328,1.78867173194885,1.927490234375,2.06821274757385,2.30444312095642,2.29780268669128,2.27548813819885,2.24238276481628,2.21328163146973,2.16777300834656,2.16772437095642,2.16762709617615,2.16757845878601,2.16748046875,2.16743183135986,1.93588864803314,1.74018561840057,1.52836894989014,1.30761742591858,1.12075185775757,0.999707043170929,1.00039064884186,1.00126957893372,1.00214838981628,1.00307607650757,1.00356411933899,1.00400364398956,0.997705161571503,0.967138469219208,0.960107624530792,0.986230552196503,0.998730778694153,1.02568387985229,1.07470726966858,1.07998073101044,1.07470726966858,1.04667985439301,1.03198230266571],[3.75830054283142,3.75434541702271,3.75820326805115,3.70532202720642,3.62753915786743,3.40039038658142,3.30463886260986,3.22363305091858,3.21708965301514,3.26464819908142,3.29350566864014,3.33242177963257,3.42290019989014,3.45058608055115,3.46762704849243,3.48237323760986,3.57998061180115,3.66884756088257,3.73593759536743,3.75830054283142],[35.5354499816895,35.5281753540039,35.5294418334961,35.535400390625,35.5938453674316,35.59521484375,35.5374488830566,35.5108413696289,35.4958000183105,35.483642578125,35.4595222473145,35.4686012268066,35.4231452941895,35.385986328125,35.3638191223145,35.3594741821289,35.3660163879395,35.3807640075684,35.409912109375,35.4248046875,35.4098625183105,35.346923828125,35.33935546875,35.3062019348145,35.3280792236328,35.3485870361328,35.3263664245605,35.1840324401855,35.1427230834961,35.1228523254395,35.132568359375,35.17919921875,35.2152824401855,35.215087890625,35.3097648620605,35.3151397705078,35.2686042785645,35.1592292785645,35.0951652526855,35.044677734375,35.0186004638672,35.01416015625,35.0251960754395,35.00732421875,34.9592781066895,34.9344711303711,34.9506340026855,35.0143585205078,35.0583038330078,35.0890617370605,35.1153297424316,35.1603507995605,35.221923828125,35.24609375,35.2334976196289,35.2351570129395,35.2572288513184,35.295166015625,35.4155769348145,35.5347671508789,35.5662612915039,35.5303726196289,35.513916015625,35.5501480102539,35.604736328125,35.655517578125,35.6342277526855,35.5562019348145,35.5354499816895],[35.4652824401855,35.4090843200684,35.4564437866211,35.5111351013184,35.5977554321289,35.7262725830078,35.7886734008789,35.8204574584961,35.7144546508789,35.6294898986816,35.5589332580566,35.4785652160645,35.4652824401855],[36.1897964477539,36.1463851928711,36.1762199401855,36.2209968566895,36.3203125,36.3687477111816,36.3839378356934,36.328125,36.24658203125,36.1897964477539],[35.9292984008789,35.9083023071289,35.9110336303711,35.9573249816895,36.0691909790039,36.14111328125,36.1715812683105,36.2137680053711,36.2769508361816,36.3453102111816,36.4262237548828,36.4336433410645,36.3702621459961,36.2098617553711,36.1296844482422,36.0653305053711,36.0475082397461,35.9292984008789],[36.3926277160645,36.340087890625,36.3589324951172,36.3789520263672,36.4048843383789,36.4422874450684,36.46533203125,36.4737281799316,36.43505859375,36.3926277160645],[36.5853996276855,36.5615234375,36.5113754272461,36.5469245910645,36.5954093933105,36.58056640625,36.6078605651855,36.6385726928711,36.6242179870605,36.5853996276855],[36.5539054870605,36.5379867553711,36.5836906433105,36.6075172424316,36.6348648071289,36.6411628723145,36.6225128173828,36.5826683044434,36.5539054870605],[36.7637710571289,36.7050285339355,36.6839828491211,36.6556167602539,36.7229957580566,36.7442855834961,36.7129402160645,36.7289581298828,36.7474632263184,36.7637710571289],[36.6740226745605,36.6583480834961,36.7215309143066,36.7584457397461,36.7891616821289,36.7173347473145,36.6740226745605],[36.7271003723145,36.7259254455566,36.7742195129395,36.8403816223145,36.8986320495605,36.9051284790039,36.868896484375,36.8091316223145,36.777587890625,36.770751953125,36.7271003723145],[36.7904281616211,36.7822265625,36.7897453308105,36.8070335388184,36.8253936767578,36.8475608825684,36.8865737915039,36.9374046325684,36.9027328491211,36.8796882629395,36.7904281616211],[36.9214363098145,36.9170913696289,36.9592781066895,36.9985847473145,37.0216293334961,37.0238304138184,36.94921875,36.9214363098145],[36.9590339660645,36.9452133178711,37.0246086120605,37.0872573852539,37.0521011352539,37.0096664428711,37.0015640258789,36.9759788513184,36.9590339660645],[37.0684051513672,36.9913063049316,36.9996566772461,37.0349617004395,37.107421875,37.1485366821289,37.1378440856934,37.0841827392578,37.0684051513672],[36.9675788879395,36.9296875,36.984375,37.0704116821289,37.1963882446289,37.1851081848145,37.1525382995605,37.039306640625,36.9675788879395],[37.1250953674316,37.1100578308105,37.1319808959961,37.1868667602539,37.210205078125,37.1923332214355,37.1676750183105,37.1250953674316],[37.3444366455078,37.3141136169434,37.3834457397461,37.4196281433105,37.4503898620605,37.4751968383789,37.4495620727539,37.4080085754395,37.3444366455078],[37.4935073852539,37.3896980285645,37.3905754089355,37.4065933227539,37.4463386535645,37.5088844299316,37.4935073852539],[37.4191398620605,37.4129905700684,37.4893074035645,37.5091781616211,37.4710922241211,37.4470672607422,37.4191398620605],[37.5293922424316,37.5255851745605,37.5655746459961,37.6349143981934,37.6382789611816,37.6730461120605,37.67431640625,37.6195793151855,37.5685043334961,37.5293922424316],[37.599609375,37.5351066589355,37.5450668334961,37.6144523620605,37.6459465026855,37.6769027709961,37.6806640625,37.6479988098145,37.6306648254395,37.599609375],[37.5768547058105,37.5282745361328,37.6011238098145,37.677734375,37.6827125549316,37.6490249633789,37.5768547058105],[37.8114242553711,37.7784652709961,37.781982421875,37.77001953125,37.7092781066895,37.7004890441895,37.6447257995605,37.656982421875,37.7054672241211,37.7104988098145,37.7237281799316,37.7808609008789,37.8097648620605,37.8114242553711],[37.8053703308105,37.7080078125,37.7321319580078,37.6647453308105,37.7434577941895,37.8231468200684,37.8550300598145,37.9295387268066,37.8529815673828,37.8407249450684,37.8053703308105],[37.7596168518066,37.6923828125,37.7705078125,37.8240242004395,37.8707046508789,37.8988761901855,37.961669921875,37.9874992370605,37.9901390075684,37.9136734008789,37.90478515625,37.857666015625,37.7969245910645,37.7596168518066],[37.9258804321289,37.9011688232422,37.9023933410645,37.9114723205566,37.9312515258789,37.940673828125,37.9803695678711,37.9911155700684,37.9860343933105,37.97021484375,37.9258804321289],[38.3833503723145,38.2678680419922,38.2461891174316,38.14208984375,38.0888214111328,38.070556640625,38.1197280883789,38.09765625,38.1066398620605,38.1641616821289,38.1843757629395,38.2187995910645,38.2341804504395,38.1884269714355,38.1798820495605,38.2217292785645,38.3367691040039,38.356201171875,38.3182106018066,38.3323211669922,38.39453125,38.45654296875,38.4749488830566,38.3833503723145],[38.3294448852539,38.318603515625,38.414306640625,38.4803237915039,38.4839820861816,38.476318359375,38.451416015625,38.4259262084961,38.3986320495605,38.3657684326172,38.3294448852539],[38.2180671691895,38.1615219116211,38.2433128356934,38.2696266174316,38.3026351928711,38.353515625,38.416015625,38.5083999633789,38.5740242004395,38.6017074584961,38.5446281433105,38.5407218933105,38.4861831665039,38.4684562683105,38.3029289245605,38.2796363830566,38.2341804504395,38.2180671691895],[38.6086921691895,38.6009788513184,38.6062507629395,38.6017074584961,38.5825691223145,38.661865234375,38.7601547241211,38.8175773620605,38.84423828125,38.7991714477539,38.6383285522461,38.6086921691895],[38.8092269897461,38.7848167419434,38.7886734008789,38.8194351196289,38.8323745727539,38.8886222839355,38.961669921875,38.9802703857422,38.9422340393066,38.8788566589355,38.8092269897461],[38.9586448669434,38.85009765625,38.8134765625,38.7702178955078,38.7646980285645,38.6865692138672,38.6709976196289,38.6484870910645,38.5882797241211,38.541015625,38.5042495727539,38.4634284973145,38.338623046875,38.2200202941895,38.1624984741211,38.1451187133789,38.1475105285645,38.1239738464355,38.0201644897461,37.979736328125,37.9699249267578,37.9805183410645,38.0049781799316,38.0165519714355,38.0185546875,38.0603523254395,38.1175308227539,38.1516609191895,38.2042961120605,38.2430648803711,38.3168449401855,38.3372077941895,38.3741188049316,38.3899917602539,38.4007339477539,38.4012222290039,38.4430656433105,38.5525398254395,38.5819816589355,38.6129417419434,38.6558570861816,38.7350120544434,38.8012237548828,38.8448219299316,38.8733901977539,38.8396492004395,38.84765625,38.8705062866211,38.9159164428711,39.002685546875,39.0313491821289,39.034912109375,38.9586448669434],[39.1144027709961,39.08056640625,39.0953598022461,39.2085952758789,39.1144027709961],[39.1583023071289,39.1465835571289,39.2263679504395,39.2677230834961,39.2005348205566,39.1583023071289],[39.3294448852539,39.2701187133789,39.1717758178711,39.1095237731934,39.048828125,39.0314445495605,39.0643539428711,39.0748062133789,39.0314445495605,38.994140625,38.9728050231934,38.9739265441895,39.02587890625,39.0810546875,39.1641082763672,39.1975593566895,39.1942863464355,39.0956077575684,39.1389656066895,39.1786613464355,39.2000503540039,39.2875480651855,39.2846183776855,39.3042945861816,39.3319816589355,39.37353515625,39.3830070495605,39.3294448852539],[39.4327125549316,39.3766098022461,39.4114227294922,39.4615249633789,39.5853042602539,39.7261734008789,39.76708984375,39.798095703125,39.8201141357422,39.7972640991211,39.7737312316895,39.7467269897461,39.692626953125,39.6681175231934,39.6194839477539,39.5999984741211,39.5059089660645,39.4704093933105,39.4420890808105,39.4327125549316],[39.9832992553711,39.9495620727539,39.8913116455078,39.80810546875,39.8061027526855,39.8229484558105,39.8543968200684,39.8941421508789,39.892578125,39.8494148254395,39.8299293518066,39.8258285522461,39.8523902893066,39.9098625183105,39.9763679504395,39.9996566772461,40.0054168701172,39.956298828125,39.9847640991211,40.0155258178711,40.0348167419434,39.9832992553711],[40.4265594482422,40.4004402160645,40.4828109741211,40.515869140625,40.4919891357422,40.4638671875,40.4265594482422],[40.6151847839355,40.5794448852539,40.6470222473145,40.6872062683105,40.7687492370605,40.7929191589355,40.7862815856934,40.7302742004395,40.703857421875,40.6588859558105,40.6151847839355],[40.7267570495605,40.7691383361816,40.8440895080566,40.8877944946289,40.943115234375,40.9328117370605,40.9947280883789,40.967529296875,40.8575172424316,40.8694801330566,40.9355964660645,40.94775390625,40.9127426147461,40.7861328125,40.7240753173828,40.7483367919922,40.747802734375,40.677001953125,40.6279754638672,40.5443840026855,40.4815444946289,40.4185562133789,40.40576171875,40.4093246459961,40.3277816772461,40.2417984008789,40.147705078125,40.2151870727539,40.280029296875,40.3035621643066,40.3587875366211,40.3680152893066,40.3297348022461,40.2862777709961,40.2051277160645,40.1552238464355,40.1145515441895,40.0246086120605,39.9940414428711,39.965576171875,40.0222663879395,40.2238273620605,40.2639656066895,40.2219734191895,40.1154289245605,40.0739250183105,39.9588890075684,39.9344749450684,39.924072265625,39.9898414611816,40.0899429321289,40.2164535522461,40.3042984008789,40.39990234375,40.4906234741211,40.5242652893066,40.5908699035645,40.57861328125,40.5364761352539,40.4955558776855,40.4286155700684,40.3666038513184,40.2764167785645,40.1193351745605,40.0369110107422,39.8005828857422,39.62890625,39.5638198852539,39.4920425415039,39.3584442138672,39.288818359375,39.1749000549316,39.1043930053711,39.1014633178711,39.1327629089355,39.21044921875,39.2577629089355,39.3310546875,39.3063468933105,39.2585906982422,39.1699714660645,39.1115226745605,39.0309066772461,39.0379409790039,38.9477043151855,38.901611328125,38.89892578125,38.8905754089355,38.8674812316895,38.8506813049316,38.8491706848145,38.8003921508789,38.7418937683105,38.66796875,38.6612319946289,38.5255393981934,38.4894027709961,38.3524398803711,38.3254890441895,38.2750015258789,38.226806640625,38.1397933959961,37.955322265625,37.8179206848145,37.7745132446289,37.7096176147461,37.677734375,37.6767578125,37.7777824401855,37.8840827941895,38.0105476379395,38.0327644348145,38.0348663330078,37.9920921325684,37.9590339660645,37.912841796875,37.9026336669922,37.8783683776855,37.8531227111816,37.7953109741211,37.7162590026855,37.6202125549316,37.5954093933105,37.5975570678711,37.5797843933105,37.5415534973145,37.4969215393066,37.4638671875,37.440185546875,37.3772964477539,37.3485374450684,37.3338394165039,37.36376953125,37.4405784606934,37.4817848205566,37.51708984375,37.5322265625,37.5851058959961,37.5421371459961,37.393310546875,37.2908210754395,37.015869140625,36.853515625,36.7749519348145,36.64453125,36.547607421875,36.4480972290039,36.4518547058105,36.4869613647461,36.5283660888672,36.6871070861816,36.7861824035645,36.7939453125,36.7797355651855,36.5681648254395,36.4469223022461,36.4757804870605,36.5135726928711,36.6461906433105,36.701904296875,36.882568359375,36.90283203125,36.9639129638672,37.0289535522461,37.0165023803711,36.9900894165039,36.8917999267578,36.8036613464355,36.7373046875,36.8632316589355,37.0809555053711,37.2003898620605,37.3092765808105,37.38720703125,37.541015625,37.6399421691895,37.6693382263184,37.7745132446289,37.828857421875,37.8541526794434,37.8916015625,37.9192886352539,38.0274391174316,38.1966781616211,38.2047348022461,38.1646003723145,38.1750984191895,38.2742195129395,38.328125,38.3211898803711,38.188720703125,38.1132316589355,38.0469245910645,37.981201171875,37.9675788879395,37.9583015441895,38.0074729919434,38.0509300231934,38.0746078491211,38.0733375549316,38.0963859558105,38.1336898803711,38.1760749816895,38.1964378356934,38.2020988464355,38.2155265808105,38.2019500732422,38.2347145080566,38.26171875,38.2895011901855,38.3449211120605,38.4385261535645,38.3855476379395,38.3568382263184,38.3528327941895,38.4124526977539,38.366943359375,38.3550300598145,38.35400390625,38.3335914611816,38.3213882446289,38.4078140258789,38.4748039245605,38.4873046875,38.4243659973145,38.3739242553711,38.3455581665039,38.3846702575684,38.5032730102539,38.6540031433105,38.7757301330566,38.8075180053711,38.8744125366211,38.9278793334961,38.9411163330078,38.8851547241211,38.8962898254395,38.9220733642578,38.9791984558105,39.0299835205078,39.0322723388672,39.0262718200684,39.0367698669434,39.008544921875,39.03515625,39.0674781799316,39.147705078125,39.2552719116211,39.3270988464355,39.5458030700684,39.6412620544434,39.7094268798828,39.710693359375,39.6991233825684,39.66162109375,39.6535186767578,39.6783676147461,39.701171875,39.7385749816895,39.7822265625,39.7966804504395,39.791748046875,39.8026390075684,39.841796875,39.890625,39.9507789611816,39.9794425964355,39.9910659790039,40.0171852111816,40.0494613647461,40.0655746459961,40.0685043334961,40.0826644897461,40.1173820495605,40.1517601013184,40.2463874816895,40.2926750183105,40.3349151611328,40.3918952941895,40.4454612731934,40.4679183959961,40.494384765625,40.5633811950684,40.6224594116211,40.6586418151855,40.7177734375,40.7752952575684,40.8499031066895,40.8561515808105,40.8631324768066,40.8671379089355,40.9071769714355,40.9036140441895,40.8689460754395,40.8963394165039,40.950439453125,41.107421875,41.1309547424316,41.1405296325684,41.1586418151855,41.1551742553711,41.1233863830566,41.1185073852539,41.1401863098145,41.1785163879395,41.312744140625,41.3319816589355,41.3373527526855,41.3362770080566,41.3256378173828,41.3220710754395,41.3849601745605,41.3896484375,41.3987312316895,41.3860359191895,41.3867645263672,41.4129867553711,41.4559555053711,41.4522933959961,41.4600563049316,41.4690895080566,41.5272445678711,41.5308113098145,41.5250473022461,41.5235328674316,41.5552253723145,41.5525398254395,41.4673843383789,41.4427261352539,41.4199714660645,41.3561019897461,41.3729019165039,41.3942375183105,41.364990234375,41.3157730102539,41.2435531616211,41.2643547058105,41.2998046875,41.3101081848145,41.3150405883789,41.3304214477539,41.3119125366211,41.3506813049316,41.3857421875,41.4348640441895,41.5215339660645,41.6082038879395,41.6401863098145,41.6732406616211,41.7041511535645,41.7256851196289,41.7437973022461,41.716552734375,41.6963386535645,41.6633796691895,41.6332511901855,41.6072273254395,41.6012687683105,41.5121574401855,41.4017601013184,41.3541488647461,41.3431129455566,41.23876953125,41.1432647705078,41.0970230102539,41.0643043518066,41.0367660522461,40.9970703125,40.9544906616211,40.8832054138184,40.8265113830566,40.7496566772461,40.7402839660645,40.7267570495605],[12.0126466751099,12.0084466934204,12.0454587936401,12.1084470748901,12.1851568222046,12.2370119094849,12.2232913970947,12.0539541244507,12.0126466751099],[60.7813949584961,60.6603050231934,60.6866683959961,60.7407684326172,60.7659187316895,60.7583999633789,60.7798309326172,60.9087867736816,60.9435043334961,60.8891639709473,60.8415565490723,60.7813949584961],[65.531982421875,65.5313491821289,65.60986328125,65.6954574584961,65.7222671508789,65.663330078125,65.5755844116211,65.531982421875],[69.552490234375,69.5171432495117,69.5250015258789,69.5518569946289,69.674072265625,69.7320327758789,69.8040542602539,69.8547821044922,69.9139175415039,69.9241714477539,69.90869140625,69.8485336303711,69.8290557861328,69.7976531982422,69.7566909790039,69.6642532348633,69.6178207397461,69.5830078125,69.552490234375],[69.9447326660156,69.8634338378906,69.8072280883789,69.7815399169922,69.74267578125,69.722412109375,69.70361328125,69.686279296875,69.6639709472656,69.6366653442383,69.6047821044922,69.5500030517578,69.4891052246094,69.3639144897461,69.3426284790039,69.2566452026367,69.2601547241211,69.2642135620117,69.302001953125,69.337158203125,69.3644027709961,69.4035110473633,69.4277801513672,69.4373016357422,69.4366760253906,69.4651336669922,69.49072265625,69.5063018798828,69.5403366088867,69.53369140625,69.5531768798828,69.5654296875,69.5771942138672,69.6105499267578,69.6305236816406,69.6650390625,69.713623046875,69.9019012451172,69.9498519897461,69.9656677246094,69.923828125,69.94189453125,70.0111770629883,70.0525360107422,70.0851135253906,70.1329574584961,70.1610794067383,70.1893997192383,70.2177734375,70.2561492919922,70.3172836303711,70.2964401245117,70.2212905883789,70.2053680419922,70.140869140625,69.9447326660156],[70.855224609375,70.8525390625,70.8706588745117,70.8822784423828,70.9043884277344,70.968017578125,70.9764633178711,70.941650390625,70.8921356201172,70.8688507080078,70.855224609375],[70.9213409423828,70.8624572753906,70.8346710205078,70.811279296875,70.7945861816406,70.7796859741211,70.7405776977539,70.7143096923828,70.6933135986328,70.6526870727539,70.5989227294922,70.5730438232422,70.5091323852539,70.4540481567383,70.454345703125,70.5114212036133,70.5533676147461,70.531494140625,70.4786605834961,70.4540023803711,70.4671401977539,70.48681640625,70.5143508911133,70.5948257446289,70.6152801513672,70.6420440673828,70.7127914428711,70.7897491455078,70.8971176147461,70.9137725830078,70.8756332397461,70.8675765991211,70.8626937866211,70.8756332397461,70.9192428588867,71.0436553955078,71.0420379638672,70.9979553222656,70.976318359375,70.9425354003906,70.9213409423828],[71.0408172607422,71.0342712402344,71.0851593017578,71.1043014526367,71.1277313232422,71.155517578125,71.2072296142578,71.2830123901367,71.2970733642578,71.2496109008789,71.1858444213867,71.1534194946289,71.115234375,71.0829162597656,71.0563049316406,71.0408172607422],[72.7911148071289,72.7232894897461,72.684326171875,72.5684127807617,72.5643539428711,72.5794448852539,72.6099090576172,72.6172409057617,72.636474609375,72.6521453857422,72.662109375,72.6683578491211,72.6564483642578,72.66845703125,72.7191848754883,72.7532196044922,72.7822799682617,72.793701171875,72.7803726196289,72.7807159423828,72.7886199951172,72.8247985839844,72.8416519165039,72.8205032348633,72.7911148071289],[75.4073257446289,75.301513671875,75.2044448852539,75.1427764892578,75.1515121459961,75.0369186401367,74.9927749633789,75.0104522705078,75.0016555786133,75.0721664428711,75.1956558227539,75.3191375732422,75.3896484375,75.3279800415039,75.3720703125,75.4073257446289],[76.0423355102539,76.015869140625,76.4303665161133,76.5802688598633,76.6949691772461,76.7038116455078,76.6420364379883,76.4039077758789,76.0423355102539],[77.3252944946289,77.3164596557617,77.3554153442383,77.3855438232422,77.431640625,77.4475555419922,77.4635238647461,77.4670867919922,77.4599609375,77.431640625,77.4032669067383,77.3943862915039,77.3536605834961,77.3252944946289],[77.642333984375,77.642333984375,77.6683654785156,77.71435546875,77.8626022338867,77.8746032714844,77.8585968017578,77.782470703125,77.7063980102539,77.642333984375],[77.9737777709961,77.9385223388672,78.0443344116211,78.1854476928711,78.3441925048828,78.423583984375,78.4412155151367,78.423583984375,78.3529815673828,78.2119140625,78.1148910522461,77.9737777709961],[79.8258819580078,79.7112274169922,79.7200241088867,79.77294921875,79.8523406982422,79.9404830932617,80.0110855102539,80.05517578125,80.0287094116211,79.9404830932617,79.8258819580078],[81.8464813232422,81.8143081665039,81.8271942138672,81.9172897338867,81.9912567138672,82.078125,82.11669921875,82.1231994628906,82.0491714477539,81.949462890625,81.8464813232422],[82.0836410522461,82.0660171508789,82.1718292236328,82.2776870727539,82.3481979370117,82.5333938598633,82.5992202758789,82.6569366455078,82.6657257080078,82.692138671875,82.5775375366211,82.4805145263672,82.401123046875,82.2423858642578,82.0836410522461],[83.5648422241211,83.5047836303711,83.4349136352539,83.4372100830078,83.410400390625,83.3769073486328,83.2896499633789,83.260986328125,83.2375030517578,83.2213897705078,83.192626953125,83.1574249267578,83.0889129638672,83.0853500366211,82.9834442138672,82.9778747558594,82.9916458129883,83.0936965942383,83.1104965209961,83.1020050048828,83.063720703125,83.0771942138672,83.0671920776367,83.096435546875,83.1596145629883,83.0185546875,82.8773956298828,82.8930206298828,82.8851089477539,82.8387680053711,82.8191375732422,82.8296890258789,82.7891616821289,82.7164077758789,82.6825180053711,82.6341857910156,82.5954055786133,82.5477066040039,82.4627914428711,82.3847198486328,82.32470703125,82.2870559692383,82.1611785888672,82.1312484741211,82.0548324584961,81.9554672241211,81.93994140625,81.9958953857422,82.0488739013672,82.0463333129883,82.0011215209961,81.8828125,81.7009735107422,81.7418441772461,81.7730484008789,81.8271942138672,81.8852996826172,81.947265625,81.9894561767578,82.0118179321289,82.030517578125,82.0530319213867,82.074951171875,82.0686950683594,81.9838409423828,81.9342269897461,81.86962890625,81.7899475097656,81.6951675415039,81.6013717651367,81.4375,81.3482437133789,81.283935546875,81.1371078491211,81.097900390625,80.9267120361328,80.8473663330078,80.7892608642578,80.7781753540039,80.8328094482422,80.871826171875,80.9126510620117,81.0202178955078,81.0501937866211,81.1781692504883,81.2261734008789,81.2763671875,81.31201171875,81.5643539428711,81.639892578125,81.6400451660156,81.51220703125,81.492431640625,81.4979476928711,81.4668502807617,81.441162109375,81.4281768798828,81.397705078125,81.430419921875,81.450927734375,81.5438919067383,81.626220703125,81.7290496826172,81.7539596557617,81.776611328125,81.7854995727539,81.8335952758789,81.8369598388672,81.82177734375,81.8138732910156,81.7890625,81.72021484375,81.6825180053711,81.6491165161133,81.5775375366211,81.5026397705078,81.4806137084961,81.4568328857422,81.4240264892578,81.3092269897461,81.2325210571289,81.0877914428711,81.0380859375,81.0186004638672,81.013916015625,80.9931106567383,80.9732894897461,80.9132385253906,80.8704605102539,80.8324203491211,80.7760696411133,80.7632751464844,80.721435546875,80.650390625,80.6417007446289,80.6498031616211,80.5733871459961,80.51123046875,80.4842300415039,80.4276351928711,80.3951187133789,80.3293991088867,80.251953125,80.1982421875,80.190185546875,80.2036590576172,80.2007827758789,80.176025390625,80.1720733642578,80.2070846557617,80.2476043701172,80.2616195678711,80.2577133178711,80.2414016723633,80.1447296142578,80.0787124633789,80.01123046875,79.9376449584961,79.8579559326172,79.8033676147461,79.7737808227539,79.755859375,79.7464828491211,79.75537109375,79.7503433227539,79.7341766357422,79.6831512451172,79.635009765625,79.5673294067383,79.4883804321289,79.3981399536133,79.3488311767578,79.3380355834961,79.3415985107422,79.3253936767578,79.2894515991211,79.25146484375,79.2113800048828,79.1783676147461,79.15234375,79.1230010986328,79.0650405883789,79.04736328125,79.0121078491211,78.9591293334961,78.9109344482422,78.8676300048828,78.841796875,78.830322265625,78.8288116455078,78.8039093017578,78.6586456298828,78.6425323486328,78.5959014892578,78.5550308227539,78.3798370361328,78.2930145263672,78.1739730834961,78.0735855102539,77.9918441772461,77.8974609375,77.7906265258789,77.7085418701172,77.6513671875,77.6975631713867,77.8472213745117,77.911865234375,77.8915557861328,77.8619613647461,77.8034133911133,77.7669448852539,77.7189025878906,77.6783676147461,77.6212921142578,77.5852508544922,77.5658187866211,77.5719757080078,77.6663589477539,77.6898422241211,77.66162109375,77.6189956665039,77.4473190307617,77.368408203125,77.3323745727539,77.29443359375,77.2459945678711,77.2223663330078,77.2327651977539,77.2804718017578,77.2830581665039,77.2593765258789,77.2152786254883,77.1328582763672,77.0121078491211,76.9211959838867,76.8600540161133,76.7781753540039,76.7632827758789,76.7677230834961,76.7845230102539,76.8137664794922,76.8365783691406,76.861083984375,76.9144058227539,76.9275894165039,76.9207992553711,76.8870086669922,76.8426742553711,76.6878890991211,76.6899948120117,76.7431640625,76.7940902709961,76.793701171875,76.729248046875,76.7042922973633,76.6807632446289,76.6250534057617,76.6122131347656,76.6014633178711,76.5880889892578,76.573486328125,76.4005355834961,76.293701171875,76.2718734741211,76.2640151977539,76.2679672241211,76.3040084838867,76.275146484375,76.2398452758789,76.2310485839844,76.2324752807617,76.2190933227539,76.1206588745117,75.9966812133789,75.8973617553711,75.7949676513672,75.7575225830078,75.6895980834961,75.644775390625,75.6038589477539,75.56689453125,75.4944381713867,75.3864212036133,75.2981948852539,75.2298355102539,75.1802291870117,75.1493682861328,75.157470703125,75.2045440673828,75.2546920776367,75.3079605102539,75.3142547607422,75.1569366455078,75.1490707397461,75.1333999633789,75.0647964477539,75.0234375,75.0398483276367,75.1197280883789,75.0663528442383,75.00390625,74.9714813232422,74.9644546508789,74.9719772338867,74.99755859375,75.0685501098633,75.0793914794922,75.0743637084961,75.0534210205078,74.9925842285156,74.8917465209961,74.8059616088867,74.7352066040039,74.6898498535156,74.6698760986328,74.6541519165039,74.6359329223633,74.7282257080078,74.8429183959961,74.9751968383789,75.0192413330078,74.9751968383789,74.8517074584961,74.6245651245117,74.6009216308594,74.54638671875,74.4794921875,74.4001922607422,74.3426284790039,74.3067855834961,74.2840347290039,74.2694854736328,74.2579574584961,74.2822723388672,74.2828140258789,74.2046356201172,74.1373519897461,74.1108856201172,74.1634750366211,74.2442855834961,74.3572769165039,74.4827651977539,74.5657196044922,74.5674819946289,74.439208984375,74.3900451660156,74.3301773071289,74.3025436401367,74.286376953125,74.2724151611328,74.2455062866211,74.2057189941406,74.16552734375,74.125,74.0909652709961,74.0634307861328,74.0298767089844,73.990478515625,73.9709930419922,73.9624557495117,73.9412612915039,73.8482437133789,73.8196792602539,73.6530227661133,73.4928665161133,73.4635772705078,73.4566421508789,73.4316940307617,73.3581008911133,73.2698745727539,73.2692337036133,73.3462371826172,73.397705078125,73.5431137084961,73.7644500732422,73.6724166870117,73.6285171508789,73.6057586669922,73.6022033691406,73.617919921875,73.6527862548828,73.7336883544922,73.8136215209961,73.8516082763672,73.8408203125,73.7938003540039,73.7395935058594,73.5801696777344,73.5398941040039,73.5112762451172,73.4857940673828,73.4310073852539,73.3906784057617,73.2928237915039,73.2776336669922,73.2530288696289,73.259033203125,73.3129425048828,73.3481903076172,73.3795394897461,73.436279296875,73.3741683959961,73.2794952392578,73.2489776611328,73.192138671875,73.17138671875,73.1669921875,73.1789093017578,73.1764678955078,73.1598129272461,73.1384811401367,73.112548828125,73.0889205932617,73.067626953125,73.0678176879883,73.1324234008789,73.1370086669922,73.12109375,73.1714782714844,73.1932144165039,73.1987838745117,73.2757797241211,73.3619613647461,73.396484375,73.4229507446289,73.4093780517578,73.3982925415039,73.3167953491211,73.2590789794922,73.1932601928711,73.1715850830078,73.083984375,72.986083984375,72.9650421142578,72.91845703125,72.7208023071289,72.6354522705078,72.3992233276367,72.3447799682617,72.2202606201172,72.1195297241211,72.1577682495117,72.2239227294922,72.3265609741211,72.392578125,72.4524383544922,72.4987258911133,72.6873016357422,72.9217300415039,73.0376434326172,73.0441360473633,73.0130844116211,72.9802703857422,72.9241180419922,72.8468780517578,72.7939910888672,72.7955551147461,72.7158203125,72.6776351928711,72.6728057861328,72.6943893432617,72.721923828125,72.79736328125,72.8102569580078,72.8427734375,72.8892135620117,72.9015121459961,72.8897399902344,72.86865234375,72.58251953125,72.50634765625,72.4733428955078,72.419189453125,72.3929672241211,72.3469772338867,72.3903350830078,72.4373550415039,72.4202194213867,72.3482360839844,72.3113250732422,72.2007369995117,72.1397933959961,72.0810089111328,71.9994125366211,71.9706497192383,71.9282684326172,71.913818359375,71.7698211669922,71.7538070678711,71.7446746826172,71.6888198852539,71.5645523071289,71.52490234375,71.5004425048828,71.4566879272461,71.3834457397461,71.2486801147461,71.3734893798828,71.4323272705078,71.449951171875,71.4525451660156,71.5082015991211,71.4783172607422,71.3374557495117,71.2059555053711,71.0923843383789,70.9158630371094,70.8563003540039,70.8046340942383,70.5904769897461,70.5262145996094,70.4685592651367,70.4434585571289,70.4718780517578,70.46240234375,70.5131378173828,70.5712890625,70.6119155883789,70.648681640625,70.8600082397461,70.8078155517578,70.7649841308594,70.721435546875,70.4933090209961,70.4373016357422,70.4507751464844,70.4424819946289,70.4509735107422,70.55517578125,70.6494598388672,70.7910690307617,70.9233856201172,71.0463333129883,71.1463851928711,71.2235336303711,71.2860870361328,71.3339385986328,71.3956985473633,71.4712448120117,71.530029296875,71.5719299316406,71.5899429321289,71.5833435058594,71.6305618286133,71.6265640258789,71.6021957397461,71.5326690673828,71.5007858276367,71.4934997558594,71.498046875,71.4801788330078,71.43994140625,71.3683090209961,71.2653350830078,71.18359375,71.0928268432617,71.05029296875,70.9686965942383,70.9504852294922,70.9449234008789,70.9527816772461,70.9934539794922,71.001708984375,71.0071792602539,70.992919921875,70.9493103027344,70.9246063232422,70.8952102661133,70.8395080566406,70.7567825317383,70.6990203857422,70.6556625366211,70.5735397338867,70.5475540161133,70.4615249633789,70.4449691772461,70.4472198486328,70.477783203125,70.4769058227539,70.4022445678711,70.4066925048828,70.4755401611328,70.4742202758789,70.46337890625,70.4375457763672,70.3966293334961,70.3570785522461,70.3188934326172,70.2811965942383,70.2557144165039,70.2171401977539,70.1244583129883,70.0282211303711,69.9916000366211,70.0379409790039,70.1408157348633,70.2012252807617,70.2421875,70.2213363647461,70.24560546875,70.3469696044922,70.3531723022461,70.2950744628906,70.1811981201172,70.1393051147461,70.1145935058594,70.1258316040039,70.10791015625,70.0675811767578,70.0334014892578,69.9857482910156,69.9542541503906,69.9453582763672,69.9228515625,69.9008331298828,69.8829574584961,69.8048324584961,69.792724609375,69.7914505004883,69.7405242919922,69.7441864013672,69.7367172241211,69.7178268432617,69.6813430786133,69.6272506713867,69.588623046875,69.5655822753906,69.5580596923828,69.5903778076172,69.5855407714844,69.5623550415039,69.5269546508789,69.4792938232422,69.4393081665039,69.4070739746094,69.3184051513672,69.2930679321289,69.2721176147461,69.2605514526367,69.1924819946289,69.1651840209961,69.0915985107422,69.0457077026367,69.0209503173828,68.9544448852539,68.927978515625,68.889892578125,68.8172836303711,68.7811508178711,68.7515640258789,68.7021484375,68.6759262084961,68.6728515625,68.6543426513672,68.601806640625,68.5843276977539,68.4935073852539,68.47900390625,68.446533203125,68.3598098754883,68.3319320678711,68.2987747192383,68.289306640625,68.3108367919922,68.3115692138672,68.2985382080078,68.2719192504883,68.198974609375,68.1933135986328,68.2511672973633,68.2249526977539,68.162353515625,68.117919921875,68.0728454589844,68.0613250732422,68.079833984375,68.1284637451172,68.22998046875,68.3849105834961,68.4373092651367,68.3875961303711,68.3390121459961,68.2916564941406,68.2572784423828,68.2359848022461,68.2252426147461,68.22509765625,68.2130355834961,68.1391067504883,68.0631866455078,67.9911117553711,67.9228515625,67.8827667236328,67.8709487915039,67.70068359375,67.6792449951172,67.658203125,67.626708984375,67.4857482910156,67.4427185058594,67.38671875,67.3769989013672,67.3541946411133,67.3181686401367,67.2581558227539,67.1742172241211,66.9422912597656,66.7259292602539,66.6550750732422,66.6250457763672,66.6357879638672,66.6301727294922,66.5921401977539,66.5233383178711,66.4708938598633,66.4347610473633,66.2791519165039,66.2502975463867,66.2685546875,66.2615203857422,66.34375,66.3739776611328,66.44140625,66.40625,66.3868637084961,66.3583984375,66.3235321044922,66.1399383544922,66.1022415161133,66.0592269897461,65.9866180419922,65.8648452758789,65.830810546875,65.9300842285156,65.959716796875,66.0077133178711,65.97314453125,65.9408645629883,65.8123092651367,65.7900924682617,65.7950668334961,65.7713394165039,65.7825698852539,65.8411102294922,65.8714370727539,65.7880859375,65.7902297973633,65.7201690673828,65.6563491821289,65.6287155151367,65.630859375,65.5930633544922,65.6335983276367,65.7096176147461,65.8138198852539,65.8567886352539,65.90966796875,65.9779815673828,66.1946258544922,66.3043975830078,66.3239212036133,66.3478469848633,66.385498046875,66.3984375,66.3855895996094,66.3226547241211,66.2615203857422,66.203125,66.14111328125,65.9725570678711,65.9035186767578,65.9828567504883,66.0096664428711,65.9216766357422,65.8383331298828,65.8108901977539,65.7117233276367,65.6243591308594,65.6111297607422,65.5862731933594,65.5562057495117,65.55615234375,65.5225067138672,65.368896484375,65.3407745361328,65.287841796875,65.2549285888672,65.1416015625,65.1025390625,65.0488739013672,65.1087341308594,65.0819854736328,65.100830078125,65.0353546142578,64.987548828125,64.8688507080078,64.8780746459961,64.9153366088867,64.8254928588867,64.6731948852539,64.595947265625,64.5362777709961,64.4799270629883,64.423828125,64.3444290161133,64.3297348022461,64.2669448852539,64.2217788696289,64.2350158691406,64.2665100097656,64.2814407348633,64.29833984375,64.1773910522461,64.1210403442383,64.1544418334961,64.1625518798828,64.1317367553711,63.927734375,63.7623558044434,63.7252426147461,63.6261711120605,63.5336418151855,63.5078620910645,63.5138206481934,63.4122543334961,63.3489265441895,63.3092765808105,63.2737808227539,63.209228515625,63.1306648254395,63.0618667602539,63.0689430236816,63.0645027160645,63.0702590942383,63.1596183776855,63.1893577575684,63.2087898254395,63.1513214111328,63.1167449951172,63.0522499084473,62.9724578857422,62.9158706665039,62.8912620544434,62.7371101379395,62.7337913513184,62.6939926147461,62.7073249816895,62.72314453125,62.7130355834961,62.7266616821289,62.72021484375,62.6767120361328,62.6375007629395,62.5981903076172,62.5684585571289,62.51220703125,62.4660606384277,62.3971176147461,62.3477020263672,62.2890663146973,62.1527328491211,62.0591812133789,62.0135269165039,61.9534149169922,61.8572273254395,61.7713851928711,61.7746086120605,61.7553176879883,61.7174797058105,61.6817359924316,61.617431640625,61.5370140075684,61.3627967834473,61.0641098022461,60.7674827575684,60.5236778259277,60.5169448852539,60.5072746276855,60.5197715759277,60.5760269165039,60.5945816040039,60.5953636169434,60.5673866271973,60.52197265625,60.5029754638672,60.4729995727539,60.4449691772461,60.3906745910645,60.3328628540039,60.3010292053223,60.263427734375,60.0612258911133,59.9913063049316,59.9281272888184,59.9369125366211,59.9589385986328,60.0254936218262,59.9942359924316,59.9037590026855,59.858642578125,59.8493156433105,59.845947265625,59.8154754638672,59.8319320678711,59.8777351379395,59.9248085021973,59.9167976379395,59.8929176330566,59.9017066955566,59.8990707397461,59.922607421875,60.0145492553711,60.060791015625,60.1802711486816,60.244384765625,60.2735366821289,60.2047843933105,60.0955085754395,60.0294952392578,60.0166549682617,60.0499038696289,60.2029266357422,60.2959480285645,60.3729515075684,60.3827171325684,60.4162139892578,60.4572296142578,60.5319290161133,60.6552772521973,60.6646003723145,60.5071792602539,60.4545440673828,60.4449195861816,60.4682579040527,60.5188484191895,60.5418472290039,60.5794410705566,60.5997047424316,60.61572265625,60.7765121459961,60.9717788696289,61.0284156799316,61.0941429138184,61.18115234375,61.2183074951172,61.2055625915527,61.1757850646973,61.1292037963867,61.0968284606934,61.0223655700684,60.9620628356934,60.9049339294434,60.8603057861328,60.81640625,60.8203620910645,60.8113250732422,60.7828636169434,60.8003425598145,60.8426284790039,60.8474578857422,60.8270950317383,60.828857421875,60.8001441955566,60.7294921875,60.7247543334961,60.7219734191895,60.7424278259277,60.7692375183105,60.8068389892578,60.8559074401855,60.9457511901855,60.9977569580078,61.0156707763672,60.9994621276855,61.012939453125,61.0047378540039,61.1384773254395,61.1716804504395,61.1874046325684,61.2247085571289,61.2339859008789,61.2474136352539,61.2774391174316,61.3520050048828,61.4287147521973,61.5238800048828,61.5486793518066,61.5899391174316,61.6321296691895,61.6856460571289,61.7100563049316,61.7477989196777,61.7723121643066,61.8385238647461,61.8901901245117,61.9386215209961,61.9934043884277,62.0154800415039,62.0393524169922,62.0796852111816,62.1082000732422,62.1125984191895,62.0993156433105,62.0457534790039,62.0102081298828,61.9985847473145,62.0169410705566,62.0925788879395,62.1508750915527,62.2327156066895,62.2733421325684,62.2865219116211,62.3244667053223,62.3645057678223,62.4111328125,62.4662132263184,62.4731941223145,62.5307655334473,62.5780754089355,62.6797866821289,62.7219772338867,62.8087882995605,62.9037628173828,63.0446281433105,62.9767570495605,62.8287582397461,62.8220252990723,62.8488273620605,62.9449195861816,62.9711418151855,63.0000457763672,63.0512733459473,63.0907707214355,63.1669425964355,63.2575645446777,63.4364280700684,63.6422805786133,63.7580070495605,63.90478515625,64.006103515625,64.052978515625,64.1055679321289,64.1492691040039,64.162353515625,64.1703567504883,64.2142562866211,64.2031707763672,64.2088851928711,64.2293472290039,64.2658233642578,64.3128433227539,64.3043441772461,64.2233428955078,64.1590347290039,64.1230926513672,64.125,64.103271484375,64.0970230102539,64.1031799316406,64.1647415161133,64.205078125,64.314208984375,64.4631805419922,64.5605926513672,64.5727996826172,64.5675811767578,64.5589828491211,64.6167984008789,64.6446838378906,64.6781768798828,64.6931610107422,64.6825714111328,64.61474609375,64.4895553588867,64.447265625,64.5074234008789,64.5849075317383,64.7037582397461,64.7538528442383,64.778564453125,64.7665023803711,64.8533248901367,64.8852081298828,64.9275360107422,65.0518569946289,65.1139602661133,65.1967315673828,65.2011260986328,65.0969772338867,65.023681640625,64.862548828125,64.797607421875,64.7461395263672,64.6952209472656,64.6648406982422,64.6284637451172,64.707763671875,64.7857437133789,64.7581024169922,64.7015609741211,64.6231002807617,64.5518035888672,64.3770523071289,64.2799377441406,64.2319793701172,64.21875,64.2567901611328,64.3460922241211,64.4159164428711,64.5970687866211,64.6815414428711,64.79541015625,65.060546875,65.1549301147461,65.2051239013672,65.2213363647461,65.2750473022461,65.3288040161133,65.3484802246094,65.3626937866211,65.4419403076172,65.5307159423828,65.669921875,65.7131805419922,65.7464828491211,65.7510299682617,65.7757797241211,65.78564453125,65.7791442871094,65.7234878540039,65.7034149169922,65.6167907714844,65.5694808959961,65.4613265991211,65.4613800048828,65.5908203125,65.5660171508789,65.5745620727539,65.5940399169922,65.7708511352539,65.9771499633789,65.9790573120117,65.9874038696289,66.0343780517578,66.04833984375,66.0732879638672,66.1708984375,66.3624038696289,66.4376449584961,66.4701156616211,66.50732421875,66.587890625,66.6231460571289,66.6515579223633,66.68359375,66.7320327758789,66.8412094116211,66.8815383911133,66.8909683227539,66.853759765625,66.7540054321289,66.6978454589844,66.4466857910156,66.355224609375,66.296875,66.2411117553711,66.201416015625,66.177734375,66.1599578857422,66.1393585205078,66.1544952392578,66.2735366821289,66.3440475463867,66.4136734008789,66.5132827758789,66.5838394165039,66.6221694946289,66.6485366821289,66.721435546875,66.7538070678711,66.8268051147461,66.8527374267578,66.8501510620117,66.8599090576172,66.8811492919922,66.8975601196289,66.9090805053711,66.9068832397461,66.9193878173828,66.9319305419922,66.9246597290039,66.9459457397461,66.9864730834961,67.1355514526367,67.326904296875,67.4181594848633,67.4981918334961,67.5247039794922,67.5849609375,67.687255859375,67.7497024536133,67.7612838745117,67.7523422241211,67.6637191772461,67.6463851928711,67.667724609375,67.6365203857422,67.5088882446289,67.5279312133789,67.558837890625,67.693603515625,67.7338409423828,67.744384765625,67.7835388183594,67.806640625,67.7865753173828,67.7544937133789,67.7378387451172,67.7651901245117,67.7787094116211,67.8369140625,67.8179168701172,67.7949676513672,67.7732391357422,67.7577667236328,67.5745620727539,67.5364761352539,67.5490188598633,67.6682586669922,67.71533203125,67.7666015625,67.8368148803711,67.9705123901367,68.116943359375,68.207763671875,68.2179183959961,68.2045440673828,68.1456527709961,68.0754928588867,68.0567321777344,68.0547790527344,68.0771484375,68.1160659790039,68.1430130004883,68.1981964111328,68.2177734375,68.2418441772461,68.3255386352539,68.3852081298828,68.419921875,68.4163589477539,68.3935012817383,68.3839874267578,68.3653793334961,68.2730484008789,68.2518081665039,68.2208023071289,68.2186050415039,68.2615280151367,68.2783660888672,68.3098602294922,68.302734375,68.2932586669922,68.29736328125,68.3521499633789,68.4129943847656,68.6108932495117,68.6615219116211,68.708740234375,68.7011260986328,68.5481948852539,68.5348129272461,68.5471725463867,68.5984344482422,68.6191864013672,68.6826705932617,68.791259765625,68.8169937133789,68.7562942504883,68.7399368286133,68.7399368286133,68.9384307861328,69.0905227661133,69.1282730102539,69.1168441772461,69.1373977661133,69.1706008911133,69.1853485107422,69.2055206298828,69.2478561401367,69.2344741821289,69.2062530517578,69.20947265625,69.2748031616211,69.4117736816406,69.4742202758789,69.5990295410156,69.6630401611328,69.725341796875,69.7697296142578,69.7962417602539,69.8252410888672,69.935791015625,69.96630859375,69.994140625,70.0144500732422,70.0271530151367,70.0393524169922,70.0149459838867,70.0032272338867,70.0398941040039,70.0574188232422,70.0519027709961,69.9892044067383,69.9871597290039,70.0045394897461,70.0589370727539,70.078125,70.172119140625,70.234130859375,70.3019027709961,70.3533172607422,70.3885269165039,70.4216842651367,70.4684066772461,70.5711975097656,70.6568832397461,70.6992645263672,70.7516174316406,70.7892074584961,70.8201141357422,70.8099136352539,70.7960891723633,70.7666015625,70.7610321044922,70.7693862915039,70.7505798339844,70.7299346923828,70.686767578125,70.5032196044922,70.439453125,70.4317855834961,70.3636245727539,70.3648986816406,70.3968734741211,70.4175796508789,70.43798828125,70.453857421875,70.5290985107422,70.5887680053711,70.6875457763672,70.7428741455078,70.7680206298828,70.8526840209961,70.9030227661133,70.9188995361328,70.9922332763672,71.0104522705078,71.01904296875,71.0140151977539,70.9768524169922,70.9717330932617,70.9862823486328,71.0013198852539,71.04345703125,71.1190414428711,71.130126953125,71.1216354370117,71.1475601196289,71.189697265625,71.200439453125,71.1740188598633,71.1707000732422,71.1799850463867,71.3128890991211,71.3527297973633,71.3699722290039,71.412841796875,71.4576644897461,71.5015106201172,71.59912109375,71.6717300415039,71.6829071044922,71.66943359375,71.63671875,71.6299819946289,71.6722717285156,71.6626434326172,71.6019058227539,71.5359420776367,71.5399398803711,71.5790023803711,71.6067886352539,71.6401901245117,71.6858901977539,71.7101593017578,71.7751922607422,71.8074264526367,71.7897491455078,71.8196334838867,71.8708953857422,71.9357452392578,71.999755859375,72.0980529785156,72.1233444213867,72.15966796875,72.2925796508789,72.3258285522461,72.3510284423828,72.3626403808594,72.3417510986328,72.3188018798828,72.2849655151367,72.2398452758789,72.1834487915039,72.1340789794922,72.0800323486328,72.051513671875,71.9762649536133,71.8935546875,71.8499526977539,71.8055648803711,71.7576675415039,71.718017578125,71.6785202026367,71.6419906616211,71.6556625366211,71.6578674316406,71.5926742553711,71.5259246826172,71.458984375,71.4185104370117,71.4172821044922,71.3844757080078,71.3672409057617,71.3752899169922,71.4085998535156,71.4267578125,71.4717788696289,71.5535125732422,71.6267623901367,71.6915054321289,71.7386245727539,71.7682571411133,71.9576721191406,72.1106948852539,72.2684097290039,72.325439453125,72.3561019897461,72.37939453125,72.3941955566406,72.1995620727539,72.1788558959961,72.2226028442383,72.3004379272461,72.3185043334961,72.3111343383789,72.3543930053711,72.4198760986328,72.43701171875,72.4534683227539,72.5032730102539,72.4996109008789,72.534423828125,72.5719757080078,72.6416015625,72.7001953125,72.75048828125,72.7910690307617,72.8317413330078,72.8725128173828,72.9175796508789,72.9668426513672,73.01513671875,72.9606475830078,72.938232421875,72.9332046508789,72.9561538696289,72.9644012451172,72.9849166870117,72.9914016723633,73.0079116821289,73.0541000366211,73.1128463745117,73.1402816772461,73.1619110107422,73.2029266357422,73.2623062133789,73.3271026611328,73.3973617553711,73.4605026245117,73.3990707397461,73.3839797973633,73.4279251098633,73.4605026245117,73.504638671875,73.5368118286133,73.5581512451172,73.5907745361328,73.6274948120117,73.6703109741211,73.7595672607422,73.7516098022461,73.7597198486328,73.8334503173828,73.8954086303711,73.9306182861328,73.9459533691406,73.9638671875,74.0072784423828,74.0390625,74.1291046142578,74.1634292602539,74.1812057495117,74.1821823120117,74.1585464477539,74.1311950683594,74.1182174682617,74.1252899169922,74.1594772338867,74.195068359375,74.2191925048828,74.2783660888672,74.32958984375,74.3781204223633,74.4292449951172,74.4575653076172,74.486083984375,74.490478515625,74.5268096923828,74.6143035888672,74.6716766357422,74.6949691772461,74.7333450317383,74.7867660522461,74.8402328491211,74.8937454223633,74.9454574584961,75.0399856567383,75.1051788330078,75.2049331665039,75.2474670410156,75.2789535522461,75.3527374267578,75.3852996826172,75.4720687866211,75.5066909790039,75.6120071411133,75.6890640258789,75.7164001464844,75.73046875,75.7647018432617,75.8189010620117,75.8585968017578,75.8962936401367,75.9933166503906,76.09716796875,76.1578598022461,76.1799850463867,76.1856460571289,76.2423324584961,76.2604522705078,76.2521514892578,76.2615203857422,76.319091796875,76.35205078125,76.3394546508789,76.2778778076172,76.2171401977539,76.2089385986328,76.2644958496094,76.3033218383789,76.3165054321289,76.3040084838867,76.2530822753906,76.2162551879883,76.1725082397461,76.1515121459961,76.1463851928711,76.1305694580078,76.1298370361328,76.1442413330078,76.1727066040039,76.21533203125,76.2383346557617,76.24169921875,76.2196350097656,76.15478515625,76.13916015625,76.1459503173828,76.1661682128906,76.2178726196289,76.2129364013672,76.19482421875,76.15185546875,76.0499954223633,75.9773941040039,75.9687957763672,76.0670394897461,76.0907745361328,76.1501922607422,76.1866226196289,76.2808609008789,76.3318862915039,76.3717269897461,76.399169921875,76.436279296875,76.5613784790039,76.5866241455078,76.6167449951172,76.6356430053711,76.650634765625,76.6776809692383,76.6680221557617,76.6861343383789,76.7358932495117,76.7523956298828,76.7827606201172,76.8270568847656,76.8766174316406,76.9314498901367,76.9690399169922,76.9894561767578,76.8545913696289,76.807373046875,76.8218307495117,76.8441925048828,76.8690948486328,76.896484375,76.91650390625,76.9290008544922,76.98486328125,77.0286636352539,77.0738754272461,77.1205062866211,77.1543426513672,77.1754379272461,77.19384765625,77.22900390625,77.1953125,77.3069381713867,77.342529296875,77.3795852661133,77.3846740722656,77.3642044067383,77.3380355834961,77.2802734375,77.2977066040039,77.3498001098633,77.39306640625,77.4682083129883,77.515380859375,77.5644989013672,77.6156768798828,77.6812057495117,77.6866226196289,77.6708450317383,77.634521484375,77.5429153442383,77.5237808227539,77.5188980102539,77.5304641723633,77.544189453125,77.5927734375,77.6018600463867,77.58056640625,77.5288543701172,77.4926300048828,77.471923828125,77.4629364013672,77.4671401977539,77.5476608276367,77.58349609375,77.6377868652344,77.6903839111328,77.6995620727539,77.7171859741211,77.7982482910156,77.8313980102539,77.8413543701172,77.8431091308594,77.8000030517578,77.792724609375,77.7915496826172,77.8131332397461,77.83203125,77.8753890991211,77.8998031616211,77.9368133544922,77.9569320678711,77.9904251098633,78.085205078125,78.1548843383789,78.1943359375,78.2791061401367,78.2987289428711,78.3353042602539,78.3623046875,78.3995590209961,78.4820327758789,78.5043411254883,78.5527801513672,78.6231460571289,78.6389694213867,78.6426239013672,78.6384735107422,78.6558074951172,78.6901397705078,78.7249069213867,78.7776870727539,78.857421875,78.8667907714844,78.8819351196289,78.9230499267578,78.94287109375,78.979736328125,79.037841796875,79.0658187866211,79.06787109375,79.0803756713867,79.1168975830078,79.1233444213867,79.1376953125,79.1178207397461,79.1182098388672,79.1323699951172,79.1737289428711,79.2764587402344,79.3402328491211,79.4373092651367,79.5890121459961,79.7369613647461,79.8812484741211,79.9691848754883,80.0006332397461,80.0405731201172,80.0716781616211,80.0992660522461,80.1121063232422,80.1335983276367,80.141845703125,80.1369171142578,80.1044464111328,80.0824737548828,80.0859375,80.0777282714844,80.0479965209961,80.0240707397461,80.0294876098633,80.072265625,80.0802764892578,80.0762252807617,80.092041015625,80.1231460571289,80.1664581298828,80.22216796875,80.2800750732422,80.384521484375,80.4129867553711,80.5295867919922,80.5841751098633,80.625,80.6489715576172,80.6597213745117,80.68505859375,80.7665023803711,80.8363265991211,80.9660186767578,81,81.0564422607422,81.0573272705078,81.0432586669922,81.0138626098633,80.8855895996094,80.8895492553711,80.9380798339844,81.0833511352539,81.1431198120117,81.2069778442383,81.2183609008789,81.214111328125,81.1943893432617,81.1727523803711,81.1375961303711,81.11572265625,81.1167984008789,81.1335906982422,81.1884765625,81.281494140625,81.3960952758789,81.5323257446289,81.6318893432617,81.694580078125,81.7468719482422,81.8095703125,81.8553695678711,81.920166015625,81.9373550415039,81.9330062866211,81.9026336669922,81.884033203125,81.8251953125,81.6900405883789,81.6222152709961,81.5917434692383,81.5398941040039,81.429931640625,81.3827133178711,81.3656234741211,81.3628921508789,81.394287109375,81.4599609375,81.5321807861328,81.6620178222656,81.753662109375,81.8582077026367,81.92041015625,81.9820785522461,82.0066375732422,82.0930709838867,82.1683120727539,82.2271423339844,82.2211380004883,82.2457504272461,82.2828598022461,82.299560546875,82.3513641357422,82.3506317138672,82.3260726928711,82.2792434692383,82.2368621826172,82.1640625,82.0615768432617,81.738037109375,81.6768569946289,81.6532745361328,81.6883773803711,81.7536163330078,81.7997589111328,81.8709945678711,81.9671401977539,82.0383758544922,82.1189498901367,82.251220703125,82.3217315673828,82.3217315673828,82.0782165527344,82.025634765625,81.8952178955078,81.9090805053711,81.8930206298828,81.8977966308594,81.9180679321289,81.9721221923828,82.1207046508789,82.2373580932617,82.3828125,82.4601593017578,82.4740676879883,82.472412109375,82.4054183959961,82.1736297607422,82.0969772338867,81.8288116455078,81.7882843017578,81.7798385620117,81.8129425048828,81.8489227294922,81.8968200683594,81.9566879272461,82.02587890625,82.1044387817383,82.2600631713867,82.310791015625,82.3681640625,82.4717254638672,82.5426254272461,82.7252426147461,82.7470245361328,82.7709426879883,82.7849578857422,82.741455078125,82.7254867553711,82.7098159790039,82.6891555786133,82.6803207397461,82.7049789428711,82.75,82.7786178588867,82.8567886352539,82.883544921875,82.8855972290039,82.8650894165039,82.8548812866211,82.8588333129883,82.951904296875,83.0638732910156,83.0613327026367,83.0176773071289,83.0786666870117,83.1290512084961,83.1468200683594,83.255126953125,83.2646026611328,83.2587890625,83.2319793701172,83.2051696777344,83.1477508544922,83.1300277709961,83.1267547607422,83.1007843017578,83.1848602294922,83.2751998901367,83.3321838378906,83.2989273071289,83.2555694580078,83.2039031982422,83.1753387451172,82.9988708496094,82.9986343383789,83.0135726928711,83.0462875366211,83.0948257446289,83.1607437133789,83.1851043701172,83.2581558227539,83.2862854003906,83.3325729370117,83.370849609375,83.4010238647461,83.414794921875,83.4022903442383,83.412109375,83.4376449584961,83.4855422973633,83.4977493286133,83.4991226196289,83.4684066772461,83.4658203125,83.4799346923828,83.5099105834961,83.5289611816406,83.5369644165039,83.5386199951172,83.5457534790039,83.5684585571289,83.5711441040039,83.5575714111328,83.5286636352539,83.5299835205078,83.5772476196289,83.599609375,83.5934066772461,83.5648422241211],[17.8148441314697,17.572265625,17.2912120819092,17.0846672058105,16.8089847564697,16.5271492004395,16.142822265625,15.8944339752197,15.8886709213257,15.9006834030151,15.8898439407349,15.890625,15.8689947128296,15.8624992370605,15.8065433502197,15.76416015625,15.849609375,15.9010744094849,15.9273433685303,15.9502935409546,15.72900390625,15.6948728561401,15.6160154342651,15.481201171875,15.3604974746704,15.2510251998901,15.1524419784546,15.1426763534546,15.0723152160645,15.0398921966553,14.9005870819092,14.8660640716553,14.7887201309204,14.6692380905151,14.6068859100342,14.5299806594849,14.4607419967651,14.416015625,14.4276361465454,14.4311037063599,14.4137706756592,14.4099111557007,14.3900880813599,14.3470697402954,14.2772464752197,14.2412595748901,14.2246580123901,14.1827144622803,14.1413087844849,14.0770015716553,14.0500974655151,14.0550775527954,14.0456056594849,13.9973640441895,13.904052734375,13.8347654342651,13.7830076217651,13.7365226745605,13.9009284973145,13.9290037155151,13.9255857467651,13.9901847839355,14.1149406433105,14.2282218933105,14.54541015625,14.5709962844849,14.6300773620605,14.691014289856,14.7613277435303,14.8183603286743,14.9013175964355,14.9635753631592,15.0019540786743,15.0267572402954,15.07421875,15.2376947402954,15.2749996185303,15.3208980560303,15.4955558776855,15.7032232284546,15.9323720932007,16.0701656341553,16.0704574584961,16.0706539154053,16.07080078125,16.071044921875,16.0711917877197,16.0727062225342,16.162353515625,16.2613773345947,16.351318359375,16.3910160064697,16.4395523071289,16.4678211212158,16.5107421875,16.5651359558105,16.6309070587158,16.7081069946289,16.787109375,16.8678226470947,16.9761714935303,17.1122550964355,17.1998043060303,17.2364253997803,17.255859375,17.2541007995605,17.25244140625,17.4474601745605,17.6207523345947,17.81640625,17.8161125183105,17.8157215118408,17.8153324127197,17.8149909973145,17.8148441314697],[13.25927734375,13.2575197219849,13.2910652160645,13.3134765625,13.4287109375,13.52685546875,13.6223630905151,13.6146488189697,13.5990238189697,13.5703125,13.4111328125,13.25927734375],[5.54843711853027,5.48525428771973,5.44516658782959,5.37397432327271,5.33535146713257,5.24677705764771,5.24287128448486,5.23154306411743,5.21420860290527,5.19540977478027,5.17851591110229,5.15703105926514,5.10585975646973,5.049560546875,5.02016639709473,5.00458955764771,5.00449228286743,5.00068330764771,4.99106454849243,4.95449209213257,4.92905282974243,4.92304706573486,4.880615234375,4.82041025161743,4.77929735183716,4.72431659698486,4.66816425323486,4.57709980010986,4.50678682327271,4.453125,4.34995126724243,4.23647499084473,4.17192411422729,4.10166025161743,4.00195360183716,3.856689453125,3.78725576400757,3.67597627639771,3.58828139305115,3.51738262176514,3.42373061180115,3.39453148841858,3.37094736099243,3.35283207893372,3.35429644584656,3.36225628852844,3.37543940544128,3.37709975242615,3.35361313819885,3.21884775161743,3.164306640625,3.14228510856628,3.10888671875,3.07856464385986,3.00307655334473,2.96337890625,2.8828125,2.853271484375,2.83325242996216,2.77553701400757,2.76826167106628,2.74785161018372,2.66567397117615,2.64111304283142,2.63750004768372,2.60898447036743,2.53217768669128,2.51323246955872,2.45683622360229,2.39536118507385,2.32597661018372,2.27714848518372,2.22666001319885,2.11489272117615,2.03647470474243,2.01601552963257,2.00507855415344,1.97480452060699,1.94213843345642,1.92724609375,1.90722632408142,1.92265617847443,1.91430652141571,1.89218735694885,1.88125014305115,1.91640639305115,1.92124044895172,1.93647480010986,2.00581049919128,2.01396489143372,1.98159182071686,1.95922863483429,1.96347665786743,1.94013655185699,1.90893542766571,1.77382802963257,1.72607433795929,1.70410132408142,1.70478487014771,1.70000016689301,1.66728544235229,1.65058600902557,1.64843738079071,1.57431614398956,1.53994154930115,1.52026379108429,1.51435554027557,1.51699233055115,1.54785180091858,1.56328094005585,1.57431614398956,1.59194314479828,1.58754849433899,1.55668962001801,1.53022468090057,1.48173809051514,1.46625971794128,1.43867182731628,1.34775376319885,1.31225621700287,1.28466820716858,1.27915060520172,1.28105461597443,1.24750959873199,1.20849633216858,1.20122087001801,1.20361304283142,1.24887669086456,1.30458998680115,1.34365248680115,1.37602519989014,1.46459996700287,1.50820314884186,1.52734398841858,1.63242208957672,1.70000016689301,1.71801745891571,1.74628889560699,1.79521489143372,1.84233379364014,1.86147451400757,1.87417018413544,1.90063464641571,1.96240210533142,2.12163066864014,2.27412104606628,2.32705092430115,2.36293911933899,2.58837866783142,2.68999028205872,2.76542949676514,2.99047875404358,3.08784198760986,3.28310537338257,3.34921836853027,3.39858388900757,3.462158203125,3.58750009536743,3.66655278205872,3.69980454444885,3.75273442268372,3.81967735290527,3.88344717025757,3.93354487419128,3.96000957489014,3.97539091110229,4.02314472198486,4.160400390625,4.18818330764771,4.22675800323486,4.28764677047729,4.353515625,4.381103515625,4.41665029525757,4.47592782974243,4.48032236099243,4.501708984375,4.50458955764771,4.51118135452271,4.53325176239014,4.56962871551514,4.59765625,4.66665029525757,4.74052715301514,4.81269550323486,4.90751886367798,4.98984384536743,5.08286142349243,5.14399385452271,5.19423866271973,5.23881816864014,5.23881816864014,5.25795888900757,5.19931650161743,5.21015644073486,5.18808555603027,5.19248056411743,5.22114276885986,5.20205068588257,5.43740224838257,5.67421913146973,5.906982421875,5.93876934051514,6.04951190948486,6.12919950485229,6.17441368103027,6.21430683135986,6.38510751724243,6.446533203125,6.51337909698486,6.58837890625,6.65092802047729,6.69453096389771,6.71137714385986,6.72661161422729,6.73276329040527,6.7578125,6.78691434860229,6.78847694396973,6.76831007003784,6.80595684051514,6.85707950592041,6.94536113739014,7.00288105010986,7.092041015625,7.13398456573486,7.15000009536743,7.16455078125,7.16655302047729,7.14370107650757,7.15620088577271,7.21108388900757,7.256591796875,7.32084941864014,7.36332988739014,7.49868106842041,7.53593730926514,7.59663105010986,7.64833974838257,7.77202081680298,7.81318378448486,7.82763671875,7.85400390625,7.91943359375,7.99404335021973,8.05356502532959,8.16201210021973,8.19160079956055,8.24868106842041,8.27915000915527,8.30595779418945,8.54931640625,8.53261756896973,8.37382793426514,8.33950233459473,8.33872127532959,8.37998008728027,8.36259746551514,8.25400352478027,8.07460975646973,7.735595703125,7.60664081573486,7.5458984375,7.39804697036743,7.32578086853027,7.03813409805298,6.84365272521973,6.69731473922729,6.50253963470459,6.39077091217041,6.45151424407959,6.62724590301514,6.73398447036743,6.85117197036743,6.87929677963257,6.82939434051514,6.82060527801514,6.785888671875,6.59853458404541,6.45039033889771,6.33154249191284,6.27211904525757,6.17841815948486,6.09731435775757,5.88501024246216,5.56459951400757,5.54843711853027],[22.2105484008789,22.1951656341553,22.2683601379395,22.292236328125,22.296630859375,22.2635726928711,22.2335453033447,22.2105484008789],[22.2104988098145,22.21044921875,22.220458984375,22.2416973114014,22.2802734375,22.3333988189697,22.2775402069092,22.2104988098145],[22.5119132995605,22.5144538879395,22.54296875,22.55126953125,22.5649890899658,22.5649890899658,22.553955078125,22.5409660339355,22.5367679595947,22.4994621276855,22.4576168060303,22.4374027252197,22.396240234375,22.373779296875,22.3252925872803,22.2955551147461,22.3484363555908,22.3758792877197,22.364990234375,22.3960952758789,22.4281749725342,22.4840335845947,22.5119132995605],[-53.1371078491211,-53.1845703125,-53.1841812133789,-53.1467781066895,-53.0298805236816,-53.021484375,-52.9893569946289,-52.9757804870605,-52.96630859375,-52.9999008178711,-53.0271453857422,-53.0912094116211,-53.11279296875,-53.1298828125,-53.1371078491211],[14.9930658340454,15.008056640625,14.9770011901855,14.907567024231,14.8764162063599,14.8835439682007,14.8562507629395,14.7944812774658,14.7710933685303,14.7860841751099,14.7708988189697,14.7506351470947,14.7310056686401,14.7204103469849,14.7260265350342,14.7478523254395,14.738525390625,14.6981449127197,14.6873540878296,14.7063484191895,14.6917486190796,14.6437015533447,14.6333990097046,14.6610851287842,14.7133779525757,14.7903804779053,14.8097658157349,14.7524423599243,14.685546875,14.6447267532349,14.607666015625,14.5829591751099,14.5706052780151,14.525146484375,14.4466304779053,14.3859872817993,14.3433103561401,14.3118162155151,14.2916498184204,14.2238779067993,14.1086912155151,14.0282220840454,13.9825687408447,13.9318361282349,13.8760738372803,13.8586921691895,13.85205078125,13.8444337844849,13.9656744003296,14.0501461029053,14.0372066497803,13.9945802688599,13.8994626998901,13.7700681686401,13.7556638717651,13.7748537063599,13.7634763717651,13.7461423873901,13.69873046875,13.63525390625,13.4072265625,13.3133792877197,13.2843751907349,13.27978515625,13.2665033340454,13.2235841751099,13.1793947219849,13.1175298690796,13.0537118911743,13.0078125,12.991455078125,12.979248046875,13.084716796875,13.1274404525757,13.2154302597046,13.27490234375,13.3105955123901,13.3529300689697,13.3855953216553,13.3600587844849,13.3766603469849,13.399169921875,13.4513673782349,13.4831047058105,13.5060062408447,13.5213861465454,13.580322265625,13.6499509811401,13.8126945495605,13.8410654067993,13.8899898529053,13.8949708938599,13.879638671875,13.9046382904053,13.9605960845947,13.9873533248901,13.9426755905151,13.8753900527954,13.8509769439697,13.854248046875,13.9045400619507,13.9642086029053,13.9789543151855,14.0001459121704,14.0155277252197,14.0320796966553,14.072265625,14.124755859375,14.1636714935303,14.2527341842651,14.2969732284546,14.3291492462158,14.3702144622803,14.3603019714355,14.4113759994507,14.416015625,14.4607419967651,14.5299806594849,14.6068859100342,14.6692380905151,14.7887201309204,14.8660640716553,14.9005870819092,15.0398921966553,15.0723152160645,15.1426763534546,15.1524419784546,15.2510251998901,15.3604974746704,15.481201171875,15.6160154342651,15.6948728561401,15.72900390625,15.7010250091553,15.7648429870605,15.7861824035645,15.8625984191895,15.8793458938599,15.9106435775757,15.9098634719849,15.8323736190796,15.7901849746704,15.8264656066895,15.8344240188599,15.7623538970947,15.7942390441895,15.8010740280151,15.783203125,15.8851566314697,15.9056644439697,15.9534177780151,16.0022449493408,16.024169921875,16.0028324127197,15.8995122909546,15.9181642532349,15.9739751815796,15.9898929595947,15.8835945129395,15.8020009994507,15.7939453125,15.8125982284546,15.8294925689697,15.8472652435303,15.8727531433105,15.8226079940796,15.4368648529053,15.4054679870605,15.5196294784546,15.5108880996704,15.492431640625,15.4301261901855,15.4009275436401,15.3976078033447,15.4144048690796,15.39404296875,15.3527336120605,15.2892570495605,15.2203607559204,15.2192392349243,15.2607431411743,15.2657718658447,15.2193841934204,15.2221202850342,15.2939939498901,15.3654289245605,15.3684072494507,15.2399911880493,15.0789060592651,15.0422840118408,14.9930658340454],[16.3783683776855,16.3002452850342,16.3017578125,16.3621101379395,16.4138660430908,16.439208984375,16.4282245635986,16.3783683776855],[16.4615230560303,16.4036121368408,16.4296875,16.4833011627197,16.5139656066895,16.5108890533447,16.4877433776855,16.4615230560303],[42.76904296875,42.7003402709961,42.7903785705566,42.7986297607422,42.8003883361816,42.76904296875],[42.99658203125,42.9686508178711,42.9814910888672,42.96435546875,42.9326171875,42.9170379638672,42.9148902893066,42.9277839660645,42.8955078125,42.9127426147461,42.9336929321289,42.9599113464355,42.99658203125],[42.8971176147461,42.9154777526855,42.9022445678711,42.8450698852539,42.8074226379395,42.7412605285645,42.6905746459961,42.5994148254395,42.586669921875,42.5597190856934,42.52294921875,42.4811019897461,42.4329071044922,42.5278778076172,42.634033203125,42.7974128723145,42.837158203125,42.9684562683105,43.014892578125,43.0255851745605,43.02587890625,42.8506851196289,42.8971176147461],[43.1257820129395,43.1154289245605,43.1231422424316,43.1438980102539,43.1973648071289,43.2137718200684,43.229248046875,43.2137718200684,43.1749496459961,43.1438980102539,43.1257820129395],[43.2706565856934,43.26806640625,43.2861785888672,43.3172378540039,43.3434066772461,43.3870582580566,43.3818855285645,43.350830078125,43.3146476745605,43.2979507446289,43.2706565856934],[43.9738273620605,43.8995094299316,43.914794921875,43.9607925415039,44.0107421875,43.9738273620605],[44.0623054504395,44.0270500183105,44.0933113098145,44.1378402709961,44.1576652526855,44.0623054504395],[43.9223594665527,43.90771484375,43.8977546691895,43.9118156433105,43.9072761535645,44.1255378723145,44.16796875,44.1171875,43.9223594665527],[44.3357429504395,44.3091773986816,44.358154296875,44.3930168151855,44.434326171875,44.48583984375,44.5447235107422,44.6647453308105,44.6973609924316,44.648681640625,44.6182632446289,44.61083984375,44.5342292785645,44.4357414245605,44.3501930236816,44.3475570678711,44.3357429504395],[44.7589378356934,44.71484375,44.754638671875,44.7698707580566,44.7998046875,44.8243675231934,44.84814453125,44.8448257446289,44.8213844299316,44.7589378356934],[44.6600608825684,44.6212387084961,44.6703109741211,44.75830078125,44.900390625,44.9404296875,44.9799308776855,45.0199699401855,45.1446266174316,45.1649894714355,45.1674346923828,45.0809555053711,45.03125,44.97021484375,44.8691902160645,44.725341796875,44.693359375,44.6600608825684],[44.97705078125,44.9556159973145,44.9939460754395,45.0254402160645,45.035400390625,45.0792007446289,45.0986328125,45.1468276977539,45.2247543334961,45.1780281066895,45.0900421142578,45.0654792785645,44.97705078125],[45.9317398071289,45.9076156616211,45.8655281066895,45.8357429504395,45.76708984375,45.6558113098145,45.600830078125,45.5580062866211,45.5272483825684,45.5149917602539,45.502197265625,45.4658203125,45.3995132446289,45.3369140625,45.2779808044434,45.26806640625,45.2454109191895,45.2306175231934,45.2124977111816,45.1890640258789,45.1730003356934,45.1672859191895,45.1677742004395,45.1962394714355,45.1754875183105,45.1517105102539,45.13720703125,44.9737815856934,44.9267578125,44.9109878540039,44.9175300598145,44.9193840026855,44.9040031433105,44.8691902160645,44.8651885986328,44.8832511901855,44.9148941040039,44.9472160339355,44.9772453308105,45.0265121459961,45.0774421691895,45.0858383178711,45.1020050048828,45.1205558776855,45.13427734375,45.1329116821289,45.119384765625,45.1417961120605,45.1118659973145,45.0772476196289,45.078125,45.1584014892578,45.1634750366211,45.12255859375,45.120361328125,45.13330078125,45.1639633178711,45.1705589294434,45.1560554504395,45.1717758178711,45.2765617370605,45.1968765258789,45.2166976928711,45.1620140075684,45.058349609375,45.0088348388672,45.026611328125,45.0722160339355,45.1896018981934,45.2107925415039,45.2157211303711,45.2027816772461,45.178955078125,45.0075187683105,44.8563957214355,44.7658195495605,44.6819343566895,44.53759765625,44.52099609375,44.4737319946289,44.3520011901855,44.2151374816895,44.12451171875,44.0596160888672,44.0025901794434,43.9131851196289,43.8150367736816,43.77880859375,43.6490249633789,43.5165519714355,43.47021484375,43.4457511901855,43.3438491821289,43.3056144714355,43.1989250183105,43.0427742004395,43.006591796875,42.9800796508789,42.9597663879395,42.9383811950684,42.9622535705566,43.1148910522461,43.2111320495605,43.3924293518066,43.4640617370605,43.5433616638184,43.53125,43.5063018798828,43.5055198669434,43.519775390625,43.5689468383789,43.6069793701172,43.6566390991211,43.7359390258789,43.811279296875,43.9087905883789,44.172119140625,44.2567863464355,44.2728996276855,44.2714347839355,44.288818359375,44.2892570495605,44.27197265625,44.3282737731934,44.3834953308105,44.6029281616211,44.7065887451172,44.8182601928711,44.9713859558105,45.0810089111328,45.222900390625,45.2977066040039,45.3421401977539,45.3377914428711,45.2825164794922,45.15966796875,44.9976081848145,44.9271965026855,44.8356437683105,44.829345703125,44.83740234375,44.9915046691895,45.1082000732422,45.1634254455566,45.2313995361328,45.4817848205566,45.5168914794922,45.4767570495605,45.4283714294434,45.4498023986816,45.4826164245605,45.5033683776855,45.5094223022461,45.4778327941895,45.4851570129395,45.4866218566895,45.4814453125,45.5057640075684,45.59521484375,45.645263671875,45.6572265625,45.6512718200684,45.610107421875,45.5714836120605,45.5084953308105,45.4782218933105,45.4673347473145,45.49267578125,45.4999046325684,45.4507789611816,45.44140625,45.467041015625,45.5022926330566,45.5415534973145,45.5796852111816,45.6126441955566,45.6455078125,45.659912109375,45.7177238464355,45.7326164245605,45.797607421875,45.8340339660645,45.8621597290039,45.9044456481934,45.9836921691895,46.0484848022461,46.1092262268066,46.1399917602539,46.1719245910645,46.2007331848145,46.2132301330566,46.2339859008789,46.2578620910645,46.2776374816895,46.3053703308105,46.371337890625,46.3822250366211,46.3728523254395,46.3891105651855,46.4838409423828,46.5079078674316,46.5213890075684,46.5346183776855,46.5244140625,46.4999046325684,46.4850120544434,46.4164047241211,46.3393058776855,46.253662109375,46.1873016357422,46.1403312683105,46.0766143798828,45.9961433410645,45.9510726928711,45.9413070678711,45.9137687683105,45.8683624267578,45.8272476196289,45.7904281616211,45.770263671875,45.7654762268066,45.7644538879395,45.7530250549316,45.767333984375,45.796142578125,45.8132820129395,45.907470703125,45.8993682861328,45.9108428955078,45.9317398071289,45.9317398071289],[18.7776851654053,18.7071285247803,18.7909183502197,18.896728515625,18.9540538787842,18.96728515625,18.9320316314697,18.8614749908447,18.7776851654053],[19.7181644439697,19.6881847381592,19.486572265625,19.4219722747803,19.324462890625,19.2858390808105,19.1959476470947,19.1635246276855,19.1307621002197,19.0455074310303,18.9870128631592,18.9200210571289,18.8563976287842,18.80322265625,18.73291015625,18.6455078125,18.6141605377197,18.6103515625,18.597900390625,18.5125980377197,18.4162120819092,18.34130859375,18.2707996368408,18.2039546966553,18.0391597747803,18.119140625,18.1860847473145,18.2120113372803,18.2285633087158,18.2199230194092,18.2083988189697,18.1869144439697,18.1762218475342,18.1561527252197,18.1517562866211,18.2056140899658,18.2335453033447,18.2511730194092,18.245361328125,18.2290534973145,18.1902351379395,18.1217784881592,18.0582027435303,18.0418949127197,18.1431655883789,18.2151355743408,18.2691898345947,18.34619140625,18.39306640625,18.4500007629395,18.6247062683105,18.6566886901855,18.6626949310303,18.64111328125,18.6014156341553,18.575439453125,18.5653324127197,18.5223636627197,18.4557132720947,18.434814453125,18.442138671875,18.4682140350342,18.5153312683105,18.55078125,18.5586910247803,18.574462890625,18.6237297058105,18.6749515533447,18.7435531616211,18.8941402435303,19.0715827941895,19.13134765625,19.2406253814697,19.3418464660645,19.4410648345947,19.5260753631592,19.6107425689697,19.6373043060303,19.65869140625,19.7221183776855,19.8074226379395,19.8545894622803,19.8836917877197,19.90380859375,19.9280757904053,19.90087890625,19.8132801055908,19.74462890625,19.7216796875,19.6967296600342,19.7181644439697],[20.0375003814697,20.01416015625,19.98583984375,20.00341796875,20.0354499816895,20.0274410247803,20.0314464569092,20.062255859375,20.0858402252197,20.0936508178711,20.0918941497803,20.0375003814697],[48.4053230285645,48.4134292602539,48.4121589660645,48.4073753356934,48.3608894348145,48.3580093383789,48.3272933959961,48.28662109375,48.2560539245605,48.2433090209961,48.2053718566895,48.134033203125,48.1043930053711,48.1036109924316,48.1070289611816,48.109619140625,48.09521484375,48.060302734375,48.029541015625,47.9970703125,47.9603004455566,47.947265625,47.9225578308105,47.7990226745605,47.7663116455078,47.7595710754395,47.7725601196289,47.7626457214355,47.7362289428711,47.7278327941895,47.6963844299316,47.6290550231934,47.572021484375,47.53662109375,47.5050277709961,47.3957023620605,47.3642578125,47.3325691223145,47.3045883178711,47.1381340026855,47.0848121643066,47.0438995361328,47.0065422058105,46.9637680053711,46.8783683776855,46.7897453308105,46.7533683776855,46.7042961120605,46.6478538513184,46.6207504272461,46.6078147888184,46.5724601745605,46.4863777160645,46.44775390625,46.4123039245605,46.3915519714355,46.3526878356934,46.3043441772461,46.2824211120605,46.2422332763672,46.2597160339355,46.2462425231934,46.2174797058105,46.1944351196289,46.1728019714355,46.1456527709961,46.1334953308105,46.1669425964355,46.1330070495605,46.1085929870605,46.1260261535645,46.1418914794922,46.1614761352539,46.1458969116211,46.1519050598145,46.169189453125,46.1551742553711,46.0873527526855,46.064453125,46.0498046875,46.0285148620605,46.0028800964355,45.9844245910645,45.9870109558105,46.0161628723145,46.009521484375,45.982666015625,45.959716796875,45.931396484375,45.9317398071289,45.9317398071289,45.9108428955078,45.8993682861328,45.907470703125,45.8132820129395,45.796142578125,45.767333984375,45.7530250549316,45.7644538879395,45.7654762268066,45.770263671875,45.7904281616211,45.8272476196289,45.8683624267578,45.9137687683105,45.9413070678711,45.9510726928711,45.9961433410645,46.0766143798828,46.1403312683105,46.1873016357422,46.253662109375,46.3393058776855,46.4164047241211,46.4850120544434,46.4999046325684,46.5220718383789,46.6072273254395,46.638671875,46.6808128356934,46.7047843933105,46.7216339111328,46.7825202941895,46.8279800415039,46.8572769165039,46.86328125,46.971923828125,47.002197265625,46.9969711303711,47.0067863464355,47.0224609375,47.057861328125,47.0912590026855,47.1226577758789,47.140380859375,47.1459007263184,47.2234344482422,47.2527351379395,47.2731437683105,47.367431640625,47.3995132446289,47.404541015625,47.4246559143066,47.4475593566895,47.4766120910645,47.5360374450684,47.60888671875,47.6562995910645,47.6744651794434,47.695068359375,47.7244606018066,47.7473640441895,47.7505340576172,47.739013671875,47.686279296875,47.6786613464355,47.6939964294434,47.697265625,47.6953125,47.7075691223145,47.7637672424316,47.8045425415039,47.8371086120605,47.8729476928711,47.90087890625,47.963623046875,48.0059585571289,48.0120582580566,48.0043449401855,47.9933586120605,47.9909172058105,47.8875961303711,47.8099098205566,47.7701644897461,47.7668952941895,47.763427734375,47.7770042419434,47.7871589660645,47.8064956665039,47.8528823852539,47.8926734924316,47.939453125,48.0002899169922,48.0508270263672,48.0730476379395,48.1106948852539,48.162109375,48.2128410339355,48.2230949401855,48.1998062133789,48.155029296875,48.13134765625,48.1466331481934,48.2220191955566,48.2955589294434,48.4951171875,48.5269050598145,48.5497093200684,48.545654296875,48.5196762084961,48.5059051513672,48.5105972290039,48.553466796875,48.55224609375,48.5218772888184,48.4957008361816,48.4636726379395,48.4185066223145,48.4014663696289,48.3783683776855,48.3465805053711,48.3380851745605,48.3933601379395,48.4053230285645],[-10.9092769622803,-10.90966796875,-10.8991212844849,-10.81103515625,-10.7618169784546,-10.6984376907349,-10.6399412155151,-10.5181636810303,-10.4862308502197,-10.4724607467651,-10.4749021530151,-10.5675783157349,-10.6226568222046,-10.6512689590454,-10.6984376907349,-10.80615234375,-10.8763675689697,-10.9092769622803],[-10.59033203125,-10.6021480560303,-10.5731439590454,-10.5556640625,-10.5074224472046,-10.4388666152954,-10.4330081939697,-10.4469728469849,-10.5284185409546,-10.59033203125],[-10.3026371002197,-10.3374996185303,-10.26416015625,-10.17138671875,-10.1399412155151,-10.1769523620605,-10.1939458847046,-10.2271480560303,-10.3026371002197],[-9.37470722198486,-9.41972637176514,-9.50634765625,-9.54287147521973,-9.60312461853027,-9.64785194396973,-9.65468788146973,-9.64560508728027,-9.67402362823486,-9.71904373168945,-9.80644607543945,-9.90312576293945,-9.95703125,-10.0375003814697,-10.1084966659546,-10.2066402435303,-10.2279300689697,-10.2356443405151,-10.2940425872803,-10.2634763717651,-10.2422857284546,-10.2000980377197,-10.1228513717651,-10.0397462844849,-9.96650314331055,-9.91748046875,-9.77353572845459,-9.76054668426514,-9.77109336853027,-9.77177715301514,-9.70693302154541,-9.66904258728027,-9.62050724029541,-9.57285118103027,-9.51933574676514,-9.47207069396973,-9.44023418426514,-9.38447284698486,-9.36718654632568,-9.36982440948486,-9.35244178771973,-9.38046932220459,-9.35957050323486,-9.30146503448486,-9.32158184051514,-9.37470722198486],[-9.51191425323486,-9.56533241271973,-9.66562557220459,-9.759765625,-9.91416072845459,-9.99296855926514,-10.0861330032349,-10.1486330032349,-10.1698246002197,-10.1833009719849,-10.2948236465454,-10.3435544967651,-10.34716796875,-10.3109369277954,-10.2701177597046,-10.2150392532349,-10.1677732467651,-10.1288089752197,-10.07861328125,-10.01513671875,-9.966796875,-9.83808612823486,-9.70527362823486,-9.61484336853027,-9.453125,-9.37294864654541,-9.34160137176514,-9.37539100646973,-9.41640567779541,-9.42314529418945,-9.41386699676514,-9.42792987823486,-9.41376972198486,-9.34990215301514,-9.31435585021973,-9.23857402801514,-9.19033241271973,-9.15537071228027,-9.11669921875,-9.06181621551514,-8.96845626831055,-8.94248104095459,-9.03154277801514,-9.05341720581055,-9.06425762176514,-9.00400352478027,-9.01542949676514,-9.04257774353027,-9.12294864654541,-9.18984317779541,-9.19492149353027,-9.21376895904541,-9.25468730926514,-9.29423809051514,-9.32597732543945,-9.3818359375,-9.51191425323486],[-8.76982498168945,-8.80419921875,-8.74287128448486,-8.71543025970459,-8.67539024353027,-8.669921875,-8.71318340301514,-8.76982498168945],[-8.54521465301514,-8.60400390625,-8.58730506896973,-8.53066444396973,-8.4970703125,-8.44834041595459,-8.43984317779541,-8.45693397521973,-8.47578144073486,-8.49482440948486,-8.54521465301514],[-8.74101543426514,-8.75048828125,-8.73603534698486,-8.64707088470459,-8.58652400970459,-8.5390625,-8.45498085021973,-8.42919921875,-8.45566368103027,-8.47294998168945,-8.48105430603027,-8.48261642456055,-8.51884746551514,-8.55341815948486,-8.58935546875,-8.62822246551514,-8.67177677154541,-8.74101543426514],[-8.35478591918945,-8.39863300323486,-8.3955078125,-8.33779335021973,-8.27480506896973,-8.25380897521973,-8.23544979095459,-8.26904296875,-8.35478591918945],[-8.61386680603027,-8.82099628448486,-8.85439395904541,-8.88613319396973,-8.92900371551514,-8.90615272521973,-8.912109375,-8.87314510345459,-8.82558536529541,-8.78789138793945,-8.74277400970459,-8.75800704956055,-8.76523494720459,-8.74492168426514,-8.611328125,-8.43740272521973,-8.29521465301514,-8.23798847198486,-8.20419979095459,-8.28271484375,-8.30410099029541,-8.33603477478027,-8.38691425323486,-8.61386680603027],[-8.32949161529541,-8.39130783081055,-8.49872970581055,-8.53144454956055,-8.55166053771973,-8.44384765625,-8.44892501831055,-8.35410213470459,-8.31865215301514,-8.31777381896973,-8.35615253448486,-8.3642578125,-8.20341777801514,-8.20175743103027,-8.20869159698486,-8.22880840301514,-8.32949161529541],[-8.2724609375,-8.29960918426514,-8.42441463470459,-8.42246055603027,-8.47793006896973,-8.50166034698486,-8.52382850646973,-8.544921875,-8.56679630279541,-8.53232479095459,-8.57607460021973,-8.58662128448486,-8.53564453125,-8.53857421875,-8.53066444396973,-8.43906211853027,-8.35371112823486,-8.32226657867432,-8.31337833404541,-8.30058574676514,-8.28046894073486,-8.26708984375,-8.26523399353027,-8.29150390625,-8.29130840301514,-8.1904296875,-8.21337890625,-8.2392578125,-8.2724609375],[-8.388671875,-8.40175724029541,-8.37148475646973,-8.33027362823486,-8.30908203125,-8.26845741271973,-8.19921875,-8.17343807220459,-8.17363262176514,-8.33749961853027,-8.388671875],[-8.36728572845459,-8.36796855926514,-8.34873104095459,-8.30703163146973,-8.2392578125,-8.18339824676514,-8.15195274353027,-8.14824104309082,-8.18925762176514,-8.36728572845459],[-8.23886680603027,-8.24003887176514,-8.19990253448486,-8.15781211853027,-8.14023399353027,-8.13916015625,-8.17714881896973,-8.19707012176514,-8.22343730926514,-8.23886680603027],[-8.14082050323486,-8.20175743103027,-8.16806602478027,-8.15957069396973,-8.166015625,-8.17958927154541,-8.20478534698486,-8.32646465301514,-8.35283184051514,-8.44462871551514,-8.41513633728027,-8.38593769073486,-8.29580116271973,-8.25302791595459,-8.18320369720459,-8.13544845581055,-8.14082050323486],[-8.09882831573486,-8.13906288146973,-8.13486385345459,-8.17070388793945,-8.25537109375,-8.19023418426514,-8.12031269073486,-8.09882831573486],[-8.61171913146973,-8.64726638793945,-8.68095684051514,-8.72548770904541,-8.73466777801514,-8.73828125,-8.73027324676514,-8.74472618103027,-8.8603515625,-8.87040996551514,-8.89873027801514,-8.85380840301514,-8.82060527801514,-8.81220722198486,-8.81484413146973,-8.91689491271973,-8.8955078125,-8.90449237823486,-8.92597675323486,-8.93544864654541,-8.92832088470459,-8.84882831573486,-8.80185604095459,-8.8203125,-8.77695274353027,-8.81015586853027,-8.85761737823486,-8.8076171875,-8.76357364654541,-8.69765567779541,-8.62294960021973,-8.57050800323486,-8.52285194396973,-8.47314453125,-8.41982460021973,-8.44511699676514,-8.43554782867432,-8.37753868103027,-8.28984451293945,-8.2578125,-8.24892520904541,-8.26611328125,-8.25986289978027,-8.24042987823486,-8.30781269073486,-8.32148456573486,-8.32666015625,-8.36552810668945,-8.42353439331055,-8.47793006896973,-8.55087852478027,-8.57783222198486,-8.58515548706055,-8.57529354095459,-8.52617168426514,-8.505859375,-8.50664138793945,-8.49394512176514,-8.48212909698486,-8.45517539978027,-8.47187423706055,-8.49668025970459,-8.62490272521973,-8.62831974029541,-8.60078048706055,-8.56640625,-8.51357364654541,-8.46962928771973,-8.43154335021973,-8.40244197845459,-8.35312461853027,-8.30439472198486,-8.22187519073486,-8.18593692779541,-8.12656211853027,-8.09326171875,-8.10556602478027,-8.15195274353027,-8.3291015625,-8.35410213470459,-8.38095760345459,-8.41630840301514,-8.48115158081055,-8.56220626831055,-8.595703125,-8.61171913146973],[-8.31875038146973,-8.34990310668945,-8.27080059051514,-8.09130859375,-8.12451171875,-8.13076114654541,-8.21201133728027,-8.24570274353027,-8.31875038146973],[-8.31777381896973,-8.35722637176514,-8.353515625,-8.29326152801514,-8.271484375,-8.27041053771973,-8.28066444396973,-8.32343769073486,-8.39345741271973,-8.41494083404541,-8.33115196228027,-8.30585956573486,-8.29306602478027,-8.29765605926514,-8.33769512176514,-8.45673847198486,-8.56093788146973,-8.59980487823486,-8.62822246551514,-8.66816425323486,-8.7099609375,-8.73046779632568,-8.74960899353027,-8.7412109375,-8.71308612823486,-8.70273399353027,-8.71210956573486,-8.73544883728027,-8.77363204956055,-8.79082012176514,-8.80888748168945,-8.83339786529541,-8.83828067779541,-8.80527305603027,-8.81191349029541,-8.8564453125,-8.85546875,-8.81337928771973,-8.70371055603027,-8.67460918426514,-8.80781269073486,-8.84052753448486,-8.85595798492432,-8.85058689117432,-8.93144607543945,-8.92011737823486,-8.919921875,-9.00751972198486,-9.03193378448486,-9.03369140625,-9.02617168426514,-9.03408241271973,-9.06923866271973,-9.09902381896973,-9.07636642456055,-9.04619121551514,-9.00634765625,-8.95546913146973,-8.89433670043945,-8.81093692779541,-8.66464805603027,-8.59794902801514,-8.53242206573486,-8.50830078125,-8.50341796875,-8.44443321228027,-8.3671875,-8.37451076507568,-8.42851543426514,-8.43496131896973,-8.42636680603027,-8.45957088470459,-8.53554725646973,-8.56328105926514,-8.58261775970459,-8.71113300323486,-8.70439434051514,-8.72802734375,-8.65029239654541,-8.65214920043945,-8.59189510345459,-8.52753925323486,-8.47519493103027,-8.46425819396973,-8.46738338470459,-8.45888614654541,-8.34208965301514,-8.27900409698486,-8.20458984375,-8.14951229095459,-8.10087966918945,-8.08906173706055,-8.12226581573486,-8.14999961853027,-8.26728534698486,-8.31777381896973],[-8.15517616271973,-8.20830059051514,-8.36357402801514,-8.40712833404541,-8.4482421875,-8.51416015625,-8.61572265625,-8.66367149353027,-8.75751876831055,-8.79755878448486,-8.81953144073486,-8.83544921875,-8.84902286529541,-8.82939529418945,-8.76894474029541,-8.69687461853027,-8.62949275970459,-8.57304668426514,-8.49638652801514,-8.42851543426514,-8.39394474029541,-8.37832069396973,-8.34541034698486,-8.26083946228027,-8.21474647521973,-8.16630840301514,-8.11943340301514,-8.11660194396973,-8.12773513793945,-8.1826171875,-8.18710899353027,-8.17441368103027,-8.06572246551514,-8.06748008728027,-8.11542987823486,-8.15517616271973],[-7.95458936691284,-8.04648494720459,-8.04072284698486,-7.91738271713257,-7.83134698867798,-7.80341863632202,-7.79482460021973,-7.81972646713257,-7.88935518264771,-7.95458936691284],[-7.66787099838257,-7.71650362014771,-7.73789072036743,-7.75351572036743,-7.8076171875,-7.86992168426514,-7.95039033889771,-7.91767549514771,-7.91230440139771,-7.88398504257202,-7.88583993911743,-7.91093778610229,-7.97929668426514,-7.98457050323486,-7.88066434860229,-7.81669902801514,-7.66337823867798,-7.69736289978027,-7.70673799514771,-7.6767578125,-7.60781288146973,-7.57177686691284,-7.66220712661743,-7.66787099838257],[-7.623046875,-7.64648485183716,-7.57246065139771,-7.51279306411743,-7.53105449676514,-7.57851600646973,-7.59687471389771,-7.623046875],[-8.27363300323486,-8.40517520904541,-8.38193416595459,-8.37968730926514,-8.41171836853027,-8.38613319396973,-8.26220703125,-7.93222665786743,-7.64160203933716,-7.56621026992798,-7.49531221389771,-7.4384765625,-7.37959003448486,-7.39042949676514,-7.41464853286743,-7.51162099838257,-7.58798837661743,-7.69609403610229,-7.88212871551514,-8.05908203125,-8.19833946228027,-8.27363300323486],[-7.99951171875,-8.01083946228027,-7.99785089492798,-7.99736356735229,-7.92187452316284,-7.86503887176514,-7.78173780441284,-7.68486356735229,-7.671875,-7.61669921875,-7.47050809860229,-7.43808603286743,-7.42539072036743,-7.34013605117798,-7.31533241271973,-7.25068378448486,-7.22060537338257,-7.16513633728027,-7.1357421875,-7.11279344558716,-7.14023447036743,-7.19707012176514,-7.26689434051514,-7.43886756896973,-7.62617158889771,-7.68222713470459,-7.73066425323486,-7.77665948867798,-7.86914014816284,-7.94804716110229,-7.98144578933716,-7.99951171875],[-7.20205116271973,-7.25136661529541,-7.22499990463257,-7.16747999191284,-7.1591796875,-7.14394521713257,-7.11679697036743,-7.1044921875,-7.20205116271973],[-7.13349580764771,-7.17314481735229,-7.16328144073486,-7.09755849838257,-7.08037137985229,-7.07343721389771,-7.08066415786743,-7.13349580764771],[-7.18330049514771,-7.20859336853027,-7.15634775161743,-7.13457012176514,-7.08320331573486,-7.06874990463257,-7.09111309051514,-7.11337852478027,-7.13798809051514,-7.18330049514771],[-7.11894512176514,-7.12470722198486,-7.11581993103027,-7.01826143264771,-7.06015586853027,-7.06308603286743,-7.11894512176514],[-7.10537099838257,-7.11992216110229,-7.11171913146973,-7.19335985183716,-7.21845722198486,-7.21835994720459,-7.20732402801514,-7.20761680603027,-7.22412109375,-7.21181678771973,-7.1396484375,-7.07275390625,-7.00126981735229,-6.89990186691284,-6.87998056411743,-6.873046875,-6.96015596389771,-6.98935556411743,-7.04902362823486,-7.10537099838257],[-6.97080087661743,-6.98779296875,-6.95253896713257,-6.90517568588257,-6.86123037338257,-6.83847618103027,-6.83974647521973,-6.87021493911743,-6.90185546875,-6.93867158889771,-6.94062519073486,-6.97080087661743],[-6.74980497360229,-6.76533222198486,-6.73291015625,-6.71279335021973,-6.65771436691284,-6.62568330764771,-6.62333965301514,-6.66865205764771,-6.74980497360229],[-6.64042949676514,-6.66249990463257,-6.64306640625,-6.61494159698486,-6.54560565948486,-6.52910184860229,-6.52392625808716,-6.56142568588257,-6.64042949676514],[-6.54941415786743,-6.55888700485229,-6.47724580764771,-6.4560546875,-6.50585985183716,-6.54941415786743],[-6.43417930603027,-6.44238233566284,-6.39306640625,-6.349609375,-6.32460975646973,-6.32353496551514,-6.43417930603027],[-6.44228506088257,-6.5126953125,-6.59140634536743,-6.6796875,-6.81484365463257,-6.84873056411743,-6.90878915786743,-6.83378887176514,-6.76933622360229,-6.47158145904541,-6.4814453125,-6.47929668426514,-6.45976591110229,-6.42646503448486,-6.25537109375,-6.19082021713257,-6.17626953125,-6.22636699676514,-6.31611347198486,-6.38671875,-6.44228506088257],[-6.00761699676514,-6.12177753448486,-6.18271493911743,-6.2158203125,-6.21894550323486,-6.23330068588257,-6.27695322036743,-6.29667949676514,-6.2890625,-6.2666015625,-6.26503944396973,-6.28603506088257,-6.38281202316284,-6.47119140625,-6.51621150970459,-6.72919940948486,-6.79052734375,-6.80830097198486,-6.80839824676514,-6.81728506088257,-6.86699199676514,-6.86015605926514,-6.81015634536743,-6.84257841110229,-6.90244150161743,-6.91582012176514,-6.89873075485229,-6.89511728286743,-6.91240262985229,-6.93837881088257,-6.94775390625,-6.94726610183716,-6.89726543426514,-6.80566453933716,-6.69013643264771,-6.56982421875,-6.51806640625,-6.47236347198486,-6.44267606735229,-6.42421913146973,-6.43564462661743,-6.46474599838257,-6.66904306411743,-6.68671894073486,-6.69951200485229,-6.69287109375,-6.65185546875,-6.64824247360229,-6.69872999191284,-6.74169921875,-6.7734375,-6.80595684051514,-6.89335966110229,-6.90507841110229,-6.89443397521973,-6.90302753448486,-6.92646503448486,-7.05058622360229,-7.17802762985229,-7.22128915786743,-7.26503896713257,-7.30449199676514,-7.43164110183716,-7.55244159698486,-7.65771532058716,-7.71816396713257,-7.72382783889771,-7.70302772521973,-7.67724609375,-7.63212919235229,-7.63300800323486,-7.77109384536743,-7.79247999191284,-7.89560556411743,-8.00459003448486,-8.26328086853027,-8.33427715301514,-8.40517520904541,-8.55927753448486,-8.60380840301514,-8.68476581573486,-8.72724628448486,-8.76962947845459,-8.74052715301514,-8.70537185668945,-8.64736366271973,-8.61464881896973,-8.62646484375,-8.568359375,-8.47802734375,-8.28671932220459,-8.28828144073486,-8.31269550323486,-8.36142539978027,-8.39609336853027,-8.4091796875,-8.39960956573486,-8.35361289978027,-8.32392597198486,-8.30507755279541,-8.26171779632568,-8.23955059051514,-8.20195293426514,-8.14941501617432,-7.89052724838257,-7.82841777801514,-7.70488262176514,-7.69492197036743,-7.7041015625,-7.66787099838257,-7.66708946228027,-7.70722627639771,-7.73603439331055,-7.79697275161743,-7.79404306411743,-7.78232431411743,-7.72412109375,-7.68837928771973,-7.63554716110229,-7.56669855117798,-7.54189395904541,-7.4716796875,-7.44746065139771,-7.41552782058716,-7.39423799514771,-7.36865234375,-7.31171941757202,-7.23935604095459,-7.17675733566284,-7.11386680603027,-7.0537109375,-6.92783212661743,-6.85898447036743,-6.84580039978027,-6.84609413146973,-6.86035108566284,-6.85371065139771,-6.83320283889771,-6.826171875,-6.84101581573486,-6.83525419235229,-6.77802705764771,-6.72939443588257,-6.67412090301514,-6.66435575485229,-6.75078153610229,-6.76796865463257,-6.78691434860229,-6.78154277801514,-6.67099618911743,-6.61669921875,-6.46953105926514,-6.49794912338257,-6.48037099838257,-6.45693349838257,-6.11640644073486,-6.01699209213257,-5.93427753448486,-5.91416025161743,-5.96474599838257,-5.98408222198486,-6.017578125,-6.02187490463257,-6.03837871551514,-6.09824180603027,-6.09199237823486,-6.07343769073486,-6.00849628448486,-5.90419960021973,-5.95712900161743,-5.97812461853027,-6.00761699676514],[-5.97373104095459,-6.02158164978027,-5.96669960021973,-5.93935585021973,-5.88017606735229,-5.90214824676514,-5.97373104095459],[-6.29843711853027,-6.46484375,-6.40615224838257,-6.25400400161743,-6.18017578125,-6.09492158889771,-5.87626981735229,-5.77529287338257,-5.90380859375,-5.96923828125,-6.29843711853027],[-5.81103515625,-5.84648418426514,-5.84365224838257,-5.80361366271973,-5.73085975646973,-5.72617197036743,-5.75273466110229,-5.81103515625],[-5.85078144073486,-5.94707059860229,-5.9130859375,-5.91259765625,-5.85605478286743,-5.73886728286743,-5.60703182220459,-5.60898447036743,-5.64834022521973,-5.66171836853027,-5.78886699676514,-5.85078144073486],[-5.70703172683716,-5.74560546875,-5.81679677963257,-5.88271474838257,-5.94970750808716,-6.05009746551514,-6.09999990463257,-6.17060565948486,-6.20234346389771,-6.29511737823486,-6.328125,-6.33730459213257,-6.36533212661743,-6.33486318588257,-6.27050828933716,-6.20078134536743,-6.17167949676514,-6.09033250808716,-6.06289052963257,-6.01953125,-6.00849628448486,-6.009765625,-5.970703125,-5.8330078125,-5.74443387985229,-5.70722627639771,-5.68193340301514,-5.712890625,-5.55751895904541,-5.52509784698486,-5.43847608566284,-5.42734384536743,-5.43857431411743,-5.49238252639771,-5.53925800323486,-5.58134746551514,-5.60302734375,-5.70703172683716],[-5.90205097198486,-5.98798847198486,-5.78525400161743,-5.6826171875,-5.62138700485229,-5.31064462661743,-5.31787109375,-5.34814453125,-5.57597637176514,-5.73583984375,-5.90205097198486],[-5.47871065139771,-5.48349618911743,-5.36054658889771,-5.32255840301514,-5.28623056411743,-5.28886747360229,-5.36640644073486,-5.40478563308716,-5.47871065139771],[-5.27158164978027,-5.373046875,-5.36738252639771,-5.33183574676514,-5.29833984375,-5.27109336853027,-5.24980497360229,-5.27158164978027],[-5.43798828125,-5.46474599838257,-5.35029315948486,-5.25615215301514,-5.20292949676514,-5.15625,-5.14462852478027,-5.09599590301514,-5.072265625,-5.07558631896973,-5.14082050323486,-5.15878915786743,-5.22128963470459,-5.43798828125],[-5.26943397521973,-5.33583974838257,-5.38749980926514,-5.39121103286743,-5.38066387176514,-5.33544969558716,-5.38310527801514,-5.38095712661743,-5.31953096389771,-5.1376953125,-5.06982421875,-4.99853515625,-4.84658193588257,-4.76718759536743,-4.70712900161743,-4.63388681411743,-4.61865234375,-4.67499971389771,-4.93388652801514,-5.13867139816284,-5.26943397521973],[-4.55117130279541,-4.76621055603027,-4.82265615463257,-4.73994112014771,-4.72343778610229,-4.73994112014771,-4.74902391433716,-4.74824190139771,-4.83173799514771,-4.91025400161743,-4.96308612823486,-5.13847637176514,-5.18574237823486,-5.16240262985229,-5.15644550323486,-5.22402381896973,-5.27333974838257,-5.33300828933716,-5.39316368103027,-5.41933631896973,-5.39355516433716,-5.40576171875,-5.43671894073486,-5.47744178771973,-5.51933574676514,-5.63798809051514,-5.67128896713257,-5.63496065139771,-5.66621112823486,-5.66337919235229,-5.54462909698486,-5.48886680603027,-5.42626953125,-5.38115262985229,-5.33085918426514,-5.26191425323486,-5.21015644073486,-5.17724609375,-5.05244159698486,-5.00009822845459,-4.94443368911743,-4.83125019073486,-4.61835956573486,-4.44267559051514,-4.39472675323486,-4.38691377639771,-4.43359422683716,-4.55117130279541],[-4.24589872360229,-4.29931688308716,-4.25742197036743,-4.16962862014771,-4.11103534698486,-4.19980478286743,-4.24589872360229],[-4.11298847198486,-4.23330068588257,-4.22714853286743,-4.14804744720459,-4.06132841110229,-4.02998065948486,-3.98095703125,-3.99755859375,-4.04091787338257,-4.11298847198486],[-3.58544898033142,-3.63789057731628,-3.54042935371399,-3.51474595069885,-3.54130840301514,-3.58544898033142],[-3.67460942268372,-3.71113300323486,-3.73525404930115,-3.73271465301514,-3.69765663146973,-3.67714834213257,-3.68642592430115,-3.71455049514771,-3.77099609375,-3.74306607246399,-3.69931626319885,-3.67939472198486,-3.60087871551514,-3.58750009536743,-3.51230454444885,-3.51591777801514,-3.56367182731628,-3.59765648841858,-3.63320350646973,-3.67460942268372],[-3.54687523841858,-3.58916020393372,-3.60253930091858,-3.58779287338257,-3.58857464790344,-3.51220726966858,-3.49482417106628,-3.51669931411743,-3.52451133728027,-3.54687523841858],[-3.46445298194885,-3.63671875,-3.53906226158142,-3.42333960533142,-3.39990234375,-3.46445298194885],[-3.31513667106628,-3.35917997360229,-3.33134794235229,-3.28818345069885,-3.26152324676514,-3.25429725646973,-3.31513667106628],[-3.8681640625,-4.05410146713257,-4.00693368911743,-3.81748032569885,-3.69990229606628,-3.61240220069885,-3.45791029930115,-3.33955097198486,-3.26035141944885,-3.25107431411743,-3.39482426643372,-3.44882822036743,-3.49501943588257,-3.53476572036743,-3.71855425834656,-3.76298856735229,-3.82089877128601,-3.8681640625],[-3.08789086341858,-3.166015625,-3.21699237823486,-3.27753925323486,-3.31084012985229,-3.33808588981628,-3.39101576805115,-3.47109341621399,-3.63300800323486,-3.64726543426514,-3.6708984375,-3.76455092430115,-3.78291010856628,-3.78916025161743,-3.81367206573486,-3.82363247871399,-3.77167963981628,-3.71064424514771,-3.60517573356628,-3.57939457893372,-3.52275371551514,-3.42099618911743,-3.35585951805115,-3.17050790786743,-3.12812495231628,-3.10546851158142,-3.14814472198486,-3.10322260856628,-3.06523418426514,-3.06914067268372,-3.08789086341858],[-2.93652319908142,-3.02001929283142,-3.01630878448486,-3.00439476966858,-2.95234370231628,-2.93652319908142],[-3.00527358055115,-3.02529263496399,-3.01445293426514,-2.98681640625,-2.96044945716858,-2.93281221389771,-2.89892578125,-2.93398451805115,-3.00527358055115],[-2.89951205253601,-2.92529273033142,-2.90058565139771,-2.87294888496399,-2.83808565139771,-2.83466815948486,-2.84501957893372,-2.89951205253601],[-2.86582040786743,-2.97666001319885,-2.99296879768372,-2.97851538658142,-2.98935556411743,-3.13085913658142,-3.22802758216858,-3.31191372871399,-3.39150404930115,-3.41132807731628,-3.41874980926514,-3.53330063819885,-3.5703125,-3.85771512985229,-3.74882793426514,-3.62519502639771,-3.57929730415344,-3.47470712661743,-3.43886685371399,-3.39160132408142,-3.32714819908142,-3.31718730926514,-3.31884765625,-3.32851576805115,-3.36318397521973,-3.43369126319885,-3.45322275161743,-3.40869140625,-3.39267563819885,-3.34921836853027,-3.32607436180115,-3.30419945716858,-3.27167963981628,-3.24111342430115,-3.22929668426514,-3.23496103286743,-3.265625,-3.30048847198486,-3.39658188819885,-3.43339872360229,-3.44912099838257,-3.43984389305115,-3.416015625,-3.24052739143372,-3.20263695716858,-3.17167949676514,-3.15742206573486,-3.18408226966858,-3.23857402801514,-3.30332040786743,-3.34052729606628,-3.44433569908142,-3.50605487823486,-3.49628925323486,-3.39726567268372,-3.34140634536743,-3.28232407569885,-3.22207069396973,-2.93457007408142,-2.86591768264771,-2.84218740463257,-2.85664033889771,-2.84960961341858,-2.82851552963257,-2.83847665786743,-2.89511752128601,-2.93701148033142,-2.93349647521973,-2.88906216621399,-2.82050776481628,-2.79072260856628,-2.78574204444885,-2.79033207893372,-2.80615258216858,-2.86582040786743],[-3.18290996551514,-3.32851576805115,-3.22919917106628,-3.15878915786743,-3.11308598518372,-3.05693387985229,-2.98681640625,-2.82021450996399,-2.78554654121399,-2.78320336341858,-3.00195336341858,-3.03896450996399,-3.11689448356628,-3.14130878448486,-3.18290996551514],[-2.99765634536743,-3.10302710533142,-3.14277315139771,-3.19492220878601,-3.22685551643372,-3.22177720069885,-3.16660141944885,-3.12929654121399,-3.08632826805115,-3.09667992591858,-3.16074228286743,-3.20556664466858,-3.20937514305115,-3.12480473518372,-3.05839824676514,-2.97656273841858,-2.94072246551514,-2.92011713981628,-2.86308622360229,-2.79970741271973,-2.73154306411743,-2.56630825996399,-2.53027319908142,-2.55966782569885,-2.59697246551514,-2.69697260856628,-2.82998061180115,-2.99765634536743],[-2.74101567268372,-2.82138681411743,-2.81972646713257,-2.76982402801514,-2.64931607246399,-2.60976552963257,-2.59414076805115,-2.525390625,-2.51025414466858,-2.6796875,-2.74101567268372],[-2.12353515625,-2.13876962661743,-2.09521484375,-2.03691411018372,-2.02763676643372,-2.03076195716858,-2.0517578125,-2.12353515625],[-2.34306621551514,-2.36972641944885,-2.28173851966858,-2.25751972198486,-2.16152334213257,-2.11542963981628,-2.05117177963257,-2.02578115463257,-2.01660180091858,-2.06337928771973,-2.17773485183716,-2.28437542915344,-2.34306621551514],[-2.45126938819885,-2.46943378448486,-2.41542959213257,-2.26279306411743,-2.22216820716858,-2.0771484375,-2.03593730926514,-1.97480452060699,-1.97578144073486,-2.01181650161743,-2.16806650161743,-2.36581993103027,-2.45126938819885],[-1.95546889305115,-1.99570310115814,-1.99433600902557,-1.91865217685699,-1.87832021713257,-1.95546889305115],[-1.78974592685699,-1.82285153865814,-1.85888659954071,-1.91660165786743,-1.90615212917328,-1.94003903865814,-1.93808579444885,-1.88222658634186,-1.84306657314301,-1.80898416042328,-1.80087900161743,-1.81376957893372,-1.78974592685699],[-1.81650376319885,-1.89892601966858,-1.85488259792328,-1.81484377384186,-1.78671848773956,-1.77265644073486,-1.81650376319885],[-1.69052743911743,-1.74980449676514,-1.80458950996399,-1.88984394073486,-1.90263688564301,-1.94160163402557,-1.97128880023956,-1.99189472198486,-1.98183572292328,-2.00937509536743,-2.04775404930115,-2.06386733055115,-2.02832055091858,-1.98642587661743,-1.89443337917328,-1.86689472198486,-1.75888657569885,-1.73046886920929,-1.73222672939301,-1.69199192523956,-1.69052743911743],[-1.70546841621399,-1.74101531505585,-1.74082052707672,-1.69931662082672,-1.69257807731628,-1.71289050579071,-1.78027319908142,-1.77089858055115,-1.79394507408142,-1.81005835533142,-1.87714838981628,-1.88896501064301,-1.94306647777557,-1.89443337917328,-1.97822272777557,-2.00693368911743,-2.00517582893372,-1.85888659954071,-1.68749988079071,-1.65927720069885,-1.64433610439301,-1.63593780994415,-1.70546841621399],[-1.59179699420929,-1.64199221134186,-1.63554692268372,-1.65498054027557,-1.72158169746399,-1.73398458957672,-1.75380837917328,-1.79970681667328,-1.83769500255585,-1.87304723262787,-1.89042949676514,-1.87246096134186,-1.89365243911743,-1.85917985439301,-1.82412111759186,-1.75214850902557,-1.66835904121399,-1.61621081829071,-1.59179699420929],[-1.61962914466858,-1.66162109375,-1.56777358055115,-1.53984355926514,-1.56406247615814,-1.61962914466858],[-1.66943347454071,-1.73828125,-1.80019557476044,-1.86699211597443,-2.18867182731628,-2.46484398841858,-2.57333970069885,-2.61796832084656,-2.65800809860229,-2.70400381088257,-2.89550757408142,-2.93613243103027,-3.00117182731628,-3.07177758216858,-3.07138657569885,-3.05556631088257,-3.02900385856628,-2.99423837661743,-2.96660137176514,-2.94873070716858,-2.89404320716858,-2.85537123680115,-2.82490253448486,-2.74355435371399,-2.64326167106628,-2.49345684051514,-2.45195341110229,-2.41542959213257,-2.30742192268372,-2.18134760856628,-2.13261699676514,-2.10312509536743,-2.07900381088257,-2.12509775161743,-2.1142578125,-2.07939457893372,-2.04257798194885,-1.97265636920929,-1.91689455509186,-1.86054694652557,-1.81318342685699,-1.75078141689301,-1.70507824420929,-1.65732395648956,-1.61103534698486,-1.57470691204071,-1.52675771713257,-1.61044907569885,-1.68037116527557,-1.73105442523956,-1.65869164466858,-1.53388655185699,-1.50605452060699,-1.50498020648956,-1.53916001319885,-1.59316408634186,-1.66943347454071],[-1.70429670810699,-1.71083998680115,-1.68144547939301,-1.53486335277557,-1.50283217430115,-1.50820314884186,-1.55185556411743,-1.59091818332672,-1.62744128704071,-1.70429670810699],[-1.66054689884186,-1.701171875,-1.71240234375,-1.68515598773956,-1.69082021713257,-1.728515625,-1.69667983055115,-1.64482426643372,-1.58984386920929,-1.45371079444885,-1.35078120231628,-1.33242201805115,-1.36025393009186,-1.43906259536743,-1.53164041042328,-1.60371112823486,-1.66054689884186],[-1.17128908634186,-1.23369157314301,-1.28769516944885,-1.38935554027557,-1.25400400161743,-1.23681652545929,-1.25927758216858,-1.28603529930115,-1.33740210533142,-1.44736301898956,-1.49882829189301,-1.50712883472443,-1.44306635856628,-1.43720674514771,-1.57695293426514,-1.59833967685699,-1.61601543426514,-1.57744157314301,-1.55605483055115,-1.49277341365814,-1.30449199676514,-1.33984339237213,-1.51064443588257,-1.58720719814301,-1.54824197292328,-1.43212866783142,-1.28300786018372,-1.18222677707672,-1.18916010856628,-1.15751969814301,-1.17128908634186],[-1.18066442012787,-1.28281235694885,-1.27753889560699,-1.24121105670929,-1.04414045810699,-0.9853515625,-0.979101777076721,-1.00732398033142,-1.03935575485229,-1.10517609119415,-1.14501953125,-1.18066442012787],[-1.11601579189301,-1.13427710533142,-1.11415994167328,-1.03759789466858,-0.978808462619781,-0.938476741313934,-0.978906452655792,-1.03056657314301,-1.03408217430115,-1.11601579189301],[-1.77792978286743,-1.78349614143372,-1.66367173194885,-1.60996091365814,-1.53828108310699,-1.26132810115814,-1.19785141944885,-0.970507979393005,-0.915624976158142,-0.954003870487213,-1.05624997615814,-1.24072301387787,-1.34013676643372,-1.38417959213257,-1.41845726966858,-1.44238293170929,-1.55927717685699,-1.62773418426514,-1.73847651481628,-1.77792978286743],[-1.31552708148956,-1.34345710277557,-1.31728506088257,-1.25546860694885,-1.17255878448486,-1.10439455509186,-1.05019557476044,-0.983984589576721,-0.959765613079071,-0.890038967132568,-0.915332138538361,-0.917578160762787,-0.968262016773224,-1.18818390369415,-1.31552708148956],[-0.777441263198853,-0.828417897224426,-0.862890481948853,-0.923925995826721,-0.887402176856995,-0.851171791553497,-0.832519710063934,-0.837304472923279,-0.826269328594208,-0.821874737739563,-0.826660096645355,-0.811621069908142,-0.822460830211639,-0.813867092132568,-0.765039026737213,-0.777441263198853],[-0.65136706829071,-0.690429627895355,-0.688281059265137,-0.732519626617432,-0.711621046066284,-0.725781142711639,-0.877734124660492,-0.978320360183716,-1.06464838981628,-1.09404289722443,-1.17314422130585,-1.21474611759186,-1.21679675579071,-1.16972661018372,-1.17841780185699,-1.11943352222443,-1.02832067012787,-0.823046982288361,-0.881933271884918,-0.787304759025574,-0.785058736801147,-0.801074028015137,-0.768847525119781,-0.70488303899765,-0.65136706829071],[-0.780956923961639,-0.801562249660492,-0.775292694568634,-0.760937213897705,-0.619335889816284,-0.623437702655792,-0.758398413658142,-0.780956923961639],[-0.528711140155792,-0.529980182647705,-0.486523658037186,-0.448730260133743,-0.440917938947678,-0.418261498212814,-0.417285144329071,-0.436523377895355,-0.469140708446503,-0.528711140155792],[-0.406836122274399,-0.451269596815109,-0.502636969089508,-0.489843994379044,-0.490820288658142,-0.494726717472076,-0.525000095367432,-0.526171803474426,-0.478808790445328,-0.407031506299973,-0.417675912380219,-0.406836122274399],[-2.60976552963257,-2.80341792106628,-3.00664067268372,-3.20996117591858,-3.41328144073486,-3.61660146713257,-3.81982421875,-4.02314472198486,-4.22646522521973,-4.42978525161743,-4.63300800323486,-4.83632802963257,-5.03964853286743,-5.24296903610229,-5.44619131088257,-5.64951133728027,-5.85283231735229,-6.05615234375,-6.25937509536743,-6.34609365463257,-6.45224618911743,-6.61152315139771,-6.74003887176514,-6.84003877639771,-6.90537118911743,-7.07255840301514,-7.27587842941284,-7.47919893264771,-7.68251895904541,-7.88574266433716,-8.08906173706055,-8.29238224029541,-8.49570274353027,-8.69892597198486,-8.90224647521973,-9.10556602478027,-9.11874961853027,-9.08505916595459,-8.97373008728027,-8.84677696228027,-8.72832012176514,-8.62040996551514,-8.30058574676514,-8.19550800323486,-8.16650390625,-8.13935565948486,-8.083984375,-7.92373085021973,-8.02275371551514,-8.10117244720459,-8.10634803771973,-8.12539005279541,-8.17275333404541,-8.18906211853027,-8.16582012176514,-8.10693359375,-8.04658222198486,-7.98242139816284,-8.08613300323486,-8.14287090301514,-8.26240253448486,-8.23779296875,-8.19228553771973,-8.14511680603027,-8.0947265625,-8.04121112823486,-7.9130859375,-7.83759784698486,-7.69140625,-7.63925743103027,-7.58720731735229,-7.52832078933716,-7.50820302963257,-7.47246074676514,-7.37324237823486,-7.33964872360229,-7.29892635345459,-7.25146484375,-7.21572256088257,-7.20136737823486,-7.20361375808716,-7.22587919235229,-7.22714853286743,-7.1904296875,-7.20175838470459,-7.20058584213257,-7.13632822036743,-7.06982421875,-6.9365234375,-6.91074180603027,-6.88652372360229,-6.8583984375,-6.79042959213257,-6.73115205764771,-6.62568330764771,-6.56044912338257,-6.45380878448486,-6.34335994720459,-6.11855411529541,-5.94902324676514,-5.88750028610229,-5.84365224838257,-5.83857440948486,-5.80703163146973,-5.72441387176514,-5.67568397521973,-5.68818378448486,-5.71201181411743,-5.71650362014771,-5.70917987823486,-5.67597627639771,-5.62890672683716,-5.54580068588257,-5.46523380279541,-5.42763662338257,-5.37011766433716,-5.34882783889771,-5.35048866271973,-5.31201124191284,-5.25615215301514,-5.01435518264771,-4.94540977478027,-4.97568368911743,-4.99042987823486,-4.95078086853027,-4.92441415786743,-4.92871046066284,-4.90732431411743,-4.89511728286743,-4.89316368103027,-4.81874990463257,-4.70126962661743,-4.65068340301514,-4.58476591110229,-4.53085947036743,-4.47841787338257,-4.44306659698486,-4.44179677963257,-4.453125,-4.45068359375,-4.19540977478027,-4.07910108566284,-4.01113271713257,-3.95478487014771,-3.93847608566284,-3.92216777801514,-3.92988252639771,-3.97607421875,-3.97929692268372,-3.94863271713257,-3.90996098518372,-3.94580101966858,-3.88701152801514,-3.82509756088257,-3.79677748680115,-3.79970717430115,-3.82197284698486,-3.81796908378601,-3.77558588981628,-3.72011709213257,-3.68037104606628,-3.65000009536743,-3.57792949676514,-3.47949194908142,-3.30917978286743,-3.24814462661743,-3.14892554283142,-3.05478525161743,-3.04433608055115,-3.08749985694885,-3.1318359375,-3.185546875,-3.36435556411743,-3.41611313819885,-3.41191387176514,-3.51640629768372,-3.61552739143372,-3.73212885856628,-3.78515625,-3.84257817268372,-3.89902377128601,-4.06230449676514,-4.07011747360229,-4.06904315948486,-4.09492206573486,-4.05693387985229,-4.00742244720459,-3.94892597198486,-3.828125,-3.70361328125,-3.55097651481628,-3.41298818588257,-3.29462885856628,-3.13066434860229,-2.97509765625,-2.94345712661743,-2.92958998680115,-2.91455101966858,-2.85605478286743,-2.78857421875,-2.75957036018372,-2.68037104606628,-2.68417954444885,-2.72714853286743,-2.76621103286743,-2.78906226158142,-2.65820336341858,-2.48740220069885,-2.45029306411743,-2.43779277801514,-2.45429682731628,-2.51396465301514,-2.54169940948486,-2.54716825485229,-2.60058617591858,-2.62460970878601,-2.54404306411743,-2.45068383216858,-2.42167949676514,-2.41503882408142,-2.39091801643372,-2.30449247360229,-2.29365277290344,-2.21962881088257,-2.18359375,-2.14746069908142,-2.10205101966858,-2.18915987014771,-2.22558569908142,-2.21572303771973,-2.21445322036743,-2.27255868911743,-2.27021455764771,-2.24667978286743,-2.21845698356628,-2.24042940139771,-2.24228525161743,-2.17578148841858,-2.09238266944885,-2.033203125,-1.99033224582672,-1.93251967430115,-1.71494114398956,-1.55966794490814,-1.55654299259186,-1.54121112823486,-1.39345693588257,-1.42968726158142,-1.44833993911743,-1.45527327060699,-1.44765639305115,-1.42470717430115,-1.38398420810699,-1.28408205509186,-1.24726557731628,-1.21884739398956,-1.16582012176514,-1.00693380832672,-0.952636897563934,-0.89716762304306,-0.855468988418579,-0.833594024181366,-0.781835854053497,-0.703808724880219,-0.657128691673279,-0.582421898841858,-0.537011563777924,-0.491113066673279,-0.454101711511612,-0.35546875,-0.347461074590683,-0.358886450529099,-0.417382568120956,-0.511816322803497,-0.635741949081421,-0.726171910762787,-0.741406261920929,-0.731445372104645,-0.744335770606995,-0.769726395606995,-0.846777200698853,-0.897363066673279,-1.00185561180115,-1.10244131088257,-1.20312476158142,-1.31054699420929,-1.36298799514771,-1.47412133216858,-1.52910172939301,-1.62080085277557,-1.72099614143372,-1.84453105926514,-1.96875,-2.08291006088257,-2.19521522521973,-2.30908179283142,-2.62099599838257,-2.83232402801514,-2.71425795555115,-2.5830078125,-2.53564476966858,-2.51044893264771,-2.53671860694885,-2.58984375,-2.70585966110229,-2.93359375,-2.94404268264771,-2.90917944908142,-2.97880887985229,-3.10761713981628,-3.20986318588257,-3.24990272521973,-3.33310532569885,-3.34853506088257,-3.36855483055115,-3.37490248680115,-3.34511709213257,-3.26875019073486,-3.18603491783142,-2.99531292915344,-2.90410113334656,-2.76425790786743,-2.73427700996399,-2.58310580253601,-2.52949213981628,-2.42568373680115,-2.32519507408142,-2.27333998680115,-2.22431612014771,-2.19765639305115,-2.10507798194885,-2.02548837661743,-1.97314453125,-1.88125014305115,-1.84091806411743,-1.80214822292328,-1.68564474582672,-1.56582033634186,-1.48320317268372,-1.48378884792328,-1.55654299259186,-1.61591804027557,-1.79111289978027,-1.84550797939301,-1.91777348518372,-1.96787118911743,-1.99208998680115,-2.03886747360229,-2.09921884536743,-2.21181654930115,-2.34824228286743,-2.3564453125,-2.35000014305115,-2.37568354606628,-2.41201186180115,-2.42041039466858,-2.44580078125,-2.47207045555115,-2.50810575485229,-2.60712862014771,-2.60976552963257],[-0.334668099880219,-0.431835979223251,-0.466601371765137,-0.520508050918579,-0.596679985523224,-0.612890660762787,-0.583691596984863,-0.658593773841858,-0.539062738418579,-0.463281065225601,-0.385742098093033,-0.380175828933716,-0.382617205381393,-0.402832299470901,-0.334668099880219],[-0.318945556879044,-0.46835920214653,-0.61015647649765,-0.689453303813934,-0.694433450698853,-0.72412121295929,-0.759863495826721,-0.808691501617432,-0.847754061222076,-0.883691132068634,-0.832030951976776,-0.783984422683716,-0.766015350818634,-0.802441120147705,-0.805956840515137,-0.739062249660492,-0.642968654632568,-0.59960949420929,-0.500293016433716,-0.460254102945328,-0.390918225049973,-0.335839778184891,-0.33164057135582,-0.406347393989563,-0.335937768220901,-0.306640714406967,-0.318945556879044],[-0.495312303304672,-0.521191596984863,-0.520508050918579,-0.413867473602295,-0.278613328933716,-0.255761474370956,-0.284375011920929,-0.318652212619781,-0.391015619039536,-0.495312303304672],[-0.347949266433716,-0.38290998339653,-0.357617348432541,-0.29746076464653,-0.227539300918579,-0.230566248297691,-0.276660114526749,-0.317773282527924,-0.347949266433716],[-0.00410180492326617,-0.0465820394456387,-0.0399414636194706,-0.149804979562759,-0.204296693205833,-0.241113260388374,-0.290332049131393,-0.365722388029099,-0.374120801687241,-0.345996052026749,-0.330078125,-0.360742270946503,-0.337597638368607,-0.26845708489418,-0.226465001702309,-0.0806638598442078,-0.0859373658895493,-0.142969012260437,-0.180566504597664,-0.291406333446503,-0.302148729562759,-0.298339694738388,-0.344433575868607,-0.416015475988388,-0.443847477436066,-0.391601741313934,-0.296581864356995,-0.328613072633743,-0.361816555261612,-0.366406500339508,-0.267382770776749,-0.262304663658142,-0.209668174386024,-0.154687717556953,-0.101465091109276,-0.0728515833616257,-0.0984862819314003,-0.0601072870194912,-0.0699220448732376,-0.0454103425145149,-0.0298339128494263,-0.00410180492326617],[-0.530469000339508,-0.576855659484863,-0.531836032867432,-0.46787104010582,-0.37929692864418,-0.308984100818634,-0.226465001702309,-0.000781514449045062,0.00708003947511315,-0.0175290703773499,-0.167675524950027,-0.257617294788361,-0.379687666893005,-0.530469000339508],[-0.175976544618607,-0.192480593919754,-0.140625044703484,-0.211718782782555,-0.24726565182209,-0.282812535762787,-0.28447237610817,-0.316992342472076,-0.323339551687241,-0.208691284060478,-0.245605230331421,-0.212109550833702,-0.18916030228138,-0.12636698782444,0.0177244488149881,-0.0184572618454695,-0.0628420114517212,-0.0760251954197884,-0.103027552366257,-0.175976544618607],[-0.187011703848839,-0.189843833446503,-0.131445109844208,-0.0664063841104507,0.0454103425145149,-0.139257967472076,-0.187011703848839],[-0.00585963949561119,-0.036621168255806,0.00634779967367649,0.0166014526039362,0.124414347112179,0.14257824420929,0.0689938589930534,-0.00585963949561119],[0.0595217235386372,0.0346679575741291,0.0626953318715096,0.0796388387680054,0.0952146947383881,0.232079982757568,0.223290830850601,0.103076249361038,0.0595217235386372],[0.541943609714508,0.536181926727295,0.549072325229645,0.643115341663361,0.699755668640137,0.698632657527924,0.613964974880219,0.541943609714508],[0.642089784145355,0.631005823612213,0.634863018989563,0.675146520137787,0.743554830551147,0.744385063648224,0.733447253704071,0.683300852775574,0.642089784145355],[0.791308581829071,0.758447170257568,0.769433617591858,0.811913847923279,0.842480540275574,0.862011551856995,0.847460746765137,0.791308581829071],[0.664452850818634,0.65087890625,0.777880728244781,0.851123034954071,0.869580328464508,0.825000166893005,0.778124809265137,0.722704887390137,0.664452850818634],[0.801025390625,0.772217035293579,0.829980373382568,0.87182629108429,0.89135730266571,0.846337735652924,0.801025390625],[0.833984196186066,0.804883122444153,0.896240055561066,0.917480766773224,0.933544814586639,0.943994343280792,0.896728813648224,0.87988269329071,0.833984196186066],[0.746631026268005,0.708105325698853,0.736523449420929,0.779589831829071,0.784375190734863,0.831591546535492,0.856640696525574,0.889502108097076,0.950341641902924,0.986865341663361,1.08876943588257,1.13022458553314,1.05439472198486,0.989208698272705,0.959374904632568,0.892724394798279,0.859277367591858,0.746631026268005],[1.04833972454071,0.993359208106995,1.00683581829071,1.0712890625,1.11723625659943,1.13364279270172,1.07255864143372,1.04833972454071],[0.870165884494781,0.841650366783142,0.848144292831421,0.882031261920929,0.94267600774765,0.996777534484863,1.03095674514771,1.04125964641571,1.05371117591858,1.15883803367615,1.16547858715057,1.06777334213257,1.01474606990814,0.870165884494781],[1.18056643009186,1.13701188564301,1.16557610034943,1.12822270393372,1.09238302707672,0.989550888538361,1.01323235034943,1.04648447036743,1.07138705253601,1.08701145648956,1.13745129108429,1.13725590705872,1.18056643009186],[1.21611320972443,1.14106428623199,1.10458981990814,1.04951202869415,0.961035251617432,0.858984649181366,0.831933557987213,0.852636873722076,0.886767327785492,0.913476347923279,0.932519733905792,0.956494033336639,0.995556533336639,1.05043971538544,1.01611316204071,1.01489281654358,1.07739269733429,1.10263657569885,1.18149399757385,1.19604480266571,1.18022489547729,1.21611320972443],[0.990136981010437,0.959765613079071,1.00703120231628,1.07568359375,1.14716804027557,1.26396489143372,1.34785175323486,1.39707052707672,1.39526402950287,1.34565436840057,1.26079082489014,1.23422837257385,1.15625011920929,1.06728518009186,0.990136981010437],[1.46508800983429,1.18374001979828,1.14589834213257,1.01826179027557,0.973925769329071,0.884228527545929,0.628320395946503,0.564453423023224,0.596093833446503,0.641064703464508,0.833886802196503,0.94140636920929,0.946972668170929,1.05693352222443,1.18735361099243,1.42548799514771,1.42363286018372,1.48164081573486,1.52792942523956,1.53974616527557,1.46508800983429],[1.45917975902557,1.33090817928314,1.36445307731628,1.41547846794128,1.45312523841858,1.46542942523956,1.49858379364014,1.55820286273956,1.58564460277557,1.60795903205872,1.62539076805115,1.51005840301514,1.45917975902557],[0.995312452316284,0.825585782527924,0.743554830551147,0.655273497104645,0.557128727436066,0.470605224370956,0.444970488548279,0.398437738418579,0.38037121295929,0.37456026673317,0.305517733097076,0.29746076464653,0.300341606140137,0.31757789850235,0.326611161231995,0.41552734375,0.485839575529099,0.493505716323853,0.485986262559891,0.481054812669754,0.46801769733429,0.436572045087814,0.450878828763962,0.486132889986038,0.498437762260437,0.494824111461639,0.441699475049973,0.446777582168579,0.514697194099426,0.528320372104645,0.510302603244781,0.449218988418579,0.408251911401749,0.268505781888962,0.166552528738976,0.0397460795938969,-0.0899411961436272,-0.196191638708115,-0.307128876447678,-0.432031363248825,-0.555566251277924,-0.649902582168579,-0.868261992931366,-0.899218857288361,-0.960644543170929,-1.03945314884186,-1.25849604606628,-1.37011742591858,-1.37148451805115,-1.36367154121399,-1.37783229351044,-1.40654325485229,-1.33945322036743,-1.21250021457672,-1.11816370487213,-0.938574194908142,-0.855566382408142,-0.828515887260437,-0.840331971645355,-0.887890338897705,-0.925683796405792,-0.945996224880219,-0.933300733566284,-0.874999940395355,-0.839257657527924,-0.793749928474426,-0.757031381130219,-0.756640613079071,-0.769824385643005,-0.755175530910492,-0.722070038318634,-0.687011897563934,-0.658886551856995,-0.640722632408142,-0.599804878234863,-0.570703208446503,-0.591503858566284,-0.648534953594208,-0.707421839237213,-0.778222799301147,-0.96162086725235,-1.00410163402557,-1.02607405185699,-1.00175774097443,-0.907031178474426,-0.872363209724426,-0.900976657867432,-0.928125143051147,-0.96601539850235,-1.06425762176514,-1.17519545555115,-1.34785175323486,-1.49785149097443,-1.55527329444885,-1.59394514560699,-1.69326162338257,-1.76699209213257,-1.86279296875,-1.89541041851044,-1.90576195716858,-1.88779282569885,-1.83378922939301,-1.87822258472443,-1.94599628448486,-1.97011709213257,-2.04501914978027,-2.15087914466858,-2.17363286018372,-2.20800805091858,-2.24091815948486,-2.33154296875,-2.54238271713257,-2.65644550323486,-2.74951171875,-2.90761709213257,-2.95224571228027,-3.00419926643372,-3.05156230926514,-3.14238309860229,-3.20087909698486,-3.27509784698486,-3.38271474838257,-3.52744150161743,-3.57626962661743,-3.62041044235229,-3.66162085533142,-3.69423794746399,-3.71142578125,-3.73984408378601,-3.85263681411743,-3.88232445716858,-3.92343735694885,-3.98466825485229,-4.0205078125,-4.08447265625,-4.09999990463257,-4.08164072036743,-4.05419874191284,-4.06455039978027,-4.10908174514771,-4.16630840301514,-4.22939443588257,-4.34912061691284,-4.39199161529541,-4.38994121551514,-4.34072256088257,-4.37626934051514,-4.41074180603027,-4.42216777801514,-4.41738271713257,-4.42207050323486,-4.49638652801514,-4.54023456573486,-4.62011766433716,-4.67529296875,-4.79169893264771,-4.83242177963257,-4.84794950485229,-4.81669950485229,-4.78564405441284,-4.75957012176514,-4.68125009536743,-4.5810546875,-4.28290987014771,-4.24462890625,-4.21054697036743,-4.15634775161743,-4.09267568588257,-4.01484346389771,-3.98427724838257,-3.91943359375,-3.55576157569885,-3.52060556411743,-3.46035170555115,-3.40400409698486,-3.20517587661743,-3.16709017753601,-3.01015639305115,-2.88095736503601,-2.75166010856628,-2.67031288146973,-2.64560580253601,-2.64160132408142,-2.66757798194885,-2.73261737823486,-2.86962890625,-2.94931626319885,-3.05283212661743,-3.15429711341858,-3.24687504768372,-3.34814476966858,-3.70732378959656,-3.74785161018372,-3.85234355926514,-4.08574199676514,-4.41513681411743,-4.61738252639771,-4.72724628448486,-4.96318387985229,-5.09267568588257,-5.14609384536743,-5.39257764816284,-5.49003887176514,-5.59101581573486,-5.54160118103027,-5.54414033889771,-5.55937528610229,-5.57548856735229,-5.57763719558716,-5.59628915786743,-5.66181659698486,-5.68828153610229,-5.69335985183716,-5.61103534698486,-5.52167940139771,-5.42480421066284,-5.31415987014771,-5.20058584213257,-5.07919931411743,-4.87734365463257,-4.74189424514771,-4.630859375,-4.52314424514771,-4.42353487014771,-4.03437519073486,-3.99980449676514,-3.7685546875,-3.72978496551514,-3.66738295555115,-3.60781264305115,-3.55410122871399,-3.51298809051514,-3.47539067268372,-3.45898461341858,-3.47529268264771,-3.53759789466858,-3.48271465301514,-3.39804697036743,-3.28017592430115,-3.15664052963257,-3.04062509536743,-2.92851543426514,-2.85009765625,-2.76474595069885,-2.72080063819885,-2.68232440948486,-2.65019512176514,-2.63144493103027,-2.59746122360229,-2.48291039466858,-2.38232398033142,-2.25849604606628,-2.14003920555115,-2.03095722198486,-1.9296875,-1.82529294490814,-1.65966796875,-1.58427751064301,-1.49570286273956,-1.40820300579071,-1.24345707893372,-0.906738340854645,-0.727929711341858,-0.680761516094208,-0.763964712619781,-0.861914157867432,-0.773242115974426,-0.68632835149765,-0.483593553304672,-0.0884767174720764,-0.0510253272950649,-0.0569823756814003,-0.0221190359443426,0.0400881394743919,0.186914309859276,0.238671869039536,0.44506847858429,0.520214736461639,0.566601395606995,0.692529499530792,0.740136563777924,0.774169683456421,0.861230611801147,0.970800817012787,0.979150533676147,0.983154296875,0.887548863887787,0.848681688308716,0.817529380321503,0.854394674301147,0.902392506599426,0.943652331829071,0.986670017242432,1.03564476966858,1.14926731586456,1.25283181667328,1.28896474838257,1.31181669235229,1.32578134536743,1.32763659954071,1.26249980926514,1.24980485439301,1.25454092025757,1.24360370635986,1.21440410614014,1.15551769733429,1.10473656654358,1.07968735694885,1.06796872615814,1.08852517604828,1.03115212917328,1.01806640625,0.984472751617432,0.940576076507568,0.862890481948853,0.845702886581421,0.850000083446503,0.922997713088989,0.938965022563934,0.941797077655792,0.928075850009918,0.838183403015137,0.850439548492432,1.02226555347443,1.18510758876801,1.23046875,1.30405247211456,1.39243185520172,1.41616201400757,1.44140601158142,1.46757805347443,1.57602524757385,1.67216813564301,1.70102524757385,1.68569338321686,1.64365255832672,1.50229477882385,1.47871065139771,1.40839850902557,1.37890601158142,1.18022489547729,1.08261692523956,0.995312452316284],[2.07841825485229,1.99653327465057,1.94345688819885,1.88256859779358,1.78916037082672,1.71572268009186,1.73320305347443,1.75937521457672,1.90131866931915,2.02167987823486,2.06782245635986,2.06074237823486,2.12670874595642,2.07841825485229],[0.848144292831421,0.825927793979645,0.832128942012787,0.876806497573853,0.934717059135437,1.04257810115814,1.11562502384186,1.12705087661743,1.15781247615814,1.23789083957672,1.31660151481628,1.40063488483429,1.51752912998199,1.55922842025757,1.57255887985229,1.52773416042328,1.46372091770172,1.36728525161743,1.10639643669128,1.06943380832672,0.979247868061066,0.907129168510437,0.876806497573853,0.804980456829071,0.733789324760437,0.638818144798279,0.549951255321503,0.508251905441284,0.438476592302322,0.360351502895355,0.323242157697678,0.305371046066284,0.268359124660492,0.216259479522705,0.337890386581421,0.39155250787735,0.397948980331421,0.414257645606995,0.460888415575027,0.471874892711639,0.438085824251175,0.372265577316284,0.298339694738388,0.206298604607582,0.150537207722664,0.0495115742087364,-0.248339965939522,-0.485253989696503,-0.731640756130219,-0.816308438777924,-0.892675697803497,-0.870019793510437,-0.787695527076721,-0.706054747104645,-0.657324135303497,-0.423534989356995,-0.379883050918579,-0.300390899181366,-0.241894751787186,-0.162890747189522,-0.0799316242337227,0.0348633378744125,0.149023458361626,0.288086026906967,0.33676740527153,0.38290998339653,0.489648580551147,0.610888779163361,0.680664122104645,0.742529213428497,0.796044647693634,0.848242282867432,0.924023330211639,1.13999009132385,1.25195288658142,1.46748065948486,1.57207012176514,1.63422858715057,1.70014619827271,1.84370136260986,1.96611332893372,2.13735389709473,2.17470693588257,2.19902348518372,2.15707993507385,2.11987280845642,1.94565403461456,1.90629875659943,1.83295893669128,1.78964865207672,1.70122051239014,1.58349609375,1.45810532569885,1.33173847198486,1.28959965705872,1.16279315948486,1.01386713981628,0.977197289466858,0.936132788658142,0.886572539806366,0.848144292831421],[2.07563471794128,2.05327153205872,2.15849590301514,2.21689462661743,2.23222637176514,2.21601557731628,2.20082998275757,2.14853501319885,2.07563471794128],[2.0517578125,2.03471708297729,2.08251929283142,2.29746103286743,2.46933579444885,2.57050752639771,2.59609365463257,2.59760737419128,2.47368168830872,2.22441411018372,2.09707021713257,2.0517578125],[2.651611328125,2.62954115867615,2.746826171875,2.80537080764771,2.78388667106628,2.76298809051514,2.70703125,2.651611328125],[2.36000967025757,2.35068368911743,2.37207055091858,2.42958974838257,2.59575223922729,2.59843754768372,2.64091777801514,2.65561532974243,2.76650381088257,2.82597684860229,2.85498046875,2.91601538658142,2.88906216621399,2.78139662742615,2.7412109375,2.72089862823486,2.66132807731628,2.51518559455872,2.46562457084656,2.41582036018372,2.36000967025757],[2.90541958808899,2.85302710533142,2.88564467430115,2.99189448356628,2.99897456169128,2.90541958808899],[2.86303687095642,2.85917973518372,2.88886690139771,2.94008755683899,2.98090791702271,3.01132822036743,3.06250023841858,3.03696274757385,3.01303720474243,2.99594712257385,2.984375,2.97651338577271,2.90395522117615,2.86303687095642],[3.15712857246399,3.08823227882385,3.12856459617615,3.20488286018372,3.22959017753601,3.21630835533142,3.15712857246399],[3.28051733970642,3.24775385856628,3.32822251319885,3.38637709617615,3.43198227882385,3.43608403205872,3.40751981735229,3.28051733970642],[3.43603539466858,3.40541982650757,3.46113252639771,3.54960942268372,3.59321308135986,3.63911104202271,3.68418002128601,3.73325204849243,3.67041015625,3.57109355926514,3.47651362419128,3.43603539466858],[3.76845693588257,3.75693368911743,3.78388690948486,3.81342768669128,3.84316420555115,3.85790991783142,3.81240200996399,3.78720664978027,3.76845693588257],[3.87465786933899,3.83251953125,3.92841792106628,4.04194307327271,4.00141572952271,3.91772437095642,3.87465786933899],[4.18613290786743,4.09052753448486,4.05429697036743,4.00400352478027,4.12148427963257,4.16899394989014,4.16694355010986,4.18613290786743],[3.68964815139771,3.65307641029358,3.70454096794128,3.76796865463257,3.77216792106628,3.78457021713257,3.81035161018372,3.85209941864014,3.88896489143372,3.98286128044128,4.04257822036743,4.20048809051514,4.21713829040527,4.15175771713257,3.98618125915527,3.8759765625,3.836181640625,3.68964815139771],[4.17060565948486,4.16230440139771,4.13339805603027,4.07607460021973,3.98833012580872,3.92993211746216,3.87797832489014,3.79672837257385,3.770263671875,3.73388671875,3.68925786018372,3.64482426643372,3.63632822036743,3.67827153205872,3.73037147521973,3.66557598114014,3.62851548194885,3.63930654525757,3.63896512985229,3.62265634536743,3.61264681816101,3.59199213981628,3.42661142349243,3.36538100242615,3.24355483055115,3.19374990463257,3.16518568992615,3.10458946228027,3.09848642349243,3.06435537338257,3.0048828125,2.95082998275757,2.92929720878601,2.91494154930115,2.88730454444885,2.85927700996399,2.82529282569885,2.80693340301514,2.77558565139771,2.74677753448486,2.66894507408142,2.541748046875,2.37763667106628,2.31782245635986,2.26274442672729,2.21542954444885,2.15996098518372,2.08701181411743,2.06064438819885,2.02685546875,2.00200223922729,1.96840798854828,1.86679673194885,1.70185542106628,1.64033174514771,1.41645479202271,1.31899416446686,1.09584939479828,1.04428720474243,0.98212867975235,0.886865317821503,0.839209020137787,0.813525557518005,0.847070574760437,0.874365091323853,0.929150223731995,1.03916037082672,1.09868144989014,1.03198230266571,0.96381813287735,0.889550745487213,0.831347405910492,0.788671791553497,0.754003822803497,0.729638874530792,0.34101590514183,0.235888451337814,-0.20048825442791,-0.323730319738388,-0.554394543170929,-0.675293207168579,-0.72753894329071,-0.770898699760437,-0.796679437160492,-0.867187678813934,-0.925683796405792,-1.00898432731628,-1.11269545555115,-1.18769502639771,-1.2236328125,-1.21826183795929,-1.18378925323486,-1.1171875,-1.04423844814301,-1.09814465045929,-1.15078115463257,-1.20712864398956,-1.26660168170929,-1.32734382152557,-1.37578105926514,-1.42861354351044,-1.47392594814301,-1.55312466621399,-1.59804689884186,-1.63281226158142,-1.71249985694885,-1.74433612823486,-1.78486323356628,-1.77861332893372,-1.78486323356628,-1.86416006088257,-1.92314434051514,-2.05253887176514,-2.13984394073486,-2.158203125,-2.18671846389771,-2.20791006088257,-2.29970693588257,-2.41083979606628,-2.51054692268372,-2.53828144073486,-2.51982426643372,-2.52158212661743,-2.55185556411743,-2.60332012176514,-2.57802748680115,-2.70683598518372,-2.83271479606628,-2.90214872360229,-2.95878911018372,-2.97695279121399,-2.93457007408142,-2.98378872871399,-3.02529263496399,-3.12636709213257,-3.14853525161743,-3.18300795555115,-3.23320317268372,-3.34824228286743,-3.43281245231628,-3.52333974838257,-3.59501957893372,-3.90683579444885,-4.16972684860229,-4.15185546875,-4.11171865463257,-3.70332002639771,-3.49443340301514,-3.37666034698486,-3.48183584213257,-3.47119164466858,-3.44443368911743,-3.41005849838257,-3.36474633216858,-3.23515629768372,-3.30625009536743,-3.36113262176514,-3.35439467430115,-3.32724618911743,-3.28515601158142,-3.27890610694885,-3.39433622360229,-3.45624995231628,-3.45527362823486,-3.41992211341858,-3.33203148841858,-3.24609351158142,-3.19570302963257,-3.177734375,-3.18408226966858,-3.22890663146973,-3.22363305091858,-3.24648427963257,-2.93349647521973,-3.18710923194885,-3.32216787338257,-3.40048837661743,-3.37109398841858,-3.32099628448486,-3.38144493103027,-3.52968764305115,-3.55253887176514,-3.55185532569885,-3.53251957893372,-3.42011737823486,-3.30771470069885,-3.05722665786743,-3.00800776481628,-2.93915987014771,-2.88945293426514,-2.92578148841858,-2.97548842430115,-2.97334003448486,-2.93369102478027,-2.95644497871399,-3.05576157569885,-3.07109379768372,-3.04873037338257,-2.99511742591858,-2.94619131088257,-2.90859389305115,-2.9384765625,-2.98867154121399,-3.02089810371399,-3.00478482246399,-2.89140629768372,-2.93378901481628,-2.94677758216858,-2.98535132408142,-2.96611309051514,-2.92509794235229,-2.68867206573486,-2.23388671875,-2.00136733055115,-1.94638681411743,-1.86279296875,-1.74287116527557,-1.64257836341858,-1.52568352222443,-1.39882826805115,-1.27480471134186,-1.18115246295929,-1.10107421875,-1.01132833957672,-0.944238424301147,-0.868750095367432,-0.875390708446503,-0.845800876617432,-0.807421863079071,-0.732031524181366,-0.680175960063934,-0.667382955551147,-0.638183295726776,-0.577441215515137,-0.494922071695328,-0.445409953594208,-0.390918225049973,-0.265038818120956,-0.185546651482582,-0.142480283975601,-0.00942401029169559,0.0311522893607616,0.0557619705796242,0.073827899992466,0.117480412125587,0.167675524950027,0.252831935882568,0.355664104223251,0.532812356948853,0.793945372104645,0.912646651268005,1.13461923599243,1.20449185371399,1.22392547130585,1.25385737419128,1.25815403461456,1.24716806411743,1.23964858055115,1.43847644329071,1.49589824676514,1.60708010196686,1.70546841621399,1.82109367847443,1.92270481586456,2.02753877639771,1.89619112014771,1.84834015369415,1.80629885196686,1.77666020393372,1.61489248275757,1.52294933795929,1.43896460533142,1.39785146713257,1.33803689479828,1.28256833553314,1.23574221134186,1.19013643264771,0.995995998382568,0.939062297344208,0.88207995891571,0.861962854862213,0.878124892711639,1.01733410358429,1.02636742591858,1.05053699016571,1.043212890625,0.995751917362213,0.994336187839508,1.02260756492615,1.01420927047729,0.999462962150574,1.01166987419128,1.11328101158142,1.14335954189301,1.24360370635986,1.33818352222443,1.43906259536743,1.47963881492615,1.51474630832672,1.55908167362213,1.56699204444885,1.54755842685699,1.49624037742615,1.45712912082672,1.43388652801514,1.43178713321686,1.40810561180115,1.3271484375,1.30214858055115,1.30839836597443,1.23593759536743,1.26059591770172,1.31137716770172,1.37988293170929,1.43427717685699,1.45527327060699,1.45234382152557,1.47089838981628,1.50004875659943,1.45200169086456,1.46713864803314,1.51416015625,1.61704099178314,1.68627953529358,1.81904304027557,1.85078144073486,1.86899399757385,1.89394533634186,1.93378925323486,1.98002934455872,2.01894521713257,2.05161118507385,2.16240215301514,2.21293950080872,2.25048828125,2.26938509941101,2.35083031654358,2.44614243507385,2.49291968345642,2.523193359375,2.56689429283142,2.61240220069885,2.63422846794128,2.68701148033142,2.72343730926514,2.75781273841858,2.79121088981628,2.84111332893372,2.89487314224243,2.97446274757385,3.02592778205872,2.99394512176514,3.00873994827271,3.03432607650757,3.12812495231628,3.17314457893372,3.20864272117615,3.34238243103027,3.36167001724243,3.44575214385986,3.50229501724243,3.63369131088257,3.73305654525757,3.93876981735229,3.97553682327271,4.08198213577271,4.19301748275757,4.25375938415527,4.33330059051514,4.34804677963257,4.34868144989014,4.29067420959473,4.35517597198486,4.362548828125,4.35371065139771,4.32734394073486,4.30820322036743,4.37080049514771,4.35986328125,4.33842754364014,4.339111328125,4.35498046875,4.34013700485229,4.33706092834473,4.29931688308716,4.19287109375,4.17138671875,4.17060565948486],[4.03349590301514,4.01259803771973,4.020263671875,4.07099628448486,4.16220712661743,4.25849628448486,4.28256845474243,4.34418964385986,4.41582012176514,4.54790067672729,4.53720712661743,4.47983407974243,4.37250947952271,4.291015625,4.17998027801514,4.03349590301514],[5.22934579849243,5.22021484375,5.27446269989014,5.26914024353027,5.22993183135986,5.20732402801514,5.23603487014771,5.22832012176514,5.20585966110229,5.17036151885986,5.04013681411743,4.87998056411743,4.77748966217041,4.66225576400757,4.63520479202271,4.41455125808716,4.32231426239014,4.19453096389771,4.09287118911743,3.99755859375,3.92812466621399,3.88554692268372,3.83476567268372,3.75942373275757,3.71035122871399,3.58124971389771,3.31118154525757,3.18305659294128,2.98818349838257,2.89492177963257,2.79423832893372,2.64760756492615,2.46660161018372,2.41147446632385,2.33164095878601,2.25742173194885,2.18915987014771,2.136962890625,2.12006807327271,1.9892578125,1.94824206829071,2.05058598518372,2.14018535614014,2.19423818588257,2.24257802963257,2.28330111503601,2.29472637176514,2.25747084617615,2.10224652290344,2.01181650161743,1.88701176643372,1.75742197036743,1.69306623935699,1.67055642604828,1.66123044490814,1.62138688564301,1.44213879108429,1.35791003704071,1.25888681411743,1.14169919490814,1.01870119571686,0.990331768989563,0.841992437839508,0.779297113418579,0.74882835149765,0.715478777885437,0.664208769798279,0.578906238079071,0.491991966962814,0.415331959724426,0.297754138708115,0.216454863548279,0.24448224902153,0.278613328933716,0.331982612609863,0.399804800748825,0.49453130364418,0.513720810413361,0.506933569908142,0.480175852775574,0.38706049323082,0.288916260004044,0.174414083361626,0.0469728000462055,-0.0195802599191666,-0.0687497779726982,-0.19179704785347,-0.240429699420929,-0.27167996764183,-0.362207293510437,-0.418066680431366,-0.465527057647705,-0.533593833446503,-0.575585961341858,-0.754687368869781,-0.795703172683716,-0.88671863079071,-0.979101777076721,-1.02138674259186,-1.05429685115814,-1.05341780185699,-1.03837883472443,-1.07421851158142,-1.25068378448486,-1.36240255832672,-1.60009753704071,-1.69873046875,-1.81943368911743,-1.92177724838257,-1.98720729351044,-2.04082012176514,-2.09296894073486,-2.17177748680115,-2.23417973518372,-2.28271484375,-2.38554716110229,-2.42988324165344,-2.54335975646973,-2.59521460533142,-2.59814476966858,-2.57089829444885,-2.41884779930115,-2.39218759536743,-2.37089824676514,-2.35751962661743,-2.35625004768372,-2.38017559051514,-2.4296875,-2.49199223518372,-2.88779330253601,-3.10625004768372,-3.16064453125,-3.21718740463257,-3.26093769073486,-3.41005849838257,-3.45126962661743,-3.61367177963257,-3.73056650161743,-3.77968740463257,-3.8330078125,-3.88134813308716,-4.12177753448486,-4.16289043426514,-4.55390644073486,-4.65976572036743,-4.79365253448486,-5.00957059860229,-5.67656230926514,-5.71640586853027,-5.81826162338257,-5.81757783889771,-5.79960918426514,-5.76064443588257,-5.71230459213257,-5.67275381088257,-5.54951143264771,-5.57001972198486,-5.72285175323486,-5.74550771713257,-5.72685527801514,-5.68115186691284,-5.52041006088257,-5.57177734375,-5.64150381088257,-5.81621074676514,-5.89267587661743,-5.90790987014771,-5.90458965301514,-5.80312490463257,-5.69072294235229,-5.53886699676514,-5.46660184860229,-5.38593769073486,-5.07958984375,-5.03281259536743,-4.81640672683716,-4.76523447036743,-4.67568349838257,-4.59619140625,-4.470703125,-4.15214824676514,-3.96923851966858,-3.67451190948486,-3.59921908378601,-3.37802743911743,-3.24404287338257,-3.1669921875,-2.89882826805115,-2.80849599838257,-2.72871088981628,-2.66396498680115,-2.58779311180115,-2.34521484375,-2.24853491783142,-2.14394497871399,-1.93417942523956,-1.29912114143372,-1.10126960277557,-0.826660096645355,-0.798828065395355,-0.55292946100235,-0.474218815565109,-0.40019553899765,-0.313769459724426,-0.0329588241875172,0.0450682863593102,0.102441415190697,0.208593890070915,0.267773538827896,0.351757705211639,0.458935767412186,0.686377048492432,1.03193366527557,1.49462914466858,1.70195341110229,1.86459934711456,1.90214836597443,2.195068359375,2.23818349838257,2.26420903205872,2.28286170959473,2.35854458808899,2.49428701400757,2.67641592025757,2.78510713577271,2.84658217430115,2.97529315948486,3.07705092430115,3.18901371955872,3.27573275566101,3.57514667510986,3.65371108055115,3.70854520797729,3.76660180091858,3.81630849838257,3.986328125,4.07275438308716,4.26328134536743,4.66196250915527,4.76137638092041,4.86503934860229,4.97617197036743,5.28403329849243,5.34624004364014,5.41079092025757,5.46430683135986,5.51708936691284,5.56479454040527,5.59287118911743,5.62880849838257,5.62460899353027,5.60908174514771,5.57929658889771,5.51450204849243,5.35117197036743,5.29428720474243,5.26699256896973,5.22934579849243],[5.81240272521973,5.78413105010986,5.79853534698486,5.88950157165527,5.90703105926514,5.89775371551514,5.87675762176514,5.84267568588257,5.81240272521973],[54.1853523254395,54.0586891174316,54.0814437866211,54.0694351196289,54.0730476379395,54.1187973022461,54.2249031066895,54.2669410705566,54.376708984375,54.4071769714355,54.4029312133789,54.392578125,54.2690887451172,54.2253913879395,54.1853523254395],[6.8310546875,6.74868154525757,7.00068330764771,7.01606464385986,7.13623094558716,7.18359327316284,7.23662042617798,7.20683670043945,6.97348642349243,6.8310546875],[7.35649347305298,7.26186561584473,7.31874990463257,7.35810565948486,7.37993240356445,7.41059541702271,7.35649347305298],[7.87783241271973,7.87656211853027,7.91933631896973,7.96401357650757,8.00693321228027,8.01791954040527,7.94838857650757,7.89912080764771,7.87783241271973],[8.056640625,8.01943397521973,8.02446269989014,8.05268573760986,8.07265567779541,8.10859394073486,8.171875,8.22465896606445,8.21376991271973,8.15976619720459,8.056640625],[8.26909255981445,8.25668907165527,8.251953125,8.25351524353027,8.26591777801514,8.27529335021973,8.26484394073486,8.27456092834473,8.30634784698486,8.31650447845459,8.31103610992432,8.29306602478027,8.26909255981445],[8.24951171875,8.21205997467041,8.218505859375,8.27495098114014,8.327880859375,8.349365234375,8.24951171875],[9.13667011260986,9.13095760345459,9.16508769989014,9.20488262176514,9.23066425323486,9.243896484375,9.24052810668945,9.17338848114014,9.13667011260986],[10.5548830032349,10.520751953125,10.5474119186401,10.6505851745605,10.7511234283447,10.7935066223145,10.8655271530151,10.8974609375,10.7998046875,10.7042474746704,10.5548830032349],[11.2024908065796,11.1960935592651,11.2142581939697,11.2433109283447,11.2627449035645,11.2604484558105,11.2415533065796,11.2159185409546,11.2024908065796],[11.3811521530151,11.361328125,11.3864259719849,11.4267578125,11.5091304779053,11.4634275436401,11.4112300872803,11.3811521530151],[12.0368165969849,11.8994140625,11.95947265625,12.00244140625,12.0317869186401,12.0368165969849],[12.8648929595947,12.7999515533447,12.9392576217651,12.9567880630493,12.9615726470947,12.9485349655151,12.8648929595947],[11.5360841751099,11.5125484466553,11.5387210845947,11.7182130813599,11.8334474563599,11.8733892440796,11.9305171966553,11.9495115280151,12.0138673782349,12.1122064590454,12.1923828125,12.2146968841553,12.2155752182007,12.2257814407349,12.3025388717651,12.3359384536743,12.3573246002197,12.5412607192993,12.6156253814697,12.6690921783447,12.7796382904053,12.8208990097046,12.87890625,13.0026359558105,13.0395994186401,13.2305660247803,13.3581056594849,13.48583984375,13.5438480377197,13.5454587936401,13.4364748001099,13.4006834030151,13.336181640625,13.2520990371704,13.2215824127197,13.1548824310303,13.08935546875,13.0624990463257,12.9751949310303,12.9422855377197,12.8504877090454,12.5385246276855,12.4530754089355,12.43603515625,12.2279300689697,12.1814451217651,12.0792484283447,12.03466796875,11.9927730560303,11.9404296875,11.9175291061401,11.8746576309204,11.7646484375,11.63916015625,11.5360841751099],[35.4958992004395,35.4927749633789,35.4845199584961,35.4607925415039,35.4490242004395,35.4716300964355,35.490234375,35.4797859191895,35.4494132995605,35.3069343566895,35.2510261535645,35.1349143981934,34.9464836120605,34.7698211669922,34.6539077758789,34.6063957214355,34.5579605102539,34.5181655883789,34.4529304504395,34.3903312683105,34.3519554138184,34.3026390075684,34.2581062316895,34.2282218933105,34.18896484375,34.0876960754395,34.0555648803711,34.0133781433105,33.8875961303711,33.8087921142578,33.6503410339355,33.4997062683105,33.4311027526855,33.3867683410645,33.3465347290039,33.2914543151855,33.2503890991211,33.2262687683105,33.2189445495605,33.1719245910645,33.1081047058105,33.0525398254395,33.0226554870605,33.00146484375,32.946044921875,32.8648452758789,32.8090324401855,32.7587890625,32.7030754089355,32.5640144348145,32.507568359375,32.5010757446289,32.4972152709961,32.4757804870605,32.38818359375,32.3651390075684,32.3582038879395,32.4119644165039,32.4680671691895,32.499267578125,32.5583992004395,32.5970191955566,32.5789527893066,32.57080078125,32.5577163696289,32.5447273254395,32.5198745727539,32.4667015075684,32.3973617553711,32.3003387451172,32.2362327575684,32.2157707214355,32.0230484008789,31.9837875366211,31.9579601287842,31.8876457214355,31.8055171966553,31.7403812408447,31.6683597564697,31.6180648803711,31.55029296875,31.4718246459961,31.4365711212158,31.3237800598145,31.302490234375,31.2936515808105,31.301513671875,31.3313484191895,31.3372077941895,31.32861328125,31.4141101837158,31.4262199401855,31.4026355743408,31.2417488098145,31.105712890625,31.0799331665039,31.064208984375,30.9937019348145,30.9490718841553,30.9652328491211,30.96826171875,30.9246101379395,30.8941879272461,30.8887691497803,30.7819347381592,30.7898445129395,30.7592277526855,30.6837406158447,30.6053237915039,30.5684089660645,30.5613288879395,30.5094718933105,30.4635238647461,30.4488773345947,30.4148426055908,30.3603992462158,30.2905769348145,30.2371082305908,30.1645030975342,30.1800308227539,30.1719226837158,30.1397457122803,30.1193351745605,29.9943351745605,29.9558601379395,29.8998050689697,29.7302742004395,29.5720691680908,29.4233417510986,29.3180160522461,29.1946296691895,29.1243152618408,29.1003894805908,28.9941902160645,28.8703117370605,28.8301773071289,28.7760753631592,28.7233390808105,28.6664066314697,28.6120128631592,28.6048831939697,28.6357917785645,28.6651878356934,28.6496067047119,28.5962390899658,28.5539054870605,28.5396957397461,28.4685554504395,28.4095706939697,28.3350086212158,28.2894039154053,28.2408676147461,28.1763668060303,28.0622081756592,27.98046875,27.913818359375,27.8670902252197,27.8744621276855,27.8992671966553,27.9137687683105,27.9005870819092,27.8649425506592,27.7565441131592,27.6718273162842,27.6870613098145,27.6734371185303,27.5966796875,27.5189952850342,27.4676761627197,27.4445323944092,27.4022960662842,27.3709964752197,27.4102535247803,27.44482421875,27.46533203125,27.4563484191895,27.3959465026855,27.3778305053711,27.4351081848145,27.461669921875,27.4913578033447,27.4278316497803,27.3481941223145,27.2986812591553,27.2498531341553,27.2036609649658,27.0916996002197,27.041015625,26.9269046783447,26.8785152435303,26.8629398345947,26.8609867095947,26.8466300964355,26.7815418243408,26.7665519714355,26.7503414154053,26.7410144805908,26.7972164154053,26.83984375,26.8290042877197,26.7816410064697,26.712646484375,26.6397457122803,26.6041507720947,26.6001949310303,26.6493663787842,26.6117172241211,26.5979976654053,26.5744132995605,26.5562973022461,26.4959964752197,26.43505859375,26.4419441223145,26.555419921875,26.5416011810303,26.4332027435303,26.3942375183105,26.3603019714355,26.4229488372803,26.4049797058105,26.3991222381592,26.4292984008789,26.4369125366211,26.3823719024658,26.39501953125,26.4300289154053,26.58642578125,26.7248039245605,26.8073253631592,26.928466796875,27.0860824584961,27.1339359283447,27.4088859558105,27.5673828125,27.6424312591553,27.7492179870605,27.7986812591553,27.8433113098145,27.87060546875,27.904541015625,27.9330081939697,27.9489250183105,27.9688472747803,28.0116691589355,28.0344715118408,28.057373046875,28.0933589935303,28.0918464660645,28.0396976470947,28.0069332122803,27.9072761535645,27.86865234375,27.7673835754395,27.5218753814697,27.4298820495605,27.3628406524658,27.3160648345947,27.2974605560303,27.2181167602539,27.1755867004395,27.1342296600342,27.0990238189697,27.0655765533447,26.9614753723145,26.9322261810303,26.8650398254395,26.816162109375,26.8486328125,26.8265609741211,26.8034172058105,26.7962417602539,26.7789554595947,26.76220703125,26.7411117553711,26.7194328308105,26.7139167785645,26.7015628814697,26.7239265441895,26.7545909881592,26.8475093841553,26.8541507720947,26.8903312683105,26.8507823944092,26.7965831756592,26.7802238464355,26.7716789245605,26.7777347564697,26.8034172058105,26.7899417877197,26.8670902252197,26.8668937683105,26.8073253631592,26.802001953125,26.8307628631592,26.8529796600342,26.8600578308105,26.86083984375,26.85498046875,26.8748531341553,26.9148445129395,26.9751949310303,27.0408191680908,27.0792980194092,27.0999031066895,27.1473636627197,27.2143058776855,27.2906246185303,27.3646965026855,27.4501953125,27.4583015441895,27.4386234283447,27.4425277709961,27.4936027526855,27.5576667785645,27.6114253997803,27.677001953125,27.7373046875,27.7599620819092,27.7598152160645,27.7464370727539,27.7296867370605,27.7303714752197,27.8076171875,27.812255859375,27.8269519805908,27.8415031433105,27.8302230834961,27.8207511901855,27.8246097564697,27.845947265625,27.8791999816895,27.9489250183105,27.9889640808105,28.0257320404053,28.0498542785645,28.0615234375,28.0933589935303,28.1471195220947,28.2281246185303,28.32763671875,28.4022941589355,28.4927253723145,28.5908203125,28.6294918060303,28.6540546417236,28.6902332305908,28.7297840118408,28.80322265625,28.8607902526855,28.9075183868408,28.9595203399658,28.9758796691895,29.0599098205566,29.1446285247803,29.2162132263184,29.3124027252197,29.2970199584961,29.2516098022461,29.2012691497803,29.1758804321289,29.1440410614014,29.1491718292236,29.1040534973145,29.0495586395264,29.035888671875,29.0374011993408,29.0542964935303,29.102294921875,29.1370124816895,29.1511726379395,29.2063465118408,29.3138198852539,29.3909187316895,29.4471683502197,29.4241199493408,29.3813972473145,29.2724590301514,29.2458019256592,29.2609844207764,29.2490730285645,29.2098159790039,29.1612300872803,29.1176776885986,29.0820808410645,28.9634761810303,28.922607421875,28.9097175598145,29.0274391174316,29.0506839752197,29.0222663879395,28.9593257904053,28.82958984375,28.763671875,28.6065425872803,28.525390625,28.496826171875,28.4685554504395,28.4281730651855,28.4120597839355,28.3865242004395,28.3672847747803,28.367919921875,28.406005859375,28.4599132537842,28.4497566223145,28.3670406341553,28.3624019622803,28.3376941680908,28.3689441680908,28.3403339385986,28.23681640625,28.2179679870605,28.1552238464355,28.0859870910645,28.0308589935303,27.9823246002197,27.937744140625,27.9070796966553,27.8900394439697,27.8368644714355,27.760009765625,27.6982917785645,27.6438484191895,27.5867195129395,27.5148429870605,27.4395999908447,27.1633319854736,27.1154289245605,27.10205078125,27.13330078125,27.1778316497803,27.2961921691895,27.3314933776855,27.3392581939697,27.2783679962158,27.2612800598145,27.2170906066895,27.1280765533447,27.046630859375,27.0138187408447,26.950439453125,26.7560558319092,26.6722660064697,26.6414070129395,26.5972652435303,26.5254878997803,26.4739742279053,26.3472652435303,26.1911125183105,26.0914058685303,26.0704097747803,26.041259765625,25.9873046875,25.9413089752197,25.9129390716553,25.7704582214355,25.7002449035645,25.5972156524658,25.5193367004395,25.4588871002197,25.4100093841553,25.31982421875,25.2434577941895,25.2157230377197,25.1915035247803,25.1645984649658,25.1385746002197,25.0978507995605,25.0487308502197,24.9310073852539,24.7672367095947,24.6376457214355,24.5140628814697,24.4737300872803,24.3218746185303,24.1131839752197,23.97265625,23.87646484375,23.8720703125,23.9029293060303,23.9438953399658,23.9769039154053,24.0065441131592,24.00537109375,23.986083984375,23.9728507995605,23.9874019622803,24.0741214752197,24.064208984375,24.0218734741211,23.774169921875,23.6820793151855,23.5280284881592,23.3391590118408,23.1325187683105,23.0849628448486,23.0303707122803,23.0154781341553,23.0370121002197,23.0320320129395,22.997314453125,22.907958984375,22.8057136535645,22.7182140350342,22.6332530975342,22.547119140625,22.3602046966553,22.2918949127197,22.2306137084961,22.2051753997803,22.2094230651855,22.1839847564697,22.1457042694092,22.0037593841553,21.9889163970947,22.0101566314697,22.1047859191895,22.1324214935303,22.1309566497803,22.1060066223145,22.04931640625,22.0113277435303,21.9780769348145,22.0480461120605,22.4103031158447,22.5256843566895,22.6854019165039,22.7344245910645,22.8218250274658,22.8970203399658,22.9290027618408,23.06982421875,23.2423820495605,23.3238277435303,23.4924793243408,23.68359375,23.675537109375,23.7218761444092,23.7209968566895,23.6777839660645,23.6919918060303,23.6859855651855,23.5982418060303,23.5046863555908,23.4710445404053,23.3610343933105,23.2873039245605,23.2098140716553,23.1061038970947,23.0535163879395,23.0048332214355,22.9796886444092,22.9915523529053,23.03369140625,23.1412601470947,23.1999015808105,23.2138671875,23.197998046875,23.1304683685303,23.0745601654053,23.068359375,23.0770034790039,23.1043949127197,23.3736324310303,23.5810565948486,23.66064453125,23.7628898620605,23.9204597473145,24.018798828125,24.0604972839355,24.093505859375,24.10009765625,24.0907726287842,24.1065921783447,24.15283203125,24.1900882720947,24.205078125,24.2106437683105,24.17529296875,24.1953125,24.2606945037842,24.3255367279053,24.3567371368408,24.3708992004395,24.3743648529053,24.3861808776855,24.4080562591553,24.4939441680908,24.6857414245605,24.77099609375,24.7862300872803,24.88134765625,24.8950672149658,24.8487796783447,24.8494129180908,24.8685054779053,24.9033203125,24.9441413879395,25.01513671875,25.1109371185303,25.1694831848145,25.16064453125,25.1421375274658,25.151611328125,25.177978515625,25.174072265625,25.1594734191895,25.167724609375,25.1666011810303,25.15771484375,25.1849613189697,25.219970703125,25.2583484649658,25.2931632995605,25.2927722930908,25.3053703308105,25.3361320495605,25.3758296966553,25.5601558685303,25.839599609375,25.94140625,26.1712417602539,26.2138195037842,26.2156734466553,26.18603515625,26.13232421875,26.0724124908447,26.0052719116211,25.9835453033447,26.006103515625,26.03759765625,26.10595703125,26.2022457122803,26.308349609375,26.3769054412842,26.4102554321289,26.4195327758789,26.412109375,26.3750972747803,26.3379878997803,26.2861328125,26.2508792877197,26.245361328125,26.260498046875,26.252197265625,26.2793960571289,26.2818355560303,26.2917003631592,26.3529777526855,26.4306640625,26.5177745819092,26.571533203125,26.5641117095947,26.5047855377197,26.4825687408447,26.4715328216553,26.4371089935303,26.4010257720947,26.3694820404053,26.31201171875,26.2575206756592,26.1780757904053,26.087158203125,26.0182113647461,25.9563484191895,25.8882312774658,25.8411140441895,25.8114242553711,25.789794921875,25.6981945037842,25.5744132995605,25.5370121002197,25.4953136444092,25.490478515625,25.4562492370605,25.3655281066895,25.3335456848145,25.290771484375,25.25927734375,25.2229976654053,25.1943855285645,25.1762199401855,25.1689453125,25.1804695129395,25.1878910064697,25.1884288787842,24.9615230560303,24.8818359375,24.8819332122803,24.9206066131592,24.9146480560303,24.7130374908447,24.6644535064697,24.6278324127197,24.5499019622803,24.4857902526855,24.4606437683105,24.479736328125,24.453857421875,24.3892574310303,24.3466320037842,24.3259773254395,24.27490234375,24.2309093475342,24.1862316131592,24.0696296691895,24.0025405883789,23.8263664245605,23.6744136810303,23.6021976470947,23.57275390625,23.5500011444092,23.4930171966553,23.4366207122803,23.2928218841553,23.2549800872803,23.2296867370605,23.2103996276855,23.1866207122803,23.04052734375,22.9388675689697,22.843505859375,22.6875495910645,22.671142578125,22.6320323944092,22.5109367370605,22.2746086120605,22.1862297058105,22.0931644439697,21.9372062683105,21.9379386901855,21.8336429595947,21.7586917877197,21.6541023254395,21.641357421875,21.653076171875,21.7446765899658,21.661376953125,21.5843753814697,21.6219730377197,21.6624031066895,21.7334957122803,22.0054206848145,22.0360832214355,22.05615234375,22.0669422149658,22.1219711303711,21.8321285247803,21.7137699127197,21.6597156524658,21.6142578125,21.7233390808105,21.7582035064697,21.6968746185303,21.622314453125,21.6357898712158,21.6941413879395,21.7935543060303,22.0329093933105,22.1395511627197,22.2177238464355,22.2656745910645,22.3743171691895,22.2550296783447,22.212646484375,22.1827144622803,22.1217288970947,22.04736328125,22.0010738372803,21.825439453125,21.7273445129395,21.6535148620605,21.5448741912842,21.5007820129395,21.3653316497803,21.2367172241211,21.1063480377197,20.965576171875,20.7450695037842,20.7001476287842,20.6197757720947,20.5343265533447,20.4191398620605,20.3559093475342,20.313232421875,20.171630859375,20.0889167785645,20.0067386627197,20.0537605285645,20.0530281066895,19.98779296875,19.9446773529053,19.91943359375,19.8958015441895,19.791748046875,19.6929187774658,19.6969242095947,19.7269039154053,19.7534675598145,19.8668937683105,19.8876953125,19.8958988189697,19.7576656341553,19.6208992004395,19.5948734283447,19.601318359375,19.6789073944092,19.6568851470947,19.6265621185303,19.5083484649658,19.1253910064697,19.0500984191895,18.9646987915039,18.8843250274658,18.6897468566895,18.4005851745605,18.2926750183105,18.0698738098145,18.0036144256592,17.7866706848145,17.6089839935303,17.4618148803711,17.27392578125,17.09619140625,16.9780769348145,16.9360847473145,16.8785629272461,16.8251953125,16.7828121185303,16.70654296875,16.6643562316895,16.5598640441895,16.4853515625,16.3294925689697,16.33447265625,16.365234375,16.3370609283447,16.2639636993408,15.9617681503296,15.8814449310303,15.8087406158447,15.7583494186401,15.7596673965454,15.7822265625,15.7659177780151,15.8673334121704,15.8880853652954,15.89501953125,15.7927732467651,15.7107429504395,15.3236331939697,15.0740222930908,14.7982416152954,14.5778312683105,14.4783210754395,14.3494148254395,14.2865715026855,14.2122068405151,14.0589351654053,13.8582029342651,13.7734861373901,13.6858406066895,13.4850587844849,13.5212888717651,13.6057615280151,13.7137689590454,13.6062498092651,13.5287113189697,13.4367189407349,13.361328125,12.6903314590454,12.4520015716553,12.2958011627197,12.2354488372803,11.98876953125,11.6902341842651,11.5752925872803,11.4466800689697,11.37060546875,11.3125486373901,11.3386726379395,11.2688474655151,11.1968755722046,10.7688474655151,10.3225584030151,10.3043460845947,10.2997064590454,10.3123540878296,10.3296394348145,10.3059568405151,10.2566890716553,10.1748046875,10.0352048873901,9.68310546875,9.56577110290527,9.45287990570068,9.393798828125,9.33334922790527,9.30893611907959,9.28461837768555,9.25214862823486,9.1923828125,9.25600624084473,9.2685546875,9.10502910614014,8.99018573760986,8.890869140625,8.66337966918945,8.51132869720459,8.38457012176514,8.18984413146973,8.1298828125,8.07832050323486,8.14531230926514,8.31591796875,8.40727519989014,8.84707069396973,8.90278339385986,9.09077072143555,9.16084003448486,9.18876934051514,9.20781326293945,9.23681640625,9.45209980010986,9.67646503448486,9.92709922790527,9.90986347198486,9.82734394073486,9.70737266540527,9.53989219665527,9.52045917510986,9.53623104095459,9.922119140625,10.0179691314697,10.0242671966553,10.0861330032349,10.1637697219849,10.2006349563599,10.3270015716553,10.4022455215454,10.7840824127197,11.0575685501099,11.3617677688599,11.4684085845947,11.7031240463257,11.8122072219849,11.9584474563599,12.0233402252197,12.0575199127197,12.5645503997803,12.8445796966553,12.9768552780151,13.0773439407349,13.5069332122803,13.583740234375,13.6676273345947,13.8496580123901,14.0463380813599,14.1688470840454,14.2165031433105,14.4074220657349,14.4947271347046,14.575439453125,14.6495113372803,14.7088861465454,14.9021968841553,14.9493646621704,15.0747547149658,15.3064451217651,15.39697265625,15.39697265625,15.4824714660645,15.53857421875,15.5730466842651,15.6593751907349,15.6569328308105,15.7088871002197,15.87109375,16.0542488098145,16.152099609375,16.4598636627197,17.1985340118408,17.5274410247803,17.6219234466553,17.90673828125,18.0977039337158,18.25927734375,18.3656253814697,18.5761241912842,18.642822265625,18.6830577850342,18.7789554595947,18.9271965026855,19.0210933685303,19.1533203125,19.0145015716553,18.9755859375,19.0792961120605,19.21875,19.2520999908447,19.2989273071289,19.2774410247803,19.3629894256592,19.4131832122803,19.4505367279053,19.5198230743408,19.5782718658447,19.7571277618408,19.7979488372803,19.8309574127197,20.0780277252197,20.5631847381592,20.6727542877197,20.8285140991211,20.9524898529053,21.0835933685303,21.1171875,21.129150390625,21.1776371002197,21.3719730377197,21.4357433319092,21.4708003997803,21.4559097290039,21.4618167877197,21.55126953125,21.6199226379395,21.6996097564697,21.7504386901855,21.7046890258789,21.687255859375,21.6965827941895,21.8775882720947,21.93798828125,21.9719219207764,21.9617176055908,21.9762210845947,22.1599617004395,22.1996097564697,22.2071781158447,22.2333011627197,22.2636222839355,22.2781257629395,22.2480964660645,22.2702140808105,22.2697257995605,22.2451667785645,22.1892108917236,22.0897445678711,22.0276374816895,21.9848155975342,21.9199714660645,21.8629875183105,21.8230476379395,21.7945804595947,21.7743167877197,21.7282238006592,21.6612319946289,21.531005859375,21.2240715026855,21.1557140350342,20.9705581665039,20.8697738647461,20.7388668060303,20.7145023345947,20.7404289245605,20.8401851654053,21.0946769714355,21.1788082122803,21.5057125091553,21.6785640716553,21.8395500183105,21.9915027618408,22.19677734375,22.290283203125,22.3854007720947,22.4373054504395,22.416259765625,22.3360843658447,22.3001956939697,22.2854976654053,22.4083976745605,22.4035148620605,22.4651851654053,22.4517574310303,22.5477046966553,22.5535144805908,22.57275390625,22.8157711029053,22.9703121185303,23.0024909973145,23.0401363372803,23.0895023345947,23.0771007537842,23.0301265716553,22.9734859466553,22.9397468566895,22.9708976745605,22.9656734466553,22.947021484375,22.8564453125,22.7751960754395,22.7590808868408,22.8485355377197,23.0537109375,23.18994140625,23.3640613555908,23.5714855194092,23.6294918060303,23.7541522979736,23.8521003723145,23.8084964752197,23.7479972839355,23.70556640625,23.6168460845947,23.5969734191895,23.7289047241211,23.8573246002197,23.9005374908447,23.927978515625,23.9508800506592,23.9672355651855,23.9666004180908,23.9646968841553,24.265625,24.2919921875,24.3072261810303,24.313720703125,24.30908203125,24.2640132904053,24.2665042877197,24.2924308776855,24.2730960845947,24.2863292694092,24.2686519622803,24.2682609558105,24.275390625,24.2730960845947,24.2251949310303,24.172607421875,24.1652355194092,24.17138671875,24.1915531158447,24.2405776977539,24.2875003814697,24.3562984466553,24.4121589660645,24.4183101654053,24.3857898712158,24.3311042785645,24.279052734375,24.24609375,24.2379875183105,24.2454109191895,24.261962890625,24.34375,24.3623542785645,24.3610343933105,24.4000968933105,24.4299812316895,24.4443378448486,24.4874038696289,24.5224609375,24.5718746185303,24.6187515258789,24.6539058685303,24.687744140625,24.7576675415039,24.8916015625,25.06298828125,25.2058601379395,25.3310546875,25.4228992462158,25.6257820129395,25.6669425964355,25.69189453125,25.7059574127197,25.685302734375,25.6813468933105,25.6857433319092,25.70654296875,25.9100570678711,25.9900398254395,26.0719718933105,26.2147960662842,26.3475589752197,26.471435546875,26.5064449310303,26.5480480194092,26.5787582397461,26.5861320495605,26.6270503997803,26.6991214752197,26.74267578125,26.77099609375,26.804443359375,26.9541492462158,27.1229496002197,27.1746101379395,27.2280769348145,27.2645015716553,27.3126945495605,27.4736328125,27.6947269439697,27.8490238189697,27.8948726654053,27.9341316223145,27.9816398620605,28.0250492095947,28.0231437683105,27.9837894439697,27.9374504089355,27.8353519439697,27.7689952850342,27.72900390625,27.7096195220947,27.7144546508789,27.8316402435303,27.8552742004395,27.869873046875,27.9150867462158,27.9624996185303,28.0474605560303,28.1772956848145,28.3463382720947,28.4217758178711,28.5658206939697,28.697265625,28.7519035339355,28.8961429595947,29.0287609100342,29.0888195037842,29.3639163970947,29.5506362915039,29.6106929779053,29.7729988098145,29.9343757629395,29.9716796875,30.033203125,30.0933589935303,30.1620121002197,30.2220706939697,30.2816410064697,30.3521480560303,30.3940410614014,30.4353504180908,30.5196800231934,30.7689952850342,30.8935527801514,30.8934059143066,30.9596672058105,31.03466796875,31.0687503814697,31.1128444671631,31.1326656341553,31.1855945587158,31.2613754272461,31.4653797149658,31.52392578125,31.7129402160645,31.76513671875,31.8185558319092,31.8897476196289,31.9488277435303,32.08935546875,32.1047859191895,32.1403312683105,32.2152824401855,32.2792015075684,32.3188972473145,32.3721199035645,32.4203605651855,32.4622077941895,32.4578132629395,32.4937973022461,32.5189437866211,32.6077156066895,32.7576675415039,32.7709007263184,32.7532234191895,32.77099609375,32.7686996459961,32.8104476928711,32.86083984375,32.927978515625,32.9917984008789,33.005126953125,33.0203132629395,33.075439453125,33.1434059143066,33.189453125,33.22119140625,33.2421875,33.30126953125,33.3841323852539,33.4553680419922,33.5069808959961,33.5450668334961,33.5917015075684,33.6324234008789,33.6678237915039,33.7212905883789,33.7881851196289,33.8386726379395,33.8865737915039,33.9460945129395,33.9901885986328,34.00341796875,34.0037117004395,34.018798828125,34.0430679321289,34.07568359375,34.1080093383789,34.144775390625,34.1913070678711,34.2366218566895,34.287841796875,34.3253402709961,34.3782234191895,34.42236328125,34.4853057861328,34.529052734375,34.6534652709961,34.6806640625,34.7208976745605,34.765380859375,34.7320327758789,34.7157707214355,34.677734375,34.6458511352539,34.6368179321289,34.6390151977539,34.6013679504395,34.5367164611816,34.5027351379395,34.5030746459961,34.5602569580078,34.612548828125,34.669921875,34.667724609375,34.756103515625,34.7408676147461,34.73583984375,34.7869148254395,34.8475570678711,34.8778305053711,34.9001960754395,34.938720703125,34.9919929504395,35.0623512268066,35.1099128723145,35.1715316772461,35.2355461120605,35.3025856018066,35.3787612915039,35.4432601928711,35.4958992004395],[-12.1941404342651,-12.1998043060303,-12.1955080032349,-12.1925783157349,-12.1872072219849,-12.1865234375,-12.1818361282349,-12.17333984375,-12.1615238189697,-12.1732425689697,-12.1846675872803,-12.1941404342651],[-12.1818361282349,-12.1868162155151,-12.1814451217651,-12.1876955032349,-12.1973628997803,-12.1796875,-12.1506834030151,-12.1261720657349,-12.1261720657349,-12.1360349655151,-12.1441402435303,-12.1602535247803,-12.1712894439697,-12.1818361282349],[-10.4929685592651,-10.5641603469849,-10.5249996185303,-10.5125007629395,-10.4596681594849,-10.4522457122803,-10.4494142532349,-10.4306640625,-10.4929685592651],[-7.37744092941284,-7.41718769073486,-7.43535184860229,-7.33447313308716,-7.26337957382202,-7.26337957382202,-7.2998046875,-7.39570331573486,-7.36757802963257,-7.33779335021973,-7.30966758728027,-7.27822303771973,-7.23027324676514,-7.22040987014771,-7.26171875,-7.29482412338257,-7.37744092941284],[53.9131355285645,53.8845710754395,53.9205551147461,53.959716796875,53.9776878356934,54.016845703125,54.0052223205566,54.0036163330078,53.9872093200684,53.9131355285645],[54.0887184143066,54.0535163879395,54.0172386169434,54.0036163330078,54.0110359191895,53.9872093200684,53.9413108825684,53.8821754455566,53.8402328491211,53.7457046508789,53.640869140625,53.5775413513184,53.4989242553711,53.4603042602539,53.390869140625,53.3664054870605,53.3012237548828,53.1663093566895,53.0911636352539,52.9270973205566,52.8656272888184,52.8072776794434,52.7381324768066,52.6634750366211,52.5431137084961,52.4020042419434,52.366943359375,52.3453598022461,52.2466773986816,52.2026863098145,52.1888198852539,52.2135238647461,52.2104988098145,52.1785659790039,52.1592292785645,52.1685562133789,52.24951171875,52.1659164428711,52.1393089294434,52.1449699401855,52.1227035522461,52.098876953125,52.0616226196289,52.0185546875,51.9931144714355,51.979736328125,51.947998046875,51.935302734375,51.8657722473145,51.8255882263184,51.8135261535645,51.85400390625,51.8783187866211,51.8906745910645,51.8887710571289,51.8762702941895,51.8477058410645,51.79296875,51.7393074035645,51.7120590209961,51.70703125,51.6513671875,51.6361808776855,51.5849113464355,51.4982414245605,51.4972152709961,51.519287109375,51.529052734375,51.5221710205078,51.4737281799316,51.4833488464355,51.6037101745605,51.6644515991211,51.6811027526855,51.6892547607422,51.6470680236816,51.6111335754395,51.6006813049316,51.6555671691895,51.730712890625,51.76611328125,51.7801246643066,51.8242683410645,51.8744125366211,51.77099609375,51.7835960388184,51.8124542236328,51.7989234924316,51.8687515258789,51.9745140075684,52.02001953125,52.0445785522461,52.079833984375,52.1229476928711,52.1366729736328,52.125732421875,52.1349105834961,52.1690902709961,52.2069320678711,52.2716789245605,52.2820777893066,52.2759284973145,52.2593269348145,52.2376441955566,52.2500953674316,52.2914543151855,52.37548828125,52.4037094116211,52.4426765441895,52.4663581848145,52.5469245910645,52.5591812133789,52.5787582397461,52.6211433410645,52.6796379089355,52.7123069763184,52.7554206848145,52.6682624816895,52.6349143981934,52.6170883178711,52.6269035339355,52.6539573669434,52.6436538696289,52.6227569580078,52.5799827575684,52.5697250366211,52.648193359375,52.7811508178711,52.8231925964355,52.896240234375,52.9287605285645,52.947265625,53.0975608825684,53.1248512268066,53.1292495727539,53.153076171875,53.1531753540039,53.1620597839355,53.2070808410645,53.2357406616211,53.25048828125,53.23486328125,53.2382316589355,53.2520523071289,53.27197265625,53.3230476379395,53.33447265625,53.33447265625,53.31884765625,53.3203582763672,53.3427238464355,53.3949699401855,53.4072761535645,53.3970222473145,53.412841796875,53.4456062316895,53.4783210754395,53.5093269348145,53.5485343933105,53.5678215026855,53.5614242553711,53.5904273986816,53.6044921875,53.6331024169922,53.6576194763184,53.6951179504395,53.7271957397461,53.781494140625,53.805419921875,53.8411636352539,53.8798332214355,53.8910140991211,53.8637237548828,53.9375991821289,54.0042991638184,54.0482902526855,54.0953598022461,54.0752410888672,54.1416015625,54.1871070861816,54.15576171875,54.2158203125,54.2578125,54.2760276794434,54.2681121826172,54.2688941955566,54.3004379272461,54.3085441589355,54.2986297607422,54.2096214294434,54.2255363464355,54.2817878723145,54.2879905700684,54.2634773254395,54.2311058044434,54.2412109375,54.3036117553711,54.3468742370605,54.403564453125,54.4419403076172,54.4610824584961,54.48486328125,54.5072746276855,54.5801277160645,54.6408195495605,54.6092796325684,54.6812019348145,54.7320289611816,54.7608871459961,54.782958984375,54.8094711303711,54.83154296875,54.8894538879395,54.965087890625,55.0204124450684,55.0564422607422,55.1082038879395,55.1462898254395,55.159912109375,55.1953125,55.19189453125,55.2000503540039,55.185791015625,55.2483406066895,55.2564926147461,55.2439956665039,55.1996574401855,55.17138671875,55.1222152709961,55.084228515625,55.0549774169922,55.0250473022461,54.970947265625,54.9939956665039,55.0469741821289,55.0902824401855,55.1447257995605,55.19384765625,55.2479476928711,55.2817840576172,55.2987785339355,55.3602066040039,55.3658218383789,55.35302734375,55.30517578125,55.2676277160645,55.2378921508789,55.1783218383789,55.1370124816895,55.0919914245605,55.0276870727539,55.0033187866211,54.9051284790039,54.8771018981934,54.825439453125,54.7679672241211,54.7457046508789,54.72802734375,54.71044921875,54.7192840576172,54.7178726196289,54.6983413696289,54.683349609375,54.6660652160645,54.6396980285645,54.6158218383789,54.5949211120605,54.5712394714355,54.512451171875,54.4769554138184,54.4535140991211,54.4142608642578,54.2965812683105,54.2837905883789,54.2152824401855,54.1866722106934,54.1438484191895,54.1335945129395,54.1373062133789,54.1212425231934,54.1334457397461,54.156005859375,54.214111328125,54.239501953125,54.27490234375,54.3018074035645,54.3553695678711,54.4082527160645,54.4066886901855,54.3743171691895,54.3291015625,54.2940406799316,54.2686538696289,54.21435546875,54.1956062316895,54.1847152709961,54.1634292602539,54.0847663879395,54.0586433410645,54.0572776794434,54.0636253356934,54.0606422424316,54.0770988464355,54.0948715209961,54.0887184143066],[26.921142578125,26.8011722564697,26.7011241912842,26.7322769165039,26.7308101654053,26.6924800872803,26.6858386993408,26.6175289154053,26.5831069946289,26.5857429504395,26.5926284790039,26.6392097473145,26.6575698852539,26.64794921875,26.7100105285645,26.8119621276855,26.857177734375,26.9309577941895,26.9476566314697,26.9098148345947,26.9833488464355,27.0032730102539,26.9520988464355,26.921142578125],[38.8777809143066,38.8690414428711,38.9126472473145,38.9066886901855,38.9043960571289,39.08740234375,39.1484375,39.18017578125,39.2528839111328,39.3409690856934,39.4238739013672,39.4983367919922,39.5433578491211,39.6485824584961,39.68505859375,39.6839370727539,39.560546875,39.4483375549316,39.3990707397461,39.35498046875,39.3123512268066,39.2811050415039,39.2411117553711,39.2028350830078,39.171630859375,39.09912109375,39.0592765808105,39.0188484191895,38.9936027526855,38.9789543151855,38.9586448669434,38.9350090026855,38.9118156433105,38.88427734375,38.853759765625,38.8190422058105,38.72412109375,38.689208984375,38.6422882080078,38.6134757995605,38.6056175231934,38.5862312316895,38.4110832214355,38.3987274169922,38.437255859375,38.4355010986328,38.3925285339355,38.1436538696289,38.01513671875,37.89013671875,37.7760772705078,37.6675796508789,37.6005859375,37.5199699401855,37.4966812133789,37.4805145263672,37.4448738098145,37.4071273803711,37.3805198669434,37.339599609375,37.149169921875,37.013671875,36.8102035522461,36.7425804138184,36.614501953125,36.6217308044434,36.8687515258789,36.9303245544434,36.9303245544434,36.8812026977539,36.8531265258789,36.8184547424316,36.8126945495605,36.8183097839355,36.8496589660645,36.9013214111328,36.9524917602539,37.1817359924316,37.3435554504395,37.3324737548828,37.3536148071289,37.4075698852539,37.4402351379395,37.4447250366211,37.4701652526855,37.5019035339355,37.72265625,37.7779312133789,37.9024887084961,37.9813461303711,38.0511245727539,38.0997543334961,38.0946273803711,38.0775413513184,38.078369140625,38.0733909606934,38.0804176330566,38.0948219299316,38.1911125183105,38.2225112915039,38.2494163513184,38.2496109008789,38.2566413879395,38.2500495910645,38.2130355834961,38.2099609375,38.2164077758789,38.1795883178711,38.13037109375,38.0893096923828,38.0329093933105,37.9899406433105,37.9733390808105,37.9477043151855,37.928466796875,37.9052734375,37.86083984375,37.8304672241211,37.7830543518066,37.6658210754395,37.647216796875,37.6353530883789,37.6385269165039,37.6881828308105,37.6515617370605,37.65625,37.6834945678711,37.649658203125,37.520751953125,37.5237312316895,37.5106430053711,37.4811515808105,37.4447250366211,37.333740234375,37.2528305053711,37.1789054870605,37.1384773254395,37.0416030883789,36.962890625,36.8294410705566,36.653564453125,36.6376457214355,36.6429672241211,36.642578125,36.572265625,36.4327125549316,36.2896957397461,36.1905250549316,36.09912109375,36.0528335571289,35.9999008178711,35.9767570495605,35.9436988830566,35.8676300048828,35.7618141174316,35.70556640625,35.6592788696289,35.6195793151855,35.5534210205078,35.5137710571289,35.4740715026855,35.4244613647461,35.3616218566895,35.31201171875,35.2888641357422,35.2723121643066,35.20947265625,35.1565437316895,35.09375,35.0507316589355,35.0011215209961,34.9217300415039,34.8556175231934,34.7993659973145,34.749755859375,34.7100601196289,34.6538543701172,34.6339836120605,34.5876960754395,34.5546379089355,34.5447273254395,34.5182647705078,34.4917945861816,34.4752426147461,34.4180183410645,34.3194313049316,34.3071746826172,34.2196311950684,34.0947761535645,33.8419914245605,33.7119140625,33.638916015625,33.5883293151855,33.5604019165039,33.5586891174316,33.5389633178711,33.5052223205566,33.4562492370605,33.3638191223145,33.3235359191895,33.1378402709961,33.0587882995605,32.994873046875,32.7943840026855,32.6000022888184,32.2494125366211,32.16796875,31.9871120452881,31.877197265625,31.7344722747803,31.6605968475342,31.4951667785645,31.4832515716553,31.4511222839355,31.4216289520264,31.3824214935303,31.2853031158447,31.0725593566895,30.9132823944092,30.8319358825684,30.5993633270264,30.3637218475342,30.12841796875,29.8586902618408,29.6634254455566,29.5427227020264,29.3726062774658,29.331787109375,29.2649898529053,29.0060539245605,28.8708992004395,28.7916011810303,28.6676769256592,28.5465335845947,28.4910163879395,28.4788093566895,28.4147472381592,28.3638668060303,28.2351570129395,28.2527847290039,28.2528820037842,28.2435550689697,28.2020511627197,28.0020503997803,27.800537109375,27.4970207214355,27.4445323944092,27.3567390441895,27.3001937866211,27.265625,27.2501964569092,27.2294445037842,27.2184085845947,27.2524909973145,27.2439460754395,27.2079105377197,27.1514644622803,27.1245613098145,27.0776863098145,26.9981441497803,26.8792476654053,26.86474609375,26.8375968933105,26.6655750274658,26.6497554779053,26.63232421875,26.6438961029053,26.63916015625,26.5936527252197,26.56103515625,26.5426273345947,26.4908695220947,26.427490234375,26.3692378997803,26.3570308685303,26.3689937591553,26.3182621002197,26.242431640625,26.2259292602539,26.165283203125,25.995849609375,25.8433589935303,25.8210926055908,25.7689933776855,25.7512702941895,25.6923828125,25.5846176147461,25.2861328125,25.2023448944092,25.1955070495605,25.1536617279053,25.1021003723145,25.1419906616211,25.1838855743408,25.2822265625,25.329833984375,25.4135246276855,25.4370613098145,25.3115711212158,25.3841304779053,25.36181640625,25.40087890625,25.4032707214355,25.4814949035645,25.427734375,25.4172859191895,25.5545902252197,25.5924320220947,25.5808601379395,25.5916023254395,25.6408195495605,25.691650390625,25.6530284881592,25.7249011993408,25.7915515899658,25.9188480377197,26.0372066497803,26.1588382720947,26.3714351654053,26.6800785064697,26.8006839752197,26.9054698944092,26.9945793151855,27.0899906158447,27.127685546875,27.2002449035645,27.1906242370605,27.1431159973145,27.03759765625,26.9775390625,26.93212890625,26.8299331665039,26.7705574035645,26.7859382629395,26.7253913879395,26.5566883087158,26.5050792694092,26.5089359283447,26.5891590118408,26.6966304779053,26.7323741912842,26.7077140808105,26.7255859375,26.8517589569092,26.9432621002197,27.0044918060303,27.1419448852539,27.3233890533447,27.3919925689697,27.4933586120605,27.6165046691895,27.71728515625,27.8244152069092,27.8482418060303,27.844970703125,27.8642101287842,27.9100093841553,28.13134765625,28.2188472747803,28.4351558685303,28.5121097564697,28.7261219024658,28.7820796966553,28.8701648712158,28.9278316497803,29.00439453125,29.0626945495605,29.117431640625,29.1465816497803,29.2122058868408,29.3398418426514,29.4200706481934,29.5480003356934,29.6790542602539,29.8729019165039,29.9212417602539,30.048095703125,30.1985340118408,30.2093734741211,30.0289535522461,30.1304683685303,30.3069324493408,30.3334465026855,30.3739261627197,30.3972644805908,30.4067859649658,30.3753910064697,30.4125003814697,30.4723148345947,30.5094242095947,30.5065441131592,30.4504890441895,30.3972644805908,30.3276386260986,30.241455078125,30.1208992004395,30.0624008178711,30.0354976654053,30.0283184051514,29.9750499725342,29.9623527526855,30.0038089752197,30.03759765625,30.109619140625,30.1598644256592,30.1883296966553,30.2301750183105,30.2854480743408,30.3158226013184,30.3213367462158,30.3550300598145,30.4168472290039,30.4576644897461,30.4656257629395,30.6564464569092,30.8236331939697,30.9897937774658,30.9964370727539,31.0023918151855,31.1415023803711,31.4005851745605,31.6013660430908,31.7944355010986,31.9364242553711,32.0879898071289,32.1508293151855,32.340087890625,32.4237327575684,32.4555206298828,32.4740219116211,32.4666023254395,32.5684089660645,32.68798828125,32.8339347839355,32.9292488098145,32.9502410888672,32.9576683044434,32.9758796691895,33.0282249450684,33.0865211486816,33.1744117736816,33.2296371459961,33.4157218933105,33.4701156616211,33.4919929504395,33.545654296875,33.58154296875,33.6097679138184,33.6233406066895,33.6248054504395,33.6028327941895,33.6866722106934,33.9254875183105,33.9544906616211,33.9708518981934,34.0440444946289,34.1525382995605,34.2155303955078,34.28466796875,34.4151382446289,34.4703636169434,34.5338859558105,34.5815925598145,34.5745124816895,34.5738258361816,34.6126937866211,34.748779296875,34.7984390258789,35.0285148620605,35.0801773071289,35.1276359558105,35.1967315673828,35.2322731018066,35.2842750549316,35.3216819763184,35.4248046875,35.476806640625,35.524169921875,35.6081047058105,35.6731452941895,35.7113761901855,35.744140625,35.7732429504395,35.820556640625,35.8353996276855,35.8218269348145,35.836669921875,35.9283676147461,35.9771995544434,36.008544921875,36.0027847290039,36.0153312683105,36.0546379089355,36.35595703125,36.3971710205078,36.4073753356934,36.4092750549316,36.4300308227539,36.4716339111328,36.5260772705078,36.5975570678711,36.6588859558105,36.6983871459961,36.7376937866211,36.7659187316895,36.79931640625,37.0638694763184,37.135009765625,37.1424331665039,37.1563491821289,37.2170906066895,37.269775390625,37.2903823852539,37.3571281433105,37.4237289428711,37.4353981018066,37.5063972473145,37.5602035522461,37.608642578125,37.6363258361816,37.6581573486328,37.7103538513184,37.74462890625,37.8292503356934,37.8717765808105,37.8801727294922,37.9080543518066,37.9671859741211,38.038818359375,38.1092758178711,38.146484375,38.209716796875,38.2545890808105,38.3177757263184,38.334228515625,38.3567886352539,38.3695831298828,38.3747062683105,38.3862762451172,38.4201164245605,38.5578117370605,38.6406745910645,38.7006340026855,38.8360366821289,38.8632316589355,38.9343757629395,38.994384765625,39.0167503356934,39.0562515258789,39.1080589294434,39.1448249816895,39.1806144714355,39.2183113098145,39.2599601745605,39.3108406066895,39.3510246276855,39.37744140625,39.3929710388184,39.4052276611328,39.3967781066895,39.3960456848145,39.422119140625,39.666748046875,39.7312507629395,39.7685546875,39.6817398071289,39.6510734558105,39.6504402160645,39.6291007995605,39.4235343933105,39.3628883361816,39.3115730285645,39.2542953491211,39.2156219482422,39.1946792602539,39.13916015625,39.0958976745605,39.0062522888184,38.9728050231934,38.9079093933105,38.8777809143066],[37.1424331665039,37.135009765625,37.0638694763184,36.79931640625,36.7659187316895,36.7376937866211,36.6983871459961,36.6588859558105,36.5975570678711,36.5260772705078,36.4716339111328,36.4300308227539,36.4092750549316,36.4073753356934,36.3971710205078,36.35595703125,36.0546379089355,36.0153312683105,36.0027847290039,36.008544921875,35.9771995544434,35.9283676147461,35.836669921875,35.8218269348145,35.8353996276855,35.820556640625,35.7732429504395,35.744140625,35.7113761901855,35.6731452941895,35.6081047058105,35.524169921875,35.476806640625,35.4248046875,35.3216819763184,35.2842750549316,35.2322731018066,35.1967315673828,35.1276359558105,35.0801773071289,35.0285148620605,34.7984390258789,34.748779296875,34.6126937866211,34.5738258361816,34.5745124816895,34.5815925598145,34.5338859558105,34.4703636169434,34.4151382446289,34.28466796875,34.2155303955078,34.1525382995605,34.0440444946289,33.9708518981934,33.9544906616211,33.9254875183105,33.6866722106934,33.6028327941895,33.6248054504395,33.6233406066895,33.6097679138184,33.58154296875,33.545654296875,33.4919929504395,33.4701156616211,33.4157218933105,33.2296371459961,33.1744117736816,33.0865211486816,33.0282249450684,32.9758796691895,32.9576683044434,32.9502410888672,32.9292488098145,32.8339347839355,32.68798828125,32.5684089660645,32.4666023254395,32.4740219116211,32.4555206298828,32.4237327575684,32.340087890625,32.1508293151855,32.0879898071289,31.9364242553711,31.7944355010986,31.6013660430908,31.4005851745605,31.1415023803711,31.0023918151855,30.9964370727539,30.9897937774658,30.8236331939697,30.6564464569092,30.4656257629395,30.4576644897461,30.4168472290039,30.3550300598145,30.3213367462158,30.3158226013184,30.2854480743408,30.2301750183105,30.1883296966553,30.1598644256592,30.109619140625,30.03759765625,30.0038089752197,29.9623527526855,29.9384784698486,29.9567375183105,30.0409164428711,30.043212890625,30.0113296508789,29.9828128814697,30.0766124725342,30.0956058502197,30.0973148345947,30.0964832305908,30.0796890258789,30.04150390625,30.0009784698486,29.9613285064697,29.9399909973145,29.8229961395264,29.6728515625,29.5375022888184,29.3474617004395,29.2596664428711,29.0962409973145,29.0636711120605,29.0958480834961,29.1315422058105,29.1670894622803,29.1936054229736,29.2023448944092,29.4352531433105,29.6193370819092,29.8492183685303,30.083984375,30.3222179412842,30.4952125549316,30.7177753448486,30.92041015625,31.0803737640381,31.2203617095947,31.3297367095947,31.489013671875,31.6163578033447,31.7254390716553,31.8933601379395,31.93896484375,31.9950199127197,32.0425262451172,32.0917510986328,32.12451171875,32.2438468933105,32.3509750366211,32.3312034606934,32.3056640625,32.4725608825684,32.4931640625,32.710693359375,32.9346656799316,33.15087890625,33.3722152709961,33.5140151977539,33.6200180053711,33.7683601379395,33.911376953125,34.0476570129395,34.19775390625,34.33203125,34.3865699768066,34.4290542602539,34.6123046875,34.7689971923828,34.8053207397461,35.0273933410645,35.2881813049316,35.427490234375,35.5506362915039,35.6404304504395,35.724609375,35.8099594116211,35.93896484375,36.0733871459961,36.2030296325684,36.2724609375,36.3833503723145,36.4644012451172,36.5146484375,36.56640625,36.59716796875,36.7408218383789,36.8260231018066,36.9611320495605,37.0605964660645,37.0950202941895,37.1085968017578,37.1287117004395,37.249267578125,37.3619117736816,37.3718757629395,37.3349151611328,37.3247566223145,37.3673820495605,37.3448715209961,37.3165016174316,37.3146476745605,37.2445297241211,37.2358436584473,37.2272453308105,37.2235336303711,37.269287109375,37.3135261535645,37.3124542236328,37.3018569946289,37.282958984375,37.2498550415039,37.20263671875,37.0518074035645,37.0118675231934,36.9833030700684,36.97802734375,37.0107421875,37.0584945678711,37.1105461120605,37.1582527160645,37.176025390625,37.173583984375,37.165283203125,37.1424331665039],[66.228515625,66.2248001098633,66.2591323852539,66.2816925048828,66.3597106933594,66.3814468383789,66.3761215209961,66.3815460205078,66.3739776611328,66.34228515625,66.3314437866211,66.2842712402344,66.1778793334961,66.1256332397461,66.1024398803711,66.061279296875,66.0378875732422,66.05908203125,66.0508346557617,66.0202102661133,65.9898910522461,65.9598617553711,65.89697265625,65.8337860107422,65.7809066772461,65.7642593383789,65.7556686401367,65.7899398803711,65.7874069213867,65.7703628540039,65.7387161254883,65.7101058959961,65.6844711303711,65.6582565307617,65.5753402709961,65.6275405883789,65.6422805786133,65.6160659790039,65.5859375,65.5330047607422,65.550537109375,65.5495071411133,65.5193328857422,65.4871597290039,65.4413070678711,65.3989715576172,65.381591796875,65.3689956665039,65.3547897338867,65.322509765625,65.2909698486328,65.2895050048828,65.2750015258789,65.2574691772461,65.2369613647461,65.2228469848633,65.2151412963867,65.1925277709961,65.1430206298828,65.1246566772461,65.09765625,65.068115234375,65.0359344482422,65.016845703125,65.0137252807617,64.9928665161133,64.9580078125,64.9140167236328,64.8621597290039,64.7836380004883,64.7418975830078,64.7147903442383,64.724365234375,64.7452163696289,64.6774368286133,64.6356887817383,64.600830078125,64.5831069946289,64.538330078125,64.4939956665039,64.4459457397461,64.4159622192383,64.3798294067383,64.3196792602539,64.2958984375,64.2969207763672,64.2582015991211,64.1766586303711,64.1112289428711,64.0372085571289,63.9163551330566,63.865478515625,63.8517570495605,63.8409156799316,63.80810546875,63.7465782165527,63.7129898071289,63.6823768615723,63.6363754272461,63.6197280883789,63.6068878173828,63.5901870727539,63.5357437133789,63.4963340759277,63.4969711303711,63.5308609008789,63.5296859741211,63.5245094299316,63.5138664245605,63.4731941223145,63.4542465209961,63.4066886901855,63.4419937133789,63.478515625,63.5365715026855,63.5520477294922,63.5558128356934,63.6371116638184,63.6873550415039,63.7081985473633,63.7319831848145,63.7481918334961,63.7569808959961,63.7578620910645,63.7649383544922,63.80517578125,63.7921371459961,63.7353515625,63.7374000549316,63.7657699584961,63.7933578491211,63.8039054870605,63.8383750915527,63.8879356384277,63.9068374633789,63.9344215393066,63.9398460388184,63.9445304870605,63.9354476928711,63.872802734375,63.8583946228027,63.8437538146973,63.8372573852539,63.8277359008789,63.8685035705566,63.9594764709473,64.0193862915039,64.0483932495117,64.0832061767578,64.0772933959961,64.0496063232422,64.0103530883789,63.991455078125,64.0392150878906,64.0713424682617,64.1018600463867,64.1537628173828,64.1803207397461,64.2054138183594,64.2848663330078,64.32177734375,64.3490219116211,64.3666000366211,64.379150390625,64.3978500366211,64.3978500366211,64.3139114379883,64.3139114379883,64.3270034790039,64.3506851196289,64.3916015625,64.3946762084961,64.4131851196289,64.4522018432617,64.5149917602539,64.5978012084961,64.6100082397461,64.6263656616211,64.6397476196289,64.647705078125,64.5625457763672,64.5330581665039,64.538818359375,64.5718765258789,64.5865707397461,64.6244125366211,64.647216796875,64.7139663696289,64.7269058227539,64.7334899902344,64.7949752807617,64.788818359375,64.8243637084961,64.8092727661133,64.7565460205078,64.7391662597656,64.7506408691406,64.7785186767578,64.8161087036133,64.8634338378906,64.8964309692383,64.9152297973633,64.9241714477539,64.9127426147461,64.9458541870117,64.9527816772461,64.9580078125,64.9932556152344,65.0021438598633,64.9897994995117,64.9658737182617,65.0030288696289,65.0216827392578,65.0331039428711,65.0464859008789,65.0263671875,65.0257263183594,65.03955078125,65.045654296875,65.0487823486328,65.0791015625,65.1059112548828,65.1737365722656,65.1876907348633,65.125244140625,65.126220703125,65.1593322753906,65.19677734375,65.2268600463867,65.2916030883789,65.3435516357422,65.3997116088867,65.4215316772461,65.4473648071289,65.4934539794922,65.480712890625,65.535400390625,65.5677719116211,65.5474166870117,65.5804748535156,65.5347671508789,65.4686050415039,65.4227523803711,65.4075622558594,65.4450225830078,65.4872055053711,65.5003356933594,65.5251922607422,65.6012191772461,65.614990234375,65.6080093383789,65.5549774169922,65.5383758544922,65.6162109375,65.6461410522461,65.69091796875,65.7101593017578,65.7597122192383,65.7764663696289,65.7823257446289,65.7655792236328,65.6795883178711,65.69482421875,65.7265167236328,65.75,65.7622528076172,65.7637176513672,65.7812042236328,65.8063430786133,65.8492202758789,65.868896484375,65.8845748901367,65.8800277709961,65.9542922973633,65.9969787597656,66.017578125,66.04345703125,66.060791015625,66.0694274902344,66.0260696411133,66.0242233276367,66.05224609375,66.0934143066406,66.1088409423828,66.12158203125,66.14501953125,66.1644058227539,66.1810073852539,66.1817398071289,66.1666030883789,66.0862350463867,66.0636749267578,66.033935546875,65.9970703125,65.9821243286133,65.9948272705078,65.9793014526367,65.9834899902344,66.0390167236328,66.0259323120117,65.9999542236328,65.9764633178711,65.9441909790039,65.876953125,65.8674850463867,65.9054183959961,65.9083023071289,65.9273910522461,65.9980926513672,66.0576705932617,66.0700149536133,66.152587890625,66.1720657348633,66.2127380371094,66.2332077026367,66.25146484375,66.2587432861328,66.2577667236328,66.2663116455078,66.2877426147461,66.3015594482422,66.3139190673828,66.3376922607422,66.32470703125,66.3241729736328,66.3387145996094,66.3572311401367,66.3843765258789,66.429443359375,66.4406204223633,66.4327621459961,66.4454116821289,66.4301223754883,66.385498046875,66.30712890625,66.2569808959961,66.2412567138672,66.2001953125,66.0896987915039,66.0255889892578,66.00927734375,65.9900894165039,65.9675827026367,65.955078125,65.9387664794922,65.8953170776367,65.87646484375,65.7418975830078,65.7133331298828,65.6982421875,65.7235870361328,65.6807556152344,65.6351547241211,65.6096725463867,65.5846633911133,65.5789108276367,65.5924301147461,65.5782241821289,65.536376953125,65.5016555786133,65.4740676879883,65.462158203125,65.4658203125,65.4586868286133,65.4206008911133,65.3042449951172,65.2666015625,65.3000030517578,65.3849639892578,65.4283676147461,65.4302749633789,65.4445266723633,65.47119140625,65.565185546875,65.6364212036133,65.6582565307617,65.6630859375,65.6541976928711,65.5794906616211,65.5669403076172,65.571044921875,65.6217269897461,65.7190399169922,65.8277282714844,65.9477005004883,66.0332489013672,66.0843734741211,66.10009765625,66.0804672241211,66.0492706298828,65.9301300048828,65.8677749633789,65.80078125,65.779052734375,65.76806640625,65.7723617553711,65.787841796875,65.814453125,65.9849166870117,66.0379943847656,66.07568359375,66.097900390625,66.1215286254883,66.1603546142578,66.18115234375,66.1839370727539,66.1687927246094,66.1357421875,66.0713348388672,65.9645538330078,65.8847122192383,65.7580108642578,65.736572265625,65.7340850830078,65.7505340576172,65.77392578125,65.8302688598633,65.905029296875,66.0931625366211,66.1288070678711,66.1574249267578,66.1605453491211,66.143310546875,66.1141128540039,65.9991683959961,65.9713821411133,65.9644012451172,65.9783248901367,65.9996566772461,66.0255432128906,66.0888671875,66.2028274536133,66.2061996459961,66.1972198486328,66.1673812866211,66.1434555053711,66.125244140625,66.1316375732422,66.1715850830078,66.1959457397461,66.2525405883789,66.2783660888672,66.4467315673828,66.4811553955078,66.5229034423828,66.5260772705078,66.5146484375,66.432861328125,66.3917007446289,66.3585968017578,66.2857437133789,66.2587890625,66.228515625],[32.7349128723145,32.7289085388184,32.6820793151855,32.6680145263672,32.640869140625,32.619873046875,32.3955078125,32.4016609191895,32.4506340026855,32.4930152893066,32.5074729919434,32.512939453125,32.534423828125,32.46044921875,32.3381843566895,32.2810554504395,32.1609344482422,32.0871086120605,31.9916019439697,31.9132804870605,31.8664054870605,31.8412590026855,31.8233394622803,31.8167972564697,31.8379402160645,31.8167495727539,31.7763175964355,31.7500019073486,31.7344722747803,31.6732425689697,31.602294921875,31.5365715026855,31.3968734741211,31.3681621551514,31.3513202667236,31.3662090301514,31.4228038787842,31.48291015625,31.4792976379395,31.3253917694092,31.3248519897461,31.2305183410645,31.2144527435303,31.1324214935303,30.9822750091553,30.8602046966553,30.8022480010986,30.6734848022461,30.5239276885986,30.4208984375,30.3843250274658,30.1953144073486,30.1417007446289,29.9778804779053,29.8969230651855,29.7870597839355,29.5550270080566,29.4773445129395,29.5639171600342,29.8121089935303,29.9820308685303,30.1914539337158,30.4460468292236,30.5073738098145,30.5962905883789,30.8278331756592,30.9950199127197,31.2083015441895,31.2929210662842,31.3627433776855,31.5256366729736,31.5416488647461,31.5848636627197,31.59228515625,31.8957023620605,32.1963386535645,32.6140594482422,32.8266105651855,32.9671897888184,33.0836906433105,33.0919914245605,33.0795402526855,33.07568359375,33.1194801330566,33.25048828125,33.271484375,33.2406234741211,33.2750511169434,33.3326187133789,33.3697738647461,33.4156723022461,33.4317359924316,33.3704605102539,33.3305168151855,33.2782211303711,33.2495613098145,33.1924781799316,33.1356925964355,33.0885734558105,33.0393562316895,32.9980964660645,32.9496078491211,32.8623504638672,32.7823219299316,32.7349128723145],[36.7570304870605,36.7459983825684,36.7803688049316,36.82861328125,36.8430671691895,36.8209495544434,36.7763671875,36.7570304870605],[38.2203102111816,38.1066398620605,38.0629386901855,37.7848129272461,37.7205581665039,37.6507301330566,37.5895500183105,37.5318870544434,37.4585952758789,37.37548828125,37.334716796875,37.3080062866211,37.2828598022461,37.2443351745605,37.2091789245605,37.1387214660645,37.096923828125,37.05517578125,37.0132789611816,36.934814453125,36.8916015625,36.8392562866211,36.7852554321289,36.7364768981934,36.6878433227539,36.6938972473145,36.7235374450684,36.7104034423828,36.7666015625,36.7767562866211,36.7986831665039,36.9728507995605,37.0464363098145,37.1036605834961,37.1071319580078,37.1006355285645,37.1358871459961,37.254150390625,37.3487319946289,37.4103546142578,37.4518051147461,37.4792976379395,37.5065422058105,37.5705070495605,37.5751953125,37.5673828125,37.5718269348145,37.5943336486816,37.6695327758789,37.7737808227539,37.8197746276855,37.938720703125,38.0529289245605,38.0849609375,38.10791015625,38.1417007446289,38.1830558776855,38.063720703125,38.0348663330078,38.0413093566895,38.0840835571289,38.1309089660645,38.1903343200684,38.1914520263672,38.1805191040039,38.1268043518066,38.1102523803711,38.1031265258789,38.000732421875,37.9840354919434,37.981201171875,38.024169921875,38.04052734375,38.016845703125,38.0425796508789,38.0455093383789,38.0850601196289,38.1507835388184,38.1669921875,38.1716804504395,38.1675796508789,38.1527328491211,38.1680679321289,38.2110366821289,38.2303733825684,38.2173347473145,38.2908706665039,38.2958984375,38.267578125,38.2203102111816],[39.0675315856934,38.9686546325684,39.0386695861816,39.0987777709961,39.1159210205078,39.0906257629395,39.0675315856934],[40.7056159973145,40.6969718933105,40.708740234375,40.7240753173828,40.7618179321289,40.7394027709961,40.7181625366211,40.7056159973145],[41.0398445129395,40.994140625,40.9974632263184,41.0312995910645,41.09912109375,41.1218757629395,41.1016120910645,41.062744140625,41.0398445129395],[40.8820304870605,40.818115234375,40.5562019348145,40.4995613098145,40.4415054321289,40.4002914428711,40.2684097290039,40.1592292785645,40.091796875,40.0170402526855,39.9243659973145,39.3543968200684,39.2535667419434,39.166015625,39.1395530700684,39.1675300598145,39.216796875,39.2138175964355,39.1969718933105,39.2112808227539,39.2391586303711,39.0432624816895,38.9637222290039,38.9128875732422,38.90966796875,38.9267082214355,38.9265632629395,38.9643058776855,39.0303230285645,39.1104965209961,39.2057113647461,39.2917976379395,39.4815902709961,39.5230484008789,39.5627899169922,39.647705078125,39.7216796875,39.7480964660645,39.7216796875,39.7315940856934,39.7696762084961,39.8392105102539,39.8974609375,39.917236328125,39.9781723022461,40.0376472473145,40.0775871276855,40.1307106018066,40.2926750183105,40.3523445129395,40.4426765441895,40.5005340576172,40.5586433410645,40.60595703125,40.651611328125,40.7710456848145,40.8707046508789,40.913330078125,40.9070320129395,40.8575172424316,40.8463363647461,40.8343276977539,40.8501968383789,40.895263671875,40.9499015808105,41.1103515625,41.1429214477539,41.1851577758789,41.2421875,41.257080078125,41.20166015625,41.1958999633789,41.150146484375,41.1063461303711,41.0536613464355,41.030517578125,41.0172843933105,41.0048828125,40.9924812316895,40.93212890625,40.9147453308105,40.8820304870605],[42.858154296875,42.8191909790039,42.7965812683105,42.77099609375,42.7131805419922,42.7611351013184,42.7369155883789,42.7420387268066,42.7850608825684,42.810302734375,42.8157730102539,42.8280754089355,42.8223152160645,42.858154296875],[46.520263671875,46.462890625,46.4485359191895,46.4150848388672,46.3691902160645,46.3175315856934,46.2616233825684,46.2249526977539,46.2123069763184,46.2235336303711,46.2166023254395,46.1965827941895,46.1770515441895,46.1577644348145,46.1331062316895,46.089111328125,46.03955078125,46.0092277526855,45.9871101379395,45.9737777709961,45.9797859191895,45.961669921875,45.8341331481934,45.8123550415039,45.7919921875,45.7612800598145,45.680419921875,45.6148452758789,45.5928726196289,45.5819816589355,45.5876007080078,45.6272430419922,45.7709503173828,45.7707023620605,45.7099609375,45.7713851928711,45.74658203125,45.6978988647461,45.6375007629395,45.6107902526855,45.5442886352539,45.4972152709961,45.461669921875,45.4679222106934,45.544921875,45.5462913513184,45.4919929504395,45.446044921875,45.3688468933105,45.2415046691895,45.2077140808105,45.039794921875,44.9679679870605,44.8994140625,44.84521484375,44.79833984375,44.8331069946289,44.8322296142578,44.7225112915039,44.429443359375,44.223876953125,44.1342277526855,43.9947280883789,43.9211883544922,43.6860847473145,43.6116714477539,43.5712890625,43.389892578125,43.1803703308105,42.8515625,42.6895484924316,42.5064468383789,42.2442855834961,42.0525398254395,41.9340324401855,41.9132347106934,41.939453125,41.9281234741211,41.8961906433105,41.8140144348145,41.7584953308105,41.7007789611816,41.620849609375,41.5120582580566,41.4354019165039,41.2320327758789,41.0621566772461,40.9754371643066,40.840576171875,40.6551742553711,40.56494140625,40.370849609375,40.2210426330566,40.1048355102539,39.9868659973145,39.9036140441895,39.8213844299316,39.8525390625,39.9369621276855,40.2801780700684,40.31494140625,40.3402328491211,40.3990745544434,40.4378890991211,40.4864234924316,40.5027809143066,40.5134735107422,40.4580535888672,40.3264656066895,40.13720703125,39.8596687316895,39.74755859375,39.638916015625,39.5783195495605,39.4815902709961,39.380615234375,39.1365699768066,38.9980964660645,38.9193382263184,38.9397964477539,38.8896980285645,38.8001480102539,38.7147979736328,38.4935569763184,38.4090805053711,38.2495613098145,38.0863761901855,38.0186538696289,37.9418449401855,37.9391136169434,38.0342292785645,38.1753921508789,38.2623062133789,38.302978515625,38.4834938049316,38.6139144897461,38.6717300415039,38.7125968933105,38.7364234924316,38.7592277526855,38.9411163330078,39.0238265991211,39.1394538879395,39.3536148071289,39.6265144348145,39.8700675964355,39.9901885986328,40.0528297424316,40.0521469116211,40.0700187683105,40.239013671875,40.2647476196289,40.3095703125,40.3774909973145,40.4693336486816,40.5560531616211,40.6299819946289,40.6684074401855,40.644775390625,40.6264152526855,40.6327133178711,40.599853515625,40.5988273620605,40.7287101745605,40.7593231201172,40.8126487731934,40.8207054138184,40.8271484375,40.7939453125,40.812255859375,40.8703117370605,41.1299819946289,41.2356452941895,41.2544937133789,41.232177734375,41.2785148620605,41.2888641357422,41.2776832580566,41.2438468933105,41.2662086486816,41.3009262084961,41.4087371826172,41.4696769714355,41.8126449584961,41.9408683776855,42.08203125,42.2875480651855,42.3629417419434,42.4232902526855,42.4157257080078,42.3931159973145,42.389892578125,42.4166030883789,42.444091796875,42.4565925598145,42.53515625,42.7387199401855,42.8042984008789,42.8446769714355,42.8999519348145,42.9363288879395,42.9571762084961,42.95361328125,42.967529296875,43.0651397705078,43.14013671875,43.2038078308105,43.3711929321289,43.5130844116211,43.8521003723145,43.947509765625,44.0199699401855,44.1011734008789,44.3192405700684,44.322998046875,44.4077606201172,44.4223136901855,44.3461456298828,44.1365203857422,43.9189453125,43.8767585754395,43.8025856018066,43.7671394348145,43.8229484558105,43.8648910522461,43.9110832214355,43.9654273986816,44.0336418151855,44.0831527709961,44.1160125732422,44.1648445129395,44.1683616638184,44.1273918151855,44.1379852294922,44.2017097473145,44.280029296875,44.3357429504395,44.3920440673828,44.4281730651855,44.4632797241211,44.5106925964355,44.5645484924316,44.6316413879395,44.6771469116211,44.68896484375,44.7166976928711,44.8272972106934,44.8450202941895,44.8587379455566,44.8603019714355,44.8831520080566,44.92138671875,44.9729995727539,45.0226097106934,45.0681648254395,45.1179695129395,45.144287109375,45.1453132629395,45.1356468200684,45.215576171875,45.2226066589355,45.2399406433105,45.3490257263184,45.3817367553711,45.4009246826172,45.4236831665039,45.50048828125,45.58056640625,45.6703605651855,45.7100105285645,45.7408676147461,45.7800788879395,45.8145523071289,45.8683624267578,45.92578125,45.90380859375,45.8804206848145,45.912353515625,45.9444313049316,45.9781723022461,45.9722175598145,45.9218292236328,45.9474601745605,46.0159149169922,46.0519065856934,46.1609382629395,46.1875953674316,46.2560043334961,46.2710456848145,46.3412132263184,46.4034156799316,46.4451179504395,46.446044921875,46.431884765625,46.4027824401855,46.2828598022461,46.2458992004395,46.1598167419434,46.1107902526855,46.0771484375,46.06103515625,45.9961929321289,45.9186973571777,45.8619651794434,45.8300285339355,45.845703125,45.8755874633789,45.9281234741211,45.9831047058105,46.014892578125,46.0514640808105,46.1024398803711,46.21923828125,46.2867660522461,46.3912582397461,46.4751968383789,46.4955558776855,46.4806632995605,46.4823226928711,46.4308090209961,46.3487777709961,46.3061981201172,46.2960929870605,46.2958984375,46.3460426330566,46.3677749633789,46.36181640625,46.3276824951172,46.2380867004395,46.2279777526855,46.2382316589355,46.2535133361816,46.2879867553711,46.3628425598145,46.4207534790039,46.4478988647461,46.4832038879395,46.5467758178711,46.5999031066895,46.6143531799316,46.62109375,46.5648460388184,46.5470695495605,46.550048828125,46.5828590393066,46.6188468933105,46.6650390625,46.73486328125,46.8649406433105,46.8551292419434,46.8537101745605,46.8463859558105,46.7933120727539,46.7752418518066,46.7694816589355,46.7770004272461,46.7969741821289,46.859130859375,46.9361839294434,46.9756851196289,46.9830589294434,46.9974098205566,46.99658203125,46.9846649169922,46.986083984375,47.0396957397461,47.0821304321289,47.0750007629395,47.0608863830566,47.0281753540039,46.9847640991211,46.9352531433105,46.8358879089355,46.7598114013672,46.70263671875,46.6725082397461,46.6541023254395,46.6474609375,46.6258811950684,46.5726547241211,46.5579109191895,46.5555648803711,46.520263671875],[43.9345703125,43.8940925598145,43.9014167785645,43.9529762268066,43.98974609375,43.982421875,43.9345703125],[18.4574222564697,18.4214839935303,18.4029312133789,18.40185546875,18.3904285430908,18.3042964935303,18.2571773529053,18.15185546875,17.9703140258789,17.9135246276855,17.8798332214355,17.8682136535645,17.8662109375,17.9009761810303,17.9276371002197,17.9404296875,17.9648933410645,17.976318359375,17.9737300872803,17.9041004180908,17.8487796783447,17.8541030883789,17.9012699127197,17.8800773620605,17.8450679779053,17.7149410247803,17.779541015625,17.8336906433105,17.8560543060303,17.8597164154053,17.8773918151855,17.9874992370605,18.01904296875,18.0475597381592,18.173828125,18.191162109375,18.2180671691895,18.2872066497803,18.3497562408447,18.42626953125,18.4480953216553,18.44482421875,18.4678230285645,18.5006847381592,18.522216796875,18.467041015625,18.4664554595947,18.4574222564697],[49.2312469482422,49.1808128356934,49.1698265075684,49.1874008178711,49.1874008178711,49.1763687133789,49.266357421875,49.25537109375,49.2312469482422],[32.12451171875,32.0074729919434,31.9949207305908,31.9464855194092,31.8474617004395,31.7811527252197,31.6963367462158,31.6258792877197,31.5561027526855,31.4915027618408,31.3551750183105,31.1468276977539,31.0077648162842,30.8289527893066,30.6692867279053,30.5000019073486,30.442626953125,30.3481464385986,30.3309574127197,30.3132820129395,30.1445789337158,30.01171875,29.9950695037842,29.9462890625,29.8970699310303,29.8660163879395,29.8316402435303,29.6661148071289,29.4951171875,29.3553714752197,29.2005367279053,29.1904792785645,29.2142601013184,29.2548809051514,29.2940902709961,29.3209476470947,29.3535175323486,29.4844722747803,29.5550270080566,29.7870597839355,29.8969230651855,29.9778804779053,30.1417007446289,30.1953144073486,30.3843250274658,30.4208984375,30.5239276885986,30.6734848022461,30.8022480010986,30.8602046966553,30.9822750091553,31.1324214935303,31.2144527435303,31.2305183410645,31.3248519897461,31.3253917694092,31.4792976379395,31.5623512268066,31.67236328125,31.7655258178711,31.9849109649658,32.10302734375,32.2378921508789,32.3955078125,32.619873046875,32.640869140625,32.6680145263672,32.6820793151855,32.7289085388184,32.7349128723145,32.7137680053711,32.6666984558105,32.5337905883789,32.4951171875,32.4574737548828,32.3869132995605,32.361328125,32.3172874450684,32.4655265808105,32.5907707214355,32.7330551147461,32.829833984375,32.994873046875,33.0992202758789,33.2366218566895,33.3722152709961,33.15087890625,32.9346656799316,32.710693359375,32.4931640625,32.4725608825684,32.3056640625,32.3312034606934,32.3509750366211,32.2438468933105,32.12451171875],[24.2801265716553,24.2660636901855,24.2833003997803,24.2880363464355,24.3177738189697,24.3484859466553,24.3913097381592,24.4144535064697,24.3620128631592,24.3236312866211,24.2801265716553],[24.5159168243408,24.3580570220947,24.3350582122803,24.3476085662842,24.4358406066895,24.4696292877197,24.4518547058105,24.4586429595947,24.5871105194092,24.5663566589355,24.5159168243408],[24.7431640625,24.7170906066895,24.7325191497803,24.8719234466553,24.8523921966553,24.8046875,24.77685546875,24.7431640625],[26.6165046691895,26.6156749725342,26.7215347290039,26.7264652252197,26.7099609375,26.6487789154053,26.6165046691895],[26.6527824401855,26.60693359375,26.55224609375,26.5335922241211,26.4564933776855,26.4424819946289,26.3805656433105,26.328125,26.3189449310303,26.2550773620605,26.2086906433105,26.1712417602539,26.1525382995605,26.09716796875,26.0947265625,26.1544914245605,26.1991691589355,26.3079090118408,26.4339351654053,26.4485340118408,26.466064453125,26.5557136535645,26.5939464569092,26.6310539245605,26.6749515533447,26.693603515625,26.679443359375,26.6468753814697,26.643310546875,26.6677742004395,26.71142578125,26.796875,26.8818855285645,26.8121089935303,26.720703125,26.6527824401855],[27.7208003997803,27.7024917602539,27.727783203125,27.8424320220947,27.8979988098145,27.9102554321289,27.8111343383789,27.7702140808105,27.7208003997803],[28.1049308776855,28.0815925598145,28.1011238098145,28.1924800872803,28.1761703491211,28.1447277069092,28.1049308776855],[28.208984375,28.1277351379395,28.2008800506592,28.249755859375,28.262939453125,28.2825183868408,28.3135738372803,28.359619140625,28.395263671875,28.3975086212158,28.4310550689697,28.4612789154053,28.4758777618408,28.5174808502197,28.4696292877197,28.43212890625,28.4112796783447,28.3611831665039,28.2987308502197,28.2723140716553,28.2547855377197,28.208984375],[30.2629871368408,30.2414073944092,30.2646980285645,30.38818359375,30.4655265808105,30.3889636993408,30.3668937683105,30.2629871368408],[30.3969230651855,30.3863258361816,30.4442405700684,30.5750961303711,30.6711940765381,30.7922859191895,30.8188953399658,30.8284645080566,30.7908687591553,30.6424789428711,30.5299797058105,30.3969230651855],[31.6571273803711,31.6396484375,31.71826171875,31.787109375,31.7424812316895,31.6571273803711],[32.4237327575684,32.4193344116211,32.4310035705566,32.4627914428711,32.5271987915039,32.5157241821289,32.4577140808105,32.4237327575684],[32.2296905517578,32.1939926147461,32.2281723022461,32.2437477111816,32.2918472290039,32.3136711120605,32.34619140625,32.4688491821289,32.5216331481934,32.5412101745605,32.4916038513184,32.340576171875,32.2296905517578],[32.7838859558105,32.7568855285645,32.7723655700684,32.7759780883789,32.7628898620605,32.6933135986328,32.6521492004395,32.6463356018066,32.63671875,32.5861320495605,32.604736328125,32.62841796875,32.6620140075684,32.7838859558105],[33.0560531616211,33.0454597473145,33.0782241821289,33.107177734375,33.1251487731934,33.1292495727539,33.093505859375,33.0560531616211],[32.8402824401855,32.8294906616211,32.9196319580078,32.9518547058105,32.9690895080566,33.132568359375,33.0676727294922,33.0033187866211,32.9931144714355,32.9461936950684,32.9288558959961,32.8402824401855],[33.2230491638184,33.1758308410645,33.176025390625,33.2311019897461,33.2573738098145,33.3312530517578,33.3577613830566,33.3610343933105,33.2843246459961,33.2230491638184],[33.7488288879395,33.7073249816895,33.7396965026855,33.8289070129395,33.8583984375,33.8292007446289,33.7488288879395],[33.6025886535645,33.57275390625,33.5708999633789,33.5844230651855,33.6235847473145,33.6523933410645,33.6377944946289,33.6028327941895,33.5538101196289,33.5023460388184,33.391845703125,33.2740707397461,33.2520980834961,33.2545890808105,33.181640625,33.1180686950684,33.0877914428711,33.0470695495605,33.0101585388184,32.9736824035645,32.9190444946289,32.8823738098145,32.8439445495605,32.5928230285645,32.4656257629395,32.3254890441895,32.2230453491211,32.1167488098145,32.001953125,31.8834972381592,31.7784175872803,31.6708011627197,31.4046878814697,31.4096183776855,31.4418468475342,31.4368648529053,31.3776836395264,31.2561511993408,31.112060546875,31.0151386260986,31.0592784881592,31.0940914154053,31.1220703125,31.1558132171631,31.2690906524658,31.3832035064697,31.5260734558105,31.5774402618408,31.5981941223145,31.6041011810303,31.6240730285645,31.6712894439697,31.706298828125,31.7176742553711,31.7184085845947,31.6654300689697,31.5630855560303,31.4596672058105,31.403076171875,31.3523921966553,31.2674789428711,31.217529296875,31.1785163879395,31.2668952941895,31.273193359375,31.2918930053711,31.4084949493408,31.4365711212158,31.4506855010986,31.48779296875,31.6014652252197,31.6963367462158,31.7300777435303,31.7688484191895,31.8489742279053,31.9498519897461,32.0907745361328,32.1150398254395,32.1435050964355,32.218994140625,32.304931640625,32.4560546875,32.6192398071289,32.6263656616211,32.6569328308105,32.734130859375,32.8315925598145,32.9513664245605,33.0925788879395,33.15478515625,33.1776351928711,33.14453125,33.1048355102539,33.0129890441895,32.9317893981934,32.851318359375,32.8468284606934,32.8668479919434,32.8526344299316,32.8103523254395,32.755859375,32.7018547058105,32.6749992370605,32.6771469116211,32.706298828125,32.7478523254395,32.7708015441895,32.7217292785645,32.6217269897461,32.5709953308105,32.645263671875,32.725341796875,32.7816390991211,32.875244140625,32.9293937683105,32.9949226379395,33.0599594116211,32.9855499267578,32.8926734924316,32.8519058227539,32.8515625,32.9879913330078,33.0223617553711,33.0835914611816,33.1866226196289,33.2362785339355,33.3436508178711,33.364990234375,33.3598136901855,33.32177734375,33.375244140625,33.40380859375,33.43701171875,33.4834976196289,33.5217742919922,33.5396995544434,33.5781745910645,33.5982894897461,33.5977058410645,33.6344757080078,33.7342300415039,33.7889671325684,33.8346176147461,33.9154777526855,33.9277839660645,33.9177742004395,33.8720207214355,33.7758293151855,33.6728515625,33.6025886535645],[33.9451675415039,33.9085922241211,33.9235343933105,33.9131851196289,33.8799324035645,33.8470230102539,33.8714866638184,33.8728523254395,33.9277839660645,33.9478034973145,33.9451675415039],[34.1151847839355,34.0750961303711,34.0872535705566,34.1219711303711,34.1640625,34.1726570129395,34.1269035339355,34.1151847839355],[34.1233901977539,34.0828094482422,34.14501953125,34.3206558227539,34.2847671508789,34.2308120727539,34.1233901977539],[34.25634765625,34.2147483825684,34.2266120910645,34.0439453125,33.9826164245605,33.927734375,33.8478012084961,33.8205070495605,33.7292976379395,33.6083984375,33.5268058776855,33.439453125,33.3469696044922,33.2472152709961,33.2867660522461,33.4483413696289,33.49267578125,33.5163078308105,33.510986328125,33.3599624633789,33.2496109008789,33.0831565856934,33.0282249450684,33.012451171875,32.98388671875,32.8419914245605,32.7545890808105,32.7520027160645,32.7759246826172,32.762451171875,32.9024887084961,32.9195327758789,32.9165992736816,33.0076637268066,33.0593757629395,33.12646484375,33.18115234375,33.2112808227539,33.25537109375,33.2930679321289,33.3045883178711,33.3312530517578,33.4304695129395,33.43408203125,33.4167976379395,33.3401374816895,33.3399925231934,33.3945808410645,33.4695320129395,33.512451171875,33.6329116821289,33.68994140625,33.7909202575684,33.8522453308105,33.9061508178711,33.992431640625,34.021240234375,34.0953140258789,34.088134765625,33.9971199035645,33.9272956848145,33.9332046508789,33.968994140625,33.97705078125,33.9728012084961,34.0171394348145,34.0693855285645,34.1346702575684,34.2438468933105,34.2328605651855,34.2373542785645,34.3068351745605,34.3480491638184,34.3583984375,34.3190422058105,34.25634765625],[34.483642578125,34.4637680053711,34.4689445495605,34.4230461120605,34.467041015625,34.496337890625,34.5192375183105,34.5343742370605,34.5223655700684,34.483642578125],[34.2881317138672,34.2029304504395,34.2088851928711,34.2469711303711,34.2941398620605,34.3681640625,34.47265625,34.5190925598145,34.544921875,34.5440444946289,34.3982925415039,34.2881317138672],[34.3536605834961,34.3055152893066,34.339599609375,34.3704566955566,34.5218772888184,34.5792961120605,34.6072731018066,34.6865730285645,34.6713371276855,34.6494636535645,34.6155281066895,34.5404319763184,34.4164543151855,34.3536605834961],[34.7265129089355,34.6795425415039,34.6898918151855,34.7205085754395,34.7754402160645,34.77587890625,34.7265129089355],[36.203857421875,36.16650390625,36.1787605285645,36.2326164245605,36.2934074401855,36.3401374816895,36.2463874816895,36.203857421875],[37.8221168518066,37.819580078125,37.8293952941895,37.8541984558105,37.8958015441895,37.9695320129395,37.9908218383789,37.9945793151855,38.0784645080566,38.1611328125,38.2914543151855,38.31591796875,38.2589836120605,38.1243171691895,38.07568359375,38.0655288696289,37.9039077758789,37.8221168518066],[41.3726539611816,41.3538055419922,41.4047355651855,41.2511749267578,41.0963363647461,40.83935546875,40.7233390808105,40.6111335754395,40.5307121276855,40.4736328125,40.2911643981934,40.0672340393066,39.95849609375,39.8444328308105,39.792236328125,39.7398910522461,39.6683578491211,39.6105499267578,39.4288063049316,39.2187004089355,39.111328125,39.0900382995605,39.0404281616211,39.0174331665039,38.9996070861816,38.9951667785645,38.974853515625,38.9179191589355,38.8651351928711,38.8165054321289,38.7628440856934,38.6320304870605,38.4978523254395,38.4041481018066,38.3797340393066,38.3813972473145,38.3379402160645,38.3125495910645,38.1488761901855,38.0528793334961,37.9496116638184,37.8226051330566,37.6984367370605,37.4672355651855,37.1146507263184,37.0020523071289,36.9257354736328,36.8903350830078,36.8468780517578,36.7318840026855,36.5027809143066,36.4453125,36.3855972290039,36.3078155517578,36.2313461303711,36.1424293518066,36.0592308044434,35.845703125,35.7825202941895,35.7249526977539,35.6612777709961,35.6320304870605,35.51025390625,35.394775390625,35.2669906616211,35.2211418151855,35.1814460754395,35.155029296875,35.0964813232422,35.0382804870605,34.9473152160645,34.8996086120605,34.9148941040039,34.9569320678711,35.0098609924316,35.0721664428711,35.2323265075684,35.2966804504395,35.3452606201172,35.4229965209961,35.4852066040039,35.5404281616211,35.5851554870605,35.6333503723145,35.668212890625,35.6683578491211,35.6580581665039,35.6121063232422,35.549560546875,35.5203628540039,35.4948234558105,35.4091300964355,35.3194808959961,35.2739753723145,35.2523918151855,35.2215347290039,35.1492652893066,35.1421394348145,35.2432594299316,35.2985343933105,35.298095703125,35.2780265808105,35.2107429504395,35.1548805236816,35.0971183776855,34.9564933776855,34.8391609191895,34.7360343933105,34.6983871459961,34.62841796875,34.6192359924316,34.6510238647461,34.69921875,34.875732421875,34.9748039245605,35.0252456665039,35.095703125,35.1240730285645,35.0864715576172,35.0441398620605,34.9871597290039,34.9151878356934,34.8477020263672,34.732666015625,34.5963401794434,34.6409149169922,34.65087890625,34.6474113464355,34.6642112731934,34.6363792419434,34.5828132629395,34.6214370727539,34.7035140991211,34.7275886535645,34.7725105285645,34.7747077941895,34.759033203125,34.7659187316895,34.8141136169434,34.8349113464355,34.815185546875,34.7215347290039,34.7090339660645,34.7331085205078,34.8058586120605,34.9125022888184,34.9787101745605,35.0355491638184,35.0595207214355,35.05029296875,35.0229988098145,34.984130859375,34.78955078125,34.6783714294434,34.5890617370605,34.4642105102539,34.43359375,34.3804664611816,34.3240699768066,34.2992668151855,34.2577171325684,34.1768569946289,34.0948715209961,33.7782211303711,33.5617179870605,33.4869613647461,33.5533676147461,33.6287612915039,33.7219734191895,33.8062515258789,33.8980484008789,34.0069808959961,34.1826171875,34.2883796691895,34.3165550231934,34.3808097839355,34.4167976379395,34.5004386901855,34.5469741821289,34.6174812316895,34.654296875,34.6529312133789,34.6310043334961,34.6618156433105,34.7470703125,34.7652359008789,34.7706031799316,34.7547874450684,34.7236824035645,34.7138671875,34.6976547241211,34.5931167602539,34.5272979736328,34.49462890625,34.4858894348145,34.4646987915039,34.4301261901855,34.4331550598145,34.3853492736816,34.343994140625,34.3024444580078,34.32958984375,34.2552261352539,34.24609375,34.2870597839355,34.3533668518066,34.324951171875,34.2270050048828,34.0320320129395,33.9442367553711,33.8387680053711,33.85546875,34.0452613830566,34.0520515441895,34.0193862915039,34.0036125183105,33.9651870727539,33.947998046875,33.9756355285645,34.0206527709961,34.0072746276855,33.9757308959961,34.2618179321289,34.299560546875,34.3497085571289,34.392578125,34.4073715209961,34.3934593200684,34.4131851196289,34.4698219299316,34.5501480102539,34.615478515625,34.6670913696289,34.726318359375,34.8093757629395,34.8999977111816,34.9665031433105,35.0223121643066,35.1562995910645,35.3068351745605,35.4183120727539,35.4490242004395,35.5112800598145,35.558837890625,35.5565452575684,35.4588394165039,35.4722175598145,35.4974594116211,35.5114288330078,35.4952621459961,35.494873046875,35.5072250366211,35.5392570495605,35.5779304504395,35.6279296875,35.6632308959961,35.7470703125,35.7411117553711,35.7217750549316,35.65966796875,35.5918922424316,35.5508766174316,35.5255393981934,35.5177230834961,35.5031242370605,35.5495109558105,35.6068878173828,35.6825218200684,35.7676277160645,35.8741188049316,35.9905738830566,36.1168441772461,36.2233428955078,36.2876968383789,36.3617706298828,36.5719718933105,36.7420387268066,36.9510269165039,37.1983871459961,37.3821258544922,37.4136734008789,37.4974632263184,37.5220718383789,37.4854469299316,37.4374504089355,37.2831535339355,37.2197265625,37.2000465393066,37.1719741821289,37.11767578125,37.0267601013184,36.9596176147461,36.8951187133789,36.8372077941895,36.7740745544434,36.753173828125,36.7537612915039,36.7703628540039,36.9247589111328,36.9515609741211,37.0645980834961,37.1510772705078,37.1733894348145,37.2184066772461,37.3921356201172,37.47216796875,37.5606460571289,37.6634292602539,37.7747077941895,37.8439445495605,38.0090789794922,38.0990257263184,38.142578125,38.2687492370605,38.3998031616211,38.5025367736816,38.598876953125,38.697021484375,38.7881355285645,38.881591796875,39.1049346923828,39.2285614013672,39.2731437683105,39.3106460571289,39.3580551147461,39.4111328125,39.4637184143066,39.6244125366211,39.749267578125,39.8550796508789,39.8851089477539,39.8868675231934,39.8777351379395,39.9208488464355,39.9589347839355,39.9660148620605,39.9856948852539,40.021728515625,40.136962890625,40.2603530883789,40.3148918151855,40.414306640625,40.5338859558105,40.5984344482422,40.6727561950684,40.733154296875,40.7473602294922,40.7515602111816,40.77490234375,40.8087882995605,40.8460922241211,40.9476585388184,41.0056648254395,41.160888671875,41.2033195495605,41.2297859191895,41.2096672058105,41.2056655883789,41.2118186950684,41.1954078674316,41.1556167602539,40.8932609558105,40.8578109741211,40.830322265625,40.8343276977539,40.8751487731934,40.9295387268066,40.9407730102539,40.8822746276855,40.9240226745605,40.9884757995605,41.1026840209961,41.2056159973145,41.2436027526855,41.2367172241211,41.20849609375,41.1930656433105,41.1388168334961,41.253662109375,41.4254379272461,41.4797821044922,41.5055656433105,41.4757347106934,41.4558601379395,41.3726539611816],[42.0810089111328,42.0756340026855,42.0840797424316,42.15966796875,42.1995582580566,42.2274436950684,42.2352066040039,42.0963859558105,42.0810089111328],[45.1193351745605,45.1122093200684,45.1539077758789,45.2061996459961,45.2478523254395,45.2324714660645,45.1785621643066,45.1504859924316,45.1193351745605],[45.3328590393066,45.2693367004395,45.36376953125,45.4654769897461,45.46484375,45.4495620727539,45.4000015258789,45.3328590393066],[44.1169891357422,44.1119117736816,44.1166496276855,44.1015625,43.9495582580566,43.9302215576172,43.927978515625,43.9402313232422,43.98193359375,44.1661605834961,44.3338851928711,44.327392578125,44.2812995910645,44.2297859191895,44.076171875,43.869384765625,43.7645492553711,43.6624984741211,43.5782203674316,43.462890625,43.3025398254395,43.2822265625,43.2797355651855,43.3277816772461,43.3888664245605,43.3962898254395,43.3859405517578,43.3434600830078,43.2913093566895,43.1926765441895,43.1742210388184,43.1802749633789,43.1767082214355,43.135498046875,43.0888671875,43.0316390991211,43.0009269714355,42.9937019348145,42.9469261169434,42.943603515625,42.9844207763672,42.9736328125,42.8813972473145,42.7481460571289,42.5987281799316,42.4188957214355,42.3251457214355,42.2203598022461,42.1510276794434,42.0843276977539,42.037841796875,42.0001945495605,42.022216796875,42.1183586120605,42.2579612731934,42.4717292785645,42.5790519714355,42.5469245910645,42.3421401977539,42.3595695495605,42.5,42.5556182861328,42.5713348388672,42.5695304870605,42.5593757629395,42.4871559143066,42.4351081848145,42.3760757446289,42.3342781066895,42.2933578491211,42.2007331848145,42.1317863464355,42.11865234375,42.1234893798828,42.1163558959961,41.977783203125,41.8480453491211,41.8050765991211,41.7598152160645,41.7374000549316,41.7432594299316,41.7604026794434,41.8155746459961,41.7685546875,41.6721687316895,41.5673828125,41.519287109375,41.4560089111328,41.4232444763184,41.43408203125,41.4737777709961,41.5213394165039,41.576416015625,41.6957550048828,41.80322265625,41.9129409790039,42.0673332214355,42.0995597839355,42.1900367736816,42.278076171875,42.3875961303711,42.4481430053711,42.5817375183105,42.6492156982422,42.6714363098145,42.6847648620605,42.7329597473145,42.7955055236816,42.8668441772461,42.9541015625,43.0499038696289,43.1673355102539,43.2371101379395,43.3031272888184,43.3381843566895,43.3117179870605,43.2149887084961,43.2054672241211,43.2009773254395,43.179931640625,43.18505859375,43.1996574401855,43.2796363830566,43.3814964294434,43.5124969482422,43.6426277160645,43.7486305236816,43.9189949035645,44.0194320678711,44.2636222839355,44.3711891174316,44.4825172424316,44.7163581848145,44.9410629272461,45.0512199401855,45.1559600830078,45.2559547424316,45.3486328125,45.3765640258789,45.4012680053711,45.4188957214355,45.4387702941895,45.4833030700684,45.509521484375,45.4834976196289,45.4379386901855,45.3256340026855,45.125,44.8191871643066,44.6701164245605,44.534912109375,44.3966331481934,44.2775382995605,44.2213401794434,44.1316413879395,44.1169891357422],[35.4958992004395,35.4432601928711,35.3787612915039,35.3025856018066,35.2355461120605,35.1715316772461,35.1099128723145,35.1963386535645,35.2464370727539,35.3466300964355,35.4357414245605,35.5718269348145,35.6617202758789,35.61328125,35.5520515441895,35.5081558227539,35.4755859375,35.4734382629395,35.4718246459961,35.4805641174316,35.4958992004395],[44.8546371459961,44.8264617919922,44.8306159973145,44.9369621276855,45.0447235107422,45.0667953491211,45.0819320678711,45.0582542419434,45.010009765625,44.9491195678711,44.8815422058105,44.8546371459961],[44.9720687866211,44.9585914611816,45.0224113464355,45.075927734375,45.0830078125,44.9984359741211,44.9720687866211],[45.4118156433105,45.4013175964355,45.4281768798828,45.4739761352539,45.5280265808105,45.51806640625,45.4607429504395,45.4118156433105],[49.0857887268066,49.1058616638184,49.1099090576172,49.0975570678711,49.090576171875,49.0497055053711,49.0088386535645,48.9393577575684,48.8607406616211,48.6971702575684,48.6355476379395,48.5286140441895,48.50537109375,48.4862289428711,48.4545402526855,48.4237289428711,48.4080581665039,48.3850593566895,48.3118171691895,48.2505378723145,48.2040061950684,48.0518569946289,47.9156227111816,47.7464866638184,47.5584945678711,47.49365234375,47.3974113464355,47.33837890625,47.254638671875,47.1884765625,47.1007804870605,47.0635261535645,47.0467300415039,47.036376953125,46.9612312316895,46.9092292785645,46.8431663513184,46.8307151794434,46.8643569946289,46.9393539428711,46.9723625183105,46.9749488830566,46.9757804870605,46.9961395263672,46.9947280883789,46.9786148071289,46.9705085754395,46.9978523254395,47.0210456848145,47.043212890625,47.108642578125,47.1865730285645,47.2093772888184,47.1859397888184,47.1414566040039,47.0334968566895,46.9660148620605,46.6244621276855,46.3866691589355,46.15869140625,46.0058097839355,45.8119125366211,45.6715316772461,45.5949211120605,45.563720703125,45.5199203491211,45.4719734191895,45.4425773620605,45.4242668151855,45.3744163513184,45.2931137084961,45.2159690856934,45.1554183959961,45.12548828125,45.1235847473145,45.1624488830566,45.2058601379395,45.2190933227539,45.1948738098145,45.161865234375,45.1608390808105,45.1820793151855,45.2260246276855,45.3108406066895,45.349365234375,45.3108406066895,45.2461929321289,45.1691398620605,45.1292953491211,45.1355476379395,45.1265144348145,45.10498046875,45.0750961303711,45.0339851379395,45.0064468383789,44.944091796875,44.8837890625,44.86083984375,44.8251953125,44.7972145080566,44.8037605285645,44.80810546875,44.7703132629395,44.74609375,44.7146492004395,44.68408203125,44.6769065856934,44.6554222106934,44.6268081665039,44.552001953125,44.4383773803711,44.3265151977539,44.2232894897461,44.1712875366211,44.0972671508789,44.0471687316895,43.9517593383789,43.89208984375,43.6851081848145,43.5641593933105,43.4270515441895,43.3529815673828,43.31005859375,43.2742691040039,43.204345703125,43.1615753173828,43.1189460754395,43.1024894714355,43.1282730102539,43.0857887268066,43.0431137084961,43.0204124450684,42.99560546875,42.9737777709961,42.935546875,42.9117202758789,42.8734855651855,42.8557624816895,42.7972640991211,42.7344703674316,42.66552734375,42.6255378723145,42.5183563232422,42.3994140625,42.274169921875,42.2354011535645,42.2078132629395,42.1900367736816,42.302978515625,42.4131355285645,42.4384765625,42.4566421508789,42.4575691223145,42.4834976196289,42.5472145080566,42.6048355102539,42.666015625,42.759033203125,42.7757339477539,42.7638168334961,42.7667007446289,42.77490234375,42.7908210754395,42.8287124633789,42.8646469116211,42.8714866638184,42.8643531799316,42.8575210571289,42.8952140808105,42.9022445678711,42.9000511169434,42.9047355651855,42.9044456481934,42.9126472473145,42.9706535339355,42.9735832214355,42.9714813232422,42.9288063049316,42.9188957214355,42.9237289428711,42.9285125732422,42.9375,42.9329109191895,42.8304710388184,42.8146018981934,42.8369636535645,42.9043922424316,42.9781723022461,43.0562019348145,43.179443359375,43.2403793334961,43.2052726745605,43.194091796875,43.1886215209961,43.1950187683105,43.132568359375,43.087890625,43.0478973388672,43.0027809143066,42.7030296325684,42.593505859375,42.4090347290039,42.4197731018066,42.4669914245605,42.5041046142578,42.52685546875,42.5611305236816,42.6034660339355,42.6378898620605,42.660400390625,42.677734375,42.7576675415039,42.7606925964355,42.8221664428711,42.8214836120605,42.8188972473145,42.7786636352539,42.7669410705566,42.7831535339355,42.7335472106934,42.6674346923828,42.5865249633789,42.5354499816895,42.4590835571289,42.4193840026855,42.3399925231934,42.293701171875,42.2486801147461,42.2072257995605,42.1941909790039,42.1686515808105,42.1074714660645,42.0547332763672,42.0360336303711,42.0394515991211,42.0802726745605,42.0785636901855,42.0279769897461,41.9459953308105,41.8205070495605,41.7540512084961,41.6973152160645,41.672119140625,41.6290550231934,41.5418930053711,41.4905738830566,41.4602508544922,41.4252433776855,41.366943359375,41.2641143798828,41.2050285339355,41.1238288879395,41.0418930053711,40.9615211486816,40.8762702941895,40.8292961120605,40.76513671875,40.7112808227539,40.6599617004395,40.6226577758789,40.608642578125,40.6194305419922,40.6561050415039,40.7217750549316,40.7540512084961,40.8092765808105,40.860595703125,40.9602546691895,41.0286102294922,41.061279296875,41.0962409973145,41.1300315856934,41.1965827941895,41.1802711486816,41.1639175415039,41.187255859375,41.1771469116211,41.1695327758789,41.162353515625,41.1533203125,41.1423835754395,41.1570816040039,41.1791496276855,41.270751953125,41.3486328125,41.4943351745605,41.6069831848145,41.7412605285645,41.889404296875,41.994873046875,41.9983406066895,42.0011253356934,42.0048828125,42.0887680053711,42.1944808959961,42.3147926330566,42.4727516174316,42.6051750183105,42.7666473388672,42.8733863830566,42.9908180236816,42.95458984375,42.9145011901855,42.876953125,42.9721183776855,43.0645980834961,43.2051734924316,43.310546875,43.3720207214355,43.4175300598145,43.4941902160645,43.5736808776855,43.6490707397461,43.7146949768066,43.6939468383789,43.6529769897461,43.6134757995605,43.5716323852539,43.5511703491211,43.5589332580566,43.5657234191895,43.5778312683105,43.588134765625,43.5986328125,43.6132354736328,43.6279754638672,43.6084976196289,43.5838890075684,43.5580101013184,43.53662109375,43.5095710754395,43.4893531799316,43.4921379089355,43.5772438049316,43.7135734558105,43.7961921691895,43.877197265625,43.9939460754395,44.082275390625,44.1686019897461,44.2482414245605,44.348388671875,44.393798828125,44.455078125,44.5208511352539,44.5866203308105,44.6524391174316,44.7182121276855,44.7839813232422,44.8497581481934,44.9155769348145,44.9813003540039,45.0470733642578,45.1128425598145,45.1786155700684,45.2444343566895,45.3102035522461,45.3759765625,45.4417991638184,45.507568359375,45.5553703308105,45.5429191589355,45.5094223022461,45.474365234375,45.439697265625,45.37744140625,45.3374519348145,45.3036613464355,45.2677230834961,45.2206077575684,45.1810569763184,45.134765625,45.0937995910645,45.059326171875,45.0233879089355,44.9949226379395,44.765380859375,44.5358390808105,44.3063011169434,44.0767593383789,43.8472137451172,43.6176261901855,43.3880882263184,43.1585922241211,42.9290542602539,42.6995124816895,42.469970703125,42.2404289245605,42.0108871459961,41.7813453674316,41.5517578125,41.3222160339355,41.3241195678711,41.310791015625,41.27880859375,41.2627449035645,41.2722663879395,41.2962913513184,41.346923828125,41.4083976745605,41.4581031799316,41.5602531433105,41.6387214660645,41.8100090026855,41.864013671875,41.9190902709961,41.9651832580566,42.0782241821289,42.18017578125,42.2799797058105,42.3041954040527,42.335205078125,42.3358879089355,42.3297882080078,42.296875,42.2582511901855,42.2058601379395,42.1477546691895,42.1307144165039,42.0605964660645,41.9443855285645,41.7803688049316,41.8858909606934,42.04833984375,42.1006355285645,42.2371597290039,42.3308601379395,42.42822265625,42.5556640625,42.7601585388184,42.8054695129395,42.8202629089355,42.824462890625,42.816162109375,42.7998046875,42.8687515258789,42.8797836303711,42.8605461120605,42.8505859375,42.86962890625,42.9104499816895,42.9544410705566,43.0043449401855,43.1040534973145,43.158447265625,43.1705093383789,43.1673812866211,43.1741218566895,43.230712890625,43.3556632995605,43.4208526611328,43.4823760986328,43.5329093933105,43.5767097473145,43.6487770080566,43.7501449584961,43.9585456848145,44.1927719116211,44.22802734375,44.2650871276855,44.2947731018066,44.3254890441895,44.3551254272461,44.406494140625,44.4615211486816,44.5265617370605,44.58154296875,44.6240234375,44.6333503723145,44.6287612915039,44.5304718017578,44.5078125,44.5013656616211,44.5324249267578,44.5412101745605,44.531005859375,44.5775413513184,44.6019515991211,44.599853515625,44.6187515258789,44.708984375,44.8115730285645,44.85400390625,44.9218254089355,44.9803237915039,45.0402336120605,45.1216812133789,45.2297859191895,45.2795906066895,45.3578605651855,45.3428688049316,45.3994598388672,45.3883781433105,45.4046401977539,45.3986320495605,45.343505859375,45.3197250366211,45.3075218200684,45.3319816589355,45.4073753356934,45.4967269897461,45.57275390625,45.779541015625,45.9678688049316,46.191650390625,46.4140625,46.4752922058105,46.5474586486816,46.6083488464355,46.6690444946289,46.7420425415039,46.8560523986816,46.8929214477539,46.9543952941895,46.9571304321289,46.9906730651855,46.9636688232422,46.922119140625,46.894775390625,46.8395004272461,46.82861328125,46.839599609375,46.9019050598145,46.8948745727539,46.9337387084961,47.01806640625,47.0299301147461,47.0973129272461,47.1101570129395,47.0406723022461,46.95166015625,46.938720703125,46.8822746276855,46.873291015625,46.8829116821289,46.8794898986816,46.794921875,46.6964340209961,46.63427734375,46.5956535339355,46.5714836120605,46.5675773620605,46.5452156066895,46.5372543334961,46.5191421508789,46.4855461120605,46.4102058410645,46.4368171691895,46.3856925964355,46.337158203125,46.3488273620605,46.442138671875,46.5079612731934,46.5664520263672,46.5770988464355,46.6056137084961,46.6499481201172,46.6986312866211,46.7343254089355,46.7571296691895,46.7659187316895,46.7586936950684,46.73681640625,46.7103538513184,46.7054214477539,46.7257804870605,46.7746124267578,46.9549293518066,47.1004905700684,47.2623023986816,47.3209953308105,47.4564933776855,47.5899391174316,47.7087898254395,47.7454109191895,47.7606925964355,47.7899894714355,47.8039054870605,47.7686538696289,47.7409172058105,47.79248046875,47.8767585754395,47.9477043151855,48.0201148986816,48.1270027160645,48.2324714660645,48.2844734191895,48.3235855102539,48.4122543334961,48.5738754272461,48.8055686950684,48.9696273803711,49.038330078125,49.0983390808105,49.1502952575684,49.1999015808105,49.2525863647461,49.3038558959961,49.3670921325684,49.5022468566895,49.6969718933105,49.8527336120605,49.9390602111816,50.0008773803711,50.0584983825684,50.1402359008789,50.2174797058105,50.2735366821289,50.318115234375,50.3579597473145,50.4027366638184,50.41357421875,50.3779792785645,50.2823219299316,50.0936050415039,49.9700202941895,49.9319343566895,49.8582534790039,49.8285140991211,49.8747062683105,49.9283218383789,49.96240234375,50.0131340026855,50.099853515625,50.1564445495605,50.2284660339355,50.353759765625,50.5503425598145,50.6126937866211,50.619873046875,50.6068840026855,50.601318359375,50.6445770263672,50.72607421875,50.8517074584961,50.9346656799316,51.0270004272461,51.0835914611816,51.102294921875,51.1318855285645,51.1971702575684,51.2545890808105,51.2895011901855,51.3215789794922,51.3697280883789,51.505615234375,51.5891609191895,51.6751441955566,51.7291984558105,51.7191886901855,51.681640625,51.6474609375,51.594482421875,51.5401840209961,51.4974136352539,51.475341796875,51.4712867736816,51.4820327758789,51.4839859008789,51.5542488098145,51.6727066040039,51.7093734741211,51.6812973022461,51.59423828125,51.5121612548828,51.4816398620605,51.4807624816895,51.4795417785645,51.4981460571289,51.4979019165039,51.4945793151855,51.4669456481934,51.4637184143066,51.4849624633789,51.4936027526855,51.4823722839355,51.4445304870605,51.3995590209961,51.2518081665039,51.2137222290039,51.1611785888672,51.1151847839355,51.040771484375,50.9957008361816,50.9140625,50.7803230285645,50.6739234924316,50.5837898254395,50.5411643981934,50.5357894897461,50.5506858825684,50.5916023254395,50.66015625,50.7810516357422,50.8798828125,50.9112815856934,50.946044921875,50.990234375,51.0115699768066,50.9980964660645,50.9413566589355,50.8697776794434,50.7447280883789,50.665283203125,50.601806640625,50.5828628540039,50.60205078125,50.6537590026855,50.7135276794434,50.7762680053711,50.8446273803711,50.9360847473145,51.01953125,51.0044937133789,50.9808616638184,51.0315895080566,51.0455551147461,51.065185546875,51.0360336303711,50.946533203125,50.8888664245605,50.8955574035645,50.9251441955566,50.98095703125,51.046875,51.0890121459961,51.0916481018066,51.06884765625,51.072265625,51.0817375183105,51.0638198852539,50.9710464477539,50.8683128356934,50.7372055053711,50.6944313049316,50.6761245727539,50.668212890625,50.6479034423828,50.6204109191895,50.6042976379395,50.5828132629395,50.5110816955566,50.4928703308105,50.5439453125,50.58203125,50.690185546875,50.7992706298828,50.8396949768066,50.8502922058105,50.8341827392578,50.769775390625,50.7041511535645,50.6791496276855,50.669189453125,50.6637191772461,50.6955070495605,50.7748031616211,50.8610343933105,50.990234375,51.1370124816895,51.2296867370605,51.3246078491211,51.4147491455078,51.44189453125,51.4923820495605,51.5287132263184,51.5370597839355,51.616943359375,51.6511726379395,51.7039070129395,51.7729988098145,51.8346214294434,51.890625,51.9332504272461,51.9764671325684,52.0245132446289,52.1255874633789,52.1463394165039,52.1508293151855,52.2333984375,52.2805671691895,52.3368644714355,52.394775390625,52.56982421875,52.67578125,52.7447242736816,52.8194351196289,52.8601570129395,52.933349609375,52.9724617004395,52.9890632629395,52.9959945678711,52.978515625,52.9693832397461,52.9559097290039,52.9437484741211,52.9661140441895,53.0054206848145,53.0574226379395,53.1078643798828,53.1739234924316,53.2284660339355,53.2224578857422,53.2394065856934,53.2751960754395,53.2871589660645,53.3367652893066,53.4062004089355,53.4458999633789,53.4657249450684,53.455810546875,53.4846687316895,53.5015602111816,53.5232925415039,53.5544929504395,53.5802726745605,53.5870628356934,53.565185546875,53.5509796142578,53.5831031799316,53.6217269897461,53.6574249267578,53.71044921875,53.7534675598145,53.81298828125,53.8824691772461,53.9638175964355,54.0194816589355,54.0492668151855,53.9949226379395,53.9464836120605,53.9543914794922,53.9799308776855,54.0026397705078,54.01318359375,54.04443359375,54.0692863464355,54.105224609375,54.1392593383789,54.1710433959961,54.1704597473145,54.1832046508789,54.221923828125,54.2432136535645,54.2450218200684,54.2364768981934,54.26708984375,54.2797355651855,54.3029289245605,54.347412109375,54.3841781616211,54.3621597290039,54.3522453308105,54.3685569763184,54.3966293334961,54.3687515258789,54.3401870727539,54.3644065856934,54.4411125183105,54.5160636901855,54.5515632629395,54.564453125,54.5933074951172,54.623291015625,54.6187019348145,54.6933097839355,54.6595230102539,54.6673812866211,54.7154312133789,54.7378921508789,54.7881851196289,54.8288078308105,54.8544921875,54.8724098205566,54.9435539245605,54.9537124633789,54.9595718383789,54.9767074584961,55.0030288696289,55.0524406433105,55.115234375,55.1609382629395,55.1865234375,55.1944351196289,55.204833984375,55.3084945678711,55.3583488464355,55.3895988464355,55.3725128173828,55.3568840026855,55.307373046875,55.2456550598145,55.1990737915039,55.1767578125,55.1624526977539,55.18359375,55.2122535705566,55.253173828125,55.2823715209961,55.30517578125,55.2611351013184,55.1279792785645,54.9504890441895,54.7150421142578,54.5993156433105,54.5386238098145,54.4554214477539,54.3640632629395,54.260498046875,54.2122077941895,54.1583480834961,54.1780281066895,54.2214851379395,54.2056655883789,54.2316398620605,54.3084449768066,54.3256340026855,54.2721214294434,54.1814460754395,54.123046875,54.0536613464355,53.9644546508789,53.9418449401855,53.9757804870605,53.9959487915039,54.0230484008789,54.0564956665039,54.0904312133789,54.12158203125,54.1343269348145,54.12451171875,54.1073226928711,53.9807624816895,53.9578132629395,53.9556159973145,53.9628410339355,53.9993133544922,54.0449714660645,54.0673828125,54.0634765625,54.0423812866211,53.9961891174316,53.929443359375,53.8683128356934,53.8114738464355,53.7072257995605,53.598388671875,53.5431632995605,53.5062026977539,53.4543952941895,53.4475593566895,53.4688949584961,53.5762710571289,53.602783203125,53.6197280883789,53.6114234924316,53.5764656066895,53.5277366638184,53.4876441955566,53.5044441223145,53.5507316589355,53.6037101745605,53.6472663879395,53.75439453125,53.82568359375,53.8340339660645,53.8192367553711,53.8267097473145,53.893798828125,53.9701156616211,54.021728515625,54.0685043334961,54.0896492004395,54.1060066223145,54.1147956848145,54.16796875,54.258544921875,54.3119621276855,54.335693359375,54.35107421875,54.3871078491211,54.4368629455566,54.4423828125,54.3218727111816,54.1824684143066,54.145263671875,54.1515121459961,54.113525390625,54.0552749633789,54.0225563049316,53.9932136535645,53.9425277709961,53.8226547241211,53.6701164245605,53.498779296875,53.379150390625,53.3174285888672,53.2691879272461,53.094970703125,52.9296875,52.638427734375,52.3570289611816,52.0474090576172,51.8681144714355,51.4931144714355,51.3779754638672,51.1600074768066,50.9554672241211,50.7745590209961,50.7582015991211,50.8072738647461,50.8399887084961,50.8583488464355,50.9117698669434,50.9246101379395,50.9190902709961,50.9462890625,50.9976081848145,51.0926246643066,51.1363754272461,51.183349609375,51.2017593383789,51.2165985107422,51.2242202758789,51.27734375,51.2934074401855,51.2834968566895,51.2814445495605,51.2427749633789,51.1897964477539,51.1856956481934,51.1910667419434,51.1465797424316,51.0723648071289,51.0149421691895,50.96875,50.9462890625,50.9664077758789,50.9564933776855,50.9097633361816,50.8710441589355,50.8236808776855,50.7711410522461,50.7398414611816,50.7391128540039,50.764404296875,50.766357421875,50.7108383178711,50.7194328308105,50.7418937683105,50.7275886535645,50.771484375,50.8263168334961,50.8694839477539,50.8933601379395,50.8931159973145,50.8972663879395,50.9605941772461,50.9892082214355,50.9945793151855,50.9945793151855,50.9357414245605,50.8871612548828,50.8180160522461,50.774658203125,50.6768531799316,50.604736328125,50.5205574035645,50.4374504089355,50.2882308959961,50.2391586303711,50.2391586303711,50.21875,50.202392578125,50.09130859375,50.0879859924316,50.0614280700684,50.0103034973145,49.9510765075684,49.8941383361816,49.8216285705566,49.750732421875,49.6648445129395,49.6158180236816,49.5994644165039,49.6239280700684,49.6053733825684,49.5565452575684,49.5504417419434,49.4993171691895,49.50341796875,49.5054664611816,49.4993171691895,49.5463409423828,49.5875015258789,49.6384735107422,49.707763671875,49.7691383361816,49.7772941589355,49.7486801147461,49.695556640625,49.6566925048828,49.6097145080566,49.5626945495605,49.55859375,49.4878883361816,49.3220710754395,49.2873039245605,49.2545890808105,49.2397918701172,49.2161598205566,49.1476593017578,49.0857887268066],[-2.15839838981628,-2.16728496551514,-2.10976576805115,-2.03652358055115,-2.05302739143372,-2.06982445716858,-2.08505868911743,-2.10009765625,-2.15839838981628],[3.97773432731628,3.80161118507385,3.59047842025757,3.20166015625,2.99706983566284,2.84243178367615,2.81464838981628,2.642333984375,2.220947265625,1.378173828125,0.535400390625,-0.307324260473251,-0.728711247444153,-0.870312571525574,-1.04746079444885,-1.22050786018372,-1.44951164722443,-1.57226550579071,-1.61318385601044,-1.69531214237213,-1.86699211597443,-1.94501936435699,-1.98232388496399,-1.97519528865814,-1.95058608055115,-1.99179673194885,-2.05595707893372,-2.04248046875,-2.02353501319885,-2.13749980926514,-2.19375014305115,-2.26992177963257,-2.33632826805115,-2.39238309860229,-2.53945326805115,-2.55566382408142,-2.62861347198486,-2.68837857246399,-2.81904292106628,-3.01923847198486,-3.17333984375,-3.25058603286743,-3.35068345069885,-3.44248032569885,-3.53584003448486,-3.57675766944885,-3.78603529930115,-3.91308569908142,-3.95517611503601,-3.99326157569885,-4.06787061691284,-4.119140625,-4.15283203125,-4.47841787338257,-4.62548828125,-4.60859346389771,-4.66552734375,-4.6923828125,-4.67724561691284,-4.62353515625,-4.51298809051514,-4.40244150161743,-4.29189443588257,-4.18144512176514,-4.07089805603027,-3.96035122871399,-3.84980487823486,-3.7392578125,-3.67441391944885,-3.63613271713257,-3.55976557731628,-3.54082012176514,-3.51679658889771,-3.51152372360229,-3.4970703125,-3.46025371551514,-3.40722632408142,-3.30576181411743,-3.24619150161743,-3.17841815948486,-3.07001948356628,-3.04540991783142,-2.98857426643372,-2.86962890625,-2.75058579444885,-2.63164043426514,-2.51259756088257,-2.3935546875,-2.27460932731628,-2.15566420555115,-2.03662085533142,-1.91757798194885,-1.79863250255585,-1.67958974838257,-1.56054675579071,-1.44160139560699,-1.32255840301514,-1.20361304283142,-1.08457016944885,-1.03984391689301,-1.00205111503601,-1.00205111503601,-0.831640779972076,-0.397851586341858,-0.0169922094792128,0.173779249191284,0.294531255960464,0.382470518350601,0.50512707233429,0.605175793170929,0.686425805091858,0.731249988079071,0.867285072803497,1.04213857650757,1.10156238079071,1.15644538402557,1.18530297279358,1.21425795555115,1.23071277141571,1.24453127384186,1.27285158634186,1.38115262985229,1.41669952869415,1.48901355266571,1.55649423599243,1.59926736354828,1.64335930347443,1.71962916851044,1.77363264560699,1.86191403865814,2.06240224838257,2.23017597198486,2.41791963577271,2.47968745231628,2.58969688415527,2.59575223922729,2.61982417106628,2.72343730926514,2.818115234375,2.84194350242615,2.92475605010986,3.11997056007385,3.16347646713257,3.35751938819885,3.41269516944885,3.60625028610229,3.65058588981628,3.69150424003601,3.73315382003784,3.81298804283142,3.84086894989014,3.86977505683899,3.88916015625,4.22021484375,4.41909170150757,4.62065410614014,4.87548828125,5.10957050323486,5.31186532974243,5.49228525161743,5.45791006088257,5.41206073760986,5.38408231735229,5.36489248275757,5.38515615463257,5.41328144073486,5.41909170150757,5.34399366378784,5.278564453125,5.20810556411743,5.15693378448486,5.10556602478027,4.95048809051514,4.80800771713257,4.70263624191284,4.61982393264771,4.50380897521973,4.46811532974243,4.44970750808716,4.44472646713257,4.43725538253784,4.43012666702271,4.42734384536743,4.41147470474243,4.25454092025757,4.11083984375,3.98593735694885,3.86464834213257,3.74672842025757,3.64882802963257,3.61899399757385,3.60483384132385,3.60009789466858,3.55898475646973,3.52060556411743,3.50087881088257,3.47875952720642,3.45610356330872,3.46918940544128,3.57783222198486,3.75424814224243,3.85146474838257,3.94794917106628,4.08271503448486,4.17763710021973,4.27304697036743,4.19033193588257,4.05747032165527,3.99194383621216,3.96298813819885,3.94355440139771,3.94306659698486,3.94619131088257,3.96328139305115,3.97905254364014,3.97773432731628],[42.1900367736816,42.1298332214355,42.059814453125,42.04345703125,42.0324211120605,42.0149917602539,41.9957542419434,41.8988761901855,41.8209953308105,41.7810554504395,41.7828102111816,41.7191390991211,41.56005859375,41.4595680236816,41.4175300598145,41.3716316223145,41.3251953125,41.2814445495605,41.0756340026855,41.0506820678711,41.0556144714355,41.0243148803711,40.9927749633789,41.0143547058105,41.0107421875,41.0391616821289,41.024169921875,40.9823226928711,40.818115234375,40.7796363830566,40.7422370910645,40.662353515625,40.577880859375,40.51123046875,40.4495124816895,40.3897933959961,40.3522453308105,40.4307632446289,40.4084014892578,40.37646484375,40.3875465393066,40.3714332580566,40.30322265625,40.3058090209961,40.3292465209961,40.5166015625,40.6053199768066,40.6251983642578,40.6275405883789,40.4802742004395,40.4541015625,40.4495124816895,40.4935073852539,40.4826164245605,40.4587898254395,40.4285163879395,40.344970703125,40.3285179138184,40.3298835754395,40.3105964660645,40.2721672058105,40.13720703125,40.092041015625,40.0743179321289,40.0593719482422,40.0431137084961,39.9788093566895,39.8779296875,39.8286628723145,39.8001480102539,39.7628402709961,39.7145500183105,39.6064949035645,39.5785140991211,39.5333023071289,39.4889640808105,39.4622535705566,39.4488792419434,39.4576187133789,39.4605941772461,39.4427223205566,39.412353515625,39.3745613098145,39.3619155883789,39.3570785522461,39.3604011535645,39.385986328125,39.377197265625,39.3573722839355,39.3368644714355,39.2737312316895,39.2156753540039,39.20751953125,39.2607421875,39.3106460571289,39.3521499633789,39.3509254455566,39.2755851745605,39.2779808044434,39.3065910339355,39.3777351379395,39.4229965209961,39.4470710754395,39.4530754089355,39.4788093566895,39.51708984375,39.5538558959961,39.5821800231934,39.6036643981934,39.5978507995605,39.5686988830566,39.5353050231934,39.5198211669922,39.5135726928711,39.493408203125,39.411865234375,39.3947296142578,39.4132804870605,39.4712867736816,39.5644035339355,39.5758781433105,39.5873527526855,39.5818862915039,39.5426292419434,39.5750007629395,39.5841789245605,39.5575714111328,39.5605964660645,39.5530738830566,39.5567359924316,39.5749015808105,39.5737762451172,39.5320777893066,39.532470703125,39.5248069763184,39.6658706665039,39.7610855102539,39.8270988464355,39.9177742004395,39.9685554504395,39.9470710754395,39.9097633361816,39.9197273254395,39.950439453125,39.9906234741211,40.020751953125,40.0603485107422,40.0973167419434,40.1580085754395,40.2022476196289,40.1727561950684,40.1048355102539,40.0698699951172,40.0492172241211,39.9499015808105,39.9544944763184,39.9745101928711,39.9989738464355,40.0834465026855,40.1311531066895,40.1875953674316,40.2206535339355,40.2388687133789,40.2548828125,40.271240234375,40.2869148254395,40.2751922607422,40.2419929504395,40.208984375,40.2102508544922,40.2171363830566,40.2080078125,40.1786117553711,40.15234375,40.1880378723145,40.2343254089355,40.240966796875,40.2585945129395,40.2895050048828,40.3407211303711,40.4386215209961,40.4544410705566,40.4543952941895,40.4386215209961,40.4242172241211,40.4016571044922,40.4273910522461,40.4630851745605,40.5197257995605,40.54345703125,40.5651359558105,40.5780754089355,40.5243644714355,40.5254402160645,40.5556144714355,40.6086921691895,40.650390625,40.7860336303711,40.8106422424316,40.8285140991211,40.860107421875,40.8424301147461,40.8423347473145,40.8621559143066,40.8699226379395,40.8837890625,40.981689453125,41.0189437866211,41.04345703125,41.0399436950684,41.0142593383789,41.0259284973145,41.0668487548828,41.1184577941895,41.1737289428711,41.1865730285645,41.1647491455078,41.1870613098145,41.1950187683105,41.3113784790039,41.3610382080078,41.4131355285645,41.4280281066895,41.4540023803711,41.5155754089355,41.5330085754395,41.5412101745605,41.5341796875,41.5032730102539,41.4354476928711,41.367431640625,41.333251953125,41.3080558776855,41.3074722290039,41.3418960571289,41.1360321044922,41.1233863830566,41.1524925231934,41.1399421691895,41.1526374816895,41.1865730285645,41.1959953308105,41.2248992919922,41.2625007629395,41.4005355834961,41.4498062133789,41.4603500366211,41.4126472473145,41.4495582580566,41.496826171875,41.5144538879395,41.5399894714355,41.5714378356934,41.7250480651855,41.830810546875,41.87548828125,41.9052238464355,41.9226570129395,42.0196266174316,42.0308113098145,42.0379867553711,42.0777816772461,42.1628913879395,42.186279296875,42.2064476013184,42.25,42.28466796875,42.2665519714355,42.2486801147461,42.293701171875,42.3399925231934,42.4193840026855,42.4590835571289,42.5354499816895,42.5865249633789,42.6674346923828,42.7335472106934,42.7831535339355,42.7669410705566,42.7786636352539,42.8188972473145,42.8214836120605,42.8221664428711,42.7606925964355,42.7576675415039,42.677734375,42.660400390625,42.6378898620605,42.6034660339355,42.5611305236816,42.52685546875,42.5041046142578,42.4669914245605,42.4197731018066,42.4090347290039,42.593505859375,42.7030296325684,43.0027809143066,43.0478973388672,43.087890625,43.132568359375,43.1950187683105,43.1886215209961,43.194091796875,43.2052726745605,43.2403793334961,43.179443359375,43.0562019348145,42.9781723022461,42.9043922424316,42.8369636535645,42.8146018981934,42.8304710388184,42.9329109191895,42.9375,42.9285125732422,42.9237289428711,42.9188957214355,42.9288063049316,42.9714813232422,42.9735832214355,42.9706535339355,42.9126472473145,42.9044456481934,42.9047355651855,42.9000511169434,42.9022445678711,42.8952140808105,42.8575210571289,42.8643531799316,42.8714866638184,42.8646469116211,42.8287124633789,42.7908210754395,42.77490234375,42.7667007446289,42.7638168334961,42.7757339477539,42.759033203125,42.666015625,42.6048355102539,42.5472145080566,42.4834976196289,42.4575691223145,42.4566421508789,42.4384765625,42.4131355285645,42.302978515625,42.1900367736816],[39.8252944946289,39.8458480834961,39.85546875,39.8666000366211,39.8824195861816,39.8827133178711,39.863037109375,39.8281745910645,39.7909164428711,39.7867660522461,39.8252944946289],[39.9502449035645,39.9953117370605,39.9932594299316,39.98095703125,39.9686508178711,39.9460945129395,39.9174308776855,39.9071273803711,39.9502449035645],[39.892578125,39.9067878723145,39.9798316955566,40.0481452941895,40.0596656799316,40.0798797607422,40.149169921875,40.1522941589355,40.1332511901855,40.0879898071289,40.0388641357422,40.0057601928711,39.9949188232422,39.9925308227539,39.976318359375,39.8951187133789,39.8849143981934,39.8834495544434,39.892578125],[10.718505859375,10.6796875,10.7595701217651,10.781982421875,10.718505859375],[11.2850589752197,11.2754888534546,11.2757816314697,11.2904300689697,11.3480958938599,11.38330078125,11.3899412155151,11.2850589752197],[10.4112300872803,10.541259765625,10.5866203308105,10.6439456939697,10.6551265716553,10.58056640625,10.5089359283447,10.5521974563599,10.6046390533447,10.668701171875,10.7210445404053,10.7585935592651,10.89013671875,11.0586919784546,11.1077632904053,11.1466798782349,11.083984375,10.9767580032349,10.9215822219849,10.9092769622803,10.9137210845947,11.0737800598145,11.2110843658447,11.3677740097046,11.4606437683105,11.5886716842651,11.7105960845947,11.7734861373901,11.74169921875,11.7066898345947,11.7320804595947,12.0897951126099,12.2556638717651,12.3833980560303,12.42626953125,12.4935054779053,12.5699214935303,12.6699705123901,12.8283205032349,13.0150394439697,13.0779781341553,13.1929693222046,13.2882328033447,13.5399904251099,13.560302734375,13.5675783157349,13.585693359375,13.6263675689697,13.6599607467651,13.7169437408447,13.8418941497803,13.9724607467651,14.0548820495605,14.13671875,14.2525386810303,14.3326177597046,14.351318359375,14.3786134719849,14.4174318313599,14.4210939407349,14.3741693496704,14.3621578216553,14.36279296875,14.35791015625,14.3627433776855,14.3955078125,14.3695793151855,14.3900384902954,14.4278316497803,14.4040031433105,14.3661127090454,14.2894535064697,14.2544431686401,14.2273921966553,14.2274417877197,14.2809572219849,14.3360834121704,14.3462400436401,14.319091796875,14.2593746185303,14.2005367279053,14.1614742279053,14.1095705032349,14.1070804595947,14.1561527252197,14.0849609375,14.049072265625,13.9766120910645,13.9245119094849,13.9211912155151,14.0491209030151,14.1270999908447,14.2628908157349,14.3430185317993,14.3571767807007,14.3723640441895,14.3881349563599,14.4762201309204,14.4662113189697,14.4547853469849,14.5150384902954,14.5782222747803,14.5494146347046,14.5050783157349,14.4793949127197,14.4413080215454,14.3877439498901,14.3351068496704,14.314697265625,14.3293933868408,14.3273429870605,14.3910160064697,14.4256839752197,14.415771484375,14.4166984558105,14.4979009628296,14.5721197128296,14.5923833847046,14.5553226470947,14.5628900527954,14.6649894714355,14.705078125,14.6499519348145,14.5457515716553,14.4512205123901,14.3687009811401,14.307861328125,14.1266117095947,14.0688972473145,14.0194826126099,13.9930181503296,13.8156251907349,13.6541986465454,13.5216789245605,13.4377927780151,13.2254400253296,13.0303707122803,12.9331045150757,12.8357419967651,12.7059078216553,12.5399904251099,12.4317865371704,12.3645505905151,12.2957029342651,12.260498046875,12.3190431594849,12.3215827941895,12.3040046691895,12.2770509719849,12.1758794784546,12.0774908065796,12.0523433685303,11.9792966842651,11.9691896438599,11.9655275344849,11.9484376907349,11.9117183685303,11.7383785247803,11.6978025436401,11.6870107650757,11.6818351745605,11.7083492279053,11.7512693405151,11.7580080032349,11.6824712753296,11.6529293060303,11.6483879089355,11.635009765625,11.601318359375,11.5591306686401,11.487060546875,11.3724126815796,11.2942876815796,11.2448244094849,11.0786628723145,11.037109375,11.0123043060303,10.9219732284546,10.794921875,10.7972650527954,10.851806640625,10.8851566314697,10.8584966659546,10.8635749816895,10.9260740280151,10.989990234375,10.9940433502197,10.9688959121704,10.951416015625,10.9516115188599,10.9400873184204,10.8451652526855,10.8614740371704,10.8975591659546,10.911376953125,10.8868646621704,10.8093748092651,10.7337894439697,10.7016611099243,10.6619138717651,10.5902347564697,10.5344724655151,10.5208005905151,10.5232419967651,10.5159664154053,10.4633302688599,10.42236328125,10.4112300872803],[-11.4410152435303,-11.4568367004395,-11.4512701034546,-11.43115234375,-11.4092769622803,-11.3917970657349,-11.3926753997803,-11.4143552780151,-11.4410152435303],[-5.62666034698486,-5.6318359375,-5.63164043426514,-5.61855459213257,-5.60761690139771,-5.60751962661743,-5.60947322845459,-5.611328125,-5.619140625,-5.62666034698486],[-4.67509794235229,-4.68837881088257,-4.69472646713257,-4.69365262985229,-4.68964862823486,-4.681640625,-4.68681669235229,-4.68564414978027,-4.67402362823486,-4.66533184051514,-4.66171836853027,-4.65732431411743,-4.65947246551514,-4.67509794235229],[-4.51113271713257,-4.51796865463257,-4.51601552963257,-4.51484394073486,-4.52167987823486,-4.52441358566284,-4.50703096389771,-4.49394512176514,-4.49169921875,-4.49541044235229,-4.49951124191284,-4.49951124191284,-4.50263690948486,-4.51113271713257],[-4.46347665786743,-4.46806621551514,-4.46650409698486,-4.45976495742798,-4.44921875,-4.44160175323486,-4.44414091110229,-4.45371103286743,-4.46347665786743],[-4.08798837661743,-4.09384775161743,-4.08584022521973,-4.07109355926514,-4.05488300323486,-4.04804706573486,-4.03857421875,-4.03105449676514,-4.04160118103027,-4.05595684051514,-4.08798837661743],[-3.13544917106628,-3.14326190948486,-3.13691425323486,-3.12509751319885,-3.11503934860229,-3.12041020393372,-3.13544917106628],[-2.76640629768372,-2.78554654121399,-2.81123042106628,-2.84697294235229,-2.85585951805115,-2.85556650161743,-2.84443354606628,-2.82978534698486,-2.82568335533142,-2.82226586341858,-2.82451176643372,-2.83984351158142,-2.84667944908142,-2.82919907569885,-2.79853487014771,-2.78798866271973,-2.77910161018372,-2.77314424514771,-2.77861309051514,-2.78134727478027,-2.77412128448486,-2.76787066459656,-2.76142573356628,-2.76640629768372],[-1.21191394329071,-1.26337885856628,-1.25644540786743,-1.23642575740814,-1.18457019329071,-1.13369154930115,-1.14736342430115,-1.18710947036743,-1.21191394329071],[-0.873730301856995,-0.875976800918579,-0.865625202655792,-0.852636873722076,-0.846875131130219,-0.856542706489563,-0.873730301856995],[-0.801757574081421,-0.829003989696503,-0.804198980331421,-0.773632824420929,-0.647070467472076,-0.629784882068634,-0.591796696186066,-0.591796696186066,-0.626562595367432,-0.642187714576721,-0.725683689117432,-0.801757574081421],[1.01313483715057,0.973437488079071,0.946240305900574,0.91523414850235,0.835449278354645,0.842773377895355,0.914745986461639,0.962695121765137,0.999072253704071,0.990966618061066,0.990966618061066,1.02509772777557,1.02509772777557,1.01313483715057],[1.34209001064301,1.33837902545929,1.35874021053314,1.38134741783142,1.38754904270172,1.37514626979828,1.36337888240814,1.35751986503601,1.35708034038544,1.34633791446686,1.34209001064301],[1.71738255023956,1.71308588981628,1.72749006748199,1.747314453125,1.80932629108429,1.80439472198486,1.77875947952271,1.74155282974243,1.71738255023956],[1.84570324420929,1.82255876064301,1.88540005683899,1.92592763900757,1.94370150566101,1.93251967430115,1.91269516944885,1.89697277545929,1.84570324420929],[1.85556626319885,1.73984360694885,1.73173797130585,1.78754866123199,1.90205085277557,1.92685532569885,1.88569355010986,1.84726583957672,1.92768561840057,1.94609355926514,2.029296875,2.02504873275757,1.96855461597443,1.85556626319885],[3.05122089385986,3.01254916191101,3.03305625915527,3.03388667106628,3.07109379768372,3.073974609375,3.05351543426514,3.05122089385986],[3.12919926643372,3.09589838981628,3.10126948356628,3.14877963066101,3.14291977882385,3.12919926643372],[3.92353510856628,3.83920931816101,3.79658246040344,3.80048799514771,3.81533217430115,3.82265615463257,3.83837890625,3.86318349838257,3.88051772117615,3.88051772117615,3.84663057327271,3.87324237823486,3.89956021308899,3.9169921875,3.92353510856628],[17.1218757629395,17.1005859375,17.1295909881592,17.1991214752197,17.2010250091553,17.1701164245605,17.1218757629395],[17.239990234375,17.2244129180908,17.2860355377197,17.3028335571289,17.3392581939697,17.3470706939697,17.3653316497803,17.3864250183105,17.402587890625,17.353271484375,17.2909183502197,17.2623043060303,17.239990234375],[33.2236328125,33.2015113830566,33.21484375,33.2252426147461,33.2825698852539,33.31201171875,33.3680648803711,33.4603996276855,33.54931640625,33.55322265625,33.51513671875,33.44384765625,33.3823738098145,33.3411636352539,33.2716789245605,33.2383308410645,33.2236328125],[34.343994140625,34.2964363098145,34.305419921875,34.3511199951172,34.3903312683105,34.3958969116211,34.343994140625],[34.3705101013184,34.3551750183105,34.3895988464355,34.3987312316895,34.4439430236816,34.53271484375,34.5633277893066,34.544921875,34.4979515075684,34.4264183044434,34.3705101013184],[34.6150398254395,34.5840835571289,34.6309051513672,34.68212890625,34.6150398254395],[34.7311515808105,34.7069816589355,34.7142066955566,34.7830543518066,34.8375473022461,34.8674812316895,34.9148445129395,34.8296852111816,34.7311515808105],[34.8058586120605,34.7080574035645,34.7032203674316,34.7662620544434,34.7354965209961,34.7349624633789,34.8133316040039,34.87451171875,34.9209938049316,34.8930168151855,34.8786125183105,34.8058586120605],[34.7985382080078,34.7368659973145,34.819580078125,34.8652839660645,34.9322738647461,35.0087890625,35.0135765075684,34.7985382080078],[36.4925765991211,36.4278831481934,36.4705581665039,36.612548828125,36.5711402893066,36.4925765991211],[37.4784660339355,37.44873046875,37.4784660339355,37.5099143981934,37.5372085571289,37.5537109375,37.5297393798828,37.4784660339355],[37.73681640625,37.6046905517578,37.6103515625,37.6236343383789,37.6494140625,37.7720222473145,37.8226585388184,37.7825660705566,37.73681640625],[38.6234359741211,38.1760749816895,37.8870582580566,37.6776351928711,37.2745628356934,37.0590324401855,36.925537109375,36.7418937683105,36.6366233825684,36.470703125,36.385498046875,36.3227081298828,36.2021484375,36.1376457214355,36.0521507263184,36.018798828125,36.0064468383789,36.03759765625,36.050537109375,35.9476585388184,35.6874046325684,35.4978523254395,35.332763671875,35.1818351745605,35.1227035522461,35.1015129089355,35.0938987731934,35.1195793151855,35.1009788513184,35.0694313049316,35.0156745910645,34.9320793151855,34.870361328125,34.8750953674316,34.9109840393066,34.9158706665039,34.93359375,35.02197265625,35.0186996459961,34.96630859375,34.9546890258789,34.9263687133789,34.8896942138672,34.8434066772461,34.7825660705566,34.7210464477539,34.6902351379395,34.7659187316895,34.840087890625,34.8442878723145,34.8230934143066,34.7430191040039,34.6884765625,34.625244140625,34.5525398254395,34.5006332397461,34.4632797241211,34.546142578125,34.6050300598145,34.6616706848145,34.7203598022461,34.755126953125,34.6068840026855,34.4388656616211,34.4510765075684,34.4942855834961,34.5118675231934,34.4035148620605,34.3175315856934,34.3142585754395,34.3506317138672,34.4283676147461,34.4939460754395,34.5896492004395,34.6732444763184,34.719970703125,34.694580078125,34.6978988647461,34.737548828125,34.75634765625,34.7786598205566,34.8243637084961,34.8367691040039,34.8103523254395,34.8233871459961,34.9328117370605,35.0451164245605,35.1541519165039,35.2168960571289,35.3144035339355,35.4556159973145,35.5012702941895,35.5344734191895,35.5709953308105,35.5897445678711,35.6063461303711,35.6470718383789,35.6693344116211,35.7142105102539,35.7688484191895,35.8719711303711,35.8978996276855,35.9224128723145,35.9745101928711,36.01416015625,36.0379409790039,36.1050300598145,36.1661605834961,36.23583984375,36.3412094116211,36.4296875,36.4776344299316,36.5856437683105,36.6937980651855,36.6780281066895,36.6511726379395,36.6892585754395,36.6916007995605,36.7719268798828,36.8709449768066,36.9582023620605,36.9690437316895,37.0074691772461,37.0195808410645,36.9603538513184,36.9484367370605,36.8460922241211,36.824462890625,36.862060546875,36.9061508178711,36.9394035339355,36.9576187133789,36.9757308959961,37.1027336120605,37.158203125,37.1935577392578,37.294921875,37.4106941223145,37.4471206665039,37.5511703491211,37.617431640625,37.6537590026855,37.7165031433105,37.7447242736816,37.7554702758789,37.7818374633789,37.8007316589355,37.8279304504395,37.9171867370605,37.9789543151855,38.1060562133789,38.1755867004395,38.2405281066895,38.2838859558105,38.3045425415039,38.3132820129395,38.3049812316895,38.3125,38.3192405700684,38.3077125549316,38.3004379272461,38.3085441589355,38.3273429870605,38.3593254089355,38.4169921875,38.5237808227539,38.6078643798828,38.6234359741211],[42.2475090026855,42.2476577758789,42.2198257446289,42.1871604919434,42.1300773620605,42.1003913879395,42.0995101928711,42.1289558410645,42.1749992370605,42.1712875366211,42.071044921875,42.0183601379395,41.904296875,41.87353515625,41.8538093566895,41.8662109375,41.8736801147461,41.9174308776855,42.0130844116211,42.1714820861816,42.223388671875,42.2749519348145,42.3087921142578,42.3389663696289,42.4258766174316,42.524658203125,42.5472679138184,42.5570831298828,42.6315460205078,42.6858406066895,42.7320289611816,42.7600593566895,42.759765625,42.754638671875,42.798828125,42.8279304504395,42.85791015625,42.8790550231934,42.924560546875,42.9530258178711,43.0341796875,43.0709457397461,43.099853515625,43.13037109375,43.1784172058105,43.1986312866211,43.2263679504395,43.2585945129395,43.2610816955566,43.2379379272461,43.2139663696289,43.1734390258789,43.1516609191895,43.1160163879395,43.0916976928711,43.0430183410645,42.9561996459961,42.9132308959961,42.8747062683105,42.83154296875,42.7514190673828,42.6814918518066,42.6819801330566,42.6698226928711,42.6515121459961,42.595458984375,42.4232444763184,42.387451171875,42.3500022888184,42.3284187316895,42.2808113098145,42.2475090026855],[29.6243171691895,29.6019554138184,29.6114253997803,29.665283203125,29.7989273071289,29.8487796783447,29.8702144622803,29.8863296508789,29.8965816497803,29.9595699310303,29.9788570404053,29.9363269805908,29.7826633453369,29.719970703125,29.6947250366211,29.6243171691895],[28.5429191589355,28.54052734375,28.5375003814697,28.5354499816895,28.5331554412842,28.6279773712158,28.7315425872803,28.8378410339355,28.9895496368408,29.0261726379395,29.0456523895264,29.0746097564697,29.0962409973145,29.2596664428711,29.3474617004395,29.5375022888184,29.6728515625,29.8229961395264,29.9399909973145,29.9613285064697,30.0009784698486,30.04150390625,30.0796890258789,30.0964832305908,30.0973148345947,30.0956058502197,30.0766124725342,29.9828128814697,29.9458980560303,29.8357887268066,29.7155742645264,29.6181144714355,29.5724620819092,29.5791015625,29.5975112915039,29.6166973114014,29.4874019622803,29.4169445037842,29.3930187225342,29.36572265625,29.3666038513184,29.3855476379395,29.3553714752197,29.2754878997803,29.2107410430908,28.9793930053711,28.9012699127197,28.7632827758789,28.6918449401855,28.631591796875,28.5429191589355],[22.3791980743408,22.2840328216553,22.1781749725342,22.0271492004395,21.957763671875,21.904296875,21.8517570495605,21.7713375091553,21.71142578125,21.676025390625,21.662109375,21.6779289245605,21.7096672058105,21.7979488372803,21.807373046875,21.7347640991211,21.7222652435303,21.7129383087158,21.6813468933105,21.626220703125,21.5697746276855,21.50634765625,21.43994140625,21.3375015258789,21.2659187316895,21.2025871276855,20.8916492462158,20.8406257629395,20.7798347473145,20.7378425598145,20.6970691680908,20.7169437408447,20.8095226287842,20.8614253997803,20.9412117004395,20.9455070495605,20.9139652252197,20.8210945129395,20.7337398529053,20.68798828125,20.6466808319092,20.6002445220947,20.5548820495605,20.5295906066895,20.4857425689697,20.44140625,20.4247550964355,20.4136714935303,20.37451171875,20.3285160064697,20.2890129089355,20.2247066497803,20.205322265625,20.2168464660645,20.2024402618408,20.1690921783447,20.0828113555908,20.0181140899658,19.9471683502197,19.9040050506592,19.8361320495605,19.7547359466553,19.6187496185303,19.6105480194092,19.6854972839355,19.6808586120605,19.6784191131592,19.6751480102539,19.646484375,19.5641593933105,19.4825687408447,19.4204597473145,19.3660640716553,19.3399906158447,19.3049793243408,19.2685070037842,19.2309074401855,19.195556640625,18.9838371276855,18.93408203125,18.8606452941895,18.8034191131592,18.7283210754395,18.6788578033447,18.6509780883789,18.6167964935303,18.5730457305908,18.4962387084961,18.4500980377197,18.4052734375,18.3387222290039,18.2353515625,18.1896495819092,18.1792488098145,18.154296875,18.0774402618408,17.9836921691895,17.9182624816895,17.8344249725342,17.7378425598145,17.6444320678711,17.5286636352539,17.4469718933105,17.415283203125,17.216796875,17.1437015533447,17.0025386810303,16.9818363189697,16.9541015625,16.8766098022461,16.821044921875,16.6507320404053,16.60009765625,16.5379390716553,16.4926280975342,16.458984375,16.4525394439697,16.4903316497803,16.5262699127197,16.515625,16.3965339660645,16.3531246185303,16.3118171691895,16.2798347473145,16.1363277435303,16.0840339660645,16.0673828125,16.0430183410645,15.9978513717651,15.9516611099243,15.9217290878296,15.838623046875,15.8024892807007,15.7472658157349,15.6780757904053,15.6187009811401,15.5604972839355,15.4658212661743,15.3916006088257,15.3098640441895,15.2552242279053,15.1898441314697,15.1184568405151,15.0570306777954,15.0214347839355,14.979884147644,14.9159173965454,14.8718271255493,14.8173837661743,14.705078125,14.6649894714355,14.5628900527954,14.5553226470947,14.5923833847046,14.5721197128296,14.4979009628296,14.4166984558105,14.415771484375,14.4256839752197,14.3910160064697,14.3273429870605,14.3293933868408,14.314697265625,14.3351068496704,14.3877439498901,14.4413080215454,14.4793949127197,14.5050783157349,14.5494146347046,14.5782222747803,14.5150384902954,14.4547853469849,14.4662113189697,14.4762201309204,14.3881349563599,14.3723640441895,14.3571767807007,14.3430185317993,14.2628908157349,14.1270999908447,14.0491209030151,13.9211912155151,13.9245119094849,13.9766120910645,14.049072265625,14.0849609375,14.1561527252197,14.1070804595947,14.1095705032349,14.1614742279053,14.2005367279053,14.2593746185303,14.319091796875,14.3462400436401,14.3678712844849,14.4166984558105,14.4716310501099,14.5301275253296,14.5906744003296,14.6612300872803,14.8433103561401,14.9324703216553,15.0416011810303,15.1275882720947,15.256591796875,15.3196296691895,15.3608884811401,15.4132318496704,15.4882802963257,15.5859384536743,15.6565418243408,15.699951171875,15.7412605285645,15.7804193496704,15.8298835754395,15.8896970748901,15.9421873092651,15.9874515533447,16.0378894805908,16.0935554504395,16.1602535247803,16.2379875183105,16.3399410247803,16.466064453125,16.6475582122803,16.8843746185303,17.0771484375,17.30029296875,17.4616680145264,17.5467281341553,17.6092777252197,17.698974609375,17.8158187866211,17.9883785247803,18.2166976928711,18.3189945220947,18.2953128814697,18.3165054321289,18.382568359375,18.4181652069092,18.42333984375,18.4083976745605,18.3734855651855,18.3389663696289,18.3049812316895,18.2784671783447,18.2594718933105,18.2217292785645,18.1382331848145,18.0285167694092,17.9862308502197,17.9769039154053,17.945556640625,17.8922348022461,17.8517589569092,17.8241214752197,17.8179683685303,17.8333492279053,17.86962890625,17.9267578125,17.965087890625,17.984619140625,18.0459480285645,18.1489734649658,18.203857421875,18.2106456756592,18.1698246002197,18.0814952850342,18.0464363098145,18.0646476745605,18.0333976745605,17.9526863098145,17.889404296875,17.8205089569092,17.8123531341553,17.71875,17.625,17.4990234375,17.4795398712158,17.5099620819092,17.5411128997803,17.5838871002197,17.7971687316895,18.0335445404053,18.1426277160645,18.22216796875,18.286865234375,18.3545398712158,18.4070320129395,18.4419918060303,18.47900390625,18.5335445404053,18.6183109283447,18.7927742004395,18.9771480560303,19.0889167785645,19.2115249633789,19.3279285430908,19.4866218566895,19.54833984375,19.5791988372803,19.6107921600342,19.6053714752197,19.5850582122803,19.5419425964355,19.5147457122803,19.4998531341553,19.553466796875,19.6444835662842,19.756103515625,19.888916015625,19.996337890625,20.0886726379395,20.1323738098145,20.1779289245605,20.18408203125,20.24072265625,20.3403797149658,20.3858871459961,20.3772964477539,20.339599609375,20.2727546691895,20.2454090118408,20.2576675415039,20.316650390625,20.3722171783447,20.589111328125,20.7302722930908,20.7957019805908,20.8232421875,20.81298828125,20.8250980377197,20.8595695495605,20.8792476654053,20.8842296600342,20.9219245910645,20.9923839569092,21.0381832122803,21.059326171875,21.13037109375,21.2513675689697,21.3126449584961,21.314208984375,21.3662109375,21.4686527252197,21.5674800872803,21.5220718383789,21.4075183868408,21.38330078125,21.3424320220947,21.2782230377197,21.2237300872803,21.1973152160645,21.1841297149658,21.2308120727539,21.2342777252197,21.2035636901855,21.1844234466553,21.1696281433105,21.150146484375,21.1563968658447,21.2041492462158,21.2125968933105,21.2359867095947,21.2789058685303,21.31494140625,21.39501953125,21.5338382720947,21.6057605743408,21.7779769897461,21.8265132904053,21.8824710845947,21.9896965026855,22.0552730560303,22.1208972930908,22.1624011993408,22.2098636627197,22.253662109375,22.2763671875,22.3274402618408,22.4054203033447,22.4623050689697,22.486572265625,22.4952640533447,22.4903316497803,22.3884773254395,22.4394054412842,22.439208984375,22.4122562408447,22.3791980743408],[33.4317359924316,33.4156723022461,33.3697738647461,33.3326187133789,33.2750511169434,33.2406234741211,33.271484375,33.25048828125,33.1194801330566,33.07568359375,33.0795402526855,33.0919914245605,33.0836906433105,33.1600074768066,33.2589836120605,33.3926277160645,33.5034675598145,33.8797378540039,34.0321807861328,34.2482414245605,34.4374046325684,34.4933090209961,34.5474128723145,34.6291999816895,34.6286163330078,34.6328582763672,34.6787109375,34.6579093933105,34.6134757995605,34.56689453125,34.5133323669434,34.4996109008789,34.4951667785645,34.4661636352539,34.432373046875,34.2212409973145,34.1343231201172,34.0568351745605,34.0498504638672,34.0113258361816,33.9586410522461,33.92529296875,33.8941879272461,33.8551254272461,33.83935546875,33.8270530700684,33.8355941772461,33.8395500183105,33.8395042419434,33.8315925598145,33.783935546875,33.7526359558105,33.732421875,33.6675758361816,33.6230964660645,33.5979499816895,33.5850563049316,33.5625,33.5345687866211,33.5002899169922,33.4653778076172,33.4317359924316],[7.55849552154541,7.54702091217041,7.51640653610229,7.41181659698486,7.07402372360229,6.98095750808716,6.92001962661743,6.86039972305298,6.80156278610229,6.70512676239014,6.50781297683716,6.49052715301514,6.46806669235229,6.45639705657959,6.46250009536743,6.41318368911743,6.35127019882202,6.31904363632202,6.29072237014771,6.28754901885986,6.29838848114014,6.2861328125,6.23486328125,6.15014696121216,6.07636690139771,6.03891611099243,5.97509765625,5.91904306411743,5.90771484375,5.842041015625,5.84550809860229,5.85371112823486,5.84130907058716,5.65131807327271,5.55058622360229,5.50991249084473,5.47788095474243,5.32451152801514,5.23642539978027,5.13979482650757,5.10849618911743,5.08066415786743,5.00644493103027,4.916748046875,4.821533203125,4.57231426239014,4.38642549514771,4.351318359375,4.36679649353027,4.50869131088257,4.58999013900757,5.05463886260986,5.24106454849243,5.51870155334473,6.07763671875,6.16733360290527,6.21093797683716,6.25849580764771,6.31015586853027,6.465087890625,6.557373046875,6.688232421875,6.90654277801514,6.95122098922729,7.09467792510986,7.23261690139771,7.31440448760986,7.39858388900757,7.46303701400757,7.53823328018188,7.73642539978027,7.75937509536743,7.89643526077271,8.07114219665527,8.12529277801514,8.15761756896973,8.18793964385986,8.31084060668945,8.4541015625,8.48515605926514,8.48881912231445,8.51972675323486,8.505859375,8.464599609375,8.42988300323486,8.51918983459473,8.53769588470459,8.53456974029541,8.45395469665527,8.45888614654541,8.482177734375,8.48442363739014,8.47353553771973,8.43603515625,8.40234375,8.37861347198486,8.34609413146973,8.260009765625,8.17626953125,8.156982421875,8.10698318481445,8.05210018157959,8.02324199676514,7.96792030334473,7.90849685668945,7.86669921875,7.79462909698486,7.70380783081055,7.63955020904541,7.57187509536743,7.50996112823486,7.44252920150757,7.41586923599243,7.39843702316284,7.39492177963257,7.40869188308716,7.37773466110229,7.33330106735229,7.27841758728027,7.25058650970459,7.21591806411743,7.22548818588257,7.25888681411743,7.27461004257202,7.26616191864014,7.2626953125,7.32280302047729,7.39194297790527,7.466796875,7.49570322036743,7.54355478286743,7.60527324676514,7.65888643264771,7.68837928771973,7.68793964385986,7.67705059051514,7.62509775161743,7.58554649353027,7.55849552154541],[31.6549797058105,31.6269035339355,31.5671882629395,31.5140151977539,31.427490234375,31.3348140716553,31.1991710662842,31.0612316131592,30.9264640808105,30.7765617370605,30.6785144805908,30.5580062866211,30.4575214385986,30.2505874633789,30.2010746002197,30.1315441131592,29.8860359191895,29.8087387084961,29.5702629089355,29.3762722015381,29.2238292694092,29.181884765625,28.9573249816895,28.7327651977539,28.5082035064697,28.2836418151855,28.0590839385986,27.8345222473145,27.6099605560303,27.3854007720947,27.1608390808105,26.9362316131592,26.711669921875,26.4871101379395,26.2625484466553,26.0379886627197,25.8134269714355,25.5888671875,25.3643074035645,25.1397457122803,24.9151859283447,24.6906242370605,24.466064453125,24.2415046691895,24.0169429779053,23.7923831939697,23.5678215026855,23.3432140350342,23.11865234375,22.8940925598145,22.6695308685303,22.4449710845947,22.2204093933105,21.995849609375,21.49755859375,20.9992179870605,20.5009269714355,20.0025882720947,20.0007820129395,19.9990253448486,19.9972648620605,19.9954586029053,19.9955558776855,19.9957027435303,19.995849609375,19.9959468841553,19.87109375,19.7462882995605,19.6214847564697,19.4966316223145,19.7332038879395,19.9697761535645,20.2063484191895,20.4429206848145,20.6794929504395,20.9161148071289,21.1526374816895,21.3892574310303,21.6258296966553,21.8624019622803,22.0989742279053,22.3355464935303,22.5721187591553,22.8086910247803,23.0452632904053,23.2818355560303,23.4452152252197,23.2857418060303,23.1606941223145,22.9961910247803,22.9962882995605,22.7825202941895,22.6184558868408,22.6196765899658,22.6237316131592,22.902099609375,23.1195316314697,23.18017578125,23.291259765625,23.4016609191895,23.5178718566895,23.69482421875,23.892578125,24.1396961212158,24.2908191680908,24.3143558502197,24.4340343475342,24.5513668060303,24.480224609375,24.4855937957764,24.5302257537842,24.5910148620605,24.6762199401855,24.7902355194092,25.051025390625,25.258544921875,25.3320808410645,25.624267578125,25.89013671875,26.067138671875,26.1470699310303,26.2455081939697,26.333740234375,26.4382305145264,26.5519523620605,26.6308097839355,26.8479480743408,26.9158210754395,27.0447769165039,27.2193355560303,27.3308582305908,27.552978515625,27.7856922149658,28.0433101654053,28.5602054595947,28.9669933319092,29.1147937774658,29.1769523620605,29.3689460754395,29.5669898986816,29.6364269256592,29.7959461212158,29.99365234375,30.1152324676514,30.1792964935303,30.2293968200684,30.2823238372803,30.3422374725342,30.3873023986816,30.4253406524658,30.580078125,30.6659679412842,30.7832012176514,30.8649425506592,30.9408226013184,31.0321273803711,31.2509765625,31.4637680053711,31.5458030700684,31.5851058959961,31.6849613189697,31.704833984375,31.7360363006592,31.8025398254395,31.8857402801514,31.9295387268066,31.9753913879395,32.0211906433105,32.0806655883789,32.1727027893066,32.2567405700684,32.34521484375,32.4136734008789,32.4733390808105,32.5249519348145,32.642578125,32.7816925048828,32.8973655700684,32.9657249450684,33.1555671691895,33.1819343566895,33.118896484375,33.0937004089355,32.8585433959961,32.8291015625,32.8010749816895,32.8973655700684,32.9146461486816,32.8242683410645,32.798828125,32.7917976379395,32.7097663879395,32.6812515258789,32.5502891540527,32.5110855102539,32.3911628723145,32.3116722106934,32.15966796875,31.9711894989014,31.834228515625,31.6567859649658,31.5311031341553,31.4264183044434,31.3609886169434,31.2644538879395,31.2272930145264,31.2147464752197,31.0814933776855,30.9275875091553,30.8519039154053,30.7772941589355,30.4156723022461,30.2904300689697,30.2661113739014,30.2880840301514,30.4137706756592,30.4883804321289,30.8006839752197,30.9637203216553,31.07861328125,31.1955089569092,31.300537109375,31.4106426239014,31.5560073852539,31.8175277709961,31.9990711212158,32.1078605651855,32.21875,32.4307632446289,32.5801773071289,32.7755355834961,32.7776832580566,32.7991676330566,32.9373016357422,32.9424819946289,32.9086456298828,32.9182624816895,32.8798828125,32.7939453125,32.7405281066895,32.6871566772461,32.6187515258789,32.4481468200684,32.3974113464355,32.3314437866211,32.2138175964355,32.15869140625,32.127197265625,32.0370140075684,32.0092277526855,31.9965343475342,32.0159645080566,31.9842758178711,31.9537124633789,31.8833503723145,31.7123050689697,31.6549797058105],[13.8219728469849,13.7175779342651,13.7831058502197,13.8655757904053,13.9155759811401,14.0109376907349,14.0728511810303,14.0933589935303,14.0111322402954,13.8219728469849],[47.057373046875,47.062744140625,47.062255859375,47.0975074768066,47.1726570129395,47.270751953125,47.254638671875,47.234130859375,47.2122535705566,47.1854972839355,47.1579093933105,47.132080078125,47.1071281433105,47.0758285522461,47.057373046875],[9.05073261260986,8.97500038146973,9.02685546875,9.06977462768555,9.10459041595459,9.06572246551514,9.05073261260986],[9.63066387176514,9.61982440948486,9.68637752532959,9.71464824676514,9.73437404632568,9.74433612823486,9.74116230010986,9.67914962768555,9.63066387176514],[9.81269550323486,9.80747127532959,9.81000900268555,9.79633808135986,9.642333984375,9.36635780334473,9.08588790893555,9.02451133728027,8.97148418426514,8.95693302154541,8.9326171875,8.66196346282959,8.60839939117432,8.54941368103027,8.50551795959473,8.48359394073486,8.47206974029541,8.43144512176514,8.21523475646973,8.14785099029541,8.118896484375,7.78247117996216,7.74155235290527,7.71093797683716,7.68447256088257,7.62500047683716,7.46479463577271,7.42841815948486,7.288330078125,7.02045869827271,6.90126943588257,6.75620174407959,6.61430644989014,6.51186513900757,6.42514657974243,6.24091815948486,6.203857421875,6.08837890625,5.97905302047729,5.94936513900757,6.009765625,6.15317344665527,6.36440420150757,6.58452129364014,6.82929706573486,7.585205078125,7.79648399353027,8.065673828125,8.18232440948486,8.29423809051514,8.04887676239014,8.01845741271973,8.04999923706055,8.30405235290527,8.41157150268555,8.69150352478027,8.74116230010986,8.846435546875,8.89921951293945,9.09565448760986,9.2099609375,9.32685565948486,9.45327091217041,9.57783126831055,9.53813457489014,9.49477481842041,9.46543025970459,9.48046779632568,9.48095703125,9.54877948760986,9.61127948760986,9.64990234375,9.69936561584473,9.74233341217041,9.79262638092041,9.81269550323486],[-30.1019515991211,-30.1265640258789,-30.1287117004395,-30.123046875,-30.12890625,-30.1424827575684,-30.1475601196289,-30.2184581756592,-30.4098625183105,-30.4499015808105,-30.5250988006592,-30.5845699310303,-30.6310558319092,-30.6422843933105,-30.6238288879395,-30.5999984741211,-30.5422840118408,-30.4664077758789,-30.4112300872803,-30.3809585571289,-30.3639640808105,-30.3384780883789,-30.3252925872803,-30.31591796875,-30.2791976928711,-30.2473621368408,-30.1585941314697,-30.1056632995605,-30.0153312683105,-29.9413089752197,-29.8402366638184,-29.7537117004395,-29.6640605926514,-29.6255874633789,-29.5993175506592,-29.5541973114014,-29.5193367004395,-29.4552726745605,-29.3600597381592,-29.302734375,-29.2765617370605,-29.2361335754395,-29.1464824676514,-29.0469741821289,-28.9400405883789,-28.9090824127197,-28.8733406066895,-28.7799816131592,-28.7012672424316,-28.6158199310303,-28.5941410064697,-28.5817375183105,-28.5978527069092,-28.6467761993408,-28.6876964569092,-28.7588863372803,-28.7760753631592,-28.8814449310303,-28.9537105560303,-29.0369129180908,-29.0783214569092,-29.08984375,-29.1636714935303,-29.2184562683105,-29.2697277069092,-29.3197269439697,-29.4419918060303,-29.56689453125,-29.6188488006592,-29.6516609191895,-29.7009754180908,-29.8011703491211,-29.9190425872803,-29.9675769805908,-29.9994125366211,-30.0384750366211,-30.1019515991211],[55.2789077758789,55.2866706848145,55.4019508361816,55.476806640625,55.5831069946289,55.6165008544922,55.5681648254395,55.4877433776855,55.3504867553711,55.2789077758789],[55.6675300598145,55.6226577758789,55.5464820861816,55.4481468200684,55.3719253540039,55.3424835205078,55.3180160522461,55.3064460754395,55.2933578491211,55.2730941772461,55.2467765808105,55.2249031066895,55.2041969299316,55.1301765441895,55.1395988464355,55.12451171875,55.0901374816895,55.0503883361816,55.0032691955566,54.9623031616211,54.9471664428711,54.9192886352539,54.833251953125,54.7178726196289,54.6360359191895,54.5903816223145,54.5642585754395,54.5357933044434,54.4603996276855,54.3770523071289,54.3318367004395,54.3106918334961,54.3101043701172,54.29296875,54.2596664428711,54.22119140625,54.1797866821289,54.156982421875,54.1404762268066,54.139892578125,54.1596183776855,54.1752433776855,54.2151336669922,54.2649421691895,54.2927742004395,54.251220703125,54.2512702941895,54.2142562866211,54.1549301147461,54.133056640625,54.1451683044434,54.1189918518066,53.9982414245605,53.9746589660645,53.9798355102539,53.9318351745605,53.8929672241211,53.9199714660645,53.950439453125,53.9450187683105,53.931640625,53.93896484375,53.9356918334961,53.9122543334961,53.9198265075684,53.9397926330566,53.9589347839355,54.0059585571289,54.0790023803711,54.1434555053711,54.2004890441895,54.2403335571289,54.2814445495605,54.2994651794434,54.30419921875,54.327880859375,54.3483428955078,54.3663558959961,54.3905296325684,54.3958015441895,54.3567848205566,54.4056167602539,54.4930191040039,54.5629386901855,54.6326179504395,54.8384742736816,54.8712882995605,54.9288558959961,54.970703125,55.0591316223145,55.0642585754395,55.0593757629395,55.0636711120605,55.1007308959961,55.1603507995605,55.1953125,55.2344245910645,55.2755355834961,55.2644538879395,55.2641105651855,55.2711906433105,55.3437995910645,55.4550285339355,55.6177253723145,55.8134269714355,56.02294921875,56.070068359375,56.1881332397461,56.3145523071289,56.3259773254395,56.4007835388184,56.40673828125,56.3928718566895,56.3750991821289,56.3772926330566,56.3964347839355,56.3804206848145,56.3240737915039,56.3306617736816,56.3671417236328,56.3338356018066,56.3346176147461,56.3292465209961,56.2952613830566,56.2642555236816,56.2830085754395,56.2840805053711,56.2962913513184,56.3812980651855,56.3958969116211,56.4111824035645,56.398193359375,56.3255844116211,56.2003898620605,56.1784172058105,56.1301765441895,56.1048316955566,55.9943351745605,55.9401359558105,55.8968734741211,55.8121070861816,55.7504386901855,55.7038078308105,55.6724090576172,55.6675300598145],[50.1209983825684,50.0942344665527,50.0343742370605,49.9743156433105,49.9151344299316,49.8721694946289,49.837890625,49.8053245544434,49.7984848022461,49.75439453125,49.7078132629395,49.6820297241211,49.6449699401855,49.599609375,49.5126953125,49.4527359008789,49.4775390625,49.4943389892578,49.4989280700684,49.4852066040039,49.4546394348145,49.4454612731934,49.4546394348145,49.4775390625,49.48974609375,49.5050773620605,49.5382804870605,49.5538063049316,49.5783195495605,49.6128425598145,49.644775390625,49.732177734375,49.7588882446289,49.8083000183105,49.8333511352539,49.857177734375,49.8756370544434,49.9196281433105,49.9612312316895,50.0126953125,50.0828132629395,50.1671867370605,50.154296875,50.1545906066895,50.123779296875,50.1209983825684],[57.5281219482422,57.5240249633789,57.5081558227539,57.4297866821289,57.3681182861328,57.3169479370117,57.2933120727539,57.2477035522461,57.1944808959961,57.1668930053711,57.1351051330566,57.0546379089355,56.9780769348145,56.8456535339355,56.8432121276855,56.8670883178711,56.8534202575684,56.8241691589355,56.7410659790039,56.7037124633789,56.6453132629395,56.599853515625,56.5457000732422,56.5106925964355,56.3868675231934,56.3155746459961,56.2604026794434,56.1903305053711,56.1429176330566,56.1458015441895,56.13330078125,56.076171875,55.9415512084961,55.9117202758789,55.8091773986816,55.7987785339355,55.8035163879395,55.8059577941895,55.8039054870605,55.83056640625,55.8129386901855,55.709228515625,55.6939926147461,55.6796379089355,55.6675300598145,55.6724090576172,55.7038078308105,55.7504386901855,55.8121070861816,55.8968734741211,55.9401359558105,55.9943351745605,56.1048316955566,56.1301765441895,56.1784172058105,56.2003898620605,56.3255844116211,56.398193359375,56.4111824035645,56.3958969116211,56.3812980651855,56.2962913513184,56.2840805053711,56.2830085754395,56.2642555236816,56.2952613830566,56.3292465209961,56.3346176147461,56.3338356018066,56.3671417236328,56.3306617736816,56.3240737915039,56.3804206848145,56.3964347839355,56.3772926330566,56.3750991821289,56.3928718566895,56.40673828125,56.4007835388184,56.3259773254395,56.3145523071289,56.1881332397461,56.070068359375,56.2589378356934,56.6365699768066,56.82373046875,56.9327659606934,57.0176734924316,57.1310043334961,57.23583984375,57.3224563598633,57.5709953308105,57.5978507995605,57.6667938232422,57.7242660522461,57.6511764526367,57.5953598022461,57.3920936584473,57.3238296508789,57.0897483825684,56.9710464477539,57.0084953308105,57.0661125183105,57.1723136901855,57.2500495910645,57.3250007629395,57.6453132629395,57.7841300964355,57.8706092834473,57.8661613464355,57.9078598022461,57.9852523803711,57.9887237548828,58.0045890808105,58.0634231567383,58.0321273803711,57.9965858459473,57.9961395263672,58.0093765258789,58.0322227478027,58.0484886169434,58.0394515991211,57.9427757263184,57.9201698303223,57.9138145446777,57.8685531616211,57.8381843566895,57.8147430419922,57.7855453491211,57.6627464294434,57.60107421875,57.5444793701172,57.531005859375,57.5887184143066,57.6087875366211,57.6091270446777,57.5787620544434,57.5383338928223,57.5254859924316,57.5281219482422],[22.195556640625,22.2174797058105,22.2415523529053,22.2459487915039,22.2226085662842,22.2214851379395,22.20166015625,22.19775390625,22.195556640625],[18.0689468383789,18.0689468383789,18.0907230377197,18.1153316497803,18.1130867004395,18.1042976379395,18.0689468383789],[35.1041984558105,35.02978515625,34.9708480834961,34.8355445861816,34.7519035339355,34.7231941223145,34.6546363830566,34.6073265075684,34.5570793151855,34.49609375,34.467041015625,34.4332504272461,34.367919921875,34.1760749816895,33.9902839660645,33.8582038879395,33.7818374633789,33.7168464660645,33.5667495727539,33.3186531066895,33.183349609375,33.0735816955566,32.8776397705078,32.7848167419434,32.703369140625,32.6756820678711,32.6084938049316,32.5522956848145,32.4683113098145,32.399169921875,32.3376007080078,32.2711410522461,32.16455078125,32.1072273254395,32.0890159606934,32.0948753356934,32.0995597839355,32.1047859191895,32.1150398254395,32.121337890625,32.1299819946289,32.1256828308105,32.0957527160645,32.07470703125,32.06884765625,32.0425262451172,31.9639644622803,31.8742179870605,31.8342761993408,31.7045402526855,31.686767578125,31.7000961303711,31.6895503997803,31.6619148254395,31.619873046875,31.56640625,31.5123558044434,31.4371109008789,31.3618183135986,31.3088397979736,31.2554683685303,31.1978034973145,31.1666011810303,31.1618156433105,31.1354007720947,31.1113777160645,31.0657730102539,31.0009269714355,30.9640140533447,30.9444828033447,30.92724609375,30.9135246276855,30.8095703125,30.6988754272461,30.6255359649658,30.6047859191895,30.5523929595947,30.4653797149658,30.326416015625,30.1661624908447,30.0586414337158,29.9569339752197,29.9179706573486,29.8690414428711,29.8312511444092,29.8189468383789,29.8106918334961,29.8083019256592,29.8161144256592,29.8203601837158,29.8091316223145,29.7837886810303,29.7260246276855,29.6598644256592,29.6038570404053,29.5789546966553,29.5687999725342,29.5838375091553,29.6016101837158,29.6251945495605,29.6195793151855,29.6126480102539,29.5749034881592,29.4947280883789,29.4250011444092,29.3922367095947,29.3751964569092,29.3495101928711,29.1747570037842,29.1324234008789,28.9805164337158,28.9301738739014,28.8801784515381,28.7678718566895,28.7186031341553,28.6894035339355,28.6207523345947,28.46923828125,28.3236808776855,28.1120109558105,27.900390625,27.6564445495605,27.6564445495605,27.6559085845947,27.6138668060303,27.5308589935303,27.4605464935303,27.4165515899658,27.3609371185303,27.3082027435303,27.2505874633789,27.1910152435303,27.1509761810303,27.1207027435303,27.1041011810303,27.0904293060303,27.0904293060303,27.1001968383789,27.0982437133789,27.0982437133789,27.0884780883789,27.0503902435303,26.9908218383789,26.9107418060303,26.8609390258789,26.8501968383789,26.8501968383789,26.8902339935303,26.9107418060303,26.9087886810303,26.8804683685303,26.8609390258789,26.8609390258789,26.9009761810303,26.9605464935303,26.9908218383789,27.0005855560303,27.0201168060303,27.0103511810303,27.0103511810303,26.9703121185303,26.9410152435303,26.9107418060303,26.8833999633789,26.7935543060303,26.7447280883789,26.6841793060303,26.6333980560303,26.5835933685303,26.5201168060303,26.4703121185303,26.4009761810303,26.3609371185303,26.2955074310303,26.2134780883789,26.1626949310303,26.1041011810303,26.0865230560303,26.0708980560303,26.0503902435303,26.0308589935303,25.996337890625,25.9908199310303,25.9205074310303,25.8707027435303,25.8306636810303,25.7310543060303,25.6402339935303,25.5201168060303,25.4205074310303,25.2603015899658,25.1109371185303,24.9703121185303,24.8804683685303,24.8306636810303,24.7701168060303,24.7310543060303,24.6802730560303,24.6304702758789,24.5708980560303,24.5201187133789,24.4972648620605,24.4703121185303,24.4009780883789,24.3003902435303,24.2203121185303,24.0904312133789,24.0201168060303,23.9810543060303,23.9410152435303,23.9107418060303,23.8707027435303,23.8306636810303,23.7906246185303,23.7505855560303,23.6910171508789,23.6207027435303,23.5201168060303,23.4107418060303,23.3404293060303,23.1001949310303,22.9605464935303,22.8707027435303,22.7603511810303,22.5904312133789,22.4507808685303,22.3707027435303,22.3101558685303,22.2408199310303,22.1910171508789,22.1207027435303,22.0806636810303,22.0406265258789,21.9908695220947,21.9107418060303,21.8609371185303,21.8208980560303,21.7505855560303,21.6802730560303,21.6001949310303,21.5005874633789,21.4507808685303,21.4410152435303,21.4410152435303,21.4507808685303,21.4507808685303,21.4703121185303,21.4908199310303,21.5005874633789,21.5005874633789,21.4810543060303,21.4810543060303,21.4703121185303,21.4302730560303,21.4207515716553,21.4207019805908,21.9000015258789,22.1597175598145,22.2743644714355,22.33349609375,22.5945301055908,22.8348140716553,22.9453620910645,23.0319347381592,23.097900390625,23.2275390625,23.4254875183105,23.5526371002197,23.74951171875,23.7928714752197,23.8422355651855,23.8003425598145,23.7275867462158,23.6703128814697,23.7408199310303,23.8444328308105,23.9529304504395,24.07275390625,24.4788093566895,24.548828125,24.7197742462158,24.87158203125,25.2201175689697,25.4041500091553,25.5477046966553,25.8085441589355,25.9252452850342,26.1630382537842,26.2537097930908,26.2967281341553,26.4154281616211,26.48876953125,26.6429195404053,26.735107421875,26.8726577758789,27.1466312408447,27.4346199035645,27.65185546875,27.6557121276855,27.7698249816895,27.9141597747803,27.9784183502197,28.0094242095947,28.1292972564697,28.3101081848145,28.3820323944092,28.5260753631592,28.7137699127197,28.9392070770264,29.06494140625,29.3803691864014,29.6414051055908,29.8092308044434,29.9582042694092,30.1092777252197,30.3526363372803,30.4475593566895,30.6031227111816,30.6445789337158,30.7179183959961,30.8472652435303,31.0696296691895,31.4246101379395,31.7109870910645,32.0863761901855,32.2405776977539,32.48583984375,32.5724601745605,32.9204597473145,33.1871566772461,33.25244140625,33.3743629455566,33.6402816772461,33.830322265625,33.9690437316895,34.1329116821289,34.7760772705078,35.68115234375,35.7857933044434,35.8159675598145,35.8289031982422,35.8620109558105,35.9298820495605,35.9027366638184,35.8565444946289,35.7452125549316,35.61474609375,35.4677734375,35.2812995910645,35.2063980102539,35.1614761352539,35.243408203125,35.2449188232422,35.2799797058105,35.2283210754395,35.2118186950684,35.2391128540039,35.3172378540039,35.4072723388672,35.3630867004395,35.3151397705078,35.287109375,35.1726570129395,35.1278343200684,35.1352043151855,35.1126937866211,35.1234855651855,35.1041984558105],[43.7504386901855,43.7317390441895,43.7532234191895,43.7653312683105,43.7708969116211,43.761474609375,43.7504386901855],[45.450439453125,45.5137672424316,45.5691413879395,45.5989761352539,45.6127433776855,45.6282730102539,45.6471176147461,45.7130851745605,45.7888679504395,45.8255348205566,45.883056640625,45.9726066589355,46.138671875,46.3475608825684,46.4512710571289,46.508056640625,46.6408195495605,46.7063980102539,46.7920875549316,46.9784164428711,47.043212890625,47.114501953125,47.1683082580566,47.2275886535645,47.2864227294922,47.3405265808105,47.4756355285645,47.5366668701172,47.5531272888184,47.6397476196289,47.7179679870605,47.7822265625,47.8417510986328,47.9592781066895,48.0476570129395,48.1104965209961,48.155029296875,48.2111320495605,48.2558135986328,48.2634773254395,48.2598609924316,48.2941398620605,48.3871574401855,48.3719253540039,48.3682594299316,48.3714332580566,48.4327125549316,48.4156265258789,48.4430656433105,48.4777336120605,48.4704093933105,48.4648895263672,48.4495124816895,48.416259765625,48.365234375,48.3335418701172,48.3212890625,48.2957992553711,48.2570304870605,48.2379875183105,48.2385711669922,48.2130393981934,48.1614265441895,48.1444320678711,48.162109375,48.1468734741211,48.1086883544922,48.0905303955078,48.1502914428711,48.1443862915039,48.1195793151855,47.9956550598145,47.9511260986328,47.9330062866211,47.9523429870605,47.9754409790039,47.9645538330078,47.8824234008789,47.7750015258789,47.7315444946289,47.6585960388184,47.5808601379395,47.5303726196289,47.4896965026855,47.4556617736816,47.4444808959961,47.375732421875,47.3280258178711,47.2926254272461,47.2907218933105,47.27099609375,47.246826171875,47.185546875,47.1280288696289,47.0911140441895,47.0475120544434,46.9967269897461,46.9640159606934,46.9388198852539,46.8829116821289,46.8289070129395,46.7824211120605,46.7237815856934,46.625,46.5388679504395,46.423095703125,46.4015617370605,46.3778305053711,46.3602066040039,46.3505363464355,46.4377937316895,46.44873046875,46.416748046875,46.3988265991211,46.4077644348145,46.4346694946289,46.4537582397461,46.4559593200684,46.4369125366211,46.445068359375,46.4665985107422,46.3926239013672,46.376953125,46.3793449401855,46.5049819946289,46.5239753723145,46.5269050598145,46.4970207214355,46.45849609375,46.4241218566895,46.3622550964355,46.2884292602539,46.1764640808105,46.1276397705078,46.0499534606934,45.9786605834961,45.9371566772461,45.8520011901855,45.7938461303711,45.7357902526855,45.665771484375,45.6178207397461,45.5724105834961,45.5415534973145,45.5177268981934,45.5071754455566,45.4985847473145,45.48388671875,45.450439453125],[-16.9029312133789,-17.0865249633789,-16.9332027435303,-16.71240234375,-16.6953125,-16.9029312133789],[-13.3638668060303,-13.400390625,-13.38525390625,-13.2599611282349,-13.2560548782349,-13.20458984375,-13.1982421875,-13.3095703125,-13.3638668060303],[-12.43212890625,-12.5367193222046,-12.6371097564697,-12.8796873092651,-12.9730472564697,-13.072265625,-13.2702150344849,-13.5779294967651,-14.0402345657349,-14.5144529342651,-14.7320318222046,-14.9368162155151,-15.1493167877197,-15.3856449127197,-15.6291017532349,-15.8584957122803,-15.9015636444092,-15.96044921875,-15.9578132629395,-15.8986330032349,-15.8017568588257,-15.5735349655151,-15.4577150344849,-15.439453125,-15.4495124816895,-15.5215826034546,-15.5669918060303,-15.6957025527954,-15.8114252090454,-15.9289064407349,-16.0767574310303,-16.1214847564697,-16.1590824127197,-16.255859375,-16.4865245819092,-16.60302734375,-16.7030277252197,-16.7583980560303,-16.8151359558105,-16.8494148254395,-16.8928718566895,-16.93115234375,-17.0329113006592,-17.2406253814697,-17.3466796875,-17.6695327758789,-17.8985347747803,-18.3363285064697,-18.5440444946289,-18.7922859191895,-19.11962890625,-19.5304679870605,-19.9532222747803,-20.2073249816895,-20.45751953125,-20.8999996185303,-21.3490238189697,-21.8430652618408,-22.3939456939697,-22.4658203125,-22.7472648620605,-22.9915046691895,-23.2333984375,-23.6331043243408,-23.75634765625,-23.8746089935303,-24.1251945495605,-24.2184581756592,-24.3175773620605,-24.4431629180908,-24.5643558502197,-24.7872066497803,-24.97900390625,-25.0487308502197,-25.14990234375,-25.17041015625,-25.1727542877197,-25.2303695678711,-25.34130859375,-25.4684562683105,-25.5287113189697,-25.5631828308105,-25.5705070495605,-25.5430660247803,-25.3341789245605,-25.2997074127197,-25.2710933685303,-25.2533206939697,-25.22607421875,-25.1168956756592,-25.0246086120605,-24.9957046508789,-24.9320316314697,-24.8634757995605,-24.7867183685303,-24.640625,-24.5383777618408,-24.35791015625,-24.30029296875,-24.1087894439697,-23.9791984558105,-23.7418937683105,-23.6302738189697,-23.5296878814697,-23.4208984375,-23.3065433502197,-23.1881828308105,-23.0804691314697,-22.8863277435303,-22.7908191680908,-22.69189453125,-22.3835945129395,-22.2763671875,-22.04931640625,-21.9325199127197,-21.8511714935303,-21.7904300689697,-21.73828125,-21.6964855194092,-21.6466789245605,-21.3564453125,-21.2919921875,-21.2549800872803,-21.17919921875,-21.0768547058105,-20.8658199310303,-20.65625,-20.5460929870605,-20.3796882629395,-20.1455078125,-20.03515625,-19.9220695495605,-19.6742191314697,-19.5508785247803,-19.4287109375,-19.3031253814697,-19.0751953125,-19.0326175689697,-18.8631820678711,-18.7406253814697,-18.6185550689697,-18.5035152435303,-18.2884769439697,-17.9330081939697,-17.8044910430908,-17.6903324127197,-17.5814456939697,-17.3916015625,-16.70263671875,-16.6214847564697,-16.4113273620605,-16.2890625,-16.2437496185303,-16.21728515625,-16.2044925689697,-16.1745109558105,-16.1533203125,-16.0951175689697,-15.9828119277954,-15.9504880905151,-15.9623041152954,-16.0104484558105,-16.0367183685303,-15.9858388900757,-15.9842767715454,-15.9925775527954,-15.94580078125,-15.8831062316895,-15.8388671875,-15.8137693405151,-15.8000974655151,-15.7821283340454,-15.7382802963257,-15.7468748092651,-15.9045906066895,-15.9181642532349,-15.9246101379395,-15.8958988189697,-15.8133783340454,-15.7666997909546,-15.7136716842651,-15.60009765625,-15.5134773254395,-15.3818349838257,-15.2295904159546,-15.2190427780151,-15.2431640625,-15.4226560592651,-15.4522457122803,-15.4563474655151,-15.4341802597046,-15.3617191314697,-15.3015632629395,-15.2438468933105,-15.1950197219849,-15.1500968933105,-15.0440435409546,-14.9426765441895,-14.82177734375,-14.7661123275757,-14.7033205032349,-14.7132806777954,-14.7643556594849,-14.8183603286743,-14.8719720840454,-14.9250001907349,-14.9957027435303,-15.0093746185303,-14.9921865463257,-14.8642568588257,-14.7432622909546,-14.6803712844849,-14.6367177963257,-14.6455087661743,-14.6725597381592,-14.5448246002197,-14.3699226379395,-14.0672855377197,-14.0042972564697,-13.98486328125,-13.96044921875,-13.8582029342651,-13.8075189590454,-13.7306642532349,-13.6624021530151,-13.6146488189697,-13.5962886810303,-13.62255859375,-13.70654296875,-13.7193355560303,-13.638671875,-13.5379877090454,-13.46875,-13.4259767532349,-13.2674798965454,-12.9358406066895,-12.8390626907349,-12.7216796875,-12.6101560592651,-12.4708986282349,-12.4400386810303,-12.45849609375,-12.4390630722046,-12.3158197402954,-12.07958984375,-12.0801753997803,-12.1239252090454,-12.1886720657349,-12.236328125,-12.3484373092651,-12.43212890625],[3.23125004768372,3.22939467430115,3.24648427963257,3.27138686180115,3.28876972198486,3.28984403610229,3.27431631088257,3.25014662742615,3.23125004768372],[4.16455078125,4.15517568588257,4.158935546875,4.17070293426514,4.18813467025757,4.21044921875,4.23461961746216,4.24765634536743,4.24331045150757,4.22968721389771,4.21103525161743,4.18686485290527,4.16455078125],[18.6773433685303,18.6541996002197,18.6758785247803,18.7780265808105,18.773681640625,18.760009765625,18.7114734649658,18.6773433685303],[18.741455078125,18.7203617095947,18.7816410064697,18.8301258087158,18.8631343841553,18.8017101287842,18.741455078125],[20.3033199310303,20.2721691131592,20.38232421875,20.4897937774658,20.551513671875,20.5587882995605,20.5790519714355,20.5517578125,20.4684562683105,20.3033199310303],[21.2393074035645,21.1910152435303,21.1967792510986,21.2333011627197,21.27880859375,21.2799797058105,21.2643070220947,21.2393074035645],[21.61083984375,21.5285148620605,21.5614738464355,21.6131343841553,21.6978511810303,21.712158203125,21.6763668060303,21.65234375,21.61083984375],[24.1510734558105,24.1475582122803,24.2006340026855,24.3309078216553,24.3448238372803,24.1833972930908,24.16357421875,24.1510734558105],[24.3936042785645,24.3463859558105,24.5333995819092,24.5511226654053,24.5379886627197,24.3936042785645],[25.0034656524658,24.8915519714355,24.9080581665039,24.9688472747803,25.046630859375,25.0814456939697,25.0878410339355,25.0421390533447,25.0034656524658],[24.5457038879395,24.5345706939697,24.6502933502197,24.7295894622803,24.7631340026855,24.7896499633789,24.9511241912842,25.28564453125,25.224365234375,24.8854484558105,24.841064453125,24.7996578216553,24.7295894622803,24.6540031433105,24.5836429595947,24.5457038879395],[26.0206069946289,25.9845199584961,25.9740715026855,25.9991703033447,25.8497066497803,25.8358898162842,26.0406246185303,26.06982421875,26.07568359375,26.0206069946289],[28.0693855285645,28.0372562408447,28.1039562225342,28.2205581665039,28.3427734375,28.3683586120605,28.327880859375,28.172119140625,28.0693855285645],[28.9419422149658,28.9037590026855,29.1123046875,29.1627426147461,29.1875972747803,29.182861328125,29.1305179595947,29.08642578125,29.0433616638184,28.9419422149658],[29.0053195953369,28.7693367004395,28.7731456756592,28.8476085662842,28.8939952850342,29.167724609375,29.2036628723145,29.2404289245605,29.206787109375,29.1259765625,29.0053195953369],[29.0522480010986,29.0347652435303,29.0967292785645,29.3076171875,29.4132328033447,29.4626941680908,29.5730457305908,29.5599117279053,29.4859371185303,29.41748046875,29.3389167785645,29.3018569946289,29.1319351196289,29.0522480010986],[31.7056159973145,31.7013664245605,31.7474117279053,31.7897930145264,31.7940902709961,31.7568855285645,31.7242164611816,31.7056159973145],[25.9614734649658,25.7549324035645,25.58544921875,25.2331066131592,25.0145492553711,24.3899917602539,23.9806156158447,23.7879409790039,23.7606449127197,23.7322273254395,23.30615234375,22.9423828125,22.8860359191895,22.7763195037842,22.62451171875,22.5570812225342,22.5103015899658,22.279296875,22.1058597564697,21.8085441589355,21.704833984375,21.6149406433105,21.564208984375,21.4378910064697,21.3739261627197,21.2725601196289,21.3564929962158,21.46533203125,21.5238285064697,21.5666999816895,21.6124019622803,21.7620124816895,22.0266609191895,21.6036605834961,21.535888671875,21.5077133178711,21.4779796600342,21.4320297241211,21.3980464935303,21.10400390625,20.8000984191895,20.717041015625,20.614990234375,20.1882820129395,19.8697757720947,19.5672378540039,19.4728507995605,19.34375,19.1990718841553,19.1056652069092,19.0537605285645,18.8971672058105,18.8055171966553,18.8038578033447,18.85009765625,18.819580078125,18.754638671875,18.6905746459961,18.6904296875,18.7236804962158,18.7683601379395,18.77490234375,18.7191410064697,18.7007312774658,18.5706043243408,18.5145988464355,18.3484878540039,18.1748542785645,18.1666507720947,18.1659679412842,18.1952629089355,18.304443359375,18.35791015625,18.4304676055908,18.4437980651855,18.4234390258789,18.4686527252197,18.5241203308105,18.5745124816895,18.6116695404053,18.664794921875,18.67529296875,18.6848640441895,18.7043952941895,18.7158699035645,18.6377925872803,18.5996589660645,18.5634288787842,18.5285148620605,18.4706058502197,18.4471683502197,18.45654296875,18.5418453216553,18.6244621276855,18.7206535339355,18.7732906341553,18.7765617370605,18.8061046600342,18.8768081665039,18.9005870819092,18.8328113555908,18.8330078125,18.8646469116211,18.8897953033447,19.0374984741211,19.0981941223145,19.1516609191895,19.3522472381592,19.7298831939697,19.7959480285645,19.911865234375,19.94677734375,20.0257320404053,20.2240219116211,20.3799800872803,20.5563468933105,20.7137222290039,20.7575206756592,21.0094242095947,21.1208992004395,21.2526378631592,21.2746105194092,21.4141120910645,21.4481449127197,21.5386714935303,21.5693836212158,21.56689453125,21.5789546966553,21.5914554595947,21.603515625,21.6158676147461,21.6040515899658,21.5495109558105,21.5358390808105,21.4724597930908,21.4469738006592,21.4580078125,21.477294921875,21.5142097473145,21.5464363098145,21.5439453125,21.526611328125,21.5249519348145,21.5514640808105,21.57373046875,21.5716304779053,21.5824203491211,21.6214847564697,21.5922355651855,21.4628410339355,21.4216785430908,21.2342281341553,21.2000484466553,21.1505355834961,21.0052242279053,20.8850593566895,20.7864761352539,20.6312503814697,20.5072765350342,20.2313957214355,20.1021480560303,19.99853515625,19.8984851837158,19.8615245819092,19.8241691589355,19.8274917602539,19.7794914245605,19.6371097564697,19.5937004089355,19.5539054870605,19.572998046875,19.57470703125,19.5864734649658,19.5833492279053,19.501708984375,19.4437503814697,19.4255847930908,19.4157218933105,19.3827133178711,19.3523445129395,19.2578620910645,19.25048828125,19.3209476470947,19.3174800872803,19.2877941131592,19.04638671875,18.7985343933105,18.655029296875,18.4461421966553,18.3570785522461,18.2689933776855,18.2738780975342,18.4408683776855,18.48388671875,18.5244636535645,18.7268562316895,18.8389148712158,18.83447265625,18.7730464935303,18.7196769714355,18.6426277160645,18.5145511627197,18.472412109375,18.4823246002197,18.4767570495605,18.4458999633789,18.29052734375,18.0716304779053,17.9655265808105,17.9288082122803,17.91455078125,17.9396495819092,17.9997062683105,17.9708023071289,17.9019527435303,17.8148441314697,17.8149909973145,17.8153324127197,17.8157215118408,17.8161125183105,17.81640625,17.6207523345947,17.4474601745605,17.25244140625,17.2541007995605,17.255859375,17.2364253997803,17.1998043060303,17.1122550964355,16.9761714935303,16.8678226470947,16.787109375,16.7081069946289,16.6309070587158,16.5651359558105,16.5107421875,16.4678211212158,16.4395523071289,16.3910160064697,16.351318359375,16.2613773345947,16.162353515625,16.0727062225342,16.0711917877197,16.071044921875,16.07080078125,16.0706539154053,16.0704574584961,16.0701656341553,15.9323720932007,15.7032232284546,15.4955558776855,15.3208980560303,15.2749996185303,15.2376947402954,15.07421875,15.0267572402954,15.0019540786743,14.9635753631592,14.9013175964355,14.8183603286743,14.7613277435303,14.691014289856,14.6300773620605,14.5709962844849,14.54541015625,14.5677738189697,14.8396482467651,15.1385746002197,15.2361316680908,15.3102531433105,15.4480476379395,15.7503910064697,15.8884763717651,16.0535640716553,16.145263671875,16.205078125,16.2393550872803,16.2847652435303,16.2873554229736,16.2262706756592,16.1954097747803,16.1693363189697,16.1675281524658,16.1456050872803,16.0620613098145,16.0189456939697,16.1865711212158,16.2019062042236,16.2282218933105,16.3158187866211,16.3475589752197,16.351806640625,16.2912101745605,16.2871589660645,16.3270492553711,16.3645992279053,16.4197273254395,16.41748046875,16.3791007995605,16.3062496185303,16.2776374816895,16.24658203125,16.2291011810303,16.2096672058105,16.2100086212158,16.1769523620605,15.974705696106,15.8877935409546,15.6930665969849,15.6831064224243,15.6519041061401,15.7264165878296,15.9092779159546,15.9668464660645,16.206298828125,16.3048343658447,16.5347652435303,16.5445804595947,16.5814456939697,16.6647472381592,16.7196292877197,16.9205074310303,16.9841804504395,17.0640640258789,17.2004871368408,17.276123046875,17.3931159973145,17.5142078399658,17.6153316497803,17.6515617370605,17.8419933319092,17.9222660064697,17.9597663879395,17.9727058410645,17.9574222564697,18.0414066314697,18.0628414154053,18.1868667602539,18.3253898620605,18.484375,18.6329593658447,18.8284664154053,18.9118156433105,19.010986328125,19.0912094116211,19.1528797149658,19.3093757629395,19.4432621002197,19.5622062683105,19.7064933776855,19.97607421875,20.0753917694092,20.2278327941895,20.3163089752197,20.3855953216553,20.4359855651855,20.4979496002197,20.5118656158447,20.5790519714355,20.6341781616211,20.6685066223145,20.7529773712158,20.775390625,20.7766132354736,20.8087406158447,20.8437995910645,20.9261226654053,21.0265636444092,21.1191883087158,21.2497062683105,21.3804206848145,21.4908199310303,21.6182632446289,21.6724624633789,21.8184566497803,21.9880867004395,22.326904296875,22.6274909973145,22.7770023345947,22.8290538787842,23.0609378814697,23.1956062316895,23.449462890625,23.6106929779053,23.8812484741211,24.01611328125,24.471923828125,24.471923828125,24.3600597381592,24.369384765625,24.4239749908447,24.4891605377197,24.5047855377197,24.4901371002197,24.5035648345947,24.5250492095947,24.5390129089355,24.6148910522461,24.6935558319092,24.7833976745605,24.9748058319092,25.08154296875,25.0736808776855,25.0306644439697,25.0184097290039,25.01806640625,25.0353507995605,25.0670413970947,25.093505859375,25.1943359375,25.26513671875,25.3829116821289,25.4242191314697,25.5380363464355,25.5433101654053,25.5115718841553,25.48046875,25.5166988372803,25.5515613555908,25.5884780883789,25.696044921875,25.7334461212158,25.6902828216553,25.6419925689697,25.6150398254395,25.592529296875,25.6087894439697,25.6331539154053,25.7271480560303,26.0325679779053,26.1384773254395,26.2431144714355,26.3052253723145,26.2583503723145,26.2527332305908,26.3057117462158,26.3552742004395,26.4046859741211,26.449951171875,26.5338859558105,26.7103519439697,26.6968250274658,26.7029304504395,26.7701168060303,26.8833999633789,26.9781723022461,27.0286617279053,27.079345703125,27.1622066497803,27.2333011627197,27.3226547241211,27.3956050872803,27.4501457214355,27.5443363189697,27.6539058685303,27.795654296875,27.8642101287842,27.9151859283447,27.9175777435303,27.8888683319092,27.9259757995605,27.9669914245605,28.115234375,28.3839855194092,28.4705562591553,28.5639629364014,28.6481456756592,28.7524909973145,28.7979011535645,28.8231925964355,28.8958969116211,29.0188961029053,29.1179676055908,29.2694816589355,29.3229007720947,29.3477039337158,29.4197254180908,29.4601078033447,29.5364265441895,29.7195320129395,29.8700675964355,29.9168472290039,29.9854488372803,30.1256828308105,30.3001461029053,30.5100116729736,30.6510238647461,30.7933101654053,30.9380855560303,31.0271968841553,31.0480976104736,31.0772953033447,31.0608901977539,31.0870113372803,31.1792469024658,31.2071781158447,31.2360363006592,31.2559566497803,31.2936038970947,31.3458995819092,31.4676265716553,31.5233421325684,31.5577621459961,31.6293449401855,31.5927257537842,31.5251483917236,31.5103511810303,31.5073719024658,31.554443359375,31.7335433959961,31.7622547149658,31.7774410247803,31.8064937591553,31.9007339477539,31.8506336212158,31.7985363006592,31.6471176147461,31.5379409790039,31.1563968658447,31.0804691314697,30.958740234375,30.7651863098145,30.6211929321289,30.5068855285645,30.2381362915039,30.1562995910645,30.0222644805908,29.8964862823486,29.8302268981934,29.7343273162842,29.6095218658447,29.439453125,29.3674812316895,29.1022434234619,29.0233898162842,28.9267082214355,28.9466800689697,28.873046875,28.8390636444092,28.8131351470947,28.8188495635986,28.7987785339355,28.5117683410645,28.4726085662842,28.4558582305908,28.4242172241211,28.3506355285645,28.2919921875,28.2071285247803,28.0921878814697,27.994873046875,27.900634765625,27.8259773254395,27.657470703125,27.5234375,27.3474617004395,27.1866703033447,27.1459465026855,27.0791015625,27.0097160339355,26.9670886993408,26.8401870727539,26.6785163879395,26.5727043151855,26.5644035339355,26.5809574127197,26.687255859375,26.7562503814697,26.8650875091553,26.8796863555908,26.7076168060303,26.5791988372803,26.5066413879395,26.4084491729736,26.3499507904053,26.2650375366211,26.1254386901855,25.9313468933105,25.789794921875,25.5726070404053,25.5269527435303,25.4203128814697,25.1442375183105,24.9945812225342,24.86767578125,24.7885246276855,24.6715335845947,24.58984375,24.3414554595947,24.2141590118408,24.1833972930908,24.1651382446289,24.1309566497803,24.1004886627197,24.1394519805908,24.1948719024658,24.2591781616211,24.3059577941895,24.3394546508789,24.3445301055908,24.1746101379395,24.109375,24.0330066680908,23.9390144348145,23.8648929595947,23.8038082122803,23.6615715026855,23.597900390625,23.4801254272461,23.4055652618408,23.2147464752197,23.1598148345947,23.0786628723145,22.9818363189697,22.9221687316895,22.8858890533447,22.89404296875,23.0054683685303,23.3415031433105,23.4122562408447,23.5176753997803,23.6049327850342,23.7373046875,23.8770027160645,23.9702644348145,24.1052722930908,24.3290042877197,24.4430160522461,24.5558109283447,24.5541515350342,24.5425300598145,24.5733890533447,24.6317882537842,24.6700687408447,24.8400382995605,24.9340343475342,25.0431156158447,25.323974609375,25.4882316589355,25.5728511810303,25.5843753814697,25.63037109375,25.7655277252197,25.91259765625,26.2139167785645,26.2734851837158,26.3167476654053,26.583251953125,26.7165031433105,26.7921867370605,26.9462394714355,26.8569812774658,26.7909660339355,26.7958011627197,26.7212886810303,26.7913570404053,26.870849609375,26.9665031433105,26.9852542877197,26.9876937866211,27.10595703125,27.1435070037842,27.1580066680908,27.2181644439697,27.2835941314697,27.376220703125,27.4311027526855,27.53955078125,27.6591796875,27.7360363006592,27.7988758087158,27.8418445587158,27.8299312591553,27.783935546875,27.796875,27.8412094116211,27.8729972839355,27.8385734558105,27.7757320404053,27.7181148529053,27.6714363098145,27.6756839752197,27.7266120910645,27.83056640625,27.8679695129395,27.919677734375,27.9080085754395,27.9344730377197,28.0132808685303,28.2213382720947,28.4261722564697,28.6054191589355,28.7299308776855,29.0945816040039,29.2818851470947,29.3516101837158,29.3844223022461,29.4272441864014,29.5319347381592,29.6800308227539,29.7563953399658,29.9357433319092,29.960205078125,30.0841789245605,30.3036136627197,30.3598155975342,30.4144535064697,30.5635738372803,30.7054672241211,30.8041496276855,30.9705085754395,31.0509757995605,31.1273441314697,31.2027835845947,31.3609886169434,31.4990730285645,31.5648937225342,31.6986312866211,31.7345714569092,31.74365234375,31.7403335571289,31.7580070495605,31.85107421875,31.9973640441895,32.1985359191895,32.3050308227539,32.3436012268066,32.5333480834961,32.5547866821289,32.5762214660645,32.5976066589355,32.6190452575684,32.6404762268066,32.661865234375,32.6833038330078,32.7047348022461,32.71533203125,32.5647964477539,32.50830078125,32.3603019714355,32.2123069763184,32.0643043518066,31.9162578582764,31.7682609558105,31.6202144622803,31.4722652435303,31.3242168426514,31.3248519897461,31.3255348205566,31.3261699676514,31.3268051147461,31.3274421691895,31.328125,31.3288078308105,31.3294429779053,31.44189453125,31.5543956756592,31.6668472290039,31.7793464660645,31.7781715393066,31.7770519256592,31.7758769989014,31.7747573852539,31.7736339569092,31.7724628448486,31.7713394165039,31.7701663970947,31.7684078216553,31.7644538879395,31.6790046691895,31.5446796417236,31.450927734375,31.3977546691895,31.2410163879395,30.9807605743408,30.8072738647461,30.7205543518066,30.6459445953369,30.5833511352539,30.4476547241211,30.1343746185303,29.9905281066895,29.8542957305908,29.6776847839355,29.5737323760986,29.5424327850342,29.4798812866211,29.3861351013184,29.3231468200684,29.2910652160645,29.2068862915039,29.0707015991211,29.0011234283447,28.9981937408447,29.0418968200684,29.1322250366211,29.1903800964355,29.2164077758789,29.2580070495605,29.3153324127197,29.4439449310303,29.6439456939697,29.7523441314697,29.7690925598145,29.8066425323486,29.8649921417236,29.8711910247803,29.8252429962158,29.7957038879395,29.7824687957764,29.7869644165039,29.8092308044434,29.8080558776855,29.7835445404053,29.7731437683105,29.77685546875,29.7425785064697,29.6340808868408,29.4604015350342,29.4603023529053,29.4006824493408,29.314697265625,29.1825199127197,29.0685539245605,28.9728012084961,28.8213386535645,28.6142082214355,28.5025386810303,28.4864253997803,28.4281253814697,28.3276863098145,28.2426280975342,28.1729488372803,28.0478515625,27.8672847747803,27.7299308776855,27.6358871459961,27.54833984375,27.4673843383789,27.3980464935303,27.34033203125,27.2854976654053,27.2336902618408,27.1701164245605,27.0816898345947,27.0566902160645,27.056640625,27.0366706848145,26.8847179412842,26.7619132995605,26.56591796875,26.5641593933105,26.4469242095947,26.3989753723145,26.3812503814697,26.3404293060303,26.2764644622803,26.2378425598145,26.2245616912842,26.182373046875,26.1111812591553,26.064453125,26.0420417785645,25.9841785430908,25.8908214569092,25.871826171875,25.8705081939697,25.884765625,25.9111824035645,25.9416007995605,25.9614734649658],[5.830078125,5.7998046875,5.80190420150757,5.82441377639771,5.85581016540527,5.94511699676514,5.97705125808716,6.01416063308716,5.97568321228027,5.93520498275757,5.830078125],[7.04824209213257,7.026611328125,7.02827167510986,6.97314453125,6.97661113739014,7.00014591217041,7.01005840301514,7.00717782974243,7.01625967025757,7.04824209213257],[7.13823223114014,7.08696317672729,7.11093711853027,7.09555673599243,7.08115196228027,7.06874990463257,7.06874990463257,7.07353496551514,7.10927772521973,7.15610361099243,7.17177724838257,7.13823223114014],[7.30898475646973,7.29355430603027,7.30273389816284,7.32192373275757,7.33623123168945,7.32246065139771,7.30898475646973],[11.153076171875,11.146240234375,11.1533689498901,11.1663084030151,11.1686525344849,11.1650390625,11.153076171875],[42.31396484375,42.1650886535645,42.1048355102539,42.0591316223145,42.0256843566895,41.9936065673828,41.835205078125,41.7750968933105,41.7571792602539,41.7398414611816,41.7169456481934,41.6056175231934,41.3561019897461,41.3362770080566,41.3373527526855,41.3319816589355,41.312744140625,41.1785163879395,41.1401863098145,41.1185073852539,41.1233863830566,41.1551742553711,41.1586418151855,41.1405296325684,41.1309547424316,41.107421875,40.950439453125,40.8963394165039,40.8689460754395,40.9036140441895,40.9071769714355,40.8671379089355,40.8631324768066,40.8561515808105,40.8499031066895,40.8715362548828,40.9031257629395,40.9179229736328,40.9052734375,40.9283676147461,41.0616683959961,41.0830574035645,41.1278305053711,41.2726058959961,41.3360824584961,41.3914070129395,41.5212898254395,41.5541038513184,41.5747566223145,41.6270484924316,41.7064933776855,41.8623504638672,41.8736801147461,41.8662109375,41.8538093566895,41.87353515625,41.904296875,42.0183601379395,42.071044921875,42.1712875366211,42.1749992370605,42.1289558410645,42.0995101928711,42.1003913879395,42.1300773620605,42.1871604919434,42.2198257446289,42.2476577758789,42.2475090026855,42.2421379089355,42.2677230834961,42.3031234741211,42.3083992004395,42.3220672607422,42.320068359375,42.3046379089355,42.3249969482422,42.358154296875,42.349853515625,42.3217277526855,42.31396484375],[19.1427726745605,18.9680671691895,18.7043476104736,18.4105949401855,18.1394519805908,17.8305187225342,17.5821781158447,17.2884273529053,16.9963855743408,16.9626941680908,16.7981929779053,16.581787109375,16.3577136993408,16.1927242279053,16.0355472564697,15.9456539154053,15.8968267440796,15.8379878997803,15.7552738189697,15.7017087936401,15.6740236282349,15.6416988372803,15.4831047058105,15.3563480377197,15.3911123275757,15.4271984100342,15.4248533248901,15.4083003997803,15.3409671783447,15.3298835754395,15.3204097747803,15.3093757629395,15.3017082214355,15.2864742279053,15.2722654342651,15.1261234283447,14.9869136810303,14.9821290969849,14.9548816680908,14.9790048599243,14.9801759719849,14.9636726379395,14.9114742279053,14.9848155975342,14.987060546875,15.0594234466553,15.0124998092651,15.0285167694092,15.0596675872803,15.0778818130493,15.0697755813599,15.0477533340454,14.9374017715454,14.8413572311401,14.8195314407349,14.7615242004395,14.6260747909546,14.5268068313599,14.5084962844849,14.4860353469849,14.4814929962158,14.45654296875,14.1946287155151,14.16845703125,14.2741212844849,14.25830078125,14.2275876998901,14.07373046875,13.9507322311401,13.7867679595947,13.7363767623901,13.6794929504395,13.6484375,13.6371097564697,13.6391115188599,13.6728515625,13.6583499908447,13.5774421691895,13.4007816314697,13.28076171875,13.2437009811401,13.1963863372803,13.1827154159546,13.1941900253296,13.37353515625,13.4021968841553,13.3824224472046,13.3062009811401,13.2561521530151,13.1973142623901,13.1190423965454,13.052490234375,12.975341796875,12.8794918060303,12.793701171875,12.6722173690796,12.63037109375,12.5815916061401,12.4930667877197,12.3375978469849,12.2817869186401,12.2264642715454,12.155029296875,12.1202154159546,12.076171875,12.0321292877197,11.9933109283447,11.967529296875,11.9423828125,11.8902826309204,11.8279294967651,11.7604494094849,11.6833009719849,11.619873046875,11.5767574310303,11.5224609375,11.3757820129395,11.2059574127197,11.1302738189697,11.0887203216553,11.0423831939697,10.9310550689697,10.7713871002197,10.6439456939697,10.5650873184204,10.4834470748901,10.426025390625,10.43994140625,10.4332036972046,10.3895511627197,10.3547353744507,10.3072271347046,10.2750978469849,10.2391119003296,10.19482421875,10.201904296875,10.2321290969849,10.2616205215454,10.2791986465454,10.3223628997803,10.3694334030151,10.4002933502197,10.4762697219849,10.55810546875,10.5975093841553,10.7179203033447,10.7240724563599,10.6928215026855,10.6851081848145,10.6717767715454,10.6487302780151,10.5723638534546,10.5591306686401,10.5612306594849,10.58642578125,10.6564455032349,10.6337890625,10.5780277252197,10.5120124816895,10.3921871185303,10.3494625091553,10.3571290969849,10.3569822311401,10.3450679779053,10.3423337936401,10.299560546875,10.2518548965454,10.19873046875,10.1762208938599,10.1556644439697,10.1432609558105,10.144775390625,10.2035150527954,10.2256832122803,10.2593746185303,10.3401365280151,10.34130859375,10.3839349746704,10.4397945404053,10.4368171691895,10.4212398529053,10.4274415969849,10.34228515625,10.236572265625,10.1857423782349,10.1634769439697,10.1625003814697,10.2295408248901,10.2784175872803,10.3218746185303,10.43798828125,10.4859867095947,10.6175785064697,10.74951171875,10.8269529342651,10.8960943222046,10.9497556686401,10.9906244277954,11.0299320220947,11.0483884811401,10.9966793060303,10.9869632720947,10.990478515625,11.0094728469849,11.0358400344849,11.177001953125,11.2359371185303,11.2807130813599,11.3047361373901,11.3394041061401,11.3665523529053,11.3862791061401,11.4122066497803,11.4780759811401,11.485107421875,11.5149898529053,11.6177740097046,11.6374998092651,11.6482419967651,11.6732425689697,11.80712890625,11.9225101470947,12.1085453033447,12.2255859375,12.3458986282349,12.40234375,12.449951171875,12.4828615188599,12.4902830123901,12.4792966842651,12.4646482467651,12.4422359466553,12.3660154342651,12.3237295150757,12.2827644348145,12.2554206848145,12.25244140625,12.2286624908447,12.1824703216553,12.1431150436401,12.0424795150757,12.0299320220947,12.0424795150757,12.116455078125,12.1774415969849,12.212646484375,12.1902837753296,12.1795415878296,12.138671875,11.9902839660645,11.9412107467651,11.9255380630493,11.8994140625,11.8987302780151,11.9164552688599,11.92724609375,12.0342769622803,12.15185546875,12.2051753997803,12.20751953125,12.1708011627197,12.0950193405151,12.0248537063599,12.0040521621704,12.0154294967651,12.10400390625,12.1669425964355,12.1986331939697,12.2471685409546,12.3192386627197,12.377685546875,12.4043941497803,12.479248046875,12.5319337844849,12.5577154159546,12.6275882720947,12.7754878997803,12.8318853378296,12.9419927597046,12.9916009902954,13.0282230377197,13.0869626998901,13.1702632904053,13.2369632720947,13.2900390625,13.369873046875,13.3823728561401,13.39453125,13.3670902252197,13.3272943496704,13.3158197402954,13.3645505905151,13.4062986373901,13.4444332122803,13.5108880996704,13.633056640625,13.73388671875,13.7880859375,13.8289546966553,13.8752927780151,13.9307613372803,13.9746580123901,14.0718259811401,14.2064943313599,14.2742195129395,14.3232908248901,14.3766603469849,14.4585933685303,14.5711431503296,14.6481447219849,14.8090333938599,14.7453603744507,14.7663583755493,14.804931640625,14.8869142532349,14.9951667785645,15.12939453125,15.244873046875,15.3427238464355,15.4255361557007,15.5120601654053,15.5732431411743,15.6368169784546,15.6253900527954,15.5367679595947,15.3586416244507,15.222900390625,15.151123046875,15.1504878997803,15.2817392349243,15.3949222564697,15.4226560592651,15.4348630905151,15.4397945404053,15.4379386901855,15.416015625,15.3960456848145,15.3836917877197,15.373779296875,15.4014654159546,15.437255859375,15.4582033157349,15.5116701126099,15.5748529434204,15.623046875,15.6676273345947,15.6773929595947,15.5256843566895,15.5028324127197,15.49609375,15.49609375,15.49609375,15.4961423873901,15.4961423873901,15.4961423873901,15.4961423873901,15.4961423873901,15.4961423873901,15.4961910247803,15.4961910247803,15.4961910247803,15.4961910247803,15.4961910247803,15.4961910247803,15.4962396621704,15.4962892532349,15.4962892532349,15.4962892532349,15.7894048690796,16.0579090118408,16.2828617095947,16.4420413970947,16.5686531066895,16.8095703125,17.0582523345947,17.306884765625,17.5555667877197,17.8042469024658,18.0529289245605,18.3015613555908,18.5502433776855,18.7988758087158,19.0475082397461,19.2961902618408,19.5448722839355,19.7935066223145,20.0421886444092,20.2908687591553,20.5395030975342,20.7881851196289,21.036865234375,21.2855472564697,21.5341796875,21.7828617095947,22.0315437316895,22.2801761627197,22.5288581848145,22.7774906158447,23.026123046875,23.2748031616211,23.5234851837158,23.7721672058105,24.0207996368408,24.2694816589355,24.5181636810303,24.7667961120605,24.99462890625,24.9948253631592,24.9949722290039,24.9951667785645,24.9954109191895,24.99560546875,24.8044929504395,24.62353515625,24.4094715118408,24.1953620910645,23.9812507629395,23.7671394348145,23.5530281066895,23.3388671875,23.1248035430908,22.91064453125,22.696533203125,22.4824714660645,22.268310546875,22.05419921875,21.8512420654297,21.8401355743408,21.6259746551514,21.411865234375,21.19775390625,21.1022453308105,21.0625,20.9819831848145,20.8913078308105,20.8174304962158,20.7672863006592,20.7135753631592,20.5555667877197,20.5243644714355,20.4588375091553,20.3783702850342,20.3315906524658,20.296875,20.272705078125,20.247802734375,20.2103023529053,20.0638675689697,20.0350093841553,19.992919921875,19.9694347381592,19.9559574127197,19.9166011810303,19.8501968383789,19.7896976470947,19.770751953125,19.7183113098145,19.5604000091553,19.4735851287842,19.4109382629395,19.3726081848145,19.3454093933105,19.3120613098145,19.2681636810303,19.212158203125,19.1500988006592,19.1031742095947,19.0729007720947,19.0132808685303,18.9883785247803,18.9866218566895,18.9884281158447,18.9961433410645,19.0416011810303,19.083740234375,19.1427726745605],[35.8527336120605,35.8202133178711,35.8216781616211,35.8722648620605,35.9784164428711,35.9574203491211,35.8862800598145,35.8527336120605],[36.027587890625,36.0121574401855,36.042236328125,36.0604019165039,36.0757827758789,36.0623054504395,36.0362319946289,36.027587890625],[9.93344688415527,9.87539100646973,9.87788105010986,9.97465801239014,10.0452146530151,10.0513181686401,10.0076169967651,9.95395469665527,9.93344688415527],[10.9527339935303,10.7796869277954,10.7531251907349,10.7398929595947,10.7444334030151,10.8377447128296,10.89794921875,10.8866205215454,10.963134765625,10.9803218841553,10.9527339935303],[10.9615240097046,10.9593753814697,10.9642572402954,10.979736328125,11.0869626998901,10.9615240097046],[11.4782228469849,11.4557619094849,11.4565439224243,11.472412109375,11.5671873092651,11.6447267532349,11.7583990097046,11.7830076217651,11.722900390625,11.5940923690796,11.5236320495605,11.4782228469849],[11.744873046875,11.5386724472046,11.5671873092651,11.5670890808105,11.6835451126099,11.71484375,11.79150390625,11.804931640625,11.744873046875],[11.6923828125,11.6368160247803,11.6958990097046,11.8602533340454,11.7566890716553,11.7332038879395,11.6923828125],[11.905029296875,11.8994140625,12.0612802505493,12.0842771530151,12.0051755905151,11.9566402435303,11.9250974655151,11.905029296875],[12.1504392623901,12.1448726654053,12.1480960845947,12.1644535064697,12.2324705123901,12.2800779342651,12.2917966842651,12.2877931594849,12.2787103652954,12.26123046875,12.223388671875,12.1913089752197,12.1504392623901],[12.3897953033447,12.353515625,12.2790040969849,12.3223142623901,12.3460941314697,12.393798828125,12.3870115280151,12.3978509902954,12.3897953033447],[12.5979490280151,12.5705080032349,12.5713376998901,12.4737300872803,12.3536624908447,12.3361825942993,12.3359870910645,12.5114250183105,12.611572265625,12.6781740188599,12.6471185684204,12.5979490280151],[13.0990724563599,12.9347171783447,13.0140132904053,13.1043939590454,13.1885738372803,13.2022457122803,13.1893558502197,13.1516599655151,13.0990724563599],[15.8193359375,15.7938480377197,15.8121099472046,15.8230466842651,15.9330072402954,15.89208984375,15.8193359375],[15.9459476470947,15.848388671875,15.9941396713257,16.0753421783447,16.15283203125,16.2055168151855,16.1413078308105,16.019287109375,15.9459476470947],[16.2532215118408,16.2401371002197,16.3057117462158,16.4610347747803,16.4968757629395,16.5050792694092,16.4860343933105,16.4607925415039,16.4295425415039,16.3973636627197,16.2532215118408],[18.6842784881592,18.6756839752197,18.7595710754395,18.8675308227539,18.8888187408447,18.8655281066895,18.8080558776855,18.7157230377197,18.6842784881592],[19.5582523345947,19.4752941131592,19.4810562133789,19.4286136627197,19.4081535339355,19.3654308319092,19.33203125,19.2986831665039,19.32568359375,19.4163093566895,19.4589366912842,19.4950695037842,19.54443359375,19.5582523345947],[19.892578125,19.7547855377197,19.8064460754395,19.8775882720947,19.9503421783447,19.892578125],[19.9239253997803,19.828857421875,19.8680171966553,19.9998054504395,20.0864753723145,20.0461921691895,19.9239253997803],[21.5674800872803,21.4686527252197,21.3662109375,21.314208984375,21.3126449584961,21.2513675689697,21.13037109375,21.059326171875,21.0381832122803,20.9923839569092,20.9219245910645,20.8842296600342,20.8792476654053,20.8595695495605,20.8250980377197,20.81298828125,20.8232421875,20.7957019805908,20.7302722930908,20.589111328125,20.3722171783447,20.316650390625,20.3795890808105,20.4154300689697,20.4244136810303,20.3844718933105,20.34130859375,20.325439453125,20.3204574584961,20.3428211212158,20.363037109375,20.35205078125,20.2606449127197,20.187744140625,20.1498546600342,20.1183109283447,20.0934562683105,20.0789051055908,20.0804195404053,20.1151371002197,20.1166019439697,20.099365234375,20.0736331939697,20.0417976379395,19.8613777160645,19.8049335479736,19.7729015350342,19.7695789337158,19.7784671783447,19.7710933685303,19.7013187408447,19.6944332122803,19.6891593933105,19.687255859375,19.690673828125,19.7621593475342,19.7697257995605,19.74951171875,19.6537113189697,19.5928707122803,19.4599609375,19.265869140625,19.1304683685303,18.9964847564697,18.9317874908447,18.6208000183105,18.5881843566895,18.572021484375,18.5612297058105,18.5287094116211,18.4977531433105,18.4942378997803,18.5175285339355,18.5179691314697,18.4942874908447,18.3596687316895,18.2958984375,18.302978515625,18.2903327941895,18.2580070495605,18.1737289428711,18.0374031066895,17.935302734375,17.8335456848145,17.797119140625,17.7758312225342,17.6812515258789,17.5333003997803,17.373291015625,17.2398929595947,17.1476554870605,16.9756832122803,16.89501953125,16.7322273254395,16.63818359375,16.5709476470947,16.5147953033447,16.3304214477539,16.305419921875,16.4175777435303,16.3941879272461,16.3519058227539,16.298095703125,16.237060546875,16.1808109283447,16.0506839752197,15.9386234283447,15.7686033248901,15.5597648620605,15.4035634994507,15.3676767349243,15.3506832122803,15.3573732376099,15.278564453125,15.2715826034546,15.2413568496704,15.2041015625,15.1474113464355,14.9759273529053,14.8147468566895,14.6964836120605,14.6029787063599,14.4728994369507,14.3599128723145,14.2357425689697,14.0498533248901,13.9471673965454,13.82275390625,13.7166996002197,13.5757808685303,13.4969244003296,13.2330570220947,13.1729984283447,13.103515625,13.03076171875,12.9613285064697,12.8819341659546,12.73974609375,12.6528806686401,12.59423828125,12.5479011535645,12.4736328125,12.3948240280151,12.3090333938599,12.1902351379395,12.0896482467651,11.7812013626099,11.7496585845947,11.6871576309204,11.6306638717651,11.6125001907349,11.5543947219849,11.3894529342651,11.1052742004395,10.9199705123901,10.788330078125,10.7084474563599,10.6609373092651,10.6235837936401,10.5570306777954,10.4308595657349,10.3508310317993,10.2660150527954,10.1903810501099,10.1790523529053,10.0349607467651,10.1072273254395,10.1825199127197,10.3531246185303,10.6758308410645,10.7189455032349,10.7406740188599,10.8644046783447,10.9869146347046,11.1331062316895,11.2403812408447,11.3299798965454,11.4352540969849,11.5212898254395,11.5916996002197,11.6650876998901,11.7197265625,11.7392578125,11.7792482376099,11.7183589935303,11.7383785247803,11.8014650344849,11.869140625,11.9103031158447,11.9567375183105,12.047119140625,12.126708984375,12.2252435684204,12.2254877090454,12.2453126907349,12.3000001907349,12.3484859466553,12.4407224655151,12.5399408340454,12.6624021530151,12.7705078125,12.8482427597046,12.9860353469849,13.1619148254395,13.2930669784546,13.4837884902954,13.7334957122803,13.8403806686401,13.9344730377197,13.9801759719849,13.6476068496704,13.712890625,13.8983392715454,13.9864749908447,14.1615228652954,14.335301399231,14.4614744186401,14.6526851654053,14.695556640625,14.652587890625,14.7639150619507,14.7387199401855,14.8589344024658,15.1849126815796,15.3067865371704,15.4309577941895,15.8755369186401,16.0195808410645,16.1438465118408,16.2538566589355,16.4576644897461,16.5204582214355,16.5685558319092,16.5516109466553,16.5372066497803,16.5252933502197,16.52294921875,16.6717777252197,16.7431163787842,16.892578125,17.06201171875,17.0954093933105,17.16455078125,17.2069339752197,17.3173351287842,17.4010257720947,17.3421859741211,17.3048343658447,17.2029285430908,17.0309562683105,16.9211902618408,16.7783679962158,16.7103519439697,16.5639152526855,16.5143566131592,16.5049324035645,16.5205078125,16.5959949493408,16.6591300964355,16.7653312683105,16.7805652618408,16.768310546875,16.6312484741211,16.5674324035645,16.4444332122803,16.4100589752197,16.342529296875,16.3533706665039,16.3399410247803,16.2846183776855,16.2537097930908,16.1690444946289,16.0733890533447,15.9767580032349,15.8378429412842,15.7227535247803,15.7292976379395,15.7561521530151,15.8804206848145,15.9854497909546,16.0976066589355,16.0332527160645,15.8768072128296,15.82568359375,15.8391609191895,15.8182621002197,15.9791011810303,16.0381832122803,16.0879383087158,16.1408195495605,16.1828136444092,16.1024398803711,16.0328617095947,15.9710941314697,15.9043941497803,15.98876953125,16.0648441314697,16.13330078125,16.2420406341553,16.3987312316895,16.4524898529053,16.5119132995605,16.4255847930908,16.3361320495605,16.30908203125,16.288818359375,16.1861324310303,16.0943851470947,16.0076179504395,16.0164546966553,16.1266117095947,16.5172863006592,16.5721683502197,16.6399402618408,16.8681640625,16.9544906616211,17.1354484558105,17.1665534973145,17.3085441589355,17.5693359375,17.698974609375,17.8246097564697,18.20166015625,18.5072269439697,18.6091804504395,18.732421875,18.7411613464355,18.8492183685303,18.8934078216553,19.1461925506592,19.2874031066895,19.268798828125,19.1817874908447,19.1460933685303,18.9950695037842,18.9583988189697,18.899658203125,18.9605941772461,19.0269031524658,19.1884765625,19.3694820404053,19.3975582122803,19.4011726379395,19.2665042877197,19.2384757995605,19.2719230651855,19.329345703125,19.4408702850342,19.481689453125,19.50390625,19.5341300964355,19.6095695495605,19.6480464935303,19.697265625,19.7319831848145,19.7760753631592,19.8541507720947,19.9121589660645,19.9095687866211,20.0094242095947,20.038330078125,20.0583000183105,20.0701179504395,20.040771484375,19.8983402252197,19.8512210845947,19.8580074310303,20.0748538970947,20.1297855377197,20.1813468933105,20.1886711120605,20.1852550506592,20.3776378631592,20.4061527252197,20.3460445404053,20.2879867553711,20.1521492004395,20.1775875091553,20.21142578125,20.2826175689697,20.3017578125,20.34033203125,20.4148445129395,20.4690418243408,20.5626964569092,20.56396484375,20.453369140625,20.2956047058105,20.4698734283447,20.7175769805908,20.7918453216553,20.8644523620605,20.9315910339355,21.0046882629395,21.0614738464355,21.1126956939697,21.2022457122803,21.2931156158447,21.3578605651855,21.4275875091553,21.4397945404053,21.4090328216553,21.3629875183105,21.31982421875,21.2633304595947,21.2701644897461,21.3062019348145,21.3507328033447,21.4673347473145,21.6090335845947,21.9403324127197,21.9780769348145,22.0113277435303,22.04931640625,22.1060066223145,22.1309566497803,22.1324214935303,22.1047859191895,22.0101566314697,21.9889163970947,22.0037593841553,22.1457042694092,22.1839847564697,22.2094230651855,22.2051753997803,22.2306137084961,22.2918949127197,22.3602046966553,22.547119140625,22.6332530975342,22.7182140350342,22.8057136535645,22.907958984375,22.997314453125,23.0320320129395,23.0370121002197,23.0154781341553,23.0303707122803,23.0849628448486,23.1325187683105,23.3391590118408,23.5280284881592,23.6820793151855,23.774169921875,24.0218734741211,24.064208984375,24.0741214752197,23.9874019622803,23.9728507995605,23.986083984375,24.00537109375,24.0065441131592,23.9769039154053,23.9438953399658,23.9029293060303,23.8720703125,23.87646484375,23.97265625,24.1131839752197,24.3218746185303,24.4737300872803,24.5140628814697,24.6376457214355,24.7672367095947,24.9310073852539,25.0487308502197,25.0978507995605,25.1385746002197,25.1645984649658,25.1915035247803,25.2157230377197,25.2434577941895,25.31982421875,25.4100093841553,25.4588871002197,25.5193367004395,25.5972156524658,25.7002449035645,25.7704582214355,25.9129390716553,25.9413089752197,25.9873046875,26.041259765625,26.0704097747803,26.0914058685303,26.1911125183105,26.3472652435303,26.4739742279053,26.5254878997803,26.5972652435303,26.6414070129395,26.6722660064697,26.7560558319092,26.950439453125,27.0138187408447,27.046630859375,27.1280765533447,27.2170906066895,27.2612800598145,27.2783679962158,27.3392581939697,27.3314933776855,27.2961921691895,27.1778316497803,27.13330078125,27.10205078125,27.1154289245605,27.1633319854736,27.4395999908447,27.5148429870605,27.5867195129395,27.6438484191895,27.6982917785645,27.760009765625,27.8368644714355,27.8900394439697,27.9070796966553,27.937744140625,27.9823246002197,28.0308589935303,28.0859870910645,28.1552238464355,28.2179679870605,28.2544937133789,28.3539066314697,28.4256362915039,28.4563465118408,28.5102043151855,28.5170402526855,28.5,28.4693374633789,28.4071292877197,28.3561515808105,28.3563480377197,28.3635749816895,28.3564949035645,28.3138179779053,28.2115230560303,28.1858882904053,28.1422843933105,28.0552234649658,27.9675769805908,27.6631832122803,27.5990715026855,27.5500965118408,27.5380859375,27.5870609283447,27.6394538879395,27.6572265625,27.6476554870605,27.5988273620605,27.5724601745605,27.4219245910645,27.2453117370605,27.1906242370605,27.044921875,26.8773918151855,26.7857418060303,26.6981430053711,26.5833988189697,26.4296894073486,26.2985343933105,26.1893539428711,26.1394538879395,26.1140613555908,26.0724124908447,26.0037097930908,25.9177722930908,25.8635749816895,25.8267097473145,25.8232421875,25.7888660430908,25.677978515625,25.5867671966553,25.56884765625,25.5945301055908,25.5710945129395,25.4157238006592,25.3489742279053,25.2925281524658,25.2593269348145,25.2361335754395,25.2518539428711,25.1580562591553,25.0343265533447,24.9703598022461,24.869873046875,24.8419914245605,24.8201160430908,24.7748031616211,24.6312007904053,24.49169921875,24.44384765625,24.4229507446289,24.3799819946289,24.312744140625,24.228759765625,24.1308116912842,23.9884757995605,23.9110355377197,23.8871574401855,23.8980960845947,23.931884765625,23.9862785339355,24.0654296875,24.1106452941895,24.1190414428711,24.1156730651855,24.0988292694092,24.06982421875,24.1160640716553,24.1187000274658,24.1211929321289,24.090576171875,23.9640617370605,23.9050788879395,23.841796875,23.7831058502197,23.7378425598145,23.6243667602539,23.5204105377197,23.4825191497803,23.4400882720947,23.3803215026855,23.3074703216553,23.1912593841553,23.1305675506592,23.1033191680908,23.0958995819092,23.0692386627197,23.0462398529053,23.0045890808105,22.9591312408447,22.9272937774658,22.8250980377197,22.6886730194092,22.5865230560303,22.498046875,22.370361328125,22.2825679779053,22.1924800872803,22.1533203125,22.1259765625,22.1101551055908,22.1006336212158,22.1107902526855,22.0891590118408,22.0497074127197,22.0280284881592,21.9883308410645,21.901611328125,21.82080078125,21.7587413787842,21.7016105651855,21.6827640533447,21.6606464385986,21.6170406341553,21.5579090118408,21.5111827850342,21.4805183410645,21.4629878997803,21.5010261535645,21.4840812683105,21.4581050872803,21.4717769622803,21.5049304962158,21.6551761627197,21.7363777160645,21.755859375,21.74609375,21.7355480194092,21.7051277160645,21.5816402435303,21.5674800872803],[43.5332984924316,43.5210456848145,43.4850120544434,43.449951171875,43.4139671325684,43.3428230285645,43.2122573852539,43.1734390258789,43.1639633178711,43.1097679138184,43.0965347290039,43.0816383361816,42.968505859375,42.9354515075684,42.8928718566895,42.852783203125,42.8279304504395,42.798828125,42.754638671875,42.759765625,42.7600593566895,42.7320289611816,42.6858406066895,42.6315460205078,42.5570831298828,42.5472679138184,42.5499038696289,42.5066871643066,42.486328125,42.4761695861816,42.4969253540039,42.525146484375,42.60693359375,42.634521484375,42.64794921875,42.6285667419434,42.5654296875,42.491943359375,42.4153823852539,42.3418922424316,42.249267578125,42.1725578308105,42.1292953491211,42.069091796875,42.0240211486816,41.9977531433105,41.9188499450684,41.8690910339355,41.9486312866211,42.0605010986328,42.2494621276855,42.3780746459961,42.3983879089355,42.4231452941895,42.4427223205566,42.4441871643066,42.4285163879395,42.4329071044922,42.4811019897461,42.52294921875,42.5597190856934,42.5645027160645,42.5791969299316,42.6201171875,42.6416015625,42.6741714477539,42.7772445678711,42.8440933227539,42.9684562683105,42.9978981018066,43.0121574401855,43.0276832580566,43.1246070861816,43.1536598205566,43.1939468383789,43.2308120727539,43.2835426330566,43.3463363647461,43.3481903076172,43.3394508361816,43.285400390625,43.2924308776855,43.3573226928711,43.4423828125,43.4967269897461,43.5266609191895,43.5423316955566,43.5325164794922,43.5177230834961,43.5277328491211,43.5354499816895,43.5332984924316],[49.8237762451172,49.684814453125,49.4062004089355,49.1703605651855,49.0374526977539,48.9361343383789,48.8400382995605,48.7822761535645,48.6893539428711,48.5772476196289,48.4557113647461,48.3463401794434,48.2482414245605,48.1862297058105,48.130859375,47.9450187683105,47.8748054504395,47.7989234924316,47.7382316589355,47.6869163513184,47.7113265991211,47.78955078125,47.8582038879395,47.85986328125,47.8440437316895,47.8395500183105,47.864501953125,47.8697738647461,47.8530769348145,47.8365707397461,47.806396484375,47.7402839660645,47.6663551330566,47.6521987915039,47.6757316589355,47.7413597106934,47.8046875,47.9082984924316,47.9878921508789,47.9998550415039,47.9996070861816,48.0189437866211,48.0289077758789,47.99951171875,47.9839859008789,47.9432640075684,47.822265625,47.7576179504395,47.72509765625,47.7029304504395,47.6853523254395,47.6541519165039,47.6162605285645,47.5584945678711,47.5251922607422,47.4925804138184,47.47265625,47.4307136535645,47.41015625,47.380859375,47.2559051513672,47.2224578857422,47.1500015258789,47.0900382995605,47.0270042419434,46.9788093566895,46.9065895080566,46.8578109741211,46.7914543151855,46.7328605651855,46.6721687316895,46.627197265625,46.6060028076172,46.6039543151855,46.6266593933105,46.6138191223145,46.638671875,46.6921882629395,46.73486328125,46.7602043151855,46.7470703125,46.69189453125,46.70166015625,46.69189453125,46.7031745910645,46.717041015625,46.6785659790039,46.6666030883789,46.6193351745605,46.5376968383789,46.5181655883789,46.5220718383789,46.552001953125,46.5882797241211,46.5862312316895,46.5708961486816,46.53759765625,46.4366683959961,46.3913078308105,46.3620109558105,46.3522453308105,46.3550758361816,46.3617668151855,46.387939453125,46.3766593933105,46.3219757080078,46.3130836486816,46.289794921875,46.2090797424316,46.1587905883789,46.0965805053711,45.9630355834961,45.8869132995605,45.8457527160645,45.7959938049316,45.7393569946289,45.6867179870605,45.6769523620605,45.6261711120605,45.5348167419434,45.458251953125,45.4395027160645,45.4199676513672,45.3961944580078,45.3902320861816,45.3782691955566,45.4196281433105,45.4132804870605,45.3899917602539,45.3646011352539,45.3163070678711,45.271728515625,45.2025871276855,45.1108894348145,45.0498542785645,44.9711418151855,44.9425773620605,44.9123039245605,44.8961944580078,44.825927734375,44.7634735107422,44.7457046508789,44.7623558044434,44.7674331665039,44.7916488647461,44.7948265075684,44.8103523254395,44.8834457397461,44.9176788330078,45.0109367370605,45.0582046508789,45.0630378723145,45.0629386901855,45.0816383361816,45.0640602111816,44.9695320129395,44.899169921875,44.8271484375,44.6729011535645,44.56982421875,44.5115699768066,44.419189453125,44.3672866821289,44.3223648071289,44.2716293334961,44.19189453125,44.1071281433105,44.0411148071289,43.9346694946289,43.87890625,43.81494140625,43.75244140625,43.71142578125,43.68017578125,43.6646003723145,43.6211433410645,43.5631828308105,43.4962921142578,43.4927749633789,43.4749031066895,43.3919944763184,43.3687515258789,43.3414077758789,43.2568855285645,43.194091796875,43.1107902526855,43.0738792419434,42.9905281066895,42.895263671875,42.8441390991211,42.8135719299316,42.773681640625,42.7427253723145,42.7100105285645,42.6605949401855,42.6062469482422,42.5538101196289,42.5105476379395,42.4559555053711,42.4383811950684,42.4405746459961,42.4264640808105,42.4161148071289,42.4292945861816,42.436767578125,42.4473152160645,42.4271965026855,42.4058570861816,42.4009780883789,42.3492660522461,42.321533203125,42.3088874816895,42.2887191772461,42.2635765075684,42.2273445129395,42.2115745544434,42.1405754089355,41.9939956665039,41.8750991821289,41.854736328125,41.7709007263184,41.738037109375,41.6632804870605,41.6159210205078,41.5955085754395,41.6437492370605,41.6411628723145,41.65869140625,41.8770027160645,41.8461456298828,41.7969741821289,41.7513160705566,41.8558616638184,41.9365730285645,42.0059585571289,42.052001953125,42.0920906066895,42.1581077575684,42.2158699035645,42.2923316955566,42.4658203125,42.5000495910645,42.5235366821289,42.5387687683105,42.5378913879395,42.5513153076172,42.5877952575684,42.6167984008789,42.6707534790039,42.6773452758789,42.6294441223145,42.5682106018066,42.6162109375,42.6387176513672,42.6845245361328,42.7362785339355,42.789794921875,42.76025390625,42.7438468933105,42.7203598022461,42.7467765808105,42.8493156433105,42.9287109375,43.0145034790039,43.0961456298828,43.2064933776855,43.2759742736816,43.3836898803711,43.6640625,43.8536109924316,43.8922386169434,43.9539527893066,43.9861831665039,44.0059547424316,44.0393562316895,44.1048812866211,44.1954078674316,44.2615242004395,44.278076171875,44.2594261169434,44.3033218383789,44.350830078125,44.4725074768066,44.5194854736328,44.6451644897461,44.6749534606934,44.7242202758789,44.8319358825684,44.9009780883789,44.9444808959961,44.983154296875,45.0201683044434,45.0357437133789,45.010986328125,45.0089340209961,45.0352554321289,45.0685043334961,45.0693359375,45.0689468383789,45.0765113830566,45.0982437133789,45.124755859375,45.1181144714355,45.14453125,45.1939430236816,45.2174301147461,45.2159156799316,45.1939430236816,45.1960906982422,45.2628936767578,45.3706550598145,45.4189453125,45.4746589660645,45.5252418518066,45.5951652526855,45.7308120727539,45.853515625,45.8854026794434,45.921630859375,45.9850578308105,46.0357933044434,46.10498046875,46.1772918701172,46.2706565856934,46.3242683410645,46.3879890441895,46.5290031433105,46.5660629272461,46.5957527160645,46.6610832214355,46.7490234375,46.8832511901855,46.9544944763184,46.9851570129395,47.0038566589355,47.1002922058105,47.2140121459961,47.28515625,47.3288078308105,47.4081535339355,47.5041007995605,47.556640625,47.5969734191895,47.6551742553711,47.6761703491211,47.7020988464355,47.7454109191895,47.8035659790039,47.8504867553711,47.877685546875,47.8863258361816,47.8443336486816,47.8232917785645,47.8270034790039,47.8524894714355,47.879150390625,47.9090805053711,48.0039558410645,48.029052734375,48.0248527526855,47.9809074401855,47.9876976013184,48.0025405883789,48.0499496459961,48.0890121459961,48.1017112731934,48.1705589294434,48.2201652526855,48.3174324035645,48.3844718933105,48.4034156799316,48.4720687866211,48.5090827941895,48.537841796875,48.5551300048828,48.5810546875,48.6033210754395,48.6404304504395,48.675048828125,48.7071800231934,48.735595703125,48.7652816772461,48.7916526794434,48.8357429504395,48.8816413879395,48.9185562133789,48.9455070495605,48.9655265808105,49.0001449584961,49.0319328308105,49.0802726745605,49.1163063049316,49.1623039245605,49.162109375,49.16455078125,49.1869125366211,49.2197723388672,49.2562980651855,49.2984390258789,49.3814926147461,49.4517097473145,49.4725608825684,49.4828643798828,49.4727058410645,49.4825706481934,49.4861335754395,49.4645500183105,49.4462394714355,49.4484367370605,49.4815406799316,49.5276374816895,49.5396957397461,49.5076675415039,49.4837417602539,49.4728012084961,49.5013656616211,49.5322265625,49.595703125,49.6270523071289,49.6111335754395,49.6115226745605,49.6605453491211,49.69970703125,49.7174797058105,49.75048828125,49.8232879638672,49.9030303955078,49.9480972290039,49.9535140991211,49.9843254089355,50.0692863464355,50.09375,50.1033210754395,50.11669921875,50.1511726379395,50.1668968200684,50.2133293151855,50.2223625183105,50.2594261169434,50.3059539794922,50.32373046875,50.3641586303711,50.4154777526855,50.422607421875,50.452392578125,50.46337890625,50.4700660705566,50.468017578125,50.5221672058105,50.56201171875,50.5755348205566,50.6151390075684,50.66552734375,50.693603515625,50.6976089477539,50.6919898986816,50.7005882263184,50.7751960754395,50.8122062683105,50.8498039245605,50.8641586303711,50.8030776977539,50.7650871276855,50.7254371643066,50.6882820129395,50.6832046508789,50.7109375,50.7786636352539,50.7891120910645,50.7782211303711,50.7449226379395,50.7125015258789,50.654541015625,50.6039047241211,50.6065406799316,50.6155776977539,50.6084938049316,50.5974617004395,50.5855445861816,50.5836906433105,50.57763671875,50.56884765625,50.5728530883789,50.556396484375,50.5113754272461,50.4048843383789,50.3034172058105,50.2218246459961,50.1796379089355,50.1657218933105,50.1328125,50.0879402160645,50.0237312316895,50.028076171875,50.0432624816895,50.0481948852539,50.0437507629395,50.0082511901855,49.9615745544434,49.9354476928711,49.94384765625,49.9441413879395,49.9412078857422,49.91552734375,49.9112319946289,49.94384765625,49.9660148620605,50.0125007629395,50.012939453125,49.9905738830566,49.9600105285645,49.9735832214355,49.998779296875,49.9987297058105,49.9824714660645,49.9541015625,49.901123046875,49.8960456848145,49.9115219116211,49.918701171875,49.8925323486328,49.87841796875,49.8978500366211,49.9115715026855,49.8828125,49.8298835754395,49.8050308227539,49.76171875,49.7308082580566,49.741455078125,49.7730484008789,49.8431129455566,49.9114723205566,49.93359375,49.9446296691895,49.9445304870605,49.94677734375,49.9967765808105,50.0142593383789,50.0778312683105,50.1065940856934,50.1805648803711,50.2276840209961,50.3024406433105,50.4229965209961,50.4869613647461,50.5332527160645,50.5571746826172,50.5685539245605,50.6038093566895,50.6446304321289,50.7020530700684,50.7691421508789,50.8176765441895,50.8551788330078,50.8871612548828,50.943359375,50.9852561950684,51.0516624450684,51.1651878356934,51.2178726196289,51.250732421875,51.2804679870605,51.3484382629395,51.3770523071289,51.449951171875,51.4835433959961,51.4857406616211,51.505615234375,51.5784187316895,51.6345710754395,51.6742668151855,51.7176246643066,51.8011741638184,51.9050750732422,51.9574699401855,52.070068359375,52.1172828674316,52.1017074584961,52.035400390625,52.0348625183105,51.9988746643066,51.9235343933105,51.89990234375,51.8925285339355,51.8716316223145,51.8275413513184,51.7555198669434,51.7371101379395,51.7298355102539,51.72607421875,51.7134742736816,51.6615753173828,51.604248046875,51.5530281066895,51.5132789611816,51.4747543334961,51.45263671875,51.4714813232422,51.4671859741211,51.4210433959961,51.3822250366211,51.3534660339355,51.3137702941895,51.2608413696289,51.216064453125,51.1075172424316,51.0506858825684,50.9743156433105,50.9014663696289,50.8294448852539,50.7912101745605,50.7686996459961,50.7184562683105,50.66552734375,50.6346664428711,50.5851097106934,50.5442390441895,50.5361785888672,50.5256805419922,50.4613265991211,50.3871574401855,50.3665542602539,50.333251953125,50.3006362915039,50.2907257080078,50.2642555236816,50.2002906799316,50.1870613098145,50.1649436950684,50.1385726928711,50.1538581848145,50.1760749816895,50.1718292236328,50.1572761535645,50.1542472839355,50.16943359375,50.2144546508789,50.2752914428711,50.30615234375,50.3171882629395,50.3418426513672,50.3829116821289,50.3899421691895,50.4295883178711,50.4604988098145,50.4737319946289,50.4412612915039,50.4141578674316,50.4053726196289,50.367919921875,50.3325691223145,50.3045883178711,50.3175773620605,50.3288078308105,50.3125991821289,50.248291015625,50.1966781616211,50.0864753723145,50.0330085754395,49.9894027709961,49.9866714477539,49.9831047058105,49.9600105285645,49.9478034973145,49.9247093200684,49.8490219116211,49.74072265625,49.6910171508789,49.6535148620605,49.6468772888184,49.593994140625,49.5626449584961,49.5248031616211,49.3963851928711,49.3415069580078,49.3228034973145,49.3356437683105,49.3353500366211,49.3349113464355,49.2963371276855,49.2698745727539,49.2393035888672,49.2056159973145,49.17041015625,49.2158699035645,49.219970703125,49.1870613098145,49.1375961303711,49.1429672241211,49.1661605834961,49.3042984008789,49.3558578491211,49.3426246643066,49.3609352111816,49.3764152526855,49.3977546691895,49.4036102294922,49.416015625,49.4242172241211,49.5145988464355,49.5323219299316,49.5072746276855,49.5235824584961,49.5692367553711,49.6162605285645,49.6925277709961,49.7971687316895,49.8743133544922,49.9416007995605,50.0070304870605,50.0615234375,50.1011238098145,50.2047348022461,50.2572746276855,50.2744140625,50.2554702758789,50.241455078125,50.2457008361816,50.2336921691895,50.1830558776855,50.1385726928711,50.0594215393066,49.9488754272461,49.9117660522461,49.896484375,49.8860359191895,49.880615234375,49.9059066772461,49.9521484375,50.0107917785645,50.00927734375,49.9780769348145,49.9203109741211,49.8770484924316,49.8237762451172],[14.1581048965454,14.111328125,14.1209964752197,14.1369142532349,14.1636228561401,14.189453125,14.1802244186401,14.1581048965454],[14.9705076217651,14.9195308685303,14.9988269805908,15.0308094024658,15.0601558685303,15.0594730377197,14.9705076217651],[15.1331548690796,15.1072254180908,15.113525390625,15.1250972747803,15.2152824401855,15.2568836212158,15.265380859375,15.22265625,15.1746091842651,15.1331548690796],[16.339111328125,16.3321285247803,16.3357906341553,16.3515129089355,16.3779792785645,16.379638671875,16.3597660064697,16.339111328125],[18.0789566040039,18.0569343566895,18.1554203033447,18.1726551055908,18.1367683410645,18.0789566040039],[18.7625007629395,18.7252445220947,18.7526359558105,18.8067874908447,18.8016128540039,18.7904796600342,18.7625007629395],[-26.8493156433105,-26.8509769439697,-26.8557624816895,-26.8584957122803,-26.8616218566895,-26.83349609375,-26.8394546508789,-26.52001953125,-26.4498043060303,-26.34716796875,-26.28125,-26.2150402069092,-26.1101551055908,-26.0183582305908,-25.9722652435303,-25.9576187133789,-25.8853511810303,-25.77392578125,-25.6319332122803,-25.3594722747803,-25.2634773254395,-25.0738277435303,-24.8440437316895,-24.6382827758789,-24.4606437683105,-24.37646484375,-24.3302726745605,-24.2362308502197,-24.0402336120605,-23.8921871185303,-23.7945308685303,-23.7430667877197,-23.6742191314697,-23.5529289245605,-23.4823226928711,-23.42578125,-23.2794933319092,-23.0166988372803,-22.8250980377197,-22.6175785064697,-22.4786128997803,-22.4546871185303,-22.4020500183105,-22.298828125,-22.1535148620605,-21.9833984375,-21.83154296875,-21.6980476379395,-21.5154304504395,-21.3348617553711,-21.3118152618408,-21.2970714569092,-21.1365242004395,-20.9500980377197,-20.8289051055908,-20.7129878997803,-20.6597652435303,-20.6130847930908,-20.51611328125,-20.3615226745605,-20.2171878814697,-19.98486328125,-19.93017578125,-19.8738288879395,-19.79541015625,-19.6680660247803,-19.5582027435303,-19.3887691497803,-19.2414054870605,-19.1524410247803,-19.1043949127197,-19.0587882995605,-19.0243167877197,-19.0018558502197,-18.94091796875,-18.8684577941895,-18.8284187316895,-18.763671875,-18.728515625,-18.6890621185303,-18.6329097747803,-18.49267578125,-18.3595714569092,-18.3125972747803,-18.271484375,-18.1962890625,-18.0829105377197,-17.7654304504395,-17.4375,-17.2515621185303,-17.0377922058105,-16.8835945129395,-16.7759761810303,-16.7123050689697,-16.7041988372803,-16.6976566314697,-16.6776371002197,-16.5894546508789,-16.5157241821289,-16.44873046875,-16.4288082122803,-16.2141609191895,-16.1796875,-16.15234375,-16.0236320495605,-16.01171875,-15.9992179870605,-15.9953126907349,-15.9782218933105,-15.8007822036743,-15.64306640625,-15.505859375,-15.3497076034546,-15.2888679504395,-15.1832027435303,-15.0668954849243,-15.0105476379395,-14.9903326034546,-14.9075193405151,-14.8665046691895,-14.8191404342651,-14.753321647644,-14.6946296691895,-14.6376943588257,-14.5771484375,-14.5367193222046,-14.4960927963257,-14.4144535064697,-14.3865232467651,-14.3408203125,-14.3230466842651,-14.2295894622803,-14.1224613189697,-14.0849609375,-14.0133790969849,-14.0430669784546,-14.2894535064697,-14.43408203125,-14.5681638717651,-14.5616216659546,-14.5302724838257,-14.5172853469849,-14.4871091842651,-14.4852533340454,-14.4493160247803,-14.4237308502197,-14.4085931777954,-14.4248046875,-14.59814453125,-14.7307615280151,-14.92236328125,-15.0159177780151,-15.1409177780151,-15.2972660064697,-15.4771490097046,-15.5667972564697,-15.7052736282349,-15.7734375,-15.8293933868408,-15.8874998092651,-15.9361324310303,-16.0237312316895,-16.0802726745605,-16.130859375,-16.1992168426514,-16.2467784881592,-16.2744140625,-16.3191413879395,-16.4315433502197,-16.5670890808105,-16.7603511810303,-16.8195323944092,-16.8338871002197,-16.8985347747803,-16.9738273620605,-17.0168933868408,-17.07861328125,-17.1109371185303,-17.1272449493408,-17.1310558319092,-17.1184577941895,-17.0969734191895,-16.8078136444092,-16.6392574310303,-16.5733394622803,-16.5602550506592,-16.5048828125,-16.3753910064697,-16.2471694946289,-16.1931648254395,-16.1605472564697,-16.1258792877197,-16.0958003997803,-16.0583000183105,-15.9586906433105,-15.6803712844849,-15.4189453125,-15.265625,-15.0346689224243,-14.8917961120605,-14.8637685775757,-14.6708984375,-14.4655275344849,-14.2010736465454,-14.0586910247803,-13.8968744277954,-13.6434574127197,-13.5516605377197,-13.5160150527954,-13.4867191314697,-13.4378910064697,-13.3601560592651,-13.21630859375,-13.1086921691895,-12.92578125,-12.6667966842651,-12.5907230377197,-12.3958988189697,-12.2105464935303,-12.1647462844849,-12.1202154159546,-11.9837884902954,-11.8870115280151,-11.8340826034546,-11.6900386810303,-11.6202144622803,-11.5886716842651,-11.57568359375,-11.578125,-11.5748052597046,-11.5832033157349,-11.5895500183105,-11.6047849655151,-11.6023445129395,-11.58203125,-11.5321292877197,-11.4529294967651,-11.4546871185303,-11.5373048782349,-11.6092767715454,-11.6707029342651,-11.7063474655151,-11.7162113189697,-11.6842775344849,-11.6103515625,-11.5712890625,-11.5669918060303,-11.5921878814697,-11.6471681594849,-11.6865234375,-11.7104501724243,-11.6750974655151,-11.5806646347046,-11.48193359375,-11.3791017532349,-11.3166990280151,-11.2947273254395,-11.2821292877197,-11.2787113189697,-11.3111333847046,-11.4132814407349,-11.3453130722046,-11.2289056777954,-11.1672849655151,-11.1668949127197,-11.12255859375,-11.0345697402954,-10.978515625,-10.9547853469849,-10.9124021530151,-10.82080078125,-10.6875,-10.5515632629395,-10.46435546875,-10.5673828125,-10.6615238189697,-10.7162103652954,-10.76513671875,-10.8306646347046,-10.9295902252197,-10.9984378814697,-11.025390625,-11.0656251907349,-11.1789064407349,-11.265625,-11.3320322036743,-11.4494132995605,-11.6573247909546,-11.8444337844849,-11.9404296875,-12.0045900344849,-12.1194334030151,-12.3128900527954,-12.3927736282349,-12.4921875,-12.5265626907349,-12.6355466842651,-12.7583980560303,-12.8246088027954,-12.90478515625,-12.9359378814697,-12.9831056594849,-12.9846677780151,-13.0577144622803,-13.115234375,-13.2234373092651,-13.2937498092651,-13.3740234375,-13.4628915786743,-13.5314445495605,-13.6203126907349,-13.8450193405151,-14.1228513717651,-14.1673831939697,-14.1988277435303,-14.2144536972046,-14.2906255722046,-14.3900384902954,-14.4518556594849,-14.5386724472046,-14.4207029342651,-14.4212884902954,-14.4675788879395,-14.5355472564697,-14.5690431594849,-14.6349611282349,-14.7186508178711,-14.79150390625,-14.8424806594849,-14.9297857284546,-15.0116205215454,-15.0652341842651,-15.0824222564697,-15.1155281066895,-15.1926746368408,-15.2609376907349,-15.4734363555908,-15.7639656066895,-15.8670892715454,-15.9792976379395,-16.0252933502197,-16.0653324127197,-16.2254886627197,-16.2517566680908,-16.2945308685303,-16.4356441497803,-16.4681625366211,-16.5793952941895,-16.7925777435303,-16.8419933319092,-16.9728507995605,-17.0045890808105,-17.0416011810303,-17.05517578125,-17.0457038879395,-17.05029296875,-17.0783214569092,-17.1701164245605,-17.2427730560303,-17.2759761810303,-17.3213863372803,-17.3931636810303,-17.5707035064697,-17.7399406433105,-17.9092769622803,-17.9349613189697,-17.9934577941895,-18.080078125,-18.1290035247803,-18.3073234558105,-18.5181636810303,-18.5757808685303,-18.6929683685303,-18.7697277069092,-18.7931652069092,-18.7196292877197,-18.861328125,-18.8713855743408,-18.8423824310303,-18.9125003814697,-18.9933586120605,-19.1638660430908,-19.4939460754395,-19.8126945495605,-19.82177734375,-19.8205070495605,-19.7095718383789,-19.7013664245605,-19.7671871185303,-19.8219718933105,-19.9294929504395,-20.0908203125,-20.4043941497803,-20.4730472564697,-20.5619144439697,-20.6708011627197,-20.8062496185303,-21.1952133178711,-21.3953132629395,-21.6509780883789,-21.76171875,-22.0374031066895,-22.2603511810303,-22.3968753814697,-22.45458984375,-22.4025382995605,-22.3162097930908,-22.1776371002197,-22.1159191131592,-22.1247081756592,-22.1901378631592,-22.2481441497803,-22.3026371002197,-22.3765621185303,-22.65771484375,-22.7720699310303,-22.9630851745605,-23.1851558685303,-23.7078113555908,-23.7982425689697,-23.8376941680908,-23.8510761260986,-23.7844715118408,-23.7849597930908,-23.8244132995605,-24.0655269622803,-24.1711902618408,-24.4302730560303,-24.5414066314697,-24.6505870819092,-24.8212890625,-25.0679683685303,-25.1888675689697,-25.2609367370605,-25.4904308319092,-25.6443367004395,-25.8208980560303,-25.9017581939697,-26.0041007995605,-26.0919914245605,-26.1584949493408,-26.2030277252197,-26.2414054870605,-26.2680683135986,-26.1298828125,-26.0869140625,-26.0835933685303,-26.2523422241211,-26.8304691314697,-26.8493156433105],[-12.013671875,-11.9847660064697,-11.9710931777954,-11.9822263717651,-12.0186529159546,-12.0658206939697,-12.0666017532349,-12.013671875],[-12.1106443405151,-12.08837890625,-12.05908203125,-12.03076171875,-12.0130863189697,-12.0027341842651,-12.0088863372803,-12.0475578308105,-12.1007804870605,-12.1186513900757,-12.1106443405151],[19.7064437866211,19.6092758178711,19.6463871002197,19.7103519439697,19.8019523620605,19.8492679595947,19.8662109375,19.7064437866211],[14.8090333938599,14.8169927597046,14.8890132904053,14.9746589660645,15.0390129089355,15.0820808410645,15.1312494277954,15.1866693496704,15.2235355377197,15.2425289154053,15.2624025344849,15.2896480560303,15.3404302597046,15.4530277252197,15.5048828125,15.4966306686401,15.5104494094849,15.5352535247803,15.57177734375,15.6033191680908,15.6168947219849,15.7003898620605,15.8538579940796,15.9734869003296,16.0591793060303,16.0911121368408,16.0970211029053,16.1103019714355,16.1352043151855,16.14404296875,16.1183109283447,16.1269054412842,16.1688003540039,16.1725101470947,16.1380367279053,16.1481456756592,16.202880859375,16.2572269439697,16.3111324310303,16.4188480377197,16.5802745819092,16.6559581756592,16.6458988189697,16.6535148620605,16.6789073944092,16.6769046783447,16.6474609375,16.6409664154053,16.6573734283447,16.6449222564697,16.6036128997803,16.5826168060303,16.5819835662842,16.5565929412842,16.506591796875,16.485107421875,16.4921398162842,16.5104484558105,16.5401363372803,16.5470714569092,16.5312976837158,16.4513187408447,16.3071765899658,16.2249031066895,16.2045402526855,16.0972175598145,15.9173345565796,15.83837890625,16.2868156433105,16.4542484283447,16.6015148162842,16.9264163970947,17.1925773620605,17.5458488464355,17.887939453125,18.22314453125,18.5211925506592,18.7181644439697,19.0033206939697,19.15380859375,19.2850570678711,19.3619632720947,19.390625,19.4102535247803,19.5126457214355,19.47314453125,19.7871589660645,20.0009765625,20.1412601470947,20.2279300689697,20.4158687591553,20.65234375,20.6897964477539,20.7095203399658,20.6540031433105,20.6041507720947,20.6341781616211,20.80615234375,21.0861320495605,21.1147956848145,21.0764636993408,21.0396976470947,20.80615234375,20.8988285064697,21.0080070495605,21.1424331665039,21.3292484283447,21.3293952941895,21.3296394348145,21.3299312591553,21.3302249908447,21.3304691314697,21.3307609558105,21.3310546875,21.331298828125,21.3315906524658,21.3318843841553,21.3321285247803,21.3324222564697,21.3327140808105,21.3329582214355,21.333251953125,21.3335437774658,21.3337879180908,21.3339347839355,21.4667987823486,21.5720691680908,21.7138175964355,21.8547859191895,21.9957523345947,22.128173828125,22.2604484558105,22.3832530975342,22.4959964752197,22.5607414245605,22.6893062591553,22.7532234191895,22.8205070495605,22.8840827941895,23.000244140625,23.0895500183105,23.1927261352539,23.2713375091553,23.2908210754395,23.3180179595947,23.3774909973145,23.4354496002197,23.4675788879395,23.5764656066895,23.6978988647461,23.8340320587158,23.97021484375,24.1063480377197,24.2424812316895,24.378662109375,24.5147953033447,24.6509761810303,24.787109375,24.9232425689697,25.0593757629395,25.195556640625,25.3316898345947,25.4678707122803,25.60400390625,25.7401371002197,25.8763179779053,25.9954090118408,25.9954090118408,25.9954090118408,25.9954090118408,25.9954090118408,25.9954586029053,25.9954586029053,25.9954586029053,25.9954586029053,25.9954586029053,25.9954586029053,25.9954586029053,25.9954586029053,25.9955062866211,25.9955062866211,25.9955062866211,25.9955062866211,25.9955062866211,25.9955062866211,26.1094722747803,26.273193359375,26.4977054595947,26.72314453125,26.9213371276855,27.1192874908447,27.2859382629395,27.1753425598145,27.0647449493408,26.9541492462158,26.8435554504395,26.7329597473145,26.6223621368408,26.5117664337158,26.4011726379395,26.2905769348145,26.1799793243408,26.0694332122803,25.9587898254395,25.8481941223145,25.7375965118408,25.6270008087158,25.5164070129395,25.4237785339355,25.2745113372803,25.1354503631592,24.99560546875,24.9954109191895,24.9951667785645,24.9949722290039,24.9948253631592,24.99462890625,24.7667961120605,24.5181636810303,24.2694816589355,24.0207996368408,23.7721672058105,23.5234851837158,23.2748031616211,23.026123046875,22.7774906158447,22.5288581848145,22.2801761627197,22.0315437316895,21.7828617095947,21.5341796875,21.2855472564697,21.036865234375,20.7881851196289,20.5395030975342,20.2908687591553,20.0421886444092,19.7935066223145,19.5448722839355,19.2961902618408,19.0475082397461,18.7988758087158,18.5502433776855,18.3015613555908,18.0529289245605,17.8042469024658,17.5555667877197,17.306884765625,17.0582523345947,16.8095703125,16.5686531066895,16.4420413970947,16.2828617095947,16.0579090118408,15.7894048690796,15.4962892532349,15.4962892532349,15.4962892532349,15.4962396621704,15.4961910247803,15.4961910247803,15.4961910247803,15.4961910247803,15.4961910247803,15.4961910247803,15.4961423873901,15.4961423873901,15.4961423873901,15.4961423873901,15.4961423873901,15.4961423873901,15.49609375,15.49609375,15.49609375,15.5028324127197,15.5256843566895,15.6773929595947,15.6676273345947,15.623046875,15.5748529434204,15.5116701126099,15.4582033157349,15.437255859375,15.4014654159546,15.373779296875,15.3836917877197,15.3960456848145,15.416015625,15.4379386901855,15.4397945404053,15.4348630905151,15.4226560592651,15.3949222564697,15.2817392349243,15.1504878997803,15.151123046875,15.222900390625,15.3586416244507,15.5367679595947,15.6253900527954,15.6368169784546,15.5732431411743,15.5120601654053,15.4255361557007,15.3427238464355,15.244873046875,15.12939453125,14.9951667785645,14.8869142532349,14.804931640625,14.7663583755493,14.7453603744507,14.8090333938599],[16.7403335571289,16.6812000274658,16.6995124816895,16.7515640258789,16.8043937683105,16.8095703125,16.7403335571289],[-20.48486328125,-20.51318359375,-20.5037097930908,-20.4499988555908,-20.4276371002197,-20.4064445495605,-20.3375968933105,-20.2286128997803,-20.1837902069092,-20.1439456939697,-20.0559577941895,-19.9971675872803,-19.9899425506592,-20.0984382629395,-20.2125968933105,-20.3269538879395,-20.3688488006592,-20.4348640441895,-20.48486328125],[-12.1106443405151,-12.1186513900757,-12.1007804870605,-12.0475578308105,-12.0088863372803,-12.0027341842651,-12.0130863189697,-12.03076171875,-12.05908203125,-12.08837890625,-12.1106443405151],[-12.013671875,-12.0666017532349,-12.0658206939697,-12.0186529159546,-11.9822263717651,-11.9710931777954,-11.9847660064697,-12.013671875],[-11.578125,-11.57568359375,-11.5886716842651,-11.6202144622803,-11.6900386810303,-11.8340826034546,-11.8870115280151,-11.9837884902954,-12.1202154159546,-12.1647462844849,-12.2105464935303,-12.3958988189697,-12.5907230377197,-12.6667966842651,-12.92578125,-13.1086921691895,-13.21630859375,-13.3601560592651,-13.4378910064697,-13.4867191314697,-13.5160150527954,-13.5516605377197,-13.6434574127197,-13.8968744277954,-14.0586910247803,-14.2010736465454,-14.4655275344849,-14.6708984375,-14.8637685775757,-14.8917961120605,-15.0346689224243,-15.265625,-15.4189453125,-15.6803712844849,-15.9586906433105,-16.0583000183105,-16.0958003997803,-16.1258792877197,-16.1605472564697,-16.1931648254395,-16.2471694946289,-16.3753910064697,-16.5048828125,-16.5602550506592,-16.5733394622803,-16.6392574310303,-16.8078136444092,-17.0969734191895,-17.1184577941895,-17.1310558319092,-17.1272449493408,-17.1109371185303,-17.07861328125,-17.0168933868408,-16.9738273620605,-16.8985347747803,-16.8338871002197,-16.8195323944092,-16.7603511810303,-16.5670890808105,-16.4315433502197,-16.3191413879395,-16.2744140625,-16.2467784881592,-16.1992168426514,-16.130859375,-16.0802726745605,-16.0237312316895,-15.9361324310303,-15.8874998092651,-15.8293933868408,-15.7734375,-15.7052736282349,-15.5667972564697,-15.4771490097046,-15.2972660064697,-15.1409177780151,-15.0159177780151,-14.92236328125,-14.7307615280151,-14.59814453125,-14.4248046875,-14.4085931777954,-14.4237308502197,-14.4493160247803,-14.4852533340454,-14.4871091842651,-14.5172853469849,-14.5302724838257,-14.5616216659546,-14.5681638717651,-14.43408203125,-14.2894535064697,-14.0430669784546,-14.0133790969849,-13.94091796875,-13.9591798782349,-14.0100584030151,-14.0237302780151,-14.0221681594849,-14.0093755722046,-13.9768562316895,-13.8838872909546,-13.8173818588257,-13.7916011810303,-13.7610349655151,-13.7314453125,-13.7102537155151,-13.6884765625,-13.6565427780151,-13.6103515625,-13.5904293060303,-13.55029296875,-13.5027341842651,-13.45703125,-13.3570308685303,-13.2574214935303,-13.2249994277954,-13.1588869094849,-13.0842771530151,-12.9894533157349,-12.8996095657349,-12.8647470474243,-12.8043947219849,-12.7013673782349,-12.6304683685303,-12.5565433502197,-12.4898433685303,-12.4604501724243,-12.4034175872803,-12.3477535247803,-12.3310546875,-12.3296871185303,-12.3083009719849,-12.1125974655151,-11.8881845474243,-11.7999992370605,-11.6908206939697,-11.6111326217651,-11.57763671875,-11.5348625183105,-11.4176759719849,-11.4039058685303,-11.2491207122803,-11.1579103469849,-11.0851564407349,-10.9811525344849,-10.9150390625,-10.8933591842651,-10.8523445129395,-10.8126945495605,-10.8017578125,-10.7831058502197,-10.5905275344849,-10.5531253814697,-10.4885740280151,-10.3913087844849,-10.3515625,-10.2346677780151,-10.19970703125,-10.1208982467651,-10.0379886627197,-9.95400333404541,-9.86220645904541,-9.81181621551514,-9.75957012176514,-9.68300819396973,-9.62617111206055,-9.603515625,-9.60263729095459,-9.63818454742432,-9.63505840301514,-9.62285137176514,-9.57363319396973,-9.52031326293945,-9.48417949676514,-9.43398475646973,-9.40742206573486,-9.39970684051514,-9.39501953125,-9.49589824676514,-9.50048828125,-9.51914119720459,-9.60800743103027,-9.61972618103027,-9.60752010345459,-9.59814453125,-9.61093807220459,-9.66298770904541,-9.67011737823486,-9.67216873168945,-9.65820217132568,-9.62734413146973,-9.56533241271973,-9.53173828125,-9.49540996551514,-9.53779315948486,-9.73154354095459,-9.75654315948486,-9.94882774353027,-10.0301752090454,-10.0731449127197,-10.2411136627197,-10.31982421875,-10.3796873092651,-10.4276361465454,-10.4961919784546,-10.5250978469849,-10.6255865097046,-10.7100582122803,-10.79248046875,-10.8728513717651,-10.990234375,-11.0375003814697,-11.0804691314697,-11.1271486282349,-11.1774415969849,-11.2381839752197,-11.3094720840454,-11.3416986465454,-11.3409175872803,-11.3519525527954,-11.3935546875,-11.4634761810303,-11.5437507629395,-11.578125],[2.41533231735229,2.40219712257385,2.43759751319885,2.74116206169128,2.76831030845642,2.76445317268372,2.70932602882385,2.57631850242615,2.45893549919128,2.41533231735229],[2.73173832893372,2.72133827209473,2.72822284698486,2.76723670959473,2.85683560371399,2.87172865867615,2.77421855926514,2.73173832893372],[2.98847699165344,2.97041010856628,2.99648451805115,3.03281235694885,3.0673828125,3.04760718345642,2.98847699165344],[4.18613290786743,4.16694355010986,4.16899394989014,4.20400428771973,4.25019550323486,4.26240205764771,4.25234365463257,4.18613290786743],[5.29472637176514,5.26699256896973,5.28286123275757,5.44687461853027,5.4677734375,5.43793964385986,5.41005897521973,5.29472637176514],[6.46572256088257,6.35859394073486,6.31083965301514,6.29707050323486,6.312744140625,6.27158212661743,6.26328134536743,6.33754873275757,6.36713886260986,6.41835927963257,6.42734384536743,6.40961933135986,6.44501924514771,6.46572256088257],[6.24223661422729,6.20341825485229,6.17202186584473,5.862548828125,5.64492177963257,5.56376981735229,5.52495098114014,5.408447265625,5.26215839385986,4.85029268264771,4.66948223114014,4.39326190948486,3.97685551643372,3.76914048194885,3.67109346389771,3.52060556411743,3.37856411933899,3.26059579849243,2.93310570716858,2.83657240867615,2.77475595474243,2.58046889305115,2.50849628448486,2.26123070716858,1.72285139560699,1.48066389560699,1.41557645797729,1.38857412338257,1.36489260196686,1.41225564479828,1.44619131088257,1.48833000659943,1.52978527545929,1.57929682731628,1.62363290786743,1.55004906654358,1.45478510856628,1.44667947292328,1.47656261920929,1.44965827465057,1.33281266689301,1.32949233055115,1.42983388900757,1.49785149097443,1.54614269733429,1.79233407974243,1.85556626319885,2.04238247871399,2.24848628044128,2.44941401481628,2.57358360290527,2.68364262580872,2.8134765625,2.83896470069885,2.88520503044128,3.01113271713257,3.14248037338257,3.25327157974243,3.47202157974243,3.62470722198486,3.77670907974243,3.86445307731628,3.96621108055115,4.00180673599243,4.02338886260986,4.09721660614014,4.22573232650757,4.37343692779541,4.65224599838257,5.04428720474243,5.58769559860229,5.77797842025757,5.98417997360229,6.18251943588257,6.32421875,6.44199180603027,6.48867225646973,6.64160108566284,6.67182588577271,6.68662071228027,6.68271493911743,6.54990196228027,6.467529296875,6.447998046875,6.48066425323486,6.46005868911743,6.42617177963257,6.33164072036743,6.24540996551514,6.25766611099243,6.24531269073486,6.24257802963257,6.16606473922729,6.03369140625,5.95649385452271,5.84619140625,5.77104520797729,5.72451162338257,5.67490243911743,5.63676738739014,5.64306640625,5.66875028610229,5.73369121551514,5.78935480117798,5.85166025161743,5.90776395797729,5.90200233459473,5.87714862823486,5.79599666595459,5.77880859375,5.77060556411743,5.77934598922729,5.82529306411743,5.911376953125,5.97934579849243,6.09667921066284,6.18466806411743,6.24223661422729],[4.17060565948486,4.17138671875,4.19287109375,4.29931688308716,4.33706092834473,4.34013700485229,4.35498046875,4.339111328125,4.33842754364014,4.35986328125,4.37080049514771,4.30820322036743,4.32734394073486,4.35371065139771,4.362548828125,4.35517597198486,4.29067420959473,4.34868144989014,4.34804677963257,4.33330059051514,4.25375938415527,4.19301748275757,4.08198213577271,3.97553682327271,3.93876981735229,3.73305654525757,3.63369131088257,3.50229501724243,3.44575214385986,3.36167001724243,3.34238243103027,3.20864272117615,3.17314457893372,3.12812495231628,3.03432607650757,3.00873994827271,2.99394512176514,3.02592778205872,2.97446274757385,2.89487314224243,2.84111332893372,2.79121088981628,2.75781273841858,2.72343730926514,2.68701148033142,2.63422846794128,2.61240220069885,2.56689429283142,2.523193359375,2.49291968345642,2.44614243507385,2.35083031654358,2.26938509941101,2.25048828125,2.21293950080872,2.16240215301514,2.05161118507385,2.01894521713257,1.98002934455872,1.93378925323486,1.89394533634186,1.86899399757385,1.85078144073486,1.81904304027557,1.68627953529358,1.61704099178314,1.51416015625,1.46713864803314,1.45200169086456,1.50004875659943,1.47089838981628,1.45234382152557,1.45527327060699,1.43427717685699,1.37988293170929,1.31137716770172,1.26059591770172,1.23593759536743,1.30839836597443,1.30214858055115,1.3271484375,1.40810561180115,1.43178713321686,1.43388652801514,1.45712912082672,1.49624037742615,1.54755842685699,1.56699204444885,1.55908167362213,1.51474630832672,1.47963881492615,1.43906259536743,1.33818352222443,1.24360370635986,1.14335954189301,1.11328101158142,1.01166987419128,0.999462962150574,1.01420927047729,1.02260756492615,0.994336187839508,0.995751917362213,1.043212890625,1.05053699016571,1.02636742591858,1.01733410358429,0.878124892711639,0.861962854862213,0.88207995891571,0.939062297344208,0.995995998382568,1.19013643264771,1.23574221134186,1.28256833553314,1.33803689479828,1.39785146713257,1.43896460533142,1.52294933795929,1.61489248275757,1.77666020393372,1.80629885196686,1.84834015369415,1.89619112014771,2.02753877639771,1.88876974582672,1.85781276226044,1.76445329189301,1.71762716770172,1.69858384132385,1.69472658634186,1.701171875,1.71972644329071,1.69985353946686,1.54804658889771,1.52084946632385,1.532470703125,1.51733374595642,1.40087890625,1.38696300983429,1.39584946632385,1.44902348518372,1.48667013645172,1.55781269073486,1.63364255428314,1.68408226966858,1.73876941204071,1.90229499340057,1.985107421875,2.06386733055115,2.13974595069885,2.19765639305115,2.29716753959656,2.379638671875,2.43574213981628,2.39877963066101,2.36445283889771,2.36787128448486,2.38154315948486,2.42407202720642,2.49809598922729,2.63432645797729,2.74301767349243,2.81796860694885,2.85380887985229,2.91469693183899,3.07045912742615,3.13071298599243,3.16191387176514,3.20522451400757,3.34350609779358,3.56147480010986,3.74057626724243,4.00141572952271,4.24321317672729,4.28872108459473,4.42070293426514,4.48281288146973,4.54580068588257,4.57524394989014,4.59287118911743,4.59267568588257,4.56523418426514,4.52695322036743,4.47788047790527,4.41425800323486,4.354736328125,4.30419969558716,4.26279258728027,4.255859375,4.20356416702271,4.11357402801514,4.049072265625,4.02397489547729,4.03764677047729,4.09653329849243,4.16879892349243,4.26650381088257,4.28076171875,4.3544921875,4.39321327209473,4.42876005172729,4.46391582489014,4.55302762985229,4.66650390625,4.71806621551514,4.75483417510986,4.80175733566284,4.85624980926514,4.89970684051514,4.82114219665527,4.69135713577271,4.58266592025757,4.39042949676514,4.36420917510986,4.34721708297729,4.35258817672729,4.36528253555298,4.38076210021973,4.45634746551514,4.63398408889771,4.75058603286743,4.86669921875,4.89975595474243,4.93276405334473,4.96918964385986,5.04892587661743,5.09355449676514,5.19414091110229,5.25410175323486,5.33051729202271,5.41318368911743,5.56669950485229,5.60341739654541,5.54887676239014,5.53510761260986,5.53300762176514,5.5361328125,5.613525390625,5.72495126724243,5.88237333297729,6.00327157974243,6.12954044342041,6.52167940139771,6.58271455764771,6.97709989547729,6.990234375,6.95205068588257,6.82670927047729,6.77207040786743,6.69106483459473,6.60610342025757,6.60791015625,6.65966796875,6.79736328125,6.91684532165527,6.96889686584473,6.93999052047729,6.91923809051514,6.83339834213257,6.783447265625,6.67690372467041,6.61225605010986,6.57148408889771,6.51264619827271,6.47368144989014,6.4267578125,6.34999990463257,6.27231454849243,6.196533203125,6.07358407974243,6.00185537338257,5.94072246551514,5.88466787338257,5.9404296875,5.97226572036743,6.05332040786743,5.98344755172729,5.86250019073486,5.83207988739014,5.78750038146973,5.76918935775757,5.70625019073486,5.71210956573486,5.75419902801514,5.820556640625,5.81972694396973,5.80605459213257,5.763427734375,5.72890663146973,5.68452167510986,5.59208965301514,5.55854511260986,5.42900371551514,5.41782236099243,5.41523456573486,5.430908203125,5.41264629364014,5.36591815948486,5.30810546875,5.24589872360229,5.19872999191284,5.15981435775757,5.10048818588257,5.02290010452271,4.96406221389771,4.96811532974243,5.01850605010986,5.01201200485229,4.98886728286743,4.82851600646973,4.66870164871216,4.50214815139771,4.46064424514771,4.40966796875,4.37924814224243,4.36235332489014,4.33574199676514,4.31601524353027,4.28759765625,4.250244140625,4.262939453125,4.33754873275757,4.34282255172729,4.30449199676514,4.20000028610229,4.17060565948486],[7.168212890625,7.11528348922729,7.17885732650757,7.26069355010986,7.33701133728027,7.35166025161743,7.29062509536743,7.22080087661743,7.18476581573486,7.168212890625],[-17.640625,-17.5994129180908,-17.5601558685303,-17.5208988189697,-17.4895515441895,-17.4810543060303,-17.5177745819092,-17.54345703125,-17.5685539245605,-17.6343746185303,-17.7935543060303,-17.7875995635986,-17.8213882446289,-17.8646488189697,-18.052734375,-18.0285167694092,-17.9894523620605,-17.9782218933105,-18.0234375,-18.0775394439697,-18.1541023254395,-18.2292003631592,-18.26953125,-18.42431640625,-18.4494132995605,-18.4599609375,-18.4529285430908,-18.3864250183105,-18.2310543060303,-18.02734375,-18.0075206756592,-17.9997062683105,-18.0095691680908,-18.0671882629395,-18.11572265625,-18.1986331939697,-18.265625,-18.3068370819092,-18.31884765625,-18.5205078125,-18.9285144805908,-19.33642578125,-19.7443351745605,-20.15234375,-20.5602550506592,-20.9681625366211,-21.3760738372803,-21.7840824127197,-21.9619140625,-22.0001945495605,-22.0001945495605,-22.0001945495605,-22.0001945495605,-22.0001945495605,-22.2425785064697,-22.529296875,-22.81591796875,-23.1025390625,-23.38916015625,-23.67578125,-23.96240234375,-24.2490234375,-24.5357418060303,-24.751953125,-24.7767562866211,-25.19677734375,-25.6416015625,-26.0863285064697,-26.5311527252197,-26.9759769439697,-27.4207019805908,-27.8655281066895,-28.3103523254395,-28.4512691497803,-28.4494152069092,-28.50390625,-28.5746097564697,-28.66162109375,-28.7144527435303,-28.7333011627197,-28.7777347564697,-28.8479499816895,-28.9016590118408,-28.9387702941895,-28.9279308319092,-28.869140625,-28.8552742004395,-28.8862285614014,-28.8716812133789,-28.8113288879395,-28.7769527435303,-28.7683601379395,-28.7430648803711,-28.6981430053711,-28.62109375,-28.5626945495605,-28.5011730194092,-28.45166015625,-28.4139652252197,-28.3532238006592,-28.2694339752197,-28.2286128997803,-28.2308597564697,-28.1988296508789,-28.1325187683105,-28.0822257995605,-28.0310554504395,-28.0696296691895,-28.1279296875,-28.2189445495605,-28.2645511627197,-28.3408203125,-28.3947277069092,-28.4521484375,-28.4754886627197,-28.4649410247803,-28.4878902435303,-28.5728511810303,-28.6175765991211,-28.5365238189697,-28.2317390441895,-28.1525382995605,-27.9658203125,-27.3865222930908,-27.2749996185303,-26.9951171875,-26.78759765625,-26.6678714752197,-26.6001949310303,-26.5080070495605,-26.42578125,-26.3180656433105,-25.9582023620605,-25.7256832122803,-25.5335941314697,-25.3585929870605,-25.2463874816895,-25.033203125,-24.7879867553711,-24.5480461120605,-24.2019538879395,-24.0503902435303,-23.6428699493408,-23.4766597747803,-23.2811527252197,-23.07861328125,-22.9680671691895,-22.8805675506592,-22.908203125,-22.92138671875,-22.80517578125,-22.7025394439697,-22.4491214752197,-22.18994140625,-21.767578125,-21.6066417694092,-21.4732418060303,-20.9167003631592,-20.52392578125,-20.1846675872803,-20.0282230377197,-18.9267578125,-18.7510738372803,-18.5409164428711,-18.470703125,-18.2705078125,-18.0017566680908,-17.7509765625,-17.466796875,-17.2492198944092,-17.2265625,-17.1685543060303,-17.16455078125,-17.2099609375,-17.21337890625,-17.2058601379395,-17.2126941680908,-17.1605472564697,-17.1082038879395,-17.0625972747803,-17.0154304504395,-16.9676761627197,-16.9716796875,-16.9895515441895,-17.0078125,-17.0400390625,-17.1412124633789,-17.2334957122803,-17.2883777618408,-17.3607425689697,-17.3887710571289,-17.4041996002197,-17.4088859558105,-17.3977527618408,-17.3876953125,-17.3879890441895,-17.3885746002197,-17.38916015625,-17.3896484375,-17.3902359008789,-17.3908195495605,-17.3914070129395,-17.3919925689697,-17.392578125,-17.3927726745605,-17.39599609375,-17.3994140625,-17.4051742553711,-17.4246082305908,-17.4427738189697,-17.5700187683105,-17.7032222747803,-17.7663097381592,-17.8035163879395,-17.8176746368408,-17.8084964752197,-17.8254871368408,-17.86865234375,-17.88134765625,-17.8636722564697,-17.8874034881592,-17.9525394439697,-17.9966793060303,-18.0197257995605,-18.0060558319092,-17.9557609558105,-17.9629878997803,-17.99951171875,-18.0006828308105,-17.94775390625,-17.9051761627197,-17.8374996185303,-17.7816410064697,-17.6988277435303,-17.640625],[-22.6232414245605,-22.6611328125,-22.6533222198486,-22.63916015625,-22.6185531616211,-22.5414066314697,-22.5792007446289,-22.6232414245605],[-21.4299793243408,-21.4484367370605,-21.4452152252197,-21.6158199310303,-21.6431636810303,-21.6416015625,-21.6057605743408,-21.5821285247803,-21.5236339569092,-21.3926773071289,-21.3728523254395,-21.337890625,-21.3697280883789,-21.4069328308105,-21.4299793243408],[-21.16064453125,-21.1687507629395,-21.0967769622803,-21.0606441497803,-20.9972648620605,-20.9225578308105,-20.9041004180908,-20.8035144805908,-20.76611328125,-20.7594718933105,-20.72021484375,-20.6735343933105,-20.7005863189697,-20.7325210571289,-20.8915042877197,-20.9420890808105,-21.0552730560303,-21.0870113372803,-21.16064453125],[-20.69873046875,-20.7085933685303,-20.6170902252197,-20.5611343383789,-20.4504871368408,-20.4133796691895,-20.4182605743408,-20.4501953125,-20.4775390625,-20.525390625,-20.5853519439697,-20.5962886810303,-20.6619148254395,-20.69873046875],[-20.24609375,-20.3088855743408,-20.2822265625,-20.3811511993408,-20.6810550689697,-20.7445316314697,-20.7688484191895,-20.8179683685303,-20.8870124816895,-20.9358386993408,-20.9813480377197,-21.0427722930908,-21.08056640625,-21.1799793243408,-21.2671871185303,-21.3117179870605,-21.36376953125,-21.38916015625,-21.4423828125,-21.48388671875,-21.6372051239014,-21.7828121185303,-21.8728504180908,-21.9530277252197,-22.0169925689697,-22.0901355743408,-22.2615222930908,-22.3228511810303,-22.3533210754395,-22.35546875,-22.3761711120605,-22.2655277252197,-22.2492179870605,-22.2560558319092,-22.2315425872803,-22.1961917877197,-22.1550769805908,-22.0891590118408,-22.04443359375,-21.98876953125,-21.9566402435303,-21.9080085754395,-21.8538074493408,-21.77734375,-21.7242183685303,-21.6150379180908,-21.580078125,-21.5254878997803,-21.3268566131592,-21.2898426055908,-21.2015628814697,-20.9920902252197,-20.9058609008789,-20.8291015625,-20.7392578125,-20.6327152252197,-20.4801750183105,-20.4149417877197,-20.3479499816895,-20.3048839569092,-20.2786121368408,-20.2335929870605,-20.1728515625,-20.1415042877197,-20.24609375],[-19.3117198944092,-19.3331069946289,-19.17431640625,-19.1146488189697,-19.23828125,-19.3117198944092],[22.9961910247803,22.4183597564697,21.9220714569092,21.6058101654053,21.5233898162842,21.4674320220947,21.4115238189697,20.9543952941895,20.8749027252197,20.7333011627197,20.67236328125,20.3998546600342,20.34619140625,20.3031749725342,19.9825687408447,19.904052734375,19.4952144622803,19.206787109375,18.8108386993408,18.3370609283447,17.937255859375,17.4084949493408,16.9083976745605,16.6338882446289,16.1466312408447,15.7501468658447,15.4847640991211,14.9661121368408,14.6307621002197,14.4555177688599,14.3806648254395,14.1344232559204,13.70458984375,13.7017583847046,13.6708498001099,13.5730466842651,13.5345211029053,13.4490232467651,13.38037109375,13.3265619277954,13.1943359375,13.09375,13.0736808776855,13.0904302597046,13.1917963027954,13.2977056503296,13.3405275344849,13.3536128997803,13.3715333938599,13.3302249908447,13.281005859375,13.2701177597046,13.2061519622803,13.13525390625,12.8106451034546,12.8214845657349,12.8574705123901,12.9081544876099,13.0596685409546,13.2911624908447,13.32275390625,13.3409175872803,13.337890625,13.1071786880493,13.1122560501099,13.0863285064697,13.0291013717651,13.0001955032349,12.9955568313599,13.0082025527954,13.0432615280151,13.107666015625,13.3642578125,13.4091310501099,13.4854011535645,13.6036138534546,13.6587886810303,13.6729984283447,13.6636714935303,13.765380859375,13.8728523254395,13.8591794967651,13.8368654251099,13.7572269439697,13.7427244186401,13.7491216659546,13.759765625,13.7332029342651,13.7018070220947,13.6475095748901,13.5010747909546,13.4821290969849,13.4577150344849,13.0554685592651,12.9346685409546,12.7750482559204,12.6221685409546,12.5299806594849,12.4052734375,12.2016115188599,12.0615720748901,11.9703617095947,11.9269533157349,11.8873043060303,11.8277339935303,11.7996578216553,11.762451171875,11.7318363189697,11.6962890625,11.7874507904053,11.8519525527954,11.8804683685303,11.9271488189697,11.9918947219849,12.1180658340454,12.3677253723145,12.3736820220947,12.3838386535645,12.3536138534546,12.3127927780151,12.2967767715454,12.2943363189697,12.2627925872803,12.221923828125,12.1884269714355,11.9993162155151,11.8970699310303,11.9459953308105,12.136474609375,12.2779788970947,12.3093748092651,12.3579587936401,12.379150390625,12.3938474655151,12.41259765625,12.42724609375,12.466064453125,12.5384273529053,12.6364259719849,12.7012691497803,12.7139654159546,12.7162103652954,12.7074222564697,12.6278810501099,12.61328125,12.6198234558105,12.6353998184204,12.6764650344849,12.8342771530151,13.0011224746704,13.0248041152954,13.0418949127197,13.1703615188599,13.3245115280151,13.3648433685303,13.3407716751099,13.32958984375,13.3575201034546,13.412353515625,13.4678707122803,13.5519533157349,13.5811519622803,13.6109371185303,13.6264162063599,13.6500492095947,13.6745119094849,13.6854009628296,13.7034177780151,13.8397455215454,13.9721193313599,14.0763664245605,14.1390132904053,14.2458009719849,14.2880373001099,14.3964357376099,14.4972162246704,14.6529293060303,14.7828121185303,14.8650388717651,14.9114742279053,14.9636726379395,14.9801759719849,14.9790048599243,14.9548816680908,14.9821290969849,14.9869136810303,15.1261234283447,15.2722654342651,15.2864742279053,15.3017082214355,15.3093757629395,15.3204097747803,15.3298835754395,15.3409671783447,15.4083003997803,15.4248533248901,15.4271984100342,15.3911123275757,15.3563480377197,15.4831047058105,15.6416988372803,15.6740236282349,15.7017087936401,15.7552738189697,15.8379878997803,15.8968267440796,15.9456539154053,16.0355472564697,16.1927242279053,16.3577136993408,16.581787109375,16.7981929779053,16.9626941680908,16.9963855743408,17.2884273529053,17.5821781158447,17.8305187225342,18.1394519805908,18.4105949401855,18.7043476104736,18.9680671691895,19.1427726745605,19.1845207214355,19.227783203125,19.2910652160645,19.3595199584961,19.4342288970947,19.4791488647461,19.7319831848145,19.8461418151855,20.0729484558105,20.2480487823486,20.4705066680908,20.6944828033447,20.8730964660645,21.0755863189697,21.380859375,21.6861324310303,21.9914054870605,22.2967281341553,22.6020030975342,22.9072761535645,23.2125968933105,23.5178718566895,23.4016609191895,23.291259765625,23.18017578125,23.1195316314697,22.902099609375,22.6237316131592,22.6196765899658,22.6184558868408,22.7825202941895,22.9962882995605,22.9961910247803],[-29.0176753997803,-29.0283203125,-29.0342769622803,-29.0420913696289,-29.0589847564697,-29.0756855010986,-29.0828113555908,-29.0853538513184,-29.0884780883789,-29.0921859741211,-29.0962886810303,-29.0821285247803,-29.0729503631592,-29.0721683502197,-29.0828113555908,-29.0828113555908,-29.0718746185303,-29.0619144439697,-29.0528316497803,-29.0451164245605,-29.0358409881592,-29.0285148620605,-29.0250968933105,-29.0281257629395,-29.0139636993408,-29.0176753997803],[4.41816425323486,4.38764667510986,4.39511728286743,4.52734327316284,4.49892616271973,4.48720741271973,4.41816425323486],[13.70458984375,13.4895505905151,13.2584962844849,13.0785150527954,12.61279296875,12.5240726470947,12.4840822219849,12.4472169876099,12.3837890625,12.3564939498901,12.3441410064697,12.2982425689697,12.2220706939697,12.2094240188599,12.1509761810303,12.1086912155151,11.9866209030151,11.829833984375,11.7287111282349,11.5911626815796,11.5324220657349,11.4922847747803,11.4461421966553,11.4011716842651,11.2681636810303,11.24853515625,11.2450189590454,11.2118644714355,11.1400880813599,10.8731441497803,10.6050786972046,10.3832521438599,10.1714353561401,10.0361814498901,9.96005821228027,9.91591835021973,9.81401348114014,9.64516544342041,9.56377029418945,9.53964805603027,9.48832988739014,9.42626953125,9.30351638793945,9.17075252532959,9.01943397521973,8.88662052154541,8.81787109375,8.74565410614014,8.66777420043945,8.62412166595459,8.59555625915527,8.41972637176514,8.28232383728027,8.22739219665527,7.94248056411743,7.72778272628784,7.65200185775757,7.58974647521973,7.40073299407959,7.34506845474243,7.27226543426514,7.20195293426514,7.13798809051514,7.11640596389771,7.05620098114014,6.95156240463257,6.88886690139771,6.85463809967041,6.69726610183716,6.65502882003784,6.597412109375,6.533935546875,6.48466825485229,6.45053672790527,6.43793964385986,6.45771503448486,6.50551748275757,6.69789981842041,6.73911094665527,6.77656269073486,6.88178682327271,6.98828172683716,7.06308603286743,7.05771493911743,7.03745079040527,6.93046855926514,6.89125967025757,6.87773466110229,6.87675809860229,6.8916015625,6.91279315948486,6.95917987823486,6.99643611907959,6.92138671875,6.80327177047729,6.783935546875,6.76015567779541,6.65000009536743,6.531982421875,6.47041034698486,6.41865253448486,6.37338876724243,6.31962871551514,6.18613290786743,6.00908231735229,5.917724609375,5.781005859375,5.62968730926514,5.46376991271973,5.19746112823486,5.04687547683716,4.92700242996216,4.83281278610229,4.75522470474243,4.7578125,4.72470712661743,4.74624061584473,4.81376981735229,4.82475614547729,4.92397451400757,4.90747022628784,4.65610361099243,4.55761766433716,4.55537080764771,4.52226543426514,4.52534151077271,4.56093788146973,4.65517568588257,4.64545869827271,4.59492206573486,4.55522489547729,4.54765605926514,4.612060546875,4.68408203125,4.71616220474243,4.68583965301514,4.61557626724243,4.51440477371216,4.39731454849243,4.39067363739014,4.44111299514771,4.52348613739014,4.645263671875,4.72470712661743,4.72470712661743,4.65200185775757,4.59262704849243,4.46914100646973,4.37333965301514,4.34355449676514,4.34243154525757,4.34023427963257,4.37578105926514,4.45517635345459,4.47597646713257,4.34140634536743,4.33193349838257,4.33315420150757,4.30385732650757,4.30942392349243,4.33447265625,4.37167930603027,4.43212890625,4.385498046875,4.29228496551514,4.27739238739014,4.29062509536743,4.33857393264771,4.38774394989014,4.45595693588257,4.647216796875,4.73320341110229,4.83876943588257,4.94584989547729,5.12900400161743,5.14228534698486,5.12656259536743,5.15385770797729,5.17377901077271,5.19501972198486,5.25927782058716,5.33774423599243,5.36533212661743,5.37861299514771,5.42636728286743,5.47421884536743,5.40175819396973,5.48378896713257,5.53354501724243,5.57168006896973,5.57749032974243,5.57451152801514,5.61171865463257,5.62470722198486,5.623291015625,5.64794921875,5.70751953125,5.6943359375,5.64155292510986,5.60273408889771,5.64155292510986,5.72812509536743,5.76708984375,5.79750919342041,6.02631855010986,6.21718740463257,6.34858417510986,6.411376953125,6.408935546875,6.42705106735229,6.45727586746216,6.47744131088257,6.58383798599243,6.59794950485229,6.53134727478027,6.52499961853027,6.39692401885986,6.375732421875,6.36923837661743,6.42768573760986,6.595703125,6.66176795959473,6.71171855926514,6.77163076400757,6.85283231735229,6.98027324676514,7.01982450485229,7.06791973114014,7.14321231842041,7.39506816864014,7.42250919342041,7.44340801239014,7.47685623168945,7.54189395904541,7.61625957489014,7.72309541702271,7.82661151885986,7.87373018264771,8.0498046875,8.27299785614014,8.371826171875,8.44189453125,8.614013671875,8.78251934051514,9.04853439331055,9.06137752532959,9.08383846282959,9.18828105926514,9.32060527801514,9.45161151885986,9.49467754364014,9.56562519073486,9.66704082489014,9.77846717834473,9.81279373168945,9.83862400054932,9.85190391540527,9.90732383728027,10.0045413970947,10.16015625,10.2683591842651,10.29248046875,10.3506841659546,10.4089841842651,10.4277830123901,10.4126949310303,10.4176263809204,10.4358882904053,10.607421875,10.6537599563599,10.7687492370605,10.8504390716553,10.971923828125,11.07958984375,11.1203126907349,11.1545906066895,11.1768560409546,11.3954095840454,11.4992189407349,11.6318845748901,11.6962890625,11.7318363189697,11.762451171875,11.7996578216553,11.8277339935303,11.8873043060303,11.9269533157349,11.9703617095947,12.0615720748901,12.2016115188599,12.4052734375,12.5299806594849,12.6221685409546,12.7750482559204,12.9346685409546,13.0554685592651,13.4577150344849,13.4821290969849,13.5010747909546,13.6475095748901,13.7018070220947,13.7332029342651,13.759765625,13.7491216659546,13.7427244186401,13.7572269439697,13.8368654251099,13.8591794967651,13.8728523254395,13.765380859375,13.6636714935303,13.6729984283447,13.6587886810303,13.6036138534546,13.4854011535645,13.4091310501099,13.3642578125,13.107666015625,13.0432615280151,13.0082025527954,12.9955568313599,13.0001955032349,13.0291013717651,13.0863285064697,13.1122560501099,13.1071786880493,13.337890625,13.3409175872803,13.32275390625,13.2911624908447,13.0596685409546,12.9081544876099,12.8574705123901,12.8214845657349,12.8106451034546,13.13525390625,13.2061519622803,13.2701177597046,13.281005859375,13.3302249908447,13.3715333938599,13.3536128997803,13.3405275344849,13.2977056503296,13.1917963027954,13.0904302597046,13.0736808776855,13.09375,13.1943359375,13.3265619277954,13.38037109375,13.4490232467651,13.5345211029053,13.5730466842651,13.6708498001099,13.7017583847046,13.70458984375],[14.9930658340454,14.9563961029053,14.932373046875,14.8127937316895,14.8020992279053,14.8905277252197,14.902099609375,14.8706541061401,14.8253421783447,14.7661123275757,14.7652826309204,14.7490234375,14.340087890625,14.2671384811401,14.1536130905151,14.0569829940796,13.9964838027954,13.7388191223145,13.3203125,12.9439449310303,12.5962896347046,12.5141115188599,12.4118165969849,12.3934078216553,12.396484375,12.4593267440796,12.5145511627197,12.579345703125,12.6671390533447,12.7130861282349,12.6128902435303,12.568115234375,12.5526371002197,12.5019521713257,12.40673828125,12.3370609283447,12.2870607376099,12.2275400161743,12.0243167877197,12.0299806594849,12.0574216842651,12.0592775344849,11.9773921966553,11.931640625,11.8963871002197,11.8610353469849,11.836181640625,11.8212890625,11.8245601654053,11.723876953125,11.6420412063599,11.5665044784546,11.5039548873901,11.4281730651855,11.3536624908447,11.3000497817993,11.1305170059204,11.01025390625,10.933837890625,10.917236328125,10.8368654251099,10.785888671875,10.7432613372803,10.7353515625,10.7756834030151,10.7803707122803,10.8017091751099,10.84130859375,10.9007329940796,10.9798831939697,10.9744625091553,10.9916505813599,11.0456056594849,11.0521974563599,11.0059080123901,10.9453125,11.0399408340454,11.1064453125,11.1663084030151,11.189453125,11.1844720840454,11.153076171875,11.0974607467651,11.08154296875,11.0662593841553,11.0621099472046,11.0885734558105,11.19873046875,11.3313474655151,11.73828125,11.9815425872803,12.1566400527954,12.2477531433105,12.4341306686401,12.5083494186401,12.757568359375,12.903564453125,12.9656734466553,13.0433101654053,13.0396966934204,12.9841804504395,12.921142578125,12.920654296875,12.949951171875,12.979248046875,12.991455078125,13.0078125,13.0537118911743,13.1175298690796,13.1793947219849,13.2235841751099,13.2665033340454,13.27978515625,13.2843751907349,13.3133792877197,13.4072265625,13.63525390625,13.69873046875,13.7461423873901,13.7634763717651,13.7748537063599,13.7556638717651,13.7700681686401,13.8994626998901,13.9945802688599,14.0372066497803,14.0501461029053,13.9656744003296,13.8444337844849,13.85205078125,13.8586921691895,13.8760738372803,13.9318361282349,13.9825687408447,14.0282220840454,14.1086912155151,14.2238779067993,14.2916498184204,14.3118162155151,14.3433103561401,14.3859872817993,14.4466304779053,14.525146484375,14.5706052780151,14.5829591751099,14.607666015625,14.6447267532349,14.685546875,14.7524423599243,14.8097658157349,14.7903804779053,14.7133779525757,14.6610851287842,14.6333990097046,14.6437015533447,14.6917486190796,14.7063484191895,14.6873540878296,14.6981449127197,14.738525390625,14.7478523254395,14.7260265350342,14.7204103469849,14.7310056686401,14.7506351470947,14.7708988189697,14.7860841751099,14.7710933685303,14.7944812774658,14.8562507629395,14.8835439682007,14.8764162063599,14.907567024231,14.9770011901855,15.008056640625,14.9930658340454],[-19.0830078125,-19.1378917694092,-19.0728530883789,-18.990234375,-18.9686527252197,-18.9660167694092,-19.0425777435303,-19.0830078125],[12.1445798873901,12.0320796966553,12.0822763442993,12.1717281341553,12.2067384719849,12.2280769348145,12.2575187683105,12.3019533157349,12.2312498092651,12.1445798873901],[17.4956550598145,17.4750499725342,17.4769039154053,17.496826171875,17.5303707122803,17.5211925506592,17.5160636901855,17.50927734375,17.4956550598145],[17.6231441497803,17.6195812225342,17.628662109375,17.645263671875,17.6518058776855,17.6472187042236,17.6341304779053,17.6231441497803],[51.386474609375,51.3487319946289,51.3070793151855,51.2470703125,51.2076683044434,51.2125968933105,51.2332038879395,51.2548332214355,51.2756843566895,51.2861824035645,51.2636222839355,51.2422370910645,51.2457504272461,51.2636222839355,51.2911148071289,51.377685546875,51.3935050964355,51.3994140625,51.369140625,51.3544921875,51.3959465026855,51.3606452941895,51.386474609375],[51.7394561767578,51.6849136352539,51.66748046875,51.6487808227539,51.6270484924316,51.6939926147461,51.67822265625,51.7099113464355,51.7296867370605,51.7464370727539,51.7394561767578],[53.0707015991211,52.9998054504395,53.0196304321289,53.0360374450684,53.09130859375,53.1833038330078,53.0707015991211],[53.3080062866211,53.2345695495605,53.2462387084961,53.3102035522461,53.3080062866211],[51.386474609375,51.4015159606934,51.4432106018066,51.4093742370605,51.4499015808105,51.4539031982422,51.4861793518066,51.540771484375,51.57666015625,51.589111328125,51.5960464477539,51.57421875,51.4557647705078,51.4566879272461,51.4716300964355,51.50390625,51.519287109375,51.5511207580566,51.5958518981934,51.6103019714355,51.6334457397461,51.6729011535645,51.810546875,51.8478012084961,51.927734375,51.9940910339355,52.0119132995605,52.0589828491211,52.1968231201172,52.3091773986816,52.4425773620605,52.8097648620605,52.8721199035645,52.9413070678711,52.9282684326172,52.9083480834961,52.9606437683105,53.0964851379395,53.2140617370605,53.2686996459961,53.3751945495605,53.4070777893066,53.415283203125,53.4342765808105,53.441162109375,53.3753890991211,53.3272972106934,53.3005867004395,53.2822761535645,53.1872062683105,52.99951171875,52.9662132263184,52.8870124816895,52.7447776794434,52.6513671875,52.633544921875,52.6340827941895,52.6178703308105,52.59765625,52.5735855102539,52.5496597290039,52.5301742553711,52.4992179870605,52.4640159606934,52.4422836303711,52.4402847290039,52.444091796875,52.4189949035645,52.3802261352539,52.3314971923828,52.2660140991211,52.2055168151855,52.1357917785645,52.1112327575684,52.0986824035645,52.0802230834961,52.056884765625,52.0361824035645,51.9801750183105,51.9673805236816,51.9382820129395,51.910888671875,51.8539543151855,51.8583984375,51.8300285339355,51.8246574401855,51.8507308959961,51.8807601928711,51.8704109191895,51.8539543151855,51.833984375,51.8026847839355,51.7624015808105,51.6582489013672,51.6377944946289,51.5989265441895,51.5500984191895,51.4889183044434,51.4500007629395,51.4105949401855,51.3548316955566,51.22412109375,51.1990242004395,51.1799812316895,51.1747093200684,51.1648445129395,51.1474151611328,51.0566902160645,51.0408210754395,51.0453147888184,51.0301246643066,51.0056648254395,50.9842262268066,50.9729499816895,50.949951171875,50.9048843383789,50.7504386901855,50.752555847168,50.7545433044434,50.7595710754395,50.7747535705566,50.8059539794922,50.8436050415039,50.8666496276855,50.93212890625,50.9502449035645,50.9599113464355,50.98876953125,51.08642578125,51.1256370544434,51.153076171875,51.1694831848145,51.1984405517578,51.2393074035645,51.2749977111816,51.2850608825684,51.2729988098145,51.2597160339355,51.2789573669434,51.3464851379395,51.4068336486816,51.453125,51.4690933227539,51.4453620910645,51.4077644348145,51.4032745361328,51.4120597839355,51.4328117370605,51.4527320861816,51.4773941040039,51.4911117553711,51.4217262268066,51.4219245910645,51.4485855102539,51.4747047424316,51.4598121643066,51.4275894165039,51.3670883178711,51.3560028076172,51.3615226745605,51.386474609375],[53.3857421875,53.3777809143066,53.3917961120605,53.4314460754395,53.4435539245605,53.4380836486816,53.3857421875],[53.4588394165039,53.442626953125,53.4548835754395,53.4665031433105,53.473388671875,53.47509765625,53.4649887084961,53.4588394165039],[53.5107421875,53.476806640625,53.4839363098145,53.4937515258789,53.5149879455566,53.5107421875],[53.58251953125,53.5792007446289,53.6056632995605,53.62548828125,53.62548828125,53.58251953125],[60.3075637817383,60.1887664794922,60.1977577209473,60.2433128356934,60.2724113464355,60.3411636352539,60.4120597839355,60.4472618103027,60.4520492553711,60.3889656066895,60.3075637817383],[61.0845680236816,61.0719261169434,61.0827178955078,61.17822265625,61.1938514709473,61.1993637084961,61.1482467651367,61.0845680236816],[63.3376007080078,63.3369102478027,63.3523483276367,63.3850555419922,63.4139137268066,63.4498062133789,63.4708023071289,63.4313468933105,63.3664054870605,63.3376007080078],[63.6671371459961,63.664794921875,63.6871566772461,63.7318382263184,63.7743148803711,63.8013153076172,63.8046340942383,63.7714347839355,63.7259788513184,63.7034683227539,63.6671371459961],[64.8658676147461,64.8380355834961,64.8603973388672,64.8431167602539,64.8703155517578,64.9232406616211,64.9787139892578,64.9761734008789,64.9079055786133,64.8658676147461],[65.6265106201172,65.595703125,65.6045379638672,65.6309585571289,65.6838836669922,65.7059097290039,65.7015609741211,65.679443359375,65.6265106201172],[65.9019546508789,65.8990707397461,65.93994140625,65.9770965576172,66.0019073486328,66.008544921875,66.0113754272461,65.99169921875,65.9638671875,65.9019546508789],[66.0432662963867,66.03662109375,66.0807647705078,66.1226577758789,66.1513214111328,66.1850128173828,66.2105484008789,66.1779251098633,66.1224594116211,66.0719223022461,66.0432662963867],[67.8741226196289,67.8212356567383,67.9177780151367,68.0154800415039,68.0713424682617,68.0494155883789,68.002685546875,67.9564514160156,67.9345703125,67.8741226196289],[68.2653274536133,68.2482376098633,68.2532196044922,68.246826171875,68.2186050415039,68.1875457763672,68.1685028076172,68.12109375,68.10498046875,68.1047897338867,68.0938415527344,68.0516586303711,68.082763671875,68.0606918334961,68.021240234375,68.0096664428711,67.995361328125,68.0872497558594,68.12060546875,68.1604537963867,68.1665496826172,68.1632308959961,68.2490234375,68.2733917236328,68.276123046875,68.2653274536133],[68.943115234375,68.8424301147461,68.7835922241211,68.6724090576172,68.6163024902344,68.6063461303711,68.6109848022461,68.6379928588867,68.6682662963867,68.6772003173828,68.6383361816406,68.633056640625,68.6632308959961,68.71142578125,68.7718734741211,68.81884765625,68.814697265625,68.8001022338867,68.7909698486328,68.8475570678711,68.8866729736328,68.9138717651367,68.894287109375,69.0005416870117,69.0080108642578,69.0039520263672,68.9815444946289,68.943115234375],[68.5612258911133,68.55419921875,68.6504898071289,68.6805191040039,68.7140121459961,68.7464370727539,68.7993698120117,68.8423843383789,68.853759765625,68.8683090209961,68.8763198852539,68.841552734375,68.8029251098633,68.716552734375,68.6330032348633,68.56787109375,68.5384750366211,68.4636688232422,68.4024963378906,68.3892593383789,68.3942337036133,68.4090270996094,68.4090881347656,68.3560104370117,68.3128433227539,68.3252944946289,68.3782272338867,68.3738250732422,68.3104019165039,68.2892074584961,68.28271484375,68.3065872192383,68.198486328125,68.1782684326172,68.1907730102539,68.2569351196289,68.341552734375,68.4003448486328,68.44140625,68.6158218383789,68.8053207397461,68.8737258911133,68.9123992919922,68.919189453125,68.9785690307617,69.04345703125,69.132568359375,69.1705093383789,69.2778778076172,69.3020477294922,69.3020477294922,69.27392578125,69.2164077758789,69.1126480102539,69.0242156982422,68.9607467651367,68.9084930419922,68.8191909790039,68.7332077026367,68.6170425415039,68.5612258911133],[69.5962371826172,69.5390625,69.5565414428711,69.5630416870117,69.54296875,69.5066452026367,69.5049819946289,69.45751953125,69.395751953125,69.349609375,69.3287048339844,69.2743225097656,69.1981430053711,69.1720199584961,69.160400390625,69.1968231201172,69.1300354003906,69.0259246826172,69.013671875,69.046630859375,69.0693893432617,69.0707015991211,69.0951156616211,69.1123580932617,69.1378936767578,69.1906204223633,69.2847137451172,69.3303680419922,69.3619155883789,69.3988265991211,69.3814926147461,69.4167022705078,69.4388656616211,69.4776916503906,69.5038070678711,69.5271453857422,69.5301818847656,69.5696792602539,69.5868606567383,69.5962371826172],[69.7967758178711,69.7675247192383,69.7916030883789,69.8290557861328,69.9055633544922,69.9024429321289,69.8732452392578,69.83837890625,69.7967758178711],[70.0897445678711,70.0665054321289,70.0570297241211,70.0714340209961,70.0809097290039,70.0765609741211,70.119140625,70.1548843383789,70.2033233642578,70.2308578491211,70.2195281982422,70.2054672241211,70.0897445678711],[70.06640625,70.011962890625,70.0171813964844,70.0377426147461,70.0478973388672,70.0191345214844,69.9701690673828,69.9084014892578,69.8202667236328,69.7998046875,69.8104476928711,69.7595672607422,69.7066879272461,69.6398468017578,69.6053695678711,69.5790100097656,69.5528335571289,69.5354995727539,69.557861328125,69.6020965576172,69.6261215209961,69.6357421875,69.6767578125,69.7018051147461,69.7154846191406,69.7678680419922,69.7815399169922,69.7686538696289,69.8065872192383,69.8130340576172,69.7816390991211,69.8248519897461,69.8643035888672,69.8909225463867,69.9601058959961,70.0105438232422,70.0430145263672,70.037841796875,70.085693359375,70.1346740722656,70.1666030883789,70.244140625,70.2474594116211,70.1785583496094,70.06640625],[70.2166976928711,70.2049789428711,70.2122573852539,70.201904296875,70.1492691040039,70.1285629272461,70.10205078125,70.0762176513672,70.0684585571289,70.0774383544922,70.1104965209961,70.1653289794922,70.2190933227539,70.2661666870117,70.2735824584961,70.2166976928711],[70.54931640625,70.5025405883789,70.4639663696289,70.4081497192383,70.3349609375,70.3152847290039,70.2964859008789,70.2826156616211,70.2960968017578,70.3588409423828,70.3776321411133,70.352783203125,70.3846664428711,70.416748046875,70.444580078125,70.4869155883789,70.5160598754883,70.505126953125,70.6170883178711,70.5936508178711,70.54931640625],[70.5673828125,70.5274887084961,70.5618667602539,70.5970687866211,70.6752471923828,70.7228012084961,70.7473678588867,70.7293930053711,70.71435546875,70.6996078491211,70.6505889892578,70.5673828125],[70.8157730102539,70.784423828125,70.75390625,70.7216796875,70.5940933227539,70.5735397338867,70.5535202026367,70.5415496826172,70.5590286254883,70.5331573486328,70.515869140625,70.5091781616211,70.5148010253906,70.5621109008789,70.6133270263672,70.6571350097656,70.6562957763672,70.6668930053711,70.6576614379883,70.7025833129883,70.6971740722656,70.7284164428711,70.7109909057617,70.8154754638672,70.812744140625,70.8425750732422,70.8157730102539],[69.783447265625,69.6517562866211,69.6058120727539,69.5612258911133,69.5384292602539,69.5285186767578,69.5325698852539,69.584716796875,69.6337890625,69.6358413696289,69.6298828125,69.58056640625,69.5427703857422,69.5016174316406,69.4642639160156,69.432861328125,69.3924789428711,69.3604507446289,69.2981414794922,69.2706069946289,69.0970230102539,69.071533203125,69.0499572753906,69.02197265625,69.0605926513672,69.1189956665039,69.1769104003906,69.2879867553711,69.36669921875,69.3939437866211,69.4729995727539,69.6714324951172,69.7314910888672,69.82275390625,69.8714370727539,69.9716796875,70.0616683959961,70.0648422241211,70.042236328125,69.9600601196289,69.918701171875,69.906494140625,69.9046859741211,69.9281234741211,69.9330596923828,69.9263153076172,69.9150390625,69.7819290161133,69.7146987915039,69.6915588378906,69.6526336669922,69.588623046875,69.3665084838867,69.2826690673828,69.2314453125,69.0761184692383,68.9901351928711,68.9191436767578,68.8871536254883,68.880615234375,68.8624496459961,68.8213348388672,68.7652893066406,68.6395950317383,68.59326171875,68.6064910888672,68.65283203125,68.6886749267578,68.7115249633789,68.7608871459961,68.7984390258789,68.805908203125,68.7583999633789,68.7138671875,68.6776351928711,68.6489715576172,68.642578125,68.6743621826172,68.6953125,68.72021484375,68.7198715209961,68.776611328125,68.8558578491211,68.9927749633789,69.0411148071289,69.1544876098633,69.2707061767578,69.2735824584961,69.2774887084961,69.273681640625,69.2472686767578,69.214111328125,69.1865692138672,69.080810546875,69.054443359375,69.041748046875,69.0714340209961,69.0694808959961,69.036865234375,69.0333023071289,69.0208969116211,68.934326171875,68.899658203125,68.8487319946289,68.7540512084961,68.6731491088867,68.6073226928711,68.5420379638672,68.4775390625,68.390380859375,68.3563995361328,68.3622589111328,68.3924331665039,68.46533203125,68.4927291870117,68.5011215209961,68.5000457763672,68.5624008178711,68.555419921875,68.5284118652344,68.4677734375,68.3168411254883,68.2006378173828,68.1334457397461,68.0878372192383,67.9648895263672,68.0484390258789,68.1038131713867,68.0301284790039,67.89501953125,67.6283187866211,67.6195831298828,67.5517578125,67.5206069946289,67.5051803588867,67.4258346557617,67.3120574951172,67.2520065307617,67.1550750732422,67.0933609008789,67.0549774169922,66.9764175415039,66.7688522338867,66.5521011352539,66.4898452758789,66.3059616088867,66.2520523071289,66.1910629272461,66.1675262451172,66.1537094116211,66.1293487548828,65.9322738647461,65.8450164794922,65.7932586669922,65.7428741455078,65.6463928222656,65.3014678955078,65.2643585205078,65.1708526611328,64.9461441040039,64.7967758178711,64.58154296875,64.5135726928711,64.4640121459961,64.3877410888672,64.2603073120117,64.1735305786133,64.0955047607422,64.0407180786133,64.0140151977539,64.0406265258789,64.0748062133789,64.0750961303711,64.0504913330078,64,63.9574203491211,63.9404830932617,63.8435554504395,63.6711959838867,63.595947265625,63.4922370910645,63.2917022705078,63.0891609191895,63.08251953125,63.0006332397461,62.9478492736816,62.9194831848145,62.8259239196777,62.7213401794434,62.6600112915039,62.5918960571289,62.2855949401855,62.2137680053711,62.1674308776855,61.9768562316895,61.7207527160645,61.6534690856934,61.5730018615723,61.5413093566895,61.4457015991211,61.3522987365723,61.2902870178223,61.2218284606934,61.1739730834961,61.1082534790039,61.0598640441895,61.0468254089355,61.04150390625,61.0231895446777,61.002685546875,60.8921394348145,60.6896476745605,60.5456504821777,60.4507331848145,60.3544960021973,60.3052215576172,60.2388687133789,60.1067848205566,60.0400390625,59.9672355651855,59.9128875732422,59.8976097106934,59.8913078308105,59.8636703491211,59.7824745178223,59.6971702575684,59.5922889709473,59.5557594299316,59.4314422607422,59.2898941040039,59.1575660705566,59.0186538696289,58.9260787963867,58.8930206298828,58.9095191955566,59.0365219116211,59.0657196044922,59.1045417785645,59.1432151794434,59.1417961120605,59.1644515991211,59.1708526611328,59.1839370727539,59.2959938049316,59.3892135620117,59.4281730651855,59.6024932861328,59.6800270080566,59.7645530700684,59.6957969665527,59.5871086120605,59.54150390625,59.5193328857422,59.4556617736816,59.4435997009277,59.3774909973145,59.2796363830566,59.0620613098145,59.0386734008789,59.00927734375,59.02880859375,58.9682121276855,58.9584922790527,59.0270462036133,59.1177749633789,59.1126899719238,59.0679206848145,59.0097198486328,58.97119140625,58.9460411071777,58.9330101013184,58.8568344116211,58.8056602478027,58.74755859375,58.739013671875,58.7118606567383,58.6749992370605,58.5699729919434,58.3005905151367,58.2244606018066,58.1453132629395,58.1472663879395,58.0799827575684,58.0209465026855,58.0476531982422,58.0242195129395,58.0705070495605,58.102294921875,58.1207504272461,58.1428718566895,58.1507339477539,58.154541015625,58.1322250366211,58.0815467834473,58.0683097839355,58.0973167419434,58.1234359741211,58.1763687133789,58.2240219116211,58.2337913513184,58.2627449035645,58.2664031982422,58.2594223022461,58.2679672241211,58.3751487731934,58.4323234558105,58.5236358642578,58.6204109191895,58.7265129089355,58.8226547241211,58.9751930236816,59.0128898620605,58.9594688415527,58.8702621459961,58.8746604919434,58.9446754455566,59.0009307861328,59.0164527893066,58.9519500732422,58.9876937866211,59.0604972839355,59.0979537963867,59.1354484558105,59.1861305236816,59.2339820861816,59.299072265625,59.3681602478027,59.4380874633789,59.5055656433105,59.547119140625,59.5609855651855,59.5345230102539,59.4896507263184,59.4144554138184,59.3534698486328,59.3298377990723,59.3102531433105,59.2912139892578,59.2038078308105,59.1663551330566,59.1625518798828,59.2264633178711,59.4536628723145,59.5643043518066,59.642578125,59.65576171875,59.7130851745605,59.7130851745605,59.6866188049316,59.6609382629395,59.7339859008789,59.7446746826172,59.818359375,59.831787109375,59.8155746459961,59.8131828308105,59.794677734375,59.8079109191895,59.8630828857422,59.9127922058105,60.031494140625,60.0834922790527,60.1320762634277,60.1651344299316,60.2334976196289,60.3529777526855,60.4075660705566,60.360595703125,60.213623046875,60.1529312133789,60.3672332763672,60.4181671142578,60.4541015625,60.4782257080078,60.5119667053223,60.5007858276367,60.4190902709961,60.3462371826172,60.2901344299316,60.2055625915527,60.150634765625,60.0700187683105,60.0262222290039,60.010009765625,59.9077682495117,59.8255844116211,59.7601051330566,59.7097663879395,59.6917953491211,59.6422843933105,59.6388206481934,59.6678237915039,59.731689453125,59.8336944580078,59.9070777893066,59.978759765625,60.0457038879395,60.0879402160645,60.0864715576172,60.0672378540039,60.0703125,60.1231918334961,60.1541061401367,60.1584968566895,60.1541061401367,60.2057151794434,60.3083992004395,60.4456024169922,60.4848136901855,60.6245613098145,60.6879844665527,60.6942901611328,60.6173362731934,60.5695762634277,60.635986328125,60.70751953125,60.8585433959961,60.9361343383789,61.0381851196289,61.0713386535645,61.0537071228027,61.0471649169922,61.0561065673828,61.1173324584961,61.0809555053711,61.0842819213867,61.1370086669922,61.1424331665039,61.1021499633789,61.0559539794922,60.9941444396973,60.9529304504395,60.96630859375,61.0152816772461,61.0911636352539,61.1771507263184,61.2105484008789,61.22216796875,61.3005867004395,61.4192390441895,61.4346199035645,61.3720245361328,61.283935546875,61.213623046875,61.1809539794922,61.1659698486328,61.1605453491211,61.1903800964355,61.2065887451172,61.2290992736816,61.279296875,61.2896499633789,61.2445297241211,61.1545906066895,61.1338882446289,61.1672897338867,61.1476097106934,61.1023445129395,61.1082534790039,61.1875457763672,61.2505874633789,61.377685546875,61.4335975646973,61.4571266174316,61.4554672241211,61.4855003356934,61.5050277709961,61.5433578491211,61.6201629638672,61.6452178955078,61.7106895446777,61.8095703125,61.8783187866211,61.9004402160645,61.8853988647461,61.8969230651855,61.8270988464355,61.7875022888184,61.8074188232422,61.8697738647461,61.8870124816895,61.8509750366211,61.8524436950684,61.9228973388672,61.9456100463867,61.9355964660645,61.9569816589355,62.0266609191895,62.159912109375,62.1886749267578,62.1539077758789,62.1517105102539,62.2073745727539,62.2391090393066,62.3108863830566,62.3789100646973,62.3846664428711,62.416015625,62.4071311950684,62.3756866455078,62.3496055603027,62.352783203125,62.4072761535645,62.4232902526855,62.4680709838867,62.4480972290039,62.4163093566895,62.407470703125,62.4471702575684,62.482421875,62.5199241638184,62.5838356018066,62.6111335754395,62.6096649169922,62.6212921142578,62.6378898620605,62.6267547607422,62.6022987365723,62.5428199768066,62.5481948852539,62.5640144348145,62.5855979919434,62.6103019714355,62.6455039978027,62.6720657348633,62.7210006713867,62.7318382263184,62.771240234375,62.7117652893066,62.7523422241211,62.7520027160645,62.7288055419922,62.7007331848145,62.7207069396973,62.7896499633789,62.9027366638184,62.9304656982422,62.9576644897461,63.0232887268066,63.0995140075684,63.1091804504395,63.1038589477539,63.11279296875,63.0909690856934,62.9955101013184,62.9655265808105,62.8462371826172,62.8805656433105,63.0421867370605,63.0821800231934,63.1615257263184,63.2365226745605,63.2865715026855,63.3133811950684,63.3423347473145,63.3920936584473,63.4261207580566,63.4241676330566,63.4452667236328,63.4988746643066,63.5351104736328,63.6011695861816,63.6226043701172,63.6458969116211,63.5936508178711,63.5662612915039,63.5003929138184,63.4634284973145,63.4593276977539,63.5703582763672,63.585693359375,63.6095733642578,63.6245574951172,63.5241661071777,63.4920387268066,63.4788551330566,63.3952598571777,63.3908233642578,63.4327163696289,63.4547882080078,63.4693336486816,63.4472198486328,63.4635772705078,63.4612808227539,63.5363235473633,63.5580024719238,63.625,63.6511764526367,63.6981925964355,63.7191886901855,63.7638168334961,63.8048324584961,63.8376922607422,63.875732421875,63.8781204223633,63.8988800048828,63.9481925964355,64.0029754638672,64.0245132446289,64.0488739013672,64.030517578125,63.9881362915039,63.9210968017578,63.9015617370605,63.84521484375,63.7702102661133,63.571044921875,63.5126914978027,63.5217781066895,63.5762214660645,63.6165046691895,63.6995124816895,63.6973114013672,63.678955078125,63.7061500549316,63.7948265075684,63.8648910522461,63.9178237915039,63.9817352294922,64.0831604003906,64.1796417236328,64.4183044433594,64.4944839477539,64.5777359008789,64.6145553588867,64.6794967651367,64.6859359741211,64.744384765625,64.8139190673828,64.8182601928711,64.7730026245117,64.7547836303711,64.8293991088867,64.9059066772461,64.975830078125,65.178955078125,65.1453628540039,65.0859832763672,65.0994110107422,65.2144088745117,65.3392562866211,65.3174819946289,65.266357421875,65.1953125,65.18408203125,65.1933059692383,65.24072265625,65.256103515625,65.2454605102539,65.2791519165039,65.3623580932617,65.4862289428711,65.5681610107422,65.6301727294922,65.80615234375,65.9021987915039,65.952880859375,65.9416046142578,65.9562454223633,66.0191879272461,66.069091796875,66.1004409790039,66.1827621459961,66.1799774169922,66.2210464477539,66.2473602294922,66.2975616455078,66.3197326660156,66.2735824584961,66.2519073486328,66.2525863647461,66.2367172241211,66.2306671142578,66.4308090209961,66.5394058227539,66.5371551513672,66.6408157348633,66.7018585205078,66.7155227661133,66.7416534423828,66.7948226928711,66.782470703125,66.7943344116211,66.8193817138672,66.8516616821289,66.9070816040039,66.9380416870117,66.9607925415039,66.9648971557617,67.0730895996094,67.1192398071289,67.1112289428711,67.158935546875,67.1426773071289,67.1738815307617,67.1944885253906,67.2024459838867,67.2466812133789,67.2569351196289,67.268310546875,67.2674331665039,67.2559585571289,67.2713851928711,67.2978515625,67.3397445678711,67.3860321044922,67.4990234375,67.5742645263672,67.5550308227539,67.483154296875,67.4741668701172,67.450927734375,67.3517608642578,67.3485336303711,67.44384765625,67.5213851928711,67.5428161621094,67.5147933959961,67.5439453125,67.6021423339844,67.6553726196289,67.7079620361328,67.7344284057617,67.7652816772461,67.6825714111328,67.663330078125,67.6749038696289,67.7498550415039,67.809326171875,67.9557647705078,67.9727020263672,67.9609375,67.9196243286133,67.9262161254883,67.9482955932617,67.9878845214844,68.0036087036133,68.0364761352539,68.0687484741211,68.1028289794922,68.1643600463867,68.1821823120117,68.2287139892578,68.2181701660156,68.1999053955078,68.02734375,67.8865737915039,67.8814468383789,68.001220703125,68.03564453125,68.0618209838867,68.0916061401367,68.1017608642578,68.14453125,68.28125,68.3167495727539,68.3895492553711,68.4062957763672,68.3552780151367,68.3546905517578,68.368408203125,68.4103546142578,68.4263153076172,68.42626953125,68.4474639892578,68.4610824584961,68.4743118286133,68.48193359375,68.4592819213867,68.4664611816406,68.4906692504883,68.5325698852539,68.5926742553711,68.6257781982422,68.6854019165039,68.6934585571289,68.7993698120117,68.8787612915039,69.0011291503906,69.1000442504883,69.1563034057617,69.1812057495117,69.2326126098633,69.3252487182617,69.43310546875,69.4706039428711,69.47509765625,69.4398422241211,69.3648452758789,69.3218765258789,69.314453125,69.3355941772461,69.37841796875,69.4343795776367,69.4905776977539,69.5203628540039,69.5170440673828,69.5233383178711,69.5611343383789,69.5876922607422,69.6237335205078,69.660400390625,69.7478485107422,69.8047409057617,69.7816390991211,69.6129379272461,69.5038070678711,69.4240264892578,69.5038070678711,69.7221221923828,69.8246078491211,69.8834457397461,69.896728515625,69.9272003173828,69.9453125,69.9219207763672,69.8676223754883,69.6769485473633,69.6166458129883,69.5358428955078,69.3556671142578,69.3326644897461,69.3412170410156,69.3709487915039,69.5420913696289,69.5205078125,69.5345230102539,69.584716796875,69.6328125,69.6923370361328,69.85107421875,69.9139175415039,69.9073257446289,69.916015625,69.887451171875,69.8895034790039,70.0032272338867,70.01318359375,69.9380340576172,69.887451171875,69.814697265625,69.8345718383789,70.0042495727539,70.0660629272461,70.0981903076172,70.1744613647461,70.208251953125,70.2333984375,70.2576675415039,70.2298812866211,70.2933578491211,70.2759704589844,70.3091812133789,70.2645034790039,70.2777328491211,70.3375930786133,70.374755859375,70.3404769897461,70.3049850463867,70.2367706298828,70.1018600463867,70.029052734375,69.9933166503906,69.9833984375,70.019775390625,70.0635757446289,70.1048355102539,70.2072296142578,70.2474594116211,70.3997573852539,70.478759765625,70.4853515625,70.6624069213867,70.694580078125,70.7020034790039,70.7453079223633,70.772705078125,70.8263168334961,70.8915481567383,71.0010223388672,71.0084533691406,70.97802734375,70.9286117553711,70.8720245361328,70.843505859375,70.8494186401367,70.8919372558594,70.911865234375,70.9006805419922,70.8733444213867,70.8697280883789,70.8531723022461,70.8167953491211,70.7771453857422,70.6719741821289,70.5523910522461,70.4894027709961,70.3240203857422,70.2182083129883,70.1439971923828,70.1090316772461,70.136474609375,70.2354965209961,70.340576171875,70.6253890991211,70.7826156616211,70.9127960205078,70.9397430419922,70.8535614013672,70.740966796875,70.6691360473633,70.63623046875,70.5508804321289,70.5034637451172,70.4538116455078,70.4100112915039,70.4216842651367,70.5113754272461,70.6084442138672,70.6811981201172,70.7440490722656,70.8035659790039,70.8040008544922,70.827392578125,70.9100036621094,70.9472198486328,70.9967269897461,71.09130859375,71.0808563232422,71.0593719482422,71.0430145263672,70.9752960205078,70.8694381713867,70.8251953125,70.7979507446289,70.7175827026367,70.6779327392578,70.6642532348633,70.704345703125,70.66796875,70.576904296875,70.440185546875,70.3604049682617,70.2876434326172,70.2485809326172,70.4034194946289,70.4430694580078,70.5013732910156,70.6188049316406,70.7596664428711,70.8415069580078,70.8639602661133,70.8607406616211,70.8299255371094,70.761474609375,70.734130859375,70.7050323486328,70.6685485839844,70.6468276977539,70.6425323486328,70.6943817138672,70.7029800415039,70.6221694946289,70.5623016357422,70.5433044433594,70.5471649169922,70.523681640625,70.401123046875,70.3438415527344,70.2744140625,70.1978530883789,70.1247024536133,70.0964813232422,70.1454086303711,70.092529296875,69.9767532348633,69.9436950683594,69.8740692138672,69.8182144165039,69.7801284790039,69.7445755004883,69.7278747558594,69.7366714477539,69.7175750732422,69.7459411621094,69.8411636352539,69.8622055053711,69.8345718383789,69.7328109741211,69.7222671508789,69.7948760986328,69.7896499633789,69.7957000732422,69.783447265625],[71.14208984375,71.1038513183594,71.1046371459961,71.0331573486328,71.0394973754883,70.9958038330078,70.97509765625,70.9625015258789,70.9538116455078,70.9607849121094,71.0195846557617,71.0341339111328,71.052978515625,71.097412109375,71.14208984375],[70.8391571044922,70.8326644897461,70.8548812866211,70.9159240722656,71.0306625366211,71.14013671875,71.1776885986328,71.1168899536133,71.041259765625,70.9811477661133,70.9404296875,70.8391571044922],[74.3910217285156,74.3521499633789,74.41064453125,74.4856948852539,74.51416015625,74.5179138183594,74.4789581298828,74.4567413330078,74.3910217285156],[78.595703125,78.572021484375,78.5769577026367,78.4076690673828,78.2281723022461,78.2152328491211,78.2399368286133,78.251953125,78.2386245727539,78.186279296875,78.1494598388672,78.1205062866211,78.0880813598633,77.9915084838867,77.9578628540039,77.8754425048828,77.86474609375,77.8985366821289,77.8344268798828,77.756591796875,77.6582565307617,77.630615234375,77.5577163696289,77.4977493286133,77.4623565673828,77.4014205932617,77.3803176879883,77.3850631713867,77.3607940673828,77.3113784790039,77.2757873535156,77.2666549682617,77.31591796875,77.3311004638672,77.360107421875,77.4293441772461,77.5001449584961,77.5393524169922,77.5535125732422,77.5496139526367,77.5711441040039,77.5701141357422,77.5288543701172,77.5011749267578,77.494140625,77.4409637451172,77.4596633911133,77.5653305053711,77.6194839477539,77.7109375,77.7717819213867,77.8121109008789,77.9160690307617,77.9235382080078,78.0057601928711,78.0591812133789,78.1658630371094,78.2521438598633,78.3255844116211,78.4193801879883,78.4120101928711,78.4778366088867,78.5147933959961,78.5567398071289,78.5975646972656,78.595703125],[78.64892578125,78.646484375,78.7202682495117,78.7843246459961,78.8105010986328,78.8114776611328,78.7239761352539,78.697509765625,78.64892578125],[78.6106948852539,78.5416946411133,78.548583984375,78.47509765625,78.4360809326172,78.4093322753906,78.3749008178711,78.3056106567383,78.2325668334961,78.224853515625,78.3290023803711,78.3882293701172,78.4387664794922,78.4412612915039,78.4632797241211,78.6447219848633,78.6865234375,78.7538604736328,78.8375015258789,78.9029312133789,78.8874969482422,78.8463821411133,78.7533721923828,78.7244567871094,78.68603515625,78.6405792236328,78.6106948852539],[78.9120559692383,78.90576171875,78.921630859375,78.9047393798828,78.8521041870117,78.8800811767578,78.88720703125,78.8287124633789,78.8521499633789,78.908447265625,78.9270477294922,78.96142578125,78.9673309326172,78.9708557128906,78.9120559692383],[79.90673828125,79.90478515625,79.9154281616211,79.9434585571289,79.9589309692383,79.9407730102539,79.8846664428711,79.8570251464844,79.800048828125,79.7042541503906,79.635009765625,79.5695343017578,79.5333557128906,79.4813461303711,79.4307632446289,79.3859329223633,79.4370574951172,79.6005859375,79.6106872558594,79.6051788330078,79.5715789794922,79.4881820678711,79.4605941772461,79.4266586303711,79.384765625,79.336669921875,79.3031768798828,79.281494140625,79.26171875,79.26025390625,79.2342758178711,79.1791534423828,79.1570281982422,79.1756820678711,79.1468276977539,79.0562057495117,79.0767059326172,79.125,79.1456604003906,79.1292495727539,79.1066436767578,79.0591354370117,78.9813995361328,78.9066925048828,78.8526382446289,78.7958526611328,78.7720184326172,78.7404251098633,78.6994171142578,78.67626953125,78.6723098754883,78.6432647705078,78.6227035522461,78.6095657348633,78.5978469848633,78.5621643066406,78.4797821044922,78.37939453125,78.3189468383789,78.2342300415039,78.1824722290039,78.1322784423828,78.0814895629883,78.0417022705078,78.0400848388672,78.0479965209961,78.0250473022461,77.9905776977539,77.9420394897461,77.7939453125,77.6822738647461,77.5785675048828,77.5226058959961,77.5070343017578,77.4967727661133,77.3993606567383,77.2252502441406,77.1568832397461,77.0489273071289,77.0106430053711,76.9691925048828,76.8949203491211,76.8116226196289,76.7793960571289,76.7203598022461,76.6589813232422,76.6061553955078,76.5792922973633,76.6093292236328,76.644775390625,76.7015151977539,76.738525390625,76.7607421875,76.8864288330078,77.0851058959961,77.162353515625,77.1990280151367,77.2344741821289,77.2821273803711,77.3355941772461,77.4032211303711,77.4452133178711,77.5082015991211,77.545166015625,77.5641098022461,77.5796356201172,77.5708465576172,77.5379409790039,77.5250549316406,77.6888198852539,77.782470703125,77.7986831665039,77.7977066040039,77.8419418334961,77.8980026245117,77.9115753173828,77.8802185058594,77.8471221923828,77.8470687866211,77.869140625,77.8569869995117,77.8090286254883,77.7786636352539,77.7664566040039,77.7713851928711,77.7962341308594,77.8538055419922,77.88330078125,77.91943359375,78.0281219482422,78.0576171875,78.0746078491211,78.0850143432617,78.0855407714844,78.0668411254883,78.0050735473633,78.0713882446289,78.1512222290039,78.220947265625,78.2327117919922,78.2275924682617,78.2647018432617,78.2990188598633,78.3270492553711,78.339111328125,78.3528823852539,78.3504409790039,78.369384765625,78.4171371459961,78.4004898071289,78.3972702026367,78.4071807861328,78.4488754272461,78.5035629272461,78.6128921508789,78.6636199951172,78.6563034057617,78.6385192871094,78.5381317138672,78.4930191040039,78.4713363647461,78.4732437133789,78.487548828125,78.5541000366211,78.5890655517578,78.6082992553711,78.6631317138672,78.72119140625,78.7711944580078,78.7811965942383,78.7323226928711,78.6642608642578,78.630126953125,78.6394500732422,78.6655731201172,78.70556640625,78.720947265625,78.720947265625,78.7049789428711,78.6753921508789,78.6305160522461,78.58056640625,78.5409164428711,78.4924774169922,78.4619598388672,78.4146041870117,78.3923797607422,78.3599090576172,78.3098602294922,78.2709045410156,78.2667465209961,78.2451629638672,78.2375030517578,78.3010787963867,78.3312530517578,78.3514633178711,78.384765625,78.4829559326172,78.5946807861328,78.6055221557617,78.6423797607422,78.6742172241211,78.7164077758789,78.7662582397461,78.8116683959961,78.8318862915039,78.8829574584961,78.9503860473633,78.9729995727539,78.9829635620117,78.9044876098633,78.9142608642578,78.9532241821289,78.9663543701172,78.975341796875,78.97509765625,78.983154296875,79.0252990722656,79.0772476196289,79.1118698120117,79.15234375,79.2130813598633,79.2675247192383,79.2926788330078,79.2911605834961,79.2835006713867,79.2052688598633,79.1512680053711,79.109130859375,79.129638671875,79.2329635620117,79.3048858642578,79.3501968383789,79.4154281616211,79.4628448486328,79.5201644897461,79.5555191040039,79.5816421508789,79.6409225463867,79.6903305053711,79.7335891723633,79.7582473754883,79.7886734008789,79.7987823486328,79.7965774536133,79.7603073120117,79.7169876098633,79.720458984375,79.7848587036133,79.7994155883789,79.8206024169922,79.737548828125,79.7190933227539,79.713134765625,79.75,79.7978973388672,79.8159637451172,79.7732391357422,79.7757797241211,79.8317337036133,79.8609924316406,79.8169403076172,79.7934036254883,79.76171875,79.752197265625,79.7152786254883,79.6851501464844,79.5694808959961,79.5880813598633,79.5748062133789,79.4807586669922,79.4708938598633,79.4572296142578,79.4291534423828,79.3756866455078,79.3396453857422,79.3441390991211,79.3831100463867,79.4292907714844,79.48193359375,79.5386734008789,79.5856475830078,79.6187057495117,79.7259750366211,79.7987289428711,79.7664031982422,79.6753463745117,79.5454559326172,79.4067916870117,79.23486328125,79.1742706298828,79.159912109375,78.9810562133789,78.9761276245117,79.1121063232422,79.3423767089844,79.5192413330078,79.5868682861328,79.6818313598633,79.7090301513672,79.7335891723633,79.8351135253906,79.8844223022461,79.9539566040039,80.00732421875,80.0494613647461,80.0525894165039,80.0205078125,79.90673828125],[80.119140625,80.0814514160156,80.10791015625,80.2179718017578,80.2287139892578,80.2423324584961,80.2174301147461,80.1981430053711,80.119140625],[80.3009262084961,80.24560546875,80.2881851196289,80.331787109375,80.3583526611328,80.3483352661133,80.3009262084961],[80.2499465942383,80.2388153076172,80.242919921875,80.2184524536133,80.1591796875,80.1387634277344,80.1324691772461,80.0597152709961,80.0492172241211,80.0896530151367,80.1902770996094,80.308349609375,80.3655319213867,80.4022445678711,80.4164581298828,80.4126434326172,80.4330062866211,80.468994140625,80.4739685058594,80.4466781616211,80.4252471923828,80.380859375,80.317626953125,80.1869583129883,80.1788635253906,80.20654296875,80.244384765625,80.3045883178711,80.295166015625,80.3031311035156,80.3292922973633,80.3603973388672,80.3551712036133,80.295166015625,80.2858428955078,80.3013153076172,80.3006820678711,80.2766647338867,80.2331085205078,80.2097702026367,80.1880416870117,80.1751480102539,80.1754837036133,80.1600112915039,80.12548828125,80.0592269897461,79.9065933227539,79.8653793334961,79.6774444580078,79.6170425415039,79.5613784790039,79.4397430419922,79.4030227661133,79.3450698852539,79.3388671875,79.3672332763672,79.3646011352539,79.3016128540039,79.2634811401367,79.2154769897461,79.1942901611328,79.2056121826172,79.2306671142578,79.2643508911133,79.3290557861328,79.411865234375,79.3810501098633,79.3978576660156,79.4095230102539,79.4415054321289,79.46337890625,79.4896011352539,79.5337905883789,79.5911636352539,79.6179733276367,79.6336364746094,79.6402359008789,79.6322708129883,79.6327667236328,79.6905288696289,79.7071838378906,79.7485885620117,79.774658203125,79.778564453125,79.7441864013672,79.7286148071289,79.7265625,79.736328125,79.7607421875,79.8245086669922,79.8597183227539,79.8873519897461,79.92919921875,79.9666976928711,79.9962387084961,80.03662109375,80.0595703125,80.0934066772461,80.1431198120117,80.171142578125,80.1935043334961,80.1748046875,80.138671875,80.1164016723633,80.1632308959961,80.1854019165039,80.2632293701172,80.3018569946289,80.33154296875,80.3359909057617,80.3230895996094,80.25,80.2271957397461,80.2947235107422,80.3268051147461,80.3533706665039,80.40234375,80.4625549316406,80.4778289794922,80.47119140625,80.4299774169922,80.4009323120117,80.3716278076172,80.2986831665039,80.2499465942383],[27.87060546875,27.8433113098145,27.7986812591553,27.7492179870605,27.6424312591553,27.5673828125,27.4088859558105,27.1339359283447,27.0860824584961,26.928466796875,26.8073253631592,26.7248039245605,26.58642578125,26.4300289154053,26.39501953125,26.3823719024658,26.4369125366211,26.4292984008789,26.3991222381592,26.4049797058105,26.4229488372803,26.3603019714355,26.3942375183105,26.4332027435303,26.5416011810303,26.555419921875,26.4419441223145,26.43505859375,26.4959964752197,26.5562973022461,26.5744132995605,26.5979976654053,26.6117172241211,26.6493663787842,26.6001949310303,26.6041507720947,26.6397457122803,26.712646484375,26.7816410064697,26.8290042877197,26.83984375,26.7972164154053,26.7410144805908,26.7503414154053,26.7665519714355,26.7815418243408,26.8466300964355,26.8609867095947,26.8629398345947,26.8785152435303,26.9269046783447,27.041015625,27.0916996002197,27.2036609649658,27.2498531341553,27.2986812591553,27.3481941223145,27.4278316497803,27.4913578033447,27.461669921875,27.4351081848145,27.3778305053711,27.3959465026855,27.4563484191895,27.46533203125,27.44482421875,27.4102535247803,27.3709964752197,27.4022960662842,27.4445323944092,27.4676761627197,27.5189952850342,27.5966796875,27.6734371185303,27.6870613098145,27.6718273162842,27.7565441131592,27.8649425506592,27.9005870819092,27.9137687683105,27.8992671966553,27.8744621276855,27.8670902252197,27.913818359375,27.98046875,28.0622081756592,28.1763668060303,28.2408676147461,28.2894039154053,28.3350086212158,28.4095706939697,28.4685554504395,28.5396957397461,28.5539054870605,28.5962390899658,28.6496067047119,28.6651878356934,28.6357917785645,28.6048831939697,28.6120128631592,28.6664066314697,28.7233390808105,28.7760753631592,28.8301773071289,28.8703117370605,28.9941902160645,29.1003894805908,29.1243152618408,29.1946296691895,29.3180160522461,29.4233417510986,29.5720691680908,29.7302742004395,29.8998050689697,29.9558601379395,29.9943351745605,30.1193351745605,30.1397457122803,30.1719226837158,30.1800308227539,30.1645030975342,30.0989742279053,30.0368156433105,30.0398921966553,30.0933113098145,30.3375968933105,30.3875007629395,30.3624038696289,30.3267574310303,30.2450675964355,30.1589832305908,30.1151847839355,30.0638656616211,29.9415035247803,29.8311996459961,29.6833972930908,29.6180667877197,29.6126480102539,29.5545902252197,29.4391593933105,29.3063468933105,29.1835956573486,29.1875972747803,29.2274417877197,29.2794914245605,29.2538566589355,29.2199687957764,29.1562976837158,29.036376953125,28.9117679595947,28.8681163787842,28.8039054870605,28.7529315948486,28.6595687866211,28.6215343475342,28.5955562591553,28.5792484283447,28.5602054595947,28.5536136627197,28.6096687316895,28.6026363372803,28.5922355651855,28.5718746185303,28.4842777252197,28.3722667694092,28.3159656524658,28.2926254272461,28.2760257720947,28.2774410247803,28.2206554412842,28.1353511810303,27.9896965026855,27.92822265625,27.910400390625,27.9347171783447,27.9945812225342,28.0835933685303,28.1143550872803,28.0916996002197,28.0220699310303,27.9595203399658,27.9286613464355,27.9395503997803,27.9635257720947,28.085205078125,28.10302734375,28.0949211120605,28.0706558227539,28.0220699310303,27.9991703033447,27.9684562683105,27.9286613464355,27.8383293151855,27.8219242095947,27.8238277435303,27.8218269348145,27.815185546875,27.8213882446289,27.8860855102539,27.8908214569092,27.8833980560303,27.87060546875],[-0.51660168170929,-0.550781428813934,-0.546484291553497,-0.523730397224426,-0.499121308326721,-0.489355266094208,-0.496972739696503,-0.51660168170929],[-52.4972648620605,-52.5482406616211,-52.5703125,-52.5518531799316,-52.5285148620605,-52.4954109191895,-52.4988288879395,-52.4851531982422,-52.4972648620605],[-50.7615203857422,-50.845703125,-50.8460922241211,-50.8226547241211,-50.7867202758789,-50.7779273986816,-50.8195304870605,-50.8214836120605,-50.8077163696289,-50.7630844116211,-50.6790046691895,-50.5730476379395,-50.5389633178711,-50.5309562683105,-50.5439453125,-50.55859375,-50.5772476196289,-50.6120109558105,-50.6524391174316,-50.6943359375,-50.7146492004395,-50.7508811950684,-50.7615203857422],[-46.8622093200684,-46.9021492004395,-46.8877944946289,-46.9326171875,-46.9561538696289,-46.98828125,-46.97900390625,-47.0270500183105,-47.0700187683105,-47.1015625,-47.1174812316895,-47.1798820495605,-47.17041015625,-47.1760749816895,-47.2429656982422,-47.263671875,-47.2586898803711,-47.1990203857422,-47.1422882080078,-47.0877914428711,-47.0442390441895,-47.0135765075684,-46.9568367004395,-46.9065437316895,-46.7976570129395,-46.6998023986816,-46.6944351196289,-46.8622093200684],[-45.6558609008789,-45.7049789428711,-45.7296867370605,-45.7298812866211,-45.7244148254395,-45.7017593383789,-45.7082023620605,-45.6998062133789,-45.6444320678711,-45.6150398254395,-45.638671875,-45.6558609008789],[-45.1796875,-45.2998046875,-45.27685546875,-45.2405281066895,-45.1803703308105,-45.1796875],[-44.3216781616211,-44.33056640625,-44.2735366821289,-44.2367172241211,-44.2245101928711,-44.2684555053711,-44.3216781616211],[-43.7403335571289,-43.7663078308105,-43.7648429870605,-43.8667984008789,-43.7906227111816,-43.7609367370605,-43.76806640625,-43.7847671508789,-43.8048820495605,-43.8161125183105,-43.8601570129395,-43.9546890258789,-43.9514656066895,-44.0252914428711,-44.0484352111816,-44.0768585205078,-44.1166000366211,-44.1149406433105,-44.1072273254395,-44.0361328125,-44.0062484741211,-43.9541015625,-43.9009742736816,-43.8519515991211,-43.8202171325684,-43.8345680236816,-43.8239250183105,-43.7579116821289,-43.76513671875,-43.7175788879395,-43.7403335571289],[-40.8636741638184,-40.9217758178711,-40.8814468383789,-40.7936515808105,-40.7493133544922,-40.7462882995605,-40.7129898071289,-40.7868156433105,-40.8636741638184],[-41.279296875,-41.2841796875,-41.2109375,-41.1513671875,-41.10205078125,-40.9889640808105,-40.9723625183105,-40.9507827758789,-40.9241218566895,-40.9177742004395,-40.98486328125,-41.0072250366211,-41.0314445495605,-41.0701179504395,-41.1244163513184,-41.1920890808105,-41.27197265625,-41.2393531799316,-41.1873054504395,-41.1580047607422,-41.0999031066895,-41.072265625,-41.0281257629395,-40.9932632446289,-41.0061492919922,-41.0046882629395,-40.9909210205078,-40.9854469299316,-41.0244140625,-41.01953125,-41.0687484741211,-41.1255874633789,-41.16015625,-41.2173843383789,-41.2418937683105,-41.2482414245605,-41.1715812683105,-41.1037101745605,-41.1883811950684,-41.2642555236816,-41.3122062683105,-41.3270492553711,-41.3659172058105,-41.4294929504395,-41.4716796875,-41.5051727294922,-41.5618171691895,-41.6707992553711,-41.6572265625,-41.677734375,-41.7406234741211,-41.8130874633789,-41.8501930236816,-42.0030250549316,-42.08056640625,-42.1301727294922,-42.2116203308105,-42.2709007263184,-42.4739227294922,-42.5179672241211,-42.8408203125,-42.9765625,-43.0227546691895,-43.0602531433105,-43.1242179870605,-43.19775390625,-43.2587890625,-43.2724609375,-43.2995147705078,-43.3146476745605,-43.3547859191895,-43.3997077941895,-43.4279289245605,-43.43603515625,-43.4647445678711,-43.4443359375,-43.4678688049316,-43.5172882080078,-43.5619163513184,-43.6209945678711,-43.6585922241211,-43.6761703491211,-43.7035140991211,-43.7978515625,-43.8441429138184,-43.8746070861816,-43.8854484558105,-43.8914070129395,-43.8701171875,-43.8130836486816,-43.8313484191895,-43.8436546325684,-43.8333969116211,-43.7735366821289,-43.7394523620605,-43.7266578674316,-43.7464828491211,-43.77783203125,-43.82958984375,-43.859375,-43.8678703308105,-43.8250007629395,-43.7635726928711,-43.7017555236816,-43.7400398254395,-43.8337860107422,-43.89599609375,-43.9456062316895,-43.9842758178711,-44.0069313049316,-44.0422859191895,-44.0974617004395,-44.1171875,-44.1183586120605,-44.1358375549316,-44.2086906433105,-44.2549819946289,-44.26416015625,-44.2787094116211,-44.3018569946289,-44.5211906433105,-44.6122055053711,-44.7678718566895,-44.9123039245605,-44.9114265441895,-44.93701171875,-44.9777336120605,-45.0392570495605,-45.1514625549316,-45.2164039611816,-45.3739280700684,-45.5191383361816,-45.6842765808105,-45.7139625549316,-45.7560539245605,-45.79248046875,-45.8438453674316,-45.8708992004395,-45.8780288696289,-45.8957023620605,-45.9410171508789,-45.9917945861816,-46.0826187133789,-46.1608390808105,-46.3343734741211,-46.4797859191895,-46.5213890075684,-46.5516586303711,-46.6205062866211,-46.6306648254395,-46.6129875183105,-46.5782241821289,-46.5663108825684,-46.5875968933105,-46.6110343933105,-46.587890625,-46.6053733825684,-46.58837890625,-46.5457038879395,-46.4890594482422,-46.4471664428711,-46.4187507629395,-46.3857421875,-46.3622055053711,-46.3529281616211,-46.3677749633789,-46.3662109375,-46.2271461486816,-46.1929702758789,-46.1485366821289,-46.1546859741211,-46.2289047241211,-46.2415046691895,-46.2494125366211,-46.2254867553711,-46.1978530883789,-46.1336898803711,-45.9572257995605,-45.9808616638184,-46.052734375,-46.0416984558105,-45.9632797241211,-45.9283180236816,-45.8893585205078,-45.9553718566895,-45.9638671875,-45.9027328491211,-45.8318367004395,-45.8117179870605,-45.7745094299316,-45.7501945495605,-45.7121086120605,-45.6990242004395,-45.6456031799316,-45.6028327941895,-45.53173828125,-45.5499038696289,-45.5435562133789,-45.4684562683105,-45.40966796875,-45.4079093933105,-45.3675765991211,-45.3112297058105,-45.3074226379395,-45.3832015991211,-45.4109344482422,-45.3539085388184,-45.3179702758789,-45.3018531799316,-45.2903327941895,-45.2802734375,-45.2658195495605,-45.2224617004395,-45.1766624450684,-45.1236305236816,-45.05078125,-45.0941429138184,-45.0822257995605,-45.0481452941895,-44.9970703125,-44.9634780883789,-44.8279266357422,-44.9582977294922,-44.9150390625,-44.8739280700684,-44.8382835388184,-44.8023414611816,-44.7713890075684,-44.7408180236816,-44.6413078308105,-44.5950202941895,-44.6247062683105,-44.6647453308105,-44.625,-44.5920867919922,-44.5006828308105,-44.3587913513184,-44.2236328125,-44.08203125,-44.0305633544922,-43.97216796875,-43.9964866638184,-43.9919929504395,-43.8899421691895,-43.8634796142578,-43.89990234375,-43.9130859375,-43.81982421875,-43.7770500183105,-43.7015609741211,-43.6236343383789,-43.5912094116211,-43.5384788513184,-43.5370140075684,-43.4971694946289,-43.458984375,-43.4616203308105,-43.4465827941895,-43.4259757995605,-43.3494148254395,-43.2650375366211,-43.24755859375,-43.2220687866211,-43.1638679504395,-43.1446266174316,-43.1536140441895,-43.1822242736816,-43.1346702575684,-43.1076164245605,-43.0662117004395,-43.0376930236816,-43.0584945678711,-43.091796875,-43.0407257080078,-43.0089836120605,-42.9724578857422,-42.9612312316895,-43.02978515625,-42.9754905700684,-42.9273414611816,-42.8486328125,-42.7183609008789,-42.763671875,-42.8187484741211,-42.8850555419922,-42.8621063232422,-42.8018569946289,-42.6960945129395,-42.50048828125,-42.4786148071289,-42.46533203125,-42.4601593017578,-42.4305648803711,-42.4019546508789,-42.3025360107422,-42.1890640258789,-42.0799789428711,-41.9730453491211,-41.7947273254395,-41.7575187683105,-41.7447280883789,-41.7196273803711,-41.6551742553711,-41.5386734008789,-41.4447288513184,-41.2015609741211,-40.947265625,-40.7586936950684,-40.6221694946289,-40.5182609558105,-40.4966812133789,-40.4900398254395,-40.5187530517578,-40.5437507629395,-40.6053733825684,-40.6677742004395,-40.7236328125,-40.7734375,-40.8203125,-40.8482437133789,-40.95361328125,-41.07861328125,-41.1858406066895,-41.279296875],[-36.279296875,-36.3338890075684,-36.314453125,-36.2732429504395,-36.2306632995605,-36.2177734375,-36.134765625,-36.0948257446289,-36.0777359008789,-36.0708999633789,-36.1146507263184,-36.1769523620605,-36.279296875],[-34.9347648620605,-34.9805679321289,-34.9479522705078,-34.896484375,-34.8443374633789,-34.9285163879395,-34.9469718933105,-35.0056648254395,-35.0545883178711,-35.0685539245605,-35.0412101745605,-35.0262680053711,-35.05712890625,-35.1428718566895,-35.17236328125,-35.2164077758789,-35.2628898620605,-35.2999992370605,-35.30859375,-35.2535171508789,-35.2466773986816,-35.3245124816895,-35.3685531616211,-35.3670883178711,-35.4107437133789,-35.4541015625,-35.58203125,-35.626953125,-35.6673851013184,-35.7855453491211,-35.7937507629395,-35.7737312316895,-35.79736328125,-35.8840827941895,-36.0066413879395,-36.08056640625,-36.3094749450684,-36.3909149169922,-36.4446296691895,-36.4908218383789,-36.6121101379395,-36.6498031616211,-36.7740211486816,-36.7957992553711,-36.8412094116211,-36.8532218933105,-36.87255859375,-36.9093742370605,-36.8650360107422,-36.8529319763184,-36.9122085571289,-36.9712867736816,-36.9932594299316,-37.0409202575684,-37.1561546325684,-37.2069320678711,-37.2167015075684,-37.2013664245605,-37.1593742370605,-37.0464859008789,-36.86572265625,-36.8069343566895,-36.748046875,-36.6895484924316,-36.63427734375,-36.5918960571289,-36.5563468933105,-36.5007820129395,-36.4756813049316,-36.5226554870605,-36.5792961120605,-36.7469711303711,-36.7351531982422,-36.8045921325684,-36.8750991821289,-36.9577178955078,-37.20458984375,-37.43701171875,-37.5382843017578,-37.5867195129395,-37.5617179870605,-37.5762672424316,-37.6006813049316,-37.6451187133789,-37.6669921875,-37.6638641357422,-37.6800765991211,-37.8309555053711,-37.8896484375,-37.9857406616211,-37.9934577941895,-37.9908218383789,-37.9574203491211,-37.8974609375,-37.8078117370605,-37.70556640625,-37.6559562683105,-37.6168937683105,-37.5806655883789,-37.5548820495605,-37.56689453125,-37.6184539794922,-37.6597633361816,-37.6920890808105,-37.7576179504395,-37.8543930053711,-37.9602546691895,-38.2008781433105,-38.4440460205078,-38.5511741638184,-38.6336898803711,-38.6939468383789,-38.7222633361816,-38.8602561950684,-39.0217742919922,-39.0624008178711,-39.0945320129395,-39.1424789428711,-39.2395477294922,-39.2254905700684,-39.1447257995605,-39.1109352111816,-39.0857391357422,-39.0738296508789,-39.0811500549316,-39.1158218383789,-39.1861343383789,-39.2217788696289,-39.2668952941895,-39.3675765991211,-39.49072265625,-39.5552749633789,-39.6051750183105,-39.6731452941895,-39.9107437133789,-40.1578140258789,-40.2284164428711,-40.29345703125,-40.4419898986816,-40.5700187683105,-40.6676788330078,-40.7689437866211,-40.8768539428711,-41.0290985107422,-41.1296882629395,-41.2132797241211,-41.3201179504395,-41.4117202758789,-41.5382804870605,-41.580078125,-41.6106414794922,-41.5744171142578,-41.5349617004395,-41.4490242004395,-41.4173812866211,-41.3912086486816,-41.4329109191895,-41.4240264892578,-41.404296875,-41.2782211303711,-41.24267578125,-41.2230453491211,-41.2307624816895,-41.2628898620605,-41.2907218933105,-41.3252944946289,-41.3262672424316,-41.3126945495605,-41.2894515991211,-41.2512702941895,-41.2176742553711,-41.0587921142578,-40.84765625,-40.62158203125,-40.50537109375,-40.2893562316895,-40.1994132995605,-40.1149406433105,-39.9521484375,-39.8601570129395,-39.84716796875,-39.81298828125,-39.7351531982422,-39.6433601379395,-39.5681648254395,-39.5090827941895,-39.42578125,-39.3761749267578,-39.3187484741211,-39.2653350830078,-39.2112312316895,-39.1695289611816,-39.1393547058105,-39.03125,-38.9710922241211,-38.97216796875,-38.9626007080078,-38.92578125,-38.8416023254395,-38.7850570678711,-38.6052742004395,-38.4283218383789,-38.2255859375,-38.0998077392578,-38.0226554870605,-37.8955078125,-37.8489265441895,-37.82080078125,-37.8044929504395,-37.6851539611816,-37.5046882629395,-37.44873046875,-37.3934555053711,-37.3390617370605,-37.3252944946289,-37.2731437683105,-37.0977516174316,-37.0699234008789,-37.0887718200684,-37.2152366638184,-37.1501007080078,-37.1100578308105,-37.0892601013184,-37.0847663879395,-36.9437484741211,-36.9494132995605,-36.9718742370605,-36.9857406616211,-36.9733390808105,-36.94189453125,-36.8825187683105,-36.7682609558105,-36.7259750366211,-36.4922866821289,-36.4849624633789,-36.6019515991211,-36.5645484924316,-36.5107421875,-36.4508781433105,-36.4055671691895,-36.3759765625,-36.3228492736816,-36.2741203308105,-36.2400360107422,-36.1705055236816,-36.1630859375,-36.1956024169922,-36.2437515258789,-36.2491226196289,-36.1224594116211,-36.0206031799316,-35.90869140625,-35.9541969299316,-36.0181617736816,-36.1462898254395,-36.2894554138184,-36.32763671875,-36.376953125,-36.3910140991211,-36.3597679138184,-36.2372055053711,-36.1758766174316,-35.5425758361816,-35.458984375,-35.3885726928711,-35.3572273254395,-35.3191413879395,-35.3125953674316,-35.3298835754395,-35.3623046875,-35.3992156982422,-35.4811515808105,-35.5000953674316,-35.443359375,-35.4083023071289,-35.3663101196289,-35.3396492004395,-35.3312492370605,-35.2477531433105,-35.2052726745605,-35.1237297058105,-35.0162124633789,-34.9033203125,-34.7999038696289,-34.63232421875,-34.4551773071289,-34.4329109191895,-34.4291000366211,-34.53515625,-34.5964851379395,-34.6482429504395,-34.8069343566895,-34.8527336120605,-34.8990211486816,-34.9347648620605],[-9.35546875,-9.35830116271973,-9.35244178771973,-9.3447265625,-9.33330059051514,-9.33261775970459,-9.33876991271973,-9.34658241271973,-9.35546875],[-8.58076190948486,-8.58291053771973,-8.57158184051514,-8.55918025970459,-8.54794883728027,-8.54648399353027,-8.55615234375,-8.56748008728027,-8.58076190948486],[20.21875,20.20361328125,20.210693359375,20.3373527526855,20.49658203125,20.6805667877197,20.5161628723145,20.4239253997803,20.266845703125,20.21875],[24.9791984558105,24.7163581848145,24.4703121185303,24.3345699310303,24.1501941680908,23.980712890625,23.9227542877197,23.8036613464355,23.7591304779053,23.716552734375,23.6238288879395,23.6181640625,23.6456546783447,23.6434555053711,23.5171871185303,23.3974609375,23.3341808319092,23.2347164154053,23.1305675506592,22.9718742370605,22.7933597564697,22.6608390808105,22.5785160064697,22.546142578125,22.5089836120605,22.4205570220947,22.30517578125,22.2199211120605,22.0538082122803,21.9513683319092,21.7823238372803,21.4988288879395,21.4353523254395,21.2890625,21.11279296875,20.80712890625,20.50390625,20.4068832397461,20.3869152069092,20.3954582214355,20.4239749908447,20.5068359375,20.5992183685303,20.5895004272461,20.5703620910645,20.3436031341553,20.244140625,20.1177253723145,19.95458984375,19.8044929504395,19.6784191131592,19.60693359375,19.4322261810303,19.2533187866211,19.1459465026855,19.01708984375,18.9773445129395,18.9578609466553,18.9437999725342,18.902587890625,18.8278312683105,18.7535152435303,18.5873546600342,18.1659679412842,17.9879875183105,17.9507827758789,17.9352035522461,17.8860855102539,17.84326171875,17.5856456756592,17.5047359466553,17.4462394714355,17.381591796875,17.3208980560303,17.1576175689697,17.0389156341553,16.9646472930908,17.0088863372803,17.03125,17.0336437225342,17.0055179595947,16.9178218841553,16.855712890625,16.7599601745605,16.7233390808105,16.6483879089355,16.7802238464355,16.9120597839355,17.0438480377197,17.1756839752197,17.2679195404053,17.3003902435303,17.39794921875,17.4978523254395,17.5977535247803,17.6976070404053,17.7975101470947,17.8974132537842,17.997314453125,18.09716796875,18.1970710754395,18.2969245910645,18.3968257904053,18.4966793060303,18.5965824127197,18.6964855194092,18.79638671875,18.8962879180908,18.9961433410645,19.0431632995605,19.1004886627197,19.1578140258789,19.2151374816895,19.2724609375,19.3297863006592,19.3871574401855,19.4444828033447,19.5018062591553,19.5591316223145,19.6165027618408,19.673828125,19.7311534881592,19.7884769439697,19.8458003997803,19.9031257629395,19.9604988098145,19.9959468841553,20.1292476654053,20.2399425506592,20.3506336212158,20.4613285064697,20.572021484375,20.6827144622803,20.7934074401855,20.9041004180908,21.0147953033447,21.1255378723145,21.2362289428711,21.346923828125,21.4576168060303,21.5683097839355,21.6790027618408,21.7896976470947,21.900390625,22.0018558502197,22.0995101928711,22.2306632995605,22.3678226470947,22.4969234466553,22.5909175872803,22.7041015625,22.8500003814697,22.9229507446289,23.0347652435303,23.18994140625,23.387451171875,23.5187492370605,23.6329097747803,23.724609375,23.8190441131592,23.885498046875,23.90966796875,23.9411125183105,23.9913578033447,24.0241222381592,24.01708984375,24.0414066314697,24.0633773803711,24.0929679870605,24.1426277160645,24.2151355743408,24.22265625,24.24267578125,24.3498039245605,24.383544921875,24.4235363006592,24.4906253814697,24.5773448944092,24.63623046875,24.68359375,24.7812995910645,24.8681163787842,24.9112796783447,24.951416015625,24.9717769622803,24.9702625274658,24.9532222747803,24.90771484375,24.8764171600342,24.8720703125,24.8589344024658,24.7982406616211,24.73876953125,24.7486820220947,24.7955074310303,24.8333015441895,24.86669921875,24.9312992095947,24.9732913970947,24.9791984558105],[25.2132816314697,25.2666988372803,25.3038082122803,25.3008785247803,25.2786140441895,25.2355461120605,25.2088375091553,25.2132816314697],[25.6506824493408,25.6277332305908,25.6253910064697,25.6449222564697,25.6905269622803,25.74609375,25.8484859466553,25.9451656341553,26.0474605560303,26.0681629180908,26.0626468658447,26.20703125,26.2292003631592,26.2197742462158,26.2352066040039,26.3136234283447,26.3563480377197,26.3511714935303,26.3271980285645,26.2081546783447,26.1087398529053,25.8045902252197,25.751953125,25.7093257904053,25.6506824493408],[35.6617202758789,35.5718269348145,35.4357414245605,35.3466300964355,35.2464370727539,35.1963386535645,35.1099128723145,35.0623512268066,34.9919929504395,34.938720703125,34.9001960754395,34.8778305053711,34.8475570678711,34.7869148254395,34.73583984375,34.7408676147461,34.756103515625,34.667724609375,34.669921875,34.612548828125,34.5602569580078,34.5030746459961,34.5027351379395,34.5367164611816,34.6013679504395,34.6390151977539,34.6368179321289,34.6458511352539,34.677734375,34.7157707214355,34.7320327758789,34.765380859375,34.7208976745605,34.6806640625,34.6534652709961,34.529052734375,34.4853057861328,34.42236328125,34.3782234191895,34.3253402709961,34.287841796875,34.2366218566895,34.1913070678711,34.144775390625,34.1080093383789,34.07568359375,34.0430679321289,34.018798828125,34.0037117004395,34.00341796875,33.9901885986328,33.9460945129395,33.8865737915039,33.8386726379395,33.7881851196289,33.7212905883789,33.6678237915039,33.6324234008789,33.5917015075684,33.5450668334961,33.5069808959961,33.4553680419922,33.3841323852539,33.30126953125,33.2421875,33.22119140625,33.189453125,33.1434059143066,33.075439453125,33.0203132629395,33.005126953125,32.9917984008789,32.927978515625,32.86083984375,32.8104476928711,32.7686996459961,32.77099609375,32.7532234191895,32.7709007263184,32.7576675415039,32.6077156066895,32.5189437866211,32.4937973022461,32.4578132629395,32.4622077941895,32.4203605651855,32.3721199035645,32.3188972473145,32.2792015075684,32.2152824401855,32.1403312683105,32.1047859191895,32.08935546875,31.9488277435303,31.8897476196289,31.8185558319092,31.76513671875,31.7129402160645,31.52392578125,31.4653797149658,31.2613754272461,31.1855945587158,31.1326656341553,31.1128444671631,31.0687503814697,31.03466796875,30.9596672058105,30.8934059143066,30.8935527801514,30.7689952850342,30.5196800231934,30.4353504180908,30.3940410614014,30.3521480560303,30.2816410064697,30.2220706939697,30.1620121002197,30.0933589935303,30.033203125,29.9716796875,29.9343757629395,29.7729988098145,29.6106929779053,29.5506362915039,29.3639163970947,29.0888195037842,29.0287609100342,28.8961429595947,28.7519035339355,28.697265625,28.5658206939697,28.4217758178711,28.3463382720947,28.1772956848145,28.0474605560303,27.9624996185303,27.9150867462158,27.869873046875,27.8552742004395,27.8316402435303,27.7144546508789,27.7096195220947,27.72900390625,27.7689952850342,27.8353519439697,27.9374504089355,27.9837894439697,28.0231437683105,28.0250492095947,27.9816398620605,27.9341316223145,27.8948726654053,27.8490238189697,27.6947269439697,27.4736328125,27.3126945495605,27.2645015716553,27.2280769348145,27.1746101379395,27.1229496002197,26.9541492462158,26.804443359375,26.77099609375,26.74267578125,26.6991214752197,26.6270503997803,26.5861320495605,26.5787582397461,26.5480480194092,26.5064449310303,26.471435546875,26.3475589752197,26.2147960662842,26.0719718933105,25.9900398254395,25.9100570678711,25.70654296875,25.6857433319092,25.6813468933105,25.685302734375,25.7059574127197,25.69189453125,25.6669425964355,25.6257820129395,25.4228992462158,25.3310546875,25.2058601379395,25.06298828125,24.8916015625,24.7576675415039,24.687744140625,24.6539058685303,24.6187515258789,24.5718746185303,24.5224609375,24.4874038696289,24.4443378448486,24.4299812316895,24.4000968933105,24.3610343933105,24.3623542785645,24.34375,24.261962890625,24.2454109191895,24.2379875183105,24.24609375,24.279052734375,24.3311042785645,24.3857898712158,24.4183101654053,24.4121589660645,24.3562984466553,24.2875003814697,24.2405776977539,24.1915531158447,24.17138671875,24.1652355194092,24.172607421875,24.2251949310303,24.2730960845947,24.275390625,24.2682609558105,24.2686519622803,24.2863292694092,24.2730960845947,24.2924308776855,24.2665042877197,24.2640132904053,24.30908203125,24.313720703125,24.3072261810303,24.2919921875,24.265625,23.9646968841553,23.9666004180908,23.9672355651855,23.9508800506592,23.927978515625,23.9005374908447,23.8573246002197,23.7972183227539,23.7533702850342,23.818359375,23.8482418060303,23.8260726928711,23.82861328125,23.9026851654053,23.8280754089355,23.8109874725342,23.8672847747803,23.9198722839355,23.8818359375,23.9400386810303,24.0182609558105,24.0398921966553,24.0648422241211,24.0916023254395,24.1748046875,24.2628917694092,24.3677730560303,24.756103515625,24.7919445037842,24.8609371185303,24.9288578033447,25.111328125,25.226318359375,25.3785152435303,25.484375,25.5753402709961,25.601806640625,25.5898933410645,25.5539054870605,25.4932613372803,25.5073719024658,25.4850578308105,25.4453125,25.4468250274658,25.4657707214355,25.46435546875,25.4196281433105,25.3552722930908,25.3743171691895,25.3110847473145,25.3073253631592,25.18408203125,25.206298828125,25.2366695404053,25.3334484100342,25.3739261627197,25.4029293060303,25.3511734008789,25.342529296875,25.3858890533447,25.3531742095947,25.2975101470947,25.2108402252197,25.2275867462158,25.2548828125,25.2246589660645,25.2647953033447,25.2547359466553,25.197265625,25.1525382995605,25.1349124908447,25.1973628997803,25.224853515625,25.2066402435303,25.1553211212158,25.1312980651855,25.13818359375,25.1863269805908,25.2023448944092,25.2861328125,25.5846176147461,25.6923828125,25.7512702941895,25.7689933776855,25.8210926055908,25.8433589935303,25.995849609375,26.165283203125,26.2259292602539,26.242431640625,26.3182621002197,26.3689937591553,26.3570308685303,26.3692378997803,26.427490234375,26.4908695220947,26.5426273345947,26.56103515625,26.5936527252197,26.63916015625,26.6438961029053,26.63232421875,26.6497554779053,26.6655750274658,26.8375968933105,26.86474609375,26.8792476654053,26.9981441497803,27.0776863098145,27.1245613098145,27.1514644622803,27.2079105377197,27.2439460754395,27.2524909973145,27.2184085845947,27.2294445037842,27.2501964569092,27.265625,27.3001937866211,27.3567390441895,27.4445323944092,27.4970207214355,27.800537109375,28.0020503997803,28.2020511627197,28.2435550689697,28.2528820037842,28.2527847290039,28.2351570129395,28.3638668060303,28.4147472381592,28.4788093566895,28.4910163879395,28.5465335845947,28.6676769256592,28.7916011810303,28.8708992004395,29.0060539245605,29.2649898529053,29.331787109375,29.3726062774658,29.5427227020264,29.6634254455566,29.8586902618408,29.7494144439697,29.6656742095947,29.5304183959961,29.4253902435303,29.4083499908447,29.4979991912842,29.4300804138184,29.3919429779053,29.4142570495605,29.4603519439697,29.5069332122803,29.5443344116211,29.5645008087158,29.5671367645264,29.5641593933105,29.5527839660645,29.5594749450684,29.5776348114014,29.6515617370605,29.7013206481934,29.7789077758789,29.8355941772461,29.86572265625,29.9200191497803,29.9685554504395,30.0435066223145,30.109619140625,30.1934566497803,30.3211421966553,30.5029792785645,30.6079120635986,30.8027839660645,30.9122066497803,30.9645500183105,30.9965839385986,31.0199699401855,31.0460433959961,31.1945323944092,31.263671875,31.3056144714355,31.3002433776855,31.2429218292236,31.2178230285645,31.234619140625,31.2776870727539,31.31298828125,31.3439464569092,31.3792476654053,31.4099597930908,31.4533214569092,31.5064945220947,31.5387706756592,31.5481929779053,31.6779804229736,31.7632827758789,31.8029804229736,31.8073749542236,31.7676734924316,31.7544918060303,31.7941417694092,31.8029804229736,31.7597179412842,31.7080574035645,31.6464347839355,31.6342296600342,31.6673831939697,31.7384757995605,31.8380870819092,31.9368171691895,32.2494621276855,32.4335441589355,32.5305671691895,32.59033203125,32.6827163696289,32.7642555236816,32.8328132629395,33.0200691223145,33.0641593933105,33.0947265625,33.1124992370605,33.1980972290039,33.2890129089355,33.3690414428711,33.4546852111816,33.6207542419434,33.7198715209961,33.8976554870605,34.0072746276855,34.0518074035645,33.9759750366211,33.9611320495605,33.950439453125,33.9522933959961,33.9818840026855,34.0497093200684,34.1202659606934,34.2040557861328,34.2732429504395,34.3694343566895,34.43115234375,34.4862823486328,34.5303688049316,34.5546379089355,34.599609375,34.6815948486328,34.779541015625,34.8677253723145,34.9096183776855,34.9669418334961,35.0511245727539,35.1014137268066,35.1506843566895,35.1830062866211,35.2117652893066,35.2479972839355,35.2888641357422,35.3285140991211,35.3704109191895,35.4079093933105,35.4608383178711,35.5468254089355,35.5975074768066,35.714599609375,35.833740234375,35.8801765441895,35.9385261535645,36.0006828308105,36.0420875549316,36.1217765808105,36.1711883544922,36.2932624816895,36.377685546875,36.4364738464355,36.4265632629395,36.4318351745605,36.486083984375,36.5341796875,36.6337394714355,36.7008819580078,36.7347183227539,36.7423858642578,36.7658195495605,36.802001953125,36.82958984375,36.8350105285645,36.8516120910645,36.8685569763184,36.8816909790039,36.8877906799316,36.8884773254395,36.8529319763184,36.8230972290039,36.8257331848145,36.8968772888184,36.9836921691895,37.0221710205078,37.0366706848145,37.0357398986816,37.0127449035645,36.9791030883789,36.9524383544922,36.9683570861816,36.9871597290039,36.9732398986816,36.9134788513184,36.8836898803711,36.7382316589355,36.7250480651855,36.7593269348145,36.7419929504395,36.6949234008789,36.6497039794922,36.6007347106934,36.5215797424316,36.4581069946289,36.3824234008789,36.1688461303711,36.1339340209961,36.0884742736816,36.0489730834961,36.017578125,35.996337890625,35.9830093383789,35.94921875,35.8290023803711,35.810546875,35.8109397888184,35.837158203125,35.8782234191895,35.8870620727539,35.7729988098145,35.7293968200684,35.6786613464355,35.6617202758789],[7.33281230926514,7.32753896713257,7.370361328125,7.45332002639771,7.48017549514771,7.50766611099243,7.59238290786743,7.62163066864014,7.62119102478027,7.52343797683716,7.48554658889771,7.42500066757202,7.38017559051514,7.33281230926514],[8.25419902801514,8.20981407165527,8.25185585021973,8.29541015625,8.29580116271973,8.25419902801514],[8.27426719665527,8.23193359375,8.32626914978027,8.3505859375,8.43583965301514,8.458251953125,8.46025371551514,8.4482421875,8.347900390625,8.30244159698486,8.27426719665527],[9.38071346282959,9.33408164978027,9.41811561584473,9.43188381195068,9.43027305603027,9.38071346282959],[8.65830135345459,8.64467716217041,8.56586933135986,8.49843788146973,8.42724609375,8.35165977478027,8.26953125,8.18706035614014,8.03388595581055,7.97246074676514,7.93251943588257,7.90815448760986,7.83652353286743,7.74907255172729,7.70585966110229,7.56625986099243,7.54306602478027,7.56455039978027,7.63461875915527,7.69121026992798,7.71093797683716,7.71186494827271,7.69882774353027,7.66806650161743,7.53696250915527,7.48369121551514,7.44282197952271,7.22934627532959,7.25634765625,7.54379892349243,7.89990282058716,8.06098556518555,8.070556640625,8.06694316864014,8.091796875,8.13862323760986,8.24755859375,8.33027362823486,8.38608360290527,8.37958908081055,8.28476619720459,8.23027420043945,8.15117168426514,8.13325214385986,8.21621036529541,8.32539081573486,8.39711952209473,8.49697303771973,8.45371055603027,8.41733360290527,8.39663028717041,8.42143535614014,8.45371055603027,8.46000957489014,8.4892578125,8.50566387176514,8.44340801239014,8.40493202209473,8.35859298706055,8.35532188415527,8.4033203125,8.44668006896973,8.628173828125,8.71372032165527,8.7421875,8.7529296875,8.81108379364014,8.84218692779541,8.93251991271973,8.99716758728027,9.02006816864014,9.00600528717041,8.97006893157959,8.92446231842041,8.90327072143555,8.85097694396973,8.77534198760986,8.71157264709473,8.63920879364014,8.59550762176514,8.34965896606445,8.31396484375,8.28876876831055,8.26245212554932,8.21386814117432,8.13994121551514,8.07705020904541,8.028564453125,7.99799776077271,7.85166025161743,7.66704082489014,7.59980487823486,7.50004863739014,7.45322275161743,7.43344783782959,7.42563486099243,7.38569355010986,7.32465791702271,7.27495145797729,7.22568368911743,7.22006845474243,7.27714824676514,7.43749952316284,7.71113348007202,7.89975595474243,7.87631893157959,7.85439491271973,7.80751943588257,7.66840791702271,7.62094736099243,7.62548875808716,7.67529296875,7.72119140625,8.01591777801514,8.07138633728027,8.13754844665527,8.16543006896973,8.215087890625,8.22275352478027,8.19482421875,8.23037147521973,8.31103610992432,8.27485370635986,8.28740215301514,8.32197284698486,8.30351543426514,8.24633884429932,8.09951114654541,8.07065391540527,8.13056564331055,8.19960975646973,8.2568359375,8.31601524353027,8.33774471282959,8.36777305603027,8.45351600646973,8.48935508728027,8.56396484375,8.63530254364014,8.74033164978027,8.80532264709473,8.857421875,8.89858436584473,8.91606426239014,8.95170879364014,8.99028301239014,9.05585956573486,9.06010723114014,9.24887657165527,9.44917011260986,9.46904277801514,9.48100566864014,9.511474609375,9.57080078125,9.591796875,9.54609394073486,9.505859375,9.51923847198486,9.538818359375,9.55820369720459,9.57666015625,9.52324199676514,9.42856407165527,9.38193321228027,9.33725643157959,9.20917987823486,9.19062519073486,9.21542930603027,9.19174861907959,9.16811466217041,9.14165115356445,9.031494140625,8.98007869720459,8.93486309051514,8.95610332489014,8.944091796875,8.95722675323486,9.04560565948486,9.11103534698486,9.14042949676514,9.11870098114014,9.07412147521973,9.01894569396973,8.82700157165527,8.78056621551514,8.78671836853027,8.81264686584473,8.88720607757568,9.02187442779541,9.08193302154541,9.20991230010986,9.34370136260986,9.361328125,9.37807559967041,9.47929668426514,9.55820369720459,9.59785175323486,9.56923770904541,9.53193378448486,9.53676795959473,9.51044940948486,9.45297813415527,9.428466796875,9.43476581573486,9.40629863739014,9.23627853393555,9.068115234375,8.88945388793945,8.65830135345459],[-24.3973636627197,-24.41259765625,-24.3997058868408,-24.3707027435303,-24.3402347564697,-24.3232421875,-24.3335933685303,-24.3646488189697,-24.3973636627197],[-4.23593759536743,-4.30117177963257,-4.32724571228027,-4.33310556411743,-4.28662109375,-4.29814434051514,-4.30117177963257,-4.24697256088257,-4.19365262985229,-4.15009784698486,-4.16757822036743,-4.16865205764771,-4.15888690948486,-4.17333984375,-4.22958993911743,-4.29531240463257,-4.35048818588257,-4.38720703125,-4.38818359375,-4.42431640625,-4.43759775161743,-4.46972703933716,-4.4873046875,-4.50439500808716,-4.55332040786743,-4.57460975646973,-4.64228487014771,-4.74892616271973,-4.78603506088257,-4.90126943588257,-5.00957059860229,-5.06718778610229,-5.09375,-5.12275362014771,-5.15771484375,-5.1982421875,-5.30253934860229,-5.49521446228027,-5.58964824676514,-5.63486337661743,-5.78955030441284,-5.93339872360229,-6.02871131896973,-6.09843730926514,-6.15644502639771,-6.26064443588257,-6.34433603286743,-6.40087890625,-6.46581983566284,-6.52519512176514,-6.56406259536743,-6.57470703125,-6.67949199676514,-6.83378887176514,-6.90576171875,-6.97353506088257,-7.07988309860229,-7.13505840301514,-7.17275381088257,-7.26279306411743,-7.30927753448486,-7.33535146713257,-7.34121131896973,-7.34990262985229,-7.37314462661743,-7.36728525161743,-7.37890625,-7.41669940948486,-7.46025371551514,-7.50663995742798,-7.53574180603027,-7.55605459213257,-7.58505916595459,-7.61123085021973,-7.65478515625,-7.73896455764771,-7.75351572036743,-7.78251981735229,-7.82900428771973,-7.87539100646973,-7.89570283889771,-7.93642520904541,-7.98574209213257,-8.02060508728027,-8.07285118103027,-8.14541053771973,-8.19189453125,-8.24990177154541,-8.29931640625,-8.34580039978027,-8.39218807220459,-8.42705059051514,-8.458984375,-8.47929763793945,-8.51416015625,-8.56699180603027,-8.65400409698486,-8.71933555603027,-8.81406211853027,-8.8828125,-8.9931640625,-9.1201171875,-9.26572322845459,-9.41142654418945,-9.40742206573486,-9.41035175323486,-9.45205116271973,-9.4921875,-9.51015663146973,-9.556640625,-9.62919902801514,-9.6884765625,-9.77431678771973,-9.84404277801514,-9.91015625,-10.0037107467651,-10.0051765441895,-10.0055665969849,-10.0060548782349,-9.98857498168945,-9.96601581573486,-9.85244083404541,-9.81875038146973,-9.76572227478027,-9.66904258728027,-9.62529277801514,-9.57168006896973,-9.51796817779541,-9.47822284698486,-9.46367168426514,-9.4375,-9.48984336853027,-9.54345703125,-9.62050724029541,-9.70458984375,-9.76748085021973,-9.82373046875,-9.97177696228027,-10.1815423965454,-10.361328125,-10.5860347747803,-10.8408203125,-11.01025390625,-10.9768562316895,-10.9468746185303,-11.0248050689697,-11.05859375,-11.0666990280151,-11.0642576217651,-11.0476570129395,-10.982421875,-10.9298830032349,-10.9333982467651,-10.9541015625,-10.9517574310303,-11.1687498092651,-11.3275394439697,-11.5085935592651,-11.654296875,-11.8756828308105,-12.0667972564697,-12.2704105377197,-12.5019521713257,-12.5607423782349,-12.6077146530151,-12.6872072219849,-12.7295904159546,-12.7551755905151,-12.8220701217651,-12.8800783157349,-12.9625978469849,-13.38232421875,-13.4963874816895,-13.5944337844849,-13.6439456939697,-13.6828126907349,-13.7802724838257,-13.9759769439697,-14.0146484375,-14.0943365097046,-14.1697263717651,-14.1988277435303,-14.234375,-14.2650384902954,-14.3772459030151,-14.4175786972046,-14.4703121185303,-14.5309572219849,-14.5725584030151,-14.5970706939697,-14.6710939407349,-14.7458982467651,-14.7953128814697,-14.8875007629395,-14.9629878997803,-15.0389652252197,-15.1987295150757,-15.2366199493408,-15.3329105377197,-15.3994140625,-15.6034183502197,-15.6406240463257,-15.7369146347046,-16.1491222381592,-16.1828136444092,-16.2219734191895,-16.2176761627197,-16.2619132995605,-16.3127937316895,-16.337890625,-16.3547859191895,-16.3890628814697,-16.4336910247803,-16.4759769439697,-16.5426750183105,-16.6421871185303,-16.67431640625,-16.7130851745605,-16.7684574127197,-16.8609371185303,-17.0013675689697,-17.0400390625,-17.08837890625,-17.1047840118408,-17.2001953125,-17.24853515625,-17.29443359375,-17.3329105377197,-17.3889656066895,-17.4603500366211,-17.5048828125,-17.5060558319092,-17.5732421875,-17.6498050689697,-17.6649417877197,-17.7038097381592,-17.7851543426514,-17.8999996185303,-17.990234375,-18.0934581756592,-18.2060546875,-18.2834968566895,-18.3251934051514,-18.3253898620605,-18.3335933685303,-18.3456058502197,-18.2777347564697,-18.0525379180908,-17.9320316314697,-17.8756828308105,-17.6825199127197,-17.6205081939697,-17.4219741821289,-17.3660163879395,-17.2943363189697,-17.1988277435303,-17.1510753631592,-17.0640640258789,-17.0025386810303,-16.8761711120605,-16.7749996185303,-16.7081069946289,-16.6145515441895,-16.5208969116211,-16.3885746002197,-16.3042964935303,-16.20166015625,-16.15283203125,-15.9125003814697,-15.8339834213257,-15.6990242004395,-15.4119138717651,-15.3201179504395,-15.1781253814697,-15.0935554504395,-14.8992195129395,-14.7849607467651,-14.6335935592651,-14.4958009719849,-14.3203125,-14.2266597747803,-14.1331052780151,-13.9484376907349,-13.8630857467651,-13.8214845657349,-13.8028326034546,-13.5152339935303,-13.3711919784546,-13.1099605560303,-12.984375,-12.8234376907349,-12.7280263900757,-12.5271482467651,-12.3487310409546,-12.21923828125,-12.1727533340454,-12.1068353652954,-12.0603513717651,-11.9234371185303,-11.6633787155151,-11.5324220657349,-11.2877922058105,-11.1935548782349,-11.0220699310303,-10.8367185592651,-10.2606449127197,-10.0890626907349,-9.81035137176514,-9.65205097198486,-9.37060546875,-9.15664005279541,-8.97109317779541,-8.74042987823486,-8.61699199676514,-8.40459060668945,-8.21015644073486,-8.04716873168945,-7.92324209213257,-7.83554697036743,-7.4189453125,-7.29560613632202,-7.06650400161743,-6.90165996551514,-6.76894521713257,-6.64960956573486,-6.2822265625,-6.12939453125,-6.05673789978027,-5.94238328933716,-5.87529325485229,-5.81240272521973,-5.86093759536743,-5.8408203125,-5.75898456573486,-5.63505887985229,-5.47539043426514,-5.16708993911743,-5.10185575485229,-5.02783203125,-4.87949180603027,-4.76074266433716,-4.66953086853027,-4.322265625,-4.23427724838257,-3.88164043426514,-3.73105478286743,-3.63818359375,-3.49609398841858,-3.38789081573486,-3.40644550323486,-3.42460942268372,-3.46103525161743,-3.49248027801514,-3.52216815948486,-3.57675766944885,-3.61318373680115,-3.65449237823486,-3.71074223518372,-3.73886680603027,-3.78769516944885,-3.87773394584656,-3.90585947036743,-3.92402338981628,-3.94882822036743,-4.00507831573486,-4.00341749191284,-3.97861337661743,-4.01005887985229,-4.06953144073486,-4.119140625,-4.16552734375,-4.20517539978027,-4.20849561691284,-4.33583974838257,-4.39365243911743,-4.43007802963257,-4.46142578125,-4.46367168426514,-4.41679668426514,-4.34902334213257,-4.31103515625,-4.29609394073486,-4.32753896713257,-4.39033222198486,-4.44589853286743,-4.47636747360229,-4.46757793426514,-4.45488262176514,-4.50058603286743,-4.53916025161743,-4.67060565948486,-4.76621055603027,-4.84003925323486,-4.92783164978027,-4.95761680603027,-4.95820331573486,-4.99062490463257,-4.96914005279541,-4.90800809860229,-4.87324237823486,-4.8583984375,-4.81865215301514,-4.77070331573486,-4.71445322036743,-4.6650390625,-4.59267568588257,-4.56240224838257,-4.51767587661743,-4.45820283889771,-4.42509746551514,-4.38398408889771,-4.32587909698486,-4.24814462661743,-4.15732431411743,-4.04160118103027,-3.98691415786743,-3.95214819908142,-3.90205025672913,-3.84306621551514,-3.77685523033142,-3.70576143264771,-3.67431640625,-3.59482407569885,-3.43124985694885,-3.39736366271973,-3.38828086853027,-3.39902329444885,-3.43613290786743,-3.47255849838257,-3.48916006088257,-3.48583984375,-3.46513628959656,-3.43212890625,-3.39980506896973,-3.38046884536743,-3.35019540786743,-3.28388714790344,-3.20683622360229,-3.04697299003601,-2.98164081573486,-2.91240215301514,-2.85996079444885,-2.80966806411743,-2.73769545555115,-2.63593745231628,-2.56259775161743,-2.43232440948486,-2.33134746551514,-2.24394512176514,-2.13310527801514,-1.89345717430115,-1.72812497615814,-1.60732424259186,-1.53124976158142,-1.31630861759186,-1.07119154930115,-0.962206959724426,-0.924316167831421,-0.940234661102295,-0.966796934604645,-0.96806663274765,-0.966796934604645,-0.951855361461639,-0.707129061222076,-0.653906404972076,-0.590136766433716,-0.55537086725235,-0.506542861461639,-0.408886730670929,-0.321777105331421,-0.248339965939522,-0.200097486376762,-0.15761724114418,-0.122851319611073,-0.122851319611073,-0.157129094004631,-0.145995944738388,-0.142187505960464,-0.106543220579624,-0.0417479984462261,-0.0417479984462261,-0.050488468259573,-0.11669946461916,-0.155859425663948,-0.188183397054672,-0.199413955211639,-0.203320384025574,-0.200097486376762,-0.244531527161598,-0.29863303899765,-0.335253655910492,-0.37001958489418,-0.429882794618607,-0.470117002725601,-0.517675936222076,-0.580664098262787,-0.691406488418579,-0.766601502895355,-0.808398187160492,-0.850878953933716,-0.927831768989563,-0.970605432987213,-0.997753798961639,-1.02861332893372,-1.09814465045929,-1.12519514560699,-1.19667971134186,-1.21796846389771,-1.21416008472443,-1.24882793426514,-1.31640601158142,-1.40136706829071,-1.44970703125,-1.53662133216858,-1.63886725902557,-1.69306623935699,-1.73740231990814,-1.78388690948486,-1.77226555347443,-1.78769540786743,-1.83027362823486,-1.88037133216858,-2.00332045555115,-2.08105492591858,-2.15634799003601,-2.20839834213257,-2.27822279930115,-2.31201195716858,-2.33974599838257,-2.39404273033142,-2.40849637985229,-2.40546894073486,-2.39218759536743,-2.36103534698486,-2.35166001319885,-2.36513662338257,-2.39501976966858,-2.42890620231628,-2.40927720069885,-2.40048813819885,-2.38066434860229,-2.32460951805115,-2.32656264305115,-2.28867197036743,-2.22773432731628,-2.16630864143372,-2.15273404121399,-2.18212866783142,-2.22421860694885,-2.27919912338257,-2.29375004768372,-2.33408236503601,-2.33486318588257,-2.31308627128601,-2.24541020393372,-2.22578120231628,-2.20683574676514,-2.21855449676514,-2.34199190139771,-2.40576148033142,-2.41826176643372,-2.45312523841858,-2.49072265625,-2.529296875,-2.55253911018372,-2.60654330253601,-2.63984394073486,-2.65820336341858,-2.70165967941284,-2.73076152801514,-2.75019502639771,-2.86406254768372,-3.08730459213257,-3.28828096389771,-3.60458970069885,-3.78154325485229,-3.78896450996399,-3.86640620231628,-3.86933588981628,-3.849609375,-3.81874990463257,-3.81435537338257,-3.84423828125,-3.88271498680115,-3.9951171875,-4.05019521713257,-4.09218788146973,-4.16201162338257,-4.23593759536743],[5.256591796875,5.19623994827271,5.16831064224243,5.18408203125,5.21538066864014,5.16899442672729,5.15112352371216,5.07954120635986,5.06020498275757,5.06953096389771,5.13315391540527,5.22841787338257,5.26362276077271,5.33242177963257,5.34008836746216,5.28408193588257,5.256591796875],[6.07563447952271,6.00351572036743,6.02226543426514,6.00209951400757,5.96450185775757,5.93984365463257,5.86997079849243,5.94272470474243,5.89301776885986,5.92294979095459,5.89619112014771,5.95263671875,6.00693368911743,6.09599590301514,6.07563447952271],[6.42831993103027,6.41455078125,6.41582059860229,6.517578125,6.562744140625,6.61372041702271,6.6640625,6.67622041702271,6.74072217941284,6.63891553878784,6.60224628448486,6.57978534698486,6.48291015625,6.42831993103027],[6.96274375915527,6.90576171875,7.03999042510986,7.07319307327271,7.185546875,7.13066434860229,6.96274375915527],[7.40913152694702,7.3857421875,7.36103487014771,7.31459999084473,7.31596660614014,7.39335870742798,7.428466796875,7.39731407165527,7.43339824676514,7.40913152694702],[7.88339853286743,7.80751943588257,7.89492177963257,8.01665019989014,8.050537109375,8.06914043426514,7.88339853286743],[8.21464824676514,8.19101524353027,8.25351524353027,8.31499004364014,8.30849552154541,8.28925800323486,8.21464824676514],[9.14262676239014,9.08310604095459,9.13232421875,9.17509746551514,9.225830078125,9.24301719665527,9.2431640625,9.19008827209473,9.14262676239014],[9.2373046875,9.133544921875,9.10332012176514,9.12973594665527,9.19209003448486,9.21552658081055,9.21357440948486,9.26826190948486,9.27875995635986,9.2373046875],[9.59355449676514,9.56796932220459,9.62148475646973,9.73920822143555,9.75908184051514,9.6845703125,9.59355449676514],[9.32094764709473,9.2607421875,9.27670955657959,9.12490177154541,9.08056640625,8.95205020904541,8.8447265625,8.74394607543945,8.6962890625,8.62729454040527,8.59565448760986,8.56005859375,8.539306640625,8.48388767242432,8.32675743103027,8.20283222198486,8.14877891540527,7.92744112014771,7.83281230926514,7.75698184967041,7.72480487823486,7.67724609375,7.54677772521973,7.32514667510986,7.24775362014771,7.17582988739014,7.01235389709473,6.88232421875,6.89101552963257,6.8525390625,6.73388671875,6.4833984375,6.30966806411743,6.39755868911743,6.48964834213257,6.73334932327271,6.84316396713257,6.94355440139771,7.03320264816284,7.11699199676514,7.33330106735229,7.32216787338257,7.26303720474243,7.22231483459473,7.16059589385986,7.10507822036743,7.0166015625,6.91113233566284,6.79575204849243,6.68994092941284,6.60712862014771,6.57373046875,6.49960947036743,6.46577119827271,6.22500038146973,5.97866201400757,5.87016582489014,5.66425800323486,5.59897470474243,5.63227558135986,5.75693368911743,5.80830049514771,5.92558574676514,6.03315448760986,6.06953096389771,6.06250047683716,6.04697275161743,5.90625047683716,5.87065410614014,5.86572265625,5.87534189224243,5.99819326400757,6.11972665786743,6.23325204849243,6.40444278717041,6.53256845474243,6.66655254364014,6.86298799514771,6.9296875,6.99370098114014,7.11411142349243,7.17509746551514,7.21879863739014,7.267333984375,7.33212852478027,7.39643573760986,7.43671846389771,7.57788038253784,7.66464853286743,7.74262666702271,7.785400390625,7.81777381896973,7.83164024353027,7.83212947845459,7.80791044235229,7.75634765625,7.71025371551514,7.66538047790527,7.40751934051514,7.46411180496216,7.52944326400757,7.5751953125,7.62993192672729,7.66689395904541,7.70043897628784,7.61435556411743,7.54628944396973,7.530517578125,7.52929639816284,7.55849552154541,7.72246122360229,7.77412128448486,7.76313495635986,7.67275381088257,7.63896465301514,7.56113290786743,7.34023427963257,7.17001914978027,7.00419950485229,6.94965791702271,6.91372060775757,6.92861318588257,6.96821308135986,7.0751953125,7.19951105117798,7.27875947952271,7.36357402801514,7.659912109375,7.76538038253784,7.81049823760986,7.94511699676514,8.02841758728027,8.0458984375,8.09331035614014,8.13310623168945,8.13369178771973,8.15644550323486,8.22050857543945,8.28691387176514,8.35605525970459,8.39834022521973,8.43393516540527,8.48081111907959,8.51601600646973,8.54145431518555,8.57041072845459,8.61562442779541,8.70332050323486,8.68154239654541,8.6474609375,8.62060546875,8.54770469665527,8.43271446228027,8.37607479095459,8.18881797790527,8.14511680603027,8.05825233459473,8.04912090301514,8.12841892242432,8.15898418426514,8.20146465301514,8.22954082489014,8.27138614654541,8.385986328125,8.50844669342041,8.55942440032959,8.599853515625,8.60634708404541,8.52265644073486,8.56298828125,8.68979454040527,8.87412071228027,8.92402267456055,8.97226524353027,8.95668888092041,8.89052677154541,8.86875057220459,8.92207050323486,9.02714824676514,9.02656269073486,8.99179744720459,9.01474571228027,9.14091777801514,9.27587890625,9.669189453125,9.75678730010986,9.75913143157959,9.65449237823486,9.51313400268555,9.42665958404541,9.32094764709473],[9.76621055603027,9.76079082489014,9.83852577209473,9.92705059051514,10.0592288970947,9.94355487823486,9.89111328125,9.86518573760986,9.79995155334473,9.76777362823486,9.76621055603027],[9.982177734375,9.93271446228027,10.0629396438599,10.155029296875,10.1510744094849,10.1156730651855,9.982177734375],[9.78720664978027,9.75048732757568,9.75351524353027,9.74790096282959,9.65410232543945,9.63022422790527,9.59931659698486,9.62397480010986,9.67573261260986,9.76113224029541,9.8173828125,9.87880897521973,9.91962909698486,10.0001955032349,10.0613279342651,10.1352052688599,10.159912109375,10.141357421875,10.1295900344849,10.1264162063599,10.0654783248901,10.0267086029053,9.87919902801514,9.82958984375,9.78720664978027],[9.91445350646973,9.886474609375,9.944091796875,9.99819278717041,10.090087890625,10.1187009811401,10.1915035247803,10.3097162246704,10.3636722564697,10.3795900344849,10.4368171691895,10.4401369094849,10.3920402526855,10.2454099655151,10.07177734375,9.96318435668945,9.93901348114014,9.91445350646973],[10.4859867095947,10.4552736282349,10.5516119003296,10.64013671875,10.6047859191895,10.5701169967651,10.5387210845947,10.4859867095947],[10.6060056686401,10.6014652252197,10.7066898345947,10.6913576126099,10.6798343658447,10.632568359375,10.6060056686401],[10.4727048873901,10.4590339660645,10.4610357284546,10.4249505996704,10.4925289154053,10.6075677871704,10.6950197219849,10.7225103378296,10.7388191223145,10.7406253814697,10.7063970565796,10.6545896530151,10.4982423782349,10.4727048873901],[9.06411170959473,9.05336856842041,9.058837890625,9.10795879364014,9.31982517242432,9.371337890625,9.41035175323486,9.44321250915527,9.48281288146973,9.69389629364014,9.82304668426514,9.89609336853027,9.96152400970459,9.97919940948486,9.98154258728027,9.99013710021973,10.0869140625,10.125,10.2840328216553,10.395263671875,10.5038080215454,10.5534172058105,10.6025390625,10.6983394622803,10.7657232284546,10.836181640625,10.8866205215454,10.9118165969849,10.9886713027954,10.9939451217651,10.9230470657349,10.8160648345947,10.78076171875,10.6620111465454,10.5823249816895,10.458984375,10.3253908157349,10.12451171875,10.0590333938599,9.93330001831055,9.8642578125,9.71464824676514,9.65932655334473,9.60615253448486,9.35698223114014,9.31748008728027,9.27294921875,9.21728515625,9.12138652801514,9.087890625,9.06411170959473],[9.44960975646973,9.42294979095459,9.48896503448486,9.57807636260986,9.88925743103027,9.96708965301514,10.1403322219849,10.3029298782349,10.4736814498901,10.5622072219849,10.7310056686401,10.9638185501099,11.0409183502197,11.0791501998901,11.137451171875,11.1869144439697,11.2735357284546,11.2172365188599,11.106689453125,11.0536136627197,11.0287590026855,10.92578125,10.7678709030151,10.5855960845947,10.4000978469849,10.3166007995605,10.2577142715454,10.2208013534546,10.1283197402954,10.0202140808105,9.92172813415527,9.58930683135986,9.44960975646973],[11.2833013534546,11.1507320404053,11.1514644622803,11.2479982376099,11.2791509628296,11.2833013534546],[8.43959903717041,8.36728572845459,8.45668983459473,8.54096603393555,8.71357440948486,8.76665019989014,8.902587890625,8.96831035614014,9.09824180603027,9.24067401885986,9.25126934051514,9.25341796875,9.269775390625,9.3466796875,9.602783203125,9.79365158081055,10.0350093841553,10.1053228378296,10.1312990188599,10.3535642623901,10.3858394622803,10.4092779159546,10.439453125,10.477294921875,10.5740232467651,10.6871099472046,10.7509765625,10.8451652526855,10.9736328125,11.0329103469849,11.2937984466553,11.346435546875,11.3135251998901,11.2667970657349,11.1568355560303,11.1016111373901,11.0455074310303,10.9531736373901,10.7073726654053,10.5517091751099,10.5003414154053,10.4074220657349,10.3793458938599,10.3543939590454,10.3272943496704,10.251708984375,10.1521492004395,10.1006832122803,10.0610837936401,9.99345684051514,9.94931602478027,9.91611385345459,9.86210918426514,9.76679611206055,9.42275428771973,9.33266639709473,9.25600624084473,9.20146560668945,9.16796875,9.10136699676514,8.98354530334473,8.87709999084473,8.79824161529541,8.72861289978027,8.67783260345459,8.64199256896973,8.59560489654541,8.538330078125,8.51137733459473,8.495849609375,8.43959903717041],[11.5253410339355,11.472412109375,11.39306640625,11.3756837844849,11.4087400436401,11.4319334030151,11.4746580123901,11.4736328125,11.5154294967651,11.5253410339355],[11.3430671691895,11.30810546875,11.3220710754395,11.4014158248901,11.3728513717651,11.2559080123901,11.2117185592651,11.13525390625,10.951904296875,10.7856931686401,10.75146484375,10.7215814590454,10.6847171783447,10.637451171875,10.5848636627197,10.4572257995605,10.349609375,10.3077154159546,10.2638187408447,10.2724113464355,10.2353506088257,10.189453125,10.2183103561401,10.3234376907349,10.3675785064697,10.1970701217651,10.1152830123901,10.0331048965454,10.0958976745605,10.1346197128296,10.1680669784546,10.2745609283447,10.3275384902954,10.4397459030151,10.6822261810303,10.7317867279053,10.7813968658447,10.8797369003296,10.9619626998901,10.9622068405151,10.9044427871704,10.923583984375,11.1503419876099,11.3707036972046,11.4270992279053,11.4861812591553,11.5352048873901,11.5149898529053,11.4572267532349,11.4238777160645,11.39501953125,11.3430671691895],[11.4921875,11.48583984375,11.53173828125,11.6659173965454,11.6950197219849,11.6871089935303,11.6396980285645,11.549560546875,11.4921875],[11.5947751998901,11.5433111190796,11.5575675964355,11.5807619094849,11.6585445404053,11.7153329849243,11.6831064224243,11.6266117095947,11.5947751998901],[11.6150884628296,11.5641603469849,11.60791015625,11.5956544876099,11.529296875,11.4873533248901,11.4413089752197,11.5414543151855,11.5355463027954,11.4425287246704,11.3635740280151,11.2868165969849,11.1968755722046,11.1165037155151,11.0581541061401,11.0224609375,10.9900388717651,10.9412107467651,10.8797369003296,10.8238277435303,10.8009271621704,10.69189453125,10.6229000091553,10.5755367279053,10.5140628814697,10.4583005905151,10.4443845748901,10.4708986282349,10.49365234375,10.6385746002197,10.6988763809204,10.7573728561401,10.8716793060303,10.9791011810303,11.0973634719849,11.32568359375,11.6429195404053,11.6808595657349,11.7237300872803,11.75830078125,11.7908697128296,11.8543453216553,11.8973627090454,11.8954095840454,11.8550786972046,11.7720212936401,11.7021970748901,11.6150884628296],[11.7033195495605,11.6693849563599,11.6907224655151,11.7403316497803,11.7744636535645,11.9539546966553,11.9813480377197,11.9602537155151,11.93212890625,11.917236328125,11.8605470657349,11.8213376998901,11.7834959030151,11.7033195495605],[12.1676759719849,12.1523923873901,12.1670408248901,12.2198238372803,12.1417484283447,12.0774421691895,12.0124025344849,12.0047855377197,12.0196285247803,11.9937496185303,12.0082521438599,12.0692386627197,12.1787595748901,12.1990232467651,12.2439947128296,12.2725095748901,12.2798824310303,12.2998533248901,12.3134279251099,12.319091796875,12.2704105377197,12.19775390625,12.1676759719849],[12.3090333938599,12.2855958938599,12.3836917877197,12.4294919967651,12.455078125,12.491943359375,12.4916019439697,12.4242677688599,12.38232421875,12.3090333938599],[12.5278806686401,12.4462890625,12.38720703125,12.3218259811401,12.2927732467651,12.2848625183105,12.251953125,12.19140625,12.1357917785645,12.0545902252197,11.9525394439697,11.7715816497803,11.7137699127197,11.6554203033447,11.5943355560303,11.5442380905151,11.3782224655151,11.3230466842651,11.2794923782349,11.2382326126099,11.23388671875,11.164794921875,11.0735845565796,11.0496091842651,11.1208000183105,11.1320314407349,11.1125974655151,11.1422843933105,11.1450681686401,11.2670412063599,11.287353515625,11.3412590026855,11.4791507720947,11.5583982467651,11.6384763717651,11.7023439407349,11.7649412155151,11.754638671875,11.7754888534546,11.8520994186401,11.8963384628296,11.933349609375,12.0208988189697,12.0551271438599,12.0791997909546,12.1527833938599,12.2439947128296,12.40380859375,12.5693359375,12.5262203216553,12.5345697402954,12.5725584030151,12.5278806686401],[12.2873544692993,12.1690921783447,11.966796875,11.8115720748901,11.75244140625,11.8188972473145,11.91357421875,11.9344720840454,11.9515619277954,12.0026369094849,12.0500001907349,12.0693359375,12.0902347564697,12.1966314315796,12.2166509628296,12.1942377090454,12.036376953125,11.9256353378296,11.9679679870605,12.1065921783447,12.3280277252197,12.3954591751099,12.4946775436401,12.58349609375,12.5423831939697,12.501220703125,12.44482421875,12.4069337844849,12.2873544692993],[12.5288076400757,12.498291015625,12.5030765533447,12.5569343566895,12.5929203033447,12.5892581939697,12.5288076400757],[12.3548822402954,12.1056146621704,12.19140625,12.2453126907349,12.2903804779053,12.331298828125,12.3854007720947,12.435302734375,12.5985345840454,12.650634765625,12.6526365280151,12.6125984191895,12.5375490188599,12.3548822402954],[12.4539060592651,12.3662595748901,12.3985347747803,12.5705080032349,12.63330078125,12.6749019622803,12.6107912063599,12.4539060592651],[12.8534183502197,12.7008295059204,12.8050785064697,12.8758792877197,12.9934577941895,13.0347166061401,13.0586919784546,13.1071786880493,13.1161613464355,13.1133794784546,12.9054193496704,12.8534183502197],[13.4794921875,13.4709959030151,13.5010747909546,13.4859857559204,13.4287109375,13.4107427597046,13.3812494277954,13.4323244094849,13.3741207122803,13.2654790878296,13.1884279251099,13.1312017440796,13.0888671875,13.0195798873901,12.9315910339355,12.8371095657349,12.6381845474243,12.584228515625,12.5079593658447,12.4230470657349,12.3887691497803,12.3607416152954,12.3005857467651,12.3130855560303,12.3036136627197,12.2767095565796,12.2187986373901,12.236328125,12.25341796875,12.3036136627197,12.3389644622803,12.3599605560303,12.446533203125,12.5116205215454,12.5811033248901,12.6458492279053,12.7036619186401,12.7479982376099,12.7905759811401,12.8409175872803,12.9698247909546,13.130615234375,13.1691398620605,13.2088861465454,13.2600584030151,13.31103515625,13.3935060501099,13.4054203033447,13.4016599655151,13.412353515625,13.4729490280151,13.5170412063599,13.5224123001099,13.4976072311401,13.4794921875],[13.5403327941895,13.5374021530151,13.5467777252197,13.4631834030151,13.4208498001099,13.3651361465454,13.2686529159546,13.2361822128296,13.2049808502197,13.28173828125,13.3286142349243,13.4244623184204,13.5661630630493,13.5403327941895],[13.7506837844849,13.6829586029053,13.7823734283447,13.8169431686401,13.842529296875,13.8580570220947,13.8206539154053,13.7506837844849],[13.6322269439697,13.5673828125,13.59033203125,13.586669921875,13.5315427780151,13.6055660247803,13.6631355285645,13.7904787063599,13.9796876907349,14.0261716842651,14.0595216751099,14.0775880813599,13.9469718933105,13.9311027526855,13.8710441589355,13.7500982284546,13.6794443130493,13.6322269439697],[14.048828125,14.0080080032349,14.1560544967651,14.1814947128296,14.2051267623901,14.228759765625,14.048828125],[15.0050287246704,14.9698724746704,14.9716310501099,14.9652843475342,14.8929681777954,14.7594242095947,14.6621589660645,14.6560544967651,14.6665048599243,14.7145509719849,14.7366218566895,14.8000001907349,14.8398427963257,14.9171876907349,14.9635753631592,15.0381355285645,15.0463857650757,15.0050287246704],[18.6152820587158,18.5634288787842,18.37646484375,18.330078125,18.29541015625,18.28515625,18.3279304504395,18.3716793060303,18.4865703582764,18.5006351470947,18.4588375091553,18.4027843475342,18.3203125,18.2342777252197,18.1571292877197,18.0642566680908,17.7564945220947,17.6644039154053,17.57568359375,17.4348640441895,17.395263671875,17.3448734283447,17.3067874908447,17.2383804321289,17.1781253814697,17.1551265716553,17.1248531341553,17.0580081939697,16.9900398254395,16.8226566314697,16.4352035522461,16.3515129089355,16.184814453125,16.1579113006592,16.0774421691895,16.0147457122803,15.9332513809204,15.8267574310303,15.7780275344849,15.7260255813599,15.669825553894,15.6231937408447,15.5095224380493,15.4166507720947,15.3750476837158,15.3244142532349,15.2666015625,15.2163095474243,14.9991703033447,14.7895021438599,14.7654294967651,14.7373046875,14.6817378997803,14.5811529159546,14.4814929962158,14.2341794967651,14.1680660247803,14.1138668060303,14.0630855560303,14.0204105377197,13.9471187591553,13.9327154159546,13.93017578125,13.9794921875,13.9961910247803,14.0447263717651,14.1116695404053,14.1480474472046,14.175048828125,14.1908206939697,14.2638673782349,14.3223638534546,14.3175296783447,14.2848634719849,14.2507810592651,14.1880855560303,14.079833984375,13.9599609375,13.9027347564697,13.845458984375,13.7887697219849,13.750244140625,13.7473630905151,13.83642578125,13.9365720748901,13.9754400253296,14.0248050689697,14.0616703033447,14.0286617279053,13.9662599563599,13.8984861373901,13.8970212936401,13.8843269348145,13.8371095657349,13.7996091842651,13.7217292785645,13.7044429779053,13.645751953125,13.528076171875,13.4315919876099,13.3535146713257,13.2694826126099,13.1916007995605,13.1105470657349,13.1169919967651,13.0997066497803,13.0319337844849,13.0249996185303,13.0357904434204,12.7911615371704,12.5670900344849,12.594970703125,12.689697265625,12.8049812316895,12.9164066314697,12.93994140625,12.9306640625,12.9055662155151,12.8969240188599,12.9117670059204,13.0331058502197,13.0440921783447,13.0990238189697,13.2155771255493,13.3535146713257,13.4028806686401,13.4417486190796,13.5919427871704,13.6172361373901,13.737060546875,13.9076175689697,13.925048828125,13.9299802780151,13.88671875,13.8202152252197,13.7630853652954,13.7031736373901,13.6568365097046,13.56201171875,13.5171394348145,13.3953609466553,13.253173828125,13.1941413879395,13.260009765625,13.3136234283447,13.363525390625,13.4927730560303,13.5206050872803,13.6482419967651,13.7883787155151,13.9376468658447,13.9458494186401,13.9345703125,13.9159669876099,13.8421878814697,13.790771484375,13.7118654251099,13.6491212844849,13.6402826309204,13.6794929504395,13.7581052780151,13.7618656158447,13.8847169876099,13.9005374908447,13.8044919967651,13.9953117370605,14.1880369186401,14.2443351745605,14.2912101745605,14.4931154251099,14.5579595565796,14.6450681686401,14.7157707214355,14.7587890625,14.7766103744507,14.816162109375,14.8812503814697,14.7661123275757,14.5946283340454,14.4831056594849,14.4413576126099,14.4401865005493,14.4533681869507,14.4933109283447,14.6083011627197,14.6843757629395,14.7933101654053,14.8087892532349,14.8003911972046,14.8510732650757,14.9781246185303,15.1145515441895,15.2292470932007,15.3402347564697,15.430908203125,15.8376960754395,15.8750009536743,15.90576171875,15.9519529342651,16.0084476470947,16.0549812316895,16.2551288604736,16.3033199310303,16.3265628814697,16.2874031066895,16.23876953125,16.2154293060303,16.1845703125,16.0662117004395,16.0476551055908,16.0514163970947,16.0664558410645,16.1095695495605,16.1609382629395,16.2216300964355,16.4003410339355,16.5292472839355,16.6454582214355,16.7618656158447,16.9556159973145,17.090087890625,17.2699222564697,17.3769054412842,17.4383296966553,17.5351066589355,17.63818359375,18.1626453399658,18.2640628814697,18.3687496185303,18.5078601837158,18.5459480285645,18.6034183502197,18.5989265441895,18.5851078033447,18.6136722564697,18.6152820587158],[18.8947257995605,18.8229007720947,18.8427238464355,18.9125499725342,18.936767578125,18.9915523529053,19.0104484558105,18.9566402435303,18.8947257995605],[19.0824222564697,19.0151844024658,19.0506839752197,19.1014156341553,19.138916015625,19.18359375,19.1430168151855,19.0824222564697],[19.3619632720947,19.2713375091553,19.2733402252197,19.3284664154053,19.3562984466553,19.3796863555908,19.3993663787842,19.3619632720947],[20.3658695220947,20.3537101745605,20.3594245910645,20.4537105560303,20.4795894622803,20.4693851470947,20.3658695220947],[20.7818851470947,20.7002944946289,20.701171875,20.7466316223145,20.8412609100342,20.8392086029053,20.7818851470947],[3.02622103691101,3.02187514305115,3.02524399757385,3.03945302963257,3.05410146713257,3.06059551239014,3.05561542510986,3.04208970069885,3.02622103691101],[7.38203096389771,7.36064481735229,7.43710947036743,7.52578067779541,7.59394502639771,7.61577177047729,7.62358379364014,7.71210908889771,7.66328096389771,7.50131845474243,7.43828105926514,7.38203096389771],[-11.4761714935303,-11.5285158157349,-11.5863285064697,-11.6125974655151,-11.6305665969849,-11.59521484375,-11.5595703125,-11.4950199127197,-11.5169925689697,-11.4714841842651,-11.4203119277954,-11.3518552780151,-11.3241214752197,-11.3563480377197,-11.4761714935303],[-11.3614263534546,-11.4159183502197,-11.3974609375,-11.4256839752197,-11.4193363189697,-11.3705081939697,-11.3479490280151,-11.3655271530151,-11.3114261627197,-11.3388671875,-11.3614263534546],[-10.5653314590454,-10.6434574127197,-10.6202154159546,-10.6034183502197,-10.5541009902954,-10.5518550872803,-10.5653314590454],[-10.0201168060303,-10.0202140808105,-9.94550800323486,-9.92265605926514,-9.95673847198486,-10.1947269439697,-10.1588869094849,-10.0925779342651,-9.99843692779541,-9.96806716918945,-9.87617206573486,-9.77431678771973,-9.70908260345459,-9.73593807220459,-9.80244159698486,-9.93896484375,-9.98310565948486,-10.0201168060303],[-9.34658241271973,-9.42851543426514,-9.40449142456055,-9.41796875,-9.51269435882568,-9.58193302154541,-9.64140605926514,-9.66748046875,-9.70283222198486,-9.66259765625,-9.65478515625,-9.65654277801514,-9.63115215301514,-9.62460899353027,-9.56171798706055,-9.53613376617432,-9.43496036529541,-9.38662052154541,-9.35996150970459,-9.34560585021973,-9.34658241271973],[-9.49384784698486,-9.5185546875,-9.50039100646973,-9.36191368103027,-9.25957012176514,-9.20634841918945,-9.26416015625,-9.34902381896973,-9.396484375,-9.49384784698486],[-8.95937538146973,-8.974609375,-8.96718788146973,-9.02451133728027,-9.04423809051514,-9.07011699676514,-9.10781288146973,-9.13076210021973,-9.15078067779541,-9.16865253448486,-9.208984375,-9.20302772521973,-9.22431659698486,-9.17714881896973,-9.16650390625,-9.12607479095459,-9.05839824676514,-9.00986289978027,-8.97001934051514,-8.95937538146973],[-8.63339805603027,-8.67714786529541,-8.61708927154541,-8.51679706573486,-8.48955059051514,-8.42343711853027,-8.47275447845459,-8.51894474029541,-8.63339805603027],[-8.73349571228027,-8.80488300323486,-8.72832012176514,-8.64179706573486,-8.59501934051514,-8.56865215301514,-8.52382850646973,-8.45058536529541,-8.42597675323486,-8.41884803771973,-8.52187538146973,-8.56806659698486,-8.73349571228027],[-8.48173809051514,-8.48476600646973,-8.41689395904541,-8.36757850646973,-8.37851524353027,-8.39091777801514,-8.4599609375,-8.48173809051514],[-5.82636690139771,-5.83398485183716,-5.78857421875,-5.74921894073486,-5.62724590301514,-5.5224609375,-5.49238252639771,-5.49082040786743,-5.61152315139771,-5.65019512176514,-5.7646484375,-5.82636690139771],[-6.68681621551514,-6.78046846389771,-6.79667997360229,-6.76152372360229,-6.79560518264771,-6.83437538146973,-6.86279249191284,-6.85595703125,-6.83027315139771,-6.78271532058716,-6.7216796875,-6.62607431411743,-6.52685594558716,-6.41162109375,-6.3076171875,-6.23369121551514,-6.20976591110229,-6.10615205764771,-6.06142568588257,-5.970703125,-5.93134784698486,-5.81650352478027,-5.7470703125,-5.54531240463257,-5.44443321228027,-5.4541015625,-5.49404335021973,-5.52138710021973,-5.53994178771973,-5.62021493911743,-5.77695322036743,-5.82832050323486,-5.86523389816284,-5.93173789978027,-5.97441434860229,-6.14511728286743,-6.18154335021973,-6.19619131088257,-6.22080087661743,-6.29570341110229,-6.38046884536743,-6.46962881088257,-6.56503915786743,-6.68681621551514],[-5.43193387985229,-5.43740224838257,-5.34238290786743,-5.30703115463257,-5.25742149353027,-5.19082021713257,-5.25156259536743,-5.38154268264771,-5.43193387985229],[-5.43271493911743,-5.44062519073486,-5.31445264816284,-5.22089862823486,-5.15195274353027,-5.11083984375,-5.03496122360229,-5.01386737823486,-5.05400419235229,-5.14267587661743,-5.21806621551514,-5.3828125,-5.43271493911743],[-4.72617197036743,-4.75576162338257,-4.73300790786743,-4.66747999191284,-4.60419940948486,-4.55429697036743,-4.53925752639771,-4.57314443588257,-4.64013624191284,-4.72617197036743],[-4.29677724838257,-4.31699228286743,-4.21220731735229,-4.28515672683716,-4.32070350646973,-4.34072256088257,-4.49082040786743,-4.56025457382202,-4.62929677963257,-4.73124980926514,-4.82216835021973,-4.95468759536743,-4.97919940948486,-4.99316358566284,-5.00380897521973,-5.07441377639771,-5.14902353286743,-5.24706983566284,-5.35703086853027,-5.45830106735229,-5.52880859375,-5.56484413146973,-5.54355430603027,-5.55234336853027,-5.59091854095459,-5.65458965301514,-5.703125,-5.74736309051514,-5.83906269073486,-5.91992235183716,-5.99667930603027,-6.01503944396973,-6.02724599838257,-6.07138681411743,-6.11445331573486,-6.1494140625,-6.18779277801514,-6.26337862014771,-6.27617168426514,-6.28935575485229,-6.29296827316284,-6.30087900161743,-6.29042959213257,-6.26093721389771,-6.12480497360229,-6.07812452316284,-6.07949256896973,-6.12763690948486,-6.11699199676514,-5.91640615463257,-5.86738300323486,-5.83076190948486,-5.80537080764771,-5.76503849029541,-5.66943359375,-5.544921875,-5.47177743911743,-5.50791025161743,-5.50742244720459,-5.48662090301514,-5.49326181411743,-5.51162147521973,-5.48457002639771,-5.52265644073486,-5.57304716110229,-5.58398389816284,-5.5732421875,-5.51601552963257,-5.52353477478027,-5.52412080764771,-5.44775390625,-5.13955020904541,-5.03466796875,-5.01181650161743,-5.01816415786743,-5.07060527801514,-5.13603496551514,-5.18642616271973,-5.30956983566284,-5.42900371551514,-5.52363300323486,-5.53564405441284,-5.47314453125,-5.46025371551514,-5.52089881896973,-5.51044893264771,-5.47089815139771,-5.45371103286743,-5.44716787338257,-5.42373037338257,-5.32070350646973,-5.20449209213257,-5.11289072036743,-4.96035194396973,-4.94130849838257,-4.93095731735229,-4.9375,-4.88330078125,-4.76103496551514,-4.63701152801514,-4.34550809860229,-4.29921865463257,-4.24736309051514,-4.20078086853027,-4.20000028610229,-4.21699190139771,-4.26083993911743,-4.29677724838257],[-4.09931659698486,-4.123046875,-4.09599637985229,-4.04472637176514,-4.04121112823486,-4.09931659698486],[-3.13339829444885,-3.22119164466858,-3.16982412338257,-3.09560561180115,-3.06250023841858,-3.04277300834656,-3.13339829444885],[-2.94736289978027,-2.99785161018372,-2.99492216110229,-2.89609360694885,-2.87050747871399,-2.84560513496399,-2.91845679283142,-2.94736289978027],[-2.83017587661743,-2.83037090301514,-2.75058579444885,-2.70859384536743,-2.73779296875,-2.80917930603027,-2.83017587661743],[-9.11874961853027,-9.10556602478027,-8.90224647521973,-8.69892597198486,-8.49570274353027,-8.29238224029541,-8.08906173706055,-7.88574266433716,-7.68251895904541,-7.47919893264771,-7.27587842941284,-7.07255840301514,-6.90537118911743,-6.84003877639771,-6.74003887176514,-6.61152315139771,-6.45224618911743,-6.34609365463257,-6.25937509536743,-6.05615234375,-5.85283231735229,-5.64951133728027,-5.44619131088257,-5.24296903610229,-5.03964853286743,-4.83632802963257,-4.63300800323486,-4.42978525161743,-4.22646522521973,-4.02314472198486,-3.81982421875,-3.61660146713257,-3.41328144073486,-3.20996117591858,-3.00664067268372,-2.80341792106628,-2.60976552963257,-2.61015605926514,-2.61132788658142,-2.62783193588257,-2.84501957893372,-2.93212866783142,-2.95253920555115,-2.95332026481628,-2.96357417106628,-3.08349609375,-3.20458960533142,-3.32070279121399,-3.34492182731628,-3.35507822036743,-3.39531254768372,-3.43115258216858,-3.57333993911743,-3.61728525161743,-3.69746088981628,-3.78359389305115,-3.80517601966858,-3.81523442268372,-3.81826186180115,-3.80273461341858,-3.80966830253601,-3.82529306411743,-3.85527372360229,-3.91308569908142,-3.99306654930115,-4.02910184860229,-4.10146522521973,-4.18818330764771,-4.27548837661743,-4.34912061691284,-4.38027334213257,-4.38525390625,-4.82304716110229,-4.890625,-5.17792987823486,-5.40244150161743,-5.47128915786743,-5.49707078933716,-5.54511737823486,-5.61660146713257,-5.91923809051514,-5.94501924514771,-5.95478534698486,-5.95078134536743,-5.96621084213257,-6.02109384536743,-6.05693340301514,-6.15478563308716,-6.26113271713257,-6.29150390625,-6.31523418426514,-6.373046875,-6.55117225646973,-6.66240262985229,-6.70361280441284,-6.7236328125,-6.74238300323486,-6.7216796875,-6.83408212661743,-6.88310527801514,-6.92880868911743,-7.16699171066284,-7.37812471389771,-7.46406221389771,-7.53378915786743,-7.6162109375,-7.71093797683716,-7.87626934051514,-7.93750047683716,-7.97539091110229,-8.10361289978027,-8.16025447845459,-8.33867263793945,-8.45966815948486,-8.50957012176514,-8.55429744720459,-8.66396427154541,-8.69453144073486,-8.93857383728027,-9.0517578125,-9.09199237823486,-9.08945274353027,-9.01689434051514,-9.01455020904541,-9.03125,-9.07099533081055,-9.18076229095459,-9.2958984375,-9.40683650970459,-9.49785232543945,-9.56884765625,-9.58828067779541,-9.61093807220459,-9.63007831573486,-9.66074275970459,-9.68818378448486,-9.73701190948486,-9.76083946228027,-9.77060604095459,-9.80585956573486,-9.86865234375,-9.93417930603027,-10.0129880905151,-10.0416011810303,-10.0607423782349,-10.0880851745605,-10.1255855560303,-10.1628904342651,-10.1896486282349,-10.2067384719849,-10.2571287155151,-10.2360353469849,-10.31787109375,-10.3379878997803,-10.3073244094849,-10.3392581939697,-10.42578125,-10.4840812683105,-10.5179691314697,-10.5576171875,-10.6369142532349,-10.6485357284546,-10.6548833847046,-10.6207036972046,-10.5771493911743,-10.5176763534546,-10.4826164245605,-10.3988285064697,-10.35302734375,-10.3374996185303,-10.3384761810303,-10.2897462844849,-10.2551755905151,-10.2339839935303,-10.1668949127197,-10.1573247909546,-10.1784181594849,-10.19140625,-10.1854486465454,-10.1282224655151,-10.1073246002197,-10.12451171875,-10.1283197402954,-10.08740234375,-10.0701169967651,-10.0130863189697,-9.95976543426514,-9.91240215301514,-9.79043006896973,-9.67470645904541,-9.57958984375,-9.42607402801514,-9.38789081573486,-9.24716758728027,-9.15390586853027,-9.0595703125,-9.09169960021973,-9.08769512176514,-9.025390625,-8.951171875,-8.74970722198486,-8.6435546875,-8.45556545257568,-8.34394550323486,-8.24638748168945,-8.21025371551514,-8.16845703125,-8.11415958404541,-8.07636737823486,-7.99277353286743,-7.96640634536743,-7.95244121551514,-7.94384813308716,-7.93007755279541,-7.86162137985229,-7.84111261367798,-7.828125,-7.80214834213257,-7.77665948867798,-7.73359346389771,-7.63154268264771,-7.64248037338257,-7.62480497360229,-7.58896446228027,-7.56738328933716,-7.59814500808716,-7.67939424514771,-7.68359375,-7.6669921875,-7.6767578125,-7.71425819396973,-7.76494169235229,-7.75722599029541,-7.70595693588257,-7.67382860183716,-7.61591768264771,-7.55009698867798,-7.49824237823486,-7.46035194396973,-7.54980516433716,-7.94423818588257,-7.95185565948486,-7.94189453125,-8.01767539978027,-8.02910137176514,-8.02822303771973,-7.99550819396973,-7.98466777801514,-8.00068378448486,-8.02910137176514,-8.11269474029541,-8.20039081573486,-8.23984336853027,-8.26386737823486,-8.31123065948486,-8.314453125,-8.28749942779541,-8.27226543426514,-8.28749942779541,-8.32168006896973,-8.31621170043945,-8.25468730926514,-8.2080078125,-8.16748046875,-8.17392539978027,-8.19580078125,-8.19833946228027,-8.25,-8.31269550323486,-8.36943340301514,-8.33564376831055,-8.34502029418945,-8.44384765625,-8.45517539978027,-8.47451210021973,-8.57216835021973,-8.66093730926514,-8.76220703125,-8.80185604095459,-8.90820217132568,-8.96103477478027,-9.03593730926514,-9.09247970581055,-9.20263671875,-9.32783222198486,-9.30332088470459,-9.23701190948486,-9.21904277801514,-9.18290996551514,-9.169921875,-9.19814491271973,-9.21259784698486,-9.21132755279541,-9.19013690948486,-9.15068340301514,-9.16816425323486,-9.21445369720459,-9.22128868103027,-9.11874961853027],[-4.75634765625,-4.83242177963257,-4.76152324676514,-4.69941425323486,-4.63583993911743,-4.49843788146973,-4.42919921875,-4.35595703125,-4.28203105926514,-4.1318359375,-3.99482440948486,-3.66816425323486,-3.58242177963257,-3.505859375,-3.48710942268372,-3.46875,-3.45341777801514,-3.40009760856628,-3.337890625,-3.15351557731628,-3.10136723518372,-3.03691411018372,-2.82900381088257,-2.77988290786743,-2.77978491783142,-2.73886728286743,-2.712890625,-2.64355444908142,-2.57294964790344,-2.68828129768372,-2.78906226158142,-2.87031221389771,-2.87529301643372,-2.94248032569885,-3.00302720069885,-3.07285165786743,-3.1728515625,-3.25136709213257,-3.27988266944885,-3.41035151481628,-3.52099585533142,-3.58193349838257,-4.10566425323486,-4.25234365463257,-4.35244131088257,-4.39169931411743,-4.47636747360229,-4.57636737823486,-4.66630887985229,-4.75634765625],[-2.66181635856628,-2.67548847198486,-2.66025376319885,-2.60253953933716,-2.51250028610229,-2.49150395393372,-2.47382831573486,-2.40498065948486,-2.38418006896973,-2.47041010856628,-2.51328110694885,-2.54111313819885,-2.63232398033142,-2.66181635856628],[-2.28310561180115,-2.33574199676514,-2.33125019073486,-2.31552767753601,-2.30556631088257,-2.26210951805115,-2.24677729606628,-2.28310561180115],[-1.96015632152557,-2.02509760856628,-2.02431654930115,-2.00107407569885,-1.99453139305115,-2.01152348518372,-2.05898475646973,-2.07060527801514,-2.06601595878601,-2.09042954444885,-2.18193387985229,-2.16660141944885,-2.18710947036743,-2.18906259536743,-2.14882826805115,-2.18271470069885,-2.17334008216858,-2.21044945716858,-2.20859360694885,-2.15410113334656,-2.12617182731628,-2.1025390625,-2.01689434051514,-1.97402346134186,-1.97773468494415,-1.94853496551514,-1.96015632152557],[-1.55302727222443,-1.58916032314301,-1.57089841365814,-1.57626938819885,-1.47167992591858,-1.40771484375,-1.35322260856628,-1.36201179027557,-1.43066418170929,-1.55302727222443],[54.4591789245605,54.4470710754395,54.4339866638184,54.4207496643066,54.4066390991211,54.3917999267578,54.37646484375,54.35986328125,54.35009765625,54.3567848205566,54.3958015441895,54.3905296325684,54.3663558959961,54.3483428955078,54.327880859375,54.30419921875,54.2994651794434,54.2814445495605,54.2403335571289,54.2004890441895,54.1434555053711,54.0790023803711,54.0059585571289,53.9589347839355,53.9397926330566,53.5992202758789,53.2709465026855,53.1121063232422,53.0275382995605,52.9048843383789,52.8187522888184,52.770263671875,52.70361328125,52.6642074584961,52.5515594482422,52.5162124633789,52.4283676147461,52.337890625,52.3069801330566,52.28662109375,52.2569351196289,52.2084465026855,52.1695327758789,52.140380859375,52.1030769348145,52.069580078125,52.0403785705566,51.9729995727539,51.8797836303711,51.8093223571777,51.7624015808105,51.7102546691895,51.618896484375,51.5179214477539,51.448974609375,51.3949241638184,51.3524894714355,51.31005859375,51.26513671875,51.126220703125,50.9925308227539,50.9404296875,50.8727531433105,50.844970703125,50.8195304870605,50.816162109375,50.8093757629395,50.7855949401855,50.7601547241211,50.7228012084961,50.6170387268066,50.5304718017578,50.5084457397461,50.45703125,50.4100570678711,50.3773460388184,50.3270492553711,50.2298316955566,50.1739234924316,50.0728492736816,49.8990707397461,49.8263664245605,49.7662620544434,49.606201171875,49.5390129089355,49.4836921691895,49.3538093566895,49.295166015625,49.240966796875,49.1927223205566,49.1711921691895,49.13623046875,49.0812492370605,49.062744140625,49.0389175415039,49.020751953125,49.0399436950684,49.0771980285645,49.0727081298828,49.081298828125,49.1532249450684,49.2095222473145,49.24609375,49.299072265625,49.3434600830078,49.3819313049316,49.4119606018066,49.4287605285645,49.429443359375,49.4170417785645,49.4182624816895,49.33984375,49.3170890808105,49.314697265625,49.3286590576172,49.3699226379395,49.3917007446289,49.3812026977539,49.3901863098145,49.392333984375,49.3840827941895,49.38525390625,49.3655281066895,49.337646484375,49.31640625,49.270751953125,49.1813011169434,49.2213859558105,49.2040023803711,49.1923332214355,49.2043952941895,49.2352066040039,49.2699699401855,49.3185539245605,49.3721694946289,49.3895988464355,49.3960456848145,49.4066429138184,49.4243659973145,49.4471206665039,49.5047836303711,49.5763664245605,49.5977058410645,49.5636215209961,49.5248565673828,49.5114250183105,49.3999977111816,49.396240234375,49.4482917785645,49.498291015625,49.5107917785645,49.5401382446289,49.6137237548828,49.7578125,49.8179206848145,49.8411140441895,49.8793449401855,49.9023933410645,49.9298324584961,49.9140625,49.9302749633789,49.9647445678711,49.9927749633789,50.0072746276855,50.0319366455078,50.0352554321289,50.020263671875,49.9990730285645,49.9722671508789,49.9832992553711,50.006591796875,50.0567855834961,50.1007804870605,50.1164054870605,50.1394996643066,50.157470703125,50.1935539245605,50.2307624816895,50.2986335754395,50.3071784973145,50.2842292785645,50.2640609741211,50.2547874450684,50.2547874450684,50.3783226013184,50.4161109924316,50.4270515441895,50.4145011901855,50.34521484375,50.2597198486328,50.2369155883789,50.2019500732422,50.1867179870605,50.1570281982422,50.1160659790039,50.0974617004395,50.1021499633789,50.1219215393066,50.2483901977539,50.34521484375,50.3668937683105,50.3718757629395,50.3940925598145,50.4237327575684,50.4546852111816,50.4830055236816,50.50048828125,50.5168952941895,50.5416526794434,50.5736351013184,50.5851554870605,50.6213874816895,50.6556167602539,50.6299324035645,50.6116180419922,50.6354484558105,50.6702651977539,50.6769027709961,50.7086906433105,50.7396965026855,50.7488746643066,50.7938499450684,50.7962875366211,50.811767578125,50.8457527160645,50.8830070495605,50.9585456848145,50.9928703308105,51.0143547058105,51.00341796875,50.92724609375,50.8865737915039,50.8613777160645,50.8589859008789,50.8716316223145,51.0087394714355,51.0951194763184,51.2527351379395,51.3771476745605,51.4353523254395,51.4633293151855,51.5238761901855,51.544921875,51.6271476745605,51.6617202758789,51.6981925964355,51.7708015441895,51.8323745727539,51.9048347473145,51.9580078125,52.0308570861816,52.07080078125,52.0818367004395,52.1102027893066,52.1500473022461,52.2074699401855,52.25,52.2776374816895,52.3141593933105,52.3596649169922,52.4311027526855,52.5285148620605,52.6456069946289,52.7825202941895,52.8782196044922,52.932861328125,52.9823226928711,53.0267562866211,53.1055679321289,53.1990242004395,53.2167510986328,53.2834930419922,53.5564460754395,53.624755859375,53.7071304321289,53.7296371459961,53.671875,53.6393547058105,53.6758766174316,53.7073249816895,53.7535171508789,53.8231925964355,53.8587379455566,53.8707504272461,53.9190444946289,53.9503440856934,53.9319305419922,53.9247093200684,54.018310546875,54.139892578125,54.2539558410645,54.266357421875,54.2903785705566,54.3330535888672,54.3616218566895,54.4368629455566,54.5538101196289,54.5963859558105,54.65185546875,54.7295417785645,54.8166999816895,54.8358383178711,54.8381843566895,54.7694320678711,54.6845703125,54.6333465576172,54.665283203125,54.7415046691895,54.7447280883789,54.5128898620605,54.430908203125,54.3695793151855,54.3489265441895,54.3860855102539,54.4346199035645,54.4591789245605],[18.0598640441895,18.0589351654053,18.0599117279053,18.0627937316895,18.0718746185303,18.0869140625,18.1006355285645,18.1151371002197,18.1208992004395,18.12255859375,18.1211433410645,18.1110363006592,18.1039543151855,18.0970211029053,18.07958984375,18.0751953125,18.0706558227539,18.0627937316895,18.0598640441895],[18.1056156158447,18.0995121002197,18.107666015625,18.1373043060303,18.1650390625,18.1610832214355,18.1443843841553,18.1333503723145,18.1056156158447],[18.4449214935303,18.4251956939697,18.4280281066895,18.468994140625,18.468994140625,18.44384765625,18.401611328125,18.3813972473145,18.2423343658447,18.1866703033447,18.1286125183105,18.0573253631592,17.974365234375,17.9494647979736,17.947265625,17.949951171875,17.9641609191895,17.9505863189697,17.9870109558105,17.9778804779053,17.986572265625,17.955078125,17.9478988647461,17.9537601470947,17.9678707122803,17.9666996002197,17.9941902160645,18.1525382995605,18.2242183685303,18.2833995819092,18.3206539154053,18.3645992279053,18.3936042785645,18.435791015625,18.4992198944092,18.5147953033447,18.5221691131592,18.4925289154053,18.4757804870605,18.4706535339355,18.4449214935303],[39.5362815856934,39.507568359375,39.5588874816895,39.6021003723145,39.6078109741211,39.5362815856934],[42.535400390625,42.4747047424316,42.4156265258789,42.3725090026855,42.3277816772461,42.3025360107422,42.2748565673828,42.2916984558105,42.3017082214355,42.214111328125,42.1832046508789,42.0969734191895,42.0457534790039,41.9911613464355,41.896728515625,41.8055191040039,41.7122573852539,41.594970703125,41.4943351745605,41.3915023803711,41.3038597106934,41.1236801147461,40.9322738647461,40.8573226928711,40.726318359375,40.6610336303711,40.4910659790039,40.4278793334961,40.3584938049316,40.3175277709961,40.1978988647461,40.1302223205566,40.0889663696289,40.0359382629395,40.0325698852539,39.99560546875,39.8959465026855,39.781982421875,39.6957054138184,39.5627899169922,39.4610824584961,39.3773918151855,39.4009780883789,39.3735847473145,39.296142578125,39.2079124450684,39.1794929504395,39.1432609558105,39.1250495910645,39.0841331481934,38.8979988098145,38.81640625,38.7861328125,38.7452163696289,38.680908203125,38.6234359741211,38.6078643798828,38.5237808227539,38.4169921875,38.3593254089355,38.3273429870605,38.3085441589355,38.3004379272461,38.3077125549316,38.3192405700684,38.3125,38.3049812316895,38.3132820129395,38.3045425415039,38.2838859558105,38.2405281066895,38.1755867004395,38.1060562133789,37.9789543151855,37.9171867370605,37.8279304504395,37.8007316589355,37.7818374633789,37.7901878356934,37.7968254089355,37.8783683776855,37.8285140991211,37.7637214660645,37.742919921875,37.8698234558105,37.8736839294434,37.9853515625,37.9626960754395,37.917724609375,37.8150367736816,37.730224609375,37.7190437316895,37.7248039245605,37.7482414245605,37.843505859375,37.882080078125,37.922607421875,37.9314460754395,38.0377960205078,38.0568389892578,38.08154296875,38.0936508178711,38.0778350830078,38.1126480102539,38.1015129089355,38.1291999816895,38.2333984375,38.2949714660645,38.3416481018066,38.4801292419434,38.5567398071289,38.6653785705566,38.680419921875,38.6761245727539,38.6862297058105,38.727783203125,38.7468757629395,38.7429656982422,38.8055191040039,38.8715324401855,39.2883796691895,39.3262672424316,39.4276351928711,39.526611328125,39.58349609375,39.59033203125,39.7018051147461,39.758056640625,39.7415046691895,39.6521987915039,39.6323738098145,39.6150856018066,39.7169418334961,39.7905731201172,39.8655242919922,39.9068832397461,39.9961433410645,40.0040512084961,40.104248046875,40.181640625,40.3192367553711,40.3837394714355,40.4598121643066,40.4581527709961,40.4647445678711,40.4978523254395,40.5238761901855,40.5474624633789,40.5894050598145,40.6446304321289,40.659912109375,40.7425765991211,40.7789535522461,40.7789535522461,40.7958984375,40.8386726379395,40.86669921875,40.8720245361328,40.8922348022461,40.9046363830566,40.9740715026855,41.023681640625,41.0782699584961,41.1377906799316,41.2256813049316,41.3213348388672,41.3518524169922,41.3580551147461,41.3939933776855,41.4955558776855,41.5943374633789,41.6409645080566,41.69189453125,41.716552734375,41.724853515625,41.7182121276855,41.7479972839355,41.7810554504395,41.7694816589355,41.7420425415039,41.6873512268066,41.643798828125,41.607421875,41.5545387268066,41.5313491821289,41.5198249816895,41.4837875366211,41.4817352294922,41.4547386169434,41.4399909973145,41.4611320495605,41.4486808776855,41.4156227111816,41.3892555236816,41.3877449035645,41.4330062866211,41.5065422058105,41.5627937316895,41.607421875,41.6553688049316,41.7000503540039,41.7691421508789,41.840576171875,41.86376953125,41.8984870910645,41.9516143798828,41.9874992370605,42.0116195678711,42.025634765625,42.0107421875,42.0208511352539,42.0406761169434,42.037841796875,42.0382347106934,42.0687980651855,42.1423835754395,42.1685066223145,42.2184562683105,42.2705535888672,42.3126945495605,42.3578605651855,42.41357421875,42.439208984375,42.4358901977539,42.4103050231934,42.3846664428711,42.39208984375,42.4358901977539,42.4442863464355,42.4481430053711,42.4749984741211,42.6038055419922,42.7054672241211,42.7765617370605,42.8942375183105,42.965087890625,42.9981460571289,42.9956550598145,42.974853515625,42.9625968933105,42.9741706848145,42.9560050964355,42.91796875,42.8917999267578,42.8726081848145,42.7448234558105,42.6849594116211,42.630859375,42.5816917419434,42.5705070495605,42.535400390625],[32.8686027526855,32.8158683776855,32.8414077758789,32.7735366821289,32.7580070495605,32.709716796875,32.6482887268066,32.6627960205078,32.7218742370605,32.766845703125,32.807373046875,32.8686027526855],[36.9599609375,36.9415512084961,36.9488792419434,36.943359375,36.9965324401855,37.0185546875,37.0240249633789,37.0001983642578,36.9599609375],[37.8409194946289,37.8340339660645,37.8486328125,37.837890625,37.7643547058105,37.7350082397461,37.71533203125,37.7628898620605,37.8260765075684,37.8724136352539,37.89404296875,37.9111328125,37.8409194946289],[38.4526824951172,38.4127464294434,38.4041481018066,38.3846702575684,38.4128913879395,38.4086418151855,38.4625511169434,38.5185546875,38.5530281066895,38.5533676147461,38.4526824951172],[38.5249977111816,38.5223617553711,38.5984382629395,38.6384773254395,38.6140594482422,38.5863265991211,38.5507316589355,38.5249977111816],[38.5556144714355,38.5435523986816,38.6205558776855,38.6553688049316,38.743896484375,38.6363258361816,38.5556144714355],[38.6434593200684,38.634033203125,38.6610374450684,38.6978492736816,38.7658195495605,38.7889671325684,38.8026885986328,38.7898445129395,38.7412109375,38.6789054870605,38.6434593200684],[39.4069328308105,39.3589324951172,39.3759765625,39.3940925598145,39.4967765808105,39.5208511352539,39.4794425964355,39.4069328308105],[37.179443359375,37.1683120727539,37.0057106018066,37.0054168701172,37.0770492553711,37.1000480651855,37.121337890625,37.0746078491211,37.07568359375,37.0160140991211,37.0322761535645,37.1660652160645,37.4308090209961,37.5924339294434,37.7328147888184,37.8718757629395,37.9586906433105,38.183837890625,38.2997550964355,38.4466819763184,38.42431640625,38.482421875,38.5181655883789,38.5099639892578,38.5121116638184,38.4552268981934,38.4381828308105,38.4480934143066,38.5389671325684,38.65673828125,38.6877937316895,38.6966781616211,38.7468757629395,38.8029289245605,38.9030265808105,38.9980964660645,39.0781707763672,39.0659675598145,39.0160675048828,38.8346710205078,38.7427711486816,38.7127914428711,38.6979026794434,38.70751953125,38.7308616638184,38.7987785339355,38.8529281616211,38.96044921875,39.1121101379395,39.2481460571289,39.2842788696289,39.3382797241211,39.39111328125,39.426025390625,39.5425758361816,39.820556640625,40.1156730651855,40.1518058776855,40.179443359375,40.2590827941895,40.6056632995605,40.6509246826172,40.7525405883789,40.91650390625,41.0294914245605,41.0862808227539,41.1544914245605,41.28466796875,41.5600090026855,41.6519508361816,41.6983909606934,41.7051773071289,41.7645988464355,41.8320808410645,41.9410629272461,42.0084953308105,42.052734375,42.0693359375,42.1150856018066,42.1374053955078,42.1336936950684,42.1118659973145,42.0693855285645,42.0399398803711,42.0181617736816,41.9270973205566,41.8958511352539,41.8369598388672,41.8199691772461,41.8119621276855,41.814208984375,41.8519058227539,41.8836402893066,41.8705558776855,41.8884773254395,41.8739738464355,41.8579559326172,41.8359870910645,41.8336944580078,41.8644065856934,41.895263671875,41.9293937683105,41.9552268981934,41.9716796875,41.9811515808105,41.9642066955566,41.9506378173828,41.9452629089355,41.9641609191895,41.95849609375,41.9345703125,41.9423828125,41.9130859375,41.8741226196289,41.78955078125,41.7040519714355,41.6725120544434,41.6644020080566,41.6653785705566,41.6421890258789,41.601806640625,41.5604476928711,41.5320320129395,41.5159187316895,41.4550323486328,41.3753890991211,41.3037109375,41.2145004272461,41.1077156066895,41.0624046325684,41.0380363464355,41.0091323852539,40.8783187866211,40.7774887084961,40.654052734375,40.6190910339355,40.483154296875,40.4432601928711,40.4109840393066,40.3762702941895,40.3431167602539,40.3007316589355,40.2516136169434,40.2083511352539,40.1679191589355,40.1426277160645,40.0568351745605,40.0218238830566,39.9371109008789,39.7983894348145,39.7139625549316,39.70556640625,39.6816864013672,39.6806640625,39.6615715026855,39.6447257995605,39.5361824035645,39.4783210754395,39.4651374816895,39.338134765625,39.1352043151855,39.1070823669434,39.0564460754395,38.9852561950684,38.9070320129395,38.8269500732422,38.7705078125,38.7145500183105,38.6493682861328,38.5668449401855,38.50146484375,38.4574203491211,38.1810073852539,38.1944351196289,38.1878929138184,38.1219749450684,38.0447273254395,38.030029296875,38.00634765625,37.9064483642578,37.786376953125,37.728271484375,37.5854988098145,37.5235824584961,37.4280281066895,37.179443359375],[-20.1646480560303,-20.2373046875,-20.29345703125,-20.3332042694092,-20.3861312866211,-20.4158191680908,-20.4654293060303,-20.5216789245605,-20.5944347381592,-20.6573257446289,-20.673828125,-20.6903324127197,-20.7474613189697,-20.7763671875,-20.8093757629395,-20.8416996002197,-20.873046875,-20.8970699310303,-20.9185543060303,-20.9979496002197,-21.1335945129395,-21.2062492370605,-21.2658195495605,-21.30224609375,-21.3550777435303,-21.41796875,-21.4940433502197,-21.5469722747803,-21.5965824127197,-21.6495132446289,-21.6991214752197,-21.751953125,-21.79833984375,-21.8511714935303,-21.9107418060303,-21.9669914245605,-22.0066413879395,-22.04638671875,-22.1091785430908,-22.1356449127197,-22.1422843933105,-22.1091785430908,-22.0992183685303,-22.1290035247803,-22.1819343566895,-22.1885738372803,-22.1984367370605,-22.2150402069092,-22.1952152252197,-22.2150402069092,-22.2448253631592,-22.2712879180908,-22.2646484375,-22.2613277435303,-22.2315425872803,-22.23486328125,-22.1819343566895,-22.1356449127197,-22.1025390625,-22.076171875,-22.0926761627197,-22.1786136627197,-22.2282218933105,-22.2646484375,-22.2811527252197,-22.2844715118408,-22.2811527252197,-22.3076171875,-22.3076171875,-22.3539066314697,-22.41015625,-22.5126953125,-22.5920906066895,-22.6218757629395,-22.6714839935303,-22.7409191131592,-22.8103523254395,-22.8864250183105,-22.9558601379395,-23.0252933502197,-23.0947265625,-23.154296875,-23.2501964569092,-23.3196277618408,-23.359375,-23.4156246185303,-23.4619140625,-23.5247058868408,-23.5809574127197,-23.6272468566895,-23.6867179870605,-23.7925777435303,-23.8653316497803,-23.9513683319092,-23.9910163879395,-24.0042972564697,-24.0174808502197,-23.9976558685303,-23.9745121002197,-23.9513683319092,-23.8884773254395,-23.8521480560303,-23.8290042877197,-23.8125,-23.8521480560303,-23.9017562866211,-23.97119140625,-24.0472660064697,-24.0658206939697,-24.1281242370605,-24.2012691497803,-24.3060550689697,-24.5281238555908,-24.8674812316895,-25.0652351379395,-25.1212882995605,-25.22021484375,-25.4327144622803,-25.5760746002197,-26.0057621002197,-26.3087882995605,-26.5329093933105,-26.6522464752197,-26.6667976379395,-26.7025394439697,-26.7593746185303,-26.806640625,-26.8445320129395,-26.8860359191895,-26.93115234375,-26.9601573944092,-26.97314453125,-27.00927734375,-27.068359375,-27.1153316497803,-27.1499996185303,-27.2076168060303,-27.2880859375,-27.3571300506592,-27.4148445129395,-27.4164066314697,-27.3619136810303,-27.32568359375,-27.3077144622803,-27.3214836120605,-27.3667964935303,-27.4387683868408,-27.5374011993408,-27.5538082122803,-27.4878921508789,-27.4678707122803,-27.4937496185303,-27.4846668243408,-27.4406242370605,-27.4357414245605,-27.4701175689697,-27.4304676055908,-27.3166027069092,-27.2734375,-27.3143558502197,-27.1960945129395,-27.1321296691895,-27.083984375,-27.0294914245605,-26.9684581756592,-26.9219722747803,-26.8900394439697,-26.8576164245605,-26.8249015808105,-26.7958984375,-26.7707023620605,-26.7310543060303,-26.6768550872803,-26.6499996185303,-26.6299800872803,-26.5925788879395,-26.4765625,-26.3814449310303,-26.3074207305908,-26.2625980377197,-26.25146484375,-26.2249031066895,-26.18017578125,-26.1385746002197,-26.0529308319092,-26.0065422058105,-25.9642562866211,-25.9069328308105,-25.78369140625,-25.7259750366211,-25.6970710754395,-25.6671867370605,-25.5987319946289,-25.5341815948486,-25.4737300872803,-25.4050788879395,-25.3284187316895,-25.1364269256592,-25.0492191314697,-24.9771480560303,-24.9538097381592,-24.9791011810303,-24.9592761993408,-24.8941402435303,-24.8428707122803,-24.78662109375,-24.5623054504395,-24.4539051055908,-24.3870105743408,-24.2667961120605,-24.0935554504395,-24.0091800689697,-24.0139656066895,-23.9635734558105,-23.8581047058105,-23.7556648254395,-23.6564464569092,-23.5570316314697,-23.45751953125,-23.3919925689697,-23.3604488372803,-23.3194332122803,-23.2687511444092,-23.1820316314697,-23.0592765808105,-22.8694343566895,-22.6124019622803,-22.4391613006592,-22.349609375,-22.2904300689697,-22.2615222930908,-22.2336921691895,-22.1839847564697,-21.9886722564697,-21.7052745819092,-21.4117183685303,-21.0660152435303,-20.82080078125,-20.5625,-20.3499031066895,-20.1990222930908,-20.0553722381592,-19.8094730377197,-19.6453132629395,-19.6064453125,-19.52099609375,-19.478515625,-19.3887691497803,-19.2975597381592,-19.2917957305908,-19.2862300872803,-19.490234375,-19.6460933685303,-19.81787109375,-19.8548831939697,-19.9988288879395,-20.1646480560303],[31.2083015441895,31.2922859191895,31.3226089477539,31.4837875366211,31.5848636627197,31.5416488647461,31.5256366729736,31.3627433776855,31.2929210662842,31.2083015441895],[31.4792976379395,31.48291015625,31.4228038787842,31.3662090301514,31.3513202667236,31.3681621551514,31.3968734741211,31.5365715026855,31.602294921875,31.6732425689697,31.7344722747803,31.7500019073486,31.7763175964355,31.8167495727539,31.8379402160645,31.8167972564697,31.8233394622803,31.8412590026855,31.8664054870605,31.9132804870605,31.9916019439697,32.0871086120605,32.1609344482422,32.2810554504395,32.3381843566895,32.46044921875,32.534423828125,32.512939453125,32.5074729919434,32.4930152893066,32.4506340026855,32.4016609191895,32.3955078125,32.2378921508789,32.10302734375,31.9849109649658,31.7655258178711,31.67236328125,31.5623512268066,31.4792976379395],[-20.8572273254395,-20.8758792877197,-20.8505859375,-20.7951164245605,-20.787109375,-20.7711906433105,-20.8133792877197,-20.8572273254395],[-18.5443363189697,-18.5663089752197,-18.5452156066895,-18.5348644256592,-18.513671875,-18.4890632629395,-18.4850578308105,-18.4708003997803,-18.4631824493408,-18.4743156433105,-18.4967765808105,-18.5193347930908,-18.5443363189697],[-18.3798828125,-18.4161128997803,-18.3991203308105,-18.36376953125,-18.3341789245605,-18.3798828125],[-18.3419933319092,-18.3609371185303,-18.2653312683105,-18.2728519439697,-18.3419933319092],[-18.1893558502197,-18.2168941497803,-18.19873046875,-18.14794921875,-18.0850582122803,-18.0591812133789,-18.0837898254395,-18.1205062866211,-18.1893558502197],[-17.8566398620605,-17.8756828308105,-17.8731460571289,-17.8666019439697,-17.8310546875,-17.7879886627197,-17.7516593933105,-17.7177734375,-17.6837882995605,-17.6697273254395,-17.6780261993408,-17.68310546875,-17.7541007995605,-17.8037109375,-17.8384761810303,-17.8492183685303,-17.8566398620605],[-17.6900405883789,-17.7366218566895,-17.8121089935303,-17.8623046875,-17.8499011993408,-17.8224601745605,-17.732421875,-17.7527332305908,-17.7349605560303,-17.6175785064697,-17.5642585754395,-17.5316410064697,-17.4963874816895,-17.5223636627197,-17.5889644622803,-17.6900405883789],[-17.5450191497803,-17.5710945129395,-17.5528316497803,-17.5277347564697,-17.5011711120605,-17.4695320129395,-17.4739265441895,-17.48779296875,-17.5450191497803],[-16.8777332305908,-16.8792972564697,-16.8636722564697,-16.7607421875,-16.7396488189697,-16.7744140625,-16.8642578125,-16.8777332305908],[-16.6197261810303,-16.6688480377197,-16.6354503631592,-16.62109375,-16.6123046875,-16.6135749816895,-16.6197261810303],[-16.634765625,-16.6404285430908,-16.5808601379395,-16.5808601379395,-16.6180648803711,-16.634765625],[-16.6575202941895,-16.6651382446289,-16.64697265625,-16.6190433502197,-16.5740222930908,-16.6037101745605,-16.6234378814697,-16.6575202941895],[-16.3297843933105,-16.3467769622803,-16.3458003997803,-16.2951164245605,-16.2511711120605,-16.2014656066895,-16.1652336120605,-16.1318359375,-16.0791988372803,-16.1598644256592,-16.224609375,-16.27783203125,-16.3297843933105],[-16.0962886810303,-16.1071281433105,-16.0277347564697,-16.0177745819092,-16.0962886810303],[-15.8560552597046,-15.9010744094849,-15.8576173782349,-15.7880868911743,-15.7570314407349,-15.7620124816895,-15.8560552597046],[-10.5153322219849,-10.5324211120605,-10.4255867004395,-10.4458980560303,-10.4629878997803,-10.4921875,-10.5153322219849],[-9.93134784698486,-10.0095701217651,-9.92626953125,-9.91542911529541,-9.91542911529541,-9.93134784698486],[-9.69521427154541,-9.74716758728027,-9.74160194396973,-9.79287052154541,-9.82070255279541,-9.845703125,-9.82949161529541,-9.77021503448486,-9.69521427154541],[-9.42597675323486,-9.44414138793945,-9.38437557220459,-9.359375,-9.328125,-9.3447265625,-9.42597675323486],[-8.94023418426514,-8.94794940948486,-8.89853477478027,-8.87236309051514,-8.86005783081055,-8.87539005279541,-8.89706993103027,-8.91562557220459,-8.94023418426514],[-8.91044998168945,-8.93398380279541,-8.9296875,-8.84804630279541,-8.79755878448486,-8.78154373168945,-8.80146503448486,-8.83847618103027,-8.87363338470459,-8.91044998168945],[24.6072273254395,24.5867195129395,24.5646495819092,24.5652351379395,24.5739269256592,24.5951175689697,24.6796379089355,24.7892589569092,24.8503894805908,24.8885746002197,25.1774425506592,25.3992671966553,25.4447269439697,25.4970703125,25.6126956939697,25.7240715026855,25.9814453125,26.08056640625,26.1532726287842,26.0111331939697,25.9023933410645,25.781005859375,25.68212890625,25.5247077941895,25.4523429870605,25.3897457122803,25.2844715118408,25.14794921875,25.0528812408447,24.96484375,24.890869140625,24.6682605743408,24.6451168060303,24.6072273254395],[45.450439453125,45.3471183776855,45.2921867370605,45.234130859375,45.240966796875,45.2519073486328,45.2668952941895,45.2862319946289,45.3098640441895,45.3110809326172,45.2899436950684,45.320556640625,45.4029312133789,45.419677734375,45.3708038330078,45.3139152526855,45.2599143981934,45.1932144165039,45.1516571044922,44.9796371459961,44.9154777526855,44.8434104919434,44.757568359375,44.798828125,44.8711433410645,44.9256858825684,44.9750480651855,44.992919921875,44.9658203125,44.9186553955078,44.8100090026855,44.7499504089355,44.71630859375,44.6368637084961,44.6024894714355,44.5650367736816,44.5747566223145,44.506103515625,44.3742179870605,44.295654296875,43.9838371276855,43.7974128723145,43.7422370910645,43.740478515625,43.7447776794434,43.7728500366211,43.8224105834961,43.9186058044434,43.9873542785645,43.956298828125,43.964599609375,43.997802734375,44.0200653076172,44.0205078125,44.1461410522461,44.1673851013184,44.1461906433105,44.083984375,44.0072746276855,43.8705558776855,43.766845703125,43.7117652893066,43.6708030700684,43.6863288879395,43.7384262084961,43.7943840026855,43.7634773254395,43.7866706848145,43.8535652160645,43.8738784790039,43.83447265625,43.8645515441895,43.8990211486816,43.9478988647461,43.9872055053711,44.0169944763184,44.0472183227539,44.0779762268066,44.1272926330566,44.1952133178711,44.23779296875,44.248291015625,44.2864761352539,44.3164558410645,44.3383293151855,44.3779792785645,44.4354476928711,44.4895973205566,44.5403327941895,44.5623550415039,44.5555152893066,44.5699195861816,44.6055183410645,44.6509742736816,44.7062492370605,44.6761245727539,44.5606918334961,44.5419464111328,44.619873046875,44.6661148071289,44.6806640625,44.71044921875,44.7554206848145,44.7900886535645,44.82666015625,44.86181640625,44.8700675964355,44.8733901977539,44.8808097839355,44.9006843566895,44.9188499450684,44.9419937133789,44.9577140808105,44.9734382629395,44.9907684326172,45.0089836120605,45.0213851928711,45.022216796875,45.032958984375,45.0751457214355,45.10986328125,45.1222648620605,45.1478996276855,45.171875,45.1925315856934,45.205078125,45.2413101196289,45.291748046875,45.2935562133789,45.321533203125,45.3653297424316,45.4275398254395,45.4678688049316,45.4844207763672,45.5000991821289,45.5174789428711,45.5364761352539,45.5974617004395,45.6620101928711,45.7225074768066,45.7498054504395,45.7581062316895,45.7489776611328,45.7374038696289,45.7352523803711,45.7793922424316,45.8694801330566,45.8995094299316,45.9407691955566,45.9754905700684,46.0506820678711,46.1085929870605,46.1330070495605,46.1669425964355,46.1334953308105,46.1456527709961,46.1728019714355,46.1944351196289,46.2174797058105,46.2462425231934,46.2597160339355,46.2422332763672,46.2824211120605,46.3043441772461,46.3526878356934,46.3915519714355,46.4123039245605,46.44775390625,46.4863777160645,46.5724601745605,46.6078147888184,46.6207504272461,46.6478538513184,46.7042961120605,46.7533683776855,46.7897453308105,46.8783683776855,46.9637680053711,47.0065422058105,47.0438995361328,47.0848121643066,47.1381340026855,47.3045883178711,47.3325691223145,47.3642578125,47.3957023620605,47.5050277709961,47.53662109375,47.572021484375,47.6290550231934,47.6963844299316,47.7278327941895,47.7362289428711,47.7626457214355,47.7725601196289,47.7595710754395,47.7663116455078,47.7990226745605,47.9225578308105,47.947265625,47.9642601013184,48.0065460205078,48.0491218566895,48.08740234375,48.0845222473145,47.989990234375,47.995849609375,47.9923362731934,47.9903793334961,47.9826202392578,47.935791015625,47.9410171508789,47.9447746276855,47.9060554504395,47.9111824035645,47.9380378723145,47.9471168518066,47.9310531616211,47.8765144348145,47.7608375549316,47.7177734375,47.72412109375,47.7457046508789,47.8230972290039,47.9107933044434,47.9324722290039,47.9675788879395,47.9925308227539,48.0643539428711,48.1132316589355,48.2037582397461,48.22998046875,48.2484855651855,48.2598609924316,48.2634773254395,48.2558135986328,48.2111320495605,48.155029296875,48.1104965209961,48.0476570129395,47.9592781066895,47.8417510986328,47.7822265625,47.7179679870605,47.6397476196289,47.5531272888184,47.5366668701172,47.4756355285645,47.3405265808105,47.2864227294922,47.2275886535645,47.1683082580566,47.114501953125,47.043212890625,46.9784164428711,46.7920875549316,46.7063980102539,46.6408195495605,46.508056640625,46.4512710571289,46.3475608825684,46.138671875,45.9726066589355,45.883056640625,45.8255348205566,45.7888679504395,45.7130851745605,45.6471176147461,45.6282730102539,45.6127433776855,45.5989761352539,45.5691413879395,45.5137672424316,45.450439453125],[43.4093246459961,43.4071311950684,43.4203643798828,43.43359375,43.4490242004395,43.440185546875,43.42919921875,43.4214820861816,43.4093246459961],[43.4595222473145,43.4545402526855,43.4553718566895,43.45703125,43.4454612731934,43.4264640808105,43.4256362915039,43.4223175048828,43.4198226928711,43.4330558776855,43.4437980651855,43.450439453125,43.4578590393066,43.4595222473145],[43.6253929138184,43.6199188232422,43.6253929138184,43.6298332214355,43.6386222839355,43.65185546875,43.6474609375,43.6441421508789,43.6253929138184],[43.7438011169434,43.7163543701172,43.740478515625,43.7970199584961,43.81298828125,43.8604965209961,43.8291511535645,43.8041496276855,43.7438011169434],[44.4976539611816,44.4246101379395,44.4404296875,44.3746604919434,44.3756828308105,44.2809562683105,44.2686500549316,44.2459487915039,44.1037101745605,44.0477523803711,43.9407234191895,43.8451194763184,43.6646003723145,43.737060546875,43.8103523254395,43.8708992004395,43.9990730285645,44.071533203125,44.1290054321289,44.1930160522461,44.2485847473145,44.2726593017578,44.5001487731934,44.4976539611816],[45.317626953125,45.2471694946289,45.2168464660645,45.0701637268066,44.9903793334961,44.9585914611816,44.9771461486816,44.9447288513184,44.8865737915039,44.8355484008789,44.6776351928711,44.5535621643066,44.53125,44.404296875,44.5130882263184,44.5657196044922,44.663330078125,44.7662124633789,44.8560562133789,44.9452171325684,45.0624504089355,45.093017578125,45.1907234191895,45.2256317138672,45.30029296875,45.38330078125,45.3777351379395,45.2621116638184,45.2582015991211,45.2824211120605,45.4846687316895,45.5206527709961,45.5264663696289,45.510009765625,45.486083984375,45.4559097290039,45.4135246276855,45.3626937866211,45.323974609375,45.317626953125],[45.4882316589355,45.469970703125,45.5620613098145,45.6181640625,45.6470680236816,45.5540542602539,45.4882316589355],[45.6420440673828,45.5913581848145,45.5933609008789,45.8397941589355,45.8760757446289,46.0219230651855,46.2003402709961,46.2134284973145,46.2085456848145,46.0123023986816,45.9332046508789,45.849365234375,45.8224601745605,45.7831535339355,45.6420440673828],[46.8971672058105,46.787109375,46.788330078125,46.8288078308105,46.8526878356934,46.8689918518066,47.0149879455566,47.1104469299316,47.1434059143066,47.1421890258789,46.8971672058105],[47.762939453125,47.7061042785645,47.7134742736816,47.7279319763184,47.7970199584961,47.8087387084961,47.762939453125],[48.790283203125,48.73876953125,48.7557144165039,48.7725105285645,48.8321304321289,48.9044456481934,48.9049339294434,48.89208984375,48.857177734375,48.790283203125],[49.31201171875,49.2676773071289,49.2940406799316,49.380615234375,49.6469230651855,49.63037109375,49.56640625,49.46826171875,49.347900390625,49.31201171875],[50.3022003173828,50.2020492553711,50.17724609375,50.1456069946289,50.0777816772461,50.041259765625,50.0611801147461,50.0946273803711,50.2127914428711,50.2645492553711,50.2978515625,50.2932624816895,50.3689460754395,50.4007339477539,50.482421875,50.6841278076172,50.7569313049316,50.7718772888184,50.6712875366211,50.5592765808105,50.4517593383789,50.3022003173828],[50.6576194763184,50.6337890625,50.6390647888184,50.7021484375,50.7318840026855,50.7847175598145,50.8621101379395,50.8595695495605,50.8429679870605,50.751220703125,50.6576194763184],[50.8219261169434,50.8105964660645,50.8373069763184,50.86962890625,50.91357421875,50.9344711303711,50.9104995727539,50.8453636169434,50.8219261169434],[54.3939476013184,54.1409645080566,54.085693359375,54.02880859375,53.9556159973145,53.8783683776855,53.8109397888184,53.7942390441895,53.4886741638184,53.2960433959961,53.21728515625,53.1343727111816,52.9630851745605,52.7000503540039,52.6135749816895,52.5291519165039,52.4786605834961,52.4429206848145,52.349365234375,52.083740234375,51.9444847106934,51.847900390625,51.7443351745605,51.6323738098145,51.583251953125,51.5206069946289,51.4714813232422,51.4019050598145,51.2992172241211,51.2770500183105,51.2462882995605,50.5067405700684,50.2826156616211,49.895751953125,49.6614761352539,49.5497550964355,49.4320335388184,49.3113288879395,49.1805191040039,49.0510749816895,48.9358406066895,48.8712387084961,48.8195304870605,48.6402854919434,48.6785621643066,48.8148422241211,48.8935546875,48.9863777160645,49.0697746276855,49.2085456848145,49.2491722106934,49.2763175964355,49.30859375,49.31201171875,49.2906761169434,49.2628402709961,49.1988296508789,49.1054191589355,48.9177742004395,48.2468757629395,48.0721702575684,47.8849143981934,47.7378883361816,47.6839866638184,47.5369148254395,47.452392578125,47.4161643981934,47.3917961120605,47.3618659973145,47.32275390625,47.2227058410645,47.0007820129395,46.8440437316895,46.7948722839355,46.807373046875,46.8056640625,46.7919921875,46.7520523071289,46.5750961303711,46.4060554504395,46.3606948852539,46.2301750183105,46.1746101379395,46.1158180236816,46.0694808959961,46.0286636352539,46.2220230102539,46.3584938049316,46.4762191772461,46.5589866638184,46.5926284790039,46.6052742004395,46.6202125549316,46.670654296875,46.7108421325684,46.7162094116211,46.7007789611816,46.6442413330078,46.5546875,46.4586906433105,46.3575706481934,46.0888671875,45.999267578125,45.9170379638672,45.9616203308105,46.0134773254395,46.0882835388184,46.1707496643066,46.4510765075684,46.6941871643066,47.0303230285645,47.1402816772461,47.2446784973145,47.3477058410645,47.5438003540039,47.5874519348145,47.7006340026855,47.808349609375,47.9021492004395,48.0133781433105,48.2900848388672,48.4771003723145,48.6546897888184,48.7019538879395,48.7500953674316,48.97216796875,49.0784683227539,49.3120613098145,49.4396476745605,49.5691413879395,50.2167510986328,50.3121109008789,50.5149879455566,50.6304664611816,50.7764625549316,50.8901863098145,50.9984855651855,51.2225608825684,51.4293937683105,51.5205078125,51.6300277709961,51.690185546875,51.736328125,51.7518081665039,51.7892074584961,51.8467788696289,51.933349609375,52.27294921875,52.359130859375,52.454833984375,52.5556144714355,52.7935066223145,53.0389137268066,53.1384773254395,53.3395004272461,53.3894538879395,53.4563980102539,53.49560546875,53.4840316772461,53.4054679870605,53.4025421142578,53.4107437133789,53.4474601745605,53.5367660522461,53.5875968933105,53.6526336669922,53.6743659973145,53.7301750183105,53.7367668151855,53.8160171508789,53.8957061767578,53.9684066772461,54.1485328674316,54.2807121276855,54.2789573669434,54.3036117553711,54.3582038879395,54.4161148071289,54.3939476013184],[54.5649909973145,54.5127449035645,54.6976585388184,54.7949714660645,54.8558578491211,54.85693359375,54.7977523803711,54.7701683044434,54.6904792785645,54.5649909973145],[55.1004409790039,54.9267578125,54.9239768981934,54.9606437683105,54.946044921875,54.9561538696289,55.0093727111816,55.0927276611328,55.0917472839355,55.1078147888184,55.1004409790039],[55.0926284790039,55.0533180236816,55.06005859375,55.0335464477539,54.9909172058105,54.90087890625,54.8207015991211,54.7890129089355,54.7495613098145,54.6969223022461,54.6632347106934,54.6532707214355,54.8258285522461,54.8733863830566,54.7923812866211,54.7905769348145,54.8910179138184,55.0006828308105,55.0160179138184,55.1630859375,55.197021484375,55.1100578308105,55.0926284790039],[55.2641105651855,55.2644538879395,55.2755355834961,55.2344245910645,55.1953125,55.1603507995605,55.1007308959961,55.0636711120605,55.0593757629395,55.0642585754395,55.0591316223145,54.970703125,54.9288558959961,54.8712882995605,54.8384742736816,54.6326179504395,54.5629386901855,54.4930191040039,54.4056167602539,54.3567848205566,54.35009765625,54.35986328125,54.37646484375,54.3917999267578,54.4066390991211,54.4207496643066,54.4339866638184,54.4470710754395,54.4591789245605,54.5448265075684,54.6338348388672,54.75,54.8304672241211,54.9211921691895,54.9564933776855,54.9512710571289,54.994873046875,55.1026382446289,55.2320327758789,55.2866706848145,55.2789077758789,55.1836433410645,54.9823722839355,54.9556655883789,54.947021484375,54.9094734191895,54.9026870727539,54.9352035522461,55.1077613830566,55.2641105651855],[54.8390617370605,54.694091796875,54.7676277160645,54.8268585205078,54.8380851745605,54.8645515441895,54.9365234375,55.0303688049316,55.0765609741211,55.1356964111328,55.1904792785645,55.2945289611816,55.3069343566895,55.3514633178711,55.323974609375,55.3119621276855,55.2423324584961,55.1654281616211,55.005615234375,54.9499015808105,54.90771484375,54.8390617370605],[59.0187492370605,59.0074234008789,59.0347671508789,59.0540542602539,59.0972137451172,59.16015625,59.1224594116211,59.09521484375,59.0187492370605],[58.6033706665039,58.5093727111816,58.5246620178223,58.546142578125,58.5789527893066,58.640869140625,58.7985382080078,58.9297370910645,58.9723663330078,59.0150413513184,59.09619140625,59.2267570495605,59.2211418151855,59.1122055053711,58.9707527160645,58.8855934143066,58.8380851745605,58.7437515258789,58.6033706665039],[65.1820831298828,65.1426773071289,65.0779266357422,65.0364761352539,65.00146484375,64.9766616821289,65.0576171875,65.0587844848633,65.0936050415039,65.1510772705078,65.1670913696289,65.1571273803711,65.1975555419922,65.1820831298828],[66.502197265625,66.48974609375,66.5653305053711,66.7159729003906,66.7510757446289,66.739013671875,66.7364730834961,66.711669921875,66.6958923339844,66.627197265625,66.5994567871094,66.569091796875,66.517578125,66.502197265625],[66.7017059326172,66.6880874633789,66.7350616455078,66.7703552246094,66.7855453491211,66.7955093383789,66.7822265625,66.7353057861328,66.7017059326172],[69.1855926513672,69.0888671875,69.0487823486328,69.0375518798828,69.09814453125,69.1255340576172,69.0532760620117,68.9737777709961,68.8597183227539,68.7784194946289,68.7430648803711,68.733154296875,68.8048782348633,68.9423828125,68.9842300415039,69.0403289794922,69.0966339111328,69.1838912963867,69.2692413330078,69.3456573486328,69.43603515625,69.4947280883789,69.50927734375,69.51123046875,69.3094253540039,69.257080078125,69.1855926513672],[69.5298385620117,69.4425277709961,69.4471740722656,69.467041015625,69.4832000732422,69.5753860473633,69.5721206665039,69.5298385620117],[68.9009780883789,68.899658203125,68.9660186767578,68.99560546875,69.04443359375,69.0815887451172,69.1102523803711,69.197021484375,69.3335952758789,69.4056625366211,69.4698257446289,69.5409164428711,69.595703125,69.6394577026367,69.634033203125,69.5924301147461,69.5009231567383,69.4365234375,69.4136657714844,69.3693313598633,69.2928237915039,69.1944351196289,69.1064453125,69.0160140991211,68.9695816040039,68.9275817871094,68.9009780883789],[69.5804672241211,69.5714340209961,69.6643600463867,69.7133331298828,69.7758331298828,69.8196258544922,69.8368682861328,69.873046875,69.9010696411133,69.9749069213867,70.0083999633789,70.0156784057617,69.8826141357422,69.8560562133789,69.8321762084961,69.7791976928711,69.7695846557617,69.7347640991211,69.6287078857422,69.6011199951172,69.5804672241211],[69.9348602294922,69.885498046875,69.793701171875,69.7259216308594,69.7152862548828,69.6876983642578,69.717041015625,69.6969680786133,69.6956558227539,69.7061996459961,69.7210464477539,69.7386245727539,69.7908630371094,69.8662109375,69.8904266357422,69.8984375,69.9219207763672,69.9107894897461,69.88330078125,69.8927688598633,70.051025390625,70.0880355834961,70.1291961669922,70.1556701660156,70.266845703125,70.3183059692383,70.350830078125,70.3595657348633,70.4329605102539,70.4651870727539,70.4604949951172,70.4371109008789,70.3616714477539,70.3109359741211,70.2489700317383,70.197021484375,70.1083450317383,70.0228500366211,69.96240234375,69.9348602294922],[70.541748046875,70.50146484375,70.514892578125,70.56103515625,70.6157684326172,70.727294921875,70.7571258544922,70.7726593017578,70.7693405151367,70.6987838745117,70.541748046875],[70.8227005004883,70.8058624267578,70.8197250366211,70.8510284423828,70.9226531982422,70.9340362548828,70.9237823486328,70.883544921875,70.8227005004883],[71.364990234375,71.291259765625,71.2379379272461,71.2418899536133,71.2152786254883,71.1597213745117,71.0650405883789,71.0309524536133,70.999267578125,70.9820327758789,70.9686965942383,71.0116195678711,71.0536117553711,71.0858459472656,71.1149368286133,71.1806640625,71.2504348754883,71.2392578125,71.2703628540039,71.284912109375,71.3250503540039,71.3568344116211,71.3850631713867,71.3833541870117,71.3551330566406,71.3897933959961,71.4037094116211,71.3998031616211,71.364990234375],[70.826416015625,70.8220672607422,71.0005874633789,71.04736328125,71.1056671142578,71.1778793334961,71.2311096191406,71.3245086669922,71.4476547241211,71.4662094116211,71.5233383178711,71.5377426147461,71.5509796142578,71.5779800415039,71.5824279785156,71.566650390625,71.5716781616211,71.59619140625,71.5932159423828,71.5770492553711,71.5411605834961,71.5291976928711,71.4816360473633,71.4654769897461,71.4375991821289,71.3905334472656,71.3399887084961,71.2816925048828,71.2630844116211,71.2191390991211,71.1668930053711,71.0675811767578,71.0419464111328,71.0147933959961,70.9398422241211,70.9189910888672,70.9234313964844,70.9716796875,70.9930191040039,70.9756851196289,70.89892578125,70.8802719116211,70.826416015625],[71.5076675415039,71.4232406616211,71.4339370727539,71.474609375,71.4834976196289,71.477294921875,71.4605484008789,71.4559097290039,71.502197265625,71.5298767089844,71.542724609375,71.55615234375,71.55712890625,71.5799331665039,71.587890625,71.5830612182617,71.5427780151367,71.5076675415039],[72.291259765625,72.2818832397461,72.2976531982422,72.3170394897461,72.3655776977539,72.439208984375,72.4861373901367,72.5652847290039,72.630859375,72.6312026977539,72.55322265625,72.5042953491211,72.482421875,72.4169921875,72.3924789428711,72.3085403442383,72.291259765625],[72.721923828125,72.710693359375,72.7516098022461,72.8034133911133,72.8507308959961,72.8922805786133,73.0943374633789,73.0386199951172,72.9831085205078,72.9186553955078,72.7693328857422,72.721923828125],[72.8734359741211,72.86376953125,72.8811492919922,72.9076690673828,72.975341796875,73.0215301513672,73.0743637084961,73.1090850830078,73.1304702758789,73.1217803955078,73.108154296875,73.0625,73.03271484375,72.9690399169922,72.9292984008789,72.90771484375,72.8734359741211],[73.08984375,73.0448684692383,73.04541015625,73.1243209838867,73.1554718017578,73.1676788330078,73.15673828125,73.1072769165039,73.08984375],[73.3083038330078,73.2686004638672,73.2561569213867,73.2255325317383,73.201171875,73.13916015625,73.1136703491211,73.0330047607422,72.9832000732422,72.9049835205078,72.8480911254883,72.8065948486328,72.7894058227539,72.7895050048828,72.7664031982422,72.7215347290039,72.6717300415039,72.5990676879883,72.5753936767578,72.549072265625,72.5013198852539,72.465087890625,72.40869140625,72.3778381347656,72.3136215209961,72.2206497192383,72.1823272705078,72.1068878173828,72.014892578125,71.9353561401367,71.8692398071289,71.7833480834961,71.6898880004883,71.507568359375,71.3456115722656,71.1073684692383,70.927001953125,70.8760223388672,70.7922897338867,70.7658233642578,70.7288055419922,70.6610336303711,70.6360321044922,70.6051254272461,70.589111328125,70.6465301513672,70.6553726196289,70.6974639892578,70.7288055419922,70.734130859375,70.7147445678711,70.6767044067383,70.6649398803711,70.5935592651367,70.56640625,70.56298828125,70.6578598022461,70.6461410522461,70.6183547973633,70.6492691040039,70.6263198852539,70.6155776977539,70.6418914794922,70.6752471923828,70.6921844482422,70.666015625,70.666748046875,70.6781311035156,70.7418441772461,70.7132339477539,70.6800765991211,70.6933135986328,70.7446746826172,70.764892578125,70.814453125,70.87353515625,70.9005889892578,70.9146499633789,70.9508285522461,71.0006866455078,71.0522918701172,71.0869140625,71.0704116821289,71.126708984375,71.1375961303711,71.105224609375,71.12548828125,71.1962890625,71.2966766357422,71.332763671875,71.342529296875,71.3401336669922,71.399169921875,71.4772491455078,71.5301284790039,71.5416488647461,71.4950180053711,71.5056686401367,71.536865234375,71.490234375,71.4747085571289,71.4913101196289,71.525146484375,71.5711441040039,71.6480941772461,71.7768020629883,71.8255386352539,71.9343795776367,71.9797821044922,72.0711898803711,72.099365234375,72.1421356201172,72.1532211303711,72.1311569213867,72.1297378540039,72.1539535522461,72.1967239379883,72.2523422241211,72.2840347290039,72.3009719848633,72.3368682861328,72.3909683227539,72.4369659423828,72.4829559326172,72.5498504638672,72.5912551879883,72.6192855834961,72.6688995361328,72.6823272705078,72.7040481567383,72.7373580932617,72.7685546875,72.7913513183594,72.875244140625,72.8999481201172,72.91357421875,72.9132308959961,72.9037551879883,72.916748046875,72.97314453125,73.0111846923828,73.10400390625,73.1475524902344,73.1829528808594,73.2245559692383,73.2383804321289,73.26025390625,73.2932662963867,73.2989730834961,73.2764587402344,73.2813491821289,73.2994689941406,73.3700256347656,73.3876419067383,73.3832550048828,73.3568420410156,73.3083038330078],[73.0950164794922,73.048095703125,73.0444793701172,73.0562973022461,73.0371627807617,73.0845184326172,73.1266174316406,73.1692428588867,73.2243118286133,73.359375,73.4447250366211,73.4776382446289,73.514404296875,73.5042037963867,73.4779815673828,73.44775390625,73.3722152709961,73.3420944213867,73.2831573486328,73.2248077392578,73.1739730834961,73.1624526977539,73.11962890625,73.0950164794922],[73.44580078125,73.4395065307617,73.4762191772461,73.5234909057617,73.5542907714844,73.5552673339844,73.44580078125],[73.4566421508789,73.4322738647461,73.4773941040039,73.5406265258789,73.5736312866211,73.5599136352539,73.5492630004883,73.4814987182617,73.4591293334961,73.4566421508789],[73.8958969116211,73.8515625,73.8030776977539,73.5687561035156,73.5208435058594,73.4588928222656,73.2464370727539,73.2313003540039,73.2207565307617,73.2448272705078,73.2533187866211,73.2528839111328,73.2816848754883,73.3108367919922,73.3892059326172,73.446044921875,73.4520034790039,73.4353561401367,73.3614196777344,73.355224609375,73.355224609375,73.4257354736328,73.4485855102539,73.45751953125,73.4830093383789,73.5645523071289,73.629150390625,73.7775344848633,73.83154296875,73.8658676147461,73.87646484375,73.8718795776367,73.9041976928711,73.9149398803711,73.8958969116211],[73.85009765625,73.847900390625,73.8746109008789,73.9103012084961,73.9283676147461,73.9430236816406,73.933837890625,73.900390625,73.8880386352539,73.85009765625],[74.07177734375,74.0484390258789,74.0516586303711,74.03955078125,74.0753402709961,74.0916061401367,74.12890625,74.151611328125,74.1223602294922,74.0894546508789,74.07177734375],[74.0908660888672,74.0564422607422,74.0757751464844,74.0992202758789,74.131103515625,74.1492614746094,74.1614303588867,74.1485366821289,74.1112289428711,74.0908660888672],[73.9136199951172,73.885009765625,73.9291000366211,74.0595245361328,74.1214370727539,74.1796875,74.2017059326172,74.2533721923828,74.2199249267578,74.09033203125,73.9849624633789,73.9136199951172],[73.9994659423828,73.9186477661133,73.9216842651367,74.0045852661133,74.1842727661133,74.2367172241211,74.2572326660156,74.2664566040039,74.2737808227539,74.2646484375,74.2427215576172,74.2093276977539,74.1678237915039,74.0503921508789,73.9994659423828],[74.4594268798828,74.3998031616211,74.4303207397461,74.4544448852539,74.4904327392578,74.5123519897461,74.5155258178711,74.4594268798828],[74.4004364013672,74.3529815673828,74.317138671875,74.272705078125,74.2393035888672,74.1968231201172,74.1029281616211,74.0950698852539,74.146240234375,74.1632308959961,74.21923828125,74.27294921875,74.3530807495117,74.3683090209961,74.3743209838867,74.3638229370117,74.3887710571289,74.456298828125,74.4962387084961,74.5267562866211,74.5489730834961,74.4795913696289,74.4410171508789,74.4004364013672],[74.9813003540039,74.9629821777344,74.991943359375,74.9825668334961,74.9398956298828,74.8619155883789,74.8307571411133,74.8482971191406,74.8508834838867,74.8935089111328,74.93896484375,74.9659652709961,74.9928207397461,74.9813003540039],[75.4193878173828,75.386962890625,75.3505401611328,75.3389663696289,75.3409652709961,75.2471237182617,75.2627868652344,75.3165054321289,75.280517578125,75.2889175415039,75.3309555053711,75.3395462036133,75.367919921875,75.451416015625,75.4070281982422,75.4099578857422,75.4600143432617,75.4977111816406,75.5134811401367,75.515625,75.4193878173828],[75.3707580566406,75.3643035888672,75.43798828125,75.4405288696289,75.4135284423828,75.387451171875,75.3364715576172,75.3093719482422,75.2724151611328,75.2363815307617,75.2281265258789,75.2620620727539,75.2445831298828,75.21923828125,75.1640090942383,75.1343307495117,75.099853515625,75.1201705932617,75.1553268432617,75.1623992919922,75.1565399169922,74.944580078125,74.9189453125,74.8667984008789,74.7953109741211,74.7726058959961,74.7724609375,74.8004379272461,74.82568359375,74.8573226928711,74.9319839477539,74.9589385986328,74.9842758178711,74.9984359741211,75.0625,75.1142044067383,75.1982955932617,75.2955551147461,75.3937530517578,75.4809036254883,75.5582046508789,75.581787109375,75.5104446411133,75.4286575317383,75.3707580566406],[75.4095687866211,75.3819808959961,75.3895492553711,75.4632339477539,75.495849609375,75.5764694213867,75.636474609375,75.7099609375,75.7663116455078,75.8452606201172,75.7984924316406,75.729248046875,75.6943817138672,75.6255798339844,75.6055603027344,75.5219192504883,75.4861297607422,75.4383773803711,75.4095687866211],[76.1217269897461,76.0857925415039,76.1436080932617,76.1748046875,76.19482421875,76.1851577758789,76.1634292602539,76.1217269897461],[75.8289489746094,75.809814453125,75.8224182128906,75.7958526611328,75.6897964477539,75.6631851196289,75.6439437866211,75.6341323852539,75.6307144165039,75.6520004272461,75.7004928588867,75.74951171875,75.7989273071289,75.8668441772461,75.9273452758789,75.9645004272461,75.9889602661133,76.0637664794922,76.1371536254883,76.1080627441406,76.0435562133789,75.9036102294922,75.8634338378906,75.826904296875,75.8136215209961,75.8223114013672,75.8603973388672,75.8636703491211,75.5856018066406,75.5640640258789,75.5304718017578,75.48974609375,75.4160690307617,75.3655776977539,75.3245162963867,75.2689437866211,75.102294921875,75.0591812133789,75.044677734375,75.083984375,75.0828628540039,75.1168975830078,75.2174301147461,75.267822265625,75.337646484375,75.4488754272461,75.5445785522461,75.57177734375,75.6332550048828,75.6598663330078,75.7132797241211,75.7209014892578,75.6916961669922,75.66064453125,75.4575653076172,75.3926696777344,75.3461456298828,75.1332473754883,75.1030731201172,75.0624008178711,74.9703140258789,74.8677749633789,74.83740234375,74.8204116821289,74.8285598754883,74.8499069213867,74.8996047973633,74.9509811401367,74.9912643432617,74.9825668334961,74.9471664428711,74.9231948852539,74.8818359375,74.8560562133789,74.846923828125,74.894775390625,74.9637680053711,74.9640655517578,74.9456024169922,74.904052734375,74.8377990722656,74.7492218017578,74.6868133544922,74.65966796875,74.6565399169922,74.6736831665039,74.700927734375,74.7974624633789,74.8270034790039,74.870849609375,75.008544921875,75.0405731201172,75.05419921875,75.1237258911133,75.2350082397461,75.2703552246094,75.3255386352539,75.3653335571289,75.3465805053711,75.3486328125,75.5543899536133,75.7494125366211,75.7816390991211,75.7595672607422,75.8233947753906,75.90966796875,75.9552230834961,75.9881820678711,76.0156784057617,76.0277862548828,76.0472640991211,76.0805206298828,76.1149368286133,76.1300811767578,76.19970703125,76.1967315673828,76.1601028442383,76.1083450317383,76.0807113647461,76.013427734375,75.9530792236328,75.8289489746094],[76.2781219482422,76.2638168334961,76.2212448120117,76.2337417602539,76.212158203125,76.177490234375,76.1217269897461,76.1554718017578,76.1602554321289,76.1936492919922,76.2147445678711,76.2616271972656,76.2890625,76.2496109008789,76.2938995361328,76.2718734741211,76.3053741455078,76.2781219482422],[76.1991653442383,76.1884307861328,76.1957550048828,76.2069320678711,76.2578582763672,76.3241653442383,76.3448257446289,76.355224609375,76.3434066772461,76.3153839111328,76.3025894165039,76.2365188598633,76.1991653442383],[76.6208953857422,76.552978515625,76.5095748901367,76.499267578125,76.4829635620117,76.452392578125,76.4500503540039,76.4837875366211,76.5379867553711,76.54931640625,76.6029739379883,76.6261672973633,76.6328659057617,76.6183547973633,76.643798828125,76.6208953857422],[76.599365234375,76.5844268798828,76.5907211303711,76.6288528442383,76.6896057128906,76.7066955566406,76.599365234375],[76.659912109375,76.6482391357422,76.6769561767578,76.74658203125,76.7820739746094,76.7472152709961,76.677001953125,76.659912109375],[76.2375946044922,76.1612777709961,76.108154296875,76.072265625,76.0470199584961,75.9836883544922,75.9046401977539,75.8394546508789,75.7882308959961,75.7196731567383,75.672607421875,75.6687469482422,75.6030807495117,75.5757369995117,75.427734375,75.3196334838867,75.3108367919922,75.3148422241211,75.281005859375,75.2225570678711,75.1636734008789,75.11083984375,75.068603515625,75.0550308227539,75.0592803955078,75.0547332763672,75.007568359375,74.9707489013672,74.9461441040039,74.9046325683594,74.8753433227539,74.8370132446289,74.7965850830078,74.755859375,74.74462890625,74.7458953857422,74.6954574584961,74.6644515991211,74.6370162963867,74.6101608276367,74.6137161254883,74.6929702758789,74.665771484375,74.61083984375,74.5519027709961,74.5075225830078,74.4855499267578,74.4626922607422,74.4989242553711,74.4642105102539,74.4218215942383,74.3280258178711,74.2892532348633,74.2273941040039,74.1288528442383,74.0138168334961,73.9739303588867,73.8978500366211,73.8504333496094,73.8050765991211,73.7691955566406,73.7681655883789,73.7754898071289,73.8256378173828,73.8380432128906,73.8145523071289,73.7460479736328,73.658203125,73.6102981567383,73.50439453125,73.3665542602539,73.3042984008789,73.2972183227539,73.3141098022461,73.3458938598633,73.3568420410156,73.3920440673828,73.453857421875,73.4494171142578,73.4185028076172,73.3509826660156,73.4810028076172,73.5420379638672,73.6971206665039,73.7661590576172,73.800537109375,73.8223190307617,73.8857421875,73.9356460571289,73.9513168334961,73.9595718383789,74.0339813232422,74.0957489013672,74.1291046142578,74.1866149902344,74.4196319580078,74.4361343383789,74.4813003540039,74.49609375,74.5421829223633,74.5412063598633,74.5561065673828,74.5905303955078,74.627685546875,74.6562957763672,74.7960891723633,74.8975143432617,74.9570770263672,74.9729537963867,75.0134735107422,75.0033645629883,75.0587387084961,75.0906219482422,75.1249008178711,75.1683578491211,75.1942367553711,75.1865692138672,75.164306640625,75.13818359375,75.0960922241211,75.0977554321289,75.244384765625,75.2777328491211,75.3284149169922,75.3514175415039,75.3750991821289,75.3838348388672,75.3732376098633,75.3412628173828,75.3564453125,75.4544906616211,75.5066909790039,75.592529296875,75.6189880371094,75.6630859375,75.7197799682617,75.7768096923828,75.8547821044922,75.8717269897461,75.8737335205078,75.9070358276367,75.9458465576172,75.9838409423828,76.0665512084961,76.0962371826172,76.108642578125,76.1040496826172,76.0688018798828,76.0712890625,76.0892562866211,76.119873046875,76.1690368652344,76.2329559326172,76.2735366821289,76.2820281982422,76.2984848022461,76.291015625,76.2416076660156,76.23046875,76.2452163696289,76.2366714477539,76.3095245361328,76.378173828125,76.426025390625,76.4843292236328,76.4967269897461,76.4996566772461,76.5179214477539,76.5678176879883,76.5786590576172,76.5792922973633,76.6133270263672,76.6879348754883,76.74609375,76.821044921875,76.923828125,76.9637680053711,77.0077667236328,77.0115737915039,76.9906234741211,76.9336929321289,76.8706588745117,76.7895965576172,76.7605514526367,76.7076721191406,76.6597137451172,76.6104965209961,76.5729522705078,76.4494094848633,76.3134765625,76.2848587036133,76.2375946044922],[77.0266571044922,77.0072784423828,76.9921417236328,77.0025405883789,76.9749450683594,76.9906234741211,77.0213394165039,77.0115203857422,77.0188446044922,77.0564956665039,77.0975570678711,77.2055130004883,77.154052734375,77.1295928955078,77.0711898803711,77.0266571044922],[77.1888198852539,77.1839828491211,77.2099151611328,77.226806640625,77.27197265625,77.3014678955078,77.31103515625,77.2803268432617,77.2544937133789,77.1888198852539],[77.24267578125,77.2415008544922,77.2890167236328,77.3466339111328,77.3471145629883,77.3300323486328,77.3196716308594,77.2998046875,77.2682647705078,77.24267578125],[42.3025360107422,42.3277816772461,42.3725090026855,42.4156265258789,42.4747047424316,42.535400390625,42.5673332214355,42.6232414245605,42.67431640625,42.685546875,42.6998519897461,42.72705078125,42.7554206848145,42.7790985107422,42.8116226196289,42.8358383178711,42.8568344116211,42.8633308410645,42.8517570495605,42.8831024169922,42.9022445678711,42.956298828125,43.0380859375,43.0624504089355,43.0976104736328,43.1421852111816,43.2577629089355,43.337646484375,43.3780784606934,43.4330558776855,43.4690399169922,43.4904289245605,43.5055656433105,43.5670890808105,43.65087890625,43.7047386169434,44.0029296875,44.0715789794922,44.4691886901855,44.5956535339355,44.65966796875,44.7532234191895,44.7999534606934,44.8443336486816,44.8888664245605,44.9100112915039,44.920166015625,44.9361305236816,44.9840316772461,45.0131340026855,45.0836448669434,45.1365699768066,45.2053718566895,45.2426261901855,45.3052749633789,45.3268547058105,45.2737312316895,45.2439918518066,45.2259788513184,45.2032699584961,45.1599617004395,45.122802734375,45.0937004089355,45.08056640625,45.0611305236816,45.0460433959961,45.0299301147461,45.0745582580566,45.1307144165039,45.220458984375,45.3214378356934,45.4948234558105,45.5452613830566,45.5530776977539,45.5722160339355,45.6046867370605,45.6512222290039,45.705078125,45.7576675415039,45.8104476928711,45.8788070678711,45.8978004455566,45.9203109741211,45.9552230834961,46.0089378356934,46.0696296691895,46.1397438049316,46.1859359741211,46.2242698669434,46.2477531433105,46.30908203125,46.3360328674316,46.366943359375,46.4305686950684,46.4991226196289,46.6142578125,46.7131843566895,46.858154296875,46.8819808959961,46.9507827758789,46.9781265258789,47.0689964294434,47.1280746459961,47.1942367553711,47.2587432861328,47.3022003173828,47.3526344299316,47.3777351379395,47.41357421875,47.4294929504395,47.438232421875,47.4473648071289,47.4851570129395,47.5238761901855,47.6248550415039,47.6844749450684,47.7154273986816,47.8014183044434,47.874267578125,47.9751930236816,48.0225105285645,48.0829086303711,48.1201629638672,48.1533203125,48.21044921875,48.25390625,48.3217277526855,48.3553199768066,48.3688468933105,48.3734397888184,48.3599128723145,48.2737312316895,48.2077140808105,48.1330108642578,48.09716796875,48.1015167236328,48.1056632995605,48.0644035339355,47.9790992736816,47.9400863647461,47.947265625,47.8900871276855,47.7685050964355,47.7149925231934,47.7294921875,47.7179679870605,47.6805191040039,47.6820335388184,47.7226066589355,47.7278327941895,47.6976547241211,47.6914558410645,47.7093238830566,47.7598152160645,47.8429222106934,47.929443359375,48.0192375183105,48.1276359558105,48.2545890808105,48.3415031433105,48.388427734375,48.4303703308105,48.4833984375,48.5746574401855,48.6024894714355,48.6801261901855,48.773193359375,48.8611831665039,48.8663558959961,48.8916511535645,48.9722633361816,49.1988754272461,49.2785186767578,49.2866706848145,49.3234367370605,49.3888168334961,49.3894538879395,49.3894538879395,49.3623542785645,49.3538589477539,49.378662109375,49.3813972473145,49.362060546875,49.3746604919434,49.4192390441895,49.4489288330078,49.4637680053711,49.4947280883789,49.5418434143066,49.5769538879395,49.6001472473145,49.59423828125,49.5592765808105,49.568603515625,49.6221199035645,49.6715316772461,49.7167472839355,49.7602043151855,49.8018035888672,49.8734359741211,49.9750480651855,50.0716781616211,50.208984375,50.298583984375,50.3501472473145,50.3936042785645,50.4280738830566,50.4535140991211,50.4941902160645,50.5500984191895,50.621337890625,50.7079582214355,50.8294448852539,50.9858894348145,51.1001472473145,51.1723136901855,51.2301292419434,51.2613754272461,51.3148918151855,51.3741683959961,51.4122543334961,51.4480476379395,51.5056648254395,51.5450668334961,51.5663108825684,51.6099128723145,51.7030258178711,51.7812995910645,51.9258308410645,52.0312995910645,52.12646484375,52.1729965209961,52.2145004272461,52.2865219116211,52.3062515258789,52.3316383361816,52.3620109558105,52.3997573852539,52.44482421875,52.4838409423828,52.5191421508789,52.546630859375,52.5733375549316,52.6102066040039,52.6430168151855,52.6734848022461,52.6919937133789,52.7158660888672,52.7394523620605,52.7678680419922,52.8006858825684,52.8715324401855,52.8907203674316,52.8907203674316,52.9308090209961,52.956298828125,53.0037117004395,53.0422859191895,53.0574684143066,53.047607421875,53.083740234375,53.1658210754395,53.2036628723145,53.1973152160645,53.1726570129395,53.1297378540039,53.1338386535645,53.2106437683105,53.2296371459961,53.2709465026855,53.3408660888672,53.3701171875,53.3586921691895,53.4056129455566,53.510986328125,53.5465354919434,53.5266609191895,53.5264663696289,53.5294418334961,53.53076171875,53.5556144714355,53.5445823669434,53.4977073669434,53.468505859375,53.4569816589355,53.4625015258789,53.4850082397461,53.4514656066895,53.3835945129395,53.3170394897461,53.2845687866211,53.1718292236328,52.9680671691895,52.8398933410645,52.7872047424316,52.7182159423828,52.6329116821289,52.6024894714355,52.6269989013672,52.6150398254395,52.566650390625,52.4936027526855,52.3958969116211,52.2999038696289,52.2054672241211,52.0965347290039,51.9730453491211,51.8485374450684,51.7229957580566,51.6006813049316,51.422119140625,51.2670402526855,51.1794929504395,51.1077117919922,51.0301246643066,50.94677734375,50.8631362915039,50.7792510986328,50.7028350830078,50.6338844299316,50.5609893798828,50.4841804504395,50.4325180053711,50.406005859375,50.3798332214355,50.3539047241211,50.2789535522461,50.1549339294434,50.06640625,50.0133781433105,49.9788589477539,49.9628410339355,49.844482421875,49.6927757263184,49.5134773254395,49.5135269165039,49.5358428955078,49.6094245910645,49.6248512268066,49.6929702758789,49.73779296875,49.8237762451172,49.8770484924316,49.9203109741211,49.9780769348145,50.00927734375,50.0107917785645,49.9521484375,49.9059066772461,49.880615234375,49.8860359191895,49.896484375,49.9117660522461,49.9488754272461,50.0594215393066,50.1385726928711,50.1830558776855,50.2336921691895,50.2457008361816,50.241455078125,50.2554702758789,50.2744140625,50.2572746276855,50.2047348022461,50.1011238098145,50.0615234375,50.0070304870605,49.9416007995605,49.8743133544922,49.7971687316895,49.6925277709961,49.6162605285645,49.5692367553711,49.5235824584961,49.5072746276855,49.5323219299316,49.5145988464355,49.4242172241211,49.416015625,49.4036102294922,49.3977546691895,49.3764152526855,49.3609352111816,49.3426246643066,49.3558578491211,49.3042984008789,49.1661605834961,49.1429672241211,49.1375961303711,49.1870613098145,49.219970703125,49.2158699035645,49.17041015625,49.2056159973145,49.2393035888672,49.2698745727539,49.2963371276855,49.3349113464355,49.3353500366211,49.3356437683105,49.3228034973145,49.3415069580078,49.3963851928711,49.5248031616211,49.5626449584961,49.593994140625,49.6468772888184,49.6535148620605,49.6910171508789,49.74072265625,49.8490219116211,49.9247093200684,49.9478034973145,49.9600105285645,49.9831047058105,49.9866714477539,49.9894027709961,50.0330085754395,50.0864753723145,50.1966781616211,50.248291015625,50.3125991821289,50.3288078308105,50.3175773620605,50.3045883178711,50.3325691223145,50.367919921875,50.4053726196289,50.4141578674316,50.4412612915039,50.4737319946289,50.4604988098145,50.4295883178711,50.3899421691895,50.3829116821289,50.3418426513672,50.3171882629395,50.30615234375,50.2752914428711,50.2144546508789,50.16943359375,50.1542472839355,50.1572761535645,50.1718292236328,50.1760749816895,50.1538581848145,50.1385726928711,50.1649436950684,50.1870613098145,50.2002906799316,50.2642555236816,50.2907257080078,50.3006362915039,50.333251953125,50.3665542602539,50.3871574401855,50.4613265991211,50.5256805419922,50.5361785888672,50.5442390441895,50.5851097106934,50.6346664428711,50.66552734375,50.7184562683105,50.7686996459961,50.7912101745605,50.8294448852539,50.9014663696289,50.9743156433105,51.0506858825684,51.1075172424316,51.216064453125,51.2608413696289,51.3137702941895,51.3534660339355,51.3822250366211,51.4210433959961,51.4671859741211,51.4714813232422,51.45263671875,51.4747543334961,51.5132789611816,51.5530281066895,51.604248046875,51.6615753173828,51.7134742736816,51.72607421875,51.7298355102539,51.7371101379395,51.7555198669434,51.8275413513184,51.8716316223145,51.8925285339355,51.89990234375,51.9235343933105,51.9988746643066,52.0348625183105,52.035400390625,52.1017074584961,52.1172828674316,52.070068359375,51.9574699401855,51.9050750732422,51.8011741638184,51.7176246643066,51.6742668151855,51.6345710754395,51.5784187316895,51.505615234375,51.4857406616211,51.4835433959961,51.449951171875,51.3770523071289,51.3484382629395,51.2804679870605,51.250732421875,51.2178726196289,51.1651878356934,51.0516624450684,50.9852561950684,50.943359375,50.8871612548828,50.8551788330078,50.8176765441895,50.7691421508789,50.7020530700684,50.6446304321289,50.6038093566895,50.5685539245605,50.5571746826172,50.5332527160645,50.4869613647461,50.4229965209961,50.3024406433105,50.2276840209961,50.1805648803711,50.1065940856934,50.0778312683105,50.0142593383789,49.9967765808105,49.94677734375,49.9445304870605,49.9446296691895,49.93359375,49.9114723205566,49.8431129455566,49.7730484008789,49.741455078125,49.7308082580566,49.76171875,49.8050308227539,49.8298835754395,49.8828125,49.9115715026855,49.8978500366211,49.87841796875,49.8925323486328,49.918701171875,49.9115219116211,49.8960456848145,49.901123046875,49.9541015625,49.9824714660645,49.9987297058105,49.998779296875,49.9735832214355,49.9600105285645,49.9905738830566,50.012939453125,50.0125007629395,49.9660148620605,49.94384765625,49.9112319946289,49.91552734375,49.9412078857422,49.9441413879395,49.94384765625,49.9354476928711,49.9615745544434,50.0082511901855,50.0437507629395,50.0481948852539,50.0432624816895,50.028076171875,50.0237312316895,50.0879402160645,50.1328125,50.1657218933105,50.1796379089355,50.2218246459961,50.3034172058105,50.4048843383789,50.5113754272461,50.556396484375,50.5728530883789,50.56884765625,50.57763671875,50.5836906433105,50.5855445861816,50.5974617004395,50.6084938049316,50.6155776977539,50.6065406799316,50.6039047241211,50.654541015625,50.7125015258789,50.7449226379395,50.7782211303711,50.7891120910645,50.7786636352539,50.7109375,50.6832046508789,50.6882820129395,50.7254371643066,50.7650871276855,50.8030776977539,50.8641586303711,50.8498039245605,50.8122062683105,50.7751960754395,50.7005882263184,50.6919898986816,50.6976089477539,50.693603515625,50.66552734375,50.6151390075684,50.5755348205566,50.56201171875,50.5221672058105,50.468017578125,50.4700660705566,50.46337890625,50.452392578125,50.422607421875,50.4154777526855,50.3641586303711,50.32373046875,50.3059539794922,50.2594261169434,50.2223625183105,50.2133293151855,50.1668968200684,50.1511726379395,50.11669921875,50.1033210754395,50.09375,50.0692863464355,49.9843254089355,49.9535140991211,49.9480972290039,49.9030303955078,49.8232879638672,49.75048828125,49.7174797058105,49.69970703125,49.6605453491211,49.6115226745605,49.6111335754395,49.6270523071289,49.595703125,49.5322265625,49.5013656616211,49.4728012084961,49.4837417602539,49.5076675415039,49.5396957397461,49.5276374816895,49.4815406799316,49.4484367370605,49.4462394714355,49.4645500183105,49.4861335754395,49.4825706481934,49.4727058410645,49.4828643798828,49.4725608825684,49.4517097473145,49.3814926147461,49.2984390258789,49.2562980651855,49.2197723388672,49.1869125366211,49.16455078125,49.162109375,49.1623039245605,49.1658172607422,49.147216796875,49.1323699951172,49.1224136352539,49.0914535522461,49.0766105651855,49.0857887268066,49.1476593017578,49.2161598205566,49.2397918701172,49.2545890808105,49.2873039245605,49.3220710754395,49.4878883361816,49.55859375,49.5626945495605,49.6097145080566,49.6566925048828,49.695556640625,49.7486801147461,49.7772941589355,49.7691383361816,49.707763671875,49.6384735107422,49.5875015258789,49.5463409423828,49.4993171691895,49.5054664611816,49.50341796875,49.4993171691895,49.5504417419434,49.5565452575684,49.6053733825684,49.6239280700684,49.5994644165039,49.6158180236816,49.6648445129395,49.750732421875,49.8216285705566,49.8941383361816,49.9510765075684,50.0103034973145,50.0614280700684,50.0879859924316,50.09130859375,50.202392578125,50.21875,50.2391586303711,50.2391586303711,50.2882308959961,50.4374504089355,50.5205574035645,50.604736328125,50.6768531799316,50.774658203125,50.8180160522461,50.8871612548828,50.9357414245605,50.9945793151855,50.9945793151855,50.9892082214355,50.9605941772461,50.8972663879395,50.8931159973145,50.8933601379395,50.8694839477539,50.8263168334961,50.771484375,50.7275886535645,50.7418937683105,50.7194328308105,50.7108383178711,50.766357421875,50.764404296875,50.7391128540039,50.7398414611816,50.7711410522461,50.8236808776855,50.8710441589355,50.9097633361816,50.9564933776855,50.9664077758789,50.9462890625,50.96875,51.0149421691895,51.0723648071289,51.1465797424316,51.1910667419434,51.1856956481934,51.1897964477539,51.2427749633789,51.2814445495605,51.2834968566895,51.2934074401855,51.27734375,51.2242202758789,51.2165985107422,51.2017593383789,51.183349609375,51.1363754272461,51.0926246643066,50.9976081848145,50.9462890625,50.9190902709961,50.9246101379395,50.9117698669434,50.8583488464355,50.8399887084961,50.8072738647461,50.7582015991211,50.7745590209961,50.9554672241211,51.1600074768066,51.3779754638672,51.4931144714355,51.8681144714355,52.0474090576172,52.3570289611816,52.638427734375,52.9296875,53.094970703125,53.2691879272461,53.3174285888672,53.379150390625,53.498779296875,53.6701164245605,53.8226547241211,53.9425277709961,53.9932136535645,54.0225563049316,54.0552749633789,54.113525390625,54.1515121459961,54.145263671875,54.1824684143066,54.3218727111816,54.4423828125,54.4368629455566,54.3871078491211,54.35107421875,54.335693359375,54.3119621276855,54.258544921875,54.16796875,54.1147956848145,54.1060066223145,54.0896492004395,54.0685043334961,54.021728515625,53.9701156616211,53.893798828125,53.8267097473145,53.8192367553711,53.8340339660645,53.82568359375,53.75439453125,53.6472663879395,53.6037101745605,53.5507316589355,53.5044441223145,53.4876441955566,53.5277366638184,53.5764656066895,53.6114234924316,53.6197280883789,53.602783203125,53.5762710571289,53.4688949584961,53.4475593566895,53.4543952941895,53.5062026977539,53.5431632995605,53.598388671875,53.7072257995605,53.8114738464355,53.8683128356934,53.929443359375,53.9961891174316,54.0423812866211,54.0634765625,54.0673828125,54.0449714660645,53.9993133544922,53.9628410339355,53.9556159973145,53.9578132629395,53.9807624816895,54.1073226928711,54.12451171875,54.1343269348145,54.12158203125,54.0904312133789,54.0564956665039,54.0230484008789,53.9959487915039,53.9757804870605,53.9418449401855,53.9644546508789,54.0536613464355,54.123046875,54.1814460754395,54.2721214294434,54.3256340026855,54.3084449768066,54.2316398620605,54.2056655883789,54.2214851379395,54.1780281066895,54.1583480834961,54.2122077941895,54.260498046875,54.3640632629395,54.4554214477539,54.5386238098145,54.5993156433105,54.7150421142578,54.9504890441895,55.1279792785645,55.2611351013184,55.30517578125,55.2823715209961,55.253173828125,55.2122535705566,55.18359375,55.1624526977539,55.1767578125,55.1990737915039,55.2456550598145,55.307373046875,55.3568840026855,55.3725128173828,55.3895988464355,55.3583488464355,55.3084945678711,55.204833984375,55.1944351196289,55.1865234375,55.1609382629395,55.115234375,55.0524406433105,55.0030288696289,54.9767074584961,54.9595718383789,54.9537124633789,54.9435539245605,54.8724098205566,54.8544921875,54.8288078308105,54.7881851196289,54.7378921508789,54.7154312133789,54.6673812866211,54.6595230102539,54.6933097839355,54.6187019348145,54.623291015625,54.5933074951172,54.564453125,54.5515632629395,54.5160636901855,54.4411125183105,54.3644065856934,54.3401870727539,54.3687515258789,54.3966293334961,54.3685569763184,54.3522453308105,54.3621597290039,54.3841781616211,54.347412109375,54.3029289245605,54.2797355651855,54.26708984375,54.2364768981934,54.2450218200684,54.2432136535645,54.221923828125,54.1832046508789,54.1704597473145,54.1710433959961,54.1392593383789,54.105224609375,54.0692863464355,54.04443359375,54.01318359375,54.0026397705078,53.9799308776855,53.9543914794922,53.9464836120605,53.9949226379395,54.0492668151855,54.0194816589355,53.9638175964355,53.8824691772461,53.81298828125,53.7534675598145,53.71044921875,53.6574249267578,53.6217269897461,53.5831031799316,53.5509796142578,53.565185546875,53.5870628356934,53.5802726745605,53.5544929504395,53.5232925415039,53.5015602111816,53.4846687316895,53.455810546875,53.4657249450684,53.4458999633789,53.4062004089355,53.3367652893066,53.2871589660645,53.2751960754395,53.2394065856934,53.2224578857422,53.2284660339355,53.1739234924316,53.1078643798828,53.0574226379395,53.0054206848145,52.9661140441895,52.9437484741211,52.9559097290039,52.9693832397461,52.978515625,52.9959945678711,52.9890632629395,52.9724617004395,52.933349609375,52.8601570129395,52.8194351196289,52.7447242736816,52.67578125,52.56982421875,52.394775390625,52.3368644714355,52.2805671691895,52.2333984375,52.1508293151855,52.1463394165039,52.1255874633789,52.0245132446289,51.9764671325684,51.9332504272461,51.890625,51.8346214294434,51.7729988098145,51.7039070129395,51.6511726379395,51.616943359375,51.5370597839355,51.5287132263184,51.4923820495605,51.44189453125,51.4147491455078,51.3246078491211,51.2296867370605,51.1370124816895,50.990234375,50.8610343933105,50.7748031616211,50.6955070495605,50.6637191772461,50.669189453125,50.6791496276855,50.7041511535645,50.769775390625,50.8341827392578,50.8502922058105,50.8396949768066,50.7992706298828,50.690185546875,50.58203125,50.5439453125,50.4928703308105,50.5110816955566,50.5828132629395,50.6042976379395,50.6204109191895,50.6479034423828,50.668212890625,50.6761245727539,50.6944313049316,50.7372055053711,50.8683128356934,50.9710464477539,51.0638198852539,51.0817375183105,51.072265625,51.06884765625,51.0916481018066,51.0890121459961,51.046875,50.98095703125,50.9251441955566,50.8955574035645,50.8888664245605,50.946533203125,51.0360336303711,51.065185546875,51.0455551147461,51.0315895080566,50.9808616638184,51.0044937133789,51.01953125,50.9360847473145,50.8446273803711,50.7762680053711,50.7135276794434,50.6537590026855,50.60205078125,50.5828628540039,50.601806640625,50.665283203125,50.7447280883789,50.8697776794434,50.9413566589355,50.9980964660645,51.0115699768066,50.990234375,50.946044921875,50.9112815856934,50.8798828125,50.7810516357422,50.66015625,50.5916023254395,50.5506858825684,50.5357894897461,50.5411643981934,50.5837898254395,50.6739234924316,50.7803230285645,50.9140625,50.9957008361816,51.040771484375,51.1151847839355,51.1611785888672,51.2137222290039,51.2518081665039,51.3995590209961,51.4445304870605,51.4823722839355,51.4936027526855,51.4849624633789,51.4637184143066,51.4669456481934,51.4945793151855,51.4979019165039,51.4981460571289,51.4795417785645,51.4807624816895,51.4816398620605,51.5121612548828,51.59423828125,51.6812973022461,51.7093734741211,51.6727066040039,51.5542488098145,51.4839859008789,51.4820327758789,51.4712867736816,51.475341796875,51.4974136352539,51.5401840209961,51.594482421875,51.6474609375,51.681640625,51.7191886901855,51.7291984558105,51.6751441955566,51.5891609191895,51.505615234375,51.3697280883789,51.3215789794922,51.2895011901855,51.2545890808105,51.1971702575684,51.1318855285645,51.102294921875,51.0835914611816,51.0270004272461,50.9346656799316,50.8517074584961,50.72607421875,50.6445770263672,50.601318359375,50.6068840026855,50.619873046875,50.6126937866211,50.5503425598145,50.353759765625,50.2284660339355,50.1564445495605,50.099853515625,50.0131340026855,49.96240234375,49.9283218383789,49.8747062683105,49.8285140991211,49.8582534790039,49.9319343566895,49.9700202941895,50.0936050415039,50.2823219299316,50.3779792785645,50.41357421875,50.4027366638184,50.3579597473145,50.318115234375,50.2735366821289,50.2174797058105,50.1402359008789,50.0584983825684,50.0008773803711,49.9390602111816,49.8527336120605,49.6969718933105,49.5022468566895,49.3670921325684,49.3038558959961,49.2525863647461,49.1999015808105,49.1502952575684,49.0983390808105,49.038330078125,48.9696273803711,48.8055686950684,48.5738754272461,48.4122543334961,48.3235855102539,48.2844734191895,48.2324714660645,48.1270027160645,48.0201148986816,47.9477043151855,47.8767585754395,47.79248046875,47.7409172058105,47.7686538696289,47.8039054870605,47.7899894714355,47.7606925964355,47.7454109191895,47.7087898254395,47.5899391174316,47.4564933776855,47.3209953308105,47.2623023986816,47.1004905700684,46.9549293518066,46.7746124267578,46.7257804870605,46.7054214477539,46.7103538513184,46.73681640625,46.7586936950684,46.7659187316895,46.7571296691895,46.7343254089355,46.6986312866211,46.6499481201172,46.6056137084961,46.5770988464355,46.5664520263672,46.5079612731934,46.442138671875,46.3488273620605,46.337158203125,46.2915992736816,46.28173828125,46.2284660339355,46.189208984375,46.1004867553711,46.1007347106934,46.086181640625,46.0287628173828,45.9762191772461,45.9205589294434,45.8968238830566,45.8888702392578,45.90576171875,45.9348640441895,45.942138671875,45.9348640441895,45.7777824401855,45.7370109558105,45.7209968566895,45.6630401611328,45.6659698486328,45.6861839294434,45.65673828125,45.5840339660645,45.63427734375,45.6741714477539,45.6875953674316,45.6796875,45.6017112731934,45.5302238464355,45.4909172058105,45.455078125,45.4330558776855,45.4210433959961,45.2947769165039,45.2177238464355,45.1494636535645,45.0242691040039,44.9696273803711,44.9059562683105,44.8169937133789,44.837890625,44.8760757446289,44.8255844116211,44.7825698852539,44.71826171875,44.6565399169922,44.5606918334961,44.5033187866211,44.45166015625,44.420654296875,44.3871574401855,44.34326171875,44.2616729736328,44.1923828125,44.1031265258789,43.9933586120605,43.7798843383789,43.5550308227539,43.8346672058105,43.8846168518066,43.8059577941895,43.6849594116211,43.5097160339355,43.3816909790039,43.270751953125,43.21875,43.0350570678711,42.999755859375,42.9671401977539,42.9034690856934,42.8109397888184,42.6807098388672,42.644775390625,42.6134757995605,42.3537139892578,42.1809577941895,42.0802268981934,41.9534187316895,41.9239730834961,41.9051246643066,41.844482421875,41.7793464660645,41.663330078125,41.6019020080566,41.5450172424316,41.4847640991211,41.4586906433105,41.333984375,41.2127418518066,41.1992683410645,41.2181167602539,41.2290534973145,41.2824211120605,41.3150863647461,41.4556159973145,41.5160636901855,41.5546875,41.5874977111816,41.6213874816895,41.67041015625,41.743408203125,41.8125991821289,41.8313484191895,41.8069305419922,41.8000984191895,41.8123054504395,41.8704109191895,41.8909683227539,41.9046363830566,41.9603500366211,41.9898910522461,41.9920387268066,42.0087394714355,42.035400390625,42.0707054138184,42.1099586486816,42.1588859558105,42.205078125,42.2347183227539,42.3573722839355,42.4750480651855,42.4980964660645,42.5176734924316,42.5357398986816,42.52978515625,42.6482391357422,42.6750030517578,42.6941413879395,42.7302742004395,42.7563972473145,42.746826171875,42.6167984008789,42.7096214294434,42.7347183227539,42.7484855651855,42.7486343383789,42.7035140991211,42.6536140441895,42.6163558959961,42.5956039428711,42.5665512084961,42.571533203125,42.5938491821289,42.6169929504395,42.6575202941895,42.7029762268066,42.727783203125,42.7470207214355,42.8077163696289,42.844482421875,42.8966789245605,42.9890632629395,43.0496559143066,43.0915031433105,43.1326179504395,43.1695785522461,43.1590805053711,43.1551246643066,43.2242164611816,43.2280769348145,43.2073249816895,43.1991233825684,43.1901359558105,43.21923828125,43.2763175964355,43.3333969116211,43.3744659423828,43.4180641174316,43.4799308776855,43.5338859558105,43.5120086669922,43.542724609375,43.5697784423828,43.5531234741211,43.48486328125,43.4198226928711,43.4728012084961,43.7278823852539,43.8972663879395,44.2880859375,44.3180160522461,44.3744621276855,44.419677734375,44.6988258361816,44.661376953125,44.6708526611328,44.6952629089355,44.7353515625,44.7883796691895,44.905029296875,44.9719734191895,45.069580078125,45.12646484375,45.1513175964355,45.1854972839355,45.2517585754395,45.2896957397461,45.3400382995605,45.3483390808105,45.3718757629395,45.4097175598145,45.4270515441895,45.3835945129395,45.3028793334961,45.2723121643066,45.3109397888184,45.377197265625,45.4297370910645,45.4883804321289,45.486328125,45.49951171875,45.564697265625,45.654052734375,45.799560546875,46.001708984375,46.0477523803711,46.01708984375,45.9698753356934,45.934814453125,46.0028305053711,46.0948219299316,46.0953636169434,46.0800285339355,46.0905265808105,46.241943359375,46.3943367004395,46.3828620910645,46.406494140625,46.5320816040039,46.6361312866211,46.6337890625,46.6180191040039,46.690673828125,46.7012672424316,46.6783180236816,46.6636695861816,46.732177734375,46.8130836486816,46.873046875,46.9061546325684,47.0234375,47.0441398620605,47.0708961486816,47.1057624816895,47.1995086669922,47.2688484191895,47.272216796875,47.1756820678711,47.1439476013184,47.1503410339355,47.2122077941895,47.23583984375,47.2616195678711,47.2391128540039,47.175537109375,47.0914573669434,47.1355972290039,47.1752471923828,47.2127418518066,47.2369651794434,47.259033203125,47.2766609191895,47.2876930236816,47.2965316772461,47.32080078125,47.3427734375,47.3736801147461,47.408935546875,47.4795417785645,47.5591812133789,47.6099586486816,47.6224098205566,47.6659164428711,47.714111328125,47.8370132446289,47.8551254272461,47.8484878540039,47.83740234375,47.833740234375,47.8412094116211,47.8448219299316,47.8875503540039,47.9644546508789,48.0353050231934,48.1683578491211,48.2379417419434,48.2688941955566,48.2819328308105,48.2884254455566,48.3027839660645,48.3319358825684,48.3604507446289,48.4190902709961,48.4842262268066,48.5427742004395,48.5718765258789,48.5912132263184,48.6624526977539,48.7393569946289,48.7820777893066,48.8077163696289,48.7937507629395,48.807373046875,48.8220710754395,48.8514137268066,48.8779792785645,48.9144554138184,48.9595718383789,49.0079078674316,49.0365715026855,49.0640640258789,49.1298332214355,49.2002944946289,49.2515602111816,49.3072280883789,49.3688468933105,49.4315452575684,49.4970703125,49.5426750183105,49.5768547058105,49.5967292785645,49.5907745361328,49.5676765441895,49.572021484375,49.6506843566895,49.72802734375,49.7306671142578,49.7420425415039,49.7819328308105,49.8332023620605,49.85595703125,49.8417472839355,49.8184089660645,49.82470703125,49.8843269348145,49.952880859375,49.95458984375,49.9640617370605,50.0523452758789,50.0514640808105,50.025390625,49.9545402526855,49.9394035339355,49.9278335571289,49.9200172424316,49.9642105102539,50.1090812683105,50.2149429321289,50.2918434143066,50.3407249450684,50.4114761352539,50.4176254272461,50.3949699401855,50.3608856201172,50.3515129089355,50.3395500183105,50.2918434143066,50.2462387084961,50.209228515625,50.23486328125,50.2804679870605,50.2968254089355,50.2804679870605,50.311767578125,50.3678245544434,50.4085464477539,50.419677734375,50.4371070861816,50.40576171875,50.3459968566895,50.3687477111816,50.4399909973145,50.4599113464355,50.5396995544434,50.6109352111816,50.6422348022461,50.6820793151855,50.7276840209961,50.767578125,50.7989234924316,50.904296875,50.9499015808105,50.9869155883789,51.0211448669434,51.0438957214355,51.0467796325684,51.0438957214355,51.0609855651855,51.120849609375,51.1806640625,51.2034187316895,51.2017593383789,51.189208984375,51.1693344116211,51.1722183227539,51.203125,51.237060546875,51.2437973022461,51.25537109375,51.27685546875,51.3116683959961,51.3401870727539,51.3632316589355,51.419921875,51.4840812683105,51.5538063049316,51.6079597473145,51.6449699401855,51.6791496276855,51.6922378540039,51.7165069580078,51.7415046691895,51.7804183959961,51.9796371459961,52.1559562683105,52.25146484375,52.3156242370605,52.3447761535645,52.3326148986816,52.3337898254395,52.3535652160645,52.3404312133789,52.25634765625,52.2526359558105,52.2791023254395,52.3085479736328,52.3072280883789,52.2948226928711,52.2721214294434,52.114013671875,52.0829582214355,52.0505867004395,52.0450210571289,52.046630859375,52.0708999633789,52.0994110107422,52.10107421875,52.1258316040039,52.2206535339355,52.26220703125,52.2848129272461,52.3123054504395,52.426025390625,52.532470703125,52.5461921691895,52.6330070495605,52.69873046875,52.7314453125,52.7592277526855,52.7982444763184,52.86181640625,52.9334487915039,52.9897956848145,53.0167007446289,53.0608863830566,53.1389656066895,53.184814453125,53.196044921875,53.2024917602539,53.200927734375,53.1841812133789,53.1468772888184,53.106201171875,53.0894508361816,53.0911636352539,53.1283721923828,53.2105941772461,53.2703132629395,53.3124046325684,53.3289031982422,53.3363304138184,53.3714370727539,53.41943359375,53.4481468200684,53.5469703674316,53.5792465209961,53.6172866821289,53.6533203125,53.6929206848145,53.78125,53.796875,53.7919425964355,53.8104515075684,53.85498046875,53.9350090026855,54.0007820129395,54.0307121276855,54.055908203125,54.1111793518066,54.1959457397461,54.2916984558105,54.3916511535645,54.4529800415039,54.4917945861816,54.51708984375,54.6109390258789,54.6253433227539,54.6484870910645,54.6958999633789,54.7832527160645,54.8060035705566,54.8609352111816,54.9149894714355,54.9407234191895,55.0504875183105,55.0877914428711,55.1375961303711,55.2234382629395,55.2787132263184,55.3069839477539,55.3302726745605,55.3604011535645,55.3974151611328,55.5253448486328,55.5700187683105,55.5963859558105,55.6075210571289,55.6011238098145,55.6221199035645,55.6554679870605,55.666259765625,55.7002906799316,55.768798828125,55.7868156433105,55.84521484375,55.83642578125,55.8452606201172,55.8323249816895,55.7951202392578,55.7704086303711,55.7697257995605,55.7511749267578,55.6845703125,55.724853515625,55.7843742370605,55.834716796875,55.8810577392578,55.9122047424316,55.938720703125,55.9678726196289,56.0211448669434,56.0217781066895,56.0021018981934,55.9425773620605,55.9553718566895,56.0026359558105,56.061767578125,56.0919914245605,56.0890159606934,56.0867195129395,56.0525398254395,56.055908203125,56.1429176330566,56.1903305053711,56.2604026794434,56.3155746459961,56.3868675231934,56.5106925964355,56.5457000732422,56.599853515625,56.6453132629395,56.7037124633789,56.7410659790039,56.8241691589355,56.8534202575684,56.8670883178711,56.8432121276855,56.8456535339355,56.9780769348145,57.0546379089355,57.1351051330566,57.1668930053711,57.1944808959961,57.2477035522461,57.2933120727539,57.3169479370117,57.3681182861328,57.4297866821289,57.5081558227539,57.5240249633789,57.5281219482422,57.55029296875,57.612548828125,57.6667938232422,57.7249526977539,57.7642059326172,57.7994155883789,57.8410148620605,57.8567352294922,57.8707046508789,57.8841323852539,57.9054679870605,57.9346199035645,58.0139198303223,58.1380844116211,58.2213401794434,58.2700729370117,58.3262710571289,58.3814926147461,58.4352531433105,58.7330551147461,58.7872543334961,58.84130859375,58.8862762451172,58.9449691772461,59.0519981384277,59.1926727294922,59.2776336669922,59.2970199584961,59.3017120361328,59.3278274536133,59.3432655334473,59.3575668334961,59.3744125366211,59.403076171875,59.4531784057617,59.4842758178711,59.5540046691895,59.6471672058105,59.7247581481934,59.7815437316895,59.7865257263184,59.7246551513672,59.6925277709961,59.7340812683105,59.8142547607422,59.8495597839355,59.8180656433105,59.8066864013672,59.8119125366211,59.8287620544434,59.8547859191895,59.9015655517578,59.9609870910645,59.999755859375,59.9556655883789,59.8735809326172,59.904296875,59.9571304321289,60.0025901794434,60.0263671875,60.120849609375,60.1953125,60.2018547058105,60.1759262084961,60.1914520263672,60.3315467834473,60.3752937316895,60.4829559326172,60.5401382446289,60.4916038513184,60.5428733825684,60.6109886169434,60.6525382995605,60.6772956848145,60.5709991455078,60.5361289978027,60.745849609375,60.8969230651855,60.9196319580078,60.960205078125,61.0028305053711,61.0587425231934,61.1690444946289,61.2877922058105,61.4442367553711,61.4934577941895,61.5460968017578,61.7115707397461,61.7573738098145,61.96484375,62.0682106018066,62.1275863647461,62.3237838745117,62.4813957214355,62.5678253173828,62.691650390625,62.7761268615723,62.8854026794434,62.9216270446777,62.955322265625,63.0077171325684,63.0680694580078,63.1418952941895,63.2082977294922,63.3006362915039,63.41748046875,63.5040512084961,63.6890144348145,63.7351570129395,63.7473182678223,63.8033180236816,63.947509765625,64.0206069946289,64.0772933959961,64.14111328125,64.2000045776367,64.2365188598633,64.2824172973633,64.3661117553711,64.443359375,64.5242691040039,64.5577087402344,64.6446228027344,64.6880874633789,64.7325744628906,64.7650451660156,64.8042984008789,64.8457565307617,64.9117660522461,64.9684143066406,65.001953125,65.0394973754883,65.080322265625,65.10791015625,65.1450653076172,65.185302734375,65.2047348022461,65.223876953125,65.2347717285156,65.2486801147461,65.2653350830078,65.3369598388672,65.4734344482422,65.5687484741211,65.6245574951172,65.6343765258789,65.6636199951172,65.6707077026367,65.6816864013672,65.7262649536133,65.7865219116211,66.02294921875,66.091064453125,66.1770553588867,66.23486328125,66.276123046875,66.3568420410156,66.439697265625,66.5321731567383,66.6170425415039,66.6955032348633,66.8492202758789,66.8917541503906,66.9302291870117,66.970947265625,67.0965805053711,67.201416015625,67.3243637084961,67.4264221191406,67.5474624633789,67.6682586669922,67.6885757446289,67.7539978027344,67.9291000366211,68.0618667602539,68.1179656982422,68.1897964477539,68.3513717651367,68.4883728027344,68.5376510620117,68.7714309692383,68.8138198852539,68.8400421142578,68.8564453125,68.8655242919922,68.8722686767578,68.9041519165039,68.92822265625,68.9610290527344,69.0096664428711,69.02197265625,69.0499572753906,69.071533203125,69.0970230102539,69.2706069946289,69.2981414794922,69.3604507446289,69.3924789428711,69.432861328125,69.4642639160156,69.5016174316406,69.5427703857422,69.58056640625,69.6298828125,69.6358413696289,69.6337890625,69.584716796875,69.5325698852539,69.5285186767578,69.5384292602539,69.5612258911133,69.6058120727539,69.6517562866211,69.783447265625,69.7692337036133,69.6895980834961,69.6969223022461,69.7209930419922,69.8157653808594,69.8319854736328,69.8099136352539,69.8353042602539,69.9139175415039,69.9536666870117,69.8687057495117,69.8064956665039,69.7518615722656,69.7221221923828,69.6705017089844,69.6261672973633,69.6017074584961,69.605712890625,69.6740188598633,69.632568359375,69.5966262817383,69.5542449951172,69.4791030883789,69.4894485473633,69.4608383178711,69.4701232910156,69.4456100463867,69.3833465576172,69.3673324584961,69.427734375,69.4442901611328,69.4281768798828,69.378173828125,69.3152770996094,69.2674331665039,69.15185546875,69.1168441772461,69.0687026977539,69.0981979370117,69.13037109375,69.2891616821289,69.3102493286133,69.3131332397461,69.3029327392578,69.2280807495117,69.2212448120117,69.2308120727539,69.2655792236328,69.2754364013672,69.1917495727539,69.0034713745117,68.692138671875,68.4151382446289,68.3556213378906,68.3218765258789,68.3447265625,68.3249053955078,68.0717315673828,68.05859375,68.1121520996094,68.1622085571289,68.1508255004883,68.114501953125,68.015380859375,67.94189453125,67.8318862915039,67.7896957397461,67.7740783691406,67.7430191040039,67.7134780883789,67.4441909790039,67.3860321044922,67.2669372558594,67.2184600830078,67.2096710205078,67.1214370727539,66.914306640625,66.826171875,66.4466247558594,66.2999496459961,66.1320266723633,66.0690383911133,66.064453125,66.0956039428711,66.1295928955078,66.2250518798828,66.2725601196289,66.2935562133789,66.3022918701172,66.3957977294922,66.4286651611328,66.6111297607422,66.5596160888672,66.5503463745117,66.6131820678711,66.6297836303711,66.6512222290039,66.7032699584961,66.7067337036133,66.7509765625,66.7846221923828,66.7643508911133,66.7645492553711,66.8439483642578,66.9083023071289,67.0215301513672,67.0611343383789,67.0868148803711,67.1526870727539,67.1614303588867,67.1298294067383,67.1132354736328,67.06787109375,67.0038528442383,66.9163055419922,66.8295440673828,66.7469253540039,66.7213821411133,66.7041015625,66.679931640625,66.6038589477539,66.5738754272461,66.5316467285156,66.4842758178711,66.4713897705078,66.442626953125,66.3845672607422,66.3468704223633,66.3295364379883,66.3156280517578,66.3209915161133,66.2252426147461,66.12841796875,65.9518585205078,65.8645553588867,65.8163604736328,65.7682571411133,65.71630859375,65.6640625,65.5099182128906,65.4566879272461,65.395751953125,65.2779312133789,65.1681137084961,64.9859848022461,64.9126968383789,64.8001937866211,64.7559585571289,64.7386703491211,64.7066955566406,64.5600051879883,64.4402313232422,64.362548828125,64.3467712402344,64.3783264160156,64.3353576660156,64.1890106201172,64.0343780517578,64.0028305053711,63.9450721740723,63.9095191955566,63.8167457580566,63.8133773803711,63.8934059143066,63.9491195678711,64.0258331298828,64.0910186767578,64.20703125,64.3201141357422,64.3663101196289,64.39697265625,64.3735809326172,64.3779296875,64.4084930419922,64.4891586303711,64.6852493286133,64.7505340576172,64.7909698486328,64.8473663330078,64.9386215209961,64.9354476928711,64.9871597290039,65.17236328125,65.1958999633789,65.1942825317383,65.1082534790039,64.8787612915039,64.8545913696289,64.8512191772461,64.8570785522461,64.8271484375,64.791259765625,64.7866668701172,64.7139205932617,64.570556640625,64.5770492553711,64.6563949584961,64.6905212402344,64.770751953125,64.7840270996094,64.7548828125,64.7787094116211,64.8962936401367,64.9980926513672,65.0632858276367,65.2547836303711,65.349853515625,65.4479522705078,65.5347137451172,65.5979461669922,65.751708984375,65.8437957763672,65.9634246826172,65.9878921508789,66.0210952758789,66.1234359741211,66.2593231201172,66.4659194946289,66.5196762084961,66.5147476196289,66.482421875,66.4225082397461,66.4113311767578,66.4209442138672,66.41552734375,66.3212890625,66.2912139892578,66.2509765625,66.1733856201172,66.1233901977539,66.146728515625,66.1584014892578,66.1423873901367,66.0986862182617,66.0497512817383,66.0085906982422,66.0645523071289,66.1127395629883,66.2350540161133,66.4070816040039,66.481689453125,66.6717758178711,66.7946243286133,66.937744140625,67.0041961669922,67.0996551513672,67.1673355102539,67.1886291503906,67.2544937133789,67.32958984375,67.4393081665039,67.670654296875,67.99560546875,68.0712432861328,68.1126022338867,68.1544418334961,68.2537612915039,68.3271026611328,68.6085433959961,68.6357955932617,68.6733856201172,68.6817398071289,68.6798324584961,68.548828125,68.541748046875,68.5781707763672,68.5465316772461,68.4796905517578,68.2913589477539,68.1188507080078,67.970458984375,67.8488235473633,67.8236846923828,67.8181610107422,67.757568359375,67.6888656616211,67.4774398803711,67.4131317138672,67.3507843017578,67.2847213745117,67.1855926513672,66.9891662597656,66.8910598754883,66.8531265258789,66.843505859375,66.8428192138672,66.8189926147461,66.8001937866211,66.8189926147461,66.8255386352539,66.9298400878906,66.9759292602539,67.0450210571289,67.275634765625,67.355712890625,67.4546890258789,67.5153350830078,67.5841827392578,67.650390625,67.6952590942383,67.6814956665039,67.7313461303711,67.8270034790039,67.8742218017578,67.8959426879883,67.8697280883789,67.8538131713867,67.8704071044922,68.0651397705078,68.1753387451172,68.2183609008789,68.3177261352539,68.3499526977539,68.3633270263672,68.4024353027344,68.476318359375,68.5387725830078,68.5413131713867,68.5320281982422,68.4593734741211,68.4186096191406,68.374267578125,68.3509292602539,68.3396987915039,68.3517074584961,68.3821258544922,68.4267578125,68.4840316772461,68.5061569213867,68.5924377441406,68.6102066040039,68.608154296875,68.731201171875,68.9125518798828,68.9959030151367,69.0033187866211,68.9923324584961,68.9647445678711,68.9266052246094,68.907470703125,68.8846740722656,68.8712463378906,68.8442840576172,68.8118667602539,68.801513671875,68.7089385986328,68.6339797973633,68.5369567871094,68.435546875,68.3826675415039,68.4025421142578,68.3670959472656,68.3432159423828,68.3116760253906,68.2674789428711,68.2568359375,68.2596664428711,68.231201171875,68.2273483276367,68.259033203125,68.2663040161133,68.2750930786133,68.2941360473633,68.2730484008789,68.1841812133789,68.2018585205078,68.3738250732422,68.4800262451172,68.5678253173828,68.5758743286133,68.6372985839844,68.6488800048828,68.6240692138672,68.6190414428711,68.5666961669922,68.5540008544922,68.6415023803711,68.8897476196289,68.8339385986328,68.9162139892578,69.0038070678711,69.0060577392578,68.9725646972656,68.8962860107422,68.849609375,68.7383804321289,68.7084426879883,68.6913070678711,68.6163024902344,68.4443359375,68.4002914428711,68.3511199951172,68.3516082763672,68.3803253173828,68.3960494995117,68.4219284057617,68.4713363647461,68.510498046875,68.6049346923828,68.7063446044922,68.6995086669922,68.7289581298828,68.7870101928711,68.8952178955078,68.9867706298828,69.1455078125,69.1102523803711,69.45703125,69.5909194946289,69.6526336669922,69.6923370361328,69.8211441040039,69.8471221923828,69.8514633178711,69.7630386352539,69.7431182861328,69.67529296875,69.53466796875,69.4356384277344,69.3253860473633,69.247802734375,69.2698287963867,69.2013626098633,69.1734390258789,69.13232421875,69.0770034790039,69.0363311767578,68.9613265991211,68.9478530883789,68.8919906616211,68.8735885620117,68.7539520263672,68.5792999267578,68.513671875,68.4036636352539,68.3142547607422,68.3484420776367,68.5674285888672,68.8179702758789,68.9506378173828,68.9561996459961,68.9173812866211,68.9273910522461,68.9670944213867,69.0675735473633,69.2362289428711,69.4207992553711,69.4800338745117,69.5299835205078,69.5844268798828,69.6937026977539,69.6555633544922,69.5966796875,69.5538101196289,69.6091842651367,69.6592254638672,69.7401351928711,69.8421859741211,70.0142517089844,70.005615234375,70.0306091308594,70.1080551147461,70.171630859375,70.2199249267578,70.2951202392578,70.5000991821289,70.7387161254883,70.7984313964844,70.8375549316406,70.79736328125,70.8185119628906,70.8646926879883,70.9005889892578,70.9623565673828,71.063720703125,71.0417022705078,71.0813980102539,71.139892578125,71.2823715209961,71.3478469848633,71.4120101928711,71.5483932495117,71.6828155517578,71.8526306152344,72.0127487182617,72.3915557861328,72.669921875,72.9555206298828,72.98193359375,72.9775390625,72.9563980102539,72.9318389892578,72.8975601196289,72.8849639892578,72.882568359375,72.9011688232422,72.890380859375,72.9136657714844,72.902099609375,72.8196792602539,72.8290023803711,72.7903289794922,72.7444763183594,72.69140625,72.4829559326172,72.3431625366211,72.0794448852539,72.0125503540039,71.8216323852539,71.6955032348633,71.6091766357422,71.5479507446289,71.5113754272461,71.4573745727539,71.3066864013672,71.151123046875,70.9632339477539,70.8228530883789,70.4573211669922,70.4034194946289,70.3455581665039,70.2749481201172,70.1725082397461,69.793212890625,69.4840316772461,69.37841796875,69.1542434692383,69.0805206298828,68.9687042236328,68.8748550415039,68.8152313232422,68.706787109375,68.5745086669922,68.5326156616211,68.4818878173828,68.4307632446289,68.2944793701172,68.1813430786133,68.0909194946289,67.9730453491211,67.8650360107422,67.7669448852539,67.6962356567383,67.5869598388672,67.0076217651367,66.939697265625,66.9615249633789,66.8789596557617,66.7604446411133,66.68310546875,66.6867141723633,66.6044921875,66.5481414794922,66.5194320678711,66.5486831665039,66.6024932861328,66.6476058959961,66.6682586669922,66.7008819580078,66.7453155517578,66.7541961669922,66.7537612915039,66.6973190307617,66.685791015625,66.7543487548828,66.8299789428711,66.845458984375,66.8145980834961,66.82861328125,66.81591796875,66.788330078125,66.766357421875,66.7235870361328,66.6407241821289,66.5786590576172,66.5107421875,66.4845733642578,66.4014205932617,66.3423843383789,66.3666534423828,66.3594284057617,66.333740234375,66.2467269897461,66.2533187866211,66.3321228027344,66.5065460205078,66.560791015625,66.8068389892578,66.861083984375,66.9953079223633,67.0849609375,67.3276901245117,67.4141082763672,67.6946258544922,67.766357421875,67.8975067138672,67.9859313964844,68.0735397338867,68.2183074951172,68.3030700683594,68.4206085205078,68.6588821411133,68.751220703125,68.8617248535156,68.9011688232422,68.9757308959961,68.9915008544922,68.978271484375,68.8976058959961,68.7769012451172,68.59619140625,68.4695816040039,68.3155746459961,67.9410171508789,67.7785110473633,67.7356414794922,67.6986846923828,67.6439437866211,67.589599609375,67.5702590942383,67.5591735839844,67.5784683227539,67.589111328125,67.6131362915039,67.6312026977539,67.6391143798828,67.6783676147461,67.7519073486328,67.9096221923828,68.0076675415039,68.1903839111328,68.2223587036133,68.2347183227539,68.2594680786133,68.3770523071289,68.4822769165039,68.6304702758789,68.9030303955078,68.9051284790039,68.9586410522461,69.1173858642578,69.2350616455078,69.2518081665039,69.2386245727539,69.1163101196289,69.090576171875,69.1445770263672,69.1146469116211,69.1432113647461,69.1982421875,69.41796875,69.50390625,69.6170959472656,69.7072219848633,69.802978515625,70.1756820678711,70.2728500366211,70.4454574584961,70.5787048339844,70.6536178588867,71.0686950683594,71.2165069580078,71.2635269165039,71.3197784423828,71.3852005004883,71.4449157714844,71.8450622558594,71.9147491455078,71.9578094482422,71.9970245361328,72.077392578125,72.1448211669922,72.19921875,72.2631378173828,72.420654296875,72.5487823486328,72.6194305419922,72.7101058959961,72.8118667602539,72.8384246826172,72.8538131713867,72.8527374267578,72.796630859375,72.6850051879883,72.5810546875,72.5121536254883,72.4572296142578,72.3822708129883,72.3499984741211,72.2962417602539,72.2535171508789,72.2323226928711,72.1707992553711,71.9832000732422,71.9589309692383,71.8133773803711,71.6546401977539,71.5343780517578,71.4946746826172,71.4300765991211,71.3784713745117,71.3417510986328,71.2659225463867,71.2185516357422,71.2020492553711,71.1278839111328,71.1810531616211,71.1679229736328,70.986328125,70.930419921875,70.9118194580078,70.9337921142578,70.9501953125,71.0019989013672,70.9971694946289,70.9735336303711,70.9759826660156,70.993896484375,71.025390625,71.0871047973633,71.2663116455078,71.3240737915039,71.3005828857422,71.3115768432617,71.4093780517578,71.4465789794922,71.552490234375,71.595458984375,71.682861328125,71.8290023803711,71.910400390625,71.9266128540039,72.0060043334961,72.0330047607422,72.0041961669922,71.8420867919922,71.83642578125,71.9070739746094,71.9522933959961,72.044677734375,72.092041015625,72.1143035888672,72.0717239379883,72.1077651977539,72.1565399169922,72.192138671875,72.201416015625,72.2291946411133,72.3287124633789,72.37744140625,72.3949737548828,72.3807678222656,72.2230453491211,72.1531295776367,72.0982894897461,72.0891647338867,72.0542984008789,71.97021484375,71.7461471557617,71.7159652709961,71.7068328857422,71.7585983276367,71.7641143798828,71.7486801147461,71.7205123901367,71.6681671142578,71.6021957397461,71.5624542236328,71.4513702392578,71.419921875,71.2928695678711,71.260009765625,71.0934600830078,71.0561981201172,70.9977035522461,70.8794403076172,70.807373046875,70.7067413330078,70.59814453125,70.511474609375,70.3956985473633,70.4302749633789,70.48291015625,70.5436019897461,70.6900863647461,70.8899383544922,70.9420852661133,70.954833984375,70.8954086303711,70.8152389526367,70.6947250366211,70.5805206298828,70.4074172973633,70.2864685058594,70.2177276611328,70.154052734375,70.1045455932617,70.0882797241211,70.093017578125,70.1095733642578,70.1571731567383,70.2210922241211,70.2767105102539,70.3213424682617,70.3452682495117,70.4183654785156,70.4664077758789,70.5464859008789,70.6722106933594,70.7659149169922,70.988525390625,71.1036148071289,71.27587890625,71.467529296875,71.5142593383789,71.5436477661133,71.5943832397461,71.6498031616211,71.6839370727539,71.8275375366211,71.8747100830078,71.90283203125,71.9252471923828,72.0718307495117,72.1051254272461,72.211181640625,72.237548828125,72.2654266357422,72.3266067504883,72.3517074584961,72.3588409423828,72.3897476196289,72.48828125,72.5199737548828,72.6479034423828,72.7120132446289,72.7591781616211,72.8607940673828,72.9491729736328,73.0252456665039,73.0491638183594,73.0860824584961,73.1552276611328,73.2311477661133,73.2896499633789,73.3568420410156,73.4137191772461,73.4740219116211,73.5149917602539,73.5684585571289,73.6404266357422,73.6588363647461,73.66650390625,73.6864776611328,73.7220230102539,73.7628402709961,73.7195281982422,73.7215270996094,73.734619140625,73.8215789794922,73.85693359375,73.894287109375,73.8871078491211,73.8607406616211,73.8241653442383,73.7168426513672,73.6197738647461,73.5783233642578,73.4927749633789,73.4589385986328,73.4383316040039,73.3716812133789,73.3269500732422,73.2726058959961,73.1957473754883,73.1404800415039,73.1067886352539,73.1258316040039,73.3067398071289,73.3470687866211,73.3904266357422,73.4564895629883,73.48583984375,73.5191421508789,73.53466796875,73.56884765625,73.6150436401367,73.7046890258789,73.7559051513672,73.8107376098633,73.8324737548828,73.8460464477539,73.878662109375,74.1953125,74.2437515258789,74.2793960571289,74.3160171508789,74.4230422973633,74.4501037597656,74.4442367553711,74.4142532348633,74.3253402709961,74.3638687133789,74.403564453125,74.44970703125,74.5224609375,74.5854949951172,74.6285629272461,74.6451187133789,74.740234375,74.7282257080078,74.7574234008789,74.8162155151367,74.6824188232422,74.7178726196289,74.7788543701172,74.94091796875,75.0132369995117,75.0525360107422,75.072265625,75.068115234375,75.11279296875,75.1698226928711,75.1917495727539,75.1295852661133,75.2904815673828,75.3691940307617,75.4701156616211,75.458251953125,75.591064453125,75.6495666503906,75.649658203125,75.7236862182617,75.7496566772461,75.7791061401367,75.8541030883789,75.9128875732422,75.959228515625,75.9215850830078,75.956298828125,75.932861328125,75.9012680053711,75.958984375,75.9446258544922,75.912841796875,75.9026870727539,75.9099578857422,75.9794921875,76.0750961303711,76.0258331298828,76.0987777709961,76.1007308959961,76.0541458129883,76.101318359375,76.1235885620117,76.102783203125,76.1079559326172,76.1517562866211,76.113525390625,76.1396026611328,76.1373062133789,76.1131362915039,76.0819854736328,76.0096740722656,75.8921890258789,75.872314453125,75.926025390625,76.0055694580078,75.9898910522461,75.921630859375,75.8912048339844,75.9310531616211,76.0186996459961,76.0334014892578,75.980224609375,76.029052734375,76.0780258178711,76.0886688232422,76.1336975097656,76.1664047241211,76.1805725097656,76.24267578125,76.2240219116211,76.2075729370117,76.1776428222656,76.1093215942383,76.0823211669922,76.0780258178711,76.0287628173828,75.9563446044922,75.85205078125,75.8031768798828,75.798583984375,75.811279296875,75.8806610107422,75.9302673339844,76.1359329223633,76.2401809692383,76.2751007080078,76.3843307495117,76.4308090209961,76.4806594848633,76.5095748901367,76.471435546875,76.4898986816406,76.4791488647461,76.5251922607422,76.4772491455078,76.4789047241211,76.439208984375,76.4854965209961,76.5356903076172,76.530517578125,76.5567321777344,76.6150894165039,76.7040023803711,76.7813491821289,76.822509765625,76.9006805419922,76.990478515625,77.028564453125,77.1015625,77.1980972290039,77.508544921875,77.62646484375,77.6410598754883,77.6319351196289,77.7304229736328,77.73046875,77.652099609375,77.5947265625,77.5492172241211,77.5252456665039,77.4888687133789,77.4476089477539,77.3905258178711,77.3520050048828,77.2378387451172,77.1747131347656,77.1326675415039,77.101806640625,77.0923309326172,77.1006851196289,77.00146484375,76.9975051879883,77.0453109741211,77.0478515625,77.0137710571289,77.0317916870117,77.0343780517578,76.990966796875,76.9265594482422,76.822021484375,76.73046875,76.5733871459961,76.5862808227539,76.5894546508789,76.5122604370117,76.5146942138672,76.4800796508789,76.5240707397461,76.5101089477539,76.5223083496094,76.5696792602539,76.66064453125,76.7184524536133,76.7378463745117,76.7195358276367,76.7201232910156,76.7492141723633,76.7118606567383,76.7583999633789,76.7230453491211,76.6866760253906,76.622314453125,76.6035614013672,76.5534133911133,76.4803161621094,76.4205551147461,76.3804702758789,76.4239807128906,76.4346694946289,76.4083023071289,76.383544921875,76.2188491821289,76.1869125366211,76.129638671875,76.0771942138672,76.0535659790039,76.05859375,76.1141128540039,76.1329116821289,76.1745147705078,76.2152328491211,76.23974609375,76.2581024169922,76.2516555786133,76.1788558959961,76.1121139526367,75.8916549682617,75.9212875366211,75.8560028076172,75.7047882080078,75.5926742553711,75.5684051513672,75.56396484375,75.6218719482422,75.6566848754883,75.6778793334961,75.6986770629883,75.8353958129883,75.8498992919922,75.8436508178711,75.8301773071289,75.7376480102539,75.5719223022461,75.6205062866211,75.6114273071289,75.5342788696289,75.5020523071289,75.4506378173828,75.29296875,75.0150375366211,74.8531799316406,74.7400436401367,74.658447265625,74.548095703125,74.466064453125,74.378662109375,74.3219757080078,74.2930679321289,74.2613296508789,74.2088851928711,74.169189453125,74.0888137817383,74.0323257446289,73.694091796875,73.625,73.6210479736328,73.5894088745117,73.3766632080078,73.3306655883789,73.3080139160156,72.9592742919922,72.8410186767578,72.7770538330078,72.7899398803711,72.836669921875,72.9498596191406,73.0020065307617,73.060546875,73.1063995361328,73.139404296875,73.1772918701172,73.1631317138672,73.1731414794922,73.235595703125,73.2579116821289,73.265869140625,73.3102035522461,73.3190383911133,73.37841796875,73.3996124267578,73.4874496459961,73.4540023803711,73.4724578857422,73.62890625,73.6891632080078,73.730712890625,73.759765625,73.7799301147461,73.7260284423828,73.7085494995117,73.7225570678711,73.7437515258789,73.8002395629883,73.8812484741211,73.9306182861328,73.994384765625,74.0174331665039,73.9479064941406,73.9393539428711,74.0528259277344,74.04736328125,74.0285110473633,74.0048370361328,73.9685516357422,73.8848648071289,73.8277359008789,73.7452621459961,73.7089385986328,73.7111358642578,73.74609375,73.7711486816406,73.8356399536133,73.9620590209961,73.9457015991211,73.9138717651367,73.83740234375,73.7074203491211,73.6476058959961,73.582763671875,73.4595718383789,73.3915023803711,73.3460922241211,73.1451187133789,73.0478515625,72.94189453125,72.897216796875,72.8306655883789,72.7694854736328,72.7301788330078,72.6573715209961,72.634521484375,72.6541519165039,72.6771011352539,72.7110366821289,72.7388610839844,72.8058624267578,72.8783264160156,72.9321823120117,73.0543441772461,73.1422348022461,73.2326202392578,73.2735900878906,73.3179702758789,73.3265609741211,73.3458023071289,73.367431640625,73.3785629272461,73.4336471557617,73.5049819946289,73.5334014892578,73.5846710205078,73.607177734375,73.7025909423828,73.6760787963867,73.5991668701172,73.5897979736328,73.5378952026367,73.5183563232422,73.481201171875,73.4645004272461,73.4644012451172,73.3672332763672,73.2465286254883,73.1172866821289,73.06396484375,72.9791030883789,72.9713363647461,72.9811019897461,72.9367218017578,72.9708557128906,72.9696807861328,72.9608840942383,72.897216796875,72.8805618286133,72.8777847290039,72.8908233642578,72.906494140625,72.9312973022461,72.9706573486328,73.0166931152344,73.0279312133789,72.9646530151367,72.9548797607422,73.0018081665039,73.0856475830078,73.1441879272461,73.1729049682617,73.1773452758789,73.1932601928711,73.2616195678711,73.3473129272461,73.4024963378906,73.4308090209961,73.5329055786133,73.6368637084961,73.6663589477539,73.6267547607422,73.6893081665039,73.7123031616211,73.7548370361328,73.7512664794922,73.7117691040039,73.5206069946289,73.4474182128906,73.4684600830078,73.4980926513672,73.5174789428711,73.5481948852539,73.5366744995117,73.5062942504883,73.4636688232422,73.4197692871094,73.3941879272461,73.3887710571289,73.3349151611328,73.4341812133789,73.5282211303711,73.5474624633789,73.4815444946289,73.445556640625,73.4256362915039,73.3907699584961,73.3523941040039,73.33056640625,73.3007278442383,73.2674789428711,73.2624053955078,73.2333984375,73.1902389526367,73.1393508911133,73.110595703125,73.1075134277344,73.1123504638672,73.04541015625,72.9726104736328,72.9432678222656,72.8951644897461,72.8858947753906,72.8724594116211,72.7757339477539,72.7051773071289,72.6769561767578,72.5858917236328,72.5501480102539,72.5473098754883,72.53515625,72.495849609375,72.4857482910156,72.4376907348633,72.3154830932617,72.1663131713867,72.092041015625,72.0794906616211,72.2455520629883,72.3096237182617,72.4340362548828,72.4131774902344,72.3082504272461,72.25,72.0883331298828,71.7553253173828,71.7824172973633,71.8246078491211,71.878662109375,71.9532241821289,71.9169387817383,71.8501968383789,71.7393035888672,71.7448272705078,71.7075653076172,71.6634750366211,71.6017608642578,71.5928649902344,71.5088348388672,71.4048843383789,71.1195297241211,71.0653762817383,70.9473114013672,70.8925247192383,70.8883285522461,70.9623565673828,70.9358825683594,70.8035659790039,70.74609375,70.7421875,70.7655258178711,70.8282699584961,70.9010238647461,71.1014175415039,71.20263671875,71.2440414428711,71.293212890625,71.3501968383789,71.4839782714844,71.6427764892578,71.7262191772461,71.8953170776367,71.9259796142578,71.8714828491211,71.7987289428711,71.767578125,71.7551727294922,71.606689453125,71.490966796875,71.4342269897461,71.3789520263672,71.3868179321289,71.4605941772461,71.5150375366211,71.5435028076172,71.6103515625,71.6305618286133,71.6195831298828,71.5707550048828,71.4156723022461,71.3594284057617,71.2990264892578,71.2081604003906,71.1639175415039,71.1333999633789,71.1427230834961,71.1686553955078,71.1940460205078,71.226806640625,71.2608413696289,71.2858352661133,71.3074188232422,71.3255310058594,71.3585891723633,71.3634262084961,71.3840866088867,71.4297866821289,71.4635238647461,71.5259704589844,71.566162109375,71.5963363647461,71.6028366088867,71.562744140625,71.6348114013672,71.6290054321289,71.5560531616211,71.4447784423828,71.4447250366211,71.4892578125,71.4915008544922,71.5576629638672,71.700439453125,71.8849639892578,71.9267044067383,71.9513626098633,71.9983444213867,72.1485824584961,72.162109375,72.1913070678711,72.2096176147461,72.2256851196289,72.2076644897461,72.1634750366211,72.1634750366211,72.264404296875,72.3297348022461,72.4665069580078,72.49609375,72.4931106567383,72.5189437866211,72.5869140625,72.6300354003906,72.7169952392578,72.8428192138672,72.8716278076172,72.8900375366211,72.8909683227539,72.8577194213867,72.7886734008789,72.7208023071289,72.6982421875,72.6731948852539,72.6430130004883,72.6099090576172,72.5702133178711,72.5420837402344,72.4973602294922,72.4713897705078,72.4422378540039,72.3497085571289,72.362060546875,72.3926773071289,72.396240234375,72.3822708129883,72.3055191040039,72.2653350830078,72.2587890625,72.192626953125,72.1747589111328,72.2598648071289,72.3024444580078,72.29541015625,72.236572265625,72.12353515625,72.0354995727539,71.9449691772461,71.9454574584961,72.1374969482422,72.146484375,72.1422882080078,72.2218704223633,72.2258758544922,72.2063522338867,72.1775894165039,72.0669937133789,72.0206527709961,71.9413070678711,71.89013671875,71.8946228027344,71.9271545410156,71.9260711669922,71.9010238647461,71.8546447753906,71.8308563232422,71.793701171875,71.7533721923828,71.7073745727539,71.69580078125,71.7464828491211,71.808349609375,71.9220657348633,72.197509765625,72.2920379638672,72.327880859375,72.3409194946289,72.3119659423828,72.2523422241211,72.164306640625,72.0912628173828,71.9921875,71.9504928588867,71.8956604003906,71.843017578125,71.8255386352539,71.7957534790039,71.7627944946289,71.744140625,71.7146530151367,71.6904754638672,71.6879348754883,71.6640090942383,71.6014709472656,71.5807113647461,71.5213317871094,71.5108337402344,71.5201187133789,71.4988784790039,71.4552230834961,71.3858413696289,71.3388214111328,71.2265625,71.2671813964844,71.3628921508789,71.3804702758789,71.3737258911133,71.2869644165039,71.2178192138672,71.0232925415039,71.0024871826172,70.9824676513672,70.83447265625,70.8356475830078,70.8786163330078,70.8799819946289,70.9744644165039,71.0342254638672,71.0386276245117,71.0955047607422,71.09375,71.0745086669922,71.0392608642578,70.9350128173828,70.7907257080078,70.649658203125,70.6049270629883,70.506103515625,70.4236373901367,70.3096694946289,70.2213897705078,70.1587905883789,70.0814437866211,69.989990234375,69.8702087402344,69.7849578857422,69.7297821044922,69.6551742553711,69.6063461303711,69.4585418701172,69.33447265625,69.0981979370117,69.0388717651367,68.9822769165039,68.9051742553711,68.6538543701172,68.6075210571289,68.538330078125,68.5625,68.6539077758789,68.822998046875,68.849853515625,68.9051742553711,69.06396484375,69.2017059326172,69.3000946044922,69.3795394897461,69.5451126098633,69.611572265625,69.6490707397461,69.6827621459961,69.7147445678711,69.6932601928711,69.7018051147461,69.7351608276367,69.7192840576172,69.609130859375,69.5844268798828,69.5459976196289,69.4995574951172,69.4999008178711,69.554443359375,69.7403259277344,69.7282180786133,69.6991729736328,69.6256332397461,69.577392578125,69.447021484375,69.271484375,69.239501953125,69.2283706665039,69.163330078125,69.0795440673828,68.9196243286133,68.7860336303711,68.7986755371094,68.8253936767578,69.0453109741211,69.1347198486328,69.2636260986328,69.3882369995117,69.5833511352539,69.6265563964844,69.6832046508789,69.7509765625,69.8565444946289,69.9378890991211,70.1075668334961,70.0960464477539,70.0761184692383,70.0003433227539,69.9683609008789,69.9197692871094,69.8649368286133,69.8238296508789,69.9240188598633,69.9468231201172,69.89111328125,69.8741226196289,69.8816375732422,69.8556671142578,69.8600540161133,69.9041519165039,69.8953170776367,69.8602981567383,69.7685012817383,69.64599609375,69.6116180419922,69.49560546875,69.4529800415039,69.38720703125,69.3621063232422,69.3259735107422,69.2957992553711,69.2596664428711,69.0126953125,68.9834518432617,68.9404296875,68.906494140625,68.9171447753906,68.9123992919922,68.8529815673828,68.8251953125,68.7541046142578,68.6751480102539,68.5856475830078,68.6030731201172,68.6604461669922,68.593017578125,68.5459976196289,68.5017547607422,68.4666519165039,68.4248046875,68.388427734375,68.32275390625,68.264892578125,68.2865219116211,68.3379898071289,68.3782272338867,68.5185546875,68.565673828125,68.5414047241211,68.36279296875,68.2943878173828,68.2811584472656,68.2412109375,68.2242202758789,68.2368698120117,68.2451705932617,68.2225112915039,68.1746597290039,68.119140625,67.6780776977539,67.60205078125,67.5665054321289,67.5210952758789,67.4466781616211,67.3573684692383,67.3653869628906,67.376953125,67.4134292602539,67.4375,67.4075698852539,67.348876953125,67.2685012817383,67.2034683227539,67.093017578125,67.000244140625,66.9613800048828,66.9168014526367,66.7843246459961,66.7249069213867,66.6231460571289,66.6131362915039,66.6034164428711,66.5854034423828,66.5379409790039,66.4921875,66.4298782348633,66.3719711303711,66.34423828125,66.3483428955078,66.428466796875,66.4523468017578,66.4730911254883,66.3825225830078,66.2295913696289,66.2296905517578,66.2457962036133,66.2867660522461,66.3104934692383,66.3660659790039,66.4346694946289,66.48828125,66.5217819213867,66.5406265258789,66.5807113647461,66.6318817138672,66.6526412963867,66.6897964477539,66.7786178588867,66.827392578125,66.87548828125,66.94287109375,66.9820327758789,67.0015640258789,67.0397491455078,67.0376434326172,67.049072265625,67.0630416870117,67.0906295776367,67.1031265258789,67.1064453125,67.144775390625,67.1327590942383,67.1051712036133,67.069091796875,67.05224609375,67.0351104736328,66.954833984375,66.9092330932617,66.8513641357422,66.840087890625,66.8646011352539,66.9112319946289,66.9685592651367,66.9936065673828,66.9989700317383,66.9558563232422,66.942138671875,66.9250030517578,66.9305191040039,66.9524917602539,66.9778289794922,67.0339813232422,67.0648880004883,67.0268020629883,66.9917449951172,66.965576171875,66.9732894897461,66.9317321777344,66.8187026977539,66.6767578125,66.5927276611328,66.5297393798828,66.3572311401367,66.3436508178711,66.3202590942383,66.2910614013672,66.2489318847656,66.278076171875,66.2978973388672,66.2940444946289,66.2718811035156,66.2364273071289,66.2012634277344,66.1692886352539,66.1634750366211,66.1431121826172,66.05810546875,65.9989242553711,66.006103515625,66.031005859375,66.0335006713867,66.008056640625,65.9285202026367,65.8654251098633,65.8235855102539,65.7102508544922,65.65625,65.6215362548828,65.6426239013672,65.6648941040039,65.6950225830078,65.7368621826172,65.803955078125,65.8103561401367,65.7942352294922,65.7517623901367,65.698486328125,65.6280746459961,65.5499572753906,65.5110321044922,65.5020980834961,65.5027847290039,65.5271987915039,65.5331039428711,65.5104446411133,65.4959487915039,65.5079650878906,65.5420913696289,65.5669403076172,65.5704574584961,65.5823211669922,65.6175308227539,65.6696243286133,65.6900405883789,65.6924285888672,65.6810531616211,65.6120147705078,65.4959945678711,65.4745635986328,65.4495620727539,65.4477996826172,65.4557113647461,65.4251937866211,65.302734375,65.275634765625,65.24853515625,65.2282257080078,65.2218704223633,65.2267074584961,65.2057113647461,65.1287078857422,65.0481491088867,65.0021438598633,64.9647445678711,64.907958984375,64.8829116821289,64.8892059326172,64.8766098022461,64.84716796875,64.8173370361328,64.8371047973633,64.8260803222656,64.79052734375,64.7611846923828,64.7291946411133,64.7049331665039,64.6640167236328,64.6288528442383,64.6099166870117,64.603271484375,64.577880859375,64.544189453125,64.5153274536133,64.4746551513672,64.4315414428711,64.4139175415039,64.4070816040039,64.4122543334961,64.4599609375,64.4989242553711,64.5260772705078,64.5073699951172,64.3694305419922,64.3276824951172,64.2974624633789,64.2797393798828,64.2896499633789,64.3548812866211,64.410400390625,64.4426803588867,64.4874496459961,64.53955078125,64.47900390625,64.4286117553711,64.365478515625,64.3573303222656,64.364501953125,64.4097137451172,64.448974609375,64.5777893066406,64.6376419067383,64.7177734375,64.7759780883789,64.8136749267578,64.8092727661133,64.7939987182617,64.8023910522461,64.8166961669922,64.8485794067383,64.8670883178711,64.9460983276367,65.0108413696289,65.05419921875,65.1055221557617,65.2328109741211,65.3524932861328,65.4710464477539,65.5475540161133,65.6013641357422,65.6136245727539,65.6016616821289,65.5037155151367,65.4897003173828,65.48486328125,65.4955596923828,65.5372085571289,65.593017578125,65.6966323852539,65.7404251098633,65.7552185058594,65.7540054321289,65.7953643798828,65.86474609375,65.9364776611328,66.0327682495117,66.0375442504883,66.013671875,66.0372543334961,66.1242141723633,66.166015625,66.1984405517578,66.3165512084961,66.4015655517578,66.3551788330078,66.2372589111328,66.2026901245117,66.1870574951172,66.179931640625,66.2035140991211,66.23193359375,66.3460922241211,66.3750457763672,66.3533172607422,66.3125534057617,66.3050765991211,66.2874984741211,66.2198257446289,66.16259765625,66.1410675048828,66.1278839111328,66.1841278076172,66.1057586669922,66.0179748535156,65.90087890625,65.8038101196289,65.757568359375,65.6878433227539,65.6386260986328,65.5752487182617,65.5167465209961,65.4453125,65.3862838745117,65.244140625,65.1872100830078,65.0672378540039,65.0341796875,64.9209518432617,64.822021484375,64.7815933227539,64.631103515625,64.6029739379883,64.6722640991211,64.717041015625,64.7778854370117,64.9313507080078,64.95361328125,65.0141143798828,65.0819320678711,65.03759765625,65.0712432861328,65.0473098754883,65.0252456665039,65.0071792602539,65.0160140991211,64.9996566772461,64.947021484375,64.8616714477539,64.8048324584961,64.7866668701172,64.8492202758789,64.8399887084961,64.8551712036133,64.9608917236328,64.8440399169922,64.78369140625,64.77685546875,64.6838836669922,64.6814498901367,64.7466278076172,64.7466278076172,64.782470703125,64.8252944946289,64.8651885986328,64.9047317504883,64.884765625,64.843017578125,64.7838363647461,64.7051315307617,64.663818359375,64.6419448852539,64.5858383178711,64.6824264526367,64.6248550415039,64.6337890625,64.71923828125,64.7740249633789,64.7633819580078,64.73681640625,64.5727996826172,64.4444808959961,64.3047332763672,64.2222671508789,64.2195816040039,64.2352523803711,64.30908203125,64.3644027709961,64.3144073486328,64.2608871459961,64.1278839111328,64.0890121459961,64.0113754272461,63.9756355285645,63.9652824401855,63.8423309326172,63.6670875549316,63.6507301330566,63.6507301330566,63.6055679321289,63.5740699768066,63.5566368103027,63.5215301513672,63.4399452209473,63.4022979736328,63.394775390625,63.4424324035645,63.51025390625,63.5403327941895,63.4002456665039,63.3450202941895,63.2824211120605,63.190185546875,63.147216796875,63.0777320861816,63.0579109191895,63.0083045959473,62.9398460388184,62.8836936950684,62.86279296875,62.7734870910645,62.6874961853027,62.6130905151367,62.5103492736816,62.4691886901855,62.3964347839355,62.3203620910645,62.3236808776855,62.3552780151367,62.5469741821289,62.5828094482422,62.5874481201172,62.5990257263184,62.6444816589355,62.6852569580078,62.7369613647461,62.7813491821289,62.7842254638672,62.7504386901855,62.7503395080566,62.7895545959473,62.7772445678711,62.7222137451172,62.6932601928711,62.6586456298828,62.6265640258789,62.5916023254395,62.5609893798828,62.5360832214355,62.5057601928711,62.4108390808105,62.3460426330566,62.1843795776367,62.1279258728027,62.1213340759277,62.1023941040039,62.0344200134277,61.9388656616211,61.9479026794434,61.8676300048828,61.8236351013184,61.8175277709961,61.795166015625,61.6793937683105,61.716064453125,61.5567398071289,61.4066390991211,61.4062004089355,61.4691886901855,61.4361305236816,61.3755874633789,61.3116226196289,61.2930641174316,61.3106956481934,61.3144035339355,61.2951622009277,61.2493171691895,61.1904296875,61.1858863830566,61.1673851013184,61.1166000366211,61.061767578125,60.9978523254395,60.9156723022461,60.9006843566895,60.8641090393066,60.8373565673828,60.8431129455566,60.7257308959961,60.52294921875,60.4964828491211,60.4349136352539,60.3936996459961,60.3428688049316,60.259521484375,60.0478057861328,60.0097618103027,59.9655303955078,59.9860801696777,60.0670890808105,60.1042518615723,60.1478538513184,60.2179183959961,60.2502479553223,60.265380859375,60.4380340576172,60.556640625,60.5959510803223,60.5638198852539,60.5628929138184,60.5922355651855,60.5739288330078,60.5093307495117,60.4689407348633,60.4062957763672,60.3070335388184,59.947021484375,59.8724136352539,59.8562469482422,59.8494644165039,59.9220695495605,59.9793472290039,60.0888214111328,60.1783180236816,60.3460960388184,60.4142608642578,60.4848136901855,60.4803695678711,60.3568840026855,60.2364768981934,60.2051734924316,60.1349105834961,60.124755859375,60.0985832214355,59.9456062316895,59.8607406616211,59.8436050415039,59.8409690856934,59.8742179870605,59.9974594116211,60.0612831115723,60.0727005004883,60.0580596923828,59.9737815856934,59.8975563049316,59.984375,60.0173377990723,60.0370635986328,60.0411109924316,60.0280265808105,59.9784164428711,59.9140625,59.8867721557617,59.8349647521973,59.7814407348633,59.7054214477539,59.5200157165527,59.3025856018066,59.1314010620117,59.1482925415039,59.1370620727539,59.1143035888672,59.0201644897461,58.9864768981934,58.9639625549316,58.9392585754395,58.7999038696289,58.7085914611816,58.4474105834961,58.2728500366211,58.0769081115723,57.98095703125,57.9182624816895,57.8746566772461,57.8291473388672,57.7783660888672,57.7450218200684,57.7172355651855,57.7662086486816,57.9041023254395,57.9482383728027,57.9460945129395,57.8373031616211,57.7903823852539,57.6868133544922,57.6374053955078,57.5648422241211,57.4774894714355,57.3576202392578,57.2840843200684,57.2439460754395,57.102783203125,57.0233879089355,56.8753929138184,56.8114738464355,56.7568359375,56.72265625,56.7413063049316,56.7254905700684,56.6880378723145,56.5645484924316,56.4477043151855,56.2325210571289,56.1737289428711,56.1370124816895,56.044677734375,56.0337905883789,56.0656242370605,56.2322731018066,56.3308563232422,56.3994636535645,56.4490242004395,56.5218734741211,56.508056640625,56.4763679504395,56.4900856018066,56.4549331665039,56.3991241455078,56.2606964111328,56.2355003356934,56.187744140625,56.1282730102539,56.0896453857422,55.8403816223145,55.6548347473145,55.4961433410645,55.3580055236816,55.205322265625,55.138916015625,54.9979972839355,54.8861312866211,54.7521476745605,54.6886711120605,54.532958984375,54.5162620544434,54.5205574035645,54.5982398986816,54.578369140625,54.5413589477539,54.4308586120605,54.2882308959961,54.1891593933105,54.130859375,54.0083999633789,53.7836418151855,53.6726570129395,53.620849609375,53.5521965026855,53.4477043151855,53.3807640075684,53.27490234375,53.1295890808105,53.1250991821289,53.2296905517578,53.2376937866211,53.1171379089355,53.0475578308105,52.908935546875,52.9354019165039,53.0147933959961,53.0499992370605,53.0323715209961,52.9574203491211,52.9221687316895,52.8736305236816,52.6884269714355,52.6266593933105,52.4603042602539,52.3831558227539,52.3049812316895,52.0908699035645,51.8096160888672,51.6053237915039,51.5345687866211,51.4798851013184,51.408935546875,51.2127456665039,51.006591796875,50.9692878723145,51.0470733642578,51.1241188049316,51.2268562316895,51.3116226196289,51.3802719116211,51.47509765625,51.9130363464355,52.3665504455566,52.5093727111816,52.6262702941895,52.7472648620605,52.8661613464355,53.0064964294434,53.7442855834961,53.9281234741211,54.521484375,54.8645515441895,55.1991195678711,55.3484840393066,55.7935562133789,56.0722160339355,56.6952133178711,56.7520027160645,56.7815933227539,57.0211906433105,57.1522483825684,57.2901840209961,57.46630859375,57.5609359741211,57.6157722473145,57.6769027709961,57.7479515075684,57.7796363830566,57.8036575317383,57.8301734924316,57.7768096923828,57.7992668151855,58.0197715759277,57.9859352111816,58.0252914428711,58.0089797973633,58.0834503173828,58.1628456115723,58.2813491821289,58.4239273071289,58.5194358825684,58.6105499267578,58.6959495544434,58.8036613464355,59.1271514892578,59.3940391540527,59.54736328125,59.6016578674316,59.6268539428711,59.8456039428711,60.02734375,60.1522941589355,60.2322273254395,60.420166015625,60.4664077758789,60.5367164611816,60.6594696044922,60.7829132080078,60.8004379272461,60.8497581481934,60.8771476745605,60.9167976379395,61.025634765625,61.0843772888184,61.111328125,61.240478515625,61.3437995910645,61.3882293701172,61.4198760986328,61.4613761901855,61.5582542419434,61.64013671875,61.7106895446777,61.8738784790039,62.0450210571289,62.2922325134277,62.3466300964355,62.4705543518066,62.4737815856934,62.431884765625,62.4115219116211,62.3739776611328,62.4057579040527,62.4481925964355,62.4629898071289,62.4470672607422,62.493896484375,62.5169944763184,62.5710983276367,62.6754875183105,62.7046394348145,62.6965789794922,62.5509262084961,62.5114288330078,62.4553718566895,62.3730010986328,62.3369178771973,62.3134269714355,62.2595710754395,62.1529312133789,62.0499038696289,61.8910636901855,61.79150390625,61.736572265625,61.699462890625,61.6447715759277,61.6114273071289,61.570556640625,61.5540542602539,61.544189453125,61.5977058410645,61.7050285339355,61.7112770080566,61.6951179504395,61.652587890625,61.6500511169434,61.6701202392578,61.662109375,61.5406723022461,60.9628868103027,60.8926734924316,60.7533187866211,60.7398414611816,60.7085456848145,60.6670379638672,60.638427734375,60.6907234191895,60.7296371459961,60.83154296875,61.0254859924316,61.0447731018066,61.0476570129395,61.0074195861816,60.9434089660645,60.9566383361816,61.0139656066895,61.1286163330078,61.2344703674316,61.2917938232422,61.3239250183105,61.5374984741211,61.6476058959961,61.7933578491211,61.8385734558105,61.8943824768066,61.9038543701172,61.90283203125,61.7583999633789,61.719482421875,61.7814445495605,61.8080558776855,61.9141616821289,61.9293937683105,61.9222640991211,61.8502464294434,61.8108901977539,61.82568359375,61.7648468017578,61.7536125183105,61.7952613830566,61.7989234924316,61.7470703125,61.6756858825684,61.565185546875,61.5296363830566,61.4806175231934,61.2724609375,61.2060089111328,61.1550750732422,60.99560546875,60.7771492004395,60.682373046875,60.5498542785645,60.3766593933105,60.0950164794922,59.8837852478027,59.8767585754395,59.8333511352539,59.7303733825684,59.6583976745605,59.600341796875,59.5285148620605,59.4833984375,59.4751510620117,59.4814453125,59.5400924682617,59.4496116638184,59.3601570129395,59.2702178955078,59.1901397705078,59.1956024169922,59.1875495910645,59.1413116455078,59.216552734375,59.1878433227539,59.1085929870605,59.0755348205566,59.1141586303711,59.2247543334961,59.2147979736328,59.0913047790527,59.0944328308105,59.0818862915039,58.9390602111816,58.92626953125,58.9541015625,59.0264205932617,59.0307655334473,58.9970207214355,58.9104499816895,58.8667030334473,58.8750953674316,59.0825233459473,59.1640129089355,59.1466789245605,59.1600608825684,59.2235870361328,59.2779273986816,59.2905769348145,59.2840805053711,59.3232383728027,59.5241203308105,59.5611343383789,59.583251953125,59.5856399536133,59.5713386535645,59.5230484008789,59.4754371643066,59.4607429504395,59.4691390991211,59.5065422058105,59.494384765625,59.5249481201172,59.5563468933105,59.5907249450684,59.6388702392578,59.6512718200684,59.7704124450684,59.760986328125,59.7284660339355,59.6305160522461,59.5587921142578,59.5267601013184,59.4881820678711,59.4805183410645,59.4749984741211,59.5323257446289,59.4485321044922,59.37353515625,59.3999977111816,59.3691368103027,59.28271484375,59.2579078674316,59.2623062133789,59.4142112731934,59.3880386352539,59.2906723022461,59.2685585021973,59.36572265625,59.3729476928711,59.4569816589355,59.4304656982422,59.2214813232422,59.1705589294434,59.1983909606934,59.330322265625,59.3737297058105,59.4135284423828,59.3762702941895,59.4083023071289,59.411376953125,59.3436546325684,59.3701133728027,59.2401351928711,59.1526336669922,58.9996604919434,58.7452621459961,58.6490249633789,58.528076171875,58.4168472290039,58.3034629821777,58.212158203125,57.8654327392578,57.8136711120605,57.6874961853027,57.54931640625,57.51416015625,57.4557151794434,57.3583030700684,57.3296852111816,57.2615242004395,57.088134765625,56.9655265808105,56.6290016174316,56.5885276794434,56.4986839294434,56.43310546875,56.1393585205078,56.1121101379395,55.9747543334961,55.8922843933105,55.7952613830566,55.6941871643066,55.5767097473145,55.510009765625,55.3522453308105,55.16064453125,55.11376953125,54.9433135986328,54.9032211303711,54.8408203125,54.7314949035645,54.7074203491211,54.6924819946289,54.5839347839355,54.6140632629395,54.6136207580566,54.6243133544922,54.6209945678711,54.5614738464355,54.4523429870605,54.3533210754395,54.0606422424316,53.9313011169434,53.8041000366211,53.781982421875,53.83935546875,53.84814453125,53.8216781616211,54.0252418518066,54.056884765625,54.1285667419434,54.1822280883789,54.2823257446289,54.2912101745605,54.2833023071289,54.1563949584961,54.1304664611816,54.12353515625,54.1005401611328,54.027587890625,53.970458984375,53.9467277526855,53.9033203125,53.8658180236816,53.7070770263672,53.631591796875,53.5792007446289,53.546142578125,53.5389633178711,53.560302734375,53.6035652160645,53.7264137268066,53.9092750549316,53.9596710205078,53.9598655700684,53.9471664428711,53.8188018798828,53.6741714477539,53.5924301147461,53.5240211486816,53.5228996276855,53.5370101928711,53.5700187683105,53.7447738647461,53.8697242736816,54.0437507629395,54.1476554870605,54.22265625,54.29833984375,54.3200187683105,54.2178230285645,54.1929664611816,54.2771492004395,54.2564430236816,54.205322265625,54.0515632629395,54.0010261535645,53.8125991821289,53.596435546875,53.4945793151855,53.4542503356934,53.33447265625,53.2927742004395,53.1839866638184,53.0972671508789,53.0152816772461,53.0915031433105,53.087890625,53.0398445129395,52.8975563049316,52.8401374816895,52.652587890625,52.5501480102539,52.4356918334961,52.368408203125,52.2711448669434,52.2343254089355,52.1785163879395,52.0572242736816,51.920654296875,51.8606948852539,51.727783203125,51.6199226379395,51.4141616821289,51.2322769165039,51.0513153076172,50.9867668151855,50.8001937866211,50.5459938049316,50.1307601928711,50.106689453125,50.0824241638184,50.0537109375,50.0333480834961,49.9114723205566,49.8255844116211,49.7616691589355,49.5961418151855,49.3314971923828,49.289794921875,49.2208480834961,49.1591796875,49.1200180053711,49.0539054870605,48.9948234558105,48.964111328125,48.7728500366211,48.523681640625,48.4226570129395,48.3237800598145,48.1805686950684,48.0898933410645,47.9752922058105,47.8873558044434,47.6348609924316,47.38330078125,47.0572280883789,46.9762191772461,46.8898429870605,46.7450675964355,46.5434074401855,46.4629402160645,46.250732421875,45.9285125732422,45.818359375,45.6399917602539,45.3935050964355,45.171142578125,45.0800323486328,44.9781723022461,44.8221206665039,44.6667976379395,44.56201171875,44.4891090393066,44.4398422241211,44.37353515625,43.9714851379395,43.8988265991211,43.8350105285645,43.6846199035645,43.5257339477539,43.4265594482422,43.2905769348145,43.0421371459961,42.9474601745605,42.8299293518066,42.8282241821289,42.7638664245605,42.6969757080078,42.7228012084961,42.8080101013184,42.8052749633789,42.7937507629395,42.8758277893066,42.87158203125,42.9097671508789,42.88330078125,43.2386741638184,43.3135261535645,43.2450714111328,43.1189460754395,43.0954093933105,43.0951690673828,43.1707496643066,43.280029296875,43.2960472106934,43.3019523620605,43.2552757263184,43.20263671875,42.9964370727539,42.8223152160645,42.7721214294434,42.6974105834961,42.6260261535645,42.6451683044434,42.6339340209961,42.6732902526855,42.6563949584961,42.52294921875,42.3257789611816,42.3025360107422],[78.1309127807617,78.0825653076172,78.0577621459961,78.0985794067383,78.0956497192383,78.1398468017578,78.1675796508789,78.1893997192383,78.185546875,78.1309127807617],[78.2061996459961,78.1986312866211,78.2201156616211,78.2646484375,78.2726058959961,78.3400344848633,78.3362350463867,78.316650390625,78.294189453125,78.2649917602539,78.2649917602539,78.2616653442383,78.2450180053711,78.2061996459961],[79.2539596557617,79.1764144897461,79.1060562133789,78.8766555786133,78.83544921875,78.8712921142578,78.9495620727539,79.0143508911133,79.0557556152344,79.0564956665039,79.0712890625,79.1261215209961,79.1500015258789,79.1492691040039,79.1232376098633,79.0625457763672,79.01318359375,78.9771041870117,78.9639129638672,78.92333984375,78.8800277709961,78.8351593017578,78.8548812866211,78.8433151245117,78.81884765625,78.7799377441406,78.7330093383789,78.6661605834961,78.5939483642578,78.4999008178711,78.3527374267578,78.3397445678711,78.3492126464844,78.3350601196289,78.2582550048828,78.255859375,78.1878967285156,78.1898956298828,78.2017135620117,78.224609375,78.205322265625,78.1943359375,78.1919479370117,78.1429748535156,78.0475158691406,77.9749984741211,77.9568328857422,77.97607421875,78.0006866455078,78.0380859375,78.0839385986328,78.1785659790039,78.2334976196289,78.3389129638672,78.38037109375,78.470458984375,78.5039520263672,78.5357894897461,78.5738296508789,78.6314926147461,78.67919921875,78.753173828125,78.7877960205078,78.7974166870117,78.7835922241211,78.7884826660156,78.8124542236328,78.8977584838867,78.9258270263672,78.9800796508789,79.0065383911133,79.0232849121094,79.0962371826172,79.1232376098633,79.1568832397461,79.2044448852539,79.2326202392578,79.2544403076172,79.3126449584961,79.3504409790039,79.3613739013672,79.3719787597656,79.3702163696289,79.3116226196289,79.263671875,79.2524871826172,79.2560501098633,79.3125991821289,79.3733901977539,79.4129333496094,79.4300765991211,79.4332046508789,79.3921356201172,79.33154296875,79.2991256713867,79.2825241088867,79.2711868286133,79.2539596557617],[79.6510772705078,79.615234375,79.6431579589844,79.5568313598633,79.5244140625,79.5019073486328,79.489501953125,79.493408203125,79.54443359375,79.5454559326172,79.5787582397461,79.6256332397461,79.6447219848633,79.6644515991211,79.6510772705078],[79.6142044067383,79.6016159057617,79.6714401245117,79.6903305053711,79.7010269165039,79.6836929321289,79.6702651977539,79.6142044067383],[79.6852111816406,79.6754913330078,79.6846694946289,79.7905807495117,79.8355026245117,79.9049377441406,79.9814910888672,80.0307159423828,80.0492172241211,80.0522994995117,80.0454559326172,79.9965362548828,79.9411163330078,79.904541015625,79.8167495727539,79.7838821411133,79.7375946044922,79.7044906616211,79.6852111816406],[79.9442367553711,79.9319381713867,79.9205093383789,79.9319839477539,79.9244155883789,79.9805679321289,80.0354461669922,80.0485305786133,80.09423828125,80.0486373901367,80.0103530883789,79.9912643432617,79.9723129272461,79.9442367553711],[79.955810546875,79.9230499267578,79.9329605102539,79.9482955932617,79.9641571044922,79.984619140625,80.0423278808594,80.0539093017578,80.1188507080078,80.0826644897461,79.9942855834961,79.955810546875],[80.1582565307617,80.0950164794922,80.0228500366211,80.0037612915039,79.956298828125,79.8958511352539,79.8504333496094,79.7749481201172,79.7606506347656,79.7813949584961,79.8526382446289,79.9010772705078,79.8741226196289,79.8843307495117,79.953125,80.0091323852539,80.0436019897461,80.0522003173828,80.04541015625,80.016357421875,79.9863815307617,79.9701614379883,79.9413070678711,79.919921875,79.898193359375,79.8489761352539,79.777099609375,79.7383346557617,79.6689453125,79.653076171875,79.6282730102539,79.5677185058594,79.5151824951172,79.4918441772461,79.4634780883789,79.3851089477539,79.3233413696289,79.2765655517578,79.2747573852539,79.3062973022461,79.3053741455078,79.2930221557617,79.2271957397461,79.1301727294922,79.107666015625,79.0958480834961,79.0063934326172,78.96142578125,78.8527374267578,78.834228515625,78.8182601928711,78.7877960205078,78.7950210571289,78.8209991455078,78.8102035522461,78.8273391723633,78.8265609741211,78.8680191040039,78.9339370727539,78.9638137817383,78.9849624633789,79.0030288696289,79.015869140625,79.0014190673828,79.0120086669922,79.0980987548828,79.0993118286133,79.0496139526367,79.0526809692383,79.0866241455078,79.1274871826172,79.140869140625,79.1923828125,79.2186050415039,79.3075180053711,79.40234375,79.451416015625,79.4627380371094,79.4583969116211,79.4953155517578,79.631591796875,79.70166015625,79.7560043334961,79.8297348022461,79.9419403076172,80.01123046875,80.0348129272461,80.0892562866211,80.0968322753906,80.030517578125,80.0421371459961,80.0727996826172,80.1056137084961,80.1100082397461,80.0968322753906,80.1100616455078,80.1043395996094,80.1530303955078,80.1682586669922,80.1582565307617],[80.0743179321289,80.0607452392578,80.1361312866211,80.158935546875,80.230224609375,80.219482421875,80.1856384277344,80.17236328125,80.1094741821289,80.0743179321289],[80.2738265991211,80.226806640625,80.2283706665039,80.2564468383789,80.2955551147461,80.3253860473633,80.3176803588867,80.30224609375,80.2738265991211],[80.3509292602539,80.3169937133789,80.1939468383789,80.139404296875,80.0946807861328,80.0714797973633,80.0764617919922,80.087158203125,80.104736328125,80.1632766723633,80.2039031982422,80.320068359375,80.34130859375,80.330322265625,80.36328125,80.3661651611328,80.3509292602539],[80.1852035522461,80.1732482910156,80.1788635253906,80.191162109375,80.2018585205078,80.2132263183594,80.2637176513672,80.2763214111328,80.2969284057617,80.3185043334961,80.3475570678711,80.4023895263672,80.4126434326172,80.4023895263672,80.3663101196289,80.3233871459961,80.2683639526367,80.2283172607422,80.2225570678711,80.1852035522461],[80.1232452392578,80.1040573120117,80.13916015625,80.1581039428711,80.1938934326172,80.3282699584961,80.3684616088867,80.396240234375,80.4452133178711,80.4683074951172,80.4939422607422,80.4753952026367,80.4647445678711,80.4158706665039,80.388427734375,80.3432159423828,80.3187484741211,80.2977981567383,80.2481460571289,80.2018127441406,80.196533203125,80.1232452392578],[80.4728012084961,80.4213409423828,80.4762191772461,80.5038528442383,80.5154342651367,80.54248046875,80.5630340576172,80.6052780151367,80.5743713378906,80.561767578125,80.5401382446289,80.4986801147461,80.4728012084961],[80.8536605834961,80.8126983642578,80.823486328125,80.8189926147461,80.8060073852539,80.779052734375,80.7177734375,80.6332550048828,80.6292953491211,80.6681594848633,80.7562484741211,80.7651824951172,80.741943359375,80.6879425048828,80.6745147705078,80.606201171875,80.6149368286133,80.6090316772461,80.5621109008789,80.5613327026367,80.5406723022461,80.4755325317383,80.456787109375,80.4467239379883,80.4837875366211,80.5408630371094,80.5694885253906,80.5369567871094,80.560302734375,80.5987319946289,80.6112823486328,80.6522445678711,80.7351608276367,80.7552261352539,80.8143997192383,80.8529281616211,80.8536605834961],[80.8347625732422,80.7943801879883,80.730224609375,80.6836929321289,80.616943359375,80.6010284423828,80.5863265991211,80.5349578857422,80.5047378540039,80.4186096191406,80.4346694946289,80.4944381713867,80.4460906982422,80.4312515258789,80.5050277709961,80.5215301513672,80.5726547241211,80.6177749633789,80.7125473022461,80.7665023803711,80.7835922241211,80.8165054321289,80.8363723754883,80.8485870361328,80.8377456665039,80.8014678955078,80.8042449951172,80.8265609741211,80.8626403808594,80.8929214477539,80.8859329223633,80.8666000366211,80.8347625732422],[80.92724609375,80.910888671875,80.9141540527344,80.8904266357422,80.7446823120117,80.7407760620117,80.6876449584961,80.6039581298828,80.5404739379883,80.52734375,80.4976577758789,80.4720764160156,80.4253387451172,80.3765563964844,80.3691864013672,80.3537139892578,80.300048828125,80.290283203125,80.2768020629883,80.2656707763672,80.2423782348633,80.2074203491211,80.16259765625,80.1553192138672,80.1611328125,80.1953582763672,80.1832962036133,80.1582565307617,80.110107421875,80.0957946777344,80.1327590942383,80.122314453125,80.0994644165039,80.088623046875,80.0816879272461,80.1119613647461,80.1513671875,80.2125473022461,80.2392578125,80.2453079223633,80.2301254272461,80.1885223388672,80.1802291870117,80.1827697753906,80.2372131347656,80.2576675415039,80.3003387451172,80.4447708129883,80.5005340576172,80.529052734375,80.5438995361328,80.5615768432617,80.5687942504883,80.5580596923828,80.50830078125,80.5157699584961,80.5586471557617,80.656005859375,80.7121047973633,80.8213882446289,80.8653335571289,80.9238739013672,80.92724609375],[80.84814453125,80.8120574951172,80.8348159790039,80.8482360839844,80.923583984375,80.9603500366211,80.9753875732422,80.9498062133789,80.9276885986328,80.8826141357422,80.867919921875,80.84814453125],[80.9503402709961,80.9296875,80.9555206298828,80.984619140625,80.9991760253906,81.0166473388672,81.0876998901367,81.1039581298828,81.05029296875,81.0110397338867,80.9503402709961],[81.115966796875,81.0198745727539,81.0291519165039,80.9982452392578,80.9128952026367,80.8197326660156,80.7922897338867,80.7554702758789,80.6636276245117,80.6328659057617,80.62841796875,80.6372985839844,80.7033233642578,80.7519073486328,80.7386703491211,80.7652282714844,80.7830123901367,80.7869644165039,80.8136215209961,80.8719711303711,80.90185546875,80.90380859375,80.9865264892578,81.1131820678711,81.115966796875],[81.0416488647461,80.990966796875,80.93359375,80.8976516723633,80.8316879272461,80.7793960571289,80.7679672241211,80.764892578125,80.7933578491211,80.8890609741211,80.9151306152344,81.01708984375,81.0467758178711,81.0315399169922,81.1184539794922,81.1142578125,81.0945816040039,81.061767578125,81.0416488647461],[81.0474166870117,81.041259765625,81.0455551147461,81.0843734741211,81.1027297973633,81.1225128173828,81.126220703125,81.1442413330078,81.158203125,81.16943359375,81.170654296875,81.15087890625,81.108154296875,81.0718231201172,81.0474166870117],[80.7000961303711,80.6976089477539,80.7128448486328,80.7626953125,80.8218765258789,80.85302734375,80.893798828125,80.966796875,80.9809112548828,80.9811553955078,80.9983367919922,81.0357437133789,81.1063461303711,81.1444396972656,81.1751937866211,81.198486328125,81.197265625,81.1694793701172,81.14404296875,81.096435546875,81.0567398071289,81.0082015991211,80.968017578125,80.9307098388672,80.8188934326172,80.7554244995117,80.7000961303711],[81.1412124633789,81.0638198852539,81.1131362915039,81.1487350463867,81.1707077026367,81.2137222290039,81.1991195678711,81.1839370727539,81.1412124633789],[81.0755920410156,81.0300827026367,80.9941864013672,80.9578628540039,80.8418426513672,80.8267135620117,80.7982864379883,80.7632751464844,80.6980972290039,80.6986846923828,80.6780776977539,80.6524429321289,80.6140594482422,80.5355453491211,80.5198669433594,80.49658203125,80.3629913330078,80.342529296875,80.3231506347656,80.2727508544922,80.2410125732422,80.1769561767578,80.150390625,80.122802734375,80.1260757446289,80.0760269165039,80.0101089477539,80.0096206665039,80.1021041870117,80.1792984008789,80.2233352661133,80.249267578125,80.2699203491211,80.3585433959961,80.4185028076172,80.4775390625,80.4991226196289,80.5332489013672,80.6185531616211,80.7029800415039,80.791259765625,80.7686538696289,80.7808609008789,80.8100128173828,80.8721618652344,80.89306640625,80.9258270263672,80.9884796142578,81.0316925048828,81.0392074584961,81.0381317138672,81.0583953857422,81.0894546508789,81.1073760986328,81.1146469116211,81.139404296875,81.1880874633789,81.27099609375,81.2804641723633,81.2605972290039,81.21142578125,81.1927719116211,81.1839370727539,81.0992660522461,81.0755920410156],[81.3052291870117,81.2922821044922,81.3135223388672,81.3372497558594,81.3603515625,81.391845703125,81.397705078125,81.3661117553711,81.3250503540039,81.3052291870117],[81.5460433959961,81.5064468383789,81.4837875366211,81.4641571044922,81.4184112548828,81.386962890625,81.3680648803711,81.3032760620117,81.2548294067383,81.197509765625,81.1697311401367,81.1355438232422,81.178466796875,81.2379455566406,81.1982955932617,81.1752395629883,81.1786117553711,81.2239761352539,81.1884765625,81.2280731201172,81.3111801147461,81.3294448852539,81.3030776977539,81.3870086669922,81.4233856201172,81.5105514526367,81.5412063598633,81.5352554321289,81.5428695678711,81.5646514892578,81.5460433959961],[81.6093292236328,81.5965805053711,81.60888671875,81.633056640625,81.6470260620117,81.6591339111328,81.6793441772461,81.70654296875,81.7189483642578,81.6873092651367,81.6641616821289,81.6498031616211,81.6093292236328],[81.7151870727539,81.6956558227539,81.7104949951172,81.721923828125,81.7478485107422,81.7970199584961,81.8279800415039,81.8541946411133,81.825439453125,81.7809524536133,81.7589797973633,81.7151870727539],[-1.06728518009186,-1.08300769329071,-1.13115227222443,-1.20820295810699,-1.36748075485229,-1.39677715301514,-1.45869147777557,-1.56308615207672,-1.69365239143372,-1.85068356990814,-1.96748042106628,-2.04404306411743,-2.14335942268372,-2.26542997360229,-2.33847689628601,-2.36269521713257,-2.37167978286743,-2.36347651481628,-2.37382841110229,-2.39677762985229,-2.40009784698486,-2.39560532569885,-2.37607431411743,-2.31298828125,-2.34785151481628,-2.34706997871399,-2.37705063819885,-2.41396450996399,-2.41660141944885,-2.41152310371399,-2.33710908889771,-2.33955049514771,-2.54863262176514,-2.66464853286743,-2.71640610694885,-2.76640629768372,-2.79472661018372,-2.79277348518372,-2.80839848518372,-2.80859375,-2.79150366783142,-2.67304682731628,-2.62031245231628,-2.59570336341858,-2.60253953933716,-2.66455054283142,-2.72021508216858,-2.68203163146973,-2.63505864143372,-2.55556654930115,-2.44667983055115,-2.40029335021973,-2.37031245231628,-2.31279277801514,-2.23320341110229,-2.19511723518372,-2.13183617591858,-1.98457062244415,-1.86025404930115,-1.81601560115814,-1.71992194652557,-1.62158226966858,-1.51757788658142,-1.50742161273956,-1.46806621551514,-1.40976560115814,-1.38789057731628,-1.38710963726044,-1.33554685115814,-1.35166013240814,-1.45175755023956,-1.46630847454071,-1.46992218494415,-1.44697284698486,-1.36865246295929,-1.32109403610229,-1.25419950485229,-1.17880845069885,-1.11308574676514,-1.07460927963257,-1.06308591365814,-1.06601548194885,-1.06728518009186],[27.2859382629395,27.1192874908447,26.9213371276855,26.72314453125,26.4977054595947,26.273193359375,26.1094722747803,25.9955062866211,25.9955062866211,25.9955062866211,25.9955062866211,25.9955062866211,25.9955062866211,25.9954586029053,25.9954586029053,25.9954586029053,25.9954586029053,25.9954586029053,25.9954586029053,25.9954586029053,25.9954586029053,25.9954090118408,25.9954090118408,25.9954090118408,25.9954090118408,25.9954090118408,25.8763179779053,25.7401371002197,25.60400390625,25.4678707122803,25.3316898345947,25.195556640625,25.0593757629395,24.9232425689697,24.787109375,24.6509761810303,24.5147953033447,24.378662109375,24.2424812316895,24.1063480377197,23.97021484375,23.8340320587158,23.6978988647461,23.5764656066895,23.4675788879395,23.4354496002197,23.3774909973145,23.3180179595947,23.2908210754395,23.2713375091553,23.1927261352539,23.0895500183105,23.000244140625,22.8840827941895,22.8205070495605,22.7532234191895,22.6893062591553,22.5607414245605,22.4959964752197,22.3832530975342,22.2604484558105,22.128173828125,21.9957523345947,21.8547859191895,21.7138175964355,21.5720691680908,21.4667987823486,21.3339347839355,21.3337879180908,21.3335437774658,21.333251953125,21.3329582214355,21.3327140808105,21.3324222564697,21.3321285247803,21.3318843841553,21.3315906524658,21.331298828125,21.3310546875,21.3307609558105,21.3304691314697,21.3302249908447,21.3299312591553,21.3296394348145,21.3293952941895,21.3292484283447,21.1424331665039,21.0080070495605,20.8988285064697,20.80615234375,20.8568840026855,21.3770980834961,21.4207019805908,21.4207515716553,21.4302730560303,21.4703121185303,21.4810543060303,21.4810543060303,21.5005874633789,21.5005874633789,21.4908199310303,21.4703121185303,21.4507808685303,21.4507808685303,21.4410152435303,21.4410152435303,21.4507808685303,21.5005874633789,21.6001949310303,21.6802730560303,21.7505855560303,21.8208980560303,21.8609371185303,21.9107418060303,21.9908695220947,22.0406265258789,22.0806636810303,22.1207027435303,22.1910171508789,22.2408199310303,22.3101558685303,22.3707027435303,22.4507808685303,22.5904312133789,22.7603511810303,22.8707027435303,22.9605464935303,23.1001949310303,23.3404293060303,23.4107418060303,23.5201168060303,23.6207027435303,23.6910171508789,23.7505855560303,23.7906246185303,23.8306636810303,23.8707027435303,23.9107418060303,23.9410152435303,23.9810543060303,24.0201168060303,24.0904312133789,24.2203121185303,24.3003902435303,24.4009780883789,24.4703121185303,24.4972648620605,24.5201187133789,24.5708980560303,24.6304702758789,24.6802730560303,24.7310543060303,24.7701168060303,24.8306636810303,24.8804683685303,24.9703121185303,25.1109371185303,25.2603015899658,25.4205074310303,25.5201168060303,25.6402339935303,25.7310543060303,25.8306636810303,25.8707027435303,25.9205074310303,25.9908199310303,25.996337890625,26.0308589935303,26.0503902435303,26.0708980560303,26.0865230560303,26.1041011810303,26.1626949310303,26.2134780883789,26.2955074310303,26.3609371185303,26.4009761810303,26.4703121185303,26.5201168060303,26.5835933685303,26.6333980560303,26.6841793060303,26.7447280883789,26.7935543060303,26.8833999633789,26.9107418060303,26.9410152435303,26.9703121185303,27.0103511810303,27.0103511810303,27.0201168060303,27.0005855560303,26.9908218383789,26.9605464935303,26.9009761810303,26.8609390258789,26.8609390258789,26.8804683685303,26.9087886810303,26.9107418060303,26.8902339935303,26.8501968383789,26.8501968383789,26.8609390258789,26.9107418060303,26.9908218383789,27.0503902435303,27.0884780883789,27.0982437133789,27.0982437133789,27.1001968383789,27.0904293060303,27.0904293060303,27.1041011810303,27.1207027435303,27.1509761810303,27.1910152435303,27.2505874633789,27.3082027435303,27.3609371185303,27.4165515899658,27.4605464935303,27.5308589935303,27.6138668060303,27.6559085845947,27.6564445495605,27.6564445495605,27.490234375,27.2859382629395],[16.7156238555908,16.7100582122803,16.741943359375,16.7576656341553,16.8035144805908,16.7086429595947,16.5963859558105,16.5707035064697,16.5948238372803,16.6184558868408,16.6439456939697,16.6714839935303,16.6534671783447,16.6842765808105,16.7787590026855,16.8468742370605,16.8601570129395,16.8929195404053,16.9468269348145,17.0025386810303,16.99365234375,16.9364261627197,16.8062515258789,16.7786617279053,16.7489242553711,16.7156238555908],[25.383056640625,25.383056640625,25.4507331848145,25.5005855560303,25.5340347290039,25.6015625,25.6451168060303,25.688720703125,25.6453609466553,25.6198253631592,25.5587406158447,25.425537109375,25.4146480560303,25.383056640625],[25.7127933502197,25.69921875,25.7342777252197,25.8116207122803,25.8555164337158,25.79541015625,25.73486328125,25.7127933502197],[29.0962409973145,29.0746097564697,29.0456523895264,29.0261726379395,28.9895496368408,28.8378410339355,28.7315425872803,28.6279773712158,28.5331554412842,28.5354499816895,28.5375003814697,28.54052734375,28.5429191589355,28.4488773345947,28.3550300598145,28.1325664520264,27.9590816497803,27.8958988189697,27.8006839752197,27.7652816772461,27.7243175506592,27.6290531158447,27.548583984375,27.5282211303711,27.4927234649658,27.4376468658447,27.3104972839355,27.1809558868408,27.1517581939697,26.9558582305908,26.8289070129395,26.6626453399658,26.6595211029053,26.6764144897461,26.69921875,26.6785163879395,26.6087894439697,26.52685546875,26.4559574127197,26.4049320220947,26.3084964752197,26.1005363464355,26.1006832122803,26.1187019348145,26.1228504180908,26.1109867095947,25.9613761901855,25.8466320037842,25.7558097839355,25.6228523254395,25.5661144256592,25.4248046875,25.3066902160645,25.086669921875,24.9638195037842,24.869384765625,24.7892589569092,24.6796379089355,24.5951175689697,24.5739269256592,24.5652351379395,24.5646495819092,24.5867195129395,24.6072273254395,24.5643558502197,24.57080078125,24.5309581756592,24.4769058227539,24.3403816223145,24.31884765625,24.3082027435303,24.2863292694092,24.2861824035645,24.2579097747803,24.1283187866211,24.078857421875,24.0350112915039,23.9695320129395,23.9040031433105,23.8384761810303,23.7729969024658,23.70751953125,23.6419925689697,23.5764656066895,23.510986328125,23.4454593658447,23.3799800872803,23.314453125,23.2489738464355,23.1834964752197,23.1179695129395,23.0524425506592,22.9869632720947,22.9328117370605,22.9225101470947,22.9192867279053,22.9100093841553,22.8956050872803,22.8770008087158,22.8549308776855,22.830322265625,22.8040046691895,22.7768058776855,22.7496585845947,22.7233390808105,22.69873046875,22.6766605377197,22.6580085754395,22.6436538696289,22.6343746185303,22.6311531066895,22.6214847564697,22.6239261627197,22.7041015625,22.5909175872803,22.4969234466553,22.3678226470947,22.2306632995605,22.0995101928711,22.0018558502197,21.900390625,21.7896976470947,21.6790027618408,21.5683097839355,21.4576168060303,21.346923828125,21.2362289428711,21.1255378723145,21.0147953033447,20.9041004180908,20.7934074401855,20.6827144622803,20.572021484375,20.4613285064697,20.3506336212158,20.2399425506592,20.1292476654053,19.9959468841553,19.9604988098145,19.9031257629395,19.8458003997803,19.7884769439697,19.7311534881592,19.673828125,19.6165027618408,19.5591316223145,19.5018062591553,19.4444828033447,19.3871574401855,19.3297863006592,19.2724609375,19.2151374816895,19.1578140258789,19.1004886627197,19.0431632995605,18.9961433410645,18.9645519256592,18.9338874816895,18.8993663787842,18.8578624725342,18.8252925872803,18.7777824401855,18.7352523803711,18.6953125,18.6553230285645,18.621337890625,18.581787109375,18.4952144622803,18.3624019622803,18.22705078125,18.1569328308105,17.9769535064697,17.8858394622803,17.7210922241211,17.5968265533447,17.4483394622803,17.3161125183105,17.1118659973145,17.0604000091553,16.9939460754395,16.9466800689697,16.9534664154053,17.0790042877197,17.2121086120605,17.2655754089355,17.2685546875,17.2516593933105,17.2312984466553,17.253173828125,17.2784175872803,17.3020515441895,17.3197746276855,17.4062004089355,17.4233894348145,17.4274406433105,17.4295883178711,17.4316902160645,17.4043464660645,17.4143562316895,17.3985366821289,17.3655281066895,17.3674812316895,17.3383312225342,17.32470703125,17.349609375,17.3441410064697,17.3655281066895,17.421875,17.471435546875,17.4987316131592,17.5159168243408,17.5162582397461,17.4860363006592,17.456787109375,17.359375,17.32470703125,17.2664546966553,17.2392578125,17.2050304412842,17.1129875183105,17.0624504089355,16.9419918060303,16.8467769622803,16.8118152618408,16.77099609375,16.6894054412842,16.6641597747803,16.5866203308105,16.5503902435303,16.5090827941895,16.3717765808105,16.4515628814697,16.56982421875,16.6533203125,16.7369632720947,16.8013668060303,16.8684558868408,16.9596691131592,17.0498523712158,17.1224613189697,17.2566394805908,17.4349613189697,17.6693363189697,17.8857421875,18.0076656341553,18.256103515625,18.4524402618408,18.6784172058105,18.7652339935303,18.8711910247803,18.9890632629395,19.0821781158447,19.4901351928711,19.5552730560303,19.6463871002197,19.7168941497803,19.7554683685303,19.8223628997803,19.9934558868408,20.2659168243408,20.29296875,20.3903331756592,20.5176773071289,20.7370109558105,20.9739742279053,21.3103523254395,21.4327621459961,21.5189933776855,21.6639652252197,21.7759761810303,21.8817386627197,22.033447265625,22.203369140625,22.2936515808105,22.3927726745605,22.5921878814697,22.698974609375,22.7700672149658,22.8047866821289,22.8820304870605,22.8818359375,22.9890632629395,23.048583984375,23.1942882537842,23.3055171966553,23.5579090118408,23.7118663787842,23.9109859466553,24.0580081939697,24.1245613098145,24.1854019165039,24.1875,24.2744140625,24.2777347564697,24.2916488647461,24.4590339660645,24.6158199310303,24.8200187683105,24.8733386993408,24.9600582122803,25.0734386444092,25.1506843566895,25.2911128997803,25.6411628723145,25.6924800872803,25.7513179779053,25.9028816223145,26.0388660430908,26.1048831939697,26.5947761535645,26.7658214569092,26.8810043334961,27.0704593658447,27.2587890625,27.4324703216553,27.7337875366211,28.0348625183105,28.0870113372803,28.1085929870605,28.1306629180908,28.0645008087158,28.1483402252197,28.2641124725342,28.50732421875,28.7205085754395,29.3535175323486,29.3209476470947,29.2940902709961,29.2548809051514,29.2142601013184,29.1904792785645,29.2005367279053,29.3553714752197,29.4951171875,29.6661148071289,29.8316402435303,29.8660163879395,29.8970699310303,29.9462890625,29.9950695037842,30.01171875,30.1445789337158,30.3132820129395,30.3309574127197,30.3481464385986,30.442626953125,30.5000019073486,30.6692867279053,30.8289527893066,31.0077648162842,31.1468276977539,31.3551750183105,31.4915027618408,31.5561027526855,31.6258792877197,31.6963367462158,31.7811527252197,31.8474617004395,31.9464855194092,31.9949207305908,32.0074729919434,32.12451171875,32.0917510986328,32.0425262451172,31.9950199127197,31.93896484375,31.8933601379395,31.7254390716553,31.6163578033447,31.489013671875,31.3297367095947,31.2203617095947,31.0803737640381,30.92041015625,30.7177753448486,30.4952125549316,30.3222179412842,30.083984375,29.8492183685303,29.6193370819092,29.4352531433105,29.2023448944092,29.1936054229736,29.1670894622803,29.1315422058105,29.0958480834961,29.0636711120605,29.0962409973145],[21.9967288970947,21.768798828125,21.5865249633789,21.3260269165039,21.1858406066895,21.1085433959961,21.0726566314697,21.0394039154053,21.07763671875,21.103759765625,20.9817886352539,20.8949222564697,20.7319831848145,20.5567378997803,20.3949222564697,20.1207027435303,19.7918930053711,19.5818843841553,19.0919914245605,18.8201160430908,18.7531242370605,18.7174320220947,18.6943359375,18.555908203125,18.4097652435303,18.3332996368408,18.2494125366211,18.264404296875,18.2867183685303,18.2190437316895,18.0729484558105,18.0050792694092,17.9385261535645,17.8239250183105,17.7783699035645,17.7512683868408,17.7173328399658,17.68359375,17.6370124816895,17.61669921875,17.5847663879395,17.56396484375,17.5628433227539,17.5485343933105,17.5264644622803,17.5377922058105,17.5176773071289,17.4923324584961,17.4702625274658,17.4655284881592,17.4580078125,17.4205093383789,17.3682613372803,17.3350105285645,17.3241214752197,17.2881336212158,17.1086921691895,17.0617179870605,17.0570812225342,17.056884765625,17.0414047241211,17.061279296875,17.0588874816895,17.0205574035645,16.8665523529053,16.8005847930908,16.7223644256592,16.6246585845947,16.4595222473145,16.2961902618408,16.05029296875,15.9939451217651,15.7988767623901,15.7263669967651,15.3621101379395,15.2501459121704,15.1320810317993,14.9400873184204,14.7364749908447,14.5443363189697,14.2568359375,13.9884281158447,13.842041015625,13.6260747909546,13.5262689590454,13.4668455123901,13.40576171875,13.2710933685303,13.0933103561401,12.9111328125,12.8053226470947,12.7570314407349,12.7264642715454,12.706298828125,12.6848630905151,12.6610355377197,12.6237306594849,12.5373048782349,12.3005857467651,12.1555662155151,11.9570302963257,11.8165521621704,11.748291015625,11.6210451126099,11.4198732376099,11.2767572402954,11.1617670059204,10.9621095657349,10.8647947311401,10.810546875,10.7591791152954,10.7461910247803,10.804931640625,10.8645505905151,10.8801755905151,10.8428707122803,10.787841796875,10.6586427688599,10.5281257629395,10.2515621185303,10.1908693313599,10.1247568130493,9.91855525970459,9.85341739654541,9.72968769073486,9.51347637176514,9.46152400970459,9.46152400970459,9.46220684051514,9.46352577209473,9.46640682220459,9.47119140625,9.47773456573486,9.50615215301514,9.55034160614014,9.62675762176514,9.71762657165527,9.84526348114014,9.85581016540527,9.86176681518555,9.86870098114014,9.89145565032959,9.9111328125,9.94091796875,10.0071773529053,10.0277347564697,10.05419921875,10.0709476470947,10.1814451217651,10.198974609375,10.5508298873901,10.6461906433105,10.6527347564697,10.6578130722046,10.7378902435303,10.7459468841553,10.7577142715454,10.77294921875,10.8191890716553,10.8314456939697,10.8501462936401,11.5915040969849,11.6061029434204,11.6157712936401,11.6217288970947,11.6374998092651,11.6538572311401,11.6824216842651,11.6931638717651,11.8255853652954,11.9416017532349,12.135009765625,12.21728515625,12.2230958938599,12.2188472747803,12.2137689590454,12.2080078125,12.2018070220947,12.1888179779053,12.1648435592651,12.1521978378296,12.1392574310303,12.0929203033447,12.0580568313599,12.0464353561401,12.0337409973145,12.0096683502197,12.0067386627197,11.7160158157349,11.7101078033447,11.6942873001099,11.6827154159546,11.5804204940796,11.4185552597046,11.3145017623901,11.2469244003296,11.1139650344849,11.089111328125,11.0577640533447,10.6624994277954,10.6438474655151,10.4790525436401,10.3831548690796,10.3557138442993,10.2211427688599,9.79926681518555,9.77094745635986,9.75937557220459,9.75629901885986,9.75312519073486,9.745849609375,9.73496150970459,9.726806640625,9.731201171875,9.74267578125,9.97895526885986,10.2773923873901,10.250244140625,10.1219234466553,10.088623046875,10.0650882720947,9.92138671875,9.84829139709473,9.76860332489014,9.71806621551514,9.67465782165527,9.61015605926514,9.593994140625,9.59418964385986,9.54946327209473,9.45908164978027,9.38881778717041,9.32607460021973,9.32861328125,9.37880897521973,9.59965801239014,9.60161113739014,9.587890625,9.61381816864014,9.59062480926514,9.49921894073486,9.484130859375,9.52583026885986,9.96591758728027,10.0184564590454,10.0467767715454,10.1234378814697,10.1693353652954,10.2027339935303,10.2496089935303,10.3460941314697,10.406494140625,10.4205083847046,10.3185062408447,10.3299322128296,10.3118162155151,10.2937984466553,10.2357902526855,10.1758785247803,10.115234375,10.0552740097046,9.98886775970459,9.839599609375,9.774658203125,9.61030197143555,9.52734375,9.48891639709473,9.42568302154541,9.38930606842041,9.338134765625,9.22993183135986,9.17910194396973,9.05170917510986,9.00678730010986,8.91484355926514,8.88691329956055,8.81425762176514,8.767822265625,8.6962890625,8.66562461853027,8.69130802154541,8.709716796875,8.73247146606445,8.76582050323486,8.81581974029541,8.94321346282959,8.97060585021973,8.99331092834473,9.04848575592041,9.11474609375,9.26191425323486,9.34062480926514,9.61391639709473,9.71035194396973,9.82290077209473,10.0300779342651,10.1742677688599,10.3879404067993,10.4578123092651,10.7518072128296,10.7953128814697,10.919677734375,11.0290031433105,11.1920413970947,11.2671880722046,11.3448734283447,11.4032716751099,11.4099607467651,11.4398441314697,11.482666015625,11.5159177780151,11.5798826217651,11.6695308685303,11.9901371002197,12.032958984375,12.0447273254395,12.0677738189697,12.1292476654053,12.3119144439697,12.4629878997803,12.54638671875,12.6604490280151,12.70947265625,12.694580078125,12.671875,12.6781244277954,12.6993656158447,12.7412109375,12.79052734375,12.8647470474243,13.0009765625,13.1130857467651,13.2150392532349,13.2693357467651,13.32958984375,13.3987789154053,13.4716300964355,13.5380859375,13.6264162063599,13.7303218841553,13.7998046875,13.8501462936401,13.9105958938599,13.9787111282349,13.9923343658447,14.0288572311401,14.0555171966553,14.12744140625,14.161865234375,14.2032232284546,14.237060546875,14.2842283248901,14.3421392440796,14.4412107467651,14.5041990280151,14.5504884719849,14.585205078125,14.6333494186401,14.6627445220947,14.6880865097046,14.7224607467651,14.7886228561401,14.8514642715454,14.8983898162842,14.9986820220947,15.0444345474243,15.0966300964355,15.162109375,15.2381353378296,15.3113288879395,15.5331048965454,15.6258296966553,15.702540397644,15.6972179412842,15.7139654159546,15.745997428894,15.7449712753296,15.7035150527954,15.7134275436401,15.7215328216553,15.7801752090454,15.9281244277954,16.3742198944092,16.8202629089355,17.266357421875,17.71240234375,18.1584949493408,18.6045398712158,19.0505867004395,19.4966316223145,19.6214847564697,19.7462882995605,19.87109375,19.9959468841553,19.995849609375,19.9957027435303,19.9955558776855,19.9954586029053,19.9972648620605,19.9990253448486,20.0007820129395,20.0025882720947,20.5009269714355,20.9992179870605,21.49755859375,21.995849609375,21.9958019256592,21.9957523345947,21.9956550598145,21.9956073760986,21.9955577850342,21.9955081939697,21.9954605102539,21.9953632354736,21.9953136444092,21.9952640533447,21.9952163696289,21.9951171875,21.9951171875,21.9950199127197,21.9949226379395,21.994873046875,21.994873046875,22.0022945404053,22.1886234283447,22.2024421691895,22.1915035247803,22.1478023529053,22.0846691131592,21.995849609375,21.995849609375,21.9958992004395,21.9959964752197,21.9959964752197,21.99609375,21.9961433410645,21.9961910247803,21.9962406158447,21.9962902069092,21.996337890625,21.9963874816895,21.9964351654053,21.9964847564697,21.9965343475342,21.99658203125,21.9966316223145,21.9967288970947],[9.46152400970459,9.42099571228027,9.218505859375,9.041259765625,8.75185489654541,8.67636680603027,8.58222675323486,8.54526424407959,8.49209022521973,8.44350624084473,8.43110370635986,8.43256855010986,8.44340801239014,8.44775390625,8.437255859375,8.39638614654541,8.25107383728027,8.04047775268555,7.95151329040527,7.89951181411743,7.86855459213257,7.82373046875,7.76064443588257,7.72373056411743,7.707763671875,7.69043016433716,7.67099666595459,7.509521484375,7.4345703125,7.36796855926514,7.29697275161743,7.22573232650757,7.08457040786743,7.00283241271973,6.89838886260986,6.77983379364014,6.72216749191284,6.66030263900757,6.56787157058716,6.30014657974243,6.15981435775757,6.04506874084473,5.85830116271973,5.77490234375,5.67314434051514,5.58120155334473,5.51103496551514,5.49228525161743,5.31186532974243,5.10957050323486,4.87548828125,4.62065410614014,4.41909170150757,4.22021484375,3.98525357246399,3.81171846389771,3.787109375,3.75507831573486,3.75434541702271,3.77470755577087,3.88017582893372,3.79848623275757,3.77270531654358,3.76318311691284,3.74995112419128,3.70620107650757,3.65131831169128,3.60781264305115,3.52802705764771,3.51972675323486,3.52919960021973,3.56118154525757,3.60756850242615,3.70908164978027,3.77045869827271,3.80263662338257,3.70146489143372,3.67758822441101,3.68046879768372,3.73759770393372,3.78593707084656,3.78559565544128,3.72499990463257,3.63408231735229,3.54414057731628,3.49072241783142,3.53334927558899,3.57314467430115,3.62421870231628,3.64409160614014,3.63413095474243,3.62421870231628,3.65278291702271,3.72294902801514,3.78720664978027,3.835693359375,3.88388681411743,3.98193359375,4.17763710021973,4.26850557327271,4.3271484375,4.48095655441284,4.5869140625,4.63603544235229,4.61181640625,4.49838876724243,4.39189434051514,4.38818359375,4.44594717025757,4.487060546875,4.50498008728027,4.45449209213257,4.37285184860229,4.32416963577271,4.31865215301514,4.33803701400757,4.34853553771973,4.35024404525757,4.4248046875,4.47939443588257,4.53208017349243,4.56791973114014,4.59775400161743,4.64467763900757,4.70322275161743,4.7783203125,4.84599637985229,4.96757793426514,5.03920888900757,5.10917997360229,5.18632793426514,5.28964853286743,5.44077157974243,5.5625,5.61879873275757,5.67514657974243,5.72294950485229,5.77685594558716,5.85493135452271,5.94550800323486,5.99824237823486,6.01752948760986,6.06923818588257,6.18300724029541,6.274169921875,6.34492206573486,6.39623975753784,6.455322265625,6.63530302047729,6.69902372360229,6.78173828125,6.87211942672729,6.95522451400757,7.06494140625,7.22871112823486,7.33339834213257,7.427490234375,7.51933574676514,7.55722713470459,7.64897441864014,7.72456073760986,7.80791044235229,7.96484422683716,8.13754844665527,8.19155216217041,8.23945331573486,8.25844764709473,8.29140663146973,8.369140625,8.46113300323486,8.60825252532959,8.65336894989014,8.66562461853027,8.6962890625,8.767822265625,8.81425762176514,8.88691329956055,8.91484355926514,9.00678730010986,9.05170917510986,9.17910194396973,9.22993183135986,9.338134765625,9.38930606842041,9.42568302154541,9.48891639709473,9.52734375,9.61030197143555,9.774658203125,9.839599609375,9.98886775970459,10.0552740097046,10.115234375,10.1758785247803,10.2357902526855,10.2937984466553,10.3118162155151,10.3299322128296,10.3185062408447,10.4205083847046,10.406494140625,10.3460941314697,10.2496089935303,10.2027339935303,10.1693353652954,10.1234378814697,10.0467767715454,10.0184564590454,9.96591758728027,9.52583026885986,9.484130859375,9.49921894073486,9.59062480926514,9.61381816864014,9.587890625,9.60161113739014,9.59965801239014,9.37880897521973,9.32861328125,9.32607460021973,9.38881778717041,9.45908164978027,9.54946327209473,9.59418964385986,9.593994140625,9.61015605926514,9.67465782165527,9.71806621551514,9.76860332489014,9.84829139709473,9.92138671875,10.0650882720947,10.088623046875,10.1219234466553,10.250244140625,10.2773923873901,9.97895526885986,9.74267578125,9.731201171875,9.726806640625,9.73496150970459,9.745849609375,9.75312519073486,9.75629901885986,9.75937557220459,9.77094745635986,9.79926681518555,10.2211427688599,10.3557138442993,10.3831548690796,10.4790525436401,10.6438474655151,10.6624994277954,11.0577640533447,11.089111328125,11.1139650344849,11.2469244003296,11.3145017623901,11.4185552597046,11.5804204940796,11.6827154159546,11.6942873001099,11.7101078033447,11.7160158157349,12.0067386627197,12.0096683502197,12.0337409973145,12.0464353561401,12.0580568313599,12.0929203033447,12.1392574310303,12.1521978378296,12.1648435592651,12.1888179779053,12.2018070220947,12.2080078125,12.2137689590454,12.2188472747803,12.2230958938599,12.21728515625,12.135009765625,11.9416017532349,11.8255853652954,11.6931638717651,11.6824216842651,11.6538572311401,11.6374998092651,11.6217288970947,11.6157712936401,11.6061029434204,11.5915040969849,10.8501462936401,10.8314456939697,10.8191890716553,10.77294921875,10.7577142715454,10.7459468841553,10.7378902435303,10.6578130722046,10.6527347564697,10.6461906433105,10.5508298873901,10.198974609375,10.1814451217651,10.0709476470947,10.05419921875,10.0277347564697,10.0071773529053,9.94091796875,9.9111328125,9.89145565032959,9.86870098114014,9.86176681518555,9.85581016540527,9.84526348114014,9.71762657165527,9.62675762176514,9.55034160614014,9.50615215301514,9.47773456573486,9.47119140625,9.46640682220459,9.46352577209473,9.46220684051514,9.46152400970459,9.46152400970459],[14.8090333938599,14.6481447219849,14.5711431503296,14.4585933685303,14.3766603469849,14.3232908248901,14.2742195129395,14.2064943313599,14.0718259811401,13.9746580123901,13.9307613372803,13.8752927780151,13.8289546966553,13.7880859375,13.73388671875,13.633056640625,13.5108880996704,13.4444332122803,13.4062986373901,13.3645505905151,13.3158197402954,13.3272943496704,13.3670902252197,13.39453125,13.3823728561401,13.369873046875,13.2900390625,13.2369632720947,13.1702632904053,13.0869626998901,13.0282230377197,12.9916009902954,12.9419927597046,12.8318853378296,12.7754878997803,12.6275882720947,12.5577154159546,12.5319337844849,12.479248046875,12.4043941497803,12.4175777435303,12.4263181686401,12.3873043060303,12.4033203125,12.3980464935303,12.3766117095947,12.3280277252197,12.3400869369507,12.3783693313599,12.3757810592651,12.3961915969849,12.4331550598145,12.451904296875,12.52001953125,12.5322751998901,12.5143556594849,12.4916505813599,12.4776363372803,12.489990234375,12.5362796783447,12.5810546875,12.6150388717651,12.6335458755493,12.6397466659546,12.6395998001099,12.6536130905151,12.6622562408447,12.6739253997803,12.6752939224243,12.6764163970947,12.677978515625,12.678955078125,12.679931640625,12.5889654159546,12.4903802871704,12.4378910064697,12.4574222564697,12.4433107376099,12.3995113372803,12.36767578125,12.3486328125,12.3643550872803,12.3548336029053,12.3997068405151,12.4725093841553,12.5257806777954,12.56005859375,12.6048831939697,12.5818357467651,12.5807123184204,12.6094722747803,12.6248044967651,12.663818359375,12.7152833938599,12.6851558685303,12.6220216751099,12.6031732559204,12.58544921875,12.62841796875,12.670166015625,12.7023439407349,12.88330078125,12.9797849655151,13.0641603469849,13.1197271347046,13.1541500091553,13.1573247909546,13.1603021621704,13.1583490371704,13.1564455032349,13.3251466751099,13.33837890625,13.3558111190796,13.3763675689697,13.39599609375,13.4291009902954,13.4850587844849,13.5352544784546,13.5564947128296,13.5396480560303,13.5133304595947,13.4726076126099,13.4348630905151,13.4111328125,13.3517084121704,13.2688970565796,13.23583984375,13.2963876724243,13.3353033065796,13.4078121185303,13.4785642623901,13.54345703125,13.5361328125,13.5187501907349,13.4971675872803,13.4885740280151,13.5037107467651,13.5597171783447,13.6161623001099,13.6426267623901,13.6690921783447,13.7852048873901,13.8060054779053,13.8121089935303,13.7891111373901,13.7270021438599,13.5862302780151,13.5882816314697,13.5927734375,13.5968751907349,13.5873050689697,13.6895513534546,13.77099609375,13.8404302597046,13.9049320220947,13.961181640625,14.0074710845947,14.04052734375,14.0355958938599,14.0058107376099,14.004150390625,14.09326171875,14.2083501815796,14.4032230377197,14.4830560684204,14.640625,14.7010736465454,14.7292976379395,14.723484992981,14.651611328125,14.7551279067993,14.7921886444092,14.9220218658447,15.2939939498901,15.7344236373901,15.83837890625,15.9173345565796,16.0972175598145,16.2045402526855,16.2249031066895,16.3071765899658,16.4513187408447,16.5312976837158,16.5470714569092,16.5401363372803,16.5104484558105,16.4921398162842,16.485107421875,16.506591796875,16.5565929412842,16.5819835662842,16.5826168060303,16.6036128997803,16.6449222564697,16.6573734283447,16.6409664154053,16.6474609375,16.6769046783447,16.6789073944092,16.6535148620605,16.6458988189697,16.6559581756592,16.5802745819092,16.4188480377197,16.3111324310303,16.2572269439697,16.202880859375,16.1481456756592,16.1380367279053,16.1725101470947,16.1688003540039,16.1269054412842,16.1183109283447,16.14404296875,16.1352043151855,16.1103019714355,16.0970211029053,16.0911121368408,16.0591793060303,15.9734869003296,15.8538579940796,15.7003898620605,15.6168947219849,15.6033191680908,15.57177734375,15.5352535247803,15.5104494094849,15.4966306686401,15.5048828125,15.4530277252197,15.3404302597046,15.2896480560303,15.2624025344849,15.2425289154053,15.2235355377197,15.1866693496704,15.1312494277954,15.0820808410645,15.0390129089355,14.9746589660645,14.8890132904053,14.8169927597046,14.8090333938599],[1.33144509792328,1.26538074016571,1.32553732395172,1.42343747615814,1.44707024097443,1.41596662998199,1.39223647117615,1.36523413658142,1.33144509792328],[-58.4351577758789,-58.4922866821289,-58.4398422241211,-58.4153289794922,-58.3832015991211,-58.3822250366211,-58.4017562866211,-58.4351577758789],[-54.0656242370605,-54.1142578125,-54.0811500549316,-54.0850563049316,-54.1014633178711,-54.1077117919922,-54.1081047058105,-54.1898422241211,-54.2623062133789,-54.248046875,-54.3083992004395,-54.3033218383789,-54.2789039611816,-54.2511711120605,-54.2886734008789,-54.3604469299316,-54.3822250366211,-54.4583015441895,-54.5541000366211,-54.5676765441895,-54.5680694580078,-54.5547828674316,-54.6375007629395,-54.7108383178711,-54.7634773254395,-54.7923812866211,-54.8342781066895,-54.8667984008789,-54.8529281616211,-54.7798843383789,-54.6937522888184,-54.5707015991211,-54.5344734191895,-54.51123046875,-54.49609375,-54.4666023254395,-54.404296875,-54.3660125732422,-54.3394508361816,-54.3409156799316,-54.3115234375,-54.271484375,-54.1558609008789,-54.16748046875,-54.134765625,-54.0767593383789,-54.0420875549316,-54.0289077758789,-54.0080070495605,-53.99560546875,-53.9937477111816,-53.9840850830078,-54.0091819763184,-54.060546875,-54.0656242370605],[-15.9977540969849,-16.0040035247803,-15.9567375183105,-15.9061527252197,-15.9127941131592,-15.9708976745605,-15.9977540969849],[-7.97431659698486,-7.97578096389771,-7.96748065948486,-7.94374990463257,-7.90576124191284,-7.88261699676514,-7.88593769073486,-7.91259765625,-7.93544960021973,-7.95615196228027,-7.97431659698486],[-11.6651363372803,-11.6813478469849,-11.6773443222046,-11.5908203125,-11.5712890625,-11.5788078308105,-11.6296882629395,-11.6651363372803],[-11.7978515625,-11.8322257995605,-11.81494140625,-11.7888669967651,-11.7119140625,-11.6639642715454,-11.6439447402954,-11.6107416152954,-11.5943355560303,-11.57958984375,-11.5379886627197,-11.4947271347046,-11.4719724655151,-11.4928703308105,-11.6958990097046,-11.7587900161743,-11.7978515625],[-10.7578125,-10.7750968933105,-10.7702140808105,-10.7794923782349,-10.8514652252197,-10.8414058685303,-10.8440427780151,-10.7847652435303,-10.7560548782349,-10.7606449127197,-10.7030277252197,-10.6743173599243,-10.6611328125,-10.6798830032349,-10.6930665969849,-10.7452154159546,-10.7578125],[-10.3873043060303,-10.4460935592651,-10.4364261627197,-10.4768552780151,-10.4538087844849,-10.5060548782349,-10.7099609375,-10.7761716842651,-10.8232421875,-10.8321285247803,-10.8078117370605,-10.8244142532349,-10.7848625183105,-10.7643547058105,-10.7168951034546,-10.56640625,-10.4913091659546,-10.45458984375,-10.3614263534546,-10.3319330215454,-10.3264646530151,-10.2824220657349,-10.2043943405151,-10.2055673599243,-10.2379884719849,-10.3518552780151,-10.3712892532349,-10.3873043060303],[-9.62568283081055,-9.73271465301514,-9.76972675323486,-9.69111251831055,-9.71894550323486,-9.68164157867432,-9.60039138793945,-9.51377010345459,-9.44814395904541,-9.35341835021973,-9.36845779418945,-9.62568283081055],[-9.27265644073486,-9.43330097198486,-9.41865253448486,-9.42158222198486,-9.53623104095459,-9.5888671875,-9.69160175323486,-9.71503925323486,-9.76738357543945,-9.86279296875,-9.87831974029541,-9.91386699676514,-9.92861270904541,-9.89472675323486,-9.8212890625,-9.81240272521973,-9.79150390625,-9.76347637176514,-9.72607421875,-9.63681602478027,-9.53212833404541,-9.470703125,-9.35380840301514,-9.31123065948486,-9.26865196228027,-9.27265644073486],[-8.99550724029541,-9.00957012176514,-9.00732421875,-9.06113338470459,-9.14033222198486,-9.18124961853027,-9.16035175323486,-9.16865253448486,-9.16318416595459,-9.12343692779541,-9.08408164978027,-9.08076095581055,-9.03398418426514,-8.99550724029541],[-9.12353515625,-9.1259765625,-9.11376953125,-9.10966873168945,-9.07499980926514,-9.025390625,-8.99306678771973,-9.00136756896973,-9.02206993103027,-9.02998065948486,-9.09375,-9.12353515625],[-8.75888729095459,-8.79404258728027,-8.78310585021973,-8.73017501831055,-8.70595645904541,-8.69707012176514,-8.70371055603027,-8.73457050323486,-8.75888729095459],[-8.82197284698486,-8.82578086853027,-8.78593826293945,-8.67812538146973,-8.76484394073486,-8.79736328125,-8.82197284698486],[-8.68418025970459,-8.76308536529541,-8.73642539978027,-8.66874980926514,-8.58720779418945,-8.56562519073486,-8.56093788146973,-8.54423809051514,-8.50820350646973,-8.53681564331055,-8.556640625,-8.60664081573486,-8.62265586853027,-8.646484375,-8.68418025970459],[-8.71347713470459,-8.72812557220459,-8.69999980926514,-8.56503963470459,-8.51992130279541,-8.43242168426514,-8.4208984375,-8.47509765625,-8.55507850646973,-8.57548904418945,-8.65068435668945,-8.71347713470459],[-8.50790977478027,-8.521484375,-8.48476600646973,-8.45136737823486,-8.39921855926514,-8.37949180603027,-8.41445350646973,-8.45038986206055,-8.50790977478027],[-8.31396484375,-8.61201095581055,-8.66484355926514,-8.69892597198486,-8.79902362823486,-8.83749961853027,-8.85507774353027,-8.96181583404541,-9.09247970581055,-9.13261699676514,-9.19199275970459,-9.31689453125,-9.49033164978027,-9.57373046875,-9.61123085021973,-9.58955097198486,-9.39287090301514,-9.30800724029541,-9.271484375,-9.15683555603027,-8.96386814117432,-8.62060546875,-8.53925800323486,-8.37275409698486,-8.32822322845459,-8.33837985992432,-8.33632755279541,-8.31650447845459,-8.31396484375],[-8.2421875,-8.32402324676514,-8.50634765625,-8.56914043426514,-8.57265663146973,-8.61201095581055,-8.52363300323486,-8.49970722198486,-8.44540977478027,-8.33779335021973,-8.26992225646973,-8.25830078125,-8.27529335021973,-8.33330154418945,-8.31484317779541,-8.26279354095459,-8.21162128448486,-8.16123008728027,-8.09638690948486,-7.98466777801514,-7.96572256088257,-8.00595664978027,-8.16484355926514,-8.216796875,-8.2421875],[-8.17158222198486,-8.1962890625,-8.07294845581055,-8.01083946228027,-7.97099637985229,-7.95878934860229,-8.09619140625,-8.17158222198486],[-8.10810565948486,-8.12324237823486,-8.11748027801514,-8.01435565948486,-7.93798780441284,-7.86787176132202,-7.85546875,-7.88261699676514,-7.94121074676514,-8.01591777801514,-8.08183574676514,-8.10810565948486],[-7.92304658889771,-7.93681621551514,-7.8828125,-7.86591815948486,-7.80576133728027,-7.70781278610229,-7.64023447036743,-7.57402324676514,-7.61259746551514,-7.69570302963257,-7.72285127639771,-7.77792978286743,-7.876953125,-7.92304658889771],[-8.53427791595459,-8.55742168426514,-8.47382831573486,-8.37167930603027,-8.26044940948486,-8.20341777801514,-8.1962890625,-8.10332012176514,-8.04072284698486,-7.95976591110229,-7.92666006088257,-7.90693330764771,-7.81806659698486,-7.75908231735229,-7.65136766433716,-7.57714891433716,-7.54472684860229,-7.60429716110229,-7.72236394882202,-7.78916072845459,-7.83740282058716,-7.90351581573486,-7.90956974029541,-7.97617197036743,-7.994140625,-8.02900314331055,-8.32695293426514,-8.40605449676514,-8.46347618103027,-8.53427791595459],[-7.09716796875,-7.12109375,-7.08896493911743,-7.0126953125,-6.97294902801514,-7.04326200485229,-7.09716796875],[-7.33037090301514,-7.36562490463257,-7.42568349838257,-7.39306688308716,-7.359375,-7.34150409698486,-7.35302734375,-7.32363271713257,-7.18046855926514,-6.91093730926514,-6.76162099838257,-6.71523475646973,-6.63828086853027,-6.60888624191284,-6.64101552963257,-6.76406240463257,-6.89199256896973,-6.95722627639771,-7.11376953125,-7.16035175323486,-7.28046846389771,-7.30859375,-7.31367158889771,-7.33037090301514],[7.43632841110229,7.41025400161743,7.47451162338257,7.57084989547729,7.62202167510986,7.63720655441284,7.60737323760986,7.58242177963257,7.53510761260986,7.43632841110229],[8.48515605926514,8.4541015625,8.31084060668945,8.18793964385986,8.15761756896973,8.12529277801514,8.07114219665527,7.89643526077271,7.75937509536743,7.73642539978027,7.53823328018188,7.46303701400757,7.39858388900757,7.31440448760986,7.23261690139771,7.09467792510986,6.95122098922729,6.90654277801514,6.94697284698486,7.08857440948486,7.18354463577271,7.341796875,7.38627910614014,7.44248056411743,7.54501962661743,7.66572284698486,7.75327205657959,7.75336933135986,7.70058584213257,7.71586942672729,7.79111289978027,7.81870126724243,7.85664129257202,8.05517578125,8.14531230926514,8.200927734375,8.214599609375,8.33583927154541,8.42973613739014,8.48759746551514,8.48427772521973,8.44228553771973,8.42475605010986,8.52646541595459,8.58154296875,8.62978553771973,8.65625,8.61513710021973,8.625732421875,8.58876991271973,8.576904296875,8.69589900970459,8.76596641540527,8.84311485290527,8.85634803771973,8.88115215301514,8.897705078125,8.98740196228027,9.04921817779541,9.07011699676514,9.06088924407959,9.04755878448486,9.06962871551514,9.10356426239014,9.14692401885986,9.26333045959473,9.30224609375,9.37358379364014,9.48417949676514,9.56191349029541,9.60063552856445,9.63422870635986,9.671142578125,9.70497989654541,9.78720664978027,9.86215782165527,9.89814472198486,9.92978477478027,9.87539100646973,9.92275333404541,9.99301719665527,9.99418926239014,9.99545955657959,9.99653339385986,9.97773456573486,9.92534160614014,9.84316444396973,9.78632831573486,9.66162109375,9.51645469665527,9.38535213470459,9.31425857543945,9.28935527801514,9.26113319396973,9.19448184967041,9.12236309051514,9.09526348114014,9.08168888092041,9.05917930603027,8.97880840301514,8.86757850646973,8.76376914978027,8.68754863739014,8.66030216217041,8.52997970581055,8.40058517456055,8.36420917510986,8.33525371551514,8.32168006896973,8.33027362823486,8.31948184967041,8.315673828125,8.36210918426514,8.48095703125,8.49550819396973,8.48515605926514],[14.416015625,14.4113759994507,14.3603019714355,14.3702144622803,14.3291492462158,14.2969732284546,14.2527341842651,14.1636714935303,14.124755859375,14.072265625,14.0320796966553,14.0155277252197,14.0001459121704,13.9789543151855,13.9642086029053,13.9045400619507,13.854248046875,13.8509769439697,13.8753900527954,13.9426755905151,13.9873533248901,13.9605960845947,13.9046382904053,13.879638671875,13.8949708938599,13.8899898529053,13.8410654067993,13.8126945495605,13.6499509811401,13.580322265625,13.5213861465454,13.5060062408447,13.4831047058105,13.4513673782349,13.399169921875,13.3857908248901,13.2851572036743,13.2244138717651,13.1806640625,13.1687498092651,13.1640138626099,13.2135248184204,13.2810544967651,13.281494140625,13.2591800689697,13.2449703216553,13.1971673965454,13.1839351654053,13.2832517623901,13.4780759811401,13.5091304779053,13.5601081848145,13.6831541061401,13.7365226745605,13.7830076217651,13.8347654342651,13.904052734375,13.9973640441895,14.0456056594849,14.0550775527954,14.0500974655151,14.0770015716553,14.1413087844849,14.1827144622803,14.2246580123901,14.2412595748901,14.2772464752197,14.3470697402954,14.3900880813599,14.4099111557007,14.4137706756592,14.4311037063599,14.4276361465454,14.416015625],[43.9345703125,43.982421875,43.98974609375,43.9529762268066,43.9014167785645,43.8940925598145,43.9345703125],[11.2584466934204,10.9823246002197,10.7142086029053,10.4332523345947,9.97348594665527,9.8076171875,9.56411170959473,9.45175743103027,9.23271465301514,8.964599609375,8.67959022521973,8.443359375,8.22216796875,7.99707078933716,7.99707078933716,7.99707078933716,7.99707078933716,8.026123046875,8.1181640625,8.23496055603027,8.3798828125,8.48300743103027,8.5908203125,8.7001953125,8.78608417510986,8.89306640625,8.98603534698486,9.00883769989014,9.15078067779541,9.33740234375,9.34072208404541,9.37949275970459,9.48027324676514,9.60908222198486,9.77016639709473,9.87998008728027,9.92622089385986,10.0125980377197,10.1408205032349,10.2030754089355,10.2573728561401,10.36962890625,10.4917488098145,10.5675783157349,10.6000003814697,10.6213865280151,10.7869138717651,10.8459959030151,10.9032220840454,10.9602546691895,10.9993171691895,11.1943359375,11.36572265625,11.4998054504395,11.346435546875,11.0354490280151,10.7842779159546,10.55078125,10.4718751907349,10.4302244186401,10.4367198944092,10.6497554779053,10.8039064407349,10.8358888626099,10.793701171875,10.7811031341553,10.7341804504395,10.7459955215454,10.9253902435303,11.0999021530151,11.1740236282349,11.1748046875,11.1120119094849,11.1393556594849,11.2901372909546,11.3205070495605,11.3226566314697,11.2548828125,11.2584466934204],[-1.69531214237213,-1.61318385601044,-1.57226550579071,-1.44951164722443,-1.22050786018372,-1.04746079444885,-0.870312571525574,-0.728711247444153,-0.307324260473251,0.535400390625,1.378173828125,2.220947265625,2.642333984375,2.81464838981628,2.84243178367615,2.99706983566284,3.20166015625,3.59047842025757,3.80161118507385,3.97773432731628,4.03129911422729,4.137939453125,4.20166015625,4.21225595474243,4.2919921875,4.32421827316284,4.361083984375,4.4453125,4.56332969665527,4.64448261260986,4.75039052963257,4.84033203125,4.85498046875,4.91142559051514,4.93076181411743,4.95053672790527,4.95097637176514,4.93120145797729,4.915771484375,4.89990234375,4.91201210021973,5.12167978286743,5.45542001724243,5.66826152801514,5.99721670150757,6.23466730117798,6.49726581573486,6.73725605010986,7.02602577209473,7.20786094665527,7.49047803878784,7.75932645797729,7.99707078933716,8.22216796875,8.443359375,8.67959022521973,8.964599609375,9.23271465301514,9.45175743103027,9.56411170959473,9.8076171875,9.97348594665527,10.4332523345947,10.7142086029053,10.9823246002197,11.2584466934204,11.2708492279053,11.3427248001099,11.450927734375,11.529296875,11.7275390625,11.8231935501099,11.9437990188599,11.9836912155151,11.8419914245605,11.8307123184204,11.7450189590454,11.6576662063599,11.5051279067993,11.3356447219849,11.0767574310303,10.6568841934204,10.5958976745605,10.5358390808105,10.488525390625,10.4719724655151,10.4447755813599,10.4339351654053,10.479736328125,10.5298337936401,10.5546388626099,10.4986810684204,10.4752445220947,10.422607421875,10.3865232467651,10.4031248092651,10.4310550689697,10.3851566314697,10.3355464935303,10.2531251907349,9.92417049407959,9.71049785614014,9.42817401885986,9.24116230010986,9.10927772521973,8.84526348114014,8.61958026885986,8.50942325592041,8.19980430603027,7.96254825592041,7.65952205657959,7.46953058242798,7.29697275161743,6.99052762985229,6.77734375,6.40786170959473,6.17363309860229,5.49438428878784,4.95268535614014,4.49702119827271,3.96826124191284,3.28564476966858,2.47514677047729,2.30986332893372,1.81015634536743,1.39096677303314,1.10590827465057,0.857861042022705,0.621631145477295,-0.175683751702309,-0.250781327486038,-0.321484327316284,-0.45654308795929,-0.510058462619781,-0.737988531589508,-0.856152474880219,-0.973046779632568,-1.05556643009186,-1.15058577060699,-1.20341813564301,-1.43007802963257,-1.57851529121399,-1.69531214237213],[46.7624015808105,46.7528343200684,46.7671852111816,46.7982444763184,46.8072776794434,46.8110847473145,46.8015632629395,46.7786598205566,46.7624015808105],[46.8384780883789,46.7953147888184,46.8194351196289,46.84765625,46.9159698486328,46.9356422424316,47.0679664611816,47.0895500183105,47.0989723205566,47.0709953308105,47.0350074768066,46.953857421875,46.89990234375,46.8609848022461,46.8384780883789],[46.1085929870605,46.0506820678711,45.9754905700684,45.9407691955566,45.8995094299316,45.8694801330566,45.7793922424316,45.7352523803711,45.7374038696289,45.7489776611328,45.7581062316895,45.7498054504395,45.7225074768066,45.6620101928711,45.5974617004395,45.5364761352539,45.5174789428711,45.5000991821289,45.4844207763672,45.4678688049316,45.4275398254395,45.3653297424316,45.321533203125,45.2935562133789,45.291748046875,45.2413101196289,45.205078125,45.1925315856934,45.171875,45.1478996276855,45.1222648620605,45.10986328125,45.0751457214355,45.032958984375,45.022216796875,45.0213851928711,45.0089836120605,44.9907684326172,44.9734382629395,44.9577140808105,44.9419937133789,44.9188499450684,44.9006843566895,44.8808097839355,44.8733901977539,44.8700675964355,44.86181640625,44.82666015625,44.7900886535645,44.7554206848145,44.71044921875,44.6806640625,44.6661148071289,44.619873046875,44.5419464111328,44.5606918334961,44.6761245727539,44.7062492370605,44.6509742736816,44.6055183410645,44.5699195861816,44.5555152893066,44.5623550415039,44.5403327941895,44.4895973205566,44.4354476928711,44.3779792785645,44.3383293151855,44.3164558410645,44.2864761352539,44.248291015625,44.23779296875,44.22021484375,44.194091796875,44.1485824584961,44.0752944946289,44.0180168151855,44.0074195861816,43.9695320129395,43.8621101379395,43.7812957763672,43.7401351928711,43.7066383361816,43.6654777526855,43.6022491455078,43.5188484191895,43.4544944763184,43.3910636901855,43.3541526794434,43.3007316589355,43.2523422241211,43.18798828125,43.1420402526855,43.0970687866211,43.0759773254395,43.0182609558105,42.9857406616211,42.8839378356934,42.8784675598145,42.8703117370605,42.8424797058105,42.7916526794434,42.7507820129395,42.70947265625,42.6291007995605,42.5433120727539,42.50390625,42.481201171875,42.4409675598145,42.359130859375,42.328857421875,42.31396484375,42.3217277526855,42.349853515625,42.358154296875,42.3249969482422,42.3046379089355,42.320068359375,42.3220672607422,42.3083992004395,42.3031234741211,42.2677230834961,42.2421379089355,42.2475090026855,42.2808113098145,42.3284187316895,42.3500022888184,42.387451171875,42.4232444763184,42.595458984375,42.6515121459961,42.6698226928711,42.6819801330566,42.6814918518066,42.7514190673828,42.83154296875,42.8747062683105,42.9132308959961,42.9561996459961,43.0430183410645,43.0916976928711,43.1160163879395,43.1516609191895,43.1734390258789,43.2139663696289,43.2379379272461,43.2610816955566,43.2585945129395,43.2263679504395,43.1986312866211,43.1784172058105,43.13037109375,43.099853515625,43.0709457397461,43.0341796875,42.9530258178711,42.924560546875,42.8790550231934,42.85791015625,42.8279304504395,42.852783203125,42.8928718566895,42.9354515075684,42.968505859375,43.0816383361816,43.0965347290039,43.1097679138184,43.1639633178711,43.1734390258789,43.2122573852539,43.3428230285645,43.4139671325684,43.449951171875,43.4850120544434,43.5210456848145,43.5332984924316,43.5843772888184,43.591796875,43.5934562683105,43.5675773620605,43.5620613098145,43.5951652526855,43.6428718566895,43.7035636901855,43.8447723388672,43.9433097839355,43.9650382995605,43.9834480285645,43.9933586120605,43.985107421875,43.97802734375,43.9871063232422,44.0110855102539,44.04345703125,44.073486328125,44.1544914245605,44.225830078125,44.2805671691895,44.3025398254395,44.3302726745605,44.3599624633789,44.41455078125,44.4837875366211,44.52734375,44.6095695495605,44.69677734375,44.7806625366211,44.8585433959961,44.8809051513672,44.8974609375,44.9142570495605,44.8996124267578,44.871337890625,44.8691902160645,44.9040031433105,44.9193840026855,44.9175300598145,44.9109878540039,44.9267578125,44.9737815856934,45.13720703125,45.1517105102539,45.1754875183105,45.1962394714355,45.1677742004395,45.1672859191895,45.1730003356934,45.1890640258789,45.2124977111816,45.2306175231934,45.2454109191895,45.26806640625,45.2779808044434,45.3369140625,45.3995132446289,45.4658203125,45.502197265625,45.5149917602539,45.5272483825684,45.5580062866211,45.600830078125,45.6558113098145,45.76708984375,45.8357429504395,45.8655281066895,45.9076156616211,45.9317398071289,45.931396484375,45.959716796875,45.982666015625,46.009521484375,46.0161628723145,45.9870109558105,45.9844245910645,46.0028800964355,46.0285148620605,46.0498046875,46.064453125,46.0873527526855,46.1551742553711,46.169189453125,46.1519050598145,46.1458969116211,46.1614761352539,46.1418914794922,46.1260261535645,46.1085929870605],[0.12065402418375,0.0473635569214821,0.0663084015250206,0.117383003234863,0.227343916893005,0.280127078294754,0.340283066034317,0.400244265794754,0.404394775629044,0.325634837150574,0.243457213044167,0.12065402418375],[1.56772482395172,1.54155266284943,1.56357431411743,1.60336911678314,1.68017590045929,1.69912123680115,1.68305671215057,1.66196262836456,1.63110375404358,1.56772482395172],[5.35898447036743,5.28823232650757,5.18740224838257,5.01347637176514,4.95878934860229,4.91469717025757,4.83652353286743,4.74931669235229,4.69199228286743,4.5830078125,4.48501014709473,4.42802762985229,4.337646484375,4.24140596389771,4.20249032974243,4.17094707489014,4.14003896713257,4.05410146713257,3.90107417106628,3.83442378044128,3.76943373680115,3.70595693588257,3.62939429283142,3.62041044235229,3.58955073356628,3.53051781654358,3.44853496551514,3.35332012176514,3.17875957489014,3.13818383216858,2.99360370635986,2.87485361099243,2.81787109375,2.71372056007385,2.46152353286743,2.41611313819885,2.34331035614014,2.34257817268372,2.33579087257385,2.32675743103027,2.32753920555115,2.35981440544128,2.39794921875,2.441650390625,2.45473623275757,2.43955063819885,2.45039033889771,2.49736332893372,2.54833984375,2.59765601158142,2.59296870231628,2.54833984375,2.53920912742615,2.55078101158142,2.54750967025757,2.49965834617615,2.48876929283142,2.44062495231628,2.41874980926514,2.40615224838257,2.48950219154358,2.51660132408142,2.52045917510986,2.51596665382385,2.49750995635986,2.39277315139771,2.36440420150757,2.34130835533142,2.29951167106628,2.25903344154358,2.23676753044128,2.15815424919128,2.09511709213257,2.03955101966858,1.97666037082672,1.88749992847443,1.85707998275757,1.84223628044128,1.88535141944885,1.92387712001801,1.93232417106628,1.94213843345642,1.97480452060699,2.00507855415344,2.01601552963257,2.03647470474243,2.11489272117615,2.22666001319885,2.27714848518372,2.32597661018372,2.39536118507385,2.45683622360229,2.51323246955872,2.53217768669128,2.60898447036743,2.63750004768372,2.64111304283142,2.66567397117615,2.74785161018372,2.76826167106628,2.77553701400757,2.83325242996216,2.853271484375,2.8828125,2.96337890625,3.00307655334473,3.07856464385986,3.10888671875,3.14228510856628,3.164306640625,3.21884775161743,3.35361313819885,3.37709975242615,3.37543940544128,3.36225628852844,3.35429644584656,3.35283207893372,3.37094736099243,3.39453148841858,3.42373061180115,3.51738262176514,3.58828139305115,3.67597627639771,3.78725576400757,3.856689453125,4.00195360183716,4.10166025161743,4.17192411422729,4.23647499084473,4.34995126724243,4.453125,4.50678682327271,4.57709980010986,4.66816425323486,4.72431659698486,4.77929735183716,4.82041025161743,4.880615234375,4.92304706573486,4.92905282974243,4.95449209213257,4.99106454849243,5.00068330764771,5.00449228286743,5.00458955764771,5.02016639709473,5.049560546875,5.10585975646973,5.15703105926514,5.17851591110229,5.19540977478027,5.21420860290527,5.23154306411743,5.24287128448486,5.24677705764771,5.33535146713257,5.37397432327271,5.44516658782959,5.48525428771973,5.54843711853027,5.52890634536743,5.64379835128784,5.73720693588257,5.82939481735229,5.93867206573486,5.99287128448486,5.937744140625,5.88535165786743,5.79545927047729,5.69931650161743,5.79545927047729,5.89262723922729,5.961669921875,5.98588895797729,5.95263671875,5.99345731735229,5.98833036422729,5.90986299514771,5.85634756088257,5.80790996551514,5.72050809860229,5.60888671875,5.50224590301514,5.35898447036743],[49.0727081298828,49.0313987731934,48.9832496643066,48.9335441589355,48.8734855651855,48.7450675964355,48.6858406066895,48.5685043334961,48.4053230285645,48.3933601379395,48.3380851745605,48.3465805053711,48.3783683776855,48.4014663696289,48.4185066223145,48.4636726379395,48.4957008361816,48.5218772888184,48.55224609375,48.553466796875,48.5105972290039,48.5059051513672,48.5196762084961,48.545654296875,48.5497093200684,48.5269050598145,48.4951171875,48.2955589294434,48.2220191955566,48.1466331481934,48.13134765625,48.155029296875,48.1998062133789,48.2230949401855,48.2128410339355,48.162109375,48.1106948852539,48.0730476379395,48.0508270263672,48.0002899169922,47.939453125,47.8926734924316,47.8528823852539,47.8064956665039,47.7871589660645,47.7770042419434,47.763427734375,47.7668952941895,47.7701644897461,47.8099098205566,47.8875961303711,47.9909172058105,47.9933586120605,48.0043449401855,48.0120582580566,48.0059585571289,48.03955078125,48.083251953125,48.1980934143066,48.3869132995605,48.44140625,48.5035171508789,48.5509262084961,48.5885772705078,48.5988273620605,48.6769027709961,48.78076171875,48.841064453125,48.8609352111816,48.8428230285645,48.8277816772461,48.8418464660645,48.8881340026855,48.9286117553711,48.9711418151855,48.9987335205078,49.011962890625,49.0365219116211,49.0651359558105,49.1193351745605,49.1797866821289,49.2245597839355,49.2573738098145,49.3362274169922,49.3639144897461,49.3909187316895,49.4210929870605,49.4646987915039,49.491455078125,49.4884757995605,49.4939956665039,49.5092277526855,49.5107917785645,49.498291015625,49.4482917785645,49.396240234375,49.3999977111816,49.5114250183105,49.5248565673828,49.5636215209961,49.5977058410645,49.5763664245605,49.5047836303711,49.4471206665039,49.4243659973145,49.4066429138184,49.3960456848145,49.3895988464355,49.3721694946289,49.3185539245605,49.2699699401855,49.2352066040039,49.2043952941895,49.1923332214355,49.2040023803711,49.2213859558105,49.1813011169434,49.270751953125,49.31640625,49.337646484375,49.3655281066895,49.38525390625,49.3840827941895,49.392333984375,49.3901863098145,49.3812026977539,49.3917007446289,49.3699226379395,49.3286590576172,49.314697265625,49.3170890808105,49.33984375,49.4182624816895,49.4170417785645,49.429443359375,49.4287605285645,49.4119606018066,49.3819313049316,49.3434600830078,49.299072265625,49.24609375,49.2095222473145,49.1532249450684,49.081298828125,49.0727081298828],[46.4999046325684,46.5244140625,46.5346183776855,46.5213890075684,46.5079078674316,46.4838409423828,46.3891105651855,46.3728523254395,46.3822250366211,46.371337890625,46.3053703308105,46.2776374816895,46.2578620910645,46.2339859008789,46.2132301330566,46.2007331848145,46.1719245910645,46.1399917602539,46.1092262268066,46.0484848022461,45.9836921691895,45.9044456481934,45.8621597290039,45.8340339660645,45.797607421875,45.7326164245605,45.7177238464355,45.659912109375,45.6455078125,45.6126441955566,45.5796852111816,45.5415534973145,45.5022926330566,45.467041015625,45.44140625,45.4507789611816,45.4999046325684,45.49267578125,45.4673347473145,45.4782218933105,45.5084953308105,45.5714836120605,45.610107421875,45.6512718200684,45.6572265625,45.645263671875,45.59521484375,45.5057640075684,45.4814453125,45.4866218566895,45.4851570129395,45.4778327941895,45.5094223022461,45.5033683776855,45.4826164245605,45.4498023986816,45.4283714294434,45.4767570495605,45.5168914794922,45.5359382629395,45.5876007080078,45.5819816589355,45.5928726196289,45.6148452758789,45.680419921875,45.7612800598145,45.7919921875,45.8123550415039,45.8341331481934,45.961669921875,45.9797859191895,45.9737777709961,45.9871101379395,46.0092277526855,46.03955078125,46.089111328125,46.1331062316895,46.1577644348145,46.1770515441895,46.1965827941895,46.2166023254395,46.2235336303711,46.2123069763184,46.2249526977539,46.2616233825684,46.3175315856934,46.3691902160645,46.4150848388672,46.4485359191895,46.462890625,46.520263671875,46.5143051147461,46.51123046875,46.4981918334961,46.482177734375,46.4619140625,46.4407234191895,46.4279327392578,46.4161148071289,46.4170417785645,46.3997077941895,46.4129371643066,46.4360809326172,46.4634284973145,46.4991226196289,46.5445785522461,46.5804710388184,46.6059074401855,46.6132354736328,46.6259765625,46.6429672241211,46.629638671875,46.6546401977539,46.6984367370605,46.7107429504395,46.7112770080566,46.6776351928711,46.6972160339355,46.7058563232422,46.8013687133789,46.8448257446289,46.86328125,46.8572769165039,46.8279800415039,46.7825202941895,46.7216339111328,46.7047843933105,46.6808128356934,46.638671875,46.6072273254395,46.5220718383789,46.4999046325684],[56.29052734375,56.2401847839355,56.2437477111816,56.3108901977539,56.483642578125,56.5689926147461,56.8768577575684,56.9020004272461,57.0906715393066,57.1746101379395,57.2501945495605,57.3177719116211,57.3450660705566,57.3322715759277,57.3198204040527,57.2804679870605,57.229248046875,57.2080078125,56.9852027893066,56.8405265808105,56.8052253723145,56.29052734375],[57.8359375,57.8121070861816,57.7416000366211,57.7296867370605,57.7061996459961,57.4831047058105,57.3983421325684,57.3864784240723,57.361083984375,57.3235359191895,57.2427253723145,57.1969223022461,57.1630363464355,57.087646484375,56.9782218933105,56.9315414428711,56.9205093383789,57.0101547241211,57.0832023620605,57.13330078125,57.2117156982422,57.2718734741211,57.3390617370605,57.4491729736328,57.5566368103027,57.6108894348145,57.6551284790039,57.7568359375,57.8305702209473,57.8637237548828,57.8331527709961,57.9001960754395,57.9154739379883,57.8999977111816,57.8359375],[57.922607421875,57.8602523803711,57.8649864196777,57.9110374450684,57.9813461303711,57.9775428771973,57.962890625,57.922607421875],[59.0291023254395,59.0195808410645,59.0226097106934,59.0690383911133,59.0891151428223,59.1078643798828,59.1045913696289,59.0291023254395],[59.4703598022461,59.437255859375,59.4778327941895,59.4857902526855,59.5258293151855,59.5478019714355,59.5346183776855,59.5246086120605,59.4921913146973,59.4703598022461],[65.8052749633789,65.7822265625,65.8285140991211,65.8053207397461,65.8043441772461,65.7861328125,65.7499008178711,65.7353515625,65.7864761352539,65.8709487915039,65.8065414428711,65.7943344116211,65.8526306152344,65.8621063232422,65.8426742553711,65.7911605834961,65.7506332397461,65.7249984741211,65.6215362548828,65.5975570678711,65.6109390258789,65.5837936401367,65.5701141357422,65.5528869628906,65.5302200317383,65.5323715209961,65.5083541870117,65.4703598022461,65.4371109008789,65.4240264892578,65.4033737182617,65.4081039428711,65.3865661621094,65.3585891723633,65.3311538696289,65.3165512084961,65.2991256713867,65.2613754272461,65.2545394897461,65.3208465576172,65.3174362182617,65.282958984375,65.245361328125,65.2069854736328,65.1607894897461,65.1257781982422,65.0126953125,64.9412612915039,64.8769073486328,64.8086929321289,64.7743148803711,64.7247085571289,64.6293487548828,64.5443420410156,64.4630889892578,64.4161148071289,64.3795852661133,64.2991714477539,64.177978515625,63.8678207397461,63.8262672424316,63.7737274169922,63.722900390625,63.6624526977539,63.6105499267578,63.5381813049316,63.4633293151855,63.4580116271973,63.4872589111328,63.509033203125,63.4602012634277,63.4243659973145,63.4774894714355,63.4287567138672,63.3473625183105,63.2377433776855,63.2574691772461,63.2381324768066,63.2241249084473,63.2065925598145,63.1972694396973,63.1982460021973,63.1765670776367,63.1782722473145,63.1264190673828,63.0635261535645,63.0375022888184,63.0321273803711,62.9963836669922,62.9888648986816,62.9585952758789,62.9283218383789,62.8958511352539,62.8490715026855,62.8122100830078,62.7893562316895,62.7906723022461,62.8119621276855,62.8360328674316,62.8338890075684,62.8867683410645,62.8731956481934,62.8305130004883,62.7861328125,62.7210464477539,62.6798820495605,62.6594734191895,62.6406288146973,62.6262702941895,62.6005363464355,62.5784645080566,62.5027351379395,62.5008735656738,62.4508781433105,62.4510231018066,62.4825210571289,62.5083961486816,62.4627914428711,62.426513671875,62.334716796875,62.2636680603027,62.2330093383789,62.2123069763184,62.1663055419922,62.0226516723633,61.9661140441895,61.8663101196289,61.7820816040039,61.7406768798828,61.6844711303711,61.6916961669922,61.7245635986328,61.6563491821289,61.5757331848145,61.504638671875,61.4583015441895,61.3816871643066,61.3576164245605,61.3119659423828,61.2782707214355,61.2492713928223,61.1465301513672,60.9858360290527,60.9518585205078,60.8121604919434,60.7631874084473,60.7007789611816,60.6408233642578,60.6417922973633,60.6427230834961,60.6276817321777,60.5852546691895,60.5351524353027,60.539306640625,60.5800743103027,60.5897941589355,60.5114250183105,60.4079055786133,60.3615226745605,60.3371124267578,60.2535667419434,60.1528816223145,60.1192359924316,60.0794906616211,60.0258827209473,59.9801750183105,59.9422874450684,59.8277854919434,59.7572288513184,59.7329635620117,59.6573715209961,59.6009292602539,59.5657730102539,59.4903793334961,59.4768562316895,59.4376449584961,59.4205093383789,59.4303703308105,59.3593788146973,59.3290519714355,59.3162117004395,59.3314437866211,59.3671379089355,59.3753395080566,59.3686065673828,59.3967323303223,59.407958984375,59.3944854736328,59.3270492553711,59.2919425964355,59.2903327941895,59.1797370910645,59.1322250366211,59.1093711853027,59.0623054504395,59.0026359558105,58.9545860290527,58.9650382995605,58.9162063598633,58.8583984375,58.780517578125,58.7108383178711,58.6541481018066,58.6511688232422,58.6636199951172,58.6366691589355,58.6283226013184,58.6018562316895,58.6128921508789,58.5996551513672,58.5852508544922,58.4925804138184,58.4596176147461,58.4343223571777,58.3028755187988,58.2142562866211,58.1607894897461,57.9175300598145,57.9128952026367,57.8122596740723,57.7609367370605,57.6417503356934,57.5683097839355,57.5006866455078,57.43017578125,57.26513671875,57.1876945495605,57.1417007446289,57.0681648254395,56.9268074035645,56.8086929321289,56.7092781066895,56.5899887084961,56.5008316040039,56.2226066589355,56.1673812866211,56.1249504089355,56.1642074584961,56.1855964660645,56.1830101013184,56.1508331298828,56.1722183227539,56.1619148254395,56.1341323852539,56.0331535339355,56.0199203491211,56.0486297607422,56.0143547058105,55.9767570495605,55.8875465393066,55.8326148986816,55.7291488647461,55.6363792419434,55.5277328491211,55.3966331481934,55.3921890258789,55.4285621643066,55.3463859558105,55.411376953125,55.4815940856934,55.533203125,55.6125984191895,55.6937980651855,55.7481422424316,55.8060531616211,55.8818359375,56.1375961303711,56.2455558776855,56.29052734375,56.29296875,56.2350120544434,56.2421379089355,56.263916015625,56.3468742370605,56.3844223022461,56.4405746459961,56.4557609558105,56.452392578125,56.5155754089355,56.6177253723145,56.649169921875,56.662841796875,56.8232917785645,56.9063987731934,57.2269515991211,57.4469718933105,57.4260711669922,57.5219268798828,57.6126937866211,57.679443359375,57.7176742553711,57.7644538879395,57.9731903076172,58.001220703125,58.1183586120605,58.3399925231934,58.3803253173828,58.3691444396973,58.424072265625,58.4756317138672,58.679931640625,58.8664093017578,58.9227027893066,58.9886207580566,59.0455589294434,59.0782737731934,59.0868682861328,59.0365219116211,58.9095191955566,58.8930206298828,58.9260787963867,59.0186538696289,59.1575660705566,59.2898941040039,59.4314422607422,59.5557594299316,59.5922889709473,59.6971702575684,59.7824745178223,59.8636703491211,59.8913078308105,59.8976097106934,59.9128875732422,59.9672355651855,60.0400390625,60.1067848205566,60.2388687133789,60.3052215576172,60.3544960021973,60.4507331848145,60.5456504821777,60.6896476745605,60.8921394348145,61.002685546875,61.0231895446777,61.04150390625,61.0468254089355,61.0598640441895,61.1082534790039,61.1739730834961,61.2218284606934,61.2902870178223,61.3522987365723,61.4457015991211,61.5413093566895,61.5730018615723,61.6534690856934,61.7207527160645,61.9768562316895,62.1674308776855,62.2137680053711,62.2855949401855,62.5918960571289,62.6600112915039,62.7213401794434,62.8259239196777,62.9194831848145,62.9478492736816,63.0006332397461,63.08251953125,63.0891609191895,63.2917022705078,63.4922370910645,63.595947265625,63.6711959838867,63.8435554504395,63.9404830932617,63.9574203491211,64,64.0504913330078,64.0750961303711,64.0748062133789,64.0406265258789,64.0140151977539,64.0407180786133,64.0955047607422,64.1735305786133,64.2603073120117,64.3877410888672,64.4640121459961,64.5135726928711,64.58154296875,64.7967758178711,64.9461441040039,65.1708526611328,65.2643585205078,65.3014678955078,65.6463928222656,65.7428741455078,65.7932586669922,65.8450164794922,65.9322738647461,66.1293487548828,66.1537094116211,66.1675262451172,66.1910629272461,66.2520523071289,66.3059616088867,66.4898452758789,66.5521011352539,66.7688522338867,66.9764175415039,67.0549774169922,67.0933609008789,67.1550750732422,67.2520065307617,67.3120574951172,67.4258346557617,67.5051803588867,67.5206069946289,67.5517578125,67.6195831298828,67.6283187866211,67.89501953125,68.0301284790039,68.1038131713867,68.0484390258789,67.9648895263672,68.0878372192383,68.1334457397461,68.2006378173828,68.3168411254883,68.4677734375,68.5284118652344,68.555419921875,68.5624008178711,68.5000457763672,68.5011215209961,68.4927291870117,68.46533203125,68.3924331665039,68.3622589111328,68.3563995361328,68.390380859375,68.4775390625,68.5420379638672,68.6073226928711,68.6731491088867,68.7540512084961,68.8487319946289,68.899658203125,68.934326171875,69.0208969116211,69.0333023071289,69.036865234375,68.9798355102539,68.9674835205078,68.937744140625,68.9069290161133,68.8288040161133,68.7874526977539,68.724609375,68.690673828125,68.6509780883789,68.6085433959961,68.5741195678711,68.5206069946289,68.4779815673828,68.4640579223633,68.3910217285156,68.3673324584961,68.3164520263672,68.257568359375,68.1366195678711,68.1303253173828,68.0886764526367,68.017333984375,67.9543914794922,67.9332046508789,67.8751983642578,67.7965774536133,67.6961898803711,67.6143035888672,67.5903854370117,67.5621566772461,67.5178680419922,67.4792022705078,67.4602584838867,67.449951171875,67.4491729736328,67.4400405883789,67.4228973388672,67.32861328125,67.3104934692383,67.267822265625,67.2339401245117,67.1841278076172,67.12939453125,67.068115234375,67.0025863647461,66.9340286254883,66.8778305053711,66.8382263183594,66.810546875,66.7757339477539,66.7068862915039,66.6280212402344,66.5766143798828,66.505859375,66.4807586669922,66.4434127807617,66.3807067871094,66.3042907714844,66.2526397705078,66.2154312133789,66.191162109375,66.1482467651367,66.0603561401367,65.9898452758789,65.8052749633789],[-25.9576187133789,-25.9722652435303,-26.0183582305908,-26.1101551055908,-26.2150402069092,-26.28125,-26.34716796875,-26.4498043060303,-26.52001953125,-26.8394546508789,-26.8248043060303,-26.8111324310303,-26.8174800872803,-26.9606456756592,-27.1736335754395,-27.3058605194092,-27.3099613189697,-27.2955074310303,-27.2383785247803,-27.1123046875,-26.9158210754395,-26.7923812866211,-26.7852554321289,-26.7642593383789,-26.6136722564697,-26.4554691314697,-26.4134769439697,-26.21875,-26.0977535247803,-25.9806632995605,-25.8433589935303,-25.7555656433105,-25.7429695129395,-25.74658203125,-25.8672847747803,-25.9816398620605,-25.96875,-25.9576187133789],[18.0689468383789,18.0689468383789,18.04541015625,18.0191898345947,18.0414066314697,18.0643062591553,18.0689468383789],[-4.69306659698486,-4.78554677963257,-4.75459003448486,-4.69482421875,-4.65029287338257,-4.60927772521973,-4.55878925323486,-4.69306659698486],[37.1085968017578,37.0950202941895,37.0605964660645,36.9611320495605,36.8260231018066,36.7408218383789,36.59716796875,36.56640625,36.5146484375,36.4644012451172,36.3833503723145,36.2724609375,36.2030296325684,36.0733871459961,35.93896484375,35.8099594116211,35.724609375,35.6404304504395,35.5506362915039,35.427490234375,35.2881813049316,35.0273933410645,34.8053207397461,34.7689971923828,34.6123046875,34.4290542602539,34.3865699768066,34.33203125,34.19775390625,34.0476570129395,33.911376953125,33.7683601379395,33.6200180053711,33.5140151977539,33.3722152709961,33.2366218566895,33.0992202758789,32.994873046875,32.829833984375,32.7330551147461,32.5907707214355,32.4655265808105,32.3172874450684,32.361328125,32.3869132995605,32.4574737548828,32.4951171875,32.5337905883789,32.6666984558105,32.7137680053711,32.7349128723145,32.7823219299316,32.8623504638672,32.9496078491211,32.9980964660645,33.0393562316895,33.0885734558105,33.1356925964355,33.1924781799316,33.2495613098145,33.2782211303711,33.3305168151855,33.3704605102539,33.4317359924316,33.4653778076172,33.5002899169922,33.5345687866211,33.5625,33.5850563049316,33.5979499816895,33.6230964660645,33.6675758361816,33.732421875,33.7526359558105,33.783935546875,33.8315925598145,33.8395042419434,33.8395500183105,33.8355941772461,33.8270530700684,33.83935546875,33.8551254272461,33.8941879272461,33.92529296875,33.9586410522461,34.0113258361816,34.0498504638672,34.0568351745605,34.1343231201172,34.2212409973145,34.432373046875,34.4661636352539,34.4951667785645,34.4996109008789,34.5133323669434,34.56689453125,34.6134757995605,34.6579093933105,34.6787109375,34.6328582763672,34.6286163330078,34.6291999816895,34.8521003723145,34.9486351013184,35.060302734375,35.2238273620605,35.2995109558105,35.3505363464355,35.4207000732422,35.5715789794922,35.8492164611816,35.9165534973145,35.9100570678711,35.8314437866211,35.8338890075684,35.9375495910645,35.9727058410645,36.0035133361816,36.1712417602539,36.2034645080566,36.220703125,36.2239265441895,36.2339820861816,36.2635269165039,36.4574203491211,36.50634765625,36.7013664245605,36.7776870727539,36.8025398254395,36.7926750183105,36.7583961486816,36.702392578125,36.6526374816895,36.6559066772461,36.6465797424316,36.643310546875,36.6783218383789,36.7436981201172,36.7655754089355,36.7946281433105,36.9015617370605,36.8933601379395,36.8792495727539,36.8622550964355,36.7891082763672,36.715087890625,36.693115234375,36.6946792602539,36.6805686950684,36.6815910339355,36.7022476196289,36.7386207580566,36.8260765075684,37.0088882446289,37.0977058410645,37.108154296875,37.1091804504395,37.0858879089355,37.0693359375,37.07080078125,37.0891609191895,37.1261215209961,37.1563949584961,37.2060546875,37.2886238098145,37.2972640991211,37.2822265625,37.2765617370605,37.2295875549316,37.1085968017578],[21.8513660430908,21.7962894439697,21.8044929504395,21.792724609375,21.780029296875,21.769775390625,21.7552242279053,21.7580089569092,21.7953128814697,21.8513660430908],[21.7652339935303,21.751708984375,21.7902355194092,21.7906265258789,21.8434581756592,21.8520984649658,21.8334484100342,21.7875480651855,21.7652339935303],[21.8404293060303,21.8298816680908,21.8644027709961,21.8920421600342,21.8934078216553,21.9182605743408,21.950439453125,21.9519062042236,21.8624992370605,21.8404293060303],[19.4966316223145,19.0505867004395,18.6045398712158,18.1584949493408,17.71240234375,17.266357421875,16.8202629089355,16.3742198944092,15.9281244277954,15.7801752090454,15.7215328216553,15.7134275436401,15.7035150527954,15.7449712753296,15.745997428894,15.7139654159546,15.6972179412842,15.702540397644,15.6258296966553,15.5331048965454,15.3113288879395,15.2381353378296,15.162109375,15.0966300964355,15.0444345474243,14.9986820220947,14.8983898162842,14.8514642715454,14.7886228561401,14.7224607467651,14.6880865097046,14.6627445220947,14.6333494186401,14.585205078125,14.5504884719849,14.5041990280151,14.4412107467651,14.3421392440796,14.2842283248901,14.237060546875,14.2032232284546,14.161865234375,14.12744140625,14.0555171966553,14.0288572311401,13.9923343658447,13.9787111282349,13.9105958938599,13.8501462936401,13.7998046875,13.7303218841553,13.6264162063599,13.5380859375,13.4716300964355,13.3987789154053,13.32958984375,13.2693357467651,13.2150392532349,13.1130857467651,13.0009765625,12.8647470474243,12.79052734375,12.7412109375,12.6993656158447,12.6781244277954,12.671875,12.694580078125,12.70947265625,12.6604490280151,12.54638671875,12.4629878997803,12.3119144439697,12.1292476654053,12.0677738189697,12.0447273254395,12.032958984375,11.9901371002197,11.6695308685303,11.5798826217651,11.5159177780151,11.482666015625,11.4398441314697,11.4099607467651,11.4032716751099,11.3448734283447,11.2671880722046,11.1920413970947,11.0290031433105,10.919677734375,10.9271974563599,10.9540529251099,10.9773435592651,10.9962396621704,10.9515132904053,10.8941402435303,10.8513669967651,10.8260746002197,10.830078125,10.8227052688599,10.7820320129395,10.7366695404053,10.642822265625,10.6086912155151,10.5748043060303,10.5378904342651,10.4616203308105,10.3665533065796,10.2898435592651,10.23828125,10.2185544967651,10.2078123092651,10.1756830215454,10.0013675689697,9.96914100646973,9.974609375,9.71323204040527,9.63627910614014,9.52714824676514,9.40566444396973,9.34711933135986,9.32451152801514,9.30136680603027,9.27495098114014,9.12709999084473,9.13320350646973,9.07514667510986,9.04936504364014,9.02089786529541,9.02358436584473,9.01162052154541,9.01596641540527,8.99501895904541,8.93886661529541,8.88974666595459,8.87319374084473,8.85249042510986,8.83603572845459,8.71543025970459,8.65615272521973,8.59882831573486,8.59028339385986,8.5869140625,8.54121017456055,8.40507793426514,8.24379825592041,8.19770526885986,8.167724609375,8.060791015625,8.0458984375,8.03203105926514,8.02036190032959,7.98544883728027,7.97382736206055,7.98359394073486,7.90981483459473,7.89091825485229,7.88457059860229,7.81298828125,7.70190382003784,7.68081045150757,7.63369178771973,7.55732440948486,7.55097627639771,7.65175771713257,7.74336004257202,7.83588838577271,7.865478515625,7.85996150970459,7.81899404525757,7.77236318588257,7.68354511260986,7.62343692779541,7.57211923599243,7.50756883621216,7.47529268264771,7.48842811584473,7.51503944396973,7.52377939224243,7.60439443588257,7.66450166702271,7.73803758621216,7.78788995742798,7.812744140625,7.85185575485229,8.08383750915527,8.32236385345459,8.55732345581055,8.707275390625,8.79863262176514,8.810302734375,8.83915996551514,8.86567306518555,9.02524471282959,9.20351600646973,9.28505802154541,9.406494140625,9.53173828125,9.58872032165527,9.69155216217041,9.78437519073486,9.90180587768555,9.979736328125,9.98505878448486,9.95307636260986,9.94169902801514,9.96596622467041,9.98286151885986,9.98149394989014,9.95429706573486,9.96030235290527,10.0078125,10.0884761810303,10.2168951034546,10.3573732376099,10.4845209121704,10.6484870910645,10.8510732650757,11.1136722564697,11.2625007629395,11.3685541152954,11.5412588119507,11.642578125,11.724365234375,11.8455076217651,11.9071283340454,12.1083498001099,12.13037109375,12.2693843841553,12.5020990371704,12.6556148529053,12.7299318313599,12.8202142715454,12.979736328125,13.0217771530151,13.0773439407349,13.0785150527954,13.2584962844849,13.4895505905151,13.70458984375,14.1344232559204,14.3806648254395,14.4555177688599,14.6307621002197,14.9661121368408,15.4847640991211,15.7501468658447,16.1466312408447,16.6338882446289,16.9083976745605,17.4084949493408,17.937255859375,18.3370609283447,18.8108386993408,19.206787109375,19.4952144622803,19.904052734375,19.9825687408447,20.3031749725342,20.34619140625,20.3998546600342,20.67236328125,20.7333011627197,20.8749027252197,20.9543952941895,21.4115238189697,21.4674320220947,21.5233898162842,21.6058101654053,21.9220714569092,22.4183597564697,22.9961910247803,23.1606941223145,23.2857418060303,23.4452152252197,23.2818355560303,23.0452632904053,22.8086910247803,22.5721187591553,22.3355464935303,22.0989742279053,21.8624019622803,21.6258296966553,21.3892574310303,21.1526374816895,20.9161148071289,20.6794929504395,20.4429206848145,20.2063484191895,19.9697761535645,19.7332038879395,19.4966316223145],[10.9932613372803,10.8857421875,10.7525873184204,10.7102537155151,10.3866701126099,10.3595695495605,10.3515625,10.2420415878296,10.098388671875,9.99697303771973,9.96294021606445,9.75019454956055,9.56752967834473,9.46298789978027,9.36166954040527,9.28500938415527,9.13725662231445,9.050048828125,8.77099609375,8.55927753448486,8.27099609375,8.03022480010986,7.72587871551514,7.36918926239014,6.997314453125,6.99243116378784,6.87700223922729,6.77226543426514,6.73808574676514,6.68740224838257,6.61020517349243,6.58154249191284,6.42626905441284,6.29462909698486,6.25083017349243,6.216796875,6.14687538146973,6.08940410614014,6.14501905441284,6.15502977371216,6.17377948760986,6.20263671875,6.2685546875,6.32031202316284,6.32856464385986,6.38637733459473,6.45258760452271,6.51875019073486,6.54931688308716,6.58076190948486,6.592529296875,6.7421875,6.802490234375,6.85092830657959,6.88832998275757,6.93886709213257,6.97968769073486,7.00410127639771,7.03398466110229,7.09663057327271,7.22656202316284,7.35366249084473,7.38881826400757,7.39873075485229,7.43510723114014,7.49511671066284,7.546875,7.72822237014771,8.14580059051514,8.20956993103027,8.25346660614014,8.30424785614014,8.35488319396973,8.47963809967041,8.57529354095459,8.65273380279541,8.72202110290527,8.75927734375,8.81377029418945,8.85146522521973,8.89492225646973,8.97421932220459,9.11533260345459,9.22124004364014,9.35830116271973,9.39848613739014,9.48027324676514,9.49145412445068,9.48554611206055,9.43183517456055,9.426025390625,9.44189453125,9.46352577209473,9.49560546875,9.53564453125,9.57060527801514,9.58657264709473,9.60415077209473,9.62094688415527,9.64472675323486,9.66791915893555,9.67231464385986,9.67099571228027,9.68759822845459,9.803955078125,9.84458065032959,9.92490196228027,10.2364749908447,10.2685546875,10.2918453216553,10.3069343566895,10.3905267715454,10.4547853469849,10.5206050872803,10.5638666152954,10.5909690856934,10.630615234375,10.6730461120605,10.7155265808105,10.8005857467651,10.891357421875,10.9685430526733,11.02099609375,11.0440034866333,11.0555658340454,11.1156244277954,11.1017780303955,11.0696287155151,10.9919929504395,10.9781742095947,10.9549808502197,10.9554204940796,10.9830570220947,10.9932613372803],[6.52192354202271,6.51611328125,6.59682655334473,6.714111328125,6.570556640625,6.52192354202271],[7.59184598922729,7.47128868103027,7.49589872360229,7.54848623275757,7.62573289871216,7.63652372360229,7.59184598922729],[7.93393564224243,7.91704034805298,8.10854530334473,8.05732440948486,7.93393564224243],[7.90205097198486,7.82841777801514,7.82944345474243,7.78232431411743,7.77607488632202,7.92607402801514,8.13623046875,8.16630840301514,8.11064434051514,8.08564472198486,7.96455049514771,7.90205097198486],[9.05146503448486,9.04082012176514,9.09541034698486,9.1298828125,9.13911056518555,9.08037090301514,9.05146503448486],[9.58603572845459,9.52944278717041,9.46142578125,9.421630859375,9.47607421875,9.55996036529541,9.58100605010986,9.57685565948486,9.58603572845459],[9.69667911529541,9.67998027801514,9.71171855926514,9.74760723114014,9.79355430603027,9.79165077209473,9.74912071228027,9.69667911529541],[11.676513671875,11.5721673965454,11.6149415969849,11.6677732467651,11.6916990280151,11.676513671875],[11.9887208938599,11.9647464752197,11.9829587936401,11.9744138717651,11.9808101654053,12.1193361282349,12.15185546875,12.1416511535645,12.0728521347046,12.0250978469849,11.9887208938599],[20.316650390625,20.2576675415039,20.2454090118408,20.2727546691895,20.339599609375,20.3772964477539,20.3858871459961,20.3403797149658,20.24072265625,20.18408203125,20.1779289245605,20.1323738098145,20.0886726379395,19.996337890625,19.888916015625,19.756103515625,19.6444835662842,19.553466796875,19.4998531341553,19.5147457122803,19.5419425964355,19.5850582122803,19.6053714752197,19.6107921600342,19.5791988372803,19.54833984375,19.4866218566895,19.3279285430908,19.2115249633789,19.0889167785645,18.9771480560303,18.7927742004395,18.6183109283447,18.5335445404053,18.47900390625,18.4419918060303,18.4070320129395,18.3545398712158,18.286865234375,18.22216796875,18.1426277160645,18.0335445404053,17.7971687316895,17.5838871002197,17.5411128997803,17.5099620819092,17.4795398712158,17.4990234375,17.625,17.71875,17.8123531341553,17.8205089569092,17.889404296875,17.9526863098145,18.0333976745605,18.0646476745605,18.0464363098145,18.0814952850342,18.1698246002197,18.2106456756592,18.203857421875,18.1489734649658,18.0459480285645,17.984619140625,17.965087890625,17.9267578125,17.86962890625,17.8333492279053,17.8179683685303,17.8241214752197,17.8517589569092,17.8922348022461,17.945556640625,17.9769039154053,17.9862308502197,18.0285167694092,18.1382331848145,18.2217292785645,18.2594718933105,18.2784671783447,18.3049812316895,18.3389663696289,18.3734855651855,18.4083976745605,18.42333984375,18.4181652069092,18.382568359375,18.3165054321289,18.2953128814697,18.3189945220947,18.2166976928711,17.9883785247803,17.8158187866211,17.698974609375,17.6092777252197,17.5467281341553,17.4616680145264,17.30029296875,17.0771484375,16.8843746185303,16.6475582122803,16.466064453125,16.3399410247803,16.2379875183105,16.1602535247803,16.0935554504395,16.0378894805908,15.9874515533447,15.9421873092651,15.8896970748901,15.8298835754395,15.7804193496704,15.7412605285645,15.699951171875,15.6565418243408,15.5859384536743,15.4882802963257,15.4132318496704,15.3608884811401,15.3196296691895,15.256591796875,15.1275882720947,15.0416011810303,14.9324703216553,14.8433103561401,14.6612300872803,14.5906744003296,14.5301275253296,14.4716310501099,14.4166984558105,14.3678712844849,14.3462400436401,14.3360834121704,14.2809572219849,14.2274417877197,14.2273921966553,14.2544431686401,14.2894535064697,14.3661127090454,14.4040031433105,14.4278316497803,14.3900384902954,14.3695793151855,14.3955078125,14.3627433776855,14.35791015625,14.36279296875,14.3621578216553,14.3741693496704,14.4210939407349,14.4174318313599,14.3786134719849,14.351318359375,14.3326177597046,14.2525386810303,14.13671875,14.0548820495605,13.9724607467651,13.8418941497803,13.7169437408447,13.6599607467651,13.6263675689697,13.585693359375,13.5675783157349,13.560302734375,13.5399904251099,13.2882328033447,13.1929693222046,13.0779781341553,13.0150394439697,12.8283205032349,12.6699705123901,12.5699214935303,12.4935054779053,12.42626953125,12.3833980560303,12.2556638717651,12.0897951126099,11.7320804595947,11.7066898345947,11.703857421875,11.7727537155151,11.8886232376099,12.012451171875,12.1488285064697,12.2030277252197,12.1578130722046,12.1092290878296,12.1792488098145,12.2525873184204,12.3943357467651,12.3614253997803,12.3316402435303,12.4430179595947,12.5318851470947,12.5636720657349,12.59326171875,12.640380859375,12.6893548965454,12.6189451217651,12.6736326217651,12.6212406158447,12.65380859375,12.7145023345947,12.8181638717651,13.034912109375,13.187255859375,13.3030271530151,13.3575687408447,13.4319829940796,13.46240234375,13.5212888717651,13.5681638717651,13.5144529342651,13.4844732284546,13.4395503997803,13.3531732559204,13.2434568405151,13.1712398529053,13.045654296875,12.7714834213257,12.6900386810303,12.354736328125,12.1708011627197,12.0474605560303,11.9366216659546,11.748779296875,11.6617679595947,11.462890625,11.2151851654053,11.1005859375,10.8895511627197,10.5691404342651,10.3881349563599,10.31982421875,10.265869140625,10.1754388809204,9.93417930603027,9.73403263092041,9.62714862823486,9.41459941864014,9.35297870635986,9.26523399353027,9.22543907165527,9.21372032165527,9.314208984375,9.28837871551514,9.19462871551514,9.11289024353027,8.67123985290527,8.58920860290527,8.51113319396973,8.42807579040527,8.44296836853027,8.47377967834473,8.50839805603027,8.42470741271973,8.26850605010986,7.44228506088257,7.33730506896973,7.22690343856812,7.28076124191284,7.46430635452271,7.54150390625,7.55288124084473,7.551513671875,7.58432579040527,7.64418888092041,7.71596717834473,7.77490186691284,7.72812509536743,7.59926748275757,7.50053739547729,7.28012704849243,7.18784189224243,7.161376953125,7.17597675323486,7.08198261260986,6.99467802047729,6.86093759536743,6.87514638900757,6.90830039978027,6.89956092834473,6.86528348922729,6.75395488739014,6.47460889816284,6.24223661422729,6.18466806411743,6.09667921066284,5.97934579849243,5.911376953125,5.82529306411743,5.77934598922729,5.77060556411743,5.77880859375,5.79599666595459,5.87714862823486,5.90200233459473,5.90776395797729,5.85166025161743,5.78935480117798,5.73369121551514,5.66875028610229,5.64306640625,5.63676738739014,5.67490243911743,5.72451162338257,5.77104520797729,5.84619140625,5.95649385452271,6.03369140625,6.16606473922729,6.24257802963257,6.24531269073486,6.25766611099243,6.24540996551514,6.33164072036743,6.42617177963257,6.46005868911743,6.48066425323486,6.447998046875,6.467529296875,6.54990196228027,6.68271493911743,6.68662071228027,6.67182588577271,6.64160108566284,6.48867225646973,6.44199180603027,6.74990224838257,6.87665987014771,7.106201171875,7.15087890625,7.15532207489014,7.21879863739014,7.35561513900757,7.32949209213257,7.33437490463257,7.37221670150757,7.56132745742798,7.61904335021973,7.71806573867798,7.71806573867798,7.76562452316284,7.88784122467041,7.96279239654541,8.02392482757568,8.059814453125,8.25673866271973,8.30502891540527,8.34428691864014,8.31782245635986,8.24692344665527,8.17822265625,8.18696308135986,8.22622108459473,8.42309474945068,8.54365253448486,8.76787090301514,8.96894550323486,9.29052734375,9.49282264709473,9.56142520904541,9.83750057220459,10.1903810501099,10.2660150527954,10.3508310317993,10.4308595657349,10.5570306777954,10.6235837936401,10.6609373092651,10.7084474563599,10.788330078125,10.9199705123901,11.1052742004395,11.3894529342651,11.5543947219849,11.6125001907349,11.6306638717651,11.6871576309204,11.7496585845947,11.7812013626099,12.0896482467651,12.1902351379395,12.3090333938599,12.3948240280151,12.4736328125,12.5479011535645,12.59423828125,12.6528806686401,12.73974609375,12.8819341659546,12.9613285064697,13.03076171875,13.103515625,13.1729984283447,13.2330570220947,13.4969244003296,13.5757808685303,13.7166996002197,13.82275390625,13.9471673965454,14.0498533248901,14.2357425689697,14.3599128723145,14.4728994369507,14.6029787063599,14.6964836120605,14.8147468566895,14.9759273529053,15.1474113464355,15.2041015625,15.2413568496704,15.2715826034546,15.278564453125,15.3573732376099,15.3506832122803,15.3676767349243,15.4035634994507,15.5597648620605,15.7686033248901,15.9386234283447,16.0506839752197,16.1808109283447,16.237060546875,16.298095703125,16.3519058227539,16.3941879272461,16.4175777435303,16.305419921875,16.3304214477539,16.5147953033447,16.5709476470947,16.63818359375,16.7322273254395,16.89501953125,16.9756832122803,17.1476554870605,17.2398929595947,17.373291015625,17.5333003997803,17.6812515258789,17.7758312225342,17.797119140625,17.8335456848145,17.935302734375,18.0374031066895,18.1737289428711,18.2580070495605,18.2903327941895,18.302978515625,18.2958984375,18.3596687316895,18.4942874908447,18.5179691314697,18.5175285339355,18.4942378997803,18.4977531433105,18.5287094116211,18.5612297058105,18.572021484375,18.5881843566895,18.6208000183105,18.9317874908447,18.9964847564697,19.1304683685303,19.265869140625,19.4599609375,19.5928707122803,19.6537113189697,19.74951171875,19.7697257995605,19.7621593475342,19.690673828125,19.687255859375,19.6891593933105,19.6944332122803,19.7013187408447,19.7710933685303,19.7784671783447,19.7695789337158,19.7729015350342,19.8049335479736,19.8613777160645,20.0417976379395,20.0736331939697,20.099365234375,20.1166019439697,20.1151371002197,20.0804195404053,20.0789051055908,20.0934562683105,20.1183109283447,20.1498546600342,20.187744140625,20.2606449127197,20.35205078125,20.363037109375,20.3428211212158,20.3204574584961,20.325439453125,20.34130859375,20.3844718933105,20.4244136810303,20.4154300689697,20.3795890808105,20.316650390625],[39.8252944946289,39.7867660522461,39.7909164428711,39.8281745910645,39.863037109375,39.8827133178711,39.8824195861816,39.8666000366211,39.85546875,39.8458480834961,39.8252944946289],[40.9366188049316,40.9344215393066,40.9818344116211,41.014892578125,41.0248031616211,41.0016593933105,40.9608383178711,40.9366188049316],[40.2388687133789,40.2206535339355,40.1875953674316,40.1311531066895,40.0834465026855,39.9989738464355,39.9745101928711,39.9544944763184,39.9499015808105,40.0492172241211,40.0698699951172,40.1048355102539,40.1727561950684,40.2022476196289,40.1580085754395,40.0973167419434,40.0603485107422,40.020751953125,39.9906234741211,39.950439453125,39.9197273254395,39.9097633361816,39.9470710754395,39.9685554504395,39.9177742004395,39.8270988464355,39.7610855102539,39.6658706665039,39.5248069763184,39.532470703125,39.5320777893066,39.5737762451172,39.5749015808105,39.5567359924316,39.5530738830566,39.5605964660645,39.5575714111328,39.5841789245605,39.5750007629395,39.5426292419434,39.5818862915039,39.5873527526855,39.5758781433105,39.5644035339355,39.4712867736816,39.4132804870605,39.3947296142578,39.411865234375,39.493408203125,39.5135726928711,39.5198211669922,39.5353050231934,39.5686988830566,39.5978507995605,39.6036643981934,39.5821800231934,39.5538558959961,39.51708984375,39.4788093566895,39.4530754089355,39.4470710754395,39.4229965209961,39.3777351379395,39.3065910339355,39.2779808044434,39.2755851745605,39.3509254455566,39.3521499633789,39.3106460571289,39.2607421875,39.20751953125,39.2156753540039,39.2737312316895,39.3368644714355,39.3573722839355,39.377197265625,39.385986328125,39.3604011535645,39.3570785522461,39.3619155883789,39.3745613098145,39.412353515625,39.4427223205566,39.4605941772461,39.4576187133789,39.4488792419434,39.3966789245605,39.2978515625,39.2291984558105,39.1045417785645,39.0445289611816,39.0021476745605,38.9686546325684,38.9413108825684,38.9146957397461,38.88623046875,38.8542976379395,38.8172340393066,38.6989250183105,38.6068878173828,38.5628890991211,38.53369140625,38.5398445129395,38.6084938049316,38.6611824035645,38.6575164794922,38.6597671508789,38.6000022888184,38.510009765625,38.4603042602539,38.404296875,38.2747535705566,38.19189453125,38.1036148071289,38.0380859375,37.92578125,37.8327140808105,37.8049812316895,37.7725105285645,37.6873054504395,37.6014175415039,37.5728034973145,37.5303688049316,37.4512710571289,37.3856925964355,37.3440437316895,37.2935523986816,37.25,37.2316398620605,37.2419929504395,37.2859382629395,37.3570289611816,37.3944816589355,37.3823738098145,37.3956069946289,37.4187507629395,37.4154319763184,37.3724594116211,37.3294410705566,37.3162117004395,37.2831535339355,37.2317886352539,37.2393569946289,37.2615699768066,37.2912101745605,37.3294410705566,37.3757820129395,37.4187507629395,37.4304695129395,37.4372062683105,37.446044921875,37.4716796875,37.4622573852539,37.4084968566895,37.2675285339355,37.1727066040039,37.029052734375,36.98291015625,36.9005393981934,36.7664527893066,36.6942863464355,36.6840324401855,36.6969223022461,36.73291015625,36.8451156616211,37.0150871276855,37.1275405883789,37.2718276977539,37.43603515625,37.6029319763184,37.795654296875,37.8642578125,37.9101066589355,37.9331512451172,37.931884765625,37.9062995910645,37.90185546875,37.9184112548828,38.0079078674316,38.1702613830566,38.3069839477539,38.4178695678711,38.4563980102539,38.4225616455078,38.3344230651855,38.1919937133789,38.075439453125,37.9848136901855,37.9412078857422,37.9244117736816,37.8860359191895,37.765380859375,37.6641578674316,37.5824699401855,37.5435028076172,37.5472183227539,37.566162109375,37.6002922058105,37.6095695495605,37.5940437316895,37.5530776977539,37.4867172241211,37.3993148803711,37.2908706665039,37.207763671875,37.1500473022461,37.116943359375,37.1083984375,37.1583023071289,37.2665023803711,37.3250465393066,37.3339385986328,37.3280754089355,37.3168449401855,37.3028335571289,37.2707023620605,37.2580108642578,37.2680206298828,37.2583999633789,37.2244644165039,37.1834487915039,37.1375007629395,37.0884284973145,37.0363273620605,37.0130844116211,37.0215339660645,36.9498023986816,36.9720230102539,37.064208984375,37.14013671875,37.1722145080566,37.2449684143066,37.4870109558105,37.5707015991211,37.720947265625,37.83544921875,37.9284172058105,37.9596672058105,38.0329093933105,38.1167945861816,38.1695327758789,38.2110366821289,38.23779296875,38.2945327758789,38.3831062316895,38.4735336303711,38.5889167785645,38.6692886352539,38.890625,38.9276351928711,38.9620132446289,38.9835968017578,38.992919921875,38.9830093383789,38.9822273254395,38.99462890625,39.0084953308105,39.1091766357422,39.1310539245605,39.1502914428711,39.1966781616211,39.2166976928711,39.2420921325684,39.465576171875,39.482421875,39.5187492370605,39.5576171875,39.6213874816895,39.5937957763672,39.5641593933105,39.5482902526855,39.5376930236816,39.5288543701172,39.5367164611816,39.5627899169922,39.6349639892578,39.7432632446289,39.8388671875,39.8462905883789,39.8362312316895,39.8555679321289,39.8818359375,39.9041976928711,39.9091300964355,39.8843269348145,39.8909645080566,39.907470703125,39.9273414611816,39.9607887268066,40.0133323669434,40.031494140625,40.0503425598145,40.0682106018066,40.0713386535645,40.0899429321289,40.1195831298828,40.1363296508789,40.1270980834961,40.1291999816895,40.1472663879395,40.1670875549316,40.1826667785645,40.2226066589355,40.208740234375,40.1875953674316,40.1980934143066,40.2881355285645,40.2965812683105,40.327392578125,40.5665512084961,40.587646484375,40.634765625,40.7239265441895,40.7673835754395,40.7971687316895,40.76708984375,40.6790542602539,40.6619644165039,40.656982421875,40.6842765808105,40.7714347839355,40.8204116821289,40.8917007446289,40.9192390441895,41.0276374816895,41.0351066589355,41.0234375,40.9114723205566,40.8396492004395,40.8150863647461,40.7965850830078,40.778564453125,40.7395973205566,40.7217750549316,40.6877937316895,40.6690902709961,40.6611824035645,40.5627937316895,40.4535179138184,40.4392585754395,40.4120140075684,40.3841323852539,40.3613777160645,40.3453598022461,40.3245124816895,40.2671394348145,40.2141609191895,40.201171875,40.2345695495605,40.2388687133789],[38.8030776977539,38.7561531066895,38.897216796875,39.052734375,39.0965843200684,39.0940895080566,39.0379409790039,38.8030776977539],[37.3484840393066,37.3447265625,37.3681640625,37.4147453308105,37.5081062316895,37.5691413879395,37.5608406066895,37.5191421508789,37.4678230285645,37.368408203125,37.2511711120605,37.2467765808105,37.2379379272461,37.1935577392578,37.132080078125,37.0592765808105,36.9647941589355,36.7501945495605,36.5545425415039,36.4275894165039,36.3406753540039,36.22607421875,36.14892578125,36.1126976013184,36.067626953125,36.0250968933105,36.0121116638184,36.0197257995605,36.0123519897461,35.9678192138672,35.9131355285645,35.8583984375,35.8584480285645,35.84619140625,35.8186988830566,35.7667465209961,35.728271484375,35.6781234741211,35.6375503540039,35.5680656433105,35.44580078125,35.4091796875,35.3496589660645,35.2713394165039,35.2553215026855,35.233154296875,35.2398948669434,35.2513694763184,35.2312507629395,35.1707992553711,35.1891098022461,35.250244140625,35.2899398803711,35.3796882629395,35.4437026977539,35.4478988647461,35.4314918518066,35.41943359375,35.4323234558105,35.4578628540039,35.5457992553711,35.5931167602539,35.6294898986816,35.6195793151855,35.6592788696289,35.70556640625,35.7618141174316,35.8676300048828,35.9436988830566,35.9767570495605,35.9999008178711,36.0528335571289,36.09912109375,36.1905250549316,36.2896957397461,36.4327125549316,36.572265625,36.642578125,36.6429672241211,36.6376457214355,36.653564453125,36.8294410705566,36.962890625,37.0416030883789,37.1384773254395,37.1789054870605,37.2528305053711,37.333740234375,37.4447250366211,37.4811515808105,37.5106430053711,37.5237312316895,37.520751953125,37.649658203125,37.6834945678711,37.65625,37.6515617370605,37.6881828308105,37.6385269165039,37.6353530883789,37.647216796875,37.6658210754395,37.7830543518066,37.8304672241211,37.86083984375,37.9052734375,37.928466796875,37.9477043151855,37.9733390808105,37.9899406433105,38.0329093933105,38.0893096923828,38.13037109375,38.1795883178711,38.2164077758789,38.2099609375,38.2130355834961,38.2500495910645,38.2566413879395,38.2496109008789,38.2494163513184,38.2225112915039,38.1911125183105,38.0948219299316,38.0804176330566,38.0733909606934,38.078369140625,38.0775413513184,38.0946273803711,38.0997543334961,38.0511245727539,37.9813461303711,37.9024887084961,37.7779312133789,37.72265625,37.5019035339355,37.4701652526855,37.4447250366211,37.4402351379395,37.4075698852539,37.3536148071289,37.3324737548828,37.3435554504395,37.41357421875,37.6695785522461,37.9279327392578,38.0469245910645,38.28564453125,38.4059066772461,38.5149421691895,38.6217765808105,38.741943359375,38.8640632629395,38.9492683410645,39.0180168151855,39.1030769348145,39.1534156799316,39.2095718383789,39.2159690856934,39.2740745544434,39.3057136535645,39.3408203125,39.3426284790039,39.3167953491211,39.2649879455566,39.3466796875,39.4320793151855,39.6085433959961,39.5570831298828,39.5364265441895,39.5333023071289,39.5469741821289,39.607421875,39.6417503356934,39.6688003540039,39.74853515625,39.8311996459961,39.9093742370605,39.9408683776855,39.9603538513184,39.9580078125,39.9786605834961,39.9875984191895,39.8954582214355,39.7744140625,39.8338851928711,39.9125022888184,40.0540046691895,40.2197723388672,40.3987312316895,40.5467300415039,40.6856422424316,40.8634796142578,41.0380859375,40.9598617553711,40.8897476196289,40.824951171875,40.8094711303711,40.78271484375,40.7927703857422,40.8310546875,40.8185043334961,40.7464370727539,40.6656723022461,40.648681640625,40.7070808410645,40.7204093933105,40.6936988830566,40.6887702941895,40.6932640075684,40.7649421691895,40.8345680236816,40.8713874816895,40.832275390625,40.8583488464355,40.873046875,40.89111328125,40.9512710571289,41.0129890441895,41.0711441040039,41.1221694946289,41.1935539245605,41.3637199401855,41.4315910339355,41.5193862915039,41.6433601379395,41.7725601196289,41.8684577941895,42.0911598205566,42.1176261901855,42.12939453125,42.1363754272461,42.1201629638672,42.0818367004395,42.0937995910645,42.070068359375,41.9762229919434,41.895751953125,41.7118148803711,41.6525382995605,41.6136703491211,41.3418960571289,41.2100601196289,41.2002906799316,41.2308616638184,41.3654289245605,41.5294418334961,41.7803688049316,41.9443855285645,42.0605964660645,42.1307144165039,42.1477546691895,42.2058601379395,42.2582511901855,42.296875,42.3297882080078,42.3358879089355,42.335205078125,42.3041954040527,42.2799797058105,42.18017578125,42.0782241821289,41.9651832580566,41.9190902709961,41.864013671875,41.8100090026855,41.6387214660645,41.5602531433105,41.4581031799316,41.4083976745605,41.346923828125,41.2962913513184,41.2722663879395,41.2627449035645,41.27880859375,41.310791015625,41.3241195678711,41.3222160339355,41.3108367919922,41.3006324768066,41.2879867553711,41.276123046875,41.26513671875,41.2634773254395,41.3072776794434,41.331298828125,41.3502922058105,41.3717765808105,41.3899917602539,41.4505844116211,41.6693344116211,41.8565444946289,41.9148406982422,41.9571266174316,42.08447265625,42.123779296875,42.1562995910645,42.164794921875,42.1898422241211,42.2310562133789,42.335205078125,42.4200210571289,42.4587898254395,42.4876441955566,42.4865226745605,42.4615707397461,42.4477081298828,42.4288597106934,42.39892578125,42.3467750549316,42.3124504089355,42.29248046875,42.2920913696289,42.2917976379395,42.2993659973145,42.3168449401855,42.3401374816895,42.4066925048828,42.5272941589355,42.5763664245605,42.6029777526855,42.6280746459961,42.6663093566895,42.6880836486816,42.6717300415039,42.662841796875,42.6819343566895,42.7784652709961,42.6761741638184,42.5614738464355,42.540283203125,42.5281257629395,42.5237770080566,42.5114250183105,42.481689453125,42.3561515808105,42.3232917785645,42.2995109558105,42.3015594482422,42.295166015625,42.2360343933105,42.2117195129395,42.1908187866211,42.1647453308105,42.1317367553711,42.0680656433105,42.018798828125,41.9954109191895,41.9735336303711,41.9543952941895,41.9074211120605,41.8570327758789,41.8344230651855,41.8031234741211,41.7822761535645,41.7926788330078,41.7596664428711,41.7005386352539,41.6449699401855,41.5941429138184,41.5452156066895,41.4762229919434,41.4273452758789,41.3994140625,41.3489761352539,41.2216300964355,41.2161636352539,41.2457504272461,41.2486801147461,41.22900390625,41.2108879089355,41.1905746459961,41.189208984375,41.1951179504395,41.2521476745605,41.26513671875,41.2746086120605,41.2760734558105,41.2398452758789,41.1634292602539,41.0937004089355,41.0306167602539,40.893798828125,40.6833000183105,40.5412101745605,40.4674797058105,40.3320808410645,40.0362281799316,39.9756355285645,39.944091796875,39.8584938049316,39.716796875,39.6331558227539,39.49951171875,39.3770980834961,39.1881370544434,39.1605491638184,39.058349609375,38.95361328125,38.977294921875,38.8162117004395,38.7564468383789,38.7360343933105,38.6724586486816,38.5394515991211,38.3488273620605,38.2385749816895,38.2257308959961,38.2263679504395,38.2500495910645,38.2687492370605,38.2442359924316,38.2001457214355,38.1667022705078,38.1180648803711,38.0721664428711,38.0509300231934,38.0107917785645,37.9867172241211,37.9598617553711,37.9320335388184,37.7857437133789,37.5991706848145,37.4586906433105,37.3484840393066],[-9.34160137176514,-9.25615215301514,-9.19033241271973,-9.23857402801514,-9.31435585021973,-9.34990215301514,-9.41376972198486,-9.42792987823486,-9.41386699676514,-9.42314529418945,-9.41640567779541,-9.37539100646973,-9.34160137176514],[-9.51191425323486,-9.3818359375,-9.32597732543945,-9.29423809051514,-9.25468730926514,-9.21376895904541,-9.19492149353027,-9.18984317779541,-9.12294864654541,-9.04257774353027,-9.01542949676514,-9.00400352478027,-9.06425762176514,-9.05341720581055,-9.03154277801514,-8.94248104095459,-8.85908222198486,-8.7080078125,-8.64785099029541,-8.59130859375,-8.57539081573486,-8.4921875,-8.48652362823486,-8.48896503448486,-8.47080039978027,-8.45947265625,-8.42275333404541,-8.37734413146973,-8.34160137176514,-8.31572246551514,-8.34824180603027,-8.37294864654541,-8.39453125,-8.42451190948486,-8.58359336853027,-8.71523475646973,-8.75507831573486,-8.78203105926514,-8.83290958404541,-8.91269493103027,-8.95761680603027,-8.97275352478027,-8.99668025970459,-9.04355430603027,-9.12392616271973,-9.13212966918945,-9.13017559051514,-9.16093730926514,-9.27578067779541,-9.40351581573486,-9.51191425323486],[-8.13994121551514,-8.31181621551514,-8.27509784698486,-8.17861366271973,-8.14999961853027,-8.13994121551514],[-21.3004875183105,-21.4505863189697,-21.3817386627197,-21.3498058319092,-21.3034172058105,-21.3004875183105],[-21.1693363189697,-21.16943359375,-21.1397457122803,-21.1290035247803,-21.1607418060303,-21.263671875,-21.2234363555908,-21.15771484375,-21.1068344116211,-21.0682621002197,-21.0993175506592,-21.1133785247803,-21.1187496185303,-21.146484375,-21.1556644439697,-21.1693363189697],[-18.6393566131592,-18.6986331939697,-18.69775390625,-18.6633796691895,-18.6402339935303,-18.5707015991211,-18.5653324127197,-18.5885734558105,-18.6084976196289,-18.6393566131592],[10.1343269348145,10.0780277252197,10.0646486282349,10.0850582122803,10.0691404342651,10.1916990280151,10.243408203125,10.2531251907349,10.2685546875,10.5389652252197,10.6033687591553,10.6388673782349,10.6644535064697,10.6993646621704,10.7180671691895,10.7479486465454,10.7644529342651,10.7968263626099,10.8033208847046,10.8319339752197,10.8402338027954,10.7161617279053,10.6698732376099,10.55810546875,10.4822750091553,10.3863773345947,10.3233890533447,10.261474609375,10.1678228378296,10.1343269348145],[11.1785154342651,11.1686029434204,11.2083988189697,11.2772455215454,11.3235349655151,11.3253908157349,11.2637205123901,11.1785154342651],[33.7220726013184,33.7174301147461,33.690185546875,33.6871604919434,33.7176780700684,33.7174797058105,33.7389144897461,33.8556137084961,33.888671875,33.8931159973145,33.8233375549316,33.8050270080566,33.7850608825684,33.7459449768066,33.7220726013184],[34.7538108825684,34.6816902160645,34.7445831298828,34.8203125,34.8021965026855,34.7538108825684],[33.1819343566895,33.1555671691895,32.9657249450684,32.8973655700684,32.7816925048828,32.642578125,32.5249519348145,32.4733390808105,32.4136734008789,32.34521484375,32.2567405700684,32.1727027893066,32.0806655883789,32.0211906433105,31.9753913879395,31.9295387268066,31.8857402801514,31.8025398254395,31.7360363006592,31.704833984375,31.6849613189697,31.5851058959961,31.5458030700684,31.4637680053711,31.2509765625,31.0321273803711,30.9408226013184,30.8649425506592,30.7832012176514,30.6659679412842,30.580078125,30.4253406524658,30.3873023986816,30.3422374725342,30.2823238372803,30.2293968200684,30.4653797149658,30.6667976379395,30.8329124450684,31.1253433227539,31.3736820220947,31.621337890625,31.8461437225342,32.0723609924316,32.1053733825684,32.2121086120605,32.3104476928711,32.4223136901855,32.5436058044434,32.6962890625,32.9267082214355,33.0553207397461,33.0890617370605,33.172119140625,33.2331047058105,33.2685050964355,33.3623046875,33.5486335754395,33.7179183959961,33.8324699401855,33.9765129089355,34.0805168151855,34.125,34.2544898986816,34.4103012084961,34.4687004089355,34.5126953125,34.5639152526855,34.6462898254395,34.7340850830078,34.8289566040039,34.9794921875,35.0846176147461,35.203857421875,35.2996101379395,35.4031257629395,35.5822257995605,35.6549339294434,35.7192878723145,35.8018074035645,35.8705558776855,36.0509796142578,36.1887702941895,36.3679695129395,36.4181671142578,36.4556159973145,36.4951171875,36.5189437866211,36.5452651977539,36.6325225830078,36.7607421875,36.7875022888184,36.8339347839355,36.8838844299316,36.9372062683105,36.9976081848145,37.1558609008789,37.1946296691895,37.3403816223145,37.3302726745605,37.3089828491211,37.254638671875,37.21142578125,37.1353530883789,37.181640625,37.2128410339355,37.254150390625,37.2577629089355,37.2512664794922,37.2058601379395,37.0338859558105,36.8653831481934,36.781494140625,36.7318344116211,36.7913551330566,36.8794441223145,36.9302711486816,37.0592765808105,37.072509765625,36.9666976928711,36.8740730285645,36.8414573669434,36.7430152893066,36.4931182861328,36.4196319580078,36.3233375549316,36.2548828125,36.1751480102539,36.0324211120605,35.8872566223145,35.7995109558105,35.7720718383789,35.6338386535645,35.5516128540039,35.453857421875,35.3351058959961,35.2402839660645,35.0336418151855,34.8843269348145,34.678466796875,34.5447273254395,34.3460426330566,34.2800788879395,34.2116203308105,34.1403312683105,34.0563011169434,33.8500480651855,33.728271484375,33.6624984741211,33.6890144348145,33.6096687316895,33.514404296875,33.5188980102539,33.53369140625,33.6263198852539,33.5628890991211,33.3692359924316,33.308837890625,33.2863311767578,33.2715835571289,33.2492179870605,33.2335929870605,33.20947265625,33.2249031066895,33.1819343566895],[40.1363296508789,40.10546875,40.1358871459961,40.1962890625,40.2336921691895,40.2379913330078,40.1778335571289,40.1363296508789],[41.5174789428711,41.497314453125,41.4715843200684,41.4405250549316,41.432373046875,41.4956550598145,41.4940910339355,41.4867210388184,41.4749984741211,41.4540023803711,41.4398422241211,41.4700660705566,41.5592765808105,41.5707015991211,41.5788078308105,41.5857429504395,41.5789070129395,41.563720703125,41.4923820495605,41.4668464660645,41.3528289794922,41.30712890625,41.2879409790039,41.2648429870605,41.2364273071289,41.1991691589355,41.1852035522461,41.1901359558105,41.1765632629395,41.155517578125,41.1259765625,41.1071281433105,41.064697265625,41.0048332214355,40.9682121276855,40.9290046691895,40.7941398620605,40.7195320129395,40.6477546691895,40.5740737915039,40.4823722839355,40.4440422058105,40.393310546875,40.3565940856934,40.2393074035645,40.16650390625,40.149658203125,40.1263694763184,40.0702629089355,40.0231437683105,40.0141105651855,40.0357398986816,40.0403823852539,39.9957504272461,39.8875961303711,39.7464866638184,39.7035140991211,39.6846694946289,39.6504402160645,39.6510734558105,39.6817398071289,39.7685546875,39.7312507629395,39.666748046875,39.422119140625,39.3960456848145,39.3967781066895,39.4052276611328,39.3929710388184,39.37744140625,39.3510246276855,39.3108406066895,39.2599601745605,39.2183113098145,39.1806144714355,39.1448249816895,39.1080589294434,39.0562515258789,39.0167503356934,38.994384765625,38.9343757629395,38.8632316589355,38.8360366821289,38.7006340026855,38.6406745910645,38.5578117370605,38.4201164245605,38.3862762451172,38.3747062683105,38.3695831298828,38.3567886352539,38.334228515625,38.3177757263184,38.2545890808105,38.209716796875,38.146484375,38.1092758178711,38.038818359375,37.9671859741211,37.9080543518066,37.8801727294922,37.8717765808105,37.8292503356934,37.74462890625,37.7103538513184,37.6581573486328,37.6363258361816,37.608642578125,37.5602035522461,37.5063972473145,37.4353981018066,37.4237289428711,37.3571281433105,37.2903823852539,37.269775390625,37.2170906066895,37.1563491821289,37.1424331665039,37.165283203125,37.173583984375,37.176025390625,37.1582527160645,37.1105461120605,37.0584945678711,37.0107421875,36.97802734375,36.9833030700684,37.0118675231934,37.0518074035645,37.20263671875,37.2498550415039,37.282958984375,37.3018569946289,37.3124542236328,37.3135261535645,37.269287109375,37.2235336303711,37.2272453308105,37.2358436584473,37.2445297241211,37.3146476745605,37.3165016174316,37.3448715209961,37.3673820495605,37.3247566223145,37.3349151611328,37.3718757629395,37.3619117736816,37.249267578125,37.1287117004395,37.1085968017578,37.2295875549316,37.2765617370605,37.2822265625,37.2972640991211,37.2886238098145,37.2060546875,37.1563949584961,37.1261215209961,37.0891609191895,37.07080078125,37.0693359375,37.0858879089355,37.1091804504395,37.108154296875,37.0977058410645,37.0088882446289,36.8260765075684,36.7386207580566,36.7022476196289,36.6815910339355,36.6805686950684,36.6946792602539,36.693115234375,36.715087890625,36.7891082763672,36.8622550964355,36.8792495727539,36.8933601379395,36.9015617370605,36.7946281433105,36.7655754089355,36.7436981201172,36.6783218383789,36.643310546875,36.6465797424316,36.6559066772461,36.6526374816895,36.702392578125,36.7583961486816,36.7926750183105,36.8025398254395,36.7776870727539,36.7013664245605,36.50634765625,36.4574203491211,36.2635269165039,36.2339820861816,36.2239265441895,36.220703125,36.2034645080566,36.1712417602539,36.0035133361816,35.9727058410645,35.9375495910645,35.8338890075684,35.8314437866211,35.9100570678711,35.9165534973145,35.9981422424316,36.1590843200684,36.3098640441895,36.4063491821289,36.522705078125,36.6589851379395,36.7430686950684,36.8072280883789,36.8516120910645,36.9105949401855,36.8476066589355,36.778076171875,36.7639656066895,36.7243156433105,36.6527824401855,36.5970191955566,36.5751953125,36.6348648071289,36.7256851196289,36.7992668151855,36.8167953491211,36.7844734191895,36.6041984558105,36.3407745361328,36.2952156066895,36.1819801330566,36.1439933776855,36.15283203125,36.1029815673828,36.095703125,36.035888671875,36.1007308959961,36.1836433410645,36.2678718566895,36.4491195678711,36.5353050231934,36.61279296875,36.8010711669922,36.8217277526855,36.8486824035645,36.8656730651855,36.7971687316895,36.5258293151855,36.4511222839355,36.3103981018066,36.2698707580566,36.2432632446289,36.2876968383789,36.3073272705078,36.2493629455566,36.1680641174316,36.1566886901855,36.2588386535645,36.324462890625,36.397216796875,36.5201148986816,36.590087890625,36.6381340026855,36.6934547424316,36.71533203125,36.673583984375,36.67529296875,36.7008819580078,36.8038101196289,36.8119621276855,36.6863288879395,36.6463851928711,36.6344757080078,36.6702156066895,36.75146484375,36.7364768981934,36.6746101379395,36.6842269897461,36.712158203125,36.746337890625,36.7588844299316,36.7866706848145,36.8092765808105,36.8319816589355,36.9202613830566,36.9963874816895,37.029052734375,37.0294914245605,37.0074234008789,37.0195808410645,36.9818840026855,36.9765625,37.0791511535645,37.1268577575684,37.1224136352539,37.1638679504395,37.2491188049316,37.3067398071289,37.3407211303711,37.3486824035645,37.38916015625,37.4914054870605,37.6036148071289,37.6579093933105,37.6876945495605,37.7254409790039,37.88232421875,37.9786605834961,37.9868659973145,38.0628890991211,38.0547866821289,38.1383285522461,38.1983375549316,38.1763648986816,38.1492652893066,38.1622543334961,38.21435546875,38.2424812316895,38.2771949768066,38.3700714111328,38.3678703308105,38.4406242370605,38.5619125366211,38.6241683959961,38.6412124633789,38.6294937133789,38.5570335388184,38.4869155883789,38.4186058044434,38.3524398803711,38.3357429504395,38.4053726196289,38.4186058044434,38.38818359375,38.3729476928711,38.4157218933105,38.4519538879395,38.4478530883789,38.4817352294922,38.5575675964355,38.6264152526855,38.6602058410645,38.7096176147461,38.736083984375,38.7757797241211,38.8868675231934,38.9190444946289,38.9342308044434,38.9229507446289,38.9609870910645,39.013916015625,39.0567398071289,39.1156272888184,39.2606430053711,39.292236328125,39.3396492004395,39.4190444946289,39.517333984375,39.5496559143066,39.5628890991211,39.5207023620605,39.4840812683105,39.4673843383789,39.5207977294922,39.5689468383789,39.6566429138184,39.8728523254395,39.9900894165039,40.0250015258789,40.197265625,40.4002456665039,40.3963394165039,40.4523468017578,40.4556159973145,40.4148902893066,40.3759307861328,40.3199195861816,40.3288078308105,40.3508796691895,40.3817367553711,40.4814949035645,40.5096206665039,40.512939453125,40.4894523620605,40.4665985107422,40.435302734375,40.3804206848145,40.369873046875,40.4030265808105,40.37646484375,40.390869140625,40.3897476196289,40.4241714477539,40.4673805236816,40.482421875,40.5034675598145,40.5340347290039,40.6305656433105,40.6491203308105,40.7083969116211,40.7380867004395,40.7601051330566,40.7601585388184,40.8092765808105,40.8473129272461,40.9378433227539,40.9634284973145,41.007568359375,41.1016616821289,41.17724609375,41.2210464477539,41.2277374267578,41.1508293151855,41.1969261169434,41.0848655700684,41.1076164245605,41.1579093933105,41.3200187683105,41.5892105102539,41.7295913696289,41.806396484375,41.8917503356934,42.0045928955078,42.017578125,41.9636726379395,41.9568328857422,42.0632781982422,42.0275421142578,41.989501953125,41.9569816589355,41.89111328125,41.7943840026855,41.728515625,41.634033203125,41.7137222290039,41.704833984375,41.6825675964355,41.4265632629395,41.3361358642578,41.2746086120605,41.2625007629395,41.32666015625,41.3525390625,41.3634757995605,41.275390625,41.1844253540039,41.1141128540039,41.0789070129395,41.0019073486328,40.9245109558105,40.9365234375,41.0176773071289,41.1064453125,40.9825172424316,40.9664535522461,40.9771499633789,40.9430160522461,40.9613265991211,41.107421875,41.1902351379395,41.2116203308105,41.2611846923828,41.4236335754395,41.5174789428711],[41.9690437316895,41.8548851013184,41.7291488647461,41.5544929504395,41.4663581848145,41.2483901977539,41.229736328125,41.1404800415039,41.0611343383789,41.0082015991211,40.9741668701172,41.0714836120605,41.0807113647461,41.0613288879395,40.9905776977539,41.0132827758789,40.97314453125,40.8399429321289,40.6873550415039,40.5640144348145,40.498046875,40.261474609375,40.1233901977539,40.0965805053711,40.0753898620605,40.1416969299316,40.202392578125,40.2481422424316,40.314697265625,40.3902359008789,40.4450187683105,40.5442390441895,40.6266098022461,40.6246566772461,40.6063499450684,40.6180648803711,40.611328125,40.6833953857422,40.7267570495605,40.7402839660645,40.7496566772461,40.8265113830566,40.8832054138184,40.9544906616211,40.9970703125,41.0367660522461,41.0643043518066,41.0970230102539,41.1432647705078,41.23876953125,41.3431129455566,41.3541488647461,41.4017601013184,41.5121574401855,41.6012687683105,41.6072273254395,41.6332511901855,41.6633796691895,41.6963386535645,41.716552734375,41.7446784973145,41.7728004455566,41.8015632629395,41.8263664245605,41.8466796875,41.896728515625,41.9479484558105,41.9648933410645,41.9633293151855,41.9751472473145,41.9918441772461,42.02685546875,42.0586433410645,42.0770988464355,42.09326171875,42.0795402526855,42.0250473022461,41.9468765258789,41.9208030700684,41.9329109191895,41.9613304138184,41.9615249633789,41.95654296875,41.9813003540039,41.9866218566895,41.9690437316895],[24.5221195220947,24.45556640625,24.4143562316895,24.4363288879395,24.4766120910645,24.4691410064697,24.5221195220947],[22.620361328125,22.5030765533447,22.379150390625,22.26220703125,22.1415519714355,22.0326652526855,21.9249992370605,21.9560050964355,22.0331058502197,22.15966796875,22.3125495910645,22.3563957214355,22.44189453125,22.4845218658447,22.547607421875,22.5424308776855,22.62744140625,22.7179203033447,22.9749011993408,23.0370121002197,23.0937004089355,23.1497573852539,23.2120590209961,23.30517578125,23.3990726470947,23.526611328125,23.6529293060303,23.7090320587158,24.478515625,24.6422843933105,24.72265625,24.8132820129395,24.927978515625,25.0328121185303,25.0650882720947,25.1591796875,25.2490234375,25.2769050598145,25.2753429412842,25.232421875,25.1815929412842,25.1541004180908,25.1044445037842,25.0564441680908,24.9737300872803,24.895263671875,24.8245124816895,24.746337890625,24.6405277252197,24.5343742370605,24.2852535247803,24.1300773620605,24.052734375,23.8608894348145,23.6682624816895,23.424072265625,23.1725101470947,23.0672855377197,22.9666023254395,22.7763671875,22.620361328125],[-7.97744131088257,-7.99052715301514,-7.97783279418945,-7.93613243103027,-7.90058565139771,-7.83154249191284,-7.73027324676514,-7.66347646713257,-7.64921855926514,-7.72812509536743,-7.90068387985229,-7.91191434860229,-7.97744131088257],[-6.17460918426514,-6.38740205764771,-6.42724657058716,-6.45166015625,-6.45371103286743,-6.41972684860229,-6.34785127639771,-6.36494112014771,-6.27910184860229,-6.27499961853027,-6.17255830764771,-6.08320283889771,-5.93105459213257,-5.85312461853027,-5.72197246551514,-5.8115234375,-5.95117139816284,-6.11542940139771,-6.16621065139771,-6.17460918426514],[-4.90615224838257,-4.95654296875,-5.00400352478027,-5.15517568588257,-5.25546836853027,-5.39443397521973,-5.44384765625,-5.42949247360229,-5.40664052963257,-5.36855506896973,-5.11367177963257,-4.92705106735229,-4.94492197036743,-4.90615224838257],[-1.00205111503601,-1.00205111503601,-1.03984391689301,-1.08457016944885,-1.20361304283142,-1.32255840301514,-1.44160139560699,-1.56054675579071,-1.67958974838257,-1.79863250255585,-1.91757798194885,-2.03662085533142,-2.15566420555115,-2.27460932731628,-2.3935546875,-2.51259756088257,-2.63164043426514,-2.75058579444885,-2.86962890625,-2.98857426643372,-3.04540991783142,-3.07001948356628,-3.17841815948486,-3.24619150161743,-3.30576181411743,-3.40722632408142,-3.46025371551514,-3.4970703125,-3.51152372360229,-3.51679658889771,-3.54082012176514,-3.55976557731628,-3.63613271713257,-3.67441391944885,-3.7392578125,-3.84980487823486,-3.96035122871399,-4.07089805603027,-4.18144512176514,-4.29189443588257,-4.40244150161743,-4.51298809051514,-4.62353515625,-4.67724561691284,-4.6923828125,-4.77646493911743,-4.98046875,-5.06543016433716,-5.16543006896973,-5.23154306411743,-5.5185546875,-5.6259765625,-5.87763643264771,-6.07011699676514,-6.20488214492798,-6.33125019073486,-6.455078125,-6.49931621551514,-6.55595684051514,-6.68525362014771,-6.81494140625,-6.87861347198486,-7.02402353286743,-7.12412118911743,-7.20703125,-7.34140682220459,-7.51787042617798,-7.78769540786743,-7.74668025970459,-7.81279277801514,-8.01152324676514,-8.24287128448486,-8.35097694396973,-8.44384765625,-8.72080135345459,-8.86181640625,-8.90595722198486,-8.94296932220459,-9.19248008728027,-9.40947246551514,-9.57841777801514,-9.72480487823486,-9.83710956573486,-9.91455078125,-10.00048828125,-10.02197265625,-10.0922861099243,-10.1595697402954,-10.1566410064697,-10.20263671875,-10.2406244277954,-10.353515625,-10.4102544784546,-10.4429693222046,-10.46435546875,-10.5515632629395,-10.6875,-10.82080078125,-10.9124021530151,-10.9547853469849,-10.978515625,-11.0345697402954,-11.12255859375,-11.1668949127197,-11.1672849655151,-11.2289056777954,-11.3453130722046,-11.4132814407349,-11.3111333847046,-11.2787113189697,-11.2821292877197,-11.2947273254395,-11.3166990280151,-11.3791017532349,-11.48193359375,-11.5806646347046,-11.6750974655151,-11.7104501724243,-11.6865234375,-11.6471681594849,-11.5921878814697,-11.5669918060303,-11.5712890625,-11.6103515625,-11.6842775344849,-11.7162113189697,-11.7063474655151,-11.6707029342651,-11.6092767715454,-11.5373048782349,-11.4546871185303,-11.4529294967651,-11.5321292877197,-11.58203125,-11.6023445129395,-11.6047849655151,-11.5895500183105,-11.5832033157349,-11.5748052597046,-11.578125,-11.5437507629395,-11.4634761810303,-11.3935546875,-11.3519525527954,-11.3409175872803,-11.3416986465454,-11.3094720840454,-11.2381839752197,-11.1774415969849,-11.1271486282349,-11.0804691314697,-11.0375003814697,-10.990234375,-10.8728513717651,-10.79248046875,-10.7100582122803,-10.6255865097046,-10.5250978469849,-10.4961919784546,-10.4276361465454,-10.3796873092651,-10.31982421875,-10.2411136627197,-10.0731449127197,-10.0301752090454,-9.94882774353027,-9.75654315948486,-9.73154354095459,-9.53779315948486,-9.49540996551514,-9.53173828125,-9.56533241271973,-9.62734413146973,-9.65820217132568,-9.67216873168945,-9.67011737823486,-9.66298770904541,-9.61093807220459,-9.59814453125,-9.60752010345459,-9.61972618103027,-9.60800743103027,-9.51914119720459,-9.50048828125,-9.49589824676514,-9.39501953125,-9.39970684051514,-9.40742206573486,-9.380859375,-9.322265625,-9.27050685882568,-9.21269607543945,-9.15634727478027,-9.13486289978027,-9.12558650970459,-9.07333946228027,-9.0673828125,-9.05400371551514,-9.01943397521973,-8.94218730926514,-8.92197322845459,-8.90224647521973,-8.90322208404541,-8.91435527801514,-8.90878868103027,-8.86328125,-8.80546855926514,-8.71328163146973,-8.65390586853027,-8.60703086853027,-8.61191368103027,-8.59765625,-8.55097675323486,-8.47373104095459,-8.38554668426514,-8.26582050323486,-8.19365215301514,-8.10439395904541,-7.97089815139771,-7.78193330764771,-7.62714862823486,-7.46064472198486,-7.33867216110229,-7.20371103286743,-7.03789043426514,-6.97304677963257,-6.9150390625,-6.80312490463257,-6.69189453125,-6.61689472198486,-6.39443397521973,-6.31386709213257,-6.17207050323486,-6.02500009536743,-5.96542978286743,-5.77597665786743,-5.72265625,-5.65078163146973,-5.49980449676514,-5.40097618103027,-5.31660127639771,-5.17617177963257,-4.98310565948486,-4.89882802963257,-4.83564472198486,-4.66884756088257,-4.49667978286743,-4.44931650161743,-4.45585966110229,-4.41806697845459,-4.30732440948486,-4.08535194396973,-3.99287128448486,-3.85048842430115,-3.77978539466858,-3.73076200485229,-3.65390610694885,-3.58886742591858,-3.49248027801514,-3.41865253448486,-3.388671875,-3.36640596389771,-3.34736323356628,-3.30937504768372,-3.27460956573486,-3.20058584213257,-3.11640644073486,-3.06933617591858,-3.01513671875,-2.98486304283142,-2.97724604606628,-2.93525409698486,-2.91757774353027,-2.89316415786743,-2.87451148033142,-2.82402372360229,-2.76904320716858,-2.75322246551514,-2.69433569908142,-2.65888667106628,-2.64160132408142,-2.61347651481628,-2.42626953125,-2.40009784698486,-2.39677762985229,-2.37382841110229,-2.36347651481628,-2.37167978286743,-2.36269521713257,-2.33847689628601,-2.26542997360229,-2.14335942268372,-2.04404306411743,-1.96748042106628,-1.85068356990814,-1.69365239143372,-1.56308615207672,-1.45869147777557,-1.39677715301514,-1.36748075485229,-1.20820295810699,-1.13115227222443,-1.08300769329071,-1.06728518009186,-1.06249988079071,-1.06972646713257,-1.05136728286743,-1.00751936435699,-0.994921743869781,-0.999023497104645,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601],[-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-1.00205111503601,-0.999023497104645,-0.994921743869781,-1.00751936435699,-1.05136728286743,-1.06972646713257,-1.06249988079071,-1.06728518009186,-1.06601548194885,-1.06308591365814,-1.07460927963257,-1.11308574676514,-1.17880845069885,-1.25419950485229,-1.32109403610229,-1.36865246295929,-1.44697284698486,-1.46992218494415,-1.46630847454071,-1.45175755023956,-1.35166013240814,-1.33554685115814,-1.38710963726044,-1.38789057731628,-1.35673832893372,-1.12138664722443,-0.977343916893005,-0.887109398841858,-0.783105492591858,-0.691308498382568,-0.535253763198853,-0.441699475049973,-0.113573983311653,-0.0602052547037601,0.0983396098017693,0.147216930985451,0.166357159614563,0.263623058795929,0.418945580720901,0.49902331829071,0.673925578594208,0.792871057987213,0.819238543510437,0.863525331020355,0.973486244678497,1.10278332233429,1.18530297279358,1.23881840705872,1.23906242847443,1.68281257152557,1.92202115058899,2.04457974433899,2.08847641944885,2.14628887176514,2.19135737419128,2.23227572441101,2.27006840705872,2.28886723518372,2.2880859375,2.31552767753601,2.36938500404358,2.40327143669128,2.40043926239014,2.45537114143372,2.53027319908142,2.677978515625,2.84702157974243,2.89365243911743,2.93349647521973,2.96757817268372,3.00136709213257,3.04179668426514,3.16337895393372,3.28261733055115,3.34213829040527,3.40893578529358,3.46367192268372,3.49072241783142,3.54414057731628,3.63408231735229,3.72499990463257,3.78559565544128,3.78593707084656,3.73759770393372,3.68046879768372,3.67758822441101,3.70146489143372,3.80263662338257,3.77045869827271,3.70908164978027,3.60756850242615,3.56118154525757,3.52919960021973,3.51972675323486,3.52802705764771,3.60781264305115,3.65131831169128,3.70620107650757,3.74995112419128,3.76318311691284,3.77270531654358,3.79848623275757,3.88017582893372,3.77470755577087,3.75434541702271,3.75507831573486,3.787109375,3.81171846389771,3.98525357246399,4.22021484375,3.88916015625,3.86977505683899,3.84086894989014,3.81298804283142,3.73315382003784,3.69150424003601,3.65058588981628,3.60625028610229,3.41269516944885,3.35751938819885,3.16347646713257,3.11997056007385,2.92475605010986,2.84194350242615,2.818115234375,2.72343730926514,2.61982417106628,2.59575223922729,2.58969688415527,2.47968745231628,2.41791963577271,2.23017597198486,2.06240224838257,1.86191403865814,1.77363264560699,1.71962916851044,1.64335930347443,1.59926736354828,1.55649423599243,1.48901355266571,1.41669952869415,1.38115262985229,1.27285158634186,1.24453127384186,1.23071277141571,1.21425795555115,1.18530297279358,1.15644538402557,1.10156238079071,1.04213857650757,0.867285072803497,0.731249988079071,0.686425805091858,0.605175793170929,0.50512707233429,0.382470518350601,0.294531255960464,0.173779249191284,-0.0169922094792128,-0.397851586341858,-0.831640779972076,-1.00205111503601],[46.2039070129395,46.1546859741211,46.1678237915039,46.2140655517578,46.2577629089355,46.3065910339355,46.3731460571289,46.3031730651855,46.2725563049316,46.2039070129395],[47.0914573669434,47.0802230834961,47.0958480834961,47.0745620727539,46.9168930053711,46.9173355102539,46.876220703125,46.8251457214355,46.7144012451172,46.7641143798828,46.7626953125,46.7325668334961,46.6585922241211,46.6455078125,46.6667976379395,46.6243171691895,46.3813972473145,46.2039070129395,46.169189453125,46.1258811950684,46.10400390625,46.1060066223145,46.232177734375,46.2794914245605,46.3144035339355,46.3707046508789,46.4406242370605,46.2672386169434,46.2420883178711,46.1898918151855,46.0735855102539,45.9873542785645,45.8788070678711,45.7009773254395,45.4469261169434,45.3536148071289,45.3163070678711,45.3108863830566,45.3893547058105,45.401611328125,45.3716812133789,45.4241218566895,45.4530754089355,45.4567375183105,45.4332542419434,45.3935546875,45.3037605285645,45.2323226928711,45.1532707214355,45.0653800964355,45.0259780883789,45.0308113098145,45.0053215026855,45.0396003723145,45.0708503723145,45.1020050048828,45.1193351745605,45.0984840393066,44.9784164428711,44.8963394165039,44.8026351928711,44.8235855102539,44.80712890625,44.7216796875,44.5384254455566,44.423828125,44.3875961303711,44.39892578125,44.4332008361816,44.5536613464355,44.5968246459961,44.6186027526855,44.6805152893066,44.9078140258789,44.9814949035645,45.09765625,45.1878433227539,45.1707534790039,45.1947746276855,45.34814453125,45.3589859008789,45.3280754089355,45.3503913879395,45.40380859375,45.593017578125,45.7492179870605,45.7652359008789,45.8379402160645,45.9470710754395,46.0328636169434,46.0962409973145,46.078857421875,46.0576171875,46.1256828308105,46.1757316589355,46.123779296875,46.1314926147461,46.0836944580078,46.13037109375,46.260986328125,46.2872581481934,46.2816886901855,46.3246574401855,46.3461418151855,46.3486785888672,46.3644027709961,46.3871574401855,46.4299812316895,46.4624519348145,46.4717788696289,46.51025390625,46.5542945861816,46.5549774169922,46.5216789245605,46.5093727111816,46.4749488830566,46.5177764892578,46.5919914245605,46.6156234741211,46.5648460388184,46.5972175598145,46.6424789428711,46.7087898254395,46.7843742370605,46.8548355102539,46.9819831848145,47.0951156616211,47.1572265625,47.2128410339355,47.0870132446289,47.0032730102539,46.9261207580566,46.7216339111328,46.6497535705566,46.6316413879395,46.6424331665039,46.6647453308105,46.7772941589355,46.7383804321289,46.6288108825684,46.6124992370605,46.6244621276855,46.552001953125,46.4730491638184,46.3662109375,46.3040046691895,46.2665023803711,46.1053733825684,46.0901374816895,45.8667488098145,45.8499488830566,45.7979469299316,45.7523918151855,45.7320823669434,45.7546882629395,45.7224617004395,45.6825180053711,45.5999984741211,45.5406723022461,45.3433113098145,45.2599143981934,45.3139152526855,45.3708038330078,45.419677734375,45.4029312133789,45.320556640625,45.2899436950684,45.3110809326172,45.3098640441895,45.2862319946289,45.2668952941895,45.2519073486328,45.240966796875,45.234130859375,45.2921867370605,45.3471183776855,45.450439453125,45.48388671875,45.4985847473145,45.5071754455566,45.5177268981934,45.5415534973145,45.5724105834961,45.6178207397461,45.665771484375,45.7357902526855,45.7938461303711,45.8520011901855,45.9371566772461,45.9786605834961,46.0499534606934,46.1276397705078,46.1764640808105,46.2884292602539,46.3622550964355,46.4241218566895,46.45849609375,46.4970207214355,46.5269050598145,46.5239753723145,46.5049819946289,46.3793449401855,46.376953125,46.3926239013672,46.4665985107422,46.445068359375,46.4369125366211,46.4559593200684,46.4537582397461,46.4346694946289,46.4077644348145,46.3988265991211,46.416748046875,46.44873046875,46.4377937316895,46.3505363464355,46.3602066040039,46.3778305053711,46.4015617370605,46.423095703125,46.5388679504395,46.625,46.7237815856934,46.7824211120605,46.8289070129395,46.8829116821289,46.9388198852539,46.9640159606934,46.9967269897461,47.0475120544434,47.0911140441895,47.1280288696289,47.185546875,47.246826171875,47.27099609375,47.2907218933105,47.2926254272461,47.3280258178711,47.375732421875,47.4444808959961,47.4556617736816,47.4896965026855,47.5303726196289,47.5808601379395,47.6585960388184,47.7315444946289,47.7750015258789,47.8824234008789,47.9645538330078,47.9754409790039,47.9523429870605,47.9330062866211,47.9511260986328,47.9956550598145,48.1195793151855,48.1443862915039,48.1502914428711,48.0905303955078,48.1086883544922,48.1468734741211,48.162109375,48.1444320678711,48.1614265441895,48.2130393981934,48.2385711669922,48.2379875183105,48.2570304870605,48.2957992553711,48.3212890625,48.3335418701172,48.365234375,48.416259765625,48.4495124816895,48.4648895263672,48.4704093933105,48.4777336120605,48.4430656433105,48.4156265258789,48.4327125549316,48.3714332580566,48.3682594299316,48.3719253540039,48.3871574401855,48.2941398620605,48.2598609924316,48.2484855651855,48.22998046875,48.2037582397461,48.1132316589355,48.0643539428711,47.9925308227539,47.9675788879395,47.9324722290039,47.9107933044434,47.8230972290039,47.7457046508789,47.72412109375,47.7177734375,47.7608375549316,47.8765144348145,47.9310531616211,47.9471168518066,47.9380378723145,47.9111824035645,47.9060554504395,47.9447746276855,47.9410171508789,47.935791015625,47.9826202392578,47.9903793334961,47.9923362731934,47.995849609375,47.989990234375,48.0845222473145,48.08740234375,48.0491218566895,48.0065460205078,47.9642601013184,47.947265625,47.9603004455566,47.9970703125,48.029541015625,48.060302734375,48.09521484375,48.109619140625,48.1070289611816,48.1036109924316,48.1043930053711,48.134033203125,48.2053718566895,48.2433090209961,48.2560539245605,48.28662109375,48.3272933959961,48.3580093383789,48.3608894348145,48.4073753356934,48.4121589660645,48.4134292602539,48.4053230285645,48.5685043334961,48.6858406066895,48.7450675964355,48.8734855651855,48.9335441589355,48.9832496643066,49.0313987731934,49.0727081298828,49.0771980285645,49.0399436950684,49.020751953125,49.0389175415039,49.062744140625,49.0812492370605,49.13623046875,49.1711921691895,49.1927223205566,49.240966796875,49.295166015625,49.3538093566895,49.4836921691895,49.5390129089355,49.606201171875,49.7662620544434,49.8263664245605,49.8990707397461,50.0728492736816,50.1739234924316,50.2298316955566,50.3270492553711,50.3773460388184,50.4100570678711,50.45703125,50.5084457397461,50.5304718017578,50.6170387268066,50.7228012084961,50.7601547241211,50.7855949401855,50.8093757629395,50.816162109375,50.8195304870605,50.844970703125,50.8727531433105,50.9404296875,50.9925308227539,51.126220703125,51.26513671875,51.31005859375,51.3524894714355,51.3949241638184,51.448974609375,51.5179214477539,51.525390625,51.6104965209961,51.6288566589355,51.6413078308105,51.6371078491211,51.6239738464355,51.5850601196289,51.59130859375,51.6646461486816,51.7747077941895,51.8384284973145,51.8675270080566,51.883056640625,51.8895034790039,51.8882827758789,51.8991203308105,51.9111328125,51.9305152893066,51.937744140625,51.9247550964355,51.923828125,51.9135246276855,51.8550300598145,51.8444366455078,51.8134307861328,51.8019027709961,51.7707023620605,51.7540016174316,51.7608413696289,51.7520484924316,51.6135749816895,51.597412109375,51.5941390991211,51.6061019897461,51.6016120910645,51.5724105834961,51.4899406433105,51.4779777526855,51.4825668334961,51.5291481018066,51.5774421691895,51.5923843383789,51.5597648620605,51.5650405883789,51.6016578674316,51.6078605651855,51.5818328857422,51.5636215209961,51.5624504089355,51.5426254272461,51.45654296875,51.4388694763184,51.4333992004395,51.5103492736816,51.5401840209961,51.5621566772461,51.57177734375,51.5989265441895,51.6254386901855,51.6275405883789,51.6172866821289,51.5806159973145,51.4970207214355,51.4130363464355,51.382568359375,51.4083480834961,51.4345703125,51.4395523071289,51.4580078125,51.4820327758789,51.4778785705566,51.4512214660645,51.3996086120605,51.3255386352539,51.2743148803711,51.2650413513184,51.318359375,51.3554191589355,51.4063453674316,51.4712409973145,51.531494140625,51.5963401794434,51.68896484375,51.7700691223145,51.8141136169434,51.8951644897461,51.9530754089355,52.0461921691895,52.0769538879395,52.0629386901855,52.0502471923828,52.1053733825684,52.1081047058105,52.10107421875,52.0994110107422,52.0708999633789,52.046630859375,52.0450210571289,52.0505867004395,52.0829582214355,52.114013671875,52.2721214294434,52.2948226928711,52.3072280883789,52.3085479736328,52.2791023254395,52.2526359558105,52.25634765625,52.3404312133789,52.3535652160645,52.3337898254395,52.3326148986816,52.3447761535645,52.3156242370605,52.25146484375,52.1559562683105,51.9796371459961,51.7804183959961,51.7415046691895,51.7165069580078,51.6922378540039,51.6791496276855,51.6449699401855,51.6079597473145,51.5538063049316,51.4840812683105,51.419921875,51.3632316589355,51.3401870727539,51.3116683959961,51.27685546875,51.25537109375,51.2437973022461,51.237060546875,51.203125,51.1722183227539,51.1693344116211,51.189208984375,51.2017593383789,51.2034187316895,51.1806640625,51.120849609375,51.0609855651855,51.0438957214355,51.0467796325684,51.0438957214355,51.0211448669434,50.9869155883789,50.9499015808105,50.904296875,50.7989234924316,50.767578125,50.7276840209961,50.6820793151855,50.6422348022461,50.6109352111816,50.5396995544434,50.4599113464355,50.4399909973145,50.3687477111816,50.3459968566895,50.40576171875,50.4371070861816,50.419677734375,50.4085464477539,50.3678245544434,50.311767578125,50.2804679870605,50.2968254089355,50.2804679870605,50.23486328125,50.209228515625,50.2462387084961,50.2918434143066,50.3395500183105,50.3515129089355,50.3608856201172,50.3949699401855,50.4176254272461,50.4114761352539,50.3407249450684,50.2918434143066,50.2149429321289,50.1090812683105,49.9642105102539,49.9200172424316,49.9278335571289,49.9394035339355,49.9545402526855,50.025390625,50.0514640808105,50.0523452758789,49.9640617370605,49.95458984375,49.952880859375,49.8843269348145,49.82470703125,49.8184089660645,49.8417472839355,49.85595703125,49.8332023620605,49.7819328308105,49.7420425415039,49.7306671142578,49.72802734375,49.6506843566895,49.572021484375,49.5676765441895,49.5907745361328,49.5967292785645,49.5768547058105,49.5426750183105,49.4970703125,49.4315452575684,49.3688468933105,49.3072280883789,49.2515602111816,49.2002944946289,49.1298332214355,49.0640640258789,49.0365715026855,49.0079078674316,48.9595718383789,48.9144554138184,48.8779792785645,48.8514137268066,48.8220710754395,48.807373046875,48.7937507629395,48.8077163696289,48.7820777893066,48.7393569946289,48.6624526977539,48.5912132263184,48.5718765258789,48.5427742004395,48.4842262268066,48.4190902709961,48.3604507446289,48.3319358825684,48.3027839660645,48.2884254455566,48.2819328308105,48.2688941955566,48.2379417419434,48.1683578491211,48.0353050231934,47.9644546508789,47.8875503540039,47.8448219299316,47.8412094116211,47.833740234375,47.83740234375,47.8484878540039,47.8551254272461,47.8370132446289,47.714111328125,47.6659164428711,47.6224098205566,47.6099586486816,47.5591812133789,47.4795417785645,47.408935546875,47.3736801147461,47.3427734375,47.32080078125,47.2965316772461,47.2876930236816,47.2766609191895,47.259033203125,47.2369651794434,47.2127418518066,47.1752471923828,47.1355972290039,47.0914573669434],[-33.7421875,-33.7791976928711,-33.8493194580078,-34.0174789428711,-34.24951171875,-34.38037109375,-34.5169944763184,-34.6707038879395,-34.6668968200684,-34.7327156066895,-34.9328117370605,-34.8951187133789,-34.8958015441895,-34.8076171875,-34.7756843566895,-34.8109359741211,-34.9079132080078,-34.9064483642578,-34.9012680053711,-34.8610343933105,-34.775390625,-34.6766586303711,-34.4523468017578,-34.4480438232422,-34.4773445129395,-34.4476547241211,-34.39013671875,-34.3069343566895,-34.1090850830078,-33.9124031066895,-33.7191429138184,-33.5088882446289,-33.2600555419922,-33.1823234558105,-33.1379890441895,-33.1291007995605,-33.0646476745605,-32.9673843383789,-32.8936500549316,-32.7572250366211,-32.5665016174316,-32.4716796875,-32.3218765258789,-32.2489242553711,-32.1848640441895,-32.1190414428711,-32.0515632629395,-31.9865245819092,-31.9242191314697,-31.8726577758789,-31.8318367004395,-31.7692375183105,-31.6849613189697,-31.6206073760986,-31.5761737823486,-31.5343761444092,-31.4949207305908,-31.4166030883789,-31.2994136810303,-31.1953105926514,-31.1043949127197,-31.0310554504395,-30.9751949310303,-30.9374027252197,-30.91748046875,-30.8585929870605,-30.7120113372803,-30.5910129547119,-30.4952125549316,-30.3844718933105,-30.2950210571289,-30.2269535064697,-30.1877937316895,-30.2612323760986,-30.2806644439697,-30.2833995819092,-30.2648448944092,-30.1444358825684,-30.1099605560303,-30.10107421875,-30.1072273254395,-30.1869125366211,-30.4474601745605,-30.6284198760986,-30.7137699127197,-30.7776374816895,-30.8372077941895,-30.9918937683105,-31.0596675872803,-31.0791988372803,-31.0808582305908,-31.0696296691895,-31.0367183685303,-30.9871101379395,-30.9465827941895,-30.9249038696289,-30.8920879364014,-30.8581047058105,-30.8507823944092,-30.8759765625,-30.9644546508789,-31.0461902618408,-31.0929679870605,-31.1417007446289,-31.1841773986816,-31.2255859375,-31.2795886993408,-31.31396484375,-31.2790050506592,-31.3912105560303,-31.4851570129395,-31.5419902801514,-31.6227550506592,-31.7450180053711,-31.8551750183105,-31.9015636444092,-31.9281253814697,-31.9523429870605,-31.9945316314697,-32.0399436950684,-32.0568389892578,-32.0974578857422,-32.1863288879395,-32.2987327575684,-32.4030303955078,-32.5032234191895,-32.5811538696289,-32.6253929138184,-32.6800765991211,-32.7367172241211,-32.8210945129395,-32.9270515441895,-33.0103530883789,-33.0685539245605,-33.1086921691895,-33.1708984375,-33.5002899169922,-33.6228523254395,-33.6554718017578,-33.67724609375,-33.7098617553711,-33.7373046875,-33.7421875],[19.0120105743408,18.9639167785645,18.9676761627197,19.0705070495605,19.1258316040039,19.3825187683105,19.5908203125,19.7499504089355,19.8315906524658,19.8947257995605,20.01416015625,20.1673831939697,20.2598152160645,20.2758293151855,20.1634273529053,19.994384765625,19.8756351470947,19.7481937408447,19.7319831848145,19.6446285247803,19.5681648254395,19.5244636535645,19.4541015625,19.3191890716553,19.2601566314697,19.1090812683105,19.0120105743408],[20.7726554870605,20.7444839477539,20.7575206756592,20.82568359375,20.9124507904053,20.9300289154053,20.9048347473145,20.8777828216553,20.8311538696289,20.7726554870605],[20.9325675964355,20.9147453308105,20.9414558410645,20.9512710571289,20.885498046875,20.84033203125,20.7920894622803,20.7571296691895,20.7147960662842,20.6447734832764,20.6286125183105,20.5987796783447,20.6051769256592,20.6178722381592,20.7062492370605,20.8012199401855,20.7899894714355,20.8218269348145,20.9014148712158,20.9490737915039,21.0245113372803,21.0343265533447,20.99267578125,20.9325675964355],[21.2153797149658,21.1879405975342,21.19970703125,21.1772956848145,21.1635246276855,21.1550788879395,21.1035652160645,21.0563488006592,21.0978012084961,21.1125984191895,21.15234375,21.1805667877197,21.2297859191895,21.2153797149658],[21.4566421508789,21.450927734375,21.4577159881592,21.3780765533447,21.3339347839355,21.3076171875,21.2797355651855,21.2686023712158,21.2908191680908,21.340576171875,21.326904296875,21.3668937683105,21.3785152435303,21.3677234649658,21.3161144256592,21.312255859375,21.318603515625,21.3771476745605,21.4893569946289,21.5330581665039,21.5852546691895,21.6002445220947,21.6917953491211,21.7013664245605,21.5533695220947,21.5119152069092,21.471435546875,21.4566421508789],[21.841064453125,21.796875,21.8036613464355,21.8430652618408,21.8972663879395,21.9440441131592,22.0152339935303,22.004638671875,21.9581050872803,21.9074211120605,21.8787593841553,21.841064453125],[21.932373046875,21.8761234283447,21.900390625,21.9095230102539,21.9517593383789,21.9898433685303,22.0417976379395,22.1401844024658,22.22314453125,22.2195796966553,22.154052734375,22.1052742004395,22.0506839752197,21.9736328125,21.932373046875],[24.5445804595947,24.5423336029053,24.5578117370605,24.5767097473145,24.575439453125,24.5544929504395,24.5445804595947],[24.5999031066895,24.5900382995605,24.62939453125,24.6891593933105,24.6424808502197,24.6141605377197,24.5999031066895],[24.6504878997803,24.6299324035645,24.6362781524658,24.666259765625,24.7326164245605,24.75,24.68505859375,24.6676273345947,24.6504878997803],[24.716796875,24.693115234375,24.7104969024658,24.7341785430908,24.7594738006592,24.7278804779053,24.716796875],[24.8036632537842,24.8036632537842,24.81787109375,24.8462886810303,24.8352546691895,24.821044921875,24.8036632537842],[24.9031734466553,24.8984375,24.9411144256592,24.9379386901855,24.9031734466553],[25.1422843933105,24.9542484283447,25.0013179779053,25.1019535064697,25.1493167877197,25.1793460845947,25.233642578125,25.2969722747803,25.3412609100342,25.3476066589355,25.1422843933105],[26.45361328125,26.4275379180908,26.4466800689697,26.48095703125,26.5480480194092,26.4770011901855,26.4609375,26.45361328125],[26.5523433685303,26.4936027526855,26.5919933319092,26.7007331848145,26.66552734375,26.5523433685303],[26.1593761444092,26.1129398345947,26.3297843933105,26.8205089569092,27.1001968383789,27.1964855194092,26.8014640808105,26.2998046875,26.1593761444092],[27.300048828125,27.2425289154053,27.3282718658447,27.5230960845947,27.7791519165039,27.822021484375,27.5412120819092,27.300048828125],[27.2784175872803,27.2047843933105,27.3755855560303,27.6434078216553,27.8505363464355,27.7945308685303,27.6786117553711,27.2784175872803],[27.901611328125,27.899169921875,27.9810543060303,28.0138664245605,28.1174812316895,28.1329097747803,28.0888175964355,28.0160160064697,27.901611328125],[28.152587890625,28.1484375,28.2024421691895,28.2296867370605,28.3334465026855,28.340576171875,28.3763179779053,28.381591796875,28.3377914428711,28.2755870819092,28.152587890625],[29.1458988189697,29.1363277435303,29.2901363372803,29.3413066864014,29.3390636444092,29.2528800964355,29.1458988189697],[29.5007305145264,29.4864768981934,29.5730972290039,29.6103038787842,29.6439456939697,29.6409645080566,29.5969715118408,29.5847187042236,29.56689453125,29.5390148162842,29.5007305145264],[29.6426277160645,29.6066398620605,29.6328125,29.6377925872803,29.6271991729736,29.6786613464355,29.7176265716553,29.732421875,29.6426277160645],[29.7125968933105,29.6602535247803,29.6802253723145,29.7326164245605,29.7529792785645,29.7125968933105],[29.8077144622803,29.77587890625,29.9283695220947,30.0567398071289,30.0003910064697,29.9333477020264,29.8077144622803],[30.0840797424316,30.03759765625,30.0607414245605,30.0628414154053,30.0787124633789,30.0941886901855,30.1108417510986,30.1686515808105,30.1262187957764,30.0840797424316],[30.2159175872803,30.2047843933105,30.2255840301514,30.2449207305908,30.2642574310303,30.2291507720947,30.2159175872803],[30.2523422241211,30.2309093475342,30.23291015625,30.2404308319092,30.2547359466553,30.2737312316895,30.2523422241211],[30.971435546875,30.7277851104736,30.8140640258789,30.8978538513184,30.9474124908447,30.971435546875],[32.8275871276855,32.8185043334961,32.8389167785645,32.9355964660645,33.0111808776855,33.0326652526855,32.9599113464355,32.8494644165039,32.8275871276855],[33.2171859741211,33.21533203125,33.224609375,33.2783203125,33.2820320129395,33.2746086120605,33.232421875,33.2171859741211],[33.3857421875,33.3121109008789,33.3212394714355,33.3170890808105,33.3571281433105,33.4127922058105,33.427001953125,33.4319839477539,33.4370613098145,33.4641609191895,33.4771003723145,33.4150886535645,33.3857421875],[33.9188499450684,33.9048805236816,33.9180679321289,34.0138664245605,34.0265121459961,33.9849090576172,33.9733390808105,33.9188499450684],[34.0248527526855,34.0222663879395,34.0329093933105,34.0563011169434,34.0732917785645,34.0605964660645,34.0248527526855],[34.0796852111816,34.0284652709961,34.0529823303223,34.0281715393066,34.006591796875,33.9677734375,33.9949188232422,34.0321807861328,34.0678215026855,34.0796852111816],[34.6548843383789,34.6525382995605,34.66357421875,34.6846656799316,34.7001495361328,34.6945304870605,34.6548843383789],[34.6429672241211,34.6314926147461,34.75634765625,34.9146957397461,34.9389190673828,34.8036613464355,34.6429672241211],[35.190185546875,35.1188468933105,35.1230964660645,35.1741218566895,35.190185546875],[35.2400894165039,35.2128410339355,35.2215805053711,35.2786140441895,35.4794921875,35.5721206665039,35.7354011535645,35.7691421508789,35.7165031433105,35.5641593933105,35.4486351013184,35.2803192138672,35.2400894165039],[35.8559074401855,35.8355941772461,35.9461402893066,35.910400390625,35.8806648254395,35.8559074401855],[37.8882827758789,37.8720703125,38.0723152160645,38.2400894165039,38.298095703125,38.1805191040039,38.0724143981934,37.8882827758789],[39.6807632446289,39.5293960571289,39.5584945678711,39.7464332580566,39.6807632446289],[40.5228500366211,40.5186996459961,40.6146011352539,40.658447265625,40.6493186950684,40.6154289245605,40.5864715576172,40.5418434143066,40.5228500366211],[40.9860343933105,40.9213371276855,40.914794921875,40.9337882995605,40.97216796875,41.0240745544434,41.0467758178711,41.0514640808105,41.0150413513184,41.0442886352539,41.0606918334961,40.8941421508789,40.8753890991211,40.8257827758789,40.7906227111816,40.77783203125,40.6541976928711,40.6515121459961,40.66357421875,40.5998992919922,40.5927276611328,40.5705070495605,40.6217765808105,40.6409683227539,40.6559562683105,40.651611328125,40.5988273620605,40.5811996459961,40.638671875,40.6831550598145,40.725341796875,40.7916526794434,40.8336906433105,40.8700180053711,40.8380355834961,40.8812484741211,40.9062004089355,40.9196281433105,40.9199714660645,40.9267578125,40.9411125183105,40.9437980651855,40.9242172241211,40.9298324584961,40.9568862915039,40.9659652709961,40.9720726013184,40.9918441772461,41.0270042419434,41.1255340576172,41.1530265808105,41.0385246276855,40.9860343933105],[41.2655754089355,41.2494621276855,41.2863273620605,41.3175811767578,41.3284683227539,41.3744125366211,41.3974609375,41.2986335754395,41.2655754089355],[41.3763198852539,41.3274421691895,41.3589859008789,41.3735847473145,41.4485359191895,41.4572257995605,41.414794921875,41.3763198852539],[41.4852523803711,41.4667472839355,41.5150413513184,41.570556640625,41.5718269348145,41.5422859191895,41.4852523803711],[41.491943359375,41.464599609375,41.4693832397461,41.5062980651855,41.5604972839355,41.6200218200684,41.6382331848145,41.654296875,41.491943359375],[44.196044921875,44.17626953125,44.1826667785645,44.2319831848145,44.2487335205078,44.2561988830566,44.2423362731934,44.196044921875],[44.3324699401855,44.31298828125,44.3214836120605,44.2686996459961,44.2497100830078,44.27685546875,44.2943382263184,44.3642578125,44.4303703308105,44.4564971923828,44.4383773803711,44.3643569946289,44.3324699401855],[47.2047348022461,47.18505859375,47.1861343383789,47.2261238098145,47.2543449401855,47.2747077941895,47.21630859375,47.2047348022461],[47.395263671875,47.3725090026855,47.3547859191895,47.3593254089355,47.3861351013184,47.3902320861816,47.3580093383789,47.4216804504395,47.48876953125,47.489990234375,47.4461402893066,47.395263671875],[47.5945816040039,47.575439453125,47.5982894897461,47.6194801330566,47.6668434143066,47.69775390625,47.7039527893066,47.6905784606934,47.6826667785645,47.5945816040039],[48.1566429138184,48.0254402160645,48.080078125,47.9854507446289,47.9388198852539,47.9231910705566,47.917724609375,47.9313468933105,47.9640121459961,47.9813003540039,47.9924812316895,48.0296363830566,48.0550270080566,48.1285629272461,48.1514167785645,48.156494140625,48.1738777160645,48.2252922058105,48.239013671875,48.2809066772461,48.3510246276855,48.3842277526855,48.380615234375,48.3595695495605,48.3211898803711,48.2939949035645,48.2410659790039,48.2286605834961,48.2137680053711,48.200439453125,48.1566429138184],[48.4313468933105,48.4215316772461,48.4346694946289,48.4569358825684,48.4847679138184,48.5379867553711,48.5516090393066,48.5486335754395,48.5018577575684,48.4523429870605,48.4313468933105],[48.5008811950684,48.468017578125,48.4890632629395,48.5079574584961,48.5263175964355,48.5867195129395,48.6063957214355,48.61328125,48.5384750366211,48.5008811950684],[48.6727066040039,48.6509780883789,48.6298332214355,48.6265602111816,48.66064453125,48.6646995544434,48.6123046875,48.594482421875,48.626708984375,48.6521949768066,48.6791496276855,48.7069854736328,48.7103500366211,48.6727066040039],[45.0038566589355,45.00390625,45.0041999816895,45.0046882629395,45.0051765441895,45.005615234375,45.006103515625,45.006591796875,45.007080078125,45.007568359375,45.2003402709961,45.2900848388672,45.2603492736816,45.2628402709961,45.3091316223145,45.3372573852539,45.3331069946289,45.2907218933105,45.262451171875,45.2707023620605,45.3106918334961,45.3661613464355,45.40478515625,45.4106941223145,45.4094696044922,45.4283180236816,45.4553718566895,45.4989242553711,45.5513687133789,45.6439933776855,45.7068367004395,45.7382316589355,45.8019027709961,45.8680686950684,45.9061050415039,45.9391593933105,45.9798316955566,46.057373046875,46.1500015258789,46.2508773803711,46.3418464660645,46.4410667419434,46.5714378356934,46.7089347839355,46.8429183959961,46.994873046875,47.0813484191895,47.2386703491211,47.3506355285645,47.4020004272461,47.4629898071289,47.4447746276855,47.4266090393066,47.338134765625,47.2736320495605,47.2364273071289,47.2112312316895,47.2028312683105,47.2033195495605,47.2534675598145,47.2857933044434,47.316162109375,47.3445320129395,47.3544921875,47.345947265625,47.2748527526855,47.1676254272461,47.0828132629395,46.9357414245605,46.7798843383789,46.6156234741211,46.4983863830566,46.33740234375,46.2093276977539,46.0421371459961,45.9527816772461,45.927001953125,45.8917999267578,45.8741683959961,45.8601570129395,45.842529296875,45.81787109375,45.7955589294434,45.769775390625,45.7275390625,45.7017059326172,45.6864776611328,45.6864776611328,45.6711921691895,45.6441917419434,45.6207542419434,45.612548828125,45.618408203125,45.6031227111816,45.5655746459961,45.5304183959961,45.5139656066895,45.5010261535645,45.4740715026855,45.4458999633789,45.4212379455566,45.3779296875,45.3403816223145,45.3086929321289,45.27587890625,45.2476577758789,45.2101554870605,45.1737785339355,45.15380859375,45.1679229736328,45.1867179870605,45.2007827758789,45.1925315856934,45.1819801330566,45.16943359375,45.1390113830566,45.0877456665039,44.9891586303711,44.9443817138672,44.8850593566895,44.8677711486816,44.849609375,44.8276863098145,44.6755828857422,44.6968727111816,44.6565399169922,44.644775390625,44.5768051147461,44.5624008178711,44.5665016174316,44.585693359375,44.5762710571289,44.45361328125,44.4643058776855,44.420166015625,44.40087890625,44.3843231201172,44.4388198852539,44.4906272888184,44.5020027160645,44.5152359008789,44.5147933959961,44.5073738098145,44.473876953125,44.4451179504395,44.4690895080566,44.5076179504395,44.4456520080566,44.3802223205566,44.3039054870605,44.2586441040039,44.2708511352539,44.310546875,44.34228515625,44.33935546875,44.3817405700684,44.4425773620605,44.4544906616211,44.446044921875,44.4544906616211,44.509765625,44.5707511901855,44.5494117736816,44.4850578308105,44.433837890625,44.3480987548828,44.17236328125,44.0975608825684,44.037841796875,43.9864768981934,44.0009307861328,43.956298828125,43.9050788879395,43.8973617553711,43.9625968933105,43.9827651977539,43.8865699768066,43.880615234375,43.9488296508789,43.993896484375,43.9550285339355,43.8520050048828,43.8606948852539,43.8990211486816,43.91064453125,43.8668441772461,43.8052253723145,43.7723159790039,43.7898941040039,43.8195343017578,43.7970199584961,43.7878913879395,43.8180694580078,43.8550758361816,43.8346214294434,43.766357421875,43.6719207763672,43.6562042236328,43.6261215209961,43.480224609375,43.3488273620605,43.1344223022461,43.1093254089355,43.0700187683105,42.9405746459961,42.8253402709961,42.7740211486816,42.7212371826172,42.6692886352539,42.6646003723145,42.6739730834961,42.6717758178711,42.6497039794922,42.6232414245605,42.6166496276855,42.5703620910645,42.5525856018066,42.4966316223145,42.4319839477539,42.3311042785645,42.2999992370605,42.2649421691895,42.2288551330566,42.0404281616211,42.0215797424316,41.987060546875,41.9386215209961,41.8033180236816,41.7572784423828,41.7289543151855,41.7698707580566,41.826171875,41.872314453125,41.9796867370605,42.0301246643066,42.0627937316895,42.0351066589355,42.071044921875,42.0912132263184,42.1010246276855,42.0971183776855,42.0783195495605,41.9612808227539,41.807861328125,41.71044921875,41.6771469116211,41.6839866638184,41.6773414611816,41.6269073486328,41.5824699401855,41.5342292785645,41.5583000183105,41.6081047058105,41.7101058959961,41.71484375,41.5485343933105,41.5380859375,41.4894027709961,41.5164070129395,41.64111328125,41.7457046508789,41.7440414428711,41.6812515258789,41.7198753356934,41.7622566223145,41.7862281799316,41.7953147888184,41.7027359008789,41.63330078125,41.4537124633789,41.3789558410645,41.3309097290039,41.341064453125,41.3261260986328,41.2916488647461,41.3121566772461,41.2757835388184,41.265869140625,41.28515625,41.2164573669434,41.1758308410645,41.0218734741211,40.9658699035645,40.87841796875,40.8313980102539,40.8161125183105,40.7769546508789,40.7513656616211,40.8387718200684,40.9124488830566,41.05517578125,41.1706047058105,41.2180671691895,41.2497100830078,41.1357917785645,40.99609375,40.9142570495605,40.7563972473145,40.7196273803711,40.6873054504395,40.6732406616211,40.6479988098145,40.6080093383789,40.5286140441895,40.4562492370605,40.4298324584961,40.4521484375,40.4003410339355,40.328369140625,40.2505378723145,40.1713371276855,40.072998046875,39.9230461120605,39.7881317138672,39.8291015625,39.9931144714355,39.9759788513184,39.9381332397461,39.7266120910645,39.6138687133789,39.535888671875,39.5487785339355,39.48681640625,39.4545402526855,39.38720703125,39.3425788879395,39.3468742370605,39.3161125183105,39.2925796508789,39.2475090026855,39.2078590393066,39.0019035339355,38.9411163330078,38.949951171875,39.0471687316895,39.1454620361328,39.188232421875,39.2108383178711,39.2078590393066,39.2842788696289,39.33984375,39.4901847839355,39.5318832397461,39.6018562316895,39.71240234375,39.7896957397461,39.8297348022461,39.8705101013184,39.9318351745605,39.9834976196289,39.894775390625,39.8646965026855,39.8315887451172,39.7809562683105,39.7173843383789,39.6407737731934,39.5894546508789,39.552978515625,39.4769515991211,39.40283203125,39.2813949584961,39.0927734375,38.966552734375,38.8193817138672,38.7775382995605,38.7228012084961,38.6324195861816,38.5911102294922,38.599365234375,38.5787124633789,38.5033187866211,38.4263687133789,38.3830070495605,38.36572265625,38.4100074768066,38.4253921508789,38.4062004089355,38.3843231201172,38.2981452941895,38.2550773620605,38.2422828674316,38.1291999816895,38.0650405883789,37.6311988830566,37.5586929321289,37.5353507995605,37.516357421875,37.4729995727539,37.4251937866211,37.296630859375,37.1519050598145,37.2122077941895,37.2638168334961,37.3984375,37.619140625,37.75634765625,37.8213882446289,37.9539527893066,37.9737319946289,37.9715843200684,38.0327644348145,38.086669921875,38.140380859375,38.147216796875,38.1692390441895,38.2139663696289,38.26123046875,38.3187522888184,38.362060546875,38.3555183410645,38.3096656799316,38.2913589477539,38.3319320678711,38.32275390625,38.2948722839355,38.279541015625,38.3176765441895,38.361328125,38.4364242553711,38.49462890625,38.5999488830566,38.61865234375,38.6015625,38.6017074584961,38.6250991821289,38.6212387084961,38.7066879272461,38.75830078125,38.7724609375,38.7228507995605,38.7096672058105,38.8182106018066,38.8226547241211,38.8527336120605,38.9155769348145,38.9430694580078,38.9085922241211,38.9527854919434,39.0093765258789,39.0091819763184,38.99072265625,39.0821304321289,39.1229476928711,39.0636253356934,39.1916007995605,39.3150405883789,39.3688468933105,39.3672828674316,39.3759765625,39.3985824584961,39.4108390808105,39.4596176147461,39.4683609008789,39.5108871459961,39.5045890808105,39.5850563049316,39.5686988830566,39.5611305236816,39.5270004272461,39.4703102111816,39.43310546875,39.4032211303711,39.3799285888672,39.4203147888184,39.4386215209961,39.3521499633789,39.32275390625,39.4039077758789,39.3875503540039,39.364501953125,39.3246574401855,39.3039054870605,39.2528305053711,39.2250022888184,39.2693367004395,39.2542953491211,39.15869140625,39.1260261535645,39.0738754272461,39.0306129455566,39.0679664611816,39.0652313232422,39.0011711120605,38.9452171325684,38.8983421325684,38.8406257629395,38.7882804870605,38.7426261901855,38.5321769714355,38.4749488830566,38.4202117919922,38.3689956665039,38.3615226745605,38.4036598205566,38.435791015625,38.5385246276855,38.5795402526855,38.6119651794434,38.5375022888184,38.454345703125,38.2682609558105,38.1968727111816,38.1407699584961,38.0870132446289,38.1250495910645,38.1735343933105,38.2283210754395,38.262939453125,38.3902854919434,38.337158203125,38.2920913696289,38.3470230102539,38.3938980102539,38.4452629089355,38.4417495727539,38.3971176147461,38.40771484375,38.4948234558105,38.5409698486328,38.6500968933105,38.705810546875,38.7777328491211,38.8892555236816,38.7757797241211,38.7195320129395,38.6765594482422,38.6000022888184,38.5291976928711,38.3966331481934,38.3517570495605,38.3400421142578,38.3701171875,38.356689453125,38.1970710754395,38.1339378356934,38.094482421875,38.0111808776855,37.9632339477539,37.8935546875,37.8480949401855,37.7943344116211,37.7215805053711,37.6756858825684,37.67041015625,37.6822242736816,37.93798828125,37.9615211486816,38.095458984375,38.1671905517578,38.1656723022461,38.0330085754395,37.9402351379395,37.8101539611816,37.7550811767578,37.6691436767578,37.6288566589355,37.5714836120605,37.5302734375,37.4951667785645,37.4306144714355,37.3570289611816,37.3861351013184,37.3319320678711,37.2999496459961,37.2735328674316,37.3093757629395,37.5054206848145,37.4791984558105,37.4487800598145,37.3225593566895,37.2468757629395,37.2126960754395,37.1492652893066,37.1108894348145,37.0526885986328,37.0131340026855,36.9913063049316,37.03076171875,37.0723152160645,37.1428680419922,37.2217254638672,37.2176780700684,37.3176727294922,37.3292007446289,37.30908203125,37.2957038879395,37.2710456848145,37.2249984741211,37.1841316223145,37.1729507446289,37.0474090576172,36.9610366821289,36.8970184326172,36.8898429870605,36.95263671875,36.9306144714355,36.9126472473145,36.8619651794434,36.7655296325684,36.6570320129395,36.229248046875,35.8793449401855,35.819091796875,35.8719711303711,36.1037101745605,36.2710418701172,36.5665054321289,36.6326675415039,36.6590843200684,36.6375999450684,36.5999526977539,36.571044921875,36.4737815856934,36.4291534423828,36.3830070495605,36.2678718566895,36.1128425598145,36.1756820678711,36.208984375,36.2345237731934,36.279296875,36.215087890625,36.145751953125,36.1668930053711,36.1898956298828,36.1160163879395,36.13818359375,36.133544921875,36.0679664611816,36.0281753540039,36.0153312683105,36.0752944946289,36.1480979919434,36.2291488647461,36.13330078125,36.0334968566895,35.9576148986816,35.9436531066895,35.9560546875,35.952880859375,35.9670906066895,35.9912109375,35.9703140258789,35.878662109375,35.7875480651855,35.6905288696289,35.691162109375,35.72216796875,35.8959465026855,35.9601554870605,35.9557609558105,35.89990234375,35.84326171875,35.7654800415039,35.64697265625,35.5083999633789,35.3802757263184,35.3541526794434,35.3690414428711,35.4012718200684,35.4077644348145,35.3970222473145,35.4364776611328,35.5084457397461,35.5323257446289,35.5296859741211,35.4532241821289,35.4314918518066,35.4630851745605,35.5273933410645,35.4583969116211,35.3296852111816,35.3056182861328,35.2704086303711,35.2151832580566,35.1529808044434,35.1041488647461,35.0733375549316,34.9903335571289,35.0049781799316,35.1546401977539,35.0251960754395,34.9702644348145,34.9409675598145,34.9893531799316,34.9365234375,34.8429183959961,34.7772483825684,34.7699203491211,34.7521476745605,34.7069816589355,34.7041511535645,34.7014656066895,34.6973648071289,34.7079086303711,34.6156234741211,34.6029319763184,34.6202659606934,34.6943855285645,34.7308120727539,34.5921401977539,34.5547828674316,34.526611328125,34.4513664245605,34.3575172424316,34.3319816589355,34.2849617004395,34.149169921875,34.0501441955566,33.9397468566895,33.9894523620605,34.0731430053711,34.1689949035645,33.993408203125,33.9118156433105,33.9175796508789,33.8732414245605,33.7240715026855,33.65869140625,33.4059104919434,33.244140625,33.3121566772461,33.3631820678711,33.4048843383789,33.3154296875,33.1851577758789,33.1353988647461,33.0425300598145,33.0272941589355,33.0008773803711,32.9092750549316,32.8248023986816,32.7873992919922,32.81005859375,32.7287139892578,32.6671371459961,32.6199226379395,32.589111328125,32.5928688049316,32.5765113830566,32.537353515625,32.500732421875,32.5213394165039,32.53369140625,32.51171875,32.4753913879395,32.4227561950684,32.351806640625,32.3244132995605,32.2873039245605,32.2928237915039,32.3262710571289,32.3959465026855,32.381103515625,32.3486328125,32.3370590209961,32.4480476379395,32.3633804321289,32.29833984375,32.2653312683105,32.2458992004395,32.2157211303711,32.1421890258789,32.1258277893066,32.1139183044434,32.068603515625,32.0295867919922,31.9449214935303,31.8920421600342,31.8940906524658,31.8786144256592,31.8409175872803,31.8134784698486,31.7879867553711,31.7533683776855,31.7437000274658,31.7041988372803,31.6669425964355,31.6461410522461,31.6103038787842,31.5743141174316,31.5389175415039,31.5284671783447,31.5389175415039,31.5312976837158,31.4721183776855,31.43603515625,31.3711929321289,31.3532733917236,31.3323249816895,31.3137702941895,31.2639179229736,31.1718730926514,31.1794452667236,31.19970703125,31.1270503997803,31.0882816314697,31.0090351104736,30.9137706756592,30.8746585845947,30.8018074035645,30.7314453125,30.6407718658447,30.2699699401855,30.1412124633789,29.7937965393066,29.4569835662842,29.0498523712158,28.556396484375,28.486083984375,28.4264659881592,28.3646984100342,28.2715816497803,28.1808586120605,28.070068359375,27.9006843566895,27.9344730377197,28.1775875091553,28.3203620910645,28.5229015350342,28.5180168151855,28.4521980285645,28.3749027252197,28.344970703125,28.462890625,28.5162105560303,28.5785160064697,28.6009273529053,28.6328125,28.6829586029053,28.7324714660645,28.7589340209961,28.7576637268066,28.6355953216553,28.5606441497803,28.3810043334961,28.2721672058105,28.180908203125,27.2070331573486,27.0830078125,26.9939460754395,26.8077163696289,26.568603515625,26.131591796875,25.83349609375,25.8426265716553,25.8740234375,25.8783206939697,25.7417488098145,25.6185531616211,25.4271011352539,25.3312511444092,25.2298336029053,25.232421875,25.1563472747803,25.1761703491211,25.1332511901855,25.1380367279053,25.228515625,25.2689952850342,25.3096675872803,25.3191413879395,25.2243156433105,25.2642097473145,25.3116703033447,25.338134765625,25.3672370910645,25.5833988189697,25.7318363189697,25.8310546875,25.8915538787842,25.983154296875,26.1460933685303,26.4350090026855,26.4674797058105,26.4899406433105,26.59716796875,26.6870594024658,26.6646957397461,26.6314449310303,26.5520515441895,26.5398445129395,26.5520515441895,26.704345703125,26.8915519714355,26.9615726470947,26.9634265899658,26.936767578125,26.8743648529053,26.8400897979736,26.8488750457764,26.8708000183105,26.9357414245605,27.0596675872803,27.4010753631592,27.44921875,27.4996109008789,27.5152835845947,27.5245609283447,27.6782703399658,27.7711429595947,27.8354015350342,27.8628902435303,27.90283203125,27.867919921875,27.8778820037842,27.9584465026855,27.981201171875,27.9637699127197,27.8977546691895,27.8732414245605,27.7772464752197,27.7459964752197,27.7184085845947,27.7331047058105,27.7093753814697,27.734375,27.7765617370605,27.8459968566895,28.23681640625,28.48583984375,28.7699222564697,28.81201171875,28.8874988555908,29.0515632629395,29.4519062042236,29.9259757995605,30.1038093566895,30.0647468566895,30.0290050506592,29.9822769165039,29.9293937683105,29.9073734283447,29.91015625,29.8978519439697,29.7730464935303,29.7776374816895,29.7453136444092,29.7210941314697,29.7079105377197,29.6802253723145,29.6952133178711,29.7675800323486,29.8424797058105,29.7850589752197,29.7401351928711,29.7581062316895,29.7978515625,29.875732421875,29.9757804870605,30.1219234466553,30.1170902252197,30.1483898162842,30.189453125,30.2369136810303,30.2867660522461,30.279296875,30.244384765625,30.2012691497803,30.1669921875,30.1719722747803,30.2144031524658,30.3325176239014,30.3991203308105,30.4291000366211,30.4058094024658,30.4415512084961,30.4642581939697,30.4930171966553,30.4820785522461,30.49560546875,30.4670906066895,30.4247055053711,30.4028797149658,30.3723621368408,30.3392562866211,30.3742198944092,30.3966808319092,30.4308586120605,30.5019035339355,30.5703105926514,30.5539054870605,30.5004386901855,30.5389633178711,30.5387687683105,30.4537105560303,30.3966808319092,30.3392562866211,30.2942867279053,30.3092784881592,30.3638172149658,30.3941402435303,30.3681144714355,30.2647457122803,30.2309093475342,30.25439453125,30.2590827941895,30.2917976379395,30.3468761444092,30.4074211120605,30.4141597747803,30.4496593475342,30.5615234375,30.6269035339355,30.6941909790039,30.6812515258789,30.5662097930908,30.4153308868408,30.3666019439697,30.3631820678711,30.3734855651855,30.3553714752197,30.4064922332764,30.4163074493408,30.4151363372803,30.3682613372803,30.3323707580566,30.3436508178711,30.3453121185303,30.2231426239014,30.1659679412842,30.2687511444092,30.3514175415039,30.3690910339355,30.3792953491211,30.2775859832764,30.1403312683105,30.0650882720947,30.0291004180908,30.0592765808105,30.1258773803711,30.1236820220947,30.1372051239014,30.1719722747803,30.14453125,30.1170387268066,30.0783195495605,30.0457057952881,30.0072746276855,29.9298343658447,29.90380859375,29.9150409698486,30.0021018981934,30.0581531524658,30.0460472106934,30.0108890533447,29.9776859283447,29.9209957122803,29.8397960662842,29.8202133178711,29.7843742370605,29.772216796875,29.7252941131592,29.6980476379395,29.6741218566895,29.6836910247803,29.6748523712158,29.6460437774658,29.6192855834961,29.5386734008789,29.4860363006592,29.4200706481934,29.3332042694092,29.335693359375,29.2482433319092,29.2181625366211,29.202880859375,29.1427230834961,29.0986824035645,29.0461406707764,29.0165996551514,29.0540027618408,29.0811042785645,28.9986820220947,28.9813461303711,29.0702133178711,29.1050300598145,29.1941413879395,29.249267578125,29.2675285339355,29.3023929595947,29.3165035247803,29.3128910064697,29.3332042694092,29.380615234375,29.4161109924316,29.4580078125,29.5371570587158,29.50439453125,29.4797344207764,29.4633293151855,29.4313983917236,29.3368167877197,29.2967777252197,29.23974609375,29.1817893981934,29.1360836029053,29.1049327850342,29.1310062408447,29.2558116912842,29.2951183319092,29.2997570037842,29.2715339660645,29.1506366729736,29.130859375,29.1935062408447,29.2889652252197,29.3207530975342,29.3309574127197,29.3179206848145,29.3506851196289,29.4573249816895,29.5054683685303,29.5642070770264,29.5628890991211,29.5135746002197,29.5553722381592,29.6053218841553,29.7460956573486,29.7506809234619,29.8360347747803,29.80029296875,29.7607421875,29.699462890625,29.6676750183105,29.6171894073486,29.5928230285645,29.5568351745605,29.5970706939697,29.6346683502197,29.7141609191895,29.7789554595947,29.7894058227539,29.7765617370605,29.7699222564697,29.7526874542236,29.7251472473145,29.7556171417236,29.8100090026855,29.81884765625,29.850830078125,29.9140625,29.9522953033447,29.9772453308105,29.979736328125,29.8149909973145,29.7226581573486,29.6893577575684,29.67041015625,29.4845237731934,29.3842754364014,29.4180183410645,29.5479488372803,29.5678215026855,29.5353507995605,29.5478534698486,29.6552734375,29.7499980926514,29.7525882720947,29.6769542694092,29.6801776885986,29.7125968933105,29.7023448944092,29.5309581756592,29.46044921875,29.3705577850342,29.2594718933105,29.1678199768066,29.0792465209961,28.9638671875,28.8984375,28.74462890625,28.7117176055908,28.6403312683105,28.5868167877197,28.5018558502197,28.4889640808105,28.5608882904053,28.6319332122803,28.6222171783447,28.6551265716553,28.6570320129395,28.631103515625,28.594482421875,28.6482887268066,28.6844730377197,28.7157230377197,28.7232913970947,28.7087879180908,28.4887199401855,28.4792003631592,28.4573249816895,28.4060554504395,28.3671379089355,28.34130859375,28.421630859375,28.3208503723145,28.22021484375,28.1943836212158,28.1575698852539,28.1853504180908,28.2242679595947,28.1895523071289,28.1634750366211,28.1582527160645,28.1443367004395,28.1026382446289,28.0607433319092,28.0938472747803,27.986083984375,27.8795909881592,27.8544445037842,27.8700199127197,27.8593254089355,27.8372077941895,27.6706066131592,27.4193344116211,27.3282718658447,27.3166027069092,27.31396484375,27.3949222564697,27.45751953125,27.2871570587158,27.2374019622803,27.1729488372803,27.1178703308105,27.05322265625,26.9673328399658,26.9075202941895,26.7596187591553,26.6917476654053,26.48583984375,26.3965339660645,26.06787109375,26.0653324127197,26.0297374725342,25.9614734649658,25.9416007995605,25.9111824035645,25.884765625,25.8705081939697,25.871826171875,25.8908214569092,25.9841785430908,26.0420417785645,26.064453125,26.1111812591553,26.182373046875,26.2245616912842,26.2378425598145,26.2764644622803,26.3404293060303,26.3812503814697,26.3989753723145,26.4469242095947,26.5641593933105,26.56591796875,26.7619132995605,26.8847179412842,27.0366706848145,27.056640625,27.0566902160645,27.0816898345947,27.1701164245605,27.2336902618408,27.2854976654053,27.34033203125,27.3980464935303,27.4673843383789,27.54833984375,27.6358871459961,27.7299308776855,27.8672847747803,28.0478515625,28.1729488372803,28.2426280975342,28.3276863098145,28.4281253814697,28.4864253997803,28.5025386810303,28.6142082214355,28.8213386535645,28.9728012084961,29.0685539245605,29.1825199127197,29.314697265625,29.4006824493408,29.4603023529053,29.4604015350342,29.6340808868408,29.7425785064697,29.77685546875,29.7731437683105,29.7835445404053,29.8080558776855,29.8092308044434,29.7869644165039,29.7824687957764,29.7957038879395,29.8252429962158,29.8711910247803,29.8649921417236,29.8066425323486,29.7690925598145,29.7523441314697,29.6439456939697,29.4439449310303,29.3153324127197,29.2580070495605,29.2164077758789,29.1903800964355,29.1322250366211,29.0418968200684,28.9981937408447,29.0011234283447,29.0707015991211,29.2068862915039,29.2910652160645,29.3231468200684,29.3861351013184,29.4798812866211,29.5424327850342,29.5737323760986,29.6776847839355,29.8542957305908,29.9905281066895,30.1343746185303,30.4476547241211,30.5833511352539,30.6459445953369,30.7205543518066,30.8072738647461,30.9807605743408,31.2410163879395,31.3977546691895,31.450927734375,31.5446796417236,31.6790046691895,31.7644538879395,31.7684078216553,31.7701663970947,31.7713394165039,31.7724628448486,31.7736339569092,31.7747573852539,31.7758769989014,31.7770519256592,31.7781715393066,31.7793464660645,31.6668472290039,31.5543956756592,31.44189453125,31.3294429779053,31.3288078308105,31.328125,31.3274421691895,31.3268051147461,31.3261699676514,31.3255348205566,31.3248519897461,31.3242168426514,31.4722652435303,31.6202144622803,31.7682609558105,31.9162578582764,32.0643043518066,32.2123069763184,32.3603019714355,32.50830078125,32.5647964477539,32.71533203125,32.7047348022461,32.6833038330078,32.661865234375,32.6404762268066,32.6190452575684,32.5976066589355,32.5762214660645,32.5547866821289,32.5333480834961,32.5397453308105,32.649169921875,32.6878929138184,32.6640129089355,32.8062477111816,32.8733901977539,32.9388656616211,33.1000480651855,33.2955093383789,33.5384750366211,33.61962890625,33.72216796875,33.7506828308105,33.7585906982422,33.7123031616211,33.7439460754395,33.8582992553711,34.0173797607422,34.0350112915039,34.0244598388672,34.1120109558105,34.1641120910645,34.2574234008789,34.3385734558105,34.4180183410645,34.399658203125,34.4119606018066,34.4692878723145,34.4764671325684,34.4595718383789,34.4716300964355,34.5438957214355,34.5799789428711,34.6689453125,34.7493667602539,34.8119621276855,34.9492683410645,35.0764656066895,35.1224098205566,35.1576652526855,35.2096672058105,35.2749519348145,35.3654289245605,35.4250984191895,35.4807624816895,35.6071281433105,35.6763191223145,35.792236328125,35.8638687133789,35.9273910522461,36.154052734375,36.3310546875,36.4329071044922,36.5723648071289,36.657470703125,36.7322769165039,36.8009757995605,36.8512191772461,36.9389152526855,36.990966796875,37.20751953125,37.3731460571289,37.5426254272461,37.6527824401855,37.77197265625,37.7979965209961,37.7885284423828,37.7410621643066,37.6558609008789,37.5918464660645,37.5639152526855,37.5016593933105,37.4828109741211,37.478271484375,37.5182113647461,37.5437965393066,37.62646484375,37.7320327758789,37.7903327941895,37.8965797424316,37.9211921691895,37.9605979919434,38.00732421875,38.0406265258789,38.0496063232422,38.0340843200684,38.061279296875,38.0523948669434,38.055908203125,38.0839347839355,38.0748023986816,38.0804710388184,38.0749969482422,38.0868148803711,38.1201171875,38.12353515625,38.0655288696289,38.0725555419922,38.1358909606934,38.1448249816895,38.1088371276855,37.9535636901855,37.8381843566895,37.826416015625,37.8740730285645,37.90234375,37.9456558227539,38.0260734558105,38.0554695129395,37.9886245727539,38.019287109375,38.0970230102539,38.2273445129395,38.1233406066895,38.1965827941895,38.277099609375,38.3050765991211,38.4492645263672,38.5358390808105,38.6756362915039,38.9072761535645,39.1109886169434,39.368408203125,39.5149421691895,39.6187019348145,39.7754859924316,39.8607940673828,40.0945320129395,40.251953125,40.37109375,40.4912109375,40.5980949401855,40.7105484008789,40.7402839660645,40.7278823852539,40.7039031982422,40.6964836120605,40.74609375,40.771728515625,40.7750473022461,40.7907218933105,40.8220710754395,40.9697761535645,41.1559104919434,41.3841819763184,41.4595222473145,41.6217269897461,41.718994140625,41.7879409790039,41.8885726928711,41.984619140625,42.1229019165039,42.3043441772461,42.3810043334961,42.5836906433105,42.6702117919922,42.8128890991211,42.9368629455566,43.0123558044434,43.3416519165039,43.3682136535645,43.3673858642578,43.42333984375,43.4363746643066,43.4097175598145,43.5400390625,43.6515617370605,43.6917457580566,44.0556640625,44.3337898254395,44.4254875183105,44.5200691223145,44.6482429504395,44.7777328491211,45.4008293151855,45.47607421875,45.5769538879395,45.8429679870605,46.1405754089355,46.1783218383789,46.2193832397461,46.2254409790039,46.1821784973145,46.1826171875,46.22265625,46.2094230651855,46.1549797058105,46.1439933776855,46.1536140441895,46.1672821044922,46.1708488464355,46.2209968566895,46.2710914611816,46.2677268981934,46.2998542785645,46.2677726745605,46.3007316589355,46.2794418334961,46.3729019165039,46.4905242919922,46.6050796508789,46.5213890075684,46.4325675964355,46.5333480834961,46.6600112915039,46.7086944580078,46.7447776794434,46.8626976013184,46.9631805419922,46.9844703674316,47.0296859741211,47.0352058410645,47.0003433227539,46.9546890258789,47.0153312683105,47.086669921875,47.2085456848145,47.4045906066895,47.6586418151855,47.7842292785645,47.9041481018066,47.97412109375,48.1516571044922,48.285888671875,48.38037109375,48.3750495910645,48.30078125,48.242431640625,48.2000007629395,48.16845703125,48.1195297241211,48.1242179870605,48.154541015625,48.1509284973145,48.0815925598145,48.0732917785645,48.076904296875,48.0900421142578,48.1375999450684,48.1200180053711,48.0759773254395,48.0132331848145,47.9317893981934,47.8811531066895,47.7384300231934,47.7353515625,47.7836418151855,47.7931632995605,47.5519561767578,47.437744140625,47.3860816955566,47.3558082580566,47.348388671875,47.3602027893066,47.4076652526855,47.41796875,47.4010734558105,47.399658203125,47.4536628723145,47.4793472290039,47.5593757629395,47.6073760986328,47.6585464477539,47.7005386352539,47.7621078491211,47.8354988098145,47.85595703125,47.9164085388184,47.9278831481934,47.9197273254395,47.8157234191895,47.7693367004395,47.7127914428711,47.6928253173828,47.6156272888184,47.608154296875,47.6172370910645,47.6123542785645,47.5284194946289,47.4631843566895,47.4049301147461,47.2931671142578,47.2746086120605,47.2814445495605,47.31640625,47.3051300048828,47.2183570861816,47.2259788513184,47.3289566040039,47.3365707397461,47.2896461486816,47.2445793151855,47.1725578308105,47.138916015625,47.1314964294434,47.14599609375,47.11181640625,47.1108894348145,47.1442375183105,47.1669921875,47.2755851745605,47.2950210571289,47.2958030700684,47.3121109008789,47.37158203125,47.3952140808105,47.5283699035645,47.6039047241211,47.6278305053711,47.6526374816895,47.6767578125,47.7164573669434,47.7523422241211,47.7842750549316,47.820556640625,47.8986358642578,47.9330558776855,48.0107421875,48.0420417785645,48.0801734924316,48.1138191223145,48.1663589477539,48.1839332580566,48.1759262084961,48.0899429321289,48.0841331481934,48.1304664611816,48.15966796875,48.1993179321289,48.2291030883789,48.2584953308105,48.2691879272461,48.2938957214355,48.3743171691895,48.4109344482422,48.4286651611328,48.4333000183105,48.4463882446289,48.4652328491211,48.489990234375,48.4979019165039,48.4879875183105,48.5055656433105,48.5374984741211,48.55517578125,48.66943359375,48.7623062133789,48.7779769897461,48.7795906066895,48.76708984375,48.7638664245605,48.7942886352539,48.85302734375,48.9930191040039,48.9930191040039,48.9930191040039,48.9930191040039,48.9930191040039,48.9930191040039,48.9930191040039,48.9930191040039,48.9930191040039,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9930648803711,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931144714355,48.9931640625,48.9931640625,48.9931640625,48.9931640625,48.9931640625,48.9931640625,48.9931640625,48.9931640625,48.9931640625,48.9917488098145,49.2030792236328,49.3696746826172,49.3494148254395,49.3190422058105,49.3045921325684,49.2585945129395,49.1191902160645,49.0029296875,48.8634300231934,48.8629875183105,48.8084983825684,48.7744140625,48.7426261901855,48.7041015625,48.6590309143066,48.6072769165039,48.5489273071289,48.5254402160645,48.5369148254395,48.561279296875,48.6165504455566,48.6288566589355,48.6253433227539,48.619873046875,48.61181640625,48.5677757263184,48.5318374633789,48.465087890625,48.4353523254395,48.3658714294434,48.276611328125,48.276611328125,48.3289070129395,48.33837890625,48.3018531799316,48.1975593566895,48.1045875549316,48.0583000183105,48.0585479736328,48.1045875549316,48.1937026977539,48.2091331481934,48.2005348205566,48.1310539245605,48.1045875549316,48.1125984191895,48.0991706848145,48.1181144714355,48.0781745910645,48.0153312683105,47.9954605102539,48.0153312683105,47.9999008178711,47.9962425231934,48.0199699401855,48.0474128723145,48.0937995910645,48.1557121276855,48.2640113830566,48.3030776977539,48.2253913879395,48.1568832397461,48.13037109375,48.060546875,47.9617652893066,47.8484878540039,47.7352066040039,47.6364288330078,47.5666007995605,47.5400886535645,47.4600563049316,47.3800277709961,47.2999992370605,47.219970703125,47.1399421691895,47.0599632263184,46.9799346923828,46.89990234375,46.766845703125,46.6373023986816,46.5432624816895,46.4573707580566,46.4618682861328,46.4981460571289,46.515625,46.5185050964355,46.549560546875,46.5427742004395,46.5272445678711,46.5029296875,46.4835929870605,46.4447746276855,46.3708000183105,46.2886238098145,46.2265167236328,46.1470222473145,46.0849113464355,46.0729026794434,46.1090812683105,46.1227531433105,46.1168479919434,46.0586891174316,46.0237312316895,45.9946784973145,45.817138671875,45.7290534973145,45.6327629089355,45.5179672241211,45.4477081298828,45.3473625183105,45.2043952941895,45.083740234375,44.91552734375,44.7439460754395,44.572998046875,44.3915557861328,44.1922340393066,44.0153350830078,43.8222160339355,43.5708999633789,43.4740715026855,43.2632293701172,43.0726547241211,43.0173835754395,42.739501953125,42.6247062683105,42.5580558776855,42.4934577941895,42.3852043151855,42.3317375183105,42.30029296875,42.2506828308105,42.1419410705566,41.9758796691895,41.8329582214355,41.7530288696289,41.6751937866211,41.6748542785645,41.7787094116211,41.8887214660645,41.98681640625,42.1034698486328,42.2091789245605,42.2471694946289,42.2997550964355,42.3660125732422,42.4414520263672,42.5389633178711,42.6514663696289,42.74853515625,42.8023452758789,42.8637199401855,42.9091300964355,42.9352035522461,42.9613304138184,42.9806137084961,42.9970207214355,43.017333984375,43.061767578125,43.0873069763184,43.1061019897461,43.278076171875,43.3313980102539,43.466552734375,43.5271492004395,43.5833473205566,43.6249504089355,43.6314926147461,43.6306648254395,43.6295394897461,43.6286125183105,43.6274909973145,43.6268577575684,43.6288108825684,43.7848167419434,43.92431640625,44.0576171875,44.214111328125,44.2422370910645,44.303955078125,44.3625984191895,44.4169921875,44.468017578125,44.4970703125,44.7722663879395,44.8993644714355,44.9701156616211,45.00390625,44.9990730285645,45.0038566589355],[51.3726081848145,51.3722152709961,51.5350570678711,51.5867195129395,51.6438980102539,51.6559562683105,51.6155738830566,51.5276832580566,51.4699211120605,51.4208488464355,51.40087890625,51.3726081848145],[51.7919921875,51.7332992553711,51.7598648071289,51.840576171875,51.8611297607422,51.8028335571289,51.7919921875],[51.8123550415039,51.7904815673828,51.8348159790039,51.8862800598145,51.8433113098145,51.8123550415039],[51.6497077941895,51.6164054870605,51.6173820495605,51.672607421875,51.6912612915039,51.7174835205078,51.73779296875,51.7642593383789,51.8010749816895,51.8482437133789,51.8822250366211,51.9030265808105,51.9158668518066,51.91845703125,51.8604011535645,51.8400382995605,51.8262710571289,51.8016624450684,51.7778816223145,51.6858901977539,51.6497077941895],[51.7167472839355,51.7037124633789,51.6935577392578,51.704833984375,51.7012710571289,51.6941909790039,51.6765632629395,51.7010726928711,51.72119140625,51.7762222290039,51.804931640625,51.8412590026855,51.909423828125,51.9297866821289,51.9287605285645,51.9019050598145,51.8665542602539,51.8357925415039,51.8069343566895,51.7167472839355],[51.86669921875,51.8332023620605,51.83740234375,51.8201141357422,51.7542953491211,51.7356948852539,51.7311515808105,51.74560546875,51.7120590209961,51.6299324035645,51.6758804321289,51.6036643981934,51.7904815673828,51.8187484741211,51.8399391174316,51.8946762084961,51.9860343933105,51.9817848205566,51.9440422058105,51.9195823669434,51.86669921875],[51.916259765625,51.8991203308105,51.9425315856934,51.9677238464355,51.9946784973145,51.9775390625,51.9530296325684,51.916259765625],[51.9054222106934,51.8802261352539,51.89404296875,51.9328155517578,51.9795913696289,52.0304222106934,51.9668464660645,51.9054222106934],[52.00244140625,51.9729995727539,52.0042991638184,52.0298347473145,52.0560531616211,52.0994110107422,52.0999526977539,52.0823249816895,52.0494651794434,52.0289573669434,52.00244140625],[51.8828125,51.841064453125,51.8836898803711,51.9029273986816,51.9757804870605,51.9916000366211,52.0182113647461,52.1104965209961,52.1138191223145,52.10302734375,52.0597648620605,51.9938468933105,51.9475593566895,51.8828125],[52.1362800598145,52.0956535339355,52.1003875732422,52.0905265808105,52.0791511535645,52.0625,52.0679702758789,52.0456047058105,52.0415534973145,52.0626449584961,52.0481948852539,52.0536613464355,52.0721664428711,52.1036148071289,52.1233406066895,52.1312980651855,52.1042976379395,52.1183586120605,52.1437492370605,52.1362800598145],[52.2722625732422,52.2574691772461,52.2728500366211,52.3256340026855,52.3538093566895,52.3880348205566,52.3729476928711,52.32958984375,52.2722625732422],[52.0350112915039,51.993896484375,52.0221672058105,52.0382308959961,52.047119140625,52.0941925048828,52.1349601745605,52.1840324401855,52.2161598205566,52.26904296875,52.2959938049316,52.3172378540039,52.3419418334961,52.3779296875,52.420166015625,52.3672332763672,52.331787109375,52.289794921875,52.2459945678711,52.223388671875,52.2003440856934,52.1352043151855,52.0777816772461,52.0350112915039],[52.3595695495605,52.3566398620605,52.3912582397461,52.40478515625,52.4376487731934,52.451416015625,52.5041007995605,52.4951171875,52.4466323852539,52.3595695495605],[52.5814933776855,52.5497550964355,52.5615234375,52.6007308959961,52.6312484741211,52.6975593566895,52.68505859375,52.6675796508789,52.6424331665039,52.609619140625,52.5931167602539,52.5929679870605,52.5814933776855],[52.8473625183105,52.80712890625,52.7923355102539,52.8137702941895,52.8060073852539,52.829833984375,52.8510284423828,52.8833999633789,52.8836402893066,52.8667449951172,52.8473625183105],[53.0129890441895,52.9802742004395,52.99560546875,52.9426727294922,52.85205078125,52.8347625732422,52.8248519897461,52.825927734375,52.810791015625,52.814453125,52.7520980834961,52.7969245910645,52.8855476379395,52.9074211120605,52.9378890991211,53.007568359375,53.0129890441895],[53.3451156616211,53.238037109375,53.1597671508789,53.0844230651855,53.0431632995605,53.0360832214355,53.01611328125,52.9634284973145,52.9568862915039,52.8339347839355,52.83203125,52.8641624450684,52.90966796875,52.951171875,53.0197257995605,53.044921875,53.079345703125,53.1487808227539,53.175048828125,53.2272491455078,53.2557601928711,53.265625,53.2568855285645,53.283447265625,53.3035659790039,53.3219261169434,53.3538093566895,53.4087905883789,53.4575691223145,53.5001487731934,53.5333023071289,53.5569801330566,53.5582046508789,53.5079574584961,53.4849624633789,53.4345703125,53.3873062133789,53.3451156616211],[53.7232894897461,53.7204093933105,53.7451667785645,53.7677764892578,53.7841796875,53.8224601745605,53.8361320495605,53.8430671691895,53.8328132629395,53.7874031066895,53.7568855285645,53.7232894897461],[53.9009246826172,53.8534660339355,53.883544921875,53.9248046875,53.9781227111816,53.9989738464355,53.9708976745605,53.9326171875,53.873779296875,53.7854995727539,53.7264671325684,53.7009773254395,53.7205085754395,53.7176742553711,53.697509765625,53.6735343933105,53.6518058776855,53.6096649169922,53.5366706848145,53.4760246276855,53.452880859375,53.4473648071289,53.4078598022461,53.37060546875,53.3504867553711,53.3409652709961,53.3419914245605,53.32568359375,53.2919921875,53.2762184143066,53.272705078125,53.2594261169434,53.2599601745605,53.3002433776855,53.3237800598145,53.3708953857422,53.3865737915039,53.3937034606934,53.437255859375,53.4949722290039,53.5264663696289,53.6359405517578,53.6545867919922,53.6983909606934,53.6971206665039,53.6480484008789,53.641357421875,53.6461448669434,53.6853981018066,53.7129402160645,53.733154296875,53.7585945129395,53.7705535888672,53.7691421508789,53.7833976745605,53.8133811950684,53.8431167602539,53.8726081848145,53.9056625366211,53.9421882629395,53.9629402160645,53.9778823852539,54.002197265625,54.0059585571289,53.9956550598145,53.9009246826172],[54.0706520080566,54.0530281066895,54.0491676330566,54.0591773986816,54.0471687316895,54.0543441772461,54.1139678955078,54.1448249816895,54.1691398620605,54.1912612915039,54.2110328674316,54.2069816589355,54.152099609375,54.1199226379395,54.0999031066895,54.0811042785645,54.0706520080566],[54.13671875,54.129150390625,54.1395530700684,54.1835441589355,54.2533187866211,54.2786636352539,54.2845191955566,54.2738800048828,54.2218780517578,54.2080078125,54.1968269348145,54.180908203125,54.13671875],[54.4013671875,54.3795433044434,54.4022979736328,54.4443855285645,54.4945297241211,54.4620590209961,54.4466323852539,54.4013671875],[54.8470191955566,54.8423805236816,54.8729972839355,54.8958969116211,54.9315452575684,54.9828605651855,54.9834976196289,54.9547348022461,54.9320335388184,54.8671875,54.8470191955566],[54.980712890625,54.8155250549316,54.783203125,54.7655754089355,54.7477531433105,54.7232894897461,54.6860847473145,54.6689949035645,54.7356948852539,54.6383323669434,54.6256828308105,54.6209945678711,54.60302734375,54.5713348388672,54.482421875,54.4478492736816,54.4273414611816,54.4043464660645,54.407470703125,54.4190902709961,54.4613761901855,54.5447731018066,54.5679702758789,54.6078147888184,54.6629409790039,54.6919937133789,54.880859375,54.9068336486816,54.9131851196289,54.9000511169434,54.9551277160645,55.0391120910645,55.049072265625,55.0508308410645,55.037841796875,55.0143089294434,54.980712890625],[54.972412109375,54.9673347473145,54.9781265258789,55.0349578857422,55.0587882995605,55.0408668518066,54.99951171875,54.972412109375],[55.15185546875,55.0721664428711,55.0596199035645,55.0809097290039,55.0573234558105,55.0745620727539,55.1023445129395,55.1239738464355,55.1256866455078,55.1653289794922,55.1820297241211,55.2177238464355,55.2259750366211,55.15185546875],[55.079833984375,54.9495086669922,54.9037590026855,54.8877449035645,54.894287109375,54.9093246459961,54.9305686950684,54.9964866638184,55.0352516174316,55.0484886169434,55.0256843566895,55.0907211303711,55.1374015808105,55.2008323669434,55.248779296875,55.2641105651855,55.2627410888672,55.2133293151855,55.079833984375],[55.1287574768066,55.1068382263184,55.0789566040039,55.0671882629395,55.0444793701172,54.9416999816895,54.9227066040039,55.0104484558105,55.038330078125,55.056884765625,55.123046875,55.1201667785645,55.1338882446289,55.1925277709961,55.19775390625,55.2675323486328,55.2729988098145,55.2635726928711,55.2212905883789,55.182373046875,55.1446762084961,55.1287574768066],[54.8944358825684,54.8809547424316,54.8904304504395,54.9260711669922,54.9190444946289,54.9070816040039,54.8924293518066,54.8404769897461,54.7561531066895,54.7262229919434,54.6841773986816,54.7091331481934,54.7626457214355,54.8548355102539,54.9213409423828,54.9494132995605,54.9698219299316,55.08447265625,55.1751441955566,55.1854972839355,55.210693359375,55.2603530883789,55.3038101196289,55.3257293701172,55.2137184143066,55.1662101745605,55.110595703125,55.0330047607422,55.0025863647461,54.8944358825684],[55.3376922607422,55.2587890625,55.3089828491211,55.3338394165039,55.3523445129395,55.3569831848145,55.3376922607422],[55.3147964477539,55.3142547607422,55.3219223022461,55.3782730102539,55.3807601928711,55.3633804321289,55.3076171875,55.2332038879395,55.1974105834961,55.1848602294922,55.1590347290039,55.1776351928711,55.1711921691895,55.1452140808105,55.1739730834961,55.3113288879395,55.3353996276855,55.3831062316895,55.4046401977539,55.3983383178711,55.37939453125,55.3594245910645,55.3147964477539],[55.5437507629395,55.515625,55.4978523254395,55.4314460754395,55.417724609375,55.3766593933105,55.3616714477539,55.3172378540039,55.269287109375,55.413330078125,55.4969215393066,55.5392570495605,55.5271987915039,55.5223121643066,55.55908203125,55.5437507629395],[55.8211936950684,55.78955078125,55.7918472290039,55.8021965026855,55.8297843933105,55.9130859375,55.92431640625,55.9210929870605,55.8866691589355,55.8211936950684],[55.4891586303711,55.3792953491211,55.2667961120605,55.206298828125,55.2197761535645,55.2685050964355,55.2658195495605,55.27587890625,55.3163070678711,55.368408203125,55.4087905883789,55.3734893798828,55.341064453125,55.2989273071289,55.2183609008789,55.1658210754395,55.2230987548828,55.3586883544922,55.416259765625,55.5030784606934,55.5855484008789,55.8316917419434,55.9553718566895,55.948974609375,55.8566398620605,55.7276382446289,55.6695327758789,55.5680198669434,55.4891586303711],[56.3392066955566,56.3177719116211,56.31982421875,56.278564453125,56.235107421875,56.1941909790039,56.1558570861816,56.1287117004395,56.0998039245605,56.0900382995605,55.9950218200684,55.89501953125,55.8424797058105,55.798095703125,55.68701171875,55.55810546875,55.5188484191895,55.4806175231934,55.4791488647461,55.5074729919434,55.5939445495605,55.5904808044434,55.543701171875,55.5026359558105,55.4731941223145,55.46435546875,55.48291015625,55.3986320495605,55.383544921875,55.322998046875,55.2998046875,55.2549819946289,55.2367668151855,55.2244148254395,55.218017578125,55.2306137084961,55.2085914611816,55.0338401794434,54.969482421875,54.9402351379395,54.9014167785645,54.8686027526855,54.8349151611328,54.8048324584961,54.726318359375,54.713134765625,54.7125473022461,54.73486328125,54.8023452758789,54.9072265625,54.9222183227539,54.9379386901855,54.9503898620605,54.9525871276855,54.9957504272461,55.0523414611816,55.1100578308105,55.1359367370605,55.1467781066895,55.1305160522461,55.0739250183105,55.0300788879395,55.0484886169434,55.1884765625,55.3009262084961,55.32763671875,55.3602523803711,55.3775367736816,55.3761711120605,55.3955574035645,55.5041007995605,55.534912109375,55.5896987915039,55.6125984191895,55.5954093933105,55.6068840026855,55.6508293151855,55.68896484375,55.6958999633789,55.691162109375,55.748779296875,55.78515625,55.8038101196289,55.8365249633789,55.8319320678711,55.7970199584961,55.79833984375,55.8446273803711,55.886474609375,55.9207992553711,55.9570808410645,56.0186996459961,56.035888671875,55.999267578125,55.9427719116211,55.96484375,55.9994659423828,56.0936508178711,56.1456527709961,56.176513671875,56.2163581848145,56.3162574768066,56.3392066955566],[56.2472686767578,56.244140625,56.2594261169434,56.3409194946289,56.3648452758789,56.3919906616211,56.4417991638184,56.4537620544434,56.448486328125,56.4351539611816,56.4119148254395,56.3882827758789,56.3643569946289,56.3393058776855,56.3131866455078,56.2873039245605,56.2472686767578],[56.109375,55.9432601928711,55.95263671875,55.9529762268066,55.9293937683105,55.9397430419922,55.9582023620605,55.9795417785645,56.0288581848145,56.0563468933105,56.06640625,56.0781745910645,56.1300773620605,56.1981925964355,56.2236328125,56.2416496276855,56.3241691589355,56.3352546691895,56.498779296875,56.4874992370605,56.387939453125,56.2442359924316,56.109375],[56.435791015625,56.4121589660645,56.4202651977539,56.4398918151855,56.4711418151855,56.5021476745605,56.5613288879395,56.6005363464355,56.5981903076172,56.5731925964355,56.5214347839355,56.435791015625],[56.514892578125,56.5126953125,56.5390167236328,56.5706062316895,56.6087417602539,56.6029281616211,56.5816421508789,56.5578117370605,56.5438957214355,56.5295906066895,56.514892578125],[56.6350593566895,56.6151390075684,56.6281242370605,56.6177253723145,56.5940437316895,56.5424308776855,56.5457000732422,56.6079597473145,56.6350593566895],[56.5256843566895,56.5110359191895,56.512451171875,56.5244636535645,56.5672378540039,56.6068344116211,56.6374053955078,56.6963844299316,56.7947731018066,56.6847152709961,56.6357421875,56.5758285522461,56.553466796875,56.5425300598145,56.5256843566895],[56.844970703125,56.7756843566895,56.7812995910645,56.7289085388184,56.6504402160645,56.60009765625,56.611328125,56.5821800231934,56.4854965209961,56.388671875,56.2921371459961,56.1936531066895,56.1277351379395,56.10791015625,56.1011199951172,56.1189918518066,56.1330070495605,56.077392578125,56.0769538879395,56.2032699584961,56.4135246276855,56.4563484191895,56.5134773254395,56.5800285339355,56.6170921325684,56.7240257263184,56.7494621276855,56.8386726379395,56.9181671142578,56.9323234558105,56.8982887268066,56.8691902160645,56.844970703125],[57.0035171508789,56.97216796875,57.0049819946289,57.0034675598145,56.9742698669434,56.930419921875,56.8802719116211,56.8504409790039,56.7825698852539,56.7130851745605,56.6770515441895,56.6472663879395,56.6237297058105,56.6207504272461,56.6832504272461,56.7958488464355,56.830078125,56.8185081481934,56.7862319946289,56.7256851196289,56.6892585754395,56.6448249816895,56.6111297607422,56.56689453125,56.5282249450684,56.4951667785645,56.4739761352539,56.464599609375,56.473876953125,56.4517593383789,56.464111328125,56.4840316772461,56.5167961120605,56.5962905883789,56.7100105285645,56.7975120544434,56.8766593933105,56.9243621826172,56.9670906066895,57.0095710754395,57.04345703125,57.0687026977539,57.0628433227539,57.0035171508789],[57.1248550415039,57.0925788879395,57.0939445495605,57.0286140441895,57.0004386901855,57.0519027709961,57.1319351196289,57.18505859375,57.1673355102539,57.1524429321289,57.1397476196289,57.1292495727539,57.1248550415039],[57.1839370727539,57.1367683410645,57.1542015075684,57.1885719299316,57.2030296325684,57.2417984008789,57.1839370727539],[57.3514175415039,57.24169921875,57.1565437316895,57.05419921875,56.84228515625,56.7621078491211,56.7183113098145,56.6034164428711,56.5787124633789,56.5560569763184,56.5232391357422,56.4356422424316,56.3024406433105,56.2274894714355,56.2161636352539,56.2407722473145,56.28125,56.323486328125,56.4568328857422,56.5189437866211,56.5636253356934,56.5961418151855,56.6163558959961,56.6704597473145,56.6790542602539,56.6669921875,56.66015625,56.7028312683105,56.7253913879395,56.8023414611816,56.8241195678711,56.8003425598145,56.8218727111816,56.8507843017578,56.8939933776855,56.9318351745605,57.02734375,57.044921875,57.0488777160645,57.0815925598145,57.1884307861328,57.2494125366211,57.2438468933105,57.0714340209961,57.0337409973145,57.009521484375,57.0575180053711,57.1003913879395,57.2304229736328,57.2804183959961,57.3172874450684,57.3325653076172,57.3543968200684,57.3899879455566,57.4247055053711,57.5165023803711,57.5343780517578,57.5331077575684,57.431640625,57.4167022705078,57.3514175415039],[57.85009765625,57.8223648071289,57.8294906616211,57.8619613647461,57.9093780517578,57.94189453125,57.9557609558105,57.9710464477539,57.9328117370605,57.8979034423828,57.85009765625],[57.8239250183105,57.7689971923828,57.7756805419922,57.832275390625,57.8488731384277,57.8514633178711,57.8256874084473,57.8059043884277,57.7823219299316,57.7344207763672,57.7033233642578,57.6460914611816,57.6148910522461,57.5977096557617,57.5769996643066,57.4822235107422,57.4601058959961,57.4547882080078,57.4718284606934,57.5028762817383,57.5081558227539,57.4980964660645,57.4689445495605,57.4603538513184,57.4534187316895,57.4108390808105,57.3795928955078,57.3451156616211,57.3309555053711,57.3206558227539,57.3207969665527,57.281982421875,57.2376480102539,57.2263679504395,57.1671867370605,57.1379890441895,57.1030769348145,57.0776863098145,57.0523452758789,57.0295906066895,57.0103530883789,56.9836921691895,56.9607429504395,56.8583488464355,56.7742195129395,56.7779808044434,56.7884788513184,56.8045425415039,56.8206558227539,56.989501953125,56.997802734375,57.0035171508789,57.0499496459961,57.0200691223145,57.021240234375,57.06103515625,57.1084976196289,57.121826171875,57.1336936950684,57.1407699584961,57.1430206298828,57.1317863464355,57.1070289611816,57.0965309143066,57.0994644165039,57.0868644714355,57.0633316040039,57.0361328125,57.0053215026855,56.9638175964355,56.9117698669434,56.9208984375,57.0365753173828,57.2059097290039,57.3353538513184,57.3662605285645,57.4460906982422,57.5594253540039,57.5904808044434,57.6380882263184,57.6524391174316,57.6512222290039,57.6307106018066,57.5873069763184,57.5668983459473,57.55615234375,57.5300750732422,57.4390144348145,57.3582000732422,57.3253402709961,57.3051300048828,57.3668479919434,57.4432640075684,57.5956039428711,57.6358413696289,57.6466789245605,57.6407241821289,57.6634292602539,57.7147445678711,57.7571792602539,57.790771484375,57.8198738098145,57.8628425598145,57.8750953674316,57.8803672790527,57.8712348937988,57.8578147888184,57.7610855102539,57.7310066223145,57.73095703125,57.7470245361328,57.7791519165039,57.7983856201172,57.8046913146973,57.7904777526855,57.7957496643066,57.8200187683105,57.86328125,57.8788604736328,57.9106483459473,57.9576187133789,57.9719734191895,57.93603515625,57.8967742919922,57.8239250183105],[58.2442398071289,58.1467819213867,58.1244125366211,58.0574684143066,58.0385284423828,58.0119094848633,57.9918441772461,58.008544921875,58.1023902893066,58.1221160888672,58.1241226196289,58.0958480834961,58.0510749816895,58.0153312683105,57.9527816772461,57.8172378540039,57.7936515808105,57.7960891723633,57.8158226013184,57.7686538696289,57.7325706481934,57.724365234375,57.6479988098145,57.5892105102539,57.4811515808105,57.5110321044922,57.5736312866211,57.6622543334961,57.6664047241211,57.6507301330566,57.5969696044922,57.4803733825684,57.4199256896973,57.4465827941895,57.5348625183105,57.674560546875,57.8399887084961,57.8730926513672,57.97216796875,58.0505828857422,58.0959968566895,58.1080055236816,58.1430625915527,58.2188949584961,58.157470703125,58.0984840393066,58.1539039611816,58.1981430053711,58.1965293884277,58.2058143615723,58.2471694946289,58.2685089111328,58.2442398071289],[58.2289047241211,58.2040977478027,58.20654296875,58.2431144714355,58.2909164428711,58.3124046325684,58.33251953125,58.2872085571289,58.2289047241211],[58.1616706848145,58.1388168334961,58.1439933776855,57.9945297241211,57.873779296875,57.7892074584961,57.7075157165527,57.628662109375,57.6175765991211,57.6707992553711,57.7784690856934,57.8206062316895,57.839599609375,57.8793449401855,57.982177734375,58.0111312866211,58.0379409790039,58.0491714477539,58.0447235107422,58.0343742370605,57.9634323120117,57.884521484375,57.7122573852539,57.6141624450684,57.5815887451172,57.4919891357422,57.4513626098633,57.3686981201172,57.3525390625,57.3367691040039,57.3000946044922,57.1467781066895,57.0569839477539,57.0425758361816,57.0575714111328,57.0919914245605,57.137939453125,57.1955070495605,57.2317390441895,57.4201622009277,57.4820327758789,57.5678176879883,57.6380882263184,57.7360343933105,57.9950218200684,58.0778274536133,58.1468772888184,58.2020988464355,58.2627906799316,58.3289566040039,58.3546371459961,58.3201637268066,58.2249984741211,58.1616706848145],[58.3602027893066,58.3521041870117,58.41162109375,58.4134788513184,58.3630867004395,58.374267578125,58.3123512268066,58.3066902160645,58.314208984375,58.3098640441895,58.2443351745605,58.1779327392578,58.1611289978027,58.1782722473145,58.1846656799316,58.2140159606934,58.2511253356934,58.2517127990723,58.2080535888672,58.1865234375,58.154052734375,58.1338844299316,58.124267578125,58.1292457580566,58.1009750366211,58.1186065673828,58.168212890625,58.16259765625,58.1018104553223,58.0633277893066,58.0313987731934,58.0159187316895,58.0138168334961,57.9937019348145,57.9970703125,58.0630836486816,58.0872077941895,58.2385215759277,58.2970237731934,58.2938461303711,58.2756309509277,58.2785606384277,58.3456077575684,58.3956031799316,58.4164085388184,58.4505882263184,58.4457015991211,58.4281768798828,58.3997077941895,58.3602027893066],[58.4850120544434,58.4786109924316,58.5092315673828,58.5417022705078,58.56640625,58.6185073852539,58.6193809509277,58.611083984375,58.594970703125,58.5708541870117,58.5408668518066,58.4850120544434],[58.5771026611328,58.5610389709473,58.5691375732422,58.6682090759277,58.6712913513184,58.7364234924316,58.7892074584961,58.7952194213867,58.5771026611328],[59.8184089660645,59.8126487731934,59.8782272338867,59.9506797790527,59.9961891174316,60.0151863098145,59.9821281433105,59.9210929870605,59.8184089660645],[60.0653343200684,60.0347213745117,60.0532684326172,60.113525390625,60.1516609191895,60.0923309326172,60.0653343200684],[59.8132286071777,59.7988243103027,59.8283729553223,59.9019546508789,59.9437484741211,59.9536094665527,60.0366172790527,60.0970230102539,60.1853523254395,60.3113288879395,60.3582534790039,60.3630867004395,60.3322296142578,60.2888641357422,60.2543449401855,60.0752906799316,60.0519523620605,59.9911651611328,59.969970703125,59.9602584838867,59.9336929321289,59.8901863098145,59.8675308227539,59.8655776977539,59.8536109924316,59.83154296875,59.8132286071777],[60.383544921875,60.3339347839355,60.3311500549316,60.3462371826172,60.3355979919434,60.314208984375,60.2815437316895,60.22412109375,60.1728515625,60.1005821228027,60.0693359375,60.0283660888672,59.9728050231934,59.9131355285645,59.8932113647461,59.8900337219238,59.849609375,59.8197746276855,59.7754402160645,59.7641143798828,59.7738265991211,59.8148880004883,59.8344230651855,59.8646469116211,59.9838905334473,60.008544921875,60.0957069396973,60.2066421508789,60.2244606018066,60.2335472106934,60.2169914245605,60.2964401245117,60.3162612915039,60.3387718200684,60.3827667236328,60.3816871643066,60.36474609375,60.3911590576172,60.3967742919922,60.383544921875],[60.3371124267578,60.3126449584961,60.3213348388672,60.3368186950684,60.4011268615723,60.3882293701172,60.3371124267578],[60.4496612548828,60.4221687316895,60.4287605285645,60.4239234924316,60.4111785888672,60.3925323486328,60.3680191040039,60.3250503540039,60.2684555053711,60.273681640625,60.3356437683105,60.3606910705566,60.3956069946289,60.4085426330566,60.4326171875,60.4678230285645,60.4805679321289,60.4496612548828],[60.4504852294922,60.4241218566895,60.3988761901855,60.3524856567383,60.2727508544922,60.2220726013184,60.1902351379395,60.1779289245605,60.1851539611816,60.2027359008789,60.2306632995605,60.2297821044922,60.2994155883789,60.3214378356934,60.3512725830078,60.3712921142578,60.4598655700684,60.4660682678223,60.4449691772461,60.4387664794922,60.4474143981934,60.48681640625,60.4914054870605,60.4504852294922],[60.3617210388184,60.3580551147461,60.4074211120605,60.5037612915039,60.4903831481934,60.4727058410645,60.3739738464355,60.3617210388184],[60.4573707580566,60.3917465209961,60.3984870910645,60.3436546325684,60.2991218566895,60.3311004638672,60.3288536071777,60.4627952575684,60.4932098388672,60.5683059692383,60.6068382263184,60.5056610107422,60.4573707580566],[60.8261756896973,60.8179168701172,60.8306159973145,60.8443374633789,60.8948249816895,60.9161148071289,60.9241180419922,60.900146484375,60.8753929138184,60.8261756896973],[63.6400413513184,63.6156730651855,63.6196250915527,63.6091270446777,63.5854949951172,63.593994140625,63.6688461303711,63.6884727478027,63.6988296508789,63.6806144714355,63.6409149169922,63.6175308227539,63.5766563415527,63.5271987915039,63.4917449951172,63.447998046875,63.4305648803711,63.4065933227539,63.3734855651855,63.3483390808105,63.3575172424316,63.3485832214355,63.3473129272461,63.3105926513672,63.2137680053711,63.1712379455566,63.1849136352539,63.171142578125,63.1212882995605,63.0582046508789,62.9967727661133,62.9685516357422,62.9561004638672,62.9900894165039,63.09375,63.1223602294922,63.140380859375,63.1731452941895,63.19384765625,63.1963386535645,63.2322731018066,63.2842788696289,63.3111343383789,63.3492660522461,63.3792991638184,63.4443855285645,63.4529304504395,63.4458999633789,63.4162101745605,63.3721694946289,63.3392601013184,63.3319778442383,63.3512191772461,63.3942375183105,63.4247055053711,63.4772453308105,63.5298347473145,63.5805168151855,63.7030792236328,63.7270050048828,63.6400413513184],[66.2274398803711,66.2218246459961,66.2371597290039,66.2777404785156,66.3280792236328,66.317138671875,66.2781753540039,66.2274398803711],[69.6507797241211,69.3585968017578,69.0663604736328,68.774169921875,68.4819869995117,68.1897430419922,67.8975601196289,67.6053695678711,67.3131332397461,67.0209426879883,66.728759765625,66.4365234375,66.1443405151367,65.8521499633789,65.5599136352539,65.2677230834961,64.9755325317383,64.6832962036133,64.39111328125,64.098876953125,63.8066902160645,63.5144538879395,63.2222671508789,62.9300765991211,62.6378898620605,62.3456993103027,62.0534629821777,61.7612838745117,61.4690437316895,61.1768569946289,60.8846664428711,60.5924301147461,60.3002433776855,60.2591323852539,60.2183570861816,60.2997093200684,60.2374992370605,60.1831550598145,60.2528839111328,60.3283233642578,60.3336906433105,60.3397483825684,60.3437042236328,60.2794456481934,60.1727027893066,60.0835952758789,59.9932594299316,59.9457511901855,59.9013175964355,59.7782745361328,59.6833953857422,59.6111335754395,59.5419425964355,59.4429168701172,59.3735847473145,59.2811508178711,59.2262725830078,59.1194343566895,58.9912109375,58.9153823852539,58.9031295776367,58.9881858825684,59.0409698486328,59.1061019897461,59.1500473022461,59.1522445678711,59.2799301147461,59.4590835571289,59.4560508728027,59.4803199768066,59.5329132080078,59.6048355102539,59.6383781433105,59.6626434326172,59.728759765625,59.7932586669922,59.7433090209961,59.6950187683105,59.5786590576172,59.5506858825684,59.4960441589355,59.4414558410645,59.2882804870605,59.2711944580078,59.25,59.1992645263672,59.1553192138672,59.0853500366211,59.0562477111816,59.0091819763184,58.9687538146973,58.939697265625,58.8984832763672,58.8498992919922,58.7955055236816,58.7578620910645,58.7050323486328,58.59716796875,58.5034675598145,58.410888671875,58.3370628356934,58.2228546142578,58.0777320861816,57.9489784240723,57.8770027160645,57.772705078125,57.6451148986816,57.4999008178711,57.4067344665527,57.2763175964355,57.1985359191895,57.1453628540039,57.0794448852539,57.0481910705566,57.0265655517578,56.953369140625,56.8567848205566,56.8187026977539,56.7928199768066,56.7421379089355,56.684814453125,56.5899887084961,56.5960960388184,56.5988273620605,56.5567398071289,56.501220703125,56.44921875,56.4048309326172,56.3786163330078,56.3408203125,56.263671875,56.2305679321289,56.1225090026855,56.0828132629395,56.1092758178711,56.0652351379395,56.0144996643066,55.9505348205566,55.8882293701172,55.8360366821289,55.77978515625,55.7193870544434,55.6544914245605,55.5850067138672,55.5244140625,55.4123039245605,55.3436012268066,55.2979011535645,55.1370124816895,55.0602569580078,55.02587890625,54.9459495544434,54.8341789245605,54.7696800231934,54.7909164428711,54.8076171875,54.9503898620605,55.0611801147461,55.1576690673828,55.1795883178711,55.2439460754395,55.2969741821289,55.3180160522461,55.3320808410645,55.3551292419434,55.4595222473145,55.5511245727539,55.61181640625,55.7359848022461,55.8119621276855,55.9601554870605,55.9975090026855,56.0443840026855,56.0880851745605,56.0121116638184,55.9322280883789,55.8765602111816,55.8541984558105,55.8353500366211,55.7828102111816,55.7659683227539,55.7349090576172,55.6471672058105,55.5541496276855,55.5350112915039,55.5697784423828,55.5995597839355,55.7210464477539,55.75341796875,55.7806625366211,55.8395500183105,55.9300765991211,56.1601066589355,56.1612319946289,56.206787109375,56.2296371459961,56.2416496276855,56.2729988098145,56.3236846923828,56.3800811767578,56.3998527526855,56.420654296875,56.4891090393066,56.5198745727539,56.5579109191895,56.6031265258789,56.6258773803711,56.6341323852539,56.649658203125,56.7664070129395,56.7964363098145,56.822265625,56.8951644897461,56.9306144714355,56.960205078125,57.0407218933105,57.0558128356934,57.0474624633789,57.1721649169922,57.3368644714355,57.5541534423828,57.6422805786133,57.676513671875,57.6946754455566,57.6950645446777,57.6310997009277,57.5551300048828,57.5662155151367,57.72705078125,57.7751502990723,57.8329620361328,57.8638687133789,57.8801307678223,57.8656768798828,57.877685546875,57.9246559143066,57.9244651794434,57.8569831848145,57.8410148620605,57.84423828125,57.8545913696289,57.9363746643066,57.9932632446289,58.0721664428711,58.1283645629883,58.2110862731934,58.2892570495605,58.4678726196289,58.4987335205078,58.5181655883789,58.5152359008789,58.4982872009277,58.4153289794922,58.2793464660645,58.2329559326172,58.2441940307617,58.2996139526367,58.3671913146973,58.3847160339355,58.453857421875,58.6462936401367,58.7421875,58.7656211853027,58.7967758178711,58.8428726196289,59.0766143798828,59.2390632629395,59.3248977661133,59.4100570678711,59.4194374084473,59.3530731201172,59.3184585571289,59.3086891174316,59.2415008544922,59.2079124450684,59.2106895446777,59.2022972106934,59.0875511169434,58.9096183776855,58.7777328491211,58.6709022521973,58.5897445678711,58.51220703125,58.3408660888672,58.3067855834961,58.2789039611816,58.2458534240723,58.2334022521973,58.2559051513672,58.2982940673828,58.3761253356934,58.4120559692383,58.3942375183105,58.4001922607422,58.4638175964355,58.5770492553711,58.6227073669434,58.7891120910645,58.8216285705566,58.89794921875,58.9037628173828,58.8739776611328,58.8932113647461,58.9998512268066,59.0395545959473,59.0480995178223,58.9467811584473,58.8934555053711,58.8625984191895,58.8196296691895,58.788818359375,58.7701644897461,58.7654800415039,58.7869110107422,58.8272933959961,58.8463363647461,58.8625030517578,58.9070816040039,58.94091796875,58.9838409423828,59.0344734191895,59.0211410522461,58.9669418334961,58.9349098205566,58.9251480102539,58.8737297058105,58.8666496276855,58.883544921875,58.8815460205078,58.8501930236816,58.8092727661133,58.7863235473633,58.7523918151855,58.740234375,58.7006340026855,58.6797866821289,58.6176719665527,58.6244621276855,58.6022453308105,58.5063018798828,58.4527320861816,58.3841781616211,58.3642044067383,58.3503875732422,58.3279800415039,58.2452163696289,58.2439956665039,58.2664527893066,58.3324241638184,58.3952140808105,58.5811996459961,58.5899429321289,58.6258773803711,58.6442375183105,58.659912109375,58.7070808410645,58.7855453491211,58.8468742370605,58.8910179138184,58.9414520263672,59.0468292236328,59.0873031616211,59.1101112365723,59.1150894165039,59.1291542053223,59.1522445678711,59.1659202575684,59.1875495910645,59.2369117736816,59.3756332397461,59.4624519348145,59.5039520263672,59.5272979736328,59.5462379455566,59.5660667419434,59.5868148803711,59.6103057861328,59.6980934143066,59.7263145446777,59.7901840209961,59.8482894897461,59.8805160522461,59.9123573303223,59.9332962036133,59.9535636901855,59.9637680053711,59.9568367004395,59.8770027160645,59.8479461669922,59.7387199401855,59.6109390258789,59.6203575134277,59.66259765625,59.7433090209961,59.7820777893066,59.8198776245117,59.8398399353027,59.8282241821289,59.8350105285645,59.8927726745605,60.0009765625,60.0122566223145,60.01708984375,59.9734382629395,59.8307151794434,59.8056640625,59.7266578674316,59.7107429504395,59.7231941223145,59.7488746643066,59.873779296875,59.9027824401855,59.9800300598145,60.0041465759277,60.0828094482422,60.1052742004395,60.1176795959473,60.1088409423828,60.0854949951172,60.0422859191895,60.0194320678711,59.9947776794434,59.9698753356934,60.033447265625,60.0860328674316,60.0969696044922,60.0550308227539,60.0128898620605,60.0087928771973,60.0164070129395,60.0457992553711,60.0630378723145,60.0843276977539,60.1507301330566,60.1910133361816,60.2052230834961,60.2246589660645,60.2492218017578,60.272705078125,60.2950668334961,60.3351593017578,60.4591827392578,60.5335960388184,60.5846176147461,60.6690940856934,60.6628456115723,60.6008796691895,60.5369148254395,60.4536628723145,60.4153785705566,60.3801307678223,60.3885726928711,60.4407196044922,60.4675750732422,60.4692344665527,60.4781761169434,60.5246543884277,60.56201171875,60.6219711303711,60.651123046875,60.6606941223145,60.6922836303711,60.7155265808105,60.7347679138184,60.7490768432617,60.7381324768066,60.7007789611816,60.7291526794434,60.7451133728027,60.7567863464355,60.8108406066895,60.8388633728027,60.8709449768066,60.8973121643066,60.9925308227539,61.0535163879395,61.1126480102539,61.1358413696289,61.1278305053711,61.0775375366211,61.0048789978027,60.977783203125,60.9961929321289,61.0025367736816,60.996826171875,60.9782676696777,60.9467735290527,60.9254837036133,60.9145011901855,60.9180183410645,60.9502944946289,60.9703102111816,60.9949226379395,60.9794425964355,60.9330062866211,60.9095230102539,60.8853988647461,60.889892578125,60.9482917785645,60.9685554504395,61.0190467834473,61.2189445495605,61.2178230285645,61.1863784790039,61.1064910888672,61.0826683044434,61.0796890258789,61.0882835388184,61.081787109375,61.0604019165039,61.0369606018066,61.0114784240723,61.0071258544922,61.0362319946289,61.0440444946289,61.0299339294434,60.939697265625,60.8535652160645,60.8318862915039,60.8354988098145,60.8269996643066,60.8029327392578,60.7340354919434,60.7243156433105,60.69970703125,60.6752967834473,60.6093215942383,60.5833473205566,60.56982421875,60.5720710754395,60.5652351379395,60.5237808227539,60.4894523620605,60.4864273071289,60.5147933959961,60.5299835205078,60.5320816040039,60.547119140625,60.5751457214355,60.5671882629395,60.5233383178711,60.48486328125,60.4518547058105,60.4283218383789,60.4142112731934,60.39306640625,60.3649368286133,60.3231430053711,60.2676773071289,60.1677703857422,60.1542472839355,60.1468238830566,60.1454582214355,60.1220245361328,59.9891128540039,59.9747085571289,59.9889640808105,59.9874000549316,59.9700698852539,59.9568367004395,59.9477500915527,59.9512214660645,59.9799842834473,60.0002403259277,60.0334968566895,59.9982948303223,60.0136680603027,60.1057586669922,60.1002426147461,60.0010261535645,59.9662590026855,59.8943405151367,59.7704582214355,59.766845703125,59.78466796875,59.8953170776367,59.9195823669434,59.8558082580566,59.8327178955078,59.7503433227539,59.7379417419434,59.7822761535645,59.7844276428223,59.7769546508789,59.7130393981934,59.6900405883789,59.5665512084961,59.5709495544434,59.583251953125,59.5813446044922,59.5353012084961,59.5373039245605,59.5645980834961,59.5633811950684,59.5350570678711,59.4796371459961,59.4269561767578,59.3418464660645,59.3026885986328,59.2491188049316,59.2439956665039,59.2784194946289,59.30078125,59.2896499633789,59.2569313049316,59.2369117736816,59.2293930053711,59.2323226928711,59.24560546875,59.2305679321289,59.1873054504395,59.1885261535645,59.259765625,59.2650871276855,59.2851066589355,59.3427238464355,59.3863296508789,59.4063491821289,59.4622077941895,59.4827156066895,59.5163078308105,59.5855941772461,59.6376953125,59.7718238830566,59.7821807861328,59.7894058227539,59.6622543334961,59.650390625,59.6512718200684,59.7000007629395,59.7209014892578,59.7820777893066,59.9211387634277,59.9883270263672,60.0920372009277,60.2026405334473,60.2744674682617,60.4664573669434,60.5535621643066,60.6598663330078,60.7229499816895,60.742919921875,60.8412094116211,60.914794921875,61.0235824584961,61.0226593017578,60.9852027893066,60.9552230834961,60.9328155517578,60.9351577758789,60.9622535705566,60.9519996643066,60.8804206848145,60.8764190673828,60.8855476379395,60.9356956481934,60.9647445678711,60.9938507080078,61.1217308044434,61.1710929870605,61.1942405700684,61.2132835388184,61.2317390441895,61.2637214660645,61.3075180053711,61.3633270263672,61.4172859191895,61.4973640441895,61.5007781982422,61.4860305786133,61.4707069396973,61.4133796691895,61.3730010986328,61.2942390441895,61.2793426513672,61.2679176330566,61.2599601745605,61.3002433776855,61.3067893981934,61.3011245727539,61.1982421875,61.1457061767578,61.0858383178711,61.0419387817383,61.0141143798828,60.9796333312988,60.9107398986816,60.8579635620117,60.8331565856934,60.7548828125,60.7402381896973,60.7340812683105,60.6822280883789,60.5281257629395,60.4722137451172,60.4530258178711,60.409423828125,60.3811073303223,60.3363265991211,60.2654304504395,60.2384300231934,60.2370643615723,60.2471656799316,60.2928733825684,60.295654296875,60.2892608642578,60.2403831481934,60.177490234375,60.1252937316895,60.0837860107422,60.0411109924316,59.9972152709961,59.9209022521973,59.8980941772461,59.8750495910645,59.8568801879883,59.8427238464355,59.8104972839355,59.7939949035645,59.7300300598145,59.7091293334961,59.6709518432617,59.6598663330078,59.6671867370605,59.7174758911133,59.7298316955566,59.7401351928711,59.720947265625,59.6470260620117,59.6348114013672,59.6150398254395,59.5984878540039,59.5452613830566,59.5098609924316,59.4737281799316,59.3632774353027,59.3363723754883,59.2401351928711,59.1555633544922,59.1198692321777,59.0780296325684,59.0679206848145,59.0386734008789,58.9599609375,58.9085426330566,58.8843231201172,58.8578643798828,58.8222160339355,58.7548294067383,58.6547355651855,58.6263694763184,58.6041030883789,58.5878410339355,58.4929695129395,58.4417495727539,58.3971672058105,58.3658180236816,58.3043441772461,58.29345703125,58.2887725830078,58.2346153259277,58.159423828125,58.1467819213867,58.1473121643066,58.1180648803711,58.1097679138184,58.0556640625,58.0160636901855,57.913330078125,57.8818359375,57.80712890625,57.7770500183105,57.7588882446289,57.7335929870605,57.7010765075684,57.6730461120605,57.6266136169434,57.5682144165039,57.5590362548828,57.5449714660645,57.5265159606934,57.4475593566895,57.445068359375,57.4634246826172,57.44921875,57.3599624633789,57.327880859375,57.3106956481934,57.2936515808105,57.2405738830566,57.204833984375,57.1051750183105,57.0897941589355,57.0650863647461,57.0099601745605,57.0160636901855,57.005615234375,56.9688491821289,56.9476547241211,56.9420890808105,56.9129409790039,56.8602027893066,56.8265609741211,56.8120613098145,56.8084983825684,56.8158683776855,56.809814453125,56.7903327941895,56.759765625,56.6731948852539,56.6344718933105,56.627685546875,56.6334457397461,56.6516609191895,56.6452140808105,56.5921401977539,56.5520515441895,56.5431632995605,56.5316886901855,56.5204582214355,56.5095710754395,56.5103530883789,56.5010261535645,56.4781761169434,56.4535140991211,56.4358367919922,56.33544921875,56.3126983642578,56.3076667785645,56.3182640075684,56.3015632629395,56.2803230285645,56.254150390625,56.2036628723145,56.1962394714355,56.1114768981934,56.0754890441895,56.0621109008789,56.0724601745605,56.1668434143066,56.182861328125,56.1845207214355,56.1546859741211,56.0431175231934,55.9869155883789,55.8427238464355,55.8100090026855,55.7484855651855,55.6952133178711,55.6527824401855,55.6259269714355,55.64501953125,55.7948722839355,55.8246574401855,55.84375,55.8411140441895,55.8327140808105,55.80029296875,55.7921867370605,55.7948722839355,55.762939453125,55.6605453491211,55.6351089477539,55.6138191223145,55.5578155517578,55.5373039245605,55.5354957580566,55.5523948669434,55.5404319763184,55.4996566772461,55.4835472106934,55.5136222839355,55.4930686950684,55.4404296875,55.4057121276855,55.3888664245605,55.3712882995605,55.3825187683105,55.3978042602539,55.4648933410645,55.5132827758789,55.5361328125,55.5562973022461,55.5586433410645,55.5435562133789,55.5597648620605,55.5790061950684,55.6122055053711,55.6291465759277,55.618408203125,55.5928230285645,55.5633811950684,55.5299339294434,55.4207000732422,55.3911590576172,55.1986312866211,55.1393089294434,55.1437492370605,55.121337890625,55.0732421875,55.0502471923828,55.0523414611816,55.0614738464355,55.0776863098145,55.0928230285645,55.1068840026855,55.1454124450684,55.2084465026855,55.2427253723145,55.2466773986816,55.218017578125,55.0714836120605,55.0384292602539,54.99658203125,54.9499969482422,54.9545402526855,55.0464859008789,55.064697265625,55.0347633361816,54.9736328125,54.9165496826172,54.8633766174316,54.8375968933105,54.8391571044922,54.8763656616211,54.9492683410645,55.0099601745605,55.0585441589355,55.0958518981934,55.121826171875,55.1939468383789,55.2047386169434,55.1868667602539,55.183837890625,55.195556640625,55.2186012268066,55.2530288696289,55.2970733642578,55.3507804870605,55.4500007629395,55.5947265625,55.7194328308105,55.8241691589355,55.9072265625,56.0214385986328,56.0144538879395,55.9774398803711,55.9542961120605,55.9513168334961,55.9696311950684,55.99365234375,55.9705085754395,55.9413070678711,55.9117202758789,55.8871574401855,55.7718772888184,55.7544441223145,55.756591796875,55.771484375,55.8546409606934,55.8704605102539,55.8743171691895,55.8634757995605,55.837890625,55.8166999816895,55.7999038696289,55.8050765991211,55.8321800231934,55.8644523620605,55.9354476928711,55.9650382995605,56.0062980651855,56.0591278076172,56.1375007629395,56.241455078125,56.3141098022461,56.3963394165039,56.43701171875,56.5616188049316,56.6885757446289,56.7700691223145,56.8600578308105,56.8821792602539,56.847412109375,56.81640625,56.7957534790039,56.78857421875,56.7948722839355,56.8279304504395,56.8877449035645,56.9442405700684,56.9973602294922,57.0394058227539,57.1140632629395,57.1990699768066,57.2979049682617,57.3426742553711,57.3664054870605,57.4094734191895,57.5113754272461,57.5280723571777,57.5481414794922,57.5392570495605,57.5137252807617,57.497802734375,57.4834442138672,57.5062026977539,57.5182113647461,57.5258827209473,57.5406761169434,57.6014633178711,57.6380882263184,57.679443359375,57.7438926696777,57.8952178955078,58.0508308410645,58.1399421691895,58.1721687316895,58.1941871643066,58.2345199584961,58.2348136901855,58.2537117004395,58.3507308959961,58.4213409423828,58.5030288696289,58.6409187316895,58.736328125,58.7441902160645,58.7725639343262,58.9636688232422,59.1342735290527,58.9888687133789,58.8776359558105,58.7484855651855,58.6401863098145,58.6142578125,58.6417999267578,58.7456512451172,58.7826156616211,58.8503456115723,58.9383773803711,58.9993171691895,59.0093231201172,58.9799346923828,58.9774398803711,59.0374984741211,59.08984375,59.0728530883789,58.9877967834473,58.9293975830078,58.9500999450684,58.9738731384277,58.9025421142578,58.7939491271973,58.7187461853027,58.5203094482422,58.4409713745117,58.4045372009277,58.4697723388672,58.7212867736816,58.7929229736328,58.9111328125,58.8942909240723,58.8359870910645,58.8198738098145,58.9059066772461,58.9715309143066,59.0511741638184,59.00732421875,58.955078125,58.8716812133789,58.8724136352539,58.8009757995605,58.7994651794434,58.7609367370605,58.7437057495117,58.6695289611816,58.6120109558105,58.6442375183105,58.6850128173828,58.7170867919922,58.7942886352539,58.8974113464355,58.9499473571777,59.0164070129395,59.1096649169922,59.1094741821289,59.0760765075684,59.1461448669434,59.2839851379395,59.365478515625,59.4282722473145,59.5145034790039,59.5886268615723,59.714111328125,59.9800300598145,60.1783180236816,60.2839889526367,60.4568862915039,60.6143569946289,60.6848182678223,60.6953620910645,60.6948699951172,60.685546875,60.6343269348145,60.5952110290527,60.3946762084961,60.2969703674316,60.2689476013184,60.2310523986816,60.1991233825684,60.1265640258789,60.0383796691895,59.9897499084473,59.9936485290527,59.9227523803711,59.8456039428711,59.8015174865723,59.8067893981934,59.8967781066895,59.9488792419434,59.9942359924316,60.1493148803711,60.3038101196289,60.3072242736816,60.3484382629395,60.4125518798828,60.4642562866211,60.5006332397461,60.5260772705078,60.5235824584961,60.5412139892578,60.7400398254395,60.8731460571289,60.8920364379883,60.8715324401855,60.8190422058105,60.7958946228027,60.7712860107422,60.7246589660645,60.6466293334961,60.5918426513672,60.6067428588867,60.6915054321289,60.7660636901855,60.7583045959473,60.7451667785645,60.6682586669922,60.5899925231934,60.66455078125,60.7574234008789,60.7981452941895,60.8222198486328,60.8538093566895,60.8804206848145,60.8774909973145,60.9029808044434,60.938232421875,60.9697227478027,60.8646965026855,60.8699684143066,60.9312973022461,60.9206581115723,60.9328155517578,60.9656753540039,61.0436553955078,61.0867652893066,61.1117668151855,61.1148948669434,61.0941925048828,61.1301231384277,61.1924285888672,61.1868629455566,61.1528358459473,61.1696319580078,61.1977043151855,61.2276382446289,61.2687492370605,61.2748565673828,61.26611328125,61.2123069763184,61.1687507629395,61.1063003540039,61.0948753356934,61.1023445129395,61.1651840209961,61.2999038696289,61.335693359375,61.40380859375,61.4911613464355,61.5362281799316,61.5508766174316,61.5067405700684,61.5459480285645,61.5890121459961,61.6508293151855,61.6573257446289,61.6450653076172,61.6793937683105,61.6960945129395,61.748291015625,61.8031234741211,61.8341789245605,61.8368186950684,61.8692893981934,61.9274406433105,61.9593772888184,62.01171875,62.1004409790039,62.3039054870605,62.4735336303711,62.5126991271973,62.5337905883789,62.5175743103027,62.4811515808105,62.4967308044434,62.5116195678711,62.5810546875,62.6082534790039,62.6866683959961,62.7093734741211,62.6767578125,62.6231918334961,62.6770515441895,62.8009719848633,62.9180641174316,62.9706077575684,63.0204620361328,63.0404319763184,63.0304679870605,63.0540046691895,63.1276359558105,63.1852073669434,63.2150382995605,63.26171875,63.2472152709961,63.1928215026855,63.1251487731934,63.0904312133789,63.0703163146973,63.0478019714355,63.0303230285645,63.0164031982422,63.0567855834961,63.1058578491211,63.0845222473145,63.0457534790039,63.0464363098145,63.0797348022461,63.114990234375,63.2065925598145,63.2658233642578,63.4525871276855,63.5292015075684,63.5409698486328,63.5341796875,63.4713363647461,63.4529304504395,63.4681663513184,63.4969711303711,63.5579071044922,63.6605453491211,63.7293472290039,63.8189430236816,63.9349136352539,64.0311965942383,64.2512664794922,64.3965835571289,64.43994140625,64.4337921142578,64.5263671875,64.5164108276367,64.5344772338867,64.5791015625,64.6128921508789,64.6819305419922,64.755615234375,64.7955627441406,64.8395538330078,64.9039993286133,64.9254455566406,64.9240188598633,64.7948684692383,64.79248046875,64.8162612915039,64.74267578125,64.6780700683594,64.6128463745117,64.4508361816406,64.3775405883789,64.3742218017578,64.5164108276367,64.6520004272461,64.6059036254883,64.5632781982422,64.532958984375,64.5197296142578,64.4786148071289,64.423828125,64.4751968383789,64.5498046875,64.5882263183594,64.5839309692383,64.5074234008789,64.5232849121094,64.5296401977539,64.5113754272461,64.4803237915039,64.4606475830078,64.4536590576172,64.4652328491211,64.5128479003906,64.582763671875,64.625732421875,64.7280807495117,64.7975616455078,64.8269500732422,64.926513671875,64.9529800415039,65.0960922241211,65.1570816040039,65.163818359375,65.1472625732422,65.1349105834961,65.1547393798828,65.247314453125,65.2737808227539,65.2607421875,65.2858352661133,65.3055648803711,65.3527297973633,65.3382797241211,65.422119140625,65.5677719116211,65.5956039428711,65.6577606201172,65.7191390991211,65.7481460571289,65.71435546875,65.6812057495117,65.75830078125,65.8593292236328,65.8770523071289,65.9049301147461,65.9591751098633,66.0518569946289,66.1006393432617,66.1444320678711,66.1702651977539,66.1272506713867,66.121337890625,66.112548828125,66.1312026977539,66.1451187133789,66.1670913696289,66.2450714111328,66.2884750366211,66.3190383911133,66.409912109375,66.43994140625,66.4378433227539,66.5550308227539,66.5884323120117,66.6107406616211,66.616455078125,66.5746612548828,66.58349609375,66.5758743286133,66.5615768432617,66.5311050415039,66.4926300048828,66.3783645629883,66.2869110107422,66.25732421875,66.2155303955078,66.0838317871094,66.075439453125,66.0992202758789,66.059814453125,66.0508346557617,66.071044921875,66.0428695678711,66.0536575317383,66.2505416870117,66.2814025878906,66.2471694946289,66.2193908691406,66.239501953125,66.1888122558594,66.2946319580078,66.3343276977539,66.4070358276367,66.370849609375,66.4118118286133,66.4930648803711,66.693115234375,66.7336959838867,66.7356491088867,66.8051223754883,66.8943862915039,66.9308166503906,66.9473114013672,66.9186553955078,66.8013687133789,66.7841262817383,66.6672821044922,66.5596160888672,66.4595260620117,66.496337890625,66.4953155517578,66.4742202758789,66.3843765258789,66.3730926513672,66.4202651977539,66.508544921875,66.5724639892578,66.6124954223633,66.60498046875,66.6708526611328,66.6527862548828,66.5518569946289,66.5915985107422,66.6455078125,66.7003402709961,66.803955078125,66.9227981567383,66.9793472290039,67.0205535888672,67.049560546875,67.0198745727539,67.060302734375,67.1039581298828,67.0185089111328,67.0364303588867,67.0272979736328,67.1025848388672,67.195556640625,67.27099609375,67.4775924682617,67.6067352294922,68.0456008911133,68.1559066772461,68.2779312133789,68.3079605102539,68.3202667236328,68.359619140625,68.4080047607422,68.4243621826172,68.3738250732422,68.361083984375,68.3902359008789,68.4251480102539,68.5732421875,68.7972183227539,68.8853530883789,68.8675842285156,68.8824691772461,68.9024429321289,68.9364776611328,68.961181640625,69.0366668701172,69.1701202392578,69.3453598022461,69.3925323486328,69.3804702758789,69.387939453125,69.454345703125,69.6106948852539,69.7581100463867,70.0941390991211,70.2271957397461,70.2876434326172,70.3317337036133,70.2898483276367,70.27734375,70.2576675415039,70.2484436035156,70.2054672241211,70.1766662597656,70.1619644165039,70.1652374267578,70.196533203125,70.2345199584961,70.3045883178711,70.4205551147461,70.4463882446289,70.5912170410156,70.5855941772461,70.5681610107422,70.4725570678711,70.4475631713867,70.3332977294922,70.3314514160156,70.2788543701172,70.3241653442383,70.3892593383789,70.4530258178711,70.4970703125,70.4771499633789,70.5245132446289,70.5304641723633,70.6340789794922,70.6348648071289,70.7867660522461,70.8785171508789,70.8767547607422,70.8596649169922,70.8319320678711,70.8138656616211,70.78125,70.7525329589844,70.7484359741211,70.7720718383789,70.8016128540039,70.7990188598633,70.8201141357422,70.841064453125,70.8453140258789,70.860107421875,70.9412612915039,71.0396041870117,71.09326171875,71.2300262451172,71.3189468383789,71.4076690673828,71.3966751098633,71.3791046142578,71.341552734375,71.2915573120117,71.1884231567383,71.1827621459961,71.12109375,71.0615768432617,70.9954147338867,70.9278335571289,70.9022445678711,70.8419876098633,70.8346710205078,70.8572769165039,70.8943405151367,71.0149917602539,71.0822296142578,71.0992126464844,71.0830535888672,71.0484848022461,70.9871139526367,70.9277877807617,70.894287109375,70.8479919433594,70.8383331298828,70.8011245727539,70.8773422241211,70.8936004638672,70.8910675048828,70.9325637817383,70.8760223388672,70.8907241821289,70.8809585571289,70.8467788696289,70.8103561401367,70.7332534790039,70.6536178588867,70.6204605102539,70.61474609375,70.5682678222656,70.556640625,70.5601577758789,70.5380325317383,70.5113296508789,70.4520950317383,70.4187545776367,70.4516143798828,70.4646987915039,70.5099105834961,70.4901351928711,70.4438934326172,70.434326171875,70.4437026977539,70.5096664428711,70.5128936767578,70.4914016723633,70.5007858276367,70.4251937866211,70.4163055419922,70.3179168701172,70.3149871826172,70.3567428588867,70.35546875,70.3154830932617,70.3032760620117,70.2401351928711,70.2172317504883,70.17041015625,70.1917495727539,70.1861343383789,70.15625,70.1600570678711,70.0509262084961,70.033935546875,70.0086975097656,69.9821319580078,70.0390167236328,70.0541000366211,70.1019515991211,70.1014633178711,70.0895538330078,70.0953140258789,70.1162567138672,70.0337905883789,69.9395065307617,69.869873046875,69.7703628540039,69.7146987915039,69.6533660888672,69.6467742919922,69.6646957397461,69.659423828125,69.6507797241211],[39.9502449035645,39.9071273803711,39.9174308776855,39.9460945129395,39.9686508178711,39.98095703125,39.9932594299316,39.9953117370605,39.9502449035645],[39.892578125,39.8834495544434,39.8849143981934,39.8951187133789,39.976318359375,39.9925308227539,39.9949188232422,40.0057601928711,40.0388641357422,40.0879898071289,40.1332511901855,40.1522941589355,40.149169921875,40.0798797607422,40.0596656799316,40.0481452941895,39.9798316955566,39.9067878723145,39.892578125],[42.2486801147461,42.2665519714355,42.28466796875,42.25,42.2064476013184,42.186279296875,42.1628913879395,42.0777816772461,42.0379867553711,42.0308113098145,42.0196266174316,41.9226570129395,41.9052238464355,41.87548828125,41.830810546875,41.7250480651855,41.5714378356934,41.5399894714355,41.5144538879395,41.496826171875,41.4495582580566,41.4126472473145,41.4603500366211,41.4498062133789,41.4005355834961,41.2625007629395,41.2248992919922,41.1959953308105,41.1865730285645,41.1526374816895,41.1399421691895,41.1524925231934,41.1233863830566,41.1360321044922,41.3418960571289,41.3074722290039,41.3080558776855,41.333251953125,41.367431640625,41.4354476928711,41.5032730102539,41.5341796875,41.5412101745605,41.5330085754395,41.5155754089355,41.4540023803711,41.4280281066895,41.4131355285645,41.3610382080078,41.3113784790039,41.1950187683105,41.1870613098145,41.1647491455078,41.1865730285645,41.1737289428711,41.1184577941895,41.0668487548828,41.0259284973145,41.0142593383789,41.0399436950684,41.04345703125,41.0189437866211,40.981689453125,40.8837890625,40.8699226379395,40.8621559143066,40.8423347473145,40.8424301147461,40.860107421875,40.8285140991211,40.8106422424316,40.7860336303711,40.650390625,40.6086921691895,40.5556144714355,40.5254402160645,40.5243644714355,40.5780754089355,40.5651359558105,40.54345703125,40.5197257995605,40.4630851745605,40.4273910522461,40.4016571044922,40.4242172241211,40.4386215209961,40.4543952941895,40.4544410705566,40.4386215209961,40.3407211303711,40.2895050048828,40.2585945129395,40.240966796875,40.2343254089355,40.1880378723145,40.15234375,40.1786117553711,40.2080078125,40.2171363830566,40.2102508544922,40.208984375,40.2419929504395,40.2751922607422,40.2869148254395,40.271240234375,40.2548828125,40.2388687133789,40.2345695495605,40.201171875,40.2141609191895,40.2671394348145,40.3245124816895,40.3453598022461,40.3613777160645,40.3841323852539,40.4120140075684,40.4392585754395,40.4535179138184,40.5627937316895,40.6611824035645,40.6690902709961,40.6877937316895,40.7217750549316,40.7395973205566,40.778564453125,40.7965850830078,40.8150863647461,40.8396492004395,40.9114723205566,41.0234375,41.0351066589355,41.0276374816895,40.9192390441895,40.8917007446289,40.8204116821289,40.7714347839355,40.6842765808105,40.656982421875,40.6619644165039,40.6790542602539,40.76708984375,40.7971687316895,40.7673835754395,40.7239265441895,40.634765625,40.587646484375,40.5665512084961,40.327392578125,40.2965812683105,40.2881355285645,40.1980934143066,40.1875953674316,40.208740234375,40.2226066589355,40.1826667785645,40.1670875549316,40.1472663879395,40.1291999816895,40.1270980834961,40.1363296508789,40.1195831298828,40.0899429321289,40.0713386535645,40.0682106018066,40.0503425598145,40.031494140625,40.0133323669434,39.9607887268066,39.9273414611816,39.907470703125,39.8909645080566,39.8843269348145,39.9091300964355,39.9041976928711,39.8818359375,39.8555679321289,39.8362312316895,39.8462905883789,39.8388671875,39.7432632446289,39.6349639892578,39.5627899169922,39.5367164611816,39.5288543701172,39.5376930236816,39.5482902526855,39.5641593933105,39.5937957763672,39.6213874816895,39.5576171875,39.5187492370605,39.482421875,39.465576171875,39.2420921325684,39.2166976928711,39.1966781616211,39.1502914428711,39.1310539245605,39.1091766357422,39.0084953308105,38.99462890625,38.9822273254395,38.9830093383789,38.992919921875,38.9835968017578,38.9620132446289,38.9276351928711,38.890625,38.6692886352539,38.5889167785645,38.4735336303711,38.3831062316895,38.2945327758789,38.23779296875,38.2110366821289,38.1695327758789,38.1167945861816,38.0329093933105,37.9596672058105,37.9284172058105,37.83544921875,37.720947265625,37.5707015991211,37.4870109558105,37.2449684143066,37.1722145080566,37.1998062133789,37.2272453308105,37.2225074768066,37.2356452941895,37.2666511535645,37.2580108642578,37.2095680236816,37.2352066040039,37.3348159790039,37.3712882995605,37.3484840393066,37.4586906433105,37.5991706848145,37.7857437133789,37.9320335388184,37.9598617553711,37.9867172241211,38.0107917785645,38.0509300231934,38.0721664428711,38.1180648803711,38.1667022705078,38.2001457214355,38.2442359924316,38.2687492370605,38.2500495910645,38.2263679504395,38.2257308959961,38.2385749816895,38.3488273620605,38.5394515991211,38.6724586486816,38.7360343933105,38.7564468383789,38.8162117004395,38.977294921875,38.95361328125,39.058349609375,39.1605491638184,39.1881370544434,39.3770980834961,39.49951171875,39.6331558227539,39.716796875,39.8584938049316,39.944091796875,39.9756355285645,40.0362281799316,40.3320808410645,40.4674797058105,40.5412101745605,40.6833000183105,40.893798828125,41.0306167602539,41.0937004089355,41.1634292602539,41.2398452758789,41.2760734558105,41.2746086120605,41.26513671875,41.2521476745605,41.1951179504395,41.189208984375,41.1905746459961,41.2108879089355,41.22900390625,41.2486801147461,41.2457504272461,41.2161636352539,41.2216300964355,41.3489761352539,41.3994140625,41.4273452758789,41.4762229919434,41.5452156066895,41.5941429138184,41.6449699401855,41.7005386352539,41.7596664428711,41.7926788330078,41.7822761535645,41.8031234741211,41.8344230651855,41.8570327758789,41.9074211120605,41.9543952941895,41.9735336303711,41.9954109191895,42.018798828125,42.0680656433105,42.1317367553711,42.1647453308105,42.1908187866211,42.2117195129395,42.2360343933105,42.295166015625,42.3015594482422,42.2995109558105,42.3232917785645,42.3561515808105,42.481689453125,42.5114250183105,42.5237770080566,42.5281257629395,42.540283203125,42.5614738464355,42.6761741638184,42.7784652709961,42.6819343566895,42.662841796875,42.6717300415039,42.6880836486816,42.6663093566895,42.6280746459961,42.6029777526855,42.5763664245605,42.5272941589355,42.4066925048828,42.3401374816895,42.3168449401855,42.2993659973145,42.2917976379395,42.2920913696289,42.29248046875,42.3124504089355,42.3467750549316,42.39892578125,42.4288597106934,42.4477081298828,42.4615707397461,42.4865226745605,42.4876441955566,42.4587898254395,42.4200210571289,42.335205078125,42.2310562133789,42.1898422241211,42.164794921875,42.1562995910645,42.123779296875,42.08447265625,41.9571266174316,41.9148406982422,41.8565444946289,41.6693344116211,41.4505844116211,41.3899917602539,41.3717765808105,41.3502922058105,41.331298828125,41.3072776794434,41.2634773254395,41.26513671875,41.276123046875,41.2879867553711,41.3006324768066,41.3108367919922,41.3222160339355,41.5517578125,41.7813453674316,42.0108871459961,42.2404289245605,42.469970703125,42.6995124816895,42.9290542602539,43.1585922241211,43.3880882263184,43.6176261901855,43.8472137451172,44.0767593383789,44.3063011169434,44.5358390808105,44.765380859375,44.9949226379395,45.0233879089355,45.059326171875,45.0937995910645,45.134765625,45.1810569763184,45.2206077575684,45.2677230834961,45.3036613464355,45.3374519348145,45.37744140625,45.439697265625,45.474365234375,45.5094223022461,45.5429191589355,45.5553703308105,45.507568359375,45.4417991638184,45.3759765625,45.3102035522461,45.2444343566895,45.1786155700684,45.1128425598145,45.0470733642578,44.9813003540039,44.9155769348145,44.8497581481934,44.7839813232422,44.7182121276855,44.6524391174316,44.5866203308105,44.5208511352539,44.455078125,44.393798828125,44.348388671875,44.2482414245605,44.1686019897461,44.082275390625,43.9939460754395,43.877197265625,43.7961921691895,43.7135734558105,43.5772438049316,43.4921379089355,43.4893531799316,43.5095710754395,43.53662109375,43.5580101013184,43.5838890075684,43.6084976196289,43.6279754638672,43.6132354736328,43.5986328125,43.588134765625,43.5778312683105,43.5657234191895,43.5589332580566,43.5511703491211,43.5716323852539,43.6134757995605,43.6529769897461,43.6939468383789,43.7146949768066,43.6490707397461,43.5736808776855,43.4941902160645,43.4175300598145,43.3720207214355,43.310546875,43.2051734924316,43.0645980834961,42.9721183776855,42.876953125,42.9145011901855,42.95458984375,42.9908180236816,42.8733863830566,42.7666473388672,42.6051750183105,42.4727516174316,42.3147926330566,42.1944808959961,42.0887680053711,42.0048828125,42.0011253356934,41.9983406066895,41.994873046875,41.889404296875,41.7412605285645,41.6069831848145,41.4943351745605,41.3486328125,41.270751953125,41.1791496276855,41.1570816040039,41.1423835754395,41.1533203125,41.162353515625,41.1695327758789,41.1771469116211,41.187255859375,41.1639175415039,41.1802711486816,41.1965827941895,41.1300315856934,41.0962409973145,41.061279296875,41.0286102294922,40.9602546691895,40.860595703125,40.8092765808105,40.7540512084961,40.7217750549316,40.6561050415039,40.6194305419922,40.608642578125,40.6226577758789,40.6599617004395,40.7112808227539,40.76513671875,40.8292961120605,40.8762702941895,40.9615211486816,41.0418930053711,41.1238288879395,41.2050285339355,41.2641143798828,41.366943359375,41.4252433776855,41.4602508544922,41.4905738830566,41.5418930053711,41.6290550231934,41.672119140625,41.6973152160645,41.7540512084961,41.8205070495605,41.9459953308105,42.0279769897461,42.0785636901855,42.0802726745605,42.0394515991211,42.0360336303711,42.0547332763672,42.1074714660645,42.1686515808105,42.1941909790039,42.2072257995605,42.2486801147461],[40.9366188049316,40.9608383178711,41.0016593933105,41.0248031616211,41.014892578125,40.9818344116211,40.9344215393066,40.9366188049316],[41.8983879089355,41.8975601196289,41.9007301330566,41.9054679870605,41.9062004089355,41.8983879089355],[12.6952142715454,12.6947269439697,12.6981449127197,12.701171875,12.70361328125,12.710205078125,12.7190437316895,12.7288084030151,12.7348146438599,12.7354497909546,12.731689453125,12.7225580215454,12.7155275344849,12.7098636627197,12.7011232376099,12.6952142715454],[12.99462890625,12.9836912155151,12.9836912155151,12.9762697219849,12.983642578125,12.9881839752197,12.9901857376099,12.9898920059204,12.9973630905151,12.9927730560303,12.9972658157349,13.0053720474243,13.0126953125,13.017822265625,13.0257329940796,13.0432615280151,13.0525388717651,13.0487308502197,13.0451173782349,13.0416994094849,13.0382814407349,13.0247068405151,12.99462890625],[13.1581048965454,13.1422853469849,13.2095699310303,13.2876949310303,13.3306636810303,13.35595703125,13.3587398529053,13.2940425872803,13.202880859375,13.1581048965454],[8.86733436584473,8.84702110290527,8.94731426239014,8.974365234375,9.05502891540527,9.0703125,9.05336856842041,9.03188514709473,8.99570274353027,8.94960975646973,8.89926815032959,8.86733436584473],[9.13837909698486,9.10556602478027,9.13232421875,9.1787109375,9.19179725646973,9.20766639709473,9.21835899353027,9.21645450592041,9.20332050323486,9.17719745635986,9.13837909698486],[10.9064455032349,10.8839845657349,10.9064455032349,10.9378910064697,10.9738273620605,10.9738273620605,10.9302244186401,10.9064455032349],[11.1310052871704,11.0003414154053,10.975830078125,10.8875484466553,10.8812017440796,10.8843269348145,10.9014158248901,10.9587888717651,10.9416007995605,10.9615240097046,10.9815921783447,11.0519046783447,11.080322265625,11.0861330032349,11.04296875,11.0056638717651,11.0018548965454,11.0684576034546,11.167236328125,11.1310052871704],[8.54931640625,8.30595779418945,8.27915000915527,8.24868106842041,8.19160079956055,8.16201210021973,8.05356502532959,7.99404335021973,7.91943359375,7.85400390625,7.82763671875,7.81318378448486,7.77202081680298,7.64833974838257,7.59663105010986,7.53593730926514,7.49868106842041,7.36332988739014,7.32084941864014,7.256591796875,7.21108388900757,7.15620088577271,7.14370107650757,7.16655302047729,7.16455078125,7.15000009536743,7.13398456573486,7.092041015625,7.00288105010986,6.94536113739014,6.85707950592041,6.80595684051514,6.76831007003784,6.78847694396973,6.78691434860229,6.7578125,6.73276329040527,6.72661161422729,6.71137714385986,6.69453096389771,6.65092802047729,6.58837890625,6.51337909698486,6.446533203125,6.38510751724243,6.21430683135986,6.17441368103027,6.12919950485229,6.04951190948486,5.93876934051514,5.906982421875,5.67421913146973,5.43740224838257,5.20205068588257,5.19155311584473,5.16435575485229,5.08198261260986,4.99458026885986,4.94936513900757,4.89252901077271,4.82709980010986,4.77412080764771,4.72919940948486,4.68681669235229,4.57470703125,4.53525352478027,4.51933574676514,4.50468778610229,4.508056640625,4.51689481735229,4.43300771713257,4.40224599838257,4.28779268264771,4.197021484375,4.12631845474243,4.098388671875,4.15673828125,4.13852500915527,4.08432626724243,4.04228544235229,4.03964900970459,4.01792001724243,3.94033217430115,3.67294955253601,3.59345722198486,3.59394526481628,3.68647456169128,3.75644516944885,3.92226552963257,3.94389629364014,3.94287109375,3.89370131492615,3.91503882408142,3.94082021713257,3.93256855010986,3.9306640625,3.92910170555115,3.97441411018372,4.06699180603027,4.10014677047729,4.12685585021973,4.14033222198486,4.13999032974243,4.13989305496216,4.15771436691284,4.23710966110229,4.2744140625,4.27602529525757,4.23227548599243,4.08930683135986,4.01181650161743,3.89980459213257,3.66269540786743,3.58740234375,3.49121117591858,3.343994140625,3.20468759536743,3.0048828125,2.801513671875,2.67187523841858,2.57607436180115,2.52509760856628,2.50239253044128,2.48188471794128,2.45244145393372,2.43403315544128,2.43393564224243,2.41191387176514,2.34042930603027,2.22250962257385,2.15556621551514,2.13603544235229,2.12050747871399,2.04814481735229,1.97670900821686,1.96699225902557,1.95302748680115,1.93159198760986,1.90444350242615,1.7705078125,1.61928701400757,1.52949190139771,1.45527327060699,1.44687485694885,1.45278310775757,1.43100559711456,1.36987328529358,1.29384756088257,1.25332045555115,1.25712895393372,1.21972632408142,1.15844738483429,1.10810554027557,1.02221691608429,0.93188488483429,0.868652164936066,0.790478348731995,0.691259801387787,0.68798828125,0.747509956359863,0.843408167362213,0.926073968410492,0.970361351966858,0.983447134494781,0.97802746295929,0.937255859375,0.863134562969208,0.809765815734863,0.785351514816284,0.763281226158142,0.75195324420929,0.767187595367432,0.821679949760437,0.992138862609863,1.22304654121399,1.28989243507385,1.35825204849243,1.45800805091858,1.56420910358429,1.60078108310699,1.68017590045929,1.82319355010986,1.94033193588257,1.99985361099243,2.05058598518372,2.10126948356628,2.142578125,2.27548813819885,2.33276391029358,2.39013695716858,2.42944359779358,2.47167992591858,2.55991172790527,2.64365243911743,2.67675805091858,2.68994116783142,2.76933574676514,2.79360342025757,2.80019545555115,2.83330106735229,2.79360342025757,2.85532188415527,2.89282202720642,3.187255859375,3.32265615463257,3.34262704849243,3.37397503852844,3.41586899757385,3.46376919746399,3.69111323356628,3.73383784294128,3.768798828125,3.86425757408142,4.08652353286743,4.1982421875,4.28388690948486,4.38071250915527,4.455078125,4.50688505172729,4.66547870635986,4.93081045150757,5.13251924514771,5.27045869827271,5.37548828125,5.447509765625,5.55878925323486,5.70937490463257,5.83310556411743,5.92998027801514,6.02553701400757,6.11977529525757,6.18027353286743,6.24179697036743,6.28496074676514,6.28989315032959,6.24194288253784,6.197509765625,6.15654325485229,6.15678739547729,6.19819307327271,6.18437528610229,6.11533212661743,6.09970664978027,6.13759756088257,6.14799785614014,6.12397432327271,6.13491201400757,6.32148456573486,6.49438524246216,6.70024394989014,6.93793964385986,6.95361375808716,6.95205068588257,6.94794940948486,6.97260761260986,7.00712919235229,7.04052686691284,7.08276414871216,7.09003925323486,7.07758760452271,6.99443387985229,6.98671913146973,6.98520517349243,7.0263671875,7.03291034698486,7.00581026077271,6.99033212661743,7.03261661529541,7.09687519073486,7.24970722198486,7.37026405334473,7.39453172683716,7.41508817672729,7.45488262176514,7.52426719665527,7.61323213577271,7.75795936584473,7.80986356735229,7.96611356735229,8.04770565032959,8.08730411529541,8.15278339385986,8.28706073760986,8.38198280334473,8.48969745635986,8.62758827209473,8.84829044342041,9.10898399353027,9.13515567779541,9.1220703125,9.13515567779541,9.23994159698486,9.25957012176514,9.22280311584473,9.19414043426514,9.16792011260986,9.19414043426514,9.22685527801514,9.322021484375,9.443603515625,9.55463790893555,9.668212890625,9.78916072845459,10.0297355651855,10.1957521438599,10.4912605285645,10.7271966934204,10.8358402252197,10.9771480560303,11.0539064407349,11.1142578125,11.1964349746704,11.6019535064697,11.6664066314697,11.7268562316895,11.7740726470947,11.8235359191895,11.8497562408447,11.8619146347046,11.81494140625,11.7551755905151,11.71875,11.6273431777954,11.6079587936401,11.5699224472046,11.4828128814697,11.4144535064697,11.1903314590454,11.1350584030151,11.0135259628296,10.9967279434204,10.9946775436401,10.8354978561401,10.7262201309204,10.6573734283447,10.4437494277954,10.3159666061401,10.167236328125,10.1080570220947,9.81557559967041,9.64150428771973,9.55322265625,9.42763710021973,9.38642597198486,9.33574199676514,9.25000095367432,9.13388729095459,9.072509765625,9.04794883728027,9.04829120635986,9.12563514709473,9.16044902801514,9.22246074676514,9.34824275970459,9.51079082489014,9.705810546875,9.83320331573486,10.0145998001099,10.1436033248901,10.2637701034546,10.46923828125,10.533203125,10.621826171875,10.7787103652954,10.8356447219849,10.9641599655151,10.99951171875,11.2084474563599,11.2613763809204,11.3729982376099,11.4280757904053,11.519775390625,11.5303220748901,11.4443349838257,11.4742193222046,11.5413084030151,11.6720705032349,11.672119140625,11.6246099472046,11.6808595657349,11.7300777435303,11.8860349655151,12.0035152435303,12.0983896255493,12.1366214752197,12.1778802871704,12.1146001815796,12.05419921875,11.99560546875,11.8368654251099,11.676025390625,11.564208984375,11.4799318313599,11.4854488372803,11.49951171875,11.5184564590454,11.4610347747803,11.4317378997803,11.3093748092651,11.1609869003296,11.0528326034546,10.9493169784546,10.8800296783447,10.8087406158447,10.6893548965454,10.5691404342651,10.4927244186401,10.4720706939697,10.5237312316895,10.5704097747803,10.6106443405151,10.6322269439697,10.5746088027954,10.51708984375,10.4729490280151,10.2577629089355,10.2284669876099,10.159423828125,10.1223630905151,10.070068359375,10.07666015625,10.0950193405151,10.0980958938599,10.4578123092651,10.4485349655151,10.471923828125,10.50341796875,10.5581541061401,10.5792484283447,10.5425777435303,10.6351556777954,10.6326656341553,10.6637697219849,10.6432619094849,10.7091798782349,10.7201166152954,10.7070798873901,10.7498054504395,10.6995601654053,10.7410154342651,10.6814451217651,10.6453609466553,10.6339845657349,10.546875,10.56298828125,10.531494140625,10.5072269439697,10.4179191589355,10.39990234375,10.3992195129395,10.3330564498901,10.289794921875,10.1985836029053,10.10009765625,10.05615234375,10.0706548690796,10.116943359375,10.1634283065796,10.21728515625,10.200439453125,10.1761226654053,9.91840839385986,9.78305625915527,9.78818321228027,9.79296875,9.81889629364014,9.882568359375,9.84218692779541,9.82670974731445,9.82177734375,9.87949180603027,9.95341777801514,9.97924900054932,9.98486328125,9.97504901885986,9.95468711853027,9.86992168426514,9.78207969665527,9.73305606842041,9.70551776885986,9.676513671875,9.63120079040527,9.70248985290527,9.75444316864014,9.81381797790527,9.81645488739014,9.89453125,9.84750938415527,9.63305759429932,9.59760761260986,9.55644512176514,9.45331954956055,9.36074161529541,9.263671875,9.21518516540527,9.15458965301514,9.09511756896973,9.03525447845459,8.96577167510986,8.94130802154541,8.84335899353027,8.72538948059082,8.600341796875,8.59746074676514,8.546142578125,8.50869083404541,8.410400390625,8.48759746551514,8.49311447143555,8.57880878448486,8.59213924407959,8.54726505279541,8.61025428771973,8.62875938415527,8.61699199676514,8.54931640625],[18.40283203125,18.3991203308105,18.4116687774658,18.4425296783447,18.4462890625,18.4381351470947,18.40283203125],[18.464599609375,18.4574222564697,18.458984375,18.473388671875,18.5130863189697,18.5174789428711,18.464599609375],[18.7405757904053,18.7071285247803,18.7077159881592,18.730712890625,18.7326164245605,18.7385749816895,18.7511730194092,18.752685546875,18.7405757904053],[17.7943363189697,17.7501945495605,17.7501945495605,17.7061023712158,17.7017097473145,17.7722644805908,17.7943363189697],[18.3543453216553,18.327880859375,18.3315906524658,18.3411140441895,18.3719730377197,18.3543453216553],[18.330078125,18.3212890625,18.3675765991211,18.3852043151855,18.3742179870605,18.330078125],[8.68281269073486,8.68051719665527,8.700927734375,8.766357421875,8.72299861907959,8.70112323760986,8.68281269073486],[10.3908205032349,10.3411140441895,10.224853515625,10.1107416152954,10.06103515625,10.0291996002197,10.242919921875,10.3354005813599,10.37109375,10.3685064315796,10.4269533157349,10.4283695220947,10.3908205032349],[10.3971672058105,10.3365726470947,10.3874998092651,10.4203128814697,10.4461908340454,10.4457025527954,10.3971672058105],[20.7470207214355,20.7430667877197,20.82421875,20.8670406341553,20.8368167877197,20.8172836303711,20.7997570037842,20.7610340118408,20.7470207214355],[20.8157215118408,20.79638671875,20.84375,20.8642101287842,20.9279308319092,20.8582515716553,20.8157215118408],[20.9266128540039,20.9005374908447,20.9034652709961,20.9523429870605,21.0128421783447,21.0340328216553,20.981201171875,20.9266128540039],[21.2167949676514,21.0916500091553,21.0936527252197,21.2353019714355,21.2689437866211,21.2204113006592,21.2167949676514],[21.5079574584961,21.4989261627197,21.4971179962158,21.4058589935303,21.3680648803711,21.3362312316895,21.2848148345947,21.1941394805908,21.1284656524658,21.05517578125,20.94873046875,20.9595718383789,20.999267578125,20.9912109375,20.9713878631592,20.9740734100342,20.9500007629395,20.95751953125,20.9911136627197,20.9999027252197,21.0002918243408,20.9604988098145,20.80615234375,20.7350597381592,20.5265617370605,20.3921871185303,20.2888660430908,20.2059097290039,19.9920406341553,19.9873523712158,19.9390621185303,19.5874500274658,19.4669914245605,19.3788566589355,19.294189453125,19.1277847290039,19.0571765899658,18.96630859375,18.779296875,18.7462882995605,18.6458492279053,18.5741691589355,18.5024909973145,18.3163585662842,18.2594242095947,18.220703125,18.0531730651855,17.9464340209961,17.8736820220947,17.7195796966553,17.7650394439697,17.7468757629395,17.6627922058105,17.3671875,17.2213859558105,17.0555191040039,16.89794921875,16.7937507629395,16.642578125,16.608642578125,16.5680675506592,16.4878425598145,16.403076171875,16.3224620819092,16.3096199035645,16.3293933868408,16.3371086120605,16.3311042785645,16.2427234649658,16.1636714935303,16.0910625457764,16.1007804870605,16.0897941589355,16.0290546417236,15.9890632629395,15.8724613189697,15.7626953125,15.5847177505493,15.4835929870605,15.4266109466553,15.3779296875,15.1805171966553,15.0014638900757,14.8028316497803,14.7161617279053,14.5525875091553,14.3841304779053,14.2704591751099,14.154296875,14.0966796875,14.0534172058105,13.8564453125,13.7650394439697,13.854736328125,13.5905275344849,13.4507808685303,13.2793464660645,13.2191886901855,13.0254878997803,12.9559574127197,12.7190437316895,12.599609375,12.6707525253296,12.7519044876099,12.7090330123901,12.6458005905151,12.3911628723145,12.4153804779053,12.0729007720947,11.9928712844849,11.9545412063599,11.90869140625,11.9588384628296,12.0104484558105,11.9990234375,11.9724607467651,11.9120121002197,11.8371086120605,11.7734375,11.724853515625,11.6647462844849,11.60107421875,11.5926761627197,11.4683589935303,11.3363761901855,11.3154296875,11.1992673873901,11.1559572219849,11.0407228469849,10.9342775344849,10.920166015625,10.8972654342651,10.7203607559204,10.7000970840454,10.5554685592651,10.48583984375,10.4586429595947,10.3983888626099,10.4198732376099,10.4715814590454,10.4983396530151,10.5562992095947,10.6309576034546,10.6605463027954,10.6183099746704,10.4407224655151,10.4003419876099,10.3828125,10.4333009719849,10.53564453125,10.4649410247803,10.4562501907349,10.4620599746704,10.4443845748901,10.3761224746704,10.3389644622803,10.2958011627197,10.2889156341553,10.3041019439697,10.2982912063599,10.2317390441895,10.1933107376099,10.1511716842651,10.116455078125,10.0602054595947,9.99140644073486,9.94873046875,9.90107345581055,9.85986328125,9.86806583404541,9.93964862823486,10.2216806411743,10.14208984375,9.82124042510986,9.71562576293945,9.64111328125,9.60356426239014,9.55942344665527,9.55610275268555,9.67543983459473,9.96171951293945,10.000732421875,9.673583984375,9.59414005279541,9.50234413146973,9.44780254364014,9.396728515625,9.09321308135986,8.96240234375,8.80112266540527,8.71132850646973,8.62919998168945,8.583251953125,8.59765625,8.74663066864014,8.80185604095459,9.18549823760986,9.60615253448486,9.81626033782959,9.86865234375,9.90097618103027,9.94526386260986,9.99570274353027,10.0674314498901,10.1005859375,10.1147947311401,10.2027339935303,10.1991214752197,10.169921875,10.2076654434204,10.2668943405151,10.3399906158447,10.4112300872803,10.42236328125,10.4633302688599,10.5159664154053,10.5232419967651,10.5208005905151,10.5344724655151,10.5902347564697,10.6619138717651,10.7016611099243,10.7337894439697,10.8093748092651,10.8868646621704,10.911376953125,10.8975591659546,10.8614740371704,10.8451652526855,10.9400873184204,10.9516115188599,10.951416015625,10.9688959121704,10.9940433502197,10.989990234375,10.9260740280151,10.8635749816895,10.8584966659546,10.8851566314697,10.851806640625,10.7972650527954,10.794921875,10.9219732284546,11.0123043060303,11.037109375,11.0786628723145,11.2448244094849,11.2942876815796,11.3724126815796,11.487060546875,11.5591306686401,11.601318359375,11.635009765625,11.6483879089355,11.6529293060303,11.6824712753296,11.7580080032349,11.7512693405151,11.7083492279053,11.6818351745605,11.6870107650757,11.6978025436401,11.7383785247803,11.9117183685303,11.9484376907349,11.9655275344849,11.9691896438599,11.9792966842651,12.0523433685303,12.0774908065796,12.1758794784546,12.2770509719849,12.3040046691895,12.3215827941895,12.3190431594849,12.260498046875,12.2957029342651,12.3645505905151,12.4317865371704,12.5399904251099,12.7059078216553,12.8357419967651,12.9331045150757,13.0303707122803,13.2254400253296,13.4377927780151,13.5216789245605,13.6541986465454,13.8156251907349,13.9930181503296,14.0194826126099,14.0688972473145,14.1266117095947,14.307861328125,14.3687009811401,14.4512205123901,14.5457515716553,14.6499519348145,14.705078125,14.8173837661743,14.8718271255493,14.9159173965454,14.979884147644,15.0214347839355,15.0570306777954,15.1184568405151,15.1898441314697,15.2552242279053,15.3098640441895,15.3916006088257,15.4658212661743,15.5604972839355,15.6187009811401,15.6780757904053,15.7472658157349,15.8024892807007,15.838623046875,15.9217290878296,15.9516611099243,15.9978513717651,16.0430183410645,16.0673828125,16.0840339660645,16.1363277435303,16.2798347473145,16.3118171691895,16.3531246185303,16.3965339660645,16.515625,16.5262699127197,16.4903316497803,16.4525394439697,16.458984375,16.4926280975342,16.5379390716553,16.60009765625,16.6507320404053,16.821044921875,16.8766098022461,16.9541015625,16.9818363189697,17.0025386810303,17.1437015533447,17.216796875,17.415283203125,17.4469718933105,17.5286636352539,17.6444320678711,17.7378425598145,17.8344249725342,17.9182624816895,17.9836921691895,18.0774402618408,18.154296875,18.1792488098145,18.1896495819092,18.2353515625,18.3387222290039,18.4052734375,18.4500980377197,18.4962387084961,18.5730457305908,18.6167964935303,18.6509780883789,18.6788578033447,18.7283210754395,18.8034191131592,18.8606452941895,18.93408203125,18.9838371276855,19.195556640625,19.2309074401855,19.2685070037842,19.3049793243408,19.3399906158447,19.3660640716553,19.4204597473145,19.4825687408447,19.5641593933105,19.646484375,19.6751480102539,19.6784191131592,19.6808586120605,19.6854972839355,19.6105480194092,19.6187496185303,19.7547359466553,19.8361320495605,19.9040050506592,19.9471683502197,20.0181140899658,20.0828113555908,20.1690921783447,20.2024402618408,20.2168464660645,20.205322265625,20.2247066497803,20.2890129089355,20.3285160064697,20.37451171875,20.4136714935303,20.4247550964355,20.44140625,20.4857425689697,20.5295906066895,20.5548820495605,20.6002445220947,20.6466808319092,20.68798828125,20.7337398529053,20.8210945129395,20.9139652252197,20.9455070495605,20.9412117004395,20.8614253997803,20.8095226287842,20.7169437408447,20.6970691680908,20.7378425598145,20.7798347473145,20.8406257629395,20.8916492462158,21.2025871276855,21.2659187316895,21.3375015258789,21.43994140625,21.50634765625,21.5697746276855,21.626220703125,21.6813468933105,21.7129383087158,21.7222652435303,21.7347640991211,21.807373046875,21.7979488372803,21.7096672058105,21.6779289245605,21.662109375,21.676025390625,21.71142578125,21.7713375091553,21.8517570495605,21.904296875,21.957763671875,22.0271492004395,22.1781749725342,22.2840328216553,22.3791980743408,22.4146480560303,22.4660148620605,22.5459957122803,22.6466312408447,22.7080078125,22.7328128814697,22.7509288787842,22.7410163879395,22.7003917694092,22.6484870910645,22.587158203125,22.525390625,22.4661617279053,22.4482440948486,22.4529781341553,22.4975090026855,22.542236328125,22.5929679870605,22.6385250091553,22.7135257720947,22.764404296875,22.769775390625,22.7546882629395,22.5974102020264,22.5879878997803,22.611572265625,22.7344245910645,22.7820320129395,22.77001953125,22.5382328033447,22.5400886535645,22.5504875183105,22.5861320495605,22.6663589477539,22.7522945404053,22.8001461029053,22.8094234466553,22.7685050964355,22.7120113372803,22.7040519714355,22.8041019439697,22.8200206756592,22.8182125091553,22.8222160339355,22.8604984283447,22.9111328125,23.0107917785645,23.1001949310303,23.1363754272461,23.1605472564697,23.1943359375,23.2810554504395,23.3221206665039,23.34521484375,23.3076648712158,23.2353515625,23.1808586120605,23.1219730377197,23.0726547241211,23.0299320220947,22.9693355560303,22.9228038787842,22.9249515533447,22.9374504089355,22.9747543334961,22.9755363464355,22.9700679779053,22.9551277160645,22.8694343566895,22.8574714660645,22.8634757995605,22.8938961029053,22.9083499908447,22.874267578125,22.7789058685303,22.7109375,22.6377429962158,22.5860347747803,22.5732421875,22.5013675689697,22.3954105377197,22.3416996002197,22.3245105743408,22.2886238098145,22.241455078125,22.136474609375,22.0182132720947,21.9789066314697,21.9861812591553,22.0003414154053,21.9819812774658,21.9512691497803,21.9201183319092,21.9239253997803,21.8934078216553,21.8348636627197,21.794189453125,21.7170906066895,21.7106437683105,21.60888671875,21.6422843933105,21.5983390808105,21.6139163970947,21.655029296875,21.6451663970947,21.5604000091553,21.5079574584961],[-20.1866207122803,-20.2417964935303,-20.2411136627197,-20.2021484375,-20.1533203125,-20.1447257995605,-20.14794921875,-20.1866207122803],[-19.5401363372803,-19.6488285064697,-19.62353515625,-19.5450191497803,-19.4763660430908,-19.3447265625,-19.32177734375,-19.3292961120605,-19.4578113555908,-19.5401363372803],[-18.9402332305908,-18.9885749816895,-18.9832992553711,-18.8712882995605,-18.8251953125,-18.7076168060303,-18.6437492370605,-18.6174812316895,-18.6310558319092,-18.72509765625,-18.7633781433105,-18.7957019805908,-18.8667964935303,-18.9402332305908],[-17.5421886444092,-17.6846675872803,-17.6958980560303,-17.7980480194092,-17.8072261810303,-17.78076171875,-17.7457046508789,-17.7060546875,-17.6980457305908,-17.7169914245605,-17.7105464935303,-17.6448230743408,-17.55224609375,-17.544921875,-17.5439453125,-17.5520515441895,-17.5421886444092],[-16.77880859375,-16.7936515808105,-16.8350582122803,-16.7877922058105,-16.80615234375,-16.8040027618408,-16.7657222747803,-16.6900386810303,-16.6369152069092,-16.5999031066895,-16.5938472747803,-16.6396484375,-16.6708011627197,-16.6841793060303,-16.7587890625,-16.77880859375],[-16.3365230560303,-16.3467769622803,-16.3156242370605,-16.2722644805908,-16.2287101745605,-16.1964836120605,-16.1812496185303,-16.0816402435303,-16.1198234558105,-16.2313461303711,-16.2649402618408,-16.2987308502197,-16.3365230560303],[-16.0958976745605,-16.1175785064697,-16.1175785064697,-16.1662101745605,-16.2632808685303,-16.2605457305908,-16.3136711120605,-16.3405265808105,-16.3946285247803,-16.44970703125,-16.5164070129395,-16.4986324310303,-16.5743160247803,-16.5549793243408,-16.5152359008789,-16.501953125,-16.4005851745605,-16.2457046508789,-16.1544914245605,-16.1155281066895,-16.1496105194092,-16.1552734375,-16.0804672241211,-15.9285154342651,-15.8850584030151,-15.8767576217651,-15.9166984558105,-16.0958976745605],[-15.7241220474243,-15.7500972747803,-15.6852531433105,-15.62255859375,-15.64501953125,-15.7241220474243],[-15.9704093933105,-15.9716806411743,-15.9256839752197,-15.6808595657349,-15.4618167877197,-15.508204460144,-15.8922843933105,-15.9551753997803,-15.9704093933105],[-15.4359369277954,-15.4818363189697,-15.4774408340454,-15.4515628814697,-15.31201171875,-15.2832021713257,-15.4359369277954],[-15.3287115097046,-15.3906259536743,-15.3189449310303,-15.0166015625,-14.9885730743408,-14.9864253997803,-15.1968755722046,-15.3287115097046],[-14.8268547058105,-15.1574230194092,-15.15673828125,-15.13916015625,-15.0617179870605,-14.940037727356,-14.9226551055908,-14.9356451034546,-14.9744148254395,-15.07177734375,-15.1255855560303,-15.1353511810303,-15.3897466659546,-15.4430665969849,-15.4857425689697,-15.5808601379395,-15.5780277252197,-15.6348628997803,-15.6311521530151,-15.5667972564697,-15.515625,-15.4060554504395,-15.2115230560303,-14.8500986099243,-14.759765625,-14.6417961120605,-14.6365232467651,-14.7350587844849,-14.8268547058105],[-14.2609376907349,-14.3116216659546,-14.294921875,-14.2815427780151,-14.1974611282349,-14.16845703125,-14.1421871185303,-14.1837892532349,-14.2609376907349],[-13.9072265625,-13.9170904159546,-13.9093742370605,-13.7883787155151,-13.7480459213257,-13.70947265625,-13.7766599655151,-13.81396484375,-13.845703125,-13.8731451034546,-13.8845710754395,-13.9072265625],[-14.318359375,-14.3249025344849,-14.3119134902954,-14.2554683685303,-14.2316408157349,-14.2425775527954,-14.2841796875,-14.30322265625,-14.318359375],[-13.3328123092651,-13.3409175872803,-13.3016605377197,-13.2425785064697,-13.2216796875,-13.2681636810303,-13.3328123092651],[-14.0464849472046,-14.0472660064697,-14.0020503997803,-14.0016593933105,-13.9068365097046,-13.8571290969849,-13.8244142532349,-13.80712890625,-13.8791990280151,-13.9430665969849,-13.9499025344849,-13.9776363372803,-14.0224609375,-14.0464849472046],[-13.4652338027954,-13.5595703125,-13.6846685409546,-13.8042964935303,-13.7747068405151,-13.8001956939697,-13.7916994094849,-13.6448240280151,-13.5787105560303,-13.5167970657349,-13.5238285064697,-13.4828119277954,-13.4652338027954],[12.6368160247803,12.6248044967651,12.6594247817993,12.6640138626099,12.5527839660645,12.5234375,12.4833011627197,12.4466304779053,12.3606443405151,12.3189945220947,12.34228515625,12.4253416061401,12.5331544876099,12.6018552780151,12.6333503723145,12.66357421875,12.7157716751099,12.7073736190796,12.6368160247803],[13.7042961120605,13.6736326217651,13.7529783248901,13.769287109375,13.76611328125,13.7042961120605],[13.9714841842651,13.9502439498901,13.9548826217651,14.0079107284546,14.0674810409546,14.0122556686401,13.9714841842651],[15.3034181594849,15.2812013626099,15.320068359375,15.4073238372803,15.432520866394,15.3679685592651,15.332275390625,15.3034181594849],[16.6483879089355,16.4703617095947,16.3912601470947,16.2935543060303,16.17138671875,15.9568367004395,15.7605962753296,15.655517578125,15.5859384536743,15.5356941223145,15.4592781066895,15.4401359558105,15.3791017532349,15.3368158340454,15.2262687683105,15.1407709121704,15.0381841659546,14.9271974563599,14.8510255813599,14.8281259536743,14.7224130630493,14.637791633606,14.5000495910645,14.4564456939697,14.3550291061401,14.2674808502197,14.1238775253296,14.0501461029053,14.0462408065796,14.0059080123901,13.9976568222046,14.048095703125,14.0128421783447,13.9569330215454,13.8584470748901,13.66162109375,13.609375,13.5474605560303,13.465576171875,13.4327144622803,13.4155769348145,13.423828125,13.394287109375,13.3387212753296,13.2334957122803,13.0670404434204,12.998291015625,12.9385738372803,12.8158693313599,12.7841796875,12.7637691497803,12.8172359466553,12.6691408157349,12.6446294784546,12.638671875,12.607666015625,12.6165037155151,12.6744136810303,12.7444820404053,12.6988277435303,12.8390140533447,13.26708984375,13.6398429870605,13.6925296783447,13.8589353561401,14.010986328125,14.2036619186401,14.341552734375,14.4831056594849,14.5208015441895,14.5548839569092,14.7731437683105,14.8173837661743,14.8630857467651,14.8980464935303,14.9385747909546,15.0055665969849,15.1329584121704,15.2328128814697,15.3263177871704,15.2935552597046,15.2657222747803,15.3262701034546,15.3716306686401,15.6546382904053,16.0320320129395,16.3717765808105,16.5090827941895,16.5503902435303,16.5866203308105,16.6641597747803,16.6894054412842,16.77099609375,16.8118152618408,16.8467769622803,16.9419918060303,17.0624504089355,17.1129875183105,17.2050304412842,17.2392578125,17.2664546966553,17.32470703125,17.359375,17.456787109375,17.4860363006592,17.5162582397461,17.5159168243408,17.4987316131592,17.471435546875,17.421875,17.3655281066895,17.3441410064697,17.349609375,17.32470703125,17.3383312225342,17.3674812316895,17.3655281066895,17.3985366821289,17.4143562316895,17.4043464660645,17.4316902160645,17.4295883178711,17.4274406433105,17.4233894348145,17.4062004089355,17.3197746276855,17.3020515441895,17.2784175872803,17.253173828125,17.2312984466553,17.2516593933105,17.2685546875,17.2655754089355,17.2121086120605,17.0790042877197,16.9534664154053,16.9466800689697,16.9939460754395,17.0604000091553,17.1118659973145,17.3161125183105,17.4483394622803,17.5968265533447,17.7210922241211,17.8858394622803,17.9769535064697,18.1569328308105,18.22705078125,18.3624019622803,18.4952144622803,18.581787109375,18.621337890625,18.6553230285645,18.6953125,18.7352523803711,18.7777824401855,18.8252925872803,18.8578624725342,18.8993663787842,18.9338874816895,18.9645519256592,18.9961433410645,18.8962879180908,18.79638671875,18.6964855194092,18.5965824127197,18.4966793060303,18.3968257904053,18.2969245910645,18.1970710754395,18.09716796875,17.997314453125,17.8974132537842,17.7975101470947,17.6976070404053,17.5977535247803,17.4978523254395,17.39794921875,17.3003902435303,17.2679195404053,17.1756839752197,17.0438480377197,16.9120597839355,16.7802238464355,16.6483879089355],[-46.9442367553711,-46.962890625,-46.9464874267578,-46.9080085754395,-46.8489265441895,-46.8240242004395,-46.8375015258789,-46.8854484558105,-46.9016609191895,-46.9442367553711],[-22.4020500183105,-22.4546871185303,-22.4786128997803,-22.6175785064697,-22.8250980377197,-23.0166988372803,-23.2794933319092,-23.42578125,-23.4823226928711,-23.5529289245605,-23.6742191314697,-23.7430667877197,-23.7945308685303,-23.8921871185303,-24.0402336120605,-24.2362308502197,-24.3302726745605,-24.37646484375,-24.4606437683105,-24.6382827758789,-24.8440437316895,-25.0738277435303,-25.2634773254395,-25.3594722747803,-25.6319332122803,-25.77392578125,-25.8853511810303,-25.9576187133789,-25.96875,-25.9816398620605,-25.8672847747803,-25.74658203125,-25.7429695129395,-25.7555656433105,-25.8433589935303,-25.9806632995605,-26.0977535247803,-26.21875,-26.4134769439697,-26.4554691314697,-26.6136722564697,-26.7642593383789,-26.7852554321289,-26.7923812866211,-26.9158210754395,-27.1123046875,-27.2383785247803,-27.2955074310303,-27.3099613189697,-27.3058605194092,-27.1736335754395,-26.9606456756592,-26.8174800872803,-26.8111324310303,-26.8248043060303,-26.8394546508789,-26.83349609375,-26.8616218566895,-26.8584957122803,-26.8557624816895,-26.8509769439697,-26.8493156433105,-27.0801753997803,-27.4416007995605,-27.6073246002197,-28.1997089385986,-28.4982414245605,-28.6214847564697,-28.8395519256592,-28.8837871551514,-28.912109375,-28.9371109008789,-29.3781261444092,-29.5908222198486,-29.9008769989014,-30.0710926055908,-30.4341793060303,-30.7145519256592,-30.9701156616211,-31.3220691680908,-31.423828125,-31.4704113006592,-31.6747093200684,-32.0031242370605,-32.2942390441895,-32.6246109008789,-32.7692413330078,-33.0539054870605,-33.0959968566895,-33.3605461120605,-33.5211906433105,-33.7074203491211,-33.7595710754395,-33.7113265991211,-33.7371063232422,-33.849609375,-34.0111312866211,-34.0353507995605,-34.0281257629395,-33.9607429504395,-33.9736328125,-34.0597648620605,-34.1689453125,-34.1745147705078,-34.0615234375,-33.9927711486816,-33.9851570129395,-34.0689468383789,-34.0811500549316,-34.0631866455078,-34.01025390625,-34.0100593566895,-34.0538063049316,-34.0691413879395,-34.3726577758789,-34.373046875,-34.408203125,-34.4070320129395,-34.3646469116211,-34.3674812316895,-34.3865242004395,-34.43994140625,-34.4630851745605,-34.5085945129395,-34.7857437133789,-34.7747077941895,-34.7566413879395,-34.7533226013184,-34.6056671142578,-34.6150398254395,-34.57080078125,-34.4923820495605,-34.43701171875,-34.4123039245605,-34.4168968200684,-34.35009765625,-34.34375,-34.3606414794922,-34.3640632629395,-34.2964820861816,-34.25390625,-34.1884765625,-34.1082038879395,-34.0826148986816,-34.0718765258789,-34.0773429870605,-34.0859375,-34.1092796325684,-34.1680641174316,-34.3468780517578,-34.2956047058105,-34.1884765625,-34.07421875,-33.9390640258789,-33.8877906799316,-33.7964820861816,-33.71728515625,-33.5144538879395,-33.4216804504395,-33.3587875366211,-33.2073249816895,-33.15234375,-33.04638671875,-32.9615211486816,-32.8274421691895,-32.75048828125,-32.7085952758789,-32.7750968933105,-32.7491226196289,-32.6521492004395,-32.5049781799316,-32.26953125,-32.1224632263184,-31.7424812316895,-31.6551742553711,-31.3832035064697,-31.0190448760986,-30.44482421875,-30.0998039245605,-29.4034175872803,-29.0093746185303,-28.6415042877197,-28.6175765991211,-28.5728511810303,-28.4878902435303,-28.4649410247803,-28.4754886627197,-28.4521484375,-28.3947277069092,-28.3408203125,-28.2645511627197,-28.2189445495605,-28.1279296875,-28.0696296691895,-28.0310554504395,-28.0822257995605,-28.1325187683105,-28.1988296508789,-28.2308597564697,-28.2286128997803,-28.2694339752197,-28.3532238006592,-28.4139652252197,-28.45166015625,-28.5011730194092,-28.5626945495605,-28.62109375,-28.6981430053711,-28.7430648803711,-28.7683601379395,-28.7769527435303,-28.8113288879395,-28.8716812133789,-28.8862285614014,-28.8552742004395,-28.869140625,-28.9279308319092,-28.9387702941895,-28.9016590118408,-28.8479499816895,-28.7777347564697,-28.7333011627197,-28.7144527435303,-28.66162109375,-28.5746097564697,-28.50390625,-28.4494152069092,-28.4512691497803,-28.3103523254395,-27.8655281066895,-27.4207019805908,-26.9759769439697,-26.5311527252197,-26.0863285064697,-25.6416015625,-25.19677734375,-24.7767562866211,-24.8070316314697,-25.0298824310303,-25.1470699310303,-25.2212905883789,-25.4912109375,-25.7332038879395,-25.9156246185303,-25.9990234375,-26.08056640625,-26.12060546875,-26.1649417877197,-26.26416015625,-26.3401355743408,-26.44384765625,-26.5808601379395,-26.7421875,-26.8224620819092,-26.8488273620605,-26.8087902069092,-26.8210945129395,-26.8517589569092,-26.8328132629395,-26.8426761627197,-26.8541984558105,-26.8409175872803,-26.8068370819092,-26.7100601196289,-26.6783218383789,-26.6619148254395,-26.6358394622803,-26.5801753997803,-26.3888683319092,-26.2190437316895,-26.1784172058105,-26.1327152252197,-26.0711898803711,-25.8573246002197,-25.6791019439697,-25.5951175689697,-25.4579105377197,-25.3703117370605,-25.3241214752197,-25.3123054504395,-25.2886714935303,-25.2666015625,-25.2914066314697,-25.3444328308105,-25.4339847564697,-25.5446300506592,-25.6008796691895,-25.6260738372803,-25.6348628997803,-25.6329097747803,-25.7428703308105,-25.7498054504395,-25.7831058502197,-25.8173828125,-25.8134765625,-25.7540035247803,-25.75146484375,-25.7562503814697,-25.7399425506592,-25.7144527435303,-25.6627922058105,-25.6062488555908,-25.4378910064697,-25.3023433685303,-25.146484375,-24.9352531433105,-24.7879867553711,-24.7474613189697,-24.7024421691895,-24.6714839935303,-24.6135730743408,-24.5827140808105,-24.5132808685303,-24.3955078125,-24.2971687316895,-24.2408218383789,-23.7634754180908,-23.70458984375,-23.5779285430908,-23.5244140625,-23.5234375,-23.4900379180908,-23.4242191314697,-23.3835926055908,-23.3683586120605,-23.3246097564697,-23.2526359558105,-23.2176761627197,-23.2196292877197,-23.19677734375,-23.14892578125,-23.1080074310303,-23.0739269256592,-23.0335941314697,-22.9870109558105,-22.8737297058105,-22.6936511993408,-22.5933589935303,-22.5729484558105,-22.5354499816895,-22.4808597564697,-22.3951187133789,-22.2784194946289,-22.2132816314697,-22.1939449310303,-22.1927738189697,-22.1462898254395,-22.1841793060303,-22.2911128997803,-22.3290042877197,-22.2978534698486,-22.2907218933105,-22.3078136444092,-22.3449211120605,-22.4020500183105],[-30.1019515991211,-30.0384750366211,-29.9994125366211,-29.9675769805908,-29.9190425872803,-29.8011703491211,-29.7009754180908,-29.6516609191895,-29.6188488006592,-29.56689453125,-29.4419918060303,-29.3197269439697,-29.2697277069092,-29.2184562683105,-29.1636714935303,-29.08984375,-29.0783214569092,-29.0369129180908,-28.9537105560303,-28.8814449310303,-28.7760753631592,-28.7588863372803,-28.6876964569092,-28.6467761993408,-28.5978527069092,-28.5817375183105,-28.5941410064697,-28.6158199310303,-28.7012672424316,-28.7799816131592,-28.8733406066895,-28.9090824127197,-28.9400405883789,-29.0469741821289,-29.1464824676514,-29.2361335754395,-29.2765617370605,-29.302734375,-29.3600597381592,-29.4552726745605,-29.5193367004395,-29.5541973114014,-29.5993175506592,-29.6255874633789,-29.6640605926514,-29.7537117004395,-29.8402366638184,-29.9413089752197,-30.0153312683105,-30.1056632995605,-30.1585941314697,-30.2473621368408,-30.2791976928711,-30.31591796875,-30.3252925872803,-30.3384780883789,-30.3639640808105,-30.3809585571289,-30.4112300872803,-30.4664077758789,-30.5422840118408,-30.5999984741211,-30.6238288879395,-30.6422843933105,-30.6310558319092,-30.5845699310303,-30.5250988006592,-30.4499015808105,-30.4098625183105,-30.2184581756592,-30.1475601196289,-30.1424827575684,-30.12890625,-30.123046875,-30.1287117004395,-30.1265640258789,-30.1019515991211],[-9.40742206573486,-9.43398475646973,-9.48417949676514,-9.52031326293945,-9.57363319396973,-9.62285137176514,-9.63505840301514,-9.63818454742432,-9.60263729095459,-9.603515625,-9.62617111206055,-9.68300819396973,-9.75957012176514,-9.81181621551514,-9.86220645904541,-9.95400333404541,-10.0379886627197,-10.1208982467651,-10.19970703125,-10.2346677780151,-10.3515625,-10.3913087844849,-10.4885740280151,-10.5531253814697,-10.5905275344849,-10.7831058502197,-10.8017578125,-10.8126945495605,-10.8523445129395,-10.8933591842651,-10.9150390625,-10.9811525344849,-11.0851564407349,-11.1579103469849,-11.2491207122803,-11.4039058685303,-11.4176759719849,-11.5348625183105,-11.57763671875,-11.6111326217651,-11.6908206939697,-11.7999992370605,-11.8881845474243,-12.1125974655151,-12.3083009719849,-12.3296871185303,-12.3310546875,-12.3477535247803,-12.4034175872803,-12.4604501724243,-12.4898433685303,-12.5565433502197,-12.6304683685303,-12.7013673782349,-12.8043947219849,-12.8647470474243,-12.8996095657349,-12.9894533157349,-13.0842771530151,-13.1588869094849,-13.2249994277954,-13.2574214935303,-13.3570308685303,-13.45703125,-13.5027341842651,-13.55029296875,-13.5904293060303,-13.6103515625,-13.6565427780151,-13.6884765625,-13.7102537155151,-13.7314453125,-13.7610349655151,-13.7916011810303,-13.8173818588257,-13.8838872909546,-13.9768562316895,-14.0093755722046,-14.0221681594849,-14.0237302780151,-14.0100584030151,-13.9591798782349,-13.94091796875,-14.0133790969849,-14.0849609375,-14.1224613189697,-14.2295894622803,-14.3230466842651,-14.3408203125,-14.3865232467651,-14.4144535064697,-14.4960927963257,-14.5367193222046,-14.5771484375,-14.6376943588257,-14.6946296691895,-14.753321647644,-14.8191404342651,-14.8665046691895,-14.9075193405151,-14.9903326034546,-15.0105476379395,-15.0668954849243,-15.1832027435303,-15.2888679504395,-15.3497076034546,-15.505859375,-15.64306640625,-15.6434574127197,-15.64404296875,-15.6446285247803,-15.69677734375,-15.7764644622803,-15.9011726379395,-15.9500970840454,-15.98779296875,-16.0361328125,-16.1422843933105,-16.30615234375,-16.4241199493408,-16.5321273803711,-16.5319328308105,-16.6627922058105,-16.7697257995605,-16.8961906433105,-17.0603523254395,-17.2621097564697,-17.5119132995605,-17.7283191680908,-17.9583969116211,-18.04150390625,-18.0225582122803,-17.9292964935303,-17.9117183685303,-17.9698238372803,-17.9519538879395,-17.8582019805908,-17.8241214752197,-17.8495101928711,-17.8451175689697,-17.7935543060303,-17.6343746185303,-17.5685539245605,-17.54345703125,-17.5177745819092,-17.4810543060303,-17.4895515441895,-17.5208988189697,-17.5601558685303,-17.5994129180908,-17.640625,-17.4744148254395,-17.2857418060303,-17.0752944946289,-16.9102535247803,-16.8151359558105,-16.6895503997803,-16.6281242370605,-16.59716796875,-16.2627925872803,-15.95556640625,-15.7241220474243,-15.4032220840454,-15.0823249816895,-14.7614250183105,-14.4405279159546,-14.1196279525757,-13.7987308502197,-13.4777336120605,-13.1568365097046,-13.0009765625,-13.0009765625,-13.0009765625,-13.0009765625,-13.0009765625,-13.0009765625,-13.0009765625,-13.0009765625,-12.9982414245605,-12.9884767532349,-12.9569339752197,-12.7990236282349,-12.7432613372803,-12.6361322402954,-12.5437498092651,-12.4221677780151,-12.3506832122803,-12.1177730560303,-11.9878902435303,-11.8529300689697,-11.7249994277954,-11.6358404159546,-11.5872068405151,-11.5176753997803,-11.4391593933105,-11.4053707122803,-11.3741207122803,-11.3156251907349,-11.1847658157349,-11.0028324127197,-10.8717775344849,-10.8791017532349,-10.8915033340454,-10.9556646347046,-11.0259761810303,-11.0299816131592,-11.07177734375,-11.1298828125,-11.2551755905151,-11.3193359375,-11.3712892532349,-11.4170894622803,-11.4476556777954,-11.4384765625,-11.3529291152954,-11.3377933502197,-11.3211917877197,-11.2991209030151,-11.2600584030151,-11.2429685592651,-11.21240234375,-11.21240234375,-11.2369146347046,-11.3254880905151,-11.4049806594849,-11.5535163879395,-11.623046875,-11.6735353469849,-11.6998043060303,-11.75341796875,-11.744140625,-11.8201169967651,-11.8552742004395,-11.89013671875,-11.9032220840454,-11.9298820495605,-11.9478521347046,-11.9720706939697,-11.9759769439697,-11.9652347564697,-11.9435548782349,-11.9193353652954,-11.8988285064697,-11.8246097564697,-11.6637697219849,-11.6159172058105,-11.59375,-11.5791988372803,-11.6050777435303,-11.7834959030151,-11.9445314407349,-12.0796871185303,-12.1953134536743,-12.22705078125,-12.2667970657349,-12.2808589935303,-12.2848625183105,-12.3681650161743,-12.4345703125,-12.4820308685303,-12.5180654525757,-12.5774412155151,-12.6233396530151,-12.7421875,-12.8361320495605,-12.8541011810303,-12.861328125,-12.9254884719849,-12.9819326400757,-13.1194334030151,-13.2146482467651,-13.30712890625,-13.3688478469849,-13.3951168060303,-13.3983402252197,-13.3708009719849,-13.3228521347046,-13.2679681777954,-13.2489261627197,-13.2605466842651,-13.2985343933105,-13.3729486465454,-13.4143562316895,-13.4538087844849,-13.4380855560303,-13.3927736282349,-13.3697261810303,-13.16748046875,-12.9920892715454,-12.8270511627197,-12.6258783340454,-12.4505863189697,-12.30615234375,-12.1554679870605,-12.1640625,-12.1983394622803,-12.2024412155151,-12.2282228469849,-12.2668943405151,-12.3175773620605,-12.3861331939697,-12.41845703125,-12.4312496185303,-12.4047842025757,-12.3702154159546,-12.3488283157349,-12.2578125,-12.1205081939697,-12.05126953125,-11.9081048965454,-11.8791990280151,-11.8121099472046,-11.6983394622803,-11.6228513717651,-11.5666990280151,-11.4830083847046,-11.3543949127197,-11.1095695495605,-10.9332036972046,-10.8023433685303,-10.6692390441895,-10.5501956939697,-10.3973627090454,-10.31298828125,-10.0988283157349,-9.91875076293945,-9.83124923706055,-9.67880821228027,-9.51005840301514,-9.27499961853027,-9.22480487823486,-9.16943454742432,-9.07226657867432,-9.0146484375,-8.9326171875,-8.89101600646973,-8.78584003448486,-8.70058631896973,-8.59023380279541,-8.48544979095459,-8.46494102478027,-8.42783164978027,-8.38691425323486,-8.34375,-8.30029296875,-8.25820255279541,-8.22002029418945,-8.19365215301514,-8.26582050323486,-8.38554668426514,-8.47373104095459,-8.55097675323486,-8.59765625,-8.61191368103027,-8.60703086853027,-8.65390586853027,-8.71328163146973,-8.80546855926514,-8.86328125,-8.90878868103027,-8.91435527801514,-8.90322208404541,-8.90224647521973,-8.92197322845459,-8.94218730926514,-9.01943397521973,-9.05400371551514,-9.0673828125,-9.07333946228027,-9.12558650970459,-9.13486289978027,-9.15634727478027,-9.21269607543945,-9.27050685882568,-9.322265625,-9.380859375,-9.40742206573486],[-22.4020500183105,-22.3449211120605,-22.3078136444092,-22.2907218933105,-22.2978534698486,-22.3290042877197,-22.2911128997803,-22.1841793060303,-22.1462898254395,-22.1927738189697,-22.1939449310303,-22.15771484375,-22.0794925689697,-22.0657215118408,-22.0474605560303,-22.0183601379395,-21.9812488555908,-21.93994140625,-21.8113288879395,-21.796875,-21.7814464569092,-21.7660160064697,-21.7076168060303,-21.6512699127197,-21.58935546875,-21.5730457305908,-21.55419921875,-21.5067367553711,-21.3590831756592,-21.2615242004395,-21.1110363006592,-21.0642566680908,-20.94482421875,-20.8483390808105,-20.7664070129395,-20.6897468566895,-20.5945320129395,-20.5306644439697,-20.5030269622803,-20.4835929870605,-20.4748058319092,-20.4787101745605,-20.3818359375,-20.2320308685303,-20.1457996368408,-20.1009769439697,-20.05419921875,-19.9901371002197,-19.8927745819092,-19.7486324310303,-19.5693359375,-19.5382823944092,-19.3699207305908,-19.0817394256592,-18.9856452941895,-18.9386711120605,-18.7970695495605,-18.7235355377197,-18.6492176055908,-18.4417972564697,-18.3512706756592,-18.2349605560303,-18.1419906616211,-18.1044921875,-18.0412101745605,-17.9690418243408,-17.9152336120605,-17.8430671691895,-17.7935543060303,-17.8451175689697,-17.8495101928711,-17.8241214752197,-17.8582019805908,-17.9519538879395,-17.9698238372803,-17.9117183685303,-17.9292964935303,-18.0225582122803,-18.04150390625,-17.9583969116211,-17.7283191680908,-17.5119132995605,-17.2621097564697,-17.0603523254395,-16.8961906433105,-16.7697257995605,-16.6627922058105,-16.5319328308105,-16.5321273803711,-16.4241199493408,-16.30615234375,-16.1422843933105,-16.0361328125,-15.98779296875,-15.9500970840454,-15.9011726379395,-15.7764644622803,-15.69677734375,-15.6446285247803,-15.64404296875,-15.6434574127197,-15.64306640625,-15.8007822036743,-15.9782218933105,-15.9953126907349,-15.9992179870605,-16.01171875,-16.0236320495605,-16.15234375,-16.1796875,-16.2141609191895,-16.4288082122803,-16.44873046875,-16.5157241821289,-16.5894546508789,-16.6776371002197,-16.6976566314697,-16.7041988372803,-16.7123050689697,-16.7759761810303,-16.8835945129395,-17.0377922058105,-17.2515621185303,-17.4375,-17.7654304504395,-18.0829105377197,-18.1962890625,-18.271484375,-18.3125972747803,-18.3595714569092,-18.49267578125,-18.6329097747803,-18.6890621185303,-18.728515625,-18.763671875,-18.8284187316895,-18.8684577941895,-18.94091796875,-19.0018558502197,-19.0243167877197,-19.0587882995605,-19.1043949127197,-19.1524410247803,-19.2414054870605,-19.3887691497803,-19.5582027435303,-19.6680660247803,-19.79541015625,-19.8738288879395,-19.93017578125,-19.98486328125,-20.2171878814697,-20.3615226745605,-20.51611328125,-20.6130847930908,-20.6597652435303,-20.7129878997803,-20.8289051055908,-20.9500980377197,-21.1365242004395,-21.2970714569092,-21.3118152618408,-21.3348617553711,-21.5154304504395,-21.6980476379395,-21.83154296875,-21.9833984375,-22.1535148620605,-22.298828125,-22.4020500183105],[52.4276847839355,52.4853515625,52.5631828308105,52.6092758178711,52.629638671875,52.6584968566895,52.6843757629395,52.7576637268066,52.7818374633789,52.8173332214355,52.8514175415039,52.8456535339355,52.857177734375,52.8744621276855,52.9026374816895,53.0419464111328,53.064697265625,53.0272445678711,52.9667015075684,52.9234390258789,52.8861808776855,52.8359870910645,52.7857398986816,52.7303733825684,52.6698265075684,52.6266098022461,52.6237335205078,52.5655746459961,52.5204086303711,52.4513664245605,52.4307136535645,52.4045906066895,52.3643074035645,52.3441429138184,52.29150390625,52.2519035339355,52.4276847839355],[52.3018569946289,52.2976570129395,52.333740234375,52.3583488464355,52.3787612915039,52.4903297424316,52.5577163696289,52.5875968933105,52.5822257995605,52.5645980834961,52.5354499816895,52.5022468566895,52.4657707214355,52.4290046691895,52.3759307861328,52.3244132995605,52.3018569946289],[41.8983879089355,41.9062004089355,41.9054679870605,41.9007301330566,41.8975601196289,41.8983879089355]]}}},{"type":"Patches","id":"c9f1eaa89be444042d91237f331ac494","attributes":{"id":"c9f1eaa89be444042d91237f331ac494","tags":[],"line_color":{"units":"data","value":"gray"},"fill_color":{"units":"data","value":"gray"},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"xs":{"units":"data","field":"xs"},"ys":{"units":"data","field":"ys"}}},{"type":"Patches","id":"550dcc9b740049499f5d8fa3f7aceb1f","attributes":{"id":"550dcc9b740049499f5d8fa3f7aceb1f","tags":[],"line_color":{"units":"data","value":"#e1e1e1"},"fill_color":{"units":"data","value":"#e1e1e1"},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"xs":{"units":"data","field":"xs"},"ys":{"units":"data","field":"ys"}}},{"type":"Patches","id":"743f97210278929d542a8e0ca085443f","attributes":{"id":"743f97210278929d542a8e0ca085443f","tags":[],"line_color":{"units":"data","value":"gray"},"fill_color":{"units":"data","value":"gray"},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":1},"xs":{"units":"data","field":"xs"},"ys":{"units":"data","field":"ys"}}},{"type":"GlyphRenderer","id":"d74047225abb80aa550c08706d1e8ee5","attributes":{"id":"d74047225abb80aa550c08706d1e8ee5","tags":[],"selection_glyph":null,"nonselection_glyph":{"type":"Patches","id":"550dcc9b740049499f5d8fa3f7aceb1f"},"hover_glyph":{"type":"Patches","id":"743f97210278929d542a8e0ca085443f"},"name":null,"data_source":{"type":"ColumnDataSource","id":"c28bc5d1124875495f93d9a639613f8a"},"glyph":{"type":"Patches","id":"c9f1eaa89be444042d91237f331ac494"}}},{"type":"HoverTool","id":"33b3db0d39b742e31fada3b78ac8a341","attributes":{"id":"33b3db0d39b742e31fada3b78ac8a341","tags":[],"plot":{"type":"Plot","id":"f77af90710cde2b4e2ddd44c50c3e2ba","subtype":"Figure"},"renderers":[{"type":"GlyphRenderer","id":"d0d5df621e402ab18060410f73ca9d28"}],"names":[],"anchor":"center","attachment":"horizontal","line_policy":"prev","mode":"mouse","point_policy":"snap_to_data","tooltips":[["name","@hover_col_1"],["country.etc","@hover_col_2"],["population","@hover_col_3"]]}},{"type":"ColumnDataSource","id":"2a1071dfa6c0806319defe035a7a31f9","attributes":{"id":"2a1071dfa6c0806319defe035a7a31f9","tags":[],"column_names":["x","y","hover_col_1","hover_col_2","hover_col_3"],"selected":[],"data":{"x":[35.93,54.37,7.17,-0.2,-130.1,38.74,144.75,3.04,-169.92,4.89,1.51,32.85,47.51,-171.76,58.38,38.94,71.47,-57.63,23.73,-159.76,44.44,172.99,49.86,-7.99,114.95,100.5,18.56,-16.6,-61.72,-62.73,35.5,116.4,20.5,-88.79,13.38,7.44,74.57,-15.6,-74.09,-47.91,17.13,15.26,-59.61,4.33,26.1,19.08,-58.37,29.35,31.25,149.13,-66.93,-60.98,-52.34,-64.94,28.83,-71.14,79.85,-13.67,12.57,-17.48,36.32,90.39,125.58,35.74,51.51,-4.48,-6.25,68.78,45.25,-171.22,-61.08,-13.24,25.91,-81.39,-58.16,-5.35,-90.55,105.84,-64.79,31.05,-82.39,24.94,159.91,73.06,106.83,-5.71,35.22,43.15,69.17,32.58,85.31,32.52,30.52,30.06,-76.8,168.05,-61.23,15.32,134.51,101.71,9.45,33.8,-77.05,-9.14,14.51,1.35,-0.1,15.61,13.24,28.29,6.12,-3.71,8.79,73.5,-86.27,50.58,120.97,32.57,27.49,-176.13,31.14,-99.14,27.55,45.33,7.42,-10.8,-56.17,43.23,37.62,58.54,15.05,36.82,-77.33,77.22,2.12,33.38,33.37,-15.98,166.44,-175.22,-51.73,-70.03,10.75,-75.71,-1.53,-170.72,158.16,-79.53,-149.56,-55.2,2.34,104.92,57.51,147.18,-57.82,-61.51,-72.34,2.63,14.43,-23.54,28.22,125.75,-78.5,-6.84,96.15,-21.92,24.13,171.06,46.77,-64.63,12.5,-61.39,-61.74,-2.11,-61.85,-2.55,55.46,-56.18,-85.13,-84.08,-66.13,12.43,-89.19,44.21,-70.64,-69.91,6.73,18.38,103.85,21.47,23.31,126.99,18.07,-65.26,145.7,121.45,24.74,69.3,44.79,-87.22,51.43,-63.05,89.7,19.82,139.77,-6.8,13.18,10.22,106.91,9.53,179.2,14.52,12.46,55.45,16.37,102.61,168.31,25.27,21.02,-77.02,174.78,-68.93,17.09,-5.28,11.52,166.91,44.52,15.97,-13.2,47.99],"y":[31.95,24.48,9.18,5.56,-25.05,9.03,13.47,36.77,-19.05,52.37,42.51,39.93,-18.89,-13.83,37.95,15.33,51.17,-25.3,37.98,-21.2,33.33,1.33,40.39,12.65,4.93,13.73,4.36,13.46,16,17.31,33.88,39.93,44.83,17.25,52.52,46.95,42.87,11.87,4.63,-15.78,48.16,-4.25,13.11,50.83,44.44,47.51,-34.61,-3.37,30.06,-35.31,10.54,14.03,4.92,18.35,47.03,21.46,6.93,9.55,55.68,14.72,33.5,23.7,-8.57,-6.17,25.3,54.15,53.33,38.57,-12.77,-9.38,14.6,8.49,-24.65,19.28,6.79,36.14,14.63,21.03,32.3,-17.82,23.13,60.17,-9.43,33.72,-6.18,-15.92,31.78,11.56,34.53,0.32,27.71,15.58,50.43,-1.94,17.99,-29.03,13.16,-4.31,7.35,3.16,0.39,-13.97,-12.07,38.72,46.06,6.17,51.52,78.21,-8.82,-15.42,49.62,40.42,3.74,4.17,12.15,26.21,14.62,-25.95,-29.31,-13.28,-26.32,19.43,53.91,2.05,43.74,6.31,-34.87,-11.74,55.75,23.61,12.11,-1.29,25.06,28.6,13.52,35.16,35.18,18.09,-22.27,-21.14,64.18,12.53,59.91,45.42,12.37,-14.24,6.92,8.97,-17.52,5.85,48.86,11.57,-20.17,-9.48,-51.7,10.66,18.54,6.48,50.08,14.93,-25.73,39.02,-0.19,34.02,16.79,64.14,56.97,7.12,24.65,18.43,41.89,15.3,12.06,49.19,17.11,49.47,-20.87,46.79,10.97,9.93,18.44,43.94,13.69,15.38,-33.46,18.48,0.37,43.85,1.3,42,42.69,37.56,59.33,-19.06,15.14,25.02,59.44,41.31,41.72,14.09,35.67,18.22,27.48,41.33,35.67,62.03,32.87,36.84,47.93,47.14,-8.52,35.91,41.9,-4.62,48.22,17.97,-17.74,54.7,52.26,38.91,-41.28,12.1,-22.56,6.82,3.87,-0.55,40.17,45.8,27.16,29.38],"hover_col_1":["'Amman ","Abu Dhabi ","Abuja ","Accra ","Adamstown ","Addis Abeba ","Agana ","Algiers ","Alofi ","Amsterdam ","Andorra la Vella ","Ankara ","Antananarivo ","Apia ","Asgabat ","Asmara ","Astana ","Asuncion ","Athens ","Avarua ","Baghdad ","Bairiki ","Baku ","Bamako ","Bandar Seri Begawan","Bangkok ","Bangui ","Banjul ","Basse-Terre ","Basseterre ","Bayrut ","Beijing ","Belgrade ","Belmopan ","Berlin ","Bern ","Biskek ","Bissau ","Bogota ","Brasilia ","Bratislava ","Brazzaville ","Bridgetown ","Brussels ","Bucharest ","Budapest ","Buenos Aires ","Bujumbura ","Cairo ","Canberra ","Caracas ","Castries ","Cayenne ","Charlotte Amalie ","Chisinau ","Cockburn Town ","Colombo ","Conakry ","Copenhagen ","Dakar ","Damascus ","Dhaka ","Dili ","Dodoma ","Doha ","Douglas ","Dublin ","Dushanbe ","Dzaoudzi ","Fakaofo ","Fort-de-France ","Freetown ","Gaborone ","George Town ","Georgetown ","Gibraltar ","Guatemala ","Ha Noi ","Hamilton ","Harare ","Havanna ","Helsinki ","Honiara ","Islamabad ","Jakarta ","Jamestown ","Jerusalem ","Jibuti ","Kabul ","Kampala ","Kathmandu ","Khartoum ","Kiev ","Kigali ","Kingston ","Kingston ","Kingstown ","Kinshasa ","Koror ","Kuala Lumpur ","Libreville ","Lilongwe ","Lima ","Lisbon ","Ljubljana ","Lome ","London ","Longyearbyen ","Luanda ","Lusaka ","Luxemburg ","Madrid ","Malabo ","Male ","Managua ","Manama ","Manila ","Maputo ","Maseru ","Mata'utu ","Mbabane ","Mexico City ","Minsk ","Mogadishu ","Monaco-Ville ","Monrovia ","Montevideo ","Moroni ","Moscow ","Muscat ","N'Djamena ","Nairobi ","Nassau ","Ni Dilli ","Niamey ","Nicosia ","Nicosia ","Nouakchott ","Noumea ","Nuku'alofa ","Nuuk ","Oranjestad ","Oslo ","Ottawa ","Ouagadougou ","Pago Pago ","Palikir ","Panama ","Papeete ","Paramaribo ","Paris ","Phnum Penh ","Port Louis ","Port Moresby ","Port Stanley ","Port of Spain ","Port-au-Prince ","Porto Novo ","Prague ","Praia ","Pretoria ","Pyongyang ","Quito ","Rabat ","Rangoon ","Reykjavik ","Riga ","Rita ","Riyadh ","Road Town ","Rome ","Roseau ","Saint George's ","Saint Helier ","Saint John's ","Saint Peter Port ","Saint-Denis ","Saint-Pierre ","San Jose ","San Jose ","San Juan ","San Marino ","San Salvador ","San'a ","Santiago ","Santo Domingo ","Sao Tome ","Sarajevo ","Singapore ","Skopje ","Sofia ","Soul ","Stockholm ","Sucre ","Susupe ","Taipei ","Tallinn ","Tashkent ","Tbilisi ","Tegucigalpa ","Tehran ","The Valley ","Thimphu ","Tirana ","Tokyo ","Torshavn ","Tripoli ","Tunis ","Ulaanbaatar ","Vaduz ","Vaiaku ","Valletta ","Vatican City ","Victoria ","Vienna ","Vientiane ","Vila ","Vilnius ","Warsaw ","Washington ","Wellington ","Willemstad ","Windhoek ","Yamoussoukro ","Yaounde ","Yaren ","Yerevan ","Zagreb ","al-'Ayun ","al-Kuwayt "],"hover_col_2":["Jordan ","United Arab Emirates ","Nigeria ","Ghana ","Pitcairn ","Ethiopia ","Guam ","Algeria ","Niue ","Netherlands ","Andorra ","Turkey ","Madagascar ","Samoa ","Turkmenistan ","Eritrea ","Kazakhstan ","Paraguay ","Greece ","Cook Islands ","Iraq ","Kiribati ","Azerbaijan ","Mali ","Brunei ","Thailand ","Central African Republic ","Gambia ","Guadeloupe ","Saint Kitts and Nevis ","Lebanon ","China ","Serbia and Montenegro ","Belize ","Germany ","Switzerland ","Kyrgyzstan ","Guinea-Bissau ","Colombia ","Brazil ","Slovakia ","Congo ","Barbados ","Belgium ","Romania ","Hungary ","Argentina ","Burundi ","Egypt ","Australia ","Venezuela ","Saint Lucia ","French Guiana ","US Virgin Islands ","Moldova ","Turks and Caicos ","Sri Lanka ","Guinea ","Denmark ","Senegal ","Syria ","Bangladesh ","East Timor ","Tanzania ","Qatar ","Isle of Man ","Ireland ","Tajikistan ","Mayotte ","Tokelau ","Martinique ","Sierra Leone ","Botswana ","Cayman Islands ","Guyana ","Gibraltar ","Guatemala ","Vietnam ","Bermuda ","Zimbabwe ","Cuba ","Finland ","Solomon Islands ","Pakistan ","Indonesia ","Saint Helena ","Israel ","Djibouti ","Afghanistan ","Uganda ","Nepal ","Sudan ","Ukraine ","Rwanda ","Jamaica ","Norfolk Island ","Saint Vincent and The Grenadines","Congo Democratic Republic ","Palau ","Malaysia ","Gabon ","Malawi ","Peru ","Portugal ","Slovenia ","Togo ","UK ","Svalbard and Jan Mayen ","Angola ","Zambia ","Luxembourg ","Spain ","Equatorial Guinea ","Maldives ","Nicaragua ","Bahrain ","Philippines ","Mozambique ","Lesotho ","Wallis and Futuna ","Swaziland ","Mexico ","Belarus ","Somalia ","Monaco ","Liberia ","Uruguay ","Comoros ","Russia ","Oman ","Chad ","Kenya ","Bahamas ","India ","Niger ","Cyprus ","Cyprus ","Mauritania ","New Caledonia ","Tonga ","Greenland ","Aruba ","Norway ","Canada ","Burkina Faso ","American Samoa ","Micronesia ","Panama ","French Polynesia ","Suriname ","France ","Cambodia ","Mauritius ","Papua New Guinea ","Falkland Islands ","Trinidad and Tobago ","Haiti ","Benin ","Czech Republic ","Cape Verde ","South Africa ","Korea North ","Ecuador ","Morocco ","Myanmar ","Iceland ","Latvia ","Marshall Islands ","Saudi Arabia ","British Virgin Islands ","Italy ","Dominica ","Grenada ","Jersey ","Antigua and Barbuda ","Guernsey and Alderney ","Reunion ","Saint Pierre and Miquelon ","Costa Rica ","Costa Rica ","Puerto Rico ","San Marino ","El Salvador ","Yemen ","Chile ","Dominican Republic ","Sao Tome and Principe ","Bosnia and Herzegovina ","Singapore ","Macedonia ","Bulgaria ","Korea South ","Sweden ","Bolivia ","Northern Mariana Islands ","Taiwan ","Estonia ","Uzbekistan ","Georgia ","Honduras ","Iran ","Anguilla ","Bhutan ","Albania ","Japan ","Faroe Islands ","Libya ","Tunisia ","Mongolia ","Liechtenstein ","Tuvalu ","Malta ","Vatican City ","Seychelles ","Austria ","Laos ","Vanuatu ","Lithuania ","Poland ","USA ","New Zealand ","Netherlands Antilles ","Namibia ","Ivory Coast ","Cameroon ","Nauru ","Armenia ","Croatia ","Western Sahara ","Kuwait "],"hover_col_3":["1,303,197 ","619,316 ","178,462 ","2,029,143 ","51 ","2,823,167 ","1,041 ","2,029,936 ","627 ","744,159 ","20,314 ","3,579,706 ","1,463,754 ","40,805 ","823,013 ","578,860 ","351,343 ","507,574 ","725,049 ","13,645 ","5,753,612 ","45,982 ","1,118,725 ","1,342,519 ","67,077 ","4,935,988 ","547,668 ","34,388 ","11,298 ","12,883 ","1,273,440 ","7,602,069 ","1,113,589 ","14,590 ","3,378,275 ","120,596 ","915,625 ","404,119 ","7,235,084 ","2,260,541 ","422,452 ","1,326,975 ","98,725 ","1,031,925 ","1,862,930 ","1,700,019 ","11,595,183","336,561 ","7,836,243 ","324,736 ","1,808,937 ","12,904 ","62,926 ","10,415 ","623,671 ","174 ","649,496 ","1,970,382 ","1,091,978 ","2,406,598 ","1,580,909 ","6,724,976 ","163,305 ","188,150 ","351,381 ","25,621 ","1,030,431 ","538,456 ","14,558 ","267 ","89,233 ","818,709 ","214,412 ","30,570 ","236,878 ","26,404 ","1,010,253 ","1,452,055 ","889 ","1,575,127 ","2,163,132 ","558,341 ","57,410 ","794,431 ","8,556,798 ","603 ","731,731 ","633,884 ","3,120,963 ","1,403,619 ","822,930 ","2,090,001 ","2,491,404 ","800,003 ","585,300 ","890 ","18,160 ","8,096,254 ","11,458 ","1,482,359 ","591,356 ","683,477 ","7,857,121 ","508,209 ","254,188 ","737,751 ","7,489,022 ","1,263 ","2,875,277 ","1,306,577 ","76,380 ","3,146,804 ","161,409 ","87,154 ","990,417 ","147,894 ","10,546,511","1,220,167 ","116,268 ","1,310 ","78,740 ","8,659,409 ","1,747,482 ","2,723,378 ","975 ","954,458 ","1,271,664 ","43,704 ","10,472,629","24,122 ","737,281 ","2,864,667 ","231,519 ","321,883 ","801,297 ","202,488 ","42,372 ","731,242 ","94,751 ","23,733 ","15,243 ","30,710 ","821,445 ","885,542 ","1,119,775 ","4,180 ","4,552 ","406,070 ","26,400 ","224,925 ","2,141,839 ","1,673,131 ","156,760 ","289,861 ","2,269 ","49,764 ","1,277,104 ","238,199 ","1,168,374 ","117,342 ","1,687,779 ","2,992,272 ","1,399,814 ","1,688,738 ","4,572,948 ","114,576 ","738,386 ","21,270 ","4,328,067 ","8,613 ","2,561,181 ","16,577 ","4,315 ","28,910 ","25,321 ","16,702 ","137,787 ","6,254 ","32,187 ","339,588 ","417,154 ","4,624 ","534,409 ","1,921,589 ","4,893,495 ","2,253,437 ","63,772 ","737,350 ","3,601,745 ","477,493 ","1,166,143 ","10,409,345","1,260,712 ","232,669 ","2,402 ","2,491,662 ","392,386 ","1,967,879 ","1,038,343 ","872,403 ","7,160,094 ","1,435 ","74,175 ","380,403 ","8,372,440 ","13,313 ","1,164,634 ","693,294 ","862,842 ","5,248 ","4,835 ","6,748 ","767 ","22,611 ","1,570,976 ","199,863 ","37,141 ","542,014 ","1,634,441 ","548,359 ","182,254 ","98,339 ","277,349 ","200,103 ","1,344,617 ","4,587 ","1,090,537 ","700,717 ","188,084 ","63,596 "]}}},{"type":"Circle","id":"47f536056493d90ce45056103c946e20","attributes":{"id":"47f536056493d90ce45056103c946e20","tags":[],"size":{"units":"screen","value":5},"line_color":{"units":"data","value":"#1F77B4"},"fill_color":{"units":"data","value":"#1F77B4"},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"Circle","id":"796f3d61319d5ded5d967e860fdee551","attributes":{"id":"796f3d61319d5ded5d967e860fdee551","tags":[],"size":{"units":"screen","value":5},"line_color":{"units":"data","value":"#e1e1e1"},"fill_color":{"units":"data","value":"#e1e1e1"},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":0.5},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"Circle","id":"95b75c38189e60b0919574d57854b25c","attributes":{"id":"95b75c38189e60b0919574d57854b25c","tags":[],"size":{"units":"screen","value":5},"line_color":{"units":"data","value":"#1F77B4"},"fill_color":{"units":"data","value":"#1F77B4"},"line_alpha":{"units":"data","value":1},"fill_alpha":{"units":"data","value":1},"x":{"units":"data","field":"x"},"y":{"units":"data","field":"y"}}},{"type":"GlyphRenderer","id":"d0d5df621e402ab18060410f73ca9d28","attributes":{"id":"d0d5df621e402ab18060410f73ca9d28","tags":[],"selection_glyph":null,"nonselection_glyph":{"type":"Circle","id":"796f3d61319d5ded5d967e860fdee551"},"hover_glyph":{"type":"Circle","id":"95b75c38189e60b0919574d57854b25c"},"name":null,"data_source":{"type":"ColumnDataSource","id":"2a1071dfa6c0806319defe035a7a31f9"},"glyph":{"type":"Circle","id":"47f536056493d90ce45056103c946e20"}}},{"type":"Range1d","id":"fd4371919ebaacd5d415338d5ce9c7fb","attributes":{"id":"fd4371919ebaacd5d415338d5ce9c7fb","tags":[],"start":-180,"end":190.270843505859}},{"type":"Range1d","id":"19e287b0f3e3f36a6926be14219eb799","attributes":{"id":"19e287b0f3e3f36a6926be14219eb799","tags":[],"start":-85.1921844482422,"end":83.599609375}},{"type":"LinearAxis","id":"eae859109634ea0b96d0a5268118226b","attributes":{"id":"eae859109634ea0b96d0a5268118226b","tags":[],"plot":{"type":"Plot","id":"f77af90710cde2b4e2ddd44c50c3e2ba","subtype":"Figure"},"axis_label":"longitude","formatter":{"type":"BasicTickFormatter","id":"1e681a950e10cd7a3101cb2aa32f89a3"},"ticker":{"type":"BasicTicker","id":"423759cdaeb2159d421519185f604176"},"visible":true,"axis_label_text_font_size":"12pt"}},{"type":"BasicTickFormatter","id":"1e681a950e10cd7a3101cb2aa32f89a3","attributes":{"id":"1e681a950e10cd7a3101cb2aa32f89a3","tags":[]}},{"type":"BasicTicker","id":"423759cdaeb2159d421519185f604176","attributes":{"id":"423759cdaeb2159d421519185f604176","tags":[],"num_minor_ticks":5}},{"type":"Grid","id":"810e26aab35b527dac544cc889fca40e","attributes":{"id":"810e26aab35b527dac544cc889fca40e","tags":[],"dimension":0,"plot":{"type":"Plot","id":"f77af90710cde2b4e2ddd44c50c3e2ba","subtype":"Figure"},"ticker":{"type":"BasicTicker","id":"423759cdaeb2159d421519185f604176"}}},{"type":"LinearAxis","id":"54bc5a7168a4b9e88594538b8e5c725b","attributes":{"id":"54bc5a7168a4b9e88594538b8e5c725b","tags":[],"plot":{"type":"Plot","id":"f77af90710cde2b4e2ddd44c50c3e2ba","subtype":"Figure"},"axis_label":"latitude","formatter":{"type":"BasicTickFormatter","id":"0074f4a18ed8b588843bea5729dd682f"},"ticker":{"type":"BasicTicker","id":"6ef52bd4050acbdaaffe8eb9574956ce"},"visible":true,"axis_label_text_font_size":"12pt"}},{"type":"BasicTickFormatter","id":"0074f4a18ed8b588843bea5729dd682f","attributes":{"id":"0074f4a18ed8b588843bea5729dd682f","tags":[]}},{"type":"BasicTicker","id":"6ef52bd4050acbdaaffe8eb9574956ce","attributes":{"id":"6ef52bd4050acbdaaffe8eb9574956ce","tags":[],"num_minor_ticks":5}},{"type":"Grid","id":"2c01b0f5c3bec758e9da037873c4f0b2","attributes":{"id":"2c01b0f5c3bec758e9da037873c4f0b2","tags":[],"dimension":1,"plot":{"type":"Plot","id":"f77af90710cde2b4e2ddd44c50c3e2ba","subtype":"Figure"},"ticker":{"type":"BasicTicker","id":"6ef52bd4050acbdaaffe8eb9574956ce"}}}]}}},"debug":false},"evals":[],"jsHooks":[]}</script> ] .pull-right[ **highcharter** <div id="htmlwidget-b3fa877a772977675b74" style="width:400px;height:250px;" class="highchart html-widget"></div> <script type="application/json" data-for="htmlwidget-b3fa877a772977675b74">{"x":{"hc_opts":{"chart":{"reflow":true,"width":400,"height":250},"title":{"text":"Europe Time Zones"},"yAxis":{"title":{"text":null}},"credits":{"enabled":true},"exporting":{"enabled":false},"boost":{"enabled":false},"plotOptions":{"series":{"label":{"enabled":false},"turboThreshold":0},"treemap":{"layoutAlgorithm":"squarified"}},"series":[{"mapData":{"title":"Europe","version":"1.1.3","type":"FeatureCollection","copyright":"Copyright (c) 2020 Highsoft AS, Based on data from Natural Earth","copyrightShort":"Natural Earth","copyrightUrl":"http://www.naturalearthdata.com","crs":{"type":"name","properties":{"name":"urn:ogc:def:crs:EPSG:102014"}},"hc-transform":{"default":{"crs":"+proj=lcc +lat_1=43 +lat_2=62 +lat_0=30 +lon_0=10 +x_0=0 +y_0=0 +ellps=intl +units=m +no_defs","scale":0.000169255094964,"jsonres":15.5,"jsonmarginX":-999,"jsonmarginY":9851,"xoffset":-1706316.2597,"yoffset":4824494.18154}},"features":[{"type":"Feature","id":"DK","properties":{"hc-group":"admin0","hc-middle-x":0.19,"hc-middle-y":0.44,"hc-key":"dk","hc-a2":"DK","name":"Denmark","labelrank":"4","country-abbrev":"Den.","subregion":"Northern Europe","region-wb":"Europe & Central Asia","iso-a3":"DNK","iso-a2":"DK","woe-id":"23424796","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[3686,4530],[3743,4490],[3764,4508],[3787,4471],[3773,4443],[3724,4436],[3644,4481],[3658,4526],[3686,4530]]],[[[3536,4512],[3564,4499],[3541,4493],[3513,4524],[3536,4512]]],[[[3898,4549],[3847,4516],[3831,4533],[3864,4557],[3898,4549]]],[[[3443,4565],[3482,4530],[3472,4501],[3447,4513],[3443,4565]]],[[[3616,4524],[3599,4467],[3584,4496],[3635,4587],[3616,4524]]],[[[4282,4636],[4328,4610],[4317,4568],[4251,4596],[4260,4656],[4282,4636]]],[[[3217,4671],[3225,4667],[3223,4642],[3210,4667],[3217,4671]]],[[[3533,4717],[3608,4645],[3599,4563],[3567,4550],[3511,4568],[3458,4614],[3459,4642],[3424,4684],[3533,4717]]],[[[3899,4708],[3889,4727],[3902,4751],[3914,4724],[3899,4708]]],[[[3578,4795],[3579,4771],[3562,4769],[3572,4800],[3578,4795]]],[[[3306,5107],[3300,5063],[3263,5028],[3247,5060],[3264,5089],[3306,5107]]],[[[3664,5214],[3644,5204],[3639,5178],[3611,5196],[3664,5214]]],[[[3282,4516],[3255,4513],[3260,4582],[3227,4563],[3225,4595],[3261,4584],[3251,4670],[3201,4707],[3166,4708],[3184,4779],[3216,4802],[3200,4855],[3172,4868],[3178,4993],[3188,5027],[3208,5000],[3249,4999],[3274,4971],[3277,5015],[3312,5063],[3347,5035],[3326,5010],[3362,5013],[3345,5088],[3389,5126],[3417,5111],[3444,5148],[3362,5120],[3338,5136],[3269,5107],[3237,5058],[3245,5003],[3199,5057],[3260,5156],[3385,5166],[3411,5189],[3472,5289],[3508,5292],[3570,5336],[3544,5283],[3563,5244],[3562,5187],[3548,5172],[3519,5092],[3535,4992],[3603,4985],[3632,4960],[3598,4893],[3578,4899],[3563,4859],[3558,4910],[3542,4916],[3513,4872],[3517,4805],[3460,4782],[3487,4757],[3444,4736],[3454,4722],[3395,4683],[3430,4613],[3394,4588],[3434,4536],[3435,4494],[3409,4521],[3384,4487],[3384,4487],[3282,4516]]],[[[3717,4815],[3747,4806],[3772,4741],[3805,4817],[3779,4826],[3839,4872],[3901,4850],[3884,4814],[3901,4753],[3845,4704],[3842,4672],[3879,4649],[3871,4621],[3837,4612],[3834,4545],[3818,4538],[3838,4500],[3805,4457],[3768,4508],[3790,4538],[3750,4596],[3685,4601],[3656,4648],[3675,4656],[3665,4703],[3634,4751],[3681,4753],[3717,4786],[3717,4815]]]]}},{"type":"Feature","id":"FO","properties":{"hc-group":"admin0","hc-middle-x":0.56,"hc-middle-y":0.16,"hc-key":"fo","hc-a2":"FO","name":"Faroe Islands","labelrank":"6","country-abbrev":"Faeroe Is.","subregion":"Northern Europe","region-wb":"Europe & Central Asia","iso-a3":"FRO","iso-a2":"FO","woe-id":"23424816","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[1151,6735],[1169,6717],[1165,6663],[1146,6700],[1151,6735]]],[[[1176,6812],[1202,6782],[1190,6763],[1169,6796],[1176,6812]]],[[[1150,6888],[1163,6872],[1139,6859],[1115,6902],[1150,6888]]],[[[1158,6940],[1202,6846],[1193,6819],[1166,6880],[1143,6903],[1158,6940]]],[[[1199,6933],[1230,6870],[1208,6857],[1181,6898],[1172,6936],[1199,6933]]],[[[1236,6908],[1230,6914],[1227,6927],[1232,6941],[1236,6908]]],[[[1226,6902],[1222,6904],[1215,6942],[1217,6946],[1226,6902]]],[[[1254,6915],[1248,6943],[1254,6940],[1257,6921],[1255,6914],[1263,6903],[1246,6901],[1239,6895],[1243,6931],[1254,6915]]]]}},{"type":"Feature","id":"HR","properties":{"hc-group":"admin0","hc-middle-x":0.09,"hc-middle-y":0.39,"hc-key":"hr","hc-a2":"HR","name":"Croatia","labelrank":"6","country-abbrev":"Cro.","subregion":"Southern Europe","region-wb":"Europe & Central Asia","iso-a3":"HRV","iso-a2":"HR","woe-id":"23424843","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[4965,1080],[4964,1071],[4945,1066],[4938,1078],[4965,1080]]],[[[5060,1097],[5137,1081],[5142,1075],[5062,1089],[5060,1097]]],[[[4924,1135],[4980,1143],[5017,1129],[4944,1116],[4924,1135]]],[[[4799,1153],[4804,1142],[4785,1134],[4768,1148],[4799,1153]]],[[[4873,1205],[4915,1189],[5010,1189],[4904,1178],[4834,1192],[4873,1205]]],[[[4922,1243],[4925,1221],[4841,1228],[4840,1250],[4922,1243]]],[[[4813,1251],[4834,1236],[4825,1233],[4794,1244],[4813,1251]]],[[[4621,1380],[4603,1384],[4580,1409],[4607,1399],[4621,1380]]],[[[4572,1425],[4565,1419],[4550,1431],[4536,1450],[4572,1425]]],[[[4496,1441],[4573,1380],[4557,1376],[4526,1418],[4496,1441]]],[[[4482,1574],[4494,1578],[4572,1505],[4495,1543],[4482,1574]]],[[[4392,1569],[4391,1562],[4374,1601],[4388,1593],[4392,1569]]],[[[4458,1644],[4474,1623],[4481,1604],[4444,1643],[4458,1644]]],[[[4398,1672],[4394,1585],[4363,1666],[4381,1672],[4355,1720],[4372,1729],[4398,1672]]],[[[4412,1758],[4435,1713],[4468,1683],[4386,1710],[4412,1758]]],[[[5311,1013],[5150,1111],[5070,1120],[5023,1159],[5115,1130],[5161,1114],[5293,1053],[5298,1030],[5311,1013]]],[[[5272,2037],[5265,2001],[5308,1936],[5313,1878],[5390,1853],[5399,1831],[5332,1816],[5333,1734],[5308,1729],[5259,1787],[5229,1777],[5164,1799],[5120,1791],[5085,1763],[5051,1788],[5030,1773],[4966,1787],[4950,1773],[4893,1807],[4867,1774],[4805,1774],[4781,1716],[4754,1721],[4705,1771],[4662,1750],[4668,1659],[4747,1571],[4775,1482],[4881,1385],[4890,1365],[4976,1298],[5010,1291],[5019,1256],[5062,1201],[5095,1184],[5098,1143],[4981,1218],[4936,1263],[4860,1282],[4742,1274],[4729,1323],[4625,1387],[4544,1481],[4583,1510],[4492,1601],[4491,1679],[4468,1723],[4415,1769],[4362,1786],[4296,1624],[4219,1709],[4192,1822],[4209,1816],[4271,1801],[4286,1828],[4369,1822],[4403,1876],[4428,1842],[4525,1818],[4570,1837],[4542,1900],[4581,1934],[4624,1943],[4625,2002],[4602,2023],[4614,2053],[4734,2110],[4722,2143],[4777,2150],[4851,2116],[4877,2075],[4924,2054],[4959,2009],[5008,2009],[5070,1968],[5138,1979],[5180,1971],[5227,2026],[5272,2037]]]]}},{"type":"Feature","id":"NL","properties":{"hc-group":"admin0","hc-middle-x":0.34,"hc-middle-y":0.59,"hc-key":"nl","hc-a2":"NL","name":"Netherlands","labelrank":"5","country-abbrev":"Neth.","subregion":"Western Europe","region-wb":"Europe & Central Asia","iso-a3":"NLD","iso-a2":"NL","woe-id":"-90","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[2408,3636],[2399,3614],[2350,3647],[2395,3648],[2408,3636]]],[[[2593,4028],[2574,3998],[2561,4015],[2588,4051],[2593,4028]]],[[[2603,4061],[2593,4060],[2613,4079],[2625,4085],[2603,4061]]],[[[2691,4107],[2652,4094],[2648,4104],[2719,4119],[2691,4107]]],[[[2776,4121],[2783,4117],[2735,4111],[2734,4122],[2776,4121]]],[[[2817,4118],[2812,4122],[2827,4129],[2847,4126],[2817,4118]]],[[[2280,3553],[2316,3561],[2371,3536],[2395,3553],[2437,3537],[2380,3495],[2330,3523],[2285,3515],[2280,3553]]],[[[2444,3537],[2389,3565],[2367,3550],[2316,3574],[2305,3602],[2372,3613],[2422,3559],[2444,3580],[2400,3604],[2436,3629],[2402,3678],[2414,3716],[2438,3721],[2510,3811],[2565,3989],[2593,3969],[2677,4019],[2717,4076],[2786,4101],[2916,4110],[2993,4045],[2988,3965],[2969,3933],[2959,3874],[2912,3874],[2897,3827],[2945,3818],[2952,3754],[2887,3701],[2910,3678],[2832,3644],[2788,3666],[2751,3642],[2779,3594],[2794,3519],[2766,3475],[2780,3452],[2737,3419],[2761,3382],[2746,3338],[2690,3344],[2723,3456],[2660,3498],[2614,3492],[2581,3557],[2561,3534],[2537,3566],[2511,3546],[2486,3564],[2467,3530],[2444,3537]],[[2720,3812],[2735,3822],[2736,3825],[2736,3825],[2736,3825],[2750,3836],[2750,3869],[2717,3876],[2748,3862],[2736,3825],[2736,3825],[2736,3825],[2720,3812],[2720,3812],[2720,3812],[2720,3812],[2670,3782],[2621,3817],[2717,3877],[2712,3924],[2732,3940],[2678,3949],[2676,4013],[2616,3976],[2626,3929],[2657,3907],[2608,3887],[2614,3833],[2591,3815],[2674,3776],[2720,3812],[2720,3812],[2720,3812],[2720,3812]]]]}},{"type":"Feature","id":"EE","properties":{"hc-group":"admin0","hc-middle-x":0.6,"hc-middle-y":0.53,"hc-key":"ee","hc-a2":"EE","name":"Estonia","labelrank":"6","country-abbrev":"Est.","subregion":"Northern Europe","region-wb":"Europe & Central Asia","iso-a3":"EST","iso-a2":"EE","woe-id":"23424805","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[5483,5775],[5469,5746],[5441,5761],[5452,5784],[5483,5775]]],[[[5440,5891],[5460,5893],[5468,5876],[5435,5869],[5440,5891]]],[[[5370,5868],[5403,5866],[5426,5828],[5383,5810],[5362,5771],[5337,5813],[5275,5835],[5334,5850],[5348,5887],[5370,5868]]],[[[6143,6139],[6134,6134],[6119,6126],[6130,6125],[6130,6125],[6131,6125],[6133,6126],[6124,6119],[6106,6106],[6108,6072],[6106,6046],[6102,6021],[6074,6020],[6012,5988],[6005,5937],[6044,5907],[6076,5839],[6106,5812],[6161,5739],[6194,5705],[6159,5682],[6151,5595],[6071,5605],[6022,5561],[5934,5617],[5926,5637],[5861,5642],[5801,5677],[5699,5617],[5669,5588],[5681,5651],[5674,5732],[5642,5736],[5618,5688],[5557,5706],[5535,5759],[5512,5760],[5504,5815],[5545,5818],[5543,5838],[5488,5817],[5479,5858],[5495,5879],[5471,5891],[5470,5946],[5555,5992],[5541,6009],[5694,6088],[5735,6082],[5770,6142],[5882,6132],[5912,6141],[5940,6125],[6095,6143],[6111,6169],[6126,6160],[6143,6139]]],[[[5280,5633],[5297,5662],[5272,5700],[5306,5706],[5369,5757],[5380,5746],[5417,5762],[5487,5731],[5463,5728],[5448,5692],[5403,5642],[5351,5628],[5342,5561],[5313,5556],[5336,5608],[5280,5633]]]]}},{"type":"Feature","id":"BG","properties":{"hc-group":"admin0","hc-middle-x":0.54,"hc-middle-y":0.49,"hc-key":"bg","hc-a2":"BG","name":"Bulgaria","labelrank":"4","country-abbrev":"Bulg.","subregion":"Eastern Europe","region-wb":"Europe & Central Asia","iso-a3":"BGR","iso-a2":"BG","woe-id":"23424771","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[7353,1794],[7373,1738],[7359,1683],[7291,1678],[7265,1596],[7264,1551],[7287,1467],[7255,1461],[7212,1374],[7250,1382],[7288,1337],[7352,1291],[7366,1265],[7320,1259],[7271,1219],[7200,1259],[7149,1241],[7136,1215],[7058,1175],[7060,1145],[7026,1104],[6967,1083],[7003,1040],[7004,992],[6956,965],[6884,953],[6831,918],[6735,946],[6714,926],[6671,941],[6645,975],[6547,945],[6462,882],[6390,877],[6357,845],[6313,844],[6303,916],[6313,957],[6262,1037],[6189,1058],[6140,1104],[6170,1136],[6144,1181],[6146,1270],[6192,1283],[6229,1382],[6118,1443],[6066,1533],[6067,1588],[6105,1620],[6108,1660],[6115,1666],[6116,1666],[6193,1640],[6168,1606],[6183,1561],[6264,1592],[6444,1587],[6512,1615],[6610,1620],[6698,1611],[6780,1652],[6838,1749],[7006,1837],[7055,1833],[7089,1809],[7144,1825],[7163,1808],[7199,1830],[7223,1793],[7278,1781],[7353,1794]]]}},{"type":"Feature","id":"ES","properties":{"hc-group":"admin0","hc-middle-x":0.38,"hc-middle-y":0.53,"hc-key":"es","hc-a2":"ES","name":"Spain","labelrank":"2","country-abbrev":"Sp.","subregion":"Southern Europe","region-wb":"Europe & Central Asia","iso-a3":"ESP","iso-a2":"ES","woe-id":"23424950","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[1502,-54],[1511,-71],[1486,-84],[1488,-64],[1502,-54]]],[[[1546,46],[1554,23],[1497,-29],[1456,-3],[1495,44],[1546,46]]],[[[1936,261],[1927,203],[1997,190],[1929,81],[1890,58],[1826,103],[1806,154],[1767,130],[1740,175],[1879,250],[1936,261]]],[[[2150,269],[2183,239],[2186,195],[2124,231],[2082,238],[2083,271],[2150,269]]],[[[-240,-615],[-227,-613],[-212,-627],[-226,-637],[-240,-615]]],[[[1743,1010],[1741,1016],[1746,1022],[1751,1009],[1743,1010]]],[[[-212,-550],[-280,-577],[-390,-461],[-390,-378],[-423,-351],[-410,-302],[-426,-252],[-503,-171],[-621,-134],[-621,-20],[-559,40],[-533,84],[-473,83],[-444,129],[-481,131],[-520,224],[-479,302],[-423,324],[-393,365],[-430,404],[-449,478],[-438,511],[-476,588],[-361,562],[-304,645],[-328,703],[-262,740],[-270,768],[-238,881],[-249,936],[-220,941],[-168,993],[-134,993],[-54,1057],[-122,1124],[-107,1176],[-226,1223],[-257,1193],[-338,1204],[-354,1228],[-457,1228],[-463,1252],[-423,1288],[-441,1327],[-534,1320],[-566,1304],[-596,1288],[-587,1350],[-515,1391],[-569,1392],[-513,1429],[-557,1437],[-523,1496],[-578,1477],[-586,1499],[-540,1545],[-582,1564],[-574,1593],[-606,1604],[-593,1640],[-553,1673],[-514,1671],[-482,1695],[-455,1674],[-372,1686],[-356,1731],[-254,1763],[-154,1714],[-134,1675],[75,1634],[238,1587],[284,1556],[443,1491],[623,1495],[629,1468],[712,1434],[786,1445],[860,1392],[909,1376],[984,1392],[989,1363],[1059,1338],[1034,1294],[1135,1241],[1182,1231],[1213,1182],[1270,1188],[1324,1134],[1370,1139],[1390,1120],[1470,1114],[1478,1158],[1615,1104],[1634,1067],[1645,1016],[1691,1033],[1760,981],[1811,998],[1848,968],[1953,995],[2005,979],[2032,944],[1994,929],[1984,895],[2004,842],[1931,775],[1785,717],[1727,667],[1609,655],[1483,627],[1414,570],[1439,525],[1381,514],[1313,426],[1239,361],[1131,223],[1138,92],[1175,24],[1224,-23],[1143,-71],[1081,-84],[1038,-115],[959,-245],[961,-319],[864,-330],[834,-311],[724,-358],[683,-399],[650,-471],[586,-524],[536,-485],[484,-484],[458,-514],[327,-472],[280,-482],[239,-459],[132,-437],[41,-430],[12,-464],[-40,-480],[-81,-467],[-154,-480],[-207,-551],[-212,-550]]]]}},{"type":"Feature","id":"IT","properties":{"hc-group":"admin0","hc-middle-x":0.44,"hc-middle-y":0.38,"hc-key":"it","hc-a2":"IT","name":"Italy","labelrank":"2","country-abbrev":"Italy","subregion":"Southern Europe","region-wb":"Europe & Central Asia","iso-a3":"ITA","iso-a2":"IT","woe-id":"23424853","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[3951,-735],[3968,-746],[3964,-760],[3941,-747],[3951,-735]]],[[[4632,-225],[4627,-220],[4615,-212],[4628,-199],[4632,-225]]],[[[3101,-59],[3124,-77],[3111,-105],[3098,-75],[3101,-59]]],[[[3088,-62],[3076,-61],[3070,-47],[3089,-36],[3088,-62]]],[[[4361,430],[4336,430],[4339,444],[4362,437],[4361,430]]],[[[3108,525],[3108,513],[3089,508],[3096,526],[3108,525]]],[[[3570,1021],[3559,994],[3500,1004],[3511,1022],[3570,1021]]],[[[4232,1848],[4234,1880],[4197,1904],[4172,1883],[4130,1896],[4104,1856],[3969,1791],[3987,1831],[3941,1798],[3929,1753],[3956,1693],[3999,1657],[3982,1613],[3951,1618],[3952,1515],[3972,1447],[4046,1372],[4097,1349],[4164,1295],[4243,1249],[4295,1134],[4324,1039],[4355,983],[4508,842],[4595,804],[4666,801],[4791,822],[4831,780],[4776,731],[4777,691],[4803,671],[4934,618],[5040,591],[5140,527],[5263,487],[5321,437],[5395,344],[5373,247],[5302,273],[5278,332],[5239,378],[5164,374],[5094,397],[5054,427],[5026,402],[4967,290],[4976,256],[4949,206],[4957,172],[5012,157],[5111,103],[5111,19],[5133,-6],[5113,-47],[5052,-47],[4988,-111],[5003,-197],[4918,-289],[4904,-348],[4805,-351],[4795,-270],[4825,-257],[4848,-197],[4830,-143],[4905,-111],[4910,-66],[4874,-17],[4845,89],[4813,123],[4787,211],[4743,268],[4685,243],[4632,288],[4582,305],[4599,352],[4538,431],[4446,398],[4475,442],[4433,472],[4389,467],[4300,585],[4268,571],[4211,594],[4151,571],[4115,620],[4058,632],[4015,684],[3968,713],[3946,762],[3876,798],[3837,869],[3780,903],[3714,902],[3734,937],[3690,992],[3634,1019],[3644,1045],[3589,1053],[3594,1118],[3570,1188],[3547,1214],[3533,1311],[3500,1371],[3440,1384],[3317,1469],[3212,1491],[3157,1458],[3107,1400],[3093,1358],[3048,1329],[2951,1315],[2947,1336],[2994,1398],[2987,1425],[2929,1412],[2858,1449],[2825,1488],[2823,1539],[2868,1577],[2845,1630],[2803,1651],[2780,1703],[2830,1708],[2885,1743],[2898,1787],[2863,1817],[2860,1855],[2827,1879],[2830,1910],[2876,1937],[2908,1923],[2981,1950],[3039,1928],[3097,1988],[3091,2025],[3166,2075],[3162,2022],[3196,1984],[3243,1967],[3255,1898],[3287,1913],[3272,1939],[3331,2036],[3330,2087],[3365,2089],[3389,2031],[3470,2049],[3486,2010],[3509,2022],[3486,2071],[3495,2121],[3562,2101],[3551,2141],[3567,2197],[3676,2170],[3694,2212],[3737,2234],[3823,2232],[3913,2265],[3895,2240],[3956,2154],[4122,2120],[4216,2116],[4152,2046],[4209,2017],[4176,1965],[4209,1955],[4203,1909],[4269,1861],[4232,1848]],[[3995,1344],[3998,1370],[3986,1365],[3979,1352],[3995,1344]],[[4013,764],[4013,764],[4013,764],[4013,764],[4013,764]]],[[[4478,-359],[4561,-334],[4623,-299],[4666,-313],[4761,-256],[4763,-319],[4702,-424],[4681,-501],[4688,-552],[4748,-640],[4712,-662],[4694,-749],[4551,-718],[4496,-647],[4450,-628],[4404,-635],[4296,-582],[4228,-529],[4153,-506],[4105,-511],[4048,-440],[4064,-380],[4122,-328],[4157,-371],[4202,-322],[4252,-310],[4270,-344],[4366,-378],[4418,-353],[4478,-359]]],[[[3266,-34],[3251,-99],[3213,-132],[3155,-125],[3146,-78],[3104,-26],[3106,18],[3148,164],[3115,173],[3137,231],[3132,293],[3100,371],[3066,368],[3061,416],[3084,469],[3148,443],[3203,467],[3307,568],[3353,545],[3381,525],[3369,495],[3439,349],[3393,274],[3417,221],[3396,45],[3376,-56],[3315,-32],[3266,-34]]]]}},{"type":"Feature","id":"SM","properties":{"hc-group":"admin0","hc-middle-x":0.68,"hc-middle-y":0.41,"hc-key":"sm","hc-a2":"SM","name":"San Marino","labelrank":"6","country-abbrev":"S.M.","subregion":"Southern Europe","region-wb":"Europe & Central Asia","iso-a3":"SMR","iso-a2":"SM","woe-id":"23424947","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[3995,1344],[3979,1352],[3986,1365],[3998,1370],[3995,1344]]]}},{"type":"Feature","id":"VA","properties":{"hc-group":"admin0","hc-middle-x":0.62,"hc-middle-y":0.44,"hc-key":"va","hc-a2":"VA","name":"Vatican","labelrank":"6","country-abbrev":"Vat.","subregion":"Southern Europe","region-wb":"Europe & Central Asia","iso-a3":"VAT","iso-a2":"VA","woe-id":"23424986","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[4013,764],[4013,764],[4013,764],[4013,764],[4013,764]]]}},{"type":"Feature","id":"TR","properties":{"hc-group":"admin0","hc-middle-x":0.62,"hc-middle-y":0.51,"hc-key":"tr","hc-a2":"TR","name":"Turkey","labelrank":"2","country-abbrev":"Tur.","subregion":"Western Asia","region-wb":"Europe & Central Asia","iso-a3":"TUR","iso-a2":"TR","woe-id":"23424969","continent":"Asia"},"geometry":{"type":"MultiPolygon","coordinates":[[[[7048,641],[7008,614],[6984,617],[7031,663],[7048,641]]],[[[7377,872],[7400,872],[7378,846],[7356,860],[7377,872]]],[[[7026,1104],[7060,1145],[7058,1175],[7136,1215],[7149,1241],[7200,1259],[7271,1219],[7320,1259],[7366,1265],[7367,1220],[7404,1176],[7482,1136],[7608,1115],[7653,1117],[7644,1044],[7562,1015],[7478,1025],[7426,979],[7331,959],[7321,909],[7281,839],[7186,763],[7178,732],[7097,621],[7110,666],[7091,700],[7208,812],[7195,827],[7137,795],[7043,777],[7028,811],[7081,892],[7056,973],[7113,1024],[7089,1088],[7026,1104]]],[[[9732,429],[9709,427],[9717,324],[9732,308],[9732,245],[9709,236],[9731,170],[9699,144],[9700,102],[9638,110],[9642,143],[9568,212],[9628,325],[9607,380],[9556,400],[9498,327],[9502,279],[9445,237],[9287,266],[9243,235],[9170,98],[9163,32],[9133,54],[9107,-9],[8982,-55],[8915,-110],[8801,-100],[8689,-21],[8509,5],[8352,-16],[8335,-55],[8356,-128],[8340,-219],[8301,-212],[8181,-304],[8158,-284],[8099,-292],[8040,-273],[8000,-223],[8004,-185],[7961,-170],[7948,-209],[7892,-205],[7879,-172],[7800,-182],[7811,-212],[7781,-263],[7741,-246],[7754,-221],[7682,-240],[7674,-274],[7600,-284],[7653,-235],[7752,-207],[7737,-175],[7792,-123],[7626,-189],[7594,-180],[7558,-209],[7560,-159],[7597,-159],[7617,-99],[7583,-114],[7568,-76],[7518,-98],[7500,-29],[7450,-20],[7499,16],[7480,85],[7391,80],[7355,125],[7323,84],[7232,113],[7280,153],[7234,219],[7268,237],[7305,205],[7329,146],[7417,196],[7380,197],[7311,268],[7348,280],[7369,348],[7329,331],[7324,377],[7249,421],[7305,498],[7291,525],[7117,451],[7123,498],[7103,589],[7133,619],[7143,674],[7201,751],[7262,762],[7321,796],[7363,765],[7436,783],[7400,829],[7479,841],[7456,812],[7644,862],[7687,862],[7724,893],[7635,896],[7672,943],[7792,990],[7866,1032],[7743,998],[7655,1041],[7651,1092],[7679,1126],[7825,1139],[7879,1153],[7902,1180],[8032,1178],[8116,1207],[8142,1249],[8133,1281],[8262,1408],[8278,1451],[8340,1507],[8402,1540],[8471,1606],[8686,1658],[8761,1684],[8797,1744],[8844,1737],[8842,1713],[8888,1673],[8964,1661],[9041,1721],[9086,1704],[9106,1660],[9194,1625],[9215,1675],[9298,1682],[9322,1656],[9449,1661],[9460,1698],[9520,1672],[9616,1692],[9691,1735],[9724,1777],[9732,1781],[9732,429]]]]}},{"type":"Feature","id":"MT","properties":{"hc-group":"admin0","hc-middle-x":0.67,"hc-middle-y":0.77,"hc-key":"mt","hc-a2":"MT","name":"Malta","labelrank":"5","country-abbrev":"Malta","subregion":"Southern Europe","region-wb":"Middle East & North Africa","iso-a3":"MLT","iso-a2":"MT","woe-id":"23424897","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[4588,-999],[4533,-992],[4526,-965],[4572,-975],[4588,-999]]],[[[4514,-952],[4492,-948],[4508,-935],[4528,-946],[4514,-952]]]]}},{"type":"Feature","id":"FR","properties":{"hc-group":"admin0","hc-middle-x":0.47,"hc-middle-y":0.47,"hc-key":"fr","hc-a2":"FR","name":"France","labelrank":"2","country-abbrev":"Fr.","subregion":"Western Europe","region-wb":"Europe & Central Asia","iso-a3":"FRA","iso-a2":"FR","woe-id":"-90","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[1209,2125],[1227,2092],[1216,2069],[1190,2121],[1209,2125]]],[[[1184,2202],[1214,2184],[1223,2170],[1167,2205],[1184,2202]]],[[[1077,2433],[1065,2431],[1059,2452],[1077,2447],[1077,2433]]],[[[915,2566],[923,2556],[890,2567],[893,2589],[915,2566]]],[[[2720,2051],[2736,2073],[2835,2080],[2844,1995],[2876,1937],[2830,1910],[2827,1879],[2860,1855],[2863,1817],[2898,1787],[2885,1743],[2830,1708],[2780,1703],[2803,1651],[2845,1630],[2868,1577],[2823,1539],[2825,1488],[2858,1449],[2929,1412],[2987,1425],[2994,1398],[2947,1336],[2951,1315],[2945,1307],[2937,1301],[2931,1307],[2922,1295],[2876,1277],[2872,1248],[2784,1209],[2754,1175],[2770,1150],[2592,1113],[2547,1158],[2487,1170],[2478,1211],[2429,1206],[2454,1234],[2402,1241],[2378,1215],[2324,1226],[2315,1252],[2250,1258],[2231,1287],[2195,1284],[2072,1222],[2010,1176],[1989,1130],[1981,1017],[2005,979],[1953,995],[1848,968],[1811,998],[1760,981],[1691,1033],[1697,1064],[1634,1067],[1615,1104],[1478,1158],[1470,1114],[1390,1120],[1370,1139],[1324,1134],[1270,1188],[1213,1182],[1182,1231],[1135,1241],[1034,1294],[1059,1338],[989,1363],[984,1392],[1013,1392],[1059,1437],[1169,1739],[1152,1731],[1216,1974],[1230,1995],[1284,1915],[1291,1943],[1214,2039],[1247,2091],[1256,2193],[1130,2285],[1126,2324],[1083,2386],[1119,2448],[1091,2486],[1101,2527],[1073,2513],[1026,2561],[1060,2567],[1044,2600],[981,2603],[996,2638],[936,2634],[860,2708],[799,2726],[788,2753],[700,2747],[698,2797],[656,2833],[733,2829],[736,2855],[696,2895],[737,2918],[659,2913],[664,2956],[711,2987],[760,2981],[828,2994],[845,2960],[858,2988],[901,2967],[920,3008],[1000,3002],[1059,2885],[1137,2923],[1151,2890],[1227,2916],[1224,2890],[1313,2887],[1283,2917],[1297,2997],[1296,3054],[1264,3110],[1258,3206],[1317,3176],[1357,3187],[1384,3157],[1366,3140],[1383,3100],[1447,3071],[1523,3053],[1556,3030],[1633,3059],[1681,3063],[1625,3083],[1659,3147],[1746,3178],[1841,3190],[1906,3248],[1919,3273],[1946,3441],[2022,3473],[2124,3485],[2123,3485],[2129,3413],[2161,3376],[2227,3382],[2244,3311],[2310,3281],[2310,3245],[2377,3244],[2408,3217],[2390,3138],[2443,3122],[2486,3135],[2492,3162],[2529,3175],[2510,3129],[2519,3074],[2536,3076],[2629,2982],[2688,2991],[2719,2961],[2752,2975],[2791,2962],[2819,2954],[2854,2877],[2879,2886],[2960,2853],[2984,2871],[3032,2830],[3085,2829],[3136,2805],[3116,2766],[3059,2708],[3045,2634],[3008,2560],[3016,2533],[2996,2470],[3006,2412],[2964,2369],[2892,2392],[2862,2351],[2897,2346],[2805,2247],[2773,2237],[2768,2184],[2706,2139],[2688,2089],[2704,2071],[2664,2007],[2731,2035],[2720,2051]],[[1743,1010],[1751,1009],[1746,1022],[1741,1016],[1743,1010]]],[[[3204,798],[3163,857],[3201,949],[3285,996],[3325,981],[3337,1073],[3354,1080],[3367,1026],[3360,972],[3377,942],[3380,816],[3347,770],[3346,698],[3306,600],[3212,657],[3241,695],[3192,706],[3203,766],[3169,777],[3204,798]]]]}},{"type":"Feature","id":"NO","properties":{"hc-group":"admin0","hc-middle-x":0.43,"hc-middle-y":0.8,"hc-key":"no","hc-a2":"NO","name":"Norway","labelrank":"3","country-abbrev":"Nor.","subregion":"Northern Europe","region-wb":"Europe & Central Asia","iso-a3":"NOR","iso-a2":"NO","woe-id":"-90","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[2823,5756],[2848,5748],[2843,5737],[2818,5753],[2823,5756]]],[[[2800,5766],[2790,5779],[2796,5789],[2803,5778],[2800,5766]]],[[[2884,5781],[2873,5785],[2877,5796],[2899,5793],[2884,5781]]],[[[2777,5805],[2777,5769],[2759,5778],[2769,5841],[2777,5805]]],[[[2778,5961],[2792,5938],[2757,5944],[2763,5974],[2778,5961]]],[[[2819,5951],[2791,5950],[2785,5984],[2810,5991],[2819,5951]]],[[[2847,6026],[2842,5982],[2822,5979],[2818,6024],[2847,6026]]],[[[2789,6032],[2789,6009],[2776,6024],[2782,6042],[2789,6032]]],[[[2770,6105],[2770,6070],[2748,6098],[2755,6142],[2770,6105]]],[[[2764,6164],[2782,6157],[2786,6132],[2759,6160],[2764,6164]]],[[[2744,6285],[2757,6268],[2750,6265],[2737,6285],[2744,6285]]],[[[2734,6314],[2729,6303],[2722,6324],[2730,6330],[2734,6314]]],[[[2758,6346],[2771,6338],[2743,6323],[2745,6350],[2758,6346]]],[[[2782,6565],[2805,6551],[2775,6531],[2760,6550],[2782,6565]]],[[[2800,6596],[2811,6592],[2802,6572],[2789,6604],[2800,6596]]],[[[2893,6675],[2901,6648],[2869,6646],[2873,6674],[2893,6675]]],[[[2880,6686],[2899,6693],[2896,6683],[2868,6687],[2880,6686]]],[[[2943,6687],[2915,6661],[2907,6693],[2927,6711],[2943,6687]]],[[[2964,6713],[2978,6701],[2947,6703],[2938,6713],[2964,6713]]],[[[3061,6794],[3062,6784],[3029,6773],[3032,6794],[3061,6794]]],[[[3050,6830],[3065,6830],[3059,6806],[3046,6817],[3050,6830]]],[[[3200,6891],[3183,6874],[3169,6884],[3176,6896],[3200,6891]]],[[[3168,6874],[3147,6853],[3131,6886],[3153,6875],[3168,6874]]],[[[3253,6915],[3238,6915],[3235,6931],[3244,6931],[3253,6915]]],[[[3237,6913],[3214,6907],[3203,6920],[3223,6938],[3237,6913]]],[[[3227,6967],[3188,6977],[3215,7006],[3238,6981],[3227,6967]]],[[[3321,7101],[3324,7076],[3313,7067],[3257,7062],[3321,7101]]],[[[3651,7346],[3663,7333],[3660,7319],[3649,7325],[3651,7346]]],[[[3633,7339],[3658,7299],[3628,7308],[3615,7351],[3633,7339]]],[[[3648,7351],[3644,7336],[3627,7357],[3642,7363],[3648,7351]]],[[[3629,7427],[3613,7424],[3581,7411],[3614,7439],[3629,7427]]],[[[3580,7414],[3570,7409],[3576,7431],[3617,7446],[3580,7414]]],[[[3699,7488],[3672,7467],[3667,7471],[3675,7487],[3699,7488]]],[[[3759,7501],[3747,7473],[3727,7478],[3750,7520],[3759,7501]]],[[[3773,7608],[3785,7573],[3736,7520],[3740,7562],[3773,7608]]],[[[3716,7662],[3713,7631],[3693,7638],[3701,7661],[3716,7662]]],[[[3809,7748],[3780,7719],[3765,7722],[3777,7749],[3809,7748]]],[[[3791,7809],[3797,7789],[3783,7773],[3755,7758],[3791,7809]]],[[[3814,7846],[3823,7829],[3809,7822],[3803,7832],[3814,7846]]],[[[3830,7885],[3824,7862],[3812,7870],[3813,7887],[3830,7885]]],[[[3971,8096],[3969,8064],[3943,8083],[3950,8101],[3971,8096]]],[[[3984,8190],[3983,8182],[3965,8175],[3980,8195],[3984,8190]]],[[[4054,8360],[4075,8347],[4039,8341],[4039,8358],[4054,8360]]],[[[4072,8359],[4062,8358],[4059,8367],[4066,8380],[4072,8359]]],[[[3826,8372],[3831,8341],[3794,8296],[3813,8365],[3826,8372]]],[[[3859,8374],[3843,8351],[3835,8373],[3863,8394],[3859,8374]]],[[[4024,8439],[4013,8439],[4022,8456],[4030,8450],[4024,8439]]],[[[3950,8456],[3946,8436],[3940,8449],[3930,8460],[3950,8456]]],[[[3920,8440],[3929,8414],[3900,8406],[3887,8379],[3868,8396],[3868,8430],[3920,8440]]],[[[4049,8500],[4009,8443],[3948,8407],[3978,8458],[3965,8474],[3993,8493],[4049,8500]]],[[[4187,8508],[4154,8495],[4161,8532],[4178,8540],[4187,8508]]],[[[4004,8514],[3985,8532],[4029,8533],[4018,8515],[4004,8514]]],[[[4255,8614],[4227,8601],[4226,8616],[4242,8644],[4255,8614]]],[[[4265,8646],[4263,8664],[4288,8650],[4267,8626],[4265,8646]]],[[[4018,8630],[4005,8632],[4008,8645],[4017,8645],[4018,8630]]],[[[4174,8672],[4190,8672],[4192,8647],[4157,8669],[4174,8672]]],[[[4280,8687],[4273,8700],[4282,8716],[4293,8708],[4280,8687]]],[[[4108,8681],[4067,8660],[4079,8704],[4131,8772],[4131,8720],[4108,8681]]],[[[4402,8972],[4427,8934],[4405,8912],[4411,8885],[4347,8859],[4324,8875],[4355,8879],[4340,8913],[4355,8940],[4400,8943],[4402,8972]]],[[[4594,8976],[4583,8973],[4580,9002],[4595,9008],[4594,8976]]],[[[5553,9194],[5561,9190],[5528,9163],[5532,9212],[5553,9194]]],[[[4490,8977],[4471,8969],[4472,8994],[4504,9015],[4490,8977]]],[[[4616,9050],[4630,9045],[4617,9023],[4596,9036],[4616,9050]]],[[[4434,9036],[4474,9017],[4462,8968],[4435,8949],[4388,8992],[4397,9014],[4434,9036]]],[[[4391,9028],[4409,9033],[4384,9003],[4379,9041],[4391,9028]]],[[[4606,9062],[4599,9059],[4596,9083],[4607,9071],[4606,9062]]],[[[4467,9047],[4447,9050],[4446,9063],[4454,9064],[4467,9047]]],[[[4592,9071],[4576,9048],[4553,9062],[4568,9105],[4594,9096],[4592,9071]]],[[[4425,9091],[4432,9055],[4420,9048],[4408,9057],[4425,9091]]],[[[4476,9096],[4491,9074],[4528,9061],[4496,9041],[4471,9086],[4476,9096]]],[[[4683,9132],[4676,9137],[4675,9169],[4684,9161],[4683,9132]]],[[[4774,9185],[4804,9172],[4813,9150],[4754,9152],[4753,9181],[4774,9185]]],[[[4843,9269],[4867,9228],[4848,9178],[4821,9162],[4791,9208],[4843,9269]]],[[[4869,9303],[4905,9283],[4873,9246],[4852,9298],[4869,9303]]],[[[4876,9399],[4883,9370],[4863,9379],[4864,9402],[4876,9399]]],[[[4940,9438],[4944,9418],[4926,9424],[4933,9441],[4940,9438]]],[[[5023,9477],[5051,9449],[5083,9441],[5024,9405],[4993,9434],[5023,9477]]],[[[1712,9534],[1697,9493],[1622,9482],[1587,9453],[1577,9474],[1647,9502],[1675,9535],[1712,9534]]],[[[2837,6137],[2847,6128],[2861,6143],[2864,6225],[2838,6190],[2804,6173],[2756,6246],[2808,6216],[2808,6249],[2774,6257],[2774,6317],[2832,6307],[2898,6331],[2951,6306],[2999,6341],[3049,6311],[3103,6327],[3106,6341],[3048,6320],[2986,6367],[2975,6324],[2939,6346],[2852,6338],[2825,6323],[2809,6354],[2778,6366],[2772,6425],[2831,6461],[2782,6487],[2782,6518],[2834,6545],[2839,6563],[2889,6540],[2997,6528],[2848,6572],[2810,6563],[2834,6599],[2804,6641],[2820,6655],[2854,6608],[2853,6645],[2921,6644],[2937,6675],[3031,6713],[2977,6718],[3009,6739],[2972,6738],[2981,6761],[3020,6745],[3043,6771],[3086,6769],[3142,6735],[3144,6774],[3186,6789],[3226,6784],[3219,6802],[3125,6778],[3116,6795],[3070,6791],[3065,6852],[3114,6874],[3143,6843],[3207,6860],[3223,6899],[3256,6854],[3254,6831],[3293,6829],[3234,6900],[3284,6904],[3282,6916],[3268,6913],[3248,6934],[3277,6938],[3279,6933],[3276,6952],[3314,6967],[3323,6991],[3380,6970],[3367,7013],[3438,7053],[3472,6989],[3455,6960],[3491,6959],[3490,6992],[3517,7000],[3583,6986],[3598,7001],[3566,7025],[3612,7076],[3668,7101],[3620,7126],[3671,7165],[3640,7169],[3602,7143],[3617,7110],[3596,7079],[3532,7034],[3467,7015],[3451,7067],[3481,7094],[3418,7059],[3418,7091],[3503,7139],[3469,7145],[3499,7216],[3555,7265],[3555,7291],[3591,7298],[3599,7336],[3626,7290],[3653,7276],[3670,7328],[3654,7368],[3710,7397],[3684,7402],[3717,7441],[3664,7398],[3633,7401],[3653,7431],[3707,7452],[3725,7476],[3747,7468],[3802,7496],[3840,7559],[3789,7501],[3768,7537],[3803,7582],[3793,7633],[3773,7616],[3765,7646],[3815,7745],[3843,7777],[3805,7768],[3823,7796],[3901,7831],[3954,7833],[3906,7846],[3842,7816],[3875,7848],[3836,7863],[3856,7880],[3862,7946],[3889,7960],[3894,7999],[3925,8002],[3898,8039],[3971,8056],[3984,8110],[4032,8130],[4047,8109],[4068,8132],[4100,8120],[4110,8135],[4064,8142],[4074,8162],[4019,8130],[4019,8148],[3979,8135],[4013,8192],[4034,8179],[4058,8201],[4020,8211],[4051,8236],[4106,8209],[4149,8246],[4108,8224],[4081,8226],[4071,8262],[4114,8296],[4088,8311],[4030,8258],[4023,8284],[4048,8291],[4021,8312],[4043,8335],[4077,8336],[4131,8359],[4149,8378],[4143,8378],[4136,8369],[4120,8376],[4106,8379],[4072,8380],[4095,8411],[4142,8414],[4143,8449],[4160,8428],[4157,8385],[4183,8343],[4187,8376],[4208,8350],[4209,8385],[4190,8403],[4197,8447],[4164,8472],[4214,8458],[4169,8483],[4220,8504],[4236,8481],[4313,8520],[4310,8551],[4258,8516],[4201,8512],[4206,8574],[4269,8615],[4315,8661],[4285,8660],[4307,8721],[4347,8749],[4335,8783],[4360,8847],[4398,8795],[4432,8809],[4379,8837],[4379,8860],[4415,8874],[4438,8829],[4476,8818],[4476,8781],[4494,8783],[4475,8829],[4454,8825],[4426,8878],[4434,8933],[4453,8950],[4503,8963],[4497,8924],[4518,8910],[4514,8952],[4528,8955],[4537,9010],[4565,8998],[4562,8851],[4539,8804],[4572,8846],[4587,8905],[4581,8963],[4644,9005],[4660,9055],[4723,9003],[4718,9051],[4704,9071],[4669,9076],[4632,9106],[4666,9144],[4703,9099],[4692,9135],[4736,9150],[4749,9138],[4812,9136],[4828,9084],[4863,9066],[4880,9092],[4839,9106],[4860,9124],[4842,9148],[4868,9173],[4858,9194],[4891,9245],[4932,9245],[4908,9261],[4923,9308],[4953,9294],[4939,9329],[4913,9329],[4917,9362],[4944,9340],[4936,9401],[4987,9394],[5005,9357],[5018,9381],[5061,9402],[5068,9391],[5007,9267],[5022,9278],[5032,9211],[5015,9183],[5022,9138],[5040,9142],[5067,9216],[5053,9221],[5092,9324],[5092,9351],[5111,9405],[5136,9438],[5151,9400],[5149,9363],[5123,9338],[5155,9343],[5159,9257],[5203,9350],[5195,9385],[5226,9426],[5181,9438],[5201,9450],[5186,9473],[5220,9459],[5211,9501],[5245,9491],[5272,9513],[5278,9486],[5314,9491],[5314,9454],[5298,9425],[5252,9415],[5293,9408],[5255,9357],[5312,9402],[5312,9374],[5284,9325],[5323,9308],[5342,9231],[5329,9315],[5333,9413],[5347,9465],[5372,9475],[5414,9455],[5416,9427],[5455,9448],[5454,9414],[5477,9450],[5582,9386],[5590,9401],[5620,9356],[5571,9330],[5547,9266],[5450,9255],[5399,9256],[5388,9238],[5508,9223],[5489,9196],[5519,9206],[5527,9154],[5560,9161],[5576,9184],[5564,9210],[5601,9202],[5638,9209],[5659,9172],[5664,9128],[5628,9120],[5599,9138],[5597,9081],[5581,9050],[5529,9010],[5532,8960],[5509,8926],[5490,8946],[5485,8979],[5515,9065],[5476,9121],[5392,9139],[5361,9160],[5320,9205],[5290,9182],[5252,9135],[5225,9140],[5190,9125],[5173,9058],[5157,9048],[5153,8993],[5162,8948],[5159,8892],[5178,8840],[5171,8799],[5127,8767],[5129,8714],[5106,8686],[5079,8712],[5041,8717],[4991,8740],[4976,8706],[4923,8666],[4900,8683],[4835,8679],[4823,8711],[4776,8761],[4725,8830],[4687,8832],[4662,8795],[4680,8769],[4669,8753],[4635,8767],[4625,8745],[4570,8732],[4600,8702],[4600,8626],[4575,8586],[4608,8572],[4579,8529],[4475,8562],[4432,8556],[4406,8575],[4379,8560],[4391,8447],[4367,8387],[4298,8421],[4246,8357],[4219,8248],[4188,8230],[4182,8204],[4224,8141],[4225,8096],[4186,8043],[4149,7954],[4123,7920],[4134,7856],[4086,7816],[4024,7804],[4042,7721],[4035,7678],[4036,7564],[4015,7494],[3940,7341],[4002,7302],[4012,7233],[3986,7174],[3894,7193],[3825,7151],[3762,7043],[3767,7001],[3741,6952],[3774,6868],[3759,6845],[3767,6790],[3760,6746],[3794,6644],[3777,6495],[3814,6454],[3841,6445],[3880,6395],[3857,6297],[3797,6278],[3816,6212],[3850,6144],[3836,6080],[3842,6028],[3790,5963],[3748,5954],[3756,5924],[3725,5862],[3747,5773],[3727,5677],[3695,5669],[3693,5697],[3682,5728],[3612,5755],[3600,5745],[3566,5820],[3574,5870],[3560,5908],[3573,5946],[3555,5954],[3549,5910],[3565,5863],[3549,5847],[3547,5769],[3531,5776],[3524,5734],[3471,5685],[3431,5685],[3368,5649],[3395,5649],[3249,5484],[3226,5481],[3190,5441],[3177,5477],[3158,5436],[3106,5419],[3017,5426],[2946,5455],[2973,5484],[2870,5537],[2865,5560],[2827,5578],[2797,5647],[2813,5729],[2839,5695],[2883,5686],[2862,5740],[2921,5792],[2897,5830],[2835,5805],[2825,5838],[2806,5799],[2767,5869],[2816,5932],[2853,5912],[2876,5935],[2841,5964],[2890,5991],[2919,6065],[2984,6134],[2928,6105],[2878,6052],[2875,6012],[2850,6018],[2855,6053],[2837,6076],[2815,6056],[2787,6072],[2784,6124],[2805,6163],[2820,6150],[2808,6168],[2862,6203],[2857,6144],[2837,6137]]],[[[3351,7049],[3352,7056],[3362,7056],[3368,7050],[3353,7046],[3369,7031],[3280,6993],[3251,7003],[3287,7042],[3342,7057],[3351,7049]]],[[[4021,8662],[4042,8649],[4054,8642],[4064,8566],[4036,8541],[4007,8557],[4044,8579],[4041,8595],[3986,8552],[3957,8569],[3990,8595],[4025,8605],[4021,8662]]],[[[4773,9266],[4803,9297],[4786,9313],[4816,9315],[4819,9336],[4816,9285],[4784,9234],[4713,9205],[4726,9247],[4694,9238],[4726,9274],[4773,9266]]],[[[4199,8593],[4192,8544],[4150,8536],[4140,8493],[4115,8501],[4104,8462],[4073,8476],[4050,8462],[4035,8461],[4061,8507],[4055,8538],[4084,8562],[4073,8614],[4091,8659],[4119,8663],[4119,8609],[4133,8605],[4118,8557],[4146,8599],[4158,8649],[4180,8635],[4199,8593]]],[[[4220,8743],[4241,8781],[4218,8805],[4266,8805],[4276,8820],[4250,8833],[4254,8851],[4291,8832],[4307,8874],[4333,8847],[4341,8804],[4323,8792],[4334,8751],[4284,8737],[4215,8700],[4238,8726],[4220,8743]]]]}},{"type":"Feature","id":"DE","properties":{"hc-group":"admin0","hc-middle-x":0.47,"hc-middle-y":0.53,"hc-key":"de","hc-a2":"DE","name":"Germany","labelrank":"2","country-abbrev":"Ger.","subregion":"Western Europe","region-wb":"Europe & Central Asia","iso-a3":"DEU","iso-a2":"DE","woe-id":"23424829","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[3422,2394],[3345,2433],[3297,2473],[3318,2430],[3264,2425],[3205,2468],[3164,2431],[3174,2408],[3006,2412],[2996,2470],[3016,2533],[3008,2560],[3045,2634],[3059,2708],[3116,2766],[3136,2805],[3085,2829],[3032,2830],[2984,2871],[2960,2853],[2879,2886],[2854,2877],[2819,2954],[2791,2962],[2825,3058],[2774,3087],[2759,3155],[2765,3182],[2809,3209],[2804,3257],[2779,3261],[2778,3297],[2746,3338],[2761,3382],[2737,3419],[2780,3452],[2766,3475],[2794,3519],[2779,3594],[2751,3642],[2788,3666],[2832,3644],[2910,3678],[2887,3701],[2952,3754],[2945,3818],[2897,3827],[2912,3874],[2959,3874],[2969,3933],[2988,3965],[2993,4045],[3013,4070],[2965,4084],[2978,4144],[3003,4166],[3128,4178],[3163,4128],[3144,4114],[3169,4087],[3189,4115],[3179,4143],[3231,4125],[3218,4165],[3235,4214],[3283,4206],[3343,4213],[3309,4223],[3280,4265],[3282,4328],[3250,4334],[3247,4367],[3286,4374],[3312,4400],[3259,4467],[3255,4513],[3282,4516],[3384,4487],[3384,4487],[3408,4503],[3483,4447],[3481,4410],[3608,4345],[3659,4353],[3658,4307],[3605,4266],[3628,4244],[3677,4260],[3725,4229],[3731,4256],[3770,4302],[3833,4308],[3872,4347],[3897,4397],[3961,4368],[3987,4389],[4002,4345],[4064,4296],[4103,4313],[4119,4289],[4122,4320],[4193,4256],[4187,4247],[4192,4239],[4137,4226],[4146,4291],[4124,4280],[4141,4255],[4127,4235],[4157,4207],[4206,4188],[4238,4067],[4231,4002],[4198,3974],[4196,3943],[4292,3870],[4276,3824],[4307,3777],[4322,3727],[4295,3647],[4322,3611],[4327,3566],[4367,3549],[4385,3489],[4354,3378],[4318,3372],[4293,3419],[4256,3423],[4269,3379],[4136,3323],[4109,3288],[4070,3282],[4030,3255],[3940,3221],[3900,3155],[3860,3197],[3894,3121],[3947,3082],[3923,3034],[4004,2918],[4043,2910],[4079,2860],[4139,2811],[4163,2815],[4205,2765],[4189,2693],[4151,2710],[4132,2649],[4062,2613],[4008,2569],[4059,2492],[4041,2456],[4066,2454],[4072,2390],[4021,2406],[4016,2438],[3965,2426],[3902,2446],[3900,2416],[3794,2409],[3724,2353],[3670,2350],[3632,2385],[3558,2401],[3566,2362],[3538,2323],[3467,2387],[3422,2394]],[[3343,4213],[3357,4210],[3357,4210],[3357,4210],[3357,4210],[3403,4138],[3374,4205],[3357,4210],[3357,4210],[3357,4210],[3357,4210],[3354,4215],[3343,4213]]],[[[3731,4262],[3727,4247],[3711,4250],[3726,4265],[3731,4262]]],[[[3291,4389],[3278,4399],[3284,4404],[3304,4405],[3291,4389]]],[[[3688,4393],[3698,4374],[3646,4389],[3665,4409],[3688,4393]]],[[[4044,4454],[4086,4437],[4077,4404],[4111,4369],[4066,4367],[4028,4339],[4002,4362],[4024,4377],[4008,4423],[4044,4454]]],[[[3239,4455],[3210,4462],[3221,4473],[3236,4473],[3239,4455]]],[[[3210,4514],[3193,4513],[3208,4554],[3204,4535],[3210,4514]]],[[[3319,2429],[3321,2426],[3321,2426],[3319,2429]]]]}},{"type":"Feature","id":"IE","properties":{"hc-group":"admin0","hc-middle-x":0.61,"hc-middle-y":0.6,"hc-key":"ie","hc-a2":"IE","name":"Ireland","labelrank":"3","country-abbrev":"Ire.","subregion":"Northern Europe","region-wb":"Europe & Central Asia","iso-a3":"IRL","iso-a2":"IE","woe-id":"23424803","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[135,4725],[124,4687],[114,4713],[82,4730],[135,4725]]],[[[746,4592],[766,4559],[727,4568],[735,4525],[725,4463],[742,4439],[725,4388],[719,4263],[663,4188],[656,4154],[613,4108],[605,4057],[529,4093],[512,4071],[418,4086],[382,4055],[338,4055],[299,4028],[255,4032],[161,3989],[135,4002],[56,3997],[36,4014],[-44,4006],[16,4039],[29,4070],[-68,4058],[-72,4082],[1,4110],[-65,4105],[-114,4157],[-78,4178],[-14,4185],[-5,4200],[-102,4220],[-96,4241],[-38,4256],[9,4227],[26,4264],[10,4280],[61,4287],[79,4310],[137,4294],[196,4300],[197,4332],[162,4301],[95,4324],[45,4325],[112,4353],[151,4397],[130,4404],[181,4452],[237,4436],[245,4471],[130,4499],[143,4528],[92,4528],[114,4544],[74,4553],[62,4608],[121,4610],[123,4652],[182,4646],[190,4674],[130,4679],[153,4697],[142,4783],[178,4806],[276,4765],[279,4723],[304,4760],[397,4731],[372,4762],[448,4791],[481,4819],[390,4835],[376,4862],[424,4888],[462,4881],[447,4915],[493,4969],[574,4962],[602,4975],[614,4903],[622,4975],[655,4972],[651,4996],[711,4932],[652,4902],[622,4890],[581,4822],[532,4831],[538,4795],[459,4770],[496,4684],[576,4637],[639,4710],[662,4684],[668,4641],[690,4626],[685,4588],[746,4592]]]]}},{"type":"Feature","id":"UA","properties":{"hc-group":"admin0","hc-middle-x":0.58,"hc-middle-y":0.41,"hc-key":"ua","hc-a2":"UA","name":"Ukraine","labelrank":"3","country-abbrev":"Ukr.","subregion":"Eastern Europe","region-wb":"Europe & Central Asia","iso-a3":"UKR","iso-a2":"UA","woe-id":"23424976","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[7455,2386],[7493,2450],[7501,2459],[7470,2406],[7455,2386]]],[[[7966,2676],[7994,2681],[8031,2696],[8047,2688],[7966,2676]]],[[[7864,2673],[7779,2665],[7752,2669],[7846,2675],[7864,2673]]],[[[5798,2837],[5793,2890],[5819,2928],[5819,2970],[5841,3040],[5904,3029],[5867,3073],[5861,3130],[5837,3173],[5900,3310],[5994,3455],[6035,3467],[6051,3510],[6010,3586],[6038,3595],[5989,3635],[5976,3670],[5922,3719],[5907,3769],[5926,3810],[5970,3801],[6008,3848],[6025,3898],[6082,3911],[6159,3947],[6262,3963],[6378,3967],[6462,3957],[6519,3969],[6542,3930],[6616,3948],[6634,3914],[6639,3956],[6687,3955],[6704,3988],[6737,3959],[6780,3978],[6813,3942],[6825,3992],[6865,4022],[6918,3960],[6961,4007],[7005,4004],[7053,4033],[7103,3994],[7140,3984],[7149,4017],[7106,4084],[7114,4151],[7144,4237],[7196,4244],[7228,4275],[7278,4286],[7305,4275],[7359,4306],[7358,4378],[7461,4389],[7495,4437],[7544,4450],[7559,4434],[7598,4468],[7657,4432],[7685,4375],[7746,4356],[7755,4326],[7716,4303],[7775,4229],[7767,4189],[7843,4209],[7871,4200],[7929,4227],[7978,4187],[8043,4111],[8032,4093],[8058,4058],[8114,4038],[8131,4069],[8177,4085],[8220,4059],[8238,4073],[8284,4059],[8325,4117],[8405,4175],[8447,4151],[8492,4103],[8559,4075],[8583,4097],[8571,4130],[8675,4127],[8730,4116],[8766,4149],[8806,4125],[8846,4141],[8909,4111],[8957,4148],[8960,4107],[9002,4059],[8986,3990],[8948,3965],[8976,3938],[9015,3941],[8984,3894],[8990,3846],[9021,3859],[9090,3791],[9081,3739],[9100,3657],[8932,3589],[8934,3537],[8873,3489],[8881,3393],[8905,3344],[8790,3293],[8767,3220],[8748,3228],[8710,3193],[8685,3138],[8660,3145],[8577,3079],[8526,3055],[8457,2941],[8432,2953],[8409,2894],[8369,2847],[8405,2786],[8463,2728],[8547,2667],[8583,2657],[8621,2681],[8638,2731],[8673,2711],[8692,2749],[8781,2772],[8770,2707],[8791,2664],[8756,2632],[8717,2630],[8685,2601],[8631,2618],[8538,2498],[8491,2476],[8444,2431],[8433,2370],[8377,2301],[8328,2290],[8246,2312],[8278,2340],[8252,2382],[8259,2429],[8232,2470],[8195,2485],[8168,2467],[8079,2498],[8006,2470],[7998,2503],[8048,2572],[8195,2713],[8166,2718],[8144,2762],[8135,2728],[8065,2759],[8051,2728],[7988,2706],[7938,2672],[7887,2672],[7783,2705],[7811,2745],[7721,2746],[7876,2771],[7896,2816],[7798,2795],[7766,2837],[7766,2785],[7693,2768],[7604,2727],[7565,2690],[7579,2673],[7551,2554],[7503,2462],[7472,2461],[7470,2423],[7440,2382],[7406,2436],[7425,2339],[7451,2343],[7468,2301],[7457,2266],[7445,2302],[7373,2312],[7282,2223],[7186,2225],[7150,2258],[7203,2283],[7190,2327],[7228,2391],[7223,2426],[7259,2448],[7256,2502],[7235,2519],[7222,2574],[7258,2609],[7279,2560],[7319,2610],[7355,2583],[7369,2620],[7405,2591],[7448,2613],[7404,2644],[7384,2725],[7302,2743],[7303,2774],[7273,2849],[7248,2829],[7174,2884],[7170,2984],[7141,3008],[7112,2987],[7070,3029],[7015,3026],[6981,3041],[6926,3031],[6850,3073],[6803,3068],[6749,3021],[6685,3015],[6652,2968],[6596,2941],[6585,2873],[6419,2808],[6400,2767],[6360,2740],[6283,2792],[6223,2767],[6132,2776],[6052,2762],[6000,2782],[5962,2732],[5928,2776],[5901,2766],[5872,2803],[5848,2799],[5824,2842],[5798,2837]],[[7457,2615],[7536,2553],[7547,2572],[7470,2626],[7457,2615]],[[8226,2751],[8230,2774],[8290,2767],[8336,2746],[8341,2716],[8384,2771],[8375,2732],[8448,2722],[8511,2641],[8573,2639],[8552,2659],[8454,2726],[8380,2809],[8334,2833],[8341,2785],[8309,2776],[8319,2817],[8239,2836],[8226,2787],[8209,2801],[8147,2787],[8226,2751]]]]}},{"type":"Feature","id":"FI","properties":{"hc-group":"admin0","hc-middle-x":0.58,"hc-middle-y":0.72,"hc-key":"fi","hc-a2":"FI","name":"Finland","labelrank":"3","country-abbrev":"Fin.","subregion":"Northern Europe","region-wb":"Europe & Central Asia","iso-a3":"FIN","iso-a2":"FI","woe-id":"23424812","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[4993,6096],[4976,6089],[4970,6088],[4977,6101],[4993,6096]]],[[[4992,6109],[5004,6120],[5016,6114],[5011,6102],[4992,6109]]],[[[4939,6118],[4961,6095],[4955,6084],[4930,6101],[4939,6118]]],[[[4959,6128],[4966,6130],[4964,6111],[4954,6118],[4959,6128]]],[[[4873,6139],[4861,6117],[4851,6132],[4858,6155],[4873,6139]]],[[[4902,6194],[4957,6174],[4947,6141],[4922,6163],[4931,6115],[4891,6105],[4867,6158],[4902,6194]]],[[[5277,6163],[5280,6143],[5272,6140],[5263,6163],[5277,6163]]],[[[5158,6164],[5156,6153],[5142,6149],[5147,6172],[5158,6164]]],[[[5191,6183],[5175,6160],[5167,6161],[5170,6182],[5191,6183]]],[[[5210,6170],[5199,6182],[5212,6195],[5210,6182],[5210,6170]]],[[[5722,6296],[5708,6311],[5710,6317],[5727,6305],[5722,6296]]],[[[5326,6209],[5314,6149],[5284,6144],[5272,6198],[5326,6209]]],[[[5243,6212],[5215,6212],[5220,6227],[5248,6226],[5243,6212]]],[[[5261,6239],[5254,6226],[5245,6233],[5256,6244],[5261,6239]]],[[[5222,6241],[5239,6247],[5240,6243],[5227,6236],[5222,6241]]],[[[5196,6222],[5169,6235],[5179,6259],[5200,6229],[5196,6222]]],[[[5114,6275],[5115,6255],[5107,6259],[5095,6277],[5114,6275]]],[[[5079,6373],[5065,6389],[5072,6396],[5084,6386],[5079,6373]]],[[[4937,7061],[4954,7049],[4977,7043],[4964,7024],[4937,7061]]],[[[5064,7083],[5069,7094],[5088,7090],[5076,7072],[5064,7083]]],[[[4972,7074],[4968,7065],[4954,7075],[4968,7088],[4972,7074]]],[[[5135,7240],[5138,7228],[5131,7222],[5121,7233],[5135,7240]]],[[[5322,7664],[5299,7629],[5286,7647],[5295,7662],[5322,7664]]],[[[5345,6190],[5331,6207],[5327,6241],[5281,6204],[5269,6214],[5287,6256],[5239,6250],[5173,6282],[5163,6310],[5142,6267],[5098,6304],[5093,6379],[5067,6421],[5089,6423],[5097,6469],[5073,6573],[5103,6565],[5073,6596],[5037,6676],[5017,6687],[5013,6796],[4984,6806],[4983,6864],[4964,6884],[4975,6939],[5002,6971],[5001,7006],[5028,6998],[4995,7046],[5044,7070],[5050,7045],[5104,7098],[5073,7134],[5102,7148],[5112,7214],[5146,7203],[5158,7255],[5202,7289],[5195,7332],[5255,7403],[5254,7443],[5289,7495],[5290,7539],[5322,7603],[5356,7625],[5385,7602],[5393,7628],[5369,7651],[5398,7650],[5360,7689],[5364,7730],[5355,7797],[5315,7825],[5266,7823],[5238,7871],[5223,7856],[5191,7866],[5161,7928],[5117,7968],[5095,8033],[5116,8076],[5106,8131],[5115,8151],[5047,8242],[5053,8324],[5012,8336],[5020,8369],[4997,8455],[5012,8473],[4945,8521],[4925,8569],[4893,8592],[4833,8607],[4808,8603],[4787,8627],[4732,8654],[4661,8729],[4625,8745],[4635,8767],[4669,8753],[4680,8769],[4662,8795],[4687,8832],[4725,8830],[4776,8761],[4823,8711],[4835,8679],[4900,8683],[4923,8666],[4976,8706],[4991,8740],[5041,8717],[5079,8712],[5106,8686],[5129,8714],[5127,8767],[5171,8799],[5178,8840],[5159,8892],[5162,8948],[5153,8993],[5157,9048],[5173,9058],[5190,9125],[5225,9140],[5252,9135],[5290,9182],[5320,9205],[5361,9160],[5392,9139],[5476,9121],[5515,9065],[5485,8979],[5490,8946],[5509,8926],[5467,8875],[5508,8866],[5495,8771],[5543,8678],[5621,8661],[5680,8597],[5730,8568],[5735,8514],[5708,8441],[5685,8340],[5696,8295],[5773,8209],[5793,8170],[5847,8111],[5891,8036],[5907,7989],[5863,7968],[5889,7883],[5880,7852],[5913,7851],[5920,7819],[5900,7782],[5939,7733],[5974,7739],[6000,7703],[5979,7678],[6002,7628],[6081,7597],[6094,7531],[6072,7463],[6049,7441],[6136,7380],[6185,7369],[6251,7333],[6261,7312],[6320,7265],[6313,7140],[6287,7068],[6260,7026],[6195,6828],[6171,6788],[6144,6709],[6100,6652],[6036,6533],[6004,6463],[5961,6425],[5915,6445],[5896,6417],[5808,6365],[5760,6377],[5762,6313],[5734,6307],[5718,6339],[5702,6300],[5659,6290],[5659,6272],[5580,6239],[5570,6198],[5551,6217],[5449,6162],[5424,6157],[5404,6119],[5356,6102],[5399,6143],[5377,6144],[5394,6176],[5356,6176],[5346,6188],[5344,6176],[5342,6158],[5333,6178],[5345,6190]]]]}},{"type":"Feature","id":"SE","properties":{"hc-group":"admin0","hc-middle-x":0.52,"hc-middle-y":0.34,"hc-key":"se","hc-a2":"SE","name":"Sweden","labelrank":"3","country-abbrev":"Swe.","subregion":"Northern Europe","region-wb":"Europe & Central Asia","iso-a3":"SWE","iso-a2":"SE","woe-id":"23424954","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[4581,5272],[4583,5238],[4535,5043],[4529,4978],[4507,4942],[4497,5032],[4523,5130],[4538,5137],[4566,5259],[4581,5272]]],[[[4891,5495],[4910,5490],[4866,5467],[4871,5487],[4891,5495]]],[[[3744,5428],[3733,5400],[3712,5397],[3709,5423],[3744,5428]]],[[[3751,5482],[3754,5447],[3710,5435],[3696,5456],[3732,5493],[3751,5482]]],[[[4523,5546],[4520,5537],[4508,5566],[4529,5555],[4523,5546]]],[[[4726,5760],[4721,5748],[4707,5740],[4707,5752],[4726,5760]]],[[[4703,5774],[4693,5756],[4681,5760],[4687,5772],[4703,5774]]],[[[4628,5737],[4620,5738],[4617,5776],[4629,5757],[4628,5737]]],[[[4738,5788],[4732,5773],[4722,5779],[4737,5805],[4738,5788]]],[[[4740,5856],[4743,5848],[4741,5838],[4713,5856],[4740,5856]]],[[[4755,5930],[4740,5895],[4731,5912],[4741,5929],[4755,5930]]],[[[4779,5953],[4788,5967],[4786,5948],[4777,5952],[4779,5953]]],[[[4790,6015],[4791,6010],[4776,6006],[4783,6029],[4790,6015]]],[[[4688,6191],[4710,6150],[4685,6166],[4681,6205],[4688,6191]]],[[[4563,6851],[4572,6851],[4569,6834],[4556,6831],[4563,6851]]],[[[4889,7192],[4898,7199],[4898,7178],[4891,7169],[4889,7192]]],[[[4984,7737],[4981,7732],[4983,7723],[4969,7739],[4984,7737]]],[[[5014,7760],[5021,7760],[5017,7746],[5009,7757],[5014,7760]]],[[[4737,5337],[4781,5427],[4820,5465],[4829,5444],[4861,5466],[4876,5446],[4837,5414],[4844,5332],[4867,5322],[4830,5286],[4840,5271],[4792,5232],[4785,5177],[4761,5169],[4776,5205],[4743,5265],[4752,5301],[4737,5337]]],[[[4625,8745],[4661,8729],[4732,8654],[4787,8627],[4808,8603],[4833,8607],[4893,8592],[4925,8569],[4945,8521],[5012,8473],[4997,8455],[5020,8369],[5012,8336],[5053,8324],[5047,8242],[5115,8151],[5106,8131],[5116,8076],[5095,8033],[5117,7968],[5161,7928],[5191,7866],[5167,7842],[5135,7857],[5097,7848],[5074,7811],[5013,7844],[5017,7815],[4988,7838],[4970,7786],[4995,7744],[4913,7774],[4968,7739],[4929,7731],[4946,7697],[4891,7682],[4922,7655],[4919,7613],[4879,7513],[4863,7506],[4936,7408],[4882,7303],[4872,7224],[4838,7205],[4820,7152],[4789,7151],[4754,7104],[4749,7076],[4715,7110],[4726,7067],[4695,7077],[4698,7043],[4673,6991],[4656,6999],[4622,6975],[4596,6913],[4609,6891],[4560,6858],[4538,6870],[4560,6814],[4521,6754],[4482,6780],[4472,6739],[4501,6688],[4488,6609],[4501,6542],[4458,6497],[4477,6471],[4490,6327],[4518,6257],[4502,6243],[4558,6233],[4571,6196],[4609,6224],[4619,6200],[4713,6127],[4678,6140],[4708,6106],[4754,6099],[4781,6040],[4762,6002],[4798,6008],[4748,5958],[4717,5912],[4660,5882],[4732,5872],[4721,5897],[4770,5878],[4763,5862],[4703,5859],[4729,5811],[4676,5782],[4660,5738],[4637,5738],[4632,5796],[4611,5784],[4615,5719],[4522,5639],[4417,5642],[4446,5623],[4502,5632],[4528,5600],[4497,5582],[4512,5550],[4514,5494],[4490,5435],[4465,5421],[4515,5366],[4494,5370],[4518,5303],[4498,5275],[4491,5215],[4513,5179],[4452,4992],[4446,4944],[4421,4893],[4415,4914],[4373,4924],[4334,4902],[4274,4907],[4232,4897],[4247,4864],[4201,4861],[4161,4798],[4160,4770],[4191,4714],[4163,4674],[4114,4683],[4016,4653],[3962,4674],[3956,4718],[3976,4746],[3953,4767],[3880,4916],[3927,4904],[3897,4958],[3939,4970],[3934,5026],[3912,5025],[3889,5074],[3853,5099],[3798,5256],[3779,5227],[3775,5302],[3742,5330],[3738,5364],[3762,5474],[3756,5503],[3707,5491],[3692,5535],[3665,5507],[3672,5577],[3645,5689],[3660,5725],[3693,5697],[3695,5669],[3727,5677],[3747,5773],[3725,5862],[3756,5924],[3748,5954],[3790,5963],[3842,6028],[3836,6080],[3850,6144],[3816,6212],[3797,6278],[3857,6297],[3880,6395],[3841,6445],[3814,6454],[3777,6495],[3794,6644],[3760,6746],[3767,6790],[3759,6845],[3774,6868],[3741,6952],[3767,7001],[3762,7043],[3825,7151],[3894,7193],[3986,7174],[4012,7233],[4002,7302],[3940,7341],[4015,7494],[4036,7564],[4035,7678],[4042,7721],[4024,7804],[4086,7816],[4134,7856],[4123,7920],[4149,7954],[4186,8043],[4225,8096],[4224,8141],[4182,8204],[4188,8230],[4219,8248],[4246,8357],[4298,8421],[4367,8387],[4391,8447],[4379,8560],[4406,8575],[4432,8556],[4475,8562],[4579,8529],[4608,8572],[4575,8586],[4600,8626],[4600,8702],[4570,8732],[4625,8745]]]]}},{"type":"Feature","id":"RU","properties":{"hc-group":"admin0","hc-middle-x":0.59,"hc-middle-y":0.45,"hc-key":"ru","hc-a2":"RU","name":"Russia","labelrank":"2","country-abbrev":"Rus.","subregion":"Eastern Europe","region-wb":"Europe & Central Asia","iso-a3":"RUS","iso-a2":"RU","woe-id":"23424936","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[6161,5739],[6243,5717],[6237,5759],[6194,5799],[6149,5776],[6124,5800],[6149,5876],[6122,5986],[6102,6021],[6106,6046],[6108,6072],[6115,6109],[6124,6119],[6130,6125],[6119,6126],[6130,6125],[6143,6139],[6126,6160],[6111,6169],[6091,6230],[6097,6261],[6148,6234],[6154,6294],[6188,6280],[6221,6303],[6234,6358],[6388,6364],[6381,6399],[6342,6396],[6317,6431],[6199,6407],[6158,6445],[6127,6446],[6091,6487],[6133,6467],[6112,6530],[6078,6516],[6061,6476],[6004,6463],[6036,6533],[6100,6652],[6144,6709],[6171,6788],[6195,6828],[6260,7026],[6287,7068],[6313,7140],[6320,7265],[6261,7312],[6251,7333],[6185,7369],[6136,7380],[6049,7441],[6072,7463],[6094,7531],[6081,7597],[6002,7628],[5979,7678],[6000,7703],[5974,7739],[5939,7733],[5900,7782],[5920,7819],[5913,7851],[5880,7852],[5889,7883],[5863,7968],[5907,7989],[5891,8036],[5847,8111],[5793,8170],[5773,8209],[5696,8295],[5685,8340],[5708,8441],[5735,8514],[5730,8568],[5680,8597],[5621,8661],[5543,8678],[5495,8771],[5508,8866],[5467,8875],[5509,8926],[5532,8960],[5529,9010],[5581,9050],[5597,9081],[5599,9138],[5628,9120],[5664,9128],[5659,9172],[5638,9209],[5685,9209],[5710,9193],[5747,9204],[5726,9251],[5760,9239],[5736,9277],[5746,9292],[5820,9258],[5849,9267],[5881,9239],[5863,9208],[5786,9206],[5779,9175],[5814,9193],[5836,9176],[5895,9185],[5883,9145],[5922,9186],[5944,9186],[5940,9121],[5965,9116],[5964,9160],[6046,9184],[6077,9175],[6141,9187],[6142,9207],[6217,9203],[6264,9182],[6330,9171],[6416,9139],[6441,9140],[6559,9072],[6574,9088],[6704,9042],[6761,9048],[6812,9033],[6845,8998],[6855,9022],[6905,9008],[6932,8944],[6953,8939],[6971,8891],[6994,8896],[7027,8838],[7034,8774],[7005,8628],[6980,8579],[6916,8501],[6838,8442],[6764,8422],[6626,8430],[6572,8412],[6444,8403],[6359,8434],[6325,8401],[6282,8430],[6243,8419],[6195,8445],[6200,8416],[6106,8461],[6091,8493],[6038,8497],[6028,8483],[6073,8455],[6081,8424],[6125,8392],[6171,8394],[6159,8338],[6198,8349],[6253,8343],[6252,8307],[6375,8282],[6437,8236],[6462,8184],[6427,8204],[6459,8148],[6442,8071],[6519,7996],[6512,7973],[6552,7949],[6545,7915],[6581,7883],[6562,7858],[6666,7811],[6708,7845],[6743,7813],[6782,7804],[6801,7775],[6860,7771],[6919,7785],[6942,7770],[7012,7842],[7014,7867],[6972,7910],[6972,7940],[6922,7956],[6891,7915],[6859,7919],[6783,7980],[6758,7982],[6722,8032],[6761,8060],[6742,8112],[6761,8128],[6859,8118],[6917,8089],[7033,8099],[7129,8080],[7147,8130],[7199,8115],[7240,8128],[7178,8223],[7129,8251],[7045,8320],[7033,8381],[7069,8437],[7094,8533],[7145,8580],[7172,8670],[7178,8751],[7236,8752],[7235,8729],[7303,8780],[7377,8744],[7353,8690],[7399,8745],[7436,8729],[7476,8684],[7413,8795],[7402,8910],[7374,8932],[7369,8976],[7324,9009],[7263,9008],[7238,9051],[7210,9191],[7194,9213],[7159,9318],[7139,9335],[7019,9368],[7027,9390],[7093,9384],[7223,9450],[7299,9457],[7325,9430],[7402,9407],[7463,9341],[7403,9282],[7343,9236],[7359,9198],[7338,9166],[7360,9106],[7449,9098],[7514,9042],[7542,9038],[7591,9079],[7600,9064],[7667,9142],[7689,9179],[7655,9230],[7644,9306],[7615,9329],[7639,9386],[7683,9428],[7668,9467],[7697,9484],[7728,9545],[7765,9652],[7773,9713],[7794,9714],[7870,9851],[7872,9851],[7892,9835],[7898,9795],[7929,9801],[7936,9851],[8020,9851],[8021,9833],[8055,9851],[9732,9851],[9732,2582],[9688,2568],[9659,2542],[9662,2497],[9589,2522],[9379,2615],[9285,2633],[9196,2619],[9107,2671],[9032,2637],[8968,2690],[8806,2713],[8834,2751],[8827,2795],[8896,2789],[8901,2761],[8965,2827],[9012,2828],[8978,2893],[8970,2849],[8950,2912],[8981,2954],[8977,3041],[9017,3090],[9065,3076],[9077,3107],[9018,3114],[8967,3143],[8941,3127],[8863,3194],[8924,3204],[8970,3283],[9035,3333],[9069,3391],[9105,3400],[9077,3467],[9026,3422],[8905,3344],[8881,3393],[8873,3489],[8934,3537],[8932,3589],[9100,3657],[9081,3739],[9090,3791],[9021,3859],[8990,3846],[8984,3894],[9015,3941],[8976,3938],[8948,3965],[8986,3990],[9002,4059],[8960,4107],[8957,4148],[8909,4111],[8846,4141],[8806,4125],[8766,4149],[8730,4116],[8675,4127],[8571,4130],[8583,4097],[8559,4075],[8492,4103],[8447,4151],[8405,4175],[8325,4117],[8284,4059],[8238,4073],[8220,4059],[8177,4085],[8131,4069],[8114,4038],[8058,4058],[8032,4093],[8043,4111],[7978,4187],[7929,4227],[7871,4200],[7843,4209],[7767,4189],[7775,4229],[7716,4303],[7755,4326],[7746,4356],[7685,4375],[7657,4432],[7598,4468],[7559,4434],[7544,4450],[7495,4437],[7461,4389],[7358,4378],[7359,4306],[7305,4275],[7278,4286],[7227,4334],[7171,4478],[7114,4511],[7126,4570],[7180,4585],[7246,4572],[7290,4620],[7320,4704],[7266,4719],[7254,4763],[7183,4772],[7129,4751],[7118,4826],[7024,4849],[6987,4904],[6960,4909],[6966,4950],[6889,4976],[6884,5017],[6907,5053],[6852,5115],[6861,5155],[6840,5206],[6818,5198],[6759,5240],[6715,5246],[6663,5224],[6610,5166],[6584,5189],[6576,5237],[6517,5241],[6489,5207],[6449,5241],[6400,5217],[6371,5238],[6371,5279],[6340,5347],[6284,5395],[6262,5427],[6240,5414],[6248,5510],[6241,5547],[6181,5574],[6177,5602],[6151,5595],[6159,5682],[6194,5705],[6161,5739]],[[5965,9116],[5965,9113],[5965,9113],[5965,9113],[5933,9064],[5966,9090],[5965,9113],[5965,9113],[5965,9113],[5967,9115],[5965,9116]]],[[[6120,6424],[6130,6432],[6151,6421],[6138,6413],[6120,6424]]],[[[6732,7851],[6707,7864],[6714,7869],[6731,7863],[6732,7851]]],[[[6624,8069],[6645,8030],[6594,8040],[6591,8054],[6624,8069]]],[[[6641,8087],[6652,8081],[6676,8089],[6657,8072],[6641,8087]]],[[[7211,8811],[7186,8812],[7174,8829],[7188,8834],[7211,8811]]],[[[7795,9756],[7773,9714],[7788,9768],[7821,9784],[7795,9756]]],[[[6041,9195],[6002,9182],[5996,9191],[6016,9204],[6041,9195]]],[[[5279,4758],[5218,4659],[5310,4654],[5333,4669],[5314,4740],[5326,4758],[5472,4718],[5549,4740],[5594,4700],[5595,4571],[5615,4546],[5604,4540],[5111,4489],[5209,4577],[5134,4570],[5118,4544],[5118,4544],[5118,4544],[5137,4642],[5191,4649],[5231,4685],[5268,4759],[5279,4758],[5279,4758]]],[[[5088,4491],[5086,4492],[5103,4518],[5103,4518],[5088,4491]]],[[[7186,8155],[7163,8163],[7191,8181],[7206,8168],[7216,8145],[7193,8153],[7230,8134],[7199,8120],[7145,8136],[7155,8151],[7186,8155]]],[[[5103,4518],[5118,4544],[5118,4544],[5118,4544],[5117,4538],[5103,4518],[5103,4518]]],[[[7399,9803],[7406,9851],[7585,9851],[7583,9843],[7591,9782],[7564,9727],[7510,9677],[7448,9702],[7399,9803]]]]}},{"type":"Feature","id":"GB","properties":{"hc-group":"admin0","hc-middle-x":0.18,"hc-middle-y":0.36,"hc-key":"gb","hc-a2":"GB","name":"United Kingdom","labelrank":"2","country-abbrev":"U.K.","subregion":"Northern Europe","region-wb":"Europe & Central Asia","iso-a3":"GBR","iso-a2":"GB","woe-id":"-90","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[1201,3080],[1215,3052],[1176,3062],[1178,3081],[1201,3080]]],[[[1423,3486],[1461,3456],[1411,3434],[1382,3463],[1423,3486]]],[[[968,4314],[973,4313],[980,4292],[962,4305],[968,4314]]],[[[1043,4292],[1070,4288],[1010,4250],[979,4298],[986,4332],[1036,4324],[1043,4292]]],[[[1017,4928],[983,4948],[995,5006],[1025,4996],[1017,4928]]],[[[1052,5006],[1038,5025],[1033,5063],[1050,5050],[1052,5006]]],[[[885,5073],[888,5028],[851,5030],[842,5065],[811,5045],[830,5088],[886,5105],[885,5073]]],[[[898,5153],[887,5135],[874,5134],[891,5156],[898,5153]]],[[[961,5153],[957,5129],[905,5060],[894,5095],[961,5153]]],[[[803,5273],[790,5294],[831,5297],[807,5283],[803,5273]]],[[[946,5292],[953,5262],[986,5245],[991,5217],[878,5206],[922,5254],[894,5284],[935,5305],[946,5292]]],[[[862,5305],[840,5306],[862,5321],[885,5326],[862,5305]]],[[[763,5445],[745,5435],[733,5442],[754,5460],[763,5445]]],[[[935,5412],[925,5393],[906,5415],[929,5426],[935,5412]]],[[[796,5561],[811,5523],[775,5475],[769,5517],[796,5561]]],[[[811,5581],[819,5559],[805,5558],[790,5574],[811,5581]]],[[[838,5625],[833,5583],[780,5621],[795,5637],[838,5625]]],[[[1551,5818],[1538,5794],[1529,5824],[1542,5826],[1551,5818]]],[[[1501,5850],[1508,5826],[1487,5818],[1473,5849],[1501,5850]]],[[[1527,5917],[1583,5845],[1561,5835],[1544,5861],[1509,5854],[1490,5887],[1502,5918],[1527,5917]]],[[[1607,5906],[1619,5887],[1606,5883],[1600,5908],[1607,5906]]],[[[1548,5925],[1554,5910],[1540,5912],[1538,5930],[1548,5925]]],[[[1590,5936],[1590,5908],[1582,5907],[1578,5924],[1590,5936]]],[[[1647,5941],[1626,5926],[1614,5932],[1619,5939],[1647,5941]]],[[[1569,5971],[1575,5943],[1558,5948],[1549,5970],[1569,5971]]],[[[1936,6281],[1938,6271],[1924,6271],[1919,6289],[1936,6281]]],[[[1871,6164],[1837,6082],[1824,6092],[1855,6186],[1824,6165],[1802,6213],[1845,6210],[1841,6261],[1817,6262],[1863,6293],[1847,6234],[1892,6239],[1871,6164]]],[[[1904,6323],[1916,6320],[1900,6254],[1883,6259],[1904,6323]]],[[[1949,6333],[1938,6303],[1918,6307],[1936,6350],[1949,6333]]],[[[1131,3142],[1108,3146],[1111,3154],[1140,3166],[1131,3142]]],[[[1023,4529],[989,4528],[1011,4570],[1049,4605],[1082,4612],[1081,4575],[1023,4529]]],[[[-207,-551],[-210,-560],[-212,-550],[-207,-551]]],[[[9095,-490],[9069,-492],[9034,-504],[9046,-485],[9095,-490]]],[[[9270,-328],[9261,-336],[9242,-332],[9236,-330],[9242,-332],[9237,-333],[9224,-339],[9224,-339],[9223,-335],[9219,-325],[9213,-324],[9213,-324],[9233,-318],[9262,-296],[9262,-296],[9261,-299],[9240,-313],[9270,-328]],[[9232,-327],[9232,-324],[9229,-325],[9230,-327],[9232,-327]]],[[[1830,3601],[1855,3592],[1936,3594],[1915,3521],[1853,3503],[1840,3469],[1808,3479],[1725,3455],[1705,3437],[1628,3471],[1555,3474],[1508,3461],[1496,3491],[1420,3494],[1303,3481],[1290,3456],[1211,3481],[1203,3467],[1138,3522],[1064,3519],[1015,3491],[1009,3452],[970,3405],[944,3406],[885,3466],[776,3474],[761,3448],[714,3448],[705,3410],[674,3387],[633,3446],[587,3433],[597,3463],[644,3463],[728,3514],[738,3539],[784,3551],[840,3602],[856,3659],[889,3647],[934,3701],[1004,3698],[1069,3668],[1134,3657],[1142,3691],[1204,3735],[1257,3796],[1213,3761],[1161,3761],[1116,3721],[1053,3733],[1010,3807],[985,3794],[929,3803],[923,3866],[870,3863],[817,3838],[812,3872],[776,3942],[818,3963],[861,3953],[986,4004],[1026,4074],[1039,4180],[985,4185],[961,4159],[918,4168],[955,4201],[1001,4221],[1048,4271],[1194,4279],[1224,4247],[1220,4281],[1244,4294],[1243,4326],[1286,4374],[1263,4388],[1277,4433],[1324,4481],[1291,4516],[1249,4491],[1225,4578],[1205,4617],[1266,4714],[1298,4729],[1242,4752],[1231,4719],[1190,4724],[1165,4701],[1132,4713],[1101,4753],[1094,4689],[1025,4757],[1006,4744],[1011,4691],[980,4761],[1017,4840],[1101,4927],[1065,4997],[1080,5048],[1101,5062],[1100,5109],[1065,5045],[1039,5069],[1010,5064],[1006,5031],[970,4998],[950,4923],[903,4915],[958,5036],[958,5101],[1001,5152],[978,5167],[1013,5213],[1069,5218],[1025,5232],[1071,5292],[995,5245],[951,5291],[971,5303],[922,5320],[932,5335],[982,5321],[988,5362],[1054,5463],[1027,5491],[1037,5556],[1062,5536],[1051,5587],[1058,5629],[1093,5647],[1112,5620],[1131,5643],[1167,5614],[1127,5685],[1148,5677],[1145,5729],[1192,5729],[1185,5753],[1229,5825],[1273,5794],[1400,5774],[1464,5786],[1521,5771],[1498,5737],[1500,5708],[1474,5681],[1325,5602],[1324,5566],[1362,5562],[1274,5481],[1382,5502],[1401,5520],[1462,5496],[1488,5501],[1546,5480],[1637,5469],[1654,5405],[1613,5366],[1592,5311],[1554,5245],[1512,5212],[1491,5171],[1429,5146],[1432,5118],[1469,5088],[1431,5067],[1403,5076],[1362,5041],[1296,5041],[1370,5007],[1422,5032],[1448,5025],[1519,4972],[1558,4882],[1586,4863],[1584,4742],[1603,4625],[1628,4576],[1711,4521],[1730,4457],[1778,4405],[1750,4383],[1756,4346],[1794,4266],[1763,4265],[1724,4304],[1739,4267],[1791,4218],[1813,4142],[1806,4103],[1743,4053],[1803,4017],[1843,4064],[1911,4056],[1963,4037],[2022,3985],[2034,3900],[2002,3841],[1988,3788],[1913,3738],[1898,3712],[1872,3728],[1844,3654],[1767,3648],[1809,3635],[1781,3619],[1823,3604],[1821,3625],[1842,3614],[1848,3599],[1830,3601]]],[[[989,5494],[996,5479],[1046,5460],[1035,5441],[976,5406],[984,5454],[937,5454],[922,5503],[886,5554],[922,5560],[917,5592],[950,5557],[973,5607],[990,5569],[977,5518],[989,5495],[995,5528],[1011,5545],[999,5516],[989,5494]]],[[[902,5785],[935,5767],[936,5789],[1032,5834],[1037,5783],[993,5753],[989,5703],[954,5676],[875,5640],[862,5666],[909,5691],[872,5716],[902,5785]]],[[[652,4902],[679,4888],[706,4926],[738,4913],[787,4923],[845,4898],[875,4749],[902,4722],[907,4664],[888,4631],[888,4687],[855,4642],[877,4622],[822,4614],[780,4566],[746,4592],[685,4588],[690,4626],[668,4641],[662,4684],[639,4710],[576,4637],[496,4684],[459,4770],[538,4795],[532,4831],[581,4822],[622,4890],[652,4902]]]]}},{"type":"Feature","id":"CY","properties":{"hc-group":"admin0","hc-middle-x":0.19,"hc-middle-y":0.71,"hc-key":"cy","hc-a2":"CY","name":"Cyprus","labelrank":"5","country-abbrev":"Cyp.","subregion":"Western Asia","region-wb":"Europe & Central Asia","iso-a3":"CYP","iso-a2":"CY","woe-id":"-90","continent":"Asia"},"geometry":{"type":"MultiPolygon","coordinates":[[[[9232,-327],[9230,-327],[9229,-325],[9232,-324],[9232,-327]]],[[[9291,-290],[9318,-310],[9270,-328],[9240,-313],[9261,-299],[9270,-299],[9291,-290]]],[[[9224,-339],[9216,-390],[9095,-490],[9046,-485],[9034,-504],[8942,-502],[8881,-414],[8923,-412],[8944,-370],[8955,-373],[8960,-360],[8968,-358],[9008,-383],[9083,-327],[9139,-309],[9162,-352],[9223,-335],[9224,-339],[9224,-339]]],[[[9236,-330],[9239,-329],[9245,-331],[9242,-332],[9236,-330]]]]}},{"type":"Feature","id":"PT","properties":{"hc-group":"admin0","hc-middle-x":0.48,"hc-middle-y":0.36,"hc-key":"pt","hc-a2":"PT","name":"Portugal","labelrank":"2","country-abbrev":"Port.","subregion":"Southern Europe","region-wb":"Europe & Central Asia","iso-a3":"PRT","iso-a2":"PT","woe-id":"23424925","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[-896,403],[-979,424],[-941,512],[-901,573],[-905,609],[-867,611],[-810,675],[-731,789],[-719,832],[-619,1023],[-609,1164],[-611,1246],[-566,1304],[-534,1320],[-441,1327],[-423,1288],[-463,1252],[-457,1228],[-354,1228],[-338,1204],[-257,1193],[-226,1223],[-107,1176],[-122,1124],[-54,1057],[-134,993],[-168,993],[-220,941],[-249,936],[-238,881],[-270,768],[-262,740],[-328,703],[-304,645],[-361,562],[-476,588],[-438,511],[-449,478],[-430,404],[-393,365],[-423,324],[-479,302],[-520,224],[-481,131],[-444,129],[-473,83],[-533,84],[-559,40],[-621,-20],[-621,-134],[-720,-162],[-761,-157],[-805,-120],[-906,-83],[-999,-89],[-917,33],[-889,160],[-903,172],[-858,231],[-850,282],[-874,333],[-937,322],[-929,397],[-896,403]],[[-896,403],[-894,402],[-893,403],[-893,403],[-893,404],[-850,411],[-856,453],[-893,404],[-893,404],[-893,404],[-896,403]]]}},{"type":"Feature","id":"GR","properties":{"hc-group":"admin0","hc-middle-x":0.21,"hc-middle-y":0.4,"hc-key":"gr","hc-a2":"GR","name":"Greece","labelrank":"3","country-abbrev":"Greece","subregion":"Southern Europe","region-wb":"Europe & Central Asia","iso-a3":"GRC","iso-a2":"GR","woe-id":"23424833","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[7604,-670],[7608,-677],[7582,-697],[7580,-676],[7604,-670]]],[[[7631,-545],[7629,-606],[7642,-663],[7609,-618],[7631,-545]]],[[[6616,-642],[6582,-647],[6584,-593],[6626,-630],[6616,-642]]],[[[7823,-305],[7805,-407],[7756,-489],[7713,-417],[7751,-352],[7823,-305]]],[[[7182,-475],[7175,-489],[7154,-487],[7169,-474],[7182,-475]]],[[[7617,-345],[7625,-359],[7626,-370],[7604,-352],[7617,-345]]],[[[7396,-360],[7364,-392],[7351,-370],[7370,-374],[7396,-360]]],[[[7712,-275],[7723,-272],[7731,-295],[7701,-292],[7712,-275]]],[[[6934,-404],[6941,-431],[6892,-453],[6893,-421],[6934,-404]]],[[[7139,-377],[7138,-400],[7105,-385],[7113,-363],[7139,-377]]],[[[7551,-257],[7495,-300],[7519,-256],[7566,-224],[7551,-257]]],[[[7286,-293],[7247,-333],[7238,-318],[7267,-285],[7286,-293]]],[[[6946,-321],[6976,-342],[6967,-360],[6944,-328],[6946,-321]]],[[[7485,-202],[7503,-208],[7503,-228],[7477,-234],[7485,-202]]],[[[7046,-318],[7057,-279],[7081,-269],[7077,-311],[7046,-318]]],[[[7438,-167],[7460,-189],[7449,-192],[7429,-173],[7438,-167]]],[[[7161,-259],[7139,-319],[7104,-279],[7140,-233],[7161,-259]]],[[[6896,-283],[6906,-283],[6912,-303],[6889,-307],[6896,-283]]],[[[6670,-285],[6660,-295],[6632,-304],[6650,-286],[6670,-285]]],[[[6880,-220],[6863,-259],[6857,-221],[6868,-204],[6880,-220]]],[[[7078,-153],[7107,-159],[7096,-177],[7075,-183],[7078,-153]]],[[[6996,-203],[6979,-218],[6972,-209],[6981,-174],[6996,-203]]],[[[7008,-125],[7047,-125],[7046,-153],[6989,-133],[7008,-125]]],[[[6842,-149],[6851,-166],[6828,-198],[6824,-172],[6842,-149]]],[[[7294,-67],[7241,-114],[7221,-109],[7237,-79],[7294,-67]]],[[[6639,-190],[6630,-189],[6618,-168],[6650,-162],[6639,-190]]],[[[7431,3],[7459,-6],[7400,-38],[7346,-22],[7402,12],[7431,3]]],[[[5994,-248],[6054,-276],[6018,-305],[5965,-262],[5994,-248]]],[[[6938,-60],[6962,-58],[6965,-113],[6899,-63],[6938,-60]]],[[[6631,-102],[6637,-116],[6621,-132],[6605,-127],[6631,-102]]],[[[5926,-73],[5995,-185],[5927,-181],[5891,-111],[5913,-123],[5926,-73]]],[[[7173,194],[7198,187],[7214,118],[7188,68],[7149,90],[7172,121],[7124,182],[7173,194]]],[[[5956,-15],[5922,-30],[5914,-10],[5935,41],[5956,-15]]],[[[6817,208],[6846,190],[6813,180],[6793,232],[6817,208]]],[[[6561,245],[6551,230],[6536,231],[6549,254],[6561,245]]],[[[6579,258],[6627,240],[6618,227],[6600,227],[6579,258]]],[[[6648,257],[6642,262],[6651,293],[6659,292],[6648,257]]],[[[7202,424],[7264,345],[7228,323],[7167,325],[7082,354],[7082,378],[7183,438],[7202,424]]],[[[5724,291],[5711,248],[5738,200],[5658,269],[5724,291]]],[[[6940,577],[6935,505],[6895,528],[6865,505],[6853,550],[6911,552],[6940,577]]],[[[6963,721],[6969,703],[6950,692],[6917,710],[6963,721]]],[[[6760,718],[6700,711],[6718,764],[6750,754],[6760,718]]],[[[5744,264],[5783,255],[5827,318],[5801,361],[5871,401],[5896,516],[5934,546],[5929,587],[5950,612],[5928,589],[5909,607],[5912,616],[5923,633],[5943,639],[5999,661],[6044,658],[6105,737],[6162,765],[6216,761],[6280,788],[6278,832],[6313,844],[6357,845],[6390,877],[6462,882],[6547,945],[6645,975],[6671,941],[6714,926],[6735,946],[6831,918],[6884,953],[6956,965],[7004,992],[7003,1040],[6967,1083],[7026,1104],[7089,1088],[7113,1024],[7056,973],[7081,892],[7028,811],[7001,837],[6909,825],[6792,840],[6750,785],[6693,803],[6656,791],[6651,758],[6606,719],[6537,721],[6516,691],[6698,573],[6683,554],[6645,591],[6585,607],[6539,585],[6563,553],[6615,538],[6624,490],[6572,514],[6539,553],[6475,558],[6465,536],[6562,466],[6484,465],[6455,542],[6391,557],[6335,600],[6368,621],[6348,644],[6284,580],[6307,555],[6293,492],[6303,460],[6382,390],[6406,343],[6451,320],[6522,239],[6421,274],[6413,212],[6469,187],[6414,128],[6373,132],[6413,100],[6482,95],[6494,74],[6546,82],[6561,42],[6600,46],[6643,14],[6710,7],[6747,-23],[6728,-40],[6771,-143],[6765,-175],[6694,-139],[6639,-80],[6561,-117],[6518,-163],[6560,-167],[6567,-224],[6600,-217],[6659,-253],[6559,-304],[6568,-274],[6468,-264],[6483,-304],[6544,-376],[6599,-470],[6582,-493],[6638,-568],[6536,-516],[6516,-478],[6473,-492],[6460,-548],[6474,-588],[6434,-581],[6425,-525],[6361,-440],[6312,-460],[6310,-537],[6267,-521],[6258,-476],[6217,-423],[6238,-381],[6193,-305],[6081,-240],[6125,-148],[6122,-119],[6178,-130],[6208,-82],[6250,-63],[6320,-96],[6430,-113],[6480,-142],[6472,-115],[6552,-79],[6528,-63],[6420,-38],[6387,-49],[6346,-11],[6348,-44],[6254,-40],[6153,-91],[6112,-64],[6067,-102],[6041,-37],[5962,66],[6037,101],[5958,121],[5942,72],[5926,106],[5844,158],[5781,221],[5788,250],[5744,264]]],[[[5953,-97],[5966,-114],[5953,-98],[5953,-97],[5953,-97]]],[[[6631,30],[6624,62],[6509,131],[6453,139],[6520,197],[6560,161],[6601,132],[6666,117],[6715,125],[6761,87],[6757,52],[6808,-13],[6863,-2],[6866,-56],[6825,-66],[6725,22],[6724,42],[6631,30]]],[[[7250,-790],[7315,-764],[7313,-812],[7342,-826],[7437,-752],[7449,-825],[7297,-867],[7180,-916],[7094,-936],[7076,-889],[6997,-876],[6845,-893],[6778,-888],[6780,-790],[6813,-814],[6809,-762],[6836,-803],[6921,-771],[6955,-832],[7059,-791],[7119,-780],[7144,-796],[7250,-790]]],[[[5953,-97],[5936,-73],[5949,-66],[5953,-97],[5953,-97]]]]}},{"type":"Feature","id":"LT","properties":{"hc-group":"admin0","hc-middle-x":0.6,"hc-middle-y":0.49,"hc-key":"lt","hc-a2":"LT","name":"Lithuania","labelrank":"5","country-abbrev":"Lith.","subregion":"Northern Europe","region-wb":"Europe & Central Asia","iso-a3":"LTU","iso-a2":"LT","woe-id":"23424875","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[5326,4758],[5306,4834],[5272,4895],[5254,4987],[5276,4994],[5291,5038],[5404,5112],[5500,5110],[5529,5132],[5570,5111],[5594,5133],[5661,5141],[5729,5127],[5783,5137],[5842,5202],[5891,5137],[5981,5144],[5992,5129],[6081,5080],[6104,5054],[6159,5044],[6157,4946],[6218,4942],[6193,4890],[6147,4887],[6131,4831],[6080,4791],[6092,4704],[6071,4634],[6100,4633],[6118,4585],[6086,4576],[6065,4618],[6026,4598],[6009,4557],[5972,4554],[5971,4502],[5952,4504],[5908,4464],[5875,4477],[5784,4444],[5756,4450],[5735,4513],[5628,4561],[5615,4546],[5595,4571],[5594,4700],[5549,4740],[5472,4718],[5326,4758]]],[[[5268,4759],[5285,4824],[5279,4758],[5279,4758],[5268,4759]]]]}},{"type":"Feature","id":"SI","properties":{"hc-group":"admin0","hc-middle-x":0.38,"hc-middle-y":0.62,"hc-key":"si","hc-a2":"SI","name":"Slovenia","labelrank":"6","country-abbrev":"Slo.","subregion":"Southern Europe","region-wb":"Europe & Central Asia","iso-a3":"SVN","iso-a2":"SI","woe-id":"23424945","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[4209,1816],[4208,1830],[4232,1848],[4269,1861],[4203,1909],[4209,1955],[4176,1965],[4209,2017],[4152,2046],[4216,2116],[4358,2100],[4386,2084],[4444,2153],[4485,2170],[4565,2165],[4597,2198],[4673,2197],[4685,2247],[4720,2250],[4777,2150],[4722,2143],[4734,2110],[4614,2053],[4602,2023],[4625,2002],[4624,1943],[4581,1934],[4542,1900],[4570,1837],[4525,1818],[4428,1842],[4403,1876],[4369,1822],[4286,1828],[4271,1801],[4209,1816]]]}},{"type":"Feature","id":"BA","properties":{"hc-group":"admin0","hc-middle-x":0.49,"hc-middle-y":0.42,"hc-key":"ba","hc-a2":"BA","name":"Bosnia and Herzegovina","labelrank":"5","country-abbrev":"B.H.","subregion":"Southern Europe","region-wb":"Europe & Central Asia","iso-a3":"BIH","iso-a2":"BA","woe-id":"23424761","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[5115,1130],[5093,1141],[5098,1143],[5095,1184],[5062,1201],[5019,1256],[5010,1291],[4976,1298],[4890,1365],[4881,1385],[4775,1482],[4747,1571],[4668,1659],[4662,1750],[4705,1771],[4754,1721],[4781,1716],[4805,1774],[4867,1774],[4893,1807],[4950,1773],[4966,1787],[5030,1773],[5051,1788],[5085,1763],[5120,1791],[5164,1799],[5229,1777],[5259,1787],[5308,1729],[5333,1734],[5363,1755],[5405,1749],[5402,1704],[5369,1637],[5388,1571],[5489,1511],[5411,1478],[5467,1428],[5478,1370],[5448,1380],[5419,1354],[5360,1334],[5401,1286],[5364,1294],[5321,1259],[5320,1192],[5282,1179],[5293,1114],[5315,1079],[5293,1053],[5161,1114],[5115,1130]]]}},{"type":"Feature","id":"MC","properties":{"hc-group":"admin0","hc-middle-x":0.47,"hc-middle-y":0.63,"hc-key":"mc","hc-a2":"MC","name":"Monaco","labelrank":"6","country-abbrev":"Mco.","subregion":"Western Europe","region-wb":"Europe & Central Asia","iso-a3":"MCO","iso-a2":"MC","woe-id":"23424892","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[2937,1301],[2930,1294],[2922,1295],[2931,1307],[2937,1301]]]}},{"type":"Feature","id":"AL","properties":{"hc-group":"admin0","hc-middle-x":0.3,"hc-middle-y":0.65,"hc-key":"al","hc-a2":"AL","name":"Albania","labelrank":"6","country-abbrev":"Alb.","subregion":"Southern Europe","region-wb":"Europe & Central Asia","iso-a3":"ALB","iso-a2":"AL","woe-id":"23424742","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[5907,643],[5905,642],[5902,648],[5906,645],[5907,643]]],[[[5827,689],[5837,642],[5854,641],[5874,648],[5902,648],[5899,643],[5909,635],[5909,634],[5900,609],[5912,616],[5909,607],[5928,589],[5924,584],[5929,587],[5934,546],[5896,516],[5871,401],[5801,361],[5827,318],[5783,255],[5744,264],[5742,309],[5697,364],[5606,400],[5597,470],[5552,520],[5567,589],[5586,602],[5563,670],[5575,704],[5543,743],[5572,801],[5568,869],[5519,873],[5511,904],[5511,941],[5532,960],[5508,1011],[5560,1115],[5593,1064],[5642,1094],[5686,1036],[5749,1011],[5782,938],[5778,915],[5765,818],[5792,778],[5782,752],[5827,689]]],[[[5503,1000],[5503,997],[5502,997],[5503,1000]]]]}},{"type":"Feature","id":"CNM","properties":{"hc-group":"admin0","hc-middle-x":0.66,"hc-middle-y":0.61,"hc-key":"cnm","hc-a2":"CN","name":"Cyprus No Mans Area","labelrank":"9","country-abbrev":null,"subregion":"Western Asia","region-wb":"Europe & Central Asia","iso-a3":"-99","iso-a2":null,"woe-id":"-99","continent":"Asia"},"geometry":{"type":"MultiPolygon","coordinates":[[[[9288,-289],[9288,-289],[9291,-290],[9270,-299],[9261,-299],[9262,-296],[9276,-291],[9288,-289]]],[[[8944,-370],[8947,-367],[8947,-367],[8952,-370],[8956,-361],[8958,-360],[8960,-360],[8955,-373],[8944,-370]]],[[[8968,-358],[8968,-358],[8973,-358],[9012,-377],[9084,-318],[9132,-299],[9160,-323],[9213,-324],[9213,-324],[9219,-325],[9223,-335],[9162,-352],[9139,-309],[9083,-327],[9008,-383],[8968,-358]]]]}},{"type":"Feature","id":"NC","properties":{"hc-group":"admin0","hc-middle-x":0.59,"hc-middle-y":0.61,"hc-key":"nc","hc-a2":"NC","name":"Northern Cyprus","labelrank":"6","country-abbrev":"N. Cy.","subregion":"Western Asia","region-wb":"Europe & Central Asia","iso-a3":"-99","iso-a2":"NC","woe-id":"-90","continent":"Asia"},"geometry":{"type":"MultiPolygon","coordinates":[[[[9160,-323],[9132,-299],[9084,-318],[9012,-377],[8973,-358],[9011,-353],[9001,-280],[9126,-260],[9253,-171],[9359,-64],[9360,-79],[9246,-235],[9288,-289],[9276,-291],[9262,-296],[9262,-296],[9233,-318],[9213,-324],[9213,-324],[9160,-323]]],[[[8947,-367],[8951,-364],[8956,-361],[8952,-370],[8947,-367]]]]}},{"type":"Feature","id":"RS","properties":{"hc-group":"admin0","hc-middle-x":0.47,"hc-middle-y":0.51,"hc-key":"rs","hc-a2":"RS","name":"Republic of Serbia","labelrank":"5","country-abbrev":"Serb.","subregion":"Southern Europe","region-wb":"Europe & Central Asia","iso-a3":"SRB","iso-a2":"RS","woe-id":"-90","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[5333,1734],[5332,1816],[5399,1831],[5390,1853],[5313,1878],[5308,1936],[5265,2001],[5272,2037],[5302,2065],[5342,2061],[5393,2121],[5468,2131],[5533,2124],[5557,2094],[5657,2038],[5663,1963],[5714,1924],[5826,1887],[5805,1838],[5841,1827],[5814,1802],[5881,1761],[5954,1762],[5997,1730],[6036,1801],[6113,1770],[6059,1736],[6116,1666],[6105,1620],[6067,1588],[6066,1533],[6118,1443],[6229,1382],[6192,1283],[6146,1270],[6144,1181],[6170,1136],[6140,1104],[6119,1116],[5976,1056],[5999,1185],[5913,1193],[5912,1226],[5882,1230],[5837,1284],[5780,1298],[5772,1323],[5728,1295],[5747,1275],[5690,1184],[5685,1207],[5636,1220],[5590,1254],[5522,1261],[5419,1354],[5448,1380],[5478,1370],[5467,1428],[5411,1478],[5489,1511],[5388,1571],[5369,1637],[5402,1704],[5405,1749],[5363,1755],[5333,1734]]]}},{"type":"Feature","id":"RO","properties":{"hc-group":"admin0","hc-middle-x":0.45,"hc-middle-y":0.54,"hc-key":"ro","hc-a2":"RO","name":"Romania","labelrank":"3","country-abbrev":"Rom.","subregion":"Eastern Europe","region-wb":"Europe & Central Asia","iso-a3":"ROU","iso-a2":"RO","woe-id":"23424933","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[6059,1736],[6113,1770],[6036,1801],[5997,1730],[5954,1762],[5881,1761],[5814,1802],[5841,1827],[5805,1838],[5826,1887],[5714,1924],[5663,1963],[5657,2038],[5557,2094],[5533,2124],[5575,2150],[5605,2141],[5645,2188],[5703,2199],[5724,2302],[5756,2338],[5767,2428],[5790,2458],[5814,2573],[5856,2646],[5943,2693],[5962,2732],[6000,2782],[6052,2762],[6132,2776],[6223,2767],[6283,2792],[6360,2740],[6400,2767],[6419,2808],[6585,2873],[6596,2941],[6652,2968],[6687,2976],[6729,2959],[6775,2912],[6821,2832],[6884,2789],[6953,2710],[7003,2689],[7070,2596],[7089,2531],[7089,2403],[7132,2304],[7114,2288],[7150,2258],[7186,2225],[7282,2223],[7373,2312],[7445,2302],[7457,2266],[7476,2165],[7469,2151],[7380,2113],[7337,2172],[7324,2132],[7352,2077],[7322,2035],[7356,2049],[7322,1963],[7352,1873],[7353,1794],[7278,1781],[7223,1793],[7199,1830],[7163,1808],[7144,1825],[7089,1809],[7055,1833],[7006,1837],[6838,1749],[6780,1652],[6698,1611],[6610,1620],[6512,1615],[6444,1587],[6264,1592],[6183,1561],[6168,1606],[6193,1640],[6116,1666],[6115,1666],[6059,1736]]]}},{"type":"Feature","id":"ME","properties":{"hc-group":"admin0","hc-middle-x":0.29,"hc-middle-y":0.67,"hc-key":"me","hc-a2":"ME","name":"Montenegro","labelrank":"6","country-abbrev":"Mont.","subregion":"Southern Europe","region-wb":"Europe & Central Asia","iso-a3":"MNE","iso-a2":"ME","woe-id":"20069817","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[5642,1094],[5593,1064],[5560,1115],[5508,1011],[5505,1006],[5503,1000],[5502,997],[5456,988],[5511,941],[5511,904],[5519,873],[5473,892],[5463,922],[5396,986],[5377,978],[5344,1027],[5311,1013],[5298,1030],[5293,1053],[5315,1079],[5293,1114],[5282,1179],[5320,1192],[5321,1259],[5364,1294],[5401,1286],[5360,1334],[5419,1354],[5522,1261],[5590,1254],[5636,1220],[5685,1207],[5690,1184],[5626,1144],[5642,1094]]]}},{"type":"Feature","id":"LI","properties":{"hc-group":"admin0","hc-middle-x":0.62,"hc-middle-y":0.51,"hc-key":"li","hc-a2":"LI","name":"Liechtenstein","labelrank":"6","country-abbrev":"Liech.","subregion":"Western Europe","region-wb":"Europe & Central Asia","iso-a3":"LIE","iso-a2":"LI","woe-id":"23424879","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[3383,2312],[3402,2266],[3395,2252],[3374,2254],[3383,2312]]]}},{"type":"Feature","id":"AT","properties":{"hc-group":"admin0","hc-middle-x":0.68,"hc-middle-y":0.5,"hc-key":"at","hc-a2":"AT","name":"Austria","labelrank":"4","country-abbrev":"Aust.","subregion":"Western Europe","region-wb":"Europe & Central Asia","iso-a3":"AUT","iso-a2":"AT","woe-id":"23424750","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[3395,2252],[3402,2266],[3383,2312],[3409,2354],[3390,2383],[3413,2380],[3422,2394],[3467,2387],[3538,2323],[3566,2362],[3558,2401],[3632,2385],[3670,2350],[3724,2353],[3794,2409],[3900,2416],[3902,2446],[3965,2426],[4016,2438],[4021,2406],[4072,2390],[4066,2454],[4041,2456],[4059,2492],[4008,2569],[4062,2613],[4132,2649],[4151,2710],[4189,2693],[4205,2765],[4257,2717],[4304,2710],[4329,2736],[4376,2724],[4392,2779],[4415,2776],[4422,2848],[4476,2844],[4558,2815],[4583,2821],[4627,2791],[4689,2788],[4702,2808],[4787,2794],[4804,2763],[4792,2692],[4860,2595],[4836,2551],[4854,2508],[4801,2493],[4763,2511],[4726,2482],[4770,2475],[4777,2426],[4737,2393],[4759,2293],[4717,2291],[4685,2247],[4673,2197],[4597,2198],[4565,2165],[4485,2170],[4444,2153],[4386,2084],[4358,2100],[4216,2116],[4122,2120],[3956,2154],[3895,2240],[3913,2265],[3823,2232],[3737,2234],[3694,2212],[3676,2170],[3567,2197],[3550,2234],[3500,2191],[3450,2218],[3449,2240],[3395,2252]]]}},{"type":"Feature","id":"SK","properties":{"hc-group":"admin0","hc-middle-x":0.4,"hc-middle-y":0.46,"hc-key":"sk","hc-a2":"SK","name":"Slovakia","labelrank":"6","country-abbrev":"Svk.","subregion":"Eastern Europe","region-wb":"Europe & Central Asia","iso-a3":"SVK","iso-a2":"SK","woe-id":"23424877","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[4860,2595],[4792,2692],[4804,2763],[4839,2840],[4910,2834],[4968,2872],[5005,2912],[5019,2981],[5077,3047],[5131,3062],[5160,3030],[5198,3039],[5241,3102],[5283,3050],[5310,3051],[5313,2997],[5371,2995],[5381,3035],[5411,3063],[5481,3078],[5533,3055],[5545,3087],[5585,3108],[5692,3104],[5736,3063],[5841,3040],[5819,2970],[5819,2928],[5793,2890],[5798,2837],[5724,2806],[5656,2860],[5600,2830],[5535,2843],[5479,2822],[5459,2748],[5381,2693],[5328,2714],[5305,2677],[5184,2641],[5168,2619],[5187,2581],[5078,2545],[4999,2535],[4947,2553],[4897,2596],[4860,2595]]]}},{"type":"Feature","id":"HU","properties":{"hc-group":"admin0","hc-middle-x":0.45,"hc-middle-y":0.54,"hc-key":"hu","hc-a2":"HU","name":"Hungary","labelrank":"5","country-abbrev":"Hun.","subregion":"Eastern Europe","region-wb":"Europe & Central Asia","iso-a3":"HUN","iso-a2":"HU","woe-id":"23424844","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[4777,2150],[4720,2250],[4685,2247],[4717,2291],[4759,2293],[4737,2393],[4777,2426],[4770,2475],[4726,2482],[4763,2511],[4801,2493],[4854,2508],[4836,2551],[4860,2595],[4897,2596],[4947,2553],[4999,2535],[5078,2545],[5187,2581],[5168,2619],[5184,2641],[5305,2677],[5328,2714],[5381,2693],[5459,2748],[5479,2822],[5535,2843],[5600,2830],[5656,2860],[5724,2806],[5798,2837],[5824,2842],[5848,2799],[5872,2803],[5901,2766],[5928,2776],[5962,2732],[5943,2693],[5856,2646],[5814,2573],[5790,2458],[5767,2428],[5756,2338],[5724,2302],[5703,2199],[5645,2188],[5605,2141],[5575,2150],[5533,2124],[5468,2131],[5393,2121],[5342,2061],[5302,2065],[5272,2037],[5227,2026],[5180,1971],[5138,1979],[5070,1968],[5008,2009],[4959,2009],[4924,2054],[4877,2075],[4851,2116],[4777,2150]]]}},{"type":"Feature","id":"AD","properties":{"hc-group":"admin0","hc-middle-x":0.48,"hc-middle-y":0.52,"hc-key":"ad","hc-a2":"AD","name":"Andorra","labelrank":"6","country-abbrev":"And.","subregion":"Southern Europe","region-wb":"Europe & Central Asia","iso-a3":"AND","iso-a2":"AD","woe-id":"23424744","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[1691,1033],[1645,1016],[1634,1067],[1697,1064],[1691,1033]]]}},{"type":"Feature","id":"LU","properties":{"hc-group":"admin0","hc-middle-x":0.5,"hc-middle-y":0.58,"hc-key":"lu","hc-a2":"LU","name":"Luxembourg","labelrank":"6","country-abbrev":"Lux.","subregion":"Western Europe","region-wb":"Europe & Central Asia","iso-a3":"LUX","iso-a2":"LU","woe-id":"23424881","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[2791,2962],[2752,2975],[2719,2961],[2688,2991],[2708,3020],[2680,3091],[2737,3172],[2759,3155],[2774,3087],[2825,3058],[2791,2962]]]}},{"type":"Feature","id":"CH","properties":{"hc-group":"admin0","hc-middle-x":0.51,"hc-middle-y":0.49,"hc-key":"ch","hc-a2":"CH","name":"Switzerland","labelrank":"4","country-abbrev":"Switz.","subregion":"Western Europe","region-wb":"Europe & Central Asia","iso-a3":"CHE","iso-a2":"CH","woe-id":"23424957","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[3006,2412],[3174,2408],[3164,2431],[3205,2468],[3264,2425],[3318,2430],[3319,2429],[3321,2426],[3321,2426],[3378,2377],[3390,2383],[3409,2354],[3383,2312],[3374,2254],[3395,2252],[3449,2240],[3450,2218],[3500,2191],[3550,2234],[3567,2197],[3551,2141],[3562,2101],[3495,2121],[3486,2071],[3509,2022],[3486,2010],[3470,2049],[3389,2031],[3365,2089],[3330,2087],[3331,2036],[3272,1939],[3287,1913],[3255,1898],[3243,1967],[3196,1984],[3162,2022],[3166,2075],[3091,2025],[3097,1988],[3039,1928],[2981,1950],[2908,1923],[2876,1937],[2844,1995],[2835,2080],[2852,2092],[2803,2112],[2741,2096],[2720,2051],[2731,2035],[2664,2007],[2704,2071],[2688,2089],[2706,2139],[2768,2184],[2773,2237],[2805,2247],[2897,2346],[2862,2351],[2892,2392],[2964,2369],[3006,2412],[3006,2412]]]}},{"type":"Feature","id":"BE","properties":{"hc-group":"admin0","hc-middle-x":0.53,"hc-middle-y":0.37,"hc-key":"be","hc-a2":"BE","name":"Belgium","labelrank":"2","country-abbrev":"Belg.","subregion":"Western Europe","region-wb":"Europe & Central Asia","iso-a3":"BEL","iso-a2":"BE","woe-id":"23424757","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[2759,3155],[2737,3172],[2680,3091],[2708,3020],[2688,2991],[2629,2982],[2536,3076],[2519,3074],[2510,3129],[2529,3175],[2492,3162],[2486,3135],[2443,3122],[2390,3138],[2408,3217],[2377,3244],[2310,3245],[2310,3281],[2244,3311],[2227,3382],[2161,3376],[2129,3413],[2124,3485],[2239,3543],[2280,3553],[2285,3515],[2330,3523],[2380,3495],[2437,3537],[2452,3509],[2444,3537],[2467,3530],[2486,3564],[2511,3546],[2537,3566],[2561,3534],[2581,3557],[2614,3492],[2660,3498],[2723,3456],[2690,3344],[2746,3338],[2778,3297],[2779,3261],[2804,3257],[2809,3209],[2765,3182],[2759,3155]]]}},{"type":"Feature","id":"KV","properties":{"hc-group":"admin0","hc-middle-x":0.89,"hc-middle-y":0.48,"hc-key":"kv","hc-a2":"KV","name":"Kosovo","labelrank":"6","country-abbrev":"Kos.","subregion":"Southern Europe","region-wb":"Europe & Central Asia","iso-a3":"-99","iso-a2":"KV","woe-id":"-90","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[5778,915],[5782,938],[5749,1011],[5686,1036],[5642,1094],[5626,1144],[5690,1184],[5747,1275],[5728,1295],[5772,1323],[5780,1298],[5837,1284],[5882,1230],[5912,1226],[5913,1193],[5999,1185],[5976,1056],[5945,1049],[5913,1002],[5878,1026],[5813,977],[5816,931],[5778,915]]]}},{"type":"Feature","id":"PL","properties":{"hc-group":"admin0","hc-middle-x":0.56,"hc-middle-y":0.5,"hc-key":"pl","hc-a2":"PL","name":"Poland","labelrank":"3","country-abbrev":"Pol.","subregion":"Eastern Europe","region-wb":"Europe & Central Asia","iso-a3":"POL","iso-a2":"PL","woe-id":"23424923","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[5615,4546],[5628,4561],[5735,4513],[5756,4450],[5791,4360],[5868,4241],[5895,4126],[5890,4104],[5817,4042],[5789,3973],[5882,3930],[5894,3906],[5886,3834],[5907,3769],[5922,3719],[5976,3670],[5989,3635],[6038,3595],[6010,3586],[6051,3510],[6035,3467],[5994,3455],[5900,3310],[5837,3173],[5861,3130],[5867,3073],[5904,3029],[5841,3040],[5736,3063],[5692,3104],[5585,3108],[5545,3087],[5533,3055],[5481,3078],[5411,3063],[5381,3035],[5371,2995],[5313,2997],[5310,3051],[5283,3050],[5241,3102],[5198,3039],[5160,3030],[5131,3062],[5114,3108],[5088,3112],[5067,3169],[4967,3187],[4931,3173],[4880,3222],[4880,3250],[4830,3248],[4797,3277],[4739,3283],[4762,3230],[4718,3186],[4643,3260],[4615,3272],[4648,3301],[4636,3338],[4591,3335],[4470,3369],[4453,3363],[4435,3411],[4378,3418],[4385,3380],[4354,3378],[4385,3489],[4367,3549],[4327,3566],[4322,3611],[4295,3647],[4322,3727],[4307,3777],[4276,3824],[4292,3870],[4196,3943],[4198,3974],[4231,4002],[4238,4067],[4206,4188],[4249,4181],[4266,4236],[4192,4239],[4187,4247],[4193,4256],[4220,4252],[4288,4290],[4511,4381],[4576,4464],[4637,4484],[4699,4531],[4787,4562],[4858,4575],[4910,4497],[4917,4464],[4989,4447],[5048,4464],[5048,4464],[5027,4450],[5056,4436],[5111,4489],[5604,4540],[5615,4546]]],[[[5086,4492],[5088,4491],[5060,4472],[5060,4472],[5086,4492]]],[[[5048,4464],[5060,4472],[5060,4472],[5050,4465],[5048,4464],[5048,4464]]]]}},{"type":"Feature","id":"MK","properties":{"hc-group":"admin0","hc-middle-x":0.49,"hc-middle-y":0.49,"hc-key":"mk","hc-a2":"MK","name":"Macedonia","labelrank":"6","country-abbrev":"Mkd.","subregion":"Southern Europe","region-wb":"Europe & Central Asia","iso-a3":"MKD","iso-a2":"MK","woe-id":"23424890","continent":"Europe"},"geometry":{"type":"MultiPolygon","coordinates":[[[[5909,635],[5910,635],[5909,634],[5909,635]]],[[[6140,1104],[6189,1058],[6262,1037],[6313,957],[6303,916],[6313,844],[6278,832],[6280,788],[6216,761],[6162,765],[6105,737],[6044,658],[5999,661],[5943,639],[5902,680],[5907,643],[5906,645],[5902,648],[5902,648],[5874,648],[5854,641],[5866,703],[5827,689],[5782,752],[5792,778],[5765,818],[5778,915],[5816,931],[5813,977],[5878,1026],[5913,1002],[5945,1049],[5976,1056],[6119,1116],[6140,1104]]]]}},{"type":"Feature","id":"LV","properties":{"hc-group":"admin0","hc-middle-x":0.68,"hc-middle-y":0.47,"hc-key":"lv","hc-a2":"LV","name":"Latvia","labelrank":"5","country-abbrev":"Lat.","subregion":"Northern Europe","region-wb":"Europe & Central Asia","iso-a3":"LVA","iso-a2":"LV","woe-id":"23424874","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[6371,5238],[6337,5217],[6307,5156],[6308,5118],[6209,5104],[6192,5064],[6159,5044],[6104,5054],[6081,5080],[5992,5129],[5981,5144],[5891,5137],[5842,5202],[5783,5137],[5729,5127],[5661,5141],[5594,5133],[5570,5111],[5529,5132],[5500,5110],[5404,5112],[5291,5038],[5276,4994],[5254,4987],[5233,5037],[5222,5208],[5266,5271],[5258,5343],[5288,5426],[5332,5447],[5414,5508],[5433,5460],[5517,5412],[5549,5343],[5626,5314],[5663,5335],[5716,5410],[5714,5443],[5669,5588],[5699,5617],[5801,5677],[5861,5642],[5926,5637],[5934,5617],[6022,5561],[6071,5605],[6151,5595],[6177,5602],[6181,5574],[6241,5547],[6248,5510],[6240,5414],[6262,5427],[6284,5395],[6340,5347],[6371,5279],[6371,5238]]]}},{"type":"Feature","id":"BY","properties":{"hc-group":"admin0","hc-middle-x":0.48,"hc-middle-y":0.57,"hc-key":"by","hc-a2":"BY","name":"Belarus","labelrank":"4","country-abbrev":"Bela.","subregion":"Eastern Europe","region-wb":"Europe & Central Asia","iso-a3":"BLR","iso-a2":"BY","woe-id":"23424765","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[6159,5044],[6192,5064],[6209,5104],[6308,5118],[6307,5156],[6337,5217],[6371,5238],[6400,5217],[6449,5241],[6489,5207],[6517,5241],[6576,5237],[6584,5189],[6610,5166],[6663,5224],[6715,5246],[6759,5240],[6818,5198],[6840,5206],[6861,5155],[6852,5115],[6907,5053],[6884,5017],[6889,4976],[6966,4950],[6960,4909],[6987,4904],[7024,4849],[7118,4826],[7129,4751],[7183,4772],[7254,4763],[7266,4719],[7320,4704],[7290,4620],[7246,4572],[7180,4585],[7126,4570],[7114,4511],[7171,4478],[7227,4334],[7278,4286],[7228,4275],[7196,4244],[7144,4237],[7114,4151],[7106,4084],[7149,4017],[7140,3984],[7103,3994],[7053,4033],[7005,4004],[6961,4007],[6918,3960],[6865,4022],[6825,3992],[6813,3942],[6780,3978],[6737,3959],[6704,3988],[6687,3955],[6639,3956],[6634,3914],[6616,3948],[6542,3930],[6519,3969],[6462,3957],[6378,3967],[6262,3963],[6159,3947],[6082,3911],[6025,3898],[6008,3848],[5970,3801],[5926,3810],[5907,3769],[5886,3834],[5894,3906],[5882,3930],[5789,3973],[5817,4042],[5890,4104],[5895,4126],[5868,4241],[5791,4360],[5756,4450],[5784,4444],[5875,4477],[5908,4464],[5952,4504],[5971,4502],[5972,4554],[6009,4557],[6026,4598],[6065,4618],[6086,4576],[6118,4585],[6100,4633],[6071,4634],[6092,4704],[6080,4791],[6131,4831],[6147,4887],[6193,4890],[6218,4942],[6157,4946],[6159,5044]]]}},{"type":"Feature","id":"IS","properties":{"hc-group":"admin0","hc-middle-x":0.59,"hc-middle-y":0.68,"hc-key":"is","hc-a2":"IS","name":"Iceland","labelrank":"3","country-abbrev":"Iceland","subregion":"Northern Europe","region-wb":"Europe & Central Asia","iso-a3":"ISL","iso-a2":"IS","woe-id":"23424845","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[23,7765],[-31,7764],[-42,7780],[-86,7762],[-64,7754],[-101,7722],[-209,7727],[-271,7767],[-318,7818],[-367,7843],[-388,7915],[-443,7984],[-435,8008],[-470,7988],[-517,8017],[-554,8023],[-582,8046],[-631,8058],[-594,8126],[-590,8089],[-546,8080],[-466,8107],[-487,8122],[-442,8153],[-483,8140],[-497,8153],[-458,8196],[-397,8218],[-488,8200],[-493,8230],[-466,8281],[-602,8368],[-616,8355],[-649,8375],[-644,8415],[-596,8393],[-542,8384],[-532,8401],[-506,8378],[-474,8390],[-473,8368],[-433,8366],[-370,8331],[-345,8373],[-387,8362],[-439,8405],[-423,8419],[-332,8436],[-332,8465],[-371,8459],[-387,8509],[-413,8504],[-455,8548],[-556,8542],[-556,8561],[-613,8605],[-567,8624],[-538,8578],[-544,8617],[-519,8656],[-469,8575],[-488,8630],[-478,8675],[-451,8648],[-466,8697],[-455,8712],[-432,8685],[-436,8718],[-402,8730],[-378,8700],[-381,8676],[-344,8653],[-330,8618],[-337,8579],[-309,8641],[-344,8686],[-333,8728],[-353,8755],[-336,8773],[-272,8752],[-251,8712],[-259,8681],[-233,8670],[-219,8634],[-229,8605],[-208,8600],[-197,8559],[-209,8522],[-269,8453],[-246,8455],[-266,8427],[-254,8402],[-265,8355],[-201,8434],[-164,8447],[-167,8384],[-110,8440],[-96,8526],[-87,8540],[-45,8536],[-48,8465],[-33,8447],[-39,8415],[-19,8396],[-2,8460],[31,8489],[61,8471],[71,8495],[101,8497],[121,8449],[109,8419],[128,8404],[123,8338],[146,8358],[140,8395],[160,8464],[199,8439],[214,8374],[237,8373],[282,8419],[307,8391],[363,8386],[381,8410],[391,8477],[452,8463],[449,8428],[472,8410],[453,8383],[483,8328],[504,8358],[527,8353],[569,8373],[592,8356],[537,8346],[535,8321],[502,8309],[508,8287],[554,8273],[551,8240],[502,8190],[566,8182],[553,8170],[604,8087],[624,8078],[583,8028],[604,8005],[589,7946],[569,7931],[532,7974],[550,7914],[516,7902],[505,7876],[444,7877],[405,7807],[384,7815],[347,7800],[289,7807],[204,7798],[142,7765],[77,7752],[64,7775],[45,7750],[23,7765]]]}},{"type":"Feature","id":"MD","properties":{"hc-group":"admin0","hc-middle-x":0.47,"hc-middle-y":0.3,"hc-key":"md","hc-a2":"MD","name":"Moldova","labelrank":"6","country-abbrev":"Mda.","subregion":"Eastern Europe","region-wb":"Europe & Central Asia","iso-a3":"MDA","iso-a2":"MD","woe-id":"23424885","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[7150,2258],[7114,2288],[7132,2304],[7089,2403],[7089,2531],[7070,2596],[7003,2689],[6953,2710],[6884,2789],[6821,2832],[6775,2912],[6729,2959],[6687,2976],[6652,2968],[6685,3015],[6749,3021],[6803,3068],[6850,3073],[6926,3031],[6981,3041],[7015,3026],[7070,3029],[7112,2987],[7141,3008],[7170,2984],[7174,2884],[7248,2829],[7273,2849],[7303,2774],[7302,2743],[7384,2725],[7404,2644],[7448,2613],[7405,2591],[7369,2620],[7355,2583],[7319,2610],[7279,2560],[7258,2609],[7222,2574],[7235,2519],[7256,2502],[7259,2448],[7223,2426],[7228,2391],[7190,2327],[7203,2283],[7150,2258]]]}},{"type":"Feature","id":"CZ","properties":{"hc-group":"admin0","hc-middle-x":0.39,"hc-middle-y":0.51,"hc-key":"cz","hc-a2":"CZ","name":"Czech Republic","labelrank":"5","country-abbrev":"Cz. Rep.","subregion":"Eastern Europe","region-wb":"Europe & Central Asia","iso-a3":"CZE","iso-a2":"CZ","woe-id":"23424810","continent":"Europe"},"geometry":{"type":"Polygon","coordinates":[[[5131,3062],[5077,3047],[5019,2981],[5005,2912],[4968,2872],[4910,2834],[4839,2840],[4804,2763],[4787,2794],[4702,2808],[4689,2788],[4627,2791],[4583,2821],[4558,2815],[4476,2844],[4422,2848],[4415,2776],[4392,2779],[4376,2724],[4329,2736],[4304,2710],[4257,2717],[4205,2765],[4163,2815],[4139,2811],[4079,2860],[4043,2910],[4004,2918],[3923,3034],[3947,3082],[3894,3121],[3860,3197],[3900,3155],[3940,3221],[4030,3255],[4070,3282],[4109,3288],[4136,3323],[4269,3379],[4256,3423],[4293,3419],[4318,3372],[4354,3378],[4385,3380],[4378,3418],[4435,3411],[4453,3363],[4470,3369],[4591,3335],[4636,3338],[4648,3301],[4615,3272],[4643,3260],[4718,3186],[4762,3230],[4739,3283],[4797,3277],[4830,3248],[4880,3250],[4880,3222],[4931,3173],[4967,3187],[5067,3169],[5088,3112],[5114,3108],[5131,3062]]]}}]},"data":[{"country":"PT","tz":"UTC","value":1},{"country":"IE","tz":"UTC","value":1},{"country":"GB","tz":"UTC","value":1},{"country":"IS","tz":"UTC","value":1},{"country":"NO","tz":"UTC + 1","value":2},{"country":"SE","tz":"UTC + 1","value":2},{"country":"DK","tz":"UTC + 1","value":2},{"country":"DE","tz":"UTC + 1","value":2},{"country":"NL","tz":"UTC + 1","value":2},{"country":"BE","tz":"UTC + 1","value":2},{"country":"LU","tz":"UTC + 1","value":2},{"country":"ES","tz":"UTC + 1","value":2},{"country":"FR","tz":"UTC + 1","value":2},{"country":"PL","tz":"UTC + 1","value":2},{"country":"CZ","tz":"UTC + 1","value":2},{"country":"AT","tz":"UTC + 1","value":2},{"country":"CH","tz":"UTC + 1","value":2},{"country":"LI","tz":"UTC + 1","value":2},{"country":"SK","tz":"UTC + 1","value":2},{"country":"HU","tz":"UTC + 1","value":2},{"country":"SI","tz":"UTC + 1","value":2},{"country":"IT","tz":"UTC + 1","value":2},{"country":"SM","tz":"UTC + 1","value":2},{"country":"HR","tz":"UTC + 1","value":2},{"country":"BA","tz":"UTC + 1","value":2},{"country":"YF","tz":"UTC + 1","value":2},{"country":"ME","tz":"UTC + 1","value":2},{"country":"AL","tz":"UTC + 1","value":2},{"country":"MK","tz":"UTC + 1","value":2},{"country":"FI","tz":"UCT + 2","value":3},{"country":"EE","tz":"UCT + 2","value":3},{"country":"LV","tz":"UCT + 2","value":3},{"country":"LT","tz":"UCT + 2","value":3},{"country":"BY","tz":"UCT + 2","value":3},{"country":"UA","tz":"UCT + 2","value":3},{"country":"MD","tz":"UCT + 2","value":3},{"country":"RO","tz":"UCT + 2","value":3},{"country":"BG","tz":"UCT + 2","value":3},{"country":"GR","tz":"UCT + 2","value":3},{"country":"TR","tz":"UCT + 2","value":3},{"country":"CY","tz":"UCT + 2","value":3},{"country":"RU","tz":"UTC + 3","value":4}],"joinBy":["iso-a2","country"],"name":"Time zone","tooltip":{"pointFormat":"{point.name} {point.tz}"},"dataLabels":{"enabled":true,"format":"{point.country}"}}],"colorAxis":{"dataClassColor":"category","dataClasses":[{"name":"UTC","from":1,"to":2},{"name":"UTC + 1","from":2,"to":3},{"name":"UCT + 2","from":3,"to":4},{"name":"UTC + 3","from":4,"to":5}]}},"theme":{"chart":{"backgroundColor":"transparent"},"colors":["#7cb5ec","#434348","#90ed7d","#f7a35c","#8085e9","#f15c80","#e4d354","#2b908f","#f45b5b","#91e8e1"]},"conf_opts":{"global":{"Date":null,"VMLRadialGradientURL":"http =//code.highcharts.com/list(version)/gfx/vml-radial-gradient.png","canvasToolsURL":"http =//code.highcharts.com/list(version)/modules/canvas-tools.js","getTimezoneOffset":null,"timezoneOffset":0,"useUTC":true},"lang":{"contextButtonTitle":"Chart context menu","decimalPoint":".","downloadJPEG":"Download JPEG image","downloadPDF":"Download PDF document","downloadPNG":"Download PNG image","downloadSVG":"Download SVG vector image","drillUpText":"Back to {series.name}","invalidDate":null,"loading":"Loading...","months":["January","February","March","April","May","June","July","August","September","October","November","December"],"noData":"No data to display","numericSymbols":["k","M","G","T","P","E"],"printChart":"Print chart","resetZoom":"Reset zoom","resetZoomTitle":"Reset zoom level 1:1","shortMonths":["Jan","Feb","Mar","Apr","May","Jun","Jul","Aug","Sep","Oct","Nov","Dec"],"thousandsSep":" ","weekdays":["Sunday","Monday","Tuesday","Wednesday","Thursday","Friday","Saturday"]}},"type":"map","fonts":[],"debug":false},"evals":[],"jsHooks":[]}</script> ] --- class: middle, inverse # Using D3.js in R --- ## r2d3 [D3.js](https://d3js.org) is a state-of-the-art Javascript pacman::p_load for data-driven graphics It is widely used in data journalism, including the New York Times and the Guardian However, it controls each aspect of a chart and so can require rather long JS scripts <iframe src="https://d3js.org" width="100%" height="300"></iframe> --- ## r2d3 If you know D3.js, you can use it pretty easily embed them in R using the **r2d3** package ```r pacman::p_load(r2d3) r2d3(data = read_csv('../data/flare.csv'), d3_version = 4, script = 'bubbles.js') ``` <div id="htmlwidget-d342a789753ef2ca6ae6" style="width:504px;height:504px;" class="r2d3 html-widget"></div> <script type="application/json" data-for="htmlwidget-d342a789753ef2ca6ae6">{"x":{"data":{"id":["flare","flare.analytics","flare.analytics.cluster","flare.analytics.cluster.AgglomerativeCluster","flare.analytics.cluster.CommunityStructure","flare.analytics.cluster.HierarchicalCluster","flare.analytics.cluster.MergeEdge","flare.analytics.graph","flare.analytics.graph.BetweennessCentrality","flare.analytics.graph.LinkDistance","flare.analytics.graph.MaxFlowMinCut","flare.analytics.graph.ShortestPaths","flare.analytics.graph.SpanningTree","flare.analytics.optimization","flare.analytics.optimization.AspectRatioBanker","flare.animate","flare.animate.Easing","flare.animate.FunctionSequence","flare.animate.interpolate","flare.animate.interpolate.ArrayInterpolator","flare.animate.interpolate.ColorInterpolator","flare.animate.interpolate.DateInterpolator","flare.animate.interpolate.Interpolator","flare.animate.interpolate.MatrixInterpolator","flare.animate.interpolate.NumberInterpolator","flare.animate.interpolate.ObjectInterpolator","flare.animate.interpolate.PointInterpolator","flare.animate.interpolate.RectangleInterpolator","flare.animate.ISchedulable","flare.animate.Parallel","flare.animate.Pause","flare.animate.Scheduler","flare.animate.Sequence","flare.animate.Transition","flare.animate.Transitioner","flare.animate.TransitionEvent","flare.animate.Tween","flare.data","flare.data.converters","flare.data.converters.Converters","flare.data.converters.DelimitedTextConverter","flare.data.converters.GraphMLConverter","flare.data.converters.IDataConverter","flare.data.converters.JSONConverter","flare.data.DataField","flare.data.DataSchema","flare.data.DataSet","flare.data.DataSource","flare.data.DataTable","flare.data.DataUtil","flare.display","flare.display.DirtySprite","flare.display.LineSprite","flare.display.RectSprite","flare.display.TextSprite","flare.flex","flare.flex.FlareVis","flare.physics","flare.physics.DragForce","flare.physics.GravityForce","flare.physics.IForce","flare.physics.NBodyForce","flare.physics.Particle","flare.physics.Simulation","flare.physics.Spring","flare.physics.SpringForce","flare.query","flare.query.AggregateExpression","flare.query.And","flare.query.Arithmetic","flare.query.Average","flare.query.BinaryExpression","flare.query.Comparison","flare.query.CompositeExpression","flare.query.Count","flare.query.DateUtil","flare.query.Distinct","flare.query.Expression","flare.query.ExpressionIterator","flare.query.Fn","flare.query.If","flare.query.IsA","flare.query.Literal","flare.query.Match","flare.query.Maximum","flare.query.methods","flare.query.methods.add","flare.query.methods.and","flare.query.methods.average","flare.query.methods.count","flare.query.methods.distinct","flare.query.methods.div","flare.query.methods.eq","flare.query.methods.fn","flare.query.methods.gt","flare.query.methods.gte","flare.query.methods.iff","flare.query.methods.isa","flare.query.methods.lt","flare.query.methods.lte","flare.query.methods.max","flare.query.methods.min","flare.query.methods.mod","flare.query.methods.mul","flare.query.methods.neq","flare.query.methods.not","flare.query.methods.or","flare.query.methods.orderby","flare.query.methods.range","flare.query.methods.select","flare.query.methods.stddev","flare.query.methods.sub","flare.query.methods.sum","flare.query.methods.update","flare.query.methods.variance","flare.query.methods.where","flare.query.methods.xor","flare.query.methods._","flare.query.Minimum","flare.query.Not","flare.query.Or","flare.query.Query","flare.query.Range","flare.query.StringUtil","flare.query.Sum","flare.query.Variable","flare.query.Variance","flare.query.Xor","flare.scale","flare.scale.IScaleMap","flare.scale.LinearScale","flare.scale.LogScale","flare.scale.OrdinalScale","flare.scale.QuantileScale","flare.scale.QuantitativeScale","flare.scale.RootScale","flare.scale.Scale","flare.scale.ScaleType","flare.scale.TimeScale","flare.util","flare.util.Arrays","flare.util.Colors","flare.util.Dates","flare.util.Displays","flare.util.Filter","flare.util.Geometry","flare.util.heap","flare.util.heap.FibonacciHeap","flare.util.heap.HeapNode","flare.util.IEvaluable","flare.util.IPredicate","flare.util.IValueProxy","flare.util.math","flare.util.math.DenseMatrix","flare.util.math.IMatrix","flare.util.math.SparseMatrix","flare.util.Maths","flare.util.Orientation","flare.util.palette","flare.util.palette.ColorPalette","flare.util.palette.Palette","flare.util.palette.ShapePalette","flare.util.palette.SizePalette","flare.util.Property","flare.util.Shapes","flare.util.Sort","flare.util.Stats","flare.util.Strings","flare.vis","flare.vis.axis","flare.vis.axis.Axes","flare.vis.axis.Axis","flare.vis.axis.AxisGridLine","flare.vis.axis.AxisLabel","flare.vis.axis.CartesianAxes","flare.vis.controls","flare.vis.controls.AnchorControl","flare.vis.controls.ClickControl","flare.vis.controls.Control","flare.vis.controls.ControlList","flare.vis.controls.DragControl","flare.vis.controls.ExpandControl","flare.vis.controls.HoverControl","flare.vis.controls.IControl","flare.vis.controls.PanZoomControl","flare.vis.controls.SelectionControl","flare.vis.controls.TooltipControl","flare.vis.data","flare.vis.data.Data","flare.vis.data.DataList","flare.vis.data.DataSprite","flare.vis.data.EdgeSprite","flare.vis.data.NodeSprite","flare.vis.data.render","flare.vis.data.render.ArrowType","flare.vis.data.render.EdgeRenderer","flare.vis.data.render.IRenderer","flare.vis.data.render.ShapeRenderer","flare.vis.data.ScaleBinding","flare.vis.data.Tree","flare.vis.data.TreeBuilder","flare.vis.events","flare.vis.events.DataEvent","flare.vis.events.SelectionEvent","flare.vis.events.TooltipEvent","flare.vis.events.VisualizationEvent","flare.vis.legend","flare.vis.legend.Legend","flare.vis.legend.LegendItem","flare.vis.legend.LegendRange","flare.vis.operator","flare.vis.operator.distortion","flare.vis.operator.distortion.BifocalDistortion","flare.vis.operator.distortion.Distortion","flare.vis.operator.distortion.FisheyeDistortion","flare.vis.operator.encoder","flare.vis.operator.encoder.ColorEncoder","flare.vis.operator.encoder.Encoder","flare.vis.operator.encoder.PropertyEncoder","flare.vis.operator.encoder.ShapeEncoder","flare.vis.operator.encoder.SizeEncoder","flare.vis.operator.filter","flare.vis.operator.filter.FisheyeTreeFilter","flare.vis.operator.filter.GraphDistanceFilter","flare.vis.operator.filter.VisibilityFilter","flare.vis.operator.IOperator","flare.vis.operator.label","flare.vis.operator.label.Labeler","flare.vis.operator.label.RadialLabeler","flare.vis.operator.label.StackedAreaLabeler","flare.vis.operator.layout","flare.vis.operator.layout.AxisLayout","flare.vis.operator.layout.BundledEdgeRouter","flare.vis.operator.layout.CircleLayout","flare.vis.operator.layout.CirclePackingLayout","flare.vis.operator.layout.DendrogramLayout","flare.vis.operator.layout.ForceDirectedLayout","flare.vis.operator.layout.IcicleTreeLayout","flare.vis.operator.layout.IndentedTreeLayout","flare.vis.operator.layout.Layout","flare.vis.operator.layout.NodeLinkTreeLayout","flare.vis.operator.layout.PieLayout","flare.vis.operator.layout.RadialTreeLayout","flare.vis.operator.layout.RandomLayout","flare.vis.operator.layout.StackedAreaLayout","flare.vis.operator.layout.TreeMapLayout","flare.vis.operator.Operator","flare.vis.operator.OperatorList","flare.vis.operator.OperatorSequence","flare.vis.operator.OperatorSwitch","flare.vis.operator.SortOperator","flare.vis.Visualization"],"value":[null,null,null,3938,3812,6714,743,null,3534,5731,7840,5914,3416,null,7074,null,17010,5842,null,1983,2047,1375,8746,2202,1382,1629,1675,2042,1041,5176,449,5593,5534,9201,19975,1116,6006,null,null,721,4294,9800,1314,2220,1759,2165,586,3331,772,3322,null,8833,1732,3623,10066,null,4116,null,1082,1336,319,10498,2822,9983,2213,1681,null,1616,1027,3891,891,2893,5103,3677,781,4141,933,5130,3617,3240,2732,2039,1214,3748,843,null,593,330,287,277,292,595,594,460,603,625,748,461,597,619,283,283,591,603,599,386,323,307,772,296,363,600,280,307,335,299,354,264,843,1554,970,13896,1594,4130,791,1124,1876,1101,null,2105,1316,3151,3770,2435,4839,1756,4268,1821,5833,null,8258,10001,8217,12555,2324,10993,null,9354,1233,335,383,874,null,3165,2815,3366,17705,1486,null,6367,1229,2059,2291,5559,19118,6887,6557,22026,null,null,1302,24593,652,636,6703,null,2138,3824,1353,4665,2649,2832,4896,763,5222,7862,8435,null,20544,19788,10349,3301,19382,null,698,5569,353,2247,11275,7147,9930,null,2313,1880,1701,1117,null,20859,4614,10530,null,null,4461,6314,3444,null,3179,4060,4138,1690,1830,null,5219,3165,3509,1286,null,9956,3899,3202,null,6725,3727,9317,12003,4853,8411,4864,3174,7881,12870,2728,12348,870,9121,9191,2490,5248,4190,2581,2023,16540]},"type":"data.frame","container":"svg","options":null,"script":"var d3Script = function(d3, r2d3, data, svg, width, height, options, theme, console) {\nthis.d3 = d3;\n\nsvg = d3.select(svg.node());\n/* R2D3 Source File: bubbles.js */\n// !preview r2d3 data = read.csv(\"flare.csv\"), d3_version = 4\n\n// Based on https://bl.ocks.org/mbostock/4063269\n\n// Initialization\nsvg.attr(\"font-family\", \"sans-serif\")\n .attr(\"font-size\", \"8\")\n .attr(\"text-anchor\", \"middle\");\n \nvar svgSize = 600;\nvar pack = d3.pack()\n .size([svgSize, svgSize])\n .padding(1.5);\n \nvar format = d3.format(\",d\");\nvar color = d3.scaleOrdinal(d3.schemeCategory20c);\n\nvar group = svg.append(\"g\");\n\n// Resize\nr2d3.onResize(function(width, height) {\n var minSize = Math.min(width, height);\n var scale = minSize / svgSize;\n \n group.attr(\"transform\", function(d) {\n return \"\" +\n \"translate(\" + (width - minSize) / 2 + \",\" + (height - minSize) / 2 + \"),\" +\n \"scale(\" + scale + \",\" + scale + \")\";\n });\n});\n\n// Rendering\nr2d3.onRender(function(data, svg, width, height, options) {\n var root = d3.hierarchy({children: data})\n .sum(function(d) { return d.value; })\n .each(function(d) {\n if (id = d.data.id) {\n var id, i = id.lastIndexOf(\".\");\n d.id = id;\n d.package = id.slice(0, i);\n d.class = id.slice(i + 1);\n }\n });\n\n var node = group.selectAll(\".node\")\n .data(pack(root).leaves())\n .enter().append(\"g\")\n .attr(\"class\", \"node\")\n .attr(\"transform\", function(d) { return \"translate(\" + d.x + \",\" + d.y + \")\"; });\n\n node.append(\"circle\")\n .attr(\"id\", function(d) { return d.id; })\n .attr(\"r\", function(d) { return d.r; })\n .style(\"fill\", function(d) { return color(d.package); });\n\n node.append(\"clipPath\")\n .attr(\"id\", function(d) { return \"clip-\" + d.id; })\n .append(\"use\")\n .attr(\"xlink:href\", function(d) { return \"#\" + d.id; });\n\n node.append(\"text\")\n .attr(\"clip-path\", function(d) { return \"url(#clip-\" + d.id + \")\"; })\n .selectAll(\"tspan\")\n .data(function(d) { return d.class.split(/(?=[A-Z][^A-Z])/g); })\n .enter().append(\"tspan\")\n .attr(\"x\", 0)\n .attr(\"y\", function(d, i, nodes) { return 13 + (i - nodes.length / 2 - 0.5) * 10; })\n .text(function(d) { return d; });\n\n node.append(\"title\")\n .text(function(d) { return d.id + \"\\n\" + format(d.value); });\n \n r2d3.resize(width, height);\n});\n};","style":"/* R2D3 Source File: ./styles.css */\np {\n font-size: 12pt;\n}\nul:not(.nav) {\n font-size: 12pt;\n}\n.caption {\n color: #777;\n margin-top: 10px;\n}\np code {\n white-space: inherit;\n}\npre {\n word-break: normal;\n word-wrap: normal;\n}\npre code {\n white-space: inherit;\n}\n\nh1, h2, h3, h4, p {\n font-family: Charter, serif;\n}\nblockquote {\n font-size: 90%;\n margin-left: 20px;\n margin-top: 5px;\n margin-bottom: 5px;\n background-color: beige;\n}","version":4,"theme":{"default":{"background":"#FFFFFF","foreground":"#000000"},"runtime":null},"useShadow":true},"evals":[],"jsHooks":[]}</script> --- ## r2d3 ```r r2d3(data = jsonlite::read_json('../data/miserables.json'), d3_version = 4, script = 'forcegraph.js') ``` <div id="htmlwidget-59ab95be0db3c0ca28c5" style="width:504px;height:504px;" class="r2d3 html-widget"></div> <script type="application/json" data-for="htmlwidget-59ab95be0db3c0ca28c5">{"x":{"data":{"nodes":[{"id":"Myriel","group":1},{"id":"Napoleon","group":1},{"id":"Mlle.Baptistine","group":1},{"id":"Mme.Magloire","group":1},{"id":"CountessdeLo","group":1},{"id":"Geborand","group":1},{"id":"Champtercier","group":1},{"id":"Cravatte","group":1},{"id":"Count","group":1},{"id":"OldMan","group":1},{"id":"Labarre","group":2},{"id":"Valjean","group":2},{"id":"Marguerite","group":3},{"id":"Mme.deR","group":2},{"id":"Isabeau","group":2},{"id":"Gervais","group":2},{"id":"Tholomyes","group":3},{"id":"Listolier","group":3},{"id":"Fameuil","group":3},{"id":"Blacheville","group":3},{"id":"Favourite","group":3},{"id":"Dahlia","group":3},{"id":"Zephine","group":3},{"id":"Fantine","group":3},{"id":"Mme.Thenardier","group":4},{"id":"Thenardier","group":4},{"id":"Cosette","group":5},{"id":"Javert","group":4},{"id":"Fauchelevent","group":0},{"id":"Bamatabois","group":2},{"id":"Perpetue","group":3},{"id":"Simplice","group":2},{"id":"Scaufflaire","group":2},{"id":"Woman1","group":2},{"id":"Judge","group":2},{"id":"Champmathieu","group":2},{"id":"Brevet","group":2},{"id":"Chenildieu","group":2},{"id":"Cochepaille","group":2},{"id":"Pontmercy","group":4},{"id":"Boulatruelle","group":6},{"id":"Eponine","group":4},{"id":"Anzelma","group":4},{"id":"Woman2","group":5},{"id":"MotherInnocent","group":0},{"id":"Gribier","group":0},{"id":"Jondrette","group":7},{"id":"Mme.Burgon","group":7},{"id":"Gavroche","group":8},{"id":"Gillenormand","group":5},{"id":"Magnon","group":5},{"id":"Mlle.Gillenormand","group":5},{"id":"Mme.Pontmercy","group":5},{"id":"Mlle.Vaubois","group":5},{"id":"Lt.Gillenormand","group":5},{"id":"Marius","group":8},{"id":"BaronessT","group":5},{"id":"Mabeuf","group":8},{"id":"Enjolras","group":8},{"id":"Combeferre","group":8},{"id":"Prouvaire","group":8},{"id":"Feuilly","group":8},{"id":"Courfeyrac","group":8},{"id":"Bahorel","group":8},{"id":"Bossuet","group":8},{"id":"Joly","group":8},{"id":"Grantaire","group":8},{"id":"MotherPlutarch","group":9},{"id":"Gueulemer","group":4},{"id":"Babet","group":4},{"id":"Claquesous","group":4},{"id":"Montparnasse","group":4},{"id":"Toussaint","group":5},{"id":"Child1","group":10},{"id":"Child2","group":10},{"id":"Brujon","group":4},{"id":"Mme.Hucheloup","group":8}],"links":[{"source":"Napoleon","target":"Myriel","value":1},{"source":"Mlle.Baptistine","target":"Myriel","value":8},{"source":"Mme.Magloire","target":"Myriel","value":10},{"source":"Mme.Magloire","target":"Mlle.Baptistine","value":6},{"source":"CountessdeLo","target":"Myriel","value":1},{"source":"Geborand","target":"Myriel","value":1},{"source":"Champtercier","target":"Myriel","value":1},{"source":"Cravatte","target":"Myriel","value":1},{"source":"Count","target":"Myriel","value":2},{"source":"OldMan","target":"Myriel","value":1},{"source":"Valjean","target":"Labarre","value":1},{"source":"Valjean","target":"Mme.Magloire","value":3},{"source":"Valjean","target":"Mlle.Baptistine","value":3},{"source":"Valjean","target":"Myriel","value":5},{"source":"Marguerite","target":"Valjean","value":1},{"source":"Mme.deR","target":"Valjean","value":1},{"source":"Isabeau","target":"Valjean","value":1},{"source":"Gervais","target":"Valjean","value":1},{"source":"Listolier","target":"Tholomyes","value":4},{"source":"Fameuil","target":"Tholomyes","value":4},{"source":"Fameuil","target":"Listolier","value":4},{"source":"Blacheville","target":"Tholomyes","value":4},{"source":"Blacheville","target":"Listolier","value":4},{"source":"Blacheville","target":"Fameuil","value":4},{"source":"Favourite","target":"Tholomyes","value":3},{"source":"Favourite","target":"Listolier","value":3},{"source":"Favourite","target":"Fameuil","value":3},{"source":"Favourite","target":"Blacheville","value":4},{"source":"Dahlia","target":"Tholomyes","value":3},{"source":"Dahlia","target":"Listolier","value":3},{"source":"Dahlia","target":"Fameuil","value":3},{"source":"Dahlia","target":"Blacheville","value":3},{"source":"Dahlia","target":"Favourite","value":5},{"source":"Zephine","target":"Tholomyes","value":3},{"source":"Zephine","target":"Listolier","value":3},{"source":"Zephine","target":"Fameuil","value":3},{"source":"Zephine","target":"Blacheville","value":3},{"source":"Zephine","target":"Favourite","value":4},{"source":"Zephine","target":"Dahlia","value":4},{"source":"Fantine","target":"Tholomyes","value":3},{"source":"Fantine","target":"Listolier","value":3},{"source":"Fantine","target":"Fameuil","value":3},{"source":"Fantine","target":"Blacheville","value":3},{"source":"Fantine","target":"Favourite","value":4},{"source":"Fantine","target":"Dahlia","value":4},{"source":"Fantine","target":"Zephine","value":4},{"source":"Fantine","target":"Marguerite","value":2},{"source":"Fantine","target":"Valjean","value":9},{"source":"Mme.Thenardier","target":"Fantine","value":2},{"source":"Mme.Thenardier","target":"Valjean","value":7},{"source":"Thenardier","target":"Mme.Thenardier","value":13},{"source":"Thenardier","target":"Fantine","value":1},{"source":"Thenardier","target":"Valjean","value":12},{"source":"Cosette","target":"Mme.Thenardier","value":4},{"source":"Cosette","target":"Valjean","value":31},{"source":"Cosette","target":"Tholomyes","value":1},{"source":"Cosette","target":"Thenardier","value":1},{"source":"Javert","target":"Valjean","value":17},{"source":"Javert","target":"Fantine","value":5},{"source":"Javert","target":"Thenardier","value":5},{"source":"Javert","target":"Mme.Thenardier","value":1},{"source":"Javert","target":"Cosette","value":1},{"source":"Fauchelevent","target":"Valjean","value":8},{"source":"Fauchelevent","target":"Javert","value":1},{"source":"Bamatabois","target":"Fantine","value":1},{"source":"Bamatabois","target":"Javert","value":1},{"source":"Bamatabois","target":"Valjean","value":2},{"source":"Perpetue","target":"Fantine","value":1},{"source":"Simplice","target":"Perpetue","value":2},{"source":"Simplice","target":"Valjean","value":3},{"source":"Simplice","target":"Fantine","value":2},{"source":"Simplice","target":"Javert","value":1},{"source":"Scaufflaire","target":"Valjean","value":1},{"source":"Woman1","target":"Valjean","value":2},{"source":"Woman1","target":"Javert","value":1},{"source":"Judge","target":"Valjean","value":3},{"source":"Judge","target":"Bamatabois","value":2},{"source":"Champmathieu","target":"Valjean","value":3},{"source":"Champmathieu","target":"Judge","value":3},{"source":"Champmathieu","target":"Bamatabois","value":2},{"source":"Brevet","target":"Judge","value":2},{"source":"Brevet","target":"Champmathieu","value":2},{"source":"Brevet","target":"Valjean","value":2},{"source":"Brevet","target":"Bamatabois","value":1},{"source":"Chenildieu","target":"Judge","value":2},{"source":"Chenildieu","target":"Champmathieu","value":2},{"source":"Chenildieu","target":"Brevet","value":2},{"source":"Chenildieu","target":"Valjean","value":2},{"source":"Chenildieu","target":"Bamatabois","value":1},{"source":"Cochepaille","target":"Judge","value":2},{"source":"Cochepaille","target":"Champmathieu","value":2},{"source":"Cochepaille","target":"Brevet","value":2},{"source":"Cochepaille","target":"Chenildieu","value":2},{"source":"Cochepaille","target":"Valjean","value":2},{"source":"Cochepaille","target":"Bamatabois","value":1},{"source":"Pontmercy","target":"Thenardier","value":1},{"source":"Boulatruelle","target":"Thenardier","value":1},{"source":"Eponine","target":"Mme.Thenardier","value":2},{"source":"Eponine","target":"Thenardier","value":3},{"source":"Anzelma","target":"Eponine","value":2},{"source":"Anzelma","target":"Thenardier","value":2},{"source":"Anzelma","target":"Mme.Thenardier","value":1},{"source":"Woman2","target":"Valjean","value":3},{"source":"Woman2","target":"Cosette","value":1},{"source":"Woman2","target":"Javert","value":1},{"source":"MotherInnocent","target":"Fauchelevent","value":3},{"source":"MotherInnocent","target":"Valjean","value":1},{"source":"Gribier","target":"Fauchelevent","value":2},{"source":"Mme.Burgon","target":"Jondrette","value":1},{"source":"Gavroche","target":"Mme.Burgon","value":2},{"source":"Gavroche","target":"Thenardier","value":1},{"source":"Gavroche","target":"Javert","value":1},{"source":"Gavroche","target":"Valjean","value":1},{"source":"Gillenormand","target":"Cosette","value":3},{"source":"Gillenormand","target":"Valjean","value":2},{"source":"Magnon","target":"Gillenormand","value":1},{"source":"Magnon","target":"Mme.Thenardier","value":1},{"source":"Mlle.Gillenormand","target":"Gillenormand","value":9},{"source":"Mlle.Gillenormand","target":"Cosette","value":2},{"source":"Mlle.Gillenormand","target":"Valjean","value":2},{"source":"Mme.Pontmercy","target":"Mlle.Gillenormand","value":1},{"source":"Mme.Pontmercy","target":"Pontmercy","value":1},{"source":"Mlle.Vaubois","target":"Mlle.Gillenormand","value":1},{"source":"Lt.Gillenormand","target":"Mlle.Gillenormand","value":2},{"source":"Lt.Gillenormand","target":"Gillenormand","value":1},{"source":"Lt.Gillenormand","target":"Cosette","value":1},{"source":"Marius","target":"Mlle.Gillenormand","value":6},{"source":"Marius","target":"Gillenormand","value":12},{"source":"Marius","target":"Pontmercy","value":1},{"source":"Marius","target":"Lt.Gillenormand","value":1},{"source":"Marius","target":"Cosette","value":21},{"source":"Marius","target":"Valjean","value":19},{"source":"Marius","target":"Tholomyes","value":1},{"source":"Marius","target":"Thenardier","value":2},{"source":"Marius","target":"Eponine","value":5},{"source":"Marius","target":"Gavroche","value":4},{"source":"BaronessT","target":"Gillenormand","value":1},{"source":"BaronessT","target":"Marius","value":1},{"source":"Mabeuf","target":"Marius","value":1},{"source":"Mabeuf","target":"Eponine","value":1},{"source":"Mabeuf","target":"Gavroche","value":1},{"source":"Enjolras","target":"Marius","value":7},{"source":"Enjolras","target":"Gavroche","value":7},{"source":"Enjolras","target":"Javert","value":6},{"source":"Enjolras","target":"Mabeuf","value":1},{"source":"Enjolras","target":"Valjean","value":4},{"source":"Combeferre","target":"Enjolras","value":15},{"source":"Combeferre","target":"Marius","value":5},{"source":"Combeferre","target":"Gavroche","value":6},{"source":"Combeferre","target":"Mabeuf","value":2},{"source":"Prouvaire","target":"Gavroche","value":1},{"source":"Prouvaire","target":"Enjolras","value":4},{"source":"Prouvaire","target":"Combeferre","value":2},{"source":"Feuilly","target":"Gavroche","value":2},{"source":"Feuilly","target":"Enjolras","value":6},{"source":"Feuilly","target":"Prouvaire","value":2},{"source":"Feuilly","target":"Combeferre","value":5},{"source":"Feuilly","target":"Mabeuf","value":1},{"source":"Feuilly","target":"Marius","value":1},{"source":"Courfeyrac","target":"Marius","value":9},{"source":"Courfeyrac","target":"Enjolras","value":17},{"source":"Courfeyrac","target":"Combeferre","value":13},{"source":"Courfeyrac","target":"Gavroche","value":7},{"source":"Courfeyrac","target":"Mabeuf","value":2},{"source":"Courfeyrac","target":"Eponine","value":1},{"source":"Courfeyrac","target":"Feuilly","value":6},{"source":"Courfeyrac","target":"Prouvaire","value":3},{"source":"Bahorel","target":"Combeferre","value":5},{"source":"Bahorel","target":"Gavroche","value":5},{"source":"Bahorel","target":"Courfeyrac","value":6},{"source":"Bahorel","target":"Mabeuf","value":2},{"source":"Bahorel","target":"Enjolras","value":4},{"source":"Bahorel","target":"Feuilly","value":3},{"source":"Bahorel","target":"Prouvaire","value":2},{"source":"Bahorel","target":"Marius","value":1},{"source":"Bossuet","target":"Marius","value":5},{"source":"Bossuet","target":"Courfeyrac","value":12},{"source":"Bossuet","target":"Gavroche","value":5},{"source":"Bossuet","target":"Bahorel","value":4},{"source":"Bossuet","target":"Enjolras","value":10},{"source":"Bossuet","target":"Feuilly","value":6},{"source":"Bossuet","target":"Prouvaire","value":2},{"source":"Bossuet","target":"Combeferre","value":9},{"source":"Bossuet","target":"Mabeuf","value":1},{"source":"Bossuet","target":"Valjean","value":1},{"source":"Joly","target":"Bahorel","value":5},{"source":"Joly","target":"Bossuet","value":7},{"source":"Joly","target":"Gavroche","value":3},{"source":"Joly","target":"Courfeyrac","value":5},{"source":"Joly","target":"Enjolras","value":5},{"source":"Joly","target":"Feuilly","value":5},{"source":"Joly","target":"Prouvaire","value":2},{"source":"Joly","target":"Combeferre","value":5},{"source":"Joly","target":"Mabeuf","value":1},{"source":"Joly","target":"Marius","value":2},{"source":"Grantaire","target":"Bossuet","value":3},{"source":"Grantaire","target":"Enjolras","value":3},{"source":"Grantaire","target":"Combeferre","value":1},{"source":"Grantaire","target":"Courfeyrac","value":2},{"source":"Grantaire","target":"Joly","value":2},{"source":"Grantaire","target":"Gavroche","value":1},{"source":"Grantaire","target":"Bahorel","value":1},{"source":"Grantaire","target":"Feuilly","value":1},{"source":"Grantaire","target":"Prouvaire","value":1},{"source":"MotherPlutarch","target":"Mabeuf","value":3},{"source":"Gueulemer","target":"Thenardier","value":5},{"source":"Gueulemer","target":"Valjean","value":1},{"source":"Gueulemer","target":"Mme.Thenardier","value":1},{"source":"Gueulemer","target":"Javert","value":1},{"source":"Gueulemer","target":"Gavroche","value":1},{"source":"Gueulemer","target":"Eponine","value":1},{"source":"Babet","target":"Thenardier","value":6},{"source":"Babet","target":"Gueulemer","value":6},{"source":"Babet","target":"Valjean","value":1},{"source":"Babet","target":"Mme.Thenardier","value":1},{"source":"Babet","target":"Javert","value":2},{"source":"Babet","target":"Gavroche","value":1},{"source":"Babet","target":"Eponine","value":1},{"source":"Claquesous","target":"Thenardier","value":4},{"source":"Claquesous","target":"Babet","value":4},{"source":"Claquesous","target":"Gueulemer","value":4},{"source":"Claquesous","target":"Valjean","value":1},{"source":"Claquesous","target":"Mme.Thenardier","value":1},{"source":"Claquesous","target":"Javert","value":1},{"source":"Claquesous","target":"Eponine","value":1},{"source":"Claquesous","target":"Enjolras","value":1},{"source":"Montparnasse","target":"Javert","value":1},{"source":"Montparnasse","target":"Babet","value":2},{"source":"Montparnasse","target":"Gueulemer","value":2},{"source":"Montparnasse","target":"Claquesous","value":2},{"source":"Montparnasse","target":"Valjean","value":1},{"source":"Montparnasse","target":"Gavroche","value":1},{"source":"Montparnasse","target":"Eponine","value":1},{"source":"Montparnasse","target":"Thenardier","value":1},{"source":"Toussaint","target":"Cosette","value":2},{"source":"Toussaint","target":"Javert","value":1},{"source":"Toussaint","target":"Valjean","value":1},{"source":"Child1","target":"Gavroche","value":2},{"source":"Child2","target":"Gavroche","value":2},{"source":"Child2","target":"Child1","value":3},{"source":"Brujon","target":"Babet","value":3},{"source":"Brujon","target":"Gueulemer","value":3},{"source":"Brujon","target":"Thenardier","value":3},{"source":"Brujon","target":"Gavroche","value":1},{"source":"Brujon","target":"Eponine","value":1},{"source":"Brujon","target":"Claquesous","value":1},{"source":"Brujon","target":"Montparnasse","value":1},{"source":"Mme.Hucheloup","target":"Bossuet","value":1},{"source":"Mme.Hucheloup","target":"Joly","value":1},{"source":"Mme.Hucheloup","target":"Grantaire","value":1},{"source":"Mme.Hucheloup","target":"Bahorel","value":1},{"source":"Mme.Hucheloup","target":"Courfeyrac","value":1},{"source":"Mme.Hucheloup","target":"Gavroche","value":1},{"source":"Mme.Hucheloup","target":"Enjolras","value":1}]},"type":"list","container":"svg","options":null,"script":"var d3Script = function(d3, r2d3, data, svg, width, height, options, theme, console) {\nthis.d3 = d3;\n\nsvg = d3.select(svg.node());\nforcegraph.js\n};","style":"/* R2D3 Source File: ./styles.css */\np {\n font-size: 12pt;\n}\nul:not(.nav) {\n font-size: 12pt;\n}\n.caption {\n color: #777;\n margin-top: 10px;\n}\np code {\n white-space: inherit;\n}\npre {\n word-break: normal;\n word-wrap: normal;\n}\npre code {\n white-space: inherit;\n}\n\nh1, h2, h3, h4, p {\n font-family: Charter, serif;\n}\nblockquote {\n font-size: 90%;\n margin-left: 20px;\n margin-top: 5px;\n margin-bottom: 5px;\n background-color: beige;\n}","version":4,"theme":{"default":{"background":"#FFFFFF","foreground":"#000000"},"runtime":null},"useShadow":true},"evals":[],"jsHooks":[]}</script> --- ## networkD3 You can also draw networks slightly more easily using the **networkD3** package. .pull-left[ ```r pacman::p_load(networkD3) lemis <- jsonlite::fromJSON('../data/miserables.json') lemis$links <- lemis$links %>% mutate(ID = 1:n()) %>% gather(location, nm, source, target) %>% mutate(nm = as.integer(as.factor(nm))-1) %>% spread(location, nm) forceNetwork(Links = lemis$links, Nodes = lemis$nodes, Source='source', Target='target', Value = 'value', NodeID = 'id', Group = 'group', opacity=0.7, zoom = TRUE, fontSize = 20) ``` ] .pull-right[ <div id="htmlwidget-96d1b29a5fd68edf3dd0" style="width:504px;height:504px;" class="forceNetwork html-widget"></div> <script type="application/json" data-for="htmlwidget-96d1b29a5fd68edf3dd0">{"x":{"links":{"source":[63,20,32,11,22,64,73,48,54,38,33,70,18,18,39,39,28,3,3,65,69,68,74,8,12,16,66,7,0,75,75,60,53,31,31,31,47,47,57,57,52,45,45,49,49,49,4,4,46,46,46,24,67,30,30,21,2,6,6,40,35,35,35,35,35,37,37,37,37,37,1,1,1,1,15,15,15,15,15,59,59,59,59,59,72,72,9,9,9,9,55,55,55,55,55,55,55,19,27,58,3,69,69,74,42,10,8,8,8,12,12,12,12,16,16,16,16,16,25,0,0,36,31,34,51,51,45,49,17,67,30,30,21,2,2,6,40,40,35,35,1,59,59,59,72,13,14,73,73,29,29,29,23,23,23,23,76,76,76,76,27,27,27,27,69,42,10,10,25,75,60,34,21,2,40,35,35,61,14,9,9,9,44,26,26,5,5,5,29,76,76,27,27,27,18,49,24,67,2,6,15,15,15,73,23,39,39,49,17,30,2,2,6,6,40,40,40,40,40,37,56,49,24,17,30,21,2,6,1,1,58,24,24,21,40,50,28,27,51,21,6,56,6,70,49,6,70,21,17,39,21,49,49,18],"target":[62,62,62,62,62,62,43,73,73,73,73,27,71,70,58,18,39,27,39,27,39,73,39,3,3,3,70,70,58,18,39,73,41,70,39,73,34,58,51,66,51,34,18,66,45,71,34,49,49,25,31,46,31,46,49,25,49,46,73,46,17,31,2,30,67,73,58,39,31,25,73,58,31,25,73,58,39,25,24,39,73,31,25,70,39,73,31,25,15,59,6,40,35,2,21,31,24,62,48,27,73,65,27,73,3,3,42,10,73,42,10,8,73,42,10,8,12,73,58,25,70,28,53,73,18,73,51,70,46,17,31,67,46,46,67,67,67,49,21,40,39,1,37,15,18,31,31,56,50,71,44,26,71,44,26,5,71,44,26,5,71,44,26,5,73,73,73,42,70,73,28,18,67,30,31,6,24,46,13,1,37,70,71,71,44,71,44,26,5,29,23,29,23,76,58,31,73,24,24,2,70,1,37,62,29,27,70,25,49,17,17,31,49,31,2,21,24,30,17,70,50,51,39,31,24,30,21,30,70,37,73,49,31,31,6,62,73,73,34,49,17,62,24,73,34,21,58,17,24,73,24,73,18,73],"value":[1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,5,5,5,5,5,5,5,5,5,5,5,5,5,5,5,5,5,6,6,6,6,6,6,6,6,6,6,7,7,7,7,7,8,8,9,9,9,9,10,10,12,12,12,13,13,15,17,17,19,21,31],"colour":["#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666","#666"]},"nodes":{"name":["Myriel","Napoleon","Mlle.Baptistine","Mme.Magloire","CountessdeLo","Geborand","Champtercier","Cravatte","Count","OldMan","Labarre","Valjean","Marguerite","Mme.deR","Isabeau","Gervais","Tholomyes","Listolier","Fameuil","Blacheville","Favourite","Dahlia","Zephine","Fantine","Mme.Thenardier","Thenardier","Cosette","Javert","Fauchelevent","Bamatabois","Perpetue","Simplice","Scaufflaire","Woman1","Judge","Champmathieu","Brevet","Chenildieu","Cochepaille","Pontmercy","Boulatruelle","Eponine","Anzelma","Woman2","MotherInnocent","Gribier","Jondrette","Mme.Burgon","Gavroche","Gillenormand","Magnon","Mlle.Gillenormand","Mme.Pontmercy","Mlle.Vaubois","Lt.Gillenormand","Marius","BaronessT","Mabeuf","Enjolras","Combeferre","Prouvaire","Feuilly","Courfeyrac","Bahorel","Bossuet","Joly","Grantaire","MotherPlutarch","Gueulemer","Babet","Claquesous","Montparnasse","Toussaint","Child1","Child2","Brujon","Mme.Hucheloup"],"group":[1,1,1,1,1,1,1,1,1,1,2,2,3,2,2,2,3,3,3,3,3,3,3,3,4,4,5,4,0,2,3,2,2,2,2,2,2,2,2,4,6,4,4,5,0,0,7,7,8,5,5,5,5,5,5,8,5,8,8,8,8,8,8,8,8,8,8,9,4,4,4,4,5,10,10,4,8]},"options":{"NodeID":"id","Group":"group","colourScale":"d3.scaleOrdinal(d3.schemeCategory20);","fontSize":20,"fontFamily":"serif","clickTextSize":50,"linkDistance":50,"linkWidth":"function(d) { return Math.sqrt(d.value); }","charge":-30,"opacity":0.7,"zoom":true,"legend":false,"arrows":false,"nodesize":false,"radiusCalculation":" Math.sqrt(d.nodesize)+6","bounded":false,"opacityNoHover":0,"clickAction":null}},"evals":[],"jsHooks":[]}</script> ] --- ## Crosstalk **Crosstalk** is a package that allows multiple HTML widgets to share data and interact together. Not all packages in the **htmlwidgets** family are **crosstalk**-compatible. ```r load('data/exdata.rda') pacman::p_load(leaflet); pacman::p_load(crosstalk); pacman::p_load(d3scatter) gpx1 <- gpx %>% mutate(Minutes = as.numeric(difftime(Time, min(Time), units = 'mins'))) %>% filter(Minutes <= 50) %>% left_join(run_data) shared_gpx1 <- SharedData$new(gpx1) bscols( leaflet(data = shared_gpx1, width="100%", height=450) %>% addTiles() %>% addCircleMarkers(~Longitude, ~Latitude, radius=1, color='blue'), list( d3scatter(shared_gpx1, ~Minutes, ~ HR, width="100%", height=450) ) ) bscols( filter_slider("Minutes","Time", shared_gpx1, ~Minutes, width="100%") ) ``` .center[`leaflet` + `d3scatter` + `crosstalk`] --- ## Linked graphs <iframe src="https://webbedfeet.github.io/DC_R2018/Presentation.html#23" width="100%" height="500"></iframe> --- ## Conclusions + There are many many resources today to create interactive graphics + For many of them, there are wrappers in R + You can explore the respective packages for many more details about the different kinds of charts and customizations that are possible.